Patent application title: REPLIKIN PEPTIDES AND ANTIBODIES THEREFORE
Inventors:
Samuel Bogoch (New York, NY, US)
Elenore S. Bogoch (New York, NY, US)
IPC8 Class: AC07K1400FI
USPC Class:
530324
Class name: Chemistry: natural resins or derivatives; peptides or proteins; lignins or reaction products thereof peptides of 3 to 100 amino acid residues 25 or more amino acid residues in defined sequence
Publication date: 2009-05-28
Patent application number: 20090137778
Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
Patent application title: REPLIKIN PEPTIDES AND ANTIBODIES THEREFORE
Inventors:
Samuel BOGOCH
Elenore S. BOGOCH
Agents:
KENYON & KENYON LLP
Assignees:
Origin: WASHINGTON, DC US
IPC8 Class: AC07K1400FI
USPC Class:
530324
Abstract:
The present invention provides a new class of peptides related to rapid
replication and their use in diagnosing, preventing and treating disease.Claims:
1-173. (canceled)
174. A peptide consisting of 23 to 50 amino acid residues comprising any one of the sequences disclosed in the variable amino acid sequence of SEQ ID NO: 698.
175. The peptide of claim 174 wherein said peptide is an HIV peptide.
176. The peptide of claim 175 wherein said HIV peptide is a trans-activator peptide.
177. The peptide of claim 174 consisting of 23 amino acid residues.
178. A peptide consisting of 19 to 50 amino acid residues comprising any one of the sequences beginning at position 5 and ending at position 23 of the variable amino acid sequence of SEQ ID NO: 698.
179. The peptide of claim 178 wherein said peptide is an HIV peptide.
180. The peptide of claim 179 wherein said HIV peptide is a trans-activator peptide.
181. The peptide of claim 178 consisting of 19 amino acid residues.
Description:
CROSS REFERENCE TO OTHER APPLICATIONS
[0001]This application is a Continuation-In-Part of application No. U.S. Ser. No. 10/105,232, filed Mar. 26, 2002, which is a Continuation-In-Part of U.S. Ser. No. 09/984,057, filed Oct. 26, 2001, which claims priority from Provisional Applications 60/303,396, filed Jul. 9, 2001 and 60/278,761 filed Mar. 27, 2001, the subject matter of which is incorporated herein by reference.
FIELD OF THE INVENTION
[0002]This invention relates to the identification and use of Replikins, a newly discovered class of peptides that share structural characteristics. In particular, this invention relates to Replikins which have been found in viruses, bacteria, fungus, cancer associated proteins, plants and unicellular parasites and their use as targets in the development of methods of treating or preventing diseases. Further, this invention relates to the use of Replikins in the detection of these diseases. Also this invention relates to the use of Replikins to stimulate growth of plants used for food.
INTRODUCTION AND BACKGROUND OF THE INVENTION
[0003]Rapid replication is characteristic of virulence in certain bacteria, viruses and malignancies, but no chemistry common to rapid replication in different organisms has been described previously. This patent application discloses a new class of protein structures related to rapid replication. A new family of conserved small proteins related to rapid replication, named Replikins, which are used to predict and control rapid replication in multiple organisms and diseases and to induce rapid replication in plant and animal life.
[0004]We constructed an algorithm search for Replikins. In applying the algorithm invented herein not only was the function of the epitope revealed--rapid replication, but an entire family of homologues whose function is related to rapid replication was discovered, which we named Replikins.
[0005]The algorithm is based on the following: 1) Evidence that the immune system looks to parts rather than a whole protein in recognition. Protein chains are first hydrolyzed by the immune system into smaller pieces, frequently six (6) to ten (10) amino acids long, as part of the immune systems' process of recognition of foreign structures against which it may mount an immune defense. By way of example, the immune system recognizes the presence of disease by chopping up proteins of the disease agent into smaller peptide sequences and reading them. This principle is used as a basis for the algorithm with which to search for homologues of the malignin cancer epitope, once the structure of the epitope was known; 2) The specific structure of the malignin epitope, in which two of the three lysines (K's) are eight residues apart is in accordance with the apparent `rules` used by the immune system for recognition referred to above (6-10 amino acids long); 3) The fact that the malignin cancer epitope was shown to be a very strong antigen, that is--a generator of a strong immune response; that there are three lysines (K's) in the 10-mer peptide glioma Replikin and that K's are known to bind frequently to DNA and RNA as potential anchors for the entry of viruses; and 4) One histidine (H) is included in the sequence of the malignin epitope, between the two K's which are eight (8) residues apart, suggesting a connection to the metals of redox systems which are required to provide the energy for replication.
[0006]Engineered enzymes and catalytic antibodies, possessing tailored binding pockets with appropriately positioned functional groups have been successful in catalyzing a number of chemical transformations, sometimes with impressive efficiencies. Just as two or more separate proteins with specific and quite different functions are now often recognized to be synthesized together by organisms, and then separately cleaved to `go about their separate functions`, so the Replikin structure is a unique protein with a unique function that appears to be recognized separately by the immune system and may be now rationally engineered--e.g. synthesized to produce a functional unit.
[0007]From a proteomic point of view, this template based on the newly determined glioma peptide sequence has led to the discovery of a wide class of proteins with related conserved structures and a particular function, in this case replication. Examples of the increase in Replikin concentration with virulence of a disease appear in diseases including, influenza, HIV, cancer and tomato leaf curl virus. This class of structures is related to the phenomenon of rapid replication in organisms as diverse as yeast, algae, plants, the gemini curl leaf tomato virus, HIV and cancer.
[0008]In addition to detecting the presence of Replikins in rapidly replicating organisms, we found that 1) Replikin concentration (number of Replikins per 100 amino acids) and 2) Replikin compositions in specific functional states dependant on rapid replication, provide the basis for the finding that Replikins are related quantitatively as well as qualitatively to the rate of replication of the organism in which they reside. Examples of these functional proofs include the relationship found between rapid replication and virulence in glioblastoma cells, between Replikins in influenza virus and the prediction of influenza pandemics and epidemics, and the relationship between Replikin concentration and rapid replication in HIV.
[0009]The first functional basis for Replikins' role in rapid replication was found in the properties of the glioma Replikin, a 10 KD peptide called Malignin in brain glioblastoma multiforme (glioma)--a 250 KD cell protein. Antimalignin antibody increased in concentration in serum (AMAS), measured by an early stage diagnostic test for cancer now used for most or all cell types. Malignin was so named because in tissue culture the expression of this peptide and its concentration per milligram membrane protein extractable increased with increased rate of cell division per unit time. Not only is there an increase in the amount of malignin in proportion to the cell number increase but the amount of malignin is enriched, that is--increased ten fold whereas the cell number increased only five fold.
[0010]The structure of malignin protein was determined through hydrolysis and mass spectrometry which revealed what proved to be a novel 16 mer peptide sequence. We searched for the 16 mer peptide sequence which we have named a Glioma Replikin protein in databases for the healthy human genome and found that it was not present in these databases.
[0011]As such, the fixed requirement algorithm was used to search in other organisms for the Glioma Replikin protein or homologues thereof. Over 4,000 protein sequences in the "Pub Med" database were searched and homologues were found in viruses and plant forms specifically associated with rapid replication. Homologues of such Replikin proteins occurred frequently in proteins called `replicating proteins` by their investigators.
[0012]Homologues of the Replikin sequence were found in all tumor viruses (that is viruses that cause cancer), and in `replicating proteins` of algae, plants, fungi, viruses and bacteria.
[0013]That malignin is enriched ten-fold compared to the five-fold increase in cell number and membrane protein concentration in rapid replication of glioma cells suggests an integral relationship of the Replikins to replication. When the glioma replikin was synthesized in vitro and administered as a synthetic vaccine to rabbits, abundant antimalignin antibody was produced--establishing rigourously the antigenic basis of the antimalignin antibody in serum (AMAS) test, and providing the first potential synthetic cancer vaccine and the prototype for Replikin vaccines in other organisms.
[0014]The demonstration of the relationship of the Replikins to replication and the natural immune response to cancer Replikins (overriding cell type) based upon the shared specificity of cancer Replikins, permits passive augmentation of immunity with antimalignin antibody and active augmentation with synthetic Replikin vaccines.
[0015]A study of 8,090 serum specimens from cancer patients and controls has demonstrated that the concentration of antimalignin antibody increases with age in healthy individuals, as the incidence of cancer in the population increases, and increases further two to three-fold in early malignancy, regardless of cell type. In vitro this antibody is cytotoxic to cancer cells at picograms (femtomoles) per cancer cell, and in vivo the concentration of antimalignin antibody relates quantitatively to the survival of cancer patients. As shown in glioma cells, the stage in cancer at which cells only have been transformed to the immortal malignant state but remain quiescent or dormant, now can be distinguished from the more active life-threatening replicating state which is characterized by the increased concentration of Replikins. In addition, clues to the viral pathogenesis of cancer may be found in the fact that glioma glycoprotein 10B has a 50% reduction in carbohydrate residues when compared to the normal 10B. This reduction is associated with virus entry in other instances and so may be evidence of the attachment of virus for the delivery of virus Replikins to the 10B of glial cells as a step in the transformation to the malignant state.
[0016]The sharing of immunological specificity by diverse members of the class, as demonstrated with antimalignin antibody for the glioma and related cancer Replikins, suggests that B cells and their product antibodies may recognize Replikins by means of a similar recognition `language`. With the discovery of the Replikins, this shared immunological specificity may explain what was previously difficult to understand: why the antimalignin antibody is elevated in all cancers, and is cytotoxic to cancer cells and related to survival of cancer patients in most or all cell types. Thus antimalignin antibody is produced against cancer Replikins, which share immunological specificity and which are related to the phenomenon of rapid replication, not to cell type.
[0017]A second functional basis for the Replikins' role in rapid replication is the study of data from the past 100 years on influenza virus hemagglutinin protein sequences and epidemiology of influenza epidemics and pandemics. To date, only serological hemagglutinin and antibody classification, but no strain-specific conserved peptide sequences have previously been described in influenza, and no changes in concentration and composition of any strain-specific peptide sequences have been described previously which correlate with epidemiologically documented epidemics or rapid replication.
[0018]A four to ten-fold increase in the concentration of strain-specific influenza Replikins in one of each of the four major strains, influenza B, (A)H1N1, (A)H2N2 and (A)H3N2 was found, and that such increase of Replikin concentration was related to influenza epidemics caused specifically by each strain from 1902 to 2001. These increases in concentration were then shown to be due to the reappearance of at least one specific Replikin composition from 1 to up to 64 years after its disappearance, plus the emergence of new strain-specific Replikin compositions. Previously, no strain-specific chemical structures were known with which to predict which strains would predominate in coming influenza seasons, nor to devise annual mixtures of whole-virus strains for vaccines. The recent sharp increase in H3N2 Replikin concentration (1997 to 2000), the largest in H3N2's history, and the reappearance of specific Replikin compositions which were last seen in the high mortality H3N2 pandemic of 1968 and in the two high mortality epidemics of 1975 and 1977, but were absent for 20-25 years, together may be a warning of coming epidemics.
[0019]Synthetic Replikins are new vaccines. This high degree of conservation of Replikin structures observed whereby the identical structure can persist for 100 years, or reappear after an absence of from one to 64 years reappears indicates that what was previously thought to be change in virulence due to random substitution of amino acids in influenza proteins is more likely to be change due to an organized process of conservation of Replikins. In fact, if random substitutions of each amino acid occurred, the chance against an average length influenza Replikin sequence being conserved for one year (let alone 100) is calculated to be in the order of 2 to the 27th power to 1.
[0020]The significant conservation of Replikins is not unique to influenza virus is also present in foot and mouth disease virus type O and in HIV, as well as in wheat.
[0021]A third functional basis for Replikins' role in rapid replication is the increase in Replikin concentration shown to be related to rapid replication in HIV. The Replikin concentration in the slow-growing low-titre strain of HIV (NS1, "Bru"), prevalent in early stage infection, was found to be one-sixth of the Replikin concentration in the rapidly-growing high-titre strain of HIV (SI, "Lai"), prevalent in late stage HIV infection.
[0022]Other examples are given of the relation of Replikins to rapid replication. For example, in tomato curl leaf gemini virus, which devastates tomato crops, the first 161 amino acids of the `replicating protein`, which have been shown to bind to DNA, contain five Replikins.
[0023]In malaria, legendary for rapid replication, trypanosomes are released from the liver in tens of thousands from one trypanosome. Multiple, novel, almost `flamboyant` Replikin structures with concentrations of up to 36 overlapping Replikins per 100 amino acids are found therein.
[0024]The increase in Replikin concentration in influenza epidemics is functionally comparable to the glioma Replikin's increase in concentration during rapid replication of malignant glioma cells and comparable to rapid replication in HIV and in a diverse range of other organisms. Replikins thus are associated with and appear to be part of the structural bases of rapid replication in different organisms.
[0025]Replikin concentration and composition therefore provide new methods to detect and to control the process of replication, which is central to the survival and dominance of each biological population. The discovery of these new proteins related to rapid replication provides new opportunities 1) for detection of pathogens by qualitative and quantitative determinations of Replikins, 2) for the control of a broad range of diseases in which rapid replication is a key factor by targeting native Replikins and by using synthetic Replikins as vaccines, and 3) for the use of Replikins to foster growth of algal and plant foods.
[0026]There is a significant number of diseases and pathogens which have proved difficult to detect and treat and for which there is no effective vaccine. Thus, for each disorder there is a need for developing a target that will provide effective methods of detecting, treating or preventing these diseases and pathogens.
SUMMARY OF THE INVENTION
[0027]The present invention provides a method for identifying nucleotide or amino acid sequences that include a Replikin sequence. The method is referred to herein as a 3-point-recognition method. By use of the "3-point recognition" method, namely, peptides comprising from 7 to about 50 amino acids including: [0028](1) at least one lysine residue located six to ten amino acid residues from a second lysine residue; [0029](2) at least one histidine residue; and [0030](3) at least 6% lysine residues (Replikin)--constituting a new class of peptides was revealed in algae, yeast, fungi, amoebae, bacteria, plant and virus proteins having replication, transformation, or redox functions.
[0031]In one aspect of the invention there are provided isolated or synthesized peptides containing a Replikin sequence. The peptides comprise from 7 to about 50 amino acids including: [0032](1) at least one lysine residue located six to ten amino acid residues from a second lysine residues; [0033](2) at least one histidine residue; and [0034](3) at least 6% lysine residues.
[0035]The present invention also provides methods for detecting the presence of a contaminating organism in a body sample or environmental sample comprising: [0036](1) isolating nucleic acids from the body sample or environmental sample; [0037](2) screening the nucleic acids for the presence of a Replikin structure; and [0038](3) correlating the presence of a Replikin structure with the presence of the contaminating organism.
[0039]In another aspect of the invention there is provided a process for stimulating the immune system of a subject to produce antibodies that bind specifically to a Replikin sequence, said process comprising administering to the subject an effective amount of a dosage of a composition comprising at least one Replikin peptide. One embodiment comprises at least one peptide that is present in an emerging strain of the organism if such new strain emerges.
[0040]The present invention also provides antibodies that bind specifically to a Replikin, as defined herein, as well as antibody cocktails containing a plurality of antibodies that specifically bind to Replikins. In one embodiment of the invention, there are provided compositions comprising an antibody or antibodies that specifically bind to a Replikin and a pharmaceutically acceptable carrier.
[0041]In one aspect of the invention there are provided isolated, or separated from other proteins, recombinant, or synthesized peptides or other methods containing a viral Replikin sequence. The viral Replikin peptides comprise from 7 to about 50 amino acids including: [0042](1) at least one lysine residue located six to ten amino acid residues from a second lysine residue; [0043](2) at least one histidine residue; and [0044](3) at least 6% lysine residues. (viral Replikin).
[0045]The present invention also provides methods for detecting the presence of a contaminating virus in a body sample or environmental sample comprising: [0046](1) isolating nucleic acids from the body sample or environmental sample; [0047](2) screening the nucleic acids for the presence of a viral Replikin structure; and [0048](3) correlating the presence of viral Replikin structures, their concentration and composition, with the presence of the contaminating virus.
[0049]In another aspect of the invention there is provided a process for stimulating the immune system of a subject to produce antibodies that bind specifically to a viral Replikin sequence, said process comprising administering to the subject an effective amount of a dosage of a composition comprising at least one Replikin peptide. One embodiment comprises at least one peptide that is present in an emerging strain of the virus if such new strain emerges.
[0050]The present invention also provides antibodies that bind specifically to a viral Replikin, as defined herein, as well as antibody cocktails containing a plurality of antibodies that specifically bind to viral Replikins. In one embodiment of the invention, there are provided compositions comprising an antibody or antibodies that specifically bind to a viral Replikin and a pharmaceutically acceptable carrier.
[0051]The present invention also provides therapeutic compositions comprising one or more of isolated virus peptides having from 7 to about 50 amino acids comprising: [0052](1) at least one lysine residue located six to ten residues from a second lysine residue; [0053](2) at least one histidine residue; and [0054](3) at least 6% lysine residues, and a pharmaceutically acceptable carrier.
[0055]In another aspect of the invention there is provided an antisense nucleic acid molecule complementary to a virus Replikin mRNA sequence, said Replikin mRNA sequence denoting from 7 to about 50 amino acids comprising: [0056](1) at least one lysine residue located six to ten residues from a second lysine residue; [0057](2) at least one histidine residue; and [0058](3) at least 6% lysine residues.
[0059]In yet another aspect of the invention there is provided a method of simulating the immune system of a subject to produce antibodies to viruses, said method comprising: administering an effective amount of at least one virus Replikin peptide having from 7 to about 50 amino acids comprising: [0060](1) at least one lysine residue located six to ten amino acid residues from a second lysine residue; [0061](2) at least one histidine residue; and [0062](3) at least 6% lysine residues.
[0063]In another aspect, there is provided a method of selecting a virus peptide for inclusion in a preventive or therapeutic virus vaccine comprising: [0064](1) obtaining at least one isolate of each strain of a plurality of strains of said virus; [0065](2) analyzing the amino acid sequence of the at least one isolate of each strain of the plurality of strains of the virus for the presence and concentration of Replikin sequences; [0066](3) comparing the concentration of Replikin sequences in the amino acid sequence of the at least one isolate of each strain of the plurality of strains of the virus to the concentration of Replikin sequences observed in the amino acid sequence of each of the strains at least one earlier time period to provide the concentration of Replikins for at least two time periods, said at least one earlier time period being within about six months to about three years prior to step (1); [0067](4) identifying the strain of the virus having the highest increase in concentration of Replikin sequences during the at least two time periods; and [0068](5) selecting at least one Replikin sequence present in the strain of the virus peptide identified in step (4) as a peptide for inclusion in the virus vaccine.
[0069]The present invention also provides a method of making a preventive or therapeutic virus vaccine comprising: [0070](1) identifying a strain of a virus as an emerging strain, [0071](2) selecting at least one Replikin sequence present in the emerging strain as a peptide template for the virus vaccine manufacture, [0072](3) synthesizing peptides having the amino acid sequence of the at least one Replikin sequence selected in step (2), and [0073](4) combining a therapeutically effective amount of the peptides of step (3) with a pharmaceutically acceptable carrier and/or adjuvant.
[0074]In another aspect, the invention is directed to a method of identifying an emerging strain of a virus for diagnostic, preventive or therapeutic purposes comprising: [0075](1) obtaining at least one isolate of each strain of a plurality of strains of the virus; [0076](2) analyzing the amino acid sequence of the at least one isolate of each strain of the plurality of strains of the virus for the presence and concentration of Replikin sequences; [0077](3) comparing the concentration of Replikin sequences in the amino acid sequence of the at least one isolate of each strain of the plurality of strains of the virus to the concentration of Replikin sequences observed in the amino acid sequence of each of the strains at least one earlier time period to provide the concentration of Replikins for at least two time periods, said at least one earlier time period being within about six months to about three years prior to step (1); and [0078](4) identifying the strain of the virus having the highest increase in concentration of Replikin sequences during the at least two time periods.
[0079]In yet another aspect of the invention, there is provided a preventive or therapeutic virus vaccine comprising at least one isolated Replikin present in a protein of an emerging strain of the virus and a pharmaceutically acceptable carrier and/or adjuvant.
[0080]Also provided by the present invention is a method of preventing or treating a virus infection comprising administering to a patient in need thereof a preventive or therapeutic virus vaccine comprising at least one isolated Replikin present in a protein of an emerging strain of the virus and a pharmaceutically acceptable carrier and/or adjuvant.
Influenza
[0081]Influenza is an acute respiratory illness of global importance. Despite international attempts to control influenza virus outbreaks through vaccination, influenza infections remain an important cause of morbidity and mortality. Worldwide influenza epidemics and pandemics have occurred at irregular and previously unpredictable intervals throughout history and it is expected that they will continue to occur in the future. The impact of both pandemic and epidemic influenza is substantial in terms of morbidity, mortality and economic cost.
[0082]Influenza vaccines remain the most effective defense against influenza virus, but because of the ability of the virus to mutate and the availability of non-human host reservoirs, it is expected that influenza will remain an emergent or re-emergent infection. Global influenza surveillance indicates that influenza viruses may vary within a country and between countries and continents during an influenza season. Virological surveillance is of importance in monitoring antigenic shift and drift. Disease surveillance is also important in assessing the impact of epidemics. Both types of information have provided the basis of the vaccine composition and the correct use of antivirals. However, to date there has been only annual post hoc hematological classification of the increasing number of emerging influenza virus strains, and no specific chemical structure of the viruses has been identified as an indicator of approaching influenza epidemics or pandemics. Currently, the only basis for annual classification of influenza virus as active, inactive or prevalent in a given year is the activities of the virus hemagglutinin and neuraminidase proteins. No influenza viral chemical structure has been identified prior to this application that can be used for quantitative warning of epidemics or pandemics or to design more effective and safer vaccines.
[0083]Because of the annual administration of influenza vaccines and the short period of time when a vaccine can be administered, strategies directed at improving vaccine coverage are of critical importance.
[0084]In one aspect of the invention there are provided isolated or synthesized influenza virus peptides containing a Replikin sequence. The influenza Replikin virus peptides comprise from 7 to about 50 amino acids including: [0085](1) at least one lysine residue located six to ten amino acid residues from a second lysine residue; [0086](2) at least one histidine residue; and [0087](3) at least 6% lysine residues. (Influenza Replikin).
[0088]In another aspect of the invention, there is provided a process for stimulating the immune system of a subject to produce antibodies that bind specifically to an influenza virus Replikin sequence, said process comprising administering to the subject an effective amount of dosage of a composition comprising at least one influenza virus Replikin peptide. In a preferred embodiment the composition comprises at least on peptide that is present in an emerging strain of influenza virus.
[0089]The present invention also provides antibodies that bind specifically to an influenza virus Replikin, as defined herein, as well as antibody cocktails containing a plurality of antibodies that specifically bind to influenza virus Replikins. In one embodiment of the invention, there are provided compositions comprising an antibody or antibodies that specifically bind to an influenza Replikin and a pharmaceutically acceptable carrier.
[0090]The present invention also provides therapeutic compositions comprising one or more of isolated influenza virus peptides having from 7 to about 50 amino acids comprising: [0091](1) at least one lysine residue located six to ten residues form a second lysinc residue; [0092](2) at least one histidine residue; and [0093](3) at least 6% lysine residues, and a pharmaceutical acceptable carrier.
[0094]In another aspect of the invention there is provided an antisense nucleic acid molecule complementary to an influenza virus hemagglutinin Replikin mRNA sequence, said Replikin mRNA sequence denoting from 7 to about 50 amino acids comprising: [0095](1) at least one lysine residue located six to ten residues from a second lysine residue; [0096](2) at least one histidine residue; and [0097](3) at least 6% lysine residues.
[0098]In yet another aspect of the invention there is provided a method of simulating the immune system of a subject to produce antibodies to influenza virus comprising administering an effective amount of at least one influenza virus Replikin peptide having from 7 to about 50 amino acids comprising: [0099](1) at least one lysine residue located six to ten amino acid residues from a second lysine residue; [0100](2) at least one histidine residue; and [0101](3) at least 6% lysine residues.
[0102]In another aspect, there is provided a method of selecting an influenza virus peptide for inclusion in a preventive or therapeutic influenza virus vaccine comprising: [0103](1) obtaining at least one isolate of each strain of a plurality of strains of influenza virus; [0104](2) analyzing the hemagglutinin amino acid sequence of the at least one isolate of each strain of the plurality of strains of influenza virus for the presence and concentration of Replikin sequences; [0105](3) comparing the concentration of Replikin sequences in the hemagglutinin amino acid sequence of the at least one isolate of each strain of the plurality of strains of influenza virus to the concentration of Replikin sequences observed in the hemagglutinin amino acid sequence of each of the strains at least one earlier time period to provide the concentration of Replikins for at least two time periods, said at least one earlier time period being within about six months to about three years prior to step (1); [0106](4) identifying the strain of influenza virus having the highest increase in concentration of Replikin sequences during the at least two time periods; [0107](5) selecting at least one Replikin sequence present in the strain of influenza virus peptide identified in step (4) as a peptide for inclusion in an influenza virus vaccine.
[0108]The present invention also provides a method of making a preventive or therapeutic influenza virus vaccine comprising: [0109](1) identifying a strain of influenza virus as an emerging strain; [0110](2) selecting at least one Replikin sequence present in the emerging strain as a peptide template for influenza virus vaccine manufacture, [0111](3) synthesizing peptides having the amino acid sequence of the at least one Replikin sequence selected in step (2), and [0112](4) combining a therapeutically effective amount of the peptides of step (3) with a pharmaceutically acceptable carrier and/or adjuvant.
[0113]In another aspect, the invention is directed to a method of identifying an emerging strain of influenza virus for diagnostic, preventive or therapeutic purposes comprising: [0114](1) obtaining at least one isolate of each strain of a plurality of strains of influenza virus; [0115](2) analyzing the hemagglutinin amino acid sequence of the at least one isolate of each strain of the plurality of strains of influenza virus for the presence and concentration of Replikin sequences; [0116](3) comparing the concentration of Replikin sequences in the hemagglutinin amino acid sequence of the at least one isolate of each strain of the plurality of strains of influenza virus to the concentration of Replikin sequences observed in the hemagglutinin amino acid sequence of each of the strains at least one earlier time period to provide the concentration of Replikins for at least two time periods, said at least one earlier time period being within about six months to about three years prior to step (1); and [0117](4) identifying the strain of influenza virus having the highest increase in concentration of Replikin sequences during the at least two time periods.
[0118]In yet another aspect of the invention, there is provided a preventive or therapeutic influenza virus vaccine comprising at least one isolated Replikin present in the hemagglutinin protein of an emerging strain of influenza virus and a pharmaceutically acceptable carrier and/or adjuvant.
[0119]Also provided by the present invention is a method of preventing or treating influenza virus infection comprising administering to a patient in need thereof a preventive or therapeutic vaccine comprising at least one isolated Replikin present in the hemagglutinin protein of an emerging strain of influenza virus and a pharmaceutically acceptable carrier and/or adjuvant.
Trypanosomes
[0120]In one aspect of the invention there are provided isolated or synthesized trypanosome peptides containing a Replikin sequence. The trypanosome Replikin peptides comprise from 7 to about 50 amino acids including: [0121](1) at least one lysine residue located six to ten amino acid residues from a second lysine residue; [0122](2) at least one histidine residue; and [0123](3) at least 6% lysine residues. (Trypanosome Replikins).
Malaria
[0124]One trypanosome disorder which has proved difficult to treat and for which there is no effective vaccine is malaria. Malaria causes much death, and physical and economic hardship in tropical regions. Malaria is caused mainly by Plasmodium falciparum, which has proved to be extremely resistant to treatment and to date, a vaccine for malaria has remained elusive. Thus there is a need for effective malaria vaccines and methods of treating or preventing the disease. This application provides the basis for such vaccines and methods of treatment and prevention. All of the methods described above for production of and treatment with Replikin virus vaccines and Replikin influenza virus vaccines are applicable to the production of and treatment with Replikin malaria vaccines.
[0125]In the present invention, there are provided vaccines and methods for preventing or treating malaria. The malaria vaccines comprise at least one isolated Plasmodium falciparum Replikin. The present invention also provides methods for treating or preventing malaria comprising administering to a patient an effective amount of preventive or therapeutic vaccine comprising at least one isolated Plasmodium falciparum Replikin.
[0126]Also provided by the present invention are antibodies, antibody cocktails and compositions that comprise antibodies that specifically bind to a Replikin or Replikins present in a malaria antigen of Plasmodium falciparum.
[0127]Another example of a trypanosome which may be treated under the present invention as is the case for malaria, the Replikins of Treponema Pallidum (syphilis), can be used for detection, prevention, treatment of syphilis.
Bacteria
[0128]In one aspect of the invention there are provided isolated or synthesized bacterial peptides containing a Replikin sequence (bacterial Replikins). The bacterial peptides comprise from 7 to about 50 amino acids including: [0129](1) at least one lysine residue located six to ten amino acid residues from a second lysine residue; [0130](2) at least one histidine residue; and [0131](3) at least 6% lysine residues. (bacterial Replikins). U.S. application Ser. No. 10/105,232 filed Mar. 26, 2002 is incorporated by reference in its entirety, including but not limited to the bacterial sequence listing and information.
[0132]The present invention also provides methods for detecting the presence of a contaminating bacterial organism in a body sample or environmental sample comprising: [0133](1) isolating nucleic acids from the body sample or environmental sample; [0134](2) screening the nucleic acids for the presence of a Replikin structure; and [0135](3) correlating the presence of a Replikin structure with the presence of the contaminating organism.
[0136]In another aspect of the invention there is provided a process for stimulating the immune system of a subject to produce antibodies that bind specifically to a bacterial Replikin sequence, said process comprising administering to the subject an effective amount of a dosage of a composition comprising at least one bacterial Replikin peptide. One embodiment comprises at least one bacterial peptide that is present in an emerging strain of the bacterial organism if such new strain emerges.
[0137]The present invention also provides antibodies that bind specifically to a bacterial Replikin, as defined herein, as well as antibody cocktails containing a plurality of antibodies that specifically bind to bacterial Replikins. In one embodiment of the invention, there are provided compositions comprising an antibody or antibodies that specifically bind to a bacterial Replikin and a pharmaceutically acceptable carrier.
[0138]The present invention also provides therapeutic compositions comprising one or more of isolated bacterial peptides having from 7 to about 50 amino acids comprising: [0139](1) at least one lysine residue located six to ten residues from a second lysine residue; [0140](2) at least one histidine residue; [0141](3) at least 6% lysine residues; and [0142](4) a pharmaceutically acceptable carrier.
[0143]In another aspect of the invention there is provided an antisense nucleic acid molecule complementary to a bacterial Replikin mRNA sequence, said Replikin mRNA sequence denoting from 7 to about 50 amino acids comprising: [0144](1) at least one lysine residue located six to ten residues from a second lysine residue; [0145](2) at least one histidine residue; and [0146](3) at least 6% lysine residues.
[0147]In yet another aspect of the invention there is provided a method of simulating the immune system of a subject to produce antibodies to bacteria comprising administering an effective amount of at least one bacterial Replikin peptide having from 7 to about 50 amino acids comprising: [0148](1) at least one lysine residue located six to ten amino acid residues from a second lysine residue; [0149](2) at least one histidine residue; and [0150](3) at least 6% lysine residues.
[0151]In another aspect, there is provided a method of selecting a bacterial Replikin peptide for inclusion in a preventive or therapeutic bacterial vaccine comprising: [0152](1) obtaining at least one isolate of each strain of a plurality of strains of the bacteria; [0153](2) analyzing the amino acid sequence of the at least one isolate of each strain of the plurality of strains of the bacteria for the presence and concentration of bacterial Replikin sequences; [0154](3) comparing the concentration of bacterial Replikin sequences in the amino acid sequence of the at least one isolate of each strain of the plurality of strains of the bacteria to the concentration of bacterial Replikin sequences observed in the amino acid sequence of each of the strains at least one earlier time period to provide the concentration of bacterial Replikins for at least two time periods, said at least one earlier time period being within about six months to about three years prior to step (1), or earlier in rapidly mutating bacteria; [0155](4) identifying the strain of the bacteria having the highest increase in concentration of bacterial Replikin sequences during the at least two time periods; and [0156](5) selecting at least one bacterial Replikin sequence present in the strain of the bacterial peptide identified in step (4) as a peptide for inclusion in the bacterial vaccine.
[0157]The present invention also provides a method of making a preventive or therapeutic bacterial vaccine comprising: [0158](1) identifying a strain of a bacteria as an emerging strain; [0159](2) selecting at least one bacterial Replikin sequence present in the emerging strain as a peptide template for the bacterial vaccine manufacture; [0160](3) synthesizing peptides having the amino acid sequence of the at least one bacterial Replikin sequence selected in step (2); and [0161](4) combining a therapeutically effective amount of the peptides of step (3) with a pharmaceutically acceptable carrier and/or adjuvant.
[0162]In another aspect, the invention is directed to a method of identifying an emerging strain of bacteria for diagnostic, preventive or therapeutic purposes comprising: [0163](1) obtaining at least one isolate of each strain of a plurality of strains of the bacteria; [0164](2) analyzing the amino acid sequence of the at least one isolate of each strain of the plurality of strains of the bacteria for the presence and concentration of bacterial Replikin sequences; [0165](3) comparing the concentration of bacterial Replikin sequences in the amino acid sequence of the at least one isolate of each strain of the plurality of strains of the bacteria to the concentration of bacterial Replikin sequences observed in the amino acid sequence of each of the strains at least one earlier time period to provide the concentration of bacterial Replikins for at least two time periods, said at least one earlier time period being within about six months to about three years prior to step (1); and [0166](4) identifying the strain of the bacteria having the highest increase in concentration of bacterial Replikin sequences during the at least two time periods.
[0167]In yet another aspect of the invention, there is provided a preventive or therapeutic bacterial vaccine comprising at least one isolated bacterial Replikin present in a protein of an emerging strain of the bacteria and a pharmaceutically acceptable carrier and/or adjuvant.
[0168]Two important sub-species of bacteria, classified under mycobacteria, are Mycobacterium leprae (leprosy) whose 30-s ribosomal protein has a C-terminal Replikin and Mycobacterium tuberculosis (tuberculosis) whose ATPase has three Replikins:
TABLE-US-00001 Replikin in 30s ribosomal protein s6 of Mycobacterium leprae (leprosy) is: kvmrtdkh (SEQ ID NO. 699) Replikins in the ATPase of Mycobacterium tuberculosis are: hprpkvaaalkdsyrlk (SEQ ID NO. 700) hprpkvaaalk (SEQ ID NO. 701) ksaqkwpdkflagaaqvah (SEQ ID NO. 702) Replikins in the B-D-galactosidase of E. coli: hawqhqgktlfisrk (SEQ ID NO. 703) hqgktlfisrk (SEQ ID NO. 704) Replikins in Agrobacterium tumefaciens: hsdqqlavmiaakrlddyk (SEQ ID NO. 705) hlldhpasvgqldlramlaveevkidnpvymek (SEQ ID NO. 706) hpasvgqldlramlaveevkidnpvymek (SEQ ID NO. 707) kcvmakncnikcpaglttnqeafngdpralaqylmniah (SEQ ID NO. 708) kncnikcpaglttnqeafngdpralaqylmniah (SEQ ID NO. 709) hhdtysiedlaqlihdakaarvrvivk (SEQ ID NO. 710) hdtysiedlaqlihdakaarvrvivk (SEQ ID NO. 711) hdakaarvrvivk (SEQ ID NO. 712) kigqgakpgeggqlpspkvtvciaaarggtpgvelvsppphh (SEQ ID NO. 713) kigqgakpgeggqlpspkvtveiaaarggtpgvelvsppph (SEQ ID NO. 714) kaseitktlasgamshgalvaaaheavahgtnmvggmsnsgeggeh (SEQ ID NO. 715) kaseitktlasgamshgalvaaaheavah (SEQ ID NO. 716) kaseitktlasgamshgalvaaah (SEQ ID NO. 717) kaseitktlasgamsh (SEQ ID NO. 718) kryfpnvktpvggvtfaviaqavadwh (SEQ ID NO. 719) hhiaaglgfgasavyplgvqfraeekfgadadkafkrfakaackslmk (SEQ ID NO. 720) hhiaaglgfgasavyplgvqfraeekfgadadkafkrfakaackslmk (SEQ ID NO. 721) hhiaaglgfgasavyplgvqfraeekfgadadkafkrfakaaek (SEQ ID NO. 722) hhiaaglgfgasavyplgvqfraeekfgadadkafkrfak (SEQ ID NO. 723) hhiaaglgfgasavyplgvqfraeekfgadadk (SEQ ID NO. 724) hiaaglgfgasavyplgvqfraeekfgadadkafktfakaaekslmk (SEQ ID NO. 725) hiaaglgfgasavyplgvqfraeekfgadadkafkrfakaaek (SEQ ID NO. 726) hiaaglgfgasavyplgvqfraeekfgadadkafkrfak (SEQ ID NO. 727) hiaaglgfgasavyplgvqfraeekfgadadk (SEQ ID NO. 728) kfglydaafeksscgvgfitrkdgvqth (SEQ ID NO. 729)
[0169]Also provided by the present invention is a method of preventing or treating a bacterial infection comprising administering to a patient in need thereof a preventive or therapeutic vaccine comprising at least one isolated bacterial Replikin present in a protein of an emerging strain of the bacteria and a pharmaceutically acceptable carrier and/or adjuvant.
Fungus
[0170]In one aspect of the invention there are provided isolated or synthesized fungal peptides containing a Replikin sequence. The fungal Replikin peptides comprise from 7 to about 50 amino acids including: [0171](1) at least one lysine residue located six to ten amino acid residues from a second lysine residue; [0172](2) at least one histidine residue; and [0173](3) at least 6% lysine residues (fungal Replikins).
[0174]All of the methods described above for production of and treatment with bacterial Replikin vaccines are applicable to the production of and treatment with fungal Replikin vaccines.
[0175]In another aspect of the invention there is provided a process for stimulating the immune system of a subject to produce antibodies that bind specifically to a fungal Replikin sequence, said process comprising administering to the subject an effective amount of a dosage of a composition comprising at least one fungal Replikin peptide.
[0176]The present invention also provides antibodies that bind specifically to a fungal Replikin, as defined herein, as well as antibody cocktails containing a plurality of antibodies that specifically bind to viral Replikins. In one embodiment of the invention, there are provided compositions comprising an antibody or antibodies that specifically bind to a fungal Replikin and a pharmaceutically acceptable carrier.
[0177]The present invention also provides therapeutic compositions comprising one or more of isolated fungal peptides having from 7 to about 50 amino acids comprising: [0178](1) at least one lysine residue located six to ten residues from a second lysine residue; [0179](2) at least one histidine residue; [0180](3) at least 6% lysine residues; and [0181](4) a pharmaceutically acceptable carrier.
[0182]In another aspect of the invention there is provided an antisense nucleic acid molecule complementary to an fungal Replikin mRNA sequence, said Replikin mRNA sequence having from 7 to about 50 amino acids comprising: [0183](1) at least one lysine residue located six to ten residues from a second lysine residue; [0184](2) at least one histidine residue; and [0185](3) at least 6% lysine residues.
[0186]In another aspect of the invention there is provided a process for stimulating the immune system of a subject to produce antibodies that bind specifically to a fungal Replikin sequence, said process comprising administering to the subject an effective amount of a dosage of a composition comprising at least one Replikin peptide.
Increasing Replication
[0187]In yet another aspect of the invention there is provided a method for increasing the replication rate of an organism comprising transforming a gene encoding an enzyme or other protein having a replication function in the organism with at least one Replikin structure.
DEFINITIONS
[0188]As used herein, the term "peptide" or "protein" refers to a compound of two or more amino acids in which the carboxyl group of one is united with an amino group of another, forming a peptide bond. The term peptide is also used to denote the amino acid sequence encoding such a compound. As used herein, "isolated" or "synthesized" peptide or biologically active portion thereof refers to a peptide that is substantially free of cellular material or other contaminating peptides from the cell or tissue source from which the peptide is derived, or substantially free from chemical precursors or other chemicals when chemically synthesized by any method, or substantially free from contaminating peptides when synthesized by recombinant gene techniques.
[0189]As used herein, a Replikin peptide or Replikin protein is an amino acid sequence having 7 to about 50 amino acids comprising: [0190](1) at least one lysine residue located six to ten amino acid residues from a second lysine residue; [0191](2) at least one histidine residue; [0192](3) at least 6% lysine residues.Similarly, a Replikin sequence is the amino acid sequence encoding such a peptide or protein.
[0193]As used herein, "emerging strain" as used herein refers to a strain of a virus, bacterium, fungus, or other organisms identified as having an increased increasing concentration of Replikin sequences in one or more of its protein sequences relative to the concentration of Replikins in other strains of such organism. The increase or increasing concentration of Replikins occurs over a period of at least about six months, and preferably over a period of at least about one year, most preferably over a period of at least about three years or more, for example, in influenza virus, but may be a much shorter period of time for bacteria and other organisms.
[0194]As used herein, "mutation" refers to change in this structure and properties of an organism caused by substitution of amino acids. In contrast, the term "conservation" as used herein, refers to conservation of particular amino acids due to lack of substitution.
BRIEF DESCRIPTION OF THE DRAWINGS
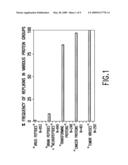
[0195]FIG. 1 is a bar graph depicting the frequency of occurrence of Replikins in various organisms.
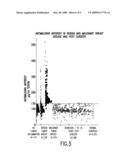
[0196]FIG. 2 is a graph depicting the percentage of malignin per milligram total membrane protein during anaerobic replication of glioblastoma cells.
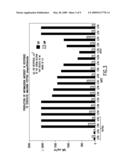
[0197]FIG. 3 is a bar graph showing amount of antimalignin antibody produced in response to exposure to the recognin 16-mer.
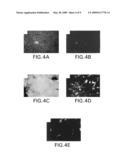
[0198]FIG. 4A is a photograph of a blood smear taken with ordinary and fluorescent light. FIG. 4B is a photograph of a blood smear taken with ordinary and fluorescent light illustrating the presence of two leukemic cells. FIG. 4C is a photograph of a dense layer of glioma cells in the presence of antimalignin antibody. FIG. 4D and FIG. 4E are photographs of the layer of cells in FIG. 4C taken at 30 and 45 minutes following addition of antimalignin antibody
[0199]FIG. 4F is a bar graph showing the inhibition of growth of small cell lung carcinoma cells in vitro by antimalignin antibody.
[0200]FIG. 5 is a plot of the amount of antimalignin antibody present in the serum of patients with benign or malignant breast disease pre- and post surgery.
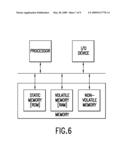
[0201]FIG. 6 is a box diagram depicting an embodiment of the invention wherein a computer is used to carry out the 3-point-recognition method of identifying Replikin sequences.
[0202]FIG. 7 is a graph showing the concentration of Replikins observed in hemagglutinin of influenza B and influenza A strain, H1N1, on a year by year basis from 1918 through 2001.
[0203]FIG. 8 is a graph of the Replikin concentration observed in hemagglutinin of influenza A strains, H2N2 and H3N2, as well as an emerging strain defined by its constituent Replikins, designated H3N2(R), on a year by year basis from 1950 to 2001.
DETAILED DESCRIPTION OF THE INVENTION
[0204]The identification of a new family of small peptides related to the phenomenon of rapid replication, referred to herein as Replikins, provides targets for detection of pathogens in a sample and developing therapies, including vaccine development. In general, knowledge of and identification of this family of peptides enables development of effective therapies and vaccines for any organism that harbors Replikins. Identification of this family of peptides also provides for the detection of viruses and virus vaccine development.
[0205]For example, identification of this family of peptides provides for the detection of influenza virus and provides new targets for influenza treatment. Identification of this family of peptides also provides for example, for the detection of malaria and provides new targets for malaria vaccine development. Further examples provided by the identification of this family of peptides include the detection of infectious disease Replikins, cancer immune Replikins and structural protein Replikins.
[0206]Rapid replication is characteristic of virulence in certain bacteria, viruses and malignancies, but no chemistry common to rapid replication in different organisms has been described. We have found a family of conserved small protein sequences related to rapid replication, which we have named Replikins. Such Replikins offer new targets for developing effective detection methods and therapies. The first Replikin found was the glioma Replikin, which was identified in brain glioblastoma multiforme (glioma) cell protein called malignin.
[0207]Hydrolysis and mass spectrometry of malignin revealed the novel 16 mer peptide sequence which contains the glioma Replikin. This Replikin was not found in databases for the normal healthy human genome and therefore appeared to be derived from some source outside the body.
[0208]We have devised an algorithm to search for the glioma Replikin or homologue thereof Homologues were not common in over 4,000 protein sequences, but were found, surprisingly, in all tumor viruses, and in the replicating proteins of algae, plants, fungi, viruses and bacteria.
[0209]We have identified that both 1) Replikin concentration (number of Replikins per 100 amino acids) and 2) Replikin composition correlate with the functional phenomenon of rapid replication. These relationships provide functional basis for the determination that Replikins are related quantitatively as well as qualitatively to the rate of replication.
[0210]The first functional basis for Replikins role to rapid replication is seen in glioma replication. The fact that glioma malignin is enriched ten-fold compared to the five-fold increase in cell number and membrane protein concentration in rapid replication of glioma cells suggests an integral relationship of the Replikins to replication. When the glioma Replikin was synthesized in vitro and administered as a synthetic vaccine to rabbits, abundant antimalignin antibody was produced. This establishes the antigenic basis of the antimalignin antibody in serum (AMAS) test, and provides the first potential synthetic cancer vaccine and the prototype for Replikin vaccines in other organisms. With the demonstration of this natural immune relationship of the Replikins to replication and this natural immune response to cancer Replikins, which overrides cell type, based upon the shared specificity of cancer Replikins and rapid replication, both passive augmentation of this immunity with antimalignin antibody and active augmentation with synthetic Replikin vaccines now is possible.
[0211]The relationship between the presence of antimalignin antibody and survival in patients was shown in a study of 8,090 serum specimens from cancer patients. The study showed that the concentration of antimalignin antibody increases with age, as the incidence of cancer in the population increases, and increases further two to three-fold in early malignancy, regardless of cell type. In vitro, the antimalignin antibody is cytotoxic to cancer cells at picograms (femtomoles) per cancer cell, and in vivo the concentration of antimalignin antibody relates quantitatively to the survival of cancer patients. As shown in glioma cells, the stage in cancer at which cells have only been transformed to the immortal malignant state but remain quiescent or dormant, now can be distinguished from the more active life-threatening replicating state, which is characterized by the increased concentration of Replikins. In addition, clues to the viral pathogenesis of cancer may be found in the fact that glioma glycoprotein 10B has a 50% reduction in carbohydrate residues when compared to the normal 1OB. This reduction is associated with virus entry in other instances, and so may be evidence of the attachment of virus for the delivery of virus Replikins to the 10B of glial cells as a step in the transformation to the malignant state.
[0212]Our study concerning influenza virus hemagglutinin protein sequences and influenza epidemiology over the past 100 years, has provided a second functional basis for the relations of Replikins to rapid replication. Only serological hemagglutinin and antibody classification, but no strain-specific conserved peptide sequences have previously been described in influenza. Further, no changes in concentration and composition of any strain-specific peptide sequences have been described previously that correlate with epidemiologically documented epidemics or rapid replication. In this study, a four to ten-fold increase in the concentration of strain-specific influenza Replikins in one of each of the four major strains, influenza B, (A)H1N1, (A)H2N2 and (A)143N2, is shown to relate to influenza epidemics caused by each strain from 1902 to 2001.
[0213]We then showed that these increases in concentration are due to the reappearance of at least one specific Replikin composition from 1 to up to 64 years after its disappearance, plus the emergence of new strain-specific Replikin compositions. Previously, no strain-specific chemical structures were known with which to predict the strains that would predominate in coming influenza seasons, nor to devise annual mixtures of whole-virus strains for vaccines. The recent sharp increase in H3N2 Replikin concentration (1997 to 2000), the largest in H3N2's history, and the reappearance of specific Replikin compositions that were last seen in the high mortality H3N2 pandemic of 1968, and in the two high mortality epidemics of 1975 and 1977, but were absent for 20-25 years, together may be a warning of coming epidemics. This high degree of conservation of Replikin structures observed, whereby the identical structure can persist for 100 years, or reappear after an absence of from one to 64 years, indicate that what was previously thought to be change due to random substitution of amino acids in influenza proteins is more likely to be change due to an organized process of conservation of Replikins.
[0214]The conservation of Replikins is not unique to influenza virus but was also observed in other sources, for example in foot and mouth disease virus, type O, HIV tat, and wheat.
[0215]A third functional basis for Replikins' role in rapid replication is seen in the increase in rapid replication in HIV. Replikin concentration was shown to be related to rapid replication in HIV. We found the Replikin concentration in the slow growing low-titre strain of HIV (NS1, "Bru"), which is prevalent in early stage infection, to be one-sixth of the Replikin concentration in the rapidly-growing high-titre strain of HIV (SI, "Lai")(prevalent in late stage HIV infection).
[0216]Further examples demonstrate the relationship of Replikins to rapid replication. In the "replicating protein," of tomato curl leaf gemini virus which devastates tomato crops, the first 161 amino acids, the sequence which has been shown to bind to DNA, was shown to contain five Replikins In malaria, legendary for rapid replication when trypanosomes are released from the liver in the tens of thousands from one trypanosome, multiple, novel, almost `flamboyant` Replikin structures have been found with concentrations of up to 36 overlapping Replikins per 100 amino acids.
[0217]The conservation of any structure is critical to whether that structure provides a stable invariant target to attack and destroy or to stimulate. When a structure is tied in some way to a basic survival mechanism of the organism, the structures tend to be conserved. A varying structure provides an inconstant target, which is a good strategy for avoiding attackers, such as antibodies that have been generated specifically against the prior structure and thus are ineffective against the modified form. This strategy is used by influenza virus, for example, so that a previous vaccine may be quite ineffective against the current virulent virus.
Replikins as Stable Targets for Treatment
[0218]Both bacteria and HIV have both Replikin and non-Replikin amino acids. In HIV, for example, there has been a recent increase in drug-resistance from 9% to 13% due to mutation, that is substitution of non-Replikin amino acids. (See detailed analysis of TAT protein of HIV discussed herein). In bacteria, the development of `resistant strains` is due to a similar mechanism. However, we have found that Replikin structures do not mutate or change to the same degree as non Replikin amino acids (see also discussion of foot and mouth disease virus conservation of Replikins discussed herein). The Replikin structures, as opposed to the non-Replikin structures are conserved and thus provide new constant targets for treatment.
[0219]Certain structures too closely related to survival functions apparently cannot change constantly. Because an essential component of the Replikin structure is histidine (h), which is know for its frequent binding to metal groups in redox enzymes and probable source of energy needed for replication, and since this histidine structure remains constant, this structure remains all the more attractive a target for destruction or stimulation.
[0220]From a proteomic point of view, inventors construction of a template based on the newly determined glioma peptide sequence led them to the discovery of a wide class of proteins with related conserved structures and a particular function, in this case replication. Examples of the increase in Replikin concentration with virulence of a disease include, influenza, HIV, cancer and tomato leaf curl virus. This newly recognized class of structures is related to the phenomenon of rapid replication in organisms as diverse as yeast, algae, plants, the gemini curl leaf tomato virus, HIV and cancer.
[0221]Replikin concentration and composition provide new quantitative methods to detect and control the process of replication, which is central to the survival and dominance of each biological population. The sharing of immunological specificity by diverse members of the class, as demonstrated with antimalignin antibody for the glioma and related cancer Replikins, suggests that B cells and their product antibodies may recognize Replikins by means of a similar recognition language.
[0222]Examples of peptide sequences of cancer Replikins or as containing a Replikin, i.e., a homologue of the glioma peptide, kagvaflhkk, may be found in such cancers of, but not limited to, the lung, brain, liver, soft-tissue, salivary gland, nasopharynx, esophagus, stomach, colon, rectum, gallbladder, breast, prostate, uterus, cervix, bladder, eye, forms of melanoma, lymphoma, leukemia, and kidney.
[0223]Replikins provide for: 1) detection of pathogens by qualitative and quantitative determinations of Replikins; 2) treatment and control of a broad range of diseases in which rapid replication is a key factor by targeting native Replikins and by using synthetic Replikins as vaccines; and 3) fostering increased growth rates of algal and plant foods.
[0224]The first Replikin sequence to be identified was the cancer cell Replikin found in a brain cancer protein, malignin, which was demonstrated to be enriched ten-fold during rapid anaerobic replication of glioblastoma multiforme (glioma) cells. (FIG. 2) Malignin is a 10 KDa portion of the 250 KDa glycoprotein 10B, which was isolated in vivo and in vitro from membranes of glioblastoma multiforme (glioma) cells. Hydrolysis and mass spectroscopy of malignin revealed a 16-mer peptide sequence, ykagvaflhkkndide (SEQ ID NO.:4), which is referred to herein as the glioma Replikin and which includes the shorter peptide, kagvaflhkk (SEQ ID NO.: 1), both of which apparently are absent in the normal human genome.
TABLE-US-00002 TABLE 1 16-mer peptide sequence ykagvaflhkkndide obtained from malignin by hydrolysis and mass spectrometry Method By Which Fragment Obtained Auto- hydrolysis of malignin Auto- immobilized Seq hydrolysis of on ID Fragment MH+ malignin free bromoace-tyl Microwaved Microwaved NO. Identified (mass) Seqence in solution cellulose 5 seconds 30 seconds 19 1-3 381.21 ( )yka(g) + 20 1-5 537.30 ( )ykagv(a) + 21 2-6 445.28 (y)kagva(f) + 22 2-7 592.35 (Y)kagvaf(l) + 23 4-11 899.55 (a)gvaflhkk(n) + 24 5-7 336.19 (g)vaf(l) + 25 6-7 237.12 (v)af(l) + 26 6-10 615.36 (v)aflhk(k) + 27 6-10 615.36 (v)aflhk(k) + 28 6-12 857.50 (v)aflhkkn(d) + 29 6-12 857.50 (v)afhkkn(d) + 30 7-8 279.17 (a)fl(h) + 31 10-16 861.43 (h)kkndide( ) + 32 11-14 489.27 (k)kndi(d) + 33 12-15 476.2 (k)ndid(e) +
[0225]When the 16-mer glioma Replikin was synthesized and injected as a synthetic vaccine into rabbits, abundant antimalignin antibody was produced. (Bogoch et al., Cancer Detection and Prevention, 26 (Suppl. 1): 402 (2002)). The concentration of antimalignin antibody in serum in vivo has been shown to relate quantitatively to the survival of cancer patients. (Bogoch et al., Protides of Biological Fluids, 31:739-747 (1984). In vitro antimalignin antibodies have been shown to be cytotoxic to cancer cells at a concentration of picograms (femtomolar) per cancer cell. (Bogoch et al., Cancer Detection and Prevention, 26 (Suppl. 1): 402 (2002).
[0226]Studies carried out by the inventors showed that the glioma Replikin is not represented in the normal healthy human genome. Consequently, a search for the origin and possible homologues of the Replikin sequence was undertaken by analysis of published sequences of various organisms.
[0227]By using the 16-mer glioma a Replikin sequence as a template and constructing a recognition proteomic system to visually scan the amino acid sequences of proteins of several different organisms, a new class of peptides, the Replikins, was identified. The present invention provides a method for identifying nucleotide or amino acid sequences that include a Replikin sequence. The method is referred to herein as a 3-point-recognition method. The three point recognition method comprises: a peptide from 7 to about 50 amino acids including: [0228](1) at least one lysine residue located six to ten amino acid residues from a second lysine residue; [0229](2) at least one histidine residue; and [0230](3) at least 6% lysine residues. (Replikin).These peptides or proteins constitute a new class of peptides in species including algae, yeast, fungi, amoebae, bacteria, plant, virus and cancer proteins having replication, transformation, or redox functions. Replikin peptides have been found to be concentrated in larger `replicating` and `transforming` proteins (so designated by their investigators, See Table 2) and cancer cell proteins. No sequences were found to be identical to the malignin 16-mer peptide.
TABLE-US-00003 [0230]TABLE 2 Examples of Replikins in various organisms - prototype: Glioma Replikin* kagvaflhkk (SEQ ID No.: 1) SEQ ID NO. Algae: 34 Caldophera prolifera kaskftkh 35 Isolepisprolifera kaqaetgeikgh Yeast: 36 Schizosaccharomyces pombe ksfkypkkhk 37 Oryza sativa kkaygnelhk 2 Sacch. cerevisiae replication binding protein hsikrelgiifdk Fungi: 38 Isocitrate lyase ICI I, Penicillium marneffei kvdivthqk 39 DNA-dependent RNA polymerase II, Discula destructiva klcedaayhrkk 40 Ophiostoma novo-ulm I, RNA in Dutch elm disease kvilplrgnikgiffkh fungus Amoeba: 41 Entamoeba invadens, histone H2B klilkgdlnkh Bacteria: 42 Pribosomal protein replication factor, Helicobacter pylori ksvhaflk 10 Replication-associated protein Staph. Aureus 43 Mycoplasma pulmonic, chromosome replication kkcktthnk 90 Macrophage infectivity potentiator, L. legionella kvhffqlkk Plants: 44 Arabidopsis thaliana, prolifera kdhdfdgdk 45 Arabidopsis thaliana, cytoplasmic ribosomal kmkglkqkkah 46 Arabidopsis thaliana, DNA binding protein kelssttqeksh Viruses: 9 Replication associated protein A [Maize streak virus] Kekkpskdeimrdiish 11 Bovine herpes virus 4, DNA replication protein hkinitngqk 12 Meleagrid herpesvirus I, replication binding protein hkdlyrllmk 47 Feline immunodeficiency hlkdyklvk 3 Foot and Mouth Disease (O) hkqkivapvk 5 HIV Type 1 kcfncgkegh 7 HIV Type 2 kcwncgkegh Tumor 48 Rous sarcoma virus tyrosine-protein kinase kklrhek Viruses: 49 v-yes, avian sarcoma kklrhdk 50 c-yes, colon cancer, malignant melanoma kklrhdk 51 v-srcC, avian sarcoma kklrhek 52 c-src, colon, mammary, panrcreatic cancer kklrhek 53 Neuroblastoma RAS viral (v-ras) oncogene kqahelak 54 VPI (major capsid protein) [Polyamavirus sp.] kthrfskh 55 Sindbis knlhekik 56 E1 [Human papilloamavirus type 71] khrpllqlk 57 v-erbB from AEV and c-erb kspnhvk 58 v-fms (feline sarcoma) knihlekk 59 c-fms (acute and chronic myclomonocytic tumors) knihlekk 60 large t-antigen I [Polyomavirus sp.] kphlaqslek 61 middle t-antigen [Polyomavirus sp,]- kqhrelkdk 62 small t-antigen [Polyomavirus sp], kqhrelkdk 63 v-abl, munine acute leukemia kvpvlisptlkh 64 Human T-cell lymphotropic virus typo 2 kslllevdkdish 65 c-kit, GI tumors, small cell lung carcinoma kagitimvkreyh 18 Hepatitis C hyppkpgcivpak Trans- 66 Transforming protein myb Ksgkhlgk Forming 67 Transforming protein myc, Burkitt lymphoma krreqlkhk Proteins: 68 Ras-related GTP-binding protein ksfevikvih 69 Transforming protein ras (teratocarcinoma) kkkhtvkk 70 TRAF-associated NF•kB activator TANK kaqkdhlsk 71 RFP transforming protein hlkrvkdlkk 72 Transforming protein D (S.C.) kygspkhrlik 73 Papilloma virus type II, transforming protein klkhilgkarfik 74 Protein tryosine kinase (BC 2.7.1.112slk kgdhvkhykirk 75 Transforming protein (axl(-)) keklrdvmvdrhk 76 Transforming protein (N-myc) klqarqqqllkkieh 77 Fibroblast growth factor 4 (Kaposi sarcoma) kkgnrvsptmkvth Cancer 78 Matrix metaloproteinase 7 (uterine) keiplhfrk Cell 79 Transcription factor 7-like kkkphikk Proteins: 80 Breast cancer antigen NY-BR-87 ktrhdplak 81 BRCA-1-Associated Ring Domain Protein (breast) khhpkdnlik 82 `Autoantigen from a breast tumor` khkrkkfrqk 83 Glioma Replikin (this study) kagvaflhkk 84 Ovarian cancer antigen khkrkkfrqk 85 EE L leukemia kkkskkhkdk 86 Proto-oncogene tyrosine-protein kinase C-ABLE hksckpalprk 87 Adenomatosis polyposis coli kkkkpsrlkgdnek 88 Gastric cancer transforming protein ktkkgnrvsptmkvth 89 Transforming protein (K-RAS 2B),lung khkekmskdgkkkkkksk
[0231]Identification of an amino acid sequence as a Replikin or as containing a Replikin, i.e., a homologue of the glioma peptide, kagvaflhkk, requires that the three following requirements be met. According to the three point recognition system the sequences have three elements: (1) at least one lysine residue located six to ten residues from another lysine residue; (2) at least one histidine residue; and (3) a composition of at least 6% lysine within an amino acid sequence of 7 to about 50 residues.
[0232]Databases were searched using the National Library of Medicine keyword "PubMed" descriptor for protein sequences containing Replikin sequences. Over 4,000 protein sequences were visually examined for homologues. Sequences of all individual proteins within each group of PubMed-classified proteins were visually scanned for peptides meeting the three above-listed requirements. An infrequent occurrence of homologues was observed in "virus peptides" as a whole (1.5%) (N=953), and in other peptides not designated as associated with malignant transformation or replication such as "brain peptides" and "neuropeptides" (together 8.5%) (N=845). However, surprisingly, homologues were significantly more frequently identified in large "replicating proteins," which were identified as having an established function in replication in bacteria, algae, and viruses. Even more surprising was the finding that Replikin homologues occurred in 100% of "tumor viruses" (N=250), in 97% of "cancer proteins" (N=401), and in 85% of "transforming viruses" (N=248). These results suggest that there are shared properties of cancer pathogenesis regardless of cell type and suggest a role of viruses in carcinogenesis, i.e., conversion of cells from a transformed albeit dormant state to a more virulent actively replicating state.
[0233]Homologues of the following amino acid sequence, kagvaflhkk, as defined by the three point recognition method, were found in such viruses, or viral peptides, as, but not limited to, adenovirus, lentivirus, a-virus, retrovirus, adeno-associated virus, human immunodeficiency virus, hepatitis virus, influenza virus, maize streak virus, herpes virus, bovine herpes virus, feline immunodeficiency virus, foot and mouth disease virus, small pox virus, rous sarcoma virus, neuroblastoma RAS viral oncogene, polyamavirus, sindbis, human papilloma virus, myelomonocytic tumor virus, murine acute leukemia, T-cell lymphotropic virus, and tomato leaf curl virus.
[0234]Replikins are present in such bacteria as, but not limited to, Acetobacter, Achromobacter, Actinomyces, Aerobacter, Alcaligenes, Arthrobacter, Azotobacter, Bacillus, Brevibacterium, Chainia, Clostridium, Corynebacterium, Erwinia, Escheria, Lebsiella, Lactobacillus, Haemophilus, Flavobacterium, Methylomonas, Micrococcus, Mycobacterium, Micronomspora, Mycoplasma, Neisseria, Nocardia, Proteus, Pseudomonas, Rhizobium, Salmonella, Serratia, Staphylococcus, Streptocossus, Streptomyces, Streptosporangium, Streptovirticillium, Vibrio, peptide, and Xanthomas.
[0235]Replikins are present in such fungi as, but not limited to, Penicillium, Diseula, Ophiostoma novo-ulim, Mycophycophta, Phytophthora infestans, Absidia, Aspergillus, Candida, Cephalosporium, Fusarium, Hansenula, Mucor, Paecilomyces, Pichia, Rhizopus, Torulopsis, Trichoderma, and Erysiphe.
[0236]Replikins are present in such yeast as, but not limited to, Saccharomyces, Cryptococcus, including Cryptococcus neoformas, Schizosaccharomyces, and Oryza.
[0237]Replikins are present in algae such as, but not limited to, Caldophera, Isolepisprolifera, Chondrus, Gracilaria, Gelidium, Caulerpa, Laurencia, Cladophexa, Sargassum, Penicillos, Halimeda, Laminaria, Fucus, Ascophyllum, Undari, Rhodymenia, Macrocystis, Eucheuma, Ahnfeltia, and Pteroclasia.
[0238]Replikins are present in amoeba such as, but not limited to, Entamoeba (including Entamoeba invadens), Amoebidae, Acanthamoeba and Naegleria.
[0239]Replikins are present in plants such as, but not limited to, Arabidopsis, wheat, rice, and maize.
Auxiliary Specifications
[0240]To permit classification of subtypes of Replikins, additional or "auxiliary specifications" to the basic "3-point-recognition" requirements may be added: (a) on a structural basis, such as the common occurrence of adjacent di- and polylysines in cancer cell proteins (e.g., transforming protein P21B(K-RAS 2B), lung, Table 2, SEQ ID NO.: 89), and other adjacent di-amino acids in TOLL-like receptors, or b) on a functional basis, such as exhibiting ATPase, tyrosine kinase or redox activity as seen in Table 2.
Functional Derivatives
[0241]"Functional derivatives" of the Replikins as described herein are fragments, variants, analogs, or chemical derivatives of the Replikins, which retain at least a portion of the immunological cross reactivity with an antibody specific for the Replikin. A fragment of the Replikin peptide refers to any subset of the molecule. Variant peptides may be made by direct chemical synthesis, for example, using methods well known in the art. An analog of a Replikin to a non-natural protein substantially similar to either the entire protein or a fragment thereof. Chemical derivatives of a Replikin contain additional chemical moieties not normally a part of the peptide or peptide fragment.
[0242]As seen in FIG. 2, during anaerobic respiration when the rate of cell replication is increased, malignin is enriched. That is, malignin is found to increase not simply in proportion to the increase in cell number and total membrane proteins, but is enriched as much as ten-fold in concentration, starting with 3% at rest and reaching 30% of total membrane protein. This clear demonstration of a marked increase in Replikin concentration with glioma cell replication points to, and is consistent with, the presence of Replikins identified with the 3-point recognition method in various organisms. For example, Replikins were identified in such proteins as "Saccharomyces cerevisiae replication binding protein" (SEQ ID NO.: 2) (hsikrelgiifdk); the "replication associated protein A of maize streak virus" (SEQ ID NO.: 8) (kyivcarealtk) and (SEQ ID NO.: 9) (kekkpskdeimrdiish); the "replication-associated protein of Staphylococcus aureus" (SEQ ID NO.: 10) (kkektthnk); the "DNA replication protein of bovine herpes virus 4" (SEQ ID NO.: 11) (hkinitngqk); and the "Mealigrid herpes virus 1 replication binding protein" (SEQ ID NO.: 12) (hkdlyrllmk). Previous studies of tomato leaf curl gemini virus show that the regulation of virus accumulation appears to involve binding of amino acids 1-160 of the "replicating protein" of that virus to leaf DNA and to other replication protein molecules during virus replication. Analysis of this sequence showed that amino acids 1-163 of this "replicating protein" contain five Replikins, namely: (SEQ ID NO.: 13) kfrinaknyfltyph, (SEQ ID NO.: 14) knletpvnklfiricrefh, (SEQ ID NO.: 15) hpniqaaksstdvk, (SEQ ID NO.: 16) ksstdvkaymdkdgdvldh, and (SEQ ID NO.: 17) kasalnilrekapkdfvlqfih.
[0243]Table 2 shows that Replikin-containing proteins also are associated frequently with redox functions, and protein synthesis or elongation, as well as with cell replication. The association with metal-based redox functions, the enrichment of the Replikin-containing glioma malignin concentration during anaerobic replication, and the cytotoxicity of antimalignin at low concentrations (picograms/cell) (FIG. 4C-f), all suggest that the Replikins are related to central respiratory survival functions, have been found less often subjected to the mutations characteristic of non-Replikin amino acids.
[0244]Of particular interest, it was observed that at least one Replikin per 100 amino acids was found to be present in the hemagglutinin proteins of almost all of the individual strains of influenza viruses examined. The Replikin sequences that were observed to occur in the hemagglutinin proteins of isolates of each of the four prevalent strains of influenza virus, influenza B, H1N1, H2N2, and H3N2, for each year that amino acid sequence data are available (1902-2001), are shown in Tables 3, 4, 5 and 6, below.
TABLE-US-00004 TABLE 3 Replikin Sequences present in hemagglutinins of Influenza B viruses in each year for which amino acid sequences were available (1902-2001). Influenza B Replikins Year Detected in Influenza B strain (Peak in FIG. 7: EB1 EB2) kshfanlk 1902, 19, 24, 38, 40, 43, 51, 59, 75, 76, 77, 89, 90, 93, 97, 98, 99, (SEQ ID NO. 91) 00, 01 kshfanlkgtk 1902, 19, 24, 38, 40, 43, 51, 59, 75, 76, 77, 89, 90, 93, 97, 98, 99, (SEQ ID NO. 92) 00, 01 kshfanlkgtktrgklcpk 1902, 19, 24, 38, 40, 43, 51, 59, 75, 76, 77, 89, 90, 93, 97, 98, 99, (SEQ ID NO. 93) 00, 01 hekygglnk 1902, 19, 24, 38, 40, 43, 51, 59, 75, 76, 77, 89, 90, 93, 97, 98, 99, (SEQ ID NO. 94) 00, 01 hekygglnksk 1902, 19, 24, 38, 40, 43, 51, 59, 75, 76, 77, 89, 90, 93, 97, 98, 99, (SEQ ID NO. 95) 00, 01 hekygglnkskpyytgehak 1902, 19, 24, 38, 40, 43, 51, 59, 75, 76, 77, 89, 90, 93, 97, 98, 99, (SEQ ID NO. 96) 00, 01 hakaigncpiwvk 1902, 19, 24, 38, 40, 43, 51, 59, 75, 76, 77, 89, 90, 93, 97, 98, 99, (SEQ ID NO. 97) 00, 01 hakaigncpiwvktplklangtk 1902, 19, 24, 38, 40, 43, 51, 59, 75, 76, 77, 89, 90, 93, 97, 98, 99, (SEQ ID NO. 98) 00, 01 hakaigncpiwvktplklangtkyrppak 1902, 19, 24, 38, 40, 43, 51, 59, 75, 76, 77, 89, 90, 93, 97, 98, 99, (SEQ ID NO. 99) 00, 01 hakaigncpiwvktplklangtkyrppakllk 1902, 19, 24, 38, 40, 43, 51, 59, 75, 76, 77, 89, 90, 93, 97, 98, 99, (SEQ ID NO. 100) 00, 01 hfanlkgtktrgk 1919, 76, 89, 90, 99, 00, 01 (SEQ ID NO. 101) hfanlkgtktrgklcpk 1919, 76, 90 00, 01 (SEQ ID NO. 102) hsdneiqmvklygdsk 1919 (SEQ ID NO. 103) hsdneiqdkmvklygdskpqk 1919 (SEQ ID NO. 104) hsdneiqmvklygdskpqk 1919, 24, 97, 98, 00 (SEQ ID NO. 105) k(a/v)silhevk 1919, 40, 59, 90, 93 (SEQ ID NO. 106) kctgtipsakasilh 1919, 00 (SEQ ID NO. 107) kctgtipsakasilhevk 1919, 93 (SEQ ID NO. 108) kygglnkskpyytgeh 1919 (SEQ ID NO. 109) kvwcasgrskvikgslpligeadclh 1919, 38, 40, 43, 59, 75, 76, 77, 89, 90, 98, 99, 00 (SEQ ID NO. 110) kpyytgehak 1919, 38, 40, 59, 89, 90, 93, 97, 98, 01 (SEQ ID NO. 111) kcmgtipsakasilhevk 1924, 43, 75, 76, 77, 93 (SEQ ID NO. 112) hnvinaekapggpyk 1938, 93, 97, 00 (SEQ ID NO. 113) hsdnetqmaklygdsk 1938, 93, 97, 00 (SEQ ID NO. 114) hgvavaadlkstqeaink 1940, 59, 00 (SEQ ID NO. 115) hgvavaadlkstqeainkdtistqeaink 1940 (SEQ ID NO. 116) klygdskpqkftssangvtth 1943, 75, 76, 77, 93, 97, 00 (SEQ ID NO. 117) hsdnetqmaklygdskpqk 1943, 75, 76, 77, 93 (SEQ ID NO. 118) hfanlkgtqtrgk 1959 (SEQ ID NO. 119) kprsalkckgfh 1988 (SEQ ID NO. 120) kskpyytgehakai(g/a)ncpiwvk 2000 (SEQ ID NO. 121) 1. Influenza B has not been responsible for any human pandemic (global distribution). 2. Abbreviation for years: eg. "19"=1919, "01"=2001. 3. The first year that a given Replikin appears is indicated at the beginning of the series of years in which that Replikin has been found. 4. Overlapping Replikin sequences are listed separately. 5. Increase in number of new Replikin structures occurs in years of epidemics (underlined): eg. 1951 and 1977 and correlates with increased total Replikin concentration (number of Replikins per 100 amino acid residues). See FIG. 7.
TABLE-US-00005 TABLE 4 H1N1 Replikin Sequences present in H1N1 hemagglutinins of Influenza viruses in each year for which amino acid sequences were available (1918-2000) H1N1 Replikin Year Detected in Influenza H1N1 Strain Peak in FIG. 7 P1 E1 E1.1, 1.2, 1.3 E1.4) hp(v/t)tigecpkyv(r/k)(s/t)(t/a)k 1918, 25, 28, 30, 31, 35, 47, 48, 51, (SEQ ID NO. 122) 52, 55, 56, 57, 59, 63, 77, 79, 80, 81, 85, 87, 88, 89, 91, 92, 95, 96, 97, 98, 99, 00 hdsnvknly(e/g)kv(k/r)(n/s)ql(k/r)nnak 1918, 28, 30, 31, 77, 79, 80, 88, 91, (SEQ ID NO. 123) 95, 98 hdsnvknly(e/g)kv(k/r)(n/s)qlk 1918, 28, 30, 31, 77, 79, 80, 88, 91, (SEQ ID NO. 124) 95, 98 hkc(nn/dd)(a/t/e)cmesv(r/k)ngtydypkyseesklnre(e/k)idgvk 1918, 30, 35, 77, 80, 98 (SEQ ID NO. 125) hkc(nn/dd)(a/t/e)cmesv(r/k)ngtydypkyseesk 1918, 30, 35, 77, 80, 98 (SEQ ID NO. 126) hqn(e/g)qgsgyaadqkstqnai(d/n)gitnkvnsviekmntqftavgkefnklek 1918, 28, 30, 31, 35, 59, 79, 95 (SEQ ID NO. 127) hqn(e/g)qgsgyaadqkstqnai(d/n)gitnkvnsviek 1918, 28, 30, 31, 35, 59, 79, 95 (SEQ ID NO. 128) hqn(e/g)qgsgyaadqkstqnai(d/n)gitnk 1918, 28, 30, 31, 35, 59, 79, 95 (SEQ ID NO. 129) kfeifpktsswpnh 1918, 77 (SEQ ID NO. 130) kg(n/s/t)sypkl(n/s)ksy(v/t)nnkgkevlvlwgvh 1918, 35, 77, 96 (SEQ ID NO. 131) ksy(v/t)nnkgkevlvlwgvh 1918, 35, 77, 96 (SEQ ID NO. 132) hkenneemesvkngtydypkyseesklnrekidgvk 1928, 31, 95 (SEQ ID NO. 133) hkenneemesvkngtydypkyseesk 1928, 31, 95 (SEQ ID NO. 134) hkenneemesvkngtydypk 1928, 31, 95 (SEQ ID NO. 135) hkennecmesvk 1928, 31, 95 (SEQ ID NO. 136) hngkssfy(k/r)nllwlt(e/g)knglypnlsksyvnnkek 1928, 95, 00 (SEQ ID NO. 137) hngkssfy(k/r)nllwlt(e/g)knglypnlsksyvnnk 1928, 31, 95, 00 (SEQ ID NO. 138) hngkssfy(k/r)nllwlt(e/g)knglypnlsk 1928, 31, 95, 00 (SEQ ID NO. 139) hngkssfy(k/r)nllwlt(e/g)k 1928, 31, 95, 00 (SEQ ID NO. 140) kssfyknllwlteknglypnlsksyvnnkekevlvlwgvh 1928, 31, 95 (SEQ ID NO. 141) knllwlteknglypnlsksyvnnkekevlvlwgvh 1928, 31, 95 (SEQ ID NO. 142) knglypnlsksyvnnkekevlvlwgvh 1928, 31, 95, 96, 00 (SEQ ID NO. 143) ksy(v/a)nnkekev(l/-)(v/-)lwgvh 1928, 31, 51, 95, 96, 98, 00 (SEQ ID NO. 144) kesswpnhtvtk 1928, 31, 95 (SEQ ID NO. 145) het(t/n)kgvtaacpyagassfyrnllwlvkkensypklsksyvnnk 1930, 35 (SEQ ID NO. 146) het(t/n)kgvtaacpyagassfymllwlvkkensypklsk 1930, 35 (SEQ ID NO. 147) kfeifpktsswpnevlvlwgvh 1930 (SEQ ID NO. 148) kerswpkh 1947, 51, 52, 55, 56, 79, 82 (SEQ ID NO. 149) klsksyvnnkekevlvlwqvh 1947, 51 (SEQ ID NO. 150) knnkekevlvlwqvh 1947 (SEQ ID NO. 151) h(k/n)(g/q)kssfy(r/k)nllwltekng(l/s)yp(n/t)lsksyannkek 1948 79, 89, 96 (SEQ ID NO. 152) h(k/n)(g/q)kssfy(r/k)nllwltek 1948 79, 89, 96 (SEQ ID NO. 153) hakkssfyk 1951, 57, 59 (SEQ ID NO. 154) hngklcrlkgk 1951, 52, 55, 56, 57, 59, 79 (SEQ ID NO. 155) hyklnn(q/g)kk 1956, 00 (SEQ ID NO. 156) hdiyrdeainnrfqiqgvkltqgyk 1956 (SEQ ID NO. 157) kgngcfeifhk 1956 (SEQ ID NO. 158) klnrliektndkyhqiek 1956 (SEQ ID NO. 159) klnrliektndkyh 1956 (SEQ ID NO. 160) kchtdkgslsttk 1956 (SEQ ID NO. 161) kinngdyaklyiwgvh 1956 (SEQ ID NO. 162) hngklcrkgiaplqlgk 1959, 82 (SEQ ID NO. 163) hetnrqvtaacpyagansffrnhiwlvkkessypklsk 1963, 81 (SEQ ID NO. 164) hetnrqvtaacpyagansffrnliwlvkkessypk 1963, 81 (SEQ ID NO. 165) hpptstdqqslyqnadayifvgsskynrkfk 1963, 81 (SEQ ID NO. 166) hpptsrdqqslyqnadayifvgsskynrkfkpeia 1963, 81 (SEQ ID NO. 167) hdiyrdeainnrfqiqgvkitqgyk 1977, 79, 91 (SEQ ID NO. 168) hqneqgsgyaadqkstqnaidgitnkvnsviekmntqftavgk 1977 (SEQ ID NO. 169) hqneqgsgyaadqkstqnaidgitnkvnsviek 1977 (SEQ ID NO. 170) hqneqgsgyaadqkstqnaingitnkvnsviekmntqftavgkefnklek 1979, 91 (SEQ ID NO. 171) hngklcrlkgiaplqlgk 1979 (SEQ ID NO. 172) hkcnnecmesvk 1979 (SEQ ID NO. 173) kfeifpkasswpnh 1981 (SEQ ID NO. 174) hdsnvknlyekvrsqlrnnak 1981 (SEQ ID NO. 175) kvnsvikkmntqfaavgkefnh 1981 (SEQ ID NO. 176) khngklck 1981 (SEQ ID NO. 177) kkgtsypklsksythnkgkevlvlwgvh 1981 (SEQ ID NO. 178) kgtsypklsksythnkgkevlvlwgvh 1981 (SEQ ID NO. 179) klsksythnkgkevlvlwgvh 1981 (SEQ ID NO. 180) ksythnkgkevlvlwgvh 1981 (SEQ ID NO. 181) kgvtascshk 1985, 87 (SEQ ID NO. 182) kgvtascshkgrssfyrnllwlteknglypnlsk 1985, 87 (SEQ ID NO. 183) kgnsypklsksyvnnkekevlvlwgih 1988 (SEQ ID NO. 184) kefnhlek 1988 (SEQ ID NO. 185) hpptstdqqslyqnadayvfvgsskynkkfkpeiatrpk 1988 (SEQ ID NO. 186) hpptstdqqslyqnadayvfvgsskynkkfk 1988 (SEQ ID NO. 187) hegkssfyrnllwltekegsypklknsyvnk 1991 (SEQ ID NO. 188) hegkssfyrnllwltekegsypk 1991 (SEQ ID NO. 189) hkcdnecmesvrngtydypkyseesk 1991 (SEQ ID NO. 190) kesswpnhtvtk 1991, 92 (SEQ ID NO. 191) knllwlteknglypnlsksyvnnkekeilvlwgvh 1991, 92, 96 (SEQ ID NO. 192) hngkssfy(k/m)(n/-)llwlt(e/g)(-/k)knglypnlsk 1991, 92, 96, 00 (SEQ ID NO. 193) hngkssfyknllwltek 1991, 92, 96 (SEQ ID NO. 194) htvtkgvtascshngkssfyknllwlteknglypnlsksyvnnkekevlvlwgvh 1995 (SEQ ID NO. 195) htvt(k/g)gv(t/s)ascshngkssfy(k/m)(n/-)llwlt(e/g)k(-n/k)glypnlsk 1995, 00 (SEQ ID NO. 196) htvtkgvtascshngkssfyknllwltek 1995 (SEQ ID NO. 197) kyvrstklrmvtglrnipsiqsrglfgaiagfieggwtgmidgwygyh 1995 (SEQ ID NO. 198) hqneqgsgyaadqkstqnaingitnkvnsiiekmntqftavgk 1995 (SEQ ID NO. 199)
hqneqgsgyaadqkstqnaingitnkvnsiiek 1995 (SEQ ID NO. 200) hqneqgsgyaadqkstqnaingitnk 1995 (SEQ ID NO. 201) hsgarsfyrnllwivkkgnsypk 1996 (SEQ ID NO. 202) hsgarsfyrnllwivkkgnsypklnk 1996 (SEQ ID NO. 203) hsgarsfyrnllwivkkgnsypklnksytndk 1996 (SEQ ID NO. 204) hsgarsfyrnllwivkkgnsypklnksytndkgk 1996 (SEQ ID NO. 205) htvskgvttscshngk 1996 (SEQ ID NO. 206) katswpnhettk 1996 (SEQ ID NO. 207) kqvttscshnqk 1996 (SEQ ID NO. 208) kgnsvpklnksytndkgkevlviwgvh 1996 (SEQ ID NO. 209) klnksytndkgkevlviwgvh 1996 (SEQ ID NO. 210) ksytndkgkevlviwgvh 1996 (SEQ ID NO. 211) hnqkssfyrnllwlt(e/q)knglypnlsksy(v/a)annkek 1997, 98, 99 (SEQ ID NO. 212) hpitigecpkyvrsak 1997 (SEQ ID NO. 213) hqneqgsgyaadqkstqnaingitnkvnsvlekmntqftavgk 1998 (SEQ ID NO. 214) hqneqgsgyaadqkstqnaingitnkvnsviek 1998 (SEQ ID NO. 215) hngkssfyrnllwlteknglypnlsksyvnnkek 1998 (SEQ ID NO. 216) 1. Influenza H1N1 was responsible for the human pandemic (global distribution) of 1918. 2. Abbreviation for years: eg. "96" = 1996. 3. The first year that a given Replikin appears is indicated at the beginning of the series of years in which that Replikin has been found in this work. 4. Overlapping Replikin sequences are listed separately. 5. Increase in number of new Replikin structures occurs in years of epidemics (underlined): eg. 1918 and 1977 and correlates with increased total Replikin concentration (number of Replikins per 100 amino acid residues). See FIG. 7.
TABLE-US-00006 TABLE 5 Replikin Sequences present in hemagglutinins of Influenza H2N2 viruses in years 1957-2000 Influenza H2N2 Reylikins Year Detected in Influenza H2N2 strain (Peak in FIG. 8: P2 E2) khfekvkilpk 1957, 58, 59, 60, 61, 64, 65, 68, 78, 83, 84, 91 (SEQ ID NO. 217) khllssvkhfekvk 1957, 58, 59, 60, 61, 83, 84, 91 (SEQ ID NO. 218) ha(k/q/m)(d/n)ilekthngk 1957, 58, 59, 60, 61, 64, 65, 68, 78, 83, 84, 91, 95 (SEQ ID NO. 219) ha(k/q/m)(d/n)ilekthngklc(k/r) 1957, 58, 59, 60, 61, 64, 65, 68, 78, 83, 84, 91, 95 (SEQ ID NO. 220) hnvhpltigecpkyvksek 1957, 58, 59, 65, 68 (SEQ ID NO. 221) hpltigecpkyvksek 1957, 58, 59, 65, 68, 64, 65, 68, 78, 83, 84, 91 (SEQ ID NO. 222) khllssvkhfekvkilpk 1957, 58, 59, 60, 61, 64, 65, 68, 78 (SEQ ID NO. 223) krqssgimktegtlencetkcqtplgainttlpfhnvh 1957, 59, 83 (SEQ ID NO. 224) kgsnyp(v/i)ak(g/r)synntsgeqmliiwq(v/i)h 1957, 58, 59, 61, 83, 91, 95 (SEQ ID NO. 225) httlgqsracavsgnpsffmmvwltekgsnypvak 1957 (SEQ ID NO. 226) khfekvk 1957, 59, 65 (SEQ ID NO. 227) kiskrgssgimktegrlencetkcqtplgainttlpfh 1957, 59, 65, 91 (SEQ ID NO. 228) krgssgimktegtlencetkcqtplgainttlpfh 1957, 59, 65, 91 (SEQ ID NO. 229) ktegtlencetkcqtplgainttlpfh 1957, 59, 65, 91 (SEQ ID NO. 230) kiskrgssgimktegtlencetkcqtplgainttlpfh 1957, 59, 65, 91 (SEQ ID NO. 231) ktegtlencetkcqtplgainttlpfhn(v/i)h 1957, 59, 65, 91 (SEQ ID NO. 232) kiskrgssgimktegtlencetkcqtplgainttlpfh 1957, 59, 65, 91 (SEQ ID NO. 233) k(e/g)snypvakgsynntsgeqmliiwgvh 1957, 60, 65 (SEQ ID NO. 234) hpltigccpkyvksek 1957, 60, 65 (SEQ ID NO. 235) kcqtplgaikttlpfh 1957, 65 (SEQ ID NO. 236) hhsndqgsgyaadkestqka(f/i)dgitnkvnsviek- 1961, 65, 68, 83, 84 -mntqfeavgklf(n/s)nleklenlnkk (SEQ ID NO. 237) hsndqgsgyaadkestqka(f/i)dgitnkvnsviek- 1961, 65, 68, 83, 84 -mntqfeavgklf(n/s)nleklenlnkk (SEQ ID NO. 238) hsndqgsgyaadkestqka(f/i)dgitnk 1961, 65, 68, 83, 84 (SEQ ID NO. 239) hdsnvrnlydkvrmqlrdnak 1964, 68, 76, 84, 91 (SEQ ID NO. 240) hkcddecmnsvkngtydypklnrneikgvk 1964, 65, 68, 76, 83, 84, 91 (SEQ ID NO. 241) hkcddecmnsvkngtydypklnrneik 1964, 65, 68, 76, 83, 84, 91 (SEQ ID NO. 242) hkcddecmnsvkngtydypk 1964, 65, 68, 76, 83, 84, 91 (SEQ ID NO. 243) hkcddecmnsvk 1964, 65, 68, 76, 83, 84, 91 (SEQ ID NO. 244) kgsnypvakgsynntngeqiliiwgvh 1976, 78 (SEQ ID NO. 245) hsndqgsgyaadkestqkavdgitnkvnsviekmntqfeavgk 1976, 91 (SEQ ID NO. 246) krgssgimktegtlencetkcqtplgainttlpfh 1976, 78, 83, 84 (SEQ ID NO. 247) hpltigecpkyvksek 1976 (SEQ ID NO. 248) hakdilekthngklck 1976 (SEQ ID NO. 249) 1. Influenza H2N2 was responsible for the human pandemic (global distribution) of 1957. 2. Abbreviation for years: eg. "58" = 1958. 3. The first year that a given Replikin appears is indicated at the beginning of the series of years in which that Replikin has been found in this work. 4. Overlapping Replikin sequences are listed separately. 5. Increase in number of new Replikin structures occurs in years of epidemics (underlined): eg. 1957 and 1965 and correlates with increased total Replikin concentration (number of Replikins per 100 amino acid residues). See FIG. 8.
TABLE-US-00007 TABLE 6 H3N2 Replikin Sequences present in H3N2 hemagglutinins of Influenza viruses in each year for which amino acid sequences were available (1968-2000) Influenza H3N2 Replikins Year Detected in Influenza H3N2 strain Influenza Replikins (Peak in FIG. 8: P3 E3 E4) hdvyrdealnnrfqikgvelksgyk 1968, 72, 75 96, 97, 98 (SEQ ID NO. 250) htidltdsemnklfertrk 1968 (SEQ ID NO. 251) kfhqiek 1968, 72, 75, 77 96, 97, 98 (SEQ ID NO. 252) ktnekfh(g/q)iek 1968 86 98 (SEQ ID NO. 253) klnr(v/l)iektnekfh 1968, 72, 75, 77 97, 98 (SEQ ID NO. 254) hqiekefsevegriqdlekyvedtk 1968, 72, 98 (SEQ ID NO. 255) kicnnphk 1975 (SEQ ID NO. 256) klnrvikktnekfh 1975 (SEQ ID NO. 257) hd(l,v)yrdealnnrfqik(g/q)ve(r/k)s(q/g)yk 1975, 76, 77, 86 (SEQ ID NO. 258) hqiekefsevegriqdlekyvedtk 1975 (SEQ ID NO. 259) kyvedtkidlwsynaellvalenqh 1975 (SEQ ID NO. 260) kyvkqnslklatgmrnvpekqtrglfgaiagfiengwegmidgwygfrh 1975 (SEQ ID NO. 261) kefsevegriqdlekyvedtkidlwsynaellvalenqh 1975 2000 (SEQ ID NO. 262) hqn(s/e)(e/q)g(t/s)g(q/y)aad(l/q)k- 1975 2000 -stq(a/n)a(i/l)d(q/g)l(n/t)(g/n)k(l/v)n(r/s)vi(e/c)k (SEQ ID NO. 263) hcd(g/q)f(q,r)nekwdlf(v,/i)er(s/t)k 1975, 76, 77, 78, 80, 81, 82, (SEQ ID NO. 264) 83, 84, 85, 86, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98 htidltdsemnkklfertrk 1977, (SEQ ID NO. 265) ksgstypvlkvtmpnndnfdklyiwgvh 1977 (SEQ ID NO. 266) klnwltksgntypvlnvtmpnndnfdklviwgvh 1982 (SEQ ID NO. 267) htidltdsemnklfektrk 1986 (SEQ ID NO. 268) klnrliektnekfhqtek 1987 (SEQ ID NO. 269) htgkssvmrsdapidfcnsecitpnqsipndkpfqnvnkitygacpk 1994 (SEQ ID NO. 270) htgkssvmrsdapidfcnsecitpnqsipndkpfqnvnk 1994 (SEQ ID NO. 271) hpstdsdqtslyvrasgrvtvstkrsqqtvipk 1994 (SEQ ID NO. 272) kyvedtkidlwsynaellvalenqh 1997, 98 (SEQ ID NO. 273) klfertrkqlrenaedmgngcfkiyh 1998 (SEQ ID NO. 274) krrsiksffsrlnwlh 1998 (SEQ ID NO. 275) hpvtigecpky(v/r)kstk 2000 (SEQ ID NO. 276) kgnsypklsklsksyiinkkkevlviwgih 2000 (SEQ ID NO. 277) klsklsks(v/y)iinkkkevlviwgih 2000 (SEQ ID NO. 278) klsks(v/y)iinkkkevlviwgih 2000 (SEQ ID NO. 279) 1. Influenza H3N2 was responsible for the human pandemic (global distribution) of 1968. 2. Abbreviation for years: eg. "77" = 1977. 3. The first year that a given Replikin appears is indicated at the beginning of the series of years in which that Replikin has been found. 4. Overlapping Replikin sequences are listed separately. 5. Increase in number of new Replikin structures occurs in years of epidemics (underlined) : eg. 1975 and correlates with increased total Replikin concentration (number of Replikins per 100 amino acid residues). See FIG. 8.
[0245]Both the concentration and type, i.e., composition of Replikins observed, were found to relate to the occurrence of influenza pandemics and epidemics. The concentration of Replikins in influenza viruses was examined by visually scanning the hemagglutinin amino acid sequences published in the National Library of Medicine "PubMed" data base for influenza strains isolated world wide from human and animal reservoirs year by year over the past century, i.e., 1900 to 2001. These Replikin concentrations (number of Replikins per 100 amino acids, mean+/-SD) were then plotted for each strain.
[0246]The concentration of Replikins was found to directly relate to the occurrence of influenza pandemics and epidemics. The concentration of Replikins found in influenza B hemagglutinin and influenza A strain, H1N1, is shown in FIG. 7, and the concentration of Replikins found in the two other common influenza virus A strains, H2N2 and H3N2 is shown in FIG. 8 (H2N2, H3N2). The data in FIG. 8 also demonstrate an emerging new strain of influenza virus as defined by its constituent Replikins (H3N2(R)).
[0247]Each influenza A strain has been responsible for one pandemic: in 1918, 1957, and 1968, respectively. The data in FIGS. 7 and 8 show that at least one Replikin per 100 amino acids is present in each of the influenza hemagglutinin proteins of all isolates of the four common influenza viruses examined, suggesting a function for Replikins in the maintenance of survival levels of replication. In the 1990s, during the decline of the H3N2 strain, there were no Replikins in many isolates of H3N2, but a high concentration of new Replikins appeared in H3N2 isolates, which define the emergence of the H3N2(R) strain.
[0248]Several properties of Replikin concentration are seen in FIG. 7 and FIG. 8 to be common to all four influenza virus strains. First, the concentration is cyclic over the years, with a single cycle of rise and fall occurring over a period of two to thirty years. This rise and fall is consistent with the known waxing and waning of individual influenza virus strain predominance by hemagglutinin and neuraminidase classification. Second, peak Replikin concentrations of each influenza virus strain previously shown to be responsible for a pandemic were observed to relate specifically and individually to each of the three years of the pandemics. For example, for the pandemic of 1918, where the influenza virus strain, H1N1, was shown to be responsible, a peak concentration of the Replikins in H1N1 independently occurred (P1); for the pandemic of 1957, where H2N2 emerged and was shown to be responsible, a peak concentration of the Replikins in H2N2 occurred (P2); and for the pandemic of 1968, where H3N2 emerged and was shown to be the cause of the pandemic, a peak concentration of the Replikins in H3N2 occurred (P3). Third, in the years immediately following each of the above three pandemics, the specific Replikin concentration decreased markedly, perhaps reflecting the broadly distributed immunity generated in each case. Thus, this post-pandemic decline is specific for H1N1 immediately following the pandemic (P1) for which it was responsible, and is not a general property of all strains at the time. An increase of Replikin concentration in influenza B repeatedly occurred simultaneously with the decrease in Replikin concentration in H1N1, e.g., EB1 in 1951 and EB2 in 1976, both associated with influenza B epidemics having the highest mortality. (Stuart-Harris, et al., Edward Arnold Ltd. (1985). Fourth, a secondary peak concentration, which exceeded the primary peak increase in concentration, occurred 15 years after each of the three pandemics, and this secondary peak was accompanied by an epidemic: 15 years after the 1918 pandemic in an H1N1 `epidemic` year (E1); eight years after the 1957 pandemic in an H2N2 `epidemic` year (E2); and occurred seven years after the 1968 pandemic in an H3N2 `epidemic` year (E3). These secondary peak concentrations of specific Replikins may reflect recovery of the strain. Fifth, peaks of each strain's specific Replikin concentration frequently appear to be associated with declines in Replikin concentration of one or both other strains, suggesting competition between strains for host sites. Sixth, there is an apparent overall tendency for the Replikin concentration of each strain to decline over a period of 35 years (H2N2) to 60 years (influenza B). This decline cannot be ascribed to the influence of vaccines because it was evident in the case of influenza B from 1901 to 1964, prior to common use of influenza vaccines. In the case of influenza B, Replikin recovery from the decline is seen to occur after 1965, but Replikin concentration declined again between 1997 and 2000 (FIG. 7). This correlates with the low occurrence of influenza B in recent case isolates. H1N1 Replikin concentration peaked in 1978-1979 (FIG. 7) together with the reappearance and prevalence of the H1N1 strain, and then peaked in 1996 coincident with an H1N1 epidemic. (FIG. 7). H1N1 Replikin concentration also declined between 1997 and 2000, and the presence of H1N1 strains decreased in isolates obtained during these years. For H2N2 Replikins, recovery from a 35 year decline has not occurred (FIG. 8), and this correlates with the absence of H2N2 from recent isolates. For H3N2, the Replikin concentration of many isolates fell to zero during the period from 1996 to 2000, but other H3N2 isolates showed a significant, sharp increase in Replikin concentration. This indicates the emergence of a substrain of H3N2, which is designated herein as H3N2(R).
[0249]FIGS. 7 and 8 demonstrate that frequently, a one to three year stepwise increase is observed before Replikin concentration reaches a peak. This stepwise increase proceeds the occurrence of an epidemic, which occurs concurrently with the Replikin peak. Thus, the stepwise increase in concentration of a particular strain is a signal that particular strain is the most likely candidate to cause an epidemic or pandemic.
[0250]Currently, Replikin concentration in the H3N2(R) strain of influenza virus is increasing (FIG. 8, 1997 to 2000). Three similar previous peak increases in H3N2 Replikin concentration are seen to have occurred in the H3N2-based pandemic of 1968 (FIG. 8), when the strain first emerged, and in the H3N2-based epidemics of 1972 and 1975 (FIG. 8). Each of these pandemic and epidemics was associated with excess mortality. (Ailing, et al., Am J. Epidemiol., 113(1):30-43 (1981). The rapid ascent in concentration of the H3N2(R) subspecies of the H3N2 Replikins in 1997-2000, therefore, statistically represents an early warning of an approaching severe epidemic or pandemic. An H3N2 epidemic occurred in Russia in 2000 (FIG. 8, E4); and the CDC report of December 2001 states that currently, H3N2 is the most frequently isolated strain of influenza virus world wide. (Morbidity and Mortality Weekly Reports (MMWR), Center for Disease Control; 50(48):1084-68 (Dec. 7, 2001).
[0251]In each case of influenza virus pandemic or epidemic new Replikins emerge. There has been no observation of two of the same Replikins in a given hemagglutinin in a given isolate. To what degree the emergence of a new Replikin represents mutations versus transfer from another animal or avian pool is unknown. In some cases, each year one or more of the original Replikin structures is conserved, while at the same time, new Replikins emerge. For example, in influenza virus B hemagglutinin, five Replikins were constantly conserved between 1919 and 2001, whereas 26 Replikins came and went during the same period (some recurred after several years absence). The disappearance and re-emergence years later of a particular Replikin structure suggests that the Replikins return from another virus host pool rather than through de novo mutation.
[0252]In the case of II1N1 Replikins, the two Replikins present in the P1 peak associated with the 1918 pandemic were not present in the recovery E1 peak of 1933, which contains 12 new Replikins. Constantly conserved Replikins, therefore, are the best choice for vaccines, either alone or in combination. However, even recently appearing Replikins accompanying one year's increase in concentration frequently persist and increase further for an additional one or more years, culminating in a concentration peak and an epidemic, thus providing both an early warning and time to vaccinate with synthetic Replikins (see for example, H1N1 in the early 1990's, FIG. 7).
[0253]The data in FIGS. 7 and 8 demonstrate a direct relationship between the presence and concentration of a particular Replikin in influenza protein sequences and the occurrence of pandemics and epidemics of influenza. Thus, analysis of the influenza virus hemagglutinin protein sequence for the presence and concentration of Replikins provides a predictor of influenza pandemics and/or epidemics, as well as a target for influenza vaccine formulation.
[0254]Composition of Replikins in Strains of Influenza Virus B: Of a total of 26 Replikins identified in this strain (Table 3), the following ten Replikins are present in every influenza B isolate examined from 1902-2001. Overlapping Replikin sequences are listed separately. Lysines and histidines are in bold type to demonstrate homology consistent with the "3-point recognition."
TABLE-US-00008 kshfanlk (SEQ ID NO. 91) kshfanlkgtk (SEQ ID NO. 92) kshfanlkgtktrgklcpk (SEQ ID NO. 93) hekygglnk (SEQ ID NO. 94) hekygglnksk (SEQ ID NO. 95) hekygglnkskpyytgehak (SEQ ID NO. 96) hakaigncpiwvk (SEQ ID NO. 97) hakaigncpiwvvkktplklangtk (SEQ ID NO. 98) hakaigncpiwvktplklangtkyrppak (SEQ ID NO. 99) hakaigncpiwvktplklangtkyrppakllk (SEQ ID NO. 100)
[0255]Tables 3 and 4 indicate that there appears to be much greater stability of the Replikin structures in influenza B hemagglutinins compared with H1N1 Replikins. Influenza B has not been responsible for any pandemic, and it appears not to have an animal or avian reservoirs. (Stuart-Harris et al., Edward Arnold Ltd., London (1985)).
[0256]Influenza H1N1 Replikins: Only one Replikin "hp(v/i)tigecpkyv(r/k)(s/t)(t/a)k" is present in every H1N1 isolate for which sequences are available from 1918, when the strain first appeared and caused the pandemic of that year, through 2000 (Table 4) ("(v/i)" indicates that the amino acid v or i is present in the same position in different years.) Although H1N1 contains only one persistent Replikin, H1N1 appears to be more prolific than influenza B. There are 95 different Replikin structures in 82 years on H1N1 versus only 31 different Replikins in 100 years of influenza B isolates (Table 4). An increase in the number of new Replikin structures occurs in years of epidemics (Tables 3, 4, 5 and 6) and correlates with increased total Replikin concentration (FIGS. 7 and 8).
[0257]Influenza H2N2 Replikins: Influenza H2N2 was responsible for the human pandemic of 1957. Three of the 20 Replikins identified in that strain for 1957 were conserved in each of the H2N2 isolates available for examination on PubMed until 1995 (Table 5).
TABLE-US-00009 (SEQ ID NO. 219) ha(k/q/m)(d/n)ilekthngk (SEQ ID NO. 220) ha(k/q/m)(d/n)ilekthngklc(k/r) (SEQ ID No. 225) kgsnyp(v/i)ak(g/r)synntsgeqmliiwq(v/i)h
[0258]However, in contrast to H1N1, only 13 additional Replikins have been found in H2N2 beginning in 1961. This paucity of appearance of new Replikins correlates with the decline in the concentration of the H2N2 Replikins and the appearance of H2N2 in isolates over the years (FIG. 8).
[0259]Influenza H3N2 Replikins: Influenza H3N2 was responsible for the human pandemic of 1968. Five Replikins which appeared in 1968 disappeared after 1977, but reappeared in the 1990s (Table 6). The only Replikin structure which persisted for 22 years was hcd(g/q)f(q/r)nekwdlf(v/i)er(s/t)k, which appeared first in 1977 and persisted through 1998. The emergence of twelve new H3N2 Replikins in the mid 1990s (Table 6) correlates with the increase in Replikin concentration at the same time (FIG. 8), and with the prevalence of the H3N2 strain in recent isolates together with the concurrent disappearance of all Replikins from some of these isolates (FIG. 8), this suggests the emergence of the new substrain H3N2(R).
[0260]FIGS. 1 and 2 show that influenza epidemics and pandemics correlate with the increased concentration of Replikins in influenza virus, which is due to the reappearance of at least one Replikin from one to 59 years after its disappearance. Also, in the A strain only, there is an emergence of new strain-specific Replikin compositions (Tables 4-6). Increase in Replikin concentration by repetition of individual Replikins within a single protein appears not to occur in influenza virus, but is seen in other organisms.
[0261]It has been believed that changes in the activity of different influenza strains are related to sequence changes in influenza hemagglutinins, which in turn are the products of substitutions effected by one of two poorly understood processes: i) antigenic drift, thought to be due to the accumulation of a series of point mutations in the hemagglutinin molecule, or ii) antigenic shift, in which the changes are so great that genetic reassortment is postulated to occur between the viruses of human and non-human hosts. First, the present data suggests that the change in activity of different influenza strains, rather than being related to non-specific sequence changes, are based upon, or relate to the increased concentration of strain-specific Replikins and strain-specific increases in the replication associated with epidemics. In addition, the data were examined for a possible insight into which sequence changes are due to "drift" or "shift", and which are due to conservation, storage in reservoirs, and reappearance. The data show that the epidemic-related increase in Replikin concentration is not due to the duplication of existing Replikins per hemagglutinin, but is due to the reappearance of at least one Replikin composition from 1 to up to 59 years after its disappearance, plus in the A strains only, the emergence of new strain-specific Replikin compositions (Tables 3-6). Thus the increase in Replikin concentration in the influenza B epidemics of 1951 and 1977 are not associated with the emergence of new Replikin compositions in the year of the epidemic but only with the reappearance of Replikin compositions which had appeared in previous years then disappeared (Table 3). In contrast, for the A strains, in addition to the reappearance of previously disappeared virus Replikins, new compositions appear (e.g. in H1N1 in the year of the epidemic of 1996, in addition to the reappearance of 6 earlier Replikins, 10 new compositions emerged). Since the A strains only, not influenza B, have access to non-human animal and avian reservoirs, totally new compositions probably derive from non-human host reservoirs rather than from mutations of existing human Replikins which appear to bear no resemblance to the new compositions other than the basic requirements of "3-point recognition" (Tables 2-5). The more prolific nature of H1N1 compared with B, and the fact that pandemics have been produced by the three A strains only, but not by the B strain, both may also be a function of the ability of the human A strains to receive new Replikin compositions from non-human viral reservoirs.
[0262]Some Replikins have appeared in only one year, disappeared, and not reappeared to date (Tables 3-6). Other Replikins disappear from one to up to 81 years, when the identical Replikin sequence reappears. Key Replikin `k` and `h` amino acids, and the spaces between them, are conserved during the constant presence of particular Replikins over many years, as shown in Tables 23-6 for the following strain-specific Replikins: ten of influenza B, the single Replikin of H1N1, and the single Replikin of H2N3, as well as for the reappearance of identical Replikins after an absence. Despite the marked replacement or substitution activity of other amino acids both inside the Replikin structure and outside it in the rest of the hemagglutinin sequences, influenza Replikin histidine (h) appears never to be, and lysine (k) is rarely replaced. Examples of this conservation are seen in the H1N1 Replikin "hp(v/i)tigecpkyv(r/k)(s/t)(t/a)k," (SEQ ID NO. 122) constant between 1918 and 2000, in the H3N2 Replikin "hcd(g/q)f(q,r)nekwdlf(v/i)er(s/t)k" (SEQ ID NO. 264) constant between 1975 and 1998 and in the H3N2 Replikin "hqn(s/e)(e/q)g(t/s)g(q/y)aad(1/q)kstq(a/n)a(i/l)d(q/g)l(n/t)(g/n)k(1/v)n- (r/s) vi(e/c)k" (SEQ ID NO. 263) which first appeared in 1975, disappeared for 25 years, and then reappeared in 2000. While many amino acids were substituted, the basic Replikin structure of 2 Lysines, 6 to 10 residues apart, one histidine, a minimum of 6% lysine in not more than approximately 50 amino acids, was conserved.
[0263]Totally random substitution would not permit the persistence of these H1N1 and H3N2 Replikins, nor from 1902 to 2001 in influenza B the persistence of 10 Replikin structures, nor the reappearance in 1993 of a 1919 18 mer Replikin after an absence of 74 years. Rather than a random type of substitution, the constancy suggests an orderly controlled process, or in the least, protection of the key Replikin residues so that they are fixed or bound in some way: lysines, perhaps bound to nucleic acids, and histidines, perhaps bound to respiratory redox enzymes. The mechanisms which control this conservation are at present unknown.
Conservation of Replikin Structures
[0264]Whether Replikin structures are conserved or are subject to extensive natural mutation was examined by scanning the protein sequences of various isolates of foot and mouth disease virus (FMDV), where mutations in proteins of these viruses have been well documented worldwide for decades. Protein sequences of FMDV isolates were visually examined for the presence of both the entire Replikin and each of the component Replikin amino acid residues observed in a particular Replikin.
[0265]Rather than being subject to extensive substitution over time as occurs in neighboring amino acids, the amino acids which comprise the Replikin structure are substituted little or not at all, that is the Replikin structure is conserved.
[0266]For example, in the protein VP1 of FMDV type O, the Replikin (SEQ ID NO.: 3) "hkqkivapvk" was found to be conserved in 78% of the 236 isolates reported in PubMed, and each amino acid was found to be conserved in individual isolates as follows: his, 95.6%; lys, 91.8%; gin 92.3%; lys, 84.1%; ile, 90.7%; val, 91.8%; ala, 97.3%; pro, 96.2%; ala, 75.4%; and lys, 88.4%. The high rate of conservation suggests structural and functional stability of the Replikin structure and provides constant targets for treatment.
[0267]Similarly, sequence conservation was found in different isolates of HIV for its Replikins, such as (SEQ ID NO.: 5) "kcfncgkegh" or (SEQ ID NO.: 6) "kvylawvpahk" in HIV Type 1 and (SEQ ID NO.: 7) "kcwncgkegh" in HIV Type 2 (Table 2). Further examples of sequence conservation were found in the HIV tat proteins, such as (SEQ ID NO.: 698) "hclvckqkkglgisygrkk," wherein the key lysine and histidine amino acids are conserved (See Table 7).
[0268]Similarly, sequence conservation was observed in plants, for example in wheat, such as in wheat ubiguitin activating enzyme E (SEQ ID NOs. 601-603). The Replikins in wheat even provided a reliable target for stimulation of plant growth as described within. Other examples of conservation are seen in the constant presence of malignin in successive generations, over ten years of tissue culture of glioma cells, and by the constancy of affinity of the glioma Replikin for antimalignin antibody isolated by immunoadsorption from 8,090 human sera from the U.S., U.K., Europe and Asia (e.g., FIG. 5 and U.S. Pat. No. 6,242,578 B1).
[0269]Similarly, conservation was observed in trans-activator (Tat) proteins in isolates of HIV. Tat (trans-activator) proteins are early RNA binding proteins regulating lentiviral transcription. These proteins are necessary components in the life cycle of all known lentiviruses, such as the human immunodeficiency viruses (HIV). Tat is a transcriptional regulator protein that acts by binding to the trans-activating response sequence (TAR) RNA element and activates transcription Initiation and/or elongation from the LTR promoter. HIV cannot replicate without tat, but the chemical basis of this has been unknown. In the HIV tat protein sequence from 89 to 102 residues, we have found a Replikin that is associated with rapid replication in other organisms. The amino acid sequence of this Replikin is "hclvcfqkkglgisygrkk." In fact, we found that this Replikin is present in every HIV tat protein. Some tat amino acids are substituted frequently, as shown in Table 8, by alternate amino acids (in small size fonts lined up below the most frequent amino acid (Table 7), the percentage of conservation for the predominant Replikin "hclvcfqkkglgisygrkk"). These substitutions have appeared for most of the individual amino acids. However, the key lysine and histidine amino acids within the Replikin sequence, which define the Replikin structure, are conserved 100% in the sequence; while substitutions are common elsewhere in other amino acids, both within and outside the Replikin, none occurs on these key histidine amino acids.
[0270]As shown in Table 7, it is not the case that lysines are not substituted in the tat protein amino acid sequence. From the left side of the table, the very first lysine in the immediate neighboring sequence, but outside the Replikin sequence, and the second lysine (k) in the sequence inside the Replikin, but "extra" in that it is not essential for the Replikin formation, are both substituted frequently. However, the 3rd, 4th and 5th lysines, and the one histidine, in parentheses, which together set up the Replikin structure, are never substituted. Thus, these key amino acid sequences are 100% conserved. As observed in the case of the influenza virus Replikins, random substitution would not permit this selective substitution and selective non-substitution to occur due to chance.
TABLE-US-00010 TABLE 7 % Replikin CONSERVATION of each constituent amino acid in the first 117 different isolates of HIV tat protein as reported in PubMed: 38 100 57 86 100 100 66 76 100 99 57 49 100 94 100 97 98 85 97 99 100 100 100% Neighboring-Amino acids [tat Replikin] k (c) s y [(h) (c) l v (c) f q k (k) g (I) g i s y g (r) (k) (k)] below are the amino acid substitutions observed for each amino acid above: h c f q i l h t a a l y h q r w p l l i h q v y s s l m r s i s m s s r n v a f p q
[0271]The conservation of the Replikin structure suggests that the Replikin structure has a specific survival function for the HIV virus which must be preserved and conserved, and cannot be sacrificed to the virus `defense` maneuver of amino acid substitution crested to avoid antibody and other `attack.` These `defense` functions, although also essential, cannot `compete` with the virus survival function of HIV replication.
[0272]Further conservation was observed in different isolates of HIV for its Replikins such as "kcfncgkegh" (SEQ ID NO. 5) or "kvylawvpahk" (SEQ ID NO. 6) in HIV Type 1 and "kcwncgkegh" (SEQ ID NO. 7) in HIV Type 2.
[0273]The high rate of conservation observed in FMVD and HIV Replikins suggests that conservation also observed in the Replikins of influenza Replikins is a general property of viral Replikins. This conservation makes them a constant and reliable tarted for either destruction, for example by using specific Replikins such as for influenza, FMVD or HIV vaccines as illustrated for the glioma Replikin, or stimulation.
[0274]Similarly, as provided in examples found in viruses including influenza viruses, FMDV, and HIV, where high rates of conservation in Replikins suggest that conservation is a general property of viral Replikins and thus making Replikins a constant and reliable target for destruction or stimulation, conservation of Replikin structures occurs in plants. For example, in wheat plants, Replikins are conserved and provide a reliable target for stimulation. Examples of conserved Replikins in wheat plants ubiquitin activating enzyme E include:
TABLE-US-00011 E3 hkdrltkkvvdiarevakvdvpeyrrh (SEQ ID NO. 601) E2 hkerldrkvvdvarevakvevpsyrrh (SEQ ID NO. 602) E1 hkerldrkvvdvarevakmevpsyrrh (SEQ ID NO. 603) * * * ** *
[0275]Similarly to conservation found in the HIV tat protein, the Replikin in the wheat ubiquitin activating enzyme E is conserved. As with the HIV tat protein, substitutions of amino acids (designated with an `*`) adjacent to the Replikin variant forms in wheat ubiquitin activating enzyme E are common. The key k and h amino acids that form the Replikin structure, however, do not vary whereas the `unessential` k that is only 5 amino acids (from the first k on the left) is substituted.
Anti-Replikin Antibodies
[0276]An anti-Replikin antibody is an antibody against a Replikin. Data on anti-Replikin antibodies also support Replikin class unity. An anti-Replikin antibody response has been quantified by immunoadsorption of serum antimalignin antibody to immobilized malignin (see Methods in U.S. Pat. No. 5,866,690). The abundant production of antimalignin antibody by administration to rabbits of the synthetic version of the 16-mer peptide whose sequence was derived from malignin, absent carbohydrate or other groups, has established rigorously that this peptide alone is an epitope, that is, provides a sufficient basis for this immune response (FIG. 3). The 16-mer peptide produced both IgM and IgG forms of the antibody. Antimalignin antibody was found to be increased in concentration in serum in 37% of 79 cases in the U.S. and Asia of hepatitis B and C, early, in the first five years of infection, long before the usual observance of liver cancer, which develops about fifteen to twenty-five years after infection. Relevant to both infectious hepatitis and HIV infections, transformed cells may be one form of safe haven for the virus: prolonging cell life and avoiding virus eviction, so that the virus remains inaccessible to anti-viral treatment.
[0277]Because administration of Replikins stimulates the immune system to produce antibodies having a cytotoxic effect, peptide vaccines based on the particular influenza virus Replikin or group of Replikins observed to be most concentrated over a given time period provide protection against the particular strain of influenza most likely to cause an outbreak in a given influenza season, e.g., an emerging strain or re-emerging strain For example, analysis of the influenza virus hemagglutinin amino acid sequence on a yearly or bi-yearly basis, provides data which are useful in formulating a specifically targeted influenza vaccine for that year. It is understood that such analysis may be conducted on a region-by-region basis or at any desired time period, so that strains emerging in different areas throughout the world can be detected and specifically targeted vaccines for each region can be formulated.
Influenza
[0278]Currently, vaccine formulations are changed twice yearly at international WHO and CDC meetings. Vaccine formulations are based on serological evidence of the most current preponderance of influenza virus strain in a given region of the world. However, prior to the present invention there has been no correlation of influenza virus strain specific amino acid sequence changes with occurrence of influenza epidemics or pandemics.
[0279]The observations of specific Replikins and their concentration in influenza virus proteins provides the first specific quantitative early chemical correlates of influenza pandemics and epidemics and provides for production and timely administration of influenza vaccines tailored specifically to treat the prevalent emerging or re-emerging strain of influenza virus in a particular region of the world. By analyzing the protein sequences of isolates of strains of influenza virus, such as the hemagglutinin protein sequence, for the presence, concentration and/or conservation of Replikins, influenza virus pandemics and epidemics can be predicted. Furthermore, the severity of such outbreaks of influenza can be significantly lessened by administering an influenza peptide vaccine based on the Replikin sequences found to be most abundant or shown to be on the rise in virus isolates over a given time period, such as about one to about three years.
[0280]An influenza peptide vaccine of the invention may include a single Replikin peptide sequence or may include a plurality of Replikin sequences observed in influenza virus strains. Preferably, the peptide vaccine is based on Replikin sequence(s) shown to be increasing in concentration over a given time period and conserved for at least that period of time. However, a vaccine may include a conserved Replikin peptide(s) in combination with a new Replikin(s) peptide or may be based on new Replikin peptide sequences. The Replikin peptides can be synthesized by any method, including chemical synthesis or recombinant gene technology, and may include non-Replikin sequences, although vaccines based on peptides containing only Replikin sequences are preferred. Preferably, vaccine compositions of the invention also contain a pharmaceutically acceptable carrier and/or adjuvant.
[0281]The influenza vaccines of the present invention can be administered alone or in combination with antiviral drugs, such as gancyclovir; interferon; interleukin; M2 inhibitors, such as, amantadine, rimantadine; neuraminidase inhibitors, such as zanamivir and oseltamivir; and the like, as well as with combinations of antiviral drugs.
Replikin Decoys in Malaria
[0282]Analysis of the primary structure of a Plasmodium falciparum malaria antigen located at the merozoite surface and/or within the parasitophorous vacuole revealed that this organism, like influenza virus, also contains numerous Replikins (Table 8). However, there are several differences between the observation of Replikins in Plasmodium falciparum and influenza virus isolates. For example, Plasmodium falciparum contains several partial Replikins, referred to herein as "Replikin decoys." These decoy structures contain an abundance of lysine residues, but lack the histidine required of Replikin structures. Specifically, these decoys contain many lysines 6 to 10 residues apart in overlapping fashion, similar to the true malaria recognins but without histidine residues. It is believed that the decoy structure maximizes the chances that an anti-malarial antibody or other agent will bind to the relatively less important structure containing the lysines, i.e., the Replikin decoys, rather than binding to histidine, which is present in Replikin structure, such as Replikins in respiratory enzymes, which could result in destruction of the trypanosome. For example, an incoming antibody, with specificity for Replikin structures, might attach to the Replikin decoy structure, leaving the true Replikin structure remains untouched.
[0283]Therefore, anti-Replikin treatment of malaria requires two phases (dual treatment): i) preliminary treatment with proteolytic enzymes that cleave the Replikin decoys, permitting `safe passage` of the specific anti-Replikin treatment; and ii) attacking malaria Replikins either with specific antibodies or by cellular immunity engendered by synthetic malaria Replikin vaccines or by organic means targeting the malaria Replikins.
Repetition and Overlapping of Replikin Structures
[0284]Another difference seen in Plasmodium falciparum is a frequent repetition of individual Replikin structures within a single protein, which was not observed with influenza virus. Repetition may occur by (a) sharing of lysine residues between Replikins, and (b) by repetition of a portion of a Replikin sequence within another Replikin sequence.
[0285]A third significant difference between Replikin structures observed in influenza virus isolates and Plasmodium falciparum is a marked overlapping of Replikin structures throughout malarial proteins, e.g., there are nine overlapping Replikins in the 39 amino acid sequence of SEQ ID NO. 380 (Replikin concentration=23.1/100 amino acids); and 15 overlapping Replikins in the 41 amino acids of SEQ ID NO. 454 (Replikin concentration=36.6/100 amino acids). Both of these overlapping Replikin structures occur in blood stage trophozoites and schizonts. In contrast, influenza virus Replikins are more scattered throughout the protein and the maximum Replikin concentration is about 7.5/100 amino acids (FIG. 7); and tomato leaf curl gemini virus, which was also observed to have overlapping Replikins has only about 3.1/100 amino acids.
[0286]This mechanism of lysinc multiples is also seen in the Replikins of cancer proteins such as in gastric cancer transforming protein, ktkkgnrvsptmkvth (SEQ ID NO. 88), and in transforming protein P21B (K-RAS 2B) of lung, khkekmskdgkkkkkks (SEQ ID NO. 89).
[0287]The relationship of higher Replikin concentration to rapid replication is also confirmed by analysis of HIV isolates. It was found that the slow-growing low titer strain of HIV (NS, "Bru," which is prevalent in early stage HIV infection) has a Replikin concentration of 1.1 (+/-1.6) Replikins per 100 amino acids, whereas the rapidly-growing high titer strain of HIV (SI, "Lai", which is prevalent in late stage HIV infection) has a Replikin concentration of 6.8 (+/-2.7) Replikins per 100 amino acid residues.
[0288]The high concentration of overlapping Replikins in malaria, influenza virus and cancer cells is consistent with the legendary high and rapid replicating ability of malaria organisms. The multitude of overlapping Replikins in malaria also provides an opportunity for the organism to flood and confuse the immune system of its host and thereby maximize the chance that the wrong antibody will be made and perpetuated, leaving key malaria antigens unharmed.
[0289]As in the case of influenza virus, for example, peptide vaccines based on the Replikin structure(s) found in the malaria organism can provide an effective means of preventing and/or treating malaria. Vaccination against malaria can be achieved by administering a composition containing one or a mixture of Replikin structures observed in Plasmodium falciparum. Furthermore, antibodies to malaria Replikins can be generated and administered for passive immunity or malaria detection
[0290]Table 8 provides a list of several Plasmodium falciparum Replikin sequences. It should be noted that this list is not meant to be complete. Different isolates of the organism may contain other Replikin structures.
TABLE-US-00012 TABLE 8 Malaria Replikins a) Primary structure of a Plasmodium falciparum malaria antigen located at the merozoite surface and within the parasitophorous vacuole a) i) DECOYS: (C-Terminal) keeeekekekekekeekekeekekeekekekeekekekeekeeekk, (SEQ ID NO. 280) or keeeekekekekekeekekeekekeekekekeekekekeekeeekkek, (SEQ ID NO. 281) or keeeekekekekekeekekeekekekeekekeekekeekeekeeekk, (SEQ ID NO. 282) or keeeekekek (SEQ ID NO. 283) ii) ReplikinS: Hkklikalkkniesiqnkk (SEQ ID NO. 284) hkklikalkkniesiqnkm (SEQ ID NO. 285) hkklikalkk (SEQ ID NO. 286) hkklikalk (SEQ ID NO. 287) katysfvntkkkiislksqghkk (SEQ ID NO. 288) katysfvntkkkiislksqghk (SEQ ID NO. 289) katysfvntkkkiislksqgh (SEQ ID NO. 290) htyvkgkkapsdpqca dikeeckellkek (SEQ ID NO. 291) kiislksqghk (SEQ ID NO. 292) kkkkfeplkngnvsetiklih (SEQ ID NO. 293) kkkfeplkngnvsetiklih (SEQ ID NO. 294) kkfeplkngnvsetiklih (SEQ ID NO. 295) kngnvsetiklih (SEQ ID NO. 296) klihlgnkdkk (SEQ ID NO. 297) kvkkigvtlkkfeplkngnvsetiklihlgnkdkkh (SEQ ID NO. 298) hliyknksynplllscvkkmnmlkenvdyiqnqnlfkelmnqkatysfvntkkkiislk (SEQ ID NO. 299) hliyknksynplllscvkkmnmlkcnvdyiqnqnlfkelmnqkatysfvntk (SEQ ID NO. 300) hliyknksynplllscvkkmnmlkenvdyiqnqnlfkelmnqk (SEQ ID NO. 301) hliyknksynplllscvkkmnmlkenvdyiqknqnlfk (SEQ ID NO. 302) hliyknksynplllscvkkmnmlk (SEQ ID NO. 303) ksannsanngkknnaeemknlvnflqshkklikalkkniesiqnkkh (SEQ ID NO. 304) kknnaeemknlvnflqshkklikalkkniesiqnkkh (SEQ ID NO. 305) knlvnflqshkklikalkkniesiqnkkh (SEQ ID NO. 306) kklikalkkniesiqnkkh (SEQ ID NO. 307) klikalkkniesiqnkkh (SEQ ID NO. 308) kkniesiqnkkh (SEQ ID NO. 309) kniesiqnkkh (SEQ ID NO. 310) knnaeemknlvnflqsh (SEQ ID NO. 311) kklikalkkniesiqnkkqghkk (SEQ ID NO. 312) kknnaeemknlvnflqshk (SEQ ID NO. 313) knnaeemknlvnflqsh (SEQ ID NO. 314) klikalkkniesiqnkkqghkk (SEQ ID NO. 315) kvkkigvtlkkfeplkngnvsetiklih (SEQ ID NO. 316) kngnvsetiklih (SEQ ID NO. 317) klihlgnkdkk (SEQ ID NO. 318) ksannsanngkknnaeemknlvnflqsh (SEQ ID NO. 319) kknnaeemknlvnflqsh (SEQ ID NO. 320) kklikalkkniesiqnkkh (SEQ ID NO. 321) kalkkniesiqnkkh (SEQ ID NO. 322) kkniesiqnkkh (SEQ ID NO. 323) kelmnqkatysfvntkkkiislksqgh (SEQ ID NO. 324) ksqghkk (SEQ ID NO. 325) kkkiislksqgh (SEQ ID NO. 326) kkiislksqgh (SEQ ID NO. 327) kkniesiqnkkh (SEQ ID NO. 328) kniesiqnkkh (SEQ ID NO. 329) htyvkgkkapsdpqcadikeeckellkek (SEQ ID NO. 330) htyvkgkkapsdpqcadikeeckellk (SEQ ID NO. 331) b) "liver stage antigen-3" gene = "LSA-3" Replikins henvlsaalentqseeekkevidvieevk (SEQ ID NO. 332) kcnvvttilekveettaesvttfsnileeiqentitndtieekleelh (SEQ ID NO. 333) hylqqmkekfskek (SEQ ID NO. 334) hylqqmkekfskeknnnvievtnkaekkgnvqvtnktekttk (SEQ ID NO. 335) hylqqmkekfskeknnnvievtnkaekkgnvqvtnktekttkvdknnk (SEQ ID NO. 336) hylqqmkekfskeknnnvievtnkaekkgnvqvtnktekttkvdknnkvpkkrrtqk (SEQ ID NO. 337) hylqqmkekfskeknnnvievtnkaekkgnvqvtnktekttkvdknnkvpkkrrtqksk (SEQ ID NO. 338) hvdevmkyvqkidkevdkevskaleskndvtnvlkqnqdffskvknfvkkyk (SEQ ID NO. 339) hvdevmkyvqkidkevdkevskaleskndvtnvlkqnqdffskvknfvkk (SEQ ID NO. 340) hvdevmkyvqkidkevdkevskaleskndvtnvlkqnqdffsk (SEQ ID NO. 341) hvdevmkyvqkidkevdkevskaleskndvtnvlk (SEQ ID NO. 342) hvdevmkyvqkidkevdkevskalesk (SEQ ID NO. 343) hvdevmkyvqkidkevdkevsk (SEQ ID NO. 344) hvdevmkyvqkidkevdk (SEQ ID NO. 345) hvdevmkyvqkidk (SEQ ID NO. 346) kdevidlivqkekriekvkakkkklekkveegvsglkkh (SEQ ID NO. 347) kvkakkkklckkveegvsglkkh (SEQ ID NO. 348) kakkkklekkvecgvsglkkh (SEQ ID NO. 349) kkkklekkveegvsglkkh (SEQ ID NO. 350) kkklekkveegvsglkkh (SEQ ID NO. 351) kklekkveegvsglkkh (SEQ ID NO. 352) klekkveegvsglkkh (SEQ ID NO. 353) kkveegvsglkkh (SEQ ID NO. 354) kveegvsglkkh (SEQ ID NO. 355) hveqnvyvdvdvpamkdqflgilneagglkemffnledvfksesdvitveeikdepvqk (SEQ ID NO. 356) hikgleeddleevddlkgsildmlkgdmelgdmdkesledvttklgerveslk (SEQ ID NO. 357) hikgleeddleevddlkgsildmlkgdmelgdmdkesledvttk (SEQ ID NO. 358) hikgleeddleevddlkgsildmlkgdmelgdmdk (SEQ ID NO. 359) hikgleeddleevddlkgsildmlk (SEQ ID NO. 360) hiisgdadvlssalgmdccqmktrkkaqrpk (SEQ ID NO. 361) hditttldevvelkdveedkick (SEQ ID NO. 362) kkleevhelk (SEQ ID NO. 363) kleevhelk (SEQ ID NO. 364) ktietdileekkkeiekdh (SEQ ID NO. 365) kkeiekdhfek (SEQ ID NO. 366) kdhfek (SEQ ID NO. 367) kfeeeaeeikh (SEQ ID NO. 368) c) 28 KDA ookinete surface antigen precursor Replikins: kdgdtkctlecaqgkkcikhksdhnhksdhnhksdpnhkkknnnnnk (SEQ ID NO. 369) kdgdtkctlecaqgkkcikhksdhnhksdhnhksdpnhkk (SEQ ID NO. 370) kdgdtkctlecaqgkkcikhksdhnhksdhnhksdpnhk (SEQ ID NO. 371) kdgdtkctlecaqgkkcikhksdhnhksdhnhk (SEQ ID NO. 372) kdgdtkctlecaqgkkcikhksdhnhk (SEQ ID NO. 373) kdgdtkctlecaqgkkcikhk (SEQ ID NO. 374) kdgdtkctleeaqgkk (SEQ ID NO. 375) kdgdtkctlecaqgk (SEQ ID NO. 376) kciqaecnykecgeqkcvwdgih (SEQ ID NO. 377) kecgeqkcvwdgih (SEQ ID NO. 378) hieckcnndyvltnryecepknkctsledtnk (SEQ ID NO. 379) d) Blood stage trophozoites and schizonts Replikins: ksdhnhksdhnhksdhnhksdhnhksdpnhkkknnnnnk (SEQ ID NO. 380) ksdhnhksdhnhksdhnhksdphnkkknnnnnk (SEQ ID NO. 381) ksdhnhksdhnhksdpnhkkknnnnnk (SEQ ID NO. 382) ksdhnhksdpnhkkknnnnnk (SEQ ID NO. 383) kkknnnnnkdnksdpnhk (SEQ ID NO. 384) kknnnnnkdnksdpnhk (SEQ ID NO. 385) knnnnnkdnksdpnhk (SEQ ID NO. 386) kdnksdpnhk (SEQ ID NO. 387) ksdpnhk (SEQ ID NO. 388) hslyalqqneeyqkvknekdqneikkikqlieknk (SEQ ID NO. 389) hslyalqqneeyqkvknekdqneikkik (SEQ ID NO. 390) hslyalqqneeyqkvknekdqneikk (SEQ ID NO. 391) hslyalqqneeyqkvknekdqneik (SEQ ID NO. 392) hklenleemdk (SEQ ID NO. 393) khfddntneqk (SEQ ID NO. 394) kkeddekh (SEQ ID NO. 395) keennkkeddekh (SEQ ID NO. 396) ktssgilnkeennkkeddekh (SEQ ID NO. 397) knihikk (SEQ ID NO. 398) hikkkegidigyk (SEQ ID NO. 399) kkmwtcklwdnkgneitknih (SEQ ID NO. 400) kkgiqwnllkkmwtcklwdnkgneitknih (SEQ ID NO. 401) kekkdsnenrkkkqkedkknpnklkkieytnkithffkaknnkqqnnvth (SEQ ID NO. 402) kkdsnenrkkkqkedkknpnklkkieytnkithffkaknnkqqnnvth (SEQ ID NO. 403) kdsnenrkkkqkedkknpnklkkieytnkithffkaknnkqqnnvth (SEQ ID NO. 404) kkqkedkknpnklkkieytnkithffkaknnkqqnnvth (SEQ ID NO. 405) kqkedkknpnklkkieytnkithffkaknnkqqnnvth (SEQ ID NO. 406) kedkknpnklkkieytnkithffkaknnkqqnnvth (SEQ ID NO. 407) knpnklkkieytnkithffkaknnkqqnnvth (SEQ ID NO. 408) kkieytnkithffkaknnkqqnnvth (SEQ ID NO. 409) kieytnkithffkaknnkqqnnvth (SEQ ID NO. 410) kithffkaknnkqqnnvth (SEQ ID NO. 411) hknnedikndnskdikndnskdikndnskdikndnnedikndnskdik (SEQ ID NO. 412) hknnedikndnskdikndnskdikndnskdikndnnedikndnsk (SEQ ID NO. 413) hknnedikndnskdikndnskdikndnskdikndnnedik (SEQ ID NO. 414) hknnedikndnskdikndnskdikndnskdik (SEQ ID NO. 415) hknnedikndnskdikndnskdikndnsk (SEQ ID NO. 416) hknnedikndnskdikndnskdik (SEQ ID NO. 417) hknnedikndnskdikndnsk (SEQ ID NO. 418) hknnedikndnskdik (SEQ ID NO. 419) hknnedik (SEQ ID NO. 420) kkyddlqnkynilnklknsleekneelkkyh (SEQ ID NO. 421) kyddlqnkynilnklknsleekneelkkyh (SEQ ID NO. 422) kynilnklknsleekneelkkyh (SEQ ID NO. 423) klknsleekneelkkyh (SEQ ID NO. 424) knsleekneelkkyh (SEQ ID NO. 425) kneelkkyh (SEQ ID NO. 426) hmgnnqdinenvynikpqefkeeeeedismvntkk (SEQ ID NO. 427) knsnelkrindnffklh (SEQ ID NO. 428) kpclykkckisqclykkckisqvwwcmpvkdtfntyernnvlnskienniekiph (SEQ ID NO. 429) hinneytnknpkncllykneernyndnnikdyinsmnfkk (SEQ ID NO. 430) hinneytnknpkncllykneernyndnnikdyinsmnfk (SEQ ID NO. 431) hinneytnknpkncllyk (SEQ ID NO. 432) knktnqskgvkgeyekkketngh (SEQ ID NO. 433) ktnqskgvkgeyekkketngh (SEQ ID NO. 434) kgvkgeyekkketngh (SEQ ID NO. 435) kgeyekkketngh (SEQ ID NO. 436) ksgmytnegnkscecsykkkssssnkvh (SEQ ID NO. 437) kscecsykkkssssnkvh (SEQ ID NO. 438) kkkssssnkvh (SEQ ID NO. 439) kkssssnkvh (SEQ ID NO. 440) kssssnkvh (SEQ ID NO. 441) himlksgmytnegnkscecsykkkssssnk (SEQ ID NO. 442) himlksgmytnegnkscecsykkk (SEQ ID NO. 443) himlksgmytnegnkscecsykk (SEQ ID NO. 444) himlksgmytnegnkscecsyk (SEQ ID NO. 445) kplaklrkrektqinktkyergdviidnteiqkiiirdyhetlnvhkldh (SEQ ID NO. 446) krektqinktkyergdviidnteiqkiiirdyhetlnvhkldh (SEQ ID NO. 447) ktqinktkyergdviidnteiqkiiirdyhetlnvhkldh (SEQ ID NO. 448) kplaklrkrektqinktkyergdviidnteiqkiiirdyhetlnvh (SEQ ID NO. 449) kplaklrkrektqinktkyergdviidnteiqkiiirdyh (SEQ ID NO. 450) klrkrektqinktkyergdviidnteiqkiiirdyh (SEQ ID NO. 451) krektqinktkyergdviidnteiqkiiirdyh (SEQ ID NO. 452) ktqinktkyergdviidnteiqkiiirdyh (SEQ ID NO. 453) kkdkekkkdsnenrkkkqkedkknpndnklkkieytnkith (SEQ ID NO. 454) kdkekkkdsnenrkkkqkedkknpndnklkkieytnkith (SEQ ID NO. 455) kekkkdsnenrkkkqkedkknpndnklkkieytnkith (SEQ ID NO. 456) kkkdsnenrkkkqkedkknpndnklkkieytnkith (SEQ ID NO. 457) kkdsnenrkkkqkedkknpndnklkkieytnkith (SEQ ID NO. 458) kdsnenrkkkqkedkknpndnklkkieytnkith (SEQ ID NO. 459) kkkqkedkknpndnklkkieytnkith (SEQ ID NO. 460) kkqkedkknpndnklkkieytnkith (SEQ ID NO. 461) kqkedkknpndnklkkieytnkith (SEQ ID NO. 462) kcdkknpndnklkkieytnkith (SEQ ID NO. 463) kknpndnklkkieytnkith (SEQ ID NO. 464) knpndnklkkieytnkith (SEQ ID NO. 465) klkkieytnkith (SEQ ID NO. 466) kkieytnkith (SEQ ID NO. 467) kieytnkith (SEQ ID NO. 468) hgqikiedvnnenfnneqmknkyndeekmdiskskslksdflek (SEQ ID NO. 469) hgqikiedvnnenfnneqmknkyndeekmdiskskslk (SEQ ID NO. 470) hgqikiedvnnenfnneqmknkyndeekmdisksk (SEQ ID NO. 471) hgqikiedvnnenfnneqmknkyndeekmdisk (SEQ ID NO. 472) kkyddlqnkynilnklknsleekneelkkyh (SEQ ID NO. 473) kyddlqnkynilnklknsleekneelkkyh (SEQ ID NO. 474) kynilnklknsleekneelkkyh (SEQ ID NO. 475) klknsleekneelkkyh (SEQ ID NO. 476) knsleekneelkkyh (SEQ ID NO. 477) kneelkkyh (SEQ ID NO. 478) hmgnnqdinenvynikpqefkeeeeedismvntkkcddiqenik (SEQ ID NO. 479) ktnlyniynnknddkdnildnenreglylcdvmknsnelkrindnffklh (SEQ ID NO. 480) knsnelkrindnffklh (SEQ ID NO. 481) krindnffklh (SEQ ID NO. 482) hinneytnknpkncllykneernyndnnikdyinsmnfkk (SEQ ID NO. 483) hinneytnknpkncllykneernyndnnikdyinsmnfk (SEQ ID NO. 484) hinneytnknpkncllyk (SEQ ID NO. 485) kpclykkckisqvwwcmpvkdtfntyernnvlnskienniekiph (SEQ ID NO. 486) kckisqvwwcmpvkdtfntyernnvlnskienniekiph (SEQ ID NO. 487) kienniekiph (SEQ ID NO. 488) knktngskgvkgeyekkketngh (SEQ ID NO. 489) ktngskgvkgeyekkketngh (SEQ ID NO. 490) kgvkgeyekkketngh (SEQ ID NO. 491) kgeyekkketngh (SEQ ID NO. 492) ktiekinkskswffeeldeidkplaklrkrektqinktkyergdviidntciqkiirdyh (SEQ ID NO. 493) kinkskswffeeldeidkplaklrkrektqinktkyergdviidnteiqkiirdyh (SEQ ID NO. 494) kplaklrkrektqinktkyergdviidnteiqkiirdyh (SEQ ID NO. 495) himlksqmytnegnkscecsykkkssssnkvh (SEQ ID NO. 496) klrkrektqinktkyergdviidnteiqkiirdyh (SEQ ID NO. 497) krektqinktkyergdviidnteiqkiirdyh (SEQ ID NO. 498) ktqinktkyergdviidnteiqkiirdyh (SEQ ID NO. 499) kplaklrkrektqinktkyergdviidnteiqkiirdyhtlnvhkldh (SEQ ID NO. 500)
klrkrektqinktkyergdviidnteiqkiirdyhtlnvhkldh (SEQ ID NO. 501) krektqinktkyergdviidnteiqkiirdyhtlnvhkldh (SEQ ID NO. 502) ktqinktkyergdviidnteiqkiirdyhtlnvhkldh (SEQ ID NO. 503) kplaklrkrektqinktkyergdviidnteiqkiirdyhtlnvh (SEQ ID NO. 504) klrkrektqinktkyergdviidnteiqkiirdyhtlnvh (SEQ ID NO. 505) krektqinktkyergdviidnteiqkiirdyhtlnvh (SEQ ID NO. 506) ktqinktkyergdviidnteiqkiirdyhtlnvh (SEQ ID NO. 507) himlksqmytnegnkscecsykkkssssnkvh (SEQ ID NO. 508) ksqmytnegnkscecsykkkssssnkvh (SEQ ID NO. 509) kscecsykkkssssnkvh (SEQ ID NO. 510) kkkssssnkvh (SEQ ID NO. 511) kkssssnkvh (SEQ ID NO. 512) kssssnkvh (SEQ ID NO. 513) himlksqmytnegnkscecsykkkssssnk (SEQ ID NO. 514) himlksqmytnegnkscecsykkk (SEQ ID NO. 515) himlksqmytnegnkscecsykk (SEQ ID NO. 516) himlksqmytnegnkscecsyk (SEQ ID NO. 517) hnnhniqiykdkrinfmnphkvmyhdnmsknertek (SEQ ID NO. 518) hnnhniqiykdkrinfmnphkvmyhdnmsk (SEQ ID NO. 519) hnnhnniqiykdkrinfmnphk (SEQ ID NO. 520) hkvmyhdnmsknertek (SEQ ID NO. 521) hkvmyhdnmsk (SEQ ID NO. 522)
Replikins in Structural Proteins
[0291]It has also been determined that some structural proteins include Replikin structures. Structural proteins are molecules involved in tissue and organ support, such as collagen in skin and connective tissue and in membrane structures, for example amyloid A4 precursor protein (APP) in brain. Overproduction of these proteins is associated with disease; specifically, scleroderma in the case of overproduction of collagen in skin (Table 9) and Alzheimer's Disease in the case of overproduction of APP in the brain (Table 10).
[0292]The association of scleroderma and malignancy has been a source of controversy during recent years. Several mechanisms of interrelationship have been suggested in earlier reports. Recent long-term studies suggest an increased association-ratio of scleroderma and malignancy. However, the underlying mechanisms remain elusive. (Wenzel, J. Eur. J. Dermatol. 20002 May-June; 12(3): 296-300).
[0293]Several proteins concerned with the excessive production of proteins in scleroderma have been found to contain Replikin structures. Thus, these provide further examples of unrecognized targets for inhibition or cessation of excessive collagen production. Table 9 provides a list of proteins in scleroderma and the associated Replikins.
[0294]The APP protein is the source of the amyloid beta A4 protein, which in excessive amounts forms placques in the extracellular spaces in the brain, producing toxic effects associated with nerve cell loss in Alzheimer's Disease. Most studies to date have focused on the inability to clear the excessive deposits of A4, but have not considered that, rather than a waste clearance problem, this may actually be a problem of overproduction of the precursor protein APP. The high concentration of the Replikins in APP (3.3 Replikins per 100 amino acids) strongly suggest that overproduction may well be the cause of Alzheimer's Disease (Table 10). Therefore, the Replikins contained in Table 10 can be blocked or inhibited by the same methods as illustrated in detail for the glioma Replikin.
TABLE-US-00013 TABLE 9 Proteins overproduced in scleroderma and associated Replikins: PMC1 HUMAN: hreictiqssggimllkdqvlrcskiagvkvaeitelilk (SEQ ID NO. 523) hreictiqssggimllkdqvlresk (SEQ ID NO. 524) 34KD nucleolar scleroderma antigen: hreictiqssggimllkdqvlrcskiagvkvaeiteliklkalendqk (SEQ ID NO. 525) hreictiqssggirmlkdqvlrcskiagvkvaeitelilk (SEQ ID NO. 526) Fibrillarin: kkmqqenmkqpeqltlepyerdh (SEQ ID NO. 527) kmqqenmkpqeqltlepyerdh (SEQ ID NO. 528) SPOP HUMAN: hemeeskknrveindvepevfkemmcfiytgkapnldk (SEQ ID NO. 529) hemeeskknrveindvepevfkemmcfiytgk (SEQ ID NO. 530) Centromere protein C: khgelkvyk (SEQ ID NO. 531) klilgpqeekgkqh (SEQ ID NO. 532) hnrihhk (SEQ ID NO. 533) hhnssrkstkktnqssk (SEQ ID NO. 534) hnssrkstkktnqssk (SEQ ID NO. 535) khhnilpktlandkhshkph (SEQ ID NO. 536) hhnilpktlandkhshk (SEQ ID NO. 537) hnilpktlandkhshk (SEQ ID NO. 538) hnilpktlandk (SEQ ID NO. 539) kntpdskkissrnindhh (SEQ ID NO. 540) kntpdskkissrnindh (SEQ ID NO. 541) kdtciqspskecqkshpksvpvsskkk (SEQ ID NO. 542) kdtciqspskecqkshpksvpvsskk (SEQ ID NO. 543) hpksvpvsskkk (SEQ ID NO. 544) hpksvpvsskk (SEQ ID NO. 545) hpksvpvssk (SEQ ID NO. 546) Factor CTCBF, KU antigen: kalqekveikqlnh (SEQ ID NO. 547) ktlfplieakkdqvtageifgdnhedgptakklktegggah (SEQ ID NO. 548) ktlfplieakkkdqvtageifqdnb (SEQ ID NO. 549) klcvfkkierhsih (SEQ ID NO. 550) klcvfkkierh (SEQ ID NO. 551) kgpsfplkgiteqqkegleivk (SEQ ID NO. 552) hgpsfplkgiteqqk (SEQ ID NO. 553) ATP synthase subunit 6: htllkilstflfk (SEQ ID NO. 554) hllgnndknllpsk (SEQ ID NO. 555) FBRL nuclear protein: hrhegvficrgkedalvtk (SEQ ID NO. 556) hegvficrgkedalvtk (SEQ ID NO. 557) hsggnrgrgrggkrghqsgk (SEQ ID NO. 558) krgnqsgknvmveph (SEQ ID NO. 559) krgnqsgknvmvephrh (SEQ ID NO. 560) kkmqqenmkpqeqltlepyerdh (SEQ ID NO. 561) kmqqenmkpqeqltlepyerdh (SEQ ID NO. 562) HP1Hs-alpha protein: haypcdaemkeketak (SEQ ID NO. 563) keanvkcpqiviafyeerltwh (SEQ ID NO. 564) kvldrrvvkgqveyllkwkgfseeh (SEQ ID NO. 565) kgqveyllkwkgfseeh (SEQ ID NO. 566) FM/Scl nucleolar protein: ksevaagvkksglpsaerlenvlfgphdcsh (SEQ ID NO. 567) ksevaagvkksgplpsaerlenvlfgph (SEQ ID NO. 568) kaaeygkkaksetfrllhakniirpqlk (SEQ ID NO. 569) kaaeygkkaksetfrllhak (SEQ ID NO. 570) ksetfrllhak (SEQ ID NO. 571) hakniirpqlk (SEQ ID NO. 572) hmnlkiaeelpk (SEQ ID NO. 573) hsldhllklycnvdsnk (SEQ ID NO. 574) hllklycnvdsnk (SEQ ID NO. 575)
TABLE-US-00014 TABLE 10 Amyloid beta A4 precursor protein (APP) Replikins: kakerleakh (SEQ ID NO. 576) kdrqhtlk (SEQ ID NO. 577) kdrqhtlkh (SEQ ID NO. 578) ketcsekstnlh (SEQ ID NO. 579) kteeisevkmdaefgh (SEQ ID NO. 580) kteeisevkmdaefghdsgfevrh (SEQ ID NO. 581) kkyvraeqkdrqhtlkh (SEQ ID NO. 582) kyvraeqkdrqhtlkh (SEQ ID NO. 583) kkyvraeqkdrqh (SEQ ID NO. 584) kyvraeqkdrqht (SEQ ID NO. 585) hhvfnmlkkyvraeqk (SEQ ID NO. 586) hvfnmlkkyvraeqk (SEQ ID NO. 587) hhvfnmlkkyvraeqkdrqhtlkh (SEQ ID NO. 588) hvfnmlkkyvraeqkdrqhtlkh (SEQ ID NO. 589) hahfqkakerleakh (SEQ ID NO. 590) hahfqkakerleak (SEQ ID NO. 591) hfqkakerleak (SEQ ID NO. 592) hqermdvcethlhwhtvaketcsekstnlh (SEQ ID NO. 593) hqermdvcethlhwhtvaketcsek (SEQ ID NO. 594) hwhtvaketcsek (SEQ ID NO. 595) htvaketcsek (SEQ ID NO. 596) hlhwhtvaketcsek (SEQ ID NO. 597) hmnvqngkwesdpsgtktcigtk (SEQ ID NO. 598) hmnvqngkwesdpsgtk (SEQ ID NO. 599)
Passive Immunity
[0295]In another embodiment of the invention, isolated Replikin peptides may be used to generate antibodies, which may be used, for example to provide passive immunity in an individual. Passive immunity to the strain of influenza identified by the method of the invention to be the most likely cause of future influenza infections may be obtained by administering antibodies to Replikin sequences of the identified strain of influenza virus to patients in need. Similarly, passive immunity to malaria may be obtained by administering antibodies to Plasmodium falciparum Replikin(s).
[0296]Various procedures known in the art may be used for the production of antibodies to Replikin sequences. Such antibodies include but are not limited to polyclonal, monoclonal, chimeric, humanized, single chain, Fab fragments and fragments produced by an Fab expression library. Antibodies that are linked to a cytotoxic agent may also be generated. Antibodies may also be administered in combination with an antiviral agent. Furthermore, combinations of antibodies to different Replikins may be administered as an antibody cocktail.
[0297]For the production of antibodies, various host animals or plants may be immunized by injection with a Replikin peptide or a combination of Replikin peptides, including but not limited to rabbits, mice, rats, and larger mammals.
[0298]Monoclonal antibodies to Replikins may be prepared by using any technique that provides for the production of antibody molecules. These include but are not limited to the hybridoma technique originally described by Kohler and Milstein, (Nature, 1975, 256:495-497), the human B-cell hybridoma technique (Kosbor et al., 1983, Immunology Today, 4:72), and the EBV hybridoma technique (Cole et al., Monoclonal Antibodies and Cancer Therapy, Alan R. Liss, Inc., pp. 77-96). In addition, techniques developed for the production of chimeric antibodies (Morrison et al., 1984, Proc. Nat. Acad. Sci USA, 81:6851-6855) or other techniques may be used. Alternatively, techniques described for the production of single chain antibodies (U.S. Pat. No. 4,946,778) can be adapted to produce Replikin-specific single chain antibodies.
[0299]Particularly useful antibodies of the invention are those that specifically bind to Replikin sequences contained in peptides and/or polypeptides of influenza virus. For example, antibodies to any of peptides observed to be present in an emerging or re-emerging strain of influenza virus and combinations of such antibodies are useful in the treatment and/or prevention of influenza. Similarly, antibodies to any Replikins present on malaria antigens and combinations of such antibodies are useful in the prevention and treatment of malaria.
[0300]Antibody fragments which contain binding sites for a Replikin may be generated by known techniques. For example, such fragments include but are not limited to F(ab')2 fragments which can be produced by pepsin digestion of the antibody molecules and the Fab fragments that can be generated by reducing the disulfide bridges of the F(ab')2 fragments. Alternatively, Fab expression libraries can be generated (Huse et al., 1989, Science, 246:1275-1281) to allow rapid and easy identification of monoclonal Fab fragments with the desired specificity.
[0301]The fact that antimalignin antibody is increased in concentration in human malignancy regardless of cancer cell type (FIG. 5), and that this antibody binds to malignant cells regardless of cell type now may be explained by the presence of the Replikin structures herein found to be present in most malignancies (FIG. 1 and Table 2). Population studies have shown that antimalignin antibody increases in concentration in healthy adults with age, and more so in high-risk families, as the frequency of cancer increases. An additional two-fold or greater antibody increase which occurs in early malignancy has been independently confirmed with a sensitivity of 97% in breast cancers 1-10 mm in size. Shown to localize preferentially in malignant cells in vivo, histochemically the antibody does not bind to normal cells but selectively binds to (FIG. 4A, b) and is highly cytotoxic to transformed cells in vitro (FIG. 4C-f). Since in these examples the same antibody is bound by several cell types, that is, brain glioma, hematopoietic cells (leukemia), and small cell carcinoma of lung, malignant Replikin class unity is again demonstrated.
[0302]Antimalignin does not increase with benign proliferation, but specifically increases only with malignant transformation and replication in breast in vivo and returns from elevated to normal values upon elimination of malignant cells (FIG. 5). Antimalignin antibody concentration has been shown to relate quantitatively to the survival of cancer patients, that is, the more antibody, the longer the survival. Taken together, these results suggest that anti-Replikin antibodies may be a part of a mechanism of control of cell transformation and replication. Augmentation of this immune response may be useful in the control of replication, either actively with synthetic Replikins as vaccines, or passively by the administration of anti-Replikin antibodies, or by the introduction of non-immune based organic agents, such as for example, carbohydrates, lipids and the like, which are similarly designed to target the Replikin specifically.
[0303]In another embodiment of the invention, immune serum containing antibodies to one or more Replikins obtained from an individual exposed to one or more Replikins may be used to induce passive immunity in another individual or animal. Immune serum may be administered via i.v. to a subject in need of treatment. Passive immunity also can be achieved by injecting a recipient with preformed antibodies to one or more Replikins. Passive immunization may be used to provide immediate protection to individuals who have been exposed to an infectious organism. Administration of immune serum or preformed antibodies is routine and the skilled practitioner can readily ascertain the amount of serum or antibodies needed to achieve the desired effect.
Synthetic Replikin Vaccine
Active Immunity
[0304]Synthetic Replikin vaccines, based on Replikins such as the glioma Replikin (SEQ ID NO.: 1) "kagvaflhkk" or the hepatitis C Replikin (SEQ ID NO.: 18) "hyppkpgcivpak", or HIV Replikins such as (SEQ ID NO.: 5) "kcfncgkegh" or (SEQ ID NO.: 6) "kvylawvpahk" or preferably, an influenza vaccine based on conserved and/or emerging or re-emerging Replikin(s) over a given time period may be used to augment antibody concentration in order to lyse the respective virus infected cells and release virus extracellularly where chemical treatment can then be effective. Similarly, a malaria vaccine, based on Replikins observed in Plasmodium falciparum malaria antigens on the merozoite surface or within the parasitophorous vacuole, for example, can be used to generate cytotoxic antibodies to malaria.
[0305]Recognin and/or Replikin peptides may be administered to a subject to induce the immune system of the subject to produce anti-Replikin antibodies. Generally, a 0.5 to about 2 mg dosage, preferably a 1 mg dosage of each peptide is administered to the subject to induce an immune response. Subsequent dosages may be administered if desired.
[0306]The Replikin sequence structure is associated with the function of replication. Thus, whether the Replikins of this invention are used for targeting sequences that contain Replikins for the purpose of diagnostic identification, promoting replication, or inhibiting or attacking replication, for example, the structure-function relationship of the Replikin is fundamental.
[0307]It is preferable to utilize only the specific Replikin structure when seeking to induce antibodies that will recognize and attach to the Replikin fragment and thereby cause destruction of the cell. Even though the larger protein sequence may be known in the art as having a "replication associated function," vaccines using the larger protein often have failed or proven ineffective.
[0308]Although the present inventors do not wish to be held to a single theory, the studies herein suggest that the prior art vaccines are ineffective because they are based on the use of the larger protein sequence. The larger protein sequence invariably has one or more epitopes (independent antigenic sequences that can induce specific antibody formation); Replikin structures usually comprise one of these potential epitopes. The presence of other epitopes within the larger protein may interfere with adequate formation of antibodies to the Replikin, by "flooding" the immune system with irrelevant antigenic stimuli that may preempt the Replikin antigens, See, e.g., Webster, R. G., J. Immunol., 97(2):177-183 (1966); and Webster et al., J. Infect. Dis., 134:48-58, 1976; Klenerman et al, Nature 394:421-422 (1998) for a discussion of this well-known phenomenon of antigenic primacy whereby the first peptide epitope presented and recognized by the immune system subsequently prevails and antibodies are made to it even though other peptide epitopes are presented at the same time. This is another reason that, in a vaccine formulation, it is important to present the constant Replikin peptide to the immune system first, before presenting other epitopes from the organism so that the Replikin is not preempted but lodged in immunological memory.
[0309]The formation of an antibody to a non-Replikin epitope may allow binding to the cell, but not necessarily lead to cell destruction. The presence of structural "decoys" on the C-termini of malaria proteins is another aspect of this ability of other epitopes to interfere with binding of effective anti-Replikin antibodies, since the decoy epitopes have many lysine residues, but no histidine residues. Thus, decoy epitopes may bind anti-Replikin antibodies, but may keep the antibodies away from histidine-bound respiratory enzymes. Treatment may therefore be most efficacious in two stages: 1) proteases to hydrolize decoys, then; 2) anti-Replikin antibodies or other anti-Replikin agents.
[0310]It is well known in the art that in the course of antibody production against a "foreign" protein, the protein is first hydrolyzed into smaller fragments. Usually fragments containing from about six to ten amino acids are selected for antibody formation. Thus, if hydrolysis of a protein does not result in Replikin-containing fragments, anti-Replikin antibodies will not be produced. In this regard, it is interesting that Replikins contain lysine residues located six to ten amino acids apart, since lysine residues are known to bind to membranes.
[0311]Furthermore, Replikin sequences contain at least one histidine residue. Histidine is frequently involved in binding to redox centers. Thus, an antibody that specifically recognizes a Replikin sequence has a better chance of inactivating or destroying the cell in which the Replikin is located, as seen with anti-malignin antibody, which is perhaps the most cytotoxic anti-cancer antibody yet described, being active at picograms per cell.
[0312]One of the reasons that vaccines directed towards a particular protein antigen of a disease causing agent have not been fully effective in providing protection against the disease (such as foot and mouth vaccine which has been developed against the VP1 protein or large segments of the VP1 protein) is that the best antibodies have not been produced, that is--it is likely that the antibodies to the Replikins have not been produced. Replikins have not been produced. That is, either epitopes other than Replikins present in the larger protein fragments may interfere according to the phenomenon of antigenic primacy referred to above, and/or because the hydrolysis of larger protein sequences into smaller sequences for processing to produce antibodies results in loss of integrity of any Replikin structure that is present, e.g., the Replikin is cut in two and/or the histidine residue is lost in the hydrolytic processing. The present studies suggest that for an effective vaccine to be produced, the Replikin sequences, and no other epitope, should be used as the vaccine. For example, a vaccine of the invention can be generated using any one of the Replikin peptides identified by the three point recognition system.
[0313]Particularly preferred peptides--for example--an influenza vaccine include peptides that have been demonstrated to be conserved over a period of one or more years, preferably about three years or more, and/or which are present in a strain of influenza virus shown to have the highest increase in concentration of Replikins relative to Replikin concentration in other influenza virus strains, e.g., an emerging strain. The increase in Replikin concentration preferably occurs over a period of at least about six months to one year, preferably at least about two years or more, and most preferably about three years or more. Among the preferred Replikin peptides for use in an influenza virus vaccine are those Replikins observed to "re-emerge" after an absence from the hemagglutinin amino acid sequence for one or more years.
[0314]The Replikin peptides of the invention, alone or in various combinations are administered to a subject, preferably by i.v. or intramuscular injection, in order to stimulate the immune system of the subject to produce antibodies to the peptide. Generally the dosage of peptides is in the range of from about 0.1 μg to about 10 mg, preferably about 10 μg to about 1 mg, and most preferably about 50 μg to about 500 ug. The skilled practitioner can readily determine the dosage and number of dosages needed to produce an effective immune response.
Quantitative Measurement Early Response(s) to Replikin Vaccines
[0315]The ability to measure quantitatively the early specific antibody response in days or a few weeks to a Replikin vaccine is a major practical advantage over other vaccines for which only a clinical response months or years later can be measured.
Adjuvants
[0316]Various adjuvants may be used to enhance the immunological response, depending on the host species, including but not limited to Freund's (complete and incomplete), mineral gels, such as aluminum hydroxide, surface active substances such as lysolecithin, pluronic polyols, polyanions, peptides, oil emulsions, key limpet hemocyanin, dinitrophenol, and potentially useful human adjuvants such as BCG and Corynebacterium parvum.
Replikin Nucleotide Sequences
[0317]Replikin DNA or RNA may have a number of uses for the diagnosis of diseases resulting from infection with a virus, bacterium or other Replikin encoding agent. For example, Replikin nucleotide sequences may be used in hybridization assays of biopsied tissue or blood, e.g., Southern or Northern analysis, including in situ hybridization assays, to diagnose the presence of a particular organism in a tissue sample or an environmental sample, for example. The present invention also contemplates kits containing antibodies specific for particular Replikins that are present in a particular pathogen of interest, or containing nucleic acid molecules (sense or antisense) that hybridize specifically to a particular Replikin, and optionally, various buffers and/or reagents needed for diagnosis.
[0318]Also within the scope of the invention are oligoribonucleotide sequences, that include antisense RNA and DNA molecules and ribozymes that function to inhibit the translation of Replikin- or recognin-containing mRNA. Both antisense RNA and DNA molecules and ribozymes may be prepared by any method known in the art. The antisense molecules can be incorporated into a wide variety of vectors for delivery to a subject. The skilled practitioner can readily determine the best route of delivery, although generally i.v. or i.m. delivery is routine. The dosage amount is also readily ascertainable.
[0319]Particularly preferred antisense nucleic acid molecules are those that are complementary to a Replikin sequence contained in a mRNA encoding, for example, an influenza virus polypeptide, wherein the Replikin sequence comprises from 7 to about 50 amino acids including: [0320](1) at least one lysine residue located six to ten residues from a second lysine residue; [0321](2) at least one histidine residue; and [0322](3) at least 6% lysine residues.More preferred are antisense nucleic acid molecules that are complementary to a Replikin present in the coding strand of the gene or to the mRNA encoding the influenza virus hemagglutinin protein, wherein the antisense nucleic acid molecule is complementary to a nucleotide sequence encoding a Replikin that has been demonstrated to be conserved over a period of six months to one or more years and/or which are present in a strain of influenza virus shown to have an increase in concentration of Replikins relative to Replikin concentration in other influenza virus strains. The increase in Replikin concentration preferably occurs over a period of at least six months, preferably about one year, most preferably about two or three years or more.
[0323]Similarly, antisense nucleic acid molecules that are complementary to mRNA those that are complementary to a mRNA encoding bacterial Replikins comprising a Replikin sequence of from 7 to about 50 amino acids including: [0324](1) at least one lysine residue located six to ten residues from a second lysine residue; [0325](2) at least one histidine residue; and [0326](3) at least 6% lysine residues.More preferred are antisense nucleic acid molecules that are complementary to the coding strand of the gene or to the mRNA encoding a protein of the bacteria.
Diagnostic Applications
[0327]For organisms such as diatom plankton, foot and mouth disease virus, tomato leaf curl gemini virus, hepatitis B and C, HIV, influenza virus and malignant cells, identified constituent Replikins are useful as vaccines, and also may be usefully targeted for diagnostic purposes. For example, blood collected for transfusions may be screened for contamination of organisms, such as HIV, by screening for the presence of Replikins shown to be specific for the contamination organism. Also, screening for Replikin structures specific for a particular pathological organism leads to diagnostic detection of the organism in body tissue or in the environment.
Replikin Stimulation of Growth
[0328]In another embodiment of the invention, Replikin structures are used to increase the replication rate of cells, tissues or organs. A method is available to increase replication rates by the addition of specific Replikin structures for other cells, tissues or organs that it is desired to replicate more rapidly, together with or without appropriate stimulae to cell division know in the art for said cells, tissues or organs to increase the rate of replication and yield. This may be accomplished, for example, by methods known in the art, by modifying or transforming a gene encoding for or associated with a protein or enzyme having a replication function in the organism with at least one Replikin structure.
[0329]In another aspect of the invention, Replikin structures are used to increase the replication of organisms. The present invention demonstrates that in influenza virus, for example, increased replication associated with epidemics is associated with increased concentration of Replikins. The increase is due to 1) the reappearance of particular Replikin structures, which were present in previous years, but which then disappeared for one or more years; and/or 2) by the appearance of new Replikin compositions. In addition, in malaria Replikins, repetition of the same Replikin in a single protein occurs.
[0330]Thus, the present invention provides methods and compositions for increasing the replication of organisms. Similarly, in the manner that Replikins of different organisms can be targeted to inhibit replication of any organism, Replikins can be used to increase the replication of any organism. For example, production of rice, maize, and wheat crops, which are critical to feeding large populations in the world, can be improved, for example, by increasing the concentration (number of Replikins/100 amino acid residues) of any particular strain of rice.
[0331]As an example, in the Oryza sativa strain of rice, catalase isolated from immature seeds was observed to contain the following different Replikins within the 491 amino acid sequence of the protein:
TABLE-US-00015 (SEQ ID NO. 625) kfpdvihafkpnprsh (SEQ ID NO. 626) kfpdvihafk (SEQ ID NO. 627) karyvkfhwk (SEQ ID NO. 628) hpkvspelraiwvnylsqedeslgvkianlnvk (SEQ ID NO. 629) hrdeevdyypsrhaplrhapptpitprpvvgrrqkatihkqndfk (SEQ ID NO. 630) katihkqndfk (SEQ ID NO. 631) happtpitprpvvgrrqkatihkqndfk (SEQ ID NO. 632) kfrpsssfdtkttttnagapvwndnealtvgprgpilledyhliekvah (SEQ ID NO. 633) kfrpsssfdtkttttnagapvwndnealtvgprgpilledyn
[0332]Thus, by using recombinant gene cloning techniques well known in the art, the concentration of Replikin structures in an organism, such as a food crop plant, can be increased, which will promote increased replication of the organism. For example, inserting additional Replikin sequences like the Replikins identified above into the Oryza sativa catalase gene by methods well know in the art will promotethis organism's replication.
[0333]Similarly, in the NBS-LRR protein of Oryza sativa (japonica cultivar group), the following Replikins were found:
TABLE-US-00016 (SEQ ID NO. 634) kvkahfqkh (SEQ ID NO. 635) kvkahfqk (SEQ ID NO. 636) kdyeidkddlih (SEQ ID NO. 637) hmkqcfafcavfpkdyeidk (SEQ ID NO. 638) hmkqcfafcavfpk (SEQ ID NO. 639) hvfwelvwrsffqnvkqigsifqrkvyrygqsdvttskihdlmhdlavh (SEQ ID NO. 640) kqigsifqrkvrygpsdvttskihdlmhdlavh (SEQ ID NO. 641) kqigsifqrkvyrygpsdvttskihdlmh (SEQ ID NO. 642) kqigsifqrkvyrygqsdvttskih
[0334]Further, for aspartic proteinase oryzasin 1 precursor protein, the following Replikins were found:
TABLE-US-00017 khgvsagik (SEQ ID NO. 643) htvfdygkmrvgfak (SEQ ID NO. 644) hsryksgqsstyqkngk (SEQ ID NO. 645)
[0335]Similarly, in the MADS-box protein FDRMADS3 transcription factor of Oryza sativa (indica cultivar-group), the following Replikins were found:
TABLE-US-00018 (SEQ ID NO. 646) kqeamvlkqeinllqkglryiygnraneh (SEQ ID NO. 647) kqeinllqkglryiygnraneh (SEQ ID NO. 648) kskegmlkaaneilqekiveqnglidvgmmvadqqngh (SEQ ID NO. 649) kaaneilqekiveqnglidvgmmvadqqngh
[0336]Similarly, in LONI MAIZE (ATP-binding redox associated Hydrolase; Serine protease; Multigene family; Mitochondrion), the following Replikins were found:
TABLE-US-00019 (SEQ ID NO. 650) kvlaahrygik (SEQ ID NO. 651) klkiamkhliprvleqh (SEQ ID NO. 652) klkiamkh (SEQ ID NO. 653) ktslassiakalnrkfirislggvkdeadirgh (SEQ ID NO. 654) kalnrkfirislggvkdeadirgh (SEQ ID NO. 655) kfirislggvkdeadirgh (SEQ ID NO. 656) kvrlskatelvdrhlqsilvaekitqkvegqlsksqk (SEQ ID NO. 657) hlqsilvaekitqkvegglsksqk (SEQ ID NO. 658) kvrlskatelvdrh (SEQ ID NO. 659) kvggsavesskqdtkngkepihwhskgvaaralh (SEQ ID NO. 660) kvggsavesskqdtkngkepihwh (SEQ ID NO. 661) kvggsavesskqdtkngkepih (SEQ ID NO. 662) kqdtkngkepihwhskgvaaralh (SEQ ID NO. 663) kqdtkngkepih
[0337]Similarly, for Glyceraldehyde 3-phospate dehydrogenase A, a chloroplast precursor, the following Replikins are found:
TABLE-US-00020 hrdlrraraaalnivptstgaakavslvlpnlk (SEQ ID NO. 664) kvlddqkfgiikgtmttth (SEQ ID NO. 665) hiqagakkvlitapgk (SEQ ID NO. 666) hgrgdaspldviaindtggvkqashllk (SEQ ID NO. 667) kqashllk (SEQ ID NO. 697)
[0338]Further, examples of rust resistance-like protein RP1-4 (Zea mays) found include the following Replikins:
TABLE-US-00021 kvrrvlskdysslkqlmtlmmdddiskhlqiiesgleeredkvwmkeniik (SEQ ID NO. 668) kvrrvlskdysslkqlmtlmmdddiskh (SEQ ID NO. 669) hlqiiesgleeredkvwmkeniik (SEQ ID NO. 670) hdlreniimkaddlask (SEQ ID NO. 671) hvqnlenvigkdealask (SEQ ID NO. 672) kkqgyelrqlkdlnelggslh (SEQ ID NO. 673) kqgyelrqlkdlnelggslh (SEQ ID NO. 674) klylksrlkelilewssengmdamnilh (SEQ ID NO. 675) hlqllqlngmverlpnkvcnlsklrylrgykdqipnigk (SEQ ID NO. 676) hlqllqlngmverlpnkvcnlskrylrgyk (SEQ ID NO. 677) hlqllqlngmverlpnkvcnlsk (SEQ ID NO. 678) hnsnklpksvgelk (SEQ ID NO. 679) klpkvgelkh (SEQ ID NO. 680) hlsvrvesmqkhkeiiyk (SEQ ID NO. 681) khkeiiyk (SEQ ID NO. 682) klrdilqesqkfllvldlalfkh (SEQ ID NO. 683) hafsgaeikdqllrmklqdtaeeiakrlgqcplaakvlgsrmcrrk (SEQ ID NO. 684) hafsgaeikdqllrmk (SEQ ID NO. 685) klqdtacciakrlgqclaakvlgsrmcrrkdiaewkaadvwfeksh (SEQ ID NO. 686) kvlgsrmcrrkdiacwkaadvwfeksh (SEQ ID NO. 687) kdiaewkaadvwfeksh (SEQ ID NO. 688) kaadvwfeksh (SEQ ID NO. 689) hvptttslptskvfgrnsdrdrivkfllgktttaeasstk (SEQ ID NO. 690) kailteakqlrdllglph (SEQ ID NO. 691) kakaksgkgpllredessstattvmkpfh (SEQ ID NO. 692) ksphrgkleswlrrlkeafydaedlldeh (SEQ ID NO. 693) ksphrgkleswlrrlk (SEQ ID NO. 694) hrgkleswlrrlk (SEQ ID NO. 695) ksphrgk (SEQ ID NO. 696)
[0339]As discussed previously, the Replikin in wheat ubiquitin activating enzyme E (SEQ ID Nos. 601-603) is conserved. This conservation of Replikin structure provides reliable targets for stimulation of plant growth.
[0340]The close relationship of Replikins to redox enzymes is also clearly indicated in this structure in wheat. Thus, this wheat ubiquitin activating enzyme E activates ubiquitin by first adenylating with ATP its carboxy-terminal glycine residue and, thereafter, linking this residue to the side chain of a cysteine residue in E1 (SEQ ID NO. 603), yielding an ubiquitin-E1 thiolester and free AMP.
[0341]A further example of the relationship of wheat Replikins to redox enzymes was also found in the PSABWheat Protein, Photosystem I P700 chlorophyll A apoprotein A2 (PsaB) (PSI-B) isolated from bread Chinese spring wheat Chloroplast Triticum aestivum. This protein functions as follows: PsaA and PsaB bind 9700, the primary electron donor of photosystem I (PSI), as well as the electron acceptors A0, A1, and FX. PSI functions as a plastocyanin/cytochrome c6-ferredoxin oxidoreductase. Cofactor P700 is a chlorophyll A dimer, A0 is chlorophyll A, A1 is a phylloquinone and FX is a 4Fe-4S iron-sulfur center. The subunit A psaA/S heterodimer binds the P700 chlorophyll special pair and subsequent electron acceptors. The PSI reaction center of higher plants and algae is composed of one at least 11 subunits. This is an integral membrane protein of the Chloroplast thylakoid membrane. The 4Fe-4S iron-sulfur "center" to which `h` bind is critical; hence the significance of `h` in Replikin structure. Next to bacterial Replikins, these wheat Replikins and plant Replikins are the most primitive evolutionary illustrations of the importance of the Replikin structure to replication and the energy source needed for replication. This basic relationship carries through algae, virus Replikins, bacteria, cancer cells, and apparently all organisms with regard to replication.
[0342]Further examples of Replikins were found in the PSAB Wheat protein, which is critical fox wheat growth. These include:
TABLE-US-00022 hlqpkwkpslswfknaesrlnhh (SEQ ID NO. 604) hlqpkwkpslswfk (SEQ ID NO. 605) kwkpslswfknaesrlnhh (SEQ ID NO. 606) kwkpslswfknaesrlnh (SEQ ID NO. 607) kpslswfknaesrlnhh (SEQ ID NO. 608) kpslswfknaesrlnh (SEQ ID NO. 609) hhaialglhtttlilvkgaldargsklmpdkk (SEQ ID NO. 610) haialglhtttlilvkgaldargsklmpdkk (SEQ ID NO. 611) hhaialglhtttlilvkgaldargsk (SEQ ID NO. 612) haialglhtttlilvkgaldargsk (SEQ ID NO. 613) htttlilvkgaldargsklmpdkk (SEQ ID NO. 614) htttlilvkgaldargsklmpdk (SEQ ID NO. 615) htttlilvkgaldargsk (SEQ ID NO. 616)
[0343]A further example of the relationship of wheat Replikins to redox is provide in the PSAA_WHEAT Photosystem I 9700 chlorophyll A apoprotein A1, that include:
TABLE-US-00023 (SEQ ID NO. 617) hhhlaiailfliaghmyrtnwgighglkdileahkgpftgqghk (SEQ ID NO. 618) hhlaiailfliaghmyrtnwgighglkdileahkgpftgqghk (SEQ ID NO. 619) hlaiailfliaghmyrtnwgighglkdileahkgpftgqghk (SEQ ID NO. 620) hmyrtnwgighglkdileahkgpftgqghk (SEQ ID NO. 621) hglkdileahkgpftgqghk (SEQ ID NO. 622) hdileahkgpftgqghk (SEQ ID NO. 623) hkgpftgqghk (SEQ ID NO. 624) kgpftgqghk
Computer Software for Identifying Replikins
[0344]The present invention also provides methods for identifying Replikin sequences in an amino acid or nucleic acid sequence. Visual scanning of over four thousand sequences was performed in developing the present 3-point-recognition methods. However, data banks comprising nucleotide and/or amino acid sequences can also be scanned by computer for the presence of sequences meeting the 3 point recognition requirements.
[0345]According to another embodiment of the invention, three-point recognition methods described herein may be performed by a computer. FIG. 6 is a block diagram of a computer available for use with the foregoing embodiments of the present invention. The computer may include a processor, an input/output device and a memory storing executable program instructions representing the 3-point-recognition methods of the foregoing embodiments. The memory may include a static memory, volatile memory and/or a nonvolatile memory. The static memory conventionally may be a read only memory ("ROM") provided on a magnetic, or an electrical or optical storage medium. The volatile memory conventionally may be a random accessmemory ("RAM") and may be integrated as a cache within the processor or provided externally from the processor as a separate integrated circuit. The non-volatile memory may be an electrical, magnetic or optical storage medium.
[0346]From a proteomic point of view the construction of a "3-point recognition" template based on the new glioma peptide sequence led directly to identification of a biology-wide class of proteins having related structures and functions. The operation of the 3-point-recognition method resembles identification by the use of a "keyword" search; but instead of using the exact spelling of the keyword "kagvaflhkk" (SEQ ID NO.: 1) as in a typical sequence homology search, or in the nucleotide specification of an amino acid, an abstraction of the keyword delimited by the "3-point-recognition" parameters is used. This delimited abstraction, although derived from a single relatively short amino acid sequence leads to identification of a class of proteins with structures that are defined by the same specifications. That particular functions, in this case transformation and replication, in addition to structures, turn out also to be shared by members of the exposed class suggests that these structures and functions are related. Thus, from this newly identified short peptide sequence, a molecular recognition `language` has been formulated, which previously has not been described. Further, the sharing of immunological specificity by diverse members of the class, as here demonstrated for the cancer Replikins, suggests that B cells and their product antibodies recognize Replikins by means of a similar recognition language.
Other Uses of the Three Point Recognition Method
[0347]Since "3-point-recognition" is a proteomic method that specifies a particular class of proteins, using three or more different recognition points for other peptides similarly should provide useful information concerning other proteins classes. Further, the "3-point-recognition" method is applicable to other recognins, for example to the TOLL `innate` recognition of lipopolyssacharides of organisms. The three point recognition method may also be modified to identify other useful compounds of covalently linked organic molecules, including other covalently linked amino acids, nucleotides, carbohydrates, lipids or combinations thereof. In this embodiment of the invention a sequence is screened for subsequences containing three or more desired structural characteristics. In the case of screening compounds composed of covalently linked amino acids, lipids or carbohydrates the subsequence of 7 to about 50 covalently linked units should contain (1) at least one first amino acid, carbohydrate or lipid residue located seven to ten residues from a second of the first amino acid, carbohydrate or lipid residue; (2) encoding at least one second amino acid, lipid or carbohydrate residue; and (3) at least 6% of the first amino acid, carbohydrate or lipid residue. In the case of screening nucleotide sequences, the subsequence of about 21 to about 150 nucleotides should contain (1) at least one codon encoding a first amino acid located within eighteen to thirty nucleotides from a second codon encoding the first amino acid residue; (2) at least one second amino acid residue; and (3) encodes at least 6% of said first amino acid residue.
[0348]Several embodiments of the present invention are specifically illustrated and described herein. However, it will be appreciated that modifications and variations of the present invention are encompassed by the above teachings and within the purview of the appended claims without departing from the spirit and intended scope of the invention.
EXAMPLE 1
Process for Extraction, Isolation and Identification of Replikins and the Use of Replikins to Target, Label or Destroy Replikin-Containing Organisms
[0349]a) Algae
[0350]The following algae were collected from Bermuda water sites and either extracted on the same day or frozen at -20 degrees C. and extracted the next day. The algae were homogenized in a cold room (at 0 to 5 degrees C.) in 1 gram aliquots in neutral buffer, for example 100 cc. of 0.005M phosphate buffer solution, pH 7 ("phosphate buffer") for 15 minutes in a Waring blender, centrifuged at 3000 rpm, and the supernatant concentrated by perevaporation and dialyzed against phosphate buffer in the cold to produce a volume of approximately 15 ml. The volume of this extract solution was noted and an aliquot taken for protein analysis, and the remainder was fractionated to obtain the protein fraction having a pK range between 1 and 4.
[0351]The preferred method of fractionation is chromatography as follows: The extract solution is fractionated in the cold room (4 degrees C.) on a DEAE cellulose (Cellex-D) column 2.5×11.0 cm, which has been equilibrated with 0.005M phosphate buffer. Stepwise eluting solvent changes are made with the following solutions: [0352]Solution 1-4.04 g. NaH2P04 and 0.5 g NaH2P04 are dissolved in 15 litres of distilled water (0.005 molar, pH 7); [0353]Solution 2-8.57 g. NaH2P04 is dissolved in 2,480 ml. of distilled water; [0354]Solution 3-17.1 g. of NaH2P04 is dissolved in 2480 ml of distilled water (0.05 molar, pH 4.7); [0355]Solution 4-59.65 g. of NaH2P04 is dissolved in 2470 ml distilled water (0.175 molar); [0356]Solution 5-101.6 g. of NaH2P04 is dissolved in 2455 ml distilled water (pH 4.3); [0357]Solution 6-340.2 g. of NaH2P04 is dissolved in 2465 of distilled water (1.0 molar, pX-i 4.1); [0358]Solution 7-283.63 g. of 80% phosphoric acid (H3P04) is made up in 2460 ml of distilled water (1.0 molar, pH 1.0).
[0359]The extract solution, in 6 to 10 ml volume, is passed onto the column and overlayed with Solution 1, and a reservoir of 300 ml of Solution 1 is attached and allowed to drip by gravity onto the column. Three ml aliquots of eluant are collected and analyzed for protein content at OD 280 until all of the protein to be removed with Solution 1 has been removed from the column. Solution 2 is then applied to the column, followed in succession by Solutions 3, 4, 5, 6 and 7 until all of the protein which can, be removed with each Solution is removed from the column. The eluates from Solution 7 are combined, dialyzed against phosphate buffer, the protein content determined of both dialysand and dialyzate, and both analyzed by gel electrophoresis. One or two bands of peptide or protein of molecular weight between 3,000 and 25,000 Daltons are obtained in Solution 7. For example the algae Caulerpa mexicana, Laurencia obtura, Cladophexa prolifera, Sargassum natans, Caulerpa verticillata, Halimeda tuna, and Penicillos capitatus, after extraction and treatment as above, all demonstrated in Solution 7 eluates sharp peptide bands in this molecular weight region with no contaminants. These Solution 7 proteins or their eluted bands are hydrolyzed, and the amino acid composition determined. The peptides so obtained, which have a lysine composition of 6% or greater are Replikin precursors. These Replikin peptide precursors are then determined for amino acid sequence and the Replikins are determined by hydrolysis and mass spectrometry as detailed in U.S. Pat. No. 6,242,578 B1. Those which fulfill the criteria defined by the "3-point-recognition" method are identified as Replikins. This procedure can also be applied to obtain yeast, bacterial and any plant Replikins.
[0360]b) Virus
[0361]Using the same extraction and column chromatography separation methods as above in a) for algae, Replikens in virus-infected cells are isolated and identified.
[0362]c) Tumor Cells In Vivo and In Vitro Tissue Culture
[0363]Using the same extraction and column chromatography separation methods as above in a) for algae, Replikins in tumor cells are isolated and identified. For example, Replikin precursors of Astrocytin isolated from malignant brain tumors, Malignin (Aglyco 1OB) isolated from glioblastoma tumor cells in tissue culture, MCF7 mammary carcinoma cells in tissue culture, and P3J Lymphoma cells in tissue culture each treated as above in a) yielded Replikin precursors with lysine content of 9.1%, 6.7%, 6.7%, and 6.5% respectively. Hydrolysis and mass spectrometry of Aglyco 1OB as described in Example 10 U.S. Pat. No. 6,242,578 B1 produced the amino acid sequence, ykagvaflhkkndiide the 16-mer Replikin.
EXAMPLE 2
[0364]As an example of diagnostic use of Replikins: Aglyco 1OB or the 16-mer Repliken may be used as antigen to capture and quantify the amount of its corresponding antibody present in serum for diagnostic purposes are as shown in FIGS. 2,3,4 and 7 of U.S. Pat. No. 6,242,578 B1.
[0365]As an example of the production of agents to attach to Replikins for labeling, nutritional or destructive purposes: Injection of the 16-mer Replikin into rabbits to produce the specific antibody to the 16-mer Replikin is shown in Example 6 and FIGS. 9A and 9B of U.S. Pat. No. 6,242,578 B1.
[0366]As an example of the use of agents to label Replikins: The use of antibodies to the 16-mer Replikin to label specific cells which contain this Replikin is shown in FIG. 5 and Example 6 of U.S. Pat. No. 6,242,578 B1.
[0367]As an example of the use of agents to destroy Replikins: The use of antibodies to the 16-mer Replikin to inhibit or destroy specific cells which contain this Replikin is shown in FIG. 6 of U.S. Pat. No. 6,242,578 B1.
EXAMPLE 3
[0368]Analysis of sequence data of isolates of influenza virus hemagglutinin protein or neuraminidase protein for the presence and concentration of Replikins is carried out by visual scanning of sequences or through use of a computer program based on the 3-point recognition system described herein. Isolates of influenza virus are obtained and the amino acid sequence of the influenza hemagglutinin and/or neuraminidase protein is obtained by any art known method, such as by sequencing the hemagglutinin or neuraminidase gene and deriving the protein sequence therefrom. Sequences are scanned for the presence of new Replikins, conservation of Replikins over time and concentration of Replikins in each isolate. Comparison of the Replikin sequences and concentrations to the amino acid sequences obtained from isolates at an earlier time, such as about six months to about three years earlier, provides data that are used to predict the emergence of strains that are most likely to be the cause of influenza in upcoming flu seasons, and that form the basis for seasonal influenza peptide vaccines or nucleic acid based vaccines. Observation of an increase in concentration, particularly a stepwise increase in concentration of Replikins in a given strain of influenza virus for a period of about six months to about three years or more is a predictor of emergence of the strain as a likely cause of influenza epidemic or pandemic in the future.
[0369]Peptide vaccines or nucleic acid-based vaccines based on the Replikins observed in the emerging strain are generated. An emerging strain is identified as the strain of influenza virus having the highest increase in concentration of Replikin sequences within the hemagglutinin and/or neuraminidase sequence during the time period. Preferably, the peptide or nucleic acid vaccine is based on or includes any Replikin sequences that are observed to be conserved in the emerging strain. Conserved Replikins are preferably those Replikin sequences which are present in the hemagglutinin or neuraminidase protein sequence for about two years and preferably longer. The vaccines may include any combination of Replikin sequences identified in the emerging strain.
[0370]For vaccine production, the Replikin peptide or peptides identified as useful for an effective vaccine are synthesized by any method, including chemical synthesis and molecular biology techniques, including cloning, expression in a host cell and purification therefrom. The peptides are preferably admixed with a pharmaceutically acceptable carrier in an amount determined to induce a therapeutic antibody reaction thereto. Generally, the dosage is about 0.1 μg to about 10 mg.
[0371]The influenza vaccine is preferably administered to a patient in need thereof prior to the onset of "flu season." Influenza flu season generally occurs in late October and lasts through late April. However, the vaccine may be administered at any time during the year. Preferably, the influenza vaccine is administered once yearly, and is based on Replikin sequences observed to be present, and preferably conserved in the emerging strain of influenza virus. Another preferred Replikin for inclusion in an influenza vaccine is a Replikin demonstrated to have re-emerged in a strain of influenza after an absence of one or more years.
EXAMPLE 4
[0372]Analysis of sequence data of isolates of Plasmodium falciparum antigens for the presence and concentration of Replikins is carried out by visual scanning of sequences or through use of a computer program based on the 3-point recognition method described herein. Isolates of Plasmodium falciparum are obtained and the amino acid sequence of the protein is obtained by any art known method, such as by sequencing the gene and deriving the protein sequence therefrom. Sequences are scanned for the presence of Replikins, conservation of Replikins over time and concentration of Replikins in each isolate. This information provides data that are used to form the basis for anti-malarial peptide vaccines or nucleic acid based vaccines.
[0373]Peptide vaccines or nucleic acid-based vaccines based on the Replikins observed in the malaria causing organism are generated. Preferably, the peptide or nucleic acid vaccine is based on or includes any Replikin sequences that are observed to be present on a surface antigen of the organism. The vaccines may include any combination of Replikin sequences identified in the malaria causing strain.
[0374]For vaccine production, the Replikin peptide or peptides identified as useful for an effective vaccine are synthesized by any method, including chemical synthesis and molecular biology techniques, including cloning, expression in a host cell and purification therefrom. The peptides are preferably admixed with a pharmaceutically acceptable carrier in an amount determined to induce a therapeutic antibody reaction thereto. Generally, the dosage is about 0.1 μg to about 10 mg.
[0375]Then malaria vaccine is preferably administered to a patient in need thereof at any time during the year, and particularly prior to travel to a tropical environment.
[0376]Another embodiment includes an antisense nucleic acid molecule complementary to the coding strand of the gene or the mRNA encoding organism for the replikins in organisms including, but not limited to, viruses, trypanosomes, bacteria, fungi, algae, amoeba, and plants, wherein said antisense nucleic acid molecules is complementary to a nucleotide sequence of a replikin containing organism.
Sequence CWU
1
729110PRTArtificial SequenceDescription of Artificial Sequence Synthetic
glioma replikin 1Lys Ala Gly Val Ala Phe Leu His Lys Lys1
5 10213PRTSaccharomyces cerevisiae 2His Ser Ile Lys
Arg Glu Leu Gly Ile Ile Phe Asp Lys1 5
10310PRTGemini vinis virus 3His Lys Gln Lys Ile Val Ala Pro Val Lys1
5 10416PRTUnknownDescription of Unknown Virus
recognin 4Tyr Lys Ala Gly Val Ala Phe Leu His Lys Lys Asn Asp Ile Asp
Glu1 5 10 15510PRTHuman
immunodeficiency virus type 1 5Lys Cys Phe Asn Cys Gly Lys Glu Gly His1
5 10611PRTHuman immunodeficiency virus type
1 6Lys Val Tyr Leu Ala Trp Val Pro Ala His Lys1 5
10710PRTHuman immunodeficiency virus type 2 7Lys Cys Trp Asn Cys
Gly Lys Glu Gly His1 5 10811PRTMaize
streak virus 8Lys Tyr Ile Val Cys Ala Arg Glu Ala His Lys1
5 10917PRTMaize streak virus 9Lys Glu Lys Lys Pro Ser
Lys Asp Glu Ile Met Arg Asp Ile Ile Ser1 5
10 15His109PRTStaphylococcus aureus 10Lys Lys Glu Lys
Thr Thr His Asn Lys1 51110PRTBovine herpesvirus 4 11His Lys
Ile Asn Ile Thr Asn Gly Gln Lys1 5
101210PRTMeleagrid herpesvirus 1 12His Lys Asp Leu Tyr Arg Leu Leu Met
Lys1 5 101315PRTUnknownDescription of
Unknown Virus recognin 13Lys Phe Arg Ile Asn Ala Lys Asn Tyr Phe Leu Thr
Tyr Pro His1 5 10
151419PRTUnknownDescription of Unknown Virus recognin 14Lys Asn Leu Glu
Thr Pro Val Asn Lys Leu Phe Ile Arg Ile Cys Arg1 5
10 15Glu Phe His1514PRTUnknownDescription of
Unknown Virus recognin 15His Pro Asn Ile Gln Ala Ala Lys Ser Ser Thr Asp
Val Lys1 5 101619PRTUnknownDescription of
Unknown Virus recognin 16Lys Ser Ser Thr Asp Val Lys Ala Tyr Met Asp Lys
Asp Gly Asp Val1 5 10
15Leu Asp His1721PRTUnknownDescription of Unknown Virus recognin 17Lys
Ala Ser Ala Leu Asn Ile Leu Arg Glu Lys Ala Pro Lys Asp Phe1
5 10 15Val Leu Gln Phe
His201813PRTHepatitis C virus 18His Tyr Pro Pro Lys Pro Gly Cys Ile Val
Pro Ala Lys1 5 10194PRTHomo sapiens 19Tyr
Lys Ala Gly1206PRTHomo sapiens 20Tyr Lys Ala Gly Val Ala1
5217PRTHomo sapiens 21Tyr Lys Ala Gly Val Ala Phe1
5227PRTHomo sapiens 22Tyr Lys Ala Gly Val Ala Phe1
5239PRTHomo sapiens 23Ala Gly Val Ala Phe His Lys Lys Asn1
5244PRTHomo sapiens 24Gly Val Ala Phe1253PRTHomo sapiens 25Val Ala
Phe1267PRTHomo sapiens 26Val Ala Phe Leu His Lys Lys1
5277PRTHomo sapiens 27Val Ala Phe Leu His Lys Lys1
5289PRTHomo sapiens 28Val Ala Phe Leu His Lys Lys Asn Asp1
5298PRTHomo sapiens 29Val Ala Phe His Lys Lys Asn Asp1
5304PRTHomo sapiens 30Ala Phe Leu His1318PRTHomo sapiens 31His Lys Lys
Asn Asp Ile Asp Glu1 5326PRTHomo sapiens 32Lys Lys Asn Asp
Ile Asp1 5336PRTHomo sapiens 33Lys Asn Asp Ile Asp Glu1
5348PRTCaldophera prolifera 34Lys Ala Ser Lys Phe Thr Lys His1
53512PRTIsolepis prolifera 35Lys Ala Gln Ala Glu Thr Gly Glu
Ile Lys Gly His1 5
103610PRTSchizosaccharomyces pombe 36Lys Ser Phe Lys Tyr Pro Lys Lys His
Lys1 5 103710PRTOryza sativa 37Lys Lys
Ala Tyr Gly Asn Glu Leu His Lys1 5
10389PRTPenicillium marneffei 38Lys Val Asp Ile Val Thr His Gln Lys1
53912PRTDiseula dcstructiva 39Lys Leu Glu Glu Asp Ala Ala Tyr
His Arg Lys Lys1 5 104017PRTOphiostoma
novo-ulmi 40Lys Val Ile Leu Pro Leu Arg Gly Asn Ile Lys Gly Ile Phe Phe
Lys1 5 10
15His4111PRTEntamoeba invadens 41Lys Leu Ile Leu Lys Gly Asp Leu Asn Lys
His1 5 10428PRTHelicobacter pylori 42Lys
Ser Val His Ala Phe Leu Lys1 5439PRTMycoplasma pulmonis
43Lys Val His Phe Phe Gln Leu Lys Lys1 5449PRTArabidopsis
thaliana 44Lys Asp His Asp Phe Asp Gly Asp Lys1
54511PRTArabidopsis thaliana 45Lys Met Lys Gly Leu Lys Gln Lys Lys Ala
His1 5 104612PRTArabidopsis thaliana
46Lys Glu Leu Ser Ser Thr Thr Gln Glu Lys Ser His1 5
10479PRTFeline immunodeficiency virus 47His Leu Lys Asp Tyr
Lys Leu Val Lys1 5487PRTRous sarcoma virus 48Lys Lys Leu
Arg His Glu Lys1 5497PRTAvian sarcoma virus 49Lys Lys Leu
Arg His Asp Lys1 5507PRTHomo sapiens 50Lys Lys Leu Arg His
Asp Lys1 5517PRTAvian sarcoma virus 51Lys Lys Leu Arg His
Glu Lys1 5527PRTHomo sapiens 52Lys Lys Leu Arg His Glu Lys1
5538PRTHomo sapiens 53Lys Gln Ala His Glu Leu Ala Lys1
5548PRTPolyama virus 54Lys Thr His Arg Phe Ser Lys His1
5558PRTSindbis virus 55Lys Asn Leu His Glu Lys Ile Lys1
5569PRTHuman papilloamavirus type 71 56Lys His Arg Pro Leu Leu Gln Leu
Lys1 5577PRTAvian encephalomyelitis virus 57Lys Ser Pro Asn
His Val Lys1 5588PRTFeline sarcoma virus 58Lys Asn Ile His
Leu Glu Lys Lys1 5598PRTHomo sapiens 59Lys Asn Ile His Leu
Glu Lys Lys1 56010PRTPolyoma virus 60Lys Pro His Leu Ala
Gln Ser Leu Glu Lys1 5 10619PRTPolyoma
virus 61Lys Gln His Arg Glu Leu Lys Asp Lys1 5629PRTPolyoma
virus 62Lys Gln His Arg Glu Leu Lys Asp Lys1 56312PRTMurine
leukemia virus 63Lys Val Pro Val Leu Ile Ser Pro Thr Leu Lys His1
5 106413PRTHuman T-cell lymphotropic virus type 2
64Lys Ser Leu Leu Leu Glu Val Asp Lys Asp Ile Ser His1 5
106513PRTHomo sapiens 65Lys Ala Gly Ile Thr Ile Met Val
Lys Arg Glu Tyr His1 5 10668PRTHomo
sapiens 66Lys Ser Gly Lys His Leu Gly Lys1 5679PRTHomo
sapiens 67Lys Arg Arg Glu Gln Leu Lys His Lys1 56810PRTHomo
sapiens 68Lys Ser Phe Glu Val Ile Lys Val Ile His1 5
10698PRTHomo sapiens 69Lys Lys Lys His Thr Val Lys Lys1
5709PRTHomo sapiens 70Lys Ala Gln Lys Asp His Leu Ser Lys1
57110PRTHomo sapiens 71His Leu Lys Arg Val Lys Asp Leu Lys Lys1
5 107211PRTHomo sapiens 72Lys Tyr Gly Ser Pro
Lys His Arg Leu Ile Lys1 5
107313PRTPapilloma virus type 11 73Lys Leu Lys His Ile Leu Gly Lys Ala
Arg Phe Ile Lys1 5 107412PRTHomo sapiens
74Lys Gly Asp His Val Lys His Tyr Lys Ile Arg Lys1 5
107513PRTHomo sapiens 75Lys Glu Lys Leu Arg Asp Val Met Val
Asp Arg His Lys1 5 107615PRTHomo sapiens
76Lys Leu Gln Ala Arg Gln Gln Gln Leu Leu Lys Lys Ile Glu His1
5 10 157714PRTHomo sapiens 77Lys
Lys Gly Asn Arg Val Ser Pro Thr Met Lys Val Thr His1 5
10789PRTHomo sapiens 78Lys Glu Ile Pro Leu His Phe Arg Lys1
5798PRTHomo sapiens 79Lys Lys Lys Pro His Ile Lys Lys1
5809PRTHomo sapiens 80Lys Thr Arg His Asp Pro Leu Ala Lys1
58110PRTHomo sapiens 81Lys His His Pro Lys Asp Asn Leu Ile Lys1
5 108210PRTHomo sapiens 82Lys His Lys Arg
Lys Lys Phe Arg Gln Lys1 5 108310PRTHomo
sapiens 83Lys Ala Gly Val Ala Phe Leu His Lys Lys1 5
108410PRTHomo sapiens 84Lys His Lys Arg Lys Lys Phe Arg Gln
Lys1 5 108510PRTHomo sapiens 85Lys Lys
Lys Ser Lys Lys His Lys Asp Lys1 5
108611PRTHomo sapiens 86His Lys Ser Glu Lys Pro Ala Leu Pro Arg Lys1
5 108714PRTHomo sapiens 87Lys Lys Lys Lys Pro
Ser Arg Leu Lys Gly Asp Asn Glu Lys1 5
108816PRTHomo sapiens 88Lys Thr Lys Lys Gly Asn Arg Val Ser Pro Thr Met
Lys Val Thr His1 5 10
158918PRTHomo sapiens 89Lys His Lys Glu Lys Met Ser Lys Asp Gly Lys Lys
Lys Lys Lys Lys1 5 10
15Ser Lys909PRTLegionella sp. 90Lys Ile His Leu Ile Ser Val Lys Lys1
5918PRTInfluenza B virus 91Lys Ser His Phe Ala Asn Leu Lys1
59211PRTInfluenza B virus 92Lys Ser His Phe Ala Asn Leu Lys Gly
Thr Lys1 5 109319PRTInfluenza B virus
93Lys Ser His Phe Ala Asn Leu Lys Gly Thr Lys Thr Arg Gly Lys Leu1
5 10 15Cys Pro
Lys949PRTInfluenza B virus 94His Glu Lys Tyr Gly Gly Leu Asn Lys1
59511PRTInfluenza B virus 95His Glu Lys Tyr Gly Gly Leu Asn Lys Ser
Lys1 5 109620PRTInfluenza B virus 96His
Glu Lys Tyr Gly Gly Leu Asn Lys Ser Lys Pro Tyr Tyr Thr Gly1
5 10 15Glu His Ala
Lys209713PRTInfluenza B virus 97His Ala Lys Ala Ile Gly Asn Cys Pro Ile
Trp Val Lys1 5 109823PRTInfluenza B virus
98His Ala Lys Ala Ile Gly Asn Cys Pro Ile Trp Val Lys Thr Pro Leu1
5 10 15Lys Leu Ala Asn Gly Thr
Lys209929PRTInfluenza B virus 99His Ala Lys Ala Ile Gly Asn Cys Pro Ile
Trp Val Lys Thr Pro Leu1 5 10
15Lys Leu Ala Asn Gly Thr Lys Tyr Arg Pro Pro Ala Lys20
2510032PRTInfluenza B virus 100His Ala Lys Ala Ile Gly Asn Cys Pro
Ile Trp Val Lys Thr Pro Leu1 5 10
15Lys Leu Ala Asn Gly Thr Lys Tyr Arg Pro Pro Ala Lys Leu Leu
Lys20 25 3010113PRTInfluenza B virus
101His Phe Ala Asn Leu Lys Gly Thr Lys Thr Arg Gly Lys1 5
1010217PRTInfluenza B virus 102His Phe Ala Asn Leu Lys
Gly Thr Lys Thr Arg Gly Lys Leu Cys Pro1 5
10 15Lys10316PRTInfluenza B virus 103His Ser Asp Asn
Glu Ile Gln Met Val Lys Leu Tyr Gly Asp Ser Lys1 5
10 1510421PRTInfluenza B virus 104His Ser Asp
Asn Glu Ile Gln Asp Lys Met Val Lys Leu Tyr Gly Asp1 5
10 15Ser Lys Pro Gln Lys2010519PRTInfluenza
B virus 105His Ser Asp Asn Glu Ile Gln Met Val Lys Leu Tyr Gly Asp Ser
Lys1 5 10 15Pro Gln
Lys1069PRTInfluenza B virusMOD_RES(2)ala or val 106Lys Xaa Ser Ile Leu
His Glu Val Lys1 510715PRTInfluenza B virus 107Lys Cys Thr
Gly Thr Ile Pro Ser Ala Lys Ala Ser Ile Leu His1 5
10 1510818PRTInfluenza B virus 108Lys Cys Thr
Gly Thr Ile Pro Ser Ala Lys Ala Ser Ile Leu His Glu1 5
10 15Val Lys10916PRTInfluenza B virus
109Lys Tyr Gly Gly Leu Asn Lys Ser Lys Pro Tyr Tyr Thr Gly Glu His1
5 10 1511026PRTInfluenza B
virus 110Lys Val Trp Cys Ala Ser Gly Arg Ser Lys Val Ile Lys Gly Ser Leu1
5 10 15Pro Leu Ile Gly
Glu Ala Asp Cys Leu His20 2511110PRTInfluenza B virus
111Lys Pro Tyr Tyr Thr Gly Glu His Ala Lys1 5
1011218PRTInfluenza B virus 112Lys Cys Met Gly Thr Ile Pro Ser Ala
Lys Ala Ser Ile Leu His Glu1 5 10
15Val Lys11315PRTInfluenza B virus 113His Asn Val Ile Asn Ala
Glu Lys Ala Pro Gly Gly Pro Tyr Lys1 5 10
1511416PRTInfluenza B virus 114His Ser Asp Asn Glu Thr
Gln Met Ala Lys Leu Tyr Gly Asp Ser Lys1 5
10 1511518PRTInfluenza B virus 115His Gly Val Ala Val
Ala Ala Asp Leu Lys Ser Thr Gln Glu Ala Ile1 5
10 15Asn Lys11629PRTInfluenza B virus 116His Gly
Val Ala Val Ala Ala Asp Leu Lys Ser Thr Gln Glu Ala Ile1 5
10 15Asn Lys Asp Thr Ile Ser Thr Gln
Glu Ala Ile Asn Lys20 2511721PRTInfluenza B virus 117Lys
Leu Tyr Gly Asp Ser Lys Pro Gln Lys Phe Thr Ser Ser Ala Asn1
5 10 15Gly Val Thr Thr
His2011819PRTInfluenza B virus 118His Ser Asp Asn Glu Thr Gln Met Ala Lys
Leu Tyr Gly Asp Ser Lys1 5 10
15Pro Gln Lys11913PRTInfluenza B virus 119His Phe Ala Asn Leu Lys
Gly Thr Gln Thr Arg Gly Lys1 5
1012012PRTInfluenza B virus 120Lys Pro Arg Ser Ala Leu Lys Cys Lys Gly
Phe His1 5 1012122PRTInfluenza B
virusMOD_RES(15)gly or ala 121Lys Ser Lys Pro Tyr Tyr Thr Gly Glu His Ala
Lys Ala Ile Xaa Asn1 5 10
15Cys Pro Ile Trp Val Lys2012216PRTInfluenza virusMOD_RES(3)val or ile
122His Pro Xaa Thr Ile Gly Glu Cys Pro Lys Tyr Val Xaa Xaa Xaa Lys1
5 10 1512321PRTInfluenza
virusMOD_RES(10)glu or gly 123His Asp Ser Asn Val Lys Asn Leu Tyr Xaa Lys
Val Xaa Xaa Gln Leu1 5 10
15Xaa Asn Asn Ala Lys2012417PRTInfluenza virusMOD_RES(10)glu or gly
124His Asp Ser Asn Val Lys Asn Leu Tyr Xaa Lys Val Xaa Xaa Gln Leu1
5 10 15Lys12536PRTInfluenza
virusMOD_RES(4)..(5)asn or asp 125His Lys Cys Xaa Xaa Xaa Cys Met Glu Ser
Val Xaa Asn Gly Thr Tyr1 5 10
15Asp Tyr Pro Lys Tyr Ser Glu Glu Ser Lys Leu Asn Arg Glu Xaa Ile20
25 30Asp Gly Val Lys3512626PRTInfluenza
virusMOD_RES(4)..(5)asn or asp 126His Lys Cys Xaa Xaa Xaa Cys Met Glu Ser
Val Xaa Asn Gly Thr Tyr1 5 10
15Asp Tyr Pro Lys Tyr Ser Glu Glu Ser Lys20
2512750PRTInfluenza virusMOD_RES(4)glu or gly 127His Gln Asn Xaa Gln Gly
Ser Gly Tyr Ala Ala Asp Gln Lys Ser Thr1 5
10 15Gln Asn Ala Ile Xaa Gly Ile Thr Asn Lys Val Asn
Ser Val Ile Glu20 25 30Lys Met Asn Thr
Gln Phe Thr Ala Val Gly Lys Glu Phe Asn Lys Leu35 40
45Glu Lys5012833PRTInfluenza virusMOD_RES(4)glu or gly
128His Gln Asn Xaa Gln Gly Ser Gly Tyr Ala Ala Asp Gln Lys Ser Thr1
5 10 15Gln Asn Ala Ile Xaa Gly
Ile Thr Asn Lys Val Asn Ser Val Ile Glu20 25
30Lys12926PRTInfluenza virusMOD_RES(4)glu or gly 129His Gln Asn Xaa
Gln Gly Ser Gly Tyr Ala Ala Asp Gln Lys Ser Thr1 5
10 15Gln Asn Ala Ile Xaa Gly Ile Thr Asn Lys20
2513014PRTInfluenza virus 130Lys Phe Glu Ile Phe Pro Lys
Thr Ser Ser Trp Pro Asn His1 5
1013127PRTInfluenza virusMOD_RES(3)asn, ser or thr 131Lys Gly Xaa Ser Tyr
Pro Lys Leu Xaa Lys Ser Tyr Xaa Asn Asn Lys1 5
10 15Gly Lys Glu Val Leu Val Leu Trp Gly Val His20
2513218PRTInfluenza virusMOD_RES(4)val or thr 132Lys Ser
Tyr Xaa Asn Asn Lys Gly Lys Glu Val Leu Val Leu Trp Gly1 5
10 15Val His13336PRTInfluenza virus
133His Lys Cys Asn Asn Glu Cys Met Glu Ser Val Lys Asn Gly Thr Tyr1
5 10 15Asp Tyr Pro Lys Tyr Ser
Glu Glu Ser Lys Leu Asn Arg Glu Lys Ile20 25
30Asp Gly Val Lys3513426PRTInfluenza virus 134His Lys Cys Asn Asn
Glu Cys Met Glu Ser Val Lys Asn Gly Thr Tyr1 5
10 15Asp Tyr Pro Lys Tyr Ser Glu Glu Ser Lys20
2513520PRTInfluenza virus 135His Lys Cys Asn Asn Glu Cys Met
Glu Ser Val Lys Asn Gly Thr Tyr1 5 10
15Asp Tyr Pro Lys2013612PRTInfluenza virus 136His Lys Cys
Asn Asn Glu Cys Met Glu Ser Val Lys1 5
1013734PRTInfluenza virusMOD_RES(9)lys or arg 137His Asn Gly Lys Ser Ser
Phe Tyr Xaa Asn Leu Leu Trp Leu Thr Xaa1 5
10 15Lys Asn Gly Leu Tyr Pro Asn Leu Ser Lys Ser Tyr
Val Asn Asn Lys20 25 30Glu
Lys13832PRTInfluenza virusMOD_RES(9)lys or arg 138His Asn Gly Lys Ser Ser
Phe Tyr Xaa Asn Leu Leu Trp Leu Thr Xaa1 5
10 15Lys Asn Gly Leu Tyr Pro Asn Leu Ser Lys Ser Tyr
Val Asn Asn Lys20 25
3013926PRTInfluenza virusMOD_RES(9)lys or arg 139His Asn Gly Lys Ser Ser
Phe Tyr Xaa Asn Leu Leu Trp Leu Thr Xaa1 5
10 15Lys Asn Gly Leu Tyr Pro Asn Leu Ser Lys20
2514017PRTInfluenza virusMOD_RES(9)lys or arg 140His Asn Gly Lys
Ser Ser Phe Tyr Xaa Asn Leu Leu Trp Leu Thr Xaa1 5
10 15Lys14140PRTInfluenza virus 141Lys Ser Ser
Phe Tyr Lys Asn Leu Leu Trp Leu Thr Glu Lys Asn Gly1 5
10 15Leu Tyr Pro Asn Leu Ser Lys Ser Tyr
Val Asn Asn Lys Glu Lys Glu20 25 30Val
Leu Val Leu Trp Gly Val His35 4014235PRTInfluenza virus
142Lys Asn Leu Leu Trp Leu Thr Glu Lys Asn Gly Leu Tyr Pro Asn Leu1
5 10 15Ser Lys Ser Tyr Val Asn
Asn Lys Glu Lys Glu Val Leu Val Leu Trp20 25
30Gly Val His3514327PRTInfluenza virus 143Lys Asn Gly Leu Tyr Pro
Asn Leu Ser Lys Ser Tyr Val Asn Asn Lys1 5
10 15Glu Lys Glu Val Leu Val Leu Trp Gly Val His20
2514418PRTInfluenza virusMOD_RES(4)val or ala 144Lys Ser Tyr
Xaa Asn Asn Lys Glu Lys Glu Val Xaa Xaa Leu Trp Gly1 5
10 15Val His14512PRTInfluenza virus 145Lys
Glu Ser Ser Trp Pro Asn His Thr Val Thr Lys1 5
1014644PRTInfluenza virusMOD_RES(4)thr or asn 146His Glu Thr Xaa Lys
Gly Val Thr Ala Ala Cys Pro Tyr Ala Gly Ala1 5
10 15Ser Ser Phe Tyr Arg Asn Leu Leu Trp Leu Val
Lys Lys Glu Asn Ser20 25 30Tyr Pro Lys
Leu Ser Lys Ser Tyr Val Asn Asn Lys35
4014738PRTInfluenza virusMOD_RES(4)thr or asn 147His Glu Thr Xaa Lys Gly
Val Thr Ala Ala Cys Pro Tyr Ala Gly Ala1 5
10 15Ser Ser Phe Tyr Arg Asn Leu Leu Trp Leu Val Lys
Lys Glu Asn Ser20 25 30Tyr Pro Lys Leu
Ser Lys3514822PRTInfluenza virus 148Lys Phe Glu Ile Phe Pro Lys Thr Ser
Ser Trp Pro Asn Glu Val Leu1 5 10
15Val Leu Trp Gly Val His201498PRTInfluenza virus 149Lys Glu Arg
Ser Trp Pro Lys His1 515021PRTInfluenza virus 150Lys Leu
Ser Lys Ser Tyr Val Asn Asn Lys Glu Lys Glu Val Leu Val1 5
10 15Leu Trp Gln Val
His2015115PRTInfluenza virus 151Lys Asn Asn Lys Glu Lys Glu Val Leu Val
Leu Trp Gln Val His1 5 10
1515234PRTInfluenza virusMOD_RES(2)lys or asn 152His Xaa Xaa Lys Ser Ser
Phe Tyr Xaa Asn Leu Leu Trp Leu Thr Glu1 5
10 15Lys Asn Gly Xaa Tyr Pro Xaa Leu Ser Lys Ser Tyr
Ala Asn Asn Lys20 25 30Glu
Lys15317PRTInfluenza virusMOD_RES(2)lys or asn 153His Xaa Xaa Lys Ser Ser
Phe Tyr Xaa Asn Leu Leu Trp Leu Thr Glu1 5
10 15Lys1549PRTInfluenza virus 154His Ala Lys Lys Ser
Ser Phe Tyr Lys1 515511PRTInfluenza virus 155His Asn Gly
Lys Leu Cys Arg Leu Lys Gly Lys1 5
101569PRTInfluenza virusMOD_RES(7)gln or gly 156His Tyr Lys Leu Asn Asn
Xaa Lys Lys1 515725PRTInfluenza virus 157His Asp Ile Tyr
Arg Asp Glu Ala Ile Asn Asn Arg Phe Gln Ile Gln1 5
10 15Gly Val Lys Leu Thr Gln Gly Tyr Lys20
2515811PRTInfluenza virus 158Lys Gly Asn Gly Cys Phe Glu Ile
Phe His Lys1 5 1015918PRTInfluenza virus
159Lys Leu Asn Arg Leu Ile Glu Lys Thr Asn Asp Lys Tyr His Gln Ile1
5 10 15Glu
Lys16014PRTInfluenza virus 160Lys Leu Asn Arg Leu Ile Glu Lys Thr Asn Asp
Lys Tyr His1 5 1016113PRTInfluenza virus
161Lys Cys His Thr Asp Lys Gly Ser Leu Ser Thr Thr Lys1 5
1016216PRTInfluenza virus 162Lys Ile Asn Asn Gly Asp Tyr
Ala Lys Leu Tyr Ile Trp Gly Val His1 5 10
1516317PRTInfluenza virus 163His Asn Gly Lys Leu Cys Arg
Lys Gly Ile Ala Pro Leu Gln Leu Gly1 5 10
15Lys16438PRTInfluenza virus 164His Glu Thr Asn Arg Gln
Val Thr Ala Ala Cys Pro Tyr Ala Gly Ala1 5
10 15Asn Ser Phe Phe Arg Asn Leu Ile Trp Leu Val Lys
Lys Glu Ser Ser20 25 30Tyr Pro Lys Leu
Ser Lys3516535PRTInfluenza virus 165His Glu Thr Asn Arg Gln Val Thr Ala
Ala Cys Pro Tyr Ala Gly Ala1 5 10
15Asn Ser Phe Phe Arg Asn Leu Ile Trp Leu Val Lys Lys Glu Ser
Ser20 25 30Tyr Pro
Lys3516631PRTInfluenza virus 166His Pro Pro Thr Ser Thr Asp Gln Gln Ser
Leu Tyr Gln Asn Ala Asp1 5 10
15Ala Tyr Ile Phe Val Gly Ser Ser Lys Tyr Asn Arg Lys Phe Lys20
25 3016735PRTInfluenza virus 167His Pro Pro
Thr Ser Thr Asp Gln Gln Ser Leu Tyr Gln Asn Ala Asp1 5
10 15Ala Tyr Ile Phe Val Gly Ser Ser Lys
Tyr Asn Arg Lys Phe Lys Pro20 25 30Glu
Ile Ala3516825PRTInfluenza virus 168His Asp Ile Tyr Arg Asp Glu Ala Ile
Asn Asn Arg Phe Gln Ile Gln1 5 10
15Gly Val Lys Ile Thr Gln Gly Tyr Lys20
2516943PRTInfluenza virus 169His Gln Asn Glu Gln Gly Ser Gly Tyr Ala Ala
Asp Gln Lys Ser Thr1 5 10
15Gln Asn Ala Ile Asp Gly Ile Thr Asn Lys Val Asn Ser Val Ile Glu20
25 30Lys Met Asn Thr Gln Phe Thr Ala Val Gly
Lys35 4017033PRTInfluenza virus 170His Gln Asn Glu Gln
Gly Ser Gly Tyr Ala Ala Asp Gln Lys Ser Thr1 5
10 15Gln Asn Ala Ile Asp Gly Ile Thr Asn Lys Val
Asn Ser Val Ile Glu20 25
30Lys17150PRTInfluenza virus 171His Gln Asn Glu Gln Gly Ser Gly Tyr Ala
Ala Asp Gln Lys Ser Thr1 5 10
15Gln Asn Ala Ile Asn Gly Ile Thr Asn Lys Val Asn Ser Val Ile Glu20
25 30Lys Met Asn Thr Gln Phe Thr Ala Val
Gly Lys Glu Phe Asn Lys Leu35 40 45Glu
Lys5017218PRTInfluenza virus 172His Asn Gly Lys Leu Cys Arg Leu Lys Gly
Ile Ala Pro Leu Gln Leu1 5 10
15Gly Lys17312PRTInfluenza virus 173His Lys Cys Asn Asn Glu Cys Met
Glu Ser Val Lys1 5 1017414PRTInfluenza
virus 174Lys Phe Glu Ile Phe Pro Lys Ala Ser Ser Trp Pro Asn His1
5 1017521PRTInfluenza virus 175His Asp Ser Asn
Val Lys Asn Leu Tyr Glu Lys Val Arg Ser Gln Leu1 5
10 15Arg Asn Asn Ala Lys2017622PRTInfluenza
virus 176Lys Val Asn Ser Val Ile Lys Lys Met Asn Thr Gln Phe Ala Ala Val1
5 10 15Gly Lys Glu Phe
Asn His201778PRTInfluenza virus 177Lys His Asn Gly Lys Leu Cys Lys1
517828PRTInfluenza virus 178Lys Lys Gly Thr Ser Tyr Pro Lys Leu
Ser Lys Ser Tyr Thr His Asn1 5 10
15Lys Gly Lys Glu Val Leu Val Leu Trp Gly Val His20
2517927PRTInfluenza virus 179Lys Gly Thr Ser Tyr Pro Lys Leu Ser Lys
Ser Tyr Thr His Asn Lys1 5 10
15Gly Lys Glu Val Leu Val Leu Trp Gly Val His20
2518021PRTInfluenza virus 180Lys Leu Ser Lys Ser Tyr Thr His Asn Lys Gly
Lys Glu Val Leu Val1 5 10
15Leu Trp Gly Val His2018118PRTInfluenza virus 181Lys Ser Tyr Thr His
Asn Lys Gly Lys Glu Val Leu Val Leu Trp Gly1 5
10 15Val His18210PRTInfluenza virus 182Lys Gly Val
Thr Ala Ser Cys Ser His Lys1 5
1018334PRTInfluenza virus 183Lys Gly Val Thr Ala Ser Cys Ser His Lys Gly
Arg Ser Ser Phe Tyr1 5 10
15Arg Asn Leu Leu Trp Leu Thr Glu Lys Asn Gly Leu Tyr Pro Asn Leu20
25 30Ser Lys18427PRTInfluenza virus 184Lys
Gly Asn Ser Tyr Pro Lys Leu Ser Lys Ser Tyr Val Asn Asn Lys1
5 10 15Glu Lys Glu Val Leu Val Leu
Trp Gly Ile His20 251858PRTInfluenza virus 185Lys Glu
Phe Asn His Leu Glu Lys1 518639PRTInfluenza virus 186His
Pro Pro Thr Ser Thr Asp Gln Gln Ser Leu Tyr Gln Asn Ala Asp1
5 10 15Ala Tyr Val Phe Val Gly Ser
Ser Lys Tyr Asn Lys Lys Phe Lys Pro20 25
30Glu Ile Ala Thr Arg Pro Lys3518731PRTInfluenza virus 187His Pro Pro
Thr Ser Thr Asp Gln Gln Ser Leu Tyr Gln Asn Ala Asp1 5
10 15Ala Tyr Val Phe Val Gly Ser Ser Lys
Tyr Asn Lys Lys Phe Lys20 25
3018831PRTInfluenza virus 188His Glu Gly Lys Ser Ser Phe Tyr Arg Asn Leu
Leu Trp Leu Thr Glu1 5 10
15Lys Glu Gly Ser Tyr Pro Lys Leu Lys Asn Ser Tyr Val Asn Lys20
25 3018923PRTInfluenza virus 189His Glu Gly Lys
Ser Ser Phe Tyr Arg Asn Leu Leu Trp Leu Thr Glu1 5
10 15Lys Glu Gly Ser Tyr Pro
Lys2019026PRTInfluenza virus 190His Lys Cys Asp Asn Glu Cys Met Glu Ser
Val Arg Asn Gly Thr Tyr1 5 10
15Asp Tyr Pro Lys Tyr Ser Glu Glu Ser Lys20
2519112PRTInfluenza virus 191Lys Glu Ser Ser Trp Pro Asn His Thr Val Thr
Lys1 5 1019235PRTInfluenza virus 192Lys
Asn Leu Leu Trp Leu Thr Glu Lys Asn Gly Leu Tyr Pro Asn Leu1
5 10 15Ser Lys Ser Tyr Val Asn Asn
Lys Glu Lys Glu Ile Leu Val Leu Trp20 25
30Gly Val His3519327PRTInfluenza virusMOD_RES(9)lys or met 193His Asn
Gly Lys Ser Ser Phe Tyr Xaa Xaa Leu Leu Trp Leu Thr Xaa1 5
10 15Xaa Lys Asn Gly Leu Tyr Pro Asn
Leu Ser Lys20 2519417PRTInfluenza virus 194His Asn Gly
Lys Ser Ser Phe Tyr Lys Asn Leu Leu Trp Leu Thr Glu1 5
10 15Lys19555PRTInfluenza virus 195His Thr
Val Thr Lys Gly Val Thr Ala Ser Cys Ser His Asn Gly Lys1 5
10 15Ser Ser Phe Tyr Lys Asn Leu Leu
Trp Leu Thr Glu Lys Asn Gly Leu20 25
30Tyr Pro Asn Leu Ser Lys Ser Tyr Val Asn Asn Lys Glu Lys Glu Val35
40 45Leu Val Leu Trp Gly Val His50
5519638PRTInfluenza virusMOD_RES(5)lys or gly 196His Thr Val Thr
Xaa Gly Val Xaa Ala Ser Cys Ser His Asn Gly Lys1 5
10 15Ser Ser Phe Tyr Xaa Xaa Leu Leu Trp Leu
Thr Xaa Lys Xaa Gly Leu20 25 30Tyr Pro
Asn Leu Ser Lys3519729PRTInfluenza virus 197His Thr Val Thr Lys Gly Val
Thr Ala Ser Cys Ser His Asn Gly Lys1 5 10
15Ser Ser Phe Tyr Lys Asn Leu Leu Trp Leu Thr Glu Lys20
2519848PRTInfluenza virus 198Lys Tyr Val Arg Ser Thr Lys
Leu Arg Met Val Thr Gly Leu Arg Asn1 5 10
15Ile Pro Ser Ile Gln Ser Arg Gly Leu Phe Gly Ala Ile
Ala Gly Phe20 25 30Ile Glu Gly Gly Trp
Thr Gly Met Ile Asp Gly Trp Tyr Gly Tyr His35 40
4519943PRTInfluenza virus 199His Gln Asn Glu Gln Gly Ser Gly Tyr
Ala Ala Asp Gln Lys Ser Thr1 5 10
15Gln Asn Ala Ile Asn Gly Ile Thr Asn Lys Val Asn Ser Ile Ile
Glu20 25 30Lys Met Asn Thr Gln Phe Thr
Ala Val Gly Lys35 4020033PRTInfluenza virus 200His Gln
Asn Glu Gln Gly Ser Gly Tyr Ala Ala Asp Gln Lys Ser Thr1 5
10 15Gln Asn Ala Ile Asn Gly Ile Thr
Asn Lys Val Asn Ser Ile Ile Glu20 25
30Lys20126PRTInfluenza virus 201His Gln Asn Glu Gln Gly Ser Gly Tyr Ala
Ala Asp Gln Lys Ser Thr1 5 10
15Gln Asn Ala Ile Asn Gly Ile Thr Asn Lys20
2520223PRTInfluenza virus 202His Ser Gly Ala Arg Ser Phe Tyr Arg Asn Leu
Leu Trp Ile Val Lys1 5 10
15Lys Gly Asn Ser Tyr Pro Lys2020326PRTInfluenza virus 203His Ser Gly
Ala Arg Ser Phe Tyr Arg Asn Leu Leu Trp Ile Val Lys1 5
10 15Lys Gly Asn Ser Tyr Pro Lys Leu Asn
Lys20 2520432PRTInfluenza virus 204His Ser Gly Ala Arg
Ser Phe Tyr Arg Asn Leu Leu Trp Ile Val Lys1 5
10 15Lys Gly Asn Ser Tyr Pro Lys Leu Asn Lys Ser
Tyr Thr Asn Asp Lys20 25
3020534PRTInfluenza virus 205His Ser Gly Ala Arg Ser Phe Tyr Arg Asn Leu
Leu Trp Ile Val Lys1 5 10
15Lys Gly Asn Ser Tyr Pro Lys Leu Asn Lys Ser Tyr Thr Asn Asp Lys20
25 30Gly Lys20616PRTInfluenza virus 206His
Thr Val Ser Lys Gly Val Thr Thr Ser Cys Ser His Asn Gly Lys1
5 10 1520712PRTInfluenza virus 207Lys
Ala Thr Ser Trp Pro Asn His Glu Thr Thr Lys1 5
1020812PRTInfluenza virus 208Lys Gln Val Thr Thr Ser Cys Ser His Asn
Gln Lys1 5 1020927PRTInfluenza virus
209Lys Gly Asn Ser Tyr Pro Lys Leu Asn Lys Ser Tyr Thr Asn Asp Lys1
5 10 15Gly Lys Glu Val Leu Val
Ile Trp Gly Val His20 2521021PRTInfluenza virus 210Lys
Leu Asn Lys Ser Tyr Thr Asn Asp Lys Gly Lys Glu Val Leu Val1
5 10 15Ile Trp Gly Val
His2021118PRTInfluenza virus 211Lys Ser Tyr Thr Asn Asp Lys Gly Lys Glu
Val Leu Val Ile Trp Gly1 5 10
15Val His21235PRTInfluenza virusMOD_RES(16)glu or gln 212His Asn Gln
Lys Ser Ser Phe Tyr Arg Asn Leu Leu Trp Leu Thr Xaa1 5
10 15Lys Asn Gly Leu Tyr Pro Asn Leu Ser
Lys Ser Tyr Xaa Ala Asn Asn20 25 30Lys
Glu Lys3521316PRTInfluenza virus 213His Pro Ile Thr Ile Gly Glu Cys Pro
Lys Tyr Val Arg Ser Ala Lys1 5 10
1521443PRTInfluenza virus 214His Gln Asn Glu Gln Gly Ser Gly Tyr
Ala Ala Asp Gln Lys Ser Thr1 5 10
15Gln Asn Ala Ile Asn Gly Ile Thr Asn Lys Val Asn Ser Val Ile
Glu20 25 30Lys Met Asn Thr Gln Phe Thr
Ala Val Gly Lys35 4021533PRTInfluenza virus 215His Gln
Asn Glu Gln Gly Ser Gly Tyr Ala Ala Asp Gln Lys Ser Thr1 5
10 15Gln Asn Ala Ile Asn Gly Ile Thr
Asn Lys Val Asn Ser Val Ile Glu20 25
30Lys21634PRTInfluenza virus 216His Asn Gly Lys Ser Ser Phe Tyr Arg Asn
Leu Leu Trp Leu Thr Glu1 5 10
15Lys Asn Gly Leu Tyr Pro Asn Leu Ser Lys Ser Tyr Val Asn Asn Lys20
25 30Glu Lys21711PRTInfluenza virus
217Lys His Phe Glu Lys Val Lys Ile Leu Pro Lys1 5
1021814PRTInfluenza virus 218Lys His Leu Leu Ser Ser Val Lys His
Phe Glu Lys Val Lys1 5
1021913PRTInfluenza virusMOD_RES(3)lys, gln or met 219His Ala Xaa Xaa Ile
Leu Glu Lys Thr His Asn Gly Lys1 5
1022016PRTInfluenza virusMOD_RES(3)lys, gln or met 220His Ala Xaa Xaa Ile
Leu Glu Lys Thr His Asn Gly Lys Leu Cys Xaa1 5
10 1522119PRTInfluenza virus 221His Asn Val His Pro
Leu Thr Ile Gly Glu Cys Pro Lys Tyr Val Lys1 5
10 15Ser Glu Lys22216PRTInfluenza virus 222His Pro
Leu Thr Ile Gly Glu Cys Pro Lys Tyr Val Lys Ser Glu Lys1 5
10 1522318PRTInfluenza virus 223Lys His
Leu Leu Ser Ser Val Lys His Phe Glu Lys Val Lys Ile Leu1 5
10 15Pro Lys22438PRTInfluenza virus
224Lys Arg Gln Ser Ser Gly Ile Met Lys Thr Glu Gly Thr Leu Glu Asn1
5 10 15Cys Glu Thr Lys Cys Gln
Thr Pro Leu Gly Ala Ile Asn Thr Thr Leu20 25
30Pro Phe His Asn Val His3522527PRTInfluenza virusMOD_RES(7)val or
ile 225Lys Gly Ser Asn Tyr Pro Xaa Ala Lys Xaa Ser Tyr Asn Asn Thr Ser1
5 10 15Gly Glu Gln Met Leu
Ile Ile Trp Gln Xaa His20 2522636PRTInfluenza virus
226His Thr Thr Leu Gly Gln Ser Arg Ala Cys Ala Val Ser Gly Asn Pro1
5 10 15Ser Phe Phe Arg Asn Met
Val Trp Leu Thr Glu Lys Gly Ser Asn Tyr20 25
30Pro Val Ala Lys352277PRTInfluenza virus 227Lys His Phe Glu Lys Val
Lys1 522838PRTInfluenza virus 228Lys Ile Ser Lys Arg Gly
Ser Ser Gly Ile Met Lys Thr Glu Gly Thr1 5
10 15Leu Glu Asn Cys Glu Thr Lys Cys Gln Thr Pro Leu
Gly Ala Ile Asn20 25 30Thr Thr Leu Pro
Phe His3522935PRTInfluenza virus 229Lys Arg Gly Ser Ser Gly Ile Met Lys
Thr Glu Gly Thr Leu Glu Asn1 5 10
15Cys Glu Thr Lys Cys Gln Thr Pro Leu Gly Ala Ile Asn Thr Thr
Leu20 25 30Pro Phe
His3523027PRTInfluenza virus 230Lys Thr Glu Gly Thr Leu Glu Asn Cys Glu
Thr Lys Cys Gln Thr Pro1 5 10
15Leu Gly Ala Ile Asn Thr Thr Leu Pro Phe His20
2523138PRTInfluenza virus 231Lys Ile Ser Lys Arg Gly Ser Ser Gly Ile Met
Lys Thr Glu Gly Thr1 5 10
15Leu Glu Asn Cys Glu Thr Lys Cys Gln Thr Pro Leu Gly Ala Ile Asn20
25 30Thr Thr Leu Pro Phe
His3523230PRTInfluenza virusMOD_RES(29)val or ile 232Lys Thr Glu Gly Thr
Leu Glu Asn Cys Glu Thr Lys Cys Gln Thr Pro1 5
10 15Leu Gly Ala Ile Asn Thr Thr Leu Pro Phe His
Asn Xaa His20 25 3023338PRTInfluenza
virus 233Lys Ile Ser Lys Arg Gly Ser Ser Gly Ile Met Lys Thr Glu Gly Thr1
5 10 15Leu Glu Asn Cys
Glu Thr Lys Cys Gln Thr Pro Leu Gly Ala Ile Asn20 25
30Thr Thr Leu Pro Phe His3523427PRTInfluenza
virusMOD_RES(2)glu or gly 234Lys Xaa Ser Asn Tyr Pro Val Ala Lys Gly Ser
Tyr Asn Asn Thr Ser1 5 10
15Gly Glu Gln Met Leu Ile Ile Trp Gly Val His20
2523516PRTInfluenza virus 235His Pro Leu Thr Ile Gly Glu Cys Pro Lys Tyr
Val Lys Ser Glu Lys1 5 10
1523616PRTInfluenza virus 236Lys Cys Gln Thr Pro Leu Gly Ala Ile Lys Thr
Thr Leu Pro Phe His1 5 10
1523758PRTInfluenza virusMOD_RES(21)phe or ile 237His His Ser Asn Asp
Gln Gly Ser Gly Tyr Ala Ala Asp Lys Glu Ser1 5
10 15Thr Gln Lys Ala Xaa Asp Gly Ile Thr Asn Lys
Val Asn Ser Val Ile20 25 30Glu Lys Met
Asn Thr Gln Phe Glu Ala Val Gly Lys Leu Phe Xaa Asn35 40
45Leu Glu Lys Leu Glu Asn Leu Asn Lys Lys50
5523857PRTInfluenza virusMOD_RES(20)phe or ile 238His Ser Asn Asp Gln
Gly Ser Gly Tyr Ala Ala Asp Lys Glu Ser Thr1 5
10 15Gln Lys Ala Xaa Asp Gly Ile Thr Asn Lys Val
Asn Ser Val Ile Glu20 25 30Lys Met Asn
Thr Gln Phe Glu Ala Val Gly Lys Leu Phe Xaa Asn Leu35 40
45Glu Lys Leu Glu Asn Leu Asn Lys Lys50
5523926PRTInfluenza virusMOD_RES(20)phe or ile 239His Ser Asn Asp Gln Gly
Ser Gly Tyr Ala Ala Asp Lys Glu Ser Thr1 5
10 15Gln Lys Ala Xaa Asp Gly Ile Thr Asn Lys20
2524021PRTInfluenza virus 240His Asp Ser Asn Val Arg Asn Leu Tyr
Asp Lys Val Arg Met Gln Leu1 5 10
15Arg Asp Asn Ala Lys2024130PRTInfluenza virus 241His Lys Cys
Asp Asp Glu Cys Met Asn Ser Val Lys Asn Gly Thr Tyr1 5
10 15Asp Tyr Pro Lys Leu Asn Arg Asn Glu
Ile Lys Gly Val Lys20 25
3024227PRTInfluenza virus 242His Lys Cys Asp Asp Glu Cys Met Asn Ser Val
Lys Asn Gly Thr Tyr1 5 10
15Asp Tyr Pro Lys Leu Asn Arg Asn Glu Ile Lys20
2524320PRTInfluenza virus 243His Lys Cys Asp Asp Glu Cys Met Asn Ser Val
Lys Asn Gly Thr Tyr1 5 10
15Asp Tyr Pro Lys2024412PRTInfluenza virus 244His Lys Cys Asp Asp Glu
Cys Met Asn Ser Val Lys1 5
1024527PRTInfluenza virus 245Lys Gly Ser Asn Tyr Pro Val Ala Lys Gly Ser
Tyr Asn Asn Thr Asn1 5 10
15Gly Glu Gln Ile Leu Ile Ile Trp Gly Val His20
2524643PRTInfluenza virus 246His Ser Asn Asp Gln Gly Ser Gly Tyr Ala Ala
Asp Lys Glu Ser Thr1 5 10
15Gln Lys Ala Val Asp Gly Ile Thr Asn Lys Val Asn Ser Val Ile Glu20
25 30Lys Met Asn Thr Gln Phe Glu Ala Val Gly
Lys35 4024735PRTInfluenza virus 247Lys Arg Gly Ser Ser
Gly Ile Met Lys Thr Glu Gly Thr Leu Glu Asn1 5
10 15Cys Glu Thr Lys Cys Gln Thr Pro Leu Gly Ala
Ile Asn Thr Thr Leu20 25 30Pro Phe
His3524816PRTInfluenza virus 248His Pro Leu Thr Ile Gly Glu Cys Pro Lys
Tyr Val Lys Ser Glu Lys1 5 10
1524916PRTInfluenza virus 249His Ala Lys Asp Ile Leu Glu Lys Thr His
Asn Gly Lys Leu Cys Lys1 5 10
1525025PRTInfluenza virus 250His Asp Val Tyr Arg Asp Glu Ala Leu Asn
Asn Arg Phe Gln Ile Lys1 5 10
15Gly Val Glu Leu Lys Ser Gly Tyr Lys20
2525119PRTInfluenza virus 251His Thr Ile Asp Leu Thr Asp Ser Glu Met Asn
Lys Leu Phe Glu Arg1 5 10
15Thr Arg Lys2527PRTInfluenza virus 252Lys Phe His Gln Ile Glu Lys1
525311PRTInfluenza virusMOD_RES(8)gly or gln 253Lys Thr Asn Glu
Lys Phe His Xaa Ile Glu Lys1 5
1025414PRTInfluenza virusMOD_RES(5)val or leu 254Lys Leu Asn Arg Xaa Ile
Glu Lys Thr Asn Glu Lys Phe His1 5
1025525PRTInfluenza virus 255His Gln Ile Glu Lys Glu Phe Ser Glu Val Glu
Gly Arg Ile Gln Asp1 5 10
15Leu Glu Lys Tyr Val Glu Asp Thr Lys20
252568PRTInfluenza virus 256Lys Ile Cys Asn Asn Pro His Lys1
525714PRTInfluenza virus 257Lys Leu Asn Arg Val Ile Lys Lys Thr Asn Glu
Lys Phe His1 5 1025824PRTInfluenza
virusMOD_RES(3)ile or val 258His Asp Xaa Tyr Arg Asp Glu Ala Leu Asn Asn
Arg Phe Gln Ile Lys1 5 10
15Xaa Val Glu Xaa Ser Xaa Tyr Lys2025925PRTInfluenza virus 259His Gln
Ile Glu Lys Glu Phe Ser Glu Val Glu Gly Arg Ile Gln Asp1 5
10 15Leu Glu Lys Tyr Val Glu Asp Thr
Lys20 2526025PRTInfluenza virus 260Lys Tyr Val Glu Asp
Thr Lys Ile Asp Leu Trp Ser Tyr Asn Ala Glu1 5
10 15Leu Leu Val Ala Leu Glu Asn Gln His20
2526149PRTInfluenza virus 261Lys Tyr Val Lys Gln Asn Ser Leu Lys
Leu Ala Thr Gly Met Arg Asn1 5 10
15Val Pro Glu Lys Gln Thr Arg Gly Leu Phe Gly Ala Ile Ala Gly
Phe20 25 30Ile Glu Asn Gly Trp Glu Gly
Met Ile Asp Gly Trp Tyr Gly Phe Arg35 40
45His26239PRTInfluenza virus 262Lys Glu Phe Ser Glu Val Glu Gly Arg Ile
Gln Asp Leu Glu Lys Tyr1 5 10
15Val Glu Asp Thr Lys Ile Asp Leu Trp Ser Tyr Asn Ala Glu Leu Leu20
25 30Val Ala Leu Glu Asn Gln
His3526333PRTInfluenza virusMOD_RES(4)ser or glu 263His Gln Asn Xaa Xaa
Gly Xaa Gly Xaa Ala Ala Asp Xaa Lys Ser Thr1 5
10 15Gln Xaa Ala Xaa Asp Xaa Ile Xaa Xaa Lys Xaa
Asn Xaa Val Ile Xaa20 25
30Lys26418PRTInfluenza virusMOD_RES(4)gly or gln 264His Cys Asp Xaa Phe
Xaa Asn Glu Lys Trp Asp Leu Phe Xaa Glu Arg1 5
10 15Xaa Lys26520PRTInfluenza virus 265His Thr Ile
Asp Leu Thr Asp Ser Glu Met Asn Lys Lys Leu Phe Glu1 5
10 15Arg Thr Arg Lys2026628PRTInfluenza
virus 266Lys Ser Gly Ser Thr Tyr Pro Val Leu Lys Val Thr Met Pro Asn Asn1
5 10 15Asp Asn Phe Asp
Lys Leu Tyr Ile Trp Gly Val His20 2526734PRTInfluenza
virus 267Lys Leu Asn Trp Leu Thr Lys Ser Gly Asn Thr Tyr Pro Val Leu Asn1
5 10 15Val Thr Met Pro
Asn Asn Asp Asn Phe Asp Lys Leu Val Ile Trp Gly20 25
30Val His26819PRTInfluenza virus 268His Thr Ile Asp Leu Thr
Asp Ser Glu Met Asn Lys Leu Phe Glu Lys1 5
10 15Thr Arg Lys26918PRTInfluenza virus 269Lys Leu Asn
Arg Leu Ile Glu Lys Thr Asn Glu Lys Phe His Gln Thr1 5
10 15Glu Lys27047PRTInfluenza virus 270His
Thr Gly Lys Ser Ser Val Met Arg Ser Asp Ala Pro Ile Asp Phe1
5 10 15Cys Asn Ser Glu Cys Ile Thr
Pro Asn Gln Ser Ile Pro Asn Asp Lys20 25
30Pro Phe Gln Asn Val Asn Lys Ile Thr Tyr Gly Ala Cys Pro Lys35
40 4527139PRTInfluenza virus 271His Thr Gly Lys
Ser Ser Val Met Arg Ser Asp Ala Pro Ile Asp Phe1 5
10 15Cys Asn Ser Glu Cys Ile Thr Pro Asn Gln
Ser Ile Pro Asn Asp Lys20 25 30Pro Phe
Gln Asn Val Asn Lys3527233PRTInfluenza virus 272His Pro Ser Thr Asp Ser
Asp Gln Thr Ser Leu Tyr Val Arg Ala Ser1 5
10 15Gly Arg Val Thr Val Ser Thr Lys Arg Ser Gln Gln
Thr Val Ile Pro20 25
30Lys27325PRTInfluenza virus 273Lys Tyr Val Glu Asp Thr Lys Ile Asp Leu
Trp Ser Tyr Asn Ala Glu1 5 10
15Leu Leu Val Ala Leu Glu Asn Gln His20
2527426PRTInfluenza virus 274Lys Leu Phe Glu Arg Thr Arg Lys Gln Leu Arg
Glu Asn Ala Glu Asp1 5 10
15Met Gly Asn Gly Cys Phe Lys Ile Tyr His20
2527516PRTInfluenza virus 275Lys Arg Arg Ser Ile Lys Ser Phe Phe Ser Arg
Leu Asn Trp Leu His1 5 10
1527616PRTInfluenza virusMOD_RES(12)val or arg 276His Pro Val Thr Ile
Gly Glu Cys Pro Lys Tyr Xaa Lys Ser Thr Lys1 5
10 1527730PRTInfluenza virus 277Lys Gly Asn Ser Tyr
Pro Lys Leu Ser Lys Leu Ser Lys Ser Tyr Ile1 5
10 15Ile Asn Lys Lys Lys Glu Val Leu Val Ile Trp
Gly Ile His20 25 3027824PRTInfluenza
virusMOD_RES(9)val or tyr 278Lys Leu Ser Lys Leu Ser Lys Ser Xaa Ile Ile
Asn Lys Lys Lys Glu1 5 10
15Val Leu Val Ile Trp Gly Ile His2027921PRTInfluenza virusMOD_RES(6)val
or tyr 279Lys Leu Ser Lys Ser Xaa Ile Ile Asn Lys Lys Lys Glu Val Leu
Val1 5 10 15Ile Trp Gly
Ile His2028046PRTPlasmodium falciparum 280Lys Glu Glu Glu Glu Lys Glu Lys
Glu Lys Glu Lys Glu Lys Glu Glu1 5 10
15Lys Glu Lys Glu Glu Lys Glu Lys Glu Glu Lys Glu Lys Glu
Lys Glu20 25 30Glu Lys Glu Lys Glu Lys
Glu Glu Lys Glu Glu Glu Lys Lys35 40
4528148PRTPlasmodium falciparum 281Lys Glu Glu Glu Glu Lys Glu Lys Glu
Lys Glu Lys Glu Lys Glu Glu1 5 10
15Lys Glu Lys Glu Glu Lys Glu Lys Glu Glu Lys Glu Lys Glu Lys
Glu20 25 30Glu Lys Glu Lys Glu Lys Glu
Glu Lys Glu Glu Glu Lys Lys Glu Lys35 40
4528247PRTPlasmodium falciparum 282Lys Glu Glu Glu Glu Lys Glu Lys Glu
Lys Glu Lys Glu Lys Glu Glu1 5 10
15Lys Glu Lys Glu Glu Lys Glu Lys Glu Lys Glu Glu Lys Glu Lys
Glu20 25 30Glu Lys Glu Lys Glu Glu Lys
Glu Glu Lys Glu Glu Glu Lys Lys35 40
4528310PRTPlasmodium falciparum 283Lys Glu Glu Glu Glu Lys Glu Lys Glu
Lys1 5 1028419PRTPlasmodium falciparum
284His Lys Lys Leu Ile Lys Ala Leu Lys Lys Asn Ile Glu Ser Ile Gln1
5 10 15Asn Lys
Lys28519PRTPlasmodium falciparum 285His Lys Lys Leu Ile Lys Ala Leu Lys
Lys Asn Ile Glu Ser Ile Gln1 5 10
15Asn Lys Met28610PRTPlasmodium falciparum 286His Lys Lys Leu
Ile Lys Ala Leu Lys Lys1 5
102879PRTPlasmodium falciparum 287His Lys Lys Leu Ile Lys Ala Leu Lys1
528823PRTPlasmodium falciparum 288Lys Ala Thr Tyr Ser Phe Val
Asn Thr Lys Lys Lys Ile Ile Ser Leu1 5 10
15Lys Ser Gln Gly His Lys Lys2028922PRTPlasmodium
falciparum 289Lys Ala Thr Tyr Ser Phe Val Asn Thr Lys Lys Lys Ile Ile Ser
Leu1 5 10 15Lys Ser Gln
Gly His Lys2029021PRTPlasmodium falciparum 290Lys Ala Thr Tyr Ser Phe Val
Asn Thr Lys Lys Lys Ile Ile Ser Leu1 5 10
15Lys Ser Gln Gly His2029129PRTPlasmodium falciparum
291His Thr Tyr Val Lys Gly Lys Lys Ala Pro Ser Asp Pro Gln Cys Ala1
5 10 15Asp Ile Lys Glu Glu Cys
Lys Glu Leu Leu Lys Glu Lys20 2529211PRTPlasmodium
falciparum 292Lys Ile Ile Ser Leu Lys Ser Gln Gly His Lys1
5 1029321PRTPlasmodium falciparum 293Lys Lys Lys Lys
Phe Glu Pro Leu Lys Asn Gly Asn Val Ser Glu Thr1 5
10 15Ile Lys Leu Ile His2029420PRTPlasmodium
falciparum 294Lys Lys Lys Phe Glu Pro Leu Lys Asn Gly Asn Val Ser Glu Thr
Ile1 5 10 15Lys Leu Ile
His2029519PRTPlasmodium falciparum 295Lys Lys Phe Glu Pro Leu Lys Asn Gly
Asn Val Ser Glu Thr Ile Lys1 5 10
15Leu Ile His29613PRTPlasmodium falciparum 296Lys Asn Gly Asn
Val Ser Glu Thr Ile Lys Leu Ile His1 5
1029711PRTPlasmodium falciparum 297Lys Leu Ile His Leu Gly Asn Lys Asp
Lys Lys1 5 1029836PRTPlasmodium
falciparum 298Lys Val Lys Lys Ile Gly Val Thr Leu Lys Lys Phe Glu Pro Leu
Lys1 5 10 15Asn Gly Asn
Val Ser Glu Thr Ile Lys Leu Ile His Leu Gly Asn Lys20 25
30Asp Lys Lys His3529959PRTPlasmodium falciparum 299His
Leu Ile Tyr Lys Asn Lys Ser Tyr Asn Pro Leu Leu Leu Ser Cys1
5 10 15Val Lys Lys Met Asn Met Leu
Lys Glu Asn Val Asp Tyr Ile Gln Asn20 25
30Gln Asn Leu Phe Lys Glu Leu Met Asn Gln Lys Ala Thr Tyr Ser Phe35
40 45Val Asn Thr Lys Lys Lys Ile Ile Ser Leu
Lys50 5530052PRTPlasmodium falciparum 300His Leu Ile Tyr
Lys Asn Lys Ser Tyr Asn Pro Leu Leu Leu Ser Cys1 5
10 15Val Lys Lys Met Asn Met Leu Lys Glu Asn
Val Asp Tyr Ile Gln Asn20 25 30Gln Asn
Leu Phe Lys Glu Leu Met Asn Gln Lys Ala Thr Tyr Ser Phe35
40 45Val Asn Thr Lys5030143PRTPlasmodium falciparum
301His Leu Ile Tyr Lys Asn Lys Ser Tyr Asn Pro Leu Leu Leu Ser Cys1
5 10 15Val Lys Lys Met Asn Met
Leu Lys Glu Asn Val Asp Tyr Ile Gln Asn20 25
30Gln Asn Leu Phe Lys Glu Leu Met Asn Gln Lys35
4030238PRTPlasmodium falciparum 302His Leu Ile Tyr Lys Asn Lys Ser Tyr
Asn Pro Leu Leu Leu Ser Cys1 5 10
15Val Lys Lys Met Asn Met Leu Lys Glu Asn Val Asp Tyr Ile Gln
Lys20 25 30Asn Gln Asn Leu Phe
Lys3530324PRTPlasmodium falciparum 303His Leu Ile Tyr Lys Asn Lys Ser Tyr
Asn Pro Leu Leu Leu Ser Cys1 5 10
15Val Lys Lys Met Asn Met Leu Lys2030447PRTPlasmodium falciparum
304Lys Ser Ala Asn Asn Ser Ala Asn Asn Gly Lys Lys Asn Asn Ala Glu1
5 10 15Glu Met Lys Asn Leu Val
Asn Phe Leu Gln Ser His Lys Lys Leu Ile20 25
30Lys Ala Leu Lys Lys Asn Ile Glu Ser Ile Gln Asn Lys Lys His35
40 4530537PRTPlasmodium falciparum 305Lys
Lys Asn Asn Ala Glu Glu Met Lys Asn Leu Val Asn Phe Leu Gln1
5 10 15Ser His Lys Lys Leu Ile Lys
Ala Leu Lys Lys Asn Ile Glu Ser Ile20 25
30Gln Asn Lys Lys His3530629PRTPlasmodium falciparum 306Lys Asn Leu Val
Asn Phe Leu Gln Ser His Lys Lys Leu Ile Lys Ala1 5
10 15Leu Lys Lys Asn Ile Glu Ser Ile Gln Asn
Lys Lys His20 2530719PRTPlasmodium falciparum 307Lys Lys
Leu Ile Lys Ala Leu Lys Lys Asn Ile Glu Ser Ile Gln Asn1 5
10 15Lys Lys His30818PRTPlasmodium
falciparum 308Lys Leu Ile Lys Ala Leu Lys Lys Asn Ile Glu Ser Ile Gln Asn
Lys1 5 10 15Lys
His30912PRTPlasmodium falciparum 309Lys Lys Asn Ile Glu Ser Ile Gln Asn
Lys Lys His1 5 1031011PRTPlasmodium
falciparum 310Lys Asn Ile Glu Ser Ile Gln Asn Lys Lys His1
5 1031117PRTPlasmodium falciparum 311Lys Asn Asn Ala
Glu Glu Met Lys Asn Leu Val Asn Phe Leu Gln Ser1 5
10 15His31223PRTPlasmodium falciparum 312Lys
Lys Leu Ile Lys Ala Leu Lys Lys Asn Ile Glu Ser Ile Gln Asn1
5 10 15Lys Lys Gln Gly His Lys
Lys2031319PRTPlasmodium falciparum 313Lys Lys Asn Asn Ala Glu Glu Met Lys
Asn Leu Val Asn Phe Leu Gln1 5 10
15Ser His Lys31417PRTPlasmodium falciparum 314Lys Asn Asn Ala
Glu Glu Met Lys Asn Leu Val Asn Phe Leu Gln Ser1 5
10 15His31522PRTPlasmodium falciparum 315Lys
Leu Ile Lys Ala Leu Lys Lys Asn Ile Glu Ser Ile Gln Asn Lys1
5 10 15Lys Gln Gly His Lys
Lys2031628PRTPlasmodium falciparum 316Lys Val Lys Lys Ile Gly Val Thr Leu
Lys Lys Phe Glu Pro Leu Lys1 5 10
15Asn Gly Asn Val Ser Glu Thr Ile Lys Leu Ile His20
2531713PRTPlasmodium falciparum 317Lys Asn Gly Asn Val Ser Glu Thr
Ile Lys Leu Ile His1 5
1031811PRTPlasmodium falciparum 318Lys Leu Ile His Leu Gly Asn Lys Asp
Lys Lys1 5 1031928PRTPlasmodium
falciparum 319Lys Ser Ala Asn Asn Ser Ala Asn Asn Gly Lys Lys Asn Asn Ala
Glu1 5 10 15Glu Met Lys
Asn Leu Val Asn Phe Leu Gln Ser His20
2532018PRTPlasmodium falciparum 320Lys Lys Asn Asn Ala Glu Glu Met Lys
Asn Leu Val Asn Phe Leu Gln1 5 10
15Ser His32119PRTPlasmodium falciparum 321Lys Lys Leu Ile Lys
Ala Leu Lys Lys Asn Ile Glu Ser Ile Gln Asn1 5
10 15Lys Lys His32215PRTPlasmodium falciparum
322Lys Ala Leu Lys Lys Asn Ile Glu Ser Ile Gln Asn Lys Lys His1
5 10 1532312PRTPlasmodium
falciparum 323Lys Lys Asn Ile Glu Ser Ile Gln Asn Lys Lys His1
5 1032427PRTPlasmodium falciparum 324Lys Glu Leu Met
Asn Gln Lys Ala Thr Tyr Ser Phe Val Asn Thr Lys1 5
10 15Lys Lys Ile Ile Ser Leu Lys Ser Gln Gly
His20 253257PRTPlasmodium falciparum 325Lys Ser Gln Gly
His Lys Lys1 532612PRTPlasmodium falciparum 326Lys Lys Lys
Ile Ile Ser Leu Lys Ser Gln Gly His1 5
1032711PRTPlasmodium falciparum 327Lys Lys Ile Ile Ser Leu Lys Ser Gln
Gly His1 5 1032812PRTPlasmodium
falciparum 328Lys Lys Asn Ile Glu Ser Ile Gln Asn Lys Lys His1
5 1032911PRTPlasmodium falciparum 329Lys Asn Ile Glu
Ser Ile Gln Asn Lys Lys His1 5
1033029PRTPlasmodium falciparum 330His Thr Tyr Val Lys Gly Lys Lys Ala
Pro Ser Asp Pro Gln Cys Ala1 5 10
15Asp Ile Lys Glu Glu Cys Lys Glu Leu Leu Lys Glu Lys20
2533127PRTPlasmodium falciparum 331His Thr Tyr Val Lys Gly Lys
Lys Ala Pro Ser Asp Pro Gln Cys Ala1 5 10
15Asp Ile Lys Glu Glu Cys Lys Glu Leu Leu Lys20
2533229PRTPlasmodium falciparum 332His Glu Asn Val Leu Ser Ala
Ala Leu Glu Asn Thr Gln Ser Glu Glu1 5 10
15Glu Lys Lys Glu Val Ile Asp Val Ile Glu Glu Val Lys20
2533348PRTPlasmodium falciparum 333Lys Glu Asn Val Val
Thr Thr Ile Leu Glu Lys Val Glu Glu Thr Thr1 5
10 15Ala Glu Ser Val Thr Thr Phe Ser Asn Ile Leu
Glu Glu Ile Gln Glu20 25 30Asn Thr Ile
Thr Asn Asp Thr Ile Glu Glu Lys Leu Glu Glu Leu His35 40
4533414PRTPlasmodium falciparum 334His Tyr Leu Gln Gln
Met Lys Glu Lys Phe Ser Lys Glu Lys1 5
1033542PRTPlasmodium falciparum 335His Tyr Leu Gln Gln Met Lys Glu Lys
Phe Ser Lys Glu Lys Asn Asn1 5 10
15Asn Val Ile Glu Val Thr Asn Lys Ala Glu Lys Lys Gly Asn Val
Gln20 25 30Val Thr Asn Lys Thr Glu Lys
Thr Thr Lys35 4033648PRTPlasmodium falciparum 336His Tyr
Leu Gln Gln Met Lys Glu Lys Phe Ser Lys Glu Lys Asn Asn1 5
10 15Asn Val Ile Glu Val Thr Asn Lys
Ala Glu Lys Lys Gly Asn Val Gln20 25
30Val Thr Asn Lys Thr Glu Lys Thr Thr Lys Val Asp Lys Asn Asn Lys35
40 4533757PRTPlasmodium falciparum 337His Tyr
Leu Gln Gln Met Lys Glu Lys Phe Ser Lys Glu Lys Asn Asn1 5
10 15Asn Val Ile Glu Val Thr Asn Lys
Ala Glu Lys Lys Gly Asn Val Gln20 25
30Val Thr Asn Lys Thr Glu Lys Thr Thr Lys Val Asp Lys Asn Asn Lys35
40 45Val Pro Lys Lys Arg Arg Thr Gln Lys50
5533859PRTPlasmodium falciparum 338His Tyr Leu Gln Gln Met
Lys Glu Lys Phe Ser Lys Glu Lys Asn Asn1 5
10 15Asn Val Ile Glu Val Thr Asn Lys Ala Glu Lys Lys
Gly Asn Val Gln20 25 30Val Thr Asn Lys
Thr Glu Lys Thr Thr Lys Val Asp Lys Asn Asn Lys35 40
45Val Pro Lys Lys Arg Arg Thr Gln Lys Ser Lys50
5533952PRTPlasmodium falciparum 339His Val Asp Glu Val Met Lys Tyr
Val Gln Lys Ile Asp Lys Glu Val1 5 10
15Asp Lys Glu Val Ser Lys Ala Leu Glu Ser Lys Asn Asp Val
Thr Asn20 25 30Val Leu Lys Gln Asn Gln
Asp Phe Phe Ser Lys Val Lys Asn Phe Val35 40
45Lys Lys Tyr Lys5034050PRTPlasmodium falciparum 340His Val Asp Glu
Val Met Lys Tyr Val Gln Lys Ile Asp Lys Glu Val1 5
10 15Asp Lys Glu Val Ser Lys Ala Leu Glu Ser
Lys Asn Asp Val Thr Asn20 25 30Val Leu
Lys Gln Asn Gln Asp Phe Phe Ser Lys Val Lys Asn Phe Val35
40 45Lys Lys5034143PRTPlasmodium falciparum 341His Val
Asp Glu Val Met Lys Tyr Val Gln Lys Ile Asp Lys Glu Val1 5
10 15Asp Lys Glu Val Ser Lys Ala Leu
Glu Ser Lys Asn Asp Val Thr Asn20 25
30Val Leu Lys Gln Asn Gln Asp Phe Phe Ser Lys35
4034235PRTPlasmodium falciparum 342His Val Asp Glu Val Met Lys Tyr Val
Gln Lys Ile Asp Lys Glu Val1 5 10
15Asp Lys Glu Val Ser Lys Ala Leu Glu Ser Lys Asn Asp Val Thr
Asn20 25 30Val Leu
Lys3534327PRTPlasmodium falciparum 343His Val Asp Glu Val Met Lys Tyr Val
Gln Lys Ile Asp Lys Glu Val1 5 10
15Asp Lys Glu Val Ser Lys Ala Leu Glu Ser Lys20
2534422PRTPlasmodium falciparum 344His Val Asp Glu Val Met Lys Tyr Val
Gln Lys Ile Asp Lys Glu Val1 5 10
15Asp Lys Glu Val Ser Lys2034518PRTPlasmodium falciparum 345His
Val Asp Glu Val Met Lys Tyr Val Gln Lys Ile Asp Lys Glu Val1
5 10 15Asp Lys34614PRTPlasmodium
falciparum 346His Val Asp Glu Val Met Lys Tyr Val Gln Lys Ile Asp Lys1
5 1034739PRTPlasmodium falciparum 347Lys Asp
Glu Val Ile Asp Leu Ile Val Gln Lys Glu Lys Arg Ile Glu1 5
10 15Lys Val Lys Ala Lys Lys Lys Lys
Leu Glu Lys Lys Val Glu Glu Gly20 25
30Val Ser Gly Leu Lys Lys His3534823PRTPlasmodium falciparum 348Lys Val
Lys Ala Lys Lys Lys Lys Leu Glu Lys Lys Val Glu Glu Gly1 5
10 15Val Ser Gly Leu Lys Lys
His2034921PRTPlasmodium falciparum 349Lys Ala Lys Lys Lys Lys Leu Glu Lys
Lys Val Glu Glu Gly Val Ser1 5 10
15Gly Leu Lys Lys His2035019PRTPlasmodium falciparum 350Lys Lys
Lys Lys Leu Glu Lys Lys Val Glu Glu Gly Val Ser Gly Leu1 5
10 15Lys Lys His35118PRTPlasmodium
falciparum 351Lys Lys Lys Leu Glu Lys Lys Val Glu Glu Gly Val Ser Gly Leu
Lys1 5 10 15Lys
His35217PRTPlasmodium falciparum 352Lys Lys Leu Glu Lys Lys Val Glu Glu
Gly Val Ser Gly Leu Lys Lys1 5 10
15His35316PRTPlasmodium falciparum 353Lys Leu Glu Lys Lys Val
Glu Glu Gly Val Ser Gly Leu Lys Lys His1 5
10 1535413PRTPlasmodium falciparum 354Lys Lys Val Glu
Glu Gly Val Ser Gly Leu Lys Lys His1 5
1035512PRTPlasmodium falciparum 355Lys Val Glu Glu Gly Val Ser Gly Leu
Lys Lys His1 5 1035659PRTPlasmodium
falciparum 356His Val Glu Gln Asn Val Tyr Val Asp Val Asp Val Pro Ala Met
Lys1 5 10 15Asp Gln Phe
Leu Gly Ile Leu Asn Glu Ala Gly Gly Leu Lys Glu Met20 25
30Phe Phe Asn Leu Glu Asp Val Phe Lys Ser Glu Ser Asp
Val Ile Thr35 40 45Val Glu Glu Ile Lys
Asp Glu Pro Val Gln Lys50 5535753PRTPlasmodium
falciparum 357His Ile Lys Gly Leu Glu Glu Asp Asp Leu Glu Glu Val Asp Asp
Leu1 5 10 15Lys Gly Ser
Ile Leu Asp Met Leu Lys Gly Asp Met Glu Leu Gly Asp20 25
30Met Asp Lys Glu Ser Leu Glu Asp Val Thr Thr Lys Leu
Gly Glu Arg35 40 45Val Glu Ser Leu
Lys5035844PRTPlasmodium falciparum 358His Ile Lys Gly Leu Glu Glu Asp Asp
Leu Glu Glu Val Asp Asp Leu1 5 10
15Lys Gly Ser Ile Leu Asp Met Leu Lys Gly Asp Met Glu Leu Gly
Asp20 25 30Met Asp Lys Glu Ser Leu Glu
Asp Val Thr Thr Lys35 4035935PRTPlasmodium falciparum
359His Ile Lys Gly Leu Glu Glu Asp Asp Leu Glu Glu Val Asp Asp Leu1
5 10 15Lys Gly Ser Ile Leu Asp
Met Leu Lys Gly Asp Met Glu Leu Gly Asp20 25
30Met Asp Lys3536025PRTPlasmodium falciparum 360His Ile Lys Gly Leu
Glu Glu Asp Asp Leu Glu Glu Val Asp Asp Leu1 5
10 15Lys Gly Ser Ile Leu Asp Met Leu Lys20
2536131PRTPlasmodium falciparum 361His Ile Ile Ser Gly Asp Ala
Asp Val Leu Ser Ser Ala Leu Gly Met1 5 10
15Asp Glu Glu Gln Met Lys Thr Arg Lys Lys Ala Gln Arg
Pro Lys20 25 3036223PRTPlasmodium
falciparum 362His Asp Ile Thr Thr Thr Leu Asp Glu Val Val Glu Leu Lys Asp
Val1 5 10 15Glu Glu Asp
Lys Ile Glu Lys2036310PRTPlasmodium falciparum 363Lys Lys Leu Glu Glu Val
His Glu Leu Lys1 5 103649PRTPlasmodium
falciparum 364Lys Leu Glu Glu Val His Glu Leu Lys1
536519PRTPlasmodium falciparum 365Lys Thr Ile Glu Thr Asp Ile Leu Glu Glu
Lys Lys Lys Glu Ile Glu1 5 10
15Lys Asp His36611PRTPlasmodium falciparum 366Lys Lys Glu Ile Glu
Lys Asp His Phe Glu Lys1 5
103676PRTPlasmodium falciparum 367Lys Asp His Phe Glu Lys1
536811PRTPlasmodium falciparum 368Lys Phe Glu Glu Glu Ala Glu Glu Ile Lys
His1 5 1036947PRTPlasmodium falciparum
369Lys Asp Gly Asp Thr Lys Cys Thr Leu Glu Cys Ala Gln Gly Lys Lys1
5 10 15Cys Ile Lys His Lys Ser
Asp His Asn His Lys Ser Asp His Asn His20 25
30Lys Ser Asp Pro Asn His Lys Lys Lys Asn Asn Asn Asn Asn Lys35
40 4537040PRTPlasmodium falciparum 370Lys
Asp Gly Asp Thr Lys Cys Thr Leu Glu Cys Ala Gln Gly Lys Lys1
5 10 15Cys Ile Lys His Lys Ser Asp
His Asn His Lys Ser Asp His Asn His20 25
30Lys Ser Asp Pro Asn His Lys Lys35
4037139PRTPlasmodium falciparum 371Lys Asp Gly Asp Thr Lys Cys Thr Leu
Glu Cys Ala Gln Gly Lys Lys1 5 10
15Cys Ile Lys His Lys Ser Asp His Asn His Lys Ser Asp His Asn
His20 25 30Lys Ser Asp Pro Asn His
Lys3537233PRTPlasmodium falciparum 372Lys Asp Gly Asp Thr Lys Cys Thr Leu
Glu Cys Ala Gln Gly Lys Lys1 5 10
15Cys Ile Lys His Lys Ser Asp His Asn His Lys Ser Asp His Asn
His20 25 30Lys37327PRTPlasmodium
falciparum 373Lys Asp Gly Asp Thr Lys Cys Thr Leu Glu Cys Ala Gln Gly Lys
Lys1 5 10 15Cys Ile Lys
His Lys Ser Asp His Asn His Lys20 2537421PRTPlasmodium
falciparum 374Lys Asp Gly Asp Thr Lys Cys Thr Leu Glu Cys Ala Gln Gly Lys
Lys1 5 10 15Cys Ile Lys
His Lys2037516PRTPlasmodium falciparum 375Lys Asp Gly Asp Thr Lys Cys Thr
Leu Glu Cys Ala Gln Gly Lys Lys1 5 10
1537615PRTPlasmodium falciparum 376Lys Asp Gly Asp Thr Lys
Cys Thr Leu Glu Cys Ala Gln Gly Lys1 5 10
1537723PRTPlasmodium falciparum 377Lys Cys Ile Gln Ala
Glu Cys Asn Tyr Lys Glu Cys Gly Glu Gln Lys1 5
10 15Cys Val Trp Asp Gly Ile
His2037814PRTPlasmodium falciparum 378Lys Glu Cys Gly Glu Gln Lys Cys Val
Trp Asp Gly Ile His1 5
1037932PRTPlasmodium falciparum 379His Ile Glu Cys Lys Cys Asn Asn Asp
Tyr Val Leu Thr Asn Arg Tyr1 5 10
15Glu Cys Glu Pro Lys Asn Lys Cys Thr Ser Leu Glu Asp Thr Asn
Lys20 25 3038039PRTPlasmodium
falciparum 380Lys Ser Asp His Asn His Lys Ser Asp His Asn His Lys Ser Asp
His1 5 10 15Asn His Lys
Ser Asp His Asn His Lys Ser Asp Pro Asn His Lys Lys20 25
30Lys Asn Asn Asn Asn Asn Lys3538133PRTPlasmodium
falciparum 381Lys Ser Asp His Asn His Lys Ser Asp His Asn His Lys Ser Asp
His1 5 10 15Asn His Lys
Ser Asp Pro Asn His Lys Lys Lys Asn Asn Asn Asn Asn20 25
30Lys38227PRTPlasmodium falciparum 382Lys Ser Asp His
Asn His Lys Ser Asp His Asn His Lys Ser Asp Pro1 5
10 15Asn His Lys Lys Lys Asn Asn Asn Asn Asn
Lys20 2538321PRTPlasmodium falciparum 383Lys Ser Asp His
Asn His Lys Ser Asp Pro Asn His Lys Lys Lys Asn1 5
10 15Asn Asn Asn Asn Lys2038418PRTPlasmodium
falciparum 384Lys Lys Lys Asn Asn Asn Asn Asn Lys Asp Asn Lys Ser Asp Pro
Asn1 5 10 15His
Lys38517PRTPlasmodium falciparum 385Lys Lys Asn Asn Asn Asn Asn Lys Asp
Asn Lys Ser Asp Pro Asn His1 5 10
15Lys38616PRTPlasmodium falciparum 386Lys Asn Asn Asn Asn Asn
Lys Asp Asn Lys Ser Asp Pro Asn His Lys1 5
10 1538710PRTPlasmodium falciparum 387Lys Asp Asn Lys
Ser Asp Pro Asn His Lys1 5
103887PRTPlasmodium falciparum 388Lys Ser Asp Pro Asn His Lys1
538935PRTPlasmodium falciparum 389His Ser Leu Tyr Ala Leu Gln Gln Asn
Glu Glu Tyr Gln Lys Val Lys1 5 10
15Asn Glu Lys Asp Gln Asn Glu Ile Lys Lys Ile Lys Gln Leu Ile
Glu20 25 30Lys Asn
Lys3539028PRTPlasmodium falciparum 390His Ser Leu Tyr Ala Leu Gln Gln Asn
Glu Glu Tyr Gln Lys Val Lys1 5 10
15Asn Glu Lys Asp Gln Asn Glu Ile Lys Lys Ile Lys20
2539126PRTPlasmodium falciparum 391His Ser Leu Tyr Ala Leu Gln Gln
Asn Glu Glu Tyr Gln Lys Val Lys1 5 10
15Asn Glu Lys Asp Gln Asn Glu Ile Lys Lys20
2539225PRTPlasmodium falciparum 392His Ser Leu Tyr Ala Leu Gln Gln Asn
Glu Glu Tyr Gln Lys Val Lys1 5 10
15Asn Glu Lys Asp Gln Asn Glu Ile Lys20
2539311PRTPlasmodium falciparum 393His Lys Leu Glu Asn Leu Glu Glu Met
Asp Lys1 5 1039411PRTPlasmodium
falciparum 394Lys His Phe Asp Asp Asn Thr Asn Glu Gln Lys1
5 103958PRTPlasmodium falciparum 395Lys Lys Glu Asp Asp
Glu Lys His1 539613PRTPlasmodium falciparum 396Lys Glu Glu
Asn Asn Lys Lys Glu Asp Asp Glu Lys His1 5
1039721PRTPlasmodium falciparum 397Lys Thr Ser Ser Gly Ile Leu Asn Lys
Glu Glu Asn Asn Lys Lys Glu1 5 10
15Asp Asp Glu Lys His203987PRTPlasmodium falciparum 398Lys Asn
Ile His Ile Lys Lys1 539913PRTPlasmodium falciparum 399His
Ile Lys Lys Lys Glu Gly Ile Asp Ile Gly Tyr Lys1 5
1040021PRTPlasmodium falciparum 400Lys Lys Met Trp Thr Cys Lys
Leu Trp Asp Asn Lys Gly Asn Glu Ile1 5 10
15Thr Lys Asn Ile His2040130PRTPlasmodium falciparum
401Lys Lys Gly Ile Gln Trp Asn Leu Leu Lys Lys Met Trp Thr Cys Lys1
5 10 15Leu Trp Asp Asn Lys Gly
Asn Glu Ile Thr Lys Asn Ile His20 25
3040250PRTPlasmodium falciparum 402Lys Glu Lys Lys Asp Ser Asn Glu Asn
Arg Lys Lys Lys Gln Lys Glu1 5 10
15Asp Lys Lys Asn Pro Asn Lys Leu Lys Lys Ile Glu Tyr Thr Asn
Lys20 25 30Ile Thr His Phe Phe Lys Ala
Lys Asn Asn Lys Gln Gln Asn Asn Val35 40
45Thr His5040348PRTPlasmodium falciparum 403Lys Lys Asp Ser Asn Glu Asn
Arg Lys Lys Lys Gln Lys Glu Asp Lys1 5 10
15Lys Asn Pro Asn Lys Leu Lys Lys Ile Glu Tyr Thr Asn
Lys Ile Thr20 25 30His Phe Phe Lys Ala
Lys Asn Asn Lys Gln Gln Asn Asn Val Thr His35 40
4540447PRTPlasmodium falciparum 404Lys Asp Ser Asn Glu Asn Arg
Lys Lys Lys Gln Lys Glu Asp Lys Lys1 5 10
15Asn Pro Asn Lys Leu Lys Lys Ile Glu Tyr Thr Asn Lys
Ile Thr His20 25 30Phe Phe Lys Ala Lys
Asn Asn Lys Gln Gln Asn Asn Val Thr His35 40
4540539PRTPlasmodium falciparum 405Lys Lys Gln Lys Glu Asp Lys Lys
Asn Pro Asn Lys Leu Lys Lys Ile1 5 10
15Glu Tyr Thr Asn Lys Ile Thr His Phe Phe Lys Ala Lys Asn
Asn Lys20 25 30Gln Gln Asn Asn Val Thr
His3540638PRTPlasmodium falciparum 406Lys Gln Lys Glu Asp Lys Lys Asn Pro
Asn Lys Leu Lys Lys Ile Glu1 5 10
15Tyr Thr Asn Lys Ile Thr His Phe Phe Lys Ala Lys Asn Asn Lys
Gln20 25 30Gln Asn Asn Val Thr
His3540736PRTPlasmodium falciparum 407Lys Glu Asp Lys Lys Asn Pro Asn Lys
Leu Lys Lys Ile Glu Tyr Thr1 5 10
15Asn Lys Ile Thr His Phe Phe Lys Ala Lys Asn Asn Lys Gln Gln
Asn20 25 30Asn Val Thr
His3540832PRTPlasmodium falciparum 408Lys Asn Pro Asn Lys Leu Lys Lys Ile
Glu Tyr Thr Asn Lys Ile Thr1 5 10
15His Phe Phe Lys Ala Lys Asn Asn Lys Gln Gln Asn Asn Val Thr
His20 25 3040926PRTPlasmodium
falciparum 409Lys Lys Ile Glu Tyr Thr Asn Lys Ile Thr His Phe Phe Lys Ala
Lys1 5 10 15Asn Asn Lys
Gln Gln Asn Asn Val Thr His20 2541025PRTPlasmodium
falciparum 410Lys Ile Glu Tyr Thr Asn Lys Ile Thr His Phe Phe Lys Ala Lys
Asn1 5 10 15Asn Lys Gln
Gln Asn Asn Val Thr His20 2541119PRTPlasmodium
falciparum 411Lys Ile Thr His Phe Phe Lys Ala Lys Asn Asn Lys Gln Gln Asn
Asn1 5 10 15Val Thr
His41248PRTPlasmodium falciparum 412His Lys Asn Asn Glu Asp Ile Lys Asn
Asp Asn Ser Lys Asp Ile Lys1 5 10
15Asn Asp Asn Ser Lys Asp Ile Lys Asn Asp Asn Ser Lys Asp Ile
Lys20 25 30Asn Asp Asn Asn Glu Asp Ile
Lys Asn Asp Asn Ser Lys Asp Ile Lys35 40
4541345PRTPlasmodium falciparum 413His Lys Asn Asn Glu Asp Ile Lys Asn
Asp Asn Ser Lys Asp Ile Lys1 5 10
15Asn Asp Asn Ser Lys Asp Ile Lys Asn Asp Asn Ser Lys Asp Ile
Lys20 25 30Asn Asp Asn Asn Glu Asp Ile
Lys Asn Asp Asn Ser Lys35 40
4541440PRTPlasmodium falciparum 414His Lys Asn Asn Glu Asp Ile Lys Asn
Asp Asn Ser Lys Asp Ile Lys1 5 10
15Asn Asp Asn Ser Lys Asp Ile Lys Asn Asp Asn Ser Lys Asp Ile
Lys20 25 30Asn Asp Asn Asn Glu Asp Ile
Lys35 4041532PRTPlasmodium falciparum 415His Lys Asn Asn
Glu Asp Ile Lys Asn Asp Asn Ser Lys Asp Ile Lys1 5
10 15Asn Asp Asn Ser Lys Asp Ile Lys Asn Asp
Asn Ser Lys Asp Ile Lys20 25
3041629PRTPlasmodium falciparum 416His Lys Asn Asn Glu Asp Ile Lys Asn
Asp Asn Ser Lys Asp Ile Lys1 5 10
15Asn Asp Asn Ser Lys Asp Ile Lys Asn Asp Asn Ser Lys20
2541724PRTPlasmodium falciparum 417His Lys Asn Asn Glu Asp Ile
Lys Asn Asp Asn Ser Lys Asp Ile Lys1 5 10
15Asn Asp Asn Ser Lys Asp Ile Lys2041821PRTPlasmodium
falciparum 418His Lys Asn Asn Glu Asp Ile Lys Asn Asp Asn Ser Lys Asp Ile
Lys1 5 10 15Asn Asp Asn
Ser Lys2041916PRTPlasmodium falciparum 419His Lys Asn Asn Glu Asp Ile Lys
Asn Asp Asn Ser Lys Asp Ile Lys1 5 10
154208PRTPlasmodium falciparum 420His Lys Asn Asn Glu Asp
Ile Lys1 542131PRTPlasmodium falciparum 421Lys Lys Tyr Asp
Asp Leu Gln Asn Lys Tyr Asn Ile Leu Asn Lys Leu1 5
10 15Lys Asn Ser Leu Glu Glu Lys Asn Glu Glu
Leu Lys Lys Tyr His20 25
3042230PRTPlasmodium falciparum 422Lys Tyr Asp Asp Leu Gln Asn Lys Tyr
Asn Ile Leu Asn Lys Leu Lys1 5 10
15Asn Ser Leu Glu Glu Lys Asn Glu Glu Leu Lys Lys Tyr His20
25 3042323PRTPlasmodium falciparum 423Lys
Tyr Asn Ile Leu Asn Lys Leu Lys Asn Ser Leu Glu Glu Lys Asn1
5 10 15Glu Glu Leu Lys Lys Tyr
His2042417PRTPlasmodium falciparum 424Lys Leu Lys Asn Ser Leu Glu Glu Lys
Asn Glu Glu Leu Lys Lys Tyr1 5 10
15His42515PRTPlasmodium falciparum 425Lys Asn Ser Leu Glu Glu
Lys Asn Glu Glu Leu Lys Lys Tyr His1 5 10
154269PRTPlasmodium falciparum 426Lys Asn Glu Glu Leu
Lys Lys Tyr His1 542735PRTPlasmodium falciparum 427His Met
Gly Asn Asn Gln Asp Ile Asn Glu Asn Val Tyr Asn Ile Lys1 5
10 15Pro Gln Glu Phe Lys Glu Glu Glu
Glu Glu Asp Ile Ser Met Val Asn20 25
30Thr Lys Lys3542817PRTPlasmodium falciparum 428Lys Asn Ser Asn Glu Leu
Lys Arg Ile Asn Asp Asn Phe Phe Lys Leu1 5
10 15His42955PRTPlasmodium falciparum 429Lys Pro Cys
Leu Tyr Lys Lys Cys Lys Ile Ser Gln Cys Leu Tyr Lys1 5
10 15Lys Cys Lys Ile Ser Gln Val Trp Trp
Cys Met Pro Val Lys Asp Thr20 25 30Phe
Asn Thr Tyr Glu Arg Asn Asn Val Leu Asn Ser Lys Ile Glu Asn35
40 45Asn Ile Glu Lys Ile Pro His50
5543040PRTPlasmodium falciparum 430His Ile Asn Asn Glu Tyr Thr Asn Lys
Asn Pro Lys Asn Cys Leu Leu1 5 10
15Tyr Lys Asn Glu Glu Arg Asn Tyr Asn Asp Asn Asn Ile Lys Asp
Tyr20 25 30Ile Asn Ser Met Asn Phe Lys
Lys35 4043139PRTPlasmodium falciparum 431His Ile Asn Asn
Glu Tyr Thr Asn Lys Asn Pro Lys Asn Cys Leu Leu1 5
10 15Tyr Lys Asn Glu Glu Arg Asn Tyr Asn Asp
Asn Asn Ile Lys Asp Tyr20 25 30Ile Asn
Ser Met Asn Phe Lys3543218PRTPlasmodium falciparum 432His Ile Asn Asn Glu
Tyr Thr Asn Lys Asn Pro Lys Asn Cys Leu Leu1 5
10 15Tyr Lys43323PRTPlasmodium falciparum 433Lys
Asn Lys Thr Asn Gln Ser Lys Gly Val Lys Gly Glu Tyr Glu Lys1
5 10 15Lys Lys Glu Thr Asn Gly
His2043421PRTPlasmodium falciparum 434Lys Thr Asn Gln Ser Lys Gly Val Lys
Gly Glu Tyr Glu Lys Lys Lys1 5 10
15Glu Thr Asn Gly His2043516PRTPlasmodium falciparum 435Lys Gly
Val Lys Gly Glu Tyr Glu Lys Lys Lys Glu Thr Asn Gly His1 5
10 1543613PRTPlasmodium falciparum
436Lys Gly Glu Tyr Glu Lys Lys Lys Glu Thr Asn Gly His1 5
1043728PRTPlasmodium falciparum 437Lys Ser Gly Met Tyr
Thr Asn Glu Gly Asn Lys Ser Cys Glu Cys Ser1 5
10 15Tyr Lys Lys Lys Ser Ser Ser Ser Asn Lys Val
His20 2543818PRTPlasmodium falciparum 438Lys Ser Cys Glu
Cys Ser Tyr Lys Lys Lys Ser Ser Ser Ser Asn Lys1 5
10 15Val His43911PRTPlasmodium falciparum
439Lys Lys Lys Ser Ser Ser Ser Asn Lys Val His1 5
1044010PRTPlasmodium falciparum 440Lys Lys Ser Ser Ser Ser Asn
Lys Val His1 5 104419PRTPlasmodium
falciparum 441Lys Ser Ser Ser Ser Asn Lys Val His1
544230PRTPlasmodium falciparum 442His Ile Met Leu Lys Ser Gly Met Tyr Thr
Asn Glu Gly Asn Lys Ser1 5 10
15Cys Glu Cys Ser Tyr Lys Lys Lys Ser Ser Ser Ser Asn Lys20
25 3044324PRTPlasmodium falciparum 443His Ile
Met Leu Lys Ser Gly Met Tyr Thr Asn Glu Gly Asn Lys Ser1 5
10 15Cys Glu Cys Ser Tyr Lys Lys
Lys2044423PRTPlasmodium falciparum 444His Ile Met Leu Lys Ser Gly Met Tyr
Thr Asn Glu Gly Asn Lys Ser1 5 10
15Cys Glu Cys Ser Tyr Lys Lys2044522PRTPlasmodium falciparum
445His Ile Met Leu Lys Ser Gly Met Tyr Thr Asn Glu Gly Asn Lys Ser1
5 10 15Cys Glu Cys Ser Tyr
Lys2044650PRTPlasmodium falciparum 446Lys Pro Leu Ala Lys Leu Arg Lys Arg
Glu Lys Thr Gln Ile Asn Lys1 5 10
15Thr Lys Tyr Glu Arg Gly Asp Val Ile Ile Asp Asn Thr Glu Ile
Gln20 25 30Lys Ile Ile Ile Arg Asp Tyr
His Glu Thr Leu Asn Val His Lys Leu35 40
45Asp His5044743PRTPlasmodium falciparum 447Lys Arg Glu Lys Thr Gln Ile
Asn Lys Thr Lys Tyr Glu Arg Gly Asp1 5 10
15Val Ile Ile Asp Asn Thr Glu Ile Gln Lys Ile Ile Ile
Arg Asp Tyr20 25 30His Glu Thr Leu Asn
Val His Lys Leu Asp His35 4044840PRTPlasmodium
falciparum 448Lys Thr Gln Ile Asn Lys Thr Lys Tyr Glu Arg Gly Asp Val Ile
Ile1 5 10 15Asp Asn Thr
Glu Ile Gln Lys Ile Ile Ile Arg Asp Tyr His Glu Thr20 25
30Leu Asn Val His Lys Leu Asp His35
4044946PRTPlasmodium falciparum 449Lys Pro Leu Ala Lys Leu Arg Lys Arg
Glu Lys Thr Gln Ile Asn Lys1 5 10
15Thr Lys Tyr Glu Arg Gly Asp Val Ile Ile Asp Asn Thr Glu Ile
Gln20 25 30Lys Ile Ile Ile Arg Asp Tyr
His Glu Thr Leu Asn Val His35 40
4545040PRTPlasmodium falciparum 450Lys Pro Leu Ala Lys Leu Arg Lys Arg
Glu Lys Thr Gln Ile Asn Lys1 5 10
15Thr Lys Tyr Glu Arg Gly Asp Val Ile Ile Asp Asn Thr Glu Ile
Gln20 25 30Lys Ile Ile Ile Arg Asp Tyr
His35 4045136PRTPlasmodium falciparum 451Lys Leu Arg Lys
Arg Glu Lys Thr Gln Ile Asn Lys Thr Lys Tyr Glu1 5
10 15Arg Gly Asp Val Ile Ile Asp Asn Thr Glu
Ile Gln Lys Ile Ile Ile20 25 30Arg Asp
Tyr His3545233PRTPlasmodium falciparum 452Lys Arg Glu Lys Thr Gln Ile Asn
Lys Thr Lys Tyr Glu Arg Gly Asp1 5 10
15Val Ile Ile Asp Asn Thr Glu Ile Gln Lys Ile Ile Ile Arg
Asp Tyr20 25 30His45330PRTPlasmodium
falciparum 453Lys Thr Gln Ile Asn Lys Thr Lys Tyr Glu Arg Gly Asp Val Ile
Ile1 5 10 15Asp Asn Thr
Glu Ile Gln Lys Ile Ile Ile Arg Asp Tyr His20 25
3045441PRTPlasmodium falciparum 454Lys Lys Asp Lys Glu Lys Lys
Lys Asp Ser Asn Glu Asn Arg Lys Lys1 5 10
15Lys Gln Lys Glu Asp Lys Lys Asn Pro Asn Asp Asn Lys
Leu Lys Lys20 25 30Ile Glu Tyr Thr Asn
Lys Ile Thr His35 4045540PRTPlasmodium falciparum 455Lys
Asp Lys Glu Lys Lys Lys Asp Ser Asn Glu Asn Arg Lys Lys Lys1
5 10 15Gln Lys Glu Asp Lys Lys Asn
Pro Asn Asp Asn Lys Leu Lys Lys Ile20 25
30Glu Tyr Thr Asn Lys Ile Thr His35
4045638PRTPlasmodium falciparum 456Lys Glu Lys Lys Lys Asp Ser Asn Glu
Asn Arg Lys Lys Lys Gln Lys1 5 10
15Glu Asp Lys Lys Asn Pro Asn Asp Asn Lys Leu Lys Lys Ile Glu
Tyr20 25 30Thr Asn Lys Ile Thr
His3545736PRTPlasmodium falciparum 457Lys Lys Lys Asp Ser Asn Glu Asn Arg
Lys Lys Lys Gln Lys Glu Asp1 5 10
15Lys Lys Asn Pro Asn Asp Asn Lys Leu Lys Lys Ile Glu Tyr Thr
Asn20 25 30Lys Ile Thr
His3545835PRTPlasmodium falciparum 458Lys Lys Asp Ser Asn Glu Asn Arg Lys
Lys Lys Gln Lys Glu Asp Lys1 5 10
15Lys Asn Pro Asn Asp Asn Lys Leu Lys Lys Ile Glu Tyr Thr Asn
Lys20 25 30Ile Thr
His3545934PRTPlasmodium falciparum 459Lys Asp Ser Asn Glu Asn Arg Lys Lys
Lys Gln Lys Glu Asp Lys Lys1 5 10
15Asn Pro Asn Asp Asn Lys Leu Lys Lys Ile Glu Tyr Thr Asn Lys
Ile20 25 30Thr His46027PRTPlasmodium
falciparum 460Lys Lys Lys Gln Lys Glu Asp Lys Lys Asn Pro Asn Asp Asn Lys
Leu1 5 10 15Lys Lys Ile
Glu Tyr Thr Asn Lys Ile Thr His20 2546126PRTPlasmodium
falciparum 461Lys Lys Gln Lys Glu Asp Lys Lys Asn Pro Asn Asp Asn Lys Leu
Lys1 5 10 15Lys Ile Glu
Tyr Thr Asn Lys Ile Thr His20 2546225PRTPlasmodium
falciparum 462Lys Gln Lys Glu Asp Lys Lys Asn Pro Asn Asp Asn Lys Leu Lys
Lys1 5 10 15Ile Glu Tyr
Thr Asn Lys Ile Thr His20 2546323PRTPlasmodium
falciparum 463Lys Glu Asp Lys Lys Asn Pro Asn Asp Asn Lys Leu Lys Lys Ile
Glu1 5 10 15Tyr Thr Asn
Lys Ile Thr His2046420PRTPlasmodium falciparum 464Lys Lys Asn Pro Asn Asp
Asn Lys Leu Lys Lys Ile Glu Tyr Thr Asn1 5
10 15Lys Ile Thr His2046519PRTPlasmodium falciparum
465Lys Asn Pro Asn Asp Asn Lys Leu Lys Lys Ile Glu Tyr Thr Asn Lys1
5 10 15Ile Thr
His46613PRTPlasmodium falciparum 466Lys Leu Lys Lys Ile Glu Tyr Thr Asn
Lys Ile Thr His1 5 1046711PRTPlasmodium
falciparum 467Lys Lys Ile Glu Tyr Thr Asn Lys Ile Thr His1
5 1046810PRTPlasmodium falciparum 468Lys Ile Glu Tyr
Thr Asn Lys Ile Thr His1 5
1046944PRTPlasmodium falciparum 469His Gly Gln Ile Lys Ile Glu Asp Val
Asn Asn Glu Asn Phe Asn Asn1 5 10
15Glu Gln Met Lys Asn Lys Tyr Asn Asp Glu Glu Lys Met Asp Ile
Ser20 25 30Lys Ser Lys Ser Leu Lys Ser
Asp Phe Leu Glu Lys35 4047038PRTPlasmodium falciparum
470His Gly Gln Ile Lys Ile Glu Asp Val Asn Asn Glu Asn Phe Asn Asn1
5 10 15Glu Gln Met Lys Asn Lys
Tyr Asn Asp Glu Glu Lys Met Asp Ile Ser20 25
30Lys Ser Lys Ser Leu Lys3547135PRTPlasmodium falciparum 471His Gly
Gln Ile Lys Ile Glu Asp Val Asn Asn Glu Asn Phe Asn Asn1 5
10 15Glu Gln Met Lys Asn Lys Tyr Asn
Asp Glu Glu Lys Met Asp Ile Ser20 25
30Lys Ser Lys3547233PRTPlasmodium falciparum 472His Gly Gln Ile Lys Ile
Glu Asp Val Asn Asn Glu Asn Phe Asn Asn1 5
10 15Glu Gln Met Lys Asn Lys Tyr Asn Asp Glu Glu Lys
Met Asp Ile Ser20 25
30Lys47331PRTPlasmodium falciparum 473Lys Lys Tyr Asp Asp Leu Gln Asn Lys
Tyr Asn Ile Leu Asn Lys Leu1 5 10
15Lys Asn Ser Leu Glu Glu Lys Asn Glu Glu Leu Lys Lys Tyr His20
25 3047430PRTPlasmodium falciparum
474Lys Tyr Asp Asp Leu Gln Asn Lys Tyr Asn Ile Leu Asn Lys Leu Lys1
5 10 15Asn Ser Leu Glu Glu Lys
Asn Glu Glu Leu Lys Lys Tyr His20 25
3047523PRTPlasmodium falciparum 475Lys Tyr Asn Ile Leu Asn Lys Leu Lys
Asn Ser Leu Glu Glu Lys Asn1 5 10
15Glu Glu Leu Lys Lys Tyr His2047617PRTPlasmodium falciparum
476Lys Leu Lys Asn Ser Leu Glu Glu Lys Asn Glu Glu Leu Lys Lys Tyr1
5 10 15His47715PRTPlasmodium
falciparum 477Lys Asn Ser Leu Glu Glu Lys Asn Glu Glu Leu Lys Lys Tyr
His1 5 10
154789PRTPlasmodium falciparum 478Lys Asn Glu Glu Leu Lys Lys Tyr His1
547944PRTPlasmodium falciparum 479His Met Gly Asn Asn Gln Asp
Ile Asn Glu Asn Val Tyr Asn Ile Lys1 5 10
15Pro Gln Glu Phe Lys Glu Glu Glu Glu Glu Asp Ile Ser
Met Val Asn20 25 30Thr Lys Lys Cys Asp
Asp Ile Gln Glu Asn Ile Lys35 4048050PRTPlasmodium
falciparum 480Lys Thr Asn Leu Tyr Asn Ile Tyr Asn Asn Lys Asn Asp Asp Lys
Asp1 5 10 15Asn Ile Leu
Asp Asn Glu Asn Arg Glu Gly Leu Tyr Leu Cys Asp Val20 25
30Met Lys Asn Ser Asn Glu Leu Lys Arg Ile Asn Asp Asn
Phe Phe Lys35 40 45Leu
His5048117PRTPlasmodium falciparum 481Lys Asn Ser Asn Glu Leu Lys Arg Ile
Asn Asp Asn Phe Phe Lys Leu1 5 10
15His48211PRTPlasmodium falciparum 482Lys Arg Ile Asn Asp Asn
Phe Phe Lys Leu His1 5
1048340PRTPlasmodium falciparum 483His Ile Asn Asn Glu Tyr Thr Asn Lys
Asn Pro Lys Asn Cys Leu Leu1 5 10
15Tyr Lys Asn Glu Glu Arg Asn Tyr Asn Asp Asn Asn Ile Lys Asp
Tyr20 25 30Ile Asn Ser Met Asn Phe Lys
Lys35 4048439PRTPlasmodium falciparum 484His Ile Asn Asn
Glu Tyr Thr Asn Lys Asn Pro Lys Asn Cys Leu Leu1 5
10 15Tyr Lys Asn Glu Glu Arg Asn Tyr Asn Asp
Asn Asn Ile Lys Asp Tyr20 25 30Ile Asn
Ser Met Asn Phe Lys3548518PRTPlasmodium falciparum 485His Ile Asn Asn Glu
Tyr Thr Asn Lys Asn Pro Lys Asn Cys Leu Leu1 5
10 15Tyr Lys48645PRTPlasmodium falciparum 486Lys
Pro Cys Leu Tyr Lys Lys Cys Lys Ile Ser Gln Val Trp Trp Cys1
5 10 15Met Pro Val Lys Asp Thr Phe
Asn Thr Tyr Glu Arg Asn Asn Val Leu20 25
30Asn Ser Lys Ile Glu Asn Asn Ile Glu Lys Ile Pro His35
40 4548739PRTPlasmodium falciparum 487Lys Cys Lys Ile
Ser Gln Val Trp Trp Cys Met Pro Val Lys Asp Thr1 5
10 15Phe Asn Thr Tyr Glu Arg Asn Asn Val Leu
Asn Ser Lys Ile Glu Asn20 25 30Asn Ile
Glu Lys Ile Pro His3548811PRTPlasmodium falciparum 488Lys Ile Glu Asn Asn
Ile Glu Lys Ile Pro His1 5
1048923PRTPlasmodium falciparum 489Lys Asn Lys Thr Asn Gly Ser Lys Gly
Val Lys Gly Glu Tyr Glu Lys1 5 10
15Lys Lys Glu Thr Asn Gly His2049021PRTPlasmodium falciparum
490Lys Thr Asn Gly Ser Lys Gly Val Lys Gly Glu Tyr Glu Lys Lys Lys1
5 10 15Glu Thr Asn Gly
His2049116PRTPlasmodium falciparum 491Lys Gly Val Lys Gly Glu Tyr Glu Lys
Lys Lys Glu Thr Asn Gly His1 5 10
1549213PRTPlasmodium falciparum 492Lys Gly Glu Tyr Glu Lys Lys
Lys Glu Thr Asn Gly His1 5
1049360PRTPlasmodium falciparum 493Lys Thr Ile Glu Lys Ile Asn Lys Ser
Lys Ser Trp Phe Phe Glu Glu1 5 10
15Leu Asp Glu Ile Asp Lys Pro Leu Ala Lys Leu Arg Lys Arg Glu
Lys20 25 30Thr Gln Ile Asn Lys Thr Lys
Tyr Glu Arg Gly Asp Val Ile Ile Asp35 40
45Asn Thr Glu Ile Gln Lys Ile Ile Arg Asp Tyr His50 55
6049456PRTPlasmodium falciparum 494Lys Ile Asn Lys Ser
Lys Ser Trp Phe Phe Glu Glu Leu Asp Glu Ile1 5
10 15Asp Lys Pro Leu Ala Lys Leu Arg Lys Arg Glu
Lys Thr Gln Ile Asn20 25 30Lys Thr Lys
Tyr Glu Arg Gly Asp Val Ile Ile Asp Asn Thr Glu Ile35 40
45Gln Lys Ile Ile Arg Asp Tyr His50
5549539PRTPlasmodium falciparum 495Lys Pro Leu Ala Lys Leu Arg Lys Arg
Glu Lys Thr Gln Ile Asn Lys1 5 10
15Thr Lys Tyr Glu Arg Gly Asp Val Ile Ile Asp Asn Thr Glu Ile
Gln20 25 30Lys Ile Ile Arg Asp Tyr
His3549632PRTPlasmodium falciparum 496His Ile Met Leu Lys Ser Gln Met Tyr
Thr Asn Glu Gly Asn Lys Ser1 5 10
15Cys Glu Cys Ser Tyr Lys Lys Lys Ser Ser Ser Ser Asn Lys Val
His20 25 3049735PRTPlasmodium
falciparum 497Lys Leu Arg Lys Arg Glu Lys Thr Gln Ile Asn Lys Thr Lys Tyr
Glu1 5 10 15Arg Gly Asp
Val Ile Ile Asp Asn Thr Glu Ile Gln Lys Ile Ile Arg20 25
30Asp Tyr His3549832PRTPlasmodium falciparum 498Lys Arg
Glu Lys Thr Gln Ile Asn Lys Thr Lys Tyr Glu Arg Gly Asp1 5
10 15Val Ile Ile Asp Asn Thr Glu Ile
Gln Lys Ile Ile Arg Asp Tyr His20 25
3049929PRTPlasmodium falciparum 499Lys Thr Gln Ile Asn Lys Thr Lys Tyr
Glu Arg Gly Asp Val Ile Ile1 5 10
15Asp Asn Thr Glu Ile Gln Lys Ile Ile Arg Asp Tyr His20
2550048PRTPlasmodium falciparum 500Lys Pro Leu Ala Lys Leu Arg
Lys Arg Glu Lys Thr Gln Ile Asn Lys1 5 10
15Thr Lys Tyr Glu Arg Gly Asp Val Ile Ile Asp Asn Thr
Glu Ile Gln20 25 30Lys Ile Ile Arg Asp
Tyr His Thr Leu Asn Val His Lys Leu Asp His35 40
4550144PRTPlasmodium falciparum 501Lys Leu Arg Lys Arg Glu Lys
Thr Gln Ile Asn Lys Thr Lys Tyr Glu1 5 10
15Arg Gly Asp Val Ile Ile Asp Asn Thr Glu Ile Gln Lys
Ile Ile Arg20 25 30Asp Tyr His Thr Leu
Asn Val His Lys Leu Asp His35 4050241PRTPlasmodium
falciparum 502Lys Arg Glu Lys Thr Gln Ile Asn Lys Thr Lys Tyr Glu Arg Gly
Asp1 5 10 15Val Ile Ile
Asp Asn Thr Glu Ile Gln Lys Ile Ile Arg Asp Tyr His20 25
30Thr Leu Asn Val His Lys Leu Asp His35
4050338PRTPlasmodium falciparum 503Lys Thr Gln Ile Asn Lys Thr Lys Tyr
Glu Arg Gly Asp Val Ile Ile1 5 10
15Asp Asn Thr Glu Ile Gln Lys Ile Ile Arg Asp Tyr His Thr Leu
Asn20 25 30Val His Lys Leu Asp
His3550444PRTPlasmodium falciparum 504Lys Pro Leu Ala Lys Leu Arg Lys Arg
Glu Lys Thr Gln Ile Asn Lys1 5 10
15Thr Lys Tyr Glu Arg Gly Asp Val Ile Ile Asp Asn Thr Glu Ile
Gln20 25 30Lys Ile Ile Arg Asp Tyr His
Thr Leu Asn Val His35 4050540PRTPlasmodium falciparum
505Lys Leu Arg Lys Arg Glu Lys Thr Gln Ile Asn Lys Thr Lys Tyr Glu1
5 10 15Arg Gly Asp Val Ile Ile
Asp Asn Thr Glu Ile Gln Lys Ile Ile Arg20 25
30Asp Tyr His Thr Leu Asn Val His35
4050637PRTPlasmodium falciparum 506Lys Arg Glu Lys Thr Gln Ile Asn Lys
Thr Lys Tyr Glu Arg Gly Asp1 5 10
15Val Ile Ile Asp Asn Thr Glu Ile Gln Lys Ile Ile Arg Asp Tyr
His20 25 30Thr Leu Asn Val
His3550734PRTPlasmodium falciparum 507Lys Thr Gln Ile Asn Lys Thr Lys Tyr
Glu Arg Gly Asp Val Ile Ile1 5 10
15Asp Asn Thr Glu Ile Gln Lys Ile Ile Arg Asp Tyr His Thr Leu
Asn20 25 30Val His50832PRTPlasmodium
falciparum 508His Ile Met Leu Lys Ser Gln Met Tyr Thr Asn Glu Gly Asn Lys
Ser1 5 10 15Cys Glu Cys
Ser Tyr Lys Lys Lys Ser Ser Ser Ser Asn Lys Val His20 25
3050928PRTPlasmodium falciparum 509Lys Ser Gln Met Tyr
Thr Asn Glu Gly Asn Lys Ser Cys Glu Cys Ser1 5
10 15Tyr Lys Lys Lys Ser Ser Ser Ser Asn Lys Val
His20 2551018PRTPlasmodium falciparum 510Lys Ser Cys Glu
Cys Ser Tyr Lys Lys Lys Ser Ser Ser Ser Asn Lys1 5
10 15Val His51111PRTPlasmodium falciparum
511Lys Lys Lys Ser Ser Ser Ser Asn Lys Val His1 5
1051210PRTPlasmodium falciparum 512Lys Lys Ser Ser Ser Ser Asn
Lys Val His1 5 105139PRTPlasmodium
falciparum 513Lys Ser Ser Ser Ser Asn Lys Val His1
551430PRTPlasmodium falciparum 514His Ile Met Leu Lys Ser Gln Met Tyr Thr
Asn Glu Gly Asn Lys Ser1 5 10
15Cys Glu Cys Ser Tyr Lys Lys Lys Ser Ser Ser Ser Asn Lys20
25 3051524PRTPlasmodium falciparum 515His Ile
Met Leu Lys Ser Gln Met Tyr Thr Asn Glu Gly Asn Lys Ser1 5
10 15Cys Glu Cys Ser Tyr Lys Lys
Lys2051623PRTPlasmodium falciparum 516His Ile Met Leu Lys Ser Gln Met Tyr
Thr Asn Glu Gly Asn Lys Ser1 5 10
15Cys Glu Cys Ser Tyr Lys Lys2051722PRTPlasmodium falciparum
517His Ile Met Leu Lys Ser Gln Met Tyr Thr Asn Glu Gly Asn Lys Ser1
5 10 15Cys Glu Cys Ser Tyr
Lys2051836PRTPlasmodium falciparum 518His Asn Asn His Asn Ile Gln Ile Tyr
Lys Asp Lys Arg Ile Asn Phe1 5 10
15Met Asn Pro His Lys Val Met Tyr His Asp Asn Met Ser Lys Asn
Glu20 25 30Arg Thr Glu
Lys3551930PRTPlasmodium falciparum 519His Asn Asn His Asn Ile Gln Ile Tyr
Lys Asp Lys Arg Ile Asn Phe1 5 10
15Met Asn Pro His Lys Val Met Tyr His Asp Asn Met Ser Lys20
25 3052021PRTPlasmodium falciparum 520His
Asn Asn His Asn Ile Gln Ile Tyr Lys Asp Lys Arg Ile Asn Phe1
5 10 15Met Asn Pro His
Lys2052117PRTPlasmodium falciparum 521His Lys Val Met Tyr His Asp Asn Met
Ser Lys Asn Glu Arg Thr Glu1 5 10
15Lys52211PRTPlasmodium falciparum 522His Lys Val Met Tyr His
Asp Asn Met Ser Lys1 5 1052340PRTHomo
sapiens 523His Arg Glu Ile Cys Thr Ile Gln Ser Ser Gly Gly Ile Met Leu
Leu1 5 10 15Lys Asp Gln
Val Leu Arg Cys Ser Lys Ile Ala Gly Val Lys Val Ala20 25
30Glu Ile Thr Glu Leu Ile Leu Lys35
4052425PRTHomo sapiens 524His Arg Glu Ile Cys Thr Ile Gln Ser Ser Gly Gly
Ile Met Leu Leu1 5 10
15Lys Asp Gln Val Leu Arg Cys Ser Lys20 2552547PRTHomo
sapiens 525His Arg Glu Ile Cys Thr Ile Gln Ser Ser Gly Gly Ile Met Leu
Leu1 5 10 15Lys Asp Gln
Val Leu Arg Cys Ser Lys Ile Ala Gly Val Lys Val Ala20 25
30Glu Ile Thr Glu Leu Ile Leu Lys Ala Leu Glu Asn Asp
Gln Lys35 40 4552640PRTHomo sapiens
526His Arg Glu Ile Cys Thr Ile Gln Ala Ala Gly Gly Ile Met Leu Leu1
5 10 15Lys Asp Gln Val Leu Arg
Cys Ser Lys Ile Ala Gly Val Lys Val Ala20 25
30Glu Ile Thr Glu Leu Ile Leu Lys35 4052723PRTHomo
sapiens 527Lys Lys Met Gln Gln Glu Asn Met Lys Pro Gln Glu Gln Leu Thr
Leu1 5 10 15Glu Pro Tyr
Glu Arg Asp His2052822PRTHomo sapiens 528Lys Met Gln Gln Glu Asn Met Lys
Pro Gln Glu Gln Leu Thr Leu Glu1 5 10
15Pro Tyr Glu Arg Asp His2052938PRTHomo sapiens 529His Glu
Met Glu Glu Ser Lys Lys Asn Arg Val Glu Ile Asn Asp Val1 5
10 15Glu Pro Glu Val Phe Lys Glu Met
Met Cys Phe Ile Tyr Thr Gly Lys20 25
30Ala Pro Asn Leu Asp Lys3553032PRTHomo sapiens 530His Glu Met Glu Glu
Ser Lys Lys Asn Arg Val Glu Ile Asn Asp Val1 5
10 15Glu Pro Glu Val Phe Lys Glu Met Met Cys Phe
Ile Tyr Thr Gly Lys20 25 305319PRTHomo
sapiens 531Lys His Gly Glu Leu Lys Val Tyr Lys1
553214PRTHomo sapiens 532Lys Leu Ile Leu Gly Pro Gln Glu Glu Lys Gly Lys
Gln His1 5 105337PRTHomo sapiens 533Lys
Asn Arg Ile His His Lys1 553417PRTHomo sapiens 534His His
Asn Ser Ser Arg Lys Ser Thr Lys Lys Thr Asn Gln Ser Ser1 5
10 15Lys53516PRTHomo sapiens 535His Asn
Ser Ser Arg Lys Ser Thr Lys Lys Thr Asn Gln Ser Ser Lys1 5
10 1553620PRTHomo sapiens 536Lys His
His Asn Ile Leu Pro Lys Thr Leu Ala Asn Asp Lys His Ser1 5
10 15His Lys Pro His2053717PRTHomo
sapiens 537His His Asn Ile Leu Pro Lys Thr Leu Ala Asn Asp Lys His Ser
His1 5 10
15Lys53816PRTHomo sapiens 538His Asn Ile Leu Pro Lys Thr Leu Ala Asn Asp
Lys His Ser His Lys1 5 10
1553912PRTHomo sapiens 539His Asn Ile Leu Pro Lys Thr Leu Ala Asn Asp
Lys1 5 1054018PRTHomo sapiens 540Lys Asn
Thr Pro Asp Ser Lys Lys Ile Ser Ser Arg Asn Ile Asn Asp1 5
10 15His His54117PRTHomo sapiens 541Lys
Asn Thr Pro Asp Ser Lys Lys Ile Ser Ser Arg Asn Ile Asn Asp1
5 10 15His54227PRTHomo sapiens 542Lys
Asp Thr Cys Ile Gln Ser Pro Ser Lys Glu Cys Gln Lys Ser His1
5 10 15Pro Lys Ser Val Pro Val Ser
Ser Lys Lys Lys20 2554326PRTHomo sapiens 543Lys Asp Thr
Cys Ile Gln Ser Pro Ser Lys Glu Cys Gln Lys Ser His1 5
10 15Pro Lys Ser Val Pro Val Ser Ser Lys
Lys20 2554412PRTHomo sapiens 544His Pro Lys Ser Val Pro
Val Ser Ser Lys Lys Lys1 5 1054511PRTHomo
sapiens 545His Pro Lys Ser Val Pro Val Ser Ser Lys Lys1 5
1054610PRTHomo sapiens 546His Pro Lys Ser Val Pro Val Ser
Ser Lys1 5 1054714PRTHomo sapiens 547Lys
Ala Leu Gln Glu Lys Val Glu Ile Lys Gln Leu Asn His1 5
1054842PRTHomo sapiens 548Lys Thr Leu Phe Pro Leu Ile Glu
Ala Lys Lys Lys Asp Gln Val Thr1 5 10
15Ala Gln Glu Ile Phe Gln Asp Asn His Glu Asp Gly Pro Thr
Ala Lys20 25 30Lys Leu Lys Thr Glu Gln
Gly Gly Ala His35 4054925PRTHomo sapiens 549Lys Thr Leu
Phe Pro Leu Ile Glu Ala Lys Lys Lys Asp Gln Val Thr1 5
10 15Ala Gln Glu Ile Phe Gln Asp Asn His20
2555014PRTHomo sapiens 550Lys Leu Cys Val Phe Lys Lys Ile
Glu Arg His Ser Ile His1 5 1055111PRTHomo
sapiens 551Lys Leu Cys Val Phe Lys Lys Ile Glu Arg His1 5
1055222PRTHomo sapiens 552His Gly Pro Ser Phe Pro Leu Lys
Gly Ile Thr Glu Gln Gln Lys Glu1 5 10
15Gly Leu Glu Ile Val Lys2055315PRTHomo sapiens 553His Gly
Pro Ser Phe Pro Leu Lys Gly Ile Thr Glu Gln Gln Lys1 5
10 1555413PRTHomo sapiens 554His Thr Leu
Leu Lys Ile Leu Ser Thr Phe Leu Phe Lys1 5
1055514PRTHomo sapiens 555His Leu Leu Gly Asn Asn Asp Lys Asn Leu Leu
Pro Ser Lys1 5 1055619PRTHomo sapiens
556His Arg His Glu Gly Val Phe Ile Cys Arg Gly Lys Glu Asp Ala Leu1
5 10 15Val Thr Lys55717PRTHomo
sapiens 557His Glu Gly Val Phe Ile Cys Arg Gly Lys Glu Asp Ala Leu Val
Thr1 5 10
15Lys55820PRTHomo sapiens 558His Ser Gly Gly Asn Arg Gly Arg Gly Arg Gly
Gly Lys Arg Gly Asn1 5 10
15Gln Ser Gly Lys2055915PRTHomo sapiens 559Lys Arg Gly Asn Gln Ser Gly
Lys Asn Val Met Val Glu Pro His1 5 10
1556017PRTHomo sapiens 560Lys Arg Gly Asn Gln Ser Gly Lys
Asn Val Met Val Glu Pro His Arg1 5 10
15His56123PRTHomo sapiens 561Lys Lys Met Gln Gln Glu Asn Met
Lys Pro Gln Glu Gln Leu Thr Leu1 5 10
15Glu Pro Tyr Glu Arg Asp His2056222PRTHomo sapiens 562Lys
Met Gln Gln Glu Asn Met Lys Pro Gln Glu Gln Leu Thr Leu Glu1
5 10 15Pro Tyr Glu Arg Asp
His2056316PRTHomo sapiens 563His Ala Tyr Pro Glu Asp Ala Glu Asn Lys Glu
Lys Glu Thr Ala Lys1 5 10
1556422PRTHomo sapiens 564Lys Glu Ala Asn Val Lys Cys Pro Gln Ile Val
Ile Ala Phe Tyr Glu1 5 10
15Glu Arg Leu Thr Trp His2056525PRTHomo sapiens 565Lys Val Leu Asp Arg
Arg Val Val Lys Gly Gln Val Glu Tyr Leu Leu1 5
10 15Lys Trp Lys Gly Phe Ser Glu Glu His20
2556617PRTHomo sapiens 566Lys Gly Gln Val Glu Tyr Leu Leu Lys Trp
Lys Gly Phe Ser Glu Glu1 5 10
15His56732PRTHomo sapiens 567Lys Ser Glu Val Ala Ala Gly Val Lys Lys
Ser Gly Pro Leu Pro Ser1 5 10
15Ala Glu Arg Leu Glu Asn Val Leu Phe Gly Pro His Asp Cys Ser His20
25 3056828PRTHomo sapiens 568Lys Ser Glu
Val Ala Ala Gly Val Lys Lys Ser Gly Pro Leu Pro Ser1 5
10 15Ala Glu Arg Leu Glu Asn Val Leu Phe
Gly Pro His20 2556928PRTHomo sapiens 569Lys Ala Ala Glu
Tyr Gly Lys Lys Ala Lys Ser Glu Thr Phe Arg Leu1 5
10 15Leu His Ala Lys Asn Ile Ile Arg Pro Gln
Leu Lys20 2557020PRTHomo sapiens 570Lys Ala Ala Glu Tyr
Gly Lys Lys Ala Lys Ser Glu Thr Phe Arg Leu1 5
10 15Leu His Ala Lys2057111PRTHomo sapiens 571Lys
Ser Glu Thr Phe Arg Leu Leu His Ala Lys1 5
1057211PRTHomo sapiens 572His Ala Lys Asn Ile Ile Arg Pro Gln Leu Lys1
5 1057312PRTHomo sapiens 573His Met Met Leu
Lys Ile Ala Glu Glu Leu Pro Lys1 5
1057417PRTHomo sapiens 574His Ser Leu Asp His Leu Leu Lys Leu Tyr Cys Asn
Val Asp Ser Asn1 5 10
15Lys57513PRTHomo sapiens 575His Leu Leu Lys Leu Tyr Cys Asn Val Asp Ser
Asn Lys1 5 1057610PRTHomo sapiens 576Lys
Ala Lys Glu Arg Leu Glu Ala Lys His1 5
105778PRTHomo sapiens 577Lys Asp Arg Gln His Thr Leu Lys1
55789PRTHomo sapiens 578Lys Asp Arg Gln His Thr Leu Lys His1
5 57912PRTHomo sapiens 579Lys Glu Thr Cys Ser Glu Lys Ser Thr Asn Leu
His1 5 1058016PRTHomo sapiens 580Lys Thr
Glu Glu Ile Ser Glu Val Lys Met Asp Ala Glu Phe Gly His1 5
10 1558124PRTHomo sapiens 581Lys Thr
Glu Glu Ile Ser Glu Val Lys Met Asp Ala Glu Phe Gly His1 5
10 15Asp Ser Gly Phe Glu Val Arg
His2058217PRTHomo sapiens 582Lys Lys Tyr Val Arg Ala Glu Gln Lys Asp Arg
Gln His Thr Leu Lys1 5 10
15His58316PRTHomo sapiens 583Lys Tyr Val Arg Ala Glu Gln Lys Asp Arg Gln
His Thr Leu Lys His1 5 10
1558413PRTHomo sapiens 584Lys Lys Tyr Val Arg Ala Glu Gln Lys Asp Arg
Gln His1 5 1058513PRTHomo sapiens 585Lys
Tyr Val Arg Ala Glu Gln Lys Asp Arg Gln His Thr1 5
1058616PRTHomo sapiens 586His His Val Phe Asn Met Leu Lys Lys
Tyr Val Arg Ala Glu Gln Lys1 5 10
1558715PRTHomo sapiens 587His Val Phe Asn Met Leu Lys Lys Tyr
Val Arg Ala Glu Gln Lys1 5 10
1558824PRTHomo sapiens 588His His Val Phe Asn Met Leu Lys Lys Tyr
Val Arg Ala Glu Gln Lys1 5 10
15Asp Arg Gln His Thr Leu Lys His2058923PRTHomo sapiens 589His Val
Phe Asn Met Leu Lys Lys Tyr Val Arg Ala Glu Gln Lys Asp1 5
10 15Arg Gln His Thr Leu Lys
His2059015PRTHomo sapiens 590His Ala His Phe Gln Lys Ala Lys Glu Arg Leu
Glu Ala Lys His1 5 10
1559114PRTHomo sapiens 591His Ala His Phe Gln Lys Ala Lys Glu Arg Leu Glu
Ala Lys1 5 1059212PRTHomo sapiens 592His
Phe Gln Lys Ala Lys Glu Arg Leu Glu Ala Lys1 5
1059330PRTHomo sapiens 593His Gln Glu Arg Met Asp Val Cys Glu Thr
His Leu His Trp His Thr1 5 10
15Val Ala Lys Glu Thr Cys Ser Glu Lys Ser Thr Asn Leu His20
25 3059425PRTHomo sapiens 594His Gln Glu Arg Met
Asp Val Cys Glu Thr His Leu His Trp His Thr1 5
10 15Val Ala Lys Glu Thr Cys Ser Glu Lys20
2559513PRTHomo sapiens 595His Trp His Thr Val Ala Lys Glu Thr Cys
Ser Glu Lys1 5 1059611PRTHomo sapiens
596His Thr Val Ala Lys Glu Thr Cys Ser Glu Lys1 5
1059715PRTHomo sapiens 597His Leu His Trp His Thr Val Ala Lys Glu
Thr Cys Ser Glu Lys1 5 10
1559823PRTHomo sapiens 598His Met Asn Val Gln Asn Gly Lys Trp Glu Ser
Asp Pro Ser Gly Thr1 5 10
15Lys Thr Cys Ile Gly Thr Lys2059917PRTHomo sapiens 599His Met Asn Val
Gln Asn Gly Lys Trp Glu Ser Asp Pro Ser Gly Thr1 5
10 15Lys60019PRTHuman immunodeficiency virus
600His Cys Leu Val Cys Phe Gln Lys Lys Gly Leu Gly Ile Ser Tyr Gly1
5 10 15Arg Lys
Lys60127PRTTriticum aestivum 601His Lys Asp Arg Leu Thr Lys Lys Val Val
Asp Ile Ala Arg Glu Val1 5 10
15Ala Lys Val Asp Val Pro Glu Tyr Arg Arg His20
2560227PRTTriticum aestivum 602His Lys Glu Arg Leu Asp Arg Lys Val Val
Asp Val Ala Arg Glu Val1 5 10
15Ala Lys Val Glu Val Pro Ser Tyr Arg Arg His20
2560327PRTTriticum aestivum 603His Lys Glu Arg Leu Asp Arg Lys Val Val
Asp Val Ala Arg Glu Val1 5 10
15Ala Lys Met Glu Val Pro Ser Tyr Arg Arg His20
2560423PRTTriticum aestivum 604His Leu Gln Pro Lys Trp Lys Pro Ser Leu
Ser Trp Phe Lys Asn Ala1 5 10
15Glu Ser Arg Leu Asn His His2060514PRTTriticum aestivum 605His Leu
Gln Pro Lys Trp Lys Pro Ser Leu Ser Trp Phe Lys1 5
1060619PRTTriticum aestivum 606Lys Trp Lys Pro Ser Leu Ser Trp
Phe Lys Asn Ala Glu Ser Arg Leu1 5 10
15Asn His His60718PRTTriticum aestivum 607Lys Trp Lys Pro
Ser Leu Ser Trp Phe Lys Asn Ala Glu Ser Arg Leu1 5
10 15Asn His60817PRTTriticum aestivum 608Lys
Pro Ser Leu Ser Trp Phe Lys Asn Ala Glu Ser Arg Leu Asn His1
5 10 15His60916PRTTriticum aestivum
609Lys Pro Ser Leu Ser Trp Phe Lys Asn Ala Glu Ser Arg Leu Asn His1
5 10 1561032PRTTriticum
aestivum 610His His Ala Ile Ala Leu Gly Leu His Thr Thr Thr Leu Ile Leu
Val1 5 10 15Lys Gly Ala
Leu Asp Ala Arg Gly Ser Lys Leu Met Pro Asp Lys Lys20 25
3061131PRTTriticum aestivum 611His Ala Ile Ala Leu Gly
Leu His Thr Thr Thr Leu Ile Leu Val Lys1 5
10 15Gly Ala Leu Asp Ala Arg Gly Ser Lys Leu Met Pro
Asp Lys Lys20 25 3061226PRTTriticum
aestivum 612His His Ala Ile Ala Leu Gly Leu His Thr Thr Thr Leu Ile Leu
Val1 5 10 15Lys Gly Ala
Leu Asp Ala Arg Gly Ser Lys20 2561325PRTTriticum
aestivum 613His Ala Ile Ala Leu Gly Leu His Thr Thr Thr Leu Ile Leu Val
Lys1 5 10 15Gly Ala Leu
Asp Ala Arg Gly Ser Lys20 2561424PRTTriticum aestivum
614His Thr Thr Thr Leu Ile Leu Val Lys Gly Ala Leu Asp Ala Arg Gly1
5 10 15Ser Lys Leu Met Pro Asp
Lys Lys2061523PRTTriticum aestivum 615His Thr Thr Thr Leu Ile Leu Val Lys
Gly Ala Leu Asp Ala Arg Gly1 5 10
15Ser Lys Leu Met Pro Asp Lys2061618PRTTriticum aestivum 616His
Thr Thr Thr Leu Ile Leu Val Lys Gly Ala Leu Asp Ala Arg Gly1
5 10 15Ser Lys61744PRTTriticum
aestivum 617His His His Leu Ala Ile Ala Ile Leu Phe Leu Ile Ala Gly His
Met1 5 10 15Tyr Arg Thr
Asn Trp Gly Ile Gly His Gly Leu Lys Asp Ile Leu Glu20 25
30Ala His Lys Gly Pro Phe Thr Gly Gln Gly His Lys35
4061843PRTTriticum aestivum 618His His Leu Ala Ile Ala Ile
Leu Phe Leu Ile Ala Gly His Met Tyr1 5 10
15Arg Thr Asn Trp Gly Ile Gly His Gly Leu Lys Asp Ile
Leu Glu Ala20 25 30His Lys Gly Pro Phe
Thr Gly Gln Gly His Lys35 4061942PRTTriticum aestivum
619His Leu Ala Ile Ala Ile Leu Phe Leu Ile Ala Gly His Met Tyr Arg1
5 10 15Thr Asn Trp Gly Ile Gly
His Gly Leu Lys Asp Ile Leu Glu Ala His20 25
30Lys Gly Pro Phe Thr Gly Gln Gly His Lys35
4062030PRTTriticum aestivum 620His Met Tyr Arg Thr Asn Trp Gly Ile Gly
His Gly Leu Lys Asp Ile1 5 10
15Leu Glu Ala His Lys Gly Pro Phe Thr Gly Gln Gly His Lys20
25 3062120PRTTriticum aestivum 621His Gly Leu
Lys Asp Ile Leu Glu Ala His Lys Gly Pro Phe Thr Gly1 5
10 15Gln Gly His Lys2062217PRTTriticum
aestivum 622Lys Asp Ile Leu Glu Ala His Lys Gly Pro Phe Thr Gly Gln Gly
His1 5 10
15Lys62311PRTTriticum aestivum 623His Lys Gly Pro Phe Thr Gly Gln Gly His
Lys1 5 1062410PRTTriticum aestivum 624Lys
Gly Pro Phe Thr Gly Gln Gly His Lys1 5
1062516PRTOryza sativa 625Lys Phe Pro Asp Val Ile His Ala Phe Lys Pro Asn
Pro Arg Ser His1 5 10
1562610PRTOryza sativa 626Lys Phe Pro Asp Val Ile His Ala Phe Lys1
5 1062710PRTOryza sativa 627Lys Ala Arg Tyr Val
Lys Phe His Trp Lys1 5 1062833PRTOryza
sativa 628His Pro Lys Val Ser Pro Glu Leu Arg Ala Ile Trp Val Asn Tyr
Leu1 5 10 15Ser Gln Cys
Asp Glu Ser Leu Gly Val Lys Ile Ala Asn Leu Asn Val20 25
30Lys62945PRTOryza sativa 629His Arg Asp Glu Glu Val Asp
Tyr Tyr Pro Ser Arg His Ala Pro Leu1 5 10
15Arg His Ala Pro Pro Thr Pro Ile Thr Pro Arg Pro Val
Val Gly Arg20 25 30Arg Gln Lys Ala Thr
Ile His Lys Gln Asn Asp Phe Lys35 40
4563011PRTOryza sativa 630Lys Ala Thr Ile His Lys Gln Asn Asp Phe Lys1
5 1063127PRTOryza sativa 631His Ala Pro Pro
Thr Pro Ile Pro Arg Pro Val Val Gly Arg Arg Gln1 5
10 15Lys Ala Thr Ile His Lys Gln Asn Asp Phe
Lys20 2563249PRTOryza sativa 632Lys Phe Arg Pro Ser Ser
Ser Phe Asp Thr Lys Thr Thr Thr Thr Asn1 5
10 15Ala Gly Ala Pro Val Trp Asn Asp Asn Glu Ala Leu
Thr Val Gly Pro20 25 30Arg Gly Pro Ile
Leu Leu Glu Asp Tyr His Leu Ile Glu Lys Val Ala35 40
45His63342PRTOryza sativa 633Lys Phe Arg Pro Ser Ser Ser Phe
Asp Thr Lys Thr Thr Thr Thr Asn1 5 10
15Ala Gly Ala Pro Val Trp Asn Asp Asn Glu Ala Leu Thr Val
Gly Pro20 25 30Arg Gly Pro Ile Leu Leu
Glu Asp Tyr His35 406349PRTOryza sativa 634Lys Val Lys
Ala His Phe Gln Lys His1 56358PRTOryza sativa 635Lys Val
Lys Ala His Phe Gln Lys1 563612PRTOryza sativa 636Lys Asp
Tyr Glu Ile Asp Lys Asp Asp Leu Ile His1 5
1063720PRTOryza sativa 637His Met Lys Gln Cys Phe Ala Phe Cys Ala Val
Phe Pro Lys Asp Tyr1 5 10
15Glu Ile Asp Lys2063814PRTOryza sativa 638His Met Lys Gln Cys Phe Ala
Phe Cys Ala Val Phe Pro Lys1 5
1063949PRTOryza sativa 639His Val Phe Trp Glu Leu Val Trp Arg Ser Phe Phe
Gln Asn Val Lys1 5 10
15Gln Ile Gly Ser Ile Phe Gln Arg Lys Val Tyr Arg Tyr Gly Gln Ser20
25 30Asp Val Thr Thr Ser Lys Ile His Asp Leu
Met His Asp Leu Ala Val35 40
45His64034PRTOryza sativa 640Lys Gln Ile Gly Ser Ile Phe Gln Arg Lys Val
Tyr Arg Tyr Gly Gln1 5 10
15Ser Asp Val Thr Thr Ser Lys Ile His Asp Leu Met His Asp Leu Ala20
25 30Val His64129PRTOryza sativa 641Lys Gln
Ile Gly Ser Ile Phe Gln Arg Lys Val Tyr Arg Tyr Gly Gln1 5
10 15Ser Asp Val Thr Thr Ser Lys Ile
His Asp Leu Met His20 2564225PRTOryza sativa 642Lys Gln
Ile Gly Ser Ile Phe Gln Arg Lys Val Tyr Arg Tyr Gly Gln1 5
10 15Ser Asp Val Thr Thr Ser Lys Ile
His20 256439PRTOryza sativa 643Lys His Gly Val Ser Ala
Gly Ile Lys1 564415PRTOryza sativa 644His Thr Val Phe Asp
Tyr Gly Lys Met Arg Val Gly Phe Ala Lys1 5
10 1564517PRTOryza sativa 645His Ser Arg Tyr Lys Ser
Gly Gln Ser Ser Thr Tyr Gln Lys Asn Gly1 5
10 15Lys64629PRTOryza sativa 646Lys Gln Glu Ala Met Val
Leu Lys Gln Glu Ile Asn Leu Leu Gln Lys1 5
10 15Gly Leu Arg Tyr Ile Tyr Gly Asn Arg Ala Asn Glu
His20 2564722PRTOryza sativa 647Lys Gln Glu Ile Asn Leu
Leu Gln Lys Gly Leu Arg Tyr Ile Tyr Gly1 5
10 15Asn Arg Ala Asn Glu His2064838PRTOryza sativa
648Lys Ser Lys Glu Gly Met Leu Lys Ala Ala Asn Glu Ile Leu Gln Glu1
5 10 15Lys Ile Val Glu Gln Asn
Gly Leu Ile Asp Val Gly Met Met Val Ala20 25
30Asp Gln Gln Asn Gly His3564931PRTOryza sativa 649Lys Ala Ala Asn
Glu Ile Leu Gln Glu Lys Ile Val Glu Gln Asn Gly1 5
10 15Leu Ile Asp Val Gly Met Met Val Ala Asp
Gln Gln Asn Gly His20 25 3065011PRTZea
mays 650Lys Val Leu Ala Ala His Arg Tyr Gly Ile Lys1 5
1065117PRTZea mays 651Lys Leu Lys Ile Ala Met Lys His Leu
Ile Pro Arg Val Leu Glu Gln1 5 10
15His6528PRTZea mays 652Lys Leu Lys Ile Ala Met Lys His1
565333PRTZea mays 653Lys Thr Ser Leu Ala Ser Ser Ile Ala Lys Ala
Leu Asn Arg Lys Phe1 5 10
15Ile Arg Ile Ser Leu Gly Gly Val Lys Asp Glu Ala Asp Ile Arg Gly20
25 30His65424PRTZea mays 654Lys Ala Leu Asn
Arg Lys Phe Ile Arg Ile Ser Leu Gly Gly Val Lys1 5
10 15Asp Glu Ala Asp Ile Arg Gly
His2065519PRTZea mays 655Lys Phe Ile Arg Ile Ser Leu Gly Gly Val Lys Asp
Glu Ala Asp Ile1 5 10
15Arg Gly His65637PRTZea mays 656Lys Val Arg Leu Ser Lys Ala Thr Glu Leu
Val Asp Arg His Leu Gln1 5 10
15Ser Ile Leu Val Ala Glu Lys Ile Thr Gln Lys Val Glu Gly Gln Leu20
25 30Ser Lys Ser Gln Lys3565724PRTZea
mays 657His Leu Gln Ser Ile Leu Val Ala Glu Lys Ile Thr Gln Lys Val Glu1
5 10 15Gly Gln Leu Ser
Lys Ser Gln Lys2065814PRTZea mays 658Lys Val Arg Leu Ser Lys Ala Thr Glu
Leu Val Asp Arg His1 5 1065934PRTZea mays
659Lys Val Gly Gly Ser Ala Val Glu Ser Ser Lys Gln Asp Thr Lys Asn1
5 10 15Gly Lys Glu Pro Ile His
Trp His Ser Lys Gly Val Ala Ala Arg Ala20 25
30Leu His66024PRTZea mays 660Lys Val Gly Gly Ser Ala Val Glu Ser Ser
Lys Gln Asp Thr Lys Asn1 5 10
15Gly Lys Glu Pro Ile His Trp His2066122PRTZea mays 661Lys Val Gly
Gly Ser Ala Val Glu Ser Ser Lys Gln Asp Thr Lys Asn1 5
10 15Gly Lys Glu Pro Ile His2066224PRTZea
mays 662Lys Gln Asp Thr Lys Asn Gly Lys Glu Pro Ile His Trp His Ser Lys1
5 10 15Gly Val Ala Ala
Arg Ala Leu His2066312PRTZea mays 663Lys Gln Asp Thr Lys Asn Gly Lys Glu
Pro Ile His1 5 1066433PRTZea mays 664His
Arg Asp Leu Arg Arg Ala Arg Ala Ala Ala Leu Asn Ile Val Pro1
5 10 15Thr Ser Thr Gly Ala Ala Lys
Ala Val Ser Leu Val Leu Pro Asn Leu20 25
30Lys66518PRTZea mays 665Lys Val Leu Asp Gln Lys Phe Gly Ile Ile Lys Gly
Thr Met Thr Thr1 5 10
15Thr His66616PRTZea mays 666His Ile Gln Ala Gly Ala Lys Lys Val Leu Ile
Thr Ala Pro Gly Lys1 5 10
1566728PRTZea mays 667His Gly Arg Gly Asp Ala Ser Pro Leu Asp Val Ile
Ala Ile Asn Asp1 5 10
15Thr Gly Gly Val Lys Gln Ala Ser His Leu Leu Lys20
2566851PRTZea mays 668Lys Val Arg Arg Val Leu Ser Lys Asp Tyr Ser Ser Leu
Lys Gln Leu1 5 10 15Met
Thr Leu Met Met Asp Asp Asp Ile Ser Lys His Leu Gln Ile Ile20
25 30Glu Ser Gly Leu Glu Glu Arg Glu Asp Lys Val
Trp Met Lys Glu Asn35 40 45Ile Ile
Lys5066928PRTZea mays 669Lys Val Arg Arg Val Leu Ser Lys Asp Tyr Ser Ser
Leu Lys Gln Leu1 5 10
15Met Thr Leu Met Met Asp Asp Asp Ile Ser Lys His20
2567024PRTZea mays 670His Leu Gln Ile Ile Glu Ser Gly Leu Glu Glu Arg Glu
Asp Lys Val1 5 10 15Trp
Met Lys Glu Asn Ile Ile Lys2067117PRTZea mays 671His Asp Leu Arg Glu Asn
Ile Ile Met Lys Ala Asp Asp Leu Ala Ser1 5
10 15Lys67218PRTZea mays 672His Val Gln Asn Leu Glu Asn
Val Ile Gly Lys Asp Glu Ala Leu Ala1 5 10
15Ser Lys67321PRTZea mays 673Lys Lys Gln Gly Tyr Glu Leu
Arg Gln Leu Lys Asp Leu Asn Glu Leu1 5 10
15Gly Gly Ser Leu His2067420PRTZea mays 674Lys Gln Gly
Tyr Glu Leu Arg Gln Leu Lys Asp Leu Asn Glu Leu Gly1 5
10 15Gly Ser Leu His2067528PRTZea mays
675Lys Leu Tyr Leu Lys Ser Arg Leu Lys Glu Leu Ile Leu Glu Trp Ser1
5 10 15Ser Glu Asn Gly Met Asp
Ala Met Asn Ile Leu His20 2567639PRTZea mays 676His Leu
Gln Leu Leu Gln Leu Asn Gly Met Val Glu Arg Leu Pro Asn1 5
10 15Lys Val Cys Asn Leu Ser Lys Leu
Arg Tyr Leu Arg Gly Tyr Lys Asp20 25
30Gln Ile Pro Asn Ile Gly Lys3567731PRTZea mays 677His Leu Gln Leu Leu
Gln Leu Asn Gly Met Val Glu Arg Leu Pro Asn1 5
10 15Lys Val Cys Asn Leu Ser Lys Leu Arg Tyr Leu
Arg Gly Tyr Lys20 25 3067823PRTZea mays
678His Leu Gln Leu Leu Gln Leu Asn Gly Met Val Glu Arg Leu Pro Asn1
5 10 15Lys Val Cys Asn Leu Ser
Lys2067914PRTZea mays 679His Asn Ser Asn Lys Leu Pro Lys Ser Val Gly Glu
Leu Lys1 5 1068011PRTZea mays 680Lys Leu
Pro Lys Ser Val Gly Glu Leu Lys His1 5
1068118PRTZea mays 681His Leu Ser Val Arg Val Glu Ser Met Gln Lys His Lys
Glu Ile Ile1 5 10 15Tyr
Lys6828PRTZea mays 682Lys His Lys Glu Ile Ile Tyr Lys1
568323PRTZea mays 683Lys Leu Arg Asp Ile Leu Gln Glu Ser Gln Lys Phe Leu
Leu Val Leu1 5 10 15Asp
Leu Ala Leu Phe Lys His2068446PRTZea mays 684His Ala Phe Ser Gly Ala Glu
Ile Lys Asp Gln Leu Leu Arg Met Lys1 5 10
15Leu Gln Asp Thr Ala Glu Cys Ile Ala Lys Arg Leu Gly
Gln Cys Pro20 25 30Leu Ala Ala Lys Val
Leu Gly Ser Arg Met Cys Arg Arg Lys35 40
4568516PRTZea mays 685His Ala Phe Ser Gly Ala Glu Ile Lys Asp Gln Leu
Leu Arg Met Lys1 5 10
1568647PRTZea mays 686Lys Leu Gln Asp Thr Ala Glu Glu Ile Ala Lys Arg Leu
Gly Gln Cys1 5 10 15Pro
Leu Ala Ala Lys Val Leu Gly Ser Arg Met Cys Arg Arg Lys Asp20
25 30Ile Ala Glu Trp Lys Ala Ala Asp Val Trp Phe
Glu Lys Ser His35 40 4568727PRTZea mays
687Lys Val Leu Gly Ser Arg Met Cys Arg Arg Lys Asp Ile Ala Glu Trp1
5 10 15Lys Ala Ala Asp Val Trp
Phe Glu Lys Ser His20 2568817PRTZea mays 688Lys Asp Ile
Ala Glu Trp Lys Ala Ala Asp Val Trp Phe Glu Lys Ser1 5
10 15His68911PRTZea mays 689Lys Ala Ala Asp
Val Trp Phe Glu Lys Ser His1 5
1069040PRTZea mays 690His Val Pro Thr Thr Thr Ser Leu Pro Thr Ser Lys Val
Phe Gly Arg1 5 10 15Asn
Ser Asp Arg Asp Arg Ile Val Lys Phe Leu Leu Gly Lys Thr Thr20
25 30Thr Ala Glu Ala Ser Ser Thr Lys35
4069118PRTZea mays 691Lys Ala Ile Leu Thr Glu Ala Lys Gln Leu Arg Asp
Leu Leu Gly Leu1 5 10
15Pro His69229PRTZea mays 692Lys Ala Lys Ala Lys Ser Gly Lys Gly Pro Leu
Leu Arg Glu Asp Glu1 5 10
15Ser Ser Ser Thr Ala Thr Thr Val Met Lys Pro Phe His20
2569329PRTZea mays 693Lys Ser Pro His Arg Gly Lys Leu Glu Ser Trp Leu Arg
Arg Leu Lys1 5 10 15Glu
Ala Phe Tyr Asp Ala Glu Asp Leu Leu Asp Glu His20
2569416PRTZea mays 694Lys Ser Pro His Arg Gly Lys Leu Glu Ser Trp Leu Arg
Arg Leu Lys1 5 10
1569513PRTZea mays 695His Arg Gly Lys Leu Glu Ser Trp Leu Arg Arg Leu
Lys1 5 106967PRTZea mays 696Lys Ser Pro
His Arg Gly Lys1 56978PRTZea mays 697Lys Gln Ala Ser His
Leu Leu Lys1 569823PRTArtificial SequenceDescription of
Artificial Sequence Replikin formula sequence 698Xaa Cys Xaa Xaa
His Cys Xaa Xaa Cys Xaa Xaa Xaa Lys Xaa Leu Xaa1 5
10 15Xaa Xaa Xaa Xaa Arg Lys
Lys206998PRTMycobacterium leprae 699Lys Val Met Arg Thr Asp Lys His1
570017PRTMycobacterium tuberculosis 700His Pro Arg Pro Lys Val
Ala Ala Ala Leu Lys Asp Ser Tyr Arg Leu1 5
10 15Lys70111PRTMycobacterium tuberculosis 701His Pro
Arg Pro Lys Val Ala Ala Ala Leu Lys1 5
1070219PRTMycobacterium tuberculosis 702Lys Ser Ala Gln Lys Trp Pro Asp
Lys Phe Leu Ala Gly Ala Ala Gln1 5 10
15Val Ala His70315PRTEscherichia coli 703His Ala Trp Gln His
Gln Gly Lys Thr Leu Phe Ile Ser Arg Lys1 5
10 1570411PRTEscherichia coli 704His Gln Gly Lys Thr
Leu Phe Ile Ser Arg Lys1 5
1070519PRTAgrobacterium tumefaciens 705His Ser Asp Gln Gln Leu Ala Val
Met Ile Ala Ala Lys Arg Leu Asp1 5 10
15Asp Tyr Lys70633PRTAgrobacterium tumefaciens 706His Leu
Leu Asp His Pro Ala Ser Val Gly Gln Leu Asp Leu Arg Ala1 5
10 15Met Leu Ala Val Glu Glu Val Lys
Ile Asp Asn Pro Val Tyr Met Glu20 25
30Lys70729PRTAgrobacterium tumefaciens 707His Pro Ala Ser Val Gly Gln Leu
Asp Leu Arg Ala Met Leu Ala Val1 5 10
15Glu Glu Val Lys Ile Asp Asn Pro Val Tyr Met Glu Lys20
2570839PRTAgrobacterium tumefaciens 708Lys Cys Val Met Ala
Lys Asn Cys Asn Ile Lys Cys Pro Ala Gly Leu1 5
10 15Thr Thr Asn Gln Glu Ala Phe Asn Gly Asp Pro
Arg Ala Leu Ala Gln20 25 30Tyr Leu Met
Asn Ile Ala His3570934PRTAgrobacterium tumefaciens 709Lys Asn Cys Asn Ile
Lys Cys Pro Ala Gly Leu Thr Thr Asn Gln Glu1 5
10 15Ala Phe Asn Gly Asp Pro Arg Ala Leu Ala Gln
Tyr Leu Met Asn Ile20 25 30Ala
His71027PRTAgrobacterium tumefaciens 710His His Asp Thr Tyr Ser Ile Glu
Asp Leu Ala Gln Leu Ile His Asp1 5 10
15Ala Lys Ala Ala Arg Val Arg Val Ile Val Lys20
2571126PRTAgrobacterium tumefaciens 711His Asp Thr Tyr Ser Ile Glu
Asp Leu Ala Gln Leu Ile His Asp Ala1 5 10
15Lys Ala Ala Arg Val Arg Val Ile Val Lys20
2571213PRTAgrobacterium tumefaciens 712His Asp Ala Lys Ala Ala Arg
Val Arg Val Ile Val Lys1 5
1071342PRTAgrobacterium tumefaciens 713Lys Ile Gly Gln Gly Ala Lys Pro
Gly Glu Gly Gly Gln Leu Pro Ser1 5 10
15Pro Lys Val Thr Val Glu Ile Ala Ala Ala Arg Gly Gly Thr
Pro Gly20 25 30Val Glu Leu Val Ser Pro
Pro Pro His His35 4071441PRTAgrobacterium tumefaciens
714Lys Ile Gly Gln Gly Ala Lys Pro Gly Glu Gly Gly Gln Leu Pro Ser1
5 10 15Pro Lys Val Thr Val Glu
Ile Ala Ala Ala Arg Gly Gly Thr Pro Gly20 25
30Val Glu Leu Val Ser Pro Pro Pro His35
4071546PRTAgrobacterium tumefaciens 715Lys Ala Ser Glu Ile Thr Lys Thr
Leu Ala Ser Gly Ala Met Ser His1 5 10
15Gly Ala Leu Val Ala Ala Ala His Glu Ala Val Ala His Gly
Thr Asn20 25 30Met Val Gly Gly Met Ser
Asn Ser Gly Glu Gly Gly Glu His35 40
4571629PRTAgrobacterium tumefaciens 716Lys Ala Ser Glu Ile Thr Lys Thr
Leu Ala Ser Gly Ala Met Ser His1 5 10
15Gly Ala Leu Val Ala Ala Ala His Glu Ala Val Ala His20
2571724PRTAgrobacterium tumefaciens 717Lys Ala Ser Glu Ile
Thr Lys Thr Leu Ala Ser Gly Ala Met Ser His1 5
10 15Gly Ala Leu Val Ala Ala Ala
His2071816PRTAgrobacterium tumefaciens 718Lys Ala Ser Glu Ile Thr Lys Thr
Leu Ala Ser Gly Ala Met Ser His1 5 10
1571927PRTAgrobacterium tumefaciens 719Lys Arg Tyr Phe Pro
Asn Val Lys Thr Pro Val Gly Gly Val Thr Phe1 5
10 15Ala Val Ile Ala Gln Ala Val Ala Asp Trp His20
2572048PRTAgrobacterium tumefaciens 720His His Ile Ala
Ala Gly Leu Gly Phe Gly Ala Ser Ala Val Tyr Pro1 5
10 15Leu Gly Val Gln Phe Arg Ala Glu Glu Lys
Phe Gly Ala Asp Ala Asp20 25 30Lys Ala
Phe Lys Arg Phe Ala Lys Ala Ala Glu Lys Ser Leu Met Lys35
40 4572148PRTAgrobacterium tumefaciens 721His His Ile
Ala Ala Gly Leu Gly Phe Gly Ala Ser Ala Val Tyr Pro1 5
10 15Leu Gly Val Gln Phe Arg Ala Glu Glu
Lys Phe Gly Ala Asp Ala Asp20 25 30Lys
Ala Phe Lys Arg Phe Ala Lys Ala Ala Glu Lys Ser Leu Met Lys35
40 4572244PRTAgrobacterium tumefaciens 722His His
Ile Ala Ala Gly Leu Gly Phe Gly Ala Ser Ala Val Tyr Pro1 5
10 15Leu Gly Val Gln Phe Arg Ala Glu
Glu Lys Phe Gly Ala Asp Ala Asp20 25
30Lys Ala Phe Lys Arg Phe Ala Lys Ala Ala Glu Lys35
4072340PRTAgrobacterium tumefaciens 723His His Ile Ala Ala Gly Leu Gly
Phe Gly Ala Ser Ala Val Tyr Pro1 5 10
15Leu Gly Val Gln Phe Arg Ala Glu Glu Lys Phe Gly Ala Asp
Ala Asp20 25 30Lys Ala Phe Lys Arg Phe
Ala Lys35 4072433PRTAgrobacterium tumefaciens 724His His
Ile Ala Ala Gly Leu Gly Phe Gly Ala Ser Ala Val Tyr Pro1 5
10 15Leu Gly Val Gln Phe Arg Ala Glu
Glu Lys Phe Gly Ala Asp Ala Asp20 25
30Lys72547PRTAgrobacterium tumefaciens 725His Ile Ala Ala Gly Leu Gly Phe
Gly Ala Ser Ala Val Tyr Pro Leu1 5 10
15Gly Val Gln Phe Arg Ala Glu Glu Lys Phe Gly Ala Asp Ala
Asp Lys20 25 30Ala Phe Lys Arg Phe Ala
Lys Ala Ala Glu Lys Ser Leu Met Lys35 40
4572643PRTAgrobacterium tumefaciens 726His Ile Ala Ala Gly Leu Gly Phe
Gly Ala Ser Ala Val Tyr Pro Leu1 5 10
15Gly Val Gln Phe Arg Ala Glu Glu Lys Phe Gly Ala Asp Ala
Asp Lys20 25 30Ala Phe Lys Arg Phe Ala
Lys Ala Ala Glu Lys35 4072739PRTAgrobacterium
tumefaciens 727His Ile Ala Ala Gly Leu Gly Phe Gly Ala Ser Ala Val Tyr
Pro Leu1 5 10 15Gly Val
Gln Phe Arg Ala Glu Glu Lys Phe Gly Ala Asp Ala Asp Lys20
25 30Ala Phe Lys Arg Phe Ala Lys3572832PRTAgrobacterium
tumefaciens 728His Ile Ala Ala Gly Leu Gly Phe Gly Ala Ser Ala Val Tyr
Pro Leu1 5 10 15Gly Val
Gln Phe Arg Ala Glu Glu Lys Phe Gly Ala Asp Ala Asp Lys20
25 3072928PRTAgrobacterium tumefaciens 729Lys Phe Gly
Leu Tyr Asp Ala Ala Phe Glu Lys Ser Ser Cys Gly Val1 5
10 15Gly Phe Ile Thr Arg Lys Asp Gly Val
Gln Thr His20 25
User Contributions:
comments("1"); ?> comment_form("1"); ?>Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
User Contributions:
Comment about this patent or add new information about this topic: