Patent application title: HUMANIZED ANTIBODIES AGAINST CXCR3
Inventors:
Rodger Smith (Jefferson, MD, US)
Palanisamy Kanakaraj (Germantown, MD, US)
Viktor Roschke (Bethesda, MD, US)
Viktor Roschke (Bethesda, MD, US)
Assignees:
Teva Biopharmaceuticals USA, Inc.
IPC8 Class: AC07K1628FI
USPC Class:
5303873
Class name: Globulins immunoglobulin, antibody, or fragment thereof, other than immunoglobulin antibody, or fragment thereof that is conjugated or adsorbed chimeric, mutated, or recombined hybrid (e.g., bifunctional, bispecific, rodent-human chimeric, single chain, rfv, immunoglobulin fusion protein, etc.)
Publication date: 2013-10-17
Patent application number: 20130274447
Abstract:
Disclosed are humanized antibodies that bind specifically to the receptor
CXCR3. The humanized antibodies may be antagonists and may be used to
treat or diagnose conditions associated with CXCR3 function.Claims:
1.-98. (canceled)
99. A humanized antibody heavy chain variable region comprising: (a) a CDR-H1 comprising the amino acid sequence {N,S,Y}MAS; (b) a CDR-H2 comprising the amino acid sequence {T,A,Y}I{S,Y}{S,G,T,Y}{G,S}{G,Y}G{F,S,Y}TYY{P,A}DS{L,Y,V}KG; and (c) a CDR-H3 comprising the amino acid sequence {H,Y}{G,Y}{A,Y}PM{T,Y}T{V,Y}ITY{A,Y}PYYFYY.
100. The humanized antibody heavy chain variable region of claim 99, comprising the amino acid sequence: TABLE-US-00003 EVQLLESGGGLVQPGGSLRLSCAASGFTFS{N, S, Y}YAMSWVRQAPGKGLEWVS{T, A, Y} I{S, Y}{S, G, T, Y}{G, S}{G, Y}G{F, S, Y}TYY{P, A}DS{L, Y, V}KGRFTISRDNSKNTLYLQMNSL RAEDTAVYYCAK{H, Y}{G, Y}{A, Y}PM{T, Y}T{V, Y}ITY{A, Y}PYYFYYWGQGTTVTVSS.
101. A polynucleotide according to claim 99, wherein the polynucleotide encodes a polypeptide comprising a humanized antibody heavy chain variable region comprising: (a) a CDR-H1 comprising an amino acid sequence selected from the group consisting of NYAIS, YYAMS, and NYAMS; (b) a CDR-H2 comprising an amino acid sequence selected from the group consisting of TYSSGGVYTYYRDSLKG, TIYSGGSYTFYPDSLEG, TIYSGGGYTFYLDSLKG, and TISSGGGYTYYPDSLKG; and (c) a CDR-H3 comprising an amino acid sequence selected from the group consisting of HGAPMTTVITYAPYYF{D,Y}Y, HGAPMTTVITYAPYYFYY, HGAPMTTVITYAPYYFDY, HGAAMTTVITYAPFYFYY, HGAPMSTEITYAPYYFYY, and HSYPMTTVITYAPYYFYY.
102. The polynucleotide of claim 101, wherein the humanized antibody heavy chain variable region comprises the amino acid sequence: TABLE-US-00004 EV{M, Q}L{V, L}ESGGGLV{K, Q}PGGSL{K, R}LSCAASGFTFSNYAMSWVRQ{T, A}P {E, G}K{R, G}LEWV{A, S}TISSGGGYTYYPDSLKGRFTISRDN{A, S}KNTL{F, Y}LQM{S, N}SLR{S, A}EDTAVYYC{V, A}{R, K}HGAPMTTVITYAPYYF{D, Y}YWGQGTT{L, V}T VSS.
103. The polynucleotide of claim 102, wherein the humanized antibody heavy chain variable region comprises the amino acid sequence: TABLE-US-00005 (SEQ ID NO: 11) EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYAMSWVRQAPKGLEWVSTI SSGGGYTYYPDSLKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKHGA PMTTVITYAPYYFYYWGQGTTVTVSS.
104. The polynucleotide of claim 101, wherein the polynucleotide encodes an amino acid sequence selected from the group consisting of: TABLE-US-00006 (a) EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYAISWVRQAPGKGLEWVS TYSSGGVYTYYRDSLKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAK HGAAMTTVITYAPFYFYYWGQGTTVTVSS; (b) EVQLLESGGGLVQPGGSLRLSCAASGFTFSYYAMSWVRQAPGKGLEWVS TIYSGGSYTFYPDSLEGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAK HGAPMSTEITYAPYYFYYWGQGTTVTVSS; (c) EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYYMSWVRQAPGKGLEWVS TIYSGGGYTFYLDSLKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAK HSYPMTTVITYAPYYFYYWGQGTTVTVSS; (d) EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYAMSWVRQAPGKGLEWVS TISSGGGYTYYPDSLKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAK HGAPMTTVITYAPYYFYYWGQGTTVTVSS; and (e) EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYAMSWVRQAPGKGLEWVS TISSGGGYTYYPDSLKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAK HGAPMTTVITYAPYYFYYWGQGTTVTVSS.
105. A polynucleotide encoding a polypeptide comprising a humanized antibody light chain variable region comprising: (a) a CDR-L1 comprising the amino acid sequence: RAS{S,Q}SV{K,S}SY{M,L}{Y,A}; (b) a CDR-L2 comprising the amino acid sequence: {Y,D}{T,A}SN{L,R}A{P,T}; and (c) a CDR-L3 comprising the amino acid sequence {Q,Y}Q{F,Y}TT{S,Y}PYT.
106. The polynucleotide of claim 105, wherein the humanized antibody light chain variable region comprises an amino acid sequence selected from the group consisting of: TABLE-US-00007 (a) {E, D}{N, V}V{L, M}TQSPAFLSVTPGEKVTITCRAS{S, Q}SV{K, S}SY{M, L}{Y, A} WYQQKPDQAPKL{W, L}I{Y, K}{Y, D}{T, A}SN{L, R}A{P, T}GVPSRFSGSGSG{N, T}D{Y, F}TFTISSLEAEDAATYYC{Q, Y}Q{F, Y}TT{S, Y}PYTFGGGTKLEIKR; and (b) EIVLTQSPATLSLSPGERATLSCRAS{S, Q}SV{K, S}SY{M, L}{Y, A}WYQQKPGQAPRLLIY {Y, D}{T, A}SN{L, R}A{P, T}GIPARFSGSGSGTDFTLTISSLEPEDFAVYYC{Q, Y}Q{F, Y}TT{S, Y}PYTFGGGTKLEIKR
107. A polynucleotide according to claim 105, wherein the polynucleotide encodes a polypeptide comprising a humanized antibody light chain variable region comprising: (a) a CDR-L1 comprising the amino acid sequence RASSSVKYMY; (b) a CDR-L2 comprising the amino acid sequence YTSNLAP; and (c) a CDR-L3 comprising an amino acid sequence selected from the group consisting of QQFTTSPYT, YQFTTSPYT, QQYTTSPYT, and QQFTTYPYT.
108. The polynucleotide of claim 107, wherein the humanized antibody light chain variable region comprises the amino acid sequence: TABLE-US-00008 {E, D}{I, N, V}V{L, M}TQSPA{T, F, I}{L, M}S{L, A, V}{S, T}{L, P}GE{R, K}{A, V}T{L, M, I} {S, T, N}CRASSSVKYMYWYQQK{S, P}{G, D}{Q, A}{A, S}P{R, K}L{L, W}I{Y, K}YTSNLAP G{I, V}P{A, S}RFSGSGSG{T, N}{D, S}{F, Y}{T, S}{L, F}TISS{M, L}E{A, G, P}ED{F, A}A {V, T}YYC{Q, Y}QFTT{S, Y}PYTFGGGTKLEIKR.
109. The polynucleotide of claim 107, wherein the humanized antibody light chain variable region comprises an amino acid sequence selected from the group consisting of TABLE-US-00009 (a) EINVLTQSPATLSLSLGERATLSCRASSSVKYMYWYQQKSGQAPRLLIYYTSNLAPGIPA RFSGSGSGTDFTLTISSMEAEDFAVYYCQQFTTSPYTFGGGTKLEIKR; (b) ENVLTQSPAFLSVTPGEKVTITCRASSSVKYMYWYQQKPDQAPKLWIYYTSNLAPGVPS RFSGSGSGNDYTFTISSLEAEDAATYYCQQFTTSPYTFGGGTKLEIKR; (c) EIVLTQSPATLSLSLGERATLSCRASSSVKYMYWYQQKSGQAPRLLIYYTSNLAPGIPARF SGSGSGTDFTLTISSMEAEDFAVYYCQQFTTSPYTFGGGTKLEIKR; (d) EIVLTQSPATLSLSPGERATLSCRASSSVKYMYWYQQKPGQAPRLLIYYTSNLAPGIPARF SGSGSGTDFTLTISSLEPEDFAVYYCQQFTTSPYTFGGGTKLEIKR; (e) EIVLTQSPATLSLSPGERATLSCRASSSVKYMYWYQQKPGQAPRLLIYYTSNLAPGIPARF SGSGSGTDFTLTISSLEPEDFAVYYCYQFTTSPYTFGGGTKLEIKR; (f) EIVLTQSPATLSLSPGERATLSCRASSSVKYMYWYQQKPGQAPRLLIYYTSNLAPGIPARF SGSGSGTDFTLTISSLEPEDFAVYYCQQYTTSPYTFGGGTKLEIKR); (g) EIVLTQSPATLSLSPGERATLSCRASSSVKYMYWYQQKPGQAPRLLIYYTSNLAPGIPARF SGSGSGTDFTLTISSLEPEDFAVYYCQQFTTYPYTFGGGTKLEIKR; and (h) ENVLTQSPAFLSVTPGEKVTITCRASSSVKYMYWYQQKPDQAPKLWIYYTSNLAPGVPS RFSGSGSGNDYTFTISSLEAEDAATYYCQQFTTSPYTFGGGTKLEIKR.
110. A polynucleotide encoding a polypeptide comprising: (a) a humanized antibody heavy chain variable region comprising: (1) a CDR-H1 comprising the amino acid sequence NYMAS; (2) a CDR-H2 comprising the amino acid sequence TISSGGGYTYYPDSLKG; and (3) a CDR-H3 comprising an amino acid sequence selected from the group consisting of HGAPMTTVITYAPYYF{D,Y}Y), HGAPMTTVITYAPYYFYY, and HGAPMTTVITYAPYYFDY; and (b) a humanized antibody light chain variable region comprising: (1) a CDR-L1 comprising the amino acid sequence RASSSVKYMY; (2) a CDR-L2 comprising the amino acid sequence YTSNLAP; and (3) a CDR-L3 comprising the amino acid sequence QQFTTSPYT, YQFTTSPYT, and QQYTTSPYT.
111. The polynucleotide of claim 110, wherein: (a) the CDR-H1 comprises the amino acid sequence NYMAS; (b) the CDR-H2 comprises the amino acid sequence TISSGGGYTYYPDSLKG; (c) the CDR-H3 comprises the amino acid sequence HGAPMTTVITYAPYYF{D,Y}Y; (d) the CDR-L1 comprises the amino acid sequence RASSSVKYMY; (e) the CDR-L2 comprises the amino acid sequence YTSNLAP; and (f) the CDR-L3 comprises the amino acid sequence QQFTTSPYT.
112. A recombinant polynucleotide comprising a promoter sequence operably linked to the polynucleotide of claim 110.
113. An isolated cell transformed with the polynucleotide of claim 112.
114. A polynucleotide according to claim 110, wherein the polypeptide comprises: (a) a humanized heavy chain variable region comprising the amino acid sequence: TABLE-US-00010 EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYAMSWVRQAPGKGLEWVST ISSGGGYTYYPDSLKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKHG APMTTVITYAPYYFYYWGQGTTVTVSS;
and (b) a humanized light chain variable region comprising the amino acid sequence TABLE-US-00011 EIVLTQSPATLSLSLGERATLSCRASSSVKYMYWYQQKSGQAPRLLIYYTSNLAPGIPARF SGSGSGTDFTLTISSMEAEDFAVYYCQQFTTSPYTFGGGTKLEIKR.
115. A polynucleotide according to claim 110, wherein the polypeptide comprises: (a) a humanized heavy chain variable region comprising the amino acid sequence: TABLE-US-00012 EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYAMSWVRQAPGKGLEWVST ISSGGGYTYYPDSLKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKHG APMTTVITYAPYYFYYWGQGTTVTVSS;
and (b) a humanized light chain variable region comprising the amino acid sequence: TABLE-US-00013 ENVLTQSPAFLSVTPGEKVTITCRASSSVKYMYWYQQKPDQAPKLWIYYT SNLAPGVPSRFSGSGSGNDYTFTISSLEAEDAATYYCQQFTTSPYTFGGG TKLEIKR.
116. A polynucleotide according to claim 110, wherein the polypeptide comprises: (a) a humanized heavy chain variable region comprising the amino acid sequence: TABLE-US-00014 EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYAMSWVRQAPGKGLEWVST ISSGGGYTYYPDSLKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKHG APMTTVITYAPYYFYYWGQGTTVTVSS;
and (b) a humanized light chain variable region comprising the amino acid sequence TABLE-US-00015 EIVLTQSPATLSLSPGERATLSCRASSSVKYMYWYQQKPGQAPRLLIYYTSNLAPGIPARF SGSGSGTDFTLTISSLEPEDFAVYYCQQFTTSPYTFGGGTKLEIKR.
117. A polynucleotide according to claim 110, wherein the polypeptide comprises: (a) a humanized heavy chain variable region comprising the amino acid sequence: TABLE-US-00016 EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYAMSWVRQAPGKGLEWVST ISSGGGYTYYPDSLKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKHG APMTTVITYAPYYFYYWGQGTTVTVSS;
and (b) a humanized light chain variable region comprising the amino acid sequence TABLE-US-00017 EIVLTQSPATLSLSPGERATLSCRASSSVKYMYWYQQKPGQAPRLLIYYTSNLAPGIPARF SGSGSGTDFTLTISSLEPEDFAVYYCYQFTTSPYTFGGGTKLEIKR.
118. A polynucleotide according to claim 110, wherein the polypeptide comprises: (a) a humanized heavy chain variable region comprising the amino acid sequence: TABLE-US-00018 EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYAMSWVRQAPGKGLEWVST ISSGGGYTYYPDSLKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKHG APMTTVITYAPYYFYYWGQGTTVTVSS;
and (b) a humanized light chain variable region comprising the amino acid sequence TABLE-US-00019 EIVLTQSPATLSLSPGERATLSCRASSSVKYMYWYQQKPGQAPRLLIYYTSNLAPGIPARF SGSGSGTDFTLTISSLEPEDFAVYYCQQYTTSPYTFGGGTKLEIKR.
119. A polynucleotide according to claim 110, wherein the polypeptide comprises: (a) a humanized heavy chain variable region comprising the amino acid sequence: TABLE-US-00020 EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYAMSWVRQAPGKGLEWVST ISSGGGYTYYPDSLKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAKHG APMTTVITYAPYYFYYWGQGTTVTVSS;
and (b) a humanized light chain variable region comprising the amino acid sequence TABLE-US-00021 EIVLTQSPATLSLSPGERATLSCRASSSVKYMYWYQQKPGQAPRLLIYYTSNLAPGIPARF SGSGSGTDFTLTISSLEPEDFAVYYCQQFTTYPYTFGGGTKLEIKR.
Description:
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application claims priority to U.S. Provisional Application No. 60/898,709 filed Feb. 1, 2007, the disclosure of which is incorporated herein by reference in its entirety.
BACKGROUND
[0002] The interaction between chemokines and their receptors is an important step in the control of leukocyte migration. Chemokines also mediate a variety of effects independent of chemotaxis, including induction and enhancement of cell-associated cytokine responses.
[0003] The human cell surface protein CD183 is a G protein-coupled receptor with selectivity for three chemokines including IP10 (interferon-g-inducible 10 kDa protein), Mig (monokine induced by interferon-g) and I-TAC (interferon-inducible T cell a-chemoattractant). These three chemokines belong to the structural subfamily of "CXC" chemokines, in which a single amino acid residue separates the first two of four highly conserved Cys residues. Historically, CD183 is the third CXC chemokine receptor discovered and, therefore, CD183 is commonly designated as "CXCR3." Binding of chemokines to CXCR3 induces cellular responses that are involved in leukocyte traffic, most notably integrin activation, cytoskeletal changes and chemotactic migration. CXCR3 is expressed on effector/memory T cells and/or in T cells present in many types of inflamed tissues (e.g., T-helper 1 cells or Th1 cells and CD8+ Tc1 cells). In addition, IP10, Mig and I-TAC are commonly produced by local cells in inflammatory lesions, suggesting that CXCR3 and its chemokines participate in the recruitment of white blood cells to sites of inflammation. Therefore, CXCR3 is a target for the development of antibodies and antagonists, which may be used in the treatment and diagnosis of diverse inflammatory and immune diseases and disorders, such as rheumatoid arthritis, multiple schlerosis, Crohn's disease, inflammatory bowel disease, chronic obstructive pulmonary disease, psoriasis, type 1 diabetes and transplant rejection. Because CXCR3 is expressed on a subset of B-cell lymphomas, CXCR3 may also be a target for treating and diagnosing lymphomas and leukemias.
SUMMARY
[0004] Disclosed are antigen-binding polypeptide molecules that bind specifically to the chemokine receptor CXCR3 (see GenBank gi:4504099). The polypeptides include a humanized heavy chain variable region and a humanized light chain variable region. For example, the polypeptides may include the framework (FR) regions of the light and heavy chain variable regions of a human antibody, while retaining substantially the antigen-binding specificity of a parental monoclonal antibody. The humanized heavy chain variable region and/or the humanized light chain variable region are at least about 90% humanized (preferably at least about 95% humanized, more preferably at least about 98% humanized, and even more preferably at least about 100% humanized), excluding the CDRs. The antigen-binding polypeptides molecules may be derived from monoclonal antibody donors (e.g., mouse monoclonal antibody donors) and may include CDRs from the monoclonal antibodies (e.g., mouse monoclonal CDRs). The polypeptides may function as antagonists for the CXCR3 receptor.
[0005] In some embodiments, the antigen-binding polypeptide binds specifically to CXCR3, and includes: (a) a humanized antibody heavy chain variable region comprising: (1) a CDR-H1 comprising an amino acid sequence of (NYMAS); (2) a CDR-H2 comprising an amino acid sequence of (TISSGGGYTYYPDSLKG); and (3) a CDR-H3 comprising an amino acid sequence of (HGAPMTTVITYAPYYF{D,Y}Y); and (b) a humanized antibody light chain variable region comprising: (1) a CDR-L1 comprising an amino acid sequence of (RASSSVKYMY); (2) a CDR-L2 comprising an amino acid sequence of (YTSNLAP); and (3) a CDR-L3 comprising an amino acid sequence of (QQFTTSPYT). The polypeptide may include a CDR-H3 comprising an amino acid sequence of (HGAPMTTVITYAPYYFYY).
[0006] In some embodiments of the polypeptides, (1) the CDR-H1 consists of the amino acid sequence of (NYMAS); (2) the CDR-H2 consists of the amino acid sequence of (TISSGGGYTYYPDSLKG); (3) the CDR-H3 consists of the amino acid sequence of (HGAPMTTVITYAPYYF{D,Y}Y); (4) the CDR-L1 consists of the amino acid sequence of (RASSSVKYMY); (5) the CDR-L2 consists of the amino acid sequence of (YTSNLAP); and (6) the CDR-L3 consists of the amino acid sequence of (QQFTTSPYT). For example, the CDR-H3 may consist of the amino acid sequence of (HGAPMTTVITYAPYYFYY).
[0007] In some embodiments, the polypeptide comprises a humanized antibody heavy chain variable region of ({E,D}{I,N,V}V{L,M}TQSPA{T,F,I}{L,M}S{L,A,V}{S,T}{L,P}GE{R,K}{A,V}T{L,M,I- }{S,T,N}CRASSSVKYMYWYQQK{S,P}{G,D}{Q,A}{A,S}P{R,K}L{L,W}I{Y,K}YTSNLAPG{I,V- }P{A,S}RFSGSGSG{T,N}{D,S}{F,Y}{T,S}{L,F}TISS{M,L}E{A,G,P}ED{F,A}A{V,T}YYC{- Q,Y}QFTT{S,Y}PYTFGGGTKLEIKR). For example, the polypeptide may comprise a humanized antibody heavy chain variable region of (EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYAMSWVRQAPKGLEWVSTISSGGGYTYYPDSLKGRFTISRD- NSKNTLYLQMNSLRAEDTAVYYCAKHGAPMTTVITYAPYYFYYWGQGTTVTVSS). In some embodiments, the polypeptide comprises a humanized antibody light chain variable region of (E {I,N}VLTQSPA{T,F,I}{L,M}S{L,A,V}{S,T}{L,P}GE{R,K}{A,V}T{L,M,I}{S,T,N}CRAS- SSVKYMYWYQQK{S,P}{G,D}{Q,A}{A,S}P{R,K}L{L,W}IYYTSNLAPG{I,V}P{A,S}RFSGSGSG{- T,N}{D,S}{F,Y}{T,S}{L,F}TISS{M,L}E{A,G}ED{F,A}A{V,T}YYCQQFTTSPYTFGGGTKLEIK- R). For example, the polypeptide may comprise a humanized antibody light chain variable region of (EIVLTQSPATLSLSLGERATLSCRASSSVKYMYWYQQKSGQAPRLLIYYTSNLAPGIPARFSGSGSGTDFTL- TISSMEAEDFAVYYCQQFTTSPYTFGGGTKLEIKR); or (ENVLTQSPAFLSVTPGEKVTITCRASSSVKYMYWYQQKPDQAPKLWIYYTSNLAPGVPSRFSGSGSGNDYTF- TISSLEAEDAATYYCQQFTTSPYTFGGGTKLEIKR).
[0008] Also disclosed are humanized antibody heavy chain variable regions. The humanized antibody heavy chain region may comprise: (1) a CDR-H1 comprising an amino acid sequence of ({N,S,Y}YAMS); (2) a CDR-H2 comprising an amino acid sequence of ({T,A,Y}I{S,Y}{S,G,T,Y}{G,S}{G,Y}G{F,S,Y}TYY{P,A}DS{L,Y,V}KG); and (3) a CDR-H3 comprising an amino acid sequence of {H,Y}{G,Y}{A,Y}PM{T,Y}T{V,Y}ITY{A,Y}PYYFYY). For example, the humanized antibody heavy chain variable region may comprise an amino acid sequence of (EVQLLESGGGLVQPGGSLRLSCAASGFTFS{N,S,Y}YAMSWVRQAPGKGLEWVS{T,A,Y}I{S,Y}{- S,G,T,Y}{G,S}{G,Y}G{F,S,Y}TYY{P,A}DS{L,Y,V}KGRFTISRDNSKNTLYLQMNSLRAEDTAVYY- CAK{H,Y}{G,Y}{A,Y}PM{T,Y}T{V,Y}ITY{A,Y}PYYFYYWGQGTTVTVSS).
[0009] In another example, a humanized antibody heavy chain variable region comprises: (1) a CDR-H1 comprising an amino acid sequence of (NYAIS); (2) a CDR-H2 comprising an amino acid sequence of (TYSSGGVYTYYRDSLKG); and (3) a CDR-H3 comprising an amino acid sequence of (HGAAMTTVITYAPFYFYY). For example, the humanized antibody heavy chain variable region may comprise an amino acid sequence of (EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYAISWVRQAPGKGLEWVSTYSSGGVYTYYRDSLKGRFTISR- DNSKNTLYLQMNSLRAEDTAVYYCAKHGAAMTTVITYAPFYFYYWGQGTTVTVSS).
[0010] In another example, a humanized antibody heavy chain variable region comprises: (1) a CDR-H1 comprising an amino acid sequence of (YYAMS); (2) a CDR-H2 comprising an amino acid sequence of (TIYSGGSYTFYPDSLEG); and (3) a CDR-H3 comprising an amino acid sequence of (HGAPMSTEITYAPYYFYY). For example, the humanized antibody heavy chain variable region may comprise an amino acid sequence of (EVQLLESGGGLVQPGGSLRLSCAASGFTFSYYAMSWVRQAPGKGLEWVSTIYSGGSYTFYPDSLEGRFTISR- DNSKNTLYLQMNSLRAEDTAVYYCAKHGAPMSTEITYAPYYFYYWGQGTTVTVSS).
[0011] In another example, a humanized antibody heavy chain variable region comprises: (1) a CDR-H1 comprising an amino acid sequence of (NYAMS); (2) a CDR-H2 comprising an amino acid sequence of (TIYSGGGYTFYLDSLKG); and (3) a CDR-H3 comprising an amino acid sequence of (HSYPMTTVITYAPYYFYY). For example, the humanized antibody heavy chain variable region may comprise an amino acid sequence of (EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYYMSWVRQAPGKGLEWVSTIYSGGGYTFYLDSLKGRFTISR- DNSKNTLYLQMNSLRAEDTAVYYCAKHSYPMTTVITYAPYYFYYWGQGTTVTVSS).
[0012] In another example, a humanized antibody heavy chain variable region comprises: (1) a CDR-H1 comprising an amino acid sequence of (NYAMS); (2) a CDR-H2 comprising an amino acid sequence of (TISSGGGYTYYPDSLKG); and (3) a CDR-H3 comprising an amino acid sequence of (HGAPMTTVITYAPYYFYY). For example, the humanized antibody heavy chain variable region may comprise an amino acid sequence of (EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYAMSWVRQAPGKGLEWVSTISSGGGYTYYPDSLKGRFTISR- DNSKNTLYLQMNSLRAEDTAVYYCAKHGAPMTTVITYAPYYFYYWGQGTTVTVSS).
[0013] In another example, a humanized antibody heavy chain variable region comprises: (1) a CDR-H1 comprising an amino acid sequence of (NYAMS); (2) a CDR-H2 comprising an amino acid sequence of (TISSGGGYTYYPDSLKG); and (3) a CDR-H3 comprising an amino acid sequence of (HGAPMTTVITYAPYYFYY). For example, the humanized antibody heavy chain variable region may comprise an amino acid sequence (EVQLLESGGGLVQPGGSLRLSCAASGFTFSNYAMSWVRQAPGKGLEWVSTISSGGGYTYYPDSLKGRFTISR- DNSKNTLYLQMNSLRAEDTAVYYCAKHGAPMTTVITYAPYYFYYWGQGTTVTVSS).
[0014] Also disclosed are humanized antibody light chain variable regions. The humanized antibody light chain region may comprise: (1) a CDR-L1 comprising an amino acid sequence of (RASSSVKYMY); (2) a CDR-L2 comprising an amino acid sequence of (YTSNLAP); and (3) a CDR-L3 comprising an amino acid sequence of (QQFTTSPYT). For example, the humanized antibody light chain variable region may comprise an amino acid sequence of (EIVLTQSPATLSLSLGERATLSCRASSSVKYMYWYQQKSGQAPRLLIYYTSNLAPGIPARFSGSGSGTDFTL- TISSMEAEDFAVYYCQOFTTSPYTFGGGTKLEIKR).
[0015] In another example, a humanized antibody light chain variable region comprises: (1) a CDR-L1 comprising an amino acid sequence of (RASSSVKYMY); (2) a CDR-L2 comprising an amino acid sequence of (YTSNLAP); and (3) a CDR-L3 comprising an amino acid sequence of (QQFTTSPYT). For example, the humanized antibody light chain variable region may comprise an amino acid sequence of (EIVLTQSPATLSLSPGERATLSCRASSSVKYMYWYQQKPGQAPRLLIYYTSNLAPGIPARFSGSGSGTDFTL- TISSLEPEDFAVYYCQQFTTSPYTFGGGTKLEIKR).
[0016] In another example, a humanized antibody light chain variable region comprises: (1) a CDR-L1 comprising an amino acid sequence of (RASSSVKYMY); (2) a CDR-L2 comprising an amino acid sequence of (YTSNLAP); and (3) a CDR-L3 comprising an amino acid sequence of (YQFTTSPYT). For example, the humanized antibody light chain variable region may comprise an amino acid sequence of (EIVLTQSPATLSLSPGERATLSCRASSSVKYMYWYQQKPGQAPRLLIYYTSNLAPGIPARFSGSGSGTDFTL- TISSLEPEDFAVYYCYQFTTSPYTFGGGTKLEIKR).
[0017] In another example, the humanized antibody light chain variable region comprises: (1) a CDR-L1 comprising an amino acid sequence of (RASSSVKYMY); (2) a CDR-L2 comprising an amino acid sequence of (YTSNLAP); and (3) a CDR-L3 comprising an amino acid sequence of (QQYTTSPYT). For example, the humanized antibody light chain variable region may comprise an amino acid sequence of (EIVLTQSPATLSLSPGERATLSCRASSSVKYMYWYQQKPGQAPRLLIYYTSNLAPGIPARFSGSGSGTDFTL- TISSLEPEDFAVYYCQQYTTSPYTFGGGTKLEIKR).
[0018] In another example, the humanized antibody light chain variable region comprises: (1) a CDR-L1 comprising an amino acid sequence of (RASSSVKYMY); (2) a CDR-L2 comprising an amino acid sequence of (YTSNLAP); and (3) a CDR-L3 comprising an amino acid sequence of (QQFTTYPYT). For example, the humanized antibody light chain variable region may comprise an amino acid sequence of (EIVLTQSPATLSLSPGERATLSCRASSSVKYMYWYQQKPGQAPRLLIYYTSNLAPGIPARFSGSGSGTDFTL- TISSLEPEDFAVYYCQQFTTYPYTFGGGTKLEIKR).
[0019] In another example, the humanized antibody light chain variable region comprises: (1) a CDR-L1 comprising an amino acid sequence of (RAS{S,Q}SV{K,S}SY{M,L}{Y,A}); (2) a CDR-L2 comprising an amino acid sequence of ({Y,D}{T,A}SN{L,R}A{P,T}); and (3) a CDR-L3 comprising an amino acid sequence of (Q,Y}Q{F,Y}TT{S,Y}PYT). For example, the humanized antibody light chain variable region may comprise an amino acid sequence of (EIVLTQSPATLSLSPGERATLSCRAS{S,Q}SV{K,S}SY{M,L}{Y,A}WYQQKPGQAPRLLIY{Y,D- }{T,A}SN{L,R}A{P,T}GIPARFSGSGSGTDFTLTISSLEPEDFAVYYC{Q,Y}Q{F,Y}TT{S,Y}PYTFG- GGTKLEIKR).
[0020] In another example, the humanized antibody light chain variable region comprises: (1) a CDR-L1 comprising an amino acid sequence of (RASSSVKYMY); (2) a CDR-L2 comprising an amino acid sequence of (YTSNLAP); and (3) a CDR-L3 comprising an amino acid sequence of (QQFTTSPYT). For example, the humanized antibody light chain variable region may comprise an amino acid sequence of (ENVLTQSPAFLSVTPGEKVTITCRASSSVKYMYWYQQKPDQAPKLWIYYTSNLAPGVPSRFSGSGSGNDYTF- TISSLEAEDAATYYCQQFTTSPYTFGGGTKLEIKR).
[0021] In another example, the humanized antibody light chain variable region comprises: (1) a CDR-L1 comprising an amino acid sequence of (RAS{S,Q}SV{K,S}SY{M,L}{Y,A}); (2) a CDR-L2 comprising an amino acid sequence of ({Y,D}{T,A}SN{L,R}A{P,T}); and (3) a CDR-L3 comprising an amino acid sequence of (Q,Y}Q{F,Y}TT{S,Y}PYT). For example, the humanized antibody light chain variable region may comprise an amino acid sequence of ((E,D)(N,V)V(L,M)TQSPAFLSVTPGEKVTITCRASSSVKYMYWYQQKPDQAPKL(W,L)I(Y,K)Y- TSNLAPGVPSRFSGSGSG(N,T)D(Y,F)TFTISSLEAEDAATYYC(Q,Y)Q(F,Y)TT(S,Y)PYTFGGGTKL- EIKR).
[0022] The aforementioned humanized heavy chains and humanized light chains may be present in the antigen binding polypeptides that binds specifically to CXCR3.
[0023] The antigen-binding polypeptide may be selected from the group consisting of an antibody molecule, a Fab fragment, a Fab' fragment, a F(ab')2 fragment, and an scFv molecule. In some embodiments, polypeptide is an antibody molecule. Antibody molecules may include chimeric antibodies that include a human heavy chain constant region and a human light chain constant region. For example, the antibody molecule may be an IgG molecule (e.g., a IgG1 or an IgG4 molecule), where the polypeptide includes the heavy chain and light chain constant domains of an IgG molecule. The polypeptide may be an scFv molecule. For example, the scFv may have a formula selected from the group consisting of NH2-L-VH-X-VK-COOH and NH2-L-VK-X-VH-COOH; wherein L is a leader sequence; VH is the humanized antibody heavy chain variable region; X is a linking polypeptide; and VK is the humanized antibody light chain variable region.
[0024] The antigen-binding polypeptide further may be conjugated or fused to a therapeutic or diagnostic agent. For example, therapeutic agents may be selected from the group consisting of a cytotoxic agent, a radioactive label, an immunomodulator, a hormone, an enzyme, an oligonucleotide, a photoactive therapeutic agent or a combination thereof. Examples of diagnostic agents may include a radioactive label, a photoactive diagnostic agent, an ultrasound-enhancing agent or a non-radioactive label.
[0025] The antigen-binding polypeptide may be an antagonist of CXCR3. Typically, the polypeptide is not an agonist of CXCR3.
[0026] The antigen-binding polypeptide binds to the CXCR3 receptor with specificity and high affinity. Typically, the polypeptide binds to CXCR3 with an affinity constant of at least about 106M-1 (preferably at least about 107M-1, more preferably at least about 108M-1, even more preferably at least about 109M-1).
[0027] Also disclosed are pharmaceutical compositions comprising the aforementioned antigen-binding polypeptides and a carrier (e.g., a diluent or excipient). The pharmaceutical may further comprise an additional therapeutic or diagnostic agent as disclosed herein.
[0028] Also disclosed are methods of treating or diagnosing a disease or condition that comprise administering the disclosed pharmaceutical compositions to a patient in need thereof. For example, the pharmaceutical compositions may be administered to treat or diagnose an inflammatory, immune, and/or malignant disease or condition. Examples of diseases and conditions may include autoimmune disease (e.g., lupus), inflammatory bowel disease (IBD), chronic obstructive pulmonary disease (COPD), arthritis (e.g., rheumatoid arthritis), multiple sclerosis, transplant rejection, central nervous system injury, Crohn's disease, psoriasis, type 1 diabetes and leukemia or lymphoma (e.g., chronic lymphocytic leukemia (CLL)).
[0029] Also disclosed are polynucleotides that encode the aforementioned polypeptides. The polynucleotides may be operably linked to a promoter for expressing the encoded polypeptides in a suitable host cell. As such, methods of producing the polypeptide encoded by the recombinant polynucleotide may include: a) culturing a cell transformed with the recombinant polynucleotide to express the encoded polypeptide; and b) recovering the polypeptide so expressed.
BRIEF DESCRIPTION OF THE DRAWINGS
[0030] FIG. 1 illustrates inhibition of 125I-IP-10 binding to Th1 cells by murine anti-CXCR3 mAb. Ab#1-5D4A; Ab#2-8A5A; Ab#3-19G2; Ab#4-V36E5A; Ab#5-V44D7A;
[0031] Ab#6-37B5A; Ab#7-21A4A; Ab#8-V15F4A; Ab#9-V3G6A; Ab#10-23E12A; Ab#11-35C4; Ab#12-39E10.
[0032] FIG. 2 illustrates inhibition of IP-10-induced Th1 cell migration by murine anti-CXCR3 mAb. Ab#1-5D4A; Ab#2-8A5A; Ab#3-19G2; Ab#4-V36E5A; Ab#5-V44D7A; Ab#6-37B5A; Ab#7-21A4A; Ab#8-V15F4A; Ab#9-V3G6A; Ab#10-23E12A.
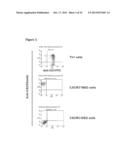
[0033] FIG. 3 illustrates a FACs analysis of murine anti-CXCR3 mAb binding to Th1 cells (top panel), CXCR3+/NSO cells (middle panel), and CXCR-/NSO cells (bottom panel).
[0034] FIG. 4 illustrates inhibition of chemokine binding to CXCR3 by murine anti-CXCR3 mAb and humanized anti-CXCR3 mAb.
[0035] FIG. 5 illustrates inhibition of chemokine mediated chemotaxis by murine anti-CXCR3 mAb and humanized anti-CXCR3 mAb.
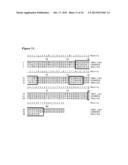
[0036] FIG. 6 illustrates an alignment of the VH Domains of 5 anti-CXCR3 Clones.
[0037] FIG. 7 illustrates an alignment of the VK Domains of 5 anti-CXCR3 Clones.
[0038] FIG. 8 illustrates an alignment of the VH Domain of anti-CXCR3 Clone V44D7 with the closest expressed human IgG and germline VH.
[0039] FIG. 9 illustrates the risk assessment of amino acid changes required for complete humanization of the VH domain of anti-CXCR3 clone V44D7. The required amino acid changes are indicated below the main sequence and were derived from an alignment to human VH3-23. The germline gene and an expressed antibody are described in GenBank accession no. AAD53829.
[0040] FIG. 10 illustrates the risk assessment of amino acid changes required for complete humanization of the VH domain of anti-CXCR3 clone V44D7. The required amino acid changes are indicated below the main sequence and were derived from an alignment to human VH3-23. The germline gene and an expressed antibody are described in GenBank accession no. AAD53829.
[0041] FIG. 11 illustrates an alignment of the VK domain of anti-CXCR3 clone V3G6 with the closest expressed human IgG and germiline VK.
[0042] FIG. 12 illustrates the risk assessment of amino acid changes required for complete humanization of the VK domain of anti-CXCR3 clone V3G6.
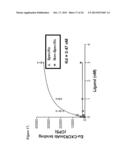
[0043] FIG. 13 illustrates inhibition of mouse CXCR3 mAb binding to CXCR3+ NSO cells by commercial CXCR3 mAbs. Approximately 0.5 nM Eu-CXCR3 mAb was incubated with CXCR3 transfected NSO cells in the presence of various concentrations of unlabeled commercial CXCRmAbs. A dose-dependent inhibition of Eu-CXCR3 mAb binding to CXCR3+ NSO cells was observed.
[0044] FIG. 14 illustrates expression of CXCR3 on Th1 cells. Th1 and Th2 cells were generated from cord blood and CXCR3 and CCR4 expression were determined by FACS. CXCR3 was present only Th1 cells.
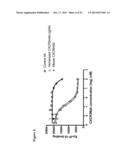
[0045] FIG. 15 illustrates an 125I-CXCL10 binding assay. Th1 cells were incubated in a 96 well plate with 125I-CXCL10 in the absence or presence of various concentrations CXCRmAbs. Cell bound 125I-CXCL10 was separated from free radioactivity by an oil column and counted using a gamma counter. IC50 values were calculated using Prizm software. Lead candidates were highlighted in green.
[0046] FIG. 16 illustrates an 125I-CXCL11 binding assay. Th1 cells were incubated in a 96 well plate with 125I-CXCL11 in the absence or presence of various concentrations CXCRmAbs. Cell bound 125I-CXCL11 was separated from free radioactivity by an oil column and counted using a gamma counter. IC50 values were calculated using Prizm software. Lead candidates were highlighted in green.
[0047] FIG. 17 illustrates Eu-CXCR3 mAb binding to Th1 cells. Th1 cells were incubated with increasing concentrations of Eu-CXCR3 mAb in the absence or presence 10-fold excess of unlabeled CXCR3 mAb. After incubation (1 hr at RT), cell bound Eu-CXCR3 mAb was separated from free Europium by washing three times and the plate was read using Vctor2 fluorometer.
[0048] FIG. 18 illustrates inhibition of 125I-CXCL11 binding to Th1 by CXCR3 mAb hybridoma supernatants. Th1 cells were incubated in a 96 well plate with 125I-CXCL11 in the absence or presence of various CXCRmAb hybridoma supernatants for 1 hr at RT. Cell bound 125I-ligands were separated from free radioactivity by an oil column and counted using a gamma counter. Seven hybridoma supernatants that inhibited CXCL11 binding to Th1 cells were selected for further development.
[0049] FIG. 19 illustrates inhibition of 125I-CXCL10 binding to Th1 by CXCR3 mAb hybridoma supernatants. Th1 cells were incubated in a 96 well plate with 125I-CXCL10 in the absence or presence of various CXCRmAb hybridoma supernatants for 1 hr at RT. Cell bound 125I-ligands were separated from free radioactivity by an oil column and counted using a gamma counter. Seven hybridoma supernatants that inhibited CXCL10 binding to Th1 cells were selected for further development.
[0050] FIG. 20 illustrates that Mouse CXCR3 mAb does not cross react with rat Th1 cells. FACS analysis was performed to determine reactivity of mouse CXCR3 mAb to polarized rat Th1 cells. Only rabbit anti-mouse CXCR3Ab bound to rat Th1 cells. Mouse anti-human CXCR3Ab did not bind to rat Th1 cells. As a control, mouse anti-CXCR3 mAb binding to human Th1 cells is also shown (bottom panel).
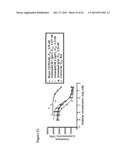
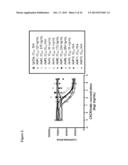
[0051] FIG. 21 illustrates inhibition Eu-CXCR3 mAb by humanized CXCR3Abs. Th1 cells were incubated in a 96 well plate with Eu-CXCR3 mAb in the absence or presence of various concentrations humanized CXCRmAbs. After incubation (1 hr at RT), cell bound Eu-CXCR3 mAb was separated from free Europium by washing three times and the plate was read using Vctor2 fluorometer. IC50 values were calculated using Prizm software. Ab1 has a heavy chain sequence of humanized anti-CXCR3 V44D7 VH Lead #5 amino acid and a light chain sequence of humanized anti-CXCR 3 V44D7 VK Lead #1 (see Informal Sequence Listing) in an IgG1 backbone. Ab2 has a heavy chain sequence of humanized anti-CXCR3 V44D7 VH Lead #5 and a light chain sequence of humanized anti-CXCR3 V44D7 VK Lead #7 (see Informal Sequence Listing) in an IgG1 backbone. Humanized Ab (IgG4) has a heavy chain sequence of humanized anti-CXCR3 V44D7 VH Lead #5 and a light chain sequence of the original mouse anti-CXCR3 V44D7 VK in an IgG4 backbone.
[0052] FIG. 22 illustrates inhibition Eu-CXC10 by humanized CXCR3Abs. Th1 cells were incubated in a 96 well plate with Eu-CXCL10 in the absence or presence of various concentrations humanized CXCRmAbs. After incubation (1 hr at RT), cell bound Eu-CXCL10 was separated from free Europium by washing three times and the plate was read using Vctor2 fluorometer. IC50 values were calculated using Prizm software.
[0053] FIG. 23 illustrates inhibition CXCL10-induced Th1 cell chemotaxis by humanized CXCR3Abs. Chemotaxis assay was performed in a ChemoTx 96-well plate (Neuro Probe, Inc). Approximately 29 μL of CXCL10 or buffer control was added to the bottom wells. 25 μL of Th1 cell suspension in the absence or presence of various concentrations of humanized antibodies was added directly on the wells of the filter. After 2 hr incubation at 37° C., cells migrated to the bottom wells were determined by cell titer glo method (Promega).
[0054] FIG. 24 illustrates an analysis of Ca++ flux in Th1 cells. Th1 cells were loaded with Fluo-4,AM (Molecular Probes) and stimulated with various mouse CXCR3 mAb antibodies as indicated. Increase in intracellular Ca++ was determined FLIPR.
DETAILED DESCRIPTION OF THE PREFERRED EMBODIMENTS
Definitions
[0055] An antibody, as described herein, refers to a full-length (i.e., naturally occurring or formed by normal immunoglobulin gene fragment recombinatorial processes) immunoglobulin molecule (e.g., an IgG antibody) or an immunologically active (i.e., specifically binding) portion of an immunoglobulin molecule, like an antibody fragment.
[0056] An antibody fragment is a portion of an antibody such as F(ab')2, F(ab)2, Fab', Fab, Fv, scFv and the like. Regardless of structure, an antibody fragment binds with the same antigen that is recognized by the intact antibody. The term "antibody fragment" also includes any synthetic or genetically engineered protein that acts like an antibody by binding to a specific antigen to form a complex. For example, antibody fragments include isolated fragments consisting of the variable regions, such as the "Fv" fragments consisting of the variable regions of the heavy and light chains, recombinant single chain polypeptide molecules in which light and heavy variable regions are connected by a peptide linker ("scFv proteins"), and minimal recognition units consisting of the amino acid residues that mimic the hypervariable region.
[0057] A humanized antibody is a recombinant protein in which the CDRs from an antibody from one species, e.g., a rodent antibody, are transferred from the heavy and light variable chains of the rodent antibody into human heavy and light variable domains or heavy and light variable domains that have been mutagenized to include at least a portion of the amino acid sequence of the human heavy and light variable domains (as represented by "percent humanization"). The constant domains of the antibody molecule may be derived from those of a human antibody.
[0058] As used herein, "percent humanization" is calculated by determining the number of framework amino acid differences (i.e., non-CDR difference) between the humanized domain and the germline domain, subtracting that number from the total number of amino acids, and then dividing that by the total number of amino acids and multiplying by 100.
[0059] As used herein, "CDR" means a "complementarity determining region" that is present in a variable domain of an antibody heavy chain (VH) or a variable domain of an antibody light chain (VL or VK). Each variable domain includes three CDRs which are designated CDR-H1, CDR-H2, and CDR-H3, for those present in the heavy chain variable domain, and CDR-L1, CDR-L2, and CDR-L3 for those present in the light chain variable domain. The Kabat numbering system is used herein. As such, CDR-H1 begins at approximately amino acid 31 (i.e., approximately 9 residues after the first cysteine residue), includes approximately 5-7 amino acids, and ends at the next tyrosine residue. CDR-H2 begins at the fifteenth residue after the end of CDR-H1, includes approximately 16-19 amino acids, and ends at the next arginine or lysine residue. CDR-H3 begins at approximately the thirty third amino acid residue after the end of CDR-H2; includes 3-25 amino acids; and ends at the sequence W-G-X-G, where X is any amino acid. CDR-L1 begins at approximately residue 24 (i.e., following a cysteine residue); includes approximately 10-17 residues; and ends at the next tyrosine residue. CDR-L2 begins at approximately the sixteenth residue after the end of CDR-L1 and includes approximately 7 residues. CDR-L3 begins at approximately the thirty third residue after the end of CDR-L2; includes approximately 7-11 residues and ends at the sequence F-G-X-G, where X is any amino acid.
[0060] The antigen-binding polypeptides disclosed herein may be conjugated or fused to a therapeutic agent, which may include radioactive labels, an immunomodulator, a hormone, a photoactive therapeutic agent, a cytotoxic agent, which may be a drug or a toxin, and a combination thereof. Drugs may include those drugs that possess the pharmaceutical property selected from the group consisting of antimitotic, antikinase, alkylating, antimetabolite, antibiotic, alkaloid, antiangiogenic, apoptotic agents and combinations thereof. More specifically, these drugs are selected from the group consisting of nitrogen mustards, ethylenimine derivatives, alkyl sulfonates, nitrosoureas, triazenes, folic acid analogs, COX-2 inhibitors, pyrimidine analogs, purine analogs, antibiotics, enzymes, epipodophyllotoxins, platinum coordination complexes, vinca alkaloids, substituted ureas, methyl hydrazine derivatives, adrenocortical suppressants, antagonists, endostatin, taxols, camptothecins, anthracyclines, taxanes, and their analogs, and a combination thereof. The toxins encompassed by the present invention may be selected from the group consisting of ricin, abrin, alpha toxin, saporin, ribonuclease (RNase), e.g., onconase, DNase I, Staphylococcal enterotoxin-A, pokeweed antiviral protein, gelonin, diphtherin toxin, Pseudomonas exotoxin, and Pseudomonas endotoxin.
[0061] Immunomodulators may be selected from the group consisting of a cytokine, a stem cell growth factor, a lymphotoxin, a hematopoietic factor, a colony stimulating factor (CSF), an interferon (IFN), erythropoietin, thrombopoietin and a combination thereof. Specifically useful are lymphotoxins such as tumor necrosis factor (TNF), hematopoietic factors, such as interleukin (IL), colony stimulating factor, such as granulocyte-colony stimulating factor (G-CSF) or granulocyte macrophage-colony stimulating factor (GM-CSF)), interferon, such as interferons-alpha, -beta, or -gamma, and stem cell growth factor, such as designated "S1 factor". More specifically, immunomodulators may include IL-1, IL-2, IL-3, IL-6, IL-10, IL-12, IL-18, IL-21 interferon-gamma, TNF-alpha or a combination thereof.
[0062] The antigen-binding polypeptides disclosed herein may be conjugated or fused to a diagnostic agent. Diagnostic agents may include photoactive diagnostic agents or radiolabels having an energy between 60 and 4,000 keV, or a non-radioactive label. The radioactive label is preferably a gamma-, beta-, and positron-emitting isotope and is selected from the group consisting of 125I, 131I, 123I, 124I, 86Y, 186Re, 188Re, 62Cu, 64Cu, 111In, 67Ga, 68Ga, 99mTc, 94mTc, 18F, 11C, 13N, 15O, 76Br and combinations thereof. Diagnostic agents may include contrast agents, for example, such as manganese, iron or gadolinium.
EXAMPLES
Isolation of Murine IgG1,k CXCR-3 Binding Antibody Using the Hybridoma Technology
[0063] BALB/c mice were immunized with CXCR-3 expressing NSO cells. In a typical procedure 5×106 cells in 50 ul of RIBI adjuvant (Sigma) were injected into rear footpads (25 ul per pad). Two additional injections in RIBI adjuvant were given at 2 week intervals followed by a final boost in PBS. Three days later mice were sacrificed, their poplietal lymph nodes were harvested and lymphocytes isolated for fusion. Lymphocytes were fused with P3X63Ag8.653 plasmacytoma cells at 5:1 ratio using PEG/DMSO (Sigma) as a fusion agent. After fusion cells were resuspended in selective HAT media and seeded at 106 cells per well in 96 well plates. The supernatants from hybridomas that survived HAT selection were screened by ELISA for the presence of mouse IgG. The IgG producing hybridomas were identified and their supernatants were further screened by FACS analysis for antibodies binding to CXCR3 expressing NSO cells (CXCR3+NSO). The hybridomas identified as positives for CXCR3+NSO cell binding were then screened for differential binding to CXCR3+NSO and PC-NSO (vector control) cells in order to identify CXCR3 specific clones. The CXCR3 specific hybridomas were subcloned twice by limiting dilutions. Hybridoma subclones were expanded in serum-free medium, the antibodies were purified on Protein-A column and further characterized in order to pick the lead candidate.
Humanization Strategy
[0064] One goal in humanizing the anti-CXCR3 antibodies was to obtain 60-80% humanized VH and VK domains that retain 90-100% of original binding affinity and specificity. Site-directed mutagenesis of individual high risk positions in VH and VK was used to further humanize the antibodies while maintaining binding affinity and specificity.
[0065] Humanization was performed by CDR grafting and structure based analysis and variable region resurfacing. (See Jones et al., NATURE (1986) May 29-Jun. 4; 321(6069):522-5; Roguska et al., PROTEIN ENGINEERING, 1996, 9(10):895-904; and Xoma, Humanizing Mouse Antibody Frameworks While Preserving 3-D Structure. PROTEIN ENGINEERING, 1994, Vol. 7, pg 805). The primary antibody sequence and 3-D structure data were utilized to identify key framework residues required to maintain the binding affinity and specificity. The 3-D structures of nine (9) different Fab and IgG molecules were analyzed (human and mouse, with or without ligand). After aligning the mouse anti-CXCR3 V44 VH and VK to the nearest human germline genes, the amino acid at every position was evaluated for potential influence on binding and immunogenicity. This information was used to assign a low, moderate, or high risk value for mutation at each position. In general, only the low and moderate risk positions were mutated while avoiding the high risk positions.
[0066] The heavy chain was 98% humanized relative to the mouse heavy chain (excluding CDR's) after this process. An affinity maturation strategy was then performed by incorporating tyrosines pair wise at each position in CDR3, including a Y115D substitution, which gave on average a 2-fold increase in affinity. The heavy chain that was used in the 2 lead candidates included 2 additional mutations at positions 97 and 98 making it 100% human, excluding the CDR's. Following the same strategy for the light chain, the VK was aligned to the A14 germline gene and low and moderate risk positions were mutated. After determining that this germline gene appears to be rarely expressed in normal humans, the process was repeated using the L6 germline as template
[0067] The "Blast for Ig sequences" website sponsored by the NCBI was used to identify the closest match to the mouse VH and VK region used in the study. The V-base website at the MRC was used to confirm the human germline sequences.
[0068] Human germline VH and VK genes were chosen as the best matches to the mouse sequence VH and VK sequences. For the mouse VH sequence, the human germline sequence VH3-23 (as designated in V-base) was identified as the best match: VH3-23 germline (EVQLLESGGGLVQPGGSLRLSCAASGFTFSSYAMSWVRQAPGKGLEQVSAIS GSGGSTYYADSVKGRFTISRDNSKNTLYLQMNSLRAEDTAVYYCAK). For the mouse VK sequence, the human germline sequence A14 and L6 (as designated in V-base) were identified as the best matches: L6 Germline (EIVLTQSPATLSLSPGERATLSCRASQSVSSYLAWYQQKPGQAPRLLIYDASN RATGIPARFSGSGSGTDFTLTISSLEPEDFAVYYCQQRSNWP); and A14 Germline (DVVMTQSPAFLSVTPGEKVTITCQASEGIGNYLYWYQQKPDQAPKLLIKYAS QSISGVPSRFSGSGSGTDFTFTISSLEAEDAATYYCQQGNKHP).
Cloning and Sequencing of Murine Anti-CXCR3 VH and VK Domains from Hybridoma Cell Lines
[0069] Hybridoma cells were pelleted, washed 3× with PBS and RNA extracted using Trizol reagent (Invitrogen, Cat. No. 15596-026) following the manufacturers protocol. Total RNA was converted to cDNA using a 5' RACE kit (Rapid Amplification of cDNA Ends, Invitrogen, Cat. No. 18374-058) following the manufacturers protocol. Briefly, RNA was ligated to random hexamer primer, Random N6, and 1st strand cDNA generated using superscript II RNAase H negative reverse transcriptase. The cDNA was purified using a GlassMax spin cartridge provided with the kit and then reacted with TdT (terminal deoxynucleotidyl transferase) in the presence of dCTP to append the cDNA with C basepairs at the 5' end. The dC-tailed cDNA was PCR amplified using an anchor primer specific for the dC tail and a gene specific primer that hybridizes to highly conserved DNA sequence in the mouse constant heavy 1 (CH1) for VH and constant kappa (CK) for VK. The resulting PCR product was analyzed by gel electrophoresis for correct size corresponding to intact VH or VK domain then purified and ligated in to a Topo TA vector (Invitrogen Cat. No. 45-0071) following manufacturers protocol. After transformation in to bacteria DNA was prepared from clones containing correct size insert and the DNA sequence determined using a Big Dye terminator sequencing reaction mix (Applied Biosystems, Part No. 4336699) and a 3700 ABI/Prism DNA analyzer following manufacturers protocol.
Humanizing Murine Anti-CXCR3 Antibodies
[0070] First, a single lead murine anti-CXCR3 antibody, V44D7, was identified based on binding data and sequence data generated as described above. The amino acid sequence of the VH and VK domains from this antibody were aligned to all known human germline VH and VK domains using currently available public databases (i.e., Blast for IgG at the NCBI and V-base at the MRC). By focusing on alignment within the framework regions a highly homologous human germline VH domain, VH3-23, and 2 different human germline VK domains, A14 and L6, were identified. At those positions in the framework where the mouse sequence differed from the human germline, an iterative process was used to convert or mutate the mouse framework so it matched the corresponding human germline framework. In addition, certain residues in CDR3 of both the VH and VK were mutated by replacement with tyrosine (i.e., affinity matured) to potentially help compensate for any losses in affinity due to the framework residues changes. The affinity matured and humanized mouse VH and VK domains were generated by a polymerase chain reaction process using a panel of overlapping synthetic DNA oligonucleotides. As part of the synthetic gene design process a codon optimization strategy was used, that is to say the triplet code for each amino acid that is preferentially utilized by mammalian cells for gene expression was incorporated at each position. The synthetic VH and VK domains were cloned in to specialized mammalian expression vectors that allowed the corresponding domains to be expressed in the context of a fully human IgG1, G4 or Kappa antibody backbone. Small-scale production of the humanized antibodies was achieved by co-tranfection of an IgG1 or G4 construct with the Kappa construct in to 293F cells with lipofectamine (Invitrogen) following manufactures protocol. Supernatants from the transient transfections were passed through Protein A or G resin and the IgG purified to homogeneity for testing in cell based assays.
Epitope Competition Studies
[0071] Various commercial CXR3 mAbs were tested in a competitive binding assay using Europium (Eu) labeled-mouse CXCR3 mAb. CXCR3 mAbs from various commercial sources inhibited Eu-CXCR3 mAb binding to CXCR3. This data indicated that mouse CXCR3 mAb and commercial antibodies bind to overlapping epitopes on CXCR3 (FIG. 13).
Antibody Affinities
[0072] Binding affinity and activity of mouse and humanized CXCR3 mAbs were determined by various competitive binding assays using 125I- and Eu-labeled chemokines and Eu-labeled CXCR3 mAb and Th1 chemotaxis assays including: 125I-CXCL10 binding assay; 125I-CXCL11 binding assay; Eu-CXCL10 binding assay; Th1 chemotaxis assay; and Eu-mouse CXCR3 mAb binding assay.
Th1 Cells
[0073] Primary Th1 cells generated from cord blood were used for all binding assays. As described in the literature, CXCR3 expression was observed only on Th1 cells but not on Th2 cells as determined by FACS analysis (FIG. 14). Th2 cells specifically expressed CCR4.
125I-CXCL10 and 125I-CXCL11 Binding Assays
[0074] The binding affinity of mouse CXCR3 mAb antibodies was determined based on their ability to inhibit radiolabeled CXCL10 and CXCL11 binding to Th1 cells (FIGS. 15, 16, 18, and 19 and Table 1). Based on these binding studies and the chemotaxis assay, three mouse CXCRmAbs were selected for further study.
TABLE-US-00001 TABLE 1 Characterization of Anti-CXCR3 mAbs Inhibition Displacement Displacement Binding of IP-10- of 125I-IP-10 of 125I-I-Tac to Th1 induced Th1 binding to binding to cells, chemotaxis, Th1 cells Th1 cells by FACS IC50 (ng/mL) (IC50, nM) (IC50, nM) IgG2 a 4.02 N/A N/A N/A Ab#1 1498 123 0.38 0.44 Ab#2 1215 575 0.41 0.47 Ab#3 681 N/A 2.6 5.7 Ab#4 1262 49 0.12 9.13 Ab#5 1119 35 0.13 0.10 Ab#6 831 172 0.94 0.76 Ab#7 4.20 N/A N/A -- Ab#8 1096 264 0.68 0.57 Ab#9 1348 39 0.20 0.14 Ab#10 66.84 N/A N/A -- Ab#11 4.80 N/A -- Ab#12 5.77 N/A N/A -- R&D mAb N/D 351 0.61 0.6 Ab#1 - 5D4A; Ab#2 - 8A5A; Ab#3 - 19G2; Ab#4 - V36E5A; Ab#5 - V44D7A; Ab#6 - 37B5A; Ab#7 - 21A4A; Ab#8 - V15F4A; Ab#9 - V3G6A; Ab#10 - 23E12A; Ab#11 - 35C4; Ab#12 - 39E10.
Eu-CXCRmAb Saturation Binding Assay
[0075] Binding affinity of mouse CXCR3 antibodies to CXCR3 was determined by direct saturation binding assay using Europium labeled mouse CXCR3 antibodies. An example of this assay using one mouse CXCR3 mAb is shown in FIG. 17. Data from this study indicated that Eu-CXCR3 mAb binding to CXCR3 was specific and saturable with binding affinity of subnanomolar Kd (0.47 nM).
Antibody Specificity
[0076] Mouse antibody hybridomas (20000) were screened by a differential screening assay with CXCR3+ and CXCR3-NSO membranes using a Eu-secondary antibody (DELFIA). Antibodies (˜2000) that bound to CXCR3+ membranes were further tested by FACS using CXCR3+/CXCR3.sup.- NSO and Th1 cells. An example for specific binding of CXCR3 mAb to CXCR3 expressing cells is shown in FIG. 3.
Species Cross Reactivity
[0077] The parental mouse CXCR3 mAb was tested for binding to polarized rat Th1 cells by FACS. Only rabbit anti-mouse CXCR3Ab (positive control) bound to rat Th1 cells. The mouse parental CXCR3 mAb did not bind to rat Th1 cells as shown in FIG. 20.
Biological Activity
[0078] Binding assay: CXCR3 mAb specifically inhibited 125I-labeled IP-10 and -I-Tac binding to primary Th1 cells. Due to unavailability of labeled Mig, it was not tested in binding assays. Chemotaxis assay: CXCR3 mAb inhibited Th1 cell migration mediated by CXCR3 chemokines.
[0079] Humanized antibodies were tested by binding and chemotaxis assays. The results of an Eu-CXCRmAb competitive binding assay are shown in FIG. 21. The results of the Eu-CXCL10 binding assay are shown in FIG. 22. The results of the Th1 chemotaxis assay are shown in FIG. 23.
Mouse Antibody Tested for Agonism of the CXCR3 Receptor
[0080] Mouse CXCR3mAbs were tested for agonistic activity in inducing ca mobilization in Th1 cells. As shown in FIG. 24, none of the antibodies s showed any agonistic activity in inducing Ca++ flux.
TABLE-US-00002 TABLE 2 IC50 Values of Different Humanized Antibodies As Determined From Binding and Chemotaxis Assays Binding Assay, Percent IC50 (nM) Chemotaxis Humanization Antibody ID Eu-IP-10 Eu-V44 IC50 (nM) H L Mouse anti- 0.14 0.29 0.09 88 79 CXCR3 V44D7 Humanized 0.8 2.6 0.31 100 96 Ab1 H5K1 Humanized 0.35 1.82 0.28 100 94 Ab2 H5K7 Humanized 1.25 1.86 N.D. 100 100 Ab3 H5K2 Humanized 0.38 0.56 N.D. 100 100 Ab4 H5K3 Humanized 0.69 1.49 N.D. 100 100 Ab5 H5K4 Humanized 0.42 0.65 N.D. 100 100 Ab6 H5K5 N.D. = not determined
[0081] In Table 2, Ab1 has a heavy chain sequence of humanized anti-CXCR3 V44D7 VH Lead #5 amino acid and a light chain sequence of humanized anti-CXCR3 V44D7 VK Lead #1 (see Informal Sequence Listing) in an IgG1 backbone. Ab2 has a heavy chain sequence of humanized anti-CXCR3 V44D7 VH Lead #5 and a light chain sequence of humanized anti-CXCR3 V44D7 VK Lead #7 (see Informal Sequence Listing) in an IgG1 backbone. Ab3 has a heavy chain sequence of humanized anti-CXCR3 V44D7 VH Lead #5 amino acid and a light chain sequence of humanized anti-CXCR3 V44D7 VK Lead #2 (see Informal Sequence Listing) in an IgG1 backbone. Ab4 has a heavy chain sequence of humanized anti-CXCR3 V44D7 VH Lead #5 and a light chain sequence of humanized anti-CXCR3 V44D7 VK Lead #3 (see Informal Sequence Listing) in an IgG1 backbone. Ab5 has a heavy chain sequence of humanized anti-CXCR3 V44D7 VH Lead #5 amino acid and a light chain sequence of humanized anti-CXCR3 V44D7 VK Lead #4 (see Informal Sequence Listing) in an IgG1 backbone. Ab6 has a heavy chain sequence of humanized anti-CXCR3 V44D7 VH Lead #5 and a light chain sequence of humanized anti-CXCR3 V44D7 VK Lead #5 (see Informal Sequence Listing) in an IgG1 backbone.
Sequence CWU
1
SEQUENCE LISTING
<160> NUMBER OF SEQ ID NOS: 84
<210> SEQ ID NO 1
<211> LENGTH: 5
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 1
Asn Tyr Met Ala Ser
1 5
<210> SEQ ID NO 2
<211> LENGTH: 17
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 2
Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu Lys
1 5 10 15
Gly
<210> SEQ ID NO 3
<211> LENGTH: 18
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (17)..(17)
<223> OTHER INFORMATION: Asp or Tyr
<400> SEQUENCE: 3
His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr Tyr Phe
1 5 10 15
Xaa Tyr
<210> SEQ ID NO 4
<211> LENGTH: 10
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 4
Arg Ala Ser Ser Ser Val Lys Tyr Met Tyr
1 5 10
<210> SEQ ID NO 5
<211> LENGTH: 7
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 5
Tyr Thr Ser Asn Leu Ala Pro
1 5
<210> SEQ ID NO 6
<211> LENGTH: 9
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 6
Gln Gln Phe Thr Thr Ser Pro Tyr Thr
1 5
<210> SEQ ID NO 7
<211> LENGTH: 18
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 7
His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr Tyr Phe
1 5 10 15
Tyr Tyr
<210> SEQ ID NO 8
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Glu or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Ile, Asn or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Leu or Met
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (10)..(10)
<223> OTHER INFORMATION: Thr, Phe or Ile
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (11)..(11)
<223> OTHER INFORMATION: Leu or Met
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (13)..(13)
<223> OTHER INFORMATION: Leu, Ala or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (14)..(14)
<223> OTHER INFORMATION: Ser or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (15)..(15)
<223> OTHER INFORMATION: Leu or Pro
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (18)..(18)
<223> OTHER INFORMATION: Arg or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (19)..(19)
<223> OTHER INFORMATION: Ala or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (21)..(21)
<223> OTHER INFORMATION: Leu, Met or Ile
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (22)..(22)
<223> OTHER INFORMATION: Ser, Thr or Asn
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (39)..(39)
<223> OTHER INFORMATION: Ser or Pro
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (40)..(40)
<223> OTHER INFORMATION: Gly or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (41)..(41)
<223> OTHER INFORMATION: Gln or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (42)..(42)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (44)..(44)
<223> OTHER INFORMATION: Arg or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (46)..(46)
<223> OTHER INFORMATION: Leu or Trp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (48)..(48)
<223> OTHER INFORMATION: Tyr or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (57)..(57)
<223> OTHER INFORMATION: Ile or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (59)..(59)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (68)..(68)
<223> OTHER INFORMATION: Thr or Asn
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (69)..(69)
<223> OTHER INFORMATION: Asp or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (70)..(70)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (71)..(71)
<223> OTHER INFORMATION: Thr or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (72)..(72)
<223> OTHER INFORMATION: Leu or Phe
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (77)..(77)
<223> OTHER INFORMATION: Met or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (79)..(79)
<223> OTHER INFORMATION: Ala, Gly or Pro
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (82)..(82)
<223> OTHER INFORMATION: Phe or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (84)..(84)
<223> OTHER INFORMATION: Val or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (88)..(88)
<223> OTHER INFORMATION: Gln or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (93)..(93)
<223> OTHER INFORMATION: Ser or Tyr
<400> SEQUENCE: 8
Xaa Xaa Val Xaa Thr Gln Ser Pro Ala Xaa Xaa Ser Xaa Xaa Xaa Gly
1 5 10 15
Glu Xaa Xaa Thr Xaa Xaa Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Xaa Xaa Xaa Xaa Pro Xaa Leu Xaa Ile Xaa
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Xaa Pro Xaa Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Xaa Xaa Xaa Xaa Xaa Thr Ile Ser Ser Xaa Glu Xaa Glu
65 70 75 80
Asp Xaa Ala Xaa Tyr Tyr Cys Xaa Gln Phe Thr Thr Xaa Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 9
<211> LENGTH: 126
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 9
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Lys Gly Leu Glu Trp Val Ser
35 40 45
Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu Lys
50 55 60
Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr Leu
65 70 75 80
Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys Ala
85 90 95
Lys His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr Tyr
100 105 110
Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 10
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Ile or Asn
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (10)..(10)
<223> OTHER INFORMATION: Thr, Phe or Ile
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (11)..(11)
<223> OTHER INFORMATION: Leu or Met
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (13)..(13)
<223> OTHER INFORMATION: Leu, Ala or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (14)..(14)
<223> OTHER INFORMATION: Ser or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (15)..(15)
<223> OTHER INFORMATION: Leu or Pro
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (18)..(18)
<223> OTHER INFORMATION: Arg or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (19)..(19)
<223> OTHER INFORMATION: Ala or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (21)..(21)
<223> OTHER INFORMATION: Leu, Met or Ile
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (22)..(22)
<223> OTHER INFORMATION: Ser, Thr or Asn
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (39)..(39)
<223> OTHER INFORMATION: Ser or Pro
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (40)..(40)
<223> OTHER INFORMATION: Gly or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (41)..(41)
<223> OTHER INFORMATION: Gln or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (42)..(42)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (44)..(44)
<223> OTHER INFORMATION: Arg or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (46)..(46)
<223> OTHER INFORMATION: Leu or Trp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (57)..(57)
<223> OTHER INFORMATION: Ile or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (59)..(59)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (68)..(68)
<223> OTHER INFORMATION: Thr or Asn
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (69)..(69)
<223> OTHER INFORMATION: Asp or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (70)..(70)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (71)..(71)
<223> OTHER INFORMATION: Thr or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (72)..(72)
<223> OTHER INFORMATION: Leu or Phe
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (77)..(77)
<223> OTHER INFORMATION: Met or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (79)..(79)
<223> OTHER INFORMATION: Ala or Gly
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (82)..(82)
<223> OTHER INFORMATION: Phe or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (84)..(84)
<223> OTHER INFORMATION: Val or Thr
<400> SEQUENCE: 10
Glu Xaa Val Leu Thr Gln Ser Pro Ala Xaa Xaa Ser Xaa Xaa Xaa Gly
1 5 10 15
Glu Xaa Xaa Thr Xaa Xaa Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Xaa Xaa Xaa Xaa Pro Xaa Leu Xaa Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Xaa Pro Xaa Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Xaa Xaa Xaa Xaa Xaa Thr Ile Ser Ser Xaa Glu Xaa Glu
65 70 75 80
Asp Xaa Ala Xaa Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 11
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 11
Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Leu Gly
1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Ser Gly Gln Ala Pro Arg Leu Leu Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Ile Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Met Glu Ala Glu
65 70 75 80
Asp Phe Ala Val Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 12
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 12
Glu Asn Val Leu Thr Gln Ser Pro Ala Phe Leu Ser Val Thr Pro Gly
1 5 10 15
Glu Lys Val Thr Ile Thr Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Asp Gln Ala Pro Lys Leu Trp Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Val Pro Ser Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Asn Asp Tyr Thr Phe Thr Ile Ser Ser Leu Glu Ala Glu
65 70 75 80
Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 13
<211> LENGTH: 5
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Asn, Ser or Tyr
<400> SEQUENCE: 13
Xaa Tyr Ala Met Ser
1 5
<210> SEQ ID NO 14
<211> LENGTH: 17
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Thr, Ala or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (3)..(3)
<223> OTHER INFORMATION: Ser or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Ser, Gly, Thr or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (5)..(5)
<223> OTHER INFORMATION: Gly or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (6)..(6)
<223> OTHER INFORMATION: Gly or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (8)..(8)
<223> OTHER INFORMATION: Phe, Ser or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (12)..(12)
<223> OTHER INFORMATION: Pro or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (15)..(15)
<223> OTHER INFORMATION: Leu, Tyr or Val
<400> SEQUENCE: 14
Xaa Ile Xaa Xaa Xaa Xaa Gly Xaa Thr Tyr Tyr Xaa Asp Ser Xaa Lys
1 5 10 15
Gly
<210> SEQ ID NO 15
<211> LENGTH: 18
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: His or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Gly or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (3)..(3)
<223> OTHER INFORMATION: Ala or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (6)..(6)
<223> OTHER INFORMATION: Thr or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (8)..(8)
<223> OTHER INFORMATION: Val or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (12)..(12)
<223> OTHER INFORMATION: Ala or Tyr
<400> SEQUENCE: 15
Xaa Xaa Xaa Pro Met Xaa Thr Xaa Ile Thr Tyr Xaa Pro Tyr Tyr Phe
1 5 10 15
Tyr Tyr
<210> SEQ ID NO 16
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (31)..(31)
<223> OTHER INFORMATION: Asn, Ser or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (50)..(50)
<223> OTHER INFORMATION: Thr, Ala or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (52)..(52)
<223> OTHER INFORMATION: Ser or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (53)..(53)
<223> OTHER INFORMATION: Ser, Gly, Thr or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (54)..(54)
<223> OTHER INFORMATION: Gly or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (55)..(55)
<223> OTHER INFORMATION: Gly or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (57)..(57)
<223> OTHER INFORMATION: Phe, Ser or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (61)..(61)
<223> OTHER INFORMATION: Pro or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (64)..(64)
<223> OTHER INFORMATION: Leu, Tyr or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (99)..(99)
<223> OTHER INFORMATION: His or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (100)..(100)
<223> OTHER INFORMATION: Gly or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (101)..(101)
<223> OTHER INFORMATION: Ala or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (104)..(104)
<223> OTHER INFORMATION: Thr or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (106)..(106)
<223> OTHER INFORMATION: Val or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (110)..(110)
<223> OTHER INFORMATION: Ala or Tyr
<400> SEQUENCE: 16
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Xaa Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Xaa Ile Xaa Xaa Xaa Xaa Gly Xaa Thr Tyr Tyr Xaa Asp Ser Xaa
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys Xaa Xaa Xaa Pro Met Xaa Thr Xaa Ile Thr Tyr Xaa Pro Tyr
100 105 110
Tyr Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 17
<211> LENGTH: 5
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 17
Asn Tyr Ala Ile Ser
1 5
<210> SEQ ID NO 18
<211> LENGTH: 17
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 18
Thr Tyr Ser Ser Gly Gly Val Tyr Thr Tyr Tyr Arg Asp Ser Leu Lys
1 5 10 15
Gly
<210> SEQ ID NO 19
<211> LENGTH: 18
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 19
His Gly Ala Ala Met Thr Thr Val Ile Thr Tyr Ala Pro Phe Tyr Phe
1 5 10 15
Tyr Tyr
<210> SEQ ID NO 20
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 20
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Thr Tyr Ser Ser Gly Gly Val Tyr Thr Tyr Tyr Arg Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys His Gly Ala Ala Met Thr Thr Val Ile Thr Tyr Ala Pro Phe
100 105 110
Tyr Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 21
<211> LENGTH: 5
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 21
Tyr Tyr Ala Met Ser
1 5
<210> SEQ ID NO 22
<211> LENGTH: 17
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 22
Thr Ile Tyr Ser Gly Gly Ser Tyr Thr Phe Tyr Pro Asp Ser Leu Glu
1 5 10 15
Gly
<210> SEQ ID NO 23
<211> LENGTH: 18
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 23
His Gly Ala Pro Met Ser Thr Glu Ile Thr Tyr Ala Pro Tyr Tyr Phe
1 5 10 15
Tyr Tyr
<210> SEQ ID NO 24
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 24
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Tyr Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Thr Ile Tyr Ser Gly Gly Ser Tyr Thr Phe Tyr Pro Asp Ser Leu
50 55 60
Glu Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys His Gly Ala Pro Met Ser Thr Glu Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 25
<211> LENGTH: 5
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 25
Asn Tyr Ala Met Ser
1 5
<210> SEQ ID NO 26
<211> LENGTH: 17
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 26
Thr Ile Tyr Ser Gly Gly Gly Tyr Thr Phe Tyr Leu Asp Ser Leu Lys
1 5 10 15
Gly
<210> SEQ ID NO 27
<211> LENGTH: 18
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 27
His Ser Tyr Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr Tyr Phe
1 5 10 15
Tyr Tyr
<210> SEQ ID NO 28
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 28
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Tyr Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Thr Ile Tyr Ser Gly Gly Gly Tyr Thr Phe Tyr Leu Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys His Ser Tyr Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 29
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 29
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 30
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 30
Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly
1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Ile Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro Glu
65 70 75 80
Asp Phe Ala Val Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 31
<211> LENGTH: 9
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 31
Tyr Gln Phe Thr Thr Ser Pro Tyr Thr
1 5
<210> SEQ ID NO 32
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 32
Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly
1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Ile Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro Glu
65 70 75 80
Asp Phe Ala Val Tyr Tyr Cys Tyr Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 33
<211> LENGTH: 9
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 33
Gln Gln Tyr Thr Thr Ser Pro Tyr Thr
1 5
<210> SEQ ID NO 34
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 34
Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly
1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Ile Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro Glu
65 70 75 80
Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 35
<211> LENGTH: 9
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 35
Gln Gln Phe Thr Thr Tyr Pro Tyr Thr
1 5
<210> SEQ ID NO 36
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 36
Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly
1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Ile Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro Glu
65 70 75 80
Asp Phe Ala Val Tyr Tyr Cys Gln Gln Phe Thr Thr Tyr Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 37
<211> LENGTH: 11
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Ser or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (7)..(7)
<223> OTHER INFORMATION: Lys or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (10)..(10)
<223> OTHER INFORMATION: Met or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (11)..(11)
<223> OTHER INFORMATION: Tyr or Ala
<400> SEQUENCE: 37
Arg Ala Ser Xaa Ser Val Xaa Ser Tyr Xaa Xaa
1 5 10
<210> SEQ ID NO 38
<211> LENGTH: 9
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Gln or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (3)..(3)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (6)..(6)
<223> OTHER INFORMATION: Ser or Tyr
<400> SEQUENCE: 38
Xaa Gln Xaa Thr Thr Xaa Pro Tyr Thr
1 5
<210> SEQ ID NO 39
<211> LENGTH: 108
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (27)..(27)
<223> OTHER INFORMATION: Ser or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (30)..(30)
<223> OTHER INFORMATION: Lys or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (33)..(33)
<223> OTHER INFORMATION: Met or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (34)..(34)
<223> OTHER INFORMATION: Tyr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (50)..(50)
<223> OTHER INFORMATION: Tyr or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (51)..(51)
<223> OTHER INFORMATION: Thr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (54)..(54)
<223> OTHER INFORMATION: Leu or Arg
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (56)..(56)
<223> OTHER INFORMATION: Pro or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (89)..(89)
<223> OTHER INFORMATION: Gln or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (91)..(91)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (94)..(94)
<223> OTHER INFORMATION: Ser or Tyr
<400> SEQUENCE: 39
Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly
1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Xaa Ser Val Xaa Ser Tyr
20 25 30
Xaa Xaa Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile
35 40 45
Tyr Xaa Xaa Ser Asn Xaa Ala Xaa Gly Ile Pro Ala Arg Phe Ser Gly
50 55 60
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro
65 70 75 80
Glu Asp Phe Ala Val Tyr Tyr Cys Xaa Gln Xaa Thr Thr Xaa Pro Tyr
85 90 95
Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 40
<211> LENGTH: 7
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Tyr or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Thr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (5)..(5)
<223> OTHER INFORMATION: Leu or Arg
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (7)..(7)
<223> OTHER INFORMATION: Pro or Thr
<400> SEQUENCE: 40
Xaa Xaa Ser Asn Xaa Ala Xaa
1 5
<210> SEQ ID NO 41
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Glu or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Asn or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Leu or Met
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (46)..(46)
<223> OTHER INFORMATION: Trp or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (48)..(48)
<223> OTHER INFORMATION: Tyr or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (68)..(68)
<223> OTHER INFORMATION: Asn or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (70)..(70)
<223> OTHER INFORMATION: Tyr or Phe
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (88)..(88)
<223> OTHER INFORMATION: Gln or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (90)..(90)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (93)..(93)
<223> OTHER INFORMATION: Ser or Tyr
<400> SEQUENCE: 41
Xaa Xaa Val Xaa Thr Gln Ser Pro Ala Phe Leu Ser Val Thr Pro Gly
1 5 10 15
Glu Lys Val Thr Ile Thr Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Asp Gln Ala Pro Lys Leu Xaa Ile Xaa
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Val Pro Ser Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Xaa Asp Xaa Thr Phe Thr Ile Ser Ser Leu Glu Ala Glu
65 70 75 80
Asp Ala Ala Thr Tyr Tyr Cys Xaa Gln Xaa Thr Thr Xaa Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 42
<211> LENGTH: 6
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Any amino acid
<400> SEQUENCE: 42
Leu Val His Xaa Val Lys
1 5
<210> SEQ ID NO 43
<211> LENGTH: 6
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Any amino acid
<400> SEQUENCE: 43
Leu Val Lys Xaa Val His
1 5
<210> SEQ ID NO 44
<211> LENGTH: 98
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 44
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Gln Val
35 40 45
Ser Ala Ile Ser Gly Ser Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys
<210> SEQ ID NO 45
<211> LENGTH: 95
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 45
Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly
1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Ser Tyr
20 25 30
Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile
35 40 45
Tyr Asp Ala Ser Asn Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser Gly
50 55 60
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro
65 70 75 80
Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Arg Ser Asn Trp Pro
85 90 95
<210> SEQ ID NO 46
<211> LENGTH: 95
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 46
Asp Val Val Met Thr Gln Ser Pro Ala Phe Leu Ser Val Thr Pro Gly
1 5 10 15
Glu Lys Val Thr Ile Thr Cys Gln Ala Ser Glu Gly Ile Gly Asn Tyr
20 25 30
Leu Tyr Trp Tyr Gln Gln Lys Pro Asp Gln Ala Pro Lys Leu Leu Ile
35 40 45
Lys Tyr Ala Ser Gln Ser Ile Ser Gly Val Pro Ser Arg Phe Ser Gly
50 55 60
Ser Gly Ser Gly Thr Asp Phe Thr Phe Thr Ile Ser Ser Leu Glu Ala
65 70 75 80
Glu Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Gly Asn Lys His Pro
85 90 95
<210> SEQ ID NO 47
<211> LENGTH: 18
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 47
His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr Tyr Phe
1 5 10 15
Asp Tyr
<210> SEQ ID NO 48
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (3)..(3)
<223> OTHER INFORMATION: Met or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (5)..(5)
<223> OTHER INFORMATION: Val or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (13)..(13)
<223> OTHER INFORMATION: Lys or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (19)..(19)
<223> OTHER INFORMATION: Lys or Arg
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (40)..(40)
<223> OTHER INFORMATION: Thr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (42)..(42)
<223> OTHER INFORMATION: Glu or Gly
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (44)..(44)
<223> OTHER INFORMATION: Arg or Gly
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (49)..(49)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (75)..(75)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (80)..(80)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (84)..(84)
<223> OTHER INFORMATION: Ser or Asn
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (88)..(88)
<223> OTHER INFORMATION: Ser or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (97)..(97)
<223> OTHER INFORMATION: Val or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (98)..(98)
<223> OTHER INFORMATION: Arg or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (115)..(115)
<223> OTHER INFORMATION: Asp or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (123)..(123)
<223> OTHER INFORMATION: Leu or Val
<400> SEQUENCE: 48
Glu Val Xaa Leu Xaa Glu Ser Gly Gly Gly Leu Val Xaa Pro Gly Gly
1 5 10 15
Ser Leu Xaa Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Xaa Pro Xaa Lys Xaa Leu Glu Trp Val
35 40 45
Xaa Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Xaa Lys Asn Thr Leu Xaa
65 70 75 80
Leu Gln Met Xaa Ser Leu Arg Xaa Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Xaa Xaa His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Xaa Tyr Trp Gly Gln Gly Thr Thr Xaa Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 49
<211> LENGTH: 108
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 49
Glu Ile Asn Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Leu
1 5 10 15
Gly Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Ser Ser Val Lys Tyr
20 25 30
Met Tyr Trp Tyr Gln Gln Lys Ser Gly Gln Ala Pro Arg Leu Leu Ile
35 40 45
Tyr Tyr Thr Ser Asn Leu Ala Pro Gly Ile Pro Ala Arg Phe Ser Gly
50 55 60
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Met Glu Ala
65 70 75 80
Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr
85 90 95
Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 50
<211> LENGTH: 108
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Glu or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Asn or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Leu or Met
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (27)..(27)
<223> OTHER INFORMATION: Ser or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (30)..(30)
<223> OTHER INFORMATION: Lys or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (33)..(33)
<223> OTHER INFORMATION: Met or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (34)..(34)
<223> OTHER INFORMATION: Tyr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (47)..(47)
<223> OTHER INFORMATION: Trp or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (49)..(49)
<223> OTHER INFORMATION: Tyr or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (50)..(50)
<223> OTHER INFORMATION: Tyr or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (51)..(51)
<223> OTHER INFORMATION: Thr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (54)..(54)
<223> OTHER INFORMATION: Leu or Arg
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (56)..(56)
<223> OTHER INFORMATION: Pro or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (69)..(69)
<223> OTHER INFORMATION: Asn or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (71)..(71)
<223> OTHER INFORMATION: Tyr or Phe
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (89)..(89)
<223> OTHER INFORMATION: Gln or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (91)..(91)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (94)..(94)
<223> OTHER INFORMATION: Ser or Tyr
<400> SEQUENCE: 50
Xaa Xaa Val Xaa Thr Gln Ser Pro Ala Phe Leu Ser Val Thr Pro Gly
1 5 10 15
Glu Lys Val Thr Ile Thr Cys Arg Ala Ser Xaa Ser Val Xaa Ser Tyr
20 25 30
Xaa Xaa Trp Tyr Gln Gln Lys Pro Asp Gln Ala Pro Lys Leu Xaa Ile
35 40 45
Xaa Xaa Xaa Ser Asn Xaa Ala Xaa Gly Val Pro Ser Arg Phe Ser Gly
50 55 60
Ser Gly Ser Gly Xaa Asp Xaa Thr Phe Thr Ile Ser Ser Leu Glu Ala
65 70 75 80
Glu Asp Ala Ala Thr Tyr Tyr Cys Xaa Gln Xaa Thr Thr Xaa Pro Tyr
85 90 95
Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 51
<211> LENGTH: 381
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 51
gaagtgatgc tggtggagtc tgggggaggc ttagtgaagc ctggagggtc cctgaaactc 60
tcctgtgcag cctctggatt cactttcagt aactatgcca tgtcttgggt tcgtcagact 120
ccggagaaga ggctggagtg ggtcgcaacc ataagtagtg gtggaggtta cacctattat 180
ccagacagtt tgaaggggcg attcaccatc tccagagaca atgccaagaa caccctgttc 240
ctgcaaatga gcagtctgag gtctgaggac acggccgtgt attattgtgt aagacatggg 300
gcccccatga ctacggtgat aacttacgct ccttactatt ttgactactg gggccagggc 360
accactctca cagtctcctc a 381
<210> SEQ ID NO 52
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 52
Glu Val Met Leu Val Glu Ser Gly Gly Gly Leu Val Lys Pro Gly Gly
1 5 10 15
Ser Leu Lys Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Thr Pro Glu Lys Arg Leu Glu Trp Val
35 40 45
Ala Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Thr Leu Phe
65 70 75 80
Leu Gln Met Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Val Arg His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Asp Tyr Trp Gly Gln Gly Thr Thr Leu Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 53
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 53
gaaaatgtgc tcacccagtc tccagcaatc atgtctgcat ctctggggga gaaggtcacc 60
atgaactgca gggccagctc aagtgtaaaa tacatgtact ggtaccagca gaagtcagat 120
gcctccccca aactatggat ttattacaca tccaacctgg ctcctggagt cccagctcgc 180
ttcagtggca gtgggtctgg gaactcttat tctctcacaa tcagcagcat ggagggtgaa 240
gatgctgcca cttattactg ccagcagttt actacttccc cgtacacgtt cggagggggg 300
accaagctgg aaataaaacg g 321
<210> SEQ ID NO 54
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 54
Glu Asn Val Leu Thr Gln Ser Pro Ala Ile Met Ser Ala Ser Leu Gly
1 5 10 15
Glu Lys Val Thr Met Asn Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Ser Asp Ala Ser Pro Lys Leu Trp Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Val Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Asn Ser Tyr Ser Leu Thr Ile Ser Ser Met Glu Gly Glu
65 70 75 80
Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 55
<211> LENGTH: 381
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (91)..(93)
<223> OTHER INFORMATION: This region may encompass 'TAC' or 'TCA'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (148)..(150)
<223> OTHER INFORMATION: This region may encompass 'TAC' or 'GCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (154)..(156)
<223> OTHER INFORMATION: This region may encompass 'TCC' or 'TAC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (157)..(159)
<223> OTHER INFORMATION: This region may encompass 'GGT' or 'ACC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (169)..(171)
<223> OTHER INFORMATION: This region may encompass 'TTC' or 'TCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (181)..(183)
<223> OTHER INFORMATION: This region may encompass 'CCC' or 'GCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (190)..(192)
<223> OTHER INFORMATION: This region may encompass 'CTG' or 'TAC'
<400> SEQUENCE: 55
gaggtgcagc tgctggagtc cggcggaggc ctggtgcagc ccggcggctc cctgcggctg 60
tcctgcgccg cctccggctt caccttctcc nnntacgcca tgtcctgggt gcggcaggcc 120
cccggcaagg gcctggagtg ggtgtccnnn atcnnnnnnt ccggaggcnn nacctactat 180
nnngactccn nnaagggccg gttcaccatc tcccgggaca actccaagaa caccctgtac 240
ctgcagatga actccctgcg ggccgaagac accgccgtgt actattgcgc caagcatgac 300
gccctcatga ccactgtgat cacctacgcc ccctcttact tctactactg gggccagggc 360
accacggtca ccgtctcctc a 381
<210> SEQ ID NO 56
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (31)..(31)
<223> OTHER INFORMATION: Tyr or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (50)..(50)
<223> OTHER INFORMATION: Tyr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (52)..(52)
<223> OTHER INFORMATION: Ser or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (53)..(53)
<223> OTHER INFORMATION: Gly or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (57)..(57)
<223> OTHER INFORMATION: Phe or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (61)..(61)
<223> OTHER INFORMATION: Pro or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (64)..(64)
<223> OTHER INFORMATION: Leu or Tyr
<400> SEQUENCE: 56
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Xaa Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Xaa Ile Xaa Xaa Ser Gly Gly Xaa Thr Tyr Tyr Xaa Asp Ser Xaa
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys His Asp Ala Leu Met Thr Thr Val Ile Thr Tyr Ala Pro Ser
100 105 110
Tyr Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 57
<211> LENGTH: 381
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 57
gaggtgcagc tgctggagtc cggcggaggc ctggtgcagc ccggcggctc cctgcggctg 60
tcctgcgccg cctccggctt caccttctcc aactacgcca tttcctgggt gcggcaggcc 120
cccggcaagg gcctggagtg ggtgtccacc tactcctccg gcggagtcta cacctactat 180
cgcgactccc tgaagggccg gttcaccatc tcccgggaca actccaagaa caccctgtac 240
ctgcagatga actccctgcg ggccgaagac accgccgtgt actattgcgc caagcacggc 300
gccgccatga ccactgtgat cacctatgcc cccttttact tctactactg gggccagggc 360
accacggtca ccgtctcctc a 381
<210> SEQ ID NO 58
<211> LENGTH: 381
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 58
gaggtgcagc tgctggagtc cggcggaggc ctggtgcagc ccggcggctc cctgcggctg 60
tcctgcgccg cctccggctt caccttctcc tactacgcca tgtcctgggt gcggcaggcc 120
cccggcaagg gcctggagtg ggtgtccacc atctactccg gcggaagcta caccttctat 180
cccgactccc tggagggccg gttcaccatc tcccgggaca actccaagaa caccctgtac 240
ctgcagatga actccctgcg ggccgaagac accgccgtgt actattgcgc caagcacggc 300
gcccccatga gcactgagat cacctacgcc ccctattact tctactactg gggccagggc 360
accacggtca ccgtctcctc a 381
<210> SEQ ID NO 59
<211> LENGTH: 381
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 59
gaggtgcagc tgctggagtc cggcggaggc ctggtgcagc ccggcggctc cctgcggctg 60
tcctgcgccg cctccggctt caccttctcc aactactaca tgtcctgggt gcggcaggcc 120
cccggcaagg gcctggagtg ggtgtccacc atctactccg gcggaggcta caccttctat 180
ctcgactccc tgaagggccg gttcaccatc tcccgggaca actccaagaa caccctgtac 240
ctgcagatga actccctgcg ggccgaagac accgccgtgt actattgcgc caagcacagc 300
taccccatga ccactgtgat cacctacgcc ccctattact tctactactg gggccagggc 360
accacggtca ccgtctcctc a 381
<210> SEQ ID NO 60
<211> LENGTH: 381
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 60
gaggtgcagc tgctggagtc cggcggaggc ctggtgcagc ccggcggctc cctgcggctg 60
tcctgcgccg cctccggctt caccttctcc aactacgcca tgtcctgggt gcggcaggcc 120
cccggcaagg gcctggagtg ggtgtccacc atctcctccg gcggaggcta cacctactat 180
cccgactccc tgaagggccg gttcaccatc tcccgggaca actccaagaa caccctgtac 240
ctgcagatga actccctgcg ggccgaagac accgccgtgt actattgcgc caagcacggc 300
gcccccatga ccactgtgat cacctacgcc ccctattact tctactactg gggccagggc 360
accacggtca ccgtctcctc a 381
<210> SEQ ID NO 61
<211> LENGTH: 381
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (91)..(93)
<223> OTHER INFORMATION: This region may encompass 'AAC' or 'TCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (148)..(150)
<223> OTHER INFORMATION: This region may encompass 'ACC' or 'GCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (157)..(159)
<223> OTHER INFORMATION: This region may encompass 'TCC' or 'GGA'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (160)..(162)
<223> OTHER INFORMATION: This region may encompass 'GGC' or 'TCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (169)..(171)
<223> OTHER INFORMATION: This region may encompass 'TAC' or 'TCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (181)..(183)
<223> OTHER INFORMATION: This region may encompass 'CCC' or 'GCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (190)..(192)
<223> OTHER INFORMATION: This region may encompass 'CTG' or 'GTC'
<400> SEQUENCE: 61
gaggtgcagc tgctggagtc cggcggaggc ctggtgcagc ccggcggctc cctgcggctg 60
tcctgcgccg cctccggctt caccttctcc nnntacgcca tgtcctgggt gcggcaggcc 120
cccggcaagg gcctggagtg ggtgtccnnn atctccnnnn nnggaggcnn nacctactat 180
nnngactccn nnaagggccg gttcaccatc tcccgggaca actccaagaa caccctgtac 240
ctgcagatga actccctgcg ggccgaagac accgccgtgt actattgcgc caagcacggc 300
gcccccatga ccactgtgat cacctacgcc ccctattact tctactactg gggccagggc 360
accacggtca ccgtctcctc a 381
<210> SEQ ID NO 62
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (31)..(31)
<223> OTHER INFORMATION: Asn or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (50)..(50)
<223> OTHER INFORMATION: Thr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (53)..(54)
<223> OTHER INFORMATION: Ser or Gly
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (57)..(57)
<223> OTHER INFORMATION: Tyr or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (61)..(61)
<223> OTHER INFORMATION: Pro or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (64)..(64)
<223> OTHER INFORMATION: Leu or Val
<400> SEQUENCE: 62
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Xaa Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Xaa Ile Ser Xaa Xaa Gly Gly Xaa Thr Tyr Tyr Xaa Asp Ser Xaa
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 63
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 63
gagatcgtgc tgacccagtc ccccgccacc ctgtccctgt ccctgggcga gcgggccacc 60
ctgtcctgcc gggcctccag ctccgtgaag tacatgtact ggtaccagca gaagtccggc 120
caggcccccc ggctgctgat ctactacacc tccaacctgg cccccggcat ccccgcccgg 180
ttctccggct ccggctccgg caccgacttc accctgacca tctccagcat ggaggccgag 240
gacttcgccg tgtactactg ccagcagttc accacctccc cctacacctt cggcggaggc 300
accaagctcg agatcaaacg t 321
<210> SEQ ID NO 64
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 64
gagatcgtgc tgacccagtc ccccgccacc ctgtccctgt cccccggcga gcgggccacc 60
ctgtcctgcc gggcctccag ctccgtgaag tacatgtact ggtaccagca gaagcccggc 120
caggcccccc ggctgctgat ctactacacc tccaacctgg cccccggcat ccccgcccgg 180
ttctccggct ccggctccgg caccgacttc accctgacca tctcctccct ggagcccgag 240
gacttcgccg tgtactactg ccagcagttc accacctccc cctacacctt cggcggaggc 300
accaagctcg agatcaagcg g 321
<210> SEQ ID NO 65
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 65
gagatcgtgc tgacccagtc ccccgccacc ctgtccctgt cccccggcga gcgggccacc 60
ctgtcctgcc gggcctccag ctccgtgaag tacatgtact ggtaccagca gaagcccggc 120
caggcccccc ggctgctgat ctactacacc tccaacctgg cccccggcat ccccgcccgg 180
ttctccggct ccggctccgg caccgacttc accctgacca tctcctccct ggagcccgag 240
gacttcgccg tgtactactg ctaccagttc accacctccc cctacacctt cggcggaggc 300
accaagctcg agatcaagcg g 321
<210> SEQ ID NO 66
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 66
gagatcgtgc tgacccagtc ccccgccacc ctgtccctgt cccccggcga gcgggccacc 60
ctgtcctgcc gggcctccag ctccgtgaag tacatgtact ggtaccagca gaagcccggc 120
caggcccccc ggctgctgat ctactacacc tccaacctgg cccccggcat ccccgcccgg 180
ttctccggct ccggctccgg caccgacttc accctgacca tctcctccct ggagcccgag 240
gacttcgccg tgtactactg ccagcagtac accacctccc cctacacctt cggcggaggc 300
accaagctcg agatcaagcg g 321
<210> SEQ ID NO 67
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 67
gagatcgtgc tgacccagtc ccccgccacc ctgtccctgt cccccggcga gcgggccacc 60
ctgtcctgcc gggcctccag ctccgtgaag tacatgtact ggtaccagca gaagcccggc 120
caggcccccc ggctgctgat ctactacacc tccaacctgg cccccggcat ccccgcccgg 180
ttctccggct ccggctccgg caccgacttc accctgacca tctcctccct ggagcccgag 240
gacttcgccg tgtactactg ccagcagttc accacctacc cctacacctt cggcggaggc 300
accaagctcg agatcaagcg g 321
<210> SEQ ID NO 68
<211> LENGTH: 324
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (79)..(81)
<223> OTHER INFORMATION: This region may encompass 'AGC' or 'CAG'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (88)..(90)
<223> OTHER INFORMATION: This region may encompass 'AGG' or 'TCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (97)..(99)
<223> OTHER INFORMATION: This region may encompass 'ATG' or 'CTG'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (100)..(102)
<223> OTHER INFORMATION: This region may encompass 'TAC' or 'GCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (148)..(150)
<223> OTHER INFORMATION: This region may encompass 'TAC' or 'GAC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (151)..(153)
<223> OTHER INFORMATION: This region may encompass 'ACC' or 'GCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (160)..(162)
<223> OTHER INFORMATION: This region may encompass 'CTG' or 'CGG'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (166)..(168)
<223> OTHER INFORMATION: This region may encompass 'CCC' or 'ACC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (265)..(267)
<223> OTHER INFORMATION: This region may encompass 'CAG' or 'TAC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (271)..(273)
<223> OTHER INFORMATION: This region may encompass 'TTC' or 'TAC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (280)..(282)
<223> OTHER INFORMATION: This region may encompass 'TCC' or 'TAC'
<400> SEQUENCE: 68
gagatcgtgc tgacccagtc ccccgccacc ctgtccctgt cccccggcga gcgggccacc 60
ctgtcctgcc gggcctccnn ntccgtgnnn tcctacnnnn nntggtacca gcagaagccc 120
ggccaggccc cccggctgct gatctacnnn nnntccaacn nngccnnngg catccccgcc 180
cggttctccg gctccggctc cggcaccgac ttcaccctga ccatctcctc cctggagccc 240
gaggacttcg ccgtgtacta ctgcnnncag nnnaccaccn nnccctacac cttcggcgga 300
ggcaccaagc tcgagatcaa gcgg 324
<210> SEQ ID NO 69
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 69
gagaacgtgc tgacccagtc ccccgccttc ctgtccgtga cccccggcga gaaggtgacc 60
atcacctgcc gggcctccag ctccgtgaag tacatgtact ggtaccagca gaagcccgac 120
caggccccca agctgtggat ctattacacc tccaacctgg cccccggcgt gccctcccgg 180
ttctccggct ccggctccgg caacgactac accttcacca tctccagcct ggaggccgag 240
gacgccgcca cctattactg ccagcagttc accacctcac cctacacctt cggaggcggg 300
accaagctcg agatcaaacg t 321
<210> SEQ ID NO 70
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (1)..(3)
<223> OTHER INFORMATION: This region may encompass 'GAG' or 'GAC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (4)..(6)
<223> OTHER INFORMATION: This region may encompass 'AAC' or 'GTG'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (10)..(12)
<223> OTHER INFORMATION: This region may encompass 'CTG' or 'ATG'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (136)..(138)
<223> OTHER INFORMATION: This region may encompass 'TGG' or 'CTG'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (142)..(144)
<223> OTHER INFORMATION: This region may encompass 'TAT' or 'AAG'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (202)..(204)
<223> OTHER INFORMATION: This region may encompass 'AAC' or 'ACC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (208)..(210)
<223> OTHER INFORMATION: This region may encompass 'TAC' or 'TTC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (262)..(264)
<223> OTHER INFORMATION: This region may encompass 'CAG' or 'TAC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (268)..(270)
<223> OTHER INFORMATION: This region may encompass 'TTC' or 'TAC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (277)..(279)
<223> OTHER INFORMATION: This region may encompass 'TCC' or 'TAC'
<400> SEQUENCE: 70
nnnnnngtgn nnacccagtc ccccgccttc ctgtccgtga cccccggcga gaaggtgacc 60
atcacctgcc gggcctccag ctccgtgaag tacatgtact ggtaccagca gaagcccgac 120
caggccccca agctgnnnat cnnntacacc tccaacctgg cccccggcgt gccctcccgg 180
ttctccggct ccggctccgg cnnngacnnn accttcacca tctccagcct ggaggccgag 240
gacgccgcca cctattactg cnnncagnnn accaccnnnc cctacacctt cggcggaggc 300
accaagctcg agatcaagcg g 321
<210> SEQ ID NO 71
<211> LENGTH: 120
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 71
Glu Val Met Leu Val Glu Ser Gly Gly Gly Val Val Lys Pro Gly Gly
1 5 10 15
Ser Leu Lys Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Ile Arg Gln Thr Pro Glu Lys Arg Leu Glu Trp Val
35 40 45
Ala Thr Ile Ser Asn Gly Gly Ser Tyr Thr Tyr Tyr Pro Asp Ser Val
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Val Lys Asn Ile Leu Tyr
65 70 75 80
Leu Gln Met Arg Ser Leu Arg Ser Glu Asp Thr Ala Met Tyr Tyr Cys
85 90 95
Ala Arg Arg Ser Ser Asp Tyr Asp Ser Trp Phe Ala Tyr Trp Gly Gln
100 105 110
Gly Thr Leu Val Thr Val Ser Ser
115 120
<210> SEQ ID NO 72
<211> LENGTH: 120
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 72
Glu Val Met Leu Val Glu Ser Gly Gly Gly Phe Val Lys Pro Gly Gly
1 5 10 15
Ser Leu Lys Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Phe
20 25 30
Ala Met Ser Trp Val Arg Gln Thr Pro Glu Lys Arg Leu Glu Trp Val
35 40 45
Ala Thr Ile Ser Asn Gly Gly Thr Tyr Thr Tyr Tyr Pro Asp Ser Val
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Ser Ser Leu Arg Ser Glu Asp Thr Ala Met Tyr Tyr Cys
85 90 95
Ala Arg Arg Ser Ser Asp Tyr Asp Ser Trp Phe Ala Tyr Trp Gly Gln
100 105 110
Gly Thr Leu Val Thr Val Ser Ser
115 120
<210> SEQ ID NO 73
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 73
Glu Val Met Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Lys Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Thr Pro Glu Lys Arg Leu Glu Trp Val
35 40 45
Ala Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Thr Leu Phe
65 70 75 80
Leu Gln Met Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Val Arg His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Asp Tyr Trp Gly Gln Gly Thr Thr Leu Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 74
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 74
Glu Val Met Leu Val Glu Ser Gly Gly Gly Val Val Lys Pro Gly Gly
1 5 10 15
Ser Leu Lys Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Thr Pro Glu Lys Arg Leu Glu Trp Val
35 40 45
Ala Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ile Leu Phe
65 70 75 80
Leu Gln Met Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Val Arg His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Asp Tyr Trp Gly Gln Gly Thr Thr Leu Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 75
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 75
Gln Ile Val Leu Thr Gln Ser Pro Ala Ile Met Ser Ala Ser Pro Gly
1 5 10 15
Glu Lys Val Thr Ile Ser Cys Ser Ala Thr Ser Ser Val Ile Phe Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Gly Ser Ser Pro Lys Pro Trp Ile Tyr
35 40 45
Arg Thr Ser Asn Leu Ala Ser Gly Val Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Thr Ser Tyr Ser Leu Thr Ile Ser Asn Met Glu Ala Glu
65 70 75 80
Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Tyr Gln Ser Tyr Pro Leu Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Leu Lys Arg
100 105
<210> SEQ ID NO 76
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 76
Gln Ile Val Leu Thr Gln Ser Pro Ala Ile Met Ser Ala Ser Pro Gly
1 5 10 15
Glu Arg Val Thr Ile Ser Cys Ser Ala Ser Ser Ser Val Ile Phe Leu
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Gly Ser Ser Pro Lys Pro Arg Ile Tyr
35 40 45
Arg Thr Ser Asn Leu Ala Ser Gly Val Pro Asp Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Thr Ser Tyr Ser Leu Thr Ile Ser Ser Met Glu Ala Glu
65 70 75 80
Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Tyr His Ser Tyr Pro Leu Thr
85 90 95
Phe Gly Ala Gly Thr Lys Leu Glu Leu Lys Arg
100 105
<210> SEQ ID NO 77
<211> LENGTH: 126
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 77
Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Ala Ile Ser Gly Ser Gly Gly Ser Ile Tyr Tyr Ala Asp Ser Val
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys Asp Val Leu Val Val Pro Ala Ala Pro Tyr Tyr Tyr Tyr Gly
100 105 110
Met Asp Val Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 78
<211> LENGTH: 98
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 78
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Ala Ile Ser Gly Ser Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys
<210> SEQ ID NO 79
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (99)..(108)
<223> OTHER INFORMATION: Any amino acid
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (110)..(111)
<223> OTHER INFORMATION: Any amino acid
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (113)..(114)
<223> OTHER INFORMATION: Any amino acid
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (116)..(116)
<223> OTHER INFORMATION: Any amino acid
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (123)..(123)
<223> OTHER INFORMATION: Any amino acid
<400> SEQUENCE: 79
Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Ala Ile Ser Gly Ser Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys Xaa Xaa Xaa Xaa Xaa Xaa Xaa Xaa Xaa Xaa Tyr Xaa Xaa Tyr
100 105 110
Xaa Xaa Asp Xaa Trp Gly Gln Gly Thr Thr Xaa Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 80
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (3)..(3)
<223> OTHER INFORMATION: Met or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (5)..(5)
<223> OTHER INFORMATION: Val or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (13)..(13)
<223> OTHER INFORMATION: Lys or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (19)..(19)
<223> OTHER INFORMATION: Lys or Arg
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (40)..(40)
<223> OTHER INFORMATION: Thr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (42)..(42)
<223> OTHER INFORMATION: Glu or Gly
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (44)..(44)
<223> OTHER INFORMATION: Arg or Gly
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (49)..(49)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (75)..(75)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (80)..(80)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (84)..(84)
<223> OTHER INFORMATION: Ser or Asn
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (88)..(88)
<223> OTHER INFORMATION: Ser or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (97)..(97)
<223> OTHER INFORMATION: Val or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (98)..(98)
<223> OTHER INFORMATION: Arg or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (123)..(123)
<223> OTHER INFORMATION: Leu or Val
<400> SEQUENCE: 80
Glu Val Xaa Leu Xaa Glu Ser Gly Gly Gly Leu Val Xaa Pro Gly Gly
1 5 10 15
Ser Leu Xaa Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Xaa Pro Xaa Lys Xaa Leu Glu Trp Val
35 40 45
Xaa Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Xaa Lys Asn Thr Leu Xaa
65 70 75 80
Leu Gln Met Xaa Ser Leu Arg Xaa Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Xaa Xaa His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Asp Tyr Trp Gly Gln Gly Thr Thr Xaa Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 81
<211> LENGTH: 106
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 81
Glu Asn Val Leu Thr Gln Ser Pro Ala Ile Met Ser Ala Ser Leu Gly
1 5 10 15
Glu Lys Val Thr Met Asn Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Ser Asp Ala Ser Pro Lys Leu Trp Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Val Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Asn Ser Tyr Ser Leu Thr Ile Ser Ser Met Glu Gly Glu
65 70 75 80
Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys
100 105
<210> SEQ ID NO 82
<211> LENGTH: 108
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 82
Asp Ile Gln Leu Thr Gln Ser Pro Ala Ile Met Ser Ala Ser Pro Gly
1 5 10 15
Glu Lys Val Thr Met Thr Cys Arg Ala Ser Ser Ser Val Ser Ser Ser
20 25 30
Tyr Leu His Trp Tyr Gln Gln Lys Ser Gly Ala Ser Pro Lys Leu Trp
35 40 45
Ile Tyr Ser Thr Ser Asn Leu Ala Ser Gly Val Pro Ala Arg Phe Ser
50 55 60
Gly Ser Gly Ser Gly Thr Ser Tyr Ser Leu Thr Ile Ser Ser Val Glu
65 70 75 80
Ala Glu Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Tyr Ser Gly Tyr Pro
85 90 95
Tyr Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys
100 105
<210> SEQ ID NO 83
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Any amino acid
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (30)..(31)
<223> OTHER INFORMATION: Any amino acid
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (78)..(78)
<223> OTHER INFORMATION: Any amino acid
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (91)..(94)
<223> OTHER INFORMATION: Any amino acid
<400> SEQUENCE: 83
Asp Xaa Val Leu Thr Gln Ser Pro Ala Ile Met Ser Ala Ser Pro Gly
1 5 10 15
Glu Lys Val Thr Met Thr Cys Arg Ala Ser Ser Ser Val Xaa Xaa Tyr
20 25 30
Leu Tyr Trp Tyr Gln Gln Lys Ser Asp Ala Ser Pro Lys Leu Trp Ile
35 40 45
Tyr Tyr Thr Ser Asn Leu Ala Ser Gly Val Pro Ala Arg Phe Ser Gly
50 55 60
Ser Gly Ser Gly Thr Ser Tyr Ser Leu Thr Ile Ser Ser Xaa Glu Ala
65 70 75 80
Glu Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Xaa Xaa Xaa Xaa Pro Tyr
85 90 95
Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys
100 105
<210> SEQ ID NO 84
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Glu or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Asn, Val or Ile
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Leu or Met
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (10)..(10)
<223> OTHER INFORMATION: Ile or Phe
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (11)..(11)
<223> OTHER INFORMATION: Met or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (13)..(13)
<223> OTHER INFORMATION: Ala or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (14)..(14)
<223> OTHER INFORMATION: Ser or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (15)..(15)
<223> OTHER INFORMATION: Leu or Pro
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (21)..(21)
<223> OTHER INFORMATION: Met or Ile
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (22)..(22)
<223> OTHER INFORMATION: Asn or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (39)..(39)
<223> OTHER INFORMATION: Ser or Pro
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (40)..(40)
<223> OTHER INFORMATION: Asp or Gly
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (41)..(41)
<223> OTHER INFORMATION: Ala or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (42)..(42)
<223> OTHER INFORMATION: Ser or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (59)..(59)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (69)..(69)
<223> OTHER INFORMATION: Ser or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (71)..(71)
<223> OTHER INFORMATION: Ser or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (72)..(72)
<223> OTHER INFORMATION: Leu or Phe
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (77)..(77)
<223> OTHER INFORMATION: Met, Leu or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (79)..(79)
<223> OTHER INFORMATION: Gly or Ala
<400> SEQUENCE: 84
Xaa Xaa Val Xaa Thr Gln Ser Pro Ala Xaa Xaa Ser Xaa Xaa Xaa Gly
1 5 10 15
Glu Lys Val Thr Xaa Xaa Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Xaa Xaa Xaa Xaa Pro Lys Leu Trp Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Val Pro Xaa Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Asn Xaa Tyr Xaa Xaa Thr Ile Ser Ser Xaa Glu Xaa Glu
65 70 75 80
Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
1
SEQUENCE LISTING
<160> NUMBER OF SEQ ID NOS: 84
<210> SEQ ID NO 1
<211> LENGTH: 5
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 1
Asn Tyr Met Ala Ser
1 5
<210> SEQ ID NO 2
<211> LENGTH: 17
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 2
Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu Lys
1 5 10 15
Gly
<210> SEQ ID NO 3
<211> LENGTH: 18
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (17)..(17)
<223> OTHER INFORMATION: Asp or Tyr
<400> SEQUENCE: 3
His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr Tyr Phe
1 5 10 15
Xaa Tyr
<210> SEQ ID NO 4
<211> LENGTH: 10
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 4
Arg Ala Ser Ser Ser Val Lys Tyr Met Tyr
1 5 10
<210> SEQ ID NO 5
<211> LENGTH: 7
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 5
Tyr Thr Ser Asn Leu Ala Pro
1 5
<210> SEQ ID NO 6
<211> LENGTH: 9
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 6
Gln Gln Phe Thr Thr Ser Pro Tyr Thr
1 5
<210> SEQ ID NO 7
<211> LENGTH: 18
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 7
His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr Tyr Phe
1 5 10 15
Tyr Tyr
<210> SEQ ID NO 8
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Glu or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Ile, Asn or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Leu or Met
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (10)..(10)
<223> OTHER INFORMATION: Thr, Phe or Ile
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (11)..(11)
<223> OTHER INFORMATION: Leu or Met
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (13)..(13)
<223> OTHER INFORMATION: Leu, Ala or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (14)..(14)
<223> OTHER INFORMATION: Ser or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (15)..(15)
<223> OTHER INFORMATION: Leu or Pro
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (18)..(18)
<223> OTHER INFORMATION: Arg or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (19)..(19)
<223> OTHER INFORMATION: Ala or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (21)..(21)
<223> OTHER INFORMATION: Leu, Met or Ile
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (22)..(22)
<223> OTHER INFORMATION: Ser, Thr or Asn
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (39)..(39)
<223> OTHER INFORMATION: Ser or Pro
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (40)..(40)
<223> OTHER INFORMATION: Gly or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (41)..(41)
<223> OTHER INFORMATION: Gln or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (42)..(42)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (44)..(44)
<223> OTHER INFORMATION: Arg or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (46)..(46)
<223> OTHER INFORMATION: Leu or Trp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (48)..(48)
<223> OTHER INFORMATION: Tyr or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (57)..(57)
<223> OTHER INFORMATION: Ile or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (59)..(59)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (68)..(68)
<223> OTHER INFORMATION: Thr or Asn
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (69)..(69)
<223> OTHER INFORMATION: Asp or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (70)..(70)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (71)..(71)
<223> OTHER INFORMATION: Thr or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (72)..(72)
<223> OTHER INFORMATION: Leu or Phe
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (77)..(77)
<223> OTHER INFORMATION: Met or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (79)..(79)
<223> OTHER INFORMATION: Ala, Gly or Pro
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (82)..(82)
<223> OTHER INFORMATION: Phe or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (84)..(84)
<223> OTHER INFORMATION: Val or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (88)..(88)
<223> OTHER INFORMATION: Gln or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (93)..(93)
<223> OTHER INFORMATION: Ser or Tyr
<400> SEQUENCE: 8
Xaa Xaa Val Xaa Thr Gln Ser Pro Ala Xaa Xaa Ser Xaa Xaa Xaa Gly
1 5 10 15
Glu Xaa Xaa Thr Xaa Xaa Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Xaa Xaa Xaa Xaa Pro Xaa Leu Xaa Ile Xaa
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Xaa Pro Xaa Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Xaa Xaa Xaa Xaa Xaa Thr Ile Ser Ser Xaa Glu Xaa Glu
65 70 75 80
Asp Xaa Ala Xaa Tyr Tyr Cys Xaa Gln Phe Thr Thr Xaa Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 9
<211> LENGTH: 126
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 9
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Lys Gly Leu Glu Trp Val Ser
35 40 45
Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu Lys
50 55 60
Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr Leu
65 70 75 80
Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys Ala
85 90 95
Lys His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr Tyr
100 105 110
Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 10
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Ile or Asn
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (10)..(10)
<223> OTHER INFORMATION: Thr, Phe or Ile
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (11)..(11)
<223> OTHER INFORMATION: Leu or Met
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (13)..(13)
<223> OTHER INFORMATION: Leu, Ala or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (14)..(14)
<223> OTHER INFORMATION: Ser or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (15)..(15)
<223> OTHER INFORMATION: Leu or Pro
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (18)..(18)
<223> OTHER INFORMATION: Arg or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (19)..(19)
<223> OTHER INFORMATION: Ala or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (21)..(21)
<223> OTHER INFORMATION: Leu, Met or Ile
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (22)..(22)
<223> OTHER INFORMATION: Ser, Thr or Asn
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (39)..(39)
<223> OTHER INFORMATION: Ser or Pro
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (40)..(40)
<223> OTHER INFORMATION: Gly or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (41)..(41)
<223> OTHER INFORMATION: Gln or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (42)..(42)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (44)..(44)
<223> OTHER INFORMATION: Arg or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (46)..(46)
<223> OTHER INFORMATION: Leu or Trp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (57)..(57)
<223> OTHER INFORMATION: Ile or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (59)..(59)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (68)..(68)
<223> OTHER INFORMATION: Thr or Asn
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (69)..(69)
<223> OTHER INFORMATION: Asp or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (70)..(70)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (71)..(71)
<223> OTHER INFORMATION: Thr or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (72)..(72)
<223> OTHER INFORMATION: Leu or Phe
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (77)..(77)
<223> OTHER INFORMATION: Met or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (79)..(79)
<223> OTHER INFORMATION: Ala or Gly
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (82)..(82)
<223> OTHER INFORMATION: Phe or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (84)..(84)
<223> OTHER INFORMATION: Val or Thr
<400> SEQUENCE: 10
Glu Xaa Val Leu Thr Gln Ser Pro Ala Xaa Xaa Ser Xaa Xaa Xaa Gly
1 5 10 15
Glu Xaa Xaa Thr Xaa Xaa Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Xaa Xaa Xaa Xaa Pro Xaa Leu Xaa Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Xaa Pro Xaa Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Xaa Xaa Xaa Xaa Xaa Thr Ile Ser Ser Xaa Glu Xaa Glu
65 70 75 80
Asp Xaa Ala Xaa Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 11
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 11
Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Leu Gly
1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Ser Gly Gln Ala Pro Arg Leu Leu Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Ile Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Met Glu Ala Glu
65 70 75 80
Asp Phe Ala Val Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 12
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 12
Glu Asn Val Leu Thr Gln Ser Pro Ala Phe Leu Ser Val Thr Pro Gly
1 5 10 15
Glu Lys Val Thr Ile Thr Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Asp Gln Ala Pro Lys Leu Trp Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Val Pro Ser Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Asn Asp Tyr Thr Phe Thr Ile Ser Ser Leu Glu Ala Glu
65 70 75 80
Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 13
<211> LENGTH: 5
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Asn, Ser or Tyr
<400> SEQUENCE: 13
Xaa Tyr Ala Met Ser
1 5
<210> SEQ ID NO 14
<211> LENGTH: 17
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Thr, Ala or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (3)..(3)
<223> OTHER INFORMATION: Ser or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Ser, Gly, Thr or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (5)..(5)
<223> OTHER INFORMATION: Gly or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (6)..(6)
<223> OTHER INFORMATION: Gly or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (8)..(8)
<223> OTHER INFORMATION: Phe, Ser or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (12)..(12)
<223> OTHER INFORMATION: Pro or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (15)..(15)
<223> OTHER INFORMATION: Leu, Tyr or Val
<400> SEQUENCE: 14
Xaa Ile Xaa Xaa Xaa Xaa Gly Xaa Thr Tyr Tyr Xaa Asp Ser Xaa Lys
1 5 10 15
Gly
<210> SEQ ID NO 15
<211> LENGTH: 18
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: His or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Gly or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (3)..(3)
<223> OTHER INFORMATION: Ala or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (6)..(6)
<223> OTHER INFORMATION: Thr or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (8)..(8)
<223> OTHER INFORMATION: Val or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (12)..(12)
<223> OTHER INFORMATION: Ala or Tyr
<400> SEQUENCE: 15
Xaa Xaa Xaa Pro Met Xaa Thr Xaa Ile Thr Tyr Xaa Pro Tyr Tyr Phe
1 5 10 15
Tyr Tyr
<210> SEQ ID NO 16
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (31)..(31)
<223> OTHER INFORMATION: Asn, Ser or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (50)..(50)
<223> OTHER INFORMATION: Thr, Ala or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (52)..(52)
<223> OTHER INFORMATION: Ser or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (53)..(53)
<223> OTHER INFORMATION: Ser, Gly, Thr or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (54)..(54)
<223> OTHER INFORMATION: Gly or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (55)..(55)
<223> OTHER INFORMATION: Gly or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (57)..(57)
<223> OTHER INFORMATION: Phe, Ser or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (61)..(61)
<223> OTHER INFORMATION: Pro or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (64)..(64)
<223> OTHER INFORMATION: Leu, Tyr or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (99)..(99)
<223> OTHER INFORMATION: His or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (100)..(100)
<223> OTHER INFORMATION: Gly or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (101)..(101)
<223> OTHER INFORMATION: Ala or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (104)..(104)
<223> OTHER INFORMATION: Thr or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (106)..(106)
<223> OTHER INFORMATION: Val or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (110)..(110)
<223> OTHER INFORMATION: Ala or Tyr
<400> SEQUENCE: 16
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Xaa Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Xaa Ile Xaa Xaa Xaa Xaa Gly Xaa Thr Tyr Tyr Xaa Asp Ser Xaa
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys Xaa Xaa Xaa Pro Met Xaa Thr Xaa Ile Thr Tyr Xaa Pro Tyr
100 105 110
Tyr Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 17
<211> LENGTH: 5
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 17
Asn Tyr Ala Ile Ser
1 5
<210> SEQ ID NO 18
<211> LENGTH: 17
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 18
Thr Tyr Ser Ser Gly Gly Val Tyr Thr Tyr Tyr Arg Asp Ser Leu Lys
1 5 10 15
Gly
<210> SEQ ID NO 19
<211> LENGTH: 18
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 19
His Gly Ala Ala Met Thr Thr Val Ile Thr Tyr Ala Pro Phe Tyr Phe
1 5 10 15
Tyr Tyr
<210> SEQ ID NO 20
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 20
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Thr Tyr Ser Ser Gly Gly Val Tyr Thr Tyr Tyr Arg Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys His Gly Ala Ala Met Thr Thr Val Ile Thr Tyr Ala Pro Phe
100 105 110
Tyr Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 21
<211> LENGTH: 5
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 21
Tyr Tyr Ala Met Ser
1 5
<210> SEQ ID NO 22
<211> LENGTH: 17
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 22
Thr Ile Tyr Ser Gly Gly Ser Tyr Thr Phe Tyr Pro Asp Ser Leu Glu
1 5 10 15
Gly
<210> SEQ ID NO 23
<211> LENGTH: 18
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 23
His Gly Ala Pro Met Ser Thr Glu Ile Thr Tyr Ala Pro Tyr Tyr Phe
1 5 10 15
Tyr Tyr
<210> SEQ ID NO 24
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 24
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Tyr Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Thr Ile Tyr Ser Gly Gly Ser Tyr Thr Phe Tyr Pro Asp Ser Leu
50 55 60
Glu Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys His Gly Ala Pro Met Ser Thr Glu Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 25
<211> LENGTH: 5
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 25
Asn Tyr Ala Met Ser
1 5
<210> SEQ ID NO 26
<211> LENGTH: 17
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 26
Thr Ile Tyr Ser Gly Gly Gly Tyr Thr Phe Tyr Leu Asp Ser Leu Lys
1 5 10 15
Gly
<210> SEQ ID NO 27
<211> LENGTH: 18
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 27
His Ser Tyr Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr Tyr Phe
1 5 10 15
Tyr Tyr
<210> SEQ ID NO 28
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 28
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Tyr Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Thr Ile Tyr Ser Gly Gly Gly Tyr Thr Phe Tyr Leu Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys His Ser Tyr Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 29
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 29
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 30
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 30
Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly
1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Ile Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro Glu
65 70 75 80
Asp Phe Ala Val Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 31
<211> LENGTH: 9
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 31
Tyr Gln Phe Thr Thr Ser Pro Tyr Thr
1 5
<210> SEQ ID NO 32
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 32
Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly
1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Ile Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro Glu
65 70 75 80
Asp Phe Ala Val Tyr Tyr Cys Tyr Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 33
<211> LENGTH: 9
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 33
Gln Gln Tyr Thr Thr Ser Pro Tyr Thr
1 5
<210> SEQ ID NO 34
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 34
Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly
1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Ile Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro Glu
65 70 75 80
Asp Phe Ala Val Tyr Tyr Cys Gln Gln Tyr Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 35
<211> LENGTH: 9
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 35
Gln Gln Phe Thr Thr Tyr Pro Tyr Thr
1 5
<210> SEQ ID NO 36
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 36
Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly
1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Ile Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro Glu
65 70 75 80
Asp Phe Ala Val Tyr Tyr Cys Gln Gln Phe Thr Thr Tyr Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 37
<211> LENGTH: 11
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Ser or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (7)..(7)
<223> OTHER INFORMATION: Lys or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (10)..(10)
<223> OTHER INFORMATION: Met or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (11)..(11)
<223> OTHER INFORMATION: Tyr or Ala
<400> SEQUENCE: 37
Arg Ala Ser Xaa Ser Val Xaa Ser Tyr Xaa Xaa
1 5 10
<210> SEQ ID NO 38
<211> LENGTH: 9
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Gln or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (3)..(3)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (6)..(6)
<223> OTHER INFORMATION: Ser or Tyr
<400> SEQUENCE: 38
Xaa Gln Xaa Thr Thr Xaa Pro Tyr Thr
1 5
<210> SEQ ID NO 39
<211> LENGTH: 108
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (27)..(27)
<223> OTHER INFORMATION: Ser or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (30)..(30)
<223> OTHER INFORMATION: Lys or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (33)..(33)
<223> OTHER INFORMATION: Met or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (34)..(34)
<223> OTHER INFORMATION: Tyr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (50)..(50)
<223> OTHER INFORMATION: Tyr or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (51)..(51)
<223> OTHER INFORMATION: Thr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (54)..(54)
<223> OTHER INFORMATION: Leu or Arg
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (56)..(56)
<223> OTHER INFORMATION: Pro or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (89)..(89)
<223> OTHER INFORMATION: Gln or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (91)..(91)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (94)..(94)
<223> OTHER INFORMATION: Ser or Tyr
<400> SEQUENCE: 39
Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly
1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Xaa Ser Val Xaa Ser Tyr
20 25 30
Xaa Xaa Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile
35 40 45
Tyr Xaa Xaa Ser Asn Xaa Ala Xaa Gly Ile Pro Ala Arg Phe Ser Gly
50 55 60
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro
65 70 75 80
Glu Asp Phe Ala Val Tyr Tyr Cys Xaa Gln Xaa Thr Thr Xaa Pro Tyr
85 90 95
Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 40
<211> LENGTH: 7
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Tyr or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Thr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (5)..(5)
<223> OTHER INFORMATION: Leu or Arg
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (7)..(7)
<223> OTHER INFORMATION: Pro or Thr
<400> SEQUENCE: 40
Xaa Xaa Ser Asn Xaa Ala Xaa
1 5
<210> SEQ ID NO 41
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Glu or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Asn or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Leu or Met
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (46)..(46)
<223> OTHER INFORMATION: Trp or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (48)..(48)
<223> OTHER INFORMATION: Tyr or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (68)..(68)
<223> OTHER INFORMATION: Asn or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (70)..(70)
<223> OTHER INFORMATION: Tyr or Phe
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (88)..(88)
<223> OTHER INFORMATION: Gln or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (90)..(90)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (93)..(93)
<223> OTHER INFORMATION: Ser or Tyr
<400> SEQUENCE: 41
Xaa Xaa Val Xaa Thr Gln Ser Pro Ala Phe Leu Ser Val Thr Pro Gly
1 5 10 15
Glu Lys Val Thr Ile Thr Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Asp Gln Ala Pro Lys Leu Xaa Ile Xaa
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Val Pro Ser Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Xaa Asp Xaa Thr Phe Thr Ile Ser Ser Leu Glu Ala Glu
65 70 75 80
Asp Ala Ala Thr Tyr Tyr Cys Xaa Gln Xaa Thr Thr Xaa Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 42
<211> LENGTH: 6
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Any amino acid
<400> SEQUENCE: 42
Leu Val His Xaa Val Lys
1 5
<210> SEQ ID NO 43
<211> LENGTH: 6
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Any amino acid
<400> SEQUENCE: 43
Leu Val Lys Xaa Val His
1 5
<210> SEQ ID NO 44
<211> LENGTH: 98
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 44
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Gln Val
35 40 45
Ser Ala Ile Ser Gly Ser Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys
<210> SEQ ID NO 45
<211> LENGTH: 95
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 45
Glu Ile Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Pro Gly
1 5 10 15
Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Ser Tyr
20 25 30
Leu Ala Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu Ile
35 40 45
Tyr Asp Ala Ser Asn Arg Ala Thr Gly Ile Pro Ala Arg Phe Ser Gly
50 55 60
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Glu Pro
65 70 75 80
Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Arg Ser Asn Trp Pro
85 90 95
<210> SEQ ID NO 46
<211> LENGTH: 95
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 46
Asp Val Val Met Thr Gln Ser Pro Ala Phe Leu Ser Val Thr Pro Gly
1 5 10 15
Glu Lys Val Thr Ile Thr Cys Gln Ala Ser Glu Gly Ile Gly Asn Tyr
20 25 30
Leu Tyr Trp Tyr Gln Gln Lys Pro Asp Gln Ala Pro Lys Leu Leu Ile
35 40 45
Lys Tyr Ala Ser Gln Ser Ile Ser Gly Val Pro Ser Arg Phe Ser Gly
50 55 60
Ser Gly Ser Gly Thr Asp Phe Thr Phe Thr Ile Ser Ser Leu Glu Ala
65 70 75 80
Glu Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Gly Asn Lys His Pro
85 90 95
<210> SEQ ID NO 47
<211> LENGTH: 18
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 47
His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr Tyr Phe
1 5 10 15
Asp Tyr
<210> SEQ ID NO 48
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (3)..(3)
<223> OTHER INFORMATION: Met or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (5)..(5)
<223> OTHER INFORMATION: Val or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (13)..(13)
<223> OTHER INFORMATION: Lys or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (19)..(19)
<223> OTHER INFORMATION: Lys or Arg
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (40)..(40)
<223> OTHER INFORMATION: Thr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (42)..(42)
<223> OTHER INFORMATION: Glu or Gly
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (44)..(44)
<223> OTHER INFORMATION: Arg or Gly
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (49)..(49)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (75)..(75)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (80)..(80)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (84)..(84)
<223> OTHER INFORMATION: Ser or Asn
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (88)..(88)
<223> OTHER INFORMATION: Ser or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (97)..(97)
<223> OTHER INFORMATION: Val or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (98)..(98)
<223> OTHER INFORMATION: Arg or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (115)..(115)
<223> OTHER INFORMATION: Asp or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (123)..(123)
<223> OTHER INFORMATION: Leu or Val
<400> SEQUENCE: 48
Glu Val Xaa Leu Xaa Glu Ser Gly Gly Gly Leu Val Xaa Pro Gly Gly
1 5 10 15
Ser Leu Xaa Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Xaa Pro Xaa Lys Xaa Leu Glu Trp Val
35 40 45
Xaa Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Xaa Lys Asn Thr Leu Xaa
65 70 75 80
Leu Gln Met Xaa Ser Leu Arg Xaa Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Xaa Xaa His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Xaa Tyr Trp Gly Gln Gly Thr Thr Xaa Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 49
<211> LENGTH: 108
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 49
Glu Ile Asn Val Leu Thr Gln Ser Pro Ala Thr Leu Ser Leu Ser Leu
1 5 10 15
Gly Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Ser Ser Val Lys Tyr
20 25 30
Met Tyr Trp Tyr Gln Gln Lys Ser Gly Gln Ala Pro Arg Leu Leu Ile
35 40 45
Tyr Tyr Thr Ser Asn Leu Ala Pro Gly Ile Pro Ala Arg Phe Ser Gly
50 55 60
Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Met Glu Ala
65 70 75 80
Glu Asp Phe Ala Val Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr
85 90 95
Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 50
<211> LENGTH: 108
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Glu or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Asn or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Leu or Met
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (27)..(27)
<223> OTHER INFORMATION: Ser or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (30)..(30)
<223> OTHER INFORMATION: Lys or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (33)..(33)
<223> OTHER INFORMATION: Met or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (34)..(34)
<223> OTHER INFORMATION: Tyr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (47)..(47)
<223> OTHER INFORMATION: Trp or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (49)..(49)
<223> OTHER INFORMATION: Tyr or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (50)..(50)
<223> OTHER INFORMATION: Tyr or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (51)..(51)
<223> OTHER INFORMATION: Thr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (54)..(54)
<223> OTHER INFORMATION: Leu or Arg
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (56)..(56)
<223> OTHER INFORMATION: Pro or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (69)..(69)
<223> OTHER INFORMATION: Asn or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (71)..(71)
<223> OTHER INFORMATION: Tyr or Phe
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (89)..(89)
<223> OTHER INFORMATION: Gln or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (91)..(91)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (94)..(94)
<223> OTHER INFORMATION: Ser or Tyr
<400> SEQUENCE: 50
Xaa Xaa Val Xaa Thr Gln Ser Pro Ala Phe Leu Ser Val Thr Pro Gly
1 5 10 15
Glu Lys Val Thr Ile Thr Cys Arg Ala Ser Xaa Ser Val Xaa Ser Tyr
20 25 30
Xaa Xaa Trp Tyr Gln Gln Lys Pro Asp Gln Ala Pro Lys Leu Xaa Ile
35 40 45
Xaa Xaa Xaa Ser Asn Xaa Ala Xaa Gly Val Pro Ser Arg Phe Ser Gly
50 55 60
Ser Gly Ser Gly Xaa Asp Xaa Thr Phe Thr Ile Ser Ser Leu Glu Ala
65 70 75 80
Glu Asp Ala Ala Thr Tyr Tyr Cys Xaa Gln Xaa Thr Thr Xaa Pro Tyr
85 90 95
Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 51
<211> LENGTH: 381
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 51
gaagtgatgc tggtggagtc tgggggaggc ttagtgaagc ctggagggtc cctgaaactc 60
tcctgtgcag cctctggatt cactttcagt aactatgcca tgtcttgggt tcgtcagact 120
ccggagaaga ggctggagtg ggtcgcaacc ataagtagtg gtggaggtta cacctattat 180
ccagacagtt tgaaggggcg attcaccatc tccagagaca atgccaagaa caccctgttc 240
ctgcaaatga gcagtctgag gtctgaggac acggccgtgt attattgtgt aagacatggg 300
gcccccatga ctacggtgat aacttacgct ccttactatt ttgactactg gggccagggc 360
accactctca cagtctcctc a 381
<210> SEQ ID NO 52
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 52
Glu Val Met Leu Val Glu Ser Gly Gly Gly Leu Val Lys Pro Gly Gly
1 5 10 15
Ser Leu Lys Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Thr Pro Glu Lys Arg Leu Glu Trp Val
35 40 45
Ala Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Thr Leu Phe
65 70 75 80
Leu Gln Met Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Val Arg His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Asp Tyr Trp Gly Gln Gly Thr Thr Leu Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 53
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 53
gaaaatgtgc tcacccagtc tccagcaatc atgtctgcat ctctggggga gaaggtcacc 60
atgaactgca gggccagctc aagtgtaaaa tacatgtact ggtaccagca gaagtcagat 120
gcctccccca aactatggat ttattacaca tccaacctgg ctcctggagt cccagctcgc 180
ttcagtggca gtgggtctgg gaactcttat tctctcacaa tcagcagcat ggagggtgaa 240
gatgctgcca cttattactg ccagcagttt actacttccc cgtacacgtt cggagggggg 300
accaagctgg aaataaaacg g 321
<210> SEQ ID NO 54
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 54
Glu Asn Val Leu Thr Gln Ser Pro Ala Ile Met Ser Ala Ser Leu Gly
1 5 10 15
Glu Lys Val Thr Met Asn Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Ser Asp Ala Ser Pro Lys Leu Trp Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Val Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Asn Ser Tyr Ser Leu Thr Ile Ser Ser Met Glu Gly Glu
65 70 75 80
Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
<210> SEQ ID NO 55
<211> LENGTH: 381
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (91)..(93)
<223> OTHER INFORMATION: This region may encompass 'TAC' or 'TCA'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (148)..(150)
<223> OTHER INFORMATION: This region may encompass 'TAC' or 'GCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (154)..(156)
<223> OTHER INFORMATION: This region may encompass 'TCC' or 'TAC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (157)..(159)
<223> OTHER INFORMATION: This region may encompass 'GGT' or 'ACC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (169)..(171)
<223> OTHER INFORMATION: This region may encompass 'TTC' or 'TCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (181)..(183)
<223> OTHER INFORMATION: This region may encompass 'CCC' or 'GCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (190)..(192)
<223> OTHER INFORMATION: This region may encompass 'CTG' or 'TAC'
<400> SEQUENCE: 55
gaggtgcagc tgctggagtc cggcggaggc ctggtgcagc ccggcggctc cctgcggctg 60
tcctgcgccg cctccggctt caccttctcc nnntacgcca tgtcctgggt gcggcaggcc 120
cccggcaagg gcctggagtg ggtgtccnnn atcnnnnnnt ccggaggcnn nacctactat 180
nnngactccn nnaagggccg gttcaccatc tcccgggaca actccaagaa caccctgtac 240
ctgcagatga actccctgcg ggccgaagac accgccgtgt actattgcgc caagcatgac 300
gccctcatga ccactgtgat cacctacgcc ccctcttact tctactactg gggccagggc 360
accacggtca ccgtctcctc a 381
<210> SEQ ID NO 56
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (31)..(31)
<223> OTHER INFORMATION: Tyr or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (50)..(50)
<223> OTHER INFORMATION: Tyr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (52)..(52)
<223> OTHER INFORMATION: Ser or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (53)..(53)
<223> OTHER INFORMATION: Gly or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (57)..(57)
<223> OTHER INFORMATION: Phe or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (61)..(61)
<223> OTHER INFORMATION: Pro or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (64)..(64)
<223> OTHER INFORMATION: Leu or Tyr
<400> SEQUENCE: 56
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Xaa Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Xaa Ile Xaa Xaa Ser Gly Gly Xaa Thr Tyr Tyr Xaa Asp Ser Xaa
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys His Asp Ala Leu Met Thr Thr Val Ile Thr Tyr Ala Pro Ser
100 105 110
Tyr Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 57
<211> LENGTH: 381
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 57
gaggtgcagc tgctggagtc cggcggaggc ctggtgcagc ccggcggctc cctgcggctg 60
tcctgcgccg cctccggctt caccttctcc aactacgcca tttcctgggt gcggcaggcc 120
cccggcaagg gcctggagtg ggtgtccacc tactcctccg gcggagtcta cacctactat 180
cgcgactccc tgaagggccg gttcaccatc tcccgggaca actccaagaa caccctgtac 240
ctgcagatga actccctgcg ggccgaagac accgccgtgt actattgcgc caagcacggc 300
gccgccatga ccactgtgat cacctatgcc cccttttact tctactactg gggccagggc 360
accacggtca ccgtctcctc a 381
<210> SEQ ID NO 58
<211> LENGTH: 381
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 58
gaggtgcagc tgctggagtc cggcggaggc ctggtgcagc ccggcggctc cctgcggctg 60
tcctgcgccg cctccggctt caccttctcc tactacgcca tgtcctgggt gcggcaggcc 120
cccggcaagg gcctggagtg ggtgtccacc atctactccg gcggaagcta caccttctat 180
cccgactccc tggagggccg gttcaccatc tcccgggaca actccaagaa caccctgtac 240
ctgcagatga actccctgcg ggccgaagac accgccgtgt actattgcgc caagcacggc 300
gcccccatga gcactgagat cacctacgcc ccctattact tctactactg gggccagggc 360
accacggtca ccgtctcctc a 381
<210> SEQ ID NO 59
<211> LENGTH: 381
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 59
gaggtgcagc tgctggagtc cggcggaggc ctggtgcagc ccggcggctc cctgcggctg 60
tcctgcgccg cctccggctt caccttctcc aactactaca tgtcctgggt gcggcaggcc 120
cccggcaagg gcctggagtg ggtgtccacc atctactccg gcggaggcta caccttctat 180
ctcgactccc tgaagggccg gttcaccatc tcccgggaca actccaagaa caccctgtac 240
ctgcagatga actccctgcg ggccgaagac accgccgtgt actattgcgc caagcacagc 300
taccccatga ccactgtgat cacctacgcc ccctattact tctactactg gggccagggc 360
accacggtca ccgtctcctc a 381
<210> SEQ ID NO 60
<211> LENGTH: 381
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 60
gaggtgcagc tgctggagtc cggcggaggc ctggtgcagc ccggcggctc cctgcggctg 60
tcctgcgccg cctccggctt caccttctcc aactacgcca tgtcctgggt gcggcaggcc 120
cccggcaagg gcctggagtg ggtgtccacc atctcctccg gcggaggcta cacctactat 180
cccgactccc tgaagggccg gttcaccatc tcccgggaca actccaagaa caccctgtac 240
ctgcagatga actccctgcg ggccgaagac accgccgtgt actattgcgc caagcacggc 300
gcccccatga ccactgtgat cacctacgcc ccctattact tctactactg gggccagggc 360
accacggtca ccgtctcctc a 381
<210> SEQ ID NO 61
<211> LENGTH: 381
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (91)..(93)
<223> OTHER INFORMATION: This region may encompass 'AAC' or 'TCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (148)..(150)
<223> OTHER INFORMATION: This region may encompass 'ACC' or 'GCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (157)..(159)
<223> OTHER INFORMATION: This region may encompass 'TCC' or 'GGA'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (160)..(162)
<223> OTHER INFORMATION: This region may encompass 'GGC' or 'TCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (169)..(171)
<223> OTHER INFORMATION: This region may encompass 'TAC' or 'TCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (181)..(183)
<223> OTHER INFORMATION: This region may encompass 'CCC' or 'GCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (190)..(192)
<223> OTHER INFORMATION: This region may encompass 'CTG' or 'GTC'
<400> SEQUENCE: 61
gaggtgcagc tgctggagtc cggcggaggc ctggtgcagc ccggcggctc cctgcggctg 60
tcctgcgccg cctccggctt caccttctcc nnntacgcca tgtcctgggt gcggcaggcc 120
cccggcaagg gcctggagtg ggtgtccnnn atctccnnnn nnggaggcnn nacctactat 180
nnngactccn nnaagggccg gttcaccatc tcccgggaca actccaagaa caccctgtac 240
ctgcagatga actccctgcg ggccgaagac accgccgtgt actattgcgc caagcacggc 300
gcccccatga ccactgtgat cacctacgcc ccctattact tctactactg gggccagggc 360
accacggtca ccgtctcctc a 381
<210> SEQ ID NO 62
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (31)..(31)
<223> OTHER INFORMATION: Asn or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (50)..(50)
<223> OTHER INFORMATION: Thr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (53)..(54)
<223> OTHER INFORMATION: Ser or Gly
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (57)..(57)
<223> OTHER INFORMATION: Tyr or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (61)..(61)
<223> OTHER INFORMATION: Pro or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (64)..(64)
<223> OTHER INFORMATION: Leu or Val
<400> SEQUENCE: 62
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Xaa Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Xaa Ile Ser Xaa Xaa Gly Gly Xaa Thr Tyr Tyr Xaa Asp Ser Xaa
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Tyr Tyr Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 63
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 63
gagatcgtgc tgacccagtc ccccgccacc ctgtccctgt ccctgggcga gcgggccacc 60
ctgtcctgcc gggcctccag ctccgtgaag tacatgtact ggtaccagca gaagtccggc 120
caggcccccc ggctgctgat ctactacacc tccaacctgg cccccggcat ccccgcccgg 180
ttctccggct ccggctccgg caccgacttc accctgacca tctccagcat ggaggccgag 240
gacttcgccg tgtactactg ccagcagttc accacctccc cctacacctt cggcggaggc 300
accaagctcg agatcaaacg t 321
<210> SEQ ID NO 64
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 64
gagatcgtgc tgacccagtc ccccgccacc ctgtccctgt cccccggcga gcgggccacc 60
ctgtcctgcc gggcctccag ctccgtgaag tacatgtact ggtaccagca gaagcccggc 120
caggcccccc ggctgctgat ctactacacc tccaacctgg cccccggcat ccccgcccgg 180
ttctccggct ccggctccgg caccgacttc accctgacca tctcctccct ggagcccgag 240
gacttcgccg tgtactactg ccagcagttc accacctccc cctacacctt cggcggaggc 300
accaagctcg agatcaagcg g 321
<210> SEQ ID NO 65
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 65
gagatcgtgc tgacccagtc ccccgccacc ctgtccctgt cccccggcga gcgggccacc 60
ctgtcctgcc gggcctccag ctccgtgaag tacatgtact ggtaccagca gaagcccggc 120
caggcccccc ggctgctgat ctactacacc tccaacctgg cccccggcat ccccgcccgg 180
ttctccggct ccggctccgg caccgacttc accctgacca tctcctccct ggagcccgag 240
gacttcgccg tgtactactg ctaccagttc accacctccc cctacacctt cggcggaggc 300
accaagctcg agatcaagcg g 321
<210> SEQ ID NO 66
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 66
gagatcgtgc tgacccagtc ccccgccacc ctgtccctgt cccccggcga gcgggccacc 60
ctgtcctgcc gggcctccag ctccgtgaag tacatgtact ggtaccagca gaagcccggc 120
caggcccccc ggctgctgat ctactacacc tccaacctgg cccccggcat ccccgcccgg 180
ttctccggct ccggctccgg caccgacttc accctgacca tctcctccct ggagcccgag 240
gacttcgccg tgtactactg ccagcagtac accacctccc cctacacctt cggcggaggc 300
accaagctcg agatcaagcg g 321
<210> SEQ ID NO 67
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 67
gagatcgtgc tgacccagtc ccccgccacc ctgtccctgt cccccggcga gcgggccacc 60
ctgtcctgcc gggcctccag ctccgtgaag tacatgtact ggtaccagca gaagcccggc 120
caggcccccc ggctgctgat ctactacacc tccaacctgg cccccggcat ccccgcccgg 180
ttctccggct ccggctccgg caccgacttc accctgacca tctcctccct ggagcccgag 240
gacttcgccg tgtactactg ccagcagttc accacctacc cctacacctt cggcggaggc 300
accaagctcg agatcaagcg g 321
<210> SEQ ID NO 68
<211> LENGTH: 324
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (79)..(81)
<223> OTHER INFORMATION: This region may encompass 'AGC' or 'CAG'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (88)..(90)
<223> OTHER INFORMATION: This region may encompass 'AGG' or 'TCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (97)..(99)
<223> OTHER INFORMATION: This region may encompass 'ATG' or 'CTG'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (100)..(102)
<223> OTHER INFORMATION: This region may encompass 'TAC' or 'GCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (148)..(150)
<223> OTHER INFORMATION: This region may encompass 'TAC' or 'GAC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (151)..(153)
<223> OTHER INFORMATION: This region may encompass 'ACC' or 'GCC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (160)..(162)
<223> OTHER INFORMATION: This region may encompass 'CTG' or 'CGG'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (166)..(168)
<223> OTHER INFORMATION: This region may encompass 'CCC' or 'ACC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (265)..(267)
<223> OTHER INFORMATION: This region may encompass 'CAG' or 'TAC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (271)..(273)
<223> OTHER INFORMATION: This region may encompass 'TTC' or 'TAC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (280)..(282)
<223> OTHER INFORMATION: This region may encompass 'TCC' or 'TAC'
<400> SEQUENCE: 68
gagatcgtgc tgacccagtc ccccgccacc ctgtccctgt cccccggcga gcgggccacc 60
ctgtcctgcc gggcctccnn ntccgtgnnn tcctacnnnn nntggtacca gcagaagccc 120
ggccaggccc cccggctgct gatctacnnn nnntccaacn nngccnnngg catccccgcc 180
cggttctccg gctccggctc cggcaccgac ttcaccctga ccatctcctc cctggagccc 240
gaggacttcg ccgtgtacta ctgcnnncag nnnaccaccn nnccctacac cttcggcgga 300
ggcaccaagc tcgagatcaa gcgg 324
<210> SEQ ID NO 69
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<400> SEQUENCE: 69
gagaacgtgc tgacccagtc ccccgccttc ctgtccgtga cccccggcga gaaggtgacc 60
atcacctgcc gggcctccag ctccgtgaag tacatgtact ggtaccagca gaagcccgac 120
caggccccca agctgtggat ctattacacc tccaacctgg cccccggcgt gccctcccgg 180
ttctccggct ccggctccgg caacgactac accttcacca tctccagcct ggaggccgag 240
gacgccgcca cctattactg ccagcagttc accacctcac cctacacctt cggaggcggg 300
accaagctcg agatcaaacg t 321
<210> SEQ ID NO 70
<211> LENGTH: 321
<212> TYPE: DNA
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polynucleotide
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (1)..(3)
<223> OTHER INFORMATION: This region may encompass 'GAG' or 'GAC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (4)..(6)
<223> OTHER INFORMATION: This region may encompass 'AAC' or 'GTG'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (10)..(12)
<223> OTHER INFORMATION: This region may encompass 'CTG' or 'ATG'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (136)..(138)
<223> OTHER INFORMATION: This region may encompass 'TGG' or 'CTG'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (142)..(144)
<223> OTHER INFORMATION: This region may encompass 'TAT' or 'AAG'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (202)..(204)
<223> OTHER INFORMATION: This region may encompass 'AAC' or 'ACC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (208)..(210)
<223> OTHER INFORMATION: This region may encompass 'TAC' or 'TTC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (262)..(264)
<223> OTHER INFORMATION: This region may encompass 'CAG' or 'TAC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (268)..(270)
<223> OTHER INFORMATION: This region may encompass 'TTC' or 'TAC'
<220> FEATURE:
<221> NAME/KEY: modified_base
<222> LOCATION: (277)..(279)
<223> OTHER INFORMATION: This region may encompass 'TCC' or 'TAC'
<400> SEQUENCE: 70
nnnnnngtgn nnacccagtc ccccgccttc ctgtccgtga cccccggcga gaaggtgacc 60
atcacctgcc gggcctccag ctccgtgaag tacatgtact ggtaccagca gaagcccgac 120
caggccccca agctgnnnat cnnntacacc tccaacctgg cccccggcgt gccctcccgg 180
ttctccggct ccggctccgg cnnngacnnn accttcacca tctccagcct ggaggccgag 240
gacgccgcca cctattactg cnnncagnnn accaccnnnc cctacacctt cggcggaggc 300
accaagctcg agatcaagcg g 321
<210> SEQ ID NO 71
<211> LENGTH: 120
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 71
Glu Val Met Leu Val Glu Ser Gly Gly Gly Val Val Lys Pro Gly Gly
1 5 10 15
Ser Leu Lys Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Ile Arg Gln Thr Pro Glu Lys Arg Leu Glu Trp Val
35 40 45
Ala Thr Ile Ser Asn Gly Gly Ser Tyr Thr Tyr Tyr Pro Asp Ser Val
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Val Lys Asn Ile Leu Tyr
65 70 75 80
Leu Gln Met Arg Ser Leu Arg Ser Glu Asp Thr Ala Met Tyr Tyr Cys
85 90 95
Ala Arg Arg Ser Ser Asp Tyr Asp Ser Trp Phe Ala Tyr Trp Gly Gln
100 105 110
Gly Thr Leu Val Thr Val Ser Ser
115 120
<210> SEQ ID NO 72
<211> LENGTH: 120
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 72
Glu Val Met Leu Val Glu Ser Gly Gly Gly Phe Val Lys Pro Gly Gly
1 5 10 15
Ser Leu Lys Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Phe
20 25 30
Ala Met Ser Trp Val Arg Gln Thr Pro Glu Lys Arg Leu Glu Trp Val
35 40 45
Ala Thr Ile Ser Asn Gly Gly Thr Tyr Thr Tyr Tyr Pro Asp Ser Val
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Ser Ser Leu Arg Ser Glu Asp Thr Ala Met Tyr Tyr Cys
85 90 95
Ala Arg Arg Ser Ser Asp Tyr Asp Ser Trp Phe Ala Tyr Trp Gly Gln
100 105 110
Gly Thr Leu Val Thr Val Ser Ser
115 120
<210> SEQ ID NO 73
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 73
Glu Val Met Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Lys Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Thr Pro Glu Lys Arg Leu Glu Trp Val
35 40 45
Ala Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Thr Leu Phe
65 70 75 80
Leu Gln Met Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Val Arg His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Asp Tyr Trp Gly Gln Gly Thr Thr Leu Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 74
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 74
Glu Val Met Leu Val Glu Ser Gly Gly Gly Val Val Lys Pro Gly Gly
1 5 10 15
Ser Leu Lys Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Thr Pro Glu Lys Arg Leu Glu Trp Val
35 40 45
Ala Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ile Leu Phe
65 70 75 80
Leu Gln Met Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Val Arg His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Asp Tyr Trp Gly Gln Gly Thr Thr Leu Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 75
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 75
Gln Ile Val Leu Thr Gln Ser Pro Ala Ile Met Ser Ala Ser Pro Gly
1 5 10 15
Glu Lys Val Thr Ile Ser Cys Ser Ala Thr Ser Ser Val Ile Phe Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Gly Ser Ser Pro Lys Pro Trp Ile Tyr
35 40 45
Arg Thr Ser Asn Leu Ala Ser Gly Val Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Thr Ser Tyr Ser Leu Thr Ile Ser Asn Met Glu Ala Glu
65 70 75 80
Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Tyr Gln Ser Tyr Pro Leu Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Leu Lys Arg
100 105
<210> SEQ ID NO 76
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 76
Gln Ile Val Leu Thr Gln Ser Pro Ala Ile Met Ser Ala Ser Pro Gly
1 5 10 15
Glu Arg Val Thr Ile Ser Cys Ser Ala Ser Ser Ser Val Ile Phe Leu
20 25 30
Tyr Trp Tyr Gln Gln Lys Pro Gly Ser Ser Pro Lys Pro Arg Ile Tyr
35 40 45
Arg Thr Ser Asn Leu Ala Ser Gly Val Pro Asp Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Thr Ser Tyr Ser Leu Thr Ile Ser Ser Met Glu Ala Glu
65 70 75 80
Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Tyr His Ser Tyr Pro Leu Thr
85 90 95
Phe Gly Ala Gly Thr Lys Leu Glu Leu Lys Arg
100 105
<210> SEQ ID NO 77
<211> LENGTH: 126
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 77
Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Ala Ile Ser Gly Ser Gly Gly Ser Ile Tyr Tyr Ala Asp Ser Val
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys Asp Val Leu Val Val Pro Ala Ala Pro Tyr Tyr Tyr Tyr Gly
100 105 110
Met Asp Val Trp Gly Gln Gly Thr Thr Val Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 78
<211> LENGTH: 98
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
peptide
<400> SEQUENCE: 78
Glu Val Gln Leu Leu Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Ala Ile Ser Gly Ser Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys
<210> SEQ ID NO 79
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (99)..(108)
<223> OTHER INFORMATION: Any amino acid
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (110)..(111)
<223> OTHER INFORMATION: Any amino acid
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (113)..(114)
<223> OTHER INFORMATION: Any amino acid
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (116)..(116)
<223> OTHER INFORMATION: Any amino acid
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (123)..(123)
<223> OTHER INFORMATION: Any amino acid
<400> SEQUENCE: 79
Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly
1 5 10 15
Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45
Ser Ala Ile Ser Gly Ser Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr
65 70 75 80
Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Ala Lys Xaa Xaa Xaa Xaa Xaa Xaa Xaa Xaa Xaa Xaa Tyr Xaa Xaa Tyr
100 105 110
Xaa Xaa Asp Xaa Trp Gly Gln Gly Thr Thr Xaa Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 80
<211> LENGTH: 127
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (3)..(3)
<223> OTHER INFORMATION: Met or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (5)..(5)
<223> OTHER INFORMATION: Val or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (13)..(13)
<223> OTHER INFORMATION: Lys or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (19)..(19)
<223> OTHER INFORMATION: Lys or Arg
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (40)..(40)
<223> OTHER INFORMATION: Thr or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (42)..(42)
<223> OTHER INFORMATION: Glu or Gly
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (44)..(44)
<223> OTHER INFORMATION: Arg or Gly
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (49)..(49)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (75)..(75)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (80)..(80)
<223> OTHER INFORMATION: Phe or Tyr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (84)..(84)
<223> OTHER INFORMATION: Ser or Asn
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (88)..(88)
<223> OTHER INFORMATION: Ser or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (97)..(97)
<223> OTHER INFORMATION: Val or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (98)..(98)
<223> OTHER INFORMATION: Arg or Lys
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (123)..(123)
<223> OTHER INFORMATION: Leu or Val
<400> SEQUENCE: 80
Glu Val Xaa Leu Xaa Glu Ser Gly Gly Gly Leu Val Xaa Pro Gly Gly
1 5 10 15
Ser Leu Xaa Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30
Ala Met Ser Trp Val Arg Gln Xaa Pro Xaa Lys Xaa Leu Glu Trp Val
35 40 45
Xaa Thr Ile Ser Ser Gly Gly Gly Tyr Thr Tyr Tyr Pro Asp Ser Leu
50 55 60
Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Xaa Lys Asn Thr Leu Xaa
65 70 75 80
Leu Gln Met Xaa Ser Leu Arg Xaa Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95
Xaa Xaa His Gly Ala Pro Met Thr Thr Val Ile Thr Tyr Ala Pro Tyr
100 105 110
Tyr Phe Asp Tyr Trp Gly Gln Gly Thr Thr Xaa Thr Val Ser Ser
115 120 125
<210> SEQ ID NO 81
<211> LENGTH: 106
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 81
Glu Asn Val Leu Thr Gln Ser Pro Ala Ile Met Ser Ala Ser Leu Gly
1 5 10 15
Glu Lys Val Thr Met Asn Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Ser Asp Ala Ser Pro Lys Leu Trp Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Val Pro Ala Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Asn Ser Tyr Ser Leu Thr Ile Ser Ser Met Glu Gly Glu
65 70 75 80
Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys
100 105
<210> SEQ ID NO 82
<211> LENGTH: 108
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<400> SEQUENCE: 82
Asp Ile Gln Leu Thr Gln Ser Pro Ala Ile Met Ser Ala Ser Pro Gly
1 5 10 15
Glu Lys Val Thr Met Thr Cys Arg Ala Ser Ser Ser Val Ser Ser Ser
20 25 30
Tyr Leu His Trp Tyr Gln Gln Lys Ser Gly Ala Ser Pro Lys Leu Trp
35 40 45
Ile Tyr Ser Thr Ser Asn Leu Ala Ser Gly Val Pro Ala Arg Phe Ser
50 55 60
Gly Ser Gly Ser Gly Thr Ser Tyr Ser Leu Thr Ile Ser Ser Val Glu
65 70 75 80
Ala Glu Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Tyr Ser Gly Tyr Pro
85 90 95
Tyr Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys
100 105
<210> SEQ ID NO 83
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Any amino acid
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (30)..(31)
<223> OTHER INFORMATION: Any amino acid
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (78)..(78)
<223> OTHER INFORMATION: Any amino acid
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (91)..(94)
<223> OTHER INFORMATION: Any amino acid
<400> SEQUENCE: 83
Asp Xaa Val Leu Thr Gln Ser Pro Ala Ile Met Ser Ala Ser Pro Gly
1 5 10 15
Glu Lys Val Thr Met Thr Cys Arg Ala Ser Ser Ser Val Xaa Xaa Tyr
20 25 30
Leu Tyr Trp Tyr Gln Gln Lys Ser Asp Ala Ser Pro Lys Leu Trp Ile
35 40 45
Tyr Tyr Thr Ser Asn Leu Ala Ser Gly Val Pro Ala Arg Phe Ser Gly
50 55 60
Ser Gly Ser Gly Thr Ser Tyr Ser Leu Thr Ile Ser Ser Xaa Glu Ala
65 70 75 80
Glu Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Xaa Xaa Xaa Xaa Pro Tyr
85 90 95
Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys
100 105
<210> SEQ ID NO 84
<211> LENGTH: 107
<212> TYPE: PRT
<213> ORGANISM: Artificial Sequence
<220> FEATURE:
<223> OTHER INFORMATION: Description of Artificial Sequence:
Synthetic
polypeptide
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (1)..(1)
<223> OTHER INFORMATION: Glu or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (2)..(2)
<223> OTHER INFORMATION: Asn, Val or Ile
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (4)..(4)
<223> OTHER INFORMATION: Leu or Met
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (10)..(10)
<223> OTHER INFORMATION: Ile or Phe
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (11)..(11)
<223> OTHER INFORMATION: Met or Leu
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (13)..(13)
<223> OTHER INFORMATION: Ala or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (14)..(14)
<223> OTHER INFORMATION: Ser or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (15)..(15)
<223> OTHER INFORMATION: Leu or Pro
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (21)..(21)
<223> OTHER INFORMATION: Met or Ile
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (22)..(22)
<223> OTHER INFORMATION: Asn or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (39)..(39)
<223> OTHER INFORMATION: Ser or Pro
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (40)..(40)
<223> OTHER INFORMATION: Asp or Gly
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (41)..(41)
<223> OTHER INFORMATION: Ala or Gln
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (42)..(42)
<223> OTHER INFORMATION: Ser or Ala
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (59)..(59)
<223> OTHER INFORMATION: Ala or Ser
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (69)..(69)
<223> OTHER INFORMATION: Ser or Asp
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (71)..(71)
<223> OTHER INFORMATION: Ser or Thr
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (72)..(72)
<223> OTHER INFORMATION: Leu or Phe
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (77)..(77)
<223> OTHER INFORMATION: Met, Leu or Val
<220> FEATURE:
<221> NAME/KEY: MOD_RES
<222> LOCATION: (79)..(79)
<223> OTHER INFORMATION: Gly or Ala
<400> SEQUENCE: 84
Xaa Xaa Val Xaa Thr Gln Ser Pro Ala Xaa Xaa Ser Xaa Xaa Xaa Gly
1 5 10 15
Glu Lys Val Thr Xaa Xaa Cys Arg Ala Ser Ser Ser Val Lys Tyr Met
20 25 30
Tyr Trp Tyr Gln Gln Lys Xaa Xaa Xaa Xaa Pro Lys Leu Trp Ile Tyr
35 40 45
Tyr Thr Ser Asn Leu Ala Pro Gly Val Pro Xaa Arg Phe Ser Gly Ser
50 55 60
Gly Ser Gly Asn Xaa Tyr Xaa Xaa Thr Ile Ser Ser Xaa Glu Xaa Glu
65 70 75 80
Asp Ala Ala Thr Tyr Tyr Cys Gln Gln Phe Thr Thr Ser Pro Tyr Thr
85 90 95
Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg
100 105
User Contributions:
Comment about this patent or add new information about this topic:
| People who visited this patent also read: | |
| Patent application number | Title |
|---|---|
| 20150115408 | ENHANCED HYDROGEN BARRIER ENCAPSULATION METHOD FOR THE CONTROL OF HYDROGEN INDUCED DEGRADATION OF FERROELECTRIC CAPACITORS IN AN F-RAM PROCESS |
| 20150115407 | Isolation Device |
| 20150115406 | SEMICONDUCTOR STRUCTURE |
| 20150115405 | WIRELESS INTERCONNECTS IN AN INTERPOSER |
| 20150115404 | INTERCONNECTION BETWEEN INDUCTOR AND METAL-INSULATOR-METAL (MIM) CAPACITOR |