Patent application title: Intermediates and enzymes of the non-mevalonate isoprenoid pathway
Inventors:
Adelbert Bacher (Garching, DE)
Felix Rohdich (Zolling, DE)
Petra Adam (Kramsach, AT)
Sabine Amslinger (Weiden, DE)
Wolfgang Eisenreich (Freising, DE)
Stefan Hecht (Bad Aibling, DE)
IPC8 Class: AC12N910FI
USPC Class:
435 6
Class name: Chemistry: molecular biology and microbiology measuring or testing process involving enzymes or micro-organisms; composition or test strip therefore; processes of forming such composition or test strip involving nucleic acid
Publication date: 2008-12-25
Patent application number: 20080318227
Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
Patent application title: Intermediates and enzymes of the non-mevalonate isoprenoid pathway
Inventors:
Adelbert Bacher
Felix Rohdich
Petra Adam
Sabine Amslinger
Wolfgang Eisenreich
Stefan Hecht
Agents:
MYERS BIGEL SIBLEY & SAJOVEC
Assignees:
Origin: RALEIGH, NC US
IPC8 Class: AC12N910FI
USPC Class:
435 6
Abstract:
The invention provides a protein in a form that is functional for the
enzymatic conversion of 2C-methyl-D-erythritol 2,4-cyclodiphosphate to
1-hydroxy-2-methyl-2-butenyl 4-diphosphate notably in its (E)-form of the
non-mevalonate biosynthetic pathway to isoprenoids. The invention also
provides a protein in a form that is functional for the enzymatic
conversion of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, notably in its
(E)-form, to isopentenyl diphosphate and/or dimethylallyl diphosphate.
Further, screening methods for inhibitors of these proteins are provided.
Further, 1-hydroxy-2-methyl-2-butenyl 4-diphosphate is provided and
chemical and enzymatic methods of its preparation.Claims:
1-103. (canceled)
104. A protein in a form that is functional for the enzymatic conversion of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, in its (E)-form, to isopentenyl diphosphate and/or dimethylallyl diphosphate.
105. The protein according to claim 104, wherein it is in a form functional for said conversion in the presence of FAD and NAD(P)H.
106. The protein according to claim 105, wherein it is in a form functional for said conversion in the presence of a metal ion selected from the group of manganese, iron, cobalt, or nickel ion.
107. The protein according to claim 104, wherein it has a sequence encoded by the ispH (formerly lytB) gene of E. coli or a function-conservative homologue of said sequence.
108. A purified isolated nucleic acid encoding a protein in a form that is functional for the enzymatic conversion of 2C-methyl-D-erythritol 2,4-cyclodiphosphate to 1-hydroxy-2-methyl-2-butenyl 4-diphosphate in its (E)-form and/or a protein in a form that is functional for the enzymatic conversion of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, in its (E)-form, to isopentenyl diphosphate and/or dimethylallyl diphosphate, with or without introns.
109. A DNA expression vector containing the sequence of the nucleic acid according to claim 108.
110. Cells, cell cultures, organisms or parts thereof recombinantly endowed with the sequence of the nucleic acid according to claim 108, wherein said cell is selected from the group consisting of bacterial, protozoal, fungal, plant, insect and mammalian cells.
111. Cells, cell cultures, organisms or parts thereof according to claim 110, wherein it is recombinantly endowed with a vector containing a nucleic acid sequence encoding a protein in a form that is functional for the enzymatic conversion of 2C-methyl-D-erythritol 2,4-cyclodiphosphate to 1-hydroxy-2-methyl-2-butenyl 4-diphosphate in its (E)-form and/or a protein in a form that is functional for the enzymatic conversion of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, in its (E)-form, to isopentenyl diphosphate and/or dimethylallyl diphosphate, and wherein said cell is optionally further endowed with at least one gene selected from the group consisting of dxs, dxr, ispD (formerly ygbP); ispE (formerly ychB); ispF (formerly ygbB) of E. coli, a function-conservative homologue thereof, and a function-conservative fusion, deletion or insertion variant of any of the above genes.
112. Cells, cell cultures, or organisms or parts thereof transformed or transfected for an increased rate of formation of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, in its (E)-form, compared to cells, cell cultures, or organisms or parts thereof absent said transformation or transfection.
113. Cells, cell cultures, or organisms or parts thereof transformed or transfected for an increased rate of conversion of (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate to isopentenyl diphosphate and/or dimethylallyl diphosphate compared to cells, cell cultures, or organisms or parts thereof absent said transformation or transfection.
114. Cells, cell cultures, or organisms or parts thereof according to claim 110 transformed or transfected for an increased expression level of a protein in a form that is functional for the enzymatic conversion of 2C-methyl-D-erythritol 2,4-cyclodiphosphate to 1-hydroxy-2-methyl-2-butenyl 4-diphosphate in its (E)-form and/or a protein in a form that is functional for the enzymatic conversion of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, in its (E)-form, to isopentenyl diphosphate and/or dimethylallyl diphosphate compared to cells, cell cultures, or organisms or parts thereof absent said transformation or transfection.
115. Cells, cell cultures or organisms or parts thereof according to claim 110, characterized by the recombinant endowment with sets of genes selected from the following group:ispC (formerly dxr), ispD, ispE, ispF, ispG (formerly gcpE); orispC, ispD, ispE, ispF, ispG, ispH (formerly lytB); ordxs, ispC, ispD, ispE, ispF, ispG; ordxs, ispC, ispD, ispE, ispF, ispG, ispH; ordxs, ispC, ispG, ordxs, ispC, ispG, ispHof E. coli or a function-conservative homologue thereof and/or a function-conservative fusion, deletion or insertion variant of any of the above genes.
116. Cells, cell cultures or organisms or parts thereof according to claim 115, characterized by further recombinant endowment(s) with gene(s) being functional for biosynthetic steps downstream from the C5 isoprenoids.
117. Cells, cell cultures or organisms or parts thereof according to claim 110, wherein at least one gene of said recombinant endowments is equipped with artificial ribosomal binding site(s) for expression of the corresponding gene product(s) at a rate enhanced compared to the rate in the absence of the artificial ribosomal binding site(s).
118. Cells, cell cultures or organisms or parts thereof according to claim 110, wherein at least one of said recombinant endowments is due to a high copy replication vector.
119. Cells, cell cultures or organisms or parts thereof according to claim 110, wherein they are of bacterial, protozoal, fungal, plant or animal origin.
120. Use of the cells, cell cultures or organisms, or parts thereof according to claim 110 or disruption products thereof for the enhanced rate of in vivo formation or for the efficient in vitro production of a biosynthetic intermediate or product of the non-mevalonate isoprenoid biosynthetic pathway.
121. Use according to claim 120, wherein said intermediate or product is a C5-isoprenoid intermediate compound; or a >C5-isoprenoid compound; or a terpenoid compound.
122. Use according to claim 120, wherein the rate of formation or production is enhanced by providing a source for CTP.
123. Use according to claim 122, wherein the source for CTP is at least one member selected from the group consisting of cytidine, uridine, cytosine, uracil, ribose, ribose 5-phosphate and any biosynthetic precursor of CTP.
124. Use according to claim 120, wherein the rate of formation or production is enhanced by providing a source for phosphorylation enhancement.
125. Use according to claim 124, wherein the source for phosphorylation enhancement is glycerol 3-phosphate, phosphoenolpyruvate, inorganic phosphate, inorganic pyrophosphate or any organic phosphate or pyrophosphate.
126. Use according to claim 120, wherein the rate of formation or production is enhanced by providing a source for reduction equivalents.
127. Use according to claim 126, wherein the source for reduction equivalents is succinate, lipids, glucose, glycerol or lactate.
128. Use of the cells, cell cultures or organisms or parts thereof according to claim 110 for the production of a protein in an enzymatically competent form for the conversion of 2C-methyl-D-erythritol 2,4-cyclodiphosphate into (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate.
129. Use of the cells, cell cultures or organisms or parts thereof according to claim 110 for the production of a protein in an enzymatically competent form for the conversion of (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate into isopentenyl diphosphate and/or dimethylallyl diphosphate.
130. Use of the cells, cell cultures or organisms or parts thereof according to claim 110 for the production of proteins in an enzymatically competent form for the conversion of 2C-methyl-D-erythritol 2,4-cyclodiphosphate into isopentenyl diphosphate and/or dimethylallyl diphosphate.
131. A method of altering the expression level of the gene product(s) of ispG and/or ispH in cells comprising:(a) transforming host cells with the ispG and/or ispH gene; and(b) growing the transformed host cells of step (a) under conditions that are suitable for the efficient expression of ispG and/or ispH, resulting in production of altered levels of the ispG and/or ispH gene product(s) in the transformed cells relative to expression levels of untransformed cells.
132. A process for the efficient in vivo synthesis of (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate; or isopentenyl diphosphate or dimethylallyl diphosphate; in salt form or in protonated form, by the following steps:(a) culturing cells, recombinantly endowed according to claim 110 for said synthesis for a predetermined period of time at a predetermined temperature;(b) optionally adding glucose to a predetermined final concentration and further culturing for a predetermined period of time;(c) harvesting the cells;(d) preparing a crude extract from the harvested cells;(e) separating and purifying optionally isotope-labelled (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate; or isopentenyl diphosphate; or dimethylallyl diphosphate; in salt form or in protonated form.
133. Cells, cell cultures or organisms or parts thereof for the efficient formation of a biosynthetic product or intermediate of the non-mevalonate pathway to isoprenoids or terpenoids, characterized by(a) first recombinant endowment with a gene functional for the biosynthesis of 1-deoxy-D-xylulose 5-phosphate from 1-deoxy-D-xylulose;(b) capability for the uptake of 1-deoxy-D-xylulose; and(c) recombinant endowment(s) with gene(s) being functional for the conversion of 1-deoxy-D-xylulose 5-phosphate into desired downstream C5-intermediate(s) of said pathway.
134. Cells, cell cultures or organisms or parts thereof according to claim 133, wherein said gene(s) of said second recombinant endowment(s) code(s) for enzyme(s) for the formation of at least one of the following C5-intermediates of the non-mevalonate isoprenoid pathway:(a) 2C-methyl-D-erythritol 4-phosphate;(b) 4-diphosphocytidyl-2C-methyl-D-erythritol;(c) 4-diphosphocytidyl-2C-methyl-D-erythritol 2-phosphate;(d) 2C-methyl-D-erythritol 2,4-cyclodiphosphate;(e) 1-hydroxy-2-methyl-2-butenyl 4-diphosphate;(f) isopentenyl diphosphate;(g) dimethylallyl diphosphate.
135. Cells, cell cultures or organisms or parts thereof according to claim 133, characterized by the recombinant endowment with sets of genes as follows:(a) xylB, dxr; or(b) xylB, dxr, ispD (formerly ygbP); or(c) xylB, dxr; ispD, ispE (formerly ychB), or(d) xylB, dxr, ispD, ispE, ispF (formerly ygbB); or(e) xylB, dxr, ispD, ispE, ispF, ispG (formerly gcpE); or(f) xylB, dxr, ispD, ispE, ispF, ispG, ispH (formerly lytB)of E. coli or a function-conservative homologue thereof and/or a function-conservative fusion, deletion or insertion variant of any of the above genes.
136. Cells, cell cultures or organisms or parts thereof according to claim 133, characterized by the recombinant endowment with xylB and ispG (formerly gcpE) and optionally at least one gene selected from the following group: dxr; ispD (formerly ygbP); ispE (formerly ychB); ispF (formerly ygbB) of E. coli or a function-conservative homologue thereof, or a function-conservative fusion, deletion or insertion variant of any of the above genes.
137. Cells, cell cultures or organisms or parts thereof according to claim 133, characterized by the recombinant endowment with xylB and ispH (formerly lytB) and optionally at least one gene selected from the following group: dxr; ispD (formerly ygbP); ispE (formerly ychB); ispF (formerly ygbB); ispG (formerly gcpE) of E. coli or a function-conservative homologue thereof, or a function-conservative fusion, deletion or insertion variant of any of the above genes.
138. Cells, cell cultures or organisms or parts thereof according to claim 133, characterized by the recombinant endowment with xyl, ispG, (formerly gcpE) and ispH (formerly lytB) and optionally at least one gene selected from the following group: dxr, ispD (formerly ygbP); ispE (formerly ychB); ispF (formerly ygbB); of E. coli or a function-conservative homologue thereof, or a function-conservative fusion, deletion or insertion variant of any of the above genes.
139. A process for the efficient in vivo synthesis of 2C-methyl-D-erythritol 4-phosphate; or 4-diphosphocytidyl-2C-methyl-D-erythritol; or 4-diphosphocytidyl-2C-methyl-D-erythritol 2-phosphate; or 2C-methyl-D-erythritol 2,4-cyclodiphosphate; or (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate; or isopentenyl diphosphate or dimethylallyl diphosphate; in salt form or in protonated form, by the following steps:(a) culturing cells, preferably bacterial cells, recombinantly endowed according to claim 133 for said synthesis for a predetermined period of time at a predetermined temperature;(b) adding 1-deoxy-D-xylulose to a predetermined final concentration and further culturing for a predetermined period of time;(c) harvesting the cells;(d) preparing a crude extract from the harvested cells;(e) separating and purifying optionally isotope-labelled 2C-methyl-D-erythritol 4-phosphate; or 4-diphosphocytidyl-2C-methyl-D-erythritol; or 4-diphosphocytidyl-2C-methyl-D-erythritol 2-phosphate; or 2C-methyl-D-erythritol 2,4-cyclodiphosphate; or 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, notably (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate; or isopentenyl diphosphate; or dimethylallyl diphosphate; in salt form or in protonated form, by preparative chromatography.
140. The process according to claim 132, wherein a source for CTP, is added in step (a).
141. The process according to claim 132, wherein a source of phosphorylation activity is added in step (a).
142. The process according to claim 132, wherein a source of reduction equivalents, is added in step (a).
143. A vector comprising a sequence coding for one of the recombinant endowments as defined according to claim 133.
144. The protein according to claim 104, wherein it is a plant protein, notably from Arabidopsis thaliana.
145. The protein according to claim 104, wherein it is a bacterial protein, notably from E. coli.
146. The protein according claim 104, wherein it is a protozoal protein, notably from Plasmodium falciparum.
147. Cells, cell cultures or organisms or parts thereof according to claim 111, characterized by the recombinant endowment with sets of genes selected from the following group:ispC (formerly dxr), ispD, ispE, ispF, ispG (formerly gcpE); orispC, ispD, ispE, ispF, ispG, ispH (formerly lytB); ordxs, ispC, ispD, ispE, ispF, ispG; ordxs, ispC, ispD, ispE, ispF, ispG, ispH; ordxs, ispC, ispG; ordxs, ispC, ispG, ispHof E. coli or a function-conservative homologue thereof and/or a function-conservative fusion, deletion or insertion variant of any of the above genes.
148. A process for the efficient in vivo synthesis of (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate; or isopentenyl diphosphate or dimethylallyl diphosphate, in salt form or in protonated form, comprising the following steps:(a) culturing cells, recombinantly endowed according to claim 112 for said synthesis for a predetermined period of time at a predetermined temperature;(b) optionally adding glucose to a predetermined final concentration and further culturing for a predetermined period of time;(c) harvesting the cells;(d) preparing a crude extract from the harvested cells; and(e) separating and purifying optionally isotope-labelled (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate; or isopentenyl diphosphate; or dimethylallyl diphosphate, in salt form or in protonated form.
149. A process for the efficient in vivo synthesis of (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate; or isopentenyl diphosphate or dimethylallyl diphosphate, in salt form or in protonated form, comprising the following steps:(a) culturing cells, recombinantly endowed according to claim 113 for said synthesis for a predetermined period of time at a predetermined temperature;(b) optionally adding glucose to a predetermined final concentration and further culturing for a predetermined period of time;(c) harvesting the cells;(d) preparing a crude extract from the harvested cells; and(e) separating and purifying optionally isotope-labelled (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate; or isopentenyl diphosphate; or dimethylallyl diphosphate, in salt form or in protonated form.
150. The process according to claim 139, wherein a source for CTP is added in step (a).
151. The process according to claim 139, wherein a source of phosphorylation activity is added in step (a).
152. The process according to claim 139, wherein a source of reduction equivalents is added in step (a).
Description:
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001]This application is a divisional application of U.S. application Ser. No. 10/474,536, having a filing date of Apr. 7, 2004, which is a 35 U.S.C. § 371 national phase application of International PCT Application Serial No. PCT/EP02/04005 filed Apr. 10, 2002, which claims priority to German Patent Application No. 101 18 166.3, filed Apr. 11, 2001, German Patent Application No. 101 30 236.3, filed Jun. 22, 2001, German Patent Application No. 101 55 084.7, filed Nov. 9, 2001, and German Patent Application No. 102 01 458.2, filed Jan. 16, 2002. The contents of these applications are hereby incorporated by reference as if recited in full herein.
FIELD OF THE INVENTION
[0002]The present invention relates to cells, cell cultures or organisms or parts thereof for the efficient formation of a biosynthetic product or intermediate or enzyme of a 1-deoxy-D-xylulose 5-phosphate-dependent biosynthetic pathway. Further, the invention relates to vectors for producing them. Further, the invention relates to their use for the formation or production of intermediates or products or enzymes of said biosynthetic pathway as well as to enzymes and intermediates. Further, the invention relates to the screening for inhibitors or enzymes for said biosynthetic pathway.
BACKGROUND OF THE INVENTION
[0003]The system of biosynthetic pathways in any organism is highly streamlined, whereby a few central trunk pathways branch into a great number of peripheral pathways. The central trunk pathways involve starting materials which are highly integrated. Therefore, central or trunk pathways are highly regulated. At the same time they are crucial for any attempts to interfere with the metabolism of any organism either by an inhibitor or by metabolic engineering.
[0004]The isoprenoid pathways are a prime example for this metabolic organisation. They are very long and highly branched, leading to some 30,000 isoprenoid or terpenoid compounds. They all seem to derive from isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP). They are produced by two alternative trunk pathways (reviewed in Eisenreich et al., 2001).
[0005]By the classical research of Bloch, Cornforth, Lynen and co-workers, isopentenyl pyrophosphate (IPP) and dimethylallyl pyrophosphate (DMAPP) have become established as key intermediates in the biosynthesis of isoprenoids via mevalonate. However, many bacteria, plastids of all plants, and the protozoon Plasmodium falciparum synthesize IPP and DMAPP by an alternative pathway via 1-deoxy-D-xylulose 5-phosphate. The discovery of the pathway was mainly based on the incorporation of isotope-labelled 1-deoxy-D-xylulose into the isoprenoid side chain of menaquinones from Escherichia coli (Arigoni and Schwarz, 1999).
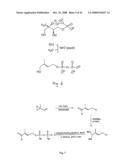
[0006]This mevalonate-independent pathway has so far only been partially explored (FIG. 1). For a better understanding of these aspects of the invention, the pathway shall be briefly explained. It can be divided into three segments:
[0007]In a first pathway segment shown in FIG. 1, pyruvate (1) is condensed with glyceraldehyde 3-phosphate (2) to 1-deoxy-D-xylulose 5-phosphate (DXP) (3). Subsequently, DXP is converted into 2C-methyl-D-erythritol 4-phosphate (MEP) (4) by a two-step reaction comprising a rearrangement and a reduction. This establishes the 5-carbon isoprenoid skeleton.
[0008]In the subsequent segment of the mevalonate-independent pathway (FIG. 1), MEP (4) is first condensed with CTP to 4-diphosphocytidyl-2C-methyl-D-erythritol (CDP-ME) (5) by 4-diphosphocytidyl-2C-methyl-D-erythritol synthase (PCT/EP00/07548). CDP-ME (5) is subsequently ATP-dependent phosphorylated by 4-diphosphocytidyl-2C-methyl-D-erythritol kinase yielding 4-diphosphocytidyl-2C-methyl-D-erythritol 2-phosphate (CDP-MEP) (6). The intermediate is subsequently converted into 2C-methyl-D-erythritol 2,4-cyclodiphosphate (cMEPP) (7) by 2C-methyl-D-erythritol 2,4-cyclodiphosphate synthase (PCT/EP00/07548). These three enzymatic steps form a biosynthetic unit which activates the isoprenoid C5-skeleton for the third pathway segment (Rohdich et al., 1999; Luttgen et al., 2000; Herz et al., 2000).
[0009]Bioinformatic studies (German Patent Application 10027821.3), as well as studies with mutants of Synechocystis sp. (Cunningham et al., 2000) and Escherichia coli (Campos et al., 2001; Altincicek et al., 2001) demonstrate the involvement of lytB and gcpE genes in the isoprenoid pathway. However, the function and the reaction catalyzed by the corresponding gene products are still unknown.
[0010]Recently, a kinase (XylB) has been described that catalyzes the conversion of 1-deoxy-D-xylulose into 1-deoxy-D-xylulose 5-phosphate at high rates (Wungsintaweekul et al., 2000).
[0011]Genes and enzymes participating in further downstream reactions have been described. However, the gene functions, the intermediates, and the mechanisms leading to the products are still unknown.
[0012]For numerous pathogenic eubacteria as well as for the malaria parasite P. falciparum, the enzymes involved in the non-mevalonate pathway are essential. The intermediates of the mevalonate-independent pathway cannot be assimilated from the environment by pathogenic eubacteria and P. falciparum. The enzymes of the alternative isoprenoid pathway do not occur in mammalia which synthesize their isoprenoids and terpenoids exclusively via the mevalonate pathway. Moreover, the idiosyncratic nature of the reactions in this pathway reduces the risk of cross-inhibitions with other, notably mammalian enzymes.
[0013]Therefore, enzymes of the alternative isoprenoid pathway seem to be specially suited as targets for novel agents against pathogenic microorganisms and herbicides. The elucidation of unknown steps and the identification of these targets, e.g. genes and cognate enzymes of these pathways is obligatory for this purpose. A further source of interest in the non-mevalonate pathway derives from the fact certain pathogens like Mycobacteria, Plasmodia, Escherichia etc. use this pathway to activate γδ T cells (Fournie and Bonneville, 1996). Therefore, γδ T cells likely act as a first line of defense against infections by such pathogens. Intermediates of the non-mevalonate pathway have been suggested to be responsible for γδ T cell activation (Jomaa et al., 1999). Recently, it was show that E. coli strains lost the ability to stimulate γδ T cells when the dxr or the gcpE gene was knocked out (Altincicek et al., 2001).
[0014]Moreover, there is a great biotechnological interest in these pathways, since they lead to valuable vitamins and isoprenoid or terpenoid products.
[0015]Previous attempts to approach these goals have been hampered by the low rate of biosynthesis along these pathways in wild-type cells studied so far.
SUMMARY OF THE INVENTION
[0016]It is an object of the invention to provide enzymes and nucleic acids coding for said enzymes as well as intermediates for the conversion of 2C-methyl-D-erythritol 2,4-cyclodiphosphate to isopentenyl diphosphate and/or dimethylallyl diphosphate.
[0017]It has surprisingly been found that the intermediate in the conversion of 2C-methyl-D-erythritol 2,4-cyclodiphosphate to isopentenyl diphosphate and/or dimethylallyl diphosphate is 1-hydroxy-2-methyl-2-butenyl 4-diphosphate. This intermediate is formed by an enzyme encoded by gcpE as designated in the E. coli genome. It has further been found that this enzyme prefers as reductant NADH or NADPH. Further, it has been found that it is promoted by Co2+.
[0018]The above intermediate is converted to isopentenyl diphosphate and/or dimethylallyl diphosphate by an enzyme encoded by lytB as designated in the E. coli genome. The latter enzyme prefers as reductant NADH or NADPH and FAD as mediator. Further it can be promoted by ions of a metal selected from manganese, iron, cobalt, nickel.
[0019]With these findings, the third segment of the trunk non-mevalonate pathway is now established. The key to these findings is the intermediate 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, notably in its E-form. This establishes the unifying principle of the invention for reactions to and from this intermediate.
[0020]Further, it is an object of the invention to provide cells, cell cultures, organisms or parts thereof for the efficient biosynthesis of isoprenoid products or intermediates of the non-mevalonate biosynthetic pathway dependent on 1-deoxy-D-xylulose 5-phosphate production from 1-deoxy-D-xylulose and/or glucose.
[0021]The present invention produces a novel in vivo system which can be used for the structure elucidation of unknown intermediates and the assignment of biological functions of putative genes or cognate enzymes in the alternative isoprenoid biosynthetic pathway. As an example, the functional assignment of the gcpE gene (now designated as ispG) and of the lytB gene (now designated ispH) in the mevalonate-independent pathway of isoprenoid biosynthesis is achieved.
[0022]More specifically, said in vivo system consists of recombinant E. coli strains harbouring vector construct(s) carrying and expressing genes for D-xylulokinase (xylB), and genes of further downstream steps of terpenoid biosynthesis, such as dxs, dxr, and/or ispD, and/or ispE, and/or ispF, and/or gcpE, and/or lytB from E. coli, and/or a carotenoid gene cluster from Erwinia uredovora.
[0023]In one aspect of the invention, the genetically modified strains can be fed with 1-deoxy-D-xylulose, notably with isotope-labelled 1-deoxy-D-xylulose, which is converted at high rates into the common intermediate of the mevalonate-independent terpenoid pathway, 1-deoxy-D-xylulose 5-phosphate, and into further intermediates of said pathway, like 2C-methyl-D-erythritol 4-phosphate, 4-diphosphocytidyl-2C-methyl-D-erythritol, 4-diphosphocytidyl-2C-methyl-D-erythritol 2-phosphate, 2C-methyl-D-erythritol 2,4-cyclodiphosphate, 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, isopentenyl diphosphate, and dimethylallyl diphosphate. Further, feeding with glucose or an intermediate of glycolysis for conversion into said further intermediates of said pathway may be performed.
[0024]Said systems are useful for the structure elucidation of hitherto elusive intermediates in the biosynthetic pathways, for in vivo screening of novel antibiotics, antimalarials, and herbicides, and as a platform for the bioconversion of exogenous 1-deoxy-D-xylulose and/or glucose into intermediates and products of the non-mevalonate pathway of terpenoid biosynthesis.
[0025]Said systems can also be used for screening chemical libraries for potential herbicides, and/or antimalarials, and/or antimicrobial substances by detecting and measuring the amount of certain intermediates formed in vivo in the presence or absence of potential inhibitors of the gene products of mevalonate-independent isoprenoid pathway genes, namely dxs, dxr, ispD, ispE, ispF, gcpE, and lytB.
[0026]Said system can further be used for the production of higher isoprenoids (e.g. isoprenoids having 10, 15, 20, 30 or 40 carbon atoms) such as carotene, α-tocopherol or vitamins by boosting the biosynthesis of isopentenyl diphosphate and/or dimethylallyl diphosphate via the non-mevalonate pathway, e.g. by using glucose as feeding material. Further feeding materials which may be used are intermediates or products of glycolysis like glyceraldehyde 3-phosphate or pyruvate.
[0027]Further, this invention provides novel compounds of formula I (see below), notably 1-hydroxy-2-methyl-2-butenyl 4-diphosphate as well as enzymatic and chemical methods for preparing said compounds. As demonstrated herein, (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate is produced from 2C-methyl-D-erythritol 2,4-cyclodiphosphate by the gcpE gene product.
[0028]It is further demonstrated herein that (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate is converted to dimethylallyl diphosphate and isopentenyl diphosphate by the lytB gene product.
BRIEF DESCRIPTION OF THE DRAWINGS
[0029]FIG. 1: Biosynthesis of both isoprenoid precursors, isopentenyl pyrophosphate and dimethylallyl pyrophosphate via the mevalonate-independent pathway.
[0030]FIG. 2: Scheme of an Escherichia coli in vivo system for generating optionally isotopically labelled intermediates of biosynthetic pathways such as the mevalonate-independent isoprenoid biosynthesis, and for the production of higher terpenoids such as carotenoids.
[0031]FIG. 3: 1H NMR spectra in D2O (pH 6) obtained according to Example 25.* indicates impurities.
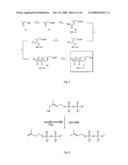
[0032]FIG. 4: Preparation of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate according to Example 24. Reagents and conditions were as follows: (a) DHP, PPTS, 25° C. (2.5 h); (b) Ph3PCHCO2Et, toluene, reflux (39 h); (c) (1) DIBAH, CH2Cl2, -78° C. (3 h), (2) 1 M NaOH/H2O; (d) p-TsCl, DMAP, CH2Cl2, 25° C. (1 h); (e) ((CH3CH2CH2CH2)4N)3HP2O7, MeCN, 25° C. (2 h); (f), HCl/H2O pH 1, 25° C. (7 min).
[0033]FIG. 5: The reaction catalyzed by the ispH (formerly lytB) gene product.
[0034]FIG. 6: The reaction catalyzed by the ispG (formerly gcpE) gene product.
[0035]FIG. 7: Chemical preparation of 3-formyl-but-2-enyl 1-diphosphate (see example 42).
[0036]FIG. 8: DNA sequence of the vector construct pBSxylBdxr (SEQ ID NO:41).
[0037]FIG. 9: DNA sequence of the vector construct pBSxylBdxrispD (SEQ ID NO:42).
[0038]FIG. 10: DNA sequence of the vector construct pBScyclo (SEQ ID NO:43).
[0039]FIG. 11: DNA sequence of the vector construct pACYCgcpE (SEQ ID NO:44).
[0040]FIG. 12: DNA sequence of the vector construct pBScaro14 (SEQ ID NO:45).
[0041]FIG. 13: DNA sequence of the vector construct pACYCcaro14 (SEQ ID NO:46).
[0042]FIG. 14: DNA sequence (SEQ ID NO:47) and corresponding amino acid sequence (SEQ ID NO:48) of the ispG (formerly gcpE) gene from Escherichia coli.
[0043]FIG. 15: DNA sequence of the vector construct pBScyclogcpE (SEQ ID NO:49).
[0044]FIG. 16: DNA sequence of the vector construct pACYClytBgcpE (SEQ ID NO:50).
[0045]FIG. 17: DNA (SEQ ID NO:51) and corresponding amino acid sequence (SEQ ID NO:52) of the ispH (formerly lytB) gene from Escherichia coli.
[0046]FIG. 18: DNA sequence of the vector construct pBScyclogcpElytB2 (SEQ ID NO:53).
[0047]FIG. 19: DNA (SEQ ID NO:54) and corresponding amino acid sequence (SEQ ID NO:55) of the ispG gene (fragment) from Arabidopsis thaliana.
[0048]FIG. 20: DNA (SEQ ID NO:56) and corresponding amino acid sequence (SEQ ID NO:57) of the ispG (formerly gcpE) gene of Arabidopsis thaliana.
[0049]FIG. 21: cDNA sequence of 1-hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate reductase (IspH) from Arabidopsis thaliana (SEQ ID NO:58)
DETAILED DESCRIPTION OF THE INVENTION
[0050]1-Deoxy-D-xylulose 5-phosphate is a common intermediate in the alternative terpenoid pathway via 2C-methyl-D-erythritol 4-phosphate. This latter pathway is operative in bacteria, certain protozoa and most significantly also in the plastids of plants, where it is in charge of the biosynthesis of a great many valuable terpenoid products, like natural rubber, carotenoids, menthol, menthone, camphor or paclitaxel. The alternative terpenoid pathway is now intensely studied. But so far only the initial steps from glyceraldehyde 3-phosphate and pyruvate via 1-deoxy-D-xylulose 5-phosphate and 2C-methyl-D-erythritol 4-phosphate, 4-diphosphocytidyl-2C-methyl-D-erythritol, 4-diphosphocytidyl-2C-methyl-D-erythritol 2-phosphate and 2C-methyl-D-erythritol 2,4-cyclodiphosphate (FIG. 1) have been elucidated. The intermediate 1-deoxy-D-xylulose 5-phosphate is of most crucial significance for a number of commercial purposes: [0051](1) It may be used as a key intermediate for commercial screening procedures regarding potential inhibitors of downstream enzymes in the biosynthesis of the alternative terpenoid pathway. [0052](2) It may be used as a key intermediate for the in vitro production of terpenoids or of intermediates thereof. [0053](3) It occurs in vivo in the biosynthesis of terpenoids as an enzymatic condensation product of glyceraldehyde 3-phosphate and pyruvate. The latter are central intermediates of the metabolism and obligatory starting materials for numerous biosynthetic pathways. Therefore, it is desirable to generate a high level of 1-deoxy-D-xylulose 5-phosphate in vivo from an exogenous source and thus independent from the pools of glyceraldehyde 3-phosphate and pyruvate for boosting the biosynthesis of terpenoids or of intermediates thereof in microorganisms or cell cultures that are either naturally or recombinantly endowed with the pathway of interest without influencing the basic intermediary metabolism of the cells. [0054](4) 1-Deoxy-D-xylulose 5-phosphate can be generated from 1-deoxy-D-xylulose by the catalytic action of the xylB gene product. Using recombinant strains comprising the xylB gene the reaction occurs in vivo and exogenous 1-deoxy-D-xylulose is converted into intracellular 1-deoxy-D-xylulose 5-phosphate at high rates. [0055](5) 1-DXP can be generated fro glucose by the catalytic action of glycolytic enzymes and DXP-synthase. Using recombinant strains comprising the dxs gene, the reaction occurs in vivo and exogeneous glucose is converted to intracellular 1-DXP at high rates.
[0056]It is an aspect of the invention to use 1-deoxy-D-xylulose as a precursor in order to boost the rates of biosynthesis of 1-deoxy-D-xylulose 5-phosphate-dependent pathways. 1-Deoxy-D-xylulose can be prepared by various published procedures (Blagg and Poulter, 1999; Kennedy et al., 1995; Piel and Boland, 1997; Shono et al., 1983; Giner, 1998).
[0057]It is an aspect of the present invention to use 1-deoxy-D-xylulose in various isotopically labelled forms. It may be labelled by radioactive isotopes or non-radioactive isotopes of C (13C or 14C), H (D or T) or 0 (170 or 180) in any combination.
[0058]Isotope-labelled 1-deoxy-D-xylulose may be prepared enzymatically using 1-deoxy-D-xylulose 5-phosphate synthase of Bacillus subtilis and commercially available glycolytic enzymes and phosphatase from isotope-labelled glucose and/or pyruvate (PCT/EP00/07548).
[0059]1-Deoxy-D-xylulose may be used as a free acid or as a salt, preferably as an alkaline (e.g., lithium, sodium, potassium) salt or as an ammonium or amine salt.
[0060]It is an aspect of the present invention to use recombinant cells, cell cultures, or organisms or parts thereof for the formation of biosynthetic products or intermediates or enzymes or for the screening for antimicrobials, antimalarials or herbicides.
[0061]For carrying out the present invention various techniques in molecular biology, microbiology and recombinant DNA technology are used which are comprehensively described in Sambrock et al., Molecular Cloning, second edition, Cold Spring Harbor Laboratory Press, Cold Sprind Harbor, N.Y.; in DNA Cloning: A Practical Approach, Vol. 1 and 2, 1985 (D. N. Glover, ed.); in Oligonucleotide Synthesis, 1984 (M. L. Gait, ed.); and in Transcription and Translation (Hames and Higgins, eds.).
[0062]Nucleic Acids
[0063]The present invention comprises nucleic acids which include prokaryotic, protozoal and plant sequences and derived sequences. A derived sequence relates to a nucleic acid sequence corresponding to a region of the sequence or orthologs thereof or complementary to "sequence-conservative" or "function-conservative" variants thereof.
[0064]Sequences may be isolated by well known techniques or are commercially available (Clontech, Palto Alto, Calif.; Stratagene, LaJolla, Calif.). Alternatively, PCR-based methods can be used for amplifying related sequence from cDNA or genomic DNA. The nucleic acids of the present invention comprise purine and pyrimidine containing polymers in various amounts, either polyribonucleotides or polydeoxyribonucleotides or mixed polyribo-polydeoxyribonucleotides. The nucleic acids may be isolated directly from cells. Alternatively, PCR may be used for the preparation of the nucleic acids by use of chemical synthesized strands or by genomic material as template. The primers used in PCR may be synthesized by using the sequence information provided by the present invention or from the database and additionally may be constructed with optionally new restriction sites in order to ease the cloning in a vector for recombinant expression.
[0065]The nucleic acids or the present invention may be flanked by natural regulation sequences or may be associated with heterologous sequences, including promoter, enhancer, response elements, signal sequences, polyadenylation sequences, introns, 5'- and 3' noncoding regions or similar. The nucleic acids may be modified on basis of well known methods. Non-limiting examples for these modifications are methylations, "Caps", substitution of one or more natural nucleotides with an analogue, and internucleotide modification, i.e. those with uncharged bond (i.e. methylphosphonates, phosphotriester, phosphoramidates, carbamates, etc.) and with charged bond (i.e. phosphorothiactes, etc.). Nucleic acids may carry additional covalent bound units such as proteins (i.e. nucleases, toxins, antibodies, signalpeptides, poly-L-lysine, etc.), intercalators (i.e. acridine, psoralene, etc.), chelators (i.e. metals, radioactive metals, iron, oxidative metals, etc.) and alkylators. The nucleic acids may be derived by formation of a methyl- or ethylphosphotriester bond or of a alkylphosphoramidate bond. Further, the nucleic acids of the present invention may be modified my labeling, which give an either directly or indirectly detectable signal. Examples for these labeling include radioisotopes, fluorescent molecules, biotin and so on.
[0066]Vectors
[0067]The invention provides nucleic acid vectors, which comprise the sequences provided by the present invention or derivatives thereof. Various vectors, including plasmids or vectors for fungi have been described for the replication and/or expression in various eucaryotic and procaryotic hosts. High copy replication vectors are preferred for the purposes of the invention. Non-limiting examples include pKK plasmids (Clontech), pUC plasmids (Invitrogen, San Diego, Calif.), pET plasmids (Novagen, Inc., Madison, Wis.) or pRSET or pREP (Invitrogen) and various suitable host cells on basis of well known techniques. Recombinant cloning vectors comprise often more than one replication system for the cloning and expression, one or more marker for the selection in the host; i.e. antibiotic resistance and one or more expression cartridge. Suitable hosts may be transformed/transfected/infected by a method as suitable including electroporation, CaCl2-mediated DNA incorporation, tungae infection, microinjection, microbombardment or other established methods.
[0068]Suitable hosts include bacteria, archaebacteriae, fungi, notable yeast, plants, notably Arabidopsis thaliana, Mentha piperita or Taxus sp. and animal cells, notably mammalian cells. Most important are E. coli, Bacillus subtilis, Saccharomyces cerevisiae, Saccharomyces carlsbergensis, Schizosaccharomyces pombe, SF9 cells, C129 cells, 293 cells, Neurospora, and CHO cells, COS cells, HeLa cells and immortalized myeloid and lymphoid mammalian cells. Preferred replication systems include M13, ColE1, SV40, baculovirus, lambda, adenovirus and so on. A great number of transcription, initiation (including ribosomal binding sites) and termination regulation regions have been isolated and there efficiency for the transcription and translation of heterologous proteins has been demonstrated in various hosts. Examples for these regions, methods for the isolation, the way for using are well known. Under suitable conditions for expression host cells may be used as source for the recombinant synthesized proteins.
[0069]Expression Systems
[0070]Preferable vectors may include a transcription element (that is a promoter), functionally connected with the enzyme domain. Optionally, the promoter may include parts of operator region and/or ribosomal binding sites. Non-limiting examples for bacterial promoters, which are compatible with E. coli, include: trc promoter, b-lactamase (penicillinase) promoter; lactose promoter, tryptophan (trp) promoter, arabinose BAD operon-promoter, lambda-derived P1 promoter and N gene ribosomal binding site and the hybrid Tac promoter, derived from sequences of trp and lac UV5 promoters. Non-limiting examples for yeast promoters include 3-phosphoglycerate kinase promoter, glyceraldehyde 3-phosphate dehydrogenase (GAPDH) promoter, galactokinase (GALI) promoter, galactoepimerase promoter and alcoholdehydrogenase (ADH) promoter. Suitable promoters for mammalian cells include without limiting viral promoters such as i.e. simian virus 40 (SV40), rous sarcoma virus (RSV), adenovirus (ADV) and bovine papilloma virus (BPV). Mammalian cells may also need terminator sequences and poly-A sequences and enhancer sequences, which may increase the expression. Sequences, which amplify the genes, may also be preferred. Further on, sequences may be included, which ease the secretion of the recombinant protein from the cell, which may be but non-limiting a bacterial, yeast or animal cell, such as i.e. a secretion signal sequence and/or prehormon sequence.
[0071]It is an important aspect of the invention that the combined recombinant endowment with xylB and other gene(s) of the alternative C5-isoprenoid pathway and optionally gene(s) for higher isoprenoids or terpenoids boost(s) these pathways. Preferably, xylB is combined with complete sets of genes to convert 1-deoxy-D-xylulose 5-phosphate into the desired intermediate or end products. For intermediates in the C5-isoprenoid pathway, cells are preferably endowed with one of the combinations of genes given in claim 76.
[0072]For the genes cited herein, the common E. coli designation were used. Other genes from E. coli or from other organisms (orthologous genes) may also be used if they have the same functions (function-conservative genes), notably if their gene products catalyze the same reaction. Further, deletion or insertion variants or fusions of these genes with other genes or nucleic acids may be used, as long as these variants are function-conservative. The above genes may be derived from bacteria, protozoa, or from higher or lower plants.
[0073]It is another important aspect of the invention that the function of gcpE as following immediately downstream from ispF has been determined. Our findings show that the gcpE gene product is involved in the formation of the novel compound 1-hydroxy-2-methyl-2-butenyl 4-diphosphate from 2C-methyl-D-erythritol 2,4-cyclodiphosphate. Therefore, we rename gcpE in ispG.
[0074]In a further aspect of the invention it was shown that the gene product of gcpE is involved in the formation of the E-isomer of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate from 2C-methyl-D-erythritol 2,4-cyclodiphosphate by comparison with chemically synthesized (E)- and (Z)-isomers of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate. Therefore, this invention further pertains to the (E) and (Z) isomers of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate salts or protonated forms thereof.
[0075]It is another important aspect of the invention that the function of lytB has been determined as following immediately downstream from ispG. Therefore, it is renamed ispH. It is our finding that ispH is involved in the conversion of (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate into isopentenyl 4-diphosphate and/or dimethylallyl 4-diphosphate.
[0076]It should be understood that "1-hydroxy-2-methyl-2-butenyl 4-phosphate" and "1-hydroxy-2-methyl-2-butenyl 4-diphosphate" comprise the free phosphoric and diphosphoric acids, respectively, and the singly or multiply deprotonated forms thereof, i.e. salts which may be salts of any cation (including Na, K, NH4.sup.+, Li, Mg, Ca, Zn, Mn, and Co cations). The protonation state of (di)phosphates and phosphate derivatives or their conjugated acids in aqueous solution depends on the pH value of the solution, as is known to persons skilled in the art. The same applies to other phosphates or phosphate derivatives.
[0077]In another aspect of the invention, (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate has been successfully incorporated into the lipid soluble fraction of Capsicum annuum chromoplasts. A 14C label of this compound was incorporated into the geranylgeraniol, b-carotene, phytoene and phytofluene fractions of C. annuum chromoplasts establishing (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate as intermediate of the non-mevalonate pathway downstream from 2C-methyl-D-erythritol 2,4-cyclodiphosphate and upstream from isopentenyl diphosphate.
[0078]It is another aspect of the invention that xylB can be combined with gcpE and optionally other genes of the alternative C5 isoprenoid pathway and/or of the higher isoprenoid pathways in vector(s) for recombinant engineering.
[0079]As a consequence of our findings regarding gcpE (now ispG) it follows that the gene lytB operates downstream of gcpE and thus in service of the conversion of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate to IPP and/or DMAPP. Therefore, it is another aspect of the invention to combine the gene lytB with xylB and optionally other genes of the common C5-isoprenoid pathway or of a higher isoprenoid pathway.
[0080]Our finding allows the efficient formation or production of intermediates or products of the isoprenoid pathway with any desired labelling, notably the following intermediates:
[0081]2C-methyl-D-erythritol 4-phosphate; 4-diphosphocytidyl-2C-methyl-D-erythritol; 4-diphosphocytidyl-2C-methyl-D-erythritol 2-phosphate; 2C-methyl-D-erythritol 2,4-cyclodiphosphate; 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, isopentenyl diphosphate; dimethylallyl diphosphate.
[0082]The formation of end products of the terpenoid pathway (e.g., b-carotene, zeaxanthine, paclitaxel, menthol, menthone, cannabinoids), may be boosted following the process of the invention.
[0083]The strains harbouring the recombinant plasmids can be cultivated in conventional culture media, preferably in terrific broth medium, at 15 to 40° C. The preferred temperature is 37° C. The E. coli strains are induced with 0.5 to 2 mM isopropyl-b-D-thiogalactoside (IPTG) at an optical density at 600 nm from 0.5 to 5. The cells are incubated after addition of 1-deoxy-D-xylulose at a concentration of 0.001 mM to 1 M preferably at a concentration of 0.01 to 30 mM for 30 min to 15 h, preferably 1 to 5 h. It has been found that the process of producing isoprenoid intermediates or products by the genetically engineered organisms of the invention can be boosted by supplying a source for CTP, for example cytidine and/or uridine and/or cytosine and/or uracil and/or ribose and/or ribose 5-phosphate and/or any biosynthetic precursors of CTP at a concentration of 0.01 to 10 mM, preferably at a concentration of 0.3 to 1 mM, and/or by supplying a source for phosphorylation activity, for example glycerol 3-phosphate and/or phosphoenolpyruvate and/or ribose 5-phosphate at a concentration of 0.1 to 100 mM, preferably at a concentration of 0.5 to 10 mM and/or inorganic phosphate and/or inorganic pyrophosphate at a concentration of 1 to 500 mM, preferably at a concentration of 10 to 100 mM and/or any organic phosphate and/or pyrophosphate, and/or by supplying a source for reduction equivalents, for example 0.1 to 1000 mM, preferably 10 to 1000 mM, lactate and/or succinate and/or glycerol and/or glucose and/or lipids at a concentration of 0.1 to 100 mM, preferably at a concentration of 0.5 to 10 mM. A particularly efficient production process is specified in claims 72 and 80 to 84.
[0084]This process can also be used with great advantages for screening for inhibitors of the enzymes involved or of downstream enzymes, dependent on the choice of the isoprenoid intermediate or product for detection. The enzymes dxs, dxr, ispD, ispE, ispF, ispG (formerly gcpE) and ispH (formerly lytB) do not occur in animals. Therefore inhibitors against dxs, dxr, ispD, ispE, ispF, ispG (formerly gcpE) and ispH (formerly lytB) have great value as (a) herbicides against weed plants or algae; (b) antibiotic agents against pathogenic bacteria; (c) agents against protozoa, like Plasmodium falciparum, the causative pathogen of malaria.
[0085]The activity of the said enzymes can be detected (in the presence or absence of a potential inhibitor) by measuring either the formation of a product or the consumption of an intermediate, preferably by TLC, HPLC or NMR.
[0086]With the finding that 1-hydroxy-2-methyl-2-butenyl 4-diphosphate is an intermediate of the non-mevalonate terpenoid pathway we have acquired essential determinants of the structure of inhibitors. Namely, the structures of a subset of inhibitors should be similar to at least a portion of the starting compound or the product or the transition state between the starting compound e.g. 2C-methyl-D-erythritol 2,4-cyclodiphosphate and the product e.g. 1-hydroxy-2-methyl-2-butenyl 4-diphosphate.
[0087]This invention discloses novel compounds, or salts thereof, of the following formula I:
whereby R1 and R2 are different from each other and one of R1 and R2 is hydrogen and the other is selected from the group consisting of --CH2--O--PO(OH)--O--PO(OH)2, --CH2--O--PO(OH)2, and --CH2OH, and whereby A stands for --CH2OH or --CHO. These compounds may be isotope-labelled.
[0088]In formula I, A preferably stands for --CH2OH.
[0089]Among R1 and R2, R1 is preferably hydrogen and R2 is preferably selected from the group consisting of --CH2--O--PO(OH)--O--PO(OH)2 and --CH2--O--PO(OH)2.
[0090]In the group consisting of --CH2--O--PO(OH)--O--PO(OH)2 and --CH2--O--PO(OH)2, --CH2--O--PO(OH)--O--PO(OH)2 is preferred.
[0091]If a compound of formula I is a salt, it may e.g. be a lithium, sodium, potassium, magnesium, ammonium, manganese salt. These salts may derive from a single or from multiple deprotonations from the (di)phosphoric acid moiety.
[0092]The novel compounds disclosed herein are useful for various applications e.g. for screening for genes, enzymes or inhibitors of the biosynthesis of isoprenoids or terpenoids, either in vitro in the presence of an electron donor or in vivo.
[0093]This invention further provides a process for the chemical preparation of a compound of formula I or a salt thereof:
wherein A represents --CH2OH and R1 and R2 are different from each other and one of R1 and R2 is hydrogen and the other is --CH2--O--PO(OH)--O--PO(OH)2, --CH2--O--PO(OH)2 or --CH2--OH by the following steps: [0094](a) converting a compound of the following formula (II):
[0094] [0095]wherein B is a protective group into a compound of the following formula (III) or (IV):
[0095] [0096]by a Wittig or Horner reagent, wherein the group D is a precursor group convertible reductively to a --CH2--OH group; [0097](b) reductively converting group D to a --CH2--OH group; [0098](c) optionally converting group --CH2--OH obtained in step (b) into --CH2--O--PO(OH)--O--PO(OH)2 or --CH2--O--PO(OH)2 or salts thereof in a manner known per se; [0099](d) optionally conversion to a desired salt; [0100](e) removing the protective group B.
[0101]In the above process, said protective group B may be any group that allows to regenerate an hydroxy group at the position it is attached to. Said protective group B is preferably stable under the conditions of step (a) to step (d). Said protective group B is removed in step (e) of said process in order to generate a hydroxy group. Protective groups for hydroxy groups are known to the skilled person. Group B may for example form an acetal group together with the remaining moiety of the compound of formula (II), (III) or (IV). Acetals can be hydrolysed under acidic conditions. Most preferably, group B is a 2-tetrahydropyranyl group.
[0102]In the above process, said group D is a precursor group convertible reductively to a --CH2--OH group. Group D may be a derivative of a carbon acid. Examples of such a group include alkoxycarbonyl and aminocarbonyl groups. Said aminocarbonyl groups may be substituted at the amino group with one or two alkyl groups. It is most preferred to use alkoxycarbonyl groups. The alkyl group of said alkoxycarbonyl groups or said alkyl groups of said aminocarbonyl groups may be a linear or branched alkyl groups which may be singly or multiply substituted. Preferred are C1-C6 alkyl groups like methyl, ethyl, propyl, butyl, pentyl or hexyl groups. Most preferred are methyl or ethyl groups. The most preferred example of said group D is an ethoxycarbonyl group.
[0103]Said compound of formula (II) may be prepared by protecting the hydroxy group of hydroxy acetone with said group B. If group B is a tetrahydropyranyl group, the compound of formula (II) may be prepared from hydroxy acetone and 3,4-dihydro-2H-pyran, preferably employing pyridinium toluene-4-sulfonate as a catalyst. A specific method for preparing acetonyl tetrahydropyranyl ether is described in example 24.
[0104]In step (a) of said process, the compound of formula (II) is converted to a compound of formula (III) or (IV) by a Wittig or a Horner reagent. Wittig-type reactions and reagents are known to skilled persons (see e.g. Watanabe et al. 1996 and references cited therein). Common Wittig reagents to be used for the above process are methylen-triphenylphosphoranes which may be substituted at the methylene group. For the above process of this invention, a methylen-triphenylphosphorane is employed which is substituted with the above-defined group D at the methylene group. Such Wittig reagents are commercially available or can be prepared according to known methods.
[0105]The olefin produced in step (a) may be formed as a mixture of the cis/trans isomers of formulas (III) and (IV). If one of said isomers is preferred, it may be enriched or separated from the other isomer by methods known in the art, preferably by chromatography. Alternatively, a separation of said isomers may be carried out after one of the following steps (b) to (e).
[0106]In step (b) of the above process, group D of the compound of formula (III) or (IV) or a mixture of said compounds is reductively converted to a --CH2--OH group. Various methods are known in the art to perform such a reduction. Conditions are chosen such that group D is reduced whereas the olefin moiety is not. Examples for reductants to be used in this step are molecular hydrogen or metal hydrides. Examples for useful metal hydrides include boron hydrides like sodium borohydride, aluminium hydrides like lithium aluminium hydride or diisobutyl aluminumhydride (DIBAH), alkali metal or metal earth hydrides like sodium hydride or calcium hydride. Aluminium hydrides are preferred. A specific example for carrying out step (b) is described in example 24.
[0107]If the desired end product of said process is a compound of formula (I), wherein R1 or R2 is --CH2--OH, the compound or mixture of compounds obtained in step (b) may be directly subjected to step (d) or step (e). Preferably, it is subjected to step (e) for removing protective group B. If the desired end product of said process is a compound of formula (I), wherein R1 or R2 is --CH2--O--PO(OH)--O--PO(OH)2 or --CH2--O--PO(OH)2, compound or mixture of compounds obtained in step (b) is subjected to step (c) of said process for converting --CH2--OH group obtained in step (b) into a --CH2--O--PO(OH)--O--PO(OH)2 or a --CH2--O--PO(OH)2 group.
[0108]Step (c) may be carried in several ways which are known to the skilled person. Step (c) may comprise substituting the hydroxy group of said --CH2--OH group obtained in step (b) by a leaving group. Step (c) may comprise converting said --CH2--OH group to a --CH2-halide group by a halogenating agent. A sulfuric, sulfonic or phosphoric acid halogenide may be employed as halogenating agent. Tosyl chloride is most preferred. Said halide may be fluoride, chloride, bromide or iodide, preferably chloride. The compound carrying said --CH2-halide group is preferably isolated. Said leaving group may further be created by reacting said --CH2--OH group obtained in step (b) with a sulfonic acid halide, preferably tosyl chloride.
[0109]Said intermediate having said leaving group may then be reacted with phosphoric or diphosphoric acid or singly or multiply deprotonated forms thereof. Preferably an alkylammonium salt of phosphoric or diphosphoric acid is used, more preferably a tetraalkylammonium salt, and most preferably a tetra-butylammonium salt. A polar aprotic solvent is preferred for this reaction. Preferably, the compound or mixture of compounds obtained is purified according to standard procedures. A specific example for carrying out step (c) is described in example 24.
[0110]In step (d), the compound or mixture of compounds obtained in step (c) may be converted to a desired salt. Methods for carrying out step (d) are well known. Such methods may comprise adjusting the pH of an aqueous solution with an appropriate acid or salt to a desired pH value.
[0111]In step (e), the protective group B of a compound obtained in one of steps (b) to (d) is removed in order to obtain a compound of formula (I) wherein A is --CH2--OH. The method for removing a protective group depends on the type of the protective group. Such methods are well known. If the protective groups forms an acetal, removing said protecting group may be achieved by acid hydrolysis (see example 24).
[0112]This invention provides protein in a form that is functional for the enzymatic conversion of 2C-methyl-D-erythritol 2,4-cyclodiphosphate to 1-hydroxy-2-methyl-2-butenyl 4-diphosphate notably in its (E)-form, preferably in the presence of NADH and/or NADPH and/or in the presence of Co2+. Said enzyme preferably has a sequence encoded by the ispG (formerly gcpE) gene of E. coli or a function-conservative homologue of said sequence, i.e. said homologue is capable of performing the same function as said protein. For many applications of said protein, it may be expressed and purified as a fusion protein, notably a fusion with maltose binding protein. In this way, enzymatically active protein may be readily obtained.
[0113]This invention further provides a protein in a form that is functional for the enzymatic conversion of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, notably in its (E)-form, to isopentenyl diphosphate and/or dimethylallyl diphosphate. Said protein preferably requires FAD and NAD(P)H for said functionality. Further, said protein may require a metal ion selected from the group of manganese, iron, cobalt, or nickel ion. Said protein preferably has a sequence encoded by the ispH (formerly lytB) gene of E. coli or a function-conservative homologue of said sequence. For many applications of said protein, it may be expressed and purified as a fusion protein, notably a fusion with maltose binding protein. In this way, enzymatically active protein may be readily obtained.
[0114]The above proteins may be plant proteins, notably from Arabidopsis thaliana, bacterial proteins, notably from E. coli, or protozoal proteins, notably from Plasmodium falciparum.
[0115]The invention further provides a purified isolated nucleic acid encoding one or both of the above proteins with or without introns. Further, the invention provides a DNA expression vector containing the sequence of said purified isolated nucleic acid.
[0116]The invention further provides cells, cell cultures, organisms or parts thereof recombinantly endowed with the sequence of said purified isolated nucleic acid or with said DNA expression vector, wherein said cell is selected from the group consisting of bacterial, protozoal, fungal, plant, insect and mammalian cells. Said cells, cell cultures, organisms or parts thereof may further be endowed with at least one gene selected from the following group: dxs, dxr, ispD (formerly ygbP); ispE (formerly ychB); ispF (formerly ygbB) of E. coli or a function-conservative homologue thereof, or a function-conservative fusion, deletion or insertion variant of any of the above genes.
[0117]The invention further provides cells, cell cultures, or organisms or parts thereof transformed or transfected for an increased rate of formation of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, notably in its (E)-form, compared to cells, cell cultures, or organisms or parts thereof absent said transformation or transfection. The transformation or transfection preferably comprises endowment with the gcpE gene of E. coli or with a function-conservative homologue from an other organism, e.g. plant or protozoal organism.
[0118]The invention also provides cells, cell cultures, or organisms or parts thereof transformed or transfected for an increased rate of conversion of (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate to isopentenyl diphosphate and/or dimethylallyl diphosphate compared to cells, cell cultures, or organisms or parts thereof absent said transformation or transfection. The transformation or transfection preferably comprises endowment with the lytB gene of E. coli or with a function-conservative homologue from an other organism, e.g. plant or protozoal organism.
[0119]The invention provides also cells, cell cultures, or organisms or parts thereof transformed or transfected for an increased expression level of the protein of one of claims 1 to 4 and/or the protein of one of claims 5 to 8 compared to cells, cell cultures, or organisms or parts thereof absent said transformation or transfection.
[0120]Moreover, the invention provides a method of altering the expression level of the gene product(s) of ispG and/or ispH or function-conservative homologues from other organisms or variants thereof in cells comprising [0121](a) transforming host cells with the ispG and/or ispH gene, [0122](b) growing the transformed host cells of step (a) under conditions that are suitable for the efficient expression of ispG and/or ispH, resulting in production of altered levels of the ispG and/or ispH gene product(s) in the transformed cells relative to expression levels of untransformed cells.
[0123]Furthermore, the invention provides a method of identifying an inhibitor of an enzyme functional for the conversion of 2C-methyl-D-erythritol 2,4-cyclodiphosphate to 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, notably its E-form, of the non-mevalonate isoprenoid pathway by the following steps: [0124](a) incubating a mixture containing said enzyme with its, optionally isotope-labeled, substrate 2C-methyl-D-erythritol-2,4-cyclodiphosphate under conditions suitable for said conversion in the presence and in the absence of a potential inhibitor, [0125](b) subsequently determining the concentration of 2C-methyl-D-erythritol 2,4-cyclodiphosphate and/or 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, and [0126](c) comparing the concentration in the presence and in the absence of said potential inhibitor.
[0127]Furthermore, the invention provides a method of identifying an inhibitor of an enzyme functional for the conversion of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, notably its E-form, to isopentenyl diphosphate or dimethylallyl diphosphate of the non-mevalonate isoprenoid pathway by the following steps: [0128](a) incubating a mixture containing said enzyme with its, optionally isotope-labeled, substrate 1-hydroxy-2-methyl-2-butenyl 4-diphosphate under conditions suitable for said conversion in the presence and in the absence of a potential inhibitor, whereby said mixture preferably contains FAD, [0129](b) determining the concentration of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate and/or isopentenyl diphosphate or dimethylallyl diphosphate, and [0130](c) comparing the concentration in the presence and in the absence of said potential inhibitor.
[0131]The above methods of identifying an inhibitor are preferably carried out by following the consumption of NADPH or NADH making use of its characteristic absorbance spectrum. Alternatively, the fluorescence of NADH or NADPH can be followed when excited around 340 nm. The above methods of identifying an inhibitor may advantageously be performed as high-throughput screening assays for inhibitors, notably in combination with photometric detection of the consumption of NADH or NADPH. Further, one or more flavin analogues (e.g. FAD, FMN) may be added to the incubation mixtures in said methods, preferably in catalytic amounts. Most preferred is the addition of FAD. Said enzymes may be employed in said methods as fusion proteins with maltose binding protein (examples 38 to 41, 44, 45), which allows straightforward expression and purification of said enzymes in enzymatically active form. Further embodiments of said methods of identifying are defined in the subclaims to these methods.
[0132]It is known that intermediates of the non-mevalonate pathway are responsible for γδ T cell activation by various pathogenic bacteria. γδ T cell activation is followed by T cell proliferation, secretion of cytokines and chemokines and is very likely crucial for regulating the immune response following pathogen infection (Altincicek et al., 2001 and references cited therein). Recently, it was shown that E. coli strains lost the ability to stimulate γδ T cells when the dxr or the gcpE gene was knocked out, strongly indicating that an intermediate downstream of gcpE and upstream of isopentenyl pyrophosphate exhibits the most potent antigenic activity (Altincicek et al., 2001). However, the intermediate produced by the gcpE gene product in the pathway has been unknown. Herein, this intermediate has surprisingly been identified as an hitherto unprecedented compound, which opens up a whole range of novel applications for this compound.
[0133]The compounds of formula I can be used as immunomodulatory or immunostimulating agents, e.g. for activating γδ T cells. Immunomodulation via γδ T cell activation by said compounds may prove useful not only to support combat against pathogens but for various conditions for which a stimulation of the immune system is desirable. The novel compounds of the invention may therefore be used for medical treatment of pathogen infections. Such a treatment stimulates the activity of the immune system against the pathogen. Preferably, the compound wherein R1=H and/or A is --CH2OH is used for this application. Alternatively, the oxidation product with A=CHO may prove to be highly active. Among the compounds of formula I, the one with the highest or most suitable γδ T cell stimulating activity may be selected in a test system known in the art (e.g. that described by Altincicek et al., 2001). Importantly, since the compounds of the invention do not act as antibiotics, development of resistancies is not a problem for the method of treatment disclosed herein.
[0134]In an advantageous embodiment, said compounds may be combined with an antibiotically active compound for treating a pathogen infection. Such a treatment combines the advantages of inhibiting pathogen proliferation by an antibiotic and stimulating the immune system against the pathogen resulting in a much faster and more efficient treatment. Such an antibiotically active compound may be a bacteriostatic antibiotic (e.g. tetracyclines).
[0135]Therefore, the novel compounds of this invention may be used for the preparation of a medicament. The invention further pertains to a pharmaceutical composition containing a compound of formula I and a pharmaceutically acceptable carrier. Said pharmaceutical composition may further contain an antibiotically active compound as mentioned above.
[0136]This invention further comprises antibodies against the compounds of formula I. Said antibodies may be polyclonal or monoclonal and may be raised according to conventional techniques. Raising such antibodies will comprise coupling of a compound of formula I has hapten to a macromolecular carrier like a protein in order to be immunogenic. Such an immunogenic compound of formula I may further be used as a vaccine.
[0137]The antibodies of the invention may be used for detecting a compound of formula I. Since said compounds are produced by organisms having the non-mevalonate isoprenoid pathway, such organisms may be detected using said antibodies. Preferably, such organisms may be detected in body fluids in a diagnostic method, thereby indicating an infection by a pathogen having the non-mevalonate pathway. A positive result in such a diagnostic method may at the same time indicate possible treatment by the compounds of the invention.
[0138]When an antibody of the invention is used for detecting a compound of formula I, it is preferably labelled to allow photometric detection and/or immobilized to a support. Such methods are well-known in the art.
[0139]This invention further provides a process for the chemical preparation of a compound of formula I or a salt thereof (see FIG. 7):
wherein A represents --CH2OH or --CHO, R1 is hydrogen, and R2 is --CH2--O--PO(OH)--O--PO(OH)2, --CH2--O--PO(OH)2 or --CH2--OH by the following steps:
[0140](a) converting 2-methyl-2-vinyl-oxiran into 4-chloro-2-methyl-2-buten-1-al;
[0141](b) converting 4-chloro-2-methyl-2-buten-1-al to its acetal;
[0142](c) substituting the chlorine atom in the product of step (b) by a hydroxyl group, a phosphate group or a pyrophosphate group;
[0143](d) hydrolysing the acetal obtained in step (c) to produce an aldehyde group;
[0144](e) optionally converting the aldehyde group of the product of step (d) to a --CH2OH group.
[0145]Preferred embodiments of this process are defined in the subclaims and are exemplified in example 42.
[0146]The invention will now be described in detail with reference to specific examples.
EXAMPLE 1
Construction of a Vector Carrying the xylB Gene of Escherichia coli Capable for Transcription and Expression of D-xylulokinase
[0147]Chromosomal DNA from Escherichia coli strain XL1-Blue (Bullock et al. 1987; commercial source: Stratagene, LaJolla, Calif., USA) is isolated according to a method described by Meade et al. 1982.
[0148]The E. coli ORF xylB (accession no. gb AE000433) from base pair (bp) position 8596 to 10144 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 pmol of the primer 5'-CCGTCGGAATTCGAGGAGAAATTAACCATGTATATCGGGATAGATCTTGG-3' (SEQ ID NO:1), 10 pmol of the primer 5'-GCAGTGAAGCTTTTACGCCATTAATGGCAGAAGTTGC-3' (SEQ ID NO:2), 20 ng of chromosomal DNA, 2 U of Taq DNA polymerase (Eurogentec, Seraing, Belgium) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0149]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 60 sec at 94° C., 60 sec at 50° C. and 75 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0150]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden, Germany).
[0151]1.0 μg of the vector pBluescript SKII.sup.- (Stratagene) and 0.5 μg of the purified PCR product are digested with EcoRI and HindIII in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (New England Biolabs, Frankfurt am Main, Germany (NEB)) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0152]20 ng of the purified vector DNA and 20 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pBSxylB. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells according to a method described by Dower et al., 1988. The plasmid pBSxylB is isolated with the plasmid isolation kit from Qiagen.
[0153]The DNA insert of the plasmid pBSxylB is sequenced by the automated dideoxynucleotide method (Sanger et al., 1992) using an ABI Prism 377® DNA sequencer from Perkin Elmer (Norwalk, USA) with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions (Foster city, USA). It is identical with the DNA sequence of the database entry (gb AE000433).
EXAMPLE 2
Construction of a Vector Carrying the XylB and dxr Genes of Escherichia coli Capable for Transcription and Expression of D-xylulokinase and DXP reductoisomerase
[0154]The E. coli ORF dxr (accession no. gb AE000126) from base pair (bp) position 9887 to 11083 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 pmol of the primer 5'-CTAGCCAAGCTTGAGGAGAAATTAACCATGAAGCAACTCACCATTCTGG-3' (SEQ ID NO:3), 10 pmol of the primer 5'-GGAGATGTCGACTCAGCTTGCGAGACGC-3' (SEQ ID NO:4), 20 ng of chromosomal DNA, 2 U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0155]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 60 sec at 94° C., 60 sec at 50° C. and 75 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0156]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden).
[0157]1.2 μg of the vector pBSxylB (Example 1) and 0.6 μg of the purified PCR product are digested with HindIII and SalI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0158]20 ng of the purified vector DNA and 18 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pBSxylBdxr. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pBSxylBdxr is isolated with the plasmid isolation kit from Qiagen.
[0159]The DNA insert of the plasmid pBSxylBdxr is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb AE000126).
[0160]The DNA sequence of the vector construct pBSxylBdxr is shown in FIG. 8.
EXAMPLE 3
Construction of a Vector Carrying the XylB, dxr and ispD Genes of Escherichia coli Capable for Transcription and Expression of D-xylulokinase, DXP Reductoisomerase and CDP-ME Synthase
[0161]The E. coli ORF ispD (accession no. gb AE000358) from base pair (bp) position 6754 to 7464 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 pmol of the primer 5'-CCGGGAGTCGACGAGGAGAAATTAACCATGGCAACCACTCATTTGGATG-3' (SEQ ID NO:5), 10 pmol of the primer 5'-GTCCAACTCGAGTTATGTATTCTCCTTGATGG-3' (SEQ ID NO:6), 20 ng of chromosomal DNA, 2 U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0162]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 30 sec at 94° C., 30 sec at 50° C. and 45 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0163]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden).
[0164]1.5 μg of the vector pBSxylBdxr (Example 2) and 0.8 μg of the purified PCR product are digested with SalI and XhoI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0165]20 ng of the purified vector DNA and 12 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pBSxylBdxrispD. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pBSxylBdxrispD is isolated with the plasmid isolation kit from Qiagen.
[0166]The DNA insert of the plasmid pBSxylBdxrispD is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb AE000126).
[0167]The DNA sequence of the vector construct pBSxylBdxrispD is shown in FIG. 9.
EXAMPLE 4
Construction of a Vector Carrying the xylB, dxr, ispD and ispF Genes of Escherichia coli Capable for Transcription and Expression of D-xylulokinase, DXP Reductoisomerase, CDP-ME Synthase, and cMEPP Synthase
[0168]The E. coli ORF's ispD and ispF (accession no. gb AE000358) from base pair (bp) position 6275 to 7464 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 pmol of the primer 5'-CCGGGAGTCGACGAGGAGAAATTAACCATGGCAACCACTCATTTGGATG-3' (SEQ ID NO:7), 10 pmol of the primer 5'-TATCAACTCGAGTCATTTTGTTGCCTTAATGAG-3' (SEQ ID NO:8), 20 ng of chromosomal DNA, 2 U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0169]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 60 sec at 94° C., 60 sec at 50° C. and 75 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0170]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden).
[0171]1.4 μg of the vector pBSxylBdxr (Example 2) and 0.7 μl of the purified PCR product are digested with SalI and XhoI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0172]20 ng of the purified vector DNA and 18 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl yielding the plasmid pBSxylBdxrispDF. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pBSxylBdxrispDF is isolated with the plasmid isolation kit from Qiagen.
[0173]The DNA insert of the plasmid pBSxylBdxrispDF is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb AE000126).
EXAMPLE 5
Construction of a Vector Carrying the xylB, dxr, ispD, ispE and ispF Genes of Escherichia coli Capable for Transcription and Expression of D-xylulokinase, DXP Reductoisomerase, CDP-ME Synthase, CDP-ME Kinase and cMEPP Synthase
[0174]The E. coli ORF ispE (accession no. gb AE000219) from base pair (bp) position 5720 to 6571 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 pmol of the primer 5'-GCGAACCTCGAGGAGGAGAAATTAACCATGCGGACACAGTGGCCC-3' (SEQ ID NO:9), 10 pmol of the primer 5'-CCTGACGGTACCTTAAAGCATGGCTCTGTGC-3' (SEQ ID NO:10), 20 ng of chromosomal DNA, 2 U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0175]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 45 sec at 94° C., 45 sec at 50° C. and 60 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0176]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden). 1.2 μg of the vector pBSxylBdxrispDF (Example 4) and 0.6 μg of the purified PCR product are digested with XhoI and KpnI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0177]20 ng of the purified vector DNA and 15 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pBScyclo. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pBScyclo is isolated with the plasmid isolation kit from Qiagen.
[0178]The DNA insert of the plasmid pBScyclo is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb AE000219). The DNA sequence of the vector construct pBScyclo is shown in FIG. 10.
EXAMPLE 6
Construction of a Vector Carrying the gcpE gene of Escherichia coli Capable for its Transcription and Expression
[0179]The E. coli ORF gcpE (accession no. gb AE000338) from base pair (bp) position 372 to 1204 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 pmol of the primer 5'-CGTACCGGATCCGAGGAGAAATTAACCATGCATAACCAGGCTCCAATTC-3' (SEQ ID NO: 11), 10 pmol of the primer 5'-CCCATCGTCGACTTATTTTTCAACCTGCTGAACGTC-3' (SEQ ID NO:12), 20 ng of chromosomal DNA, 2 U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0180]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 60 sec at 94° C., 60 sec at 50° C. and 90 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0181]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden).
[0182]2.0 μg of the vector pACYC184 (Chang and Cohen 1978, NEB) and 0.7 μg of the purified PCR product are digested with BamHI and SalI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0183]20 ng of the purified vector DNA and 20 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pACYCgcpE. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pACYCgcpE is isolated with the plasmid isolation kit from Qiagen.
[0184]The DNA insert of the plasmid pACYCgcpE is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb AE000338).
[0185]The DNA sequence of the vector construct pACYCgcpE is shown in FIG. 11.
EXAMPLE 7
Construction of Vectors Carrying a Carotenoid Operon from Erwinia uredovora Capable for the In Vivo Production of β-carotene
[0186]The open reading frames crtY, crtI and crtB of a carotenoid operon from Erwinia uredovora (accession no. gb D90087) from base pair (bp) position 2372 to 6005 is amplified by PCR using chromosomal E. uredovora DNA as template. The reaction mixture contains 10 pmol of the primer 5'-CATTGAGAAGCTTATGTGCACCG-3' (SEQ ID NO:13), 10 Pmol of the primer 5'-CTCCGGGGTCGACATGGCGC-3' (SEQ ID NO:14), 40 ng of chromosomal DNA of E. uredovora, 8 U of Taq DNA polymerase (Eurogentec), 20 nmol of dNTPs, Taq Extender (Stratagene) in a total volume of 100 μl 1× Taq Extender buffer (Stratagene).
[0187]The mixture is denaturated for 3 min at 94° C. Then 40 PCR cycles for 60 sec at 94° C., 60 sec at 50° C. and 300 sec at 72° C. followed. After further incubation for 20 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0188]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden, Germany).
[0189]1.0 μg of the vector pBluescript SKII.sup.- (Stratagene) and 2.0 μg of the purified PCR product are digested with HindIII and SalI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0190]20 ng of the purified vector DNA and 40 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pBScaro34. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pBScaro34 is isolated with the plasmid isolation kit from Qiagen.
[0191]The DNA insert of the plasmid pBScaro34 is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb D90087).
[0192]The E. uredovora ORF crtE (accession no. gb D90087) from base pair (bp) position 175 to 1148 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 pmol of the primer 5'-CCGCATCTTTCCAATTGCCG-3' (SEQ ID NO:15), 10 pmol of the primer 5'-ATGCAGCAAGCTTAACTGACGGC-3' (SEQ ID NO:16), 20 ng of chromosomal DNA, 2 U of Taq DNA polymerase (Eurogentec, Seraing, Belgium) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0193]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 45 sec at 94° C., 45 sec at 50° C. and 60 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0194]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden, Germany).
[0195]1.5 μg of the vector pBScaro34 (see above) is digested with EcoRI and HindIII and 0.6 μg of the purified PCR product are digested with MfeI and HindIII in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0196]20 ng of the purified vector DNA and 16 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pBScaro14. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pBScaro14 is isolated with the plasmid isolation kit from Qiagen.
[0197]The DNA insert of the plasmid pBScaro14 is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb D90087). The DNA sequence of the plasmid pBScaro14 is shown in FIG. 12.
[0198]5 μg of the vector pBScaro14 (see above) is digested with BamHI and SalI. The restriction mixture is prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. The restriction mixture is separated on a agarose gel and the fragments of 2237 and 2341 bp size are purified with the gel extraction kit from Qiagen.
[0199]3 μg of the vector pACYC184 (see above) is digested with BamHI and SalI. The restriction mixture is prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. The restriction mixture is separated on a agarose gel and the fragment of 3968 bp size is purified with the gel extraction kit from Qiagen.
[0200]30 ng of the purified vector DNA and each 25 ng of the purified 2237 and 2341 bp fragments are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pACYCcaro14. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pACYCcaro14 is isolated with the plasmid isolation kit from Qiagen.
[0201]The DNA insert of the plasmid pACYCcaro14 is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb D90087). The DNA sequence of the plasmid pACYCcaro14 is shown in FIG. 13.
EXAMPLE 8
Enzymatic Preparation of [U-13C5]1-deoxy-D-xylulose 5-phosphate
[0202]A reaction mixture containing 960 mg of [U-13C6]glucose (5.1 mmol), 6.1 g of ATP (10.2 mmol), 337 mg of thiamine pyrophosphate, 1.14 g of [2,3-13C2]pyruvate (10.2 mmol), 10 mM MgCl2, 5 mM dithiothreitol in 150 mM Tris hydrochloride, pH 8.0 is prepared. 410 Units of triose phosphate isomerase (from rabbit muscle, Type III-S, E. C. 5.3.1.1., Sigma), 100 U hexokinase (from Bakers Yeast, Type VI, E. C. 2.7.1.1, Sigma), 100 U phosphoglucose isomerase (from Bakers Yeast, Type III, E. C. 5.3.1.9, Sigma), 100 U phosphofructokinase (from Bacillus stearothermophilus, Type VII, E. C. 2.7.1.11, Sigma), 50 U aldolase (from rabbit muscle, E. C. 4.1.2.13, Sigma) and 12 U of recombinant DXP synthase from B. subtilis are added to a final volume of 315 ml. The reaction mixture is incubated at 37° C. overnight and during incubation the pH is hold at a constant value of 8.0. The reaction is monitored by 13C NMR spectroscopy.
EXAMPLE 9
Enzymatic Preparation of [3,4,5-13C3]1-deoxy-D-xylulose 5-phosphate
[0203]A solution containing 150 mM Tris hydrochloride, 10 mM MgCl2, 1.0 g of [U-13C6]glucose (5.4 mmol), 0.23 g (1.5 mmol) of dithiothreitol, 0.3 g (0.7 mmol) of thiamine pyrophosphate, 0.1 g (0.2 mmol) of ATP (disodium salt), and 2.2 g (11 mmol) of phosphoenol pyruvate (potassium salt) is adjusted to pH 8.0 by the addition of 8 M sodium hydroxide. 403 U (2.8 mg) of pyruvate kinase (from rabbit muscle, E. C. 2.7.1.40), 410 Units of triose phosphate isomerase (from rabbit muscle, Type III-S, E. C. 5.3.1.1., Sigma), 100 U hexokinase (from Bakers Yeast, Type VI, E. C. 2.7.1.1, Sigma), 100 U phosphoglucose isomerase (from Bakers Yeast, Type III, E. C. 5.3.1.9, Sigma), 100 U phosphofructokinase (from Bacillus stearothermophilus, Type VII, E. C. 2.7.1.11, Sigma), 50 U aldolase (from rabbit muscle, E. C. 4.1.2.13, Sigma) and 12 U recombinant DXP synthase from B. subtilis are added to a final volume of 300 ml. The reaction mixture is incubated at 37° C. for overnight.
EXAMPLE 10
Enzymatic Preparation of 1-deoxy-D-xylulose
[0204]The pH value of the reaction mixture obtained in example 8 or 9 is adjusted to 9.5. Magnesium chloride is added to a concentration of 30 mM. 50 mg (950 Units) of alkaline phosphatase from bovine intestinal mucosa (Sigma, E. C. 3.1.3.1) are added and the reaction mixture is incubated for 16 h. The conversion is monitored by 13C-NMR spectroscopy. The pH is adjusted to a value of 7.0 and the solution is centrifuged at 14,000 upm for 5 minutes. Starting from labelled glucose (examples 8 or 9) the overall yield of 1-deoxy-D-xylulose is approximately 50%. The supernatant or the lyophilised supernatant is used in incorporation experiments (see examples 11 to 17).
EXAMPLE 11
Incorporation Experiment with Recombinant Escherichia coli XL1-pBSxylB Using [3,4,5-13C3]1-deoxy-D-xylulose
[0205]0.2 litre of Luria Bertani (LB) medium containing 36 mg of ampicillin are inoculated with 10 ml of an overnight culture of E. coli strain XL1-Blue harbouring the plasmid pBSxylB (see example 1). The cells are grown in a shaking culture at 37° C. At an optical density (600 nm) of 0.6 the culture is induced with 2 mM IPTG. Two hours after induction with IPTG, 50 ml (0.9 mmol) of crude [3,4,5-13C3]1-deoxy-D-xylulose (pH 7.0) (see examples 9 and 10), are added. Aliquots of 25 ml are taken at time intervals of 30 minutes and centrifuged for 20 min at 5,000 rpm and 4° C. The cells are washed with water containing 0.9% NaCl and centrifuged as described above. The cells are suspended in 700 μl of 20 mM NaF in D2O, cooled on ice and sonified 3×10 sec with a Branson Sonifier 250 (Branson SONIC Power Company) set to 90% duty cycle output, control value of 4. The suspension is centrifuged at 15,000 rpm for 15 min. 13C NMR spectra of the supernatant are recorded directly, without further purification, with a Bruker AVANCE DRX 500 spectrometer (Karlsruhe, Germany). The NMR analysis is based on published signal assignments (Wungsintaweekul et al., 2001).
[0206]30 min after the addition of [3,4,5-13C3]1-deoxy-D-xylulose, the formation of [3,4,5-13C3]1-deoxy-D-xylulose 5-phosphate can be observed. The maximum yield of [3,4,5-13C3]1-deoxy-D-xylulose 5-phosphate is observed 3-5 h after addition of [3,4,5-13C3]1-deoxy-D-xylulose to the medium. The 13C NMR signals reveal a mixture of [3,4,5-13C3]1-deoxy-D-xylulose and [3,4,5-13C3]1-deoxy-D-xylulose-5-phosphate at a molar ratio of approximately 1:9. The intracellular concentration of [3,4,5-13C3]1-deoxy-D-xylulose 5-phosphate is estimated as 20 mM by quantitative NMR spectroscopy.
EXAMPLE 12
Incorporation Experiment with recombinant Escherichia coli XL1-pBSxylBdxr Using [U-13C5]1-deoxy-D-xylulose
[0207]0.12 litre of Luria Bertani (LB) medium containing 22 mg of ampicillin are inoculated with 10 ml of an overnight culture of E. coli strain XL1-Blue harbouring plasmid pBSxylBdxr (see example 2). The cells are grown in a shaking culture at 37° C. At an optical density (600 nm) of 0.6 the culture is induced with 2 mM IPTG. Two hours after induction with IPTG, ca. 1.0 mmol of crude [U-13C5]1-deoxy-D-xylulose (pH 7.0) (see examples 8 and 10) are added. Aliquots of 25 ml are taken in time intervals of 1 h and centrifuged for 20 min at 5,000 rpm and 4° C. The cells are washed with water containing 0.9% NaCl and centrifuged as described above. The cells are suspended in 700 μl of 20 mM NaF in D2O, cooled on ice and sonified 3×10 sec with a Branson Sonifier 250 (Branson SONIC Power Company) set to 90% duty cycle output, control value of 4. The suspension is centrifuged at 15,000 rpm for 15 min. NMR spectra of the supernatant are recorded directly, without further purification, with a Bruker AVANCE DRX 500 spectrometer (Karlsruhe, Germany).
[0208]HMQC and HMQC-TOCSY experiments reveal 1H-13C and 1H-1H spin systems (Table 1) of [U-13C5]2C-methyl-D-erythritol 4-phosphate, [U-13C5]2C-methyl-D-erythritol and [1,2,2',3,4-13C5]4-diphosphocytidyl-2C-methyl-D-erythritol at a molar ratio of approximately 6.6:7:1, respectively. The intracellular concentration of [U-13C5]2C-methyl-D-erythritol 4-phosphate is estimated as 10 mM by quantitative NMR spectroscopy.
[0209]The NMR data summarized in Table 1 are identical with published NMR data of the authentic compounds (Takahashi et al., 1998; Rohdich et al., 1999).
TABLE-US-00001 TABLE 1 NMR data of 13C-labeled products in cell extracts of E. coli XL1-pBSxylBdxr after feeding of [U-13C5]1-deoxy-D-xylulose Chemical shifts, ppm Position 1 1* 2 2-Methyl 3 4 4* [U-13C5]2C-methyl-D-erythritol 4-phosphate 13C 66.1 n.d. 18.1 73.4 648 1H 3.25 3.36 0.93 3.56 3.62 3.81 [U-13C5]2C-methyl-D-erythritol 13C 66.6 n.d. 18.0 74.6 616 1H 3.26 3.34 0.9 3.44 3.36 3.61 [1,2,2',3,4-13C5]4-diphosphocytidyl-2C-methyl-D-erythritol 13C 66.8 n.d. 18.0 73.0 667 1H 3.4 3.55 0.9 3.6 3.74 4
EXAMPLE 13
Incorporation Experiment with Recombinant Escherichia coli XL1-pBSxylBdxrispDF Using [3,4,5-13C3]1-deoxy-D-xylulose
[0210]0.1 litre of Luria Bertani (LB) medium containing 18 mg of ampicillin are inoculated with 10 ml of an overnight culture of E. coli strain XL1-Blue harbouring the plasmid pBSxylBdxrispDF (see example 4). The cells are grown in a shaking culture at 37° C. At an optical density (600 nm) of 0.5, the culture is induced with 2 mM IPTG. Two hours after induction with IPTG, ca. 1.0 mmol of crude [3,4,5-13C3]1-deoxy-D-xylulose (see examples 9 and 10) are added. After three hours, cells were harvested and centrifuged for 20 min at 5,000 rpm and 4° C. The cells are washed with water containing 0.9% NaCl and centrifuged as described above. The cells are suspended in 1.5 ml of 20 mM NaF in D2O, cooled on ice and sonified 3×15 sec. with a Branson Sonifier 250 (Branson SONIC Power Company) set to 90% duty cycle output, control value of 4. The suspension is centrifuged at 15,000 rpm for 15 min. NMR spectra of the supernatant are recorded directly, without further purification, with a Bruker AVANCE DRX 500 spectrometer (Karlsruhe, Germany).
[0211]HMQC and HMQC-TOCSY experiments reveal 1H-13C and 1H-1H spin systems of [1,3,4-13C3]2C-methyl-D-erythritol 4-phosphate, [1,3,4-13C3]2C-methyl-D-erythritol and [1,3,4-13C5]4-diphosphocytidyl-2C-methyl-D-erythritol (Table 1). The molar ratios of [1,3,4-13C3]2C-methyl-D-erythritol 4-phosphate, [1,3,4-13C3]2C-methyl-D-erythritol and [1,3,4-13C5]4-diphosphocytidyl-2C-methyl-D-erythritol are 1:0.6:0.9, respectively.
[0212]This result indicates that the intracellular amount of CTP required for the synthesis of 4-diphosphocytidyl-2C-methyl-D-erythritol is limiting. Therefore, a modified fermentation protocol was developed (see example 14).
EXAMPLE 14
Incorporation Experiment with Recombinant Escherichia coli XL1-pBSxylBdxrispDF Using [3,4,5-13C3]1-deoxy-D-xylulose
[0213]0.1 litre of Luria Bertani (LB) medium containing 18 mg of ampicillin are inoculated with 10 ml of an overnight culture of E. coli strain XL1-Blue harbouring plasmid pBSxylBispDF (see example 4). The cells are grown in a shaking culture at 37° C. At an optical density (600 nm) of 0.5, the culture is induced with 2 mM IPTG. Two hours after induction with IPTG, 10 mg (0.041 mmol) of cytidine and 5 ml of 1 M NaKHPO4, pH 7.2, and ca. 1 mmol of crude [3,4,5-13C3]1-deoxy-D-xylulose (see examples 9 and 10) are added. After three hours, the cells are harvested and centrifuged for 20 min at 5,000 rpm and 4° C. The cells are washed with water containing 0.9% NaCl and centrifuged as described above. The cells are suspended in 700 μl of 20 mM NaF in D2O, cooled on ice and sonified 3×10 sec with a Branson Sonifier 250 (Branson SONIC Power Company) set to 90% duty cycle output, control value of 4. The suspension is centrifuged at 15,000 rpm for 15 min. NMR spectra of the supernatant are recorded directly, without further purification, with a Bruker AVANCE DRX 500 spectrometer. HMQC and HMQC-TOCSY experiments reveal 1H-13C and 1H-1H spin systems (Table 1) of [1,3,4-13C3]2C-methyl-D-erythritol 4-phosphate, [1,3,4-13C3]2C-methyl-D-erythritol and [1,3,4-13C3]4-diphosphocytidyl-2C-methyl-D-erythritol at a molar ratio of approximately 1:3.4:4.2, respectively. The relative amount of [1,3,4-13C3]4-diphosphocytidyl-2C-methyl-D-erythritol is increased by a factor of 2 as compared to the relative amount in example 13. The intracellular concentration of [1,3,4-13C3]4-diphosphocytidyl-2C-methyl-D-erythritol is estimated as 10 mM by quantitative NMR spectroscopy. The relative high amount of 2C-methyl-D-erythritol indicates that unspecific phosphatases convert intermediary formed 2C-methyl-D-erythritol 4-phosphate into 2C-methyl-D-erythritol. Therefore, a modified fermentation protocol was developed to supply the cells with sufficient amounts of organic phosphates and in order to suppress the activity of phosphatases (see examples 15 to 17).
EXAMPLE 15
Incorporation Experiment with Recombinant Escherichia coli XL1-pBScyclo Using [U-13C5]1-deoxy-D-xylulose
[0214]0.2 litre of Luria Bertani (LB) medium containing 36 mg of ampicillin are inoculated with 10 ml of an overnight culture of E. coli strain. XL1-Blue harbouring the plasmid pBScyclo (see example 5). The cells are grown in a shaking culture at 37° C. At an optical density (600 nm) of 1.3, the culture is induced with 2 mM IPTG. Two hours after induction with IPTG, 30 mg (0.12 mmol) of cytidine, 300 mg (0.95 mmol) of DL-α-glycerol 3-phosphate and 10 ml of 1 M NaKHPO4, pH 7.2, are added. After 30 min, ca. 1 mmol of [U-13C5]1-deoxy-D-xylulose (see example 8 and 10) are added. Aliquots of 25 ml are taken at time intervals of 1 h and centrifuged for 20 min at 5,000 rpm and 4° C. The cells are washed with water containing 0.9% NaCl and centrifuged as described above. The cells are suspended in 700 μl of 20 mM NaF in D2O, cooled on ice and sonified 3×10 sec with a Branson Sonifier 250 (Branson SONIC Power Company) set to 90% duty cycle output, control value of 4. The suspension is centrifuged at 15,000 rpm for 15 min. NMR spectra of the cell free extract are recorded directly, without further purification, with a Bruker AVANCE DRX 500 spectrometer (Karlsruhe, Germany). 13C NMR spectra, as well as HMQC and HMQC-TOCSY spectra established [U-13C5]2C-methyl-D-erythritol 2,4-cyclodiphosphate (Herz et al., 2000) as the only product. Formation of [U-13C5]2C-methyl-D-erythritol 2,4-cyclodiphosphate can be observed 30 min after addition of [U-13C5]1-deoxy-D-xylulose, whereas the maximum yield is observed 5 h after addition of [U-13C5]1-deoxy-D-xylulose. The intracellular concentration of [U-13C5]2C-methyl-D-erythritol 2,4-cyclodiphosphate is estimated as 20 mM by quantitative NMR spectroscopy. The formation of any other isotope-labelled products, such as [U-13C5]2C-methyl-erythritol is completely suppressed.
EXAMPLE 16
Incorporation Experiment with Recombinant Escherichia coli XL1-pBScyclo-pACYCgcpE Using [2-14C]- and [U-13C5]1-deoxy-D-xylulose
[0215]0.2 litre of Terrific Broth (TB) medium containing 36 mg of ampicillin and 2.5 mg of chloramphenicol are inoculated with the E. coli strain XL1-Blue harbouring the plasmids pBScyclo and pACYCgcpE (see example 5 to 6). The cells are grown in a shaking culture at 37° C. overnight. At an optical density (600 nm) of 4.8 to 5.0, 30 mg (0.1 mmol) of cytidine, 300 mg (0.94 mmol) of DL-α-glycerol 3-phosphate and 10 ml of 1 M NaKHPO4, pH 7.2, are added. After 30 minutes, a mixture of 2.6 μmol [2-14C]1-deoxy-D-xylulose (15 μCi μmol-1) (Wungsintaweekul et al., 2001) and 1 ml of crude [U-13C5]1-deoxy-D-xylulose (0.02 mmol) (see examples 8 and 10) are added. After 1.5 h, cells are harvested and centrifuged for 10 min at 5,000 rpm and 4° C. The cells are washed with water containing 0.9% NaCl and centrifuged as described above. The cells are suspended in a mixture of 20 mM NaF (2 ml) and methanol (2 ml), cooled on ice and sonified 3×15 sec with a Branson Sonifier 250 (Branson SONIC Power Company) set to 90% duty cycle output, control value of 4. The suspension is centrifuged at 15,000 rpm for 15 min. The radioactivity of the supernatant is measured by scintillation counting (Beckmann, LS 7800). 10% of the radioactivity initially added as 14C labelled 1-deoxy-D-xylulose is detected in the supernatant. Aliquots are analysed by TLC and HPLC, as described in example 19, and the products are purified as described in example 20.
[0216]On basis of these data, 1-hydroxy-2-methyl-2-butenyl 4-diphosphate and 2-C-methyl-D-erythritol 2,4-cyclodiphosphate were identified as products at a molar ratio of 7:3 (see also examples 17 and 18).
EXAMPLE 17
Incorporation Experiment with Recombinant Escherichia coli XL1-pBScyclo-pACYCgcpE Using [U-13C5]- or [3,4,5-13C3]1-deoxy-D-xylulose
[0217]0.2 litre of Terrific Broth (TB) medium containing 36 mg of ampicillin and 2.5 mg of chloramphenicol are inoculated with the E. coli strain XL1-Blue harbouring the plasmids pBScyclo and pACYCgcpE. The cells are grown in a shaking culture at 37° C. for overnight. At an optical density (600 nm) of 4.8-5.0, 30 mg (0.1 mmol) of cytidine, 300 mg (0.93 mmol) of DL-α-glycerol 3-phosphate and 10 ml of 1 M NaKHPO4, pH 7.2, are added. After 30 minutes, 3 ml of crude [3,4,5-13C3]- or [U-13C5]1-deoxy-D-xylulose (0.05 mmol) (see examples 8, 9, and 10) are added. Aliquots of 25 ml are taken at time intervals of 1 h and centrifuged for 20 min at 5,000 rpm and 4° C. The cells are washed with water containing 0.9% NaCl and centrifuged as described above. The cells are suspended in 700 μl of 20 mM NaF in D2O or in 700 μl of a mixture of methanol and D2O (6:4; v/v) containing 10 mM NaF, cooled on ice and sonified 3×10 sec with a Branson Sonifier 250 (Branson SONIC Power Company) set to 90% duty cycle output, control value of 4 output. The suspension is centrifuged at 15,000 rpm for 15 min. NMR spectra of the cell free extracts are recorded directly with a Bruker AVANCE DRX 500 spectrometer (Karlsruhe, Germany). In order to avoid degradation during work-up, the structures of the products are determined by NMR spectroscopy without further purification (see example 18).
EXAMPLE 18
Structure Determination of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate
[0218]The 1H-decoupled 13C NMR spectrum using [U-13C5]1-deoxy-D-xylulose as starting material displays 5 13C-13C coupled signals belonging to 2C-methyl-D-erythritol 2,4-cyclodiphosphate (Herz et al., 2000) and 5 13C-13C coupled signals at 14.7, 64.5, 68.6, 122.7 and 139.5 ppm (Table 2) belonging to an unknown metabolite. The chemical shifts of the unknown metabolite suggest a double bond motif (signals at 122.7 and 139.5 ppm), a methyl group (signal at 14.7 ppm), and two carbon atoms (signals at 64.5 and 68.6 ppm) connected to OR (R=unknown). The three signals accounting for carbon atoms with sp3 hybridisation (14.7, 64.5 and 68.5 ppm) show 13C-13C coupling to one adjacent 13C atom with coupling constants of 40-50 Hz (Table 2). The signal at 122.7 ppm shows 13C couplings to two adjacent 13C neighbours (coupling constants, 74 and 50 Hz), whereas the signal at 141.5 ppm shows 13C couplings to three neighboured 13C atoms (coupling constants, 74, 43 and 43 Hz). In conjunction with the chemical shift topology, this coupling signature is indicative for a 2-methyl-2-butenyl skeleton.
[0219]HMQC and HMQC-TOCSY experiments reveal the 1H NMR chemical shifts (Table 2), as well as 13C-1H and 1H-1H spin systems (Table 3). More specifically, the 13C NMR signal at 122.7 ppm correlates to a 1H NMR signal at 5.6 ppm which is in the typical chemical shift range for H-atoms attached to CC double bonds, whereas the signal at 139.5 ppm gives no 13C-1H correlations. The signals at 64.5 and 68.6 ppm give 13C-1H correlations to 1H-signals at 4.5 and 3.9 ppm, respectively. The methyl signal at 14.7 ppm correlates to a proton signal at 1.5 ppm. In connection with 13C-13C coupling patterns (Table 2), as well as with 1H-13C long range correlations (HMBC experiment, Table 3), these data establish a 1,4-dihydroxy-2-methyl-2-butenyl system.
[0220]Starting from [3,4,5-13C3]1-deoxy-D-xylulose as feeding material three signals at 64.5, 68.6 and 122.7 ppm accounting for atoms 4, 1 and 3, respectively, of the new product are observed. It can be concluded that the carbon atoms at 1, 3 and 4 of the new product are biogenetically equivalent to the carbon atoms 3, 4 and 5 of [3,4,5-13C3]1-deoxy-D-xylulose 5-phosphate. This coupling topology is similar to the coupling pattern of 2C-methyl-D-erythritol 4-phosphate (see example 13) confirming that the new compound is derived via 2C-methyl-D-erythritol 4-phosphate.
[0221]The C-4 and C-3 13C NMR signals at 64.5 and 122.7 ppm show 13C-31P coupling of 5.5 and 8.0 Hz, respectively. These couplings indicate the presence of a phosphate or pyrophosphate group at position 4 of the 2-methyl-2-butenyl skeleton.
[0222]In line with this observation, the 1H-decoupled 31P NMR spectrum of the product displays a doublet at -9.2 (31P-31P coupling constant, 20.9 Hz) and a double-double-doublet at -10.6 ppm (31P-13C coupling constants, 5.8 and 7.4 Hz, 31P-31P coupling constant, 20.9 Hz). Without 1H-decoupling, the 31P NMR signal at -10.6 ppm is broadened whereas the signal at -9.2 ppm is not affected by 1H coupling. The chemical shifts as well as the observed coupling pattern confirm the presence of a free diphosphate moiety at position 4.
[0223]In summary, all these data establish the structure as 1-hydroxy-2-methyl-2-butenyl 4-diphosphate.
TABLE-US-00002 TABLE 2 NMR data of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate Chemical shifts, ppm Coupling constants, Hz Position 1Ha 13Cb 31Pc JPC JPP JCC 1 3.91 68.6d,e 43.0e, 5.5d, 3.5d 2 139.5d,e 74.3e, 43.3e, 43.3e 2- 1.51 14.7e 42.2e, 4.0e, 4.0e Methyl 3 5.57 122.7e 8.0d 73.9e, 49.8d, 4.0d 4 4.46 64.5d,e 5.5d 49.3d, 5.5d Pb -9.2 20.9 Pa -10.6 5.8d, 7.4d 20.9 areferenced to external trimethylsilylpropane sulfonate. breferenced to external trimethylsilylpropane sulfonate. creferenced to external 85% orthophosphoric acid. dobserved with [1,3,4-13C3]1-hydroxy-2-methyl-2-butenyl 4-diphosphate eobserved with [U-13C5]1-hydroxy-2-methyl-2-butenyl 4-diphosphate
TABLE-US-00003 TABLE 3 Correlation pattern of [1,3,4-13C3]1-hydroxy-2-methyl-2-butenyl 4- diphosphate and of [U-13C5]1-hydroxy-2-methyl-2-butenyl 4-diphosphate NMR Correlation pattern Position HMQC HMQC-TOCSY HMBC 1 1a,b 1a,b 2-methyla, 2a 2 2-methyl 2-methylb 2-methylb 3 3a,b 3a,b, 4a,b 2-methyla, 1a 4 4a,b 4a,b, 3a,b aobserved with [1,3,4-13C3]1-hydroxy-2-methyl-2-butenyl 4-diphosphate bobserved with [U-13C5]1-hydroxy-2-methyl-2-butenyl 4-diphosphate
EXAMPLE 19
Detection of Phosphorylated Metabolites of the Mevalonate-Independent Pathway
[0224]Method A) By a TLC Method
[0225]Aliquots (10 μl) of the cell-free extracts from recombinant cells prepared as described above (see example 16) are spotted on a Polygram® SIL NH--R thin layer plate (Macherey-Nagel, Duren, Germany). The TLC plate is then developed in a solvent system of n-propanol: ethyl acetate: water; 6:1:3 (v/v/v). The running time is about 4 h. The radio chromatogram is monitored and evaluated by a Phosphor Imager (Storm 860, Molecular Dynamics, USA). The Rf-values of the compounds under study are shown in Table 4.
TABLE-US-00004 TABLE 4 Rf-values of precursors and intermediates of the mevalonate-independent terpenoid pathway Chemical compound Rf-value 1-deoxy-D-xylulose 0.80 1-deoxy-D-xylulose 5-phosphate 0.5 2C-methyl-D-erythritol 4-phosphate 0.42 4-diphosphocytidyl-2C-methyl-D-erythritol 0.33 4-diphosphocytidyl-2C-methyl-D-erythritol 2-phosphate 0.27 2C-methyl-D-erythritol 2,4-cyclodiphosphate 0.47 1-hydroxy-2-methyl-2-butenyl 4-diphosphate 0.17
[0226]Method B) By a HPLC Method
[0227]Aliquots (100 μl) of the cell-free extracts from recombinant cells prepared as described above (see example 16), are analyzed by HPLC using a column of Multospher 120 RP 18-AQ-5 (4.6×250 mm, particle size 5 μm, CS-Chromatographic Service GmbH, Langerwehe, Germany) that has been equilibrated for 15 min with 10 mM tetrabutylammonium hydrogensulfate (TBAS), pH 6.0, at a flow rate of 0.75 ml min-1. After injection of the sample, the column is developed for 20 min with 10 mM TBAS, then for 60 min with a linear gradient of 0-42% (v/v) methanol in 10 mM TBAS. The effluent is monitored by a continuous-flow radio detector (Beta-RAM, Biostep GmbH, Jahnsdorf, Germany). The retention volumes of the compounds under study are shown in Table 5.
TABLE-US-00005 TABLE 5 Retention volumes of precursors and intermediates of the mevalonate- independent terpenoid pathway Retention volume Chemical compound [ml] 1-deoxy-D-xylulose 6.0 1-deoxy-D-xylulose 5-phosphate 15 2C-methyl-D-erythritol 4-phosphate 13.5 4-diphosphocytidyl-2C-methyl-D-erythritol 30.8 4-diphosphocytidyl-2C-methyl-D-erythritol 2-phosphate 41.3 2C-methyl-D-erythritol 2,4-cyclodiphosphate 31.5 1-hydroxy-2-methyl-2-butenyl 4-diphosphate 42.8
EXAMPLE 20
Purification of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate
[0228]The crude cell free extract obtained from the feeding experiment with recombinant Escherichia coli XL1-pBScyclo-pACYCgcpE using [2-14C]- and [U-13C5]1-deoxy-D-xylulose (see example 16) is lyophilized. The residue is dissolved in 600 μl of water and centrifuged for 10 min at 14,000 ppm. Aliquots of 90 μl are applied on a column of Nucleosil 10 SB (4.6×250 mm, Macherey & Nagel, Duren, Germany) which is developed with a linear gradient of 0.1-0.25 M ammonium formate in 70 ml at a flow rate of 2 ml min-1. The retention volumes for 2C-methyl-D-erythritol-2,4-cyclodiphosphate and 1-hydroxy-2-methyl-2-butenyl 4-diphosphate are 25 and 44 ml, respectively. Fractions are collected and lyophilized. NMR data of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate are identical with the data shown in example 18, Table 2.
EXAMPLE 21
Construction of a Vector Carrying the xylB, dxr, ispD, ispE, ispF and ispG Genes of Escherichia coli Capable for Transcription and Expression of D-xylulokinase, DXP Reductoisomerase, CDP-ME Synthase, CDP-ME Kinase cMEPP Synthase and 1-hydroxy-2-methyl-2-butenyl 4-diphosphate Synthase
[0229]The E. coli ORF ispG (accession no. gb AE000338) from base pair (bp) position 372 to 1204 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 μmol of the primer 5'-GCGGGAGACCGCGGGAGGAGAAATTAACCATGCATAACCAGGCTCCAATTCG-3' (SEQ ID NO:17), 10 pmol of the primer 5'-CGCTTCCCAGCGGCCGCTTATTTTTCAACCTGCTGAACG-3' (SEQ ID NO:18), 20 ng of chromosomal DNA, 2 U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0230]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 60 sec at 94° C., 60 sec at 50° C. and 90 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0231]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden).
[0232]1.4 μg of the vector pBScyclo (Example 5) and 0.8 μg of the purified PCR product are digested with SacII and NotI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0233]20 ng of the purified vector DNA and 18 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pBScyclogcpE. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pBScyclogcpE is isolated with the plasmid isolation kit from Qiagen.
[0234]The DNA insert of the plasmid pBScyclogcpE is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb AE000338). The DNA sequence of the vector construct pBScyclogcpE is shown in FIG. 15.
EXAMPLE 22
Construction of a Vector Carrying the xylB, dxr, ispD, ispE, ispF, ispG and lytB Genes of Escherichia coli Capable for Transcription and Expression of D-xylulokinase, DXP Reductoisomerase, CDP-ME Synthase, CDP-ME Kinase, cMEPP Synthase, 1-hydroxy-2-methyl-2-butenyl 4-diphosphate Synthase and LytB
[0235]The E. coli ORF lytB (accession no. gb AE005179) from base pair (bp) position 7504 to 8454 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 pmol of the primer 5'-AAATCGGAGCTCGAGGAGAAATTAACCATGCAGATCCTGTTGGCC-3' (SEQ ID NO:19), 10 pmol of the primer 5'-GCTGCTCCGCGGTTAATCGACTTCACGAATATCG-3' (SEQ ID NO:20), 20 ng of chromosomal DNA, 2. U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0236]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 45 sec at 94° C., 45 sec at 50° C. and 60 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0237]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden).
[0238]1.3 μg of the vector pBScyclogcpE (Example 21) and 0.7 μg of the purified PCR product are digested with SacI and SacII in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0239]20 ng of the purified vector DNA and 16 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pBScyclogcpElytB. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pBScyclogcpElytB is isolated with the plasmid isolation kit from Qiagen.
[0240]The DNA insert of the plasmid pBScyclogcpElytB is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb AE005179).
EXAMPLE 23
Incorporation Experiment with Recombinant Escherichia coli XL1-pBScyclogcpE Using [U-13C5]1-deoxy-D-xylulose
[0241]0.1 litre of Terrific Broth (TB) medium containing 18 mg of ampicillin are inoculated with E. coli strain Xl1-Blue harbouring the plasmid pBScyclogcpE. The cells are grown in a shaking culture at 37° C. overnight. At an optical density (600 nm) of 4.8-5.0, 30 mg (0.1 mmol) of cytidine are added. A solution containing 1.2 g of lithium lactate (12.5 mmol), 6 ml of crude [U-13C5]1-deoxy-D-xylulose (0.05 mmol) (see examples 8, 9 and 10) in 0.1 M Tris hydrochloride (pH=7.5) at a final volume of 30 ml are added continuously within 2 hours. Aliquots of 25 ml are taken at time intervals of 1 h and centrifuged for 20 min at 5,000 rpm and 4° C. The cells are washed with water containing 0.9% NaCl and centrifuged as described above. The cells are suspended in 700 μl of 20 mM NaF in D2O or in 700 μl of a mixture of methanol and D2O (6:4, v/v) containing 10 mM NaF, cooled on ice and sonified 3×10 sec with a Branson Sonifier 250 (Branson SONIC Power Company) set to 90% duty cycle output, control value of 4 output. The suspension is centrifuged at 15,000 rpm for 15 min. NMR spectra of the cell free extracts are recorded directly with a Bruker AVANCE DRX 500 spectrometer (Karlsruhe, Germany). In order to avoid degradation during work-up, the structures of the products are determined by NMR spectroscopy without further purification.
[0242]The relative amount of 1-hydroxy-2-methyl-2-butenyl 4-diphosphate to 2C-methyl-D-erythritol 2,4-cyclodiphosphate could be raised by a factor of approximately 2-3 by the addition of lithium lactate to the medium.
EXAMPLE 24
Preparation of (E)-1-hydroxy-2-methyl-2-butenyl diphosphate triammonium salt (8)
[0243]General. Chemicals are obtained from Acros Organics (Fisher Scientific GmbH, Schwerte, Germany), SIGMA-ALDRICH (Deisenhofen, Germany), MERCK (Darmstadt, Germany) and used without further purification. Solvents are used distilled and/or dried. Chromatography is performed on silica gel 60 (230-400 mesh, Fluka Riedel-de Haen, Taufkirchen, Germany), DOWEX 50 WX8 (200-400 mesh, SERVA, Heidelberg, Germany), and Cellulose (Avicel, Cellulose mikrokristallin, Merck, Darmstadt, Germany). TLC is performed on silica gel 60 F254 plastic sheets (MERCK) or cellulose F plastic sheets (MERCK), detection by anisaldeyde solution (anisaldehyde:H2SO4:HAc 0.5:1:50 v/v/v). NMR-spectra are recorded on BRUKER AMX 400, DRX 500, and AC 250 spectrometer at room temperature.
[0244]Acetonyl tetrahydropyranyl ether (12) (Hagiwara et al., 1984).
[0245]A mixture of 339 mg (1.35 mmol) of pyridinium-toluene-4-sulfonate, 9.35 ml (10.0 g, 0.135 mol) of hydroxyacetone, and 24.7 ml (22.7 g, 0.270 mol) of 3,4-dihydro-2H-pyran is stirred at room temperature for 2.5 h. Residual 3,4-dihydro-2H-pyran is removed under reduced pressure. The crude mixture is purified by FC on silicagel (hexanes/acetone 4:1, 6.5×20 cm) to yield 18.7 g (0.118 mol, 88%) of a colorless liquid.
[0246]1H NMR (CDCl3, 500 MHz) δ 4.62 (t, J=3.6 Hz, 1H), 4.22 (d, J=17.3 Hz, 1H), 4.09 (d, J=17.3 Hz, 1H), 3.83-3.79 (m, 1H), 3.51-3.47 (m, 1H), 2.15 (s, 3H), 1.87-1.49 (m, 6H); 13C NMR (CDCl3, 126 MHz) δ 206.7, 98.7, 72.3, 62.3, 30.2, 26.5, 25.2, 19.2;
[0247]MS (Cl, isobutane) m/z 159 [M+1].sup.+.
[0248](E,Z)-Ethyl-2-methyl-1-tetrahydropyranyloxy-but-2-enoate (13) (Watanabe et al., 1996).
[0249]33.0 g (94.8 mmol) of (ethoxycarbonylmethylen)-triphenylphosphorane are dissolved in 500 ml of dry toluene under nitrogen atmosphere at room temperature. Then, 10.0 g (63.2 mmol) of acetonyl tetrahydropyranyl ether 12 are added and the mixture is heated to reflux. After 39 h at this temperature the solvent is evaporated under reduced pressure to yield an orange oil. Major amounts of triphenylphosphinoxide are precipitated by the addition of 100 ml hexanes/acetone 9:1. After filtration the filtrate is concentrated and another 100 ml of hexanes/acetone 9:1 are added. The solid is filtered off and the solvent removed to yield 18 g of an orange oil that is purified by FC on silicagel (hexanes/acetone 9:1, 6.5×28 cm) to yield 12.9 g (56.5 mmol, 89%) of a mixture of (E)-13/(Z)-13=5:1.
[0250](E)-(13). 1H NMR (CDCl3, 500 MHz) δ 5.96 (q, J=1.4 Hz, 1H), 4.62 (t, J=3.5 Hz, 1H), 4.20 (dd, J=15.5 Hz, 1.3 Hz, 1H), 4.14 (q, J=7.1 Hz, 2H), 3.93 (dd, J=15.6, 1.3 Hz, 1H), 3.84-3.79 (m, 1H), 3.52-3.48 (m, 1H), 2.08 (d, J=1.4 Hz, 3H), 1.88-1.50 (m, 6H), 1.26 (t, J=7.1 Hz, 3H); 13C NMR (CDCl3, 126 MHz) δ 166.8, 154.7, 114.5, 98.0, 70.6, 62.0, 59.7, 30.3, 25.3, 19.1, 15.9, 14.3; MS (Cl, isobutane) m/z 229 [M+1].sup.+.
[0251](Z)-(13). 1H NMR (CDCl3, 500 MHz) δ 5.71 (q, J=1.4 Hz, 1H), 4.60 (t, J=3.6 Hz, 1H), 4.20 (dd, J=15.5 Hz, 1.3 Hz, 1H), 4.11 (q, J=7.1 Hz, 2H), 3.93 (dd, J=15.6, 1.3 Hz, 1H), 3.84-3.79 (m, 1H), 3.52-3.48 (m, 1H), 1.97 (d, J=1.4 Hz, 3H), 1.88-1.50 (m, 6H), 1.24 (t, J=7.1 Hz, 3H); 13C NMR (CDCl3, 126 MHz) δ 165.9, 156.8, 116.9, 98.7, 66.5, 62.3, 59.8, 30.6, 25.3, 21.9, 19.5, 14.3; MS (Cl, isobutane) m/z 229 [M+1].sup.+.
[0252](E,Z)-2-Methyl 1-tetrahydropyranyloxy-but-2-ene-4-ol (14) (Watanabe et al., 1996).
[0253]A solution of ester 13 (8.73 g, 38.2 mmol) in 100 ml of dry CH2Cl2 is cooled to -78° C. Then, 91.8 ml (91.8 mmol) of 1.0 M DIBAH in hexanes are added slowly under an atmosphere of nitrogen. The resulting solution is stirred for 3 h at -78° C. before the reaction is quenched by the addition of 1.5 ml of 1 M NaOH. After warming to room temperature the solvent is removed under reduced pressure. The resulting gummy residue is widely dissolved by adding twice 100 ml of MeOH. The resulting mixture is passed through a column of SiO2, evaporated from the solvent and then loaded on a column of SiO2/Na2SO4 that is purged with 1400 ml of MeOH. Evaporation of the solvent gives 9.5 g of a colorless liquid that is purified by FC on silica gel (hexanes/acetone 1:3, 6.5×16 cm) to yield 6.98 g (37.4 mmol, 98%) of a colorless liquid (E)-14/(Z)-14=6:1.
[0254](E)-(14). 1H NMR (CDCl3, 500 MHz) δ 5.68 (tq, J=6.6, 1.3 Hz, 1H), 4.60 (t, J=3.6 Hz, 1H), 4.20 (d, J=6.8 Hz, 2H), 4.12 (d, J=12.0 Hz, 1H), 3.87-3.82 (m, 1H), 3.85 (d, J=12.5 Hz, 1H), 3.52-3.48 (m, 1H), 1.86-1.48 (m, 6H), 1.69 (s, 3H); 13C NMR (CDCl3, 126 MHz) δ 135.7, 125.4, 97.8, 71.9, 62.1, 59.1, 30.5, 25.4, 19.4, 14.1; MS (Cl, isobutane) m/z 169 [M-H2O+1].sup.+.
[0255](Z)-(14). 1H NMR (CDCl3, 500 MHz) δ 5.64 (tq, J=6.6, 1.3 Hz, 1H), 4.63 (t, J=3.3 Hz, 1H), 4.20 (d, J=6.8 Hz, 2H), 4.15 (d, J=11.8 Hz, 1H), 3.87-3.82 (m, 1H), 3.83 (d, J=11.3 Hz, 1H), 3.52-3.48 (m, 1H), 1.86-1.48 (m, 6H), 1.79 (s, 3H); 13C NMR (CDCl3, 126 MHz) δ 136.2, 128.6, 96.6, 65.1, 61.8, 58.1, 30.3, 25.3, 21.9, 19.0; MS (Cl, isobutane) m/z 169 [M-H2O+1].sup.+.
[0256](E,Z)-4-Chloro-2-methyl 1-tetrahydropyranyloxy-but-2-en (15) (Hwang et al., 1984).
[0257]To a solution of alcohol 14 (1.00 g, 5.37 mmol) in 10 ml of dry CH2Cl2 are added 918 mg (7.52 mmol) of DMAP in 10 ml of dry CH2Cl2 and 1.23 g (6.44 mmol) of p-TsCl in 10 ml of dry CH2Cl2. The resulting solution is stirred at room temperature for 1 h. After evaporation of the solvent under reduced pressure the residue is purified by FC on silica gel (CH2Cl2, 5×20 cm) to obtain 693 mg (3.39 mmol, 63%) of a colorless liquid (E)-15/(Z)-15=6:1.
[0258](E)-(15). 1H NMR (CDCl3, 500 MHz) δ 5.77 (tq, J=8.0, 1.5 Hz, 1H), 4.64 (t, J=3.6 Hz, 1H), 4.18 (d, J=12.8 Hz, 1H), 4.15 (d, J=8.0 Hz, 2H), 3.92 (d, J=12.8 Hz, 1H), 3.90-3.86 (m, 1H), 3.59-3.52 (m, 1H), 1.92-1.52 (m, 6H), 1.77 (s, 3H); 13C NMR (CDCl3, 126 MHz) δ 138.6, 121.7, 97.8, 71.3, 62.1, 40.2, 30.5, 25.4, 19.3, 13.9; MS (Cl, isobutane) m/z 205 [M+1].sup.+.
[0259](Z)-(15). 1H NMR (CDCl3, 500 MHz) δ 5.65 (t, J=8.1 Hz, 1H), 4.61 (t, J=3.6 Hz, 1H), 4.18 (d, J=12.8 Hz, 1H), 4.15 (d, J=8.0 Hz, 2H), 3.92 (d, J=12.8 Hz, 1H), 3.90-3.86 (m, 1H), 3.59-3.52 (m, 1H), 1.92-1.52 (m, 6H), 1.86 (s, 3H); 13C NMR (CDCl3, 126 MHz) δ 138.3, 124.6, 97.5, 64.7, 62.2, 40.1, 30.5, 25.4, 21.8, 19.4; MS (Cl, isobutane) m/z 205 [M+1].sup.+.
[0260](E,Z)-2-Methyl 1-tetrahydropyranyloxy-but-2-enyl diphosphate triammonium salt (16) (Davisson et al., 1986).
[0261]To a solution of chloride 15 (260 mg, 1.27 mmol) in 1.3 ml of MeCN a solution of 1.38 g (1.52 mmol) tris(tetra-n-butylammonium) hydrogen pyrophosphate in 3.0 ml of MeCN is added slowly at room temperature, obtaining an orange-red solution. The reaction is followed by 1H-NMR, taking advantage of the up field shift of the multiplet of H-3. After 2 h the reaction is finished and the solvent removed under reduced pressure. The orange oil is dissolved in 2.5 ml of H2O and passed through a column of DOWEX 50 WX8 (2.5×3 cm) cation-exchange resin (NH4.sup.+ form) that has been equilibrated with two column volumes (40 ml) of 25 mM NH4HCO3. The column is eluted with 60 ml of 25 mM NH4HCO3. The resulting solution is lyophilized, dissolved in 5 ml of isopropanol/100 mM NH4HCO3 1:1 and loaded on a cellulose column (2×18 cm) that is eluted by isopropanol/100 mM NH4HCO3 1:1. The effluent is lyophilized obtaining 495 mg (1.25 mmol, 98%) of (E)-16/(Z)-16=6:1 as a white solid.
[0262](E)-(16). 1H NMR (D2O, 500 MHz) δ 5.52 (tq, J=6.8 Hz, 1H), 4.65 (s, 1H), 4.34 (t, J=7.0 Hz, 2H), 3.98 (d, J=12.3 Hz, 1H), 3.84 (d, J=12.1 Hz, 1H), 3.74-3.70 (m, 1H), 3.42-3.38 (m, 1H), 1.61-1.57 (m, 2H), 1.54 (s, 3H), 1.40-1.32 (m, 4H); 13C NMR (D2O, 126 MHz) δ 136.4, 123.9 (dd, J=8.0, 2.3 Hz), 98.5, 72.5, 63.2, 62.2 (d, J=5.3 Hz), 29.9, 24.5, 19.0, 13.4; 31P NMR (D2O, 101 MHz) δ -5.62 (d, J=20.9 Hz), -7.57 (d, J=20.8 Hz).
[0263](Z)-(16). 1H NMR (D2O, 500 MHz) δ 5.52 (t, J=6.8, 1H), 4.65 (s, 1H), 4.31 (t, J=7.1 Hz, 2H), 3.98 (d, J=12.3 Hz, 1H), 3.84 (d, J=12.1 Hz, 1H), 3.74-3.70 (m, 1H), 3.42-3.38 (m, 1H), 1.64 (s, 3H), 1.61-1.57 (m, 2H), 1.40-1.32 (m, 4H); 13C NMR (D2O, 126 MHz) δ 136.3, 125.8 (d, J=8.6 Hz), 98.6, 72.5, 63.2, 61.8 (d, J=5.1 Hz), 29.9, 24.5, 20.8, 19.0; 31P NMR (D2O, 101 MHz) δ -5.69 (d, J=20.8 Hz), -7.68 (d, J=20.8 Hz).
[0264](E,Z)-1-Hydroxy-2-methyl-but-2-enyl diphosphate triammonium salt (8) (Davisson et al., 1986).
[0265]268 mg (0.675 mmol) of protected pyrophosphate 16 are dissolved in 2.0 ml of D2O and the pH is adjusted to 1 by addition of 40 μl of 37% DCl in D2O. After 1 min at this pH the solution is neutralized by addition of 40 μl of 40% NaOD in D2O and an 1H NMR is measured that demonstrated 50% deprotection. The procedure is repeated until deprotection is finished and just small amounts of decomposition product are formed to get in total 7 min at pH 1. Purification is performed by loading the neutral solution that is diluted by addition of 2 ml of isopropanol/100 mM NH4HCO3 1:1 on a cellulose column (isopropanol/100 mM NH4HCO3 1:1, 2×10.5 cm) to yield 193 mg (0.616 mmol, 91%) of a white solid of (E)-8/(Z)-8=7:1.
[0266](E)-(8). 1H NMR (D2O, 500 MHz) δ 5.51 (tq, J=6.8, 1.2 Hz, 1H), 4.41 (t, J=7.2 Hz, 2H), 3.90 (s, 2H), 1.59 (s, 3H); 13C NMR (D2O, 126 MHz) δ 139.8, 120.6 (d, J=7.7 Hz), 66.5, 62.4 (d, J=5.3 Hz), 13.2; 31P NMR (D2O, 101 MHz) δ -4.48 (d, J=20.8 Hz), -7.06 (d, J=20.8 Hz).
[0267](Z)-(8). 1H NMR (D2O, 500 MHz) δ 5.49 (tm, J=6.8 Hz, 1H), 4.41 (t, J=7.3 Hz, 2H), 4.03 (s, 2H), 1.70 (s, 3H); 13C NMR (D2O, 126 MHz) δ 139.8, 123.5 (d, J=7.7 Hz), 61.7 (d, J=5.1 Hz), 59.9, 20.6; 31P NMR (D2O, 101 MHz) δ -4.48 (d, J=20.8 Hz), -7.06 (d, J=20.8 Hz).
[0268]Reagents and conditions (steps (a) to (f) in FIG. 4: 1): (a) DHP, PPTS, 25° C. (2.5 h); (b) Ph3PCHCO2Et, toluene, reflux (39 h); (c) (1) DIBAH, CH2Cl2, -78° C. (3 h), (2) 1 M NaOH/H2O; (d) p-TsCl, DMAP, CH2Cl2, 25° C. (1 h); (e) ((CH3CH2CH2CH2)4N)3HP2O7, MeCN, 25° C. (2 h); (f), HCl/H2O pH 1, 25° C. (7 min).
EXAMPLE 25
Identification of (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate
[0269]The structure of the GcpE product is further analyzed by comparison with the chemical shifts of a synthetic sample of 1-hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate.
[0270]For this purpose, [2-14C]1-hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate (0.36 μCi) is added to a cell extract obtained from bioengineered Escherichia coli cells endowed with artificial gene constructs expressing xylB, ispC, ispD, ispE, ispF and gcpE gene which are supplied with [U-13C5]-1-deoxy-D-xylulose (see example 16). The supernatant of the cell extract is purified by HPLC (Nucleosil 5 SB, 7.5×250 mm, developed with a gradient of 100 mM to 250 mM NH4HCOO, flow rate 2 ml/min, 35 min). The product is eluted at 23 min and collected. After lyophilization the residue is dissolved in D2O (pH 6) and subjected to 1H NMR analysis (FIG. 3-A). Then, 40 μl of a solution of synthetically prepared (E,Z)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate (E/Z=7:1) (D2O, pH 7) are added to the NMR sample and again analyzed by 1H NMR spectroscopy (FIG. 3-B). On the one hand, as shown in FIG. 3-B, signals accounting for (E)-1-hydroxy-2-methyl-2-butenyl are selectively increased, providing evidence that the biologically produced structure is identical with the synthetically produced one, i.e. the (E)-isomer. On the other hand, the minor (Z)-isomer raises without any correlation to signals of the biologically afforded product. FIG. 3-C shows the same effects after addition of another 40 μl of solution of the synthetically prepared (E,Z)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate.
EXAMPLE 26
Incorporation of (E)-1-hydroxy-2-methyl-2-butenyl 4-diphosphate into the Lipid Soluble Fraction of Capsicum annuum Chromoplasts
[0271]Chromoplasts are isolated by a slight modification of a method described by Camara (Camara, 1985; Camara, 1993). Pericarp of red pepper (650 g) is homogenized at 4° C. in 600 ml of 50 mM Hepes, pH 8.0, containing 1 mM DTE, 1 mM EDTA and 0.4 M sucrose (buffer A). The suspension is filtered through four layers of nylon cloth (50 μm) and centrifuged (10 min, 4,500 rpm, GSA rotor) to obtain a pellet of crude chromoplasts which is homogenized in 200 ml of buffer A. The suspension is centrifuged (10 min, 4,500 rpm, GSA rotor). The pellet is homogenized and resuspended in 3 ml of 50 mM Hepes, pH 7.6, containing 1 mM DTE. The suspension is filtered through one layer of nylon cloth (50 μm).
[0272]Reaction mixtures contain 100 mM Hepes, pH 7.6, 2 mM MnCl2, 10 mM MgCl2, 5 mM NaF, 2 mM NADP.sup.+, 1 mM NADPH, 6 mM ATP, 20 μM FAD and 2 mg of chromoplasts. 8.8 nmol of [2-14C]2C-methyl-D-erythritol 2,4-cyclodiphosphate, [2-14C]1-hydroxy-2-methyl-2-(E)-butenyl diphosphate or [2-14C]isopentenyl diphosphate (specific concentrations 15.8 μCi/μmol) are added and the mixtures are incubated at 30° C. overnight. The reaction is terminated by methylene chloride extraction. The organic phase is concentrated under a stream of nitrogen. Aliquots are spotted on silica gel plates (Polygram SIL-G, UV254, Macherey-Nagel, Duren, Germany). The plates are developed with hexane:ether=6:1 (system 1) and/or hexane:toluene=9:1 (system II), respectively. The chromatograms are monitored with a phosphor imager (Storm 860, Molecular dynamics, Sunnyvale, Calif., USA). The Rf-values of geranylgeraniol and the carotene fraction in system I are 0.35 and 0.9, respectively. The Rf-values of β-carotene, phytoene and phytofluene in system II are 0.65, 0.60 and 0.55, respectively.
[0273]The evaluation of the chromatogramms show that radioactivity can be efficiently diverted from 1-hydroxy-2-methyl-2-(E)-butenyl diphosphate into the geranylgeraniol, 3-carotene, phytoene and phytofluene fractions of C. annuum chromoplasts establishing 1-hydroxy-2-methyl-2-(E)-butenyl diphosphate as a real intermediate of the non-mevalonate pathway downstream from 2C-methyl-D-erythritol 2,4-cyclodiphosphate and upstream from isopentenyl diphosphate.
EXAMPLE 27
Construction of a Vector Carrying the ispG (gcpE) and ispH (lytB) Genes of Escherichia coli Capable for Transcription and Expression Thereof
[0274]The E. coli ORF ispH (lytB) (accession no. gb AE000113) from base pair (bp) position 5618 to 6568 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 pmol of the primer 5'-GCTTGCGTCGACGAGGAGAAATTAACCATGCAGATCCTGTTGGCCACC-3' (SEQ ID NO:21), 10 pmol of the primer 5'-GCTGCTCGGCCGTTAATCGACTTCACGAATATCG-3' (SEQ ID NO:22), 20 ng of chromosomal DNA, 2 U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0275]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 45 sec at 94° C., 45 sec at 50° C. and 60 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0276]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden).
[0277]2.4 μg of the vector pACYC184 (Chang and Cohen 1978, NEB) and 0.7 μg of the purified PCR product are digested with SalI and EagI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0278]20 ng of the purified vector DNA and 18 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pACYClytB. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pACYClytB is isolated with the plasmid isolation kit from Qiagen.
[0279]The DNA insert of the plasmid pACYClytB is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb AE000113).
[0280]The E. coli ORF ispG (gcpE) (accession no. gb AE000338) from base pair (bp) position 372 to 1204 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 pmol of the primer 5'-CGTACCGGATCCGAGGAGAAATTAACCATGCATAACCAGGCTCCAATTC-3' (SEQ ID NO:23), 10 pmol of the primer 5'-CCCATCGTCGACTTATTTTTCAACCTGCTGAACGTC-3' (SEQ ID NO:24), 20 ng of chromosomal DNA, 2 U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0281]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 60 sec at 94° C., 60 sec at 50° C. and 90 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0282]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden).
[0283]2.0 μg of the vector pACYClytB and 0.9 μg of the purified PCR product are digested with BamHI and SalI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0284]20 ng of the purified vector DNA and 23 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pACYClytBgcpE. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pACYClytBgcpE is isolated with the plasmid isolation kit from Qiagen.
[0285]The DNA insert of the plasmid pACYClytBgcpE is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb AE000338).
[0286]The DNA sequence of the vector construct pACYClytBgcpE is shown in FIG. 16.
[0287]The DNA and corresponding amino acid sequence of ispH (lytB) from Escherichia coli is shown in FIG. 17.
EXAMPLE 28
Construction of a Vector Carrying the xylB, dxr, ispD, ispE, ispF, ispG and ispH Genes of Escherichia coli capable for Transcription and Expression of D-xylulokinase, DXP Reductoisomerase, CDP-ME Synthase, CDP-ME Kinase cMEPP Synthase, 1-hydroxy-2-methyl-2-butenyl 4-diphosphate Synthase and IPP/DMAPP Synthase
[0288]The E. coli ORFs ispG (formerly gcpE) and ispH (formerly lytB) are amplified by PCR using the plasmid pACYClytBgcpE (see example 27) as template. The reaction mixture contains 10 pmol of the primer 5'-GCGGGAGACCGCGGGAGGAGAAATTAACCATGCATAACCAGGCTCCAATTCAA CG-3' (SEQ ID NO:25), 10 pmol of the primer 5'-AGGCTGGCGGCCGCTTAATCGACTTCACGAATATCG-3' (SEQ ID NO:26), 2 ng of pACYCgcpElytB DNA, 2 U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0289]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 60 sec at 94° C., 60 sec at 50° C. and 150 sec at 72° C. followed. After further incubation for 20 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0290]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden).
[0291]1.7 μg of the vector pBScyclo (Example 5) and 1.3 μg of the purified PCR product are digested with SacII and NotI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0292]22 ng of the purified vector DNA and 19 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pBScyclogcpElytB2. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pBScyclogcpElytB2 is isolated with the plasmid isolation kit from Qiagen.
[0293]The DNA insert of the plasmid pBScyclogcpElytB2 is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. The DNA sequence of the vector construct pBScyclogcpElytB2 is shown in FIG. 18.
EXAMPLE 29
Incorporation Experiment with Recombinant Escherichia coli XL1-pBScyclogcpElytB2 Using [U-13C5]1-deoxy-D-xylulose
[0294]0.1 litre of Terrific Broth (TB) medium containing 18 mg of ampicillin are inoculated with E. Coli strain Xl1-Blue harbouring the plasmid pBScyclogcpElytB2. The cells are grown in a shaking culture at 37° C. for overnight. At an optical density (600 nm) of 1.3-1.7 a solution containing 2.4 g of lithium lactate (25 mmol), 10 ml of crude [U-13C5]1-deoxy-D-xylulose (0.05 mmol) (see example 8) at a final volume of 30 ml (pH=7.4) are added continuously within 2 hours. Aliquots of 40 ml are taken at time intervals of 30 minutes and centrifuged for 20 min at 5,000 rpm and 4° C. The cells are washed with water containing 0.9% NaCl and centrifuged as described above. The cells are suspended in 700 ml of a mixture of methanol and D2O (6:4, v/v) containing 10 mM NaF, cooled on ice and sonified 3×10 sec with a Branson Sonifier 250 (Branson SONIC Power Company) set to 90% duty cycle output, control value of 4 output. The suspension is centrifuged at 15,000 rpm for 15 min. NMR spectra of the cell free extracts are recorded directly with a Bruker AVANCE DRX 500 spectrometer (Karlsruhe, Germany). In order to avoid degradation during work-up, the structures of the products are determined by NMR spectroscopy without further purification.
EXAMPLE 30
Structure Determination of isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP)
[0295]The 1H-decoupled 13C NMR spectrum using [U-13C5]1-deoxy-D-xylulose as starting material (see examples 8 and 30) displays five intense 1C-13C coupled signals belonging to 2C-methyl-D-erythritol 2,4-cyclodiphosphate (Herz et al., 2000) and five 13C-13C coupled signals with low intensities belonging to 1-hydroxy-2-methyl-2-butenyl 4-diphosphate (see example 18) (100:3 ratio for the 2-methyl 13C NMR signal intensities of 2C-methyl-D-erythritol 2,4-cyclodiphosphate and 1-hydroxy-2-methyl-2-butenyl 4-diphosphate, respectively).
[0296]In addition a set of, five 13C-13C coupled signals at 21.6 (doublet), 37.8 (triplet), 64.1 (doublet), 111.6 (doublet), and 143.3 ppm (doublet of triplets) (unknown metabolite A) accompanied by signals at 21.1 (doublet), 39.6 (triplet), 59.3 (doublet), 111.8 (doublet), and 143.2 ppm (doublet of triplets) (unknown metabolite B) is detected. The ratio of the 2-methyl signal of 2C-methyl-D-erythritol 2,4-cyclodiphosphate and the putative methyl signals of the unknown compounds at 21.6 ppm (metabolite A) and 21.1 ppm (metabolite B) is 100:24:4, respectively.
[0297]Moreover, 13C coupled signals with low intensities belonging to another unknown compound (metabolite C) at 17.1 (doublet), 24.9 (doublet), 62.7 (doublet), 119.6 (double-doublet) and 139.4 ppm (multiplet) are detected. The ratio of the intensities of the putative methyl signals at 21.6 (metabolite A), 17.1 and 24.9 (metabolite C) is 100:13:13, respectively.
[0298]The 31P NMR spectrum of the reaction mixture is characterized by intense signals for 2C-methyl-D-erythritol 2,4-cyclodiphosphate (Herz et al., 2000). Furthermore, 31P31P coupled broadened signals are observed at a chemical shift range typical for organic diphosphates (-6 to -13 ppm, 31P31P coupling constants, 20 Hz).
[0299]Metabolite A:
[0300]The signals of metabolite A at 111.6 and 143.3 ppm are conducive of a double bond motif, and the signals at 64.1, 37.8 and 21.6 ppm reflect three aliphatic carbon atoms one of which (signal at 64.1 ppm) appears to be connected to OH or OR (R=unknown).
[0301]Additional information about the structure of the unknown metabolite A can be gleaned from the 13C coupling pattern. Three of the 13C NMR signals (21.6, 64.1 and 111.6 ppm) are split into doublets indicating three 13C atoms each connected to only one 13C-labelled neighbour, one signal (37.8 ppm) displays a pseudo-triplet signature indicating a 13C atom with two adjacent 13C atoms, and one signal (143.3 ppm) is split into a doublet of triplets indicating a 13C atom with three 13C connections. In conjunction with the chemical shifts, this connectivity pattern establish metabolite A as an isopentenyl derivative.
[0302]The complex signature for the signal at 143.3 ppm deserves a more detailed analysis. The large coupling (71 Hz) is typical for 13C13C couplings between carbon atoms involved in carbon-carbon double bonds. A 71 Hz coupling is also found for the doublet signal at 111.6 ppm representing the second carbon of the double bond. Due to the coupling pattern and the chemical shifts the presence of an exo-methylene function is obvious. The two additional 13C couplings found in the triplet substructure of the signal at 143.3 ppm are both 41 Hz, and establish the respective carbon as the branching point of the structure.
[0303]HMQC experiments reveal the 1H NMR chemical shifts, as well as 13C-1H and 1-1H spin systems. More specifically, the 13C NMR signal at 111.6 ppm correlates to a 1H NMR signal at 4.73 ppm, whereas the signal at 143.3 ppm gives no 13C-1H correlation. The signals at 64.1, 37.8, and 21.6 ppm give 13C-1H correlations to 1H-signals at 4.00, 2.31, and 1.68 ppm, respectively. As shown by HMQC-TOCSY experiments, the proton signals at 2.31 and 4.00 are coupled, whereas the signals at 4.73 and 1.68 ppm are found as singlets in the HMQC-TOCSY experiment. The observed 1H NMR chemical shifts in combination with the coupling patterns demonstrate that metabolite A is an isopentenyl derivative with a single bonded heteroatom (most plausibly 0) at position 1.
[0304]The 31C and 1H chemical shifts of an authentic sample of isopentenyl diphosphate (IPP, measured in the same solvent mixture) are identical to the chemical shifts assigned to metabolite A. Thus, metabolite A is identified as [U-13C5]IPP.
[0305]Metabolite B:
[0306]As noted above, the coupling and correlation pattern of metabolite B observed in the 13C NMR signals, as well as in the HMQC and HMQC-TOCSY spectra, is virtually the same as for metabolite A (IPP) suggesting that the carbon connectivities of metabolite B and IPP are identical. As the most significant difference between the NMR data of metabolite B and IPP the 13C NMR chemical shift of one doublet signal for metabolite B (59.3 ppm) corresponding to the C-1 signal of IPP (64.1 ppm) is upfield shifted by 4.9 ppm. This suggests that a phosphate moiety is missing at C-1 in metabolite B. Therefore, metabolite B is assigned as [U-13C5]isopentene-1-ol. Presumably, isopentene-1-ol is formed from IPP by the catalytic action of pyrophosphatases and phosphatases present in the experimental system.
[0307]Metabolite C:
[0308]As described above for metabolite A (IPP), the structure of metabolite C is assigned by NMR analysis. The 13C coupling pattern of the signals attributed to metabolite C (three doublets, one double-doublet, one multiplet) suggests that the compound is another isopentane derivative. The chemical shifts observed for the double-doublet (119.6 ppm) and the multiplet (139.4 ppm) show that a carbon-carbon double bond connects C-2 (coupled to two 13C neighbours) and C-3 (coupled to three 13C neighbours) of the molecule.
[0309]The 1H NMR chemical shifts of metabolite C are revealed by HMQC and HMQC-TOCSY experiments showing two singlets at 1.75 and 1.71 ppm, and a spin system comprising signals at 5.43 and 4.45 ppm. In conjunction with the chemical shifts, this correlation pattern shows that metabolite C is a dimethylallyl derivative.
[0310]The 13C and 1H NMR chemical shifts of an authentic sample of dimethylallyl diphosphate (DMAPP) are identical to the chemical shifts of the signals attributed to metabolite C. This leaves no doubt that metabolite C is [U-13C5]dimethylallyl diphosphate (DMAPP).
[0311]The NMR data of metabolite A (IPP) and metabolite C (DMAPP) are summarized in Tables 6 and 7.
TABLE-US-00006 TABLE 6 NMR data of isopentenyl diphosphate (IPP) Chemical shifts, ppm Coupling constants, Hz Position 1Ha 13Cb 31Pc JPC JHH JPP JPH JCCd 1 4.00 64.1 4.9 6.6 6.6 34 2 2.31 37.8 8.0 6.7 40, 40 3 143.3 71, 41, 41 4 4.73 111.6 71 5 1.68 21.6 41 P -7.8 nd P -11.9 19.5 areferenced to external trimethylsilylpropane sulfonate. breferenced to external trimethylsilylpropane sulfonate. creferenced to external 85% orthophosphoric acid. dobserved with [U-13C5]IPP
TABLE-US-00007 TABLE 7 NMR data of dimethylallyl diphosphate (DMAPP) Chemical shifts, ppm Coupling constants, Hz Position 1Ha 13Cb 31Pc JPC JHH JPP JPH JCCd 1 4.45 62.7 3.6 6.6 6.6 47 2 5.43 119.6 9.0 7.2 75, 48 3 139.4 nd 4 1.75 24.9 42 5 1.71 17.1 41 P -9.1 21.7 P -6.4 21.5 areferenced to external trimethylsilylpropane sulfonate. breferenced to external trimethylsilylpropane sulfonate. creferenced to external 85% orthophosphoric acid. dobserved with [U-13C5]DMAPP
EXAMPLE 31
Cloning of the ispG Gene (Fragment) from Arabidopsis thaliana
[0312]RNA is isolated from 1 g of 2 weeks old Arabidopsis thaliana var. Columbia plants (stems and leafs) by published procedures (Logemann et al. 1987).
[0313]A mixture containing 2.75 μg RNA, 50 nmol dNTP's, 1 μg random hexameric primer, 1 μg T15-primer and 20% first strand 5× buffer (Promega) in a total volume of 50 μl is incubated for 5 min. at 95° C., cooled on ice and 500 U M-MLV reverse transcriptase (Promega) are added. The mixture is incubated for 1 h at 42° C. After incubation at 92° C. for 5 min, RNase A (20 U) and RNase H (2 U) are added and the mixture is incubated for 30 min. at 37° C.
[0314]The resulting cDNA (1 μl of this mixture) is used for the amplification of ispG by PCR.
[0315]The A. thaliana ORF ispG (accession no. dbj AB005246) without the coding region for the putative leader sequence from basepair (bp) position 2889 to 6476 is amplified by PCR using cDNA from A. thaliana as template. The reaction mixture contains 25 pmol of primer CCTGCATCCGAAGGAAGCCC (SEQ ID NO:27), 25 μmol of primer CAGTTTTCAAAGAATGGCCC (SEQ ID NO:28), 1 μl of cDNA, 2 U of Taq DNA polymerase (Eurogentec, Seraing, Belgium) and 20 nmol of dNTPs in a total volume of 100 μl in 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0316]The mixture is denaturated for 3 min at 95° C. Then 40 PCR cycles for 60 sec at 94° C., 60 sec at 50° C. and 90 sec at 72° C. followed. After further incubation for 20 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis. The PCR amplificate is purified with the PCR purification kit from Qiagen. 1.7 μg of purified PCR product are obtained.
[0317]The PCR amplificate is used as template for a second PCR reaction. The reaction mixture contains 25 pmol of primer TGAATCAGGATCCMGACGGTGAGAAGG (SEQ ID NO:29), 25 pmol of primer TCCGTTTGGTACCCTACTCATCAGCCACGG (SEQ ID NO:30), 2 μl of the first PCR amplification, 2 U of Taq DNA polymerase (Eurogentec, Seraing, Belgium) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0318]The mixture is denaturated for 3 min at 95° C. Then 40 PCR cycles for 60 sec at 94° C., 60 sec at 50° C. and 90 sec at 72° C. follow. After further incubation for 20 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0319]The PCR amplificate is purified with PCR purification kit from Qiagen. 1.4 μg of purified PCR product are obtained. 2.0 μg of the vector pQE30 and 1.4 μg of the purified PCR product are digested with BamHI and KpnI in order to produce cohesive ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0320]20 ng of vector DNA and 12 ng of PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pQEgcpEara. The ligation mixture is incubated over night at 4° C. 2 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue and M15-[pREP4] (Zamenhof et. al., 1972) cells. The plasmid pQEgcpEara is isolated as described above. 7 μg of plasmid DNA are obtained.
[0321]The DNA insert of the plasmid pQEgcpEara is sequenced as described above. The DNA sequence is found not to be identical with the sequence in the database (accession no. dbj AB005246, see FIG. 19).
EXAMPLE 32
Screening of IspG (GcpE) Enzyme Activity
[0322]0.2 g cells of XL1-pACYClytBgcpE are suspended in 1 ml 50 mM Tris hydrochloride, pH 7.4 and 2 mM DTT, cooled on ice and sonified 3×7 sec with a Branson Sonifier 250 (Branson SONIC Power Company) set to 80% duty cycle output, control value of 4 output. The suspension is centrifuged at 14000 rpm for 15 minutes. The supernatant is used as crude cell extract in assays described as follows.
[0323]The assay mixture contains 100 mM Tris hydrochloride, pH 7.4, 1.2 mM dithiothreitol, 10 mM NaF, 1 mM CoCl2, 2 mM NADH, 20 mM (18 μCi mol-1) [2-14Cquadrature2C-methyl-erythritol 2,4-cyclodiphophate, 0.5 mM pamidronate and 100 μl crude cell extract of XL1-pACYClytBgcpE in a total volume of 150 μl. The mixture is incubated for 10 to 45 min at 37° C. and cooled on ice. 10 μl of 30% (g/v) trichloroacetic acid are added and the mixture is neutralized with 20 μl of 1 M NaOH.
[0324]The mixture is centrifuged at 14.000 rpm for 10 minutes. Aliquotes of 130 μl of the supernatant are analyzed by reversed-phase ion-pair HPLC using a column of Multospher 120 RP 18-AQ-5 (4.6×250 mm, CS-Chromatographie Service GmbH, Langerwehe, Germany). The column is developed with a linear gradient of 7-21% (v/v) methanol in 20 ml of 10 mM tetra-n-butylammonium hydrogen phosphate, pH 6.0 at a flow rate of 1 ml min-1 and further with a linear gradient of 21-49% (v/v) methanol in 15 ml of 10 mM tetra-n-butylammonium hydrogen phosphate, pH 6.0. After washing the column with 49% (v/v) methanol in 5 ml of 10 mM tetra-n-butylammonium hydrogen phosphate, pH 6.0, the column is equilibrated with 7% (v/v) methanol in 20 ml of 10 mM tetra-n-butylammonium hydrogen phosphate, pH 6.0. The effluent is monitored by a continuous-flow radio detector (Beta-RAM, Biostep GmbH, Jahnsdorf, Germany). The retention volumes of 2C-methyl-erythritol 2,4-cyclodiphophate, 1-hydroxy-2-methyl-2-(E)-butenyl-4-diphosphate, DMAPP/IPP are 18, 24 and 39 ml respectively.
[0325]After 10 minutes of incubation, about 13% of 2C-methyl-erythritol 2,4-cyclodiphophate have been converted into 1-hydroxy-2-methyl-2-(E)-butenyl-4-diphosphate (5%) and into DMAPP/IPP (8%), respectively.
[0326]After 45 min, no 1-hydroxy-2-methyl-2-(E)-butenyl-4-diphosphate, but about 21% of DMAPP/IPP was found in the assay mixture.
EXAMPLE 33
Screening of IspH (LytB) Activity
[0327]Assay mixtures contain 100 mM Tris hydrochloride, pH 7.4, 1.2 mM DTT, 10 mM NaF, 0.5 mM NADH, 60 μM FAD, 0.004 μM (18 μCi μmol-1) [2-14C]1-hydroxy-2-methyl-2-(E)-butenyl-4-diphosphate, 0.5 mM pamidronate (Dunford et al., 2001) and 20 μl of crude cell extract of M15-pMALlytB cells (prepared as described in example 2) in a total volume of 150 μl. The mixture is incubated for 30 min at 37° C. The reaction is terminated by cooling on ice, addition of 10 μl of 30% (g/v) trichloroacetic acid and immediate neutralization with 20 μl 1 M sodium hydroxide. The mixtures are centrifuged and aliquots (130 μl) of the supernatant are analyzed by reversed-phase ion-pair HPLC using a column of Multospher 120 RP 18-AQ-5 (4.6×250 mm, CS-Chromatographie Service GmbH, Langerwehe, Germany) analyzed by reversed-phase ion-pair HPLC using a column of Multospher 120 RP 18-AQ-5 (4.6×250 mm, CS-Chromatographie Service GmbH, Langerwehe, Germany). The column is developed with a linear gradient of 7-21% (v/v) methanol in 20 ml of 10 mM tetra-n-butylammonium hydrogen phosphate, pH 6.0 at a flow rate of 1 ml min-1 and further with a linear gradient of 21-49% (v/v) methanol in 15 ml of 10 mM tetra-n-butylammonium hydrogen phosphate, pH 6.0. After washing the column with 49% (v/v) methanol in 5 ml of 10 mM tetra-n-butylammonium hydrogen phosphate, pH 6.0, the column is equilibrated with 7% (v/v) methanol in 20 ml of 10 mM tetra-n-butylammonium hydrogen phosphate, pH 6.0. The effluent is monitored by a continuous-flow radio detector (Beta-RAM, Biostep GmbH, Jahnsdorf, Germany). Under standard assay conditions, the HPLC peak corresponding to the substrate 1-hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate is completely diminished, whereas two new peaks corresponding to DMAPP and IPP appear, when crude cell extract of E. coli M15-pMALlytB cells is used as protein source. No conversion of 1-hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate into DMAPP and IPP can be observed, when crude cell extract of E. coli wild-type is used as protein source. This findings clearly show that the FAD and NADH- or NADPH-dependent conversion of 1-hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate into DMAPP and IPP is catalyzed by the recombinant LytB protein. The addition of pamidronate in the assay mixtures prevents a further metabolization of IPP and DMAPP by highly active prenyl transferases present in crude E. coli extracts and affects therefore the complete conversion of 1-hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate into DMAPP and IPP.
EXAMPLE 34
Construction of a Vector Carrying the dxs, xylB and ispC Genes Capable for the Transcription and Expression Thereof
[0328]The B. subtilis ORF dxs (accession no. dbj D84432) from base pair (bp) position 193991 to 195892 is amplified by PCR using pBSDXSBACSU plasmid DNA as template (see patent application PCT/EP00/07548). The reaction mixture contains 10 pmol of the primer 5'-GGCGACTCGCGAGAGGAGAAATTAACCATGGATCTTTTATCAATACAGGACC-3' (SEQ ID NO:31), 10 pmol of the primer 5'-GGCACCCGGCCGTCATGATCCAATTCCTTTTGTGTG-3' (SEQ ID NO:32), 20 ng DNA of pBSDXSBACSU plasmid, 2 U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0329]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 60 sec at 94° C., 60 sec at 50° C. and 120 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0330]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden).
[0331]2.4 μg of the vector pACYC184 (Chang and Cohen 1978, NEB) and 1.8 μg of the purified PCR product are digested with NruI and EagI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0332]20 ng of the purified vector DNA and 19 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pACYCdxs. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pACYCdxs is isolated with the plasmid isolation kit from Qiagen.
[0333]The DNA insert of the plasmid pACYCdxs is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (dbj D84432).
[0334]2.0 μg of the vector pACYCdxs and 8 μg of the vector pBScyclo (see example XXx) are digested with EagI and SalI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0335]20 ng of the digested and purified pACYCdxs vector DNA and 30 ng of a by DNA electrophoresis separated and purified 2.7 kb EagI/SalI fragment (containing the ORFs xylB and ispC from E. coli) are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pACYCdxsxylBispC. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pACYCdxsxylBispC is isolated with the plasmid isolation kit from Qiagen.
[0336]The DNA insert of the plasmid pACYCdxsxylBispC is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions.
EXAMPLE 35
Construction of a Vectors Carrying the dxs, xylB, ispC, and ispG and Optionally ispH Genes Capable for the Transcription and Expression Thereof
[0337]The E. coli ORF ispH (lytB) (accession no. gb AE000113) from base pair (bp) position 5618 to 6568 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 pmol of the primer 5'-GCTTGCGTCGACGAGGAGAAATTAACCATGCAGATCCTGTTGGCCACC-3' (SEQ ID NO:33), 10 pmol of the primer 5'-GCTGCTCTCGAGTTAATCGACTTCACGAATATCG-3' (SEQ ID NO:34), 20 ng of chromosomal DNA, 2 U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0338]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 45 sec at 94° C., 45 sec at 50° C. and 60 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0339]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden).
[0340]2.5 μg of the vector pACYCdxsxylBispC (see example 34) are linearized with SalI and 0.9 μg of the purified PCR product are digested with SalI and XhoI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0341]15 ng of the purified vector DNA and 18 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pACYCdxsxylBispClytB. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pACYCdxsxylBispClytB is isolated with the plasmid isolation kit from Qiagen.
[0342]The DNA insert of the plasmid pACYCdxsxylBispClytB is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb AE000113).
[0343]The E. coli ORF ispG (gcpE) (accession no. gb AE000338) from base pair (bp) position 372 to 1204 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 pmol of the primer 5'-GGTCGAGTCGACGAGGAGAAATTAACCATGCATAACCAGGCTCCAATTC-3' (SEQ ID NO:35), 10 pmol of the primer 5'-CCCATCCTCGAGTTATTTTTCAACCTGCTGAACGTC-3' (SEQ ID NO:36), 20 ng of chromosomal DNA, 2 U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0344]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 60 sec at 94° C., 60 sec at 50° C. and 90 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0345]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden).
[0346]Each 2.0 μg of the vectors pACYCdxsxylBispC (see example 34) and pACYCdxsxylBispClytB (see above) are linearized with SalI and 1.1 μg of the purified PCR product are digested with SalI and XhoI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0347]18 ng of the purified vector DNAs and 23 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmids pACYCdxsxylBispCgcpE and pACYCdxsxylBispClytBgcpE. The ligation mixtures are incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmids pACYCdxsxylBispCgcpE and pACYCdxsxylBispClytBgcpE are isolated with the plasmid isolation kit from Qiagen.
[0348]The DNA inserts of the plasmids pACYCdxsxylBispCgcpE and pACYCdxsxylBispClytBgcpE are sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. They are identical with the DNA sequence of the database entry (gb AE000338).
EXAMPLE 36
Incorporation Experiment with Recombinant Escherichia coli XL1-pACYCdxsxylBispCgcpE Using [U-13C6]Glucose
[0349]0.2 litre of Terrific Broth (TB) medium containing 5 mg of chloramphenicol are inoculated with E. coli strain XL1-Blue harbouring the plasmid pACYCdxsxylBispCgcpE. The cells are grown in a shaking culture at 37° C. overnight. At an optical density (600 nm) of 1.7-2.4 a solution containing 1 g of lithium lactate (10 mmol), 200 mg [U-13C6]glucose (1.1 mmol) at a final volume of 24 ml (pH=7.4) are added continuously within 2 hours. Then, after 1 hour an aliquot of 40 ml was taken and centrifuged for 20 min at 5,000 rpm and 4° C. The cells are washed with water containing 0.9% NaCl and centrifuged as described above. The cells are suspended in 700 μl of a mixture of methanol-d4 and D2O (6:4, v/v) containing 10 mM NaF, cooled on ice and sonified 3×10 sec with a Branson Sonifier 250 (Branson SONIC Power Company) set to 90% duty cycle output, control value of 4 output. The suspension is centrifuged at 15,000 rpm for 15 min. NMR spectra of the cell free extracts are recorded directly with a Bruker AVANCE DRX 500 spectrometer (Karlsruhe, Germany). In order to avoid degradation during work-up, the structures of the products are determined by NMR spectroscopy without further purification.
[0350]The 13C-NMR spectra showed signals accounting for 1-hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate (cf. Tables 2 and 4, example 18) as major product. A formation of 2C-methyl-D-erythritol 2,4-cylodiphosphate could not be detected.
EXAMPLE 37
Incorporation Experiment with Recombinant Escherichia coli XL1-pACYCdxsxylBispClytBgcpE Using Glucose
[0351]Example 36 can be carried out with recombinant Escherichia coli XL1-pACYCdxsxylBispClytBgcpE using glucose for converting glucose to isopentenyl diphosphate and/or dimethylallyl diphosphate.
EXAMPLE 38
Cloning of the ispG Gene of Escherichia coli and Expression as Maltose Binding Fusion Protein (MBP-IspG)
[0352]The E. coli ORF ispG (gcpE) (accession no. gb AE000338) from base pair (bp) position 372 to 1204 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 pmol of the primer 5'-GAACCGGAATTCATGCATAACCAGGCTCCAATTC-3' (SEQ ID NO:37), 10 pmol of the primer 5'-CGAGGCGGATCCCATCACG-3' (SEQ ID NO:38), 20 ng of chromosomal DNA, 2 U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0353]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 60-sec at 94° C., 60 sec at 50° C. and 90 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0354]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden).
[0355]2.2 μg of the vector pMAL-C2 (NEB) and 0.8 μg of the purified PCR product are digested with EcoRI and BamHI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0356]20 ng of the purified vector DNA and 15 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pMALgcpE. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pMALgcpE is isolated with the plasmid isolation kit from Qiagen.
[0357]The DNA insert of the plasmid pMALgcpE is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb AE000338).
EXAMPLE 39
Cloning of the ispH Gene of Escherichia coli and Expression as Maltose Binding Fusion Protein (MBP-IspH)
[0358]The E. coli ORF ispH (lytB) (accession no. gb AE000113) from base pair (bp) position 5618 to 6568 is amplified by PCR using chromosomal E. coli DNA as template. The reaction mixture contains 10 pmol of the primer 5'-TGGAGGGGATCCATGCAGATCCTGTTGGCCACC-3' (SEQ ID NO:39), 10 pmol of the primer 5'-GCATTTCTGCAGAACTTAGGC-3' (SEQ ID NO:40), 20 ng of chromosomal DNA, 2 U of Taq DNA polymerase (Eurogentec) and 20 nmol of dNTPs in a total volume of 100 μl containing 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-hydrochloride, pH 8.8 and 0.1% (w/w) Triton X-100.
[0359]The mixture is denaturated for 3 min at 94° C. Then 30 PCR cycles for 45 sec at 94° C., 45 sec at 50° C. and 60 sec at 72° C. followed. After further incubation for 10 min at 72° C., the mixture is cooled to 4° C. An aliquot of 2 μl is subjected to agarose gel electrophoresis.
[0360]The PCR amplificate is purified with the PCR purification kit from Qiagen (Hilden).
[0361]2.2 μg of the vector pMAL-C2 (NEB) and 0.7 μg of the purified PCR product are digested with BamHI and PstI in order to produce DNA fragments with overlapping ends. The restriction mixtures are prepared according to the conditions supplied by the customer (NEB) and are incubated for 3 h at 37° C. Digested vector DNA and PCR product are purified using the PCR purification kit from Qiagen.
[0362]20 ng of the purified vector DNA and 14 ng of the purified PCR product are ligated together with 1 U of T4-Ligase (Gibco), 2 μl of T4-Ligase buffer (Gibco) in a total volume of 10 μl, yielding the plasmid pMALlytB. The ligation mixture is incubated for 2 h at 25° C. 1 μl of the ligation mixture is transformed into electrocompetent E. coli XL1-Blue cells. The plasmid pMallytB is isolated with the plasmid isolation kit from Qiagen.
[0363]The DNA insert of the plasmid pMALlytB is sequenced by the automated dideoxynucleotide method using an ABI Prism 377® DNA sequencer from Perkin Elmer with the ABI Prism® Sequencing Analysis Software from Applied Biosystems Divisions. It is identical with the DNA sequence of the database entry (gb AE000113).
EXAMPLE 40
Preparation and Purification of Recombinant IspG Maltose Binding Fusion Protein (MRP-IspG)
[0364]0.5 liter of Luria Bertani (LB) medium containing 90 mg of ampicillin are inoculated with 10 ml of an overnight culture of E. coli strain XL1-Blue harboring plasmid pMALgcpE. The culture is grown in a shaking culture at 37° C. At an optical density (600 nm) of 0.7, the culture is induced with 2 mM IPTG. The culture is grown for further 5 h. The cells are harvested by centrifugation for 20 min at 5,000 rpm and 4° C. The cells are washed with 20 mM Tris hydrochloride pH 7.4, centrifuged as above and frozen at -20° C. for storage.
[0365]2 g of the cells are thawed in 20 ml of 20 mM Tris hydrochloride pH 7.4, 0.2 M sodium chloride and 0.02% (g/v) sodium acid (buffer A) in the presence of 1 mg ml-1 lysozyme and 100 μg ml-1 DNaseI. The mixture is incubated at 37° C. for 30 min, cooled on ice and sonified 6×10 sec with a Branson Sonifier 250 (Branson SONIC Power Company) set to 70% duty cycle output, control value of 4 output. The suspension is centrifuged at 15,000 rpm at 4° C. for 30 min. The cell free extract is applied on a column of amylose resin FF (column volume 25 ml, NEB) previously equilibrated with buffer A at a flowrate of 2 ml min-1. The column is washed with 130 ml of buffer A. MRP-IspG is eluted with a linear gradient of 0-10 mM maltose in buffer A. MRP-IspG containing fractions are combined according to SDS-PAGE and dialyzed overnight against 100 mM Tris hydrochloride pH 7.4. The homogeneity of MRP-IspG is judged by SDS-PAGE. One band at 84 kDa is visible, which is in line with the calculated molecular mass. The yield of pure MRP-IspG is 9 mg.
EXAMPLE 41
Preparation and Purification of Recombinant IspH Maltose Binding Fusion Protein (MRP-IspH)
[0366]0.5 liter of Luria Bertani (LB) medium containing 90 mg of ampicillin are inoculated with 10 ml of an overnight culture of E. coli strain XL1-Blue harboring plasmid pMALlytB. The culture is grown in a shaking culture at 37° C. At an optical density (600 nm) of 0.7, the culture is induced with 2 mM IPTG. The culture is grown for further 5 h. The cells are harvested by centrifugation for 20 min at 5,000 rpm and 4° C. The cells are washed with 20 mM Tris hydrochloride pH 7.4, centrifuged as above and frozen at -20° C. for storage.
[0367]2 g of the cells are thawed in 20 ml of 20 mM Tris hydrochloride pH 7.4, 0.2 M sodium chloride and 0.02% (g/v) sodium acid (buffer A) in the presence of 1 mg ml-1 1 lysozyme and 100 μg ml-1 DNaseI. The mixture is incubated at 37° C. for 30 min, cooled on ice and sonified 6×10 sec with a Branson Sonifier 250 (Branson SONIC Power Company) set to 70% duty cycle output, control value of 4 output. The suspension is centrifuged at 15,000 rpm at 4° C. for 30 min. The cell free extract is applied on a column of amylose resin FF (column volume 25 ml, NEB) previously equilibrated with buffer A at a flowrate of 2 ml min-1. The column is washed with 130 ml of buffer A. MRP-IspH is eluted with a linear gradient of 0-10 mM maltose in buffer A. MRP-IspH containing fractions are combined according to SDS-PAGE and dialyzed overnight against 100 mM Tris hydrochloride pH 7.4. The homogeneity of MRP-IspH is judged by SDS-PAGE. One band at 78 kDa is visible, which is in line with the calculated molecular mass. The yield of pure MRP-IspH is 14 mg.
EXAMPLE 42
Synthesis of 1-hydroxy-2-methyl-but-2-enyl-4-diphosphate (see FIG. 7)
[0368]4-Chloro-2-methyl-2-buten-1-al (Choi et al. (1999) J. Org. Chem. 64, 8051-8053) A solution containing 1.17 ml of 2-methyl-2-vinyl-oxirane (12 mmol), 1.6 g of CuCl2 (12 mmol) and 510 mg of LiCl (12 mmol) in 10 ml of ethylactetate was heated to 80° C. for 30 min. The reaction was stopped by adding 50 g of ice. The mixture was filtered through a sintered glass funnel under reduced pressure. 100 ml of CH2Cl2 was added and the organic phase was separated. The aqueous layer was extracted two times with 100 ml of CH2Cl2. The combined organic phase was dried over anhydrous MgSO4, filtered, and concentrated. The crude product was purified by chromatography over silica gel (CH2Cl2, 3×37 cm) to yield 0.755 g of a yellow liquid (6.4 mmol, 53%).
[0369]1H NMR (CDCl3, 500 MHz) δ 9.43 (s, 1H), 6.50 (t, J=7.5 Hz, 1H), 4.24 (d, J=7.5 Hz, 2H), 1.77 (s, 3H)
[0370]13C NMR (CDCl3, 125 MHz) δ 194.3, 145.7, 141.1, 38.6, 9.1
[0371]4-Chloro-2-methyl-2-buten-1-al-dimethyl-acetal
[0372]A solution of 184 mg 4-chloro-2-methyl-2-buten-1-al (1.55 mmol), 600 μl of HC(OMe)3 (5.6 mmol) and a catalytic amount of p-TsOH was incubated for 3 h at room temperature. The crude mixture was purified by chromatography over silica gel (n-hexane/ethylacetate 7:3) to yield 177 mg of a colourless liquid (1.08 mmol, 72%).
[0373]1H NMR (CDCl3, 500 MHz) δ 5.78 (t, 1H, J=7.9), 4.47 (s, 1H), 4.15 (d, J=7.9 Hz, 2H), 3.33 (s, 6H), 1.73 (s, 3H)
[0374]13C NMR (CDCl3 125 MHz) δ 137.6, 124.4, 106.0, 53.5, 39.6, 11.4
[0375](E)-3-Formyl-2-buten-1-diphosphate triammonium salt (Davisson et al. (1986) J. Org. Chem., 51, 4768) To a solution of 4-chloro-2-methyl-2-buten-1-al-dimethyl-acetal chloride (25 mg, 0.15 mmol) in 250 μl of MeCN a solution of 0.162 g (0.18 mmol) of tris(tetra-n-butylammonium)hydrogen pyrophosphate in 400 μL of MeCN was added slowly at room temperature, leading to an orange-red solution. After 2 h the reaction was finished and the solvent was removed under reduced pressure. The orange oil was dissolved in 3 mL of H2O and passed through a column of DOWEX 50 WX8 (1quadrature4 cm) cation-exchange resin (NH4.sup.+ form) that has been equilibrated with 20 mL of 25 mM NH4HCO3. The column was eluted with 20 mL of 25 mM NH4HCO3. The resulting solution was lyophilized. The obtained solid was dissolved in 2 ml water and acidified with aqueous HCl to pH=3. After 2 minutes the solution was neutralized and lyophylisized.
[0376]1H NMR (D2O, 360 MHz) δ 59.37 (s, 1H), 6.86 (t, 1H, 5.6 Hz), 4.85 (dd, J=7.9, J=5.8 Hz, 2H), 1.72 (s, 3H)
[0377]13C NMR (D2O, 90 MHz) δ 199.2, 153.1 (d, J=7.5 Hz), 138.5, 63.2 (d, J=4.9), 8.5
[0378][1-3H]1-hydroxy-2-methyl-but-2-enyl-4-diphosphate
[0379]A solution containing 50 mCi (15 μmol) NaBH3T, 15 μmol 3-formyl-2-buten-1-diphosphate triammonium salt and 100 mM Tris/HCl pH=8 was incubated for 30 minutes at room temperature. The solution was acidified by adding 1 M HCl to pH=2. After 2 minutes the solution was neutralized by adding 1 M NaOH.
[0380]The product was characterized by ion-exchange chromatography (see examples 20 and 25).
EXAMPLE 43
γδ T Cell Stimulation Assays
[0381]PBMCs from healthy donors (donor A and donor B) are isolated from heparinized peripheral blood by density centrifugation over Ficoll-Hypaque (Amersham Pharmacia Biotech, Freiburg, Germany). 5×105 PBMCs/well are cultivated in 1 mL RPMI 1640 medium supplemented with 10% human AB serum (Klinik rechts der Isar, Munchen, Germany), 2 mM L-glutamine, 10 μM mercaptoethanol. Amounts of recombinant human IL-2 (kindly provided by Eurocetus, Amsterdam, The Netherlands) and substrates are varied from 1 to 10 U and 10 to 0.1 μM, respectively. 20 μM IPP (Echelon, Research Laboratories Inc., Salt Lake City, USA) serves as a positive control whereas medium alone serves as negative control. Incubation is done for seven days at 37° C. in the presence of 7% CO2. The harvested cells are double-stained with fluorescein isothiocyanate (FITC)-conjugated mouse anti-human monoclonal antibody Vδ2 TCR and phycoerythrin (PE)-conjugated monoclonal CD3 antibody. The cells are analyzed using a FACScan supported with Cellquest (Becton Dickinson, Heidelberg, Germany).
[0382]The substrates (E)-1-hydroxy-3-methyl-but-2-enyl 4-diphosphate (HMBPP) and 3-formyl-but-2-enyl 1-diphosphate (Aldehyde) were prepared synthetically as described above.
[0383]It is found that both synthetically prepared substrates (HMBPP and Aldehyde) show at least double stimulation compared to IPP when used at a concentration that is 200-fold lower than the concentration of the IPP sample (Table 8).
TABLE-US-00008 TABLE 8 Activation of γδ T-cells by phosphororganic compounds Concentration IL-2 % γδ T-cells Substrate [μM] [U] Donor A Donor B Medium -- 1 1.51 2.75 Medium -- 5 1.45 2.23 Medium -- 10 1.32 1.68 IPP 20 1 8.19 6.24 IPP 20 5 14.42 9.32 IPP 20 10 16.6 11.86 IPP 1 1 1.56 2.22 IPP 1 5 1.59 2.67 IPP 1 10 1.71 2.19 IPP 0.1 1 1.3 2.15 IPP 0.1 5 1.3 2.26 IPP 0.1 10 1.01 2.54 HMBPP 10 1 3.3 31.42 HMBPP 10 5 17.38 63.48 HMBPP 10 10 24.94 63.34 HMBPP 1 1 5.57 35.34 HMBPP 1 5 14.4 54.12 HMBPP 1 10 19.85 55.90 HMBPP 0.1 1 11.78 32.21 HMBPP 0.1 5 22.92 44.69 HMBPP 0.1 10 34.69 36.33 HMBPP/IPP 0.5/0.5 1 7 30.35 HMBPP/IPP 0.5/0.5 5 15.38 53.76 HMBPP/IPP 0.5/0.5 10 24.19 46.58 Aldehyde 10 1 12.19 30.69 Aldehyde 10 5 34.69 30.33 Aldehyde 10 10 38.99 38.85 Aldehyde 1 1 15.91 21.18 Aldehyde 1 5 40.13 30.76 Aldehyde 1 10 48.28 36.69 Aldehyde 0.1 1 10 13.54 Aldehyde 0.1 5 19.77 18.45 Aldehyde 0.1 10 21.93 25.82 Aldehyde/IPP 0.5/0.5 1 13.98 22.11 Aldehyde/IPP 0.5/0.5 5 33.94 32.06 Aldehyde/IPP 0.5/0.5 10 42.84 36.25 IPP: isopentenyl diphosphate HMBPP: (E)-1-hydroxy-3-methyl-but-2-enyl 4-diphosphate Aldehyde: (E)-3-formyl-but-2-enyl 1-diphosphate (prepared according to example 42)
EXAMPLE 44
High Through-Put Screening Assay of 1-hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate Synthase (IspG) Activity
[0384]Assay mixtures contain 20 mM potassium phosphate, pH 7.0, 0.4 mM NADH, 0.5 mM CoCl2, 0.2 mM 2C-methyl-D-erythritol 2,4-cyclodiphosphate, and 50 μl protein in a total volume of 1 ml. The mixtures are incubated at 37° C. The oxidation of NADH is monitored photometrically at 340 nm. Alternatively, the concentration of NADH is determined by measuring the relative fluorescence of NADH at 340 nm excitation/460 nm emission.
EXAMPLE 45
High Through-Put Screening Assay of 1-hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate Reductase (IspH) Activity
[0385]Assay mixtures contain 20 mM potassium phosphate, pH 7.0, pH 8.0, 0.4 mM NADH, 20 μM FAD, 0.5 mM COCl2, 0.2 mM 1-hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate, and 50 μl protein in a total volume of 1 ml. The mixtures are incubated at 37° C. The oxidation of NADH is monitored photometrically at 340 nm. Alternatively, the concentration of NADH is determined by measuring the relative fluorescence of NADH at 340 nm excitation/460 nm emission.
REFERENCES
[0386]Altincicek, B., Kollas, A. K., Sanderbrand, S., Wiesner, J., Hintz, M., Beck, E. & Jomaa, H. (2001). GcpE Is Involved in the 2-C-Methyl-D-Erythritol 4-Phosphate Pathway of Isoprenoid Biosynthesis in Escherichia coli. J. Bacteriol. 183, 2411-2416. [0387]Altincicek, B. et al. and Jomaa, H. (2001) J. Immunology, 166, 3655-3658. [0388]Arigoni D. & Schwarz M. K. (1999) Ginkgolide biosynthesis. In Comprehensive natural product chemistry (Barton D. and Nakanishi K., eds.), Vol. 2, pp. 367-399, Pergamon. [0389]Begley, T. P., Downs, D. M., Ealick, S. E., McLafferty, F. W., VanLoon, A. P. G. M., Taylor, S., Campobasso, N., Chiu, H.-J., Kinsland, C., Reddick, J. J. & Xi, J. (1999) Thiamin biosynthesis in prokaryotes. Arch. Microbiol. 171, 293-300. Blagg, B. S. J. & Poulter, C. D. (1999) Synthesis of 1-deoxy-D-xylulose and 1-deoxy-D-xylulose 5-phosphate. J. Org. Chem. 64, 1508-1511. [0390]Bullock, W. O., Fernandez, J. M., & Short, J. M. (1987). XL1-Blue: a high efficiency plasmid transforming recA Escherichia coli with β-galactosidase selection. BioTechniques 5, 376-379. [0391]Camara, B. Methods in Enzymology 1985, eds. Law, J. H. & Rilling, H. C. (Academic Press, London) Vol. 110, pp. 267-273. [0392]Camara, B. Methods in Enzymology 1993, ed. Packer, L. (Academic Press, London) Vol. 214, pp. 352-365. [0393]Campos, N., Rodriguez-Concepcion, M., Seemann, M., Rohmer, M. and Boronat, A. (2001) Identification of gcpE as a novel gene of the 2-C-methyl-D-erythritol 4-phosphate pathway for isoprenoid biosynthesis in Escherichia coli. FEBS Lett. 488, 170-173. [0394]Chang, A. C. Y. & Cohen, S. N. (1978) Construction and characterization of amplifiable multicopy DNA cloning vehicles derived from the P15A cryptic miniplasmid. J. Bacteriol. 134, 1141-1145. [0395]Cunningham, F. X. Jr., Lafond, T. P. & Gantt E. (2000). Evidence of a role for LytB in the nonmevalonate pathway of isoprenoid biosynthesis. J. Bacteriol. 182, 5841-5848. [0396]Davisson, V. J.; Woodside, A. B.; Neal, T. R.; Stremler, K. E.; Muehibacher, M.; Poulter, C. D. J. Org. Chem. 1986, 51, 4768. [0397]Dower, W. J., Miller, J. F., & Ragsdale, C. (1988). High efficiency transformation of E. coli by high voltage electroporation. Nucleic Acids Res. 16, 6127-6145. [0398]Eisenreich, W., Rohdich, F. & Bacher, A. (2001) Deoxyxylulose phosphate pathway to terpenoids. Trends in Plant Science 6, 78-84. [0399]Fournie, J. J. and Bonneville, M. (1996) Res. Immunol. 147, 338-347. [0400]Giner, J.-L. (1998) New and efficient synthetic routes to 1-deoxy-D-xylulose. Tetrahedron Lett. 39, 2479-2482. [0401]Hagiwara, H.; Uda, H. J. Chem. Soc. Trans./1984, 91. [0402]Hwang, C. K.; Li, W. S.; Nicolaou, K. C. Tetrahedron Lett. 1984, 25, 2295. [0403]Herz, S., Wungsintaweekul, J., Schuhr, C. A., Hecht, S., Luttgen, H., Sagner, S., Fellermeier, M., Eisenreich, W., Zenk, M. H., Bacher, A. & Rohdich, F. (2000). Biosynthesis of terpenoids: YgbB protein converts 4-diphosphocytidyl-2C-methyl-D-erythritol 2-phosphate to 2C-methyl-D-erythritol 2,4-cyclodiphosphate. Proc. Natl. Acad. Sci. USA 97, 2486-2490. [0404]Jomaa, H., Feurle, J., Luhs, K., Kunzmann, V., Tony, H. P., Herderich, M. and Wilhelm, M. (1999) FEMS Immunology and Medical Microbiology 25, 371-378. [0405]Kennedy, I. A., Hemscheidt, T., Britten, J. F., Spenser, I. D. (1995) 1-Deoxy-D-xylulose Can. J. Chem. 73, 1329-1333. [0406]Luttgen, H., Rohdich, F., Herz, S., Wungsintaweekul, J., Hecht, S., Schuhr, C. A., Fellermeier, M., Sagner, S., Zenk, M. H., Bacher, A. & Eisenreich, W. (2000). Biosynthesis of terpenoids: YchB protein of Escherichia Coli phosphorylates the 2-hydroxy group of 4-diphosphocytidyl-2C-methyl-D-erythritol. Proc. Natl. Acad. Sci. USA 97, 1062-1067. [0407]Meade, H. M., Long, S. R., Ruvkun, C. B., Brown, S. E., & Auswald, F. M. (1982). Physical and genetic characterization of symbiotic and auxotrophic mutants of Rhizobium meliloti induced by transposon Tn5 mutagenis. J. Bacteriol. 149, 114-122. [0408]Piel, J. & Boland, W. (1997) Highly efficient and versatile synthesis of isotopically labeled 1-deoxy-D-xylulose Tetrahedron Lett. 38, 6387-6390. [0409]Rohdich, F., Wungsintaweekul, J., Fellermeier, M., Sagner, S., Herz, S., Kis, K., Eisenreich, W., Bacher, A & Zenk, M. H. (1999). Cytidine 5'-triphosphate biosynthesis of isoprenoids: YgbP protein of Escherichia coli catalyzes the formation of 4-diphosphocytidyl-2C-methylerythritol. Proc. Natl. Acad. Sci. USA 96, 11758-11763. [0410]Sanger, F., Nicklen, S., & Coulson, A. R. (1992). DNA sequencing with chain-terminating inhibitors. Biotechnology 24, 104-8. [0411]Shono, T., Matsamura, Y., Hamaguchi, H. & Naitoh, S. (1983) Synthesis of 2-methyl-3-hydroxy-4H-pyrane-4-one and 4-hydroxy-5-methyl-2H-furane-3-one from carbohydrates. J. Org. Chem. 48, 5126-5128. [0412]Takahashi, S., Kuzuyama, T., Watanabe H. & Seto, H. (1998) A 1-deoxy-D-xylulose 5-phosphate reductoisomerase catalysing the formation of 2-C-methyl-D-erythritol 4-phosphate in an alternative nonmevalonate pathway for terpenoid biosynthesis. Proc. Natl. Acad. Sci. USA 95, 9879-9884. [0413]Watanabe, H.; Hatakeyama, S.; Tazumi, K.; Takano, S.; Masuda, S.; Okano, T.; Kobayashi, T.; Kubodera, N. Chem. Pharm. Bull. 1996, 44, 2280.Wungsintaweekul, J., Herz, S., Hecht, S., Eisenreich, W., Feicht, R., Rohdich, F., Bacher, A. & Zenk, M. H. (2001). Phosphorylation of 1-deoxy-D-xylulose by D-xylulokinase of Escherichia coli. Eur. J. Biochem. 268, 310-316.
Sequence CWU
1
58150DNAArtificialxylB 5' PCR primer 1ccgtcggaat tcgaggagaa attaaccatg
tatatcggga tagatcttgg 50237DNAArtificialxyl B 3' PCR
primer 2gcagtgaagc ttttacgcca ttaatggcag aagttgc
37349DNAArtificialdxr PCR primer 1 3ctagccaagc ttgaggagaa attaaccatg
aagcaactca ccattctgg 49428DNAArtificialdxr PCR primer 2
4ggagatgtcg actcagcttg cgagacgc
28549DNAArtificialispD PCR primer 1 5ccgggagtcg acgaggagaa attaaccatg
gcaaccactc atttggatg 49632DNAArtificialispD PCR primer 2
6gtccaactcg agttatgtat tctccttgat gg
32749DNAArtificialispD/ispF PCR primer 1 7ccgggagtcg acgaggagaa
attaaccatg gcaaccactc atttggatg
49833DNAArtificialispD/ispF PCR primer 2 8tatcaactcg agtcattttg
ttgccttaat gag 33945DNAArtificialispE
PCR primer 1 9gcgaacctcg aggaggagaa attaaccatg cggacacagt ggccc
451031DNAArtificialispE PCR primer 2 10cctgacggta ccttaaagca
tggctctgtg c 311149DNAArtificialgcpE
PCR primer 1 11cgtaccggat ccgaggagaa attaaccatg cataaccagg ctccaattc
491236DNAArtificialgcpE PCR primer 2 12cccatcgtcg acttattttt
caacctgctg aacgtc
361323DNAArtificialcrtY/crtl/crtB PCR primer 1 13cattgagaag cttatgtgca
ccg
231420DNAArtificialcrtY/crtl/crtB PCR primer 2 14ctccggggtc gacatggcgc
201520DNAArtificialcrtE PCR
primer 1 15ccgcatcttt ccaattgccg
201623DNAArtificialcrtE PCR primer 2 16atgcagcaag cttaactgac ggc
231752DNAArtificialispG PCR
primer 1 17gcgggagacc gcgggaggag aaattaacca tgcataacca ggctccaatt cg
521839DNAArtificialispG PCR primer 2 18cgcttcccag cggccgctta
tttttcaacc tgctgaacg 391945DNAArtificiallytB
PCR primer 1 19aaatcggagc tcgaggagaa attaaccatg cagatcctgt tggcc
452034DNAArtificiallytB PCR primer 2 20gctgctccgc ggttaatcga
cttcacgaat atcg 342148DNAArtificialispH
PCR primer 1 21gcttgcgtcg acgaggagaa attaaccatg cagatcctgt tggccacc
482234DNAArtificialispH PCR primer 2 22gctgctcggc cgttaatcga
cttcacgaat atcg 342349DNAArtificialispG
(gcpE) primer 1 23cgtaccggat ccgaggagaa attaaccatg cataaccagg ctccaattc
492436DNAArtificialispG (gcpE) primer 2 24cccatcgtcg
acttattttt caacctgctg aacgtc
362555DNAArtificialispG/ispH primer 1 25gcgggagacc gcgggaggag aaattaacca
tgcataacca ggctccaatt caacg 552636DNAArtificialispG/ispH primer
2 26aggctggcgg ccgcttaatc gacttcacga atatcg
362720DNAArtificialispG primer 1 27cctgcatccg aaggaagccc
202820DNAArtificialispG primer 2
28cagttttcaa agaatggccc
202928DNAArtificialPCR primer 1 (example 31) 29tgaatcagga tccaagacgg
tgagaagg 283030DNAArtificialPCR
primer 2 (example 31) 30tccgtttggt accctactca tcagccacgg
303152DNAArtificialdxs primer 1 (example 34)
31ggcgactcgc gagaggagaa attaaccatg gatcttttat caatacagga cc
523235DNAArtificialdxs primer 2 (example 34) 32ggcacccggc cgtcatgatc
caattccttt gtgtg 353348DNAArtificialispH
primer 1 (example 35) 33gcttgcgtcg acgaggagaa attaaccatg cagatcctgt
tggccacc 483434DNAArtificialispH primer 2 (example 35)
34gctgctctcg agttaatcga cttcacgaat atcg
343549DNAArtificialispG primer 1 (example 35) 35ggtcgagtcg acgaggagaa
attaaccatg cataaccagg ctccaattc 493636DNAArtificialispG
primer 2 (example 35) 36cccatcctcg agttattttt caacctgctg aacgtc
363734DNAArtificialispG primer 1 (example 38)
37gaaccggaat tcatgcataa ccaggctcca attc
343819DNAArtificialispG primer 2 (example 38) 38cgaggcggat cccatcacg
193933DNAArtificialispH
primer 1 (example 39) 39tggaggggat ccatgcagat cctgttggcc acc
334021DNAArtificialispH primer 2 (example 39)
40gcatttctgc agaacttagg c
21415628DNAArtificialpBSxylBdxr 41gtggcacttt tcggggaaat gtgcgcggaa
cccctatttg tttatttttc taaatacatt 60caaatatgta tccgctcatg agacaataac
cctgataaat gcttcaataa tattgaaaaa 120ggaagagtat gagtattcaa catttccgtg
tcgcccttat tccctttttt gcggcatttt 180gccttcctgt ttttgctcac ccagaaacgc
tggtgaaagt aaaagatgct gaagatcagt 240tgggtgcacg agtgggttac atcgaactgg
atctcaacag cggtaagatc cttgagagtt 300ttcgccccga agaacgtttt ccaatgatga
gcacttttaa agttctgcta tgtggcgcgg 360tattatcccg tattgacgcc gggcaagagc
aactcggtcg ccgcatacac tattctcaga 420atgacttggt tgagtactca ccagtcacag
aaaagcatct tacggatggc atgacagtaa 480gagaattatg cagtgctgcc ataaccatga
gtgataacac tgcggccaac ttacttctga 540caacgatcgg aggaccgaag gagctaaccg
cttttttgca caacatgggg gatcatgtaa 600ctcgccttga tcgttgggaa ccggagctga
atgaagccat accaaacgac gagcgtgaca 660ccacgatgcc tgtagcaatg gcaacaacgt
tgcgcaaact attaactggc gaactactta 720ctctagcttc ccggcaacaa ttaatagact
ggatggaggc ggataaagtt gcaggaccac 780ttctgcgctc ggcccttccg gctggctggt
ttattgctga taaatctgga gccggtgagc 840gtgggtctcg cggtatcatt gcagcactgg
ggccagatgg taagccctcc cgtatcgtag 900ttatctacac gacggggagt caggcaacta
tggatgaacg aaatagacag atcgctgaga 960taggtgcctc actgattaag cattggtaac
tgtcagacca agtttactca tatatacttt 1020agattgattt aaaacttcat ttttaattta
aaaggatcta ggtgaagatc ctttttgata 1080atctcatgac caaaatccct taacgtgagt
tttcgttcca ctgagcgtca gaccccgtag 1140aaaagatcaa aggatcttct tgagatcctt
tttttctgcg cgtaatctgc tgcttgcaaa 1200caaaaaaacc accgctacca gcggtggttt
gtttgccgga tcaagagcta ccaactcttt 1260ttccgaaggt aactggcttc agcagagcgc
agataccaaa tactgtcctt ctagtgtagc 1320cgtagttagg ccaccacttc aagaactctg
tagcaccgcc tacatacctc gctctgctaa 1380tcctgttacc agtggctgct gccagtggcg
ataagtcgtg tcttaccggg ttggactcaa 1440gacgatagtt accggataag gcgcagcggt
cgggctgaac ggggggttcg tgcacacagc 1500ccagcttgga gcgaacgacc tacaccgaac
tgagatacct acagcgtgag ctatgagaaa 1560gcgccacgct tcccgaaggg agaaaggcgg
acaggtatcc ggtaagcggc agggtcggaa 1620caggagagcg cacgagggag cttccagggg
gaaacgcctg gtatctttat agtcctgtcg 1680ggtttcgcca cctctgactt gagcgtcgat
ttttgtgatg ctcgtcaggg gggcggagcc 1740tatggaaaaa cgccagcaac gcggcctttt
tacggttcct ggccttttgc tggccttttg 1800ctcacatgtt ctttcctgcg ttatcccctg
attctgtgga taaccgtatt accgcctttg 1860agtgagctga taccgctcgc cgcagccgaa
cgaccgagcg cagcgagtca gtgagcgagg 1920aagcggaaga gcgcccaata cgcaaaccgc
ctctccccgc gcgttggccg attcattaat 1980gcagctggca cgacaggttt cccgactgga
aagcgggcag tgagcgcaac gcaattaatg 2040tgagttagct cactcattag gcaccccagg
ctttacactt tatgcttccg gctcgtatgt 2100tgtgtggaat tgtgagcgga taacaatttc
acacaggaaa cagctatgac catgattacg 2160ccaagcgcgc aattaaccct cactaaaggg
aacaaaagct ggagctccac cgcggtggcg 2220gccgctctag aactagtgga tcccccgggc
tgcaggaatt cgaggagaaa ttaaccatgt 2280atatcgggat agatcttggc acctcgggcg
taaaagttat tttgctcaac gagcagggtg 2340aggtggttgc tgcgcaaacg gaaaagctga
ccgtttcgcg cccgcatcca ctctggtcgg 2400aacaagaccc ggaacagtgg tggcaggcaa
ctgatcgcgc aatgaaagct ctgggcgatc 2460agcattctct gcaggacgtt aaagcattgg
gtattgccgg ccagatgcac ggagcaacct 2520tgctggatgc tcagcaacgg gtgttacgcc
ctgccatttt gtggaacgac gggcgctgtg 2580cgcaagagtg cactttgctg gaagcgcgag
ttccgcaatc gcgggtgatt accggcaacc 2640tgatgatgcc cggatttact gcgcctaaat
tgctatgggt tcagcggcat gagccggaga 2700tattccgtca aatcgacaaa gtattattac
cgaaagatta cttgcgtctg cgtatgacgg 2760gggagtttgc cagcgatatg tctgacgcag
ctggcaccat gtggctggat gtcgcaaagc 2820gtgactggag tgacgtcatg ctgcaggctt
gcgacttatc tcgtgaccag atgcccgcat 2880tatacgaagg cagcgaaatt actggtgctt
tgttacctga agttgcgaaa gcgtggggta 2940tggcgacggt gccagttgtc gcaggcggtg
gcgacaatgc agctggtgca gttggtgtgg 3000gaatggttga tgctaatcag gcaatgttat
cgctggggac gtcgggggtc tattttgctg 3060tcagcgaagg gttcttaagc aagccagaaa
gcgccgtaca tagcttttgc catgcgctac 3120cgcaacgttg gcatttaatg tctgtgatgc
tgagtgcagc gtcgtgtctg gattgggccg 3180cgaaattaac cggcctgagc aatgtcccag
ctttaatcgc tgcagctcaa caggctgatg 3240aaagtgccga gccagtttgg tttctgcctt
atctttccgg cgagcgtacg ccacacaata 3300atccccaggc gaagggggtt ttctttggtt
tgactcatca acatggcccc aatgaactgg 3360cgcgagcagt gctggaaggc gtgggttatg
cgctggcaga tggcatggat gtcgtgcatg 3420cctgcggtat taaaccgcaa agtgttacgt
tgattggggg cggggcgcgt agtgagtact 3480ggcgtcagat gctggcggat atcagcggtc
agcagctcga ttaccgtacg gggggggatg 3540tggggccagc actgggcgca gcaaggctgg
cgcagatcgc ggcgaatcca gagaaatcgc 3600tcattgaatt gttgccgcaa ctaccgttag
aacagtcgca tctaccagat gcgcagcgtt 3660atgccgctta tcagccacga cgagaaacgt
tccgtcgcct ctatcagcaa cttctgccat 3720taatggcgta aaagcttgag gagaaattaa
ccatgaagca actcaccatt ctgggctcga 3780ccggctcgat tggttgcagc acgctggacg
tggtgcgcca taatcccgaa cacttccgcg 3840tagttgcgct ggtggcaggc aaaaatgtca
ctcgcatggt agaacagtgc ctggaattct 3900ctccccgcta tgccgtaatg gacgatgaag
cgagtgcgaa acttcttaaa acgatgctac 3960agcaacaggg tagccgcacc gaagtcttaa
gtgggcaaca agccgcttgc gatatggcag 4020cgcttgagga tgttgatcag gtgatggcag
ccattgttgg cgctgctggg ctgttaccta 4080cgcttgctgc gatccgcgcg ggtaaaacca
ttttgctggc caataaagaa tcactggtta 4140cctgcggacg tctgtttatg gacgccgtaa
agcagagcaa agcgcaattg ttaccggtcg 4200atagcgaaca taacgccatt tttcagagtt
taccgcaacc tatccagcat aatctgggat 4260acgctgacct tgagcaaaat ggcgtggtgt
ccattttact taccgggtct ggtggccctt 4320tccgtgagac gccattgcgc gatttggcaa
caatgacgcc ggatcaagcc tgccgtcatc 4380cgaactggtc gatggggcgt aaaatttctg
tcgattcggc taccatgatg aacaaaggtc 4440tggaatacat tgaagcgcgt tggctgttta
acgccagcgc cagccagatg gaagtgctga 4500ttcacccgca gtcagtgatt cactcaatgg
tgcgctatca ggacggcagt gttctggcgc 4560agctggggga accggatatg cgtacgccaa
ttgcccacac catggcatgg ccgaatcgcg 4620tgaactctgg cgtgaagccg ctcgattttt
gcaaactaag tgcgttgaca tttgccgcac 4680cggattatga tcgttatcca tgcctgaaac
tggcgatgga ggcgttcgaa caaggccagg 4740cagcgacgac agcattgaat gccgcaaacg
aaatcaccgt tgctgctttt cttgcgcaac 4800aaatccgctt tacggatatc gctgcgttga
atttatccgt actggaaaaa atggatatgc 4860gcgaaccaca atgtgtggac gatgtgttat
ctgttgatgc gaacgcgcgt gaagtcgcca 4920gaaaagaggt gatgcgtctc gcaagctgag
tcgacctcga gggggggccc ggtacccaat 4980tcgccctata gtgagtcgta ttacgcgcgc
tcactggccg tcgttttaca acgtcgtgac 5040tgggaaaacc ctggcgttac ccaacttaat
cgccttgcag cacatccccc tttcgccagc 5100tggcgtaata gcgaagaggc ccgcaccgat
cgcccttccc aacagttgcg cagcctgaat 5160ggcgaatgga aattgtaagc gttaatattt
tgttaaaatt cgcgttaaat ttttgttaaa 5220tcagctcatt ttttaaccaa taggccgaaa
tcggcaaaat cccttataaa tcaaaagaat 5280agaccgagat agggttgagt gttgttccag
tttggaacaa gagtccacta ttaaagaacg 5340tggactccaa cgtcaaaggg cgaaaaaccg
tctatcaggg cgatggccca ctacgtgaac 5400catcacccta atcaagtttt ttggggtcga
ggtgccgtaa agcactaaat cggaacccta 5460aagggagccc ccgatttaga gcttgacggg
gaaagccggc gaacgtggcg agaaaggaag 5520ggaagaaagc gaaaggagcg ggcgctaggg
cgctggcaag tgtagcggtc acgctgcgcg 5580taaccaccac acccgccgcg cttaatgcgc
cgctacaggg cgcgtcag 5628426354DNAArtificialpBSxylBdxrispD
42gtggcacttt tcggggaaat gtgcgcggaa cccctatttg tttatttttc taaatacatt
60caaatatgta tccgctcatg agacaataac cctgataaat gcttcaataa tattgaaaaa
120ggaagagtat gagtattcaa catttccgtg tcgcccttat tccctttttt gcggcatttt
180gccttcctgt ttttgctcac ccagaaacgc tggtgaaagt aaaagatgct gaagatcagt
240tgggtgcacg agtgggttac atcgaactgg atctcaacag cggtaagatc cttgagagtt
300ttcgccccga agaacgtttt ccaatgatga gcacttttaa agttctgcta tgtggcgcgg
360tattatcccg tattgacgcc gggcaagagc aactcggtcg ccgcatacac tattctcaga
420atgacttggt tgagtactca ccagtcacag aaaagcatct tacggatggc atgacagtaa
480gagaattatg cagtgctgcc ataaccatga gtgataacac tgcggccaac ttacttctga
540caacgatcgg aggaccgaag gagctaaccg cttttttgca caacatgggg gatcatgtaa
600ctcgccttga tcgttgggaa ccggagctga atgaagccat accaaacgac gagcgtgaca
660ccacgatgcc tgtagcaatg gcaacaacgt tgcgcaaact attaactggc gaactactta
720ctctagcttc ccggcaacaa ttaatagact ggatggaggc ggataaagtt gcaggaccac
780ttctgcgctc ggcccttccg gctggctggt ttattgctga taaatctgga gccggtgagc
840gtgggtctcg cggtatcatt gcagcactgg ggccagatgg taagccctcc cgtatcgtag
900ttatctacac gacggggagt caggcaacta tggatgaacg aaatagacag atcgctgaga
960taggtgcctc actgattaag cattggtaac tgtcagacca agtttactca tatatacttt
1020agattgattt aaaacttcat ttttaattta aaaggatcta ggtgaagatc ctttttgata
1080atctcatgac caaaatccct taacgtgagt tttcgttcca ctgagcgtca gaccccgtag
1140aaaagatcaa aggatcttct tgagatcctt tttttctgcg cgtaatctgc tgcttgcaaa
1200caaaaaaacc accgctacca gcggtggttt gtttgccgga tcaagagcta ccaactcttt
1260ttccgaaggt aactggcttc agcagagcgc agataccaaa tactgtcctt ctagtgtagc
1320cgtagttagg ccaccacttc aagaactctg tagcaccgcc tacatacctc gctctgctaa
1380tcctgttacc agtggctgct gccagtggcg ataagtcgtg tcttaccggg ttggactcaa
1440gacgatagtt accggataag gcgcagcggt cgggctgaac ggggggttcg tgcacacagc
1500ccagcttgga gcgaacgacc tacaccgaac tgagatacct acagcgtgag ctatgagaaa
1560gcgccacgct tcccgaaggg agaaaggcgg acaggtatcc ggtaagcggc agggtcggaa
1620caggagagcg cacgagggag cttccagggg gaaacgcctg gtatctttat agtcctgtcg
1680ggtttcgcca cctctgactt gagcgtcgat ttttgtgatg ctcgtcaggg gggcggagcc
1740tatggaaaaa cgccagcaac gcggcctttt tacggttcct ggccttttgc tggccttttg
1800ctcacatgtt ctttcctgcg ttatcccctg attctgtgga taaccgtatt accgcctttg
1860agtgagctga taccgctcgc cgcagccgaa cgaccgagcg cagcgagtca gtgagcgagg
1920aagcggaaga gcgcccaata cgcaaaccgc ctctccccgc gcgttggccg attcattaat
1980gcagctggca cgacaggttt cccgactgga aagcgggcag tgagcgcaac gcaattaatg
2040tgagttagct cactcattag gcaccccagg ctttacactt tatgcttccg gctcgtatgt
2100tgtgtggaat tgtgagcgga taacaatttc acacaggaaa cagctatgac catgattacg
2160ccaagcgcgc aattaaccct cactaaaggg aacaaaagct ggagctccac cgcggtggcg
2220gccgctctag aactagtgga tcccccgggc tgcaggaatt cgaggagaaa ttaaccatgt
2280atatcgggat agatcttggc acctcgggcg taaaagttat tttgctcaac gagcagggtg
2340aggtggttgc tgcgcaaacg gaaaagctga ccgtttcgcg cccgcatcca ctctggtcgg
2400aacaagaccc ggaacagtgg tggcaggcaa ctgatcgcgc aatgaaagct ctgggcgatc
2460agcattctct gcaggacgtt aaagcattgg gtattgccgg ccagatgcac ggagcaacct
2520tgctggatgc tcagcaacgg gtgttacgcc ctgccatttt gtggaacgac gggcgctgtg
2580cgcaagagtg cactttgctg gaagcgcgag ttccgcaatc gcgggtgatt accggcaacc
2640tgatgatgcc cggatttact gcgcctaaat tgctatgggt tcagcggcat gagccggaga
2700tattccgtca aatcgacaaa gtattattac cgaaagatta cttgcgtctg cgtatgacgg
2760gggagtttgc cagcgatatg tctgacgcag ctggcaccat gtggctggat gtcgcaaagc
2820gtgactggag tgacgtcatg ctgcaggctt gcgacttatc tcgtgaccag atgcccgcat
2880tatacgaagg cagcgaaatt actggtgctt tgttacctga agttgcgaaa gcgtggggta
2940tggcgacggt gccagttgtc gcaggcggtg gcgacaatgc agctggtgca gttggtgtgg
3000gaatggttga tgctaatcag gcaatgttat cgctggggac gtcgggggtc tattttgctg
3060tcagcgaagg gttcttaagc aagccagaaa gcgccgtaca tagcttttgc catgcgctac
3120cgcaacgttg gcatttaatg tctgtgatgc tgagtgcagc gtcgtgtctg gattgggccg
3180cgaaattaac cggcctgagc aatgtcccag ctttaatcgc tgcagctcaa caggctgatg
3240aaagtgccga gccagtttgg tttctgcctt atctttccgg cgagcgtacg ccacacaata
3300atccccaggc gaagggggtt ttctttggtt tgactcatca acatggcccc aatgaactgg
3360cgcgagcagt gctggaaggc gtgggttatg cgctggcaga tggcatggat gtcgtgcatg
3420cctgcggtat taaaccgcaa agtgttacgt tgattggggg cggggcgcgt agtgagtact
3480ggcgtcagat gctggcggat atcagcggtc agcagctcga ttaccgtacg gggggggatg
3540tggggccagc actgggcgca gcaaggctgg cgcagatcgc ggcgaatcca gagaaatcgc
3600tcattgaatt gttgccgcaa ctaccgttag aacagtcgca tctaccagat gcgcagcgtt
3660atgccgctta tcagccacga cgagaaacgt tccgtcgcct ctatcagcaa cttctgccat
3720taatggcgta aaagcttgag gagaaattaa ccatgaagca actcaccatt ctgggctcga
3780ccggctcgat tggttgcagc acgctggacg tggtgcgcca taatcccgaa cacttccgcg
3840tagttgcgct ggtggcaggc aaaaatgtca ctcgcatggt agaacagtgc ctggaattct
3900ctccccgcta tgccgtaatg gacgatgaag cgagtgcgaa acttcttaaa acgatgctac
3960agcaacaggg tagccgcacc gaagtcttaa gtgggcaaca agccgcttgc gatatggcag
4020cgcttgagga tgttgatcag gtgatggcag ccattgttgg cgctgctggg ctgttaccta
4080cgcttgctgc gatccgcgcg ggtaaaacca ttttgctggc caataaagaa tcactggtta
4140cctgcggacg tctgtttatg gacgccgtaa agcagagcaa agcgcaattg ttaccggtcg
4200atagcgaaca taacgccatt tttcagagtt taccgcaacc tatccagcat aatctgggat
4260acgctgacct tgagcaaaat ggcgtggtgt ccattttact taccgggtct ggtggccctt
4320tccgtgagac gccattgcgc gatttggcaa caatgacgcc ggatcaagcc tgccgtcatc
4380cgaactggtc gatggggcgt aaaatttctg tcgattcggc taccatgatg aacaaaggtc
4440tggaatacat tgaagcgcgt tggctgttta acgccagcgc cagccagatg gaagtgctga
4500ttcacccgca gtcagtgatt cactcaatgg tgcgctatca ggacggcagt gttctggcgc
4560agctggggga accggatatg cgtacgccaa ttgcccacac catggcatgg ccgaatcgcg
4620tgaactctgg cgtgaagccg ctcgattttt gcaaactaag tgcgttgaca tttgccgcac
4680cggattatga tcgttatcca tgcctgaaac tggcgatgga ggcgttcgaa caaggccagg
4740cagcgacgac agcattgaat gccgcaaacg aaatcaccgt tgctgctttt cttgcgcaac
4800aaatccgctt tacggatatc gctgcgttga atttatccgt actggaaaaa atggatatgc
4860gcgaaccaca atgtgtggac gatgtgttat ctgttgatgc gaacgcgcgt gaagtcgcca
4920gaaaagaggt gatgcgtctc gcaagctgag tcgacgagga gaaattaacc atggcaacca
4980ctcatttgga tgtttgcgcc gtggttccgg cggccggatt tggccgtcga atgcaaacgg
5040aatgtcctaa gcaatatctc tcaatcggta atcaaaccat tcttgaacac tcggtgcatg
5100cgctgctggc gcatccccgg gtgaaacgtg tcgtcattgc cataagtcct ggcgatagcc
5160gttttgcaca acttcctctg gcgaatcatc cgcaaatcac cgttgtagat ggcggtgatg
5220agcgtgccga ttccgtgctg gcaggtctga aagccgctgg cgacgcgcag tgggtattgg
5280tgcatgacgc cgctcgtcct tgtttgcatc aggatgacct cgcgcgattg ttggcgttga
5340gcgaaaccag ccgcacgggg gggatcctcg ccgcaccagt gcgcgatact atgaaacgtg
5400ccgaaccggg caaaaatgcc attgctcata ccgttgatcg caacggctta tggcacgcgc
5460tgacgccgca atttttccct cgtgagctgt tacatgactg tctgacgcgc gctctaaatg
5520aaggcgcgac tattaccgac gaagcctcgg cgctggaata ttgcggattc catcctcagt
5580tggtcgaagg ccgtgcggat aacattaaag tcacgcgccc ggaagatttg gcactggccg
5640agttttacct cacccgaacc atccatcagg agaatacata actcgagggg gggcccggta
5700cccaattcgc cctatagtga gtcgtattac gcgcgctcac tggccgtcgt tttacaacgt
5760cgtgactggg aaaaccctgg cgttacccaa cttaatcgcc ttgcagcaca tccccctttc
5820gccagctggc gtaatagcga agaggcccgc accgatcgcc cttcccaaca gttgcgcagc
5880ctgaatggcg aatggaaatt gtaagcgtta atattttgtt aaaattcgcg ttaaattttt
5940gttaaatcag ctcatttttt aaccaatagg ccgaaatcgg caaaatccct tataaatcaa
6000aagaatagac cgagataggg ttgagtgttg ttccagtttg gaacaagagt ccactattaa
6060agaacgtgga ctccaacgtc aaagggcgaa aaaccgtcta tcagggcgat ggcccactac
6120gtgaaccatc accctaatca agttttttgg ggtcgaggtg ccgtaaagca ctaaatcgga
6180accctaaagg gagcccccga tttagagctt gacggggaaa gccggcgaac gtggcgagaa
6240aggaagggaa gaaagcgaaa ggagcgggcg ctagggcgct ggcaagtgta gcggtcacgc
6300tgcgcgtaac caccacaccc gccgcgctta atgcgccgct acagggcgcg tcag
6354437691DNAArtificialpBScyclo 43gtggcacttt tcggggaaat gtgcgcggaa
cccctatttg tttatttttc taaatacatt 60caaatatgta tccgctcatg agacaataac
cctgataaat gcttcaataa tattgaaaaa 120ggaagagtat gagtattcaa catttccgtg
tcgcccttat tccctttttt gcggcatttt 180gccttcctgt ttttgctcac ccagaaacgc
tggtgaaagt aaaagatgct gaagatcagt 240tgggtgcacg agtgggttac atcgaactgg
atctcaacag cggtaagatc cttgagagtt 300ttcgccccga agaacgtttt ccaatgatga
gcacttttaa agttctgcta tgtggcgcgg 360tattatcccg tattgacgcc gggcaagagc
aactcggtcg ccgcatacac tattctcaga 420atgacttggt tgagtactca ccagtcacag
aaaagcatct tacggatggc atgacagtaa 480gagaattatg cagtgctgcc ataaccatga
gtgataacac tgcggccaac ttacttctga 540caacgatcgg aggaccgaag gagctaaccg
cttttttgca caacatgggg gatcatgtaa 600ctcgccttga tcgttgggaa ccggagctga
atgaagccat accaaacgac gagcgtgaca 660ccacgatgcc tgtagcaatg gcaacaacgt
tgcgcaaact attaactggc gaactactta 720ctctagcttc ccggcaacaa ttaatagact
ggatggaggc ggataaagtt gcaggaccac 780ttctgcgctc ggcccttccg gctggctggt
ttattgctga taaatctgga gccggtgagc 840gtgggtctcg cggtatcatt gcagcactgg
ggccagatgg taagccctcc cgtatcgtag 900ttatctacac gacggggagt caggcaacta
tggatgaacg aaatagacag atcgctgaga 960taggtgcctc actgattaag cattggtaac
tgtcagacca agtttactca tatatacttt 1020agattgattt aaaacttcat ttttaattta
aaaggatcta ggtgaagatc ctttttgata 1080atctcatgac caaaatccct taacgtgagt
tttcgttcca ctgagcgtca gaccccgtag 1140aaaagatcaa aggatcttct tgagatcctt
tttttctgcg cgtaatctgc tgcttgcaaa 1200caaaaaaacc accgctacca gcggtggttt
gtttgccgga tcaagagcta ccaactcttt 1260ttccgaaggt aactggcttc agcagagcgc
agataccaaa tactgtcctt ctagtgtagc 1320cgtagttagg ccaccacttc aagaactctg
tagcaccgcc tacatacctc gctctgctaa 1380tcctgttacc agtggctgct gccagtggcg
ataagtcgtg tcttaccggg ttggactcaa 1440gacgatagtt accggataag gcgcagcggt
cgggctgaac ggggggttcg tgcacacagc 1500ccagcttgga gcgaacgacc tacaccgaac
tgagatacct acagcgtgag ctatgagaaa 1560gcgccacgct tcccgaaggg agaaaggcgg
acaggtatcc ggtaagcggc agggtcggaa 1620caggagagcg cacgagggag cttccagggg
gaaacgcctg gtatctttat agtcctgtcg 1680ggtttcgcca cctctgactt gagcgtcgat
ttttgtgatg ctcgtcaggg gggcggagcc 1740tatggaaaaa cgccagcaac gcggcctttt
tacggttcct ggccttttgc tggccttttg 1800ctcacatgtt ctttcctgcg ttatcccctg
attctgtgga taaccgtatt accgcctttg 1860agtgagctga taccgctcgc cgcagccgaa
cgaccgagcg cagcgagtca gtgagcgagg 1920aagcggaaga gcgcccaata cgcaaaccgc
ctctccccgc gcgttggccg attcattaat 1980gcagctggca cgacaggttt cccgactgga
aagcgggcag tgagcgcaac gcaattaatg 2040tgagttagct cactcattag gcaccccagg
ctttacactt tatgcttccg gctcgtatgt 2100tgtgtggaat tgtgagcgga taacaatttc
acacaggaaa cagctatgac catgattacg 2160ccaagcgcgc aattaaccct cactaaaggg
aacaaaagct ggagctccac cgcggtggcg 2220gccgctctag aactagtgga tcccccgggc
tgcaggaatt cgaggagaaa ttaaccatgt 2280atatcgggat agatcttggc acctcgggcg
taaaagttat tttgctcaac gagcagggtg 2340aggtggttgc tgcgcaaacg gaaaagctga
ccgtttcgcg cccgcatcca ctctggtcgg 2400aacaagaccc ggaacagtgg tggcaggcaa
ctgatcgcgc aatgaaagct ctgggcgatc 2460agcattctct gcaggacgtt aaagcattgg
gtattgccgg ccagatgcac ggagcaacct 2520tgctggatgc tcagcaacgg gtgttacgcc
ctgccatttt gtggaacgac gggcgctgtg 2580cgcaagagtg cactttgctg gaagcgcgag
ttccgcaatc gcgggtgatt accggcaacc 2640tgatgatgcc cggatttact gcgcctaaat
tgctatgggt tcagcggcat gagccggaga 2700tattccgtca aatcgacaaa gtattattac
cgaaagatta cttgcgtctg cgtatgacgg 2760gggagtttgc cagcgatatg tctgacgcag
ctggcaccat gtggctggat gtcgcaaagc 2820gtgactggag tgacgtcatg ctgcaggctt
gcgacttatc tcgtgaccag atgcccgcat 2880tatacgaagg cagcgaaatt actggtgctt
tgttacctga agttgcgaaa gcgtggggta 2940tggcgacggt gccagttgtc gcaggcggtg
gcgacaatgc agctggtgca gttggtgtgg 3000gaatggttga tgctaatcag gcaatgttat
cgctggggac gtcgggggtc tattttgctg 3060tcagcgaagg gttcttaagc aagccagaaa
gcgccgtaca tagcttttgc catgcgctac 3120cgcaacgttg gcatttaatg tctgtgatgc
tgagtgcagc gtcgtgtctg gattgggccg 3180cgaaattaac cggcctgagc aatgtcccag
ctttaatcgc tgcagctcaa caggctgatg 3240aaagtgccga gccagtttgg tttctgcctt
atctttccgg cgagcgtacg ccacacaata 3300atccccaggc gaagggggtt ttctttggtt
tgactcatca acatggcccc aatgaactgg 3360cgcgagcagt gctggaaggc gtgggttatg
cgctggcaga tggcatggat gtcgtgcatg 3420cctgcggtat taaaccgcaa agtgttacgt
tgattggggg cggggcgcgt agtgagtact 3480ggcgtcagat gctggcggat atcagcggtc
agcagctcga ttaccgtacg gggggggatg 3540tggggccagc actgggcgca gcaaggctgg
cgcagatcgc ggcgaatcca gagaaatcgc 3600tcattgaatt gttgccgcaa ctaccgttag
aacagtcgca tctaccagat gcgcagcgtt 3660atgccgctta tcagccacga cgagaaacgt
tccgtcgcct ctatcagcaa cttctgccat 3720taatggcgta aaagcttgag gagaaattaa
ccatgaagca actcaccatt ctgggctcga 3780ccggctcgat tggttgcagc acgctggacg
tggtgcgcca taatcccgaa cacttccgcg 3840tagttgcgct ggtggcaggc aaaaatgtca
ctcgcatggt agaacagtgc ctggaattct 3900ctccccgcta tgccgtaatg gacgatgaag
cgagtgcgaa acttcttaaa acgatgctac 3960agcaacaggg tagccgcacc gaagtcttaa
gtgggcaaca agccgcttgc gatatggcag 4020cgcttgagga tgttgatcag gtgatggcag
ccattgttgg cgctgctggg ctgttaccta 4080cgcttgctgc gatccgcgcg ggtaaaacca
ttttgctggc caataaagaa tcactggtta 4140cctgcggacg tctgtttatg gacgccgtaa
agcagagcaa agcgcaattg ttaccggtcg 4200atagcgaaca taacgccatt tttcagagtt
taccgcaacc tatccagcat aatctgggat 4260acgctgacct tgagcaaaat ggcgtggtgt
ccattttact taccgggtct ggtggccctt 4320tccgtgagac gccattgcgc gatttggcaa
caatgacgcc ggatcaagcc tgccgtcatc 4380cgaactggtc gatggggcgt aaaatttctg
tcgattcggc taccatgatg aacaaaggtc 4440tggaatacat tgaagcgcgt tggctgttta
acgccagcgc cagccagatg gaagtgctga 4500ttcacccgca gtcagtgatt cactcaatgg
tgcgctatca ggacggcagt gttctggcgc 4560agctggggga accggatatg cgtacgccaa
ttgcccacac catggcatgg ccgaatcgcg 4620tgaactctgg cgtgaagccg ctcgattttt
gcaaactaag tgcgttgaca tttgccgcac 4680cggattatga tcgttatcca tgcctgaaac
tggcgatgga ggcgttcgaa caaggccagg 4740cagcgacgac agcattgaat gccgcaaacg
aaatcaccgt tgctgctttt cttgcgcaac 4800aaatccgctt tacggatatc gctgcgttga
atttatccgt actggaaaaa atggatatgc 4860gcgaaccaca atgtgtggac gatgtgttat
ctgttgatgc gaacgcgcgt gaagtcgcca 4920gaaaagaggt gatgcgtctc gcaagctgag
tcgacgagga gaaattaacc atggcaacca 4980ctcatttgga tgtttgcgcc gtggttccgg
cggccggatt tggccgtcga atgcaaacgg 5040aatgtcctaa gcaatatctc tcaatcggta
atcaaaccat tcttgaacac tcggtgcatg 5100cgctgctggc gcatccccgg gtgaaacgtg
tcgtcattgc cataagtcct ggcgatagcc 5160gttttgcaca acttcctctg gcgaatcatc
cgcaaatcac cgttgtagat ggcggtgatg 5220agcgtgccga ttccgtgctg gcaggtctga
aagccgctgg cgacgcgcag tgggtattgg 5280tgcatgacgc cgctcgtcct tgtttgcatc
aggatgacct cgcgcgattg ttggcgttga 5340gcgaaaccag ccgcacgggg gggatcctcg
ccgcaccagt gcgcgatact atgaaacgtg 5400ccgaaccggg caaaaatgcc attgctcata
ccgttgatcg caacggctta tggcacgcgc 5460tgacgccgca atttttccct cgtgagctgt
tacatgactg tctgacgcgc gctctaaatg 5520aaggcgcgac tattaccgac gaagcctcgg
cgctggaata ttgcggattc catcctcagt 5580tggtcgaagg ccgtgcggat aacattaaag
tcacgcgccc ggaagatttg gcactggccg 5640agttttacct cacccgaacc atccatcagg
agaatacata atgcgaattg gacacggttt 5700tgacgtacat gcctttggcg gtgaaggccc
aattatcatt ggtggcgtac gcattcctta 5760cgaaaaagga ttgctggcgc attctgatgg
cgacgtggcg ctccatgcgt tgaccgatgc 5820attgcttggc gcggcggcgc tgggggatat
cggcaagctg ttcccggata ccgatccggc 5880atttaaaggt gccgatagcc gcgagctgct
acgcgaagcc tggcgtcgta ttcaggcgaa 5940gggttatacc cttggcaacg tcgatgtcac
tatcatcgct caggcaccga agatgttgcc 6000gcacattcca caaatgcgcg tgtttattgc
cgaagatctc ggctgccata tggatgatgt 6060taacgtgaaa gccactacta cggaaaaact
gggatttacc ggacgtgggg aagggattgc 6120ctgtgaagcg gtggcgctac tcattaaggc
aacaaaatga ctcgaggagg agaaattaac 6180catgcggaca cagtggccct ctccggcaaa
acttaatctg tttttataca ttaccggtca 6240gcgtgcggat ggttaccaca cgctgcaaac
gctgtttcag tttcttgatt acggcgacac 6300catcagcatt gagcttcgtg acgatgggga
tattcgtctg ttaacgcccg ttgaaggcgt 6360ggaacatgaa gataacctga tcgttcgcgc
agcgcgattg ttgatgaaaa ctgcggcaga 6420cagcgggcgt cttccgacgg gaagcggtgc
gaatatcagc attgacaagc gtttgccgat 6480gggcggcggt ctcggcggtg gttcatccaa
tgccgcgacg gtcctggtgg cattaaatca 6540tctctggcaa tgcgggctaa gcatggatga
gctggcggaa atggggctga cgctgggcgc 6600agatgttcct gtctttgttc gggggcatgc
cgcgtttgcc gaaggcgttg gtgaaatact 6660aacgccggtg gatccgccag agaagtggta
tctggtggcg caccctggtg taagtattcc 6720gactccggtg atttttaaag atcctgaact
cccgcgcaat acgccaaaaa ggtcaataga 6780aacgttgcta aaatgtgaat tcagcaatga
ttgcgaggtt atcgcaagaa aacgttttcg 6840cgaggttgat gcggtgcttt cctggctgtt
agaatacgcc ccgtcgcgcc tgactgggac 6900aggggcctgt gtctttgctg aatttgatac
agagtctgaa gcccgccagg tgctagagca 6960agccccggaa tggctcaatg gctttgtggc
gaaaggcgct aatctttccc cattgcacag 7020agccatgctt taaggtaccc aattcgccct
atagtgagtc gtattacgcg cgctcactgg 7080ccgtcgtttt acaacgtcgt gactgggaaa
accctggcgt tacccaactt aatcgccttg 7140cagcacatcc ccctttcgcc agctggcgta
atagcgaaga ggcccgcacc gatcgccctt 7200cccaacagtt gcgcagcctg aatggcgaat
ggaaattgta agcgttaata ttttgttaaa 7260attcgcgtta aatttttgtt aaatcagctc
attttttaac caataggccg aaatcggcaa 7320aatcccttat aaatcaaaag aatagaccga
gatagggttg agtgttgttc cagtttggaa 7380caagagtcca ctattaaaga acgtggactc
caacgtcaaa gggcgaaaaa ccgtctatca 7440gggcgatggc ccactacgtg aaccatcacc
ctaatcaagt tttttggggt cgaggtgccg 7500taaagcacta aatcggaacc ctaaagggag
cccccgattt agagcttgac ggggaaagcc 7560ggcgaacgtg gcgagaaagg aagggaagaa
agcgaaagga gcgggcgcta gggcgctggc 7620aagtgtagcg gtcacgctgc gcgtaaccac
cacacccgcc gcgcttaatg cgccgctaca 7680gggcgcgtca g
7691445109DNAArtificialpACYCgcpE
44gaattccgga tgagcattca tcaggcgggc aagaatgtga ataaaggccg gataaaactt
60gtgcttattt ttctttacgg tctttaaaaa ggccgtaata tccagctgaa cggtctggtt
120ataggtacat tgagcaactg actgaaatgc ctcaaaatgt tctttacgat gccattggga
180tatatcaacg gtggtatatc cagtgatttt tttctccatt ttagcttcct tagctcctga
240aaatctcgat aactcaaaaa atacgcccgg tagtgatctt atttcattat ggtgaaagtt
300ggaacctctt acgtgccgat caacgtctca ttttcgccaa aagttggccc agggcttccc
360ggtatcaaca gggacaccag gatttattta ttctgcgaag tgatcttccg tcacaggtat
420ttattcggcg caaagtgcgt cgggtgatgc tgccaactta ctgatttagt gtatgatggt
480gtttttgagg tgctccagtg gcttctgttt ctatcagctg tccctcctgt tcagctactg
540acggggtggt gcgtaacggc aaaagcaccg ccggacatca gcgctagcgg agtgtatact
600ggcttactat gttggcactg atgagggtgt cagtgaagtg cttcatgtgg caggagaaaa
660aaggctgcac cggtgcgtca gcagaatatg tgatacagga tatattccgc ttcctcgctc
720actgactcgc tacgctcggt cgttcgactg cggcgagcgg aaatggctta cgaacggggc
780ggagatttcc tggaagatgc caggaagata cttaacaggg aagtgagagg gccgcggcaa
840agccgttttt ccataggctc cgcccccctg acaagcatca cgaaatctga cgctcaaatc
900agtggtggcg aaacccgaca ggactataaa gataccaggc gtttccccct ggcggctccc
960tcgtgcgctc tcctgttcct gcctttcggt ttaccggtgt cattccgctg ttatggccgc
1020gtttgtctca ttccacgcct gacactcagt tccgggtagg cagttcgctc caagctggac
1080tgtatgcacg aaccccccgt tcagtccgac cgctgcgcct tatccggtaa ctatcgtctt
1140gagtccaacc cggaaagaca tgcaaaagca ccactggcag cagccactgg taattgattt
1200agaggagtta gtcttgaagt catgcgccgg ttaaggctaa actgaaagga caagttttgg
1260tgactgcgct cctccaagcc agttacctcg gttcaaagag ttggtagctc agagaacctt
1320cgaaaaaccg ccctgcaagg cggttttttc gttttcagag caagagatta cgcgcagacc
1380aaaacgatct caagaagatc atcttattaa tcagataaaa tatttctaga tttcagtgca
1440atttatctct tcaaatgtag cacctgaagt cagccccata cgatataagt tgtaattctc
1500atgtttgaca gcttatcatc gataagcttt aatgcggtag tttatcacag ttaaattgct
1560aacgcagtca ggcaccgtgt atgaaatcta acaatgcgct catcgtcatc ctcggcaccg
1620tcaccctgga tgctgtaggc ataggcttgg ttatgccggt actgccgggc ctcttgcggg
1680atatcgtcca ttccgacagc atcgccagtc actatggcgt gctgctagcg ctatatgcgt
1740tgatgcaatt tctatgcgca cccgttctcg gagcactgtc cgaccgcttt ggccgccgcc
1800cagtcctgct cgcttcgcta cttggagcca ctatcgacta cgcgatcatg gcgaccacac
1860ccgtcctgtg gatccgagga gaaattaacc atgcataacc aggctccaat tcaacgtaga
1920aaatcaacac gtatttacgt tgggaatgtg ccgattggcg atggtgctcc catcgccgta
1980cagtccatga ccaatacgcg tacgacagac gtcgaagcaa cggtcaatca aatcaaggcg
2040ctggaacgcg ttggcgctga tatcgtccgt gtatccgtac cgacgatgga cgcggcagaa
2100gcgttcaaac tcatcaaaca gcaggttaac gtgccgctgg tggctgacat ccacttcgac
2160tatcgcattg cgctgaaagt agcggaatac ggcgtcgatt gtctgcgtat taaccctggc
2220aatatcggta atgaagagcg tattcgcatg gtggttgact gtgcgcgcga taaaaacatt
2280ccgatccgta ttggcgttaa cgccggatcg ctggaaaaag atctgcaaga aaagtatggc
2340gaaccgacgc cgcaggcgtt gctggaatct gccatgcgtc atgttgatca tctcgatcgc
2400ctgaacttcg atcagttcaa agtcagcgtg aaagcgtctg acgtcttcct cgctgttgag
2460tcttatcgtt tgctggcaaa acagatcgat cagccgttgc atctggggat caccgaagcc
2520ggtggtgcgc gcagcggggc agtaaaatcc gccattggtt taggtctgct gctgtctgaa
2580ggcatcggcg acacgctgcg cgtatcgctg gcggccgatc cggtcgaaga gatcaaagtc
2640ggtttcgata ttttgaaatc gctgcgtatc cgttcgcgag ggatcaactt catcgcctgc
2700ccgacctgtt cgcgtcagga atttgatgtt atcggtacgg ttaacgcgct ggagcaacgc
2760ctggaagata tcatcactcc gatggacgtt tcgattatcg gctgcgtggt gaatggccca
2820ggtgaggcgc tggtttctac actcggcgtc accggcggca acaagaaaag cggcctctat
2880gaagatggcg tgcgcaaaga ccgtctggac aacaacgata tgatcgacca gctggaagca
2940cgcattcgtg cgaaagccag tcagctggac gaagcgcgtc gaattgacgt tcagcaggtt
3000gaaaaataag tcgaccgatg cccttgagag ccttcaaccc agtcagctcc ttccggtggg
3060cgcggggcat gactatcgtc gccgcactta tgactgtctt ctttatcatg caactcgtag
3120gacaggtgcc ggcagcgctc tgggtcattt tcggcgagga ccgctttcgc tggagcgcga
3180cgatgatcgg cctgtcgctt gcggtattcg gaatcttgca cgccctcgct caagccttcg
3240tcactggtcc cgccaccaaa cgtttcggcg agaagcaggc cattatcgcc ggcatggcgg
3300ccgacgcgct gggctacgtc ttgctggcgt tcgcgacgcg aggctggatg gccttcccca
3360ttatgattct tctcgcttcc ggcggcatcg ggatgcccgc gttgcaggcc atgctgtcca
3420ggcaggtaga tgacgaccat cagggacagc ttcaaggatc gctcgcggct cttaccagcc
3480taacttcgat cactggaccg ctgatcgtca cggcgattta tgccgcctcg gcgagcacat
3540ggaacgggtt ggcatggatt gtaggcgccg ccctatacct tgtctgcctc cccgcgttgc
3600gtcgcggtgc atggagccgg gccacctcga cctgaatgga agccggcggc acctcgctaa
3660cggattcacc actccaagaa ttggagccaa tcaattcttg cggagaactg tgaatgcgca
3720aaccaaccct tggcagaaca tatccatcgc gtccgccatc tccagcagcc gcacgcggcg
3780catctcgggc agcgttgggt cctggccacg ggtgcgcatg atcgtgctcc tgtcgttgag
3840gacccggcta ggctggcggg gttgccttac tggttagcag aatgaatcac cgatacgcga
3900gcgaacgtga agcgactgct gctgcaaaac gtctgcgacc tgagcaacaa catgaatggt
3960cttcggtttc cgtgtttcgt aaagtctgga aacgcggaag tcccctacgt gctgctgaag
4020ttgcccgcaa cagagagtgg aaccaaccgg tgataccacg atactatgac tgagagtcaa
4080cgccatgagc ggcctcattt cttattctga gttacaacag tccgcaccgc tgtccggtag
4140ctccttccgg tgggcgcggg gcatgactat cgtcgccgca cttatgactg tcttctttat
4200catgcaactc gtaggacagg tgccggcagc gcccaacagt cccccggcca cggggcctgc
4260caccataccc acgccgaaac aagcgccctg caccattatg ttccggatct gcatcgcagg
4320atgctgctgg ctaccctgtg gaacacctac atctgtatta acgaagcgct aaccgttttt
4380atcaggctct gggaggcaga ataaatgatc atatcgtcaa ttattacctc cacggggaga
4440gcctgagcaa actggcctca ggcatttgag aagcacacgg tcacactgct tccggtagtc
4500aataaaccgg taaaccagca atagacataa gcggctattt aacgaccctg ccctgaaccg
4560acgaccgggt cgaatttgct ttcgaatttc tgccattcat ccgcttatta tcacttattc
4620aggcgtagca ccaggcgttt aagggcacca ataactgcct taaaaaaatt acgccccgcc
4680ctgccactca tcgcagtact gttgtaattc attaagcatt ctgccgacat ggaagccatc
4740acagacggca tgatgaacct gaatcgccag cggcatcagc accttgtcgc cttgcgtata
4800atatttgccc atggtgaaaa cgggggcgaa gaagttgtcc atattggcca cgtttaaatc
4860aaaactggtg aaactcaccc agggattggc tgagacgaaa aacatattct caataaaccc
4920tttagggaaa taggccaggt tttcaccgta acacgccaca tcttgcgaat atatgtgtag
4980aaactgccgg aaatcgtcgt ggtattcact ccagagcgat gaaaacgttt cagtttgctc
5040atggaaaacg gtgtaacaag ggtgaacact atcccatatc accagctcac cgtctttcat
5100tgccatacg
5109457494DNAArtificialpBScaro14 45gtggcacttt tcggggaaat gtgcgcggaa
cccctatttg tttatttttc taaatacatt 60caaatatgta tccgctcatg agacaataac
cctgataaat gcttcaataa tattgaaaaa 120ggaagagtat gagtattcaa catttccgtg
tcgcccttat tccctttttt gcggcatttt 180gccttcctgt ttttgctcac ccagaaacgc
tggtgaaagt aaaagatgct gaagatcagt 240tgggtgcacg agtgggttac atcgaactgg
atctcaacag cggtaagatc cttgagagtt 300ttcgccccga agaacgtttt ccaatgatga
gcacttttaa agttctgcta tgtggcgcgg 360tattatcccg tattgacgcc gggcaagagc
aactcggtcg ccgcatacac tattctcaga 420atgacttggt tgagtactca ccagtcacag
aaaagcatct tacggatggc atgacagtaa 480gagaattatg cagtgctgcc ataaccatga
gtgataacac tgcggccaac ttacttctga 540caacgatcgg aggaccgaag gagctaaccg
cttttttgca caacatgggg gatcatgtaa 600ctcgccttga tcgttgggaa ccggagctga
atgaagccat accaaacgac gagcgtgaca 660ccacgatgcc tgtagcaatg gcaacaacgt
tgcgcaaact attaactggc gaactactta 720ctctagcttc ccggcaacaa ttaatagact
ggatggaggc ggataaagtt gcaggaccac 780ttctgcgctc ggcccttccg gctggctggt
ttattgctga taaatctgga gccggtgagc 840gtgggtctcg cggtatcatt gcagcactgg
ggccagatgg taagccctcc cgtatcgtag 900ttatctacac gacggggagt caggcaacta
tggatgaacg aaatagacag atcgctgaga 960taggtgcctc actgattaag cattggtaac
tgtcagacca agtttactca tatatacttt 1020agattgattt aaaacttcat ttttaattta
aaaggatcta ggtgaagatc ctttttgata 1080atctcatgac caaaatccct taacgtgagt
tttcgttcca ctgagcgtca gaccccgtag 1140aaaagatcaa aggatcttct tgagatcctt
tttttctgcg cgtaatctgc tgcttgcaaa 1200caaaaaaacc accgctacca gcggtggttt
gtttgccgga tcaagagcta ccaactcttt 1260ttccgaaggt aactggcttc agcagagcgc
agataccaaa tactgtcctt ctagtgtagc 1320cgtagttagg ccaccacttc aagaactctg
tagcaccgcc tacatacctc gctctgctaa 1380tcctgttacc agtggctgct gccagtggcg
ataagtcgtg tcttaccggg ttggactcaa 1440gacgatagtt accggataag gcgcagcggt
cgggctgaac ggggggttcg tgcacacagc 1500ccagcttgga gcgaacgacc tacaccgaac
tgagatacct acagcgtgag ctatgagaaa 1560gcgccacgct tcccgaaggg agaaaggcgg
acaggtatcc ggtaagcggc agggtcggaa 1620caggagagcg cacgagggag cttccagggg
gaaacgcctg gtatctttat agtcctgtcg 1680ggtttcgcca cctctgactt gagcgtcgat
ttttgtgatg ctcgtcaggg gggcggagcc 1740tatggaaaaa cgccagcaac gcggcctttt
tacggttcct ggccttttgc tggccttttg 1800ctcacatgtt ctttcctgcg ttatcccctg
attctgtgga taaccgtatt accgcctttg 1860agtgagctga taccgctcgc cgcagccgaa
cgaccgagcg cagcgagtca gtgagcgagg 1920aagcggaaga gcgcccaata cgcaaaccgc
ctctccccgc gcgttggccg attcattaat 1980gcagctggca cgacaggttt cccgactgga
aagcgggcag tgagcgcaac gcaattaatg 2040tgagttagct cactcattag gcaccccagg
ctttacactt tatgcttccg gctcgtatgt 2100tgtgtggaat tgtgagcgga taacaatttc
acacaggaaa cagctatgac catgattacg 2160ccaagcgcgc aattaaccct cactaaaggg
aacaaaagct ggagctccac cgcggtggcg 2220gccgctctag aactagtgga tcccccgggc
tgcaggaatt gccgtaaatg tatccgttta 2280taaggacagc ccgaatgacg gtctgcgcaa
aaaaacacgt tcatctcact cgcgatgctg 2340cggagcagtt actggctgat attgatcgac
gccttgatca gttattgccc gtggagggag 2400aacgggatgt tgtgggtgcc gcgatgcgtg
aaggtgcgct ggcaccggga aaacgtattc 2460gccccatgtt gctgttgctg accgcccgcg
atctgggttg cgctgtcagc catgacggat 2520tactggattt ggcctgtgcg gtggaaatgg
tccacgcggc ttcgctgatc cttgacgata 2580tgccctgcat ggacgatgcg aagctgcggc
gcggacgccc taccattcat tctcattacg 2640gagagcatgt ggcaatactg gcggcggttg
ccttgctgag taaagccttt ggcgtaattg 2700ccgatgcaga tggcctcacg ccgctggcaa
aaaatcgggc ggtttctgaa ctgtcaaacg 2760ccatcggcat gcaaggattg gttcagggtc
agttcaagga tctgtctgaa ggggataagc 2820cgcgcagcgc tgaagctatt ttgatgacga
atcactttaa aaccagcacg ctgttttgtg 2880cctccatgca gatggcctcg attgttgcga
atgcctccag cgaagcgcgt gattgcctgc 2940atcgtttttc acttgatctt ggtcaggcat
ttcaactgct ggacgatttg accgatggca 3000tgaccgacac cggtaaggat agcaatcagg
acgccggtaa atcgacgctg gtcaatctgt 3060taggcccgag ggcggttgaa gaacgtctga
gacaacatct tcagcttgcc agtgagcatc 3120tctctgcggc ctgccaacac gggcacgcca
ctcaacattt tattcaggcc tggtttgaca 3180aaaaactcgc tgccgtcagt taagcttatg
tgcaccggtc agcctgtctt aagtgggagc 3240ggctatgcaa ccgcattatg atctgattct
cgtgggggct ggactcgcga atggccttat 3300cgccctgcgt cttcagcagc agcaacctga
tatgcgtatt ttgcttatcg acgccgcacc 3360ccaggcgggc gggaatcata cgtggtcatt
tcaccacgat gatttgactg agagccaaca 3420tcgttggata gctccgctgg tggttcatca
ctggcccgac tatcaggtac gctttcccac 3480acgccgtcgt aagctgaaca gcggctactt
ttgtattact tctcagcgtt tcgctgaggt 3540tttacagcga cagtttggcc cgcacttgtg
gatggatacc gcggtcgcag aggttaatgc 3600ggaatctgtt cggttgaaaa agggtcaggt
tatcggtgcc cgcgcggtga ttgacgggcg 3660gggttatgcg gcaaattcag cactgagcgt
gggcttccag gcgtttattg gccaggaatg 3720gcgattgagc cacccgcatg gtttatcgtc
tcccattatc atggatgcca cggtcgatca 3780gcaaaatggt tatcgcttcg tgtacagcct
gccgctctcg ccgaccagat tgttaattga 3840agacacgcac tatattgata atgcgacatt
agatcctgaa tgcgcgcggc aaaatatttg 3900cgactatgcc gcgcaacagg gttggcagct
tcagacactg ctgcgagaag aacagggcgc 3960cttacccatt actctgtcgg gcaatgccga
cgcattctgg cagcagcgcc ccctggcctg 4020tagtggatta cgtgccggtc tgttccatcc
taccaccggc tattcactgc cgctggcggt 4080tgccgtggcc gaccgcctga gtgcacttga
tgtctttacg tcggcctcaa ttcaccatgc 4140cattacgcat tttgcccgcg agcgctggca
gcagcagggc tttttccgca tgctgaatcg 4200catgctgttt ttagccggac ccgccgattc
acgctggcgg gttatgcagc gtttttatgg 4260tttacctgaa gatttaattg cccgttttta
tgcgggaaaa ctcacgctga ccgatcggct 4320acgtattctg agcggcaagc cgcctgttcc
ggtattagca gcattgcaag ccattatgac 4380gactcatcgt taaagagcga ctacatgaaa
ccaactacgg taattggtgc aggcttcggt 4440ggcctggcac tggcaattcg tctacaagct
gcggggatcc ccgtcttact gcttgaacaa 4500cgtgataaac ccggcggtcg ggcttatgtc
tacgaggatc aggggtttac ctttgatgca 4560ggcccgacgg ttatcaccga tcccagtgcc
attgaagaac tgtttgcact ggcaggaaaa 4620cagttaaaag agtatgtcga actgctgccg
gttacgccgt tttaccgcct gtgttgggag 4680tcagggaagg tctttaatta cgataacgat
caaacccggc tcgaagcgca gattcagcag 4740tttaatcccc gcgatgtcga aggttatcgt
cagtttctgg actattcacg cgcggtgttt 4800aaagaaggct atctaaagct cggtactgtc
ccttttttat cgttcagaga catgcttcgc 4860gccgcacctc aactggcgaa actgcaggca
tggagaagcg tttacagtaa ggttgccagt 4920tacatcgaag atgaacatct gcgccaggcg
ttttctttcc actcgctgtt ggtgggcggc 4980aatcccttcg ccacctcatc catttatacg
ttgatacacg cgctggagcg tgagtggggc 5040gtctggtttc cgcgtggcgg caccggcgca
ttagttcagg ggatgataaa gctgtttcag 5100gatctgggtg gcgaagtcgt gttaaacgcc
agagtcagcc atatggaaac gacaggaaac 5160aagattgaag ccgtgcattt agaggacggt
cgcaggttcc tgacgcaagc cgtcgcgtca 5220aatgcagatg tggttcatac ctatcgcgac
ctgttaagcc agcaccctgc cgcggttaag 5280cagtccaaca aactgcagac taagcgcatg
agtaactctc tgtttgtgct ctattttggt 5340ttgaatcacc atcatgatca gctcgcgcat
cacacggttt gtttcggccc gcgttaccgc 5400gagctgattg acgaaatttt taatcatgat
ggcctcgcag aggacttctc actttatctg 5460cacgcgccct gtgtcacgga ttcgtcactg
gcgcctgaag gttgcggcag ttactatgtg 5520ttggcgccgg tgccgcattt aggcaccgcg
aacctcgact ggacggttga ggggccaaaa 5580ctacgcgacc gtatttttgc gtaccttgag
cagcattaca tgcctggctt acggagtcag 5640ctggtcacgc accggatgtt tacgccgttt
gattttcgcg accagcttaa tgcctatcat 5700ggctcagcct tttctgtgga gcccgttctt
acccagagcg cctggtttcg gccgcataac 5760cgcgataaaa ccattactaa tctctacctg
gtcggcgcag gcacgcatcc cggcgcaggc 5820attcctggcg tcatcggctc ggcaaaagcg
acagcaggtt tgatgctgga ggatctgatt 5880tgaataatcc gtcgttactc aatcatgcgg
tcgaaacgat ggcagttggc tcgaaaagtt 5940ttgcgacagc ctcaaagtta tttgatgcaa
aaacccggcg cagcgtactg atgctctacg 6000cctggtgccg ccattgtgac gatgttattg
acgatcagac gctgggcttt caggcccggc 6060agcctgcctt acaaacgccc gaacaacgtc
tgatgcaact tgagatgaaa acgcgccagg 6120cctatgcagg atcgcagatg cacgaaccgg
cgtttgcggc ttttcaggaa gtggctatgg 6180ctcatgatat cgccccggct tacgcgtttg
atcatctgga aggcttcgcc atggatgtac 6240gcgaagcgca atacagccaa ctggatgata
cgctgcgcta ttgctatcac gttgcaggcg 6300ttgtcggctt gatgatggcg caaatcatgg
gcgtgcggga taacgccacg ctggaccgcg 6360cctgtgacct tgggctggca tttcagttga
ccaatattgc tcgcgatatt gtggacgatg 6420cgcatgcggg ccgctgttat ctgccggcaa
gctggctgga gcatgaaggt ctgaacaaag 6480agaattatgc ggcacctgaa aaccgtcagg
cgctgagccg tatcgcccgt cgtttggtgc 6540aggaagcaga accttactat ttgtctgcca
cagccggcct ggcagggttg cccctgcgtt 6600ccgcctgggc aatcgctacg gcgaagcagg
tttaccggaa aataggtgtc aaagttgaac 6660aggccggtca gcaagcctgg gatcagcggc
agtcaacgac cacgcccgaa aaattaacgc 6720tgctgctggc cgcctctggt caggccctta
cttcccggat gcgggctcat cctccccgcc 6780ctgcgcatct ctggcagcgc ccgctctagc
gccatgtcga cctcgagggg gggcccggta 6840cccaattcgc cctatagtga gtcgtattac
gcgcgctcac tggccgtcgt tttacaacgt 6900cgtgactggg aaaaccctgg cgttacccaa
cttaatcgcc ttgcagcaca tccccctttc 6960gccagctggc gtaatagcga agaggcccgc
accgatcgcc cttcccaaca gttgcgcagc 7020ctgaatggcg aatggaaatt gtaagcgtta
atattttgtt aaaattcgcg ttaaattttt 7080gttaaatcag ctcatttttt aaccaatagg
ccgaaatcgg caaaatccct tataaatcaa 7140aagaatagac cgagataggg ttgagtgttg
ttccagtttg gaacaagagt ccactattaa 7200agaacgtgga ctccaacgtc aaagggcgaa
aaaccgtcta tcagggcgat ggcccactac 7260gtgaaccatc accctaatca agttttttgg
ggtcgaggtg ccgtaaagca ctaaatcgga 7320accctaaagg gagcccccga tttagagctt
gacggggaaa gccggcgaac gtggcgagaa 7380aggaagggaa gaaagcgaaa ggagcgggcg
ctagggcgct ggcaagtgta gcggtcacgc 7440tgcgcgtaac caccacaccc gccgcgctta
atgcgccgct acagggcgcg tcag 7494468547DNAArtificialpACYCcaro14
46gaattccgga tgagcattca tcaggcgggc aagaatgtga ataaaggccg gataaaactt
60gtgcttattt ttctttacgg tctttaaaaa ggccgtaata tccagctgaa cggtctggtt
120ataggtacat tgagcaactg actgaaatgc ctcaaaatgt tctttacgat gccattggga
180tatatcaacg gtggtatatc cagtgatttt tttctccatt ttagcttcct tagctcctga
240aaatctcgat aactcaaaaa atacgcccgg tagtgatctt atttcattat ggtgaaagtt
300ggaacctctt acgtgccgat caacgtctca ttttcgccaa aagttggccc agggcttccc
360ggtatcaaca gggacaccag gatttattta ttctgcgaag tgatcttccg tcacaggtat
420ttattcggcg caaagtgcgt cgggtgatgc tgccaactta ctgatttagt gtatgatggt
480gtttttgagg tgctccagtg gcttctgttt ctatcagctg tccctcctgt tcagctactg
540acggggtggt gcgtaacggc aaaagcaccg ccggacatca gcgctagcgg agtgtatact
600ggcttactat gttggcactg atgagggtgt cagtgaagtg cttcatgtgg caggagaaaa
660aaggctgcac cggtgcgtca gcagaatatg tgatacagga tatattccgc ttcctcgctc
720actgactcgc tacgctcggt cgttcgactg cggcgagcgg aaatggctta cgaacggggc
780ggagatttcc tggaagatgc caggaagata cttaacaggg aagtgagagg gccgcggcaa
840agccgttttt ccataggctc cgcccccctg acaagcatca cgaaatctga cgctcaaatc
900agtggtggcg aaacccgaca ggactataaa gataccaggc gtttccccct ggcggctccc
960tcgtgcgctc tcctgttcct gcctttcggt ttaccggtgt cattccgctg ttatggccgc
1020gtttgtctca ttccacgcct gacactcagt tccgggtagg cagttcgctc caagctggac
1080tgtatgcacg aaccccccgt tcagtccgac cgctgcgcct tatccggtaa ctatcgtctt
1140gagtccaacc cggaaagaca tgcaaaagca ccactggcag cagccactgg taattgattt
1200agaggagtta gtcttgaagt catgcgccgg ttaaggctaa actgaaagga caagttttgg
1260tgactgcgct cctccaagcc agttacctcg gttcaaagag ttggtagctc agagaacctt
1320cgaaaaaccg ccctgcaagg cggttttttc gttttcagag caagagatta cgcgcagacc
1380aaaacgatct caagaagatc atcttattaa tcagataaaa tatttctaga tttcagtgca
1440atttatctct tcaaatgtag cacctgaagt cagccccata cgatataagt tgtaattctc
1500atgtttgaca gcttatcatc gataagcttt aatgcggtag tttatcacag ttaaattgct
1560aacgcagtca ggcaccgtgt atgaaatcta acaatgcgct catcgtcatc ctcggcaccg
1620tcaccctgga tgctgtaggc ataggcttgg ttatgccggt actgccgggc ctcttgcggg
1680atatcgtcca ttccgacagc atcgccagtc actatggcgt gctgctagcg ctatatgcgt
1740tgatgcaatt tctatgcgca cccgttctcg gagcactgtc cgaccgcttt ggccgccgcc
1800cagtcctgct cgcttcgcta cttggagcca ctatcgacta cgcgatcatg gcgaccacac
1860ccgtcctgtg gatcccccgg gctgcaggaa ttgccgtaaa tgtatccgtt tataaggaca
1920gcccgaatga cggtctgcgc aaaaaaacac gttcatctca ctcgcgatgc tgcggagcag
1980ttactggctg atattgatcg acgccttgat cagttattgc ccgtggaggg agaacgggat
2040gttgtgggtg ccgcgatgcg tgaaggtgcg ctggcaccgg gaaaacgtat tcgccccatg
2100ttgctgttgc tgaccgcccg cgatctgggt tgcgctgtca gccatgacgg attactggat
2160ttggcctgtg cggtggaaat ggtccacgcg gcttcgctga tccttgacga tatgccctgc
2220atggacgatg cgaagctgcg gcgcggacgc cctaccattc attctcatta cggagagcat
2280gtggcaatac tggcggcggt tgccttgctg agtaaagcct ttggcgtaat tgccgatgca
2340gatggcctca cgccgctggc aaaaaatcgg gcggtttctg aactgtcaaa cgccatcggc
2400atgcaaggat tggttcaggg tcagttcaag gatctgtctg aaggggataa gccgcgcagc
2460gctgaagcta ttttgatgac gaatcacttt aaaaccagca cgctgttttg tgcctccatg
2520cagatggcct cgattgttgc gaatgcctcc agcgaagcgc gtgattgcct gcatcgtttt
2580tcacttgatc ttggtcaggc atttcaactg ctggacgatt tgaccgatgg catgaccgac
2640accggtaagg atagcaatca ggacgccggt aaatcgacgc tggtcaatct gttaggcccg
2700agggcggttg aagaacgtct gagacaacat cttcagcttg ccagtgagca tctctctgcg
2760gcctgccaac acgggcacgc cactcaacat tttattcagg cctggtttga caaaaaactc
2820gctgccgtca gttaagctta tgtgcaccgg tcagcctgtc ttaagtggga gcggctatgc
2880aaccgcatta tgatctgatt ctcgtggggg ctggactcgc gaatggcctt atcgccctgc
2940gtcttcagca gcagcaacct gatatgcgta ttttgcttat cgacgccgca ccccaggcgg
3000gcgggaatca tacgtggtca tttcaccacg atgatttgac tgagagccaa catcgttgga
3060tagctccgct ggtggttcat cactggcccg actatcaggt acgctttccc acacgccgtc
3120gtaagctgaa cagcggctac ttttgtatta cttctcagcg tttcgctgag gttttacagc
3180gacagtttgg cccgcacttg tggatggata ccgcggtcgc agaggttaat gcggaatctg
3240ttcggttgaa aaagggtcag gttatcggtg cccgcgcggt gattgacggg cggggttatg
3300cggcaaattc agcactgagc gtgggcttcc aggcgtttat tggccaggaa tggcgattga
3360gccacccgca tggtttatcg tctcccatta tcatggatgc cacggtcgat cagcaaaatg
3420gttatcgctt cgtgtacagc ctgccgctct cgccgaccag attgttaatt gaagacacgc
3480actatattga taatgcgaca ttagatcctg aatgcgcgcg gcaaaatatt tgcgactatg
3540ccgcgcaaca gggttggcag cttcagacac tgctgcgaga agaacagggc gccttaccca
3600ttactctgtc gggcaatgcc gacgcattct ggcagcagcg ccccctggcc tgtagtggat
3660tacgtgccgg tctgttccat cctaccaccg gctattcact gccgctggcg gttgccgtgg
3720ccgaccgcct gagtgcactt gatgtcttta cgtcggcctc aattcaccat gccattacgc
3780attttgcccg cgagcgctgg cagcagcagg gctttttccg catgctgaat cgcatgctgt
3840ttttagccgg acccgccgat tcacgctggc gggttatgca gcgtttttat ggtttacctg
3900aagatttaat tgcccgtttt tatgcgggaa aactcacgct gaccgatcgg ctacgtattc
3960tgagcggcaa gccgcctgtt ccggtattag cagcattgca agccattatg acgactcatc
4020gttaaagagc gactacatga aaccaactac ggtaattggt gcaggcttcg gtggcctggc
4080actggcaatt cgtctacaag ctgcggggat ccccgtctta ctgcttgaac aacgtgataa
4140acccggcggt cgggcttatg tctacgagga tcaggggttt acctttgatg caggcccgac
4200ggttatcacc gatcccagtg ccattgaaga actgtttgca ctggcaggaa aacagttaaa
4260agagtatgtc gaactgctgc cggttacgcc gttttaccgc ctgtgttggg agtcagggaa
4320ggtctttaat tacgataacg atcaaacccg gctcgaagcg cagattcagc agtttaatcc
4380ccgcgatgtc gaaggttatc gtcagtttct ggactattca cgcgcggtgt ttaaagaagg
4440ctatctaaag ctcggtactg tccctttttt atcgttcaga gacatgcttc gcgccgcacc
4500tcaactggcg aaactgcagg catggagaag cgtttacagt aaggttgcca gttacatcga
4560agatgaacat ctgcgccagg cgttttcttt ccactcgctg ttggtgggcg gcaatccctt
4620cgccacctca tccatttata cgttgataca cgcgctggag cgtgagtggg gcgtctggtt
4680tccgcgtggc ggcaccggcg cattagttca ggggatgata aagctgtttc aggatctggg
4740tggcgaagtc gtgttaaacg ccagagtcag ccatatggaa acgacaggaa acaagattga
4800agccgtgcat ttagaggacg gtcgcaggtt cctgacgcaa gccgtcgcgt caaatgcaga
4860tgtggttcat acctatcgcg acctgttaag ccagcaccct gccgcggtta agcagtccaa
4920caaactgcag actaagcgca tgagtaactc tctgtttgtg ctctattttg gtttgaatca
4980ccatcatgat cagctcgcgc atcacacggt ttgtttcggc ccgcgttacc gcgagctgat
5040tgacgaaatt tttaatcatg atggcctcgc agaggacttc tcactttatc tgcacgcgcc
5100ctgtgtcacg gattcgtcac tggcgcctga aggttgcggc agttactatg tgttggcgcc
5160ggtgccgcat ttaggcaccg cgaacctcga ctggacggtt gaggggccaa aactacgcga
5220ccgtattttt gcgtaccttg agcagcatta catgcctggc ttacggagtc agctggtcac
5280gcaccggatg tttacgccgt ttgattttcg cgaccagctt aatgcctatc atggctcagc
5340cttttctgtg gagcccgttc ttacccagag cgcctggttt cggccgcata accgcgataa
5400aaccattact aatctctacc tggtcggcgc aggcacgcat cccggcgcag gcattcctgg
5460cgtcatcggc tcggcaaaag cgacagcagg tttgatgctg gaggatctga tttgaataat
5520ccgtcgttac tcaatcatgc ggtcgaaacg atggcagttg gctcgaaaag ttttgcgaca
5580gcctcaaagt tatttgatgc aaaaacccgg cgcagcgtac tgatgctcta cgcctggtgc
5640cgccattgtg acgatgttat tgacgatcag acgctgggct ttcaggcccg gcagcctgcc
5700ttacaaacgc ccgaacaacg tctgatgcaa cttgagatga aaacgcgcca ggcctatgca
5760ggatcgcaga tgcacgaacc ggcgtttgcg gcttttcagg aagtggctat ggctcatgat
5820atcgccccgg cttacgcgtt tgatcatctg gaaggcttcg ccatggatgt acgcgaagcg
5880caatacagcc aactggatga tacgctgcgc tattgctatc acgttgcagg cgttgtcggc
5940ttgatgatgg cgcaaatcat gggcgtgcgg gataacgcca cgctggaccg cgcctgtgac
6000cttgggctgg catttcagtt gaccaatatt gctcgcgata ttgtggacga tgcgcatgcg
6060ggccgctgtt atctgccggc aagctggctg gagcatgaag gtctgaacaa agagaattat
6120gcggcacctg aaaaccgtca ggcgctgagc cgtatcgccc gtcgtttggt gcaggaagca
6180gaaccttact atttgtctgc cacagccggc ctggcagggt tgcccctgcg ttccgcctgg
6240gcaatcgcta cggcgaagca ggtttaccgg aaaataggtg tcaaagttga acaggccggt
6300cagcaagcct gggatcagcg gcagtcaacg accacgcccg aaaaattaac gctgctgctg
6360gccgcctctg gtcaggccct tacttcccgg atgcgggctc atcctccccg ccctgcgcat
6420ctctggcagc gcccgctcta gcgccatgtc gaccgatgcc cttgagagcc ttcaacccag
6480tcagctcctt ccggtgggcg cggggcatga ctatcgtcgc cgcacttatg actgtcttct
6540ttatcatgca actcgtagga caggtgccgg cagcgctctg ggtcattttc ggcgaggacc
6600gctttcgctg gagcgcgacg atgatcggcc tgtcgcttgc ggtattcgga atcttgcacg
6660ccctcgctca agccttcgtc actggtcccg ccaccaaacg tttcggcgag aagcaggcca
6720ttatcgccgg catggcggcc gacgcgctgg gctacgtctt gctggcgttc gcgacgcgag
6780gctggatggc cttccccatt atgattcttc tcgcttccgg cggcatcggg atgcccgcgt
6840tgcaggccat gctgtccagg caggtagatg acgaccatca gggacagctt caaggatcgc
6900tcgcggctct taccagccta acttcgatca ctggaccgct gatcgtcacg gcgatttatg
6960ccgcctcggc gagcacatgg aacgggttgg catggattgt aggcgccgcc ctataccttg
7020tctgcctccc cgcgttgcgt cgcggtgcat ggagccgggc cacctcgacc tgaatggaag
7080ccggcggcac ctcgctaacg gattcaccac tccaagaatt ggagccaatc aattcttgcg
7140gagaactgtg aatgcgcaaa ccaacccttg gcagaacata tccatcgcgt ccgccatctc
7200cagcagccgc acgcggcgca tctcgggcag cgttgggtcc tggccacggg tgcgcatgat
7260cgtgctcctg tcgttgagga cccggctagg ctggcggggt tgccttactg gttagcagaa
7320tgaatcaccg atacgcgagc gaacgtgaag cgactgctgc tgcaaaacgt ctgcgacctg
7380agcaacaaca tgaatggtct tcggtttccg tgtttcgtaa agtctggaaa cgcggaagtc
7440ccctacgtgc tgctgaagtt gcccgcaaca gagagtggaa ccaaccggtg ataccacgat
7500actatgactg agagtcaacg ccatgagcgg cctcatttct tattctgagt tacaacagtc
7560cgcaccgctg tccggtagct ccttccggtg ggcgcggggc atgactatcg tcgccgcact
7620tatgactgtc ttctttatca tgcaactcgt aggacaggtg ccggcagcgc ccaacagtcc
7680cccggccacg gggcctgcca ccatacccac gccgaaacaa gcgccctgca ccattatgtt
7740ccggatctgc atcgcaggat gctgctggct accctgtgga acacctacat ctgtattaac
7800gaagcgctaa ccgtttttat caggctctgg gaggcagaat aaatgatcat atcgtcaatt
7860attacctcca cggggagagc ctgagcaaac tggcctcagg catttgagaa gcacacggtc
7920acactgcttc cggtagtcaa taaaccggta aaccagcaat agacataagc ggctatttaa
7980cgaccctgcc ctgaaccgac gaccgggtcg aatttgcttt cgaatttctg ccattcatcc
8040gcttattatc acttattcag gcgtagcacc aggcgtttaa gggcaccaat aactgcctta
8100aaaaaattac gccccgccct gccactcatc gcagtactgt tgtaattcat taagcattct
8160gccgacatgg aagccatcac agacggcatg atgaacctga atcgccagcg gcatcagcac
8220cttgtcgcct tgcgtataat atttgcccat ggtgaaaacg ggggcgaaga agttgtccat
8280attggccacg tttaaatcaa aactggtgaa actcacccag ggattggctg agacgaaaaa
8340catattctca ataaaccctt tagggaaata ggccaggttt tcaccgtaac acgccacatc
8400ttgcgaatat atgtgtagaa actgccggaa atcgtcgtgg tattcactcc agagcgatga
8460aaacgtttca gtttgctcat ggaaaacggt gtaacaaggg tgaacactat cccatatcac
8520cagctcaccg tctttcattg ccatacg
8547471119DNAEscherichia coliCDS(1)..(1116) 47atg cat aac cag gct cca att
caa cgt aga aaa tca aca cgt att tac 48Met His Asn Gln Ala Pro Ile
Gln Arg Arg Lys Ser Thr Arg Ile Tyr1 5 10
15gtt ggg aat gtg ccg att ggc gat ggt gct ccc atc gcc
gta cag tcc 96Val Gly Asn Val Pro Ile Gly Asp Gly Ala Pro Ile Ala
Val Gln Ser 20 25 30atg acc
aat acg cgt acg aca gac gtc gaa gca acg gtc aat caa atc 144Met Thr
Asn Thr Arg Thr Thr Asp Val Glu Ala Thr Val Asn Gln Ile 35
40 45aag gcg ctg gaa cgc gtt ggc gct gat atc
gtc cgt gta tcc gta ccg 192Lys Ala Leu Glu Arg Val Gly Ala Asp Ile
Val Arg Val Ser Val Pro 50 55 60acg
atg gac gcg gca gaa gcg ttc aaa ctc atc aaa cag cag gtt aac 240Thr
Met Asp Ala Ala Glu Ala Phe Lys Leu Ile Lys Gln Gln Val Asn65
70 75 80gtg ccg ctg gtg gct gac
atc cac ttc gac tat cgc att gcg ctg aaa 288Val Pro Leu Val Ala Asp
Ile His Phe Asp Tyr Arg Ile Ala Leu Lys 85
90 95gta gcg gaa tac ggc gtc gat tgt ctg cgt att aac
cct ggc aat atc 336Val Ala Glu Tyr Gly Val Asp Cys Leu Arg Ile Asn
Pro Gly Asn Ile 100 105 110ggt
aat gaa gag cgt att cgc atg gtg gtt gac tgt gcg cgc gat aaa 384Gly
Asn Glu Glu Arg Ile Arg Met Val Val Asp Cys Ala Arg Asp Lys 115
120 125aac att ccg atc cgt att ggc gtt aac
gcc gga tcg ctg gaa aaa gat 432Asn Ile Pro Ile Arg Ile Gly Val Asn
Ala Gly Ser Leu Glu Lys Asp 130 135
140ctg caa gaa aag tat ggc gaa ccg acg ccg cag gcg ttg ctg gaa tct
480Leu Gln Glu Lys Tyr Gly Glu Pro Thr Pro Gln Ala Leu Leu Glu Ser145
150 155 160gcc atg cgt cat
gtt gat cat ctc gat cgc ctg aac ttc gat cag ttc 528Ala Met Arg His
Val Asp His Leu Asp Arg Leu Asn Phe Asp Gln Phe 165
170 175aaa gtc agc gtg aaa gcg tct gac gtc ttc
ctc gct gtt gag tct tat 576Lys Val Ser Val Lys Ala Ser Asp Val Phe
Leu Ala Val Glu Ser Tyr 180 185
190cgt ttg ctg gca aaa cag atc gat cag ccg ttg cat ctg ggg atc acc
624Arg Leu Leu Ala Lys Gln Ile Asp Gln Pro Leu His Leu Gly Ile Thr
195 200 205gaa gcc ggt ggt gcg cgc agc
ggg gca gta aaa tcc gcc att ggt tta 672Glu Ala Gly Gly Ala Arg Ser
Gly Ala Val Lys Ser Ala Ile Gly Leu 210 215
220ggt ctg ctg ctg tct gaa ggc atc ggc gac acg ctg cgc gta tcg ctg
720Gly Leu Leu Leu Ser Glu Gly Ile Gly Asp Thr Leu Arg Val Ser Leu225
230 235 240gcg gcc gat ccg
gtc gaa gag atc aaa gtc ggt ttc gat att ttg aaa 768Ala Ala Asp Pro
Val Glu Glu Ile Lys Val Gly Phe Asp Ile Leu Lys 245
250 255tcg ctg cgt atc cgt tcg cga ggg atc aac
ttc atc gcc tgc ccg acc 816Ser Leu Arg Ile Arg Ser Arg Gly Ile Asn
Phe Ile Ala Cys Pro Thr 260 265
270tgt tcg cgt cag gaa ttt gat gtt atc ggt acg gtt aac gcg ctg gag
864Cys Ser Arg Gln Glu Phe Asp Val Ile Gly Thr Val Asn Ala Leu Glu
275 280 285caa cgc ctg gaa gat atc atc
act ccg atg gac gtt tcg att atc ggc 912Gln Arg Leu Glu Asp Ile Ile
Thr Pro Met Asp Val Ser Ile Ile Gly 290 295
300tgc gtg gtg aat ggc cca ggt gag gcg ctg gtt tct aca ctc ggc gtc
960Cys Val Val Asn Gly Pro Gly Glu Ala Leu Val Ser Thr Leu Gly Val305
310 315 320acc ggc ggc aac
aag aaa agc ggc ctc tat gaa gat ggc gtg cgc aaa 1008Thr Gly Gly Asn
Lys Lys Ser Gly Leu Tyr Glu Asp Gly Val Arg Lys 325
330 335gac cgt ctg gac aac aac gat atg atc gac
cag ctg gaa gca cgc att 1056Asp Arg Leu Asp Asn Asn Asp Met Ile Asp
Gln Leu Glu Ala Arg Ile 340 345
350cgt gcg aaa gcc agt cag ctg gac gaa gcg cgt cga att gac gtt cag
1104Arg Ala Lys Ala Ser Gln Leu Asp Glu Ala Arg Arg Ile Asp Val Gln
355 360 365cag gtt gaa aaa taa
1119Gln Val Glu Lys
37048372PRTEscherichia coli 48Met His Asn Gln Ala Pro Ile Gln Arg Arg Lys
Ser Thr Arg Ile Tyr1 5 10
15Val Gly Asn Val Pro Ile Gly Asp Gly Ala Pro Ile Ala Val Gln Ser
20 25 30Met Thr Asn Thr Arg Thr Thr
Asp Val Glu Ala Thr Val Asn Gln Ile 35 40
45Lys Ala Leu Glu Arg Val Gly Ala Asp Ile Val Arg Val Ser Val
Pro 50 55 60Thr Met Asp Ala Ala Glu
Ala Phe Lys Leu Ile Lys Gln Gln Val Asn65 70
75 80Val Pro Leu Val Ala Asp Ile His Phe Asp Tyr
Arg Ile Ala Leu Lys 85 90
95Val Ala Glu Tyr Gly Val Asp Cys Leu Arg Ile Asn Pro Gly Asn Ile
100 105 110Gly Asn Glu Glu Arg Ile
Arg Met Val Val Asp Cys Ala Arg Asp Lys 115 120
125Asn Ile Pro Ile Arg Ile Gly Val Asn Ala Gly Ser Leu Glu
Lys Asp 130 135 140Leu Gln Glu Lys Tyr
Gly Glu Pro Thr Pro Gln Ala Leu Leu Glu Ser145 150
155 160Ala Met Arg His Val Asp His Leu Asp Arg
Leu Asn Phe Asp Gln Phe 165 170
175Lys Val Ser Val Lys Ala Ser Asp Val Phe Leu Ala Val Glu Ser Tyr
180 185 190Arg Leu Leu Ala Lys
Gln Ile Asp Gln Pro Leu His Leu Gly Ile Thr 195
200 205Glu Ala Gly Gly Ala Arg Ser Gly Ala Val Lys Ser
Ala Ile Gly Leu 210 215 220Gly Leu Leu
Leu Ser Glu Gly Ile Gly Asp Thr Leu Arg Val Ser Leu225
230 235 240Ala Ala Asp Pro Val Glu Glu
Ile Lys Val Gly Phe Asp Ile Leu Lys 245
250 255Ser Leu Arg Ile Arg Ser Arg Gly Ile Asn Phe Ile
Ala Cys Pro Thr 260 265 270Cys
Ser Arg Gln Glu Phe Asp Val Ile Gly Thr Val Asn Ala Leu Glu 275
280 285Gln Arg Leu Glu Asp Ile Ile Thr Pro
Met Asp Val Ser Ile Ile Gly 290 295
300Cys Val Val Asn Gly Pro Gly Glu Ala Leu Val Ser Thr Leu Gly Val305
310 315 320Thr Gly Gly Asn
Lys Lys Ser Gly Leu Tyr Glu Asp Gly Val Arg Lys 325
330 335Asp Arg Leu Asp Asn Asn Asp Met Ile Asp
Gln Leu Glu Ala Arg Ile 340 345
350Arg Ala Lys Ala Ser Gln Leu Asp Glu Ala Arg Arg Ile Asp Val Gln
355 360 365Gln Val Glu Lys
370498823DNAArtificialpBScyclogcpE 49gtggcacttt tcggggaaat gtgcgcggaa
cccctatttg tttatttttc taaatacatt 60caaatatgta tccgctcatg agacaataac
cctgataaat gcttcaataa tattgaaaaa 120ggaagagtat gagtattcaa catttccgtg
tcgcccttat tccctttttt gcggcatttt 180gccttcctgt ttttgctcac ccagaaacgc
tggtgaaagt aaaagatgct gaagatcagt 240tgggtgcacg agtgggttac atcgaactgg
atctcaacag cggtaagatc cttgagagtt 300ttcgccccga agaacgtttt ccaatgatga
gcacttttaa agttctgcta tgtggcgcgg 360tattatcccg tattgacgcc gggcaagagc
aactcggtcg ccgcatacac tattctcaga 420atgacttggt tgagtactca ccagtcacag
aaaagcatct tacggatggc atgacagtaa 480gagaattatg cagtgctgcc ataaccatga
gtgataacac tgcggccaac ttacttctga 540caacgatcgg aggaccgaag gagctaaccg
cttttttgca caacatgggg gatcatgtaa 600ctcgccttga tcgttgggaa ccggagctga
atgaagccat accaaacgac gagcgtgaca 660ccacgatgcc tgtagcaatg gcaacaacgt
tgcgcaaact attaactggc gaactactta 720ctctagcttc ccggcaacaa ttaatagact
ggatggaggc ggataaagtt gcaggaccac 780ttctgcgctc ggcccttccg gctggctggt
ttattgctga taaatctgga gccggtgagc 840gtgggtctcg cggtatcatt gcagcactgg
ggccagatgg taagccctcc cgtatcgtag 900ttatctacac gacggggagt caggcaacta
tggatgaacg aaatagacag atcgctgaga 960taggtgcctc actgattaag cattggtaac
tgtcagacca agtttactca tatatacttt 1020agattgattt aaaacttcat ttttaattta
aaaggatcta ggtgaagatc ctttttgata 1080atctcatgac caaaatccct taacgtgagt
tttcgttcca ctgagcgtca gaccccgtag 1140aaaagatcaa aggatcttct tgagatcctt
tttttctgcg cgtaatctgc tgcttgcaaa 1200caaaaaaacc accgctacca gcggtggttt
gtttgccgga tcaagagcta ccaactcttt 1260ttccgaaggt aactggcttc agcagagcgc
agataccaaa tactgtcctt ctagtgtagc 1320cgtagttagg ccaccacttc aagaactctg
tagcaccgcc tacatacctc gctctgctaa 1380tcctgttacc agtggctgct gccagtggcg
ataagtcgtg tcttaccggg ttggactcaa 1440gacgatagtt accggataag gcgcagcggt
cgggctgaac ggggggttcg tgcacacagc 1500ccagcttgga gcgaacgacc tacaccgaac
tgagatacct acagcgtgag ctatgagaaa 1560gcgccacgct tcccgaaggg agaaaggcgg
acaggtatcc ggtaagcggc agggtcggaa 1620caggagagcg cacgagggag cttccagggg
gaaacgcctg gtatctttat agtcctgtcg 1680ggtttcgcca cctctgactt gagcgtcgat
ttttgtgatg ctcgtcaggg gggcggagcc 1740tatggaaaaa cgccagcaac gcggcctttt
tacggttcct ggccttttgc tggccttttg 1800ctcacatgtt ctttcctgcg ttatcccctg
attctgtgga taaccgtatt accgcctttg 1860agtgagctga taccgctcgc cgcagccgaa
cgaccgagcg cagcgagtca gtgagcgagg 1920aagcggaaga gcgcccaata cgcaaaccgc
ctctccccgc gcgttggccg attcattaat 1980gcagctggca cgacaggttt cccgactgga
aagcgggcag tgagcgcaac gcaattaatg 2040tgagttagct cactcattag gcaccccagg
ctttacactt tatgcttccg gctcgtatgt 2100tgtgtggaat tgtgagcgga taacaatttc
acacaggaaa cagctatgac catgattacg 2160ccaagcgcgc aattaaccct cactaaaggg
aacaaaagct ggagctccac cgcgggagga 2220gaaattaacc atgcataacc aggctccaat
tcaacgtaga aaatcaacac gtatttacgt 2280tgggaatgtg ccgattggcg atggtgctcc
catcgccgta cagtccatga ccaatacgcg 2340tacgacagac gtcgaagcaa cggtcaatca
aatcaaggcg ctggaacgcg ttggcgctga 2400tatcgtccgt gtatccgtac cgacgatgga
cgcggcagaa gcgttcaaac tcatcaaaca 2460gcaggttaac gtgccgctgg tggctgacat
ccacttcgac tatcgcattg cgctgaaagt 2520agcggaatac ggcgtcgatt gtctgcgtat
taaccctggc aatatcggta atgaagagcg 2580tattcgcatg gtggttgact gtgcgcgcga
taaaaacatt ccgatccgta ttggcgttaa 2640cgccggatcg ctggaaaaag atctgcaaga
aaagtatggc gaaccgacgc cgcaggcgtt 2700gctggaatct gccatgcgtc atgttgatca
tctcgatcgc ctgaacttcg atcagttcaa 2760agtcagcgtg aaagcgtctg acgtcttcct
cgctgttgag tcttatcgtt tgctggcaaa 2820acagatcgat cagccgttgc atctggggat
caccgaagcc ggtggtgcgc gcagcggggc 2880agtaaaatcc gccattggtt taggtctgct
gctgtctgaa ggcatcggcg acacgctgcg 2940cgtatcgctg gcggccgatc cggtcgaaga
gatcaaagtc ggtttcgata ttttgaaatc 3000gctgcgtatc cgttcgcgag ggatcaactt
catcgcctgc ccgacctgtt cgcgtcagga 3060atttgatgtt atcggtacgg ttaacgcgct
ggagcaacgc ctggaagata tcatcactcc 3120gatggacgtt tcgattatcg gctgcgtggt
gaatggccca ggtgaggcgc tggtttctac 3180actcggcgtc accggcggca acaagaaaag
cggcctctat gaagatggcg tgcgcaaaga 3240ccgtctggac aacaacgata tgatcgacca
gctggaagca cgcattcgtg cgaaagccag 3300tcagctggac gaagcgcgtc gaattgacgt
tcagcaggtt gaaaaataag cggccgctct 3360agaactagtg gatcccccgg gctgcaggaa
ttcgaggaga aattaaccat gtatatcggg 3420atagatcttg gcacctcggg cgtaaaagtt
attttgctca acgagcaggg tgaggtggtt 3480gctgcgcaaa cggaaaagct gaccgtttcg
cgcccgcatc cactctggtc ggaacaagac 3540ccggaacagt ggtggcaggc aactgatcgc
gcaatgaaag ctctgggcga tcagcattct 3600ctgcaggacg ttaaagcatt gggtattgcc
ggccagatgc acggagcaac cttgctggat 3660gctcagcaac gggtgttacg ccctgccatt
ttgtggaacg acgggcgctg tgcgcaagag 3720tgcactttgc tggaagcgcg agttccgcaa
tcgcgggtga ttaccggcaa cctgatgatg 3780cccggattta ctgcgcctaa attgctatgg
gttcagcggc atgagccgga gatattccgt 3840caaatcgaca aagtattatt accgaaagat
tacttgcgtc tgcgtatgac gggggagttt 3900gccagcgata tgtctgacgc agctggcacc
atgtggctgg atgtcgcaaa gcgtgactgg 3960agtgacgtca tgctgcaggc ttgcgactta
tctcgtgacc agatgcccgc attatacgaa 4020ggcagcgaaa ttactggtgc tttgttacct
gaagttgcga aagcgtgggg tatggcgacg 4080gtgccagttg tcgcaggcgg tggcgacaat
gcagctggtg cagttggtgt gggaatggtt 4140gatgctaatc aggcaatgtt atcgctgggg
acgtcggggg tctattttgc tgtcagcgaa 4200gggttcttaa gcaagccaga aagcgccgta
catagctttt gccatgcgct accgcaacgt 4260tggcatttaa tgtctgtgat gctgagtgca
gcgtcgtgtc tggattgggc cgcgaaatta 4320accggcctga gcaatgtccc agctttaatc
gctgcagctc aacaggctga tgaaagtgcc 4380gagccagttt ggtttctgcc ttatctttcc
ggcgagcgta cgccacacaa taatccccag 4440gcgaaggggg ttttctttgg tttgactcat
caacatggcc ccaatgaact ggcgcgagca 4500gtgctggaag gcgtgggtta tgcgctggca
gatggcatgg atgtcgtgca tgcctgcggt 4560attaaaccgc aaagtgttac gttgattggg
ggcggggcgc gtagtgagta ctggcgtcag 4620atgctggcgg atatcagcgg tcagcagctc
gattaccgta cgggggggga tgtggggcca 4680gcactgggcg cagcaaggct ggcgcagatc
gcggcgaatc cagagaaatc gctcattgaa 4740ttgttgccgc aactaccgtt agaacagtcg
catctaccag atgcgcagcg ttatgccgct 4800tatcagccac gacgagaaac gttccgtcgc
ctctatcagc aacttctgcc attaatggcg 4860taaaagcttg aggagaaatt aaccatgaag
caactcacca ttctgggctc gaccggctcg 4920attggttgca gcacgctgga cgtggtgcgc
cataatcccg aacacttccg cgtagttgcg 4980ctggtggcag gcaaaaatgt cactcgcatg
gtagaacagt gcctggaatt ctctccccgc 5040tatgccgtaa tggacgatga agcgagtgcg
aaacttctta aaacgatgct acagcaacag 5100ggtagccgca ccgaagtctt aagtgggcaa
caagccgctt gcgatatggc agcgcttgag 5160gatgttgatc aggtgatggc agccattgtt
ggcgctgctg ggctgttacc tacgcttgct 5220gcgatccgcg cgggtaaaac cattttgctg
gccaataaag aatcactggt tacctgcgga 5280cgtctgttta tggacgccgt aaagcagagc
aaagcgcaat tgttaccggt cgatagcgaa 5340cataacgcca tttttcagag tttaccgcaa
cctatccagc ataatctggg atacgctgac 5400cttgagcaaa atggcgtggt gtccatttta
cttaccgggt ctggtggccc tttccgtgag 5460acgccattgc gcgatttggc aacaatgacg
ccggatcaag cctgccgtca tccgaactgg 5520tcgatggggc gtaaaatttc tgtcgattcg
gctaccatga tgaacaaagg tctggaatac 5580attgaagcgc gttggctgtt taacgccagc
gccagccaga tggaagtgct gattcacccg 5640cagtcagtga ttcactcaat ggtgcgctat
caggacggca gtgttctggc gcagctgggg 5700gaaccggata tgcgtacgcc aattgcccac
accatggcat ggccgaatcg cgtgaactct 5760ggcgtgaagc cgctcgattt ttgcaaacta
agtgcgttga catttgccgc accggattat 5820gatcgttatc catgcctgaa actggcgatg
gaggcgttcg aacaaggcca ggcagcgacg 5880acagcattga atgccgcaaa cgaaatcacc
gttgctgctt ttcttgcgca acaaatccgc 5940tttacggata tcgctgcgtt gaatttatcc
gtactggaaa aaatggatat gcgcgaacca 6000caatgtgtgg acgatgtgtt atctgttgat
gcgaacgcgc gtgaagtcgc cagaaaagag 6060gtgatgcgtc tcgcaagctg agtcgacgag
gagaaattaa ccatggcaac cactcatttg 6120gatgtttgcg ccgtggttcc ggcggccgga
tttggccgtc gaatgcaaac ggaatgtcct 6180aagcaatatc tctcaatcgg taatcaaacc
attcttgaac actcggtgca tgcgctgctg 6240gcgcatcccc gggtgaaacg tgtcgtcatt
gccataagtc ctggcgatag ccgttttgca 6300caacttcctc tggcgaatca tccgcaaatc
accgttgtag atggcggtga tgagcgtgcc 6360gattccgtgc tggcaggtct gaaagccgct
ggcgacgcgc agtgggtatt ggtgcatgac 6420gccgctcgtc cttgtttgca tcaggatgac
ctcgcgcgat tgttggcgtt gagcgaaacc 6480agccgcacgg gggggatcct cgccgcacca
gtgcgcgata ctatgaaacg tgccgaaccg 6540ggcaaaaatg ccattgctca taccgttgat
cgcaacggct tatggcacgc gctgacgccg 6600caatttttcc ctcgtgagct gttacatgac
tgtctgacgc gcgctctaaa tgaaggcgcg 6660actattaccg acgaagcctc ggcgctggaa
tattgcggat tccatcctca gttggtcgaa 6720ggccgtgcgg ataacattaa agtcacgcgc
ccggaagatt tggcactggc cgagttttac 6780ctcacccgaa ccatccatca ggagaataca
taatgcgaat tggacacggt tttgacgtac 6840atgcctttgg cggtgaaggc ccaattatca
ttggtggcgt acgcattcct tacgaaaaag 6900gattgctggc gcattctgat ggcgacgtgg
cgctccatgc gttgaccgat gcattgcttg 6960gcgcggcggc gctgggggat atcggcaagc
tgttcccgga taccgatccg gcatttaaag 7020gtgccgatag ccgcgagctg ctacgcgaag
cctggcgtcg tattcaggcg aagggttata 7080cccttggcaa cgtcgatgtc actatcatcg
ctcaggcacc gaagatgttg ccgcacattc 7140cacaaatgcg cgtgtttatt gccgaagatc
tcggctgcca tatggatgat gttaacgtga 7200aagccactac tacggaaaaa ctgggattta
ccggacgtgg ggaagggatt gcctgtgaag 7260cggtggcgct actcattaag gcaacaaaat
gactcgagga ggagaaatta accatgcgga 7320cacagtggcc ctctccggca aaacttaatc
tgtttttata cattaccggt cagcgtgcgg 7380atggttacca cacgctgcaa acgctgtttc
agtttcttga ttacggcgac accatcagca 7440ttgagcttcg tgacgatggg gatattcgtc
tgttaacgcc cgttgaaggc gtggaacatg 7500aagataacct gatcgttcgc gcagcgcgat
tgttgatgaa aactgcggca gacagcgggc 7560gtcttccgac gggaagcggt gcgaatatca
gcattgacaa gcgtttgccg atgggcggcg 7620gtctcggcgg tggttcatcc aatgccgcga
cggtcctggt ggcattaaat catctctggc 7680aatgcgggct aagcatggat gagctggcgg
aaatggggct gacgctgggc gcagatgttc 7740ctgtctttgt tcgggggcat gccgcgtttg
ccgaaggcgt tggtgaaata ctaacgccgg 7800tggatccgcc agagaagtgg tatctggtgg
cgcaccctgg tgtaagtatt ccgactccgg 7860tgatttttaa agatcctgaa ctcccgcgca
atacgccaaa aaggtcaata gaaacgttgc 7920taaaatgtga attcagcaat gattgcgagg
ttatcgcaag aaaacgtttt cgcgaggttg 7980atgcggtgct ttcctggctg ttagaatacg
ccccgtcgcg cctgactggg acaggggcct 8040gtgtctttgc tgaatttgat acagagtctg
aagcccgcca ggtgctagag caagccccgg 8100aatggctcaa tggctttgtg gcgaaaggcg
ctaatctttc cccattgcac agagccatgc 8160tttaaggtac ccaattcgcc ctatagtgag
tcgtattacg cgcgctcact ggccgtcgtt 8220ttacaacgtc gtgactggga aaaccctggc
gttacccaac ttaatcgcct tgcagcacat 8280ccccctttcg ccagctggcg taatagcgaa
gaggcccgca ccgatcgccc ttcccaacag 8340ttgcgcagcc tgaatggcga atggaaattg
taagcgttaa tattttgtta aaattcgcgt 8400taaatttttg ttaaatcagc tcatttttta
accaataggc cgaaatcggc aaaatccctt 8460ataaatcaaa agaatagacc gagatagggt
tgagtgttgt tccagtttgg aacaagagtc 8520cactattaaa gaacgtggac tccaacgtca
aagggcgaaa aaccgtctat cagggcgatg 8580gcccactacg tgaaccatca ccctaatcaa
gttttttggg gtcgaggtgc cgtaaagcac 8640taaatcggaa ccctaaaggg agcccccgat
ttagagcttg acggggaaag ccggcgaacg 8700tggcgagaaa ggaagggaag aaagcgaaag
gagcgggcgc tagggcgctg gcaagtgtag 8760cggtcacgct gcgcgtaacc accacacccg
ccgcgcttaa tgcgccgcta cagggcgcgt 8820cag
8823505793DNAArtificialpACYClytBgcpE
50gaattccgga tgagcattca tcaggcgggc aagaatgtga ataaaggccg gataaaactt
60gtgcttattt ttctttacgg tctttaaaaa ggccgtaata tccagctgaa cggtctggtt
120ataggtacat tgagcaactg actgaaatgc ctcaaaatgt tctttacgat gccattggga
180tatatcaacg gtggtatatc cagtgatttt tttctccatt ttagcttcct tagctcctga
240aaatctcgat aactcaaaaa atacgcccgg tagtgatctt atttcattat ggtgaaagtt
300ggaacctctt acgtgccgat caacgtctca ttttcgccaa aagttggccc agggcttccc
360ggtatcaaca gggacaccag gatttattta ttctgcgaag tgatcttccg tcacaggtat
420ttattcggcg caaagtgcgt cgggtgatgc tgccaactta ctgatttagt gtatgatggt
480gtttttgagg tgctccagtg gcttctgttt ctatcagctg tccctcctgt tcagctactg
540acggggtggt gcgtaacggc aaaagcaccg ccggacatca gcgctagcgg agtgtatact
600ggcttactat gttggcactg atgagggtgt cagtgaagtg cttcatgtgg caggagaaaa
660aaggctgcac cggtgcgtca gcagaatatg tgatacagga tatattccgc ttcctcgctc
720actgactcgc tacgctcggt cgttcgactg cggcgagcgg aaatggctta cgaacggggc
780ggagatttcc tggaagatgc caggaagata cttaacaggg aagtgagagg gccgcggcaa
840agccgttttt ccataggctc cgcccccctg acaagcatca cgaaatctga cgctcaaatc
900agtggtggcg aaacccgaca ggactataaa gataccaggc gtttccccct ggcggctccc
960tcgtgcgctc tcctgttcct gcctttcggt ttaccggtgt cattccgctg ttatggccgc
1020gtttgtctca ttccacgcct gacactcagt tccgggtagg cagttcgctc caagctggac
1080tgtatgcacg aaccccccgt tcagtccgac cgctgcgcct tatccggtaa ctatcgtctt
1140gagtccaacc cggaaagaca tgcaaaagca ccactggcag cagccactgg taattgattt
1200agaggagtta gtcttgaagt catgcgccgg ttaaggctaa actgaaagga caagttttgg
1260tgactgcgct cctccaagcc agttacctcg gttcaaagag ttggtagctc agagaacctt
1320cgaaaaaccg ccctgcaagg cggttttttc gttttcagag caagagatta cgcgcagacc
1380aaaacgatct caagaagatc atcttattaa tcagataaaa tatttctaga tttcagtgca
1440atttatctct tcaaatgtag cacctgaagt cagccccata cgatataagt tgtaattctc
1500atgtttgaca gcttatcatc gataagcttt aatgcggtag tttatcacag ttaaattgct
1560aacgcagtca ggcaccgtgt atgaaatcta acaatgcgct catcgtcatc ctcggcaccg
1620tcaccctgga tgctgtaggc ataggcttgg ttatgccggt actgccgggc ctcttgcggg
1680atatcgtcca ttccgacagc atcgccagtc actatggcgt gctgctagcg ctatatgcgt
1740tgatgcaatt tctatgcgca cccgttctcg gagcactgtc cgaccgcttt ggccgccgcc
1800cagtcctgct cgcttcgcta cttggagcca ctatcgacta cgcgatcatg gcgaccacac
1860ccgtcctgtg gatccgagga gaaattaacc atgcataacc aggctccaat tcaacgtaga
1920aaatcaacac gtatttacgt tgggaatgtg ccgattggcg atggtgctcc catcgccgta
1980cagtccatga ccaatacgcg tacgacagac gtcgaagcaa cggtcaatca aatcaaggcg
2040ctggaacgcg ttggcgctga tatcgtccgt gtatccgtac cgacgatgga cgcggcagaa
2100gcgttcaaac tcatcaaaca gcaggttaac gtgccgctgg tggctgacat ccacttcgac
2160tatcgcattg cgctgaaagt agcggaatac ggcgtcgatt gtctgcgtat taaccctggc
2220aatatcggta atgaagagcg tattcgcatg gtggttgact gtgcgcgcga taaaaacatt
2280ccgatccgta ttggcgttaa cgccggatcg ctggaaaaag atctgcaaga aaagtatggc
2340gaaccgacgc cgcaggcgtt gctggaatct gccatgcgtc atgttgatca tctcgatcgc
2400ctgaacttcg atcagttcaa agtcagcgtg aaagcgtctg acgtcttcct cgctgttgag
2460tcttatcgtt tgctggcaaa acagatcgat cagccgttgc atctggggat caccgaagcc
2520ggtggtgcgc gcagcggggc agtaaaatcc gccattggtt taggtctgct gctgtctgaa
2580ggcatcggcg acacgctgcg cgtatcgctg gcggccgatc cggtcgaaga gatcaaagtc
2640ggtttcgata ttttgaaatc gctgcgtatc cgttcgcgag ggatcaactt catcgcctgc
2700ccgacctgtt cgcgtcagga atttgatgtt atcggtacgg ttaacgcgct ggagcaacgc
2760ctggaagata tcatcactcc gatggacgtt tcgattatcg gctgcgtggt gaatggccca
2820ggtgaggcgc tggtttctac actcggcgtc accggcggca acaagaaaag cggcctctat
2880gaagatggcg tgcgcaaaga ccgtctggac aacaacgata tgatcgacca gctggaagca
2940cgcattcgtg cgaaagccag tcagctggac gaagcgcgtc gaattgacgt tcagcaggtt
3000gaaaaataag tcgacgagga gaaattaacc atgcagatcc tgttggccaa cccgcgtggt
3060ttttgtgccg gggtagaccg cgctatcagc attgttgaaa acgcgctggc catttacggc
3120gcaccgatat atgtccgtca cgaagtggta cataaccgct atgtggtcga tagcttgcgt
3180gagcgtgggg ctatctttat tgagcagatt agcgaagtac cggacggcgc gatcctgatt
3240ttctccgcac acggtgtttc tcaggcggta cgtaacgaag caaaaagtcg cgatttgacg
3300gtgtttgatg ccacctgtcc gctggtgacc aaagtgcata tggaagtcgc ccgcgccagt
3360cgccgtggcg aagaatctat tctcatcggt cacgccgggc acccggaagt ggaagggaca
3420atgggccagt acagtaaccc ggaaggggga atgtatctgg tcgaatcgcc ggacgatgtg
3480tggaaactga cggtcaaaaa cgaagagaag ctctccttta tgacccagac cacgctgtcg
3540gtggatgaca cgtctgatgt gatcgacgcg ctgcgtaaac gcttcccgaa aattgtcggt
3600ccgcgcaaag atgacatctg ctacgccacg actaaccgtc aggaagcggt acgcgccctg
3660gcagaacagg cggaagttgt gttggtggtc ggttcgaaaa actcctccaa ctccaaccgt
3720ctggcggagc tggcccagcg tatgggcaaa cgcgcgtttt tgattgacga tgcgaaagac
3780atccaggaag agtgggtgaa agaggttaaa tgcgtcggcg tgactgcggg cgcatcggct
3840ccggatattc tggtgcagaa tgtggtggca cgtttgcagc agctgggcgg tggtgaagcc
3900attccgctgg aaggccgtga agaaaacatt gttttcgaag tgccgaaaga gctgcgtgtc
3960gatattcgtg aagtcgatta acggccgacg cgctgggcta cgtcttgctg gcgttcgcga
4020cgcgaggctg gatggccttc cccattatga ttcttctcgc ttccggcggc atcgggatgc
4080ccgcgttgca ggccatgctg tccaggcagg tagatgacga ccatcaggga cagcttcaag
4140gatcgctcgc ggctcttacc agcctaactt cgatcactgg accgctgatc gtcacggcga
4200tttatgccgc ctcggcgagc acatggaacg ggttggcatg gattgtaggc gccgccctat
4260accttgtctg cctccccgcg ttgcgtcgcg gtgcatggag ccgggccacc tcgacctgaa
4320tggaagccgg cggcacctcg ctaacggatt caccactcca agaattggag ccaatcaatt
4380cttgcggaga actgtgaatg cgcaaaccaa cccttggcag aacatatcca tcgcgtccgc
4440catctccagc agccgcacgc ggcgcatctc gggcagcgtt gggtcctggc cacgggtgcg
4500catgatcgtg ctcctgtcgt tgaggacccg gctaggctgg cggggttgcc ttactggtta
4560gcagaatgaa tcaccgatac gcgagcgaac gtgaagcgac tgctgctgca aaacgtctgc
4620gacctgagca acaacatgaa tggtcttcgg tttccgtgtt tcgtaaagtc tggaaacgcg
4680gaagtcccct acgtgctgct gaagttgccc gcaacagaga gtggaaccaa ccggtgatac
4740cacgatacta tgactgagag tcaacgccat gagcggcctc atttcttatt ctgagttaca
4800acagtccgca ccgctgtccg gtagctcctt ccggtgggcg cggggcatga ctatcgtcgc
4860cgcacttatg actgtcttct ttatcatgca actcgtagga caggtgccgg cagcgcccaa
4920cagtcccccg gccacggggc ctgccaccat acccacgccg aaacaagcgc cctgcaccat
4980tatgttccgg atctgcatcg caggatgctg ctggctaccc tgtggaacac ctacatctgt
5040attaacgaag cgctaaccgt ttttatcagg ctctgggagg cagaataaat gatcatatcg
5100tcaattatta cctccacggg gagagcctga gcaaactggc ctcaggcatt tgagaagcac
5160acggtcacac tgcttccggt agtcaataaa ccggtaaacc agcaatagac ataagcggct
5220atttaacgac cctgccctga accgacgacc gggtcgaatt tgctttcgaa tttctgccat
5280tcatccgctt attatcactt attcaggcgt agcaccaggc gtttaagggc accaataact
5340gccttaaaaa aattacgccc cgccctgcca ctcatcgcag tactgttgta attcattaag
5400cattctgccg acatggaagc catcacagac ggcatgatga acctgaatcg ccagcggcat
5460cagcaccttg tcgccttgcg tataatattt gcccatggtg aaaacggggg cgaagaagtt
5520gtccatattg gccacgttta aatcaaaact ggtgaaactc acccagggat tggctgagac
5580gaaaaacata ttctcaataa accctttagg gaaataggcc aggttttcac cgtaacacgc
5640cacatcttgc gaatatatgt gtagaaactg ccggaaatcg tcgtggtatt cactccagag
5700cgatgaaaac gtttcagttt gctcatggaa aacggtgtaa caagggtgaa cactatccca
5760tatcaccagc tcaccgtctt tcattgccat acg
579351951DNAEscherichia coliCDS(1)..(948) 51atg cag atc ctg ttg gcc aac
ccg cgt ggt ttt tgt gcc ggg gta gac 48Met Gln Ile Leu Leu Ala Asn
Pro Arg Gly Phe Cys Ala Gly Val Asp1 5 10
15cgc gct atc agc att gtt gaa aac gcg ctg gcc att tac
ggc gca ccg 96Arg Ala Ile Ser Ile Val Glu Asn Ala Leu Ala Ile Tyr
Gly Ala Pro 20 25 30ata tat
gtc cgt cac gaa gtg gta cat aac cgc tat gtg gtc gat agc 144Ile Tyr
Val Arg His Glu Val Val His Asn Arg Tyr Val Val Asp Ser 35
40 45ttg cgt gag cgt ggg gct atc ttt att gag
cag att agc gaa gta ccg 192Leu Arg Glu Arg Gly Ala Ile Phe Ile Glu
Gln Ile Ser Glu Val Pro 50 55 60gac
ggc gcg atc ctg att ttc tcc gca cac ggt gtt tct cag gcg gta 240Asp
Gly Ala Ile Leu Ile Phe Ser Ala His Gly Val Ser Gln Ala Val65
70 75 80cgt aac gaa gca aaa agt
cgc gat ttg acg gtg ttt gat gcc acc tgt 288Arg Asn Glu Ala Lys Ser
Arg Asp Leu Thr Val Phe Asp Ala Thr Cys 85
90 95ccg ctg gtg acc aaa gtg cat atg gaa gtc gcc cgc
gcc agt cgc cgt 336Pro Leu Val Thr Lys Val His Met Glu Val Ala Arg
Ala Ser Arg Arg 100 105 110ggc
gaa gaa tct att ctc atc ggt cac gcc ggg cac ccg gaa gtg gaa 384Gly
Glu Glu Ser Ile Leu Ile Gly His Ala Gly His Pro Glu Val Glu 115
120 125ggg aca atg ggc cag tac agt aac ccg
gaa ggg gga atg tat ctg gtc 432Gly Thr Met Gly Gln Tyr Ser Asn Pro
Glu Gly Gly Met Tyr Leu Val 130 135
140gaa tcg ccg gac gat gtg tgg aaa ctg acg gtc aaa aac gaa gag aag
480Glu Ser Pro Asp Asp Val Trp Lys Leu Thr Val Lys Asn Glu Glu Lys145
150 155 160ctc tcc ttt atg
acc cag acc acg ctg tcg gtg gat gac acg tct gat 528Leu Ser Phe Met
Thr Gln Thr Thr Leu Ser Val Asp Asp Thr Ser Asp 165
170 175gtg atc gac gcg ctg cgt aaa cgc ttc ccg
aaa att gtc ggt ccg cgc 576Val Ile Asp Ala Leu Arg Lys Arg Phe Pro
Lys Ile Val Gly Pro Arg 180 185
190aaa gat gac atc tgc tac gcc acg act aac cgt cag gaa gcg gta cgc
624Lys Asp Asp Ile Cys Tyr Ala Thr Thr Asn Arg Gln Glu Ala Val Arg
195 200 205gcc ctg gca gaa cag gcg gaa
gtt gtg ttg gtg gtc ggt tcg aaa aac 672Ala Leu Ala Glu Gln Ala Glu
Val Val Leu Val Val Gly Ser Lys Asn 210 215
220tcc tcc aac tcc aac cgt ctg gcg gag ctg gcc cag cgt atg ggc aaa
720Ser Ser Asn Ser Asn Arg Leu Ala Glu Leu Ala Gln Arg Met Gly Lys225
230 235 240cgc gcg ttt ttg
att gac gat gcg aaa gac atc cag gaa gag tgg gtg 768Arg Ala Phe Leu
Ile Asp Asp Ala Lys Asp Ile Gln Glu Glu Trp Val 245
250 255aaa gag gtt aaa tgc gtc ggc gtg act gcg
ggc gca tcg gct ccg gat 816Lys Glu Val Lys Cys Val Gly Val Thr Ala
Gly Ala Ser Ala Pro Asp 260 265
270att ctg gtg cag aat gtg gtg gca cgt ttg cag cag ctg ggc ggt ggt
864Ile Leu Val Gln Asn Val Val Ala Arg Leu Gln Gln Leu Gly Gly Gly
275 280 285gaa gcc att ccg ctg gaa ggc
cgt gaa gaa aac att gtt ttc gaa gtg 912Glu Ala Ile Pro Leu Glu Gly
Arg Glu Glu Asn Ile Val Phe Glu Val 290 295
300ccg aaa gag ctg cgt gtc gat att cgt gaa gtc gat taa
951Pro Lys Glu Leu Arg Val Asp Ile Arg Glu Val Asp305
310 31552316PRTEscherichia coli 52Met Gln Ile Leu Leu Ala
Asn Pro Arg Gly Phe Cys Ala Gly Val Asp1 5
10 15Arg Ala Ile Ser Ile Val Glu Asn Ala Leu Ala Ile
Tyr Gly Ala Pro 20 25 30Ile
Tyr Val Arg His Glu Val Val His Asn Arg Tyr Val Val Asp Ser 35
40 45Leu Arg Glu Arg Gly Ala Ile Phe Ile
Glu Gln Ile Ser Glu Val Pro 50 55
60Asp Gly Ala Ile Leu Ile Phe Ser Ala His Gly Val Ser Gln Ala Val65
70 75 80Arg Asn Glu Ala Lys
Ser Arg Asp Leu Thr Val Phe Asp Ala Thr Cys 85
90 95Pro Leu Val Thr Lys Val His Met Glu Val Ala
Arg Ala Ser Arg Arg 100 105
110Gly Glu Glu Ser Ile Leu Ile Gly His Ala Gly His Pro Glu Val Glu
115 120 125Gly Thr Met Gly Gln Tyr Ser
Asn Pro Glu Gly Gly Met Tyr Leu Val 130 135
140Glu Ser Pro Asp Asp Val Trp Lys Leu Thr Val Lys Asn Glu Glu
Lys145 150 155 160Leu Ser
Phe Met Thr Gln Thr Thr Leu Ser Val Asp Asp Thr Ser Asp
165 170 175Val Ile Asp Ala Leu Arg Lys
Arg Phe Pro Lys Ile Val Gly Pro Arg 180 185
190Lys Asp Asp Ile Cys Tyr Ala Thr Thr Asn Arg Gln Glu Ala
Val Arg 195 200 205Ala Leu Ala Glu
Gln Ala Glu Val Val Leu Val Val Gly Ser Lys Asn 210
215 220Ser Ser Asn Ser Asn Arg Leu Ala Glu Leu Ala Gln
Arg Met Gly Lys225 230 235
240Arg Ala Phe Leu Ile Asp Asp Ala Lys Asp Ile Gln Glu Glu Trp Val
245 250 255Lys Glu Val Lys Cys
Val Gly Val Thr Ala Gly Ala Ser Ala Pro Asp 260
265 270Ile Leu Val Gln Asn Val Val Ala Arg Leu Gln Gln
Leu Gly Gly Gly 275 280 285Glu Ala
Ile Pro Leu Glu Gly Arg Glu Glu Asn Ile Val Phe Glu Val 290
295 300Pro Lys Glu Leu Arg Val Asp Ile Arg Glu Val
Asp305 310
315539795DNAArtificialpBScyclogcpElytB2 53gtggcacttt tcggggaaat
gtgcgcggaa cccctatttg tttatttttc taaatacatt 60caaatatgta tccgctcatg
agacaataac cctgataaat gcttcaataa tattgaaaaa 120ggaagagtat gagtattcaa
catttccgtg tcgcccttat tccctttttt gcggcatttt 180gccttcctgt ttttgctcac
ccagaaacgc tggtgaaagt aaaagatgct gaagatcagt 240tgggtgcacg agtgggttac
atcgaactgg atctcaacag cggtaagatc cttgagagtt 300ttcgccccga agaacgtttt
ccaatgatga gcacttttaa agttctgcta tgtggcgcgg 360tattatcccg tattgacgcc
gggcaagagc aactcggtcg ccgcatacac tattctcaga 420atgacttggt tgagtactca
ccagtcacag aaaagcatct tacggatggc atgacagtaa 480gagaattatg cagtgctgcc
ataaccatga gtgataacac tgcggccaac ttacttctga 540caacgatcgg aggaccgaag
gagctaaccg cttttttgca caacatgggg gatcatgtaa 600ctcgccttga tcgttgggaa
ccggagctga atgaagccat accaaacgac gagcgtgaca 660ccacgatgcc tgtagcaatg
gcaacaacgt tgcgcaaact attaactggc gaactactta 720ctctagcttc ccggcaacaa
ttaatagact ggatggaggc ggataaagtt gcaggaccac 780ttctgcgctc ggcccttccg
gctggctggt ttattgctga taaatctgga gccggtgagc 840gtgggtctcg cggtatcatt
gcagcactgg ggccagatgg taagccctcc cgtatcgtag 900ttatctacac gacggggagt
caggcaacta tggatgaacg aaatagacag atcgctgaga 960taggtgcctc actgattaag
cattggtaac tgtcagacca agtttactca tatatacttt 1020agattgattt aaaacttcat
ttttaattta aaaggatcta ggtgaagatc ctttttgata 1080atctcatgac caaaatccct
taacgtgagt tttcgttcca ctgagcgtca gaccccgtag 1140aaaagatcaa aggatcttct
tgagatcctt tttttctgcg cgtaatctgc tgcttgcaaa 1200caaaaaaacc accgctacca
gcggtggttt gtttgccgga tcaagagcta ccaactcttt 1260ttccgaaggt aactggcttc
agcagagcgc agataccaaa tactgtcctt ctagtgtagc 1320cgtagttagg ccaccacttc
aagaactctg tagcaccgcc tacatacctc gctctgctaa 1380tcctgttacc agtggctgct
gccagtggcg ataagtcgtg tcttaccggg ttggactcaa 1440gacgatagtt accggataag
gcgcagcggt cgggctgaac ggggggttcg tgcacacagc 1500ccagcttgga gcgaacgacc
tacaccgaac tgagatacct acagcgtgag ctatgagaaa 1560gcgccacgct tcccgaaggg
agaaaggcgg acaggtatcc ggtaagcggc agggtcggaa 1620caggagagcg cacgagggag
cttccagggg gaaacgcctg gtatctttat agtcctgtcg 1680ggtttcgcca cctctgactt
gagcgtcgat ttttgtgatg ctcgtcaggg gggcggagcc 1740tatggaaaaa cgccagcaac
gcggcctttt tacggttcct ggccttttgc tggccttttg 1800ctcacatgtt ctttcctgcg
ttatcccctg attctgtgga taaccgtatt accgcctttg 1860agtgagctga taccgctcgc
cgcagccgaa cgaccgagcg cagcgagtca gtgagcgagg 1920aagcggaaga gcgcccaata
cgcaaaccgc ctctccccgc gcgttggccg attcattaat 1980gcagctggca cgacaggttt
cccgactgga aagcgggcag tgagcgcaac gcaattaatg 2040tgagttagct cactcattag
gcaccccagg ctttacactt tatgcttccg gctcgtatgt 2100tgtgtggaat tgtgagcgga
taacaatttc acacaggaaa cagctatgac catgattacg 2160ccaagcgcgc aattaaccct
cactaaaggg aacaaaagct ggagctccac cgcgggagga 2220gaaattaacc atgcataacc
aggctccaat tcaacgtaga aaatcaacac gtatttacgt 2280tgggaatgtg ccgattggcg
atggtgctcc catcgccgta cagtccatga ccaatacgcg 2340tacgacagac gtcgaagcaa
cggtcaatca aatcaaggcg ctggaacgcg ttggcgctga 2400tatcgtccgt gtatccgtac
cgacgatgga cgcggcagaa gcgttcaaac tcatcaaaca 2460gcaggttaac gtgccgctgg
tggctgacat ccacttcgac tatcgcattg cgctgaaagt 2520agcggaatac ggcgtcgatt
gtctgcgtat taaccctggc aatatcggta atgaagagcg 2580tattcgcatg gtggttgact
gtgcgcgcga taaaaacatt ccgatccgta ttggcgttaa 2640cgccggatcg ctggaaaaag
atctgcaaga aaagtatggc gaaccgacgc cgcaggcgtt 2700gctggaatct gccatgcgtc
atgttgatca tctcgatcgc ctgaacttcg atcagttcaa 2760agtcagcgtg aaagcgtctg
acgtcttcct cgctgttgag tcttatcgtt tgctggcaaa 2820acagatcgat cagccgttgc
atctggggat caccgaagcc ggtggtgcgc gcagcggggc 2880agtaaaatcc gccattggtt
taggtctgct gctgtctgaa ggcatcggcg acacgctgcg 2940cgtatcgctg gcggccgatc
cggtcgaaga gatcaaagtc ggtttcgata ttttgaaatc 3000gctgcgtatc cgttcgcgag
ggatcaactt catcgcctgc ccgacctgtt cgcgtcagga 3060atttgatgtt atcggtacgg
ttaacgcgct ggagcaacgc ctggaagata tcatcactcc 3120gatggacgtt tcgattatcg
gctgcgtggt gaatggccca ggtgaggcgc tggtttctac 3180actcggcgtc accggcggca
acaagaaaag cggcctctat gaagatggcg tgcgcaaaga 3240ccgtctggac aacaacgata
tgatcgacca gctggaagca cgcattcgtg cgaaagccag 3300tcagctggac gaagcgcgtc
gaattgacgt tcagcaggtt gaaaaataag tcgacgagga 3360gaaattaacc atgcagatcc
tgttggccaa cccgcgtggt ttttgtgccg gggtagaccg 3420cgctatcagc attgttgaaa
acgcgctggc catttacggc gcaccgatat atgtccgtca 3480cgaagtggta cataaccgct
atgtggtcga tagcttgcgt gagcgtgggg ctatctttat 3540tgagcagatt agcgaagtac
cggacggcgc gatcctgatt ttctccgcac acggtgtttc 3600tcaggcggta cgtaacgaag
caaaaagtcg cgatttgacg gtgtttgatg ccacctgtcc 3660gctggtgacc aaagtgcata
tggaagtcgc ccgcgccagt cgccgtggcg aagaatctat 3720tctcatcggt cacgccgggc
acccggaagt ggaagggaca atgggccagt acagtaaccc 3780ggaaggggga atgtatctgg
tcgaatcgcc ggacgatgtg tggaaactga cggtcaaaaa 3840cgaagagaag ctctccttta
tgacccagac cacgctgtcg gtggatgaca cgtctgatgt 3900gatcgacgcg ctgcgtaaac
gcttcccgaa aattgtcggt ccgcgcaaag atgacatctg 3960ctacgccacg actaaccgtc
aggaagcggt acgcgccctg gcagaacagg cggaagttgt 4020gttggtggtc ggttcgaaaa
actcctccaa ctccaaccgt ctggcggagc tggcccagcg 4080tatgggcaaa cgcgcgtttt
tgattgacga tgcgaaagac atccaggaag agtgggtgaa 4140agaggttaaa tgcgtcggcg
tgactgcggg cgcatcggct ccggatattc tggtgcagaa 4200tgtggtggca cgtttgcagc
agctgggcgg tggtgaagcc attccgctgg aaggccgtga 4260agaaaacatt gttttcgaag
tgccgaaaga gctgcgtgtc gatattcgtg aagtcgatta 4320agcggccgct ctagaactag
tggatccccc gggctgcagg aattcgagga gaaattaacc 4380atgtatatcg ggatagatct
tggcacctcg ggcgtaaaag ttattttgct caacgagcag 4440ggtgaggtgg ttgctgcgca
aacggaaaag ctgaccgttt cgcgcccgca tccactctgg 4500tcggaacaag acccggaaca
gtggtggcag gcaactgatc gcgcaatgaa agctctgggc 4560gatcagcatt ctctgcagga
cgttaaagca ttgggtattg ccggccagat gcacggagca 4620accttgctgg atgctcagca
acgggtgtta cgccctgcca ttttgtggaa cgacgggcgc 4680tgtgcgcaag agtgcacttt
gctggaagcg cgagttccgc aatcgcgggt gattaccggc 4740aacctgatga tgcccggatt
tactgcgcct aaattgctat gggttcagcg gcatgagccg 4800gagatattcc gtcaaatcga
caaagtatta ttaccgaaag attacttgcg tctgcgtatg 4860acgggggagt ttgccagcga
tatgtctgac gcagctggca ccatgtggct ggatgtcgca 4920aagcgtgact ggagtgacgt
catgctgcag gcttgcgact tatctcgtga ccagatgccc 4980gcattatacg aaggcagcga
aattactggt gctttgttac ctgaagttgc gaaagcgtgg 5040ggtatggcga cggtgccagt
tgtcgcaggc ggtggcgaca atgcagctgg tgcagttggt 5100gtgggaatgg ttgatgctaa
tcaggcaatg ttatcgctgg ggacgtcggg ggtctatttt 5160gctgtcagcg aagggttctt
aagcaagcca gaaagcgccg tacatagctt ttgccatgcg 5220ctaccgcaac gttggcattt
aatgtctgtg atgctgagtg cagcgtcgtg tctggattgg 5280gccgcgaaat taaccggcct
gagcaatgtc ccagctttaa tcgctgcagc tcaacaggct 5340gatgaaagtg ccgagccagt
ttggtttctg ccttatcttt ccggcgagcg tacgccacac 5400aataatcccc aggcgaaggg
ggttttcttt ggtttgactc atcaacatgg ccccaatgaa 5460ctggcgcgag cagtgctgga
aggcgtgggt tatgcgctgg cagatggcat ggatgtcgtg 5520catgcctgcg gtattaaacc
gcaaagtgtt acgttgattg ggggcggggc gcgtagtgag 5580tactggcgtc agatgctggc
ggatatcagc ggtcagcagc tcgattaccg tacggggggg 5640gatgtggggc cagcactggg
cgcagcaagg ctggcgcaga tcgcggcgaa tccagagaaa 5700tcgctcattg aattgttgcc
gcaactaccg ttagaacagt cgcatctacc agatgcgcag 5760cgttatgccg cttatcagcc
acgacgagaa acgttccgtc gcctctatca gcaacttctg 5820ccattaatgg cgtaaaagct
tgaggagaaa ttaaccatga agcaactcac cattctgggc 5880tcgaccggct cgattggttg
cagcacgctg gacgtggtgc gccataatcc cgaacacttc 5940cgcgtagttg cgctggtggc
aggcaaaaat gtcactcgca tggtagaaca gtgcctggaa 6000ttctctcccc gctatgccgt
aatggacgat gaagcgagtg cgaaacttct taaaacgatg 6060ctacagcaac agggtagccg
caccgaagtc ttaagtgggc aacaagccgc ttgcgatatg 6120gcagcgcttg aggatgttga
tcaggtgatg gcagccattg ttggcgctgc tgggctgtta 6180cctacgcttg ctgcgatccg
cgcgggtaaa accattttgc tggccaataa agaatcactg 6240gttacctgcg gacgtctgtt
tatggacgcc gtaaagcaga gcaaagcgca attgttaccg 6300gtcgatagcg aacataacgc
catttttcag agtttaccgc aacctatcca gcataatctg 6360ggatacgctg accttgagca
aaatggcgtg gtgtccattt tacttaccgg gtctggtggc 6420cctttccgtg agacgccatt
gcgcgatttg gcaacaatga cgccggatca agcctgccgt 6480catccgaact ggtcgatggg
gcgtaaaatt tctgtcgatt cggctaccat gatgaacaaa 6540ggtctggaat acattgaagc
gcgttggctg tttaacgcca gcgccagcca gatggaagtg 6600ctgattcacc cgcagtcagt
gattcactca atggtgcgct atcaggacgg cagtgttctg 6660gcgcagctgg gggaaccgga
tatgcgtacg ccaattgccc acaccatggc atggccgaat 6720cgcgtgaact ctggcgtgaa
gccgctcgat ttttgcaaac taagtgcgtt gacatttgcc 6780gcaccggatt atgatcgtta
tccatgcctg aaactggcga tggaggcgtt cgaacaaggc 6840caggcagcga cgacagcatt
gaatgccgca aacgaaatca ccgttgctgc ttttcttgcg 6900caacaaatcc gctttacgga
tatcgctgcg ttgaatttat ccgtactgga aaaaatggat 6960atgcgcgaac cacaatgtgt
ggacgatgtg ttatctgttg atgcgaacgc gcgtgaagtc 7020gccagaaaag aggtgatgcg
tctcgcaagc tgagtcgacg aggagaaatt aaccatggca 7080accactcatt tggatgtttg
cgccgtggtt ccggcggccg gatttggccg tcgaatgcaa 7140acggaatgtc ctaagcaata
tctctcaatc ggtaatcaaa ccattcttga acactcggtg 7200catgcgctgc tggcgcatcc
ccgggtgaaa cgtgtcgtca ttgccataag tcctggcgat 7260agccgttttg cacaacttcc
tctggcgaat catccgcaaa tcaccgttgt agatggcggt 7320gatgagcgtg ccgattccgt
gctggcaggt ctgaaagccg ctggcgacgc gcagtgggta 7380ttggtgcatg acgccgctcg
tccttgtttg catcaggatg acctcgcgcg attgttggcg 7440ttgagcgaaa ccagccgcac
gggggggatc ctcgccgcac cagtgcgcga tactatgaaa 7500cgtgccgaac cgggcaaaaa
tgccattgct cataccgttg atcgcaacgg cttatggcac 7560gcgctgacgc cgcaattttt
ccctcgtgag ctgttacatg actgtctgac gcgcgctcta 7620aatgaaggcg cgactattac
cgacgaagcc tcggcgctgg aatattgcgg attccatcct 7680cagttggtcg aaggccgtgc
ggataacatt aaagtcacgc gcccggaaga tttggcactg 7740gccgagtttt acctcacccg
aaccatccat caggagaata cataatgcga attggacacg 7800gttttgacgt acatgccttt
ggcggtgaag gcccaattat cattggtggc gtacgcattc 7860cttacgaaaa aggattgctg
gcgcattctg atggcgacgt ggcgctccat gcgttgaccg 7920atgcattgct tggcgcggcg
gcgctggggg atatcggcaa gctgttcccg gataccgatc 7980cggcatttaa aggtgccgat
agccgcgagc tgctacgcga agcctggcgt cgtattcagg 8040cgaagggtta tacccttggc
aacgtcgatg tcactatcat cgctcaggca ccgaagatgt 8100tgccgcacat tccacaaatg
cgcgtgttta ttgccgaaga tctcggctgc catatggatg 8160atgttaacgt gaaagccact
actacggaaa aactgggatt taccggacgt ggggaaggga 8220ttgcctgtga agcggtggcg
ctactcatta aggcaacaaa atgactcgag gaggagaaat 8280taaccatgcg gacacagtgg
ccctctccgg caaaacttaa tctgttttta tacattaccg 8340gtcagcgtgc ggatggttac
cacacgctgc aaacgctgtt tcagtttctt gattacggcg 8400acaccatcag cattgagctt
cgtgacgatg gggatattcg tctgttaacg cccgttgaag 8460gcgtggaaca tgaagataac
ctgatcgttc gcgcagcgcg attgttgatg aaaactgcgg 8520cagacagcgg gcgtcttccg
acgggaagcg gtgcgaatat cagcattgac aagcgtttgc 8580cgatgggcgg cggtctcggc
ggtggttcat ccaatgccgc gacggtcctg gtggcattaa 8640atcatctctg gcaatgcggg
ctaagcatgg atgagctggc ggaaatgggg ctgacgctgg 8700gcgcagatgt tcctgtcttt
gttcgggggc atgccgcgtt tgccgaaggc gttggtgaaa 8760tactaacgcc ggtggatccg
ccagagaagt ggtatctggt ggcgcaccct ggtgtaagta 8820ttccgactcc ggtgattttt
aaagatcctg aactcccgcg caatacgcca aaaaggtcaa 8880tagaaacgtt gctaaaatgt
gaattcagca atgattgcga ggttatcgca agaaaacgtt 8940ttcgcgaggt tgatgcggtg
ctttcctggc tgttagaata cgccccgtcg cgcctgactg 9000ggacaggggc ctgtgtcttt
gctgaatttg atacagagtc tgaagcccgc caggtgctag 9060agcaagcccc ggaatggctc
aatggctttg tggcgaaagg cgctaatctt tccccattgc 9120acagagccat gctttaaggt
acccaattcg ccctatagtg agtcgtatta cgcgcgctca 9180ctggccgtcg ttttacaacg
tcgtgactgg gaaaaccctg gcgttaccca acttaatcgc 9240cttgcagcac atcccccttt
cgccagctgg cgtaatagcg aagaggcccg caccgatcgc 9300ccttcccaac agttgcgcag
cctgaatggc gaatggaaat tgtaagcgtt aatattttgt 9360taaaattcgc gttaaatttt
tgttaaatca gctcattttt taaccaatag gccgaaatcg 9420gcaaaatccc ttataaatca
aaagaataga ccgagatagg gttgagtgtt gttccagttt 9480ggaacaagag tccactatta
aagaacgtgg actccaacgt caaagggcga aaaaccgtct 9540atcagggcga tggcccacta
cgtgaaccat caccctaatc aagttttttg gggtcgaggt 9600gccgtaaagc actaaatcgg
aaccctaaag ggagcccccg atttagagct tgacggggaa 9660agccggcgaa cgtggcgaga
aaggaaggga agaaagcgaa aggagcgggc gctagggcgc 9720tggcaagtgt agcggtcacg
ctgcgcgtaa ccaccacacc cgccgcgctt aatgcgccgc 9780tacagggcgc gtcag
9795541986DNAArabidopsis
thalianaCDS(1)..(1983) 54aag acg gtg aga agg aag act cgt act gtt atg gtt
gga aat gtc gcc 48Lys Thr Val Arg Arg Lys Thr Arg Thr Val Met Val
Gly Asn Val Ala1 5 10
15ctt gga agc gaa cat ccg ata agg att caa acg atg act act tcg gat
96Leu Gly Ser Glu His Pro Ile Arg Ile Gln Thr Met Thr Thr Ser Asp
20 25 30aca aaa gat att act gga act
gtt gat gag gtt atg aga ata gcg gat 144Thr Lys Asp Ile Thr Gly Thr
Val Asp Glu Val Met Arg Ile Ala Asp 35 40
45aaa gga gct gat att gta agg ata act gtt caa ggg aag aaa gag
gcg 192Lys Gly Ala Asp Ile Val Arg Ile Thr Val Gln Gly Lys Lys Glu
Ala 50 55 60gat gcg tgc ttt gaa ata
aaa gat aaa ctc gtt cag ctt aat tac aat 240Asp Ala Cys Phe Glu Ile
Lys Asp Lys Leu Val Gln Leu Asn Tyr Asn65 70
75 80aca ccg ctg gtt gca ggt att cat ttt gcc cct
act gta gcc tta cga 288Thr Pro Leu Val Ala Gly Ile His Phe Ala Pro
Thr Val Ala Leu Arg 85 90
95gtc gct gaa tgc ttt gac aag atc cgt gtc aac ccc gga aat ttt gcg
336Val Ala Glu Cys Phe Asp Lys Ile Arg Val Asn Pro Gly Asn Phe Ala
100 105 110gac agg cgg gcc cag ttt
gag acg ata gat tat aca gaa gat gaa tat 384Asp Arg Arg Ala Gln Phe
Glu Thr Ile Asp Tyr Thr Glu Asp Glu Tyr 115 120
125cag aaa gaa ctc cag cat atc gag cag gtc ttc act cct ttg
gtt gag 432Gln Lys Glu Leu Gln His Ile Glu Gln Val Phe Thr Pro Leu
Val Glu 130 135 140aaa tgc aaa aag tac
ggg aga gca atg cgt att ggg aca aat cat gga 480Lys Cys Lys Lys Tyr
Gly Arg Ala Met Arg Ile Gly Thr Asn His Gly145 150
155 160agt ctt tct gac cgt atc atg agc tat tac
ggg gat tct ccc cga gga 528Ser Leu Ser Asp Arg Ile Met Ser Tyr Tyr
Gly Asp Ser Pro Arg Gly 165 170
175atg gtt gaa tct gcg ttt gag ttt gca aga ata tgt cgg aaa tta gac
576Met Val Glu Ser Ala Phe Glu Phe Ala Arg Ile Cys Arg Lys Leu Asp
180 185 190tat cac aac ttt gtt ttc
tca atg aaa gcg agc aac cca gtg atc atg 624Tyr His Asn Phe Val Phe
Ser Met Lys Ala Ser Asn Pro Val Ile Met 195 200
205gtc cag gcg tac cgt tta ctt gtg gct gag atg tat gtt cat
gga tgg 672Val Gln Ala Tyr Arg Leu Leu Val Ala Glu Met Tyr Val His
Gly Trp 210 215 220gat tat cct ttg cat
ttg gga gtt act gag gca gga gaa ggc gaa gat 720Asp Tyr Pro Leu His
Leu Gly Val Thr Glu Ala Gly Glu Gly Glu Asp225 230
235 240gga cgg atg aaa tct gcg att gga att ggg
acg ctt ctt cag gac ggg 768Gly Arg Met Lys Ser Ala Ile Gly Ile Gly
Thr Leu Leu Gln Asp Gly 245 250
255ctc ggt gac aca aca aga gtt tca ctg acg gag cca cca gaa gag gag
816Leu Gly Asp Thr Thr Arg Val Ser Leu Thr Glu Pro Pro Glu Glu Glu
260 265 270ata gat ccc tgc agg cga
ttg gct aac ctc ggg aca aaa gct gcc aaa 864Ile Asp Pro Cys Arg Arg
Leu Ala Asn Leu Gly Thr Lys Ala Ala Lys 275 280
285ctt caa caa ggc gct gca ccg ttt gaa gaa aag cat agg cat
tac ttt 912Leu Gln Gln Gly Ala Ala Pro Phe Glu Glu Lys His Arg His
Tyr Phe 290 295 300gat ttt cag cgt cgg
acg ggt gat cta cct gta caa aaa gag gga gaa 960Asp Phe Gln Arg Arg
Thr Gly Asp Leu Pro Val Gln Lys Glu Gly Glu305 310
315 320gag gtt gat tac aga aat gtc ctt cac cgt
gat ggt tct gtt ctg atg 1008Glu Val Asp Tyr Arg Asn Val Leu His Arg
Asp Gly Ser Val Leu Met 325 330
335tcg att tct ctg gat caa cta aag gca cct gaa ctc ctc tac aga tca
1056Ser Ile Ser Leu Asp Gln Leu Lys Ala Pro Glu Leu Leu Tyr Arg Ser
340 345 350ctc gcc aca aag ctt gtc
gtg ggt atg cca ttc aag gat ctg gca act 1104Leu Ala Thr Lys Leu Val
Val Gly Met Pro Phe Lys Asp Leu Ala Thr 355 360
365gtt gat tca atc tta tta aga gag cta ccg cct gta gat gat
caa gtg 1152Val Asp Ser Ile Leu Leu Arg Glu Leu Pro Pro Val Asp Asp
Gln Val 370 375 380gct cgt ttg gct ctc
aaa cgg ttg att gat gtc agt atg gga gtt ata 1200Ala Arg Leu Ala Leu
Lys Arg Leu Ile Asp Val Ser Met Gly Val Ile385 390
395 400gca cct tta tca gag caa cta aca aag cca
ttg ccc aat gcc atg gtt 1248Ala Pro Leu Ser Glu Gln Leu Thr Lys Pro
Leu Pro Asn Ala Met Val 405 410
415ctt gtc aac ctc aag gaa cta tct ggt ggc gct tac aag ctt ctc cct
1296Leu Val Asn Leu Lys Glu Leu Ser Gly Gly Ala Tyr Lys Leu Leu Pro
420 425 430gaa ggt aca cgc ttg gtt
gtc tct cta cga ggc gat gag cct tac gag 1344Glu Gly Thr Arg Leu Val
Val Ser Leu Arg Gly Asp Glu Pro Tyr Glu 435 440
445gag ctt gaa ata ctc aaa aac att gat gct act atg att ctc
cat gat 1392Glu Leu Glu Ile Leu Lys Asn Ile Asp Ala Thr Met Ile Leu
His Asp 450 455 460gta cct ttc act gaa
gac aaa gtt agc aga gta cat gca gct cgg agg 1440Val Pro Phe Thr Glu
Asp Lys Val Ser Arg Val His Ala Ala Arg Arg465 470
475 480cta ttc gag ttc tta tcc gag aat tca gtt
aac ttt cct gtt att cat 1488Leu Phe Glu Phe Leu Ser Glu Asn Ser Val
Asn Phe Pro Val Ile His 485 490
495cgc ata aac ttc cca acc gga atc cac aga gac gaa ttg gtg att cat
1536Arg Ile Asn Phe Pro Thr Gly Ile His Arg Asp Glu Leu Val Ile His
500 505 510gca ggg aca tat gct gga
ggc ctt ctt gtg gat gga cta ggt gat ggc 1584Ala Gly Thr Tyr Ala Gly
Gly Leu Leu Val Asp Gly Leu Gly Asp Gly 515 520
525gta atg ctc gaa gca cct gac caa gat ttt gat ttt ctt agg
aat act 1632Val Met Leu Glu Ala Pro Asp Gln Asp Phe Asp Phe Leu Arg
Asn Thr 530 535 540tcc ttc aac tta tta
caa gga tgc aga atg cgt aac act aag acg gaa 1680Ser Phe Asn Leu Leu
Gln Gly Cys Arg Met Arg Asn Thr Lys Thr Glu545 550
555 560tat gta tcg tgc ccg tct tgt gga aga acg
ctt ttc gac ttg caa gaa 1728Tyr Val Ser Cys Pro Ser Cys Gly Arg Thr
Leu Phe Asp Leu Gln Glu 565 570
575atc agc gcc gag atc cga gaa aag act tcc cat tta cct ggc gtt tcg
1776Ile Ser Ala Glu Ile Arg Glu Lys Thr Ser His Leu Pro Gly Val Ser
580 585 590atc gca atc atg gga tgc
att gtg aat gga cca gga gaa atg gca gat 1824Ile Ala Ile Met Gly Cys
Ile Val Asn Gly Pro Gly Glu Met Ala Asp 595 600
605gct gat ttc gga tat gta ggt ggt tct ccc gga aaa atc gac
ctt tat 1872Ala Asp Phe Gly Tyr Val Gly Gly Ser Pro Gly Lys Ile Asp
Leu Tyr 610 615 620gtc gga aag acg gtg
gtg aag cgt ggg ata gct atg acg gag gca aca 1920Val Gly Lys Thr Val
Val Lys Arg Gly Ile Ala Met Thr Glu Ala Thr625 630
635 640gat gct ctg atc ggt ctg atc aaa gaa cat
ggt cgt tgg gtc gac ccg 1968Asp Ala Leu Ile Gly Leu Ile Lys Glu His
Gly Arg Trp Val Asp Pro 645 650
655ccc gtg gct gat gag tag
1986Pro Val Ala Asp Glu 66055661PRTArabidopsis thaliana
55Lys Thr Val Arg Arg Lys Thr Arg Thr Val Met Val Gly Asn Val Ala1
5 10 15Leu Gly Ser Glu His Pro
Ile Arg Ile Gln Thr Met Thr Thr Ser Asp 20 25
30Thr Lys Asp Ile Thr Gly Thr Val Asp Glu Val Met Arg
Ile Ala Asp 35 40 45Lys Gly Ala
Asp Ile Val Arg Ile Thr Val Gln Gly Lys Lys Glu Ala 50
55 60Asp Ala Cys Phe Glu Ile Lys Asp Lys Leu Val Gln
Leu Asn Tyr Asn65 70 75
80Thr Pro Leu Val Ala Gly Ile His Phe Ala Pro Thr Val Ala Leu Arg
85 90 95Val Ala Glu Cys Phe Asp
Lys Ile Arg Val Asn Pro Gly Asn Phe Ala 100
105 110Asp Arg Arg Ala Gln Phe Glu Thr Ile Asp Tyr Thr
Glu Asp Glu Tyr 115 120 125Gln Lys
Glu Leu Gln His Ile Glu Gln Val Phe Thr Pro Leu Val Glu 130
135 140Lys Cys Lys Lys Tyr Gly Arg Ala Met Arg Ile
Gly Thr Asn His Gly145 150 155
160Ser Leu Ser Asp Arg Ile Met Ser Tyr Tyr Gly Asp Ser Pro Arg Gly
165 170 175Met Val Glu Ser
Ala Phe Glu Phe Ala Arg Ile Cys Arg Lys Leu Asp 180
185 190Tyr His Asn Phe Val Phe Ser Met Lys Ala Ser
Asn Pro Val Ile Met 195 200 205Val
Gln Ala Tyr Arg Leu Leu Val Ala Glu Met Tyr Val His Gly Trp 210
215 220Asp Tyr Pro Leu His Leu Gly Val Thr Glu
Ala Gly Glu Gly Glu Asp225 230 235
240Gly Arg Met Lys Ser Ala Ile Gly Ile Gly Thr Leu Leu Gln Asp
Gly 245 250 255Leu Gly Asp
Thr Thr Arg Val Ser Leu Thr Glu Pro Pro Glu Glu Glu 260
265 270Ile Asp Pro Cys Arg Arg Leu Ala Asn Leu
Gly Thr Lys Ala Ala Lys 275 280
285Leu Gln Gln Gly Ala Ala Pro Phe Glu Glu Lys His Arg His Tyr Phe 290
295 300Asp Phe Gln Arg Arg Thr Gly Asp
Leu Pro Val Gln Lys Glu Gly Glu305 310
315 320Glu Val Asp Tyr Arg Asn Val Leu His Arg Asp Gly
Ser Val Leu Met 325 330
335Ser Ile Ser Leu Asp Gln Leu Lys Ala Pro Glu Leu Leu Tyr Arg Ser
340 345 350Leu Ala Thr Lys Leu Val
Val Gly Met Pro Phe Lys Asp Leu Ala Thr 355 360
365Val Asp Ser Ile Leu Leu Arg Glu Leu Pro Pro Val Asp Asp
Gln Val 370 375 380Ala Arg Leu Ala Leu
Lys Arg Leu Ile Asp Val Ser Met Gly Val Ile385 390
395 400Ala Pro Leu Ser Glu Gln Leu Thr Lys Pro
Leu Pro Asn Ala Met Val 405 410
415Leu Val Asn Leu Lys Glu Leu Ser Gly Gly Ala Tyr Lys Leu Leu Pro
420 425 430Glu Gly Thr Arg Leu
Val Val Ser Leu Arg Gly Asp Glu Pro Tyr Glu 435
440 445Glu Leu Glu Ile Leu Lys Asn Ile Asp Ala Thr Met
Ile Leu His Asp 450 455 460Val Pro Phe
Thr Glu Asp Lys Val Ser Arg Val His Ala Ala Arg Arg465
470 475 480Leu Phe Glu Phe Leu Ser Glu
Asn Ser Val Asn Phe Pro Val Ile His 485
490 495Arg Ile Asn Phe Pro Thr Gly Ile His Arg Asp Glu
Leu Val Ile His 500 505 510Ala
Gly Thr Tyr Ala Gly Gly Leu Leu Val Asp Gly Leu Gly Asp Gly 515
520 525Val Met Leu Glu Ala Pro Asp Gln Asp
Phe Asp Phe Leu Arg Asn Thr 530 535
540Ser Phe Asn Leu Leu Gln Gly Cys Arg Met Arg Asn Thr Lys Thr Glu545
550 555 560Tyr Val Ser Cys
Pro Ser Cys Gly Arg Thr Leu Phe Asp Leu Gln Glu 565
570 575Ile Ser Ala Glu Ile Arg Glu Lys Thr Ser
His Leu Pro Gly Val Ser 580 585
590Ile Ala Ile Met Gly Cys Ile Val Asn Gly Pro Gly Glu Met Ala Asp
595 600 605Ala Asp Phe Gly Tyr Val Gly
Gly Ser Pro Gly Lys Ile Asp Leu Tyr 610 615
620Val Gly Lys Thr Val Val Lys Arg Gly Ile Ala Met Thr Glu Ala
Thr625 630 635 640Asp Ala
Leu Ile Gly Leu Ile Lys Glu His Gly Arg Trp Val Asp Pro
645 650 655Pro Val Ala Asp Glu
660562226DNAArabidopsis thalianaCDS(1)..(2223) 56atg gcg act gga gta ttg
cca gct ccg gtt tct ggg atc aag ata ccg 48Met Ala Thr Gly Val Leu
Pro Ala Pro Val Ser Gly Ile Lys Ile Pro1 5
10 15gat tcg aaa gtc ggg ttt ggt aaa agc atg aat ctt
gtg aga att tgt 96Asp Ser Lys Val Gly Phe Gly Lys Ser Met Asn Leu
Val Arg Ile Cys 20 25 30gat
gtt agg agt cta aga tct gct agg aga aga gtt tcg gtt atc cgg 144Asp
Val Arg Ser Leu Arg Ser Ala Arg Arg Arg Val Ser Val Ile Arg 35
40 45aat tca aac caa ggc tct gat tta gct
gag ctt caa cct gca tcc gaa 192Asn Ser Asn Gln Gly Ser Asp Leu Ala
Glu Leu Gln Pro Ala Ser Glu 50 55
60gga agc cct ctc tta gtg cca aga cag aaa tat tgt gaa tca ttg cat
240Gly Ser Pro Leu Leu Val Pro Arg Gln Lys Tyr Cys Glu Ser Leu His65
70 75 80aag acg gtg aga agg
aag act cgt act gtt atg gtt gga aat gtc gcc 288Lys Thr Val Arg Arg
Lys Thr Arg Thr Val Met Val Gly Asn Val Ala 85
90 95ctt gga agc gaa cat ccg ata agg att caa acg
atg act act tcg gat 336Leu Gly Ser Glu His Pro Ile Arg Ile Gln Thr
Met Thr Thr Ser Asp 100 105
110aca aaa gat att act gga act gtt gat gag gtt atg aga ata gcg gat
384Thr Lys Asp Ile Thr Gly Thr Val Asp Glu Val Met Arg Ile Ala Asp
115 120 125aaa gga gct gat att gta agg
ata act gtt caa ggg aag aaa gag gcg 432Lys Gly Ala Asp Ile Val Arg
Ile Thr Val Gln Gly Lys Lys Glu Ala 130 135
140gat gcg tgc ttt gaa ata aaa gat aaa ctc gtt cag ctt aat tac aat
480Asp Ala Cys Phe Glu Ile Lys Asp Lys Leu Val Gln Leu Asn Tyr Asn145
150 155 160aca ccg ctg gtt
gca ggt att cat ttt gcc cct act gta gcc tta cga 528Thr Pro Leu Val
Ala Gly Ile His Phe Ala Pro Thr Val Ala Leu Arg 165
170 175gtc gct gaa tgc ttt gac aag atc cgt gtc
aac ccc gga aat ttt gcg 576Val Ala Glu Cys Phe Asp Lys Ile Arg Val
Asn Pro Gly Asn Phe Ala 180 185
190gac agg cgg gcc cag ttt gag acg ata gat tat aca gaa gat gaa tat
624Asp Arg Arg Ala Gln Phe Glu Thr Ile Asp Tyr Thr Glu Asp Glu Tyr
195 200 205cag aaa gaa ctc cag cat atc
gag cag gtc ttc act cct ttg gtt gag 672Gln Lys Glu Leu Gln His Ile
Glu Gln Val Phe Thr Pro Leu Val Glu 210 215
220aaa tgc aaa aag tac ggg aga gca atg cgt att ggg aca aat cat gga
720Lys Cys Lys Lys Tyr Gly Arg Ala Met Arg Ile Gly Thr Asn His Gly225
230 235 240agt ctt tct gac
cgt atc atg agc tat tac ggg gat tct ccc cga gga 768Ser Leu Ser Asp
Arg Ile Met Ser Tyr Tyr Gly Asp Ser Pro Arg Gly 245
250 255atg gtt gaa tct gcg ttt gag ttt gca aga
ata tgt cgg aaa tta gac 816Met Val Glu Ser Ala Phe Glu Phe Ala Arg
Ile Cys Arg Lys Leu Asp 260 265
270tat cac aac ttt gtt ttc tca atg aaa gcg agc aac cca gtg atc atg
864Tyr His Asn Phe Val Phe Ser Met Lys Ala Ser Asn Pro Val Ile Met
275 280 285gtc cag gcg tac cgt tta ctt
gtg gct gag atg tat gtt cat gga tgg 912Val Gln Ala Tyr Arg Leu Leu
Val Ala Glu Met Tyr Val His Gly Trp 290 295
300gat tat cct ttg cat ttg gga gtt act gag gca gga gaa ggc gaa gat
960Asp Tyr Pro Leu His Leu Gly Val Thr Glu Ala Gly Glu Gly Glu Asp305
310 315 320gga cgg atg aaa
tct gcg att gga att ggg acg ctt ctt cag gac ggg 1008Gly Arg Met Lys
Ser Ala Ile Gly Ile Gly Thr Leu Leu Gln Asp Gly 325
330 335ctc ggt gac aca aca aga gtt tca ctg acg
gag cca cca gaa gag gag 1056Leu Gly Asp Thr Thr Arg Val Ser Leu Thr
Glu Pro Pro Glu Glu Glu 340 345
350ata gat ccc tgc agg cga ttg gct aac ctc ggg aca aaa gct gcc aaa
1104Ile Asp Pro Cys Arg Arg Leu Ala Asn Leu Gly Thr Lys Ala Ala Lys
355 360 365ctt caa caa ggc gct gca ccg
ttt gaa gaa aag cat agg cat tac ttt 1152Leu Gln Gln Gly Ala Ala Pro
Phe Glu Glu Lys His Arg His Tyr Phe 370 375
380gat ttt cag cgt cgg acg ggt gat cta cct gta caa aaa gag gga gaa
1200Asp Phe Gln Arg Arg Thr Gly Asp Leu Pro Val Gln Lys Glu Gly Glu385
390 395 400gag gtt gat tac
aga aat gtc ctt cac cgt gat ggt tct gtt ctg atg 1248Glu Val Asp Tyr
Arg Asn Val Leu His Arg Asp Gly Ser Val Leu Met 405
410 415tcg att tct ctg gat caa cta aag gca cct
gaa ctc ctc tac aga tca 1296Ser Ile Ser Leu Asp Gln Leu Lys Ala Pro
Glu Leu Leu Tyr Arg Ser 420 425
430ctc gcc aca aag ctt gtc gtg ggt atg cca ttc aag gat ctg gca act
1344Leu Ala Thr Lys Leu Val Val Gly Met Pro Phe Lys Asp Leu Ala Thr
435 440 445gtt gat tca atc tta tta aga
gag cta ccg cct gta gat gat caa gtg 1392Val Asp Ser Ile Leu Leu Arg
Glu Leu Pro Pro Val Asp Asp Gln Val 450 455
460gct cgt ttg gct ctc aaa cgg ttg att gat gtc agt atg gga gtt ata
1440Ala Arg Leu Ala Leu Lys Arg Leu Ile Asp Val Ser Met Gly Val Ile465
470 475 480gca cct tta tca
gag caa cta aca aag cca ttg ccc aat gcc atg gtt 1488Ala Pro Leu Ser
Glu Gln Leu Thr Lys Pro Leu Pro Asn Ala Met Val 485
490 495ctt gtc aac ctc aag gaa cta tct ggt ggc
gct tac aag ctt ctc cct 1536Leu Val Asn Leu Lys Glu Leu Ser Gly Gly
Ala Tyr Lys Leu Leu Pro 500 505
510gaa ggt aca cgc ttg gtt gtc tct cta cga ggc gat gag cct tac gag
1584Glu Gly Thr Arg Leu Val Val Ser Leu Arg Gly Asp Glu Pro Tyr Glu
515 520 525gag ctt gaa ata ctc aaa aac
att gat gct act atg att ctc cat gat 1632Glu Leu Glu Ile Leu Lys Asn
Ile Asp Ala Thr Met Ile Leu His Asp 530 535
540gta cct ttc act gaa gac aaa gtt agc aga gta cat gca gct cgg agg
1680Val Pro Phe Thr Glu Asp Lys Val Ser Arg Val His Ala Ala Arg Arg545
550 555 560cta ttc gag ttc
tta tcc gag aat tca gtt aac ttt cct gtt att cat 1728Leu Phe Glu Phe
Leu Ser Glu Asn Ser Val Asn Phe Pro Val Ile His 565
570 575cgc ata aac ttc cca acc gga atc cac aga
gac gaa ttg gtg att cat 1776Arg Ile Asn Phe Pro Thr Gly Ile His Arg
Asp Glu Leu Val Ile His 580 585
590gca ggg aca tat gct gga ggc ctt ctt gtg gat gga cta ggt gat ggc
1824Ala Gly Thr Tyr Ala Gly Gly Leu Leu Val Asp Gly Leu Gly Asp Gly
595 600 605gta atg ctc gaa gca cct gac
caa gat ttt gat ttt ctt agg aat act 1872Val Met Leu Glu Ala Pro Asp
Gln Asp Phe Asp Phe Leu Arg Asn Thr 610 615
620tcc ttc aac tta tta caa gga tgc aga atg cgt aac act aag acg gaa
1920Ser Phe Asn Leu Leu Gln Gly Cys Arg Met Arg Asn Thr Lys Thr Glu625
630 635 640tat gta tcg tgc
ccg tct tgt gga aga acg ctt ttc gac ttg caa gaa 1968Tyr Val Ser Cys
Pro Ser Cys Gly Arg Thr Leu Phe Asp Leu Gln Glu 645
650 655atc agc gcc gag atc cga gaa aag act tcc
cat tta cct ggc gtt tcg 2016Ile Ser Ala Glu Ile Arg Glu Lys Thr Ser
His Leu Pro Gly Val Ser 660 665
670atc gca atc atg gga tgc att gtg aat gga cca gga gaa atg gca gat
2064Ile Ala Ile Met Gly Cys Ile Val Asn Gly Pro Gly Glu Met Ala Asp
675 680 685gct gat ttc gga tat gta ggt
ggt tct ccc gga aaa atc gac ctt tat 2112Ala Asp Phe Gly Tyr Val Gly
Gly Ser Pro Gly Lys Ile Asp Leu Tyr 690 695
700gtc gga aag acg gtg gtg aag cgt ggg ata gct atg acg gag gca aca
2160Val Gly Lys Thr Val Val Lys Arg Gly Ile Ala Met Thr Glu Ala Thr705
710 715 720gat gct ctg atc
ggt ctg atc aaa gaa cat ggt cgt tgg gtc gac ccg 2208Asp Ala Leu Ile
Gly Leu Ile Lys Glu His Gly Arg Trp Val Asp Pro 725
730 735ccc gtg gct gat gag tag
2226Pro Val Ala Asp Glu
74057741PRTArabidopsis thaliana 57Met Ala Thr Gly Val Leu Pro Ala Pro Val
Ser Gly Ile Lys Ile Pro1 5 10
15Asp Ser Lys Val Gly Phe Gly Lys Ser Met Asn Leu Val Arg Ile Cys
20 25 30Asp Val Arg Ser Leu Arg
Ser Ala Arg Arg Arg Val Ser Val Ile Arg 35 40
45Asn Ser Asn Gln Gly Ser Asp Leu Ala Glu Leu Gln Pro Ala
Ser Glu 50 55 60Gly Ser Pro Leu Leu
Val Pro Arg Gln Lys Tyr Cys Glu Ser Leu His65 70
75 80Lys Thr Val Arg Arg Lys Thr Arg Thr Val
Met Val Gly Asn Val Ala 85 90
95Leu Gly Ser Glu His Pro Ile Arg Ile Gln Thr Met Thr Thr Ser Asp
100 105 110Thr Lys Asp Ile Thr
Gly Thr Val Asp Glu Val Met Arg Ile Ala Asp 115
120 125Lys Gly Ala Asp Ile Val Arg Ile Thr Val Gln Gly
Lys Lys Glu Ala 130 135 140Asp Ala Cys
Phe Glu Ile Lys Asp Lys Leu Val Gln Leu Asn Tyr Asn145
150 155 160Thr Pro Leu Val Ala Gly Ile
His Phe Ala Pro Thr Val Ala Leu Arg 165
170 175Val Ala Glu Cys Phe Asp Lys Ile Arg Val Asn Pro
Gly Asn Phe Ala 180 185 190Asp
Arg Arg Ala Gln Phe Glu Thr Ile Asp Tyr Thr Glu Asp Glu Tyr 195
200 205Gln Lys Glu Leu Gln His Ile Glu Gln
Val Phe Thr Pro Leu Val Glu 210 215
220Lys Cys Lys Lys Tyr Gly Arg Ala Met Arg Ile Gly Thr Asn His Gly225
230 235 240Ser Leu Ser Asp
Arg Ile Met Ser Tyr Tyr Gly Asp Ser Pro Arg Gly 245
250 255Met Val Glu Ser Ala Phe Glu Phe Ala Arg
Ile Cys Arg Lys Leu Asp 260 265
270Tyr His Asn Phe Val Phe Ser Met Lys Ala Ser Asn Pro Val Ile Met
275 280 285Val Gln Ala Tyr Arg Leu Leu
Val Ala Glu Met Tyr Val His Gly Trp 290 295
300Asp Tyr Pro Leu His Leu Gly Val Thr Glu Ala Gly Glu Gly Glu
Asp305 310 315 320Gly Arg
Met Lys Ser Ala Ile Gly Ile Gly Thr Leu Leu Gln Asp Gly
325 330 335Leu Gly Asp Thr Thr Arg Val
Ser Leu Thr Glu Pro Pro Glu Glu Glu 340 345
350Ile Asp Pro Cys Arg Arg Leu Ala Asn Leu Gly Thr Lys Ala
Ala Lys 355 360 365Leu Gln Gln Gly
Ala Ala Pro Phe Glu Glu Lys His Arg His Tyr Phe 370
375 380Asp Phe Gln Arg Arg Thr Gly Asp Leu Pro Val Gln
Lys Glu Gly Glu385 390 395
400Glu Val Asp Tyr Arg Asn Val Leu His Arg Asp Gly Ser Val Leu Met
405 410 415Ser Ile Ser Leu Asp
Gln Leu Lys Ala Pro Glu Leu Leu Tyr Arg Ser 420
425 430Leu Ala Thr Lys Leu Val Val Gly Met Pro Phe Lys
Asp Leu Ala Thr 435 440 445Val Asp
Ser Ile Leu Leu Arg Glu Leu Pro Pro Val Asp Asp Gln Val 450
455 460Ala Arg Leu Ala Leu Lys Arg Leu Ile Asp Val
Ser Met Gly Val Ile465 470 475
480Ala Pro Leu Ser Glu Gln Leu Thr Lys Pro Leu Pro Asn Ala Met Val
485 490 495Leu Val Asn Leu
Lys Glu Leu Ser Gly Gly Ala Tyr Lys Leu Leu Pro 500
505 510Glu Gly Thr Arg Leu Val Val Ser Leu Arg Gly
Asp Glu Pro Tyr Glu 515 520 525Glu
Leu Glu Ile Leu Lys Asn Ile Asp Ala Thr Met Ile Leu His Asp 530
535 540Val Pro Phe Thr Glu Asp Lys Val Ser Arg
Val His Ala Ala Arg Arg545 550 555
560Leu Phe Glu Phe Leu Ser Glu Asn Ser Val Asn Phe Pro Val Ile
His 565 570 575Arg Ile Asn
Phe Pro Thr Gly Ile His Arg Asp Glu Leu Val Ile His 580
585 590Ala Gly Thr Tyr Ala Gly Gly Leu Leu Val
Asp Gly Leu Gly Asp Gly 595 600
605Val Met Leu Glu Ala Pro Asp Gln Asp Phe Asp Phe Leu Arg Asn Thr 610
615 620Ser Phe Asn Leu Leu Gln Gly Cys
Arg Met Arg Asn Thr Lys Thr Glu625 630
635 640Tyr Val Ser Cys Pro Ser Cys Gly Arg Thr Leu Phe
Asp Leu Gln Glu 645 650
655Ile Ser Ala Glu Ile Arg Glu Lys Thr Ser His Leu Pro Gly Val Ser
660 665 670Ile Ala Ile Met Gly Cys
Ile Val Asn Gly Pro Gly Glu Met Ala Asp 675 680
685Ala Asp Phe Gly Tyr Val Gly Gly Ser Pro Gly Lys Ile Asp
Leu Tyr 690 695 700Val Gly Lys Thr Val
Val Lys Arg Gly Ile Ala Met Thr Glu Ala Thr705 710
715 720Asp Ala Leu Ile Gly Leu Ile Lys Glu His
Gly Arg Trp Val Asp Pro 725 730
735Pro Val Ala Asp Glu 740581401DNAArabidopsis thaliana
58atggctgttg cgctccaatt cagccgatta tgcgttcgac cggatacttt cgtgcgggag
60aatcatctct ctggatccgg atctctccgc cgccggaaag ctttatcagt ccggtgctcg
120tctggcgatg agaacgctcc ttcgccatcg gtggtgatgg actccgattt cgacgccaag
180gtgttccgta agaacttgac gagaagcgat aattacaatc gtaaagggtt cggtcataag
240gaggagacac tcaagctcat gaatcgagag tacaccagtg atatattgga gacactgaaa
300acaaatgggt atacttattc ttggggagat gttactgtga aactcgctaa agcatatggt
360ttttgctggg gtgttgagcg tgctgttcag attgcatatg aagcacgaaa gcagtttcca
420gaggagaggc tttggattac taacgaaatc attcataacc cgaccgtcaa taagaggttg
480gaagatatgg atgttaaaat tattccggtt gaggattcaa agaaacagtt tgatgtagta
540gagaaagatg atgtggttat ccttcctgcg tttggagctg gtgttgacga gatgtatgtt
600cttaatgata aaaaggtgca aattgttgac acgacttgtc cttgggtgac aaaggtctgg
660aacacggttg agaagcacaa gaagggggaa tacacatcag taatccatgg taaatataat
720catgaagaga cgattgcaac tgcgtctttt gcaggaaagt acatcattgt aaagaacatg
780aaagaggcaa attacgtttg tgattacatt ctcggtggcc aatacgatgg atctagctcc
840acaaaagagg agttcatgga gaaattcaaa tacgcaattt cgaagggttt cgatcccgac
900aatgaccttg tcaaagttgg tattgcaaac caaacaacga tgctaaaggg agaaacagag
960gagataggaa gattactcga gacaacaatg atgcgcaagt atggagtgga aaatgtaagc
1020ggacatttca tcagcttcaa cacaatatgc gacgctactc aagagcgaca agacgcaatc
1080tatgagctag tggaagagaa gattgacctc atgctagtgg ttggcggatg gaattcaagt
1140aacacctctc accttcagga aatctcagag gcacggggaa tcccatctta ctggatcgat
1200agtgagaaac ggataggacc tgggaataaa atagcctata agctccacta tggagaactg
1260gtcgagaagg aaaactttct cccaaaggga ccaataacaa tcggtgtgac atcaggtgca
1320tcaaccccgg ataaggtcgt ggaagatgct ttggtgaagg tgttcgacat taaacgtgaa
1380gagttattgc agctggcttg a
1401
User Contributions:
comments("1"); ?> comment_form("1"); ?>Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
User Contributions:
Comment about this patent or add new information about this topic: