Patent application title: METHOD FOR THE PRODUCTION OF 2-BUTANOL
Inventors:
Michael G. Bramucci (Boothwyn, PA, US)
Butamax(tm) Advanced Biofuels Llc
Butamax(tm) Advanced Biofuels Llc
Dennis Flint (Newark, DE, US)
Edward S. Miller, Jr. (Knoxville, TN, US)
Edward S. Miller, Jr. (Knoxville, TN, US)
Vasantha Nagarajan (Wilmington, DE, US)
Natalia Sedkova (Cherry Hill, NJ, US)
Manjari Singh (West Chester, PA, US)
Manjari Singh (West Chester, PA, US)
Tina K. Van Dyk (Wilmington, DE, US)
Tina K. Van Dyk (Wilmington, DE, US)
Assignees:
BUTAMAX(TM) ADVANCED BIOFUELS LLC
IPC8 Class: AC12N1563FI
USPC Class:
435 3
Class name: Chemistry: molecular biology and microbiology condition responsive control process
Publication date: 2013-08-29
Patent application number: 20130224728
Abstract:
A method for the production of 2-butanol by fermentation using a
microbial production host is disclosed. The method employs a reduction in
temperature during the fermentation process that results in a more robust
tolerance of the production host to the butanol product.Claims:
1-38. (canceled)
39. A recombinant microbial host cell comprising DNA molecules encoding polypeptides that catalyze each of the following substrate to product conversions: i) pyruvate to alpha-acetolactate, ii) alpha-acetolactate to acetoin, iii) acetoin to 2,3-butanediol, iv) 2,3-butanediol to 2-butanone, and v) 2-butanone to 2-butanol; wherein the polypeptide that catalyzes the substrate to product conversion of pyruvate to alpha-acetolactate is acetolactate synthase, the polypeptide that catalyzes the substrate to product conversion of alpha-acetolactate to acetoin is acetolactate decarboxylase, the polypeptide that catalyzes the substrate to product conversion of acetoin to 2,3-butanediol is butanediol dehydrogenase, the polypeptide that catalyzes the substrate to product conversion of 2,3-butanediol to 2-butanone is diol dehydratase or glycerol dehydratase, and the polypeptide that catalyzes the substrate to product conversion of 2-butanone to 2-butanol is butanol dehydrogenase.
40. The recombinant microbial host cell of claim 39, wherein the acetolactate synthase has an amino acid sequence having at least 95% identity to an amino acid sequence selected from the group consisting of SEQ ID NO: 4, SEQ ID NO: 77, and SEQ ID NO: 79 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix.
41. The recombinant microbial host cell of claim 39, wherein the acetolactate decarboxylase has an amino acid sequence having at least 95% identity to an amino acid sequence selected from the group consisting of SEQ ID NO: 2, SEQ ID NO: 81, and SEQ ID NO: 83 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix.
42. The recombinant microbial host cell of claim 39, wherein the butanediol dehydrogenase has an amino acid sequence having at least 95% identity to an amino acid sequence selected from the group consisting of SEQ ID NO: 6, SEQ ID NO: 85, SEQ ID NO: 87, and SEQ ID NO: 89 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix.
43. The recombinant microbial host cell of claim 39, wherein the diol dehydratase or glycerol dehydratase comprises fused large, medium, and small subunits and has at least 95% identity to an amino acid sequence comprising all three of the amino acid sequences encoding large, medium, and small subunits, selected from the group consisting of: a) SEQ ID NO: 8, SEQ ID NO: 10, and SEQ ID NO: 12; b) SEQ ID NO: 93, SEQ ID NO: 95, and SEQ ID NO: 97; c) SEQ ID NO: 99, SEQ ID NO: 101, and SEQ ID NO: 103; d) SEQ ID NO: 105, SEQ ID NO: 107, and SEQ ID NO: 109; e) SEQ ID NO: 135, SEQ ID NO: 136, and SEQ ID NO: 137; f) SEQ ID NO: 138, SEQ ID NO: 139, and SEQ ID NO: 140; g) SEQ ID NO: 146, SEQ ID NO: 148, and SEQ ID NO: 150; h) SEQ ID NO: 141, SEQ ID NO: 142, and SEQ ID NO: 143; and i) SEQ ID NO: 164, SEQ ID NO: 165, and SEQ ID NO: 166; based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix.
44. The recombinant microbial host cell of claim 39, wherein the butanol dehydrogenase has an amino acid sequence having at least 95% identity to an amino acid sequence selected from the group consisting of SEQ ID NO: 14, SEQ ID NO: 72, SEQ ID NO: 75, and SEQ ID NO: 91 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix.
45. The recombinant microbial host cell of claim 39, wherein the acetolactate synthase has an amino acid sequence having at least 95% identity to an amino acid sequence selected from the group consisting of SEQ ID NO: 4, SEQ ID NO: 77, and SEQ ID NO: 79 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix; wherein the acetolactate decarboxylase has an amino acid sequence having at least 95% identity to an amino acid sequence selected from the group consisting of SEQ ID NO: 2, SEQ ID NO: 81, and SEQ ID NO: 83 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix; wherein the butanediol dehydrogenase has an amino acid sequence having at least 95% identity to an amino acid sequence selected from the group consisting of SEQ ID NO: 6, SEQ ID NO: 85, SEQ ID NO: 87, and SEQ ID NO: 89 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix; wherein the diol dehydratase or glycerol dehydratase comprises fused large, medium, and small subunits and has at least 95% identity to an amino acid sequence comprising all three of the amino acid sequences encoding large, medium, and small subunits, selected from the group consisting of: a) SEQ ID NO: 8, SEQ ID NO: 10, and SEQ ID NO: 12; b) SEQ ID NO: 93, SEQ ID NO: 95, and SEQ ID NO: 97; c) SEQ ID NO: 99, SEQ ID NO: 101, and SEQ ID NO: 103; d) SEQ ID NO: 105, SEQ ID NO: 107, and SEQ ID NO: 109; e) SEQ ID NO: 135, SEQ ID NO: 136, and SEQ ID NO: 137; f) SEQ ID NO: 138, SEQ ID NO: 139, and SEQ ID NO: 140; g) SEQ ID NO: 146, SEQ ID NO: 148, and SEQ ID NO: 150; h) SEQ ID NO: 141, SEQ ID NO: 142, and SEQ ID NO: 143; and i) SEQ ID NO: 164, SEQ ID NO: 165, and SEQ ID NO: 166; based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix; and wherein the butanol dehydrogenase has an amino acid sequence having at least 95% identity to an amino acid sequence selected from the group consisting of SEQ ID NO: 14, SEQ ID NO: 72, SEQ ID NO: 75, and SEQ ID NO: 91 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix.
46. The recombinant microbial host cell of claim 39, wherein the recombinant microbial host cell is selected from the group consisting of Clostridium, Zymomonas, Escherichia, Salmonella, Rhodococcus, Pseudomonas, Bacillus, Lactobacillus, Enterococcus, Alcaligenes, Klebsiella, Paenibacillus, Arthrobacter, Corynebacterium, Brevibacterium, Saccharomyces, Pichia, Candida, and Hansenula.
47. The recombinant microbial host cell of claim 46, wherein the recombinant microbial host cell is selected from the group consisting of Escherichia coli, Alcaligenes eutrophus, Bacillus licheniformis, Paenibacillus macerans, Rhodococcus erythropolis, Pseudomonas putida, Lactobacillus plantarum, Enterococcus faecium, Enterococcus gallinarium, Enterococcus faecalis, Bacillus subtilis, and Saccharomyces cerevisiae.
48. A recombinant microbial host cell comprising DNA molecules encoding polypeptides that catalyze each of the following substrate to product conversions: i) pyruvate to alpha-acetolactate, ii) alpha-acetolactate to acetoin, iii) acetoin to 3-amino-2-butanol, iv) 3-amino-2-butanol to 3-amino-2-butanol phosphate, v) 3-amino-2-butanol phosphate to 2-butanone, and vi) 2-butanone to 2-butanol; wherein the polypeptide that catalyzes the substrate to product conversion of pyruvate to alpha-acetolactate is acetolactate synthase, the polypeptide that catalyzes the substrate to product conversion of alpha-acetolactate to acetoin is acetolactate decarboxylase, the polypeptide that catalyzes the substrate to product conversion of acetoin to 3-amino-2-butanol is acetoin aminase, the polypeptide that catalyzes the substrate to product conversion of 3-amino-2-butanol to 3-amino-2-butanol phosphate is aminobutanol kinase, the polypeptide that catalyzes the substrate to product conversion of 3-amino-2-butanol phosphate to 2-butanone is aminobutanol phosphate phospho-lyase, and the polypeptide that catalyzes the substrate to product conversion of 2-butanone to 2-butanol is butanol dehydrogenase.
49. The recombinant microbial host cell of claim 48, wherein the acetolactate synthase has an amino acid sequence having at least 95% identity to an amino acid sequence selected from the group consisting of SEQ ID NO: 4, SEQ ID NO: 77, and SEQ ID NO: 79 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix.
50. The recombinant microbial host cell of claim 48, wherein the acetolactate decarboxylase has an amino acid sequence having at least 95% identity to an amino acid sequence selected from the group consisting of SEQ ID NO: 2, SEQ ID NO: 81, and SEQ ID NO: 83 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix.
51. The recombinant microbial host cell of claim 48, wherein the acetoin aminase has an amino acid sequence having at least 95% identity to an amino acid sequence as set forth in SEQ ID NO: 122 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix.
52. The recombinant microbial host cell of claim 48, wherein the aminobutanol kinase has an amino acid sequence having at least 95% identity to an amino acid sequence as set forth in SEQ ID NO: 124 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix.
53. The recombinant microbial host cell of claim 48, wherein the aminobutanol phosphate phospho-lyase has an amino acid sequence having at least 95% identity to an amino acid sequence as set forth in SEQ ID NO: 126 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix.
54. The recombinant microbial host cell of claim 48, wherein the butanol dehydrogenase has an amino acid sequence having at least 95% identity to an amino acid sequence selected from the group consisting of SEQ ID NO: 14, SEQ ID NO: 72, SEQ ID NO: 75, and SEQ ID NO: 91 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix.
55. The recombinant microbial host cell of claim 48, wherein the acetolactate synthase has an amino acid sequence having at least 95% identity to an amino acid sequence selected from the group consisting of SEQ ID NO: 4, SEQ ID NO: 77, and SEQ ID NO: 79 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix; wherein the acetolactate decarboxylase has an amino acid sequence having at least 95% identity to an amino acid sequence selected from the group consisting of SEQ ID NO: 2, SEQ ID NO: 81, and SEQ ID NO: 83 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix; wherein the acetoin aminase has an amino acid sequence having at least 95% identity to an amino acid sequence as set forth in SEQ ID NO: 122 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix; wherein the aminobutanol kinase has an amino acid sequence having at least 95% identity to an amino acid sequence as set forth in SEQ ID NO: 124 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix; wherein the aminobutanol phosphate phospho-lyase has an amino acid sequence having at least 95% identity to an amino acid sequence as set forth in SEQ ID NO: 126 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix; and wherein the butanol dehydrogenase has an amino acid sequence having at least 95% identity to an amino acid sequence selected from the group consisting of SEQ ID NO: 14, SEQ ID NO: 72, SEQ ID NO: 75, and SEQ ID NO: 91 based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix.
56. The recombinant microbial host cell of claim 48, wherein the recombinant microbial host cell is selected from the group consisting of Clostridium, Zymomonas, Escherichia, Salmonella, Rhodococcus, Pseudomonas, Bacillus, Lactobacillus, Enterococcus, Alcaligenes, Klebsiella, Paenibacillus, Arthrobacter, Corynebacterium, Brevibacterium, Saccharomyces, Pichia, Candida, and Hansenula.
57. The recombinant microbial host cell of claim 56, wherein the recombinant microbial host cell is selected from the group consisting of Escherichia coli, Alcaligenes eutrophus, Bacillus licheniformis, Paenibacillus macerans, Rhodococcus erythropolis, Pseudomonas putida, Lactobacillus plantarum, Enterococcus faecium, Enterococcus gallinarium, Enterococcus faecalis, Bacillus subtilis, and Saccharomyces cerevisiae.
58. A method to reduce the sensitivity of a recombinant microbial host cell to 2-butanol comprising: a) providing a recombinant microbial host cell which produces 2-butanol, wherein the recombinant microbial host cell comprises DNA molecules encoding polypeptides that catalyze each of the following substrate to product conversions: i) pyruvate to alpha-acetolactate, ii) alpha-acetolactate to acetoin, iii) acetoin to 2,3-butanediol, iv) 2,3-butanediol to 2-butanone, and v) 2-butanone to 2-butanol; or DNA molecules encoding polypeptides that catalyze each of the following substrate to product conversions: i) pyruvate to alpha-acetolactate, ii) alpha-acetolactate to acetoin, iii) acetoin to 3-amino-2-butanol, iv) 3-amino-2-butanol to 3-amino-2-butanol phosphate, v) 3-amino-2-butanol phosphate to 2-butanone, and vi) 2-butanone to 2-butanol; wherein at least one of the DNA molecules is heterologous to the recombinant microbial host cell; b) growing the recombinant microbial host cell in a fermentation culture; and c) determining the metabolic activity of the fermentation culture by monitoring one or more metabolic parameters selected from optical density, pH, respiratory quotient, fermentable carbon substrate utilization, CO2 production, and 1-butanol production.
59. The method of claim 58, further comprising the step adjusting the one or more metabolic parameters to support the metabolic activity.
60. The method of claim 59, wherein a decrease in one or more of the metabolic parameters indicates a decrease in metabolic activity.
61. The method of claim 60, wherein the adjusting the one or more metabolic parameters is lowering the temperature of the fermentation culture when a decrease in metabolic activity is detected.
62. The method of claim 58, wherein the polypeptide that catalyzes the substrate to product conversion of pyruvate to alpha-acetolactate is acetolactate synthase, the polypeptide that catalyzes the substrate to product conversion of alpha-acetolactate to acetoin is acetolactate decarboxylase, the polypeptide that catalyzes the substrate to product conversion of acetoin to 2,3-butanediol is butanediol dehydrogenase, the polypeptide that catalyzes the substrate to product conversion of 2,3-butanediol to 2-butanone is diol dehydratase or glycerol dehydratase, the polypeptide that catalyzes the substrate to product conversion of acetoin to 3-amino-2-butanol is acetoin aminase, the polypeptide that catalyzes the substrate to product conversion of 3-amino-2-butanol to 3-amino-2-butanol phosphate is aminobutanol kinase, the polypeptide that catalyzes the substrate to product conversion of 3-amino-2-butanol phosphate to 2-butanone is aminobutanol phosphate phospho-lyase, and the polypeptide that catalyzes the substrate to product conversion of 2-butanone to 2-butanol is butanol dehydrogenase.
Description:
FIELD OF THE INVENTION
[0001] The invention relates to a method for the production of 2-butanol by fermentation using a recombinant microbial host. Specifically, the method employs a decrease in temperature during fermentation that results in more robust tolerance of the production host to the 2-butanol product.
BACKGROUND OF THE INVENTION
[0002] Butanol is an important industrial chemical, useful as a fuel additive, as a feedstock chemical in the plastics industry, and as a foodgrade extractant in the food and flavor industry. Each year 10 to 12 billion pounds of butanol are produced by petrochemical means and the need for this commodity chemical will likely increase.
[0003] Methods for the chemical synthesis of 2-butanol are known, such as n-butene hydration (Ullmann's Encyclopedia of Industrial Chemistry, 6th edition, 2003, Wiley-VCHVerlag GmbH and Co., Weinheim, Germany, Vol. 5, pp. 716-719). These processes use starting materials derived from petrochemicals and are generally expensive, and are not environmentally friendly. The production of 2-butanol from plant-derived raw materials would minimize greenhouse gas emissions and would represent an advance in the art.
[0004] Methods for producing 2-butanol by biotransformation of other organic chemicals are also known. For example, Stampfer et al. (WO 03/078615) describe the production of secondary alcohols, such as 2-butanol, by the reduction of ketones which is catalyzed by an alcohol dehydrogenase enzyme obtained from Rhodococcus ruber. Similarly, Kojima et al. (EP 0645453) describe a method for preparing secondary alcohols, such as 2-butanol, by reduction of ketones which is catalyzed by a secondary alcohol dehydrogenase enzyme obtained from Candida parapsilosis. Additionally, Kuehnle et al. (EP 1149918) describe a process that produces both 1-butanol and 2-butanol by the oxidation of hydrocarbons by various strains of Rhodococcus ruber. The process favored 1-butanol production with a selectivity of 93.8%.
[0005] The production of 2-butanol by certain strains of Lactobacilli is also known (Speranza et. al. J. Agric. Food Chem. (1997) 45:3476-3480). The 2-butanol is produced by the transformation of meso-2,3-butanediol. The production of 2-butanol from acetolactate and acetoin by these Lactobacilli strains was also demonstrated.
[0006] Recombinant microbial production hosts expressing 2-butanol biosynthetic pathways are described in co-pending and commonly owned U.S. Patent Application Publication No. US20070259410A1. However, biological production of 2-butanol is believed to be limited by 2-butanol toxicity to the host microorganism used in the fermentation.
[0007] Some microbial strains that are tolerant to 2-butanol are known in the art (co-pending and commonly owned U.S. patent application Ser. Nos. 11/743,220 and 11/761,497). However, biological methods of producing 2-butanol to higher levels are required for cost effective commercial production.
[0008] There have been reports describing the effect of temperature on the tolerance of some microbial strains to ethanol. For example, Amartey et al. (Biotechnol. Lett. 13(9):627-632 (1991)) disclose that Bacillus stearothermophillus is less tolerant to ethanol at 70° C. than at 60° C. Herrero et al. (Appl. Environ. Microbiol. 40(3):571-577 (1980)) report that the optimum growth temperature of a wild-type strain of Clostridium thermocellum decreases as the concentration of ethanol challenge increases, whereas the optimum growth temperature of an ethanol-tolerant mutant remains constant. Brown et al. (Biotechnol. Lett. 4(4):269-274 (1982)) disclose that the yeast Saccharomyces uvarum is more resistant to growth inhibition by ethanol at temperatures 5° C. and 10° C. below its growth optimum of 35° C. However, fermentation became more resistant to ethanol inhibition with increasing temperature. Additionally, Van Uden (CRC Crit. Rev. Biotechnol. 1(3):263-273 (1984)) report that ethanol and other alkanols depress the maximum and the optimum growth temperature for growth of Saccharomyces cerevisiae while thermal death is enhanced. Moreover, Lewis et al. (U.S. Patent Application Publication No. 2004/0234649) describe methods for producing high levels of ethanol during fermentation of plant material comprising decreasing the temperature during saccharifying, fermenting, or simultaneously saccharifying and fermenting
[0009] Much less is known about the effect of temperature on the tolerance of microbial strains to butanols. Harada (Hakko Kyokaishi 20:155-156 (1962)) discloses that the yield of 1-butanol in acetone-butanol-ethanol (ABE) fermentation is increased from 18.4%-18.7% to 19.1%-21.2% by lowering the temperature from 30° C. to 28° C. when the growth of the bacteria reaches a maximum. Jones et al. (Microbiol. Rev. 50(4):484-524 (1986)) review the role of temperature in ABE fermentation. They report that the solvent yields of three different solvent producing strains remains fairly constant at 31% at 30° C. and 33° C., but decreases to 23 to 25% at 37° C. Similar results were reported for Clostridium acetobutylicum for which solvent yields decreased from 29% at 25° C. to 24% at 40° C. In the latter case, the decrease in solvent yield was attributed to a decrease in acetone production while the yield of 1-butanol was unaffected. However, Carnarius (U.S. Pat. No. 2,198,104) reports that an increase in the butanol ratio is obtained in the ABE process by decreasing the temperature of the fermentation from 30° C. to 24° C. after 16 hours. However, the effect of temperature on the production of 2-butanol by recombinant microbial hosts is not known in the art.
[0010] There is a need, therefore, for a cost-effective process for the production of 2-butanol by fermentation that provides higher yields than processes known in the art. The present invention addresses this need through the discovery of a method for producing 2-butanol by fermentation using a recombinant microbial host, which employs a decrease in temperature during fermentation, resulting in more robust tolerance of the production host to the 2-butanol product.
SUMMARY OF THE INVENTION
[0011] The invention provides a method for the production of 2-butanol by fermentation using a recombinant microbial host, which employs a decrease in temperature during fermentation that results in more robust tolerance of the production host to the 2-butanol product.
[0012] Accordingly, the invention provides a method for the production of 2-butanol comprising:
[0013] a) providing a recombinant microbial production host which produces 2-butanol;
[0014] b) seeding the production host of (a) into a fermentation
[0015] medium comprising a fermentable carbon substrate to create a fermentation culture;
[0016] c) growing the production host in the fermentation culture at a first temperature for a first period of time;
[0017] d) lowering the temperature of the fermentation culture to a second temperature; and
[0018] e) incubating the production host at the second temperature of step (d) for a second period of time;
[0019] whereby 2-butanol is produced.
BRIEF DESCRIPTION OF THE FIGURES AND SEQUENCE DESCRIPTIONS
[0020] The invention can be more fully understood from the following detailed description, FIGURE, and the accompanying sequence descriptions, which form a part of this application.
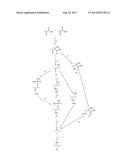
[0021] FIG. 1 shows four different pathways for biosynthesis of 2-butanone and 2-butanol.
[0022] The following sequences conform with 37 C.F.R. 1.821-1.825 ("Requirements for Patent Applications Containing Nucleotide Sequences and/or Amino Acid Sequence Disclosures--the Sequence Rules") and are consistent with World Intellectual Property Organization (WIPO) Standard ST.25 (1998) and the sequence listing requirements of the EPO and PCT (Rules 5.2 and 49.5(a-bis), and Section 208 and Annex C of the Administrative Instructions). The symbols and format used for nucleotide and amino acid sequence data comply with the rules set forth in 37 C.F.R. §1.822.
TABLE-US-00001 TABLE 1 Summary of Nucleic Acid and Protein SEQ ID Numbers SEQ ID Nucleic SEQ ID Description acid Protein budA, acetolactate decarboxylase from Klebsiella 1 2 pneumoniae ATCC 25955 alsD, acetolactate decarboxylase from Bacillus 80 81 subtilis budA, acetolactate decarboxylase from Klebsiella 82 83 terrigena budB, acetolactate synthase from Klebsiella 3 4 pneumoniae ATCC 25955 alsS, acetolactate synthase from Bacillus subtilis 76 77 budB, acetolactate synthase from Klebsiella 78 79 terrigena budC butanediol dehydrogenase from Klebsiella 5 6 pneumoniae IAM1063 butanediol dehydrogenase from Bacillus cereus 84 85 butanediol dehydrogenase from Bacillus cereus 86 87 butB, butanediol dehydrogenase from Lactococcus 88 89 lactis pddA, butanediol dehydratase alpha subunit from 7 8 Klebsiella oxytoca ATCC 8724 pddB, butanediol dehydratase beta subunit from 9 10 Klebsiella oxytoca ATCC 8724 pddC, butanediol dehydratase gamma subunit from 11 12 Klebsiella oxytoca ATCC 8724 pduC, B12 dependent diol dehydratase large 92 93 subunit from Salmonella typhimurium pduD, B12 dependent diol dehydratase medium 94 95 subunit from Salmonella typhimurium pduE, B12 dependent diol dehydratase small 96 97 subunit from Salmonella typhimurium pduC, B12 dependent diol dehydratase large 98 99 subunit from Lactobacillus collinoides pduD, B12 dependent diol dehydratase medium 100 101 subunit from Lactobacillus collinoides pduE, B12 dependent diol dehydratase small 102 103 subunit from Lactobacillus collinoides pddC, adenosylcobalamin-dependent diol 104 105 dehydratase alpha subunit from Klebsiella pneumoniae pddD, adenosylcobalamin-dependent diol 106 107 dehydratase beta subunit from Klebsiella pneumoniae pddD, adenosylcobalamin-dependent diol 108 109 dehydratase gamma subunit from Klebsiella pneumoniae ddrA, diol dehydratase reactivating factor large 110 111 subunit from Klebsiella oxytoca ddrB, diol dehydratase reactivating factor small 112 113 subunit from Klebsiella oxytoca pduG, diol dehydratase reactivating factor large 114 115 subunit from Salmonella typhimurium pduH, diol dehydratase reactivating factor small 116 117 subunit from Salmonella typhimurium pduG, diol dehydratase reactivating factor large 118 119 subunit from Lactobacillus collinoides pduH, diol dehydratase reactivating factor small 120 121 subunit from Lactobacillus collinoides sadH, butanol dehydrogenase from Rhodococcus 13 14 ruber 219 adhA, butanol dehydrogenase from Pyrococcus 90 91 furiosus chnA, cyclohexanol dehydrogenase from 71 72 Acinteobacter sp. yqhD, butanol dehydrogenase from Escherichia coli 74 75 amine: pyruvate transaminase from Vibrio fluvialis 144 122 (an acetoin aminase) codon opt. amino alcohol kinase from Erwinia carotovora 123 124 subsp. atroseptica amino alcohol O-phosphate lyase from Erwinia 125 126 carotovora subsp. atroseptica budC, acetoin reductase (butanediol 133 134 dehydrogenase) from Klebsiella terrigena (now Raoultella terrigena) glycerol dehydratase alpha subunit from Klebsiella 145 146 pneumoniae glycerol dehydratase beta subunit from Klebsiella 147 148 pneumoniae glycerol dehydratase gamma subunit from 149 150 Klebsiella pneumoniae glycerol dehydratase reactivase large subunit from 151 152 Klebsiella pneumoniae glycerol dehydratase reactivase small subunit from 153 154 Klebsiella pneumoniae
[0023] SEQ ID NOs:15-65 are the nucleotide sequences of oligonucleotide PCR, cloning, screening, and sequencing primers used in the Examples.
[0024] SEQ ID NO:66 is nucleotide sequence of the deleted region of the yqhD gene in E. coli strain MG1655 ΔyqhCD, described in Example 15.
[0025] SEQ ID NO:67 is the nucleotide sequence of a variant of the glucose isomerase promoter 1.6GI.
[0026] SEQ ID NO:68 is the nucleotide sequence of the 1.5GI promoter.
[0027] SEQ ID NO:69 is the nucleotide sequence of the diol dehydratase operon from Klebsiella oxytoca.
[0028] SEQ ID NO:70 is the nucleotide sequence of the diol dehydratase reactivating factor operon from Klebsiella oxytoca.
[0029] SEQ ID NO:73 is the nucleotide sequence of pDCQ2, which is described in Example 13.
[0030] SEQ ID NOs:127-132 are the nucleotide sequences of additional oligonucleotide PCR and cloning primers used in the Examples.
[0031] SEQ ID NO:155 is a codon optimized coding region for the amino alcohol kinase of Erwinia carotovora subsp. atroseptica.
[0032] SEQ ID NO:156 is a codon optimized coding region for the amino alcohol O-phosphate lyase of Erwinia carotovora subsp. atroseptica.
[0033] SEQ ID NOs:157-163 are the nucleotide sequences of additional oligonucleotide PCR and cloning primers used in the Examples.
[0034] SEQ ID NO:164 is the nucleotide sequence of an operon from Erwinia carotovora subsp. atroseptica.
TABLE-US-00002 TABLE 2 Additional glycerol and diol dehydratase large, medium and small subunits protein aDescription bsubunit SEQ ID Corresponding subunits from same organismc Glycerol dehydratase alpha subunit from Clostridium L 135 pasteurianum Glycerol dehydratase beta subunit from Clostridium M 136 pasteurianum Glycerol dehydratase gamma subunit from Clostridium S 137 pasteurianum Glycerol dehydratase alpha subunit from Escherichia L 138 blattae Glycerol dehydratase beta subunit from Escherichia M 139 blattae Glycerol dehydratase gamma subunit from Escherichia S 140 blattae Glycerol dehydratase alpha subunit from Citrobacter L 141 freundii Glycerol dehydratase beta subunit from Citrobacter M 142 freundii Glycerol dehydratase gamma subunit from Citrobacter S 143 freundii aDescription: from the Genbank annotation of the sequence and may not be correct including the glycerol or diol designation, or may not include subunit information. bSubunit: identified by sequence homology to the large, medium, or small subunit.of the Klebsiella oxytoca enzyme. cSubunts are listed together that are from the same organism and have annotations as the same enzyme, or have Genbank numbers close together indicating proximity in the genome.
DETAILED DESCRIPTION OF THE INVENTION
[0035] The present invention relates to a method for the production of 2-butanol using recombinant microorganisms that employs a decrease in temperature during fermentation, resulting in more robust tolerance of the production host to the 2-butanol product and therefore a higher titer of 2-butanol. The present invention meets a number of commercial and industrial needs. 2-Butanol is an important industrial commodity chemical with a variety of applications, where its potential as a fuel or fuel additive is particularly significant. Although only a four-carbon alcohol, butanol has an energy content similar to that of gasoline and can be blended with any fossil fuel. Butanol is favored as a fuel or fuel additive as it yields only CO2 and little or no SOx or NOx when burned in the standard internal combustion engine. Additionally butanol is less corrosive than ethanol, the most preferred fuel additive to date.
[0036] In addition to its utility as a biofuel or fuel additive, butanol has the potential of impacting hydrogen distribution problems in the emerging fuel cell industry. Fuel cells today are plagued by safety concerns associated with hydrogen transport and distribution. Butanol can be easily reformed for its hydrogen content and can be distributed through existing gas stations in the purity required for either fuel cells or combustion engines in vehicles.
[0037] Finally the present invention produces 2-butanol from plant derived carbon sources, avoiding the negative environmental impact associated with standard petrochemical processes for butanol production.
[0038] The following definitions and abbreviations are to be used for the interpretation of the claims and the specification.
[0039] As used herein, the terms "comprises," "comprising," "includes," "including," "has," "having," "contains" or "containing," or any other variation thereof, are intended to cover a non-exclusive inclusion. For example, a composition, a mixture, process, method, article, or apparatus that comprises a list of elements is not necessarily limited to only those elements but may include other elements not expressly listed or inherent to such composition, mixture, process, method, article, or apparatus. Further, unless expressly stated to the contrary, "or" refers to an inclusive or and not to an exclusive or. For example, a condition A or B is satisfied by any one of the following: A is true (or present) and B is false (or not present), A is false (or not present) and B is true (or present), and both A and B are true (or present).
[0040] Also, the indefinite articles "a" and "an" preceding an element or component of the invention are intended to be nonrestrictive regarding the number of instances (i.e. occurrences) of the element or component. Therefore "a" or "an" should be read to include one or at least one, and the singular word form of the element or component also includes the plural unless the number is obviously meant to be singular.
[0041] The term "invention" or "present invention" as used herein is a non-limiting term and is not intended to refer to any single embodiment of the particular invention but encompasses all possible embodiments as described in the specification and the claims.
[0042] As used herein, the term "about" modifying the quantity of an ingredient or reactant of the invention employed refers to variation in the numerical quantity that can occur, for example, through typical measuring and liquid handling procedures used for making concentrates or use solutions in the real world; through inadvertent error in these procedures; through differences in the manufacture, source, or purity of the ingredients employed to make the compositions or carry out the methods; and the like. The term "about" also encompasses amounts that differ due to different equilibrium conditions for a composition resulting from a particular initial mixture. Whether or not modified by the term "about", the claims include equivalents to the quantities. In one embodiment, the term "about" means within 10% of the reported numerical value, preferably within 5% of the reported numerical value.
[0043] The term "2-butanol biosynthetic pathway" refers to the enzyme pathways to produce 2-butanol from pyruvate.
[0044] The term "2-butanone biosynthetic pathway" refers to the enzyme pathways to produce 2-butanone from pyruvate.
[0045] The term "acetolactate synthase", also known as "acetohydroxy acid synthase", refers to a polypeptide (or polypeptides) having an enzyme activity that catalyzes the conversion of two molecules of pyruvic acid to one molecule of alpha-acetolactate. Acetolactate synthase, known as EC 2.2.1.6 [formerly 4.1.3.18] (Enzyme Nomenclature 1992, Academic Press, San Diego) may be dependent on the cofactor thiamin pyrophosphate for its activity. Suitable acetolactate synthase enzymes are available from a number of sources, for example, Bacillus subtilis [GenBank Nos: AAA22222 NCBI (National Center for Biotechnology Information) amino acid sequence (SEQ ID NO:77), L04470 NCBI nucleotide sequence (SEQ ID NO:76)], Klebsiella terrigena [GenBank Nos: AAA25055 (SEQ ID NO:79), L04507 (SEQ ID NO:78)], and Klebsiella pneumoniae [GenBank Nos: AAA25079 (SEQ ID NO:4), M73842 (SEQ ID NO:3)].
[0046] The term "acetolactate decarboxylase" refers to a polypeptide (or polypeptides) having an enzyme activity that catalyzes the conversion of alpha-acetolactate to acetoin. Acetolactate decarboxylases are known as EC 4.1.1.5 and are available, for example, from Bacillus subtilis [GenBank Nos: AAA22223 (SEQ ID NO:81), L04470 (SEQ ID NO:80)], Klebsiella terrigena [GenBank Nos: AAA25054 (SEQ ID NO:83), L04507 (SEQ ID NO:82)] and Klebsiella pneumoniae [GenBank Nos: AAU43774 (SEQ ID NO:2), AY722056 (SEQ ID NO:1)].
[0047] The term "acetoin aminase" refers to a polypeptide (or polypeptides) having an enzyme activity that catalyzes the conversion of acetoin to 3-amino-2-butanol. Acetoin aminase may utilize the cofactor pyridoxal 5'-phosphate or NADH (reduced nicotinamide adenine dinucleotide) or NADPH (reduced nicotinamide adenine dinucleotide phosphate). The resulting product may have (R) or (S) stereochemistry at the 3-position. The pyridoxal phosphate-dependent enzyme may use an amino acid such as alanine or glutamate as the amino donor. The NADH- and NADPH-dependent enzymes may use ammonia as a second substrate. A suitable example of an NADH-dependent acetoin aminase, also known as amino alcohol dehydrogenase, is described by Ito et al. (U.S. Pat. No. 6,432,688). An example of a pyridoxal-dependent acetoin aminase is the amine:pyruvate aminotransferase (also called amine:pyruvate transaminase) described by Shin and Kim (J. Org. Chem. 67:2848-2853 (2002)).
[0048] The term "butanol dehydrogenase" refers to a polypeptide (or polypeptides) having an enzyme activity that catalyzes the interconversion of 2-butanone and 2-butanol. Butanol dehydrogenases are a subset of a broad family of alcohol dehydrogenases. Butanol dehydrogenase may be NAD- or NADP-dependent. The NAD-dependent enzymes are known as EC 1.1.1.1 and are available, for example, from Rhodococcus ruber [GenBank Nos: CAD36475 (SEQ ID NO:14), AJ491307 (SEQ ID NO:13)]. The NADP-dependent enzymes are known as EC 1.1.1.2 and are available, for example, from Pyrococcus furiosus [GenBank Nos: AAC25556 (SEQ ID NO:91), AF013169 (SEQ ID NO:90)]. Additionally, a butanol dehydrogenase is available from Escherichia coli [GenBank Nos:NP--417484 (SEQ ID NO:75), NC--000913 (SEQ ID NO:74)] and a cyclohexanol dehydrogenase with activity towards 2-butanol is available from Acinetobacter sp. [GenBank Nos: AAG10026 (SEQ ID NO:72), AF282240 (SEQ ID NO:71)].
[0049] The term "acetoin kinase" refers to a polypeptide (or polypeptides) having an enzyme activity that catalyzes the conversion of acetoin to phosphoacetoin. Acetoin kinase may utilize ATP (adenosine triphosphate) or phosphoenolpyruvate as the phosphate donor in the reaction. Although there are no reports of enzymes catalyzing this reaction on acetoin, there are enzymes that catalyze the analogous reaction on the similar substrate dihydroxyacetone, for example, enzymes known as EC 2.7.1.29 (Garcia-Alles et al. (2004) Biochemistry 43:13037-13046).
[0050] The term "acetoin phosphate aminase" refers to a polypeptide (or polypeptides) having an enzyme activity that catalyzes the conversion of acetoin phosphate (also called phosphoacetoin) to 3-amino-2-butanol O-phosphate. Acetoin phosphate aminase may use the cofactor pyridoxal 5'-phosphate, NADH or NADPH. The resulting product may have (R) or (S) stereochemistry at the 3-position. The pyridoxal phosphate-dependent enzyme may use an amino acid such as alanine or glutamate. The NADH- and NADPH-dependent enzymes may use ammonia as a second substrate. Although there are no reports of enzymes catalyzing this reaction on acetoin phosphate, there is a pyridoxal phosphate-dependent enzyme that is proposed to carry out the analogous reaction on the similar substrate serinol phosphate (Yasuta et al. (2001) Appl. Environ. Microbiol. 67:4999-5009).
[0051] The term "aminobutanol phosphate phospho-lyase", also called "amino alcohol O-phosphate lyase", refers to a polypeptide (or polypeptides) having an enzyme activity that catalyzes the conversion of 3-amino-2-butanol O-phosphate to 2-butanone. Aminobutanol phosphate phospho-lyase may utilize the cofactor pyridoxal 5'-phosphate. There are reports of enzymes that catalyze the analogous reaction on the similar substrate 1-amino-2-propanol phosphate (Jones et al. (1973) Biochem J. 134:167-182). Disclosed in co-owned and co-pending US Patent Application Publication No. 20070259410A1 is an aminobutanol phosphate phospho-lyase (SEQ ID NO: 126) from the organism Erwinia carotovora, with demonstrated aminobutanol phosphate phospho-lyase activity.
[0052] The term "aminobutanol kinase" refers to a polypeptide (or polypeptides) having an enzyme activity that catalyzes the conversion of 3-amino-2-butanol to 3-amino-2-butanol O-phosphate. Aminobutanol kinase may utilize ATP as the phosphate donor. There are reports of enzymes that catalyze the analogous reaction on the similar substrates ethanolamine and 1-amino-2-propanol (Jones et al., supra). Disclosed in co-owned and co-pending US Patent Application Publication No. 20070259410A1 is an amino alcohol kinase of Erwinia carotovora subsp. atroseptica (SEQ ID NO:124).
[0053] The term "butanediol dehydrogenase" also known as "acetoin reductase" refers to a polypeptide (or polypeptides) having an enzyme activity that catalyzes the conversion of acetoin to 2,3-butanediol. Butanediol dehydrogenases are a subset of the broad family of alcohol dehydrogenases. Butanediol dehydrogenase enzymes may have specificity for production of (R)- or (S)-stereochemistry in the alcohol product. (S)-specific butanediol dehydrogenases are known as EC 1.1.1.76 and are available, for example, from Klebsiella pneumoniae (GenBank Nos: BBA13085 (SEQ ID NO:6), D86412 (SEQ ID NO:5)). (R)-specific butanediol dehydrogenases are known as EC 1.1.1.4 and are available, for example, from Bacillus cereus [GenBank Nos. NP--830481 (SEQ ID NO:85), NC--004722 (SEQ ID NO:84); AAP07682 (SEQ ID NO:87), AE017000 (SEQ ID NO:86)], and Lactococcus lactis [GenBank Nos. AAK04995 (SEQ ID NO:89), AE006323 (SEQ ID NO:88)].
[0054] The term "butanediol dehydratase", also known as "diol dehydratase" or "propanediol dehydratase" refers to a polypeptide (or polypeptides) having an enzyme activity that catalyzes the conversion of 2,3-butanediol to 2-butanone. Butanediol dehydratase may utilize the cofactor adenosyl cobalamin (vitamin B12). Adenosyl cobalamin-dependent enzymes are known as EC 4.2.1.28 and are available, for example, from Klebsiella oxytoca [GenBank Nos: BAA08099 (alpha subunit) (SEQ ID NO:8), D45071 (SEQ ID NO:7); BAA08100 (beta subunit) (SEQ ID NO:10), D45071 (SEQ ID NO:9); and BBA08101 (gamma subunit) (SEQ ID NO:12), D45071 (SEQ ID NO:11) (Note all three subunits are required for activity)], and Klebsiella pneumoniae [GenBank Nos: AAC98384 (alpha subunit) (SEQ ID NO:105), AF102064 (SEQ ID NO:104); GenBank Nos: AAC98385 (beta subunit) (SEQ ID NO:107), AF102064 (SEQ ID NO:106), GenBank Nos: AAC98386 (gamma subunit) SEQ ID NO:109), AF102064 (SEQ ID NO:108)]. Other suitable diol dehydratases include, but are not limited to, B12-dependent diol dehydratases available from Salmonella typhimurium [GenBank Nos: AAB84102 (large subunit) (SEQ ID NO:93), AF026270 (SEQ ID NO:92); GenBank Nos: AAB84103 (medium subunit) (SEQ ID NO:95), AF026270 (SEQ ID NO:94); GenBank Nos: AAB84104 (small subunit) (SEQ ID NO:97), AF026270 (SEQ ID NO:96)]; and Lactobacillus collinoides [GenBank Nos: CAC82541 (large subunit) (SEQ ID NO:99), AJ297723 (SEQ ID NO:98); GenBank Nos: CAC82542 (medium subunit) (SEQ ID NO:101); AJ297723 (SEQ ID NO:100); GenBank Nos: CAD01091 (small subunit) (SEQ ID NO:103), AJ297723 (SEQ ID NO:102)]; and enzymes from Lactobacillus brevis (particularly strains CNRZ 734 and CNRZ 735, Speranza et al., supra), and nucleotide sequences that encode the corresponding enzymes. Methods of diol dehydratase gene isolation are well known in the art (e.g., U.S. Pat. No. 5,686,276).
[0055] The term "glycerol dehydratase" refers to a polypeptide (or polypeptides) having an enzyme activity that catalyzes the conversion of glycerol to 3-hydroxypropionaldehyde. Adenosyl cobalamin-dependent glycerol dehydratases are known as EC 4.2.1.30. The glycerol dehydratases of EC 4.2.1.30 are similar to the diol dehydratases in sequence and in having three subunits. The glycerol dehydratases can also be used to convert 2,3-butanediol to 2-butanone. Some examples of glycerol dehydratases of EC 4.2.1.30 include those from Klebsiella pneumoniae (alpha subunit, SEQ ID NO:145, coding region and SEQ ID NO:146, protein; beta subunit, SEQ ID NO:147, coding region and SEQ ID NO:148, protein; and gamma subunit SEQ ID NO:149, coding region and SEQ ID NO:150, protein); from Clostridium pasteurianum [GenBank Nos: 3360389 (alpha subunit, SEQ ID NO:135), 3360390 (beta subunit, SEQ ID NO:136), and 3360391 (gamma subunit, SEQ ID NO:137)]; from Escherichia blattae [GenBank Nos: 60099613 (alpha subunit, SEQ ID NO:138), 57340191 (beta subunit, SEQ ID NO:139), and 57340192 (gamma subunit, SEQ ID NO:140)]; and from Citrobacter freundii [GenBank Nos: 1169287 (alpha subunit, SEQ ID NO:141), 1229154 (beta subunit, SEQ ID NO:142), and 1229155 (gamma subunit, SEQ ID NO:143)]. Note that all three subunits are required for activity. Additional glycerol dehydratases are listed in Table 2.
[0056] Diol and glycerol dehydratases may undergo suicide inactivation during catalysis. A reactivating factor protein, also referred to herein as "reactivase", can be used to reactivate the inactive enzymes (Mori et al., J. Biol. Chem. 272:32034 (1997)). Preferably, the reactivating factor is obtained from the same source as the diol or glycerol dehydratase used. For example, suitable diol dehydratase reactivating factors are available from Klebsiella oxytoca [GenBank Nos: AAC15871 (large subunit) (SEQ ID NO:111), AF017781 (SEQ ID NO:110); GenBank Nos: AAC15872 (small subunit) (SEQ ID NO:113), AF017781 (SEQ ID NO:112)]; Salmonella typhimurium [GenBank Nos: AAB84105 (large subunit) (SEQ ID NO:115), AF026270 (SEQ ID NO:114), GenBank Nos: AAD39008 (small subunit) (SEQ ID NO:117), AF026270 (SEQ ID NO:116)]; and Lactobacillus collinoides [GenBank Nos: CAD01092 (large subunit) (SEQ ID NO:119), AJ297723 (SEQ ID NO:118); GenBank Nos: CAD01093 (small subunit) (SEQ ID NO:121), AJ297723 (SEQ ID NO:120)]. Both the large and small subunits are required for activity. For example, suitable glycerol dehydratase reactivating factors are available from Klebsiella pneumoniae (large subunit, SEQ ID NO:151, coding region and SEQ ID NO:152, protein, and small subunit, SEQ ID NO:153, coding region and SEQ ID NO:154, protein).
[0057] The term "a facultative anaerobe" refers to a microorganism that can grow in both aerobic and anaerobic environments.
[0058] The term "carbon substrate" or "fermentable carbon substrate" refers to a carbon source capable of being metabolized by host organisms disclosed herein and particularly carbon sources selected from the group consisting of monosaccharides, oligosaccharides, polysaccharides, and one-carbon substrates or mixtures thereof.
[0059] The term "gene" refers to a nucleic acid fragment that is capable of being expressed as a specific protein, optionally including regulatory sequences preceding (5' non-coding sequences) and following (3' non-coding sequences) the coding sequence. "Native gene" refers to a gene as found in nature with its own regulatory sequences. "Chimeric gene" refers to any gene that is not a native gene, comprising regulatory and coding sequences that are not found together in nature. Accordingly, a chimeric gene may comprise regulatory sequences and coding sequences that are derived from different sources, or regulatory sequences and coding sequences derived from the same source, but arranged in a manner different than that found in nature. "Endogenous gene" refers to a native gene in its natural location in the genome of an organism. A "foreign" or "heterologous" gene refers to a gene not normally found in the host organism, but that is introduced into the host organism by gene transfer. Foreign genes can comprise native genes inserted into a non-native organism, or chimeric genes. A "transgene" is a gene that has been introduced into the genome by a transformation procedure.
[0060] As used herein, an "isolated nucleic acid fragment" or "isolated nucleic acid molecule" or "genetic construct" will be used interchangeably and will mean a polymer of RNA or DNA that is single- or double-stranded, optionally containing synthetic, non-natural or altered nucleotide bases. An isolated nucleic acid fragment in the form of a polymer of DNA may be comprised of one or more segments of cDNA, genomic DNA or synthetic DNA.
[0061] A nucleic acid fragment is "hybridizable" to another nucleic acid fragment, such as a cDNA, genomic DNA, or RNA molecule, when a single-stranded form of the nucleic acid fragment can anneal to the other nucleic acid fragment under the appropriate conditions of temperature and solution ionic strength. Hybridization and washing conditions are well known and exemplified in Sambrook, J., Fritsch, E. F. and Maniatis, T. Molecular Cloning: A Laboratory Manual, 2nd ed., Cold Spring Harbor Laboratory: Cold Spring Harbor, N.Y. (1989), particularly Chapter 11 and Table 11.1 therein (entirely incorporated herein by reference). The conditions of temperature and ionic strength determine the "stringency" of the hybridization. Stringency conditions can be adjusted to screen for moderately similar fragments (such as homologous sequences from distantly related organisms), to highly similar fragments (such as genes that duplicate functional enzymes from closely related organisms). Post-hybridization washes determine stringency conditions. One set of preferred conditions uses a series of washes starting with 6×SSC, 0.5% SDS at room temperature for 15 min, then repeated with 2×SSC, 0.5% SDS at 45° C. for 30 min, and then repeated twice with 0.2×SSC, 0.5% SDS at 50° C. for 30 min. A more preferred set of stringent conditions uses higher temperatures in which the washes are identical to those above except for the temperature of the final two 30 min washes in 0.2×SSC, 0.5% SDS was increased to 60° C. Another preferred set of highly stringent conditions uses two final washes in 0.1×SSC, 0.1% SDS at 65° C. An additional set of stringent conditions include hybridization at 0.1×SSC, 0.1% SDS, 65° C. and washes with 2×SSC, 0.1% SDS followed by 0.1×SSC, 0.1% SDS, for example.
[0062] Hybridization requires that the two nucleic acids contain complementary sequences, although depending on the stringency of the hybridization, mismatches between bases are possible. The appropriate stringency for hybridizing nucleic acids depends on the length of the nucleic acids and the degree of complementation, variables well known in the art. The greater the degree of similarity or homology between two nucleotide sequences, the greater the value of Tm for hybrids of nucleic acids having those sequences. The relative stability (corresponding to higher Tm) of nucleic acid hybridizations decreases in the following order: RNA:RNA, DNA:RNA, DNA:DNA. For hybrids of greater than 100 nucleotides in length, equations for calculating Tm have been derived (see Sambrook et al., supra, 9.50-9.51). For hybridizations with shorter nucleic acids, i.e., oligonucleotides, the position of mismatches becomes more important, and the length of the oligonucleotide determines its specificity (see Sambrook et al., supra, 11.7-11.8). In one embodiment the length for a hybridizable nucleic acid is at least about 10 nucleotides. Preferably a minimum length for a hybridizable nucleic acid is at least about 15 nucleotides; more preferably at least about 20 nucleotides; and most preferably the length is at least about 30 nucleotides. Furthermore, the skilled artisan will recognize that the temperature and wash solution salt concentration may be adjusted as necessary according to factors such as length of the probe.
[0063] A "substantial portion" of an amino acid or nucleotide sequence is that portion comprising enough of the amino acid sequence of a polypeptide or the nucleotide sequence of a gene to putatively identify that polypeptide or gene, either by manual evaluation of the sequence by one skilled in the art, or by computer-automated sequence comparison and identification using algorithms such as BLAST (Altschul, S. F., et al., J. Mol. Biol., 215:403-410 (1993)). In general, a sequence of ten or more contiguous amino acids or thirty or more nucleotides is necessary in order to putatively identify a polypeptide or nucleic acid sequence as homologous to a known protein or gene. Moreover, with respect to nucleotide sequences, gene specific oligonucleotide probes comprising 20-30 contiguous nucleotides may be used in sequence-dependent methods of gene identification (e.g., Southern hybridization) and isolation (e.g., in situ hybridization of bacterial colonies or bacteriophage plaques). In addition, short oligonucleotides of 12-15 bases may be used as amplification primers in PCR in order to obtain a particular nucleic acid fragment comprising the primers. Accordingly, a "substantial portion" of a nucleotide sequence comprises enough of the sequence to specifically identify and/or isolate a nucleic acid fragment comprising the sequence. The instant specification teaches the complete amino acid and nucleotide sequence encoding particular fungal proteins. The skilled artisan, having the benefit of the sequences as reported herein, may now use all or a substantial portion of the disclosed sequences for purposes known to those skilled in this art. Accordingly, the instant invention comprises the complete sequences as reported in the accompanying Sequence Listing, as well as substantial portions of those sequences as defined above.
[0064] The term "complementary" is used to describe the relationship between nucleotide bases that are capable of hybridizing to one another. For example, with respect to DNA, adenosine is complementary to thymine and cytosine is complementary to guanine.
[0065] The terms "homology" and "homologous" are used interchangeably herein. They refer to nucleic acid fragments wherein changes in one or more nucleotide bases do not affect the ability of the nucleic acid fragment to mediate gene expression or produce a certain phenotype. These terms also refer to modifications of the nucleic acid fragments of the instant invention such as deletion or insertion of one or more nucleotides that do not substantially alter the functional properties of the resulting nucleic acid fragment relative to the initial, unmodified fragment. It is therefore understood, as those skilled in the art will appreciate, that the invention encompasses more than the specific exemplary sequences.
[0066] Moreover, the skilled artisan recognizes that homologous nucleic acid sequences encompassed by this invention are also defined by their ability to hybridize, under moderately stringent conditions (e.g., 0.5×SSC, 0.1% SDS, 60° C.) with the sequences exemplified herein, or to any portion of the nucleotide sequences disclosed herein and which are functionally equivalent to any of the nucleic acid sequences disclosed herein.
[0067] "Codon degeneracy" refers to the nature in the genetic code permitting variation of the nucleotide sequence without effecting the amino acid sequence of an encoded polypeptide. The skilled artisan is well aware of the "codon-bias" exhibited by a specific host cell in usage of nucleotide codons to specify a given amino acid. Therefore, when synthesizing a gene for improved expression in a host cell, it is desirable to design the gene such that its frequency of codon usage approaches the frequency of preferred codon usage of the host cell.
[0068] The term "percent identity", as known in the art, is a relationship between two or more polypeptide sequences or two or more polynucleotide sequences, as determined by comparing the sequences. In the art, "identity" also means the degree of sequence relatedness between polypeptide or polynucleotide sequences, as the case may be, as determined by the match between strings of such sequences. "Identity" and "similarity" can be readily calculated by known methods, including but not limited to those described in: 1.) Computational Molecular Biology (Lesk, A. M., Ed.) Oxford University: NY (1988); 2.) Biocomputing: Informatics and Genome Projects (Smith, D. W., Ed.) Academic: NY (1993); 3.) Computer Analysis of Sequence Data, Part I (Griffin, A. M., and Griffin, H. G., Eds.) Humania: NJ (1994); 4.) Sequence Analysis in Molecular Biology (von Heinje, G., Ed.) Academic (1987); and 5.) Sequence Analysis Primer (Gribskov, M. and Devereux, J., Eds.) Stockton: NY (1991).
[0069] Preferred methods to determine identity are designed to give the best match between the sequences tested. Methods to determine identity and similarity are codified in publicly available computer programs. Sequence alignments and percent identity calculations may be performed using the MegAlign® program of the LASERGENE bioinformatics computing suite (DNASTAR Inc., Madison, Wis.). Multiple alignment of the sequences is performed using the "Clustal method of alignment" which encompasses several varieties of the algorithm including the "Clustal V method of alignment" corresponding to the alignment method labeled Clustal V (described by Higgins and Sharp, CABIOS. 5:151-153 (1989); Higgins, D. G. et al., Comput. Appl. Biosci., 8:189-191 (1992)) and found in the MegAlign® program of the LASERGENE bioinformatics computing suite (DNASTAR Inc.). For multiple alignments, the default values correspond to GAP PENALTY=10 and GAP LENGTH PENALTY=10. Default parameters for pairwise alignments and calculation of percent identity of protein sequences using the Clustal method are KTUPLE=1, GAP PENALTY=3, WINDOW=5 and DIAGONALS SAVED=5. For nucleic acids these parameters are KTUPLE=2, GAP PENALTY=5, WINDOW=4 and DIAGONALS SAVED=4. After alignment of the sequences using the Clustal V program, it is possible to obtain a "percent identity" by viewing the "sequence distances" table in the same program. Additionally the "Clustal W method of alignment" is available and corresponds to the alignment method labeled Clustal W (described by Higgins and Sharp, CABIOS. 5:151-153 (1989); Higgins, D. G. et al., Comput. Appl. Biosci. 8:189-191 (1992)) and found in the MegAlign® v6.1 program of the LASERGENE bioinformatics computing suite (DNASTAR Inc.). Default parameters for multiple alignment (GAP PENALTY=10, GAP LENGTH PENALTY=0.2, Delay Divergen Seqs(%)=30, DNA Transition Weight=0.5, Protein Weight Matrix=Gonnet Series, DNA Weight Matrix=IUB). After alignment of the sequences using the Clustal W program, it is possible to obtain a "percent identity" by viewing the "sequence distances" table in the same program.
[0070] It is well understood by one skilled in the art that many levels of sequence identity are useful in identifying polypeptides, from other species, wherein such polypeptides have the same or similar function or activity. Useful examples of percent identities include, but are not limited to: 24%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, or 95%, or any integer percentage from 24% to 100% may be useful in describing the present invention, such as 25%, 26%, 27%, 28%, 29%, 30%, 31%, 32%, 33%, 34%, 35%, 36%, 37%, 38%, 39%, 40%, 41%, 42%, 43%, 44%, 45%, 46%, 47%, 48%, 49%, 50%, 51%, 52%, 53%, 54%, 55%, 56%, 57%, 58%, 59%, 60%, 61%, 62%, 63%, 64%, 65%, 66%, 67%, 68%, 69%, 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98% or 99%. Suitable nucleic acid fragments not only have the above homologies but typically encode a polypeptide having at least 50 amino acids, preferably at least 100 amino acids, more preferably at least 150 amino acids, still more preferably at least 200 amino acids, and most preferably at least 250 amino acids.
[0071] The term "sequence analysis software" refers to any computer algorithm or software program that is useful for the analysis of nucleotide or amino acid sequences. "Sequence analysis software" may be commercially available or independently developed. Typical sequence analysis software will include, but is not limited to: 1.) the GCG suite of programs (Wisconsin Package Version 9.0, Genetics Computer Group (GCG), Madison, Wis.); 2.) BLASTP, BLASTN, BLASTX (Altschul et al., J. Mol. Biol., 215:403-410 (1990)); 3.) DNASTAR (DNASTAR, Inc. Madison, Wis.); 4.) Sequencher (Gene Codes Corporation, Ann Arbor, Mich.); and 5.) the FASTA program incorporating the Smith-Waterman algorithm (W. R. Pearson, Comput. Methods Genome Res., [Proc. Int. Symp.] (1994), Meeting Date 1992, 111-20. Editor(s): Suhai, Sandor. Plenum: New York, N.Y.). Within the context of this application it will be understood that where sequence analysis software is used for analysis, that the results of the analysis will be based on the "default values" of the program referenced, unless otherwise specified. As used herein "default values" will mean any set of values or parameters that originally load with the software when first initialized.
[0072] As used herein the term "coding sequence" or "CDS" refers to a DNA sequence that codes for a specific amino acid sequence. "Suitable regulatory sequences" refer to nucleotide sequences located upstream (5' non-coding sequences), within, or downstream (3' non-coding sequences) of a coding sequence, and which influence the transcription, RNA processing or stability, or translation of the associated coding sequence. Regulatory sequences may include promoters, translation leader sequences, introns, polyadenylation recognition sequences, RNA processing site, effector binding site and stem-loop structure.
[0073] The term "promoter" refers to a DNA sequence capable of controlling the expression of a coding sequence or functional RNA. In general, a coding sequence is located 3' to a promoter sequence. Promoters may be derived in their entirety from a native gene, or be composed of different elements derived from different promoters found in nature, or even comprise synthetic DNA segments. It is understood by those skilled in the art that different promoters may direct the expression of a gene in different tissues or cell types, or at different stages of development, or in response to different environmental or physiological conditions. Promoters which cause a gene to be expressed in most cell types at most times are commonly referred to as "constitutive promoters". It is further recognized that since in most cases the exact boundaries of regulatory sequences have not been completely defined, DNA fragments of different lengths may have identical promoter activity.
[0074] The term "operably linked" refers to the association of nucleic acid sequences on a single nucleic acid fragment so that the function of one is affected by the other. For example, a promoter is operably linked with a coding sequence when it is capable of effecting the expression of that coding sequence (i.e., that the coding sequence is under the transcriptional control of the promoter). Coding sequences can be operably linked to regulatory sequences in sense or antisense orientation.
[0075] The term "expression", as used herein, refers to the transcription and stable accumulation of sense (mRNA) or antisense RNA derived from the nucleic acid fragment disclosed herein. Expression may also refer to translation of mRNA into a polypeptide.
[0076] As used herein the term "transformation" refers to the transfer of a nucleic acid fragment into a host organism, resulting in genetically stable inheritance. Host organisms containing the transformed nucleic acid fragments are referred to as "transgenic" or "recombinant" or "transformed" organisms.
[0077] The terms "plasmid" and "vector" refer to an extra chromosomal element often carrying genes which are not part of the central metabolism of the cell, and usually in the form of circular double-stranded DNA molecules. Such elements may be autonomously replicating sequences, genome integrating sequences, phage or nucleotide sequences, linear or circular, of a single- or double-stranded DNA or RNA, derived from any source, in which a number of nucleotide sequences have been joined or recombined into a unique construction which is capable of introducing a promoter fragment and DNA sequence for a selected gene product along with appropriate 3' untranslated sequence into a cell.
[0078] "Transformation vector" refers to a specific vector containing a foreign gene and having elements in addition to the foreign gene that facilitates transformation of a particular host cell.
[0079] As used herein the term "codon degeneracy" refers to the nature in the genetic code permitting variation of the nucleotide sequence without affecting the amino acid sequence of an encoded polypeptide. The skilled artisan is well aware of the "codon-bias" exhibited by a specific host cell in usage of nucleotide codons to specify a given amino acid. Therefore, when synthesizing a gene for improved expression in a host cell, it is desirable to design the gene such that its frequency of codon usage approaches the frequency of preferred codon usage of the host cell.
[0080] The term "codon-optimized" as it refers to genes or coding regions of nucleic acid molecules for transformation of various hosts, refers to the alteration of codons in the gene or coding regions of the nucleic acid molecules to reflect the typical codon usage of the host organism without altering the polypeptide encoded by the DNA.
[0081] The term "fermentation product medium" refers to a medium in which fermentation has occurred such that product is present in the medium.
[0082] Standard recombinant DNA and molecular cloning techniques used here are well known in the art and are described by Sambrook, J., Fritsch, E. F. and Maniatis, T., Molecular Cloning: A Laboratory Manual, Second Edition, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y. (1989) (hereinafter "Maniatis"); and by Silhavy, T. J., Bennan, M. L. and Enquist, L. W., Experiments with Gene Fusions, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y. (1984); and by Ausubel, F. M. et al., Current Protocols in Molecular Biology, published by Greene Publishing Assoc. and Wiley-Interscience (1987).
2-Butanol Biosynthetic Pathways
[0083] Carbohydrate utilizing microorganisms employ the Embden-Meyerhof-Parnas (EMP) pathway, the Entner-Doudoroff pathway and the pentose phosphate cycle as the central, metabolic routes to provide energy and cellular precursors for growth and maintenance. These pathways have in common the intermediate glyceraldehyde 3-phosphate, and, ultimately, pyruvate is formed directly or in combination with the EMP pathway. The combined reactions of sugar conversion to pyruvate produce energy (e.g. adenosine 5'-triphosphate, ATP) and reducing equivalents (e.g. reduced nicotinamide adenine dinucleotide, NADH, and reduced nicotinamide adenine dinucleotide phosphate, NADPH). NADH and NADPH must be recycled to their oxidized forms (NAD.sup.+ and NADP.sup.+, respectively). In the presence of inorganic electron acceptors (e.g. O2, NO3.sup.- and SO42-), the reducing equivalents may be used to augment the energy pool; alternatively, a reduced carbon by-product may be formed.
[0084] As described in co-owned and co-pending US Patent Application Publication Nos. 20070259410A1 and 20070292927A1, 2-butanol can be produced from carbohydrate sources in recombinant microorganisms comprising a complete 2-butanol biosynthetic pathway. Four biosynthetic pathways including all steps starting with pyruvate for production of 2-butanol are shown in FIG. 1. The letters and roman numerals cited below correspond to the letters and roman numerals in FIG. 1, which are used to depict the conversion steps and products, respectively. As described below, 2-butanone is an intermediate in all of these 2-butanol biosynthetic pathways.
[0085] All of the pathways begin with the initial reaction of two pyruvate molecules to yield alpha-acetolactate (I), shown as the substrate to product conversion (a) in FIG. 1. From alpha-acetolactate, there are 4 possible pathways to 2-butanone (V), referred to herein as 2-butanone biosynthetic pathways:
[0086] Pathway 1) I-->II-->III-->IV-->V (substrate to product conversions b,c,d,e)
[0087] 2) I-->II-->VII-->IV-->V (substrate to product conversions b,g,h,e)
[0088] 3) I-->II-->VIII-->V (substrate to product conversions b,i,j)
[0089] 4) I-->IX-->X-->V (substrate to product conversions k,l,m)
[0090] The 2-butanol biosynthetic pathways conclude with the conversion of 2-butanone (V) to 2-butanol (VI). A detailed discussion of the substrate to product conversions in each pathway is given below.
Pathway 1:
[0091] (a) pyruvate to alpha-acetolactate:
[0092] The initial step in pathway 1 is the conversion of two molecules of pyruvate to one molecule of alpha-acetolactate (compound I in FIG. 1) and one molecule of carbon dioxide catalyzed by a thiamin pyrophosphate-dependent enzyme. Enzymes catalyzing this substrate to product conversion (generally called either acetolactate synthase or acetohydroxy acid synthase; EC 2.2.1.6 [switched from 4.1.3.18 in 2002]) are well-known, and they participate in the biosynthetic pathway for the proteinogenic amino acids leucine and valine, as well as in the pathway for fermentative production of 2,3-butanediol and acetoin of a number of organisms.
[0093] The skilled person will appreciate that polypeptides having acetolactate synthase activity isolated from a variety of sources will be useful in pathway 1 independent of sequence homology. Some examples of suitable acetolactate synthase enzymes are available from a number of sources, for example, Bacillus subtilis [GenBank Nos: AAA22222 NCBI (National Center for Biotechnology Information) amino acid sequence (SEQ ID NO:77), L04470 NCBI nucleotide sequence (SEQ ID NO:76)], Klebsiella terrigena [GenBank Nos: AAA25055 (SEQ ID NO:79), L04507 (SEQ ID NO:78)], and Klebsiella pneumoniae [GenBank Nos: AAA25079 (SEQ ID NO:4), M73842 (SEQ ID NO:3)]. Preferred acetolactate synthase enzymes are those that have at least 80%-85% identity to SEQ ID NO's 4, 77, and 79, where at least 85%-90% identity is more preferred and where at least 95% identity based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix, is most preferred.
[0094] (b) alpha-acetolactate to acetoin:
[0095] Alpha-acetolactate (I) is converted to acetoin (II) by the action of an enzyme such as acetolactate decarboxylase (EC 4.1.1.5). Like acetolactate synthase, this enzyme is thiamin pyrophosphate-dependent and is also involved in the production of 2,3-butanediol and acetoin by a number of organisms. The enzymes from different sources vary quite widely in size (25-50 kilodaltons), oligomerization (dimer-hexamer), localization (intracellular of extracellular), and allosteric regulation (for example, activation by branched-chain amino acids). An intracellular location is preferable to extracellular, but other variations are generally acceptable.
[0096] The skilled person will appreciate that polypeptides having acetolactate decarboxylase activity isolated from a variety of sources will be useful in pathway 1 independent of sequence homology. Some example of suitable acetolactate decarboxylase enzymes are available from a number of sources, for example, Bacillus subtilis [GenBank Nos: AAA22223 (SEQ ID NO:81), L04470 (SEQ ID NO:80)], Klebsiella terrigena [GenBank Nos: AAA25054 (SEQ ID NO:83), L04507 (SEQ ID NO:82)] and Klebsiella pneumoniae [GenBank Nos: AAU43774 (SEQ ID NO:2), AY722056 (SEQ ID NO:1)].
[0097] Preferred acetolactate decarboxylase enzymes are those that have at least 80%-85% identity to SEQ ID NO's 2, 81 and 83, where at least 85%-90% identity is more preferred and where at least 95% identity based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix, is most preferred.
[0098] (c) acetoin to 3-amino-2-butanol:
[0099] There are two known types of biochemical reactions that could effect the substrate to product conversion of acetoin (II) to 3-amino-2-butanol (III), specifically, pyridoxal phosphate-dependent transamination utilizing an accessory amino donor and direct reductive amination with ammonia. In the latter case, the reducing equivalents are supplied in the form of a reduced nicotinamide cofactor (either NADH or NADPH). An example of an NADH-dependent enzyme catalyzing this reaction with acetoin as a substrate is reported by Ito et al. (U.S. Pat. No. 6,432,688). Any stereospecificity of this enzyme has not been assessed. An example of a pyridoxal phosphate-dependent transaminase that catalyzes the conversion of acetoin to 3-amino-2-butanol has been reported by Shin and Kim (supra). This enzyme was shown in co-owned and co-pending US Patent Application Publication No. 20070259410A1 to convert both the (R) isomer of acetoin to the (2R,3S) isomer of 3-amino-2-butanol and the (S) isomer of acetoin to the (2S,3S) isomer of 3-amino-2-butanol. Either type of enzyme (i.e., transaminase or reductive aminase) is considered to be an acetoin aminase and may be utilized in the production of 2-butanol. Other enzymes in this group may have different stereospecificities.
[0100] The skilled person will appreciate that polypeptides having acetoin aminase activity isolated from a variety of sources will be useful in the present invention independent of sequence homology. One example of a protein having this activity is described in co-owned and co-pending US Patent Application Publication No. 20070259410A1 (SEQ ID NO:122). Accordingly preferred acetoin aminase enzymes are those that have at least 80%-85% identity to SEQ ID NO:122, where at least 85%-90% identity is more preferred and where at least 95% identity based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix, is most preferred.
[0101] (d) 3-amino-2-butanol to 3-amino-2-butanol O-phosphate:
[0102] There are no enzymes known in the art that catalyze the substrate to product conversion of 3-amino-2-butanol (III) to 3-amino-2-butanol phosphate (IV). However, a few Pseudomonas and Erwinia species have been shown to express an ATP-dependent ethanolamine kinase (EC 2.7.1.82) which allows them to utilize ethanolamine or 1-amino-2-propanol as a nitrogen source (Jones et al. (1973) Biochem. J. 134:167-182). It is likely that this enzyme also has activity towards 3-amino-2-butanol or could be engineered to do so, thereby providing an aminobutanol kinase. Disclosed in in co-owned and co-pending US Patent Application Publication No. 20070259410A1 is a gene of Erwinia carotovora subsp. atroseptica (SEQ ID NO:123) that encodes a protein (SEQ ID NO:24) identified as an amino alcohol kinase. This enzyme may be used to convert 3-amino-2-butanol to 3-amino-2-butanol O-phosphate.
[0103] The skilled person will appreciate that polypeptides having aminobutanol kinase activity isolated from a variety of sources will be useful in the present invention independent of sequence homology. One example of this activity is described in co-owned and co-pending US Patent Application Publication No. 20070259410A1 (SEQ ID NO:124). Accordingly preferred aminobutanol kinase enzymes are those that have at least 80%-85% identity to SEQ ID NO:124, where at least 85%-90% identity is more preferred and where at least 95% identity based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix, is most preferred.
[0104] (e) 3-amino-2-butanol phosphate to 2-butanone:
[0105] Although there are no enzymes reported to catalyze the substrate to product conversion of 3-amino-2-butanol phosphate (IV) to 2-butanone (V), the substrate is very similar to those utilized by the pyridoxal phosphate-dependent phosphoethanolamine phospho-lyase enzyme, which has been found in a small number of Pseudomonas and Erwinia species. These enzymes have activity towards phosphoethanolamine and both enantiomers of 2-phospho-1-aminopropane (Jones et al. (1973) Biochem. J. 134:167-182), and may also have activity towards 3-amino-2-butanol O-phosphate. Applicants have identified a gene of Erwinia carotovora subsp. atroseptica (SEQ ID NO:125) that encodes a protein (SEQ ID NO:126) with homology to class III aminotransferases was identified. It was shown to have activity on both aminopropanol phosphate and aminobutanol phosphate substrates. The enzyme was able to catalyze the conversion of a mixture of (R)-3-amino-(S)-2-butanol and (S)-3-amino-(R)-2-butanol O-phosphate, and a mixture of (R)-3-amino-(R)-2-butanol and (S)-3-amino-(S)-2-butanol O-phosphate to 2-butanone. The enzyme was also able to catalyze the conversion of both (R) and (S)-2-amino-1-propanol phosphate to propanone, with a preference for (S)-2-amino-1-propanol phosphate. The highest activity was with the proposed natural substrate DL-1-amino-2-propanol phosphate, which was converted to propionaldehyde.
[0106] The skilled person will appreciate that polypeptides having aminobutanol phosphate phospho-lyase activity isolated from a variety of sources will be useful in the present invention independent of sequence homology. One example of a suitable aminobutanol phosphate phospho-lyase enzyme is described in co-owned and co-pending US Patent Application Publication No. 20070259410A1 (SEQ ID NO: 126). Accordingly preferred aminobutanol phosphate phospho-lyase enzymes are those that have at least 80%-85% identity to SEQ ID NO's 126, where at least 85%-90% identity is more preferred and where at least 95% identity based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix, is most preferred.
[0107] (f) 2-butanone to 2-butanol:
[0108] The final step in all pathways to produce 2-butanol from pyruvic acid is the reduction of 2-butanone (V) to 2-butanol (VI). This substrate to product conversion is catalyzed by some members of the broad class of alcohol dehydrogenases (types utilizing either NADH or NADPH as a source of hydride, depending on the enzyme) that may be called butanol dehydrogenases. Enzymes of each type that catalyze the reduction of 2-butanone are well known, as described above in the definition for butanol dehydrogenase.
[0109] The skilled person will appreciate that polypeptides having butanol dehydrogenase activity isolated from a variety of sources will be useful in the present invention independent of sequence homology. Some example of suitable butanol dehydrogenase enzymes are available from a number of sources, for example, Rhodococcus ruber [GenBank Nos: CAD36475 (SEQ ID NO:14), AJ491307 (SEQ ID NO:13)]. The NADP-dependent enzymes are known as EC 1.1.1.2 and are available, for example, from Pyrococcus furiosus [GenBank Nos: AAC25556 (SEQ ID NO:91), AF013169 (SEQ ID NO:90)]. Additionally, a butanol dehydrogenase is available from Escherichia coli [GenBank Nos:NP--417484 (SEQ ID NO:75), NC--000913 (SEQ ID NO:74)] and a cyclohexanol dehydrogenase is available from Acinetobacter sp. [GenBank Nos: AAG10026 (SEQ ID NO:72), AF282240 (SEQ ID NO:71)]. Preferred butanol dehydrogenase enzymes are those that have at least 80%-85% identity to SEQ ID NO's 14, 91, 75, and 72, where at least 85%-90% identity is more preferred and where at least 95% identity based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix, is most preferred.
Pathway 2:
[0110] (a) pyruvate to alpha-acetolactate:
[0111] This substrate to product conversion is the same as described above for Pathway 1.
[0112] (b) alpha-acetolactate to acetoin:
[0113] This substrate to product conversion is the same as described above for Pathway 1.
[0114] (g) acetoin to phosphoacetoin:
[0115] Although enzymes that catalyze the substrate to product conversion of acetoin (II) to phosphoacetoin (VII) have not been described, the structure of the substrate acetoin is very similar to that of dihydroxyacetone, and therefore acetoin may be an acceptable substrate for dihydroxyacetone kinase (EC 2.7.1.29), an enzyme which catalyzes phosphorylation of dihydroxyacetone. Protein engineering techniques for the alteration of substrate specificity of enzymes are well known (Antikainen and Martin (2005) Bioorg. Med. Chem. 13:2701-2716) and may be used to generate an enzyme with the required specificity. In this conversion, the phosphate moiety may be supplied by any high energy biological phosphate donor, with the common substrates being phosphoenolpyruvate (as in the E. coli dihydroxyacetone kinase) and ATP (as in the Citrobacter freundii dihydroxyacetone kinase) (Garcia-Alles et al. (2004) Biochemistry 43:13037-13045).
[0116] (h) phosphoacetoin to 3-amino-2-butanol O-phosphate:
[0117] Although enzymes that catalyze the substrate to product conversion of phosphoacetoin (VII) to 3-amino-2-butanol O-phosphate (IV) have not been described, the structure of the substrate is very similar to that of dihydroxyacetone phosphate, a substrate for the proposed serinol phosphate aminotransferase encoded by the 5' portion of the rtxA gene in some species of Bradyrhizobium (Yasuta et al., supra). Thus a serinol phosphate aminotransferase may be functional in this step.
[0118] (e) 3-amino-2-butanol O-phosphate to 2-butanone:
[0119] This substrate to product conversion is the same as described above for Pathway 1.
[0120] (f) 2-butanone to 2-butanol:
[0121] This substrate to product conversion is the same as described above for Pathway 1.
Pathway 3:
[0122] (a) pyruvate to alpha-acetolactate:
[0123] This substrate to product conversion is the same as described above for Pathway 1.
[0124] (b) alpha-acetolactate to acetoin:
[0125] This substrate to product conversion is the same as described above for Pathway 1.
[0126] (i) acetoin to 2,3-butanediol:
[0127] The substrate to product conversion of acetoin (II) to 2,3-butanediol (VIII) may be catalyzed by a butanediol dehydrogenase that may either utilize NADH or NADPH as the source of reducing equivalents when carrying out reductions. Enzymes with activity towards acetoin participate in the pathway for production of 2,3-butanediol in organisms that produce that compound. The reported enzymes (e.g., BudC from Klebsiella pneumoniae (Ui et al. (2004) Letters in Applied Microbiology 39:533-537) generally utilize NADH. Either cofactor is acceptable for use in the production of 2-butanol by this pathway.
[0128] (j) 2,3-butanediol to 2-butanone:
[0129] The substrate to product conversion of 2,3-butanediol (VIII) to 2-butanone (V) may be catalyzed by diol dehydratase enzymes (EC 4.2.1.28) and glycerol dehydratase enzymes (EC 4.2.1.30). The best characterized diol dehydratase is the coenzyme B12-dependent Klebsiella oxytoca enzyme, but similar enzymes are found in many enteric bacteria. The K. oxytoca enzyme has been shown to accept meso-2,3-butanediol as a substrate (Bachovchin et al. (1977) Biochemistry 16:1082-1092), producing the desired product 2-butanone. Applicants describe a Klebsiella pneumoniae glycerol dehydratase able to convert meso-2,3-butanediol to 2-butanone. The three subunit of the Klebsiella pneumoniae glycerol dehydratase (alpha: SEQ ID NO:145 (coding region) and 146 (protein); beta: SEQ ID NO: 147 (coding region) and 148 (protein); and gamma: SEQ ID NO: 149 (coding region) and 150 (protein)) were expressed in conjunction with the two subunits of the Klebsiella pneumoniae glycerol dehydratase reactivase (large subunit, SEQ ID NO: 151 (coding region) and 152 (protein); and small subunit, SEQ ID NO: 153 (coding region) and 154 (protein)) to provide activity.
[0130] There are also reports in the literature of a B12-independent diol dehydratase from Clostridium glycolicum (Hartmanis et al. (1986) Arch. Biochem. Biophys. 245:144-152). This enzyme has activity towards 2,3-butanediol, although this activity is less than 1% of the activity towards ethanediol, but the enzyme may be engineered to improve that activity. A better-characterized B12-independent dehydratase is the glycerol dehydratase from Clostridium butyricum (O'Brien et al. (2004) Biochemistry 43:4635-4645), which has high activity towards 1,2-propanediol as well as glycerol. This enzyme uses S-adenosylmethionine as a source of adenosyl radical. There are no reports of activity towards 2,3-butanediol, but such activity, if not already present, may possibly be engineered.
[0131] The skilled person will appreciate that polypeptides having butanediol dehydrogenase activity isolated from a variety of sources will be useful in the present invention independent of sequence homology. As noted above a variety of diol and glycerol dehydratases have been described in the literature and will be suitable for use in the present invention. Accordingly, in one aspect of the invention preferred diol and glycerol dehydratase enzymes are those that have at least 80%-85% identity to enzymes having the large, medium and small subunits, respectively of the sequences listed below:
[0132] a) SEQ ID NO:8, SEQ ID NO:10, and SEQ ID NO:12;
[0133] b) SEQ ID NO:93, SEQ ID NO:95, and SEQ ID NO:97;
[0134] c) SEQ ID NO:99, SEQ ID NO:101, and SEQ ID NO:103;
[0135] d) SEQ ID NO:105, SEQ ID NO:107, and SEQ ID NO:109;
[0136] e) SEQ ID NO:135, SEQ ID NO:136, and SEQ ID NO:137;
[0137] f) SEQ ID NO:138, SEQ ID NO:139, and SEQ ID NO:140;
[0138] g) SEQ ID NO:146, SEQ ID NO:148, and SEQ ID NO:150;
[0139] h) SEQ ID NO:141, SEQ ID NO:142, and SEQ ID NO:143; and
[0140] i) SEQ ID NO:164, SEQ ID NO:165, and SEQ ID NO:166.
[0141] where at least 85%-90% identity is more preferred and where at least 95% identity based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix, is most preferred. Similarly preferred diol and glycerol dehydratase enzymes are those that have at least 80%-85% identity to enzymes having the large, medium and small subunits, respectively of the sequences listed below:
Large subunit: SEQ ID NOs: 8, 99, 105, 135, 138, 141, 146, and 164; Medium subunit: SEQ ID NOs: 10, 101, 107, 136, 139, 142, 148, and 165; Small subunit: SEQ ID NOs: 12, 103, 109, 137, 140, 143, 150, and 166; where at least 85%-90% identity is more preferred and where at least 95% identity based on the Clustal W method of alignment using the default parameters of GAP PENALTY=10, GAP LENGTH PENALTY=0.1, and Gonnet 250 series of protein weight matrix, is most preferred.
[0142] (f) 2-butanone to 2-butanol:
[0143] This substrate to product conversion is the same as described above for Pathway 1.
Pathway 4:
[0144] (a) pyruvate to alpha-acetolactate:
[0145] This substrate to product conversion is the same as described above for Pathway 1.
[0146] (k) alpha-acetolactate to 2,3-dihydroxy-2-methylbutanoic acid:
[0147] The substrate to product conversion of acetolactate (I) to 2,3-dihydroxy-2-methylbutanoic acid (IX) is not known in the art. However, the product of this conversion has been reported as a component of fermentation broths (Ziadi et al. (1973) Comptes Rendus des Seances de l'Academie des Sciences, Serie D: Sciences Naturelles 276:965-8), but the mechanism of formation is unknown. The likely mechanism of formation is reduction of acetolactate with NADH or NADPH as the electron donor. To utilize this pathway for production of 2-butanol, an enzyme catalyzing this reaction needs to be identified or engineered. However, the precedent for enzymatic reduction of ketones to alcohols is well established.
[0148] (l) 2,3-dihydroxy-2-methylbutanoic acid to 2-hydroxy-2-methyl-3-phosphobutanoic acid:
[0149] There are no enzymes known that catalyze the substrate to product conversion of 2,3-dihydroxy-2-methylbutanoic acid (IX) to 2-hydroxy-2-methyl-3-phosphobutanoic acid (X). However, there are a large number of kinases in Nature that possess varying specificity. It is therefore likely that an enzyme could be isolated or engineered with this activity.
[0150] (m) 2-hydroxy-2-methyl-3-phosphobutanoic acid to 2-butanone:
[0151] There are no known enzymes that catalyze the substrate to product conversion of 2-hydroxy-2-methyl-3-phosphobutanoic acid (X) to 2-butanone (V).
[0152] The combination of this reaction with the previous one is very similar to the multi-step reaction catalyzed by mevalonate-5-pyrophosphate (M5PP) decarboxylase, which consists of initial phosphorylation of M5PP to 3-phosphomevalonate-5-PP, followed by decarboxylation-dependent elimination of phosphate (Alvear et al. (1982) Biochemistry 21:4646-4650).
[0153] (f) 2-butanone to 2-butanol:
[0154] This substrate to product conversion is the same as described above for Pathway 1.
[0155] Thus, in providing multiple recombinant pathways from pyruvate to 2-butanol, there exists a number of choices to fulfill the individual conversion steps, and the person of skill in the art will be able to utilize publicly available sequences and sequences disclosed herein to construct the relevant pathways. A listing of a representative number of genes known in the art and useful in the construction of 2-butanol biosynthetic pathways is given above in Tables 1 and 2.
Microbial Hosts for 2-Butanol and 2-Butanone Production
[0156] Microbial hosts for 2-butanol or 2-butanone production may be selected from bacteria, cyanobacteria, filamentous fungi and yeasts. The microbial host used for 2-butanol or 2-butanone production should be tolerant to the product produced, so that the yield is not limited by toxicity of the product to the host. The selection of a microbial host for 2-butanol production is described in detail below.
[0157] Microbes that are metabolically active at high titer levels of 2-butanol are not well known in the art. Although butanol-tolerant mutants have been isolated from solventogenic Clostridia, little information is available concerning the butanol tolerance of other potentially useful bacterial strains. Most of the studies on the comparison of alcohol tolerance in bacteria suggest that butanol is more toxic than ethanol (de Cavalho et al., Microsc. Res. Tech. 64:215-22 (2004) and Kabelitz et al., FEMS Microbiol. Lett. 220:223-227 (2003)). Tomas et al. (J. Bacteriol. 186:2006-2018 (2004)) report that the yield of 1-butanol during fermentation in Clostridium acetobutylicum may be limited by butanol toxicity. The primary effect of 1-butanol on Clostridium acetobutylicum is disruption of membrane functions (Hermann et al., Appl. Environ. Microbiol. 50:1238-1243 (1985)).
[0158] The microbial hosts selected for the production of 2-butanol should be tolerant to 2-butanol and should be able to convert carbohydrates to 2-butanol using the introduced biosynthetic pathway. The criteria for selection of suitable microbial hosts include the following: intrinsic tolerance to 2-butanol, high rate of carbohydrate utilization, availability of genetic tools for gene manipulation, and the ability to generate stable chromosomal alterations.
[0159] Suitable host strains with a tolerance for 2-butanol may be identified by screening based on the intrinsic tolerance of the strain. The intrinsic tolerance of microbes to 2-butanol may be measured by determining the concentration of 2-butanol that is responsible for 50% inhibition of the growth rate (IC50) when grown in a minimal medium. The IC50 values may be determined using methods known in the art. For example, the microbes of interest may be grown in the presence of various amounts of 2-butanol and the growth rate monitored by measuring the optical density at 600 nanometers. The doubling time may be calculated from the logarithmic part of the growth curve and used as a measure of the growth rate. The concentration of 2-butanol that produces 50% inhibition of growth may be determined from a graph of the percent inhibition of growth versus the 2-butanol concentration. Preferably, the host strain should have an IC50 for 2-butanol of greater than about 0.5%. More suitable is a host strain with an IC50 for 2-butanol that is greater than about 1.5%. Particularly suitable is a host strain with an IC50 for 2-butanol that is greater than about 2.5%.
[0160] The microbial host for 2-butanol production should also utilize glucose and/or other carbohydrates at a high rate. Most microbes are capable of utilizing carbohydrates. However, certain environmental microbes cannot efficiently use carbohydrates, and therefore would not be suitable hosts.
[0161] The ability to genetically modify the host is essential for the production of any recombinant microorganism. Modes of gene transfer technology that may be used include by electroporation, conjugation, transduction or natural transformation. A broad range of host conjugative plasmids and drug resistance markers are available. The cloning vectors used with an organism are tailored to the host organism based on the nature of antibiotic resistance markers that can function in that host.
[0162] The microbial host also may be manipulated in order to inactivate competing pathways for carbon flow by inactivating various genes. This requires the availability of either transposons or chromosomal integration vectors to direct inactivation. Additionally, production hosts that are amenable to chemical mutagenesis may undergo improvements in intrinsic 2-butanol tolerance through chemical mutagenesis and mutant screening.
[0163] Based on the criteria described above, suitable microbial hosts for the production of 2-butanol include, but are not limited to, members of the genera Clostridium, Zymomonas, Escherichia, Salmonella, Rhodococcus, Pseudomonas, Bacillus, Lactobacillus, Enterococcus, Pediococcus, Alcaligenes, Klebsiella, Paenibacillus, Arthrobacter, Corynebacterium, Brevibacterium, Pichia, Candida, Hansenula and Saccharomyces. Preferred hosts include: Escherichia coli, Alcaligenes eutrophus, Bacillus licheniformis, Paenibacillus macerans, Rhodococcus erythropolis, Pseudomonas putida, Lactobacillus plantarum, Enterococcus faecium, Enterococcus gallinarium, Enterococcus faecalis, Pediococcus pentosaceus, Pediococcus acidilactici, Bacillus subtilis and Saccharomyces cerevisiae.
Construction of Production Host
[0164] Recombinant organisms containing the necessary genes that encode the enzymatic pathway for the conversion of a fermentable carbon substrate to 2-butanol may be constructed using techniques well known in the art. Genes encoding the enzymes of, for example, the 2-butanol biosynthetic Pathway 1: acetolactate synthase, acetolactate decarboxylase, acetoin aminase (or amine:pyruvate transaminase), aminobutanol kinase, aminobutanol O-phosphate lyase and butanol dehydrogenase may be isolated from various sources, as described above.
[0165] Methods of obtaining desired genes from a bacterial genome are common and well known in the art of molecular biology. For example, if the sequence of the gene is known, primers may be designed and the desired sequence amplified using standard primer-directed amplification methods such as polymerase chain reaction (U.S. Pat. No. 4,683,202) to obtain amounts of DNA suitable for cloning into expression vectors. If a gene that is heterologous to a known sequence is to be isolated, suitable genomic libraries may be created by restriction endonuclease digestion and may be screened with probes having complementary sequence to the desired gene sequence. Once the sequence is isolated, the DNA may be amplified using standard primer-directed amplification methods such as polymerase chain reaction (U.S. Pat. No. 4,683,202) to obtain amounts of DNA suitable for cloning into expression vectors, which are then tramsformed into appropriate host cells.
[0166] In addition, given the amino acid sequence of a protein with desired enzymatic activity, the coding sequence may be ascertained by reverse translating the protein sequence. A DNA fragment containing the coding sequence may be prepared synthetically and cloned into an expression vector, then transformed into the desired host cell.
[0167] In preparing a synthetic DNA fragment containing a coding sequence, this sequence may be optimized for expression in the target host cell. Tools for codon optimization for expression in a heterologous host are readily available. Some tools for codon optimization are available based on the GC content of the host organism. The GC contents of some exemplary microbial hosts are given Table 3.
TABLE-US-00003 TABLE 3 GC Contents of Microbial Hosts Strain % GC B. licheniformis 46 B. subtilis 42 C. acetobutylicum 37 E. coli 50 P. putida 61 A. eutrophus 61 Paenibacillus macerans 51 Rhodococcus erythropolis 62 Brevibacillus 50 Paenibacillus polymyxa 50
[0168] Once the relevant pathway genes are identified and isolated they may be transformed into suitable expression hosts by means well known in the art. Vectors useful for the transformation of a variety of host cells are common and commercially available from companies such as EPICENTRE® (Madison, Wis.), Invitrogen Corp. (Carlsbad, Calif.), Stratagene (La Jolla, Calif.), and New England Biolabs, Inc. (Beverly, Mass.). Typically the vector contains a selectable marker and sequences allowing autonomous replication or chromosomal integration in the desired host. In addition, suitable vectors comprise a promoter region which harbors transcriptional initiation controls and a transcriptional termination control region, between which a coding region DNA fragment may be inserted, to provide expression of the inserted coding region. Both control regions may be derived from genes homologous to the transformed host cell, although it is to be understood that such control regions may also be derived from genes that are not native to the specific species chosen as a production host.
[0169] Initiation control regions or promoters, which are useful to drive expression of the relevant pathway coding regions in the desired host cell are numerous and familiar to those skilled in the art. Virtually any promoter capable of driving these genetic elements is suitable for use including, but not limited to, promoters derived from the following genes: CYC1, HIS3, GAL1, GAL10, ADH1, PGK, PHO5, GAPDH, ADC1, TRP1, URA3, LEU2, ENO, TPI, CUP1, FBA, GPD, and GPM (useful for expression in Saccharomyces); AOX1 (useful for expression in Pichia); as well as the lac, ara, tet, trp, IPL, IPR, T7, tac, and trc promoters (useful for expression in Escherichia coli, Alcaligenes, and Pseudomonas); the amy, apr, and npr promoters, and various phage promoters useful for expression in Bacillus subtilis, Bacillus licheniformis, and Paenibacillus macerans; nisA (useful for expression Gram-positive bacteria, Eichenbaum et al. Appl. Environ. Microbiol. 64(8):2763-2769 (1998)); and the synthetic P11 promoter (useful for expression in Lactobacillus plantarum, Rud et al., Microbiology 152:1011-1019 (2006)).
[0170] Termination control regions may also be derived from various genes native to the preferred hosts. Optionally, a termination site may be unnecessary, however, it is most preferred if included.
[0171] Certain vectors are capable of replicating in a broad range of host bacteria and can be transferred by conjugation. The complete and annotated sequence of pRK404 and three related vectors: pRK437, pRK442, and pRK442(H), are available. These derivatives have proven to be valuable tools for genetic manipulation in Gram-negative bacteria (Scott et al., Plasmid 50(1):74-79 (2003)). Several plasmid derivatives of broad-host-range Inc P4 plasmid RSF1010 are also available with promoters that can function in a range of Gram-negative bacteria. Plasmid pAYC36 and pAYC37, have active promoters along with multiple cloning sites to allow for heterologous gene expression in Gram-negative bacteria.
[0172] Chromosomal gene replacement tools are also widely available. For example, a thermosensitive variant of the broad-host-range replicon pWV101 has been modified to construct a plasmid pVE6002 which can be used to effect gene replacement in a range of Gram-positive bacteria (Maguin et al., J. Bacteriol. 174(17):5633-5638 (1992)). The expression of a 2-butanol biosynthetic pathway in various preferred microbial hosts is described in more detail below.
[0173] Expression of a 2-Butanol Biosynthetic Pathway in E. Coli
[0174] Vectors useful for the transformation of E. coli are common and commercially available from the companies listed above. For example, the genes of a 2-butanol biosynthetic pathway may be isolated from various sources, as described above, cloned onto a modified pUC19 vector and transformed into E. coli NM522, as described in Examples 10 and 11. Alternatively, the genes encoding a 2-butanol biosynthetic pathway may be divided into multiple operons, cloned onto expression vectors, and transformed into various E. coli strains, as described in Examples 13, 15, and 15.
[0175] Expression of a 2-Butanol Biosynthetic Pathway in Rhodococcus erythropolis
[0176] A series of E. coli-Rhodococcus shuttle vectors are available for expression in R. erythropolis, including, but not limited to pRhBR17 and pDA71 (Kostichka et al., Appl. Microbiol. Biotechnol. 62:61-68 (2003)). Additionally, a series of promoters are available for heterologous gene expression in R. erythropolis (see for example Nakashima et al., Appl. Environ. Microbiol. 70:5557-5568 (2004), and Tao et al., Appl. Microbiol. Biotechnol. 2005, DOI 10.1007/s00253-005-0064). Targeted gene disruptions in chromosomal genes of R. erythropolis may be created using the methods described by Tao et al., supra, and Brans et al. (Appl. Envion. Microbiol. 66: 2029-2036 (2000)).
[0177] The heterologous genes required for the production of 2-butanol, as described above, may be cloned initially in pDA71 or pRhBR71 and transformed into E. coli. The vectors may then be transformed into R. erythropolis by electroporation, as described by Kostichka et al., supra. The recombinants may be grown in synthetic medium containing glucose and the production of 2-butanol can be followed using fermentation methods known in the art.
[0178] Expression of a 2-Butanol Biosynthetic Pathway in B. subtilis
[0179] Methods for gene expression and creation of mutations in B. subtilis are also well known in the art. For example, the genes of a 2-butanol biosynthetic pathway may be isolated from various sources, as described above, cloned into a modified E. coli-Bacillus shuttle vector and transformed into Bacillus subtilis BE1010, as described in Example 12. The desired genes may be cloned into a Bacillus expression vector and transformed into a strain to make a production host. Alternatively, the genes may be integrated into the Bacillus chromosome using conditional replicons or suicide vectors that are known to one skilled in the art. For example, the Bacillus Genetic Stock Center carries numerous integration vectors.
[0180] Expression of a 2-Butanol Biosynthetic Pathway in B. licheniformis
[0181] Most of the plasmids and shuttle vectors that replicate in B. subtilis may be used to transform B. licheniformis by either protoplast transformation or electroporation. The genes required for the production of 2-butanol may be cloned in plasmids pBE20 or pBE60 derivatives (Nagarajan et al., Gene 114:121-126 (1992)). Methods to transform B. licheniformis are known in the art (for example see Fleming et al. Appl. Environ. Microbiol., 61(11):3775-3780 (1995)). The plasmids constructed for expression in B. subtilis may be transformed into B. licheniformis to produce a recombinant microbial host that produces 2-butanol. Expression of a 2-butanol biosynthetic pathway in Paenibacillus macerans
[0182] Plasmids may be constructed as described above for expression in B. subtilis and used to transform Paenibacillus macerans by protoplast transformation to produce a recombinant microbial host that produces 2-butanol.
[0183] Expression of a 2-Butanol Biosynthetic Pathway in Alcaligenes (Ralstonia) eutrophus
[0184] Methods for gene expression and creation of mutations in Alcaligenes eutrophus are known in the art (see for example Taghavi et al., Appl. Environ. Microbiol., 60(10):3585-3591 (1994)). The genes for a 2-butanol biosynthetic pathway may be cloned in any of the broad host range vectors described above, and electroporated into Alcaligenes eutrophus to generate recombinants that produce 2-butanol. The poly(hydroxybutyrate) pathway in Alcaligenes has been described in detail, a variety of genetic techniques to modify the Alcaligenes eutrophus genome are known, and those tools can be applied for engineering a 2-butanol biosynthetic pathway.
[0185] Expression of a 2-Butanol Biosynthetic Pathway in Pseudomonas putida
[0186] Methods for gene expression in Pseudomonas putida are known in the art (see for example Ben-Bassat et al., U.S. Pat. No. 6,586,229, which is incorporated herein by reference). The genes of a 2-butanol biosynthetic pathway may be inserted into pPCU18, and this ligated DNA may be electroporated into electrocompetent Pseudomonas putida DOT-T1 C5aAR1 cells to generate recombinants that produce 2-butanol.
[0187] Expression of a 2-Butanol Biosynthetic Pathway in Lactobacillus plantarum
[0188] The Lactobacillus genus belongs to the Lactobacillales family and many plasmids and vectors used in the transformation of Bacillus subtilis and Streptococcus may be used for Lactobacillus. Non-limiting examples of suitable vectors include pAMβ1 and derivatives thereof (Renault et al., Gene 183:175-182 (1996); and O'Sullivan et al., Gene 137:227-231 (1993)); pMBB1 and pHW800, a derivative of pMBB1 (Wyckoff et al. Appl. Environ. Microbiol. 62:1481-1486 (1996)); pMG1, a conjugative plasmid (Tanimoto et al., J. Bacteriol. 184:5800-5804 (2002)); pNZ9520 (Kleerebezem et al., Appl. Environ. Microbiol. 63:4581-4584 (1997)); pAM401 (Fujimoto et al., Appl. Environ. Microbiol. 67:1262-1267 (2001)); and pAT392 (Arthur et al., Antimicrob. Agents Chemother. 38:1899-1903 (1994)). Several plasmids from Lactobacillus plantarum have also been reported (van Kranenburg et al., Appl. Environ. Microbiol. 71(3):1223-1230 (2005)).
[0189] The various genes for a 2-butanol biosynthetic pathway may be assembled into any suitable vector, such as those described above. The codons can be optimized for expression based on the codon index deduced from the genome sequences of Lactobacillus plantarum or Lactobacillus arizonensis. The plasmids may be introduced into the host cell using methods known in the art, such as electroporation (Cruz-Rodz et al. Molecular Genetics and Genomics 224:1252-154 (1990), Bringel, et al. Appl. Microbiol. Biotechnol. 33: 664-670 (1990), Alegre et al., FEMS Microbiology letters 241:73-77 (2004)), and conjugation (Shrago et al., Appl. Environ. Microbiol. 52:574-576 (1986)). The 2-butanol biosynthetic pathway genes can also be integrated into the chromosome of Lactobacillus using integration vectors (Hofs et al., Appl. Environ. Microbiol. 60:1401-1403 (1990), Jang et al., Micro. Lett. 24:191-195 (2003)).
[0190] Expression of a 2-Butanol Biosynthetic Pathway in Enterococcus faecium, Enterococcus gallinarium, and Enterococcus faecalis
[0191] The Enterococcus genus belongs to the Lactobacillales family and many plasmids and vectors used in the transformation of Lactobacillus, Bacillus subtilis, and Streptococcus, described above, may be used for Enterococcus. Expression vectors for E. faecalis using the nisA gene from Lactococcus may also be used (Eichenbaum et al., Appl. Environ. Microbiol. 64:2763-2769 (1998). Additionally, vectors for gene replacement in the E. faecium chromosome may be used (Nallaapareddy et al., Appl. Environ. Microbiol. 72:334-345 (2006)).
[0192] The various genes for a 2-butanol biosynthetic pathway may be assembled into any suitable vector, such as those described above. The codons can be optimized for expression based on the codon index deduced from the genome sequences of Enterococcus faecalis or Enterococcus faecium. The plasmids may be introduced into the host cell using methods known in the art, such as electroporation, as described by Cruz-Rodz et al. (Molecular Genetics and Genomics 224:1252-154 (1990)) or conjugation, as described by Tanimoto et al. (J. Bacteriol. 184:5800-5804 (2002)) and Grohamann et al. (Microbiol. Mol. Biol. Rev. 67:277-301 (2003)).
[0193] Expression of a 2-Butanol Biosynthetic Pathway in Pediococcus pentosaceus and Pediococcus acidilactici,
[0194] The Pediococcus genus belongs to the Lactobacillales family and many plasmids and vectors used in the transformation of Bacillus subtilis and Streptococcus, described above, may be used for Pediococcus. A non-limiting example of a suitable vector is pHPS9 (Bukhtiyarova et al. Appl. Environ. Microbiol. 60:3405-3408 (1994)). Several plasmids from Pediococcus have also been reported (Alegre et al., FEMS Microbiol. Lett. 250:151-156 (2005); Shareck et al. Crit. Rev Biotechnol. 24:155-208 (2004)).
[0195] The genes for a 2-butanol biosynthetic pathway may be assembled into any suitable vector, such as those described above. The codons can be optimized for expression based on the codon index deduced from the genome sequence of Pediococcus pentosaceus. The plasmids may be introduced into the host cell using methods known in the art, such as electroporation (see for example, Osmanagaoglu et al., J. Basic Microbiol. 40:233-241 (2000); Alegre et al., FEMS Microbiol. Lett. 250:151-156 (2005)) and conjugation (Gonzalez and Kunka, Appl. Environ. Microbiol. 46:81-89 (1983)). The 2-butanol biosynthetic pathway genes can also be integrated into the chromosome of Pediococcus using integration vectors (Davidson et al. Antonie van Leeuwenhoek 70:161-183 (1996)).
Fermentation Media
[0196] Fermentation media in the present invention must contain suitable carbon substrates. Suitable substrates may include but are not limited to monosaccharides such as glucose and fructose, oligosaccharides such as lactose or sucrose, polysaccharides such as starch or cellulose or mixtures thereof and unpurified mixtures from renewable feedstocks such as cheese whey permeate, cornsteep liquor, sugar beet molasses, and barley malt. Additionally the carbon substrate may also be one-carbon substrates such as carbon dioxide, or methanol for which metabolic conversion into key biochemical intermediates has been demonstrated. In addition to one and two carbon substrates, methylotrophic organisms are also known to utilize a number of other carbon containing compounds such as methylamine, glucosamine and a variety of amino acids for metabolic activity. For example, methylotrophic yeasts are known to utilize the carbon from methylamine to form trehalose or glycerol (Bellion et al., Microb. Growth C1 Compd., [Int. Symp.], 7th (1993), 415-32, Editor(s): Murrell, J. Collin; Kelly, Don P. Publisher: Intercept, Andover, UK). Similarly, various species of Candida will metabolize alanine or oleic acid (Sutter et al., Arch. Microbiol. 153:485-489 (1990)). Hence it is contemplated that the source of carbon utilized in the present invention may encompass a wide variety of carbon containing substrates and will only be limited by the choice of organism.
[0197] Although it is contemplated that all of the above mentioned carbon substrates and mixtures thereof are suitable in the present invention, preferred carbon substrates are glucose, fructose, and sucrose. Sucrose may be derived from renewable sugar sources such as sugar cane, sugar beets, cassaya, sweet sorghum, and mixtures thereof. Glucose and dextrose may be derived from renewable grain sources through saccharification of starch based feedstocks including grains such as corn, wheat, rye, barley, oats, and mixtures thereof. In addition, fermentable sugars may be derived from renewable cellulosic or lignocellulosic biomass through processes of pretreatment and saccharification, as described, for example, in co-owned and co-pending U.S. Patent Application Publication No. 2007/0031918A1, which is herein incorporated by reference. Biomass refers to any cellulosic or lignocellulosic material and includes materials comprising cellulose, and optionally further comprising hemicellulose, lignin, starch, oligosaccharides and/or monosaccharides. Biomass may also comprise additional components, such as protein and/or lipid. Biomass may be derived from a single source, or biomass can comprise a mixture derived from more than one source; for example, biomass may comprise a mixture of corn cobs and corn stover, or a mixture of grass and leaves. Biomass includes, but is not limited to, bioenergy crops, agricultural residues, municipal solid waste, industrial solid waste, sludge from paper manufacture, yard waste, wood and forestry waste. Examples of biomass include, but are not limited to, corn grain, corn cobs, crop residues such as corn husks, corn stover, grasses, wheat, wheat straw, barley, barley straw, hay, rice straw, switchgrass, waste paper, sugar cane bagasse, sorghum, soy, components obtained from milling of grains, trees, branches, roots, leaves, wood chips, sawdust, shrubs and bushes, vegetables, fruits, flowers, animal manure, and mixtures thereof.
[0198] In addition to an appropriate carbon source, fermentation media must contain suitable minerals, salts, cofactors, buffers and other components, known to those skilled in the art, suitable for the growth of the cultures and promotion of an enzymatic pathway necessary for 2-butanol production.
Culture Conditions with Temperature Lowering
[0199] In the present method, the recombinant microbial production host which produces 2-butanol is seeded into a fermentation medium comprising a fermentable carbon substrate to create a fermentation culture. The production host is grown in the fermentation culture at a first temperature for a first period of time. The first temperature is typically from about 25° C. to about 40° C.
[0200] Suitable fermentation media in the present invention include common commercially prepared media such as Luria Bertani (LB) broth, Sabouraud Dextrose (SD) broth or Yeast Medium (YM) broth. Other defined or synthetic growth media may also be used, and the appropriate medium for growth of the particular microorganism will be known by one skilled in the art of microbiology or fermentation science. The use of agents known to modulate catabolite repression directly or indirectly, e.g., cyclic adenosine 2':3'-monophosphate, may also be incorporated into the fermentation medium.
[0201] Suitable pH ranges for the fermentation are between pH 5.0 to pH 9.0, where pH 6.0 to pH 8.0 is preferred as the initial condition.
[0202] Fermentations may be performed under aerobic or anaerobic conditions, where anaerobic or microaerobic conditions are preferred.
[0203] The first period of time to grow the production host at the first temperature may be determined in a variety of ways. For example, during this period of growth a metabolic parameter of the fermentation culture may be monitored. The metabolic parameter that is monitored may be any parameter known in the art, including, but not limited to the optical density, pH, respiratory quotient, fermentable carbon substrate utilization, CO2 production, and 2-butanol production. During this period of growth, additional fermentable carbon substrate may be added, the pH may be adjusted, oxygen may be added for aerobic cells, or other culture parameters may be adjusted to support the metabolic activity of the culture. Though nutrients and culture conditions are supportive of growth, after a period of time the metabolic activity of the fermentation culture decreases as determined by the monitored parameter described above. For example, a decrease in metabolic activity may be indicated by a decrease in one or more of the following parameters: rate of optical density change, rate of pH change, rate of change in respiratory quotient (if the host cells are aerobic), rate of fermentable carbon substrate utilization, rate of 2-butanol production, rate of change in CO2 production, or rate of another metabolic parameter. The decrease in metabolic activity is related to the sensitivity of the host cells to the production of 2-butanol and/or the presence of 2-butanol in the culture. When decreased metabolic activity is detected, the temperature of the fermentation culture is lowered to reduce the sensitivity of the host cells to 2-butanol and thereby allow further production of 2-butanol. In one embodiment, the lowering of the temperature coincides with a change in the metabolic parameter that is monitored.
[0204] In one embodiment, the change in metabolic activity is a decrease in the rate of 2-butanol production. 2-Butanol production may be monitored by analyzing the amount of 2-butanol present in the fermentation culture medium as a function of time using methods well known in the art, such as using high performance liquid chromatography (HPLC) or gas chromatography (GC), which are described in the Examples herein. GC is preferred due to the short assay time.
[0205] Alternatively, the lowering of the temperature of the fermentation culture may occur at a predetermined time. The first period of time may be predetermined by establishing a correlation between a metabolic parameter of the fermentation culture and time in a series of test fermentations runs. A correlation between a metabolic parameter, as described above, and time of culture growth may be established for any 2-butanol producing host by one skilled in the art. The specific correlation may vary depending on conditions used including, but not limited to, carbon substrate, fermentation conditions, and the specific recombinant 2-butanol producing microbial production host. The correlation is most suitably made between 2-butanol production or specific glucose consumption rate and time of culture growth. Once the predetermined time has been established from the correlation, the temperature of the fermentation culture in subsequent fermentation runs is lowered at the predetermined time. For example, if it is determined by monitoring a metabolic parameter in the test fermentation runs that the rate of production of 2-butanol decreases after 12 hours, the temperature in subsequent fermentations runs is lowered after 12 hours without the need to monitor 2-butanol production in the subsequent runs.
[0206] After the first period of time, the temperature of the fermentation culture is lowered to a second temperature. Typically, the second temperature is about 3° C. to about 25° C. lower than the first temperature. Reduction in temperature to enhance tolerance of the host cells to 2-butanol is balanced with maintaining the temperature at a level where the cells continue to be metabolically active for 2-butanol production. For example, a fermentation culture that has been grown at about 35° C. may be reduced in temperature to about 28° C.; or a culture grown at about 30° C. may be reduced in temperature to about 25° C. The change in temperature may be done gradually over time or may be made as a step change. The production host is incubated at the second temperature for a second period of time, so that 2-butanol production continues. The second period of time may be determined in the same manner as the first period of time described above, e.g., by monitoring a metabolic parameter or by using a predetermined time.
[0207] Additionally, the temperature lowering and incubation steps may be repeated one or more times to more finely balance metabolic activity for 2-butanol production and 2-butanol sensitivity. For example, a culture that has been grown at about 35° C. may be reduced in temperature to about 32° C., followed by an incubation period. During this period a metabolic parameter of the fermentation culture may be monitored as described above, or a predetermined time may be used. It is particularly suitable to monitor the production of 2-butanol during this incubation period. When monitoring indicates a decrease in metabolic activity or at a predetermined time, the temperature may be reduced a second time. For example, the temperature may be reduced from about 32° C. to about 28° C. The temperature lowering and incubation steps may be repeated a third time where the temperature is reduced, for example, to about 20° C. The production host is incubated at the lowered temperature so that 2-butanol production continues. The steps may be repeated further as necessary to obtain the desired 2-butanol titer.
Industrial Batch and Continuous Fermentations
[0208] The present process employs a batch method of fermentation. A classical batch fermentation is a closed system where the composition of the medium is set at the beginning of the fermentation and not subject to artificial alterations during the fermentation. Thus, at the beginning of the fermentation the medium is inoculated with the desired organism or organisms, and fermentation is permitted to occur without adding anything to the system. Typically, however, a "batch" fermentation is batch with respect to the addition of carbon source and attempts are often made at controlling factors such as pH and oxygen concentration. In batch systems the metabolite and biomass compositions of the system change constantly up to the time the fermentation is stopped. Within batch cultures cells moderate through a static lag phase to a high growth log phase and finally to a stationary phase where growth rate is diminished or halted. If untreated, cells in the stationary phase will eventually die. Cells in log phase generally are responsible for the bulk of production of end product or intermediate.
[0209] A variation on the standard batch system is the fed-batch system. Fed-batch fermentation processes are also suitable in the present invention and comprise a typical batch system with the exception that the substrate is added in increments as the fermentation progresses. Fed-batch systems are useful when catabolite repression is apt to inhibit the metabolism of the cells and where it is desirable to have limited amounts of substrate in the media. Measurement of the actual substrate concentration in fed-batch systems is difficult and is therefore estimated on the basis of the changes of measurable factors such as pH, dissolved oxygen and the partial pressure of waste gases such as CO2. Batch and fed-batch fermentations are common and well known in the art and examples may be found in Thomas D. Brock in Biotechnology: A Textbook of Industrial Microbiology, Second Edition (1989) Sinauer Associates, Inc., Sunderland, Mass., or Deshpande, Mukund V., Appl. Biochem. Biotechnol., 36:227, (1992), herein incorporated by reference.
[0210] Although the present invention is performed in batch mode it is contemplated that the method would be adaptable to continuous fermentation methods. Continuous fermentation is an open system where a defined fermentation medium is added continuously to a bioreactor and an equal amount of conditioned media is removed simultaneously for processing. Continuous fermentation generally maintains the cultures at a constant high density.
[0211] Continuous fermentation allows for the modulation of one factor or any number of factors that affect cell growth or end product concentration. For example, one method will maintain a limiting nutrient such as the carbon source or nitrogen level at a fixed rate and allow all other parameters to moderate. In other systems a number of factors affecting growth can be altered continuously while the cell concentration, measured by the turbidity of the culture medium, is kept constant. Continuous systems strive to maintain steady state growth conditions and thus the cell loss due to the medium being drawn off must be balanced against the cell growth rate in the fermentation. Methods of modulating nutrients and growth factors for continuous fermentation processes as well as techniques for maximizing the rate of product formation are well known in the art of industrial microbiology and a variety of methods are detailed by Brock, supra.
[0212] It is contemplated that the present invention may be practiced using either batch, fed-batch or continuous processes and that any known mode of fermentation would be suitable. Additionally, it is contemplated that cells may be immobilized on a substrate as whole cell catalysts and subjected to fermentation conditions for 2-butanol production.
Methods for 2-Butanol Isolation from the Fermentation Medium
[0213] The bioproduced 2-butanol may be isolated from the fermentation medium using methods known in the art for ABE fermentations (see for example, Durre, Appl. Microbiol. Biotechnol. 49:639-648 (1998), Groot et al., Process Biochem. 27:61-75 (1992), and references therein). For example, solids may be removed from the fermentation medium by centrifugation, filtration, decantation, or the like. Then, the 2-butanol may be isolated from the fermentation medium using methods such as distillation, azeotropic distillation, liquid-liquid extraction, adsorption, gas stripping, membrane evaporation, or pervaporation.
EXAMPLES
[0214] The present invention is further defined in the following Examples. It should be understood that these Examples, while indicating a preferred embodiment of the invention, are given by way of illustration only. From the above discussion and these Examples, one skilled in the art can ascertain the essential characteristics of this invention, and, without departing from the spirit and scope thereof, can make various changes and modifications of the invention to adapt it to various uses and conditions.
General Methods
[0215] Standard recombinant DNA and molecular cloning techniques described in the Examples are well known in the art and are described by Sambrook, J., Fritsch, E. F. and Maniatis, T. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, N.Y., (1989) (Maniatis) and by T. J. Silhavy, M. L. Bennan, and L. W. Enquist, Experiments with Gene Fusions, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y. (1984) and by Ausubel, F. M. et al., Current Protocols in Molecular Biology, pub. by Greene Publishing Assoc. and Wiley-Interscience (1987).
[0216] Materials and methods suitable for the maintenance and growth of bacterial cultures are well known in the art. Techniques suitable for use in the following Examples may be found as set out in Manual of Methods for General Bacteriology (Phillipp Gerhardt, R. G. E. Murray, Ralph N. Costilow, Eugene W. Nester, Willis A. Wood, Noel R. Krieg and G. Briggs Phillips, eds), American Society for Microbiology, Washington, D.C. (1994)) or by Thomas D. Brock in Biotechnology: A Textbook of Industrial Microbiology, Second Edition, Sinauer Associates, Inc., Sunderland, Mass. (1989). All reagents, restriction enzymes and materials described for the growth and maintenance of bacterial cells were obtained from Aldrich Chemicals (Milwaukee, Wis.), BD Diagnostic Systems (Sparks, Md.), Life Technologies (Rockville, Md.), or Sigma Chemical Company (St. Louis, Mo.) unless otherwise specified. Bacterial strains are obtained from the American Type Culture Collection (ATCC, Manassas, Va.) unless otherwise noted. Oligonucleotide primers described in the following Examples are given in Table 4. All oligonucleotide primers were synthesized by Sigma-Genosys Woodlands, Tex.).
TABLE-US-00004 TABLE 4 Cloning and Screening Primers Gene SEQ ID Name Primer Sequence NO: Description budB B1 CACCATGGACAAACAG 15 budB TATCCGGTACGCC forward budB B2 CGAAGGGCGATAGCTT 16 budB TACCAATCC reverse budA B3 CACCATGAATCATTCTG 17 budA forward CTGAATGCACCTGCG budA B4 GATACTGTTTGTCCATG 18 budA reverse TGACC budC B5 CACCATGAAAAAAGTC 19 budC GCACTTGTTACC forward budC B6 TTAGTTAAATACCAT 20 budC reverse pddA B7 CACCATGAGATCGA 21 pddABC AAAGATTTG forward pddC B8 CTTAGAGAAGTTAATCG 22 pddABC TCGCC reverse sadh B9 CACCATGAAAGCCCTC 23 sadh CAGTACACC forward sadh B10 CGTCGTGTCATGCCCG 24 sadh GG reverse budA B11 GATCGAATTCGTTTAAA 25 budABC CTTAGTTTTCTACCGCA forward CG budC B12 GATCGCATGCAAGCTT 26 budABC TCATATAGTCGGAATTC reverse C pddA B13 GATCGAATTCGTTTAAA 27 pddABC CAAAGGAGGTCTGATT forward CATGAGATCG pddC B14 GATCGGATTCTTAATCG 28 pddABC TCGCC reverse sadh B15 GATCGGATCCAAAGGA 29 sadh GGTCGGGCGCATGAAA forward GCCC sadh B16 GATCTCTAGAAAGCTTT 30 sadh CAGCCCGGGACGACC reverse -- BenF ACTTTCTTTCGCCTGTT 31 -- TCAC -- BenBPR CATGAAGCTTGTTTAAA 32 -- CTCGGTGACCTTGAAA ATAATGAAAACTTATAT TGTTTTGAAAATAATGA AAACTTATATTG budAB BABC F GAGCTCGAATTCAAAG 33 budAB GAGGAAGTGTATATGA forward ATCATTC budAB BAB R GGATCCTCTAGAATTAG 34 budAB TTAAATACCATCCCGCC reverse G budC BC Spe F ACTAGTAAAGGAGGAA 40 budC forward AGAGTATGAAGAAGGT CGCACT budC BC Xba R TCTAGAAAGCAGGGGC 41 budC reverse AAGCCATGTC pddAB DDo For AAGCTTAAAGGAGGCT 44 pddABC- C-ddrAB GATTCATGAGATCGAAA ddrAB AGATT forward pddAB DDo Rev TCTAGATTATTCATCCT 45 pddABC- C-ddrAB GCTGTTCTCC ddrAB reverse chnA ChnA F CATCAATTGACTACGTA 54 chnA forward GTCGTACGTGTAAGGA GGTTTGAAATGGAAAAA ATTATG chnA ChnA R CATGCTAGCCCCGGGT 55 chnA reverse ATCTTCTACTCATTTTTT ATTTCG -- Top ter F1 CTAGAAGTCAAAAGCC 58 forward TCCGACCGGAGGCTTT TGA -- Top ter F2 CTGCTCGAGTTGCTAG 59 forward CAAGTTTAAACAAAAA AAAGCCCGCTCATTAG GCGGGCTGAGCT -- Bot ter R1 CAGCCCGCCTAATGAG 60 reverse CGGGCTTTTTTTTGTTT AAAC -- Bot ter R2 TTGCTAGCAACTCGAG 61 reverse CAGTCAAAAGCCTCCG GTCGGAGGCTTTTGAC TT KA-AT OT872 CTCCGGAATTCATGTC 127 Aminoalcohol TGACGGACGACTCACC kinase/lyase GCA operon forward KA-AT OT873 TTCCAATGCATTGGCT 128 Aminoalcohol GCAGTTATCTCTGTGC kinase/lyase ACGAGTGCCGATGA operon reverse KA OT879 AACAGCCAAGCTTGGC 129 Aminoalcohol TGCAGTCATCGCGCAT kinase TCTCCGGG reverse AT OT880 TCTCCGGAATTCATGA 130 Aminoalcohol CGTCTGAAATGACAGC lyase GACAGAAG forward pBAD. OT909 GCTAACAGGAGGAAG 131 Adds EcoRl HisB AATTCATGGGGGGTTC site to replace TC Ncol site pBAD. OT910 GAGAACCCCCCATGAA 132 Adds EcoRl HisB TTCTTCCTCCTGTTAG site to replace C Ncol site BudAB N84seqR3 GGACCTGCTTCGCTTT 159 reverse ATCG APT APTfor GCGCGCCCGGGAAGA 162 APT forward AGGAGCTCTTCACCAT GAACAAACCACAGTCTT GG APT APTrev GCGCGCCCGGGTTCAT 163 APT reverse GCCACCTCTGCG
TABLE-US-00005 TABLE 5 Sequencing Primers Gene- SEQ Name Sequence specific ID NO: M13 Forward GTAAAACGACGGCCAGT -- 35 M13 Reverse AACAGCTATGACCATG -- 36 N83 SeqF2 GCTGGATTACCAGCTCGACC -- 37 N83 SeqF3 CGGACGCATTACCGGCAAAG -- 38 N84 Seq R2 GCATCGAGATTATCGGGATG -- 65 N84 SeqR4 CGAAGCGAGAGAAGTTATCC -- 39 Trc F TTGACAATTAATCATCCGGC all 42 Trc R CTTCTCTCATCCGCCAAAAC all 43 DDko seq F2 GCATGGCGCGGATTTGACGAAC pddABC- 46 ddrAB DDko seq F5 CATTAAAGAGACCAAGTACGTG pddABC- 47 ddrAB DDko seq F7 ATATCCTGGTGGTGTCGTCGGC pddABC- 48 GT ddrAB DDko seq F9 TCTTTGTCACCAACGCCCTGCG pddABC- 49 ddrAB DDko seq R1 GCCCACCGCGCTCGCCGCCGCG pddABC- 50 ddrAB DDko seq R3 CCCCCAGGATGGCGGCTTCGGC pddABC- 51 ddrAB DDko seq R7 GGGCCGACGGCGATAATCACTT pddABC- 52 ddrAB DDko seq TTCTTCGATCCACTCCTTAACG pddABC- 53 R10 ddrAB chnSeq F1 CTCAACAGGGTGTAAGTGTAGT chnA 56 chnSeq R1 CGTTTTGATATAGCCAGGATGT chnA 57 pCL1925 CGGTATCATCAACAGGCTTACC all 62 vec F pCL1925 AGGGTTTTCCCAGTCACGACGT all 63 vec R1 pCL1925 CGCAATAGTTGGCGAAGTAATC all 64 vec R2 APTseqRev GCTAGAGATGATAGC APT 160 APTseqFor GGAAGAGACTATCCAGCG APT 161
Methods for Determining 2-Butanol and 2-Butanone Concentration in Culture Media
[0217] The concentration of 2-butanol and 2-butanone in the culture media can be determined by a number of methods known in the art. For example, a specific high performance liquid chromatography (HPLC) method utilized a Shodex SH-1011 column with a Shodex SH-G guard column, both purchased from Waters Corporation (Milford, Mass.), with refractive index (R1) detection. Chromatographic separation was achieved using 0.01 M H2SO4 as the mobile phase with a flow rate of 0.5 mL/min and a column temperature of 50° C. Under the conditions used, 2-butanone and 2-butanol had retention times of 39.5 and 44.3 min, respectively. Alternatively, gas chromatography (GC) methods are available. For example, a specific GC method utilized an HP-INNOWax column (30 m×0.53 mm id, 1 μm film thickness, Agilent Technologies, Wilmington, Del.), with a flame ionization detector (FID). The carrier gas was helium at a flow rate of 4.5 mL/min, measured at 150° C. with constant head pressure; injector split was 1:25 at 200° C.; oven temperature was 45° C. for 1 min, 45 to 220° C. at 10° C./min, and 220° C. for 5 min; and FID detection was employed at 240° C. with 26 mL/min helium makeup gas. The retention times of 2-butanone and 2-butanol were 3.61 and 5.03 min, respectively.
[0218] 2-Butanone can also be detected by derivatization with 3-methyl-2-benzothiazolinone hydrazone (MBTH). An aqueous solution containing 2-butanone is mixed with an equal volume of an aqueous solution of 6 mg/mL MBTH in 375 mM glycine-HCl (pH 2.7) and incubated at 100° C. for 3 min. The resulting MBTH-derivatized samples are analyzed on a 25 cm×4.6 mm (id) Supelosil LC-18-D5 5 μm column (Supelco) using a mobile phase of 55% acetonitrile in water at a flow rate of 1 mL/min. The 2-butanone derivative appears as two peaks (cis and trans isomers) with retention times of approximately 12.3 and 13.3 min and absorbance maxima of 230 and 307 nm.
[0219] The meaning of abbreviations is as follows: "s" means second(s), "min" means minute(s), "h" means hour(s), "psi" means pounds per square inch, "nm" means nanometers, "d" means day(s), "μL" means microliter(s), "mL" means milliliter(s), "L" means liter(s), "mm" means millimeter(s), "nm" means nanometers, "mM" means millimolar, "M" means molar, "mmol" means millimole(s), "μmol" means micromole(s)", "g" means gram(s), "μg" means microgram(s) and "ng" means nanogram(s), "PCR" means polymerase chain reaction, "OD" means optical density, "OD600" means the optical density measured at a wavelength of 600 nm, "kDa" means kilodaltons, "g" means the gravitation constant, "bp" means base pair(s), "kbp" means kilobase pair(s), "% w/v" means weight/volume percent, % v/v'' means volume/volume percent, "wt %" means percent by weight, "nt" means not tested, "HPLC" means high performance liquid chromatography, and "GC" means gas chromatography. The term "molar selectivity" is the number of moles of product produced per mole of sugar substrate consumed and is reported as a percent.
Example 1
Increased Tolerance of Lactobacillus plantarum PN0512 to 1-butanol, iso-butanol and 2-butanol at Decreased Growth Temperatures
[0220] Tolerance levels of bacterial strain Lactobacillus plantarum PN0512 (ATCC # PTA-7727) were determined at 25° C., 30° C. and 37° C. as follows. The strain was cultured in S30L medium (i.e., 10 mM ammonium sulfate, 5 mM potassium phosphate buffer, pH 7.0, 50 mM MOPS, pH 7.0, 2 mM MgCl2, 0.7 mM CaCl2, 50 μM MnCl2, 1 μM FeCl3, 1 μM ZnCl2, 1.72 μM CuCl2, 2.53 μM CoCl2, 2.42 μM Na2MoO4, 2 μM thiamine hydrochloride, 10 mM glucose, and 0.2% yeast extract). An overnight culture in the absence of any test compound was started in 15 mL of the S30L medium in a 150 mL flask, with incubation at 37° C. in a shaking water bath. The next morning, the overnight culture was diluted into three 500 mL flasks containing 150 mL of fresh medium to an initial OD600 of about 0.08. Each flask was incubated in a shaking water bath, one each at 25° C., 30° C. and 37° C. Each large culture was allowed to acclimate at the test temperature for at least 0.5 h. After the acclimation period, each large culture was split into flasks in the absence (control) and in the presence of various amounts of 1-butanol, isobutanol or 2-butanol, as listed in Tables 6, 7, and 8, respectively. Growth was followed by measuring OD600 for six hours after addition of the compounds. The results are summarized in Tables 6, 7, and 8 below.
TABLE-US-00006 TABLE 6 Growth of L. plantarum PN0512 in the presence of 1-butanol at different temperatures Concentration 1- butanol (% w/v) 37° C. 30° C. 25° C. 0.0 +1 + + 1.0 + nt3 nt 1.2 + nt nt 1.4 + nt nt 1.5 + + + 1.6 + nt nt 1.8 + nt nt 2.0 + + + 2.1 + nt nt 2.2 + nt nt 2.3 + nt nt 2.4 -2 + + 2.5 - nt nt 2.7 - + nt 2.9 - - + 3.1 - - + 3.2 nt - - 3.3 nt nt - 3.4 nt - - 1"+" = growth observed as an increase in OD600. 2"-" = no growth observed, i.e. no change in OD600. 3"nt" = not tested
TABLE-US-00007 TABLE 7 Growth of L. plantarum PN0512 in the presence of isobutanol at different temperatures Concentration isobutanol (% w/v) 37° C. 30° C. 25° C. 0.0 +1 + + 0.5 + nt3 nt 1.0 + nt nt 1.5 + + + 1.6 + nt nt 1.8 + nt nt 2.0 + + + 2.1 + nt nt 2.3 + nt nt 2.4 + + + 2.5 + nt nt 2.7 + + + 2.9 + + + 3.1 + + + 3.3 nt -2 + 3.4 - nt nt 3.5 nt nt + 3.6 nt nt - 3.8 - nt nt 4.3 - nt nt 1"+" = growth observed as an increase in OD600. 2"-" = no growth observed, i.e. no change in OD600. 3"nt" = not tested
TABLE-US-00008 TABLE 8 Growth of L. plantarum PN0512 in the presence of 2-butanol at different temperatures Concentration 2- butanol (% w/v) 37° C. 30° C. 25° C. 0.0 +1 + + 1.8 + nt3 nt 2.1 + nt nt 2.5 + nt nt 2.9 + + + 3.1 + nt nt 3.5 + nt nt 3.6 + nt nt 3.8 + + + 4.0 nt + nt 4.3 + + + 4.5 -2 + nt 4.7 - + + 4.9 nt - + 5.2 - nt + 5.6 - nt - 6.0 - nt nt 6.4 - nt nt 7.3 - nt nt 1"+" = growth observed as an increase in OD600. 2"-" = no growth observed, i.e. no change in OD600. 3"nt" = not tested
[0221] All three butanols showed a similar effect of temperature on growth inhibition of L. plantarum PN0512. The concentration that resulted in full growth inhibition was greater at 25° C. than at 37° C. In the case of 1-butanol, growth was observed at 37° C. in 2.3% 1-butanol, but not 2.4%. However, at 30° C. growth was observed in 2.7%, but not 2.9%, and at 25° C. growth was observed even in 3.1% 1-butanol. Thus, the concentration of 1-butanol that completely inhibited growth increased as growth temperature decreased. Likewise, in the case of isobutanol, growth was observed in 3.5% at 25° C. while growth was observed in 3.1% at 30° C. and 37° C., but not in 3.3% or 3.4%. Similarly, in the case of 2-butanol growth was observed at 37° C. in 4.3%, but not in 4.5%; at 30° C. in 4.7%, but not in 4.9%; and at 25° C. in 5.2%. Thus the tolerance of L. plantarum PN0512 to butanols increased with decreased growth temperature.
Example 2
Increased Tolerance of Escherichia coli to 1-Butanol at Decreased Exposure Temperature
[0222] The effect of growth and exposure temperature on survival of Escherichia coli in the presence of 1-butanol was tested using stationary phase cultures in a rich medium and log phase cultures in a defined medium. For the stationary phase studies, E. coli strain MG1655 (ATCC #700926) was grown overnight in LB medium (Teknova, Half Moon Bay, Calif.) with shaking at 250 rpm at 42° C., 29° C. or 28° C. Survival of 1-butanol shock was tested at exposure temperatures of 0° C., 28° C. or 42° C. The 1-butanol exposure at 28° C. or 42° C. was started immediately after removing the overnight cultures from the growth incubators. The 1-butanol exposure at 0° C. was done after allowing the overnight cultures to cool on ice for about 15 min. A series of solutions of 1-butanol at different concentrations in LB medium was made and 90 μL aliquots were put in microfuge tubes. To these were added 10 μL of the overnight cultures and the tubes were immediately placed in shaking incubators at 42° C. or 28° C. or left on ice for 30 min. To stop the effect of 1-butanol on the cultures, a 10-2 dilution was done by placing 2 μL of the treated culture into 198 μL of LB medium in wells of a microplate. Then, 5 μL of the undiluted treated cultures were spotted on LB agar plates. Subsequent 10-fold serial dilutions of 10-3, 10-4, 10-5 and 10-6 of the exposed cultures were done by serial pipetting of 20 μL, starting with the 10-2 dilution cultures, into 180 μL of LB medium in the microplate, using a multi-channel pipette. Prior to each transfer, the cultures were mixed by pipetting up and down six times. Each dilution (5 μL) was spotted onto an LB plate using a multi-channel pipette and allowed to soak into the plate. The plates were inverted and incubated overnight at 37° C. The number of colonies for each dilution was counted and the % growth inhibition was calculated by comparison with a control culture that had not been exposed to 1-butanol. Survival of 0% was recorded when no colonies in the spots of the undiluted or any of the serial dilutions were observed. The results are shown in Table 9.
TABLE-US-00009 TABLE 9 Survival of stationary phase E. coli in 1-butanol at 42° C., 28° C., or 0° C. Grown at Grown at Grown at Grown at Grown at Grown at 42° C. 29° C. 42° C. 28° C. 42° C. 29° C. % survival after % survival after % survival after 1-Butanol 30 min exposure 30 min exposure 30 min exposure % (w/v) at 42° C. at 28° C. at 0° C. 1.0 100 100 100 100 100 100 1.5 0.1 0.1 100 100 100 100 2.0 0 0.1 100 100 100 100 2.5 0 0 100 100 100 100 3.0 0 0 100 100 100 100 3.5 0 0 3 10 100 100 4.0 0 0 0.0004 0.0003 100 100 5.0 .sup. nt1 nt nt nt 1 1 6.0 nt nt nt nt 0 0.001 7.0 nt nt nt nt 0 0 1"nt" = not tested
[0223] A similar study was done with log-phase cultures of E. coli grown in a defined medium. E. coli strain MG1655 was allowed to grow overnight in MOPS 0.2% glucose medium (Teknova, Half Moon Bay, Calif.) at 42° C. or 28° C. The following day, the cultures were diluted into fresh medium and allowed to grow at the same temperature until in the log phase of growth. The OD600 was 0.74 for the 28° C. culture and was 0.72 for the 42° C. culture. Both of these log phase cultures were exposed to 1-butanol at 42° C., 28° C. and 0° C. as follows. A series of solutions of 1-butanol at different concentrations in MOPS 0.2% glucose medium was made and 90 μL aliquots were put in microfuge tubes. To these were added 10 μL of the log phase cultures and the tubes were immediately placed in shaking incubators at 42° C. or 28° C. or left on ice for 30 min. To stop the effect of 1-butanol on the cultures, a 10-2 dilution was done by placing 2 μL of the treated culture into 198 μL of LB medium in wells of a microplate. Then 5 μL of the undiluted treated cultures were spotted on LB agar plates. Subsequent 10-fold serial dilutions of 10-3, 10-4, 10-6 and 10-6 of the exposed cultures were done by serial pipetting of 20 μL, starting with the 10-2 dilution cultures, into 180 μL of LB medium in the microplate, using a multi-channel pipette. Prior to each transfer, the cultures were mixed by pipetting up and down six times. Each dilution (5 μL) was spotted onto an LB plate using a multi-channel pipette and allowed to soak into the plate. The plates were inverted and incubated overnight at 37° C. The number of colonies for each dilution was counted and the % growth inhibition was calculated by comparison with a control culture that had not been exposed to 1-butanol. Survival of 0% was recorded when no colonies in the spots of the undiluted or any of the serial dilutions were observed. The results are shown in Table 10.
TABLE-US-00010 TABLE 10 Survival of log-phase E. coli in 1-butanol at 42° C., 28° C., or 0° C. Grown at Grown at Grown at Grown at Grown at Grown at 42° C. 28° C. 42° C. 28° C. 42° C. 29° C. % survival after % survival after % survival after 1-Butanol 30 min exposure 30 min exposure 30 min exposure % (w/v) at 42° C. at 28° C. at 0° C. 1.0 100 100 .sup. nt1 nt nt nt 1.5 0 0 100 100 nt nt 2.0 0 0 100 100 nt nt 2.5 0 0 0.1 50 100 100 3.0 0 0 0 0 100 100 3.5 0 0 0.01 0 100 100 4.0 0 0 0.001 0 100 100 4.5 nt nt 0 0 100 100 5.0 nt nt nt nt 10 50 6.0 nt nt nt nt 1 1 1"nt" = not tested
[0224] For both the stationary phase and log-phase cultures of E. coli MG1655, the growth temperature had very little, if any, effect on the survival of a 1-butanol shock. However, the exposure temperature had a major effect on the survival of E. coli to 1-butanol shock. As can be seen from the data in Tables 9 and 10, the tolerance of E. coli MG1655 to 1-butanol increased with decreasing exposure temperature.
Example 3
Increased Tolerance of Escherichia coli to 2-Butanone at Decreased Exposure Temperature
[0225] The effect of exposure temperature on survival of Escherichia coli in the presence of 2-butanone (also referred to herein as methyl ethyl ketone or MEK) was tested as follows. E. coli strain BW25113 (The Coli Genetic Stock Center (CGSC), Yale University; #7636) was grown overnight in LB medium (Teknova, Half Moon Bay, Calif.) with shaking at 250 rpm at 37° C. Survival of MEK shock was tested at exposure temperatures of 28° C. or 37° C. A series of solutions of MEK at different concentrations in LB medium was made and 90 μL aliquots were put in microfuge tubes. To these were added 10 μL of the overnight culture and the tubes were immediately placed in shaking incubators at 37° C. or 28° C. for 30 min. To stop the effect of MEK on the cultures, a 10-2 dilution was done by placing 2 μL of the MEK treated culture into 198 μL of LB medium in wells of a microplate. Then 5 μL of the undiluted treated cultures were spotted on LB agar plates. Subsequent 10-fold serial dilutions of 10-3, 10-4, 10-6 and 10-6 of the exposed cultures were done by serial pipetting of 20 μL, starting with the 10-2 dilution cultures, into 180 μL of LB medium in the microplate, using a multi-channel pipette. Prior to each transfer, the cultures were mixed by pipetting up and down six times. Each dilution (5 μL) was spotted onto LB plates using a multi-channel pipette and allowed to soak into the plate. The plates were inverted and incubated overnight at 37° C. The number of colonies for each dilution was counted and the % growth inhibition was calculated by comparison with a control culture that had not been exposed to MEK. Survival of 0% was recorded when no colonies in the spots of the undiluted or any of the serial dilutions were observed. The results, given as the average of duplicate experiments, are shown in Table 11.
TABLE-US-00011 TABLE 11 Survival of E. coli in MEK at 37° C. and 28° C. MEK % w/v % Survival at 37° C. % Survival at 28° C. 0 100 100 4 100 100 6 0 100 8 0 0.002
[0226] Reducing the exposure temperature from 37° C. to 28° C. dramatically improved survival of E. coli to MEK treatment. At 37° C. there was full survival at 4% w/v and no survival at 6% w/v, while at 28° C. there was full survival at 6% w/v. Thus, the tolerance of E. coli to MEK increased with decreasing exposure temperature.
Example 4
[0227] Increased tolerance of E. coli and L. plantarum PN0512 to 1-Butanol at Decreased Exposure Temperature
[0228] This Example demonstrates that the toxic effects of 1-butanol and 2-butanol on various microbial cells was reduced at lower temperatures. This was demonstrated by incubating E. coli (strain MG1655; ATCC #700926), and L. plantarum (strain PN0512; ATCC #PTA-7727) with either 1-butanol or 2-butanol at different temperatures and then determining the fraction of the cells that survived the treatment at the different temperatures.
[0229] Using overnight cultures or cells from plates, 30 mL cultures of the microorganisms to be tested were started in the following culture media:
[0230] E. coli--Miller's LB medium (Teknova, Half Moon Bay, Calif.):
[0231] L. plantarum PN0512--Lactobacilli MRS Broth (BD Diagnostic Systems, Sparks, Md.). The E. coli and L. plantarum cultures were grown at 37° C. aerobically with shaking until the cultures were in log phase and the OD600 was between 0.6 and 0.8. A 50 μL aliquot of each culture was removed for a time zero sample. The remainder of the cultures was divided into six 5 mL portions and placed in six small incubation flasks or tubes. Different amounts of 1-butanol or 2-butanol were added to the six flasks to bring the concentration to predetermined values, as listed in the tables below. The flasks or tubes were incubated at a desired temperature, aerobically without shaking for 1 h. After the incubation with one of the butanols, 2 μL from each of the flasks (and in addition 2 μL of the time zero sample of the culture before exposure to one of the butanols) were pipetted into the "head" wells of a 96 well (8×12) microtiter plate, each containing 198 μL of LB medium to give a 10-2 dilution of the culture. Subsequently, 10-3, 10-4, 10-5, and 10-6 serial dilutions of the cultures were prepared as follows. The 10-3 dilution was prepared by pipetting 20 μL of the sample from the head well into the 180 μL LB medium in the next well using a multi-channel pipette. This procedure was repeated 3 more times on successive wells to prepare the 10-4, 10-5, and 10-6 dilutions. After each liquid transfer, the solution in the well was mixed by pipetting it up and down 10 times with the multi-channel pipetor. A 5 μL aliquot of each dilution was spotted onto an LB plate using a multi-channel pipette starting with the 10-6 dilution, then the 10-6, and so on working from more to less dilute without a change of tips. The spots were allowed to soak into the agar by leaving the lid of the plate slightly open for 15 to 30 min in a sterile transfer hood. The plates were covered, inverted, and incubated overnight at 37° C. The following day, the number of colonies in the spots were counted from the different dilutions. The number of living cells/mL in each of the original culture solutions from which the 2 μL was withdrawn was calculated and compared to the number of cells in the control untreated culture to determine the % of the cells surviving.
[0232] The results of experiments in which E. coli cells were treated with 1-butanol at temperatures of 0, 30, and 37° C. are shown Table 12.
TABLE-US-00012 TABLE 12 Percentage of E. coli cells surviving in 1-butanol at 0, 30 and 37° C. 1-butanol % Survival % v/v at 0° C. % Survival at 30° C. % Survival at 37° C. 0 100 100 100 1 nt1 100 72 1.5 nt 100 20 2 nt 100 0 2.5 100 23 0 3 100 0 0 3.5 100 0 nt 4 100 nt nt 4.5 100 nt nt 1"nt" = not tested
[0233] The concentration at which 1-butanol kills E. coli cells was affected by the treatment temperature. At 0° C., concentrations of 1-butanol as high as 4.5% v/v had no toxic effect on E. coli cells during a one hour treatment. At 30° C., E. coli cells were killed when treated with 3% v/v 1-butanol for one hour. At 37° C., E. coli cells were killed when treated with 2% v/v 1-butanol for one hour.
[0234] The results of experiments in which L. plantarum PN0512 cells were treated with 1-butanol at temperatures of 0, 23, and 37° C. for one hour are shown Table 13.
TABLE-US-00013 TABLE 13 Percentage of L. plantarum PN0512 cells surviving in 1-butanol at 0, 23 and 37° C. 1-butanol % Survival % v/v at 0° C. % Survival at 23° C. % Survival at 37° C. 0 100 100 100 1 nt1 nt 80 1.5 nt nt 58 2 nt 100 29 2.5 nt 100 8 3 100 82 0 3.5 100 0 0 4 100 0 nt 4.5 100 0 nt 5 0 nt nt 5.5 0 nt nt 1"nt" = not tested
[0235] The concentration at which 1-butanol kills L. plantarum PN0512 cells was affected by the treatment temperature. At 0° C., concentrations of 1-butanol as high as 4.5% v/v had no toxic effect on L. plantarum PN0512 cells during a one hour treatment. At 23° C., L. plantarum PN0512 cells were killed when treated with 3.5% v/v 1-butanol for one hour. At 37° C., L. plantarum PN0512 cells were killed when treated with 2.5% v/v 1-butanol for one hour.
Example 5
Cloning and Expression of Acetolactate Synthase
[0236] The purpose of this Example was to clone and express in E. coli the budB gene that encodes the enzyme acetolactate synthase. The budB gene was amplified from Klebsiella pneumoniae strain ATCC 25955 genomic DNA using PCR.
[0237] The budB sequence which encodes acetolactate synthase was amplified from Klebsiella pneumoniae (ATCC 25955) genomic DNA by PCR using the primer pair B1 (SEQ ID NO:15) and B2 (SEQ ID NO:16). Other PCR amplification reagents (e.g. Kod HiFi DNA Polymerase (Novagen Inc., Madison, Wis.; catalog no. 71805-3)) were supplied in manufacturers' kits and used according to the manufacturer's protocol. Klebsiella pneumoniae genomic DNA was prepared using the Gentra Puregene Puregene kit (Gentra Systems, Inc., Minneapolis, Minn.; catalog number D-5000A). Amplification was carried out in a DNA Thermocycler GeneAmp 9700 (PE Applied Biosystems, Foster city, CA). The nucleotide sequence of the open reading frame (ORF) and the predicted amino acid sequence of the enzyme are given as SEQ ID NO:3 and SEQ ID NO:4, respectively.
[0238] For expression studies the Gateway cloning technology (Invitrogen Corp., Carlsbad, Calif.) was used. The entry vector pENTR/SD/D-TOPO allows directional cloning and provided a Shine-Dalgarno sequence for the gene of interest. The destination vector pDEST14 used a T7 promoter for expression of the gene with no tag. The forward primer incorporated four bases (CACC) immediately adjacent to the translational start codon to allow directional cloning of the budB acetolactate synthase coding region PCR product into pENTR/SD/D-TOPO (Invitrogen), generating the plasmid pENTRSDD-TOPObudB. The pENTR construct was transformed into E. coli Top10 (Invitrogen) cells and plated according to the manufacturer's recommendations. Transformants were grown overnight and plasmid DNA was prepared using the QIAprep Spin Miniprep kit (Qiagen, Valencia, Calif.; catalog no. 27106) according to the manufacturer's recommendations. To create an expression clone, the budB coding region from pENTRSDD-TOPObudB was transferred to the pDEST 14 vector by in vitro recombination using the LR Clonase mix (Invitrogen, Corp., Carlsbad, Calif.). The resulting vector, pDEST14budB, was transformed into BL-21-Al cells (Invitrogen Corp.). BL-21-Al cells carry a chromosomal copy of the T7 RNA polymerase under control of the arabinose-inducible araBAD promoter.
[0239] Transformants are inoculated into LB medium supplemented with 50 μg/mL of ampicillin and grown overnight. An aliquot of the overnight culture is used to inoculate 50 mL of LB medium supplemented with 50 μg/mL of ampicillin. The culture is incubated at 37° C. with shaking until the OD600 reaches 0.6-0.8. The culture is split into two 25-mL portions and arabinose is added to one of the flasks to a final concentration of 0.2% w/v. The negative control flask is not induced with arabinose. The flasks are incubated for 4 h at 37° C. with shaking. Cells are harvested by centrifugation and the cell pellets are resuspended in 50 mM MOPS, pH 7.0 buffer. The cells are disrupted either by sonication or by passage through a French Pressure Cell. Each cell lysate is centrifuged yielding the supernatant and the pellet or the insoluble fraction. An aliquot of each fraction (whole cell lysate, from induced and control cells, is resuspended in SDS (MES) loading buffer (Invitrogen), heated to 85° C. for 10 min and subjected to SDS-PAGE analysis (NuPAGE 4-12% Bis-Tris Gel, catalog no. NP0322Box, Invitrogen). A protein of the expected molecular weight, as deduced from the nucleic acid sequence, is present in the induced culture but not in the uninduced control.
[0240] Acetolactate synthase activity in the cell free extracts is measured using the method described by Bauerle et al. (Bauerle et al. (1964) Biochim. Biophys. Acta 92:142-149). Protein concentration is measured by either the Bradford method or by the Bicinchoninic Kit (Sigma, catalog no. BCA-1; St. Louis, Mo.) using Bovine serum albumin (BSA) (Bio-Rad, Hercules, Calif.) as the standard.
Example 6
Cloning and Expression of Acetolactate Decarboxylase
[0241] The purpose of this Example was to clone and express in E. coli the budA gene that encodes the enzyme acetolactate decarboxylase. The budA gene was amplified from Klebsiella pneumoniae strain ATCC 25955 genomic DNA using PCR.
[0242] The budA sequence which encodes acetolactate decarboxylase, was cloned in the same manner as described for budB in Example 5, except that the primers used for PCR amplification were B3 (SEQ ID NO:17) and B4 (SEQ ID NO:18). The nucleotide sequence of the open reading frame (ORF) and the predicted amino acid sequence of the enzyme are given as SEQ ID NO:1 and SEQ ID NO:2, respectively. The resulting plasmid was named pENTRSDD-TOPObudA.
[0243] Acetolactate decarboxylase activity in the cell free extracts is measured using the method described by Bauerle et al., supra.
Example 7 (Prophetic)
Cloning and Expression of Butanediol Dehydrogenase
[0244] The purpose of this prophetic Example is to describe how to clone and express in E. coli the budC gene that encodes the enzyme butanediol dehydrogenase. The budC gene is amplified from Klebsiella pneumoniae strain IAM1063 genomic DNA using PCR.
[0245] The budC sequence encoding butanediol dehydrogenase is cloned and expressed in the same manner as described for budA in Example 5, except that the primers used for PCR amplification are B5 (SEQ ID NO:19) and B6 (SEQ ID NO:20) and the genomic template DNA is from Klebsiella. pneumoniae IAM1063 (which is obtained from the Institute of Applied Microbiology Culture Collection, Tokyo, Japan). Klebsiella pneumoniae IAM1063 genomic DNA is prepared using the Gentra Puregene Puregene kit (Gentra Systems, Inc., Minneapolis, Minn.; catalog number D-5000A). The nucleotide sequence of the open reading frame (ORF) and the predicted amino acid sequence of the enzyme are given as SEQ ID NO:5 and SEQ ID NO:6, respectively.
[0246] Butanediol dehydrogenase activity in the cell free extracts is measured spectrophotometrically by following NADH consumption at an absorbance of 340 nm.
Example 8 (Prophetic)
Cloning and Expression of Butanediol Dehydratase
[0247] The purpose of this prophetic Example is to describe how to clone and express in E. coli the pddA, pddB and pddC genes that encode butanediol dehydratase. The pddA, pddB and pddC genes are amplified from Klebsiella oxytoca ATCC 8724 genomic DNA using PCR.
[0248] The pddA, pddB and pddC sequences which encode butanediol dehydratase are cloned and expressed in the same manner as described for budA in Example 5, except that the genomic template DNA is from Klebsiella oxytoca ATCC 8724, and the primers are B7 (SEQ ID NO:21) and B8 (SEQ ID NO:22). Klebsiella oxytoca genomic DNA is prepared using the Gentra Puregene Puregene kit (Gentra Systems, Inc., Minneapolis, Minn.; catalog number D-5000A). A single PCR product including all three open reading frames (ORFs) is cloned, so that all three coding regions are expressed as an operon from a single promoter on the expression plasmid. The nucleotide sequences of the open reading frames for the three subunits are given as SEQ ID NOs:7, 9, and 11, respectively, and the predicted amino acid sequences of the three enzyme subunits are given as SEQ ID NOs:8, 10, and 12, respectively.
[0249] Butanediol dehydratase activity in the cell free extracts is measured by derivatizing the ketone product with 2,4-dinitrophenylhydrazine (DNPH). Briefly, 100 μL of reaction mixture, cell extract containing approximately 0.0005 units of enzyme, 40 mM potassium phosphate buffer (pH 8.0), 2 μg of adenosylcobalamin, 5 μg of 2,3,-butanediol, and 1 μg of bovine serum albumin, is quenched by addition of an equal volume of 0.05 wt % DNPH in 1.0 N HCl. After 15 min at room temperature, the color is developed by addition of 100 μL of 4 N NaOH. The amount of product is determined from the absorbance of the final solution at 550 nm compared to a standard curve prepared with 2-butanone. All reactions are carried out at 37° C. under dim red light.
Example 9 (Prophetic)
Cloning and Expression of Butanol Dehydrogenase
[0250] The purpose of this prophetic Example is to describe how to clone and express in E. coli the sadh gene that encodes butanol dehydrogenase. The sadh gene is amplified from Rhodococcus ruber strain 219 genomic DNA using PCR.
[0251] The sadh sequence encoding butanol dehydrogenase is cloned and expressed in the same manner as described for budA in Example 5, except that the genomic template DNA is from Rhodococcus ruber strain 219 (Meens, Institut fuer Mikrobiologie, Universitaet Hannover, Hannover, Germany) and the primers are B9 (SEQ ID NO:23) and B10 (SEQ ID NO:24). Rhodococcus ruber genomic DNA is prepared using the Ultra Clean® Microbial DNA Isolation Kit (MO BIO Laboratories Inc., Carlsbad, Calif.), according to the manufacturer's protocol. The nucleotide sequence of the open reading frame (ORF) and the predicted amino acid sequence of the enzyme are given as SEQ ID NO:13 and SEQ ID NO:14, respectively.
[0252] Butanol dehydrogenase activity in cell free extracts is measured by following the increase in absorbance at 340 nm resulting from the conversion of NAD to NADH when the enzyme is incubated with NAD and 2-butanol.
Example 10 (Prophetic)
Construction of a Transformation Vector for the Genes in a 2-Butanol Biosynthetic Pathway
[0253] The purpose of this prophetic Example is to describe the preparation of a transformation vector for the genes in a 2-butanol biosynthetic pathway (i.e., Pathway 3 as described above). Like most organisms, E. coli converts glucose initially to pyruvic acid. The enzymes required to convert pyruvic acid to 2-butanol following Pathway 3, i.e., acetolactate synthase, acetolactate decarboxylase, butanediol dehydrogenase, butanediol dehydratase, and butanol dehydrogenase, are encoded by the budA, budB, budC, pddA, pddB, pddC and sadh genes. To simplify building the 2-butanol biosynthetic pathway in a recombinant organism, the genes encoding the 5 steps in the pathway are divided into two operons. The upper pathway comprises the first three steps catalyzed by acetolactate synthase, acetolactate decarboxylase, and butanediol dehydrogenase. The lower pathway comprises the last two steps catalyzed by butanediol dehydratase and butanol dehydrogenase.
[0254] The coding sequences are amplified by PCR with primers that incorporate restriction sites for later cloning, and the forward primers contain an optimized E. coli ribosome binding site (AAAGGAGG). PCR products are TOPO cloned into the pCR4Blunt-TOPO vector and transformed into Top10 cells (Invitrogen). Plasmid DNA is prepared from the TOPO clones, and the sequence of the cloned PCR fragment is verified. Restriction enzymes and T4 DNA ligase (New England Biolabs, Beverly, Mass.) are used according to manufacturer's recommendations. For cloning experiments, restriction fragments are gel-purified using QIAquick Gel Extraction kit (Qiagen).
[0255] After confirmation of the sequence, the coding regions are subcloned into a modified pUC19 vector as a cloning platform. The pUC19 vector is modified by a HindIII/Sapl digest, followed by treatment with Klenow DNA polymerase to fill in the ends. The 2.4 kB vector fragment is gel-purified and religated creating pUC19dHS. Alternatively the pUC19 vector is modified by a Sphl/Sapl digest, followed by treatment with Klenow DNA polymerase to blunt the ends. The 2.4 kB vector fragment is gel-purified and religated creating pUC19dSS. The digests remove the lac promoter adjacent to the MCS (multiple cloning sites), preventing transcription of the operons from the vector.
Upper Pathway:
[0256] The budABC coding regions are amplified from Klebsiella pneumoniae genomic DNA by PCR using primer pair B11 and B12 (Table 4), given as SEQ ID NOs:25 and 26, respectively. The forward primer incorporates an EcoRI restriction site and a ribosome binding site (RBS). The reverse primer incorporates an Sphl restriction site. The PCR product is cloned into pCR4 Blunt-TOPO creating pCR4Blunt-TOPO-budABC.
[0257] To construct the upper pathway operon pCR4Blunt-TOPO-budABC is digested with EcoRI and Sphl releasing a 3.2 kbp budABC fragment. The pUC19dSS vector is also digested with EcoRI and Sphl, releasing a 2.0 kbp vector fragment. The budABC fragment and the vector fragment are ligated together using T4 DNA ligase (New England Biolabs) to form pUC19dSS-budABC.
Lower Pathway:
[0258] The pddABC coding regions are amplified from Klebsiella oxytoca ATCC 8724 genomic DNA by PCR using primers B13 and B14 (Table 4), given as SEQ ID NOs:27 and 28, respectively, creating a 2.9 kbp product. The forward primer incorporates EcoRI and Pmel restriction sites and a RBS. The reverse primer incorporates the BamHI restriction site. The PCR product is cloned into pCRBlunt II-TOPO creating pCRBluntll-pdd.
[0259] The sadh gene is amplified from Rhodococcus ruber strain 219 genomic DNA by PCR using primers B15 and B16 (Table 4), given as SEQ ID NOs:29 and 30, respectively, creating a 1.0 kbp product. The forward primer incorporates a BamHI restriction site and a RBS. The reverse primer incorporates an XbaI restriction site. The PCR product is cloned into pCRBlunt II-TOPO creating pCRBluntll-sadh.
[0260] To construct the lower pathway operon, a 2.9 kbp EcoRI and BamHI fragment from pCRBluntll-pdd, a 1.0 kbp BamHI and XbaI fragment from pCRBluntll-sadh, and the large fragment from an EcoRI and XbaI digest of pUC19dHS are ligated together. The three-way ligation creates pUC19dHS-pdd-sadh.
[0261] The pUC19dSS-budABC vector is digested with Pmel and HindIII, releasing a 3.2 kbp fragment that is cloned into pBenBP, an E. coli-B. subtilis shuttle vector. Plasmid pBenBP is created by modification of the pBE93 vector, which is described by Nagarajan (WO 93/2463, Example 4). To generate pBenBP, the Bacillus amyloliquefaciens neutral protease promoter (NPR) signal sequence and the phoA gene are removed from pBE93 with an NcoI/HindIII digest. The NPR promoter is PCR amplified from pBE93 by primers BenF and BenBPR, given by SEQ ID NOs:31 and 32, respectively. Primer BenBPR incorporates BstEII, Pmel and HindIII sites downstream of the promoter. The PCR product is digested with NcoI and HindIII, and the fragment is cloned into the corresponding sites in the vector pBE93 to create pBenBP. The upper operon fragment is subcloned into the Pmel and HindIII sites in pBenBP creating pBen-budABC.
[0262] The pUC19dHS-pdd-sadh vector is digested with Pmel and HindIII releasing a 3.9 kbp fragment that is cloned into the Pmel and HindIII sites of pBenBP, creating pBen-pdd-sadh.
Example 11 (Prophetic)
[0263] Expression of a 2-Butanol Biosynthetic Pathway in E. coli
[0264] The purpose of this prophetic Example is to describe how to express a 2-butanol biosynthetic pathway in E. coli.
[0265] The plasmids pBen-budABC and pBen-pdd-sadh, prepared as described in Example 10, are separately transformed into E. coli NM522 (ATCC No. 47000), and expression of the genes in each operon is monitored by SDS-PAGE analysis and enzyme assay. After confirmation of expression of all genes, pBen-budABC is digested with EcoRI and HindIII to release the NPR promoter-budABC fragment. The fragment is blunt ended using the Klenow fragment of DNA polymerase (New England Biolabs, catalog no. MO210S). The plasmid pBen-pdd-sadh is digested with EcoRI and similarly blunted to create a linearized, blunt-ended vector fragment. The vector and NPR-budABC fragments are ligated, creating p2BOH. This plasmid is transformed into E. coli NM522 to give E. coli NM522/p2BOH, and expression of the genes is monitored as previously described.
[0266] E. coli NM522/p2BOH is inoculated into a 250 mL shake flask containing 50 mL of medium and shaken at 250 rpm and 35° C. The medium is composed of: dextrose, 5 g/L; MOPS, 0.05 M; ammonium sulfate, 0.01 M; potassium phosphate, monobasic, 0.005 M; S10 metal mix, 1% (v/v); yeast extract, 0.1% (w/v); casamino acids, 0.1% (w/v); thiamine, 0.1 mg/L; proline, 0.05 mg/L; and biotin 0.002 mg/L, and is titrated to pH 7.0 with KOH. S10 metal mix contains: MgCl2, 200 mM; CaCl2, 70 mM; MnCl2, 5 mM; FeCl3, 0.1 mM; ZnCl2, 0.1 mM; thiamine hydrochloride, 0.2 mM; CuSO4, 172 μM; CoCl2, 253 μM; and Na2MoO4, 242 μM. After 18 h, 2-butanol is detected by HPLC or GC analysis using methods that are well known in the art, for example, as described in the General Methods section above.
Example 12 (Prophetic)
Expression of a 2-Butanol Biosynthetic Pathway in Bacillus subtilis
[0267] The purpose of this prophetic Example is to describe how to express a 2-butanol biosynthetic pathway in Bacillus subtilis.
[0268] The plasmids pBen-budABC and pBen-pdd-sadh, prepared as described in Example 10, are separately transformed into Bacillus subtilis BE1010 (J. Bacteriol. 173:2278-2282 (1991)) and expression of the genes in each operon is monitored as described in Example 11. The plasmid pBen-budABC is digested with EcoRI and HindIII to release the NPR promoter-budABC fragment. The fragment is blunt ended using the Klenow fragment of DNA polymerase (New England Biolabs, catalog no. MO210S). The plasmid pBen-pdd-sadh is digested with EcoRI and similarly blunted to create a linearized, blunt-ended vector fragment. The vector and NPR-budABC fragments are ligated, creating p2BOH. This plasmid is transformed into Bacillus subtilis BE1010 to give Bacillus subtilis BE1010/p2BOH, and expression of the genes is monitored as previously described.
[0269] Bacillus subtilis BE1010/p2BOH is inoculated into a 250 mL shake flask containing 50 mL of medium and shaken at 250 rpm and 35° C. for 18 h. The medium is composed of: dextrose, 5 g/L; MOPS, 0.05 M; glutamic acid, 0.02 M; ammonium sulfate, 0.01 M; potassium phosphate, monobasic buffer, 0.005 M; S10 metal mix (as described in Example 11), 1% (v/v); yeast extract, 0.1% (w/v); casamino acids, 0.1% (w/v); tryptophan, 50 mg/L; methionine, 50 mg/L; and lysine, 50 mg/L, and is titrated to pH 7.0 with KOH. After 18 h, 2-butanol is detected by HPLC or GC analysis using methods that are well known in the art, for example, as described in the General Methods section above.
Example 13
Construction of a Transformation Vector for the Genes in a 2-Butanol Biosynthetic Pathway
[0270] The purpose of this Example was to prepare a recombinant E. coli host carrying the genes in a 2-butanol biosynthetic pathway (i.e., Pathway 3 as described above). Like most organisms, E. coli converts glucose initially to pyruvic acid. The enzymes required to convert pyruvic acid to 2-butanone in Pathway 3, i.e., acetolactate synthase, acetolactate decarboxylase, butanediol dehydrogenase, and butanediol dehydratase are encoded by the budA, budB, budC, pddA, pddB, and pddC genes. In the last step of the pathway, a butanol dehydrogenase converts 2-butanone to 2-butanol. Dehydrogenases that carry out this last step are promiscuous and may be found in many organisms. To simplify building the 2-butanol biosynthetic pathway in a recombinant organism, the genes encoding the 5 steps in the pathway were divided into multiple operons. The upper pathway operon comprised the first three steps catalyzed by acetolactate synthase, acetolactate decarboxylase, and butanediol dehydrogenase and were cloned onto an expression vector. The lower pathway comprised the last two steps catalyzed by butanediol dehydratase including the reactivating factor (Mori et al., J. Biol. Chem. 272:32034 (1997)) and a butanol dehydrogenase. The diol dehydratase can undergo suicide inactivation during catalysis. The reactivating factor protein encoded by ddrA and ddrB (GenBank AF017781, SEQ ID NO:70) reactivates the inactive enzyme. The ddrA and ddrB genes flank the diol dehydratase operon. The operons for the dehydratase/reactivating factor and the butanol dehydrogenase were either cloned onto another expression vector or the dehydratase/reactivating factor operon was cloned singly onto another expression vector and the last step was provided by an endogenous activity in the demonstration host.
[0271] Construction of Vector pTrc99a-budABC:
[0272] The budAB coding regions were amplified from K. pneumoniae ATCC 25955 genomic DNA by PCR using primer pair BABC F and BAB R, given as SEQ ID NOs:33 and 34, respectively (see Table 4), creating a 2.5 kbp product. The forward primer incorporated SacI and EcoRI restriction sites and a ribosome binding site (RBS). The reverse primer incorporated a SpeI restriction site. The PCR product was cloned into pCR4Blunt-TOPO creating pCR4 Blunt-TOPO-budAB. Plasmid DNA was prepared from the TOPO clones and the sequence of the genes was verified with primers M13 Forward (SEQ ID NO:35), M13 Reverse (SEQ ID NO:36), N83 SeqF2 (SEQ ID NO:37), N83 SeqF3 (SEQ ID NO:38) and N84 SeqR4 (SEQ ID NO:39) (see Table 5).
[0273] The budC coding region was amplified from K. pneumoniae ATCC 25955 genomic DNA by PCR using primer pair BC Spe F and BC Xba R given as SEQ ID NOs:40 and 41, respectively, creating a 0.8 kbp product. The forward primer incorporated a SpeI restriction site, a RBS and modified the CDS by changing the second and third codons from AAA to AAG. The reverse primer incorporated an XbaI restriction site. The PCR product was cloned into pCR4 Blunt-TOPO creating pCR4 Blunt-TOPO-budC. Plasmid DNA was prepared from the TOPO clones and the sequence of the genes was verified with primers M13 Forward (SEQ ID NO:35) and M13 Reverse (SEQ ID NO:36).
[0274] To construct the budABC operon, pCR4 Blunt-TOPO-budC was digested with SnaBI and XbaI releasing a 1.0 kbp budC fragment. The vector pTrc99a (Amann et al., Gene 69(2):301-315 (1988)) was digested with SmaI and XbaI creating a 4.2 kbp linearized vector fragment. The vector and the budC fragment were ligated to create pTrc99a-budC and transformed into E. coli Top 10 cells (Invitrogen). Transformants were analyzed by PCR amplification with primers Trc F (SEQ ID NO:42) and Trc R (SEQ ID NO:43) for a 1.2 kbp product to confirm the presence of the budC insert. The budAB genes were subcloned from pCR4 Blunt-TOPO-budAB as a 2.5 kbp EcoRI/SpeI fragment. Vector pTrc99a-budC was digested with EcoRI and SpeI and the resulting 5.0 kbp vector fragment was gel-purified. The purified vector and budAB insert were ligated and transformed into E. coli Top 10 cells. Transformants were screened by PCR amplification with primers Trc F (SEQ ID NO:42) and N84 Seq R2 (SEQ ID NO:65) to confirm creation of pTrc99a-budABC. In this plasmid, the bud A, B, and C coding regions are adjacent to each other, in this order, and between the Trc promoter and the rrnB termination sequence.
Results:
[0275] Three independent isolates of E. coli Top 10/pTrc99a-budABC were examined for the production of butanediol, using E. coli Top 10/pCL1925-Kodd-ddr (described below) as a negative control. The strains were grown in LB medium containing 100 μg/mL carbenicillin. The resulting cells were used to inoculate shake flasks (approximately 175 mL total volume) containing 125 mL of TM3a/glucose medium with 100 μg/mL carbenicillin. In addition, the flasks inoculated with strains carrying pTrc99a-budABC contained 0.4 mM isopropyl β--D-1-thiogalactopyranoside (IPTG). TM3a/glucose medium contains (per liter): 10 g glucose, 13.6 g KH2PO4, 2.0 g citric acid monohydrate, 3.0 g (NH4)2SO4, 2.0 g MgSO4.7H2O, 0.2 g CaCl2.2H2O, 0.33 g ferric ammonium citrate, 1.0 mg thiamine .HCl, 0.50 g yeast extract, and 10 mL trace elements solution, adjusted to pH 6.8 with NH4OH. The solution of trace elements contained: citric acid .H2O (4.0 g/L), MnSO4.H2O (3.0 g/L), NaCl (1.0 g/L), FeSO4.7H2O (0.10 g/L), CoCl2.6H2O (0.10 g/L), ZnSO4.7H2O (0.10 g/L), CuSO4.5H2O (0.010 g/L), H3BO3 (0.010 g/L), and Na2MoO4.2H2O (0.010 g/L). The flasks, capped with vented caps, were inoculated at a starting OD600 of approximately 0.03 units and incubated at 34° C. with shaking at 300 rpm.
[0276] Approximately 23 h after induction, an aliquot of the broth was analyzed by HPLC (Shodex Sugar SH1011 column) and GC (HP-INNOWax), using the same methods described in the General Methods section for 2-butanol and 2-butanone. The results of the analysis are given in Table 14. The three E. coli clones converted glucose to acetoin and meso-2,3-butanediol, the desired intermediates of the pathway, with a molar selectivity of 14%. This selectivity was approximately 35-fold higher than that observed with the E. coli control strain lacking budABC.
TABLE-US-00014 TABLE 14 Production of Acetoin and meso-2,3-butanediol by E. coli Top 10/pTrc99a-budABC Meso-2,3- Molar Strain OD600 Acetoin, mM Butanediol, mM Selectivitya, % Negative 1.4 0.07 0.03 0.4 control Isolate #1 1.5 0.64 1.3 14 Isolate #2 1.4 0.70 1.2 14 Isolate #3 1.4 0.74 1.3 15 aMolar selectivity is (acetoin + meso-2,3-butanendiol)/(glucose consumed).
Construction of Vector DCL1925-KoDD-ddr:
[0277] The diol dehydratase (GenBank D45071, SEQ ID NO:69) and reactivating factor (GenBank AF017781, SEQ ID NO:70) operons were PCR amplified from Klebsiella oxytoca ATCC 8724 as a single unit with primers DDo For (SEQ ID NO: 44) and DDo Rev (SEQ ID NO:45). The forward primer incorporated an optimized E. coli RBS and a HindIII restriction site. The reverse primer included an XbaI restriction site. The 5318 bp PCR product was cloned into pCR4Blunt-TOPO and clones of the resulting pCR4Blunt-TOPO-Kodd-ddr were sequenced with primers M13 Forward (SEQ ID NO:35), M13 Reverse (SEQ ID NO:36), DDko seq F2 (SEQ ID NO:46), DDko seq F5 (SEQ ID NO:47), DDko seq F7 (SEQ ID NO:48), DDko seq F9 (SEQ ID NO:49), DDko seq R1 (SEQ ID NO:50), DDko seq R3 (SEQ ID NO:51), DDko seq R7 (SEQ ID NO:52), and DDko seq R10 (SEQ ID NO:53). A clone having the insert with the expected sequence was identified.
[0278] For expression, the diol dehydratase/reactivating factor genes were subcloned into pCL1925 (U.S. Pat. No. 7,074,608), a low copy plasmid carrying the glucose isomerase promoter from Streptomcyes. pCR4Blunt-TOPO-Kodd-ddr was digested with HindIII and XbaI and the resulting 5.3 kbp Kodd-ddr fragment was gel-purified. Vector pCL1925 was digested with HindIII and XbaI and the resulting 4539 bp vector fragment was gel purified. The vector and Kodd-ddr fragment were ligated and transformed into E. coli Top10. Transformants were screened by PCR with primers DDko Seq F7 (SEQ ID NO:48) and DDko Seq R7 (SEQ ID NO: 52). Amplification of the plasmid (pCL1925-Kodd-ddr) carrying the insert resulted in a product of approximately 797 bp.
[0279] Activity of diol dehydratase towards meso-2,3-butanediol was measured by incubating cell extract (total protein ˜0.8 mg/mL) with 10 mM butanediol and 12 mM coenzyme B12 in 80 mM HEPES (pH 8.2) for 17 h at room temperature. Formation of the expected product, 2-butanone, was determined by HPLC as described in the General Methods.
Construction of Vector pCL1925-KoDD-ddr::T5 chnA ter:
[0280] To provide a heterologous alcohol dehydrogenase activity, the chnA gene encoding cyclohexanol dehydrogenase from Acinetobacter sp. (Cheng et al., J. Bacteriol. 182:4744-4751 (2000)) was cloned into the pCL1925 vector with the diol dehydratase operon, pCL1925-Kodd-ddr. The chnA gene, given as SEQ ID NO:71 (Genbank No: AF282240, SEQ ID NO:73) was amplified from pDCQ2, a cosmid carrying the cyclohexanol gene cluster from Acinetobacter, with primers ChnA F (SEQ ID NO:54) and ChnA R (SEQ ID NO:55). The resulting 828 bp PCR product was cloned into pCR4Blunt-TOPO to create pCR4Blunt-TOPO-chnA and transformants were screened by colony PCR with primers M13 Forward (SEQ ID NO:35) and M13 Reverse (SEQ ID NO:36). Correct clones produced a PCR product of about 1 kbp and were sequenced with primers M13 Forward (SEQ ID NO:35) and M13 Reverse (SEQ ID NO:36).
[0281] After sequencing pCR4Blunt-TOPO-chnA to confirm the correct sequence, the chnA gene was subcloned from the plasmid as an 813 bp MfeI/SmaI fragment. The expression vector pQE30 (Qiagen) was digested with MfeI and SmaI and the resulting 3350 bp vector fragment was gel-purified. The chnA fragment and the purified vector were ligated and transformed into E. coli Top10 cells. Transformants were colony PCR screened with primers chnSeq F1 (SEQ ID NO:56) and chnseq R1 (SEQ ID NO:57) for a 494 bp PCR product. This cloning placed the chnA gene under the control of the T5 promoter in the plasmid, pQE30-chnA.
[0282] To prepare the pCL1925 vector to carry two operons, terminators were added to the vector. A tonB terminator-mcs-trpA terminator fragment was prepared by oligonucleotide annealing with primers Top ter F1 (SEQ ID NO:58), Top ter F2 (SEQ ID NO:59), Bot ter R1 (SEQ ID NO:60) and Bot ter R2 (SEQ ID NO:61). The annealed DNA was gel-purified on a 6% PAGE gel (Embi-tec, San Diego, Calif.). Vector pCL1925 was digested with SacI and XbaI and gel-purified. The annealed DNA and vector fragment were ligated to create pCL1925-ter. Transformants were screened by colony PCR amplification with primers pCL1925 vec F (SEQ ID NO:62) and pCL1925 vec R1 (SEQ ID NO:63) for the presence of a PCR product of approximately 400 bp. Positive clones from the PCR screen were sequenced with the same primers.
[0283] Vector pCL1925-ter was digested with XhoI and PmeI and the resulting 4622 bp fragment was gel-purified. pQE30-chnA was digested with NcoI and the DNA was treated with Klenow DNA polymerase to blunt the ends. pQE30-chnA was then digested with XhoI and the resulting 1.2 kbp T5 promoter-chnA fragment was gel-purified. The pCL1925-ter vector and the chnA operon fragment were ligated together to give pCL1925-ter-T5chnA and transformed into E. coli Top10. Transformants were screened by colony PCR amplification with primers pCL1925 vec F (SEQ ID NO:64) and chnseq R1 (SEQ ID NO:59) for a product of approximately 1 kbp.
[0284] To finish building the pathway vector, the pCL1925-KoDD-ddr plasmid was digested with XbaI and SacI and the resulting 9504 bp vector fragment was gel-purified. The chnA operon flanked by terminators, with the trpA terminator (Koichi et al. (1997) Volume 272, Number 51, pp. 32034-32041) 3' to the chnA coding sequence, from pCL1925-ter-T5chnA was gel-purified as a 1271 bp XbaI/SacI fragment. After ligation of the fragments and transformation into E. coli Top10, transformants were screened by colony PCR. Primers chnSeq F1 (SEQ ID NO:58) and pCL1925 vec R2 (SEQ ID NO:64) amplified the expected 1107 bp PCR product in the resulting plasmid, pCL1925-KoDD-ddr::ter-T5chnA.
Example 14
Expression of a 2-Butanol Biosynthetic Pathway in E. coli with Overexpressed Endogenous Alcohol Dehydrogenase
[0285] The purpose of this Example was to express a 2-butanol biosynthetic pathway in several E. coli strains.
Construction of E. Coli Strains Constitutively Expressing yghD:
[0286] E. coli contains a native gene (yqhD) that was identified as a 1,3-propanediol dehydrogenase (U.S. Pat. No. 6,514,733). The yqhD gene, given as SEQ ID NO:74, has 40% identity to the gene adhB in Clostridium, a probable NADH-dependent butanol dehydrogenase. The yqhD gene was placed under the constitutive expression of a variant of the glucose isomerase promoter 1.6GI (SEQ ID NO:67) in E. coli strain MG1655 1.6yqhD::Cm (WO 2004/033646) using λ Red technology (Datsenko and Wanner, Proc. Natl. Acad. Sci. U.S.A. 97:6640 (2000)). Similarly, the native promoter was replaced by the 1.5GI promoter (WO 2003/089621) (SEQ ID NO:68), creating strain MG1655 1.5yqhD::Cm, thus, replacing the 1.6GI promoter of MG1655 1.6yqhD::Cm with the 1.5G1 promoter. The 1.5GI and 1.6GI promoters differ by 1 bp in the -35 region, thereby altering the strength of the promoters (WO 2004/033646). While replacing the native yqhD promoter with either the 1.5GI or 1.6GI promoter, the yqhC gene encoding the putative transcriptional regulator for the yqh operon was deleted. Butanol dehydrogenase activity was confirmed by enzyme assay using methods that are well known in the art.
Transformation of E. coli Strains:
[0287] Pathway plasmids pCL1925-Kodd-ddr and pTrc99a-budABC, described in Example 13, were co-transformed into E. coli strains MG1655, MG1655 1.6yqhD, and MG1655 1.5yqhD. The two latter strains overexpress the 1,3-propanediol dehydrogenase, YqhD, which also has butanol dehydrogenase activity. Strains were examined for the production of 2-butanone and 2-butanol essentially as described above. Cells were inoculated into shake flasks (approximately 175 mL total volume) containing either 50 or 150 mL of TM3a/glucose medium (with 0.1 mg/L vitamin B12, appropriate antibiotics and IPTG) to represent medium and low oxygen conditions, respectively. Spectinomycin (50 μg/mL) and carbenicillin (100 μg/mL) were used for plasmids pCL1925-Kodd-ddr and pTrc99a-budABC, respectively. The flasks were inoculated at a starting OD600 of ≦0.04 units and incubated at 34° C. with shaking at 300 rpm. The flasks containing 50 mL of medium were capped with vented caps; the flasks containing 150 mL, were capped with non-vented caps to minimize air exchange. IPTG was present at time zero at a concentration of zero or 0.04 mM. Analytical results for 2-butanone and 2-butanol production are presented in Table 15. All the E. coli strains comprising a 2-butanol biosynthetic pathway produced 2-butanone under low and medium oxygen conditions and produced 2-butanol under low oxygen conditions.
TABLE-US-00015 TABLE 15 Production of 2-Butanone and 2-Butanol by E. coli MG1655 strains harboring pathway plasmids pCL1925-Kodd-ddr and pTrc99a-budABC Volume of 2-Butanone, 2-Butanol, Straina,b IPTG, mM Medium, mL mM mM MG1655 #1 0 50 0.08 Not detected MG1655 #2 0 50 0.11 Not detected MG1655 #1 0.04 50 0.12 Not detected MG1655 #2 0.04 50 0.11 Not detected MG1655 #1 0 150 0.15 0.047 MG1655 #2 0 150 0.19 0.041 MG1655 #1 0.04 150 0.10 0.015 MG1655 #2 0.04 150 0.11 0.015 MG1655 0 50 0.10 Not detected 1.5yqhD #1 MG1655 0 50 0.07 Not detected 1.5yqhD #2 MG1655 0.04 50 0.12 Not detected 1.5yqhD #1 MG1655 0.04 50 0.18 Not detected 1.5yqhD #2 MG1655 0 150 0.16 0.030 1.5yqhD #1 MG1655 0 150 0.18 0.038 1.5yqhD #2 MG1655 0.04 150 0.10 0.021 1.5yqhD #1 MG1655 0.04 150 0.09 0.017 1.5yqhD #2 MG1655 0 50 0.08 Not detected 1.6yqhD #1 MG1655 0 50 0.07 Not detected 1.6yqhD #2 MG1655 0.04 50 0.12 Not detected 1.6yqhD #1 MG1655 0.04 50 0.15 Not detected 1.6yqhD #2 MG1655 0 150 0.17 0.019 1.6yqhD #1 MG1655 0 150 0.18 0.041 1.6yqhD #2 MG1655 0.04 150 0.11 0.026 1.6yqhD #1 MG1655 0.04 150 0.11 0.038 1.6yqhD #2 Control Not detected Not detected (uninoculated medium) a#1 and #2 represent independent isolates. bMG1655 is MG1655/pCL1925-Kodd-ddr/pTrc99a-budABC MG1655 1.6yqhD is MG1655 1.6yqhD/pCL1925-Kodd-ddr/pTrc99a-budABC MG1655 1.6yqhD is MG1655 1.5yqhD/pCL1925-Kodd-ddr/pTrc99a-budABC.
Example 15
Expression of a 2-Butanol Biosynthetic Pathway in E. coli with Heteroloqous Alcohol Dehydrogenase
[0288] Plasmids pCL1925-KoDD-ddr::ter-T5chnA and pTrc99a-budABC, described in Example 13, were transformed into E. coli strains MG1655 and MG1655 ΔyqhCD for a demonstration of the production of 2-butanol.
[0289] MG1655 ΔyqhCD carries a yqhCD inactivation that was made using the method of Datsenko and Wanner (Proc. Natl. Acad. Sci. U.S.A. 97(12):6640-6645 (2000)). After replacement of the region with the FRT-CmR-FRT cassette of pKD3, the chloramphenicol resistance marker was removed using the FLP recombinase. The sequence of the deleted region is given as SEQ ID NO:66.
[0290] Strains MG1655/pTrc99a-budABC/pCL1925KoDD-ddr::ter-T5 chnA and MG1655 ΔyqhCD/pTrc99a-budABC/pCL1925KoDD-ddr::ter-T5 chnA were examined for the production of 2-butanone and 2-butanol essentially as described above. Strain MG1655 ΔyqhCD/pCL1925 was used as a negative control. Cells were inoculated into shake flasks (approximately 175 mL total volume) containing 50 or 150 mL of TM3a/glucose medium (with 0.1 mg/L vitamin B12 and appropriate antibiotics) to represent medium and low oxygen conditions, respectively. Spectinomycin (50 μg/mL) and ampicillin (100 μg/mL) were used for selection of pCL1925 based plasmids and pTrc99a-budABC, respectively. Enzyme activity derived from pTrc99a-budABC was detected by enzyme assay in the absence of IPTG inducer, thus, IPTG was not added to the medium. The flasks were inoculated at a starting OD600 of ≦0.01 units and incubated at 34° C. with shaking at 300 rpm for 24 h. The flasks containing 50 mL of medium were capped with vented caps; the flasks containing 150 mL, were capped with non-vented caps to minimize air exchange. Analytical results for 2-butanone and 2-butanol production are presented in Table 16. Both E. coli strains comprising a 2-butanol biosynthetic pathway produced 2-butanone under low and medium oxygen conditions and produced 2-butanol under low oxygen conditions, while the negative control strain did not produce detectable levels of either 2-butanone or 2-butanol.
TABLE-US-00016 TABLE 16 Production of 2-butanone and 2-butanol by E. coli strains 2- Volume, Butanone, 2-Butanol, Straina mL mM mM Negative control, MG1655 50 Not Not detected ΔyqhCD/pCL1925 detected MG1655/pTrc99a- 50 0.33 Not detected budABC/pCL1925KoDD-ddr::T5 chnA ter MG1655 ΔyqhCD/pTrc99a- 50 0.23 Not detected budABC/pCL1925KoDD-ddr::T5 chnA ter #1 MG1655 ΔyqhCD/pTrc99a- 50 0.19 Not detected budABC/pCL1925KoDD-ddr::T5 chnA #2 Negative control, MG1655 150 Not Not detected ΔyqhCD/pCL1925 detected MG1655/pTrc99a- 150 0.41 0.12 budABC/pCL1925KoDD-ddr::T5 chnA ter MG1655 ΔyqhCD/pTrc99a- 150 0.15 0.46 budABC/pCL1925KoDD-ddr::T5 chnA #1 MG1655 ΔyqhCD/pTrc99a- 150 0.44 0.14 budABC/pCL1925KoDD-ddr::T5 chnA #2 Medium Not Not detected detected a#1 and #2 represent independent isolates.
Example 16
[0291] Cloning of Amino:Pyruvate Transaminase (APT)
[0292] An amino:pyruvate transaminase (APT) from Vibrio Fluvialis JS17 was identified by Shin et al. (Appl. Microbiol. Biotechnol. (2003) 61:463-471). The amino acid sequence (SEQ ID NO:122) was found to have significant homology with ω-amino acid:pyruvate transaminases (Shin and Kim (J. Org. Chem. 67:2848-2853 (2002)). It was shown that the Vibrio Fluvialis APT has transaminase activity towards acetoin.
[0293] For expression of the APT enzyme in E. coli, a codon optimized APT coding region (SEQ ID NO:144) was designed using the preferred E. coli codons with additional considerations such as codon balance and mRNA stability, and synthesized (by DNA2.0; Redwood City, Calif.). The coding region DNA fragment was subcloned into the pBAD.HisB vector (Invitrogen) between the NcoI and HindIII sites and the resulting plasmid, hereafter referred to as pBAD.APT1, was transformed into TOP10 cells.
Example 17
Characterization of Vibrio Fluvialis APT Alanine:Acetoin Aminotransferase Activity
[0294] A 5 mL volume of LB broth+100 μg/mL ampicillin was inoculated with a fresh colony of TOP10/pBAD:APT1 cells. The culture was incubated at 37° C. for approximately 16 h with shaking (225 rpm). A 300 μL aliquot of this culture was used to inoculate 300 mL of the same medium, which was incubated at 37° C. with shaking (225 rpm). When the culture reached an OD600 of 0.8, L-arabinose was added to a final concentration of 0.2% (w/v). The culture was incubated for an additional 16 h, then harvested. The cells were washed once with 100 mM potassium phosphate buffer (pH 7.8) and then frozen and stored at -80° C.
[0295] To isolate the enzyme, the cell pellet was thawed and resuspended in 8 mL of 100 mM potassium phosphate buffer (pH 7) containing 0.2 mM ethylenediaminetetraacetate, 1 mM dithiothreitol and 1 tablet of protease inhibitor cocktail (Roche; Indianapolis, Ind.). The cells were lysed by two passes through a French pressure cell at 900 psi, and the resulting lysate was clarified by centrifugation for 30 min at 17000×g. Ammonium sulfate was added to 35% saturation, and the solution was stirred for 30 min at room temperature, at which point precipitated solids were removed by centrifugation (30 min, 17000×g). Additional ammonium sulfate was added to the supernatant to give 55% saturation, and the solution was again stirred for 30 min at room temperature. The precipitated solids were removed by centrifugation (30 min, 17000×g) and then resuspended in 5 mL of 100 mM potassium phosphate buffer (pH 7) containing 10 μM pyridoxal 5'-phosphate and 1 mM dithiothreitol. This solution was desalted by passage through a PD10 column equilibrated with Buffer A (50 mM bis-tris propane buffer (pH 6) containing 10 μM pyridoxal 5'-phosphate and 1 mM dithiothreitol). The desalted extract was then loaded onto a 20 mL Q-Fast Flow column pre-equilibrated with Buffer A. APT was eluted with a linear gradient of 0-0.1 M NaCl in Buffer A. The enzyme was detected in eluted fractions by the presence of a protein band of size ˜50 kD when analyzed by SDS-polyacrylamide gel electrophoresis and by the characteristic absorbance at 418 nm. Fractions containing the enzyme eluted at ˜0.3 M NaCl. These fractions were pooled to yield a total of 6 mL of a 5.45 mg/mL solution of enzyme, which was >90% pure, as judged by SDS-polyacrylamide gel electrophoresis.
[0296] The alanine:acetoin aminotransferase activity of APT was assayed using a lactic dehydrogenase coupled assay. Reaction mixtures contained 100 mM bis-tris propane (pH 9.0), 10 μM pyridoxal 5'-phosphate, 0-50 mM acetoin, 0-5 mM L-alanine, 0.14 or 0.28 mg/mL purified enzyme, 200 μM NADH and 20 U/mL lactic dehydrogenase (Sigma; St. Louis, Mo.). The reaction was followed by measuring the change in absorbance at 340 nm, indicative of the oxidation of NADH. Under these conditions, the kcat/Km for acetoin was 10 M-1 s-1 and that for L-alanine was 400 M-1 s-1.
[0297] The identity of the expected product 3-amino-2-butanol was confirmed by comparison to a synthetic standard. A mixture of (R,R)- and (S,S)-3-amino-2-butanol was synthesized by the method of Dickey et al. [J Amer Chem Soc 74:944 (1952)]: 5 g of trans-2,3-epoxybutane were slowly stirred into 150 mL of cold (4° C.) NH4OH. The reaction was slowly warmed to room temperature, sealed and stirred at room temperature for an additional 10 days. At this time, excess ammonia and water and residual epoxybutane were removed by rotary evaporation under vacuum at 40° C. The resulting clear oil (2.9 g) was resuspended in water to a concentration of 10% (w/v). Production of the desired product was confirmed by NMR analysis and comparison of the spectrum to that reported by Levy et al. [Org. Magnetic Resonance 14:214 (1980)]. A mixture of the corresponding (2R,3S)- and (2S,3R)-isomers was produced using the identical method with the exception that the starting material was the cis-isomer of 2,3-epoxybutane.
[0298] An analytical method for detection of 3-amino-2-butanol was developed based on the o-phthaldialdehyde derivatization method for amino acid determination reported by Roth [Anal. Chem. 43:880 (1971)]. A 200 μL aliquot of 1 mM 3-amino-2-butanol (mixture of isomers) was mixed with 200 μL of a 50 mM solution of borate (pH 9.5), to which was added 10 μL of 5 μL/mL 2-mercaptoethanol in ethanol and 10 μL of 10 mg/mL o-phthaldialdehdye in ethanol. The solution was incubated at room temperature for 10 min, at which time the derivative was extracted into 200 μL hexane. The hexane was separated from the aqueous solution by decanting, and 10 μL were injected onto a Chiracel OD HPLC column (Daicel Chemical Industries; Fort Lee, N.J.). The column was run isocratically with a mobile phase of 90:10 hexane:isopropanol at a rate of 1 mL/min. The derivatized isomers of 3-amino-2-butanol were detected by absorbance at 340 nm with retention times of approximately 15.7 and 16.8 min [(2S,3S) and (2R,3R)], and 18.4 and 21.9 min [(2R,3S) and (2S,3R)]. To differentiate the enantiomers in the first mixture, the pure (2R,3R) isomer (Bridge Organics; Vicksburg, Mich.) was also run under the identical conditions and found to be the 16.8 min peak. To differentiate the enantiomers in the second mixture, the mixture was first kinetically resolved using the alanine:acetoin aminotransferase: 0.28 mg of purified enzyme was incubated with 10 mM pyruvate and 10 mM 3-amino-2-butanol [1:1 mixture of (2R,3S) and (2S,3R) isomers] in 1 mL of 100 mM bis-tris propane (pH 9.0). After 24 h at room temperature, an aliquot was removed and analyzed as described above. Analysis revealed that the 18.4 min peak was 95% depleted, while the 21.9 min peak was >90% retained. A 100 μL aliquot of the remaining reaction mixture was mixed with 50 μL of 20 mM NADH and 10 μL of extract from the TOP10/pTrc99a-BudC strain described in Example 13. The BudC enzyme is known to reduce (R)-acetoin to meso-2,3-butanediol and (S)-acetoin to (S,S)-2,3-butanediol [Ui et al., (2004) Letters in Applied Microbiology 39:533-537]. After 3 h, samples were taken from the reaction and analyzed as described above for acetoin and butanediol. The analysis indicated that the primary product of the reduction was meso-2,3-butanediol, indicating that the product of the aminotransferase reaction was (R)-acetoin, and therefore the consumed 3-amino-2-butanol isomer was the (2R,3S) isomer. Thus the retention time of 18.4 min can be assigned to this isomer and 21.9 to the (2S,3R) isomer.
[0299] To confirm that the product of the APT-catalyzed alanine:acetoin aminotransferase reaction was 3-amino-2-butanol, 0.28 mg of purified enzyme was incubated with 10 mM acetoin, 10 mM L-alanine, 50 U lactic dehydrogenase and 200 μM NADH in 1 mL of 100 mM bis-tris propane (pH 9.0). The reaction mixture was incubated at room temperature for 20 h, after which a 200 μL aliquot was removed and derivatized as described above. The retention times of the derivatized products were 15.8 min (major product) and 18.5 min (minor product), matching that of the (2S,3S)- and (2R,3S)-3-amino-2-butanol standards.
Example 18
Identification and Cloning of Erwinia carotovora subsp. atroseptica Amino Alcohol Kinase and Amino Alcohol O-Phosphate Lyase
[0300] The purpose of this example is to describe the identification and cloning of sequences encoding an amino alcohol kinase and amino alcohol O-phosphate lyase from the bacterium Erwinia carotovora. These two enzymes are part of Pathway 1 for the conversion of 3-amino-2-butanol to 2-butanone via the intermediate 3-amino-2-butanol phosphate as shown in FIG. 1.
Prediction of the Erwinia Amino Alcohol Kinase and the Amino Alcohol O-Phosphate Lyase
[0301] ATP-dependent amino alcohol kinase and amino alcohol O-phosphate lyase activities have been detected in several Pseudomonas and Erwinia species, including Pseudomonas sp. P6 (NCIB10431), Pseudomonas putida NCIB 10558 (Jones et al. (1973) Biochem. J. 134:167-182), Erwinia carotovora, Erwinia amanas, Erwina milletiae, and Erwinia atroseptica (Jones et al. (1973) Biochem. J. 134:959-968). In these studies, the extracts of the above species were shown to have activity for the enzymatic conversion of aminopropanol through aminopropanol O-phosphate to propionaldehyde, and the conversion of ethanolamine through ethanolamine O-phosphate to acetaldehyde.
[0302] The genomic sequence of the Erwinia atroseptica strain in which these activities were reported to exist (now designated as Erwinia carotovora subsp. atroseptica strain SCRI1043 (ATCC BAA-672)) has been determined at the Sanger Institute (Bell et al. Proc. Natl. Acad. Sci. USA 101(30): 11105-11110). Analysis of the putative kinases in the Erwinia carotovora subsp. atroseptica genome revealed an operon sequence (SEQ ID NO:154) encoding a putative protein (ECA2059; SEQ ID NO:124) that is 39% identical to a Rhizobium loti homoserine kinase and a putative class-III pyridoxal phosphate (PLP)-dependent aminotransferase (ECA2060; SEQ ID NO:126) that is 58% identical to a putative aminotransferase from Rhizobium meliloti. We predicted that ECA2059 was an amino alcohol kinase and ECA2060 was an amino alcohol O-phosphate lyase which uses PLP as cofactor.
Cloning of the Putative Amino Alcohol Kinase and Putative Amino Alcohol O-Phosphase Lyase from Erwinia carotovora subsp. atroseptica
[0303] Genomic DNA of Erwinia carotovora subsp. atroseptica (ATCC #: BAA-672D) was obtained from American Type Culture Collection (ATCC). The operon encoding the putative amino alcohol kinase (KA) and amino alcohol O-phosphate lyase (AT) was named KA-AT (SEQ ID NO:154. This operon was amplified from the Erwinia genomic DNA by Phusion DNA polymerase (Finnzymes; via New England Biolabs; Ipswich, Mass.) using primers OT872 (SEQ. ID. No. 127) and OT873 (SEQ. ID. No128). A DNA fragment of 2.4 kb was obtained by the PCR reaction, which corresponds to the size of the KA-AT operon. The PCR product was digested with EcoRI and PstI restriction enzymes, and cloned into vector pKK223-3 (Amersham Biosciences; Piscataway, N.J.) which was digested with the same restriction enzymes. This produced plasmid pKK223.KA-AT, which contained the putative Erwinia amino alcohol kinase-lyase operon under control of the tac promoter. Similarly, plasmids pKK223.KA and pKK223.AT were made which placed the putative Erwinia kinase and the putative Erwinia lyase coding regions in separate vectors, each under the control of the tac promoter. For the PCR cloning of the KA coding region (SEQ ID NO:123), primers OT872 (SEQ. ID. No. 127) and OT879 (SEQ. ID. No. 129) were used; and for the PCR cloning of AT coding region (SEQ ID NO:125), primers OT873 (SEQ. ID. No. 128) and OT880 (SEQ. ID. No. 130) were used in the PCR amplifications, which generated PCR products of 1.1 kb and 1.3 kb respectively. The PCR products were each digested with EcoRI and PstI, and ligated into vector pKK223-3 to generate pKK223.KA and pKK223.AT.
In Vivo Activity of the Putative Amino Alcohol Kinase and Putative Amino Alcohol O-Phosphate Lyase from Erwinia carotovora subsp. atroseptica
[0304] Plasmids pKK223.KA-AT, pKK223.KA, pKK223.AT and pKK223-3 were transformed into the E. coli MG1655 strain. The transformants were restreaked onto a MOPS minimal media plate containing 1% glucose, 0.5% aminopropanol as a sole nitrogen source, 1 mM IPTG and 100 μg/mL ampicillin. Expression of KA-AT, KA and AT genes were induced by the IPTG. A control plate had no IPTG included. The plates were incubated at 37° C. for 7 days. On the plate with IPTG, only the strain MG1655/pKK223.KA-AT grew, while all the other three strains did not grow. On the plate without added IPTG, the strain MG1655/pKK223.KA-AT grew, but the colonies were significantly smaller than those on the IPTG-containing plate, which corresponds to the lower expression levels of KA and AT in the uninduced cells. None of the other three strains grew on this plate. This indicates that the co-expression of the putative Erwinia KA and AT genes provided sufficient enzyme activities that allowed the E. coli strain MG1655/pKK223.KA-AT to utilize aminopropanol as a sole nitrogen source. Expression of each individual enzyme of either KA or AT was not sufficient to provide such enzyme activity in vivo.
Example 19
In vitro Activity of Erwinia putative Amino Alcohol Kinase and Amino Alcohol O-Phosphate Lyase
[0305] Subcloning of the Erwinia KA-AT Operon into the pBAD.HisB Vector and Induction of Protein Expression
[0306] The protein expression levels of Erwinia putative KA and AT enzymes expressed in MG1655 cells from the pKK223.KA-AT vector were analyzed by SDS-PAGE analysis. The expression level of the Erwinia AT enzyme was relatively low, with a new protein band detected at the correct molecular weight of 46 kD in the soluble fraction of a cell extract, while no new protein band was detected at the size predicted for the KA enzyme.
[0307] In an effort to improve the expression of the Erwinia putative KA and AT genes, the KA-AT operon was subcloned into the EcoRI and HindIII sites of vector pBAD.HisB-EcoRI. pBAD.HisB-EcoRI was derived from the pBAD.HisB vector (Invitrogen), by replacing the NcoI site in pBAD.HisB with an EcoRI site via QuickChange site-directed mutagenesis (Stratagene, La Jolla, Calif.) using primers OT909 (SEQ ID.#131) & OT910 (SEQ ID.#132). In the constructed plasmid pBAD.KA-AT, the KA-AT operon was placed directly under control of the araB promoter (without His-tag).
[0308] The pBAD.KA-AT plasmid was transformed into the E. coli TOP10 strain. A 50 mL culture of TOP10/pBAD.KA-AT strain was grown to mid log phase (OD600=0.6) in LB, 100 μg/mL ampicillin media at 37° C. with shaking at 250 rpm. The culture was induced by addition of L-arabinose to a final concentration of 0.1% (w/v), and it was further incubated at 37° C. for 5 h before harvesting by centrifugation. The cell pellet was resuspended in ice cold 50 mM Tris-HCl, pH 8.0, and disrupted by sonication on ice with a Fischer Sonic Model 300 Dismembrator (Fischer, Pittsburgh, Pa.) at 50% power, repeating four cycles of 30 seconds sonication with 60 seconds rest in-between each cycle. Each sonicated sample was centrifuged (15,000×g, 4 min, 4° C.). Clarified cell free extracts were analyzed for protein expression level and amino alcohol O-phosphate lyase activity.
Chemical Synthesis of Aminobutanol O-Phosphate and Aminopropanol O-Phosphate The substrate (R,R)-3-amino-2-butanol O-phosphate was synthesized by a method based on that reported by Ferrari and Ferrari (U.S. Pat. No. 2,730,542
[1956]) for phosphoethanolamine: 10 mmol of H3PO4 in a 50% (w/v) aqueous solution was mixed with a 50% (w/v) solution of 3-amino-2-butanol (-20:1 (R,R):(S,S) isomers; Bridge Organics; Vicksburg, Mich.) while stirring on ice. After mixing, the solution was slowly warmed to room temperature and then stirred under vacuum and heated to 70° C. After 1 h at 70° C., the temperature was slowly increased to 185° C. and maintained there for an additional 2 h. At that time, the reaction was cooled to room temperature and the vacuum released. The remaining material was dissolved in water, and analysis by NMR indicated that 80% of the starting material was converted to product with 20% remaining unreacted. No additional products were observed.
[0309] The additional substrates (2R,3S)-3-amino-2-butanol O-phosphate and (2S,3R)-3-amino-2-butanol O-phosphate were synthesized by the same procedure using a 1:1 mixture of (2R,3S)-3-amino-2-butanol and (2S,3R)-3-amino-2-butanol (synthesized as described in Example 17) as the starting material. DL-1-amino-2-propanol O-phosphate, (S)-2-amino-1-propanol O-phosphate, and (R)-2-amino-1-propanol O-phosphate were synthesized by the same procedure using DL-1-amino-2-propanol, (R)-2-amino-1-propanol, or (S)-2-amino-1-propanol as the starting material.
Analysis of the Aminopropanol O-Phosphate Lyase Activity Encoded by the Putative Erwinia KA-AT Operon
[0310] The aminopropanol O-phosphate lyase assay was performed as described by Jones et al. (1973, Biochem. J. 134:167-182) and G. Gori et al. (1995, Chromatographia 40:336) The formation of propionaldehyde from aminopropanol O-phosphate was assayed colorimetrically with MBTH, which allows the detection of aldehyde formation. The reaction was performed as follows. In a 1 mL reaction, 100 μg cell free extract of E. coli TOP10/pBAD.KA-AT was added to 10 mM DL-1-amino-2-propanol O-phosphate in 100 mM Tris-HCl, pH 7.8, with 0.1 mM PLP. The reaction was incubated at 37° C. for 10 min and 30 min, with an aliquot of 100 μL reaction mixture removed at each time point and mixed with 100 μL of 6 mg/mL MBTH in 375 mM glycine-HCl, pH 2.7. This mixture was incubated at 100° C. for 3 min, cooled on ice for 15-30 s, and 1 mL of 3.3 mg/mL FeCl3.6H2O (in 10 mM HCl) was added, followed by incubation for 30 min at room temperature. The absorbance of the reaction mixture which contains the aldehyde-MBTH adduct, was measured at 670 nm. The results of the assay are listed in Table 17. In the presence of the aminopropanol phosphate substrate, PLP and cell free extract, formation of aldehyde was detected, as indicated by an Abs670 that was higher than the control background of up to 0.3. In the absence of either the substrate or the cell free extract, no aldehyde formation was detected. In the absence of added PLP, somewhat less amount aldehyde was detected, presumably due to the presence of PLP in the cell free extract. Cell free extract of the uninduced TOP10/pBAD.KA-AT-culture did not produce any detectable aldehyde in the reaction. These results indicated that the putative Erwinia amino alcohol O-phosphate lyase does catalyze the conversion of aminopropanol O-phosphate to propionaldehyde.
TABLE-US-00017 TABLE 9 Aminopropanol O-phosphate lyase assay. Sample 1 was the cell free extract of a non-induced control of E. coli TOP10/pBAD.KA-AT. Samples 2-5 contained the cell free extract of the induced culture E. coli TOP10/pBAD.KA-AT. Enzyme Induction Aminopro- extract Sample by 0.1% panol O- (100 OD670, OD670, Number arabinose phosphate PLP μg/mL) 10 min 30 min 1 uninduced (+) (+) (+) 0.262 0.255 2 induced (+) (+) (+) 1.229 2.264 3 induced (-) (+) (+) 0.303 0.223 4 induced (+) (-) (+) 0.855 1.454 5 induced (+) (+) (-) 0.156 0.065
Analysis of the Activity of the Erwinia Amino Alcohol O-Phosphate Lyase Towards Aminobutanol O-Phosphate Substrate
[0311] The activity of the amino alcohol O-phosphate lyase towards the aminobutanol O-phosphate substrates was studied under the same conditions as described above. The reaction was carried out at 37° C. overnight in a 1 mL reaction that contained 100 μg of cell free extract of E. coli TOP10/pBAD.KA-AT, 10 mM aminobutanol O-phosphate (either the mixture of (R,R)+(S,S) or the mixture of (R,S)+(S,R) isomers described in Example 19) in 100 mM Tris-HCl, pH 7.8, with 0.1 mM PLP. An aliquot of 100 μL reaction mixture was removed and the 2-butanone product was detected using the MBTH derivatization method described in the General Methods. The two peaks representing the derivatized 2-butanone isomers were observed. Therefore the Erwinia amino alcohol O-phosphate lyase is an aminobutanol phosphate phospho-lyase in addition to an aminopropanol phosphate phospho-lyase.
Analysis of the Activity of the Erwinia Amino Alcohol O-Phosphate Lyase Towards Stereoisomers of Aminopropanol O-Phosphate and Aminobutanol O-Phosphate
[0312] The activity of the Erwinia amino alcohol O-phosphate lyase towards various stereoisomers of aminopropanol O-phosphate and aminobutanol O-phosphate was studied under the same conditions as described above. In the presence of the Erwinia amino alcohol O-phosphate lyase, both (R) and (S)-2-amino-1-propanol O-phosphate were converted to propanone by the enzyme, but the product yield was much higher with the (S) isomer. The enzyme also produced butanone from both mixtures of 3-amino-2-butanol O-phosphate isomers, with a higher product yield found in the reaction containing the (R,S) and (S,R) substrate isomers. Both propanone and butanone products were derivatized by MBTH, and detected by HPLC as described in General Methods.
Optimization of the Gene Expression Level for the Erwinia Amino Alcohol Kinase and Amino Alcohol O-Phosphate Iyase
[0313] In order to improve the expression levels for the Erwinia amino alcohol kinase and the amino alcohol O-phosphate lyase in E. coli, codon optimized coding regions for both enzymes (named EKA: SEQ ID NO:155 and EAT: SEQ ID NO:156 respectively) were synthesized by DNA2.0 (Redwood City, Calif.). Each coding region was synthesized with 5' and 3' tails including restriction sites for cloning: EKA has 5' BbsI and 3' EcoRI, HindIII sites; EAT has 5' EcoRI and 3'HindIII sites. The EKA and EAT coding regions were provided from DNA2.0 as plasmids pEKA and pEAT, which were in the pJ51 vector of DNA2.0. The EKA optimized coding region was subcloned by ligating a BbsI and HindIII digested fragment of pEKA into the pBAD.HisB vector between the NcoI and HindIII sites, to generate plasmid pBAD.EKA. In the resulting plasmid the coding region is 5' to the His tag, so a coding region for an N-terminus His6 tag fused to the Erwinia amino alcohol kinase was constructed by performing a QuickChange site-directed mutagenesis reaction using primers SEQ ID NO:157 and SEQ ID NO:158 to generate vector pBAD.His-EKA. pBAD.His-EKA was transformed into E. coli strain BL21-Al (F ompT hsdSB (rB.sup.- mB.sup.-) gal dcm araB::T7RNAP-tetA; Invitrogen) to produce strain BL21-Al/pBAD.HisA-EKA. A 50 mL culture of BL21-Al/pBAD.HisA-EKA was grown to mid-log stage (OD600=0.6), induced with 0.1% arabinose, and further incubated at 30° C. overnight. Cell free extracts were prepared by sonication. The His6-tagged fusion protein of Erwinia amino alcohol kinase was purified using the ProBond® Purification System (Invitrogen) under non-denaturing purification conditions following the manufacturer's instructions.
[0314] The kinase activity of the His6-tagged Erwinia amino alcohol kinase is analyzed by the ADP Quest Assay (DiscoveRx, Fremont, Calif.) following the manufacturer's instructions. This is a biochemical assay that measures the accumulation of ADP, a product of the amino alcohol kinase reaction using either aminopropanol or aminobutanol as substrate. 10 mM substrate is mixed with His6-tagged Erwinia amino alcohol kinase, in 100 mM Tris-HCl, pH 7.8, 10 mM MgCl2, 2 mM KCl, 0.1 mM ATP, and incubated at 37° C. for 1 h in a 0.2 mL reaction. ADP reagent A (100 μL) and ADP reagent B (200 μL) are added and the mixture is incubated at room temperature for 30 min. The fluorescence signal indicating activity is measured with excitation wavelength of 530 nm and emission wavelength of 590 nm.
Example 20
Expression of Entire Pathway 3 Construction of Vector pCLBudAB-ter-T5chnA
[0315] The vector pTrc99a::BudABC (described in Example 13) is digested with EcoRI, and the DNA is treated with Klenow DNA polymerase to blunt the ends. The blunted vector is subsequently digested with SpeI to yield a 2.5 kb fragment containing the budA and budB genes. The vector pCL1925-ter-T5chnA (described in Example 13) is digested with HindIII, and the DNA was treated with Klenow DNA polymerase to blunt the ends. The blunted vector is subsequently digested with XbaI to yield a 4.6 kb fragment which is then ligated to the budAB fragment from pTrc99a::BudABC. The resulting plasmid, designated pCLBudAB-ter-T5chnA, is used to transform E. coli Top10 cells, and single colonies are screened for proper plasmid structure by PCR using primers pCL1925vecF (SEQ ID NO:62) and N84seqR3 (SEQ ID NO:159). Plasmid is prepared from a single colony which yields a PCR product of the expected size of 1.4 kb.
Construction of Vector DKK223.KA-AT-APT
[0316] The APT gene is amplified from the vector pBAD.APT (described in Example 16) by PCR using primers APTfor (SEQ ID NO:162; 5' includes RBS and SmaI site) and APTrev (SEQ ID NO:163; 3' adds SmaI site). The product of expected size of 1.7 kbp is gel purified and digested with SmaI to yield blunt ends. The vector pKK223.KA-AT (described in Example 18) is digested with PstI, and the DNA is treated with Klenow DNA polymerase to blunt the ends. The resulting DNA fragment is ligated with the SmaI-digested PCR product, and the ligation product is used to transform E. coli Top10 cells. Individual ampicillin resistant colonies are screened by PCR using primers OT872 (SEQ ID NO:127) and APTrev (SEQ ID NO:163). The presence of a PCR product of the expected size of 4.1 kbp indicates that the gene encoding APT is present and oriented in the same direction as the genes encoding KA and AT. The sequence of the insert is verified using the primers APTseqRev (SEQ ID NO:160) and APTseqFor (SEQ ID NO:161). This plasmid is named pKK223.KA-AT-APT. Proper expression of all three genes is verified by growing a 5 mL culture of Top10/pKK223.KA-AT-APT in LB+100 μg/mL ampicillin at 37° C. with shaking. When the OD600 reaches ˜0.8, expression of the genes on the plasmid is induced by addition of IPTG to 0.4 mM. The expression is evaluated by SDS PAGE and activity assays as described above.
Construction of 2-Butanol Production Strain and Production of 2-Butanone and 2-Butanol
[0317] E. coli strain MG1655 is transformed with both pKK223.KA-AT-APT and pCLBudAB-ter-T5chnA, and transformants selected for ampicillin and spectinomycin resistance, indicative of the presence of the plasmids. The cells are inoculated into shake flasks (approximately 175 mL total volume) containing 50 or 150 mL of TM3a/glucose medium (with appropriate antibiotics) to represent medium and low oxygen conditions, respectively. IPTG is added to 0.4 mM to induce expression of genes from pKK223.KA-AT-APT. As a negative control, MG1655 cells are grown in the same medium lacking antibiotics. The flasks are inoculated at a starting OD600 of 0.01 and incubated at 34° C. with shaking at 300 rpm for 24 h. The flasks containing 50 mL of medium are capped with vented caps; the flasks containing 150 mL are capped with non-vented caps to minimize air exchange. The MG1655/pKK223.KA-AT-APT/pCLBudAB-ter-T5chnA strain comprising a 2-butanol biosynthetic pathway produces both 2-butanone and 2-butanol under low and medium oxygen conditions while the negative control strain does not produce detectable levels of either 2-butanone or 2-butanol.
Example 21
Characterization of Glycerol Dehydratase Butanediol Dehydratase Activity
[0318] Glycerol dehydratase (E.C. 4.2.1.30) and diol dehydratase (E.C. 4.2.1.28), while structurally related, are often distinguished in the art based on various differences that include substrate specificity. This example demonstrates that glycerol dehydratase converts meso-2,3-butanediol to 2-butanone. The recombinant E. coli strain KLP23/pSYCO12, comprising Klebsiella pneumoniae genes encoding the multiple subunits of glycerol dehydratase (alpha: SEQ ID NO:145 (coding region) and 146 (protein); beta: SEQ ID NO: 147 (coding region) and 148 (protein); and gamma: SEQ ID NO: 149 (coding region) and 150 (protein)) and Klebsiella pneumoniae genes encoding the multiple subunits of glycerol dehydratase reactivase (large subunit, SEQ ID NO: 151 (coding region) and 152 (protein); and small subunit, SEQ ID NO: 153 (coding region) and 154 (protein)), is described in Emptage et al. U.S. Pat. No. 6,514,733 and in WO 2003089621, which are herein incorporated by reference. A crude, cell free extract of KLP23/pSYCO12 was prepared by methods known to one skilled in the art. Enzyme assay was performed in the absence of light in 80 mM HEPES buffer, pH 8.2 at 37° C. with 12 μM coenzyme B12 and 10 mM meso-2,3-butanediol. The formation of 2-butanone was monitored by HPLC (Shodex SH-1011 column and SH-G guard column with refractive index detection; 0.01 M H2SO4 as the mobile phase at a flow rate of 0.5 mL/min and a column temperature of 50° C.; 2-butanone retention time=40.2 min). The rate of 2-butanone formation by the glycerol dehydratase preparation was determined to be 0.4 nmol/min/mg of crude protein.
Example 22
[0319] Increased Tolerance of Saccharomyces cerevisiae to 2-Butanol at Decreased Growth Temperatures
[0320] Tolerance levels were determined for yeast strain Saccharomyces cerevisiae BY4741 (ATCC 201388) at 25° C. and 30° C. as follows. The strain was cultured in YPD medium. Overnight cultures in the absence of any test compound were started in 25 mL of YPD medium in 150 mL flasks with incubation at 30° C. or at 25° C. in shaking water baths. The next morning, each overnight culture was diluted into a 500 mL flask containing 300 mL of fresh medium to an initial OD600 of about 0.1. The flasks were incubated in shaking water baths at 30° C. or 25° C., using the same temperature as used for each overnight culture. The large cultures were incubated for 3 hours and then were split into flasks in the absence (control) and in the presence of 2.5% or 3.5% of 2-butanol. Growth was followed by measuring OD600 for six hours after addition of the 2-butanol. The ΔOD600 was calculated by subtracting the initial OD600 from the final OD600 at 6 hours. The percent growth inhibition relative to the control culture was calculated as follows: % Growth Inhibition=100-[100(Sample ΔOD600/Control ΔOD600)]. The results are summarized in Table 18 below and indicate that growth of strain BY4741 was less inhibited by 2.5% and 3.5% 2-butanol at 25° C. than by 2.5% and 3.5% 2-butanol at 30° C.
TABLE-US-00018 TABLE 18 Growth of Saccharomyces cerevisiae Strain BY4741 at 25° C. and 30° C. with 2-Butanol. % Growth % 2-Butanol Temperature ° C. Inhibition 2.5 30 95 2.5 25 89 3.5 30 99 3.5 25 97
Sequence CWU
1
1
1641780DNAKlebsiella pneumoniae 1atgaatcatt ctgctgaatg cacctgcgaa
gagagtctat gcgaaaccct gcgggcgttt 60tccgcgcagc atcccgagag cgtgctctat
cagacatcgc tcatgagcgc cctgctgagc 120ggggtttacg aaggcagcac caccatcgcg
gacctgctga aacacggcga tttcggcctc 180ggcaccttta atgagctgga cggggagctg
atcgccttca gcagtcaggt ctatcagctg 240cgcgccgacg gcagcgcgcg caaagcccag
ccggagcaga aaacgccgtt cgcggtgatg 300acctggttcc agccgcagta ccggaaaacc
tttgaccatc cggtgagccg ccagcagctg 360cacgaggtga tcgaccagca aatcccctct
gacaacctgt tctgcgccct gcgcatcgac 420ggccatttcc gccatgccca tacccgcacc
gtgccgcgcc agacgccgcc gtaccgggcg 480atgaccgacg tcctcgacga tcagccggtg
ttccgcttta accagcgcga aggggtgctg 540gtcggcttcc ggaccccgca gcatatgcag
gggatcaacg tcgccgggta tcacgagcac 600tttattaccg atgaccgcaa aggcggcggt
cacctgctgg attaccagct cgaccatggg 660gtgctgacct tcggcgaaat tcacaagctg
atgatcgacc tgcccgccga cagcgcgttc 720ctgcaggcta atctgcatcc cgataatctc
gatgccgcca tccgttccgt agaaagttaa 7802259PRTKlebsiella pneumoniae 2Met
Asn His Ser Ala Glu Cys Thr Cys Glu Glu Ser Leu Cys Glu Thr 1
5 10 15 Leu Arg Ala Phe Ser Ala
Gln His Pro Glu Ser Val Leu Tyr Gln Thr 20
25 30 Ser Leu Met Ser Ala Leu Leu Ser Gly Val
Tyr Glu Gly Ser Thr Thr 35 40
45 Ile Ala Asp Leu Leu Lys His Gly Asp Phe Gly Leu Gly Thr
Phe Asn 50 55 60
Glu Leu Asp Gly Glu Leu Ile Ala Phe Ser Ser Gln Val Tyr Gln Leu 65
70 75 80 Arg Ala Asp Gly Ser
Ala Arg Lys Ala Gln Pro Glu Gln Lys Thr Pro 85
90 95 Phe Ala Val Met Thr Trp Phe Gln Pro Gln
Tyr Arg Lys Thr Phe Asp 100 105
110 His Pro Val Ser Arg Gln Gln Leu His Glu Val Ile Asp Gln Gln
Ile 115 120 125 Pro
Ser Asp Asn Leu Phe Cys Ala Leu Arg Ile Asp Gly His Phe Arg 130
135 140 His Ala His Thr Arg Thr
Val Pro Arg Gln Thr Pro Pro Tyr Arg Ala 145 150
155 160 Met Thr Asp Val Leu Asp Asp Gln Pro Val Phe
Arg Phe Asn Gln Arg 165 170
175 Glu Gly Val Leu Val Gly Phe Arg Thr Pro Gln His Met Gln Gly Ile
180 185 190 Asn Val
Ala Gly Tyr His Glu His Phe Ile Thr Asp Asp Arg Lys Gly 195
200 205 Gly Gly His Leu Leu Asp Tyr
Gln Leu Asp His Gly Val Leu Thr Phe 210 215
220 Gly Glu Ile His Lys Leu Met Ile Asp Leu Pro Ala
Asp Ser Ala Phe 225 230 235
240 Leu Gln Ala Asn Leu His Pro Asp Asn Leu Asp Ala Ala Ile Arg Ser
245 250 255 Val Glu Ser
31680DNAKlebsiella pneumoniae 3atggacaaac agtatccggt acgccagtgg
gcgcacggcg ccgatctcgt cgtcagtcag 60ctggaagctc agggagtacg ccaggtgttc
ggcatccccg gcgccaaaat tgacaaggtc 120ttcgactcac tgctggattc ctcgattcgc
attattccgg tacgccacga agccaacgcc 180gcgtttatgg ccgccgccgt cggacgcatt
accggcaaag cgggcgtggc gctggtcacc 240tccggtccgg gctgttccaa cctgatcacc
ggcatggcca ccgcgaacag cgaaggcgac 300ccggtggtgg ccctgggcgg cgcggtaaaa
cgcgccgata aagcgaagca ggtccaccag 360agtatggata cggtggcgat gttcagcccg
gtcaccaaat acgccgtcga ggtgacggcg 420ccggatgcgc tggcggaagt ggtctccaac
gccttccgcg ccgccgagca gggccggccg 480ggcagcgcgt tcgttagcct gccgcaggat
gtggtcgatg gcccggtcag cggcaaagtg 540ctgccggcca gcggggcccc gcagatgggc
gccgcgccgg atgatgccat cgaccaggtg 600gcgaagctta tcgcccaggc gaagaacccg
atcttcctgc tcggcctgat ggccagccag 660ccggaaaaca gcaaggcgct gcgccgtttg
ctggagacca gccatattcc agtcaccagc 720acctatcagg ccgccggagc ggtgaatcag
gataacttct ctcgcttcgc cggccgggtt 780gggctgttta acaaccaggc cggggaccgt
ctgctgcagc tcgccgacct ggtgatctgc 840atcggctaca gcccggtgga atacgaaccg
gcgatgtgga acagcggcaa cgcgacgctg 900gtgcacatcg acgtgctgcc cgcctatgaa
gagcgcaact acaccccgga tgtcgagctg 960gtgggcgata tcgccggcac tctcaacaag
ctggcgcaaa atatcgatca tcggctggtg 1020ctctccccgc aggcggcgga gatcctccgc
gaccgccagc accagcgcga gctgctggac 1080cgccgcggcg cgcagctgaa ccagtttgcc
ctgcatccgc tgcgcatcgt tcgcgccatg 1140caggacatcg tcaacagcga cgtcacgttg
accgtggaca tgggcagctt ccatatctgg 1200attgcccgct acctgtacag cttccgcgcc
cgtcaggtga tgatctccaa cggccagcag 1260accatgggcg tcgccctgcc ctgggctatc
ggcgcctggc tggtcaatcc tgagcgaaaa 1320gtggtctccg tctccggcga cggcggcttc
ctgcagtcga gcatggagct ggagaccgcc 1380gtccgcctga aagccaacgt actgcacctg
atctgggtcg ataacggcta caacatggtg 1440gccattcagg aagagaaaaa ataccagcgc
ctgtccggcg tcgagttcgg gccgatggat 1500tttaaagcct atgccgaatc cttcggcgcg
aaagggtttg ccgtggaaag cgccgaggcg 1560ctggagccga ccctgcacgc ggcgatggac
gtcgacggcc cggcggtggt ggccattccg 1620gtggattatc gcgataaccc gctgctgatg
ggccagctgc atctgagtca gattctgtaa 16804559PRTKlebsiella pneumoniae 4Met
Asp Lys Gln Tyr Pro Val Arg Gln Trp Ala His Gly Ala Asp Leu 1
5 10 15 Val Val Ser Gln Leu Glu
Ala Gln Gly Val Arg Gln Val Phe Gly Ile 20
25 30 Pro Gly Ala Lys Ile Asp Lys Val Phe Asp
Ser Leu Leu Asp Ser Ser 35 40
45 Ile Arg Ile Ile Pro Val Arg His Glu Ala Asn Ala Ala Phe
Met Ala 50 55 60
Ala Ala Val Gly Arg Ile Thr Gly Lys Ala Gly Val Ala Leu Val Thr 65
70 75 80 Ser Gly Pro Gly Cys
Ser Asn Leu Ile Thr Gly Met Ala Thr Ala Asn 85
90 95 Ser Glu Gly Asp Pro Val Val Ala Leu Gly
Gly Ala Val Lys Arg Ala 100 105
110 Asp Lys Ala Lys Gln Val His Gln Ser Met Asp Thr Val Ala Met
Phe 115 120 125 Ser
Pro Val Thr Lys Tyr Ala Val Glu Val Thr Ala Pro Asp Ala Leu 130
135 140 Ala Glu Val Val Ser Asn
Ala Phe Arg Ala Ala Glu Gln Gly Arg Pro 145 150
155 160 Gly Ser Ala Phe Val Ser Leu Pro Gln Asp Val
Val Asp Gly Pro Val 165 170
175 Ser Gly Lys Val Leu Pro Ala Ser Gly Ala Pro Gln Met Gly Ala Ala
180 185 190 Pro Asp
Asp Ala Ile Asp Gln Val Ala Lys Leu Ile Ala Gln Ala Lys 195
200 205 Asn Pro Ile Phe Leu Leu Gly
Leu Met Ala Ser Gln Pro Glu Asn Ser 210 215
220 Lys Ala Leu Arg Arg Leu Leu Glu Thr Ser His Ile
Pro Val Thr Ser 225 230 235
240 Thr Tyr Gln Ala Ala Gly Ala Val Asn Gln Asp Asn Phe Ser Arg Phe
245 250 255 Ala Gly Arg
Val Gly Leu Phe Asn Asn Gln Ala Gly Asp Arg Leu Leu 260
265 270 Gln Leu Ala Asp Leu Val Ile Cys
Ile Gly Tyr Ser Pro Val Glu Tyr 275 280
285 Glu Pro Ala Met Trp Asn Ser Gly Asn Ala Thr Leu Val
His Ile Asp 290 295 300
Val Leu Pro Ala Tyr Glu Glu Arg Asn Tyr Thr Pro Asp Val Glu Leu 305
310 315 320 Val Gly Asp Ile
Ala Gly Thr Leu Asn Lys Leu Ala Gln Asn Ile Asp 325
330 335 His Arg Leu Val Leu Ser Pro Gln Ala
Ala Glu Ile Leu Arg Asp Arg 340 345
350 Gln His Gln Arg Glu Leu Leu Asp Arg Arg Gly Ala Gln Leu
Asn Gln 355 360 365
Phe Ala Leu His Pro Leu Arg Ile Val Arg Ala Met Gln Asp Ile Val 370
375 380 Asn Ser Asp Val Thr
Leu Thr Val Asp Met Gly Ser Phe His Ile Trp 385 390
395 400 Ile Ala Arg Tyr Leu Tyr Ser Phe Arg Ala
Arg Gln Val Met Ile Ser 405 410
415 Asn Gly Gln Gln Thr Met Gly Val Ala Leu Pro Trp Ala Ile Gly
Ala 420 425 430 Trp
Leu Val Asn Pro Glu Arg Lys Val Val Ser Val Ser Gly Asp Gly 435
440 445 Gly Phe Leu Gln Ser Ser
Met Glu Leu Glu Thr Ala Val Arg Leu Lys 450 455
460 Ala Asn Val Leu His Leu Ile Trp Val Asp Asn
Gly Tyr Asn Met Val 465 470 475
480 Ala Ile Gln Glu Glu Lys Lys Tyr Gln Arg Leu Ser Gly Val Glu Phe
485 490 495 Gly Pro
Met Asp Phe Lys Ala Tyr Ala Glu Ser Phe Gly Ala Lys Gly 500
505 510 Phe Ala Val Glu Ser Ala Glu
Ala Leu Glu Pro Thr Leu His Ala Ala 515 520
525 Met Asp Val Asp Gly Pro Ala Val Val Ala Ile Pro
Val Asp Tyr Arg 530 535 540
Asp Asn Pro Leu Leu Met Gly Gln Leu His Leu Ser Gln Ile Leu 545
550 555 5771DNAKlebsiella
pneumoniae 5atgaaaaaag tcgcacttgt taccggcgcc ggccagggga ttggtaaagc
tatcgccctt 60cgtctggtga aggatggatt tgccgtggcc attgccgatt ataacgacgc
caccgccaaa 120gcggtcgcct cggaaatcaa ccaggccggc ggacacgccg tggcggtgaa
agtggatgtc 180tccgaccgcg atcaggtatt tgccgccgtt gaacaggcgc gcaaaacgct
gggcggcttc 240gacgtcatcg tcaataacgc cggtgtggca ccgtctacgc cgatcgagtc
cattaccccg 300gagattgtcg acaaagtcta caacatcaac gtcaaagggg tgatctgggg
tattcaggcg 360gcggtcgagg cctttaagaa agaggggcac ggcgggaaaa tcatcaacgc
ctgttcccag 420gccggccacg tcggcaaccc ggagctggcg gtgtatagct ccagtaaatt
cgcggtacgc 480ggcttaaccc agaccgccgc tcgcgacctc gcgccgctgg gcatcacggt
caacggctac 540tgcccgggga ttgtcaaaac gccaatgtgg gccgaaattg accgccaggt
gtccgaagcc 600gccggtaaac cgctgggcta cggtaccgcc gagttcgcca aacgcatcac
tctcggtcgt 660ctgtccgagc cggaagatgt cgccgcctgc gtctcctatc ttgccagccc
ggattctgat 720tacatgaccg gtcagtcgtt gctgatcgac ggcgggatgg tatttaacta a
7716256PRTKlebsiella pneumoniae 6Met Lys Lys Val Ala Leu Val
Thr Gly Ala Gly Gln Gly Ile Gly Lys 1 5
10 15 Ala Ile Ala Leu Arg Leu Val Lys Asp Gly Phe
Ala Val Ala Ile Ala 20 25
30 Asp Tyr Asn Asp Ala Thr Ala Lys Ala Val Ala Ser Glu Ile Asn
Gln 35 40 45 Ala
Gly Gly His Ala Val Ala Val Lys Val Asp Val Ser Asp Arg Asp 50
55 60 Gln Val Phe Ala Ala Val
Glu Gln Ala Arg Lys Thr Leu Gly Gly Phe 65 70
75 80 Asp Val Ile Val Asn Asn Ala Gly Val Ala Pro
Ser Thr Pro Ile Glu 85 90
95 Ser Ile Thr Pro Glu Ile Val Asp Lys Val Tyr Asn Ile Asn Val Lys
100 105 110 Gly Val
Ile Trp Gly Ile Gln Ala Ala Val Glu Ala Phe Lys Lys Glu 115
120 125 Gly His Gly Gly Lys Ile Ile
Asn Ala Cys Ser Gln Ala Gly His Val 130 135
140 Gly Asn Pro Glu Leu Ala Val Tyr Ser Ser Ser Lys
Phe Ala Val Arg 145 150 155
160 Gly Leu Thr Gln Thr Ala Ala Arg Asp Leu Ala Pro Leu Gly Ile Thr
165 170 175 Val Asn Gly
Tyr Cys Pro Gly Ile Val Lys Thr Pro Met Trp Ala Glu 180
185 190 Ile Asp Arg Gln Val Ser Glu Ala
Ala Gly Lys Pro Leu Gly Tyr Gly 195 200
205 Thr Ala Glu Phe Ala Lys Arg Ile Thr Leu Gly Arg Leu
Ser Glu Pro 210 215 220
Glu Asp Val Ala Ala Cys Val Ser Tyr Leu Ala Ser Pro Asp Ser Asp 225
230 235 240 Tyr Met Thr Gly
Gln Ser Leu Leu Ile Asp Gly Gly Met Val Phe Asn 245
250 255 71665DNAKlebsiella oxytoca
7atgagatcga aaagatttga agcactggcg aaacgccctg tgaatcagga cggcttcgtt
60aaggagtgga tcgaagaagg ctttatcgcg atggaaagcc cgaacgaccc aaaaccgtcg
120attaaaatcg ttaacggcgc ggtgaccgag ctggacggga aaccggtaag cgattttgac
180ctgatcgacc actttatcgc ccgctacggt atcaacctga accgcgccga agaagtgatg
240gcgatggatt cggtcaagct ggccaacatg ctgtgcgatc cgaacgttaa acgcagcgaa
300atcgtcccgc tgaccaccgc gatgacgccg gcgaaaattg tcgaagtggt ttcgcatatg
360aacgtcgtcg agatgatgat ggcgatgcag aaaatgcgcg cccgccgcac cccgtcccag
420caggcgcacg tcaccaacgt caaagataac ccggtacaga ttgccgccga cgccgccgaa
480ggggcatggc gcggatttga cgaacaggaa accaccgttg cggtagcgcg ctatgcgccg
540ttcaacgcca tcgcgctgct ggtgggctcg caggtaggcc gtccgggcgt gctgacgcag
600tgctcgctgg aagaagccac cgagctgaag ctcggcatgc tgggccacac ctgctacgcc
660gaaaccatct ccgtctacgg caccgagccg gtctttaccg acggcgacga cacgccgtgg
720tcgaagggct tcctcgcctc gtcctacgcc tctcgcgggc tgaaaatgcg ctttacctcc
780ggctccggct cggaagtgca gatgggctac gccgaaggca aatccatgct ttatctggaa
840gcgcgctgca tctacatcac caaagccgcg ggcgtacagg gtctgcaaaa cggttccgta
900agctgcatcg gcgtgccgtc tgcggtgcct tccggcattc gcgcggtgct ggcggaaaac
960ctgatctgtt cgtcgctgga tctggagtgc gcctccagca acgaccagac cttcacccac
1020tccgatatgc gtcgtaccgc gcgcctgctg atgcagttcc tgccgggcac cgactttatc
1080tcctccggtt attccgcggt gccgaactac gacaacatgt tcgccggctc caacgaagat
1140gccgaagact ttgacgacta caacgtcatc cagcgcgacc tgaaggtgga cggcggtttg
1200cgtccggttc gcgaagagga cgtcatcgcc atccgtaaca aagccgcccg cgcgctgcag
1260gccgtgtttg ccggaatggg gctgccgccg attaccgatg aagaagttga agccgcgacc
1320tacgcccacg gttcgaaaga tatgccggag cgcaacatcg tcgaagacat caagttcgcc
1380caggaaatca tcaataaaaa ccgcaacggt ctggaagtgg tgaaagcgct ggcgcagggc
1440ggattcaccg acgtggccca ggacatgctc aacatccaga aagctaagct gaccggggac
1500tacctgcata cctccgcgat tatcgtcggc gacgggcagg tgctgtcagc cgtcaacgac
1560gtcaacgact atgccggtcc ggcaacgggc tatcgcctgc agggcgaacg ctgggaagag
1620attaaaaaca tccctggcgc tcttgatccc aacgagattg attaa
16658554PRTKlebsiella oxytoca 8Met Arg Ser Lys Arg Phe Glu Ala Leu Ala
Lys Arg Pro Val Asn Gln 1 5 10
15 Asp Gly Phe Val Lys Glu Trp Ile Glu Glu Gly Phe Ile Ala Met
Glu 20 25 30 Ser
Pro Asn Asp Pro Lys Pro Ser Ile Lys Ile Val Asn Gly Ala Val 35
40 45 Thr Glu Leu Asp Gly Lys
Pro Val Ser Asp Phe Asp Leu Ile Asp His 50 55
60 Phe Ile Ala Arg Tyr Gly Ile Asn Leu Asn Arg
Ala Glu Glu Val Met 65 70 75
80 Ala Met Asp Ser Val Lys Leu Ala Asn Met Leu Cys Asp Pro Asn Val
85 90 95 Lys Arg
Ser Glu Ile Val Pro Leu Thr Thr Ala Met Thr Pro Ala Lys 100
105 110 Ile Val Glu Val Val Ser His
Met Asn Val Val Glu Met Met Met Ala 115 120
125 Met Gln Lys Met Arg Ala Arg Arg Thr Pro Ser Gln
Gln Ala His Val 130 135 140
Thr Asn Val Lys Asp Asn Pro Val Gln Ile Ala Ala Asp Ala Ala Glu 145
150 155 160 Gly Ala Trp
Arg Gly Phe Asp Glu Gln Glu Thr Thr Val Ala Val Ala 165
170 175 Arg Tyr Ala Pro Phe Asn Ala Ile
Ala Leu Leu Val Gly Ser Gln Val 180 185
190 Gly Arg Pro Gly Val Leu Thr Gln Cys Ser Leu Glu Glu
Ala Thr Glu 195 200 205
Leu Lys Leu Gly Met Leu Gly His Thr Cys Tyr Ala Glu Thr Ile Ser 210
215 220 Val Tyr Gly Thr
Glu Pro Val Phe Thr Asp Gly Asp Asp Thr Pro Trp 225 230
235 240 Ser Lys Gly Phe Leu Ala Ser Ser Tyr
Ala Ser Arg Gly Leu Lys Met 245 250
255 Arg Phe Thr Ser Gly Ser Gly Ser Glu Val Gln Met Gly Tyr
Ala Glu 260 265 270
Gly Lys Ser Met Leu Tyr Leu Glu Ala Arg Cys Ile Tyr Ile Thr Lys
275 280 285 Ala Ala Gly Val
Gln Gly Leu Gln Asn Gly Ser Val Ser Cys Ile Gly 290
295 300 Val Pro Ser Ala Val Pro Ser Gly
Ile Arg Ala Val Leu Ala Glu Asn 305 310
315 320 Leu Ile Cys Ser Ser Leu Asp Leu Glu Cys Ala Ser
Ser Asn Asp Gln 325 330
335 Thr Phe Thr His Ser Asp Met Arg Arg Thr Ala Arg Leu Leu Met Gln
340 345 350 Phe Leu Pro
Gly Thr Asp Phe Ile Ser Ser Gly Tyr Ser Ala Val Pro 355
360 365 Asn Tyr Asp Asn Met Phe Ala Gly
Ser Asn Glu Asp Ala Glu Asp Phe 370 375
380 Asp Asp Tyr Asn Val Ile Gln Arg Asp Leu Lys Val Asp
Gly Gly Leu 385 390 395
400 Arg Pro Val Arg Glu Glu Asp Val Ile Ala Ile Arg Asn Lys Ala Ala
405 410 415 Arg Ala Leu Gln
Ala Val Phe Ala Gly Met Gly Leu Pro Pro Ile Thr 420
425 430 Asp Glu Glu Val Glu Ala Ala Thr Tyr
Ala His Gly Ser Lys Asp Met 435 440
445 Pro Glu Arg Asn Ile Val Glu Asp Ile Lys Phe Ala Gln Glu
Ile Ile 450 455 460
Asn Lys Asn Arg Asn Gly Leu Glu Val Val Lys Ala Leu Ala Gln Gly 465
470 475 480 Gly Phe Thr Asp Val
Ala Gln Asp Met Leu Asn Ile Gln Lys Ala Lys 485
490 495 Leu Thr Gly Asp Tyr Leu His Thr Ser Ala
Ile Ile Val Gly Asp Gly 500 505
510 Gln Val Leu Ser Ala Val Asn Asp Val Asn Asp Tyr Ala Gly Pro
Ala 515 520 525 Thr
Gly Tyr Arg Leu Gln Gly Glu Arg Trp Glu Glu Ile Lys Asn Ile 530
535 540 Pro Gly Ala Leu Asp Pro
Asn Glu Ile Asp 545 550 9675DNAKlebsiella
oxytoca 9atggaaatta atgaaaaatt gctgcgccag ataattgaag acgtgctcag
cgagatgaag 60ggcagcgata aaccggtctc gtttaatgcg ccggcggcct ccgcggcgcc
ccaggccacg 120ccgcccgccg gcgacggctt cctgacggaa gtgggcgaag cgcgtcaggg
aacccagcag 180gacgaagtga ttatcgccgt cggcccggct ttcggcctgg cgcagaccgt
caatatcgtc 240ggcatcccgc ataagagcat tttgcgcgaa gtcattgccg gtattgaaga
agaaggcatt 300aaggcgcgcg tgattcgctg ctttaaatcc tccgacgtgg ccttcgtcgc
cgttgaaggt 360aatcgcctga gcggctccgg catctctatc ggcatccagt cgaaaggcac
cacggtgatc 420caccagcagg ggctgccgcc gctctctaac ctggagctgt tcccgcaggc
gccgctgctg 480accctggaaa cctatcgcca gatcggcaaa aacgccgccc gctatgcgaa
acgcgaatcg 540ccgcagccgg tcccgacgct gaatgaccag atggcgcggc cgaagtacca
ggcgaaatcg 600gccattttgc acattaaaga gaccaagtac gtggtgacgg gcaaaaaccc
gcaggaactg 660cgcgtggcgc tttga
67510224PRTKlebsiella oxytoca 10Met Glu Ile Asn Glu Lys Leu
Leu Arg Gln Ile Ile Glu Asp Val Leu 1 5
10 15 Ser Glu Met Lys Gly Ser Asp Lys Pro Val Ser
Phe Asn Ala Pro Ala 20 25
30 Ala Ser Ala Ala Pro Gln Ala Thr Pro Pro Ala Gly Asp Gly Phe
Leu 35 40 45 Thr
Glu Val Gly Glu Ala Arg Gln Gly Thr Gln Gln Asp Glu Val Ile 50
55 60 Ile Ala Val Gly Pro Ala
Phe Gly Leu Ala Gln Thr Val Asn Ile Val 65 70
75 80 Gly Ile Pro His Lys Ser Ile Leu Arg Glu Val
Ile Ala Gly Ile Glu 85 90
95 Glu Glu Gly Ile Lys Ala Arg Val Ile Arg Cys Phe Lys Ser Ser Asp
100 105 110 Val Ala
Phe Val Ala Val Glu Gly Asn Arg Leu Ser Gly Ser Gly Ile 115
120 125 Ser Ile Gly Ile Gln Ser Lys
Gly Thr Thr Val Ile His Gln Gln Gly 130 135
140 Leu Pro Pro Leu Ser Asn Leu Glu Leu Phe Pro Gln
Ala Pro Leu Leu 145 150 155
160 Thr Leu Glu Thr Tyr Arg Gln Ile Gly Lys Asn Ala Ala Arg Tyr Ala
165 170 175 Lys Arg Glu
Ser Pro Gln Pro Val Pro Thr Leu Asn Asp Gln Met Ala 180
185 190 Arg Pro Lys Tyr Gln Ala Lys Ser
Ala Ile Leu His Ile Lys Glu Thr 195 200
205 Lys Tyr Val Val Thr Gly Lys Asn Pro Gln Glu Leu Arg
Val Ala Leu 210 215 220
11522DNAKlebsiella oxytoca 11atgaataccg acgcaattga atcgatggta
cgcgacgtat tgagccgcat gaacagcctg 60cagggcgagg cgcctgcggc ggctccggcg
gctggcggcg cgtcccgtag cgccagggtc 120agcgactacc cgctggcgaa caagcacccg
gaatgggtga aaaccgccac caataaaacg 180ctggacgact ttacgctgga aaacgtgctg
agcaataaag tcaccgccca ggatatgcgt 240attaccccgg aaaccctgcg cttacaggct
tctattgcca aagacgcggg ccgcgaccgg 300ctggcgatga acttcgagcg cgccgccgag
ctgaccgcgg taccggacga tcgcattctt 360gaaatctaca acgccctccg cccctatcgc
tcgacgaaag aggagctgct ggcgatcgcc 420gacgatctcg aaagccgcta tcaggcgaag
atttgcgccg ctttcgttcg cgaagcggcc 480acgctgtacg tcgagcgtaa aaaactcaaa
ggcgacgatt aa 52212173PRTKlebsiella oxytoca 12Met
Asn Thr Asp Ala Ile Glu Ser Met Val Arg Asp Val Leu Ser Arg 1
5 10 15 Met Asn Ser Leu Gln Gly
Glu Ala Pro Ala Ala Ala Pro Ala Ala Gly 20
25 30 Gly Ala Ser Arg Ser Ala Arg Val Ser Asp
Tyr Pro Leu Ala Asn Lys 35 40
45 His Pro Glu Trp Val Lys Thr Ala Thr Asn Lys Thr Leu Asp
Asp Phe 50 55 60
Thr Leu Glu Asn Val Leu Ser Asn Lys Val Thr Ala Gln Asp Met Arg 65
70 75 80 Ile Thr Pro Glu Thr
Leu Arg Leu Gln Ala Ser Ile Ala Lys Asp Ala 85
90 95 Gly Arg Asp Arg Leu Ala Met Asn Phe Glu
Arg Ala Ala Glu Leu Thr 100 105
110 Ala Val Pro Asp Asp Arg Ile Leu Glu Ile Tyr Asn Ala Leu Arg
Pro 115 120 125 Tyr
Arg Ser Thr Lys Glu Glu Leu Leu Ala Ile Ala Asp Asp Leu Glu 130
135 140 Ser Arg Tyr Gln Ala Lys
Ile Cys Ala Ala Phe Val Arg Glu Ala Ala 145 150
155 160 Thr Leu Tyr Val Glu Arg Lys Lys Leu Lys Gly
Asp Asp 165 170
131041DNARhodococcus ruber 13atgaaagccc tccagtacac cgagatcggc tccgagccgg
tcgtcgtcga cgtccccacc 60ccggcgcccg ggccgggtga gatcctgctg aaggtcaccg
cggccggctt gtgccactcg 120gacatcttcg tgatggacat gccggcagag cagtacatct
acggtcttcc cctcaccctc 180ggccacgagg gcgtcggcac cgtcgccgaa ctcggcgccg
gcgtcaccgg attcgagacg 240ggggacgccg tcgccgtgta cgggccgtgg gggtgcggtg
cgtgccacgc gtgcgcgcgc 300ggccgggaga actactgcac ccgcgccgcc gagctgggca
tcaccccgcc cggtctcggc 360tcgcccgggt cgatggccga gtacatgatc gtcgactcgg
cgcgccacct cgtcccgatc 420ggggacctcg accccgtcgc ggcggttccg ctcaccgacg
cgggcctgac gccgtaccac 480gcgatctcgc gggtcctgcc cctgctggga cccggctcga
ccgcggtcgt catcggggtc 540ggcggactcg ggcacgtcgg catccagatc ctgcgcgccg
tcagcgcggc ccgcgtgatc 600gccgtcgatc tcgacgacga ccgactcgcg ctcgcccgcg
aggtcggcgc cgacgcggcg 660gtgaagtcgg gcgccggggc ggcggacgcg atccgggagc
tgaccggcgg tgagggcgcg 720acggcggtgt tcgacttcgt cggcgcccag tcgacgatcg
acacggcgca gcaggtggtc 780gcgatcgacg ggcacatctc ggtggtcggc atccatgccg
gcgcccacgc caaggtcggc 840ttcttcatga tcccgttcgg cgcgtccgtc gtgacgccgt
actggggcac gcggtccgag 900ctgatggacg tcgtggacct ggcccgtgcc ggccggctcg
acatccacac cgagacgttc 960accctcgacg agggacccac ggcctaccgg cggctacgcg
agggcagcat ccgcggccgc 1020ggggtggtcg tcccgggctg a
104114346PRTKlebsiella oxytoca 14Met Lys Ala Leu
Gln Tyr Thr Glu Ile Gly Ser Glu Pro Val Val Val 1 5
10 15 Asp Val Pro Thr Pro Ala Pro Gly Pro
Gly Glu Ile Leu Leu Lys Val 20 25
30 Thr Ala Ala Gly Leu Cys His Ser Asp Ile Phe Val Met Asp
Met Pro 35 40 45
Ala Glu Gln Tyr Ile Tyr Gly Leu Pro Leu Thr Leu Gly His Glu Gly 50
55 60 Val Gly Thr Val Ala
Glu Leu Gly Ala Gly Val Thr Gly Phe Glu Thr 65 70
75 80 Gly Asp Ala Val Ala Val Tyr Gly Pro Trp
Gly Cys Gly Ala Cys His 85 90
95 Ala Cys Ala Arg Gly Arg Glu Asn Tyr Cys Thr Arg Ala Ala Glu
Leu 100 105 110 Gly
Ile Thr Pro Pro Gly Leu Gly Ser Pro Gly Ser Met Ala Glu Tyr 115
120 125 Met Ile Val Asp Ser Ala
Arg His Leu Val Pro Ile Gly Asp Leu Asp 130 135
140 Pro Val Ala Ala Val Pro Leu Thr Asp Ala Gly
Leu Thr Pro Tyr His 145 150 155
160 Ala Ile Ser Arg Val Leu Pro Leu Leu Gly Pro Gly Ser Thr Ala Val
165 170 175 Val Ile
Gly Val Gly Gly Leu Gly His Val Gly Ile Gln Ile Leu Arg 180
185 190 Ala Val Ser Ala Ala Arg Val
Ile Ala Val Asp Leu Asp Asp Asp Arg 195 200
205 Leu Ala Leu Ala Arg Glu Val Gly Ala Asp Ala Ala
Val Lys Ser Gly 210 215 220
Ala Gly Ala Ala Asp Ala Ile Arg Glu Leu Thr Gly Gly Glu Gly Ala 225
230 235 240 Thr Ala Val
Phe Asp Phe Val Gly Ala Gln Ser Thr Ile Asp Thr Ala 245
250 255 Gln Gln Val Val Ala Ile Asp Gly
His Ile Ser Val Val Gly Ile His 260 265
270 Ala Gly Ala His Ala Lys Val Gly Phe Phe Met Ile Pro
Phe Gly Ala 275 280 285
Ser Val Val Thr Pro Tyr Trp Gly Thr Arg Ser Glu Leu Met Asp Val 290
295 300 Val Asp Leu Ala
Arg Ala Gly Arg Leu Asp Ile His Thr Glu Thr Phe 305 310
315 320 Thr Leu Asp Glu Gly Pro Thr Ala Tyr
Arg Arg Leu Arg Glu Gly Ser 325 330
335 Ile Arg Gly Arg Gly Val Val Val Pro Gly 340
345 1529DNAArtificial SequencePrimer 15caccatggac
aaacagtatc cggtacgcc
291625DNAArtificial SequencePrimer 16cgaagggcga tagctttacc aatcc
251732DNAArtificial SequencePrimer
17caccatgaat cattctgctg aatgcacctg cg
321822DNAArtificial SequencePrimer 18gatactgttt gtccatgtga cc
221928DNAArtificial SequencePrimer
19caccatgaaa aaagtcgcac ttgttacc
282015DNAArtificial SequencePrimer 20ttagttaaat accat
152123DNAArtificial SequencePrimer
21caccatgaga tcgaaaagat ttg
232222DNAArtificial SequencePrimer 22cttagagaag ttaatcgtcg cc
222325DNAArtificial SequencePrimer
23caccatgaaa gccctccagt acacc
252418DNAArtificial SequencePrimer 24cgtcgtgtca tgcccggg
182536DNAArtificial SequencePrimer
25gatcgaattc gtttaaactt agttttctac cgcacg
362634DNAArtificial SequencePrimer 26gatcgcatgc aagctttcat atagtcggaa
ttcc 342743DNAArtificial SequencePrimer
27gatcgaattc gtttaaacaa aggaggtctg attcatgaga tcg
432822DNAArtificial SequencePrimer 28gatcggattc ttaatcgtcg cc
222936DNAArtificial SequencePrimer
29gatcggatcc aaaggaggtc gggcgcatga aagccc
363032DNAArtificial SequencePrimer 30gatctctaga aagctttcag cccgggacga cc
323121DNAArtificial SequencePrimer
31actttctttc gcctgtttca c
213279DNAArtificial SequencePrimer 32catgaagctt gtttaaactc ggtgaccttg
aaaataatga aaacttatat tgttttgaaa 60ataatgaaaa cttatattg
793339DNAArtificial SequencePrimer
BABC F 33gagctcgaat tcaaaggagg aagtgtatat gaatcattc
393435DNAArtificial SequencePrimer BAB R 34ggatcctcta gaattagtta
aataccatcc cgccg 353517DNAArtificial
SequencePrimer M13 Forward 35gtaaaacgac ggccagt
173616DNAArtificial SequencePrimer M13 Reverse
36aacagctatg accatg
163720DNAArtificial SequencePrimer N83 SeqF2 37gctggattac cagctcgacc
203820DNAArtificial
SequencePrimer N83SeqF3 38cggacgcatt accggcaaag
203920DNAArtificial SequencePrimer N84 SeqR4
39cgaagcgaga gaagttatcc
204038DNAArtificial SequencePrimer BC Spe F 40actagtaaag gaggaaagag
tatgaagaag gtcgcact 384126DNAArtificial
SequencePrimer BC Xba R 41tctagaaagc aggggcaagc catgtc
264220DNAArtificial SequencePrimer Trc F
42ttgacaatta atcatccggc
204320DNAArtificial SequencePrimer Trc R 43cttctctcat ccgccaaaac
204438DNAArtificial SequencePrimer
DDo For 44aagcttaaag gaggctgatt catgagatcg aaaagatt
384527DNAArtificial SequencePrimer DDo Rev 45tctagattat tcatcctgct
gttctcc 274622DNAArtificial
SequencePrimer DDko seq F2 46gcatggcgcg gatttgacga ac
224722DNAArtificial SequencePrimer DDko seq F5
47cattaaagag accaagtacg tg
224824DNAArtificial SequencePrimer DDko seq F7 48atatcctggt ggtgtcgtcg
gcgt 244922DNAArtificial
SequencePrimer DDko seq F9 49tctttgtcac caacgccctg cg
225022DNAArtificial SequencePrimer DDko seq R1
50gcccaccgcg ctcgccgccg cg
225122DNAArtificial SequencePrimer DDko seq R3 51cccccaggat ggcggcttcg gc
225222DNAArtificial
SequencePrimer DDko seq R7 52gggccgacgg cgataatcac tt
225322DNAArtificial SequencePrimer DDko seq R10
53ttcttcgatc cactccttaa cg
225456DNAArtificial SequencePrimer ChnA F 54catcaattga ctacgtagtc
gtacgtgtaa ggaggtttga aatggaaaaa attatg 565540DNAArtificial
SequencePrimer ChnA R 55catgctagcc ccgggtatct tctactcatt ttttatttcg
405622DNAArtificial SquencePrimer chnSeq F1
56ctcaacaggg tgtaagtgta gt
225722DNAArtificial SequencePrimer chnSeq R1 57cgttttgata tagccaggat gt
225835DNAArtificial
SequencePrimer Top ter F1 58ctagaagtca aaagcctccg accggaggct tttga
355960DNAArtificial SequencePrimer Top ter F2
59ctgctcgagt tgctagcaag tttaaacaaa aaaaagcccg ctcattaggc gggctgagct
606037DNAArtificial SequencePrimer Bot ter R1 60cagcccgcct aatgagcggg
cttttttttg tttaaac 376150DNAArtificial
SequencePrimer Bot ter R2 61ttgctagcaa ctcgagcagt caaaagcctc cggtcggagg
cttttgactt 506222DNAArtificial SequencePrimer pCL1925 vec
F 62cggtatcatc aacaggctta cc
226322DNAArtificial SequencePrimer pCL1925 vec R1 63agggttttcc cagtcacgac
gt 226422DNAArtificial
SequencePrimer pCL1925 vec R2 64cgcaatagtt ggcgaagtaa tc
226520DNAArtificial SequencePrimer N84 Seq R2
65gcatcgagat tatcgggatg
2066208DNAEscherichia coli 66atcgcccgca ttcttgccgc atcttccccc ggcgtcacac
cgaagtaacg tttaaactca 60cggctgtgta ggctggagct gcttcgaagt tcctatactt
tctagagaat aggaacttcg 120gaataggaac taaggaggat attcatatga ttacgttgga
tgtcagccgc cgtatatacg 180aagccgcccg ctaagctttt tacgcctc
2086742DNAArtificial SquencePromoter 1.6GI Variant
67gcccttgaca atgccacatc ctgagcaaat aattcaacca ct
426842DNAArtificial SequencePromoter 1.5 GI 68gcccttgact atgccacatc
ctgagcaaat aattcaacca ct 42693240DNAKlebsiella
oxytoca 69ggcgcggtcc gccaggcggt cacctccgcg cgcgaaatcg gcaaaaccgt
ccttgcgacc 60ctcggtgctg aaccgaaaaa cgatcgcccg tcctacatct gatacccacg
aggctgattc 120atgagatcga aaagatttga agcactggcg aaacgccctg tgaatcagga
cggcttcgtt 180aaggagtgga tcgaagaagg ctttatcgcg atggaaagcc cgaacgaccc
aaaaccgtcg 240attaaaatcg ttaacggcgc ggtgaccgag ctggacggga aaccggtaag
cgattttgac 300ctgatcgacc actttatcgc ccgctacggt atcaacctga accgcgccga
agaagtgatg 360gcgatggatt cggtcaagct ggccaacatg ctgtgcgatc cgaacgttaa
acgcagcgaa 420atcgtcccgc tgaccaccgc gatgacgccg gcgaaaattg tcgaagtggt
ttcgcatatg 480aacgtcgtcg agatgatgat ggcgatgcag aaaatgcgcg cccgccgcac
cccgtcccag 540caggcgcacg tcaccaacgt caaagataac ccggtacaga ttgccgccga
cgccgccgaa 600ggggcatggc gcggatttga cgaacaggaa accaccgttg cggtagcgcg
ctatgcgccg 660ttcaacgcca tcgcgctgct ggtgggctcg caggtaggcc gtccgggcgt
gctgacgcag 720tgctcgctgg aagaagccac cgagctgaag ctcggcatgc tgggccacac
ctgctacgcc 780gaaaccatct ccgtctacgg caccgagccg gtctttaccg acggcgacga
cacgccgtgg 840tcgaagggct tcctcgcctc gtcctacgcc tctcgcgggc tgaaaatgcg
ctttacctcc 900ggctccggct cggaagtgca gatgggctac gccgaaggca aatccatgct
ttatctggaa 960gcgcgctgca tctacatcac caaagccgcg ggcgtacagg gtctgcaaaa
cggttccgta 1020agctgcatcg gcgtgccgtc tgcggtgcct tccggcattc gcgcggtgct
ggcggaaaac 1080ctgatctgtt cgtcgctgga tctggagtgc gcctccagca acgaccagac
cttcacccac 1140tccgatatgc gtcgtaccgc gcgcctgctg atgcagttcc tgccgggcac
cgactttatc 1200tcctccggtt attccgcggt gccgaactac gacaacatgt tcgccggctc
caacgaagat 1260gccgaagact ttgacgacta caacgtcatc cagcgcgacc tgaaggtgga
cggcggtttg 1320cgtccggttc gcgaagagga cgtcatcgcc atccgtaaca aagccgcccg
cgcgctgcag 1380gccgtgtttg ccggaatggg gctgccgccg attaccgatg aagaagttga
agccgcgacc 1440tacgcccacg gttcgaaaga tatgccggag cgcaacatcg tcgaagacat
caagttcgcc 1500caggaaatca tcaataaaaa ccgcaacggt ctggaagtgg tgaaagcgct
ggcgcagggc 1560ggattcaccg acgtggccca ggacatgctc aacatccaga aagctaagct
gaccggggac 1620tacctgcata cctccgcgat tatcgtcggc gacgggcagg tgctgtcagc
cgtcaacgac 1680gtcaacgact atgccggtcc ggcaacgggc tatcgcctgc agggcgaacg
ctgggaagag 1740attaaaaaca tccctggcgc tcttgatccc aacgagattg attaaggggt
gagaaatgga 1800aattaatgaa aaattgctgc gccagataat tgaagacgtg ctcagcgaga
tgaagggcag 1860cgataaaccg gtctcgttta atgcgccggc ggcctccgcg gcgccccagg
ccacgccgcc 1920cgccggcgac ggcttcctga cggaagtggg cgaagcgcgt cagggaaccc
agcaggacga 1980agtgattatc gccgtcggcc cggctttcgg cctggcgcag accgtcaata
tcgtcggcat 2040cccgcataag agcattttgc gcgaagtcat tgccggtatt gaagaagaag
gcattaaggc 2100gcgcgtgatt cgctgcttta aatcctccga cgtggccttc gtcgccgttg
aaggtaatcg 2160cctgagcggc tccggcatct ctatcggcat ccagtcgaaa ggcaccacgg
tgatccacca 2220gcaggggctg ccgccgctct ctaacctgga gctgttcccg caggcgccgc
tgctgaccct 2280ggaaacctat cgccagatcg gcaaaaacgc cgcccgctat gcgaaacgcg
aatcgccgca 2340gccggtcccg acgctgaatg accagatggc gcggccgaag taccaggcga
aatcggccat 2400tttgcacatt aaagagacca agtacgtggt gacgggcaaa aacccgcagg
aactgcgcgt 2460ggcgctttga taaaggataa ctccatgaat accgacgcaa ttgaatcgat
ggtacgcgac 2520gtattgagcc gcatgaacag cctgcagggc gaggcgcctg cggcggctcc
ggcggctggc 2580ggcgcgtccc gtagcgccag ggtcagcgac tacccgctgg cgaacaagca
cccggaatgg 2640gtgaaaaccg ccaccaataa aacgctggac gactttacgc tggaaaacgt
gctgagcaat 2700aaagtcaccg cccaggatat gcgtattacc ccggaaaccc tgcgcttaca
ggcttctatt 2760gccaaagacg cgggccgcga ccggctggcg atgaacttcg agcgcgccgc
cgagctgacc 2820gcggtaccgg acgatcgcat tcttgaaatc tacaacgccc tccgccccta
tcgctcgacg 2880aaagaggagc tgctggcgat cgccgacgat ctcgaaagcc gctatcaggc
gaagatttgc 2940gccgctttcg ttcgcgaagc ggccacgctg tacgtcgagc gtaaaaaact
caaaggcgac 3000gattaacttc tctaagtaat tcgagatgca ttgaggcggc aagtgagtga
caaattcgtc 3060tggaacgaat ttgaacagcc ataggctggc tttagtgagg gacagggatg
tccctcataa 3120ccccgatgag cttactgtag taagtgattc gggtgaaaga acgcagccaa
caaaaaggca 3180gtttgaagta cgacgagaaa aggggcatgt gatgcgatat atagctggca
ttgatatcgg 3240702640DNAKlebsiella oxytoca 70acgtcgagcg taaaaaactc
aaaggcgacg attaacttct ctaagtaatt cgagatgcat 60tgaggcggca agtgagtgac
aaattcgtct ggaacgaatt tgaacagcca taggctggct 120ttagtgaggg acagggatgt
ccctcataac cccgatgagc ttactgtagt aagtgattcg 180ggtgaaagaa cgcagccaac
aaaaaggcag tttgaagtac gacgagaaaa ggggcatgtg 240atgcgatata tagctggcat
tgatatcggc aactcatcga cggaagtcgc cctggcgacc 300ctggatgagg ctggcgcgct
gacgatcacc cacagcgcgc tggcggaaac caccggaatc 360aaaggcacgt tgcgtaacgt
gttcgggatt caggaggcgc tcgccctcgt cgccagaggc 420gccgggatcg ccgtcagcga
tatttcgctc atccgcatca acgaagcgac gccggtgatt 480ggcgatgtgg cgatggaaac
cattaccgaa accatcatca ccgaatcgac catgatcggc 540cataacccga aaacgcccgg
cggcgcgggg cttggcacag gcatcaccat tacgccgcag 600gagctgctaa cccgcccggc
ggacgcgccc tatatcctgg tggtgtcgtc ggcgttcgat 660tttgccgata tcgccagcgt
gattaacgct tccctgcgcg ccgggtatca gattaccggc 720gtcattttac agcgcgacga
tggcgtgctg gtcagcaacc ggctggaaaa accgctgccg 780atcgttgacg aagtgctgta
catcgaccgc attccgctgg ggatgctggc ggcgattgag 840gtcgccgttc cggggaaggt
catcgaaacc ctctctaacc cttacggcat cgccaccgtc 900tttaacctca gccccgagga
gacgaagaac atcgtcccga tggcccgggc gctgattggc 960aaccgttccg ccgtggtggt
caaaacgcca tccggcgacg tcaaagcgcg cgcgataccc 1020gccggtaatc ttgagctgct
ggcccagggc cgtagcgtgc gcgtggatgt ggccgccggc 1080gccgaagcca tcatgaaagc
ggtcgacggc tgcggcaggc tcgataacgt caccggcgaa 1140tccggcacca atatcggcgg
catgctggaa cacgtgcgcc agaccatggc cgagctgacc 1200aacaagccga gcagcgaaat
atttattcag gacctgctgg ccgttgatac ctcggtaccg 1260gtgagcgtta ccggcggtct
ggccggggag ttctcgctgg agcaggccgt gggcatcgcc 1320tcgatggtga aatcggatcg
cctgcagatg gcaatgatcg cccgcgaaat cgagcagaag 1380ctcaatatcg acgtgcagat
cggcggcgca gaggccgaag ccgccatcct gggggcgctg 1440accacgccgg gcaccacccg
accgctggcg atcctcgacc tcggcgcggg ctccaccgat 1500gcctccatca tcaaccccaa
aggcgacatc atcgccaccc atctcgccgg cgcaggcgac 1560atggtgacga tgattattgc
ccgcgagctg gggctggaag accgctatct ggcggaagag 1620atcaagaagt acccgctggc
taaggtggaa agcctgttcc atttacgcca cgaggacggc 1680agcgtgcagt tcttctccac
gccgctgccg cccgccgtgt tcgcccgcgt ctgcgtggtg 1740aaagcggacg aactggtgcc
gctgcccggc gatttagcgc tggaaaaagt gcgcgccatt 1800cgccgcagcg ccaaagagcg
ggtctttgtc accaacgccc tgcgcgcgct gcgtcaggtc 1860agccccaccg gcaacattcg
cgatattccg ttcgtggtgc tggtcggcgg ttcgtcgctg 1920gatttcgaag tcccgcagct
ggtcaccgat gcgctggcgc actaccgcct ggttgccgga 1980cggggaaata ttcgcggcag
cgagggcccc cgaaacgcgg tggccaccgg cctgattctc 2040tcctggcata aggagtttgc
gcatgaacgg taatcacagc gccccggcca tcgcgatcgc 2100cgtcatcgac ggctgcgacg
gcctgtggcg cgaagtgctg ctgggtatcg aagaggaagg 2160tatccctttc cggctccagc
atcacccggc cggagaggtc gtggacagcg cctggcaggc 2220ggcgcgcagc tcgccgctgc
tggtgggcat cgcctgcgac cgccatatgc tggtcgtgca 2280ctacaagaat ttacccgcat
cggcgccgct ttttacgctg atgcatcatc aggacagtca 2340ggcccatcgc aacaccggta
ataacgcggc acggctggtc aaggggatcc ctttccggga 2400tctgaatagc gaagcaacag
gagaacagca ggatgaataa cgcactggga ctggttgaaa 2460caaaagggtt agtgggcgcc
attgaggccg ccgatgcgat ggtgaaatcc gccaacgtgc 2520agctggtcgg ctacgaaaaa
attggctcgg gcctcgtcac cgtgatggtg cgcggcgacg 2580tcggcgcggt caaagcggcg
gtagacgcgg gcagcgcggc ggcgagcgcg gtgggcgaag 264071756DNAAcinetobacter
sp. 71atggaaaaaa ttatgtcaaa taaattcaac aataaagtcg ctttaattac tggcgctggt
60tcaggtattg gtaaaagcac cgcactgctt ttggctcaac agggtgtaag tgtagtggtt
120tcagatatta acctggaagc agcacagaaa gttgtggacg aaattgtcgc tttaggcggg
180aaagcggctg cgaataaggc caatactgct gagcctgaag acatgaaagc tgcagtcgag
240tttgcggtca gcacttttgg tgcactgcat ttggccttca ataatgcggg aattctgggt
300gaagttaact ccaccgaaga attgagcatt gaaggatggc gtcgtgtgat tgatgtgaac
360ttgaatgcgg ttttctacag catgcattat gaagttcctg caatcttggc cgcagggggc
420ggagcgattg tcaataccgc ttctattgca ggcttgatcg ggattcaaaa tatttcaggc
480tatgtcgctg caaaacatgg cgtaacgggt ctaacgaaag cggcggcatt ggaatatgca
540gataaaggga ttcgcattaa ttcagtacat cctggctata tcaaaacgcc tttgattgca
600gaatttgaag aagcagaaat ggtaaaacta catccgattg gtcgtttggg acagccggaa
660gaagttgctc aggttgttgc cttcctactt tctgatgatg cttcatttgt gaccggtagt
720cagtatgtgg tcgatggtgc atatacctcg aaataa
75672251PRTAcinetobacter sp. 72Met Glu Lys Ile Met Ser Asn Lys Phe Asn
Asn Lys Val Ala Leu Ile 1 5 10
15 Thr Gly Ala Gly Ser Gly Ile Gly Lys Ser Thr Ala Leu Leu Leu
Ala 20 25 30 Gln
Gln Gly Val Ser Val Val Val Ser Asp Ile Asn Leu Glu Ala Ala 35
40 45 Gln Lys Val Val Asp Glu
Ile Val Ala Leu Gly Gly Lys Ala Ala Ala 50 55
60 Asn Lys Ala Asn Thr Ala Glu Pro Glu Asp Met
Lys Ala Ala Val Glu 65 70 75
80 Phe Ala Val Ser Thr Phe Gly Ala Leu His Leu Ala Phe Asn Asn Ala
85 90 95 Gly Ile
Leu Gly Glu Val Asn Ser Thr Glu Glu Leu Ser Ile Glu Gly 100
105 110 Trp Arg Arg Val Ile Asp Val
Asn Leu Asn Ala Val Phe Tyr Ser Met 115 120
125 His Tyr Glu Val Pro Ala Ile Leu Ala Ala Gly Gly
Gly Ala Ile Val 130 135 140
Asn Thr Ala Ser Ile Ala Gly Leu Ile Gly Ile Gln Asn Ile Ser Gly 145
150 155 160 Tyr Val Ala
Ala Lys His Gly Val Thr Gly Leu Thr Lys Ala Ala Ala 165
170 175 Leu Glu Tyr Ala Asp Lys Gly Ile
Arg Ile Asn Ser Val His Pro Gly 180 185
190 Tyr Ile Lys Thr Pro Leu Ile Ala Glu Phe Glu Glu Ala
Glu Met Val 195 200 205
Lys Leu His Pro Ile Gly Arg Leu Gly Gln Pro Glu Glu Val Ala Gln 210
215 220 Val Val Ala Phe
Leu Leu Ser Asp Asp Ala Ser Phe Val Thr Gly Ser 225 230
235 240 Gln Tyr Val Val Asp Gly Ala Tyr Thr
Ser Lys 245 250
7317417DNAAcinetobacter sp. 73ctagcattta cgcgtgaggt aggtgggtag gtctgtaatg
tgaagatcta cgaggaaatc 60ggcgtcatga cgtgaggtcc agcgaaccgt cttgcgtaat
ccgtcattca tggtgagtaa 120cattgcccgt atttcgcgtt cagtatatag cagaccagca
tgattaacga gatcctgggt 180attttagtcc ggacacccaa agtcccatgc ggtcgccaga
tccagtaagt cgactacgac 240ttgctcatct gtagccaacc ccgcaatcac ttccacaatt
ttcatcagtg gaaccggatt 300gaagaaatgg aaacctgcga tacggccctg atgctgacac
gcagatgcaa ttgaggtcac 360agatagtgag gatgtatttg aaaccagaat agtttcttca
gccacaatcc tttcaagctg 420tttaaacaaa gtttgcttga tttccagatt ttcaataatt
gcttctacga ccagatcaac 480gccagcaacc tcttcaatgc tttccaagat aatcaatcgg
gctaaggtat ccacaagctg 540ctgttcggtt aactttcctt tagcagctag tttgtgcaag
gttactttta atttttccaa 600gccttgctca gcagcgccgg gtttagcatc aaataaacgg
acctcaacac ccgcctgtgc 660tgcaatttgc gcaataccca ttcccattac gcctgtgcca
atcaaggcca ttttttgaat 720cgtcatgact tattttcctt gatattgagg gcttcgcttt
tcgaaaaagg cattgacgcc 780ttctttttga tcttgtgtat caaataaaat ttggaaggct
ttacgctcta atgccaaagc 840accatcgagt ggcatattgg cacctagtgt tgtgacttct
ttgatctgtt caacggcaat 900cggtgagagt tgggcaatct gtgtcgcaat ttcaaccgct
ttagcaaggg tttgatcatc 960ctcaaccact tcggaaacca accccatttt gtcagcttct
tctgcagaaa agatctttcc 1020tgttaacact atttgcatgg ctttaaactt ccctaccgca
cgcagtaagc gttgggtacc 1080accagcacct ggcatcagcc ccaatttgac ttcaggctga
ccaaactggg ctgattttcc 1140ggcaataatg atgtctgcat gcattgcaag ttcacaccca
ccacccaatg catatccatt 1200cacagcagcc acaatcggtt tagggcaatc aataatggcc
cgccagtact gttccgtatg 1260gcgtaaatac atgtctacgg tttttgcagt ggtgaagtcc
cggatatccg cacctgctgc 1320aaatactttt tcaccaccag taatgacaat tgcgcggact
gtatcagatg cagcgagctg 1380ctcaaacatt gctgcgagct gttggcgcag ttccagattc
aatgcatttc tagtatctgg 1440acgatgtagt tcaacaatgg ccacaccatt actttgaata
tctaaattca atatttcatt 1500ttccataaca acctacatgt ttcgcatagc ggtttattta
aaccaaatat acctgttttt 1560ttgcaacaat aaagcccaca ggaacatagt tttaaattaa
aaattggcta aaaatattta 1620aaaaacacaa ataaaatacc gcacagcggt atttgatatc
aatattattg catttatttt 1680tccattctgt catattattt tcattccaaa gcattagatc
acccctgcat gaagcagaga 1740tggctaaatt tacctatcta atacaagggc ttaaaaatga
ttcgcgatca agacacatta 1800aatcagctgg ttgacatgat ccgtcagttt gtcgatggcg
ttcttattcc caatgaagaa 1860attgttgcgg aaaccgatga aattccagct gaaatcgtgc
agcaaatgaa agaactgggt 1920ctttttggtc tcaccattcc tgaggaatat gagggtcttg
gcctgaccat ggaggaagag 1980gtttacattg catttgaact gggacgtacc tctcctgctt
tccgttcact gatcggcact 2040aacaatggga tcggttcatc aggcttaatt attgatggct
ccgaagagca gaaacagtat 2100tttttgccac gtctggcaag tggtgaaatt attggttcat
tctgtttaac tgaacctgat 2160tccggttcag atgctgcctc tttaaaaacc acagcggtga
aagatggtga tcattacatt 2220ttaaatggca ctaagcgtta catcaccaat gcaccgcatg
cgggtgtctt tactgtcatg 2280gcacgtacca gtaccgaaat taaaggtaca ggtggaattt
cagcctttat cgtggacagt 2340aaaactcctg gtatttcctt gggtaaacgt gataagaaga
tgggccaaaa aggtgcacat 2400acctgtgatg tgatttttga aaactgtcgt attcctgcat
ctgcactcat tggtggtgtt 2460gaaggtgtag gttttaaaac tgcaatgaag gtacttgata
aaggccgtat tcatattgct 2520gcattaagtg taggtgctgc tacgcgtatg ctggaagatt
ccctacaata tgccgttgag 2580cgcaaacagt ttggtcaagc gattgcgaac ttccagttga
ttcaaggtat gttagccgat 2640tctaaagctg aaatttacgc agcaaaatgt atggtattag
atgctgcccg acttcgtgat 2700gctggacaga atgtcagcac ggaagcatct tgtgccaaga
tgtttgccac tgaaatgtgt 2760ggccgtgtcg cagatcgtgg cgtacagatc catggtggtg
cgggttatat cagtgaatat 2820gctattgagc gtttttaccg tgatgtacgt ttattccgtt
tgtatgaagg tacaacgcaa 2880atccaacagg tcattattgc ccgcaatatg atccgtgaag
cgactcaata attgtataac 2940aggtattgag tgtatctaaa aggacgggat tagtgattta
agctataact tgaatactaa 3000tcctgacttt ttgatggcaa ggctataaaa cctcctagct
cattttatct ctaagctaat 3060cacagctgaa agatattttc agtcttcatc cttaccagac
agttcacaat acaaaattgg 3120attttatgaa tatgcaagaa caagaaatcg aacgcgaatc
aatggagttt gacgtcgtga 3180ttgtcggcgc aggaccggcc ggtctttctg cagcgatcaa
gatccgtcaa cttgcaattg 3240aaaacaacct gaacgatctg tcggtttgtg tggtggaaaa
aggctctgaa gtcggtgcgc 3300acatcttgtc cggtgcggta ctggaaccac gtgccatgaa
tgagctgttc ccgaactgga 3360aggaagaagg tgcaccttta aatgttccag tgaccgaaga
caagacctat ttcctgctct 3420cggatgaaaa atcacaagaa gcgccacact ggatggtgcc
taaaaccatg cataacgatg 3480gcaactatgt tatctcgctc ggcaacgtag tgcgctggtt
gggtcaaaaa gcggaagagc 3540tggaagtatc tattttcccg ggctttgccg ctgctgaaat
tctgtaccat gcagatggtt 3600cggtgaaagg cattcaaacc ggtgacatgg gcattggcaa
ggatggcgaa ccgacccata 3660actttactcc gggctatgaa ctgcatgcca aatacaccct
gtttgctgaa ggctgccgtg 3720gccacctcgg caagcgttta attgccaaat acaacctcga
taaagattca gatccacaac 3780attacggtat cggtatcaaa gagctgtggg aaatcgaccc
ggcgaaacac aagccaggtc 3840tggtgatgca cggtgccggc tggccattgt ctgaaaccgg
ttcttcaggc ggctggtggt 3900tgtatcatgc ggaaaacaat caggtgactt tgggcatgat
cgtcgatctg tcttacacca 3960acccgcatat gtatccgttt atggaaatgc agcgctggaa
aacccatccg ctgatcaagc 4020agtatctgga aggtggcaaa cgtatttctt atggcgcgcg
tgcggtaacc aaaggcggct 4080ttaactcgct accgaaattt accttcccgg gcggatcgct
gattggtgac gatgccggct 4140tcctgaactt tgccaaaatc aagggctcac ataccgcgat
gaaatccggc atgctctgcg 4200gtgaagcagt gtttgaagcc attgctgccg gtgtggaaaa
aggtggtgac cttgcggttg 4260cgcgtgtgac ggaaggcgaa gacttgtttg ccaaaaaact
gacttcttac accgacaagt 4320tcaataatag ctggctgaaa gaagagctgt acaactcgcg
taactttggc ccggccatgc 4380acaagtttgg tcagtggctc ggtggtgcgt ttaactttat
cgaccagaac gtgtttaagg 4440tgccgtttac cctgcatgac ctggtgacgg atttcggtgc
gctgaaaacc gtcgatgcgg 4500tgaacttcaa gccgaattat ccaaaaccgg atggcaaact
gacctttgac cgtctgtctt 4560cggtgtttgt atccaacacg gtgcatgaag aaaaccagcc
agcgcattta aaactgactg 4620acacttcgat tccggtgaat gtcaacctgc caaaatggga
tgaaccggcg cagcgctact 4680gccccgcggg tgtatacgaa atcatggaaa atgatgacgg
ttcgaaacgc ttccagatca 4740atgcagccaa ctgtgtgcac tgcaagacct gtgacatcaa
ggatccttca cagaacatca 4800cctgggtaac accggaaggt ggtggtggtc caaactatcc
gaatatgtaa gtctaatcac 4860ttcaaggaag aggtttccca tttcccttct ttctagcaga
tgaagaagct tgcaactaaa 4920agagattgtt tggatcagtt acccaaaatc gttgaaaaga
ttttaactct tcgattttta 4980ttttttaggt aatcctagcc ctctcggggg ctaggattaa
aaattttaag ttattccaac 5040acgaatgaca aattgttcaa tgcaaaataa aaacatacaa
tatataaata tattttttaa 5100ttaaaacata agattacaat aaaataagaa tttttatttg
gagtttgttt tttttctaca 5160atgatcatta tgtacaattt ttaggttcac cccatccaag
ccttgtgatt gcattcctgc 5220gattctttat tcaatgaata agcaatgcta ttaatcagca
atgaataacc agcactgcag 5280attttgaata aattcacatg tcgtaatgga gattatcatg
tcacaaaaaa tggattttga 5340tgctatcgtg attggtggtg gttttggcgg actttatgca
gtcaaaaaat taagagacga 5400gctcgaactt aaggttcagg cttttgataa agccacggat
gtcgcaggta cttggtactg 5460gaaccgttac ccaggtgcat tgtcggatac agaaacccac
ctctactgct attcttggga 5520taaagaatta ctacaatcgc tagaaatcaa gaaaaaatat
gtgcaaggcc ctgatgtacg 5580caagtattta cagcaagtgg ctgaaaagca tgatttaaag
aagagctatc aattcaatac 5640cgcggttcaa tcggctcatt acaacgaagc agatgccttg
tgggaagtca ccactgaata 5700tggtgataag tacacggcgc gtttcctcat cactgcttta
ggcttattgt ctgcgcctaa 5760cttgccaaac atcaaaggca ttaatcagtt taaaggtgag
ctgcatcata ccagccgctg 5820gccagatgac gtaagttttg aaggtaaacg tgtcggcgtg
attggtacgg gttccaccgg 5880tgttcaggtt attacggctg tggcacctct ggctaaacac
ctcactgtct tccagcgttc 5940tgcacaatac agcgttccaa ttggcaatga tccactgtct
gaagaagatg ttaaaaagat 6000caaagacaat tatgacaaaa tttgggatgg tgtatggaat
tcagcccttg cctttggcct 6060gaatgaaagc acagtgccag caatgagcgt atcagctgaa
gaacgcaagg cagtttttga 6120aaaggcatgg caaacaggtg gcggtttccg tttcatgttt
gaaactttcg gtgatattgc 6180caccaatatg gaagccaata tcgaagcgca aaatttcatt
aagggtaaaa ttgctgaaat 6240cgtcaaagat ccagccattg cacagaagct tatgccacag
gatttgtatg caaaacgtcc 6300gttgtgtgac agtggttact acaacacctt taaccgtgac
aatgtccgtt tagaagatgt 6360gaaagccaat ccgattgttg aaattaccga aaacggtgtg
aaactcgaaa atggcgattt 6420cgttgaatta gacatgctga tatgtgccac aggttttgat
gccgtcgatg gcaactatgt 6480gcgcatggac attcaaggta aaaacggctt ggccatgaaa
gactactgga aagaaggtcc 6540gtcgagctat atgggtgtca ccgtaaataa ctatccaaac
atgttcatgg tgcttggacc 6600gaatggcccg tttaccaacc tgccgccatc aattgaatca
caggtggaat ggatcagtga 6660taccattcaa tacacggttg aaaacaatgt tgaatccatt
gaagcgacaa aagaagcgga 6720agaacaatgg actcaaactt gcgccaatat tgcggaaatg
accttattcc ctaaagcgca 6780atcctggatt tttggtgcga atatcccggg caagaaaaac
acggtttact tctatctcgg 6840tggtttaaaa gaatatcgca gtgcgctagc caactgcaaa
aaccatgcct atgaaggttt 6900tgatattcaa ttacaacgtt cagatatcaa gcaacctgcc
aatgcctaaa tatatggggg 6960gcatccccca tattccattt tgtttaacat cagtcatatg
ccagggatgt cttatcatga 7020actatccaaa tataccttta tatatcaacg gtgagtttct
agatcatacc aatagagacg 7080tcaaagaagt ttttaatcca gtgaaccatg aatgtattgg
actcatggcc tgtgcatcac 7140aagcagacct ggactacgca cttgaaagtt cacaacaggc
ttttctaagg tggaaaaaaa 7200cttctcctat cacccgtagt gaaatcctca gaacctttgc
gaaactagcg cgtgaaaaag 7260cagcagaaat cgggcgcaat attacccttg atcaaggtaa
gcccctgaaa gaagccattg 7320cagaagtcac tgtctgtgca gaacatgcag aatggcatgc
agaagaatgc cgacgcattt 7380atggccgtgt tattccaccg cgtaacccaa atgtacagca
actagtagtc agagaaccgc 7440tgggcgtatg tctggcattt tcaccgtgga atttcccgtt
taatcaggca attcgtaaaa 7500tttctgctgc aattgctgcc ggctgcacca tcattgtgaa
aggttctggc gacacaccaa 7560gcgcggtata tgcgattgcc cagctatttc atgaggcggg
tttgccgaat ggtgtgctga 7620atgtgatttg gggtgactca aacttcattt ctgattacat
gatcaaatcg ccgatcatcc 7680aaaagatttc attcacaggc tcaaccccgg tgggtaaaaa
attagcctcg caagcgagtc 7740tgtatatgaa gccttgcacc atggaattgg gtggtcatgc
accggtcatc gtctgtgatg 7800atgctgatat tgatgccgct gttgaacatc tggtcggtta
taaattccgt aatgcaggac 7860aggtctgtgt atcaccaacc cgtttttatg tgcaggaagg
tatttataag gaattttctg 7920agaaagtggt gttaagagcc aaacagatca aagtgggttg
tggcttagac gcatcctcag 7980atatgggacc attggctcaa gctcgccgca tgcatgcaat
gcaacaaatt gttgaagatg 8040cggttcataa aggctcaaaa ttactgcttg gcggaaataa
aatttctgac aaaggcaatt 8100tttttgaacc aacggtactc ggtgacttgt gcaatgacac
ccagtttatg aatgacgagc 8160catttggtcc gatcattggt ttgatacctt ttgacacaat
agaccatgtc ctggaagaag 8220caaatcgatt accatttgga ttagcctctt acgcttttac
cacatccagc aaaaatgcgc 8280atcaaatctc atacggactg gaggctggca tggtttcgat
taaccacatg ggattggcgc 8340tcgctgaaac accttttggt ggtattaagg atagcggttt
tggtagtgaa gggggtatcg 8400aaacctttga cggttacctc agaaccaaat ttattacgca
actcaattag aaatggatct 8460tggtgtgcgt aggcacacca attctctttt gactttaagg
atgaaagtta aatgagcaca 8520gacaaagcaa atacgctgat caaacccgaa gatgtcgtgt
tatggattcc gggtaatgtc 8580acaattgaca gcatgaatgc cggttgggaa aacattgcaa
tcagagggta cgaatatacc 8640aacctcgatg tgcatattcc tgccatgcgt gactacatga
tcgtcaacta taaaaaaagt 8700gcggcggaaa tgcgtagaaa aggcgatgcc tcttgggata
cccaagtggt taagccgggt 8760tatgtctcct tgttgacctg tggtgaagat tcccgctggg
cgtggaatga ccatattgcc 8820gtcacccatg tctacatttc gcatgactcc atcacctcaa
tggcgaataa ggtgtttgat 8880tatgatatcg cttcgatccg aatcagagac gaagtcggtg
tggaagatca tgttttacct 8940gctctgactt cacttttaga actagaatta aagcaaggtg
gtttaggtgg aaacctgtat 9000ttagagagca ttaaaaacca gatcgccctg catttactcc
gtcagtatgc caaattagat 9060tttaaggaag gacagtgccg ttctggtttt actcccctac
aacgcagact gttattagaa 9120tttatcaatg aaaacatgag cattaaaatt accctcgaag
atttagcggg attagtcaag 9180atgagcgtgc ctcatttaat gagaaaattt aaagtcgatt
ttggtaattc ccctgctgcc 9240tacatcatga atctcagggt gcaatttgct aaacgtttgc
tcacttcaaa aaaagaaatt 9300ccactgaaag tgattgccag tgaagccggt ttttgcgatc
agagccatat gacccgagta 9360tttcaaaaat tttttgggaa aacacccatc gaaatcagac
aggaacacac caatctcgtg 9420tctgaaaatt cagtctcctc tattgttttt tgagtactaa
gagccacgca agaacctgat 9480tttcaataaa gcatccactg aaaaccagtg tggacttaca
tgcattattt atgcaaaata 9540acaaatgtca tgtgagtatc aagatatact ttctatcgct
atcaagaact tgccagtaca 9600ggcaatatgg atgcactcat caaccagagt cgcagaactc
caaatttaaa aaaccgagtg 9660gatgagcaaa ctgaataagc tgttgttgat tttgcaatcc
aatatccagc ttatggtcag 9720catcggacca gtaatgagct acgtcagatt ggcatcttcg
tatctggcag cggtgtgcgc 9780tctatctggc ttagacacaa tcttgagaat ttcaaaaagc
gattaaaggc acttgaaatt 9840aaagttgctc aagaaggcat tcagttgaat gatcagcaga
ttgccgcatt agaacgtaaa 9900catgaagatg atgttgcttg tggtgaaatt gaaacacatc
atccaggtta ccttggagca 9960caagatactt tttatgtcgg aaatctaaaa ggtgttgggc
atatttatca gcaaactttt 10020attgatactt atagcaaagt ggttcactgc aagctgtaca
caaccaagac accaatcaca 10080gccgcagatt tattgaatga ccgcgtgtta ccattctatg
agtcacaagg attgccaatg 10140cttcgcattt tgaccgacag aggcaccgaa tattgcggta
aagttgaaca tcacgattat 10200gagctttatt tggctctgaa tgatattgat cacactaaaa
ctaaagcagc atcaccacaa 10260acaaatggga tctgtgagcg cttccataag acgatcttgc
aggagtttta tcagattact 10320tttcgaaaga aactctatag ctcattagaa gagttacagc
ttgatctaga cggttggctg 10380aaattctata atactgaacg aacccatcag ggtaaggtgt
gtaatggcag atgagcagca 10440ttgctgcgca agattgcaac attacttgat ggaaaacgta
tttgggctga aaagaattta 10500gttcaaattt aacctgacag tcttaagcaa atatcggtaa
ctatcagatc aggtttgaga 10560taccgtctga aacgtcaagt aaatgattga gaattcatgc
tcaataatct gcttgataag 10620gctgttggtg tttgagcaca ccataacaaa gatgaatcaa
cttcctcatc gcggctccaa 10680tcgctatcat cttggtttta ccattcgcca ataaacgttc
attcattgcc ctgatgtgag 10740ggttatgccg agttgcgaca atggctgcca tatataaacc
agcacgtatt ttggaagagc 10800ccgctttgga taaacggctt ctgccatgaa tggaactacc
cgattgcttt tgaatgggga 10860ccaaaccgac aaaggcagcc gcttgactag ccctttcaaa
agtatggctg cgcaagaaac 10920tgagcattaa taaactggtt cgatctgcaa tggctggaat
actgctgagc agttctttat 10980cattttttaa atcaggattc tgattaatgt gatcatcaat
ttgctggtcg ataccctgaa 11040tgtgtttgtt taactgttca atactcttgt ggatagactg
aagtacaggt tccatcgtga 11100aggtcgactc tgctttttcc aaacgattct tttcacgttg
taaatcttca caaagaatag 11160ctcttctatc cagcaaagca ttcagcaatt gaatatgttt
aggtaaaggt tgccaaaaat 11220gtagatcggc agtcatcgca aatcgagcta ggacctcact
atccaccttg tctgttttat 11280tcagcttaga catactctga gcaaaatatc gagctctggc
aggattggtt acacagactt 11340gatagcccgc atcaaataaa tatttaacca agagttcatg
ataaatagat gttgcttcca 11400ttaaaataat ggtctgcgta gaagttgcag catgctgctt
tagccaggtt tgaagttgct 11460caaaaccttt tggtgtattt gaaaaagttt tggttttctt
tttatttgca gaattttcta 11520aaattaaaca gcaatcaatt ttagctttag caacatcaat
accaagataa aacataatct 11580ttacctgctt tatttatcca attattgttt tagcataacc
accgtctttt cttgtgaatg 11640cagcatcaaa gtgcttgtta ccgtccagag ttgtgcaagt
ggttagggca aattacaggt 11700tttatctcaa actctaactt tatgttttgc tagtacacga
aactctgcaa tttgcaatat 11760agtgatagct aatcactatg aatggtaaga tacaagctag
tacacataag aagatattac 11820ttcttctcag gcagattcgc agcaaagaaa aattttccct
tacaacaata gataaaagaa 11880aagagggtat cacccctctt tcctctttat atgggggtat
cttctactca ttttttattt 11940cgaggtatat gcaccatcga ccacatactg actaccggtc
acaaatgaag catcatcaga 12000aagtaggaag gcaacaacct gagcaacttc ttccggctgt
cccaaacgac caatcggatg 12060tagttttacc atttctgctt cttcaaattc tgcaatcaaa
ggcgttttga tatagccagg 12120atgtactgaa ttaatgcgaa tccctttatc tgcatattcc
aatgccgccg ctttcgttag 12180acccgttacg ccatgttttg cagcgacata gcctgaaata
ttttgaatcc cgatcaagcc 12240tgcaatagaa gcggtattga caatcgctcc gccccctgcg
gccaagattg caggaacttc 12300ataatgcatg ctgtagaaaa ccgcattcaa gttcacatca
atcacacgac gccatccttc 12360aatgctcaat tcttcggtgg agttaacttc acccagaatt
cccgcattat tgaaggccaa 12420atgcagtgca ccaaaagtgc tgaccgcaaa ctcgactgca
gctttcatgt cttcaggctc 12480agcagtattg gccttattcg cagccgcttt cccgcctaaa
gcgacaattt cgtccacaac 12540tttctgtgct gcttccaggt taatatctga aaccactaca
cttacaccct gttgagccaa 12600aagcagtgcg gtgcttttac caatacctga accagcgcca
gtaattaaag cgactttatt 12660gttgaattta tttgacataa ttttttccat ttcaaatttt
aagcatcaaa gcttgtttca 12720tattttaaga ttcaagaaac cagatccggt agatgactcg
tctgccaagc gacaacccgt 12780ctgatatcag gcttgcgatt caccctgtag acggttttca
ttcctaaatt ctgtatttcc 12840aagttatata aacaaaagtg ctaatctatg gggaattccc
aggatccaaa caaatagaat 12900gccatgaaag catcttttgc caagcgctgt gctgtatgtt
tcctagacaa accaccaacg 12960ataactgcaa ctttttgaac tccttacaat ttccttattt
tctttcccct tcatcgcata 13020aaaatagttt ttgcattcac aacaaaatca gcatgaatag
tttttaaact cactgtacat 13080attttctata ttgatgacca agctggatat tgaattgcaa
aattctatac agcctgttca 13140acatgatcga tttagaaggc atacagtaaa cgtgactgaa
gtccagaaat ttccaagcca 13200ttttcaacat tcacatcttg tcgccattgt aataatagct
gcagattcgg cttgatattg 13260gtagaagcag aaacgacaaa ggtatctttt ctatcactgc
cacgttcagt gacaccattc 13320accttttctt taccgccatc ggtatgtctc caggtgacag
ccaaattgga tttatcggtc 13380actttataga gtgcggagaa atctgtctgg aaaaaaacct
ctttctcaat gttggtatat 13440ttttgctcgc tataaagttc aaactgcccc accccctcaa
gcgcaaattt atcagttaaa 13500gcatggtaat aaccggcctg aacattatat tgatagcgat
cattactgat ggcaaaaccc 13560ttcgtttcat tactgccggt aggtacggtc aaaaaaccac
cgaaaccaaa atagcgccct 13620ttttcagcat catgcaatgg ccaggcgata ccacccacaa
ttaaatcacc gacacccgag 13680atatcatcag cgccattcat cttttgcttg gcaaaaggca
agaggaattg aggatctaca 13740atccaatccc ctacttcaat aaaacgaacg taacgcaata
ttcccaaatc aatgcttaaa 13800tcgagatcat cagcgacttt atcaccattt gcatacgcct
tatccgcttc cgtatgctgg 13860taataggcaa ccgctaagtt ggttccccct ggaagtgctt
gataatcccc ggcatcagaa 13920ctcacccctg cggcttgcag gtccaaagcg gcagttaaag
caaagaccaa agcagctatt 13980ttttgatttg aacgatgata gaaatagttt ttcatttgtt
tcatttttaa ctctccgttg 14040ttttgactca tttttttaaa atgagtcttc ctagcacaaa
gaccactcag gtctttgcgc 14100aatttcttga ttttgatttg ggtattaaat atggaaaaac
gttgggtgat cagttttcgt 14160gcataagcac aatacgcccg atgacgttgc catctttcaa
gtctccaaat gcggaattga 14220tctgcgaaat tggcagtttt ttcacgggaa tggctgacat
gtgggtttct ttcaccagct 14280ccaccagctc tcttaattcc tctaccgtcc ctacataact
gccctggatt gtgagtggtc 14340tcattggaat caccggaatg gaaagcttaa tttctccccc
catcaatccg cagatcacaa 14400tatgcccacc acgtgcagca ctcgccaagg caaggctcaa
tgttggatta ctgccaacca 14460gatcaaggat cagacgtgca ccaccgtcag ttgcctgaat
cagctgttga gcagcatcct 14520cacttcggct attgatgacc gataatgcac cggcagcacg
tgctgcttcc agtttgctgt 14580catcaatatc aactacgatt gcgcctttgg cttgcatagc
tttgagcaac tcgagtgcca 14640tcagccctaa accaccggca ccaatgatca ccaccggctc
gctttgaatc aaatcaccga 14700attttttcag tgcactgtat gttgtcacgc ctgcacatgc
caaaggtgca gcttcagcca 14760gatccagacc tgcaatatcc accagatatc gtggatgcgg
cacgatgata tattcggcaa 14820aaccacccgg cttggcgatg cctaactgtt gcggtttggc
acacaggttt tcttcgccac 14880gtttacagta gttgcattca ccgcaaccaa tccatggatg
aaccaagctg accatgccga 14940ccttgactga ttccgcatct ggaccgacag caaccacctg
acctgtaatt tcatgactta 15000aggttaaggg tggcttcagc ccacgatctg caagggataa
acgcttgccc ccacctagat 15060cataataacc ttcccataag tgtaaatccg tatggcatag
acctgcggct tttacatgga 15120gtaaaacttc agtacctttc ggttgcggaa tttctttctc
aacgtcttcg agtggttgtc 15180catgatgcgt cacgcagtaa cagtgcatga atctctcctt
tgaaacaata aaatagacgg 15240ccttgtagtg aacaaagtct tttattcact aagttttata
cgccgtgtgg gcactgattt 15300atgctttaaa ccactgcgca attttcgcta attcttgatc
agcttcactt gcacgcccag 15360ctaggaaagg aaaaacgtgc tgcatgttgt ccaccacaga
taaagtcaca tcaacaccct 15420ctttttttgc aatatcagca agacgtgttg cattgtctac
aagtgattca actgatccgg 15480cattgatata caaacgtggg aaaacctgat aattggcttt
taacggattc gccaatggat 15540ttgccggatc accatgttca cccaagaaca tttgtgacat
gcctttaagc agatccactg 15600taatcaaggc atcagtggca tcgttgctga tcagggtttc
acctttgtgc tccatatcca 15660gccaaggaga gaatgcaatc actgctcctg gcaactcaat
cccttcattt cgtagattga 15720gtacggttga tatcgccaga ttcccccccg cagaatcccc
tgcggtcagc atattttttg 15780cagtaaagcc acgctggagt agttctttat atactgctgt
cacgtcctga atttgtgccg 15840ggaagacatg ttctggtgaa cgtcggtaat caaccacaaa
tgcggatacc cctaaatact 15900tggccaaatg ccccaccagc ttacggtgac tggccgaaga
accgaccgca aatccaccgc 15960catgggtata aatgatgact ttggataagt cagcatcttt
cggataaatc caaagacctt 16020ctacacctgc cacaacatcg aatttataag acacttcttc
cggttccaat gtaggttgat 16080gccattcatc aaacatactg cgaaagtctt caatggtcat
attcggattt tcctgcatcc 16140gtcttgacca gttcgcatat aaatcgaaaa gaaattgagt
attgctttgt gtgctattca 16200ttttaaaatc cttgatttga tatttaagga ataaatccta
gttttattcc atgaagatat 16260aaaaacttga gtgccatcac tcatggctag acactcagaa
gatccaaatc taaagagtgg 16320ctttgcatca ctggtttgat acaatttttt gcatgactaa
gtaatctacg gataatctaa 16380ccgtttcaaa ttagtatttt aaaatgtaaa aaatacatac
cagcgaatgt tttctgcaaa 16440atcgcatcct gttcaatata gcttttgatc ctacttattc
tcttttctat tccagtccgt 16500tataaaaaag ctttcattca ttttcatgca atcatgagct
atgaatgttc ttaaacatta 16560aacgattgtg tgtatggctg acttgtacat tcttgtactt
atttttgtat aaaatgatca 16620ggctcatcaa tttatgggaa aaattacaat tcgggtacaa
tatctttcct gtttcatgaa 16680tctattcaac tcattaaact tacgaccctc aactgcccaa
aatcatagga tctgccgatc 16740cacttgcaga attagcaatg ctaaaacatg aactccaaag
agttactaaa aaaagagcat 16800attaaaaaaa agccgtggca tatttcgcaa gccagttcaa
gtcaggtatg tctttattca 16860gtacctcagt taaactttag attttcataa cgatggttat
tctgcatggc taaatacgct 16920aatcagcaaa aaactctcca aaagataggc acagaaacac
atatcaacca taaaaaccat 16980ctcagacagt atatttacaa gcctctaatt caccgcactc
acacttctct gcaagccttt 17040ttaaataccc tgtacaaagt tctcagcctg atgaagcttc
accttggact tagctttcag 17100ttcagcctgt acttggtcag tttctgaatt ttcatttgca
taaaactcct ccaccacatc 17160cataccctcc tcaatgtcag tttcaaaatg tgcattgtca
tagccttgcc gtgccatttg 17220aatggcttat tgaagattaa tggcatcacg taaagttaaa
tccacgtaat acacaggtgt 17280tcgatagctt tgcgtcgtag actttctcga agagtcaatt
gcagcggtag gcatgacagc 17340aagccattca atgccgcatg gtaataactc agccgtgcgg
ccaacgttcg tatgctgtta 17400aaacccggtt attctaa
17417741164DNAEscherichia coli 74atgaacaact
ttaatctgca caccccaacc cgcattctgt ttggtaaagg cgcaatcgct 60ggtttacgcg
aacaaattcc tcacgatgct cgcgtattga ttacctacgg cggcggcagc 120gtgaaaaaaa
ccggcgttct cgatcaagtt ctggatgccc tgaaaggcat ggacgtgctg 180gaatttggcg
gtattgagcc aaacccggct tatgaaacgc tgatgaacgc cgtgaaactg 240gttcgcgaac
agaaagtgac tttcctgctg gcggttggcg gcggttctgt actggacggc 300accaaattta
tcgccgcagc ggctaactat ccggaaaata tcgatccgtg gcacattctg 360caaacgggcg
gtaaagagat taaaagcgcc atcccgatgg gctgtgtgct gacgctgcca 420gcaaccggtt
cagaatccaa cgcaggcgcg gtgatctccc gtaaaaccac aggcgacaag 480caggcgttcc
attctgccca tgttcagccg gtatttgccg tgctcgatcc ggtttatacc 540tacaccctgc
cgccgcgtca ggtggctaac ggcgtagtgg acgcctttgt acacaccgtg 600gaacagtatg
ttaccaaacc ggttgatgcc aaaattcagg accgtttcgc agaaggcatt 660ttgctgacgc
taatcgaaga tggtccgaaa gccctgaaag agccagaaaa ctacgatgtg 720cgcgccaacg
tcatgtgggc ggcgactcag gcgctgaacg gtttgattgg cgctggcgta 780ccgcaggact
gggcaacgca tatgctgggc cacgaactga ctgcgatgca cggtctggat 840cacgcgcaaa
cactggctat cgtcctgcct gcactgtgga atgaaaaacg cgataccaag 900cgcgctaagc
tgctgcaata tgctgaacgc gtctggaaca tcactgaagg ttccgatgat 960gagcgtattg
acgccgcgat tgccgcaacc cgcaatttct ttgagcaatt aggcgtgccg 1020acccacctct
ccgactacgg tctggacggc agctccatcc cggctttgct gaaaaaactg 1080gaagagcacg
gcatgaccca actgggcgaa aatcatgaca ttacgttgga tgtcagccgc 1140cgtatatacg
aagccgcccg ctaa
116475387PRTEscherichia coli 75Met Asn Asn Phe Asn Leu His Thr Pro Thr
Arg Ile Leu Phe Gly Lys 1 5 10
15 Gly Ala Ile Ala Gly Leu Arg Glu Gln Ile Pro His Asp Ala Arg
Val 20 25 30 Leu
Ile Thr Tyr Gly Gly Gly Ser Val Lys Lys Thr Gly Val Leu Asp 35
40 45 Gln Val Leu Asp Ala Leu
Lys Gly Met Asp Val Leu Glu Phe Gly Gly 50 55
60 Ile Glu Pro Asn Pro Ala Tyr Glu Thr Leu Met
Asn Ala Val Lys Leu 65 70 75
80 Val Arg Glu Gln Lys Val Thr Phe Leu Leu Ala Val Gly Gly Gly Ser
85 90 95 Val Leu
Asp Gly Thr Lys Phe Ile Ala Ala Ala Ala Asn Tyr Pro Glu 100
105 110 Asn Ile Asp Pro Trp His Ile
Leu Gln Thr Gly Gly Lys Glu Ile Lys 115 120
125 Ser Ala Ile Pro Met Gly Cys Val Leu Thr Leu Pro
Ala Thr Gly Ser 130 135 140
Glu Ser Asn Ala Gly Ala Val Ile Ser Arg Lys Thr Thr Gly Asp Lys 145
150 155 160 Gln Ala Phe
His Ser Ala His Val Gln Pro Val Phe Ala Val Leu Asp 165
170 175 Pro Val Tyr Thr Tyr Thr Leu Pro
Pro Arg Gln Val Ala Asn Gly Val 180 185
190 Val Asp Ala Phe Val His Thr Val Glu Gln Tyr Val Thr
Lys Pro Val 195 200 205
Asp Ala Lys Ile Gln Asp Arg Phe Ala Glu Gly Ile Leu Leu Thr Leu 210
215 220 Ile Glu Asp Gly
Pro Lys Ala Leu Lys Glu Pro Glu Asn Tyr Asp Val 225 230
235 240 Arg Ala Asn Val Met Trp Ala Ala Thr
Gln Ala Leu Asn Gly Leu Ile 245 250
255 Gly Ala Gly Val Pro Gln Asp Trp Ala Thr His Met Leu Gly
His Glu 260 265 270
Leu Thr Ala Met His Gly Leu Asp His Ala Gln Thr Leu Ala Ile Val
275 280 285 Leu Pro Ala Leu
Trp Asn Glu Lys Arg Asp Thr Lys Arg Ala Lys Leu 290
295 300 Leu Gln Tyr Ala Glu Arg Val Trp
Asn Ile Thr Glu Gly Ser Asp Asp 305 310
315 320 Glu Arg Ile Asp Ala Ala Ile Ala Ala Thr Arg Asn
Phe Phe Glu Gln 325 330
335 Leu Gly Val Pro Thr His Leu Ser Asp Tyr Gly Leu Asp Gly Ser Ser
340 345 350 Ile Pro Ala
Leu Leu Lys Lys Leu Glu Glu His Gly Met Thr Gln Leu 355
360 365 Gly Glu Asn His Asp Ile Thr Leu
Asp Val Ser Arg Arg Ile Tyr Glu 370 375
380 Ala Ala Arg 385 761623DNABacillus subtilis
76atgtatttgg cattccaggt gcaaaaattg atgcggtatt tgacgcttta caagataaag
60gacctgaaat tatcgttgcc cggcacgaac aaaacgcagc aattcatggc ccaagcagtc
120ggccgtttaa ctggaaaacc gggagtcgtg ttagtcacat caggaccggg tgcctctaac
180ttggcaacag gcctgctgac agcgaacact gaaggagacc ctgtcgttgc gcttgctgga
240aacgtgatcc gtgcatatcg tttaaaacgg acacatcaat ctttggataa tgcggcgcta
300ttccagccga ttacaaaata cagtgtagaa gttcaagatg taaaaaatat accggaagct
360gttacaaatg catttaggat agcgtcagca gggcaggctg gggccgcttt tgtgagcttt
420ccgcaagatg ttgtgaatga agtcacaaat acgaaaaacg tgcgtgctgt tgcagcgcca
480aaactcggtc ctgcagcaga tgatgcaatc agtgcggcca tagcaaaaat ccaaacagca
540aaacttcctg tcgttttggt cggcatgaaa ggcggaagac cggaagcaat taaagcggtt
600cgcaagcttt tgaaaaaggt tcagcttcca tttgttgaaa catatcaagc tgccggtacc
660ctttctagag atttagagga tcaatatttt ggccgtatcg gtttgttccg caaccagcct
720ggcgatttac tgctagagca ggcagatgtt gttctgacga tcggctatga cccgattgaa
780tatgatccga aattctggaa tatcaatgga gaccggacaa ttatccattt agacgagatt
840atcgctgaca ttgatcatgc ttaccagcct gatcttgaat tgatcggtga cattccgtcc
900acgatcaatc atatcgaaca cgatgctgtg aaagtggaat ttgcagagcg tgagcagaaa
960atcctttctg atttaaaaca atatatgcat gaaggtgagc aggtgcctgc agattggaaa
1020tcagacagag cgcaccctct tgaaatcgtt aaagagttgc gtaatgcagt cgatgatcat
1080gttacagtaa cttgcgatat cggttcgcac tccatttgga tgtcacgtta tttccgcagc
1140tacgagccgt taacattaat gatcagtaac ggtatgcaaa cactcggcgt tgcgcttcct
1200tgggcaatcg gcgcttcatt ggtgaaaccg ggagaaaaag tggtttctgt ctctggtgac
1260ggcggtttct tattctcagc aatggaatta gagacagcag ttcgactaaa agcaccaatt
1320gtacacattg tatggaacga cagcacatat gacatggtgc atttccagca attgaaaaaa
1380tataaccgta catctgcggt cgatttcgga aatatcgata tcgtgaaata tgcggaaagc
1440ttcggagcaa ctgcgttgcg cgtagaatca ccagaccagc tggcagatgt tctgcgtcaa
1500ggcatgaacg ctgaaggtcc tgtcatcatc gatgtcccgg ttgactacag tgataacatt
1560aatttagcaa gtgacaagct tccgaaagaa ttcggggaac tcatgaaaac gaaagctctc
1620tag
162377540PRTBacillus subtilis 77Met Tyr Leu Ala Phe Gln Val Gln Lys Leu
Met Arg Tyr Leu Thr Leu 1 5 10
15 Tyr Lys Ile Lys Asp Leu Lys Leu Ser Leu Pro Gly Thr Asn Lys
Thr 20 25 30 Gln
Gln Phe Met Ala Gln Ala Val Gly Arg Leu Thr Gly Lys Pro Gly 35
40 45 Val Val Leu Val Thr Ser
Gly Pro Gly Ala Ser Asn Leu Ala Thr Gly 50 55
60 Leu Leu Thr Ala Asn Thr Glu Gly Asp Pro Val
Val Ala Leu Ala Gly 65 70 75
80 Asn Val Ile Arg Ala Tyr Arg Leu Lys Arg Thr His Gln Ser Leu Asp
85 90 95 Asn Ala
Ala Leu Phe Gln Pro Ile Thr Lys Tyr Ser Val Glu Val Gln 100
105 110 Asp Val Lys Asn Ile Pro Glu
Ala Val Thr Asn Ala Phe Arg Ile Ala 115 120
125 Ser Ala Gly Gln Ala Gly Ala Ala Phe Val Ser Phe
Pro Gln Asp Val 130 135 140
Val Asn Glu Val Thr Asn Thr Lys Asn Val Arg Ala Val Ala Ala Pro 145
150 155 160 Lys Leu Gly
Pro Ala Ala Asp Asp Ala Ile Ser Ala Ala Ile Ala Lys 165
170 175 Ile Gln Thr Ala Lys Leu Pro Val
Val Leu Val Gly Met Lys Gly Gly 180 185
190 Arg Pro Glu Ala Ile Lys Ala Val Arg Lys Leu Leu Lys
Lys Val Gln 195 200 205
Leu Pro Phe Val Glu Thr Tyr Gln Ala Ala Gly Thr Leu Ser Arg Asp 210
215 220 Leu Glu Asp Gln
Tyr Phe Gly Arg Ile Gly Leu Phe Arg Asn Gln Pro 225 230
235 240 Gly Asp Leu Leu Leu Glu Gln Ala Asp
Val Val Leu Thr Ile Gly Tyr 245 250
255 Asp Pro Ile Glu Tyr Asp Pro Lys Phe Trp Asn Ile Asn Gly
Asp Arg 260 265 270
Thr Ile Ile His Leu Asp Glu Ile Ile Ala Asp Ile Asp His Ala Tyr
275 280 285 Gln Pro Asp Leu
Glu Leu Ile Gly Asp Ile Pro Ser Thr Ile Asn His 290
295 300 Ile Glu His Asp Ala Val Lys Val
Glu Phe Ala Glu Arg Glu Gln Lys 305 310
315 320 Ile Leu Ser Asp Leu Lys Gln Tyr Met His Glu Gly
Glu Gln Val Pro 325 330
335 Ala Asp Trp Lys Ser Asp Arg Ala His Pro Leu Glu Ile Val Lys Glu
340 345 350 Leu Arg Asn
Ala Val Asp Asp His Val Thr Val Thr Cys Asp Ile Gly 355
360 365 Ser His Ser Ile Trp Met Ser Arg
Tyr Phe Arg Ser Tyr Glu Pro Leu 370 375
380 Thr Leu Met Ile Ser Asn Gly Met Gln Thr Leu Gly Val
Ala Leu Pro 385 390 395
400 Trp Ala Ile Gly Ala Ser Leu Val Lys Pro Gly Glu Lys Val Val Ser
405 410 415 Val Ser Gly Asp
Gly Gly Phe Leu Phe Ser Ala Met Glu Leu Glu Thr 420
425 430 Ala Val Arg Leu Lys Ala Pro Ile Val
His Ile Val Trp Asn Asp Ser 435 440
445 Thr Tyr Asp Met Val His Phe Gln Gln Leu Lys Lys Tyr Asn
Arg Thr 450 455 460
Ser Ala Val Asp Phe Gly Asn Ile Asp Ile Val Lys Tyr Ala Glu Ser 465
470 475 480 Phe Gly Ala Thr Ala
Leu Arg Val Glu Ser Pro Asp Gln Leu Ala Asp 485
490 495 Val Leu Arg Gln Gly Met Asn Ala Glu Gly
Pro Val Ile Ile Asp Val 500 505
510 Pro Val Asp Tyr Ser Asp Asn Ile Asn Leu Ala Ser Asp Lys Leu
Pro 515 520 525 Lys
Glu Phe Gly Glu Leu Met Lys Thr Lys Ala Leu 530 535
540 781680DNAKlebsiella terrigena 78atggacaaac cgcgtcacga
acgtcaatgg gcccacggtg ccgacttaat cgtcagccag 60cttgaggccc agggcgtacg
ccaggtcttc ggcatccccg gtgccaaaat cgacaaggtg 120tttgattccc tcctcgactc
ctcaatccgc attattccgg tgcgccacga ggctaacgcc 180gcctttatgg ccgcggcggt
cgggcggatt accggtaaag cgggcgtcgc gctggtgacc 240tccggtcccg gctgctcaaa
cctgattacc ggcatggcca ccgccaatag cgaaggcgac 300ccggtggtgg cgctgggcgg
cgcggtgaag cgcgcggata aggccaagct ggttcaccaa 360agcatggaca ccgtggcgat
gttcagcccg gtcaccaaat acgccgtcga ggtgaccgcc 420tccgacgcgc tggccgaggt
ggtctccaac gcctttcgcg ccgccgaaca ggggcgtccg 480gggagcgcgt ttgtcagcct
gccgcaggat atcgttgacg gccccgccag cggcagcacg 540ctgcccgcca gcagagcgcc
gcagatgggc gccgcgccgg atggcgccgt tgacagcgtg 600gcgcaggcga tcgccgcggc
gaagaaccct atcttcctgc tcgggctgat ggccagccag 660ccggaaaaca gccgcgccct
gcaccgccat gctggaaaaa agccatattc cggtcaccag 720cacctatcag gcgccggggc
ggtaaatcag gataacttcg cccgcttcgc cggccgggta 780ggcctgttta ataaccaggc
gggcgatcgc ctgctgcgtc aggcggacct gatcatctgc 840atcggctata gcccggttga
gtacgaaccg gcgatgtgga acagcggcac ggcaaccctg 900gtgcatatcg acgtgctgcc
ggcctatgaa gagcggaact acgtcccgga tatcgagctg 960gtgggcgaca tcgccgccac
cctcgagaag ctggcccagc gcattgaaca tcggctggtg 1020ttaactccgc aggcggcgga
catcctcgcc gaccgccagc gccagcggga gctgcttgac 1080cgccgcgggg cgcagctgaa
tcagtttgcg ctccacccgc tgcgcatcgt gcgggcgatg 1140caggatatcg tcaatagcga
cgtcaccttg accgtcgata tgggcagttt ccatatctgg 1200attgcccgct acctctacag
cttccgcgcc cgccaggtga tgatctccaa cggtcagcaa 1260acgatgggcg tcgcgctgcc
gtgggcaatc ggcgcgtggc tggtcaatcc gcagcgcaag 1320gtggtctcgg tatccggcga
tggcggcttc ctgcagtcga gcatggagct ggagaccgcc 1380gtgcgcctgc acgccaatat
tctgcacatc atctgggtcg ataacggcta caacatggtg 1440gcgattcagg aacagaagaa
atatcagcgc ctctccggcg tggagttcgg cccggtcgat 1500ttcaaagtct acgccgaagc
gttcggggcc tgcgggtttg cggtagagag cgccgaggcc 1560ctggagccga ccctgcgcgc
ggcgatggat gtcgacggcc cggcggtggt cgccattccg 1620gtcgattacc gcgataaccc
tctgctgatg ggccagctcc atctcagcca aatactgtga 168079559PRTKlebsiella
terrigena 79Met Asp Lys Pro Arg His Glu Arg Gln Trp Ala His Gly Ala Asp
Leu 1 5 10 15 Ile
Val Ser Gln Leu Glu Ala Gln Gly Val Arg Gln Val Phe Gly Ile
20 25 30 Pro Gly Ala Lys Ile
Asp Lys Val Phe Asp Ser Leu Leu Asp Ser Ser 35
40 45 Ile Arg Ile Ile Pro Val Arg His Glu
Ala Asn Ala Ala Phe Met Ala 50 55
60 Ala Ala Val Gly Arg Ile Thr Gly Lys Ala Gly Val Ala
Leu Val Thr 65 70 75
80 Ser Gly Pro Gly Cys Ser Asn Leu Ile Thr Gly Met Ala Thr Ala Asn
85 90 95 Ser Glu Gly Asp
Pro Val Val Ala Leu Gly Gly Ala Val Lys Arg Ala 100
105 110 Asp Lys Ala Lys Leu Val His Gln Ser
Met Asp Thr Val Ala Met Phe 115 120
125 Ser Pro Val Thr Lys Tyr Ala Val Glu Val Thr Ala Ser Asp
Ala Leu 130 135 140
Ala Glu Val Val Ser Asn Ala Phe Arg Ala Ala Glu Gln Gly Arg Pro 145
150 155 160 Gly Ser Ala Phe Val
Ser Leu Pro Gln Asp Ile Val Asp Gly Pro Ala 165
170 175 Ser Gly Ser Thr Leu Pro Ala Ser Arg Ala
Pro Gln Met Gly Ala Ala 180 185
190 Pro Asp Gly Ala Val Asp Ser Val Ala Gln Ala Ile Ala Ala Ala
Lys 195 200 205 Asn
Pro Ile Phe Leu Leu Gly Leu Met Ala Ser Gln Pro Glu Asn Ser 210
215 220 Arg Ala Leu His Arg His
Ala Gly Lys Lys Pro Tyr Ser Gly His Gln 225 230
235 240 His Leu Ser Gly Ala Gly Ala Val Asn Gln Asp
Asn Phe Ala Arg Phe 245 250
255 Ala Gly Arg Val Gly Leu Phe Asn Asn Gln Ala Gly Asp Arg Leu Leu
260 265 270 Arg Gln
Ala Asp Leu Ile Ile Cys Ile Gly Tyr Ser Pro Val Glu Tyr 275
280 285 Glu Pro Ala Met Trp Asn Ser
Gly Thr Ala Thr Leu Val His Ile Asp 290 295
300 Val Leu Pro Ala Tyr Glu Glu Arg Asn Tyr Val Pro
Asp Ile Glu Leu 305 310 315
320 Val Gly Asp Ile Ala Ala Thr Leu Glu Lys Leu Ala Gln Arg Ile Glu
325 330 335 His Arg Leu
Val Leu Thr Pro Gln Ala Ala Asp Ile Leu Ala Asp Arg 340
345 350 Gln Arg Gln Arg Glu Leu Leu Asp
Arg Arg Gly Ala Gln Leu Asn Gln 355 360
365 Phe Ala Leu His Pro Leu Arg Ile Val Arg Ala Met Gln
Asp Ile Val 370 375 380
Asn Ser Asp Val Thr Leu Thr Val Asp Met Gly Ser Phe His Ile Trp 385
390 395 400 Ile Ala Arg Tyr
Leu Tyr Ser Phe Arg Ala Arg Gln Val Met Ile Ser 405
410 415 Asn Gly Gln Gln Thr Met Gly Val Ala
Leu Pro Trp Ala Ile Gly Ala 420 425
430 Trp Leu Val Asn Pro Gln Arg Lys Val Val Ser Val Ser Gly
Asp Gly 435 440 445
Gly Phe Leu Gln Ser Ser Met Glu Leu Glu Thr Ala Val Arg Leu His 450
455 460 Ala Asn Ile Leu His
Ile Ile Trp Val Asp Asn Gly Tyr Asn Met Val 465 470
475 480 Ala Ile Gln Glu Gln Lys Lys Tyr Gln Arg
Leu Ser Gly Val Glu Phe 485 490
495 Gly Pro Val Asp Phe Lys Val Tyr Ala Glu Ala Phe Gly Ala Cys
Gly 500 505 510 Phe
Ala Val Glu Ser Ala Glu Ala Leu Glu Pro Thr Leu Arg Ala Ala 515
520 525 Met Asp Val Asp Gly Pro
Ala Val Val Ala Ile Pro Val Asp Tyr Arg 530 535
540 Asp Asn Pro Leu Leu Met Gly Gln Leu His Leu
Ser Gln Ile Leu 545 550 555
80768DNABacillus subtilis 80atgaaacgag aaagcaacat tcaagtgctc agccgtggtc
aaaaagatca gcctgtgagc 60cagatttatc aagtatcaac aatgacttct ctattagacg
gagtatatga cggagatttt 120gaactgtcag agattccgaa atatggagac ttcggtatcg
gaacctttaa caagcttgac 180ggagagctga ttgggtttga cggcgaattt taccgtcttc
gctcagacgg aaccgcgaca 240ccggtccaaa atggagaccg ttcaccgttc tgttcattta
cgttctttac accggacatg 300acgcacaaaa ttgatgcgaa aatgacacgc gaagactttg
aaaaagagat caacagcatg 360ctgccaagca gaaacttatt ttatgcaatt cgcattgacg
gattgtttaa aaaggtgcag 420acaagaacag tagaacttca agaaaaacct tacgtgccaa
tggttgaagc ggtcaaaaca 480cagccgattt tcaacttcga caacgtgaga ggaacgattg
taggtttctt gacaccagct 540tatgcaaacg gaatcgccgt ttctggctat cacctgcact
tcattgacga aggacgcaat 600tcaggcggac acgtttttga ctatgtgctt gaggattgca
cggttacgat ttctcaaaaa 660atgaacatga atctcagact tccgaacaca gcggatttct
ttaatgcgaa tctggataac 720cctgattttg cgaaagatat cgaaacaact gaaggaagcc
ctgaataa 76881255PRTBacillus subtilis 81Met Lys Arg Glu
Ser Asn Ile Gln Val Leu Ser Arg Gly Gln Lys Asp 1 5
10 15 Gln Pro Val Ser Gln Ile Tyr Gln Val
Ser Thr Met Thr Ser Leu Leu 20 25
30 Asp Gly Val Tyr Asp Gly Asp Phe Glu Leu Ser Glu Ile Pro
Lys Tyr 35 40 45
Gly Asp Phe Gly Ile Gly Thr Phe Asn Lys Leu Asp Gly Glu Leu Ile 50
55 60 Gly Phe Asp Gly Glu
Phe Tyr Arg Leu Arg Ser Asp Gly Thr Ala Thr 65 70
75 80 Pro Val Gln Asn Gly Asp Arg Ser Pro Phe
Cys Ser Phe Thr Phe Phe 85 90
95 Thr Pro Asp Met Thr His Lys Ile Asp Ala Lys Met Thr Arg Glu
Asp 100 105 110 Phe
Glu Lys Glu Ile Asn Ser Met Leu Pro Ser Arg Asn Leu Phe Tyr 115
120 125 Ala Ile Arg Ile Asp Gly
Leu Phe Lys Lys Val Gln Thr Arg Thr Val 130 135
140 Glu Leu Gln Glu Lys Pro Tyr Val Pro Met Val
Glu Ala Val Lys Thr 145 150 155
160 Gln Pro Ile Phe Asn Phe Asp Asn Val Arg Gly Thr Ile Val Gly Phe
165 170 175 Leu Thr
Pro Ala Tyr Ala Asn Gly Ile Ala Val Ser Gly Tyr His Leu 180
185 190 His Phe Ile Asp Glu Gly Arg
Asn Ser Gly Gly His Val Phe Asp Tyr 195 200
205 Val Leu Glu Asp Cys Thr Val Thr Ile Ser Gln Lys
Met Asn Met Asn 210 215 220
Leu Arg Leu Pro Asn Thr Ala Asp Phe Phe Asn Ala Asn Leu Asp Asn 225
230 235 240 Pro Asp Phe
Ala Lys Asp Ile Glu Thr Thr Glu Gly Ser Pro Glu 245
250 255 82780DNAKlebsiella terrigena
82gtgaatcatt atcctgaatg cacctgccag gagagcctgt gcgaaaccgt acgcggcttc
60tccgcccacc accctgatag cgttatctat cagacctctc tgatgagcgc gctgctgagc
120ggggtctatg agggtagcac caccatcgcc gacctgctga cccacggcga cttcggtctc
180ggcaccttta acgaactcga tggcgaactg attgccttta gcagcgaggt ctaccagctg
240cgcgctgacg gcagcgcgcg taaagcccgg gcggatcaaa aaacgccctt cgcggtgatg
300acctggttca gaccgcagta ccgtaaaacc tttgaccacc cggtcagccg ccagcagctg
360cacgacgtta tcgaccagca aatcccctcc gataacctgt tctgcgccct gcatattgat
420ggtcactttc gccacgccca cacccgcacc gtgccgcggc agacgccgcc ctatcgggcg
480atgaccgacg tgctcgatga ccagccggtt ttccgcttca accagcgcaa ggggacgctg
540gtcggctttc gcaccccgca gcatatgcag ggccttaacg ttgccggcta ccacgagcac
600tttattaccg acgatcgcca gggcggcggc catctgctgg actaccagct cgatagcggc
660gtgctgacct tcggcgagat ccacaagctg atgattgacc tcccggccga cagcgctttc
720ctgcaggccg acctgcatcc tgacaatctc gatgccgcta ttcgtgcggt agaaaactaa
78083259PRTKlebsiella terrigena 83Met Asn His Tyr Pro Glu Cys Thr Cys Gln
Glu Ser Leu Cys Glu Thr 1 5 10
15 Val Arg Gly Phe Ser Ala His His Pro Asp Ser Val Ile Tyr Gln
Thr 20 25 30 Ser
Leu Met Ser Ala Leu Leu Ser Gly Val Tyr Glu Gly Ser Thr Thr 35
40 45 Ile Ala Asp Leu Leu Thr
His Gly Asp Phe Gly Leu Gly Thr Phe Asn 50 55
60 Glu Leu Asp Gly Glu Leu Ile Ala Phe Ser Ser
Glu Val Tyr Gln Leu 65 70 75
80 Arg Ala Asp Gly Ser Ala Arg Lys Ala Arg Ala Asp Gln Lys Thr Pro
85 90 95 Phe Ala
Val Met Thr Trp Phe Arg Pro Gln Tyr Arg Lys Thr Phe Asp 100
105 110 His Pro Val Ser Arg Gln Gln
Leu His Asp Val Ile Asp Gln Gln Ile 115 120
125 Pro Ser Asp Asn Leu Phe Cys Ala Leu His Ile Asp
Gly His Phe Arg 130 135 140
His Ala His Thr Arg Thr Val Pro Arg Gln Thr Pro Pro Tyr Arg Ala 145
150 155 160 Met Thr Asp
Val Leu Asp Asp Gln Pro Val Phe Arg Phe Asn Gln Arg 165
170 175 Lys Gly Thr Leu Val Gly Phe Arg
Thr Pro Gln His Met Gln Gly Leu 180 185
190 Asn Val Ala Gly Tyr His Glu His Phe Ile Thr Asp Asp
Arg Gln Gly 195 200 205
Gly Gly His Leu Leu Asp Tyr Gln Leu Asp Ser Gly Val Leu Thr Phe 210
215 220 Gly Glu Ile His
Lys Leu Met Ile Asp Leu Pro Ala Asp Ser Ala Phe 225 230
235 240 Leu Gln Ala Asp Leu His Pro Asp Asn
Leu Asp Ala Ala Ile Arg Ala 245 250
255 Val Glu Asn 841053DNABacillus cereus 84atgaaagcac
tactttggca taatcaacgt gatgtacgag tagaagaagt accagaacca 60acagtaaaac
caggaacagt gaaaatcaaa gttaaatggt gtggtatttg tgggacagac 120ttgcatgaat
atttagcagg gcctattttt attccaacag aagaacatcc attaacacat 180gtgaaagcac
ctgttatttt aggtcatgag tttagtggtg aggtaataga gattggtgaa 240ggagttacat
ctcataaagt gggagaccgc gttgttgtag agccaattta ttcttgtggt 300aaatgtgaag
cttgtaaaca tggacattac aatgtttgtg aacaacttgt tttccacggt 360cttggcggag
aaggcggcgg tttctctgaa tatacagtag taccagaaga tatggttcat 420cacattccag
atgaaatgac gtatgaacaa ggtgcgcttg tagaaccagc agcagtagca 480gttcatgcag
tacgtcaaag taaattaaaa gaaggggaag ctgtagcggt atttggttgc 540ggtccaattg
gacttcttgt tatccaagca gctaaagcag caggagcaac tcctgttatt 600gcagttgaac
tttctaaaga acgtcaagag ttagcgaaat tagcaggtgc ggattatgta 660ttaaatccag
caactcaaga tgtgttagct gaaattcgta acttaacaaa tggtttaggt 720gtaaatgtta
gctttgaagt aacaggtgtt gaagttgtac tacgccaagc gattgaaagt 780acaagcttcg
aaggacaaac tgtaattgtt agtgtatggg aaaaagacgc aacaattact 840ccaaataact
tagtattaaa agaaaaagaa gttattggta ttttaggata ccgtcacatc 900ttcccagctg
ttattaaatt gattagctcc ggtcaaattc aagcagagaa attaattacg 960aaaaaaatta
cagtggatca agttgttgaa gaaggatttg aagcacttgt aaaagataaa 1020acacaagtga
aaattcttgt ttcacctaaa taa
105385350PRTBacillus cereus 85Met Lys Ala Leu Leu Trp His Asn Gln Arg Asp
Val Arg Val Glu Glu 1 5 10
15 Val Pro Glu Pro Thr Val Lys Pro Gly Thr Val Lys Ile Lys Val Lys
20 25 30 Trp Cys
Gly Ile Cys Gly Thr Asp Leu His Glu Tyr Leu Ala Gly Pro 35
40 45 Ile Phe Ile Pro Thr Glu Glu
His Pro Leu Thr His Val Lys Ala Pro 50 55
60 Val Ile Leu Gly His Glu Phe Ser Gly Glu Val Ile
Glu Ile Gly Glu 65 70 75
80 Gly Val Thr Ser His Lys Val Gly Asp Arg Val Val Val Glu Pro Ile
85 90 95 Tyr Ser Cys
Gly Lys Cys Glu Ala Cys Lys His Gly His Tyr Asn Val 100
105 110 Cys Glu Gln Leu Val Phe His Gly
Leu Gly Gly Glu Gly Gly Gly Phe 115 120
125 Ser Glu Tyr Thr Val Val Pro Glu Asp Met Val His His
Ile Pro Asp 130 135 140
Glu Met Thr Tyr Glu Gln Gly Ala Leu Val Glu Pro Ala Ala Val Ala 145
150 155 160 Val His Ala Val
Arg Gln Ser Lys Leu Lys Glu Gly Glu Ala Val Ala 165
170 175 Val Phe Gly Cys Gly Pro Ile Gly Leu
Leu Val Ile Gln Ala Ala Lys 180 185
190 Ala Ala Gly Ala Thr Pro Val Ile Ala Val Glu Leu Ser Lys
Glu Arg 195 200 205
Gln Glu Leu Ala Lys Leu Ala Gly Ala Asp Tyr Val Leu Asn Pro Ala 210
215 220 Thr Gln Asp Val Leu
Ala Glu Ile Arg Asn Leu Thr Asn Gly Leu Gly 225 230
235 240 Val Asn Val Ser Phe Glu Val Thr Gly Val
Glu Val Val Leu Arg Gln 245 250
255 Ala Ile Glu Ser Thr Ser Phe Glu Gly Gln Thr Val Ile Val Ser
Val 260 265 270 Trp
Glu Lys Asp Ala Thr Ile Thr Pro Asn Asn Leu Val Leu Lys Glu 275
280 285 Lys Glu Val Ile Gly Ile
Leu Gly Tyr Arg His Ile Phe Pro Ala Val 290 295
300 Ile Lys Leu Ile Ser Ser Gly Gln Ile Gln Ala
Glu Lys Leu Ile Thr 305 310 315
320 Lys Lys Ile Thr Val Asp Gln Val Val Glu Glu Gly Phe Glu Ala Leu
325 330 335 Val Lys
Asp Lys Thr Gln Val Lys Ile Leu Val Ser Pro Lys 340
345 350 861053DNABacillus cereus 86atgaaagcac
tactttggca taatcaacgt gatgtacgag tagaagaagt accagaacca 60acagtaaaac
caggaacagt gaaaatcaaa gttaaatggt gtggtatttg tgggacagac 120ttgcatgaat
atttagcagg gcctattttt attccaacag aagaacatcc attaacacat 180gtgaaagcac
ctgttatttt aggtcatgag tttagtggtg aggtaataga gattggtgaa 240ggagttacat
ctcataaagt gggagaccgc gttgttgtag agccaattta ttcttgtggt 300aaatgtgaag
cttgtaaaca tggacattac aatgtttgtg aacaacttgt tttccacggt 360cttggcggag
aaggcggcgg tttctctgaa tatacagtag taccagaaga tatggttcat 420cacattccag
atgaaatgac gtatgaacaa ggtgcgcttg tagaaccagc agcagtagca 480gttcatgcag
tacgtcaaag taaattaaaa gaaggggaag ctgtagcggt atttggttgc 540ggtccaattg
gacttcttgt tatccaagca gctaaagcag caggagcaac tcctgttatt 600gcagttgaac
tttctaaaga acgtcaagag ttagcgaaat tagcaggtgc ggattatgta 660ttaaatccag
caactcaaga tgtgttagct gaaattcgta acttaacaaa tggtttaggt 720gtaaatgtta
gctttgaagt aacaggtgtt gaagttgtac tacgccaagc gattgaaagt 780acaagcttcg
aaggacaaac tgtaattgtt agtgtatggg aaaaagacgc aacaattact 840ccaaataact
tagtattaaa agaaaaagaa gttattggta ttttaggata ccgtcacatc 900ttcccagctg
ttattaaatt gattagctcc ggtcaaattc aagcagagaa attaattacg 960aaaaaaatta
cagtggatca agttgttgaa gaaggatttg aagcacttgt aaaagataaa 1020acacaagtga
aaattcttgt ttcacctaaa taa
105387350PRTBacillus cereus 87Met Lys Ala Leu Leu Trp His Asn Gln Arg Asp
Val Arg Val Glu Glu 1 5 10
15 Val Pro Glu Pro Thr Val Lys Pro Gly Thr Val Lys Ile Lys Val Lys
20 25 30 Trp Cys
Gly Ile Cys Gly Thr Asp Leu His Glu Tyr Leu Ala Gly Pro 35
40 45 Ile Phe Ile Pro Thr Glu Glu
His Pro Leu Thr His Val Lys Ala Pro 50 55
60 Val Ile Leu Gly His Glu Phe Ser Gly Glu Val Ile
Glu Ile Gly Glu 65 70 75
80 Gly Val Thr Ser His Lys Val Gly Asp Arg Val Val Val Glu Pro Ile
85 90 95 Tyr Ser Cys
Gly Lys Cys Glu Ala Cys Lys His Gly His Tyr Asn Val 100
105 110 Cys Glu Gln Leu Val Phe His Gly
Leu Gly Gly Glu Gly Gly Gly Phe 115 120
125 Ser Glu Tyr Thr Val Val Pro Glu Asp Met Val His His
Ile Pro Asp 130 135 140
Glu Met Thr Tyr Glu Gln Gly Ala Leu Val Glu Pro Ala Ala Val Ala 145
150 155 160 Val His Ala Val
Arg Gln Ser Lys Leu Lys Glu Gly Glu Ala Val Ala 165
170 175 Val Phe Gly Cys Gly Pro Ile Gly Leu
Leu Val Ile Gln Ala Ala Lys 180 185
190 Ala Ala Gly Ala Thr Pro Val Ile Ala Val Glu Leu Ser Lys
Glu Arg 195 200 205
Gln Glu Leu Ala Lys Leu Ala Gly Ala Asp Tyr Val Leu Asn Pro Ala 210
215 220 Thr Gln Asp Val Leu
Ala Glu Ile Arg Asn Leu Thr Asn Gly Leu Gly 225 230
235 240 Val Asn Val Ser Phe Glu Val Thr Gly Val
Glu Val Val Leu Arg Gln 245 250
255 Ala Ile Glu Ser Thr Ser Phe Glu Gly Gln Thr Val Ile Val Ser
Val 260 265 270 Trp
Glu Lys Asp Ala Thr Ile Thr Pro Asn Asn Leu Val Leu Lys Glu 275
280 285 Lys Glu Val Ile Gly Ile
Leu Gly Tyr Arg His Ile Phe Pro Ala Val 290 295
300 Ile Lys Leu Ile Ser Ser Gly Gln Ile Gln Ala
Glu Lys Leu Ile Thr 305 310 315
320 Lys Lys Ile Thr Val Asp Gln Val Val Glu Glu Gly Phe Glu Ala Leu
325 330 335 Val Lys
Asp Lys Thr Gln Val Lys Ile Leu Val Ser Pro Lys 340
345 350 881113DNALactococcus lactis 88ttgcctgaaa
cgacaaccat cctatataga ggaggcgttt ttatgcgcgc agcacgtttt 60tacgaccgcg
gggatatccg cattgatgaa attaatgaac caatagtaaa agctggccaa 120gttggcattg
atgtggcttg gtgtggaatt tgtggaacag atctccatga atttttagat 180ggcccaattt
tttgtccgtc agcagaacat cctaatccaa ttactggaga agtaccacca 240gtcactcttg
gacatgaaat gtctggggtt gtaaatttta taggtgaagg agtaagcgga 300cttaaagtag
gtgaccatgt cgttgtcgaa ccttatatcg ttcccgaagg gactgataca 360agtgaaactg
gacattataa cctctcagaa ggctcaaact ttattggttt gggcggaaat 420ggtggaggtt
tggctgaaaa aatttctgtt gatgaacgtt gggttcacaa aattcctgat 480aacttaccat
tggatgaagc tgctctaatt gagccactat cagtcggcta tcacgctgtt 540gaacgagcaa
atttaagtga aaagagtacg gtattagttg ttggtgctgg accaattgga 600ctattaactg
ctgccgttgc aaaagcgcaa ggacatactg ttatcatcag tgaacctagt 660ggacttcgtc
gtaaaaaagc acaagaagca caagttgctg attatttctt caatccaatt 720gaagatgaca
ttcaagctaa agttcatgaa attaatgaaa aaggagtgga cgcagccttt 780gaatgtacct
ctgtccaacc gggatttgac gcttgtctag atgcgattcg tatgggtgga 840acagttgtca
ttgtcgcaat ttggggcaag cctgctagtg ttgatatggc aaaattagta 900atcaaagaag
ctaacctttt aggaacgatt gcttataata acactcatcc aaaaacaatt 960gatttagtat
caacaggtaa aataaaattg gaccaattca tcacagctaa aatcggtttg 1020gatgatttga
ttgataaagg attcgatacg ctgattcatc ataatgaaac agctgttaaa 1080attttagttt
caccaactgg taaaggtcta taa
111389370PRTLactococcus lactis 89Met Pro Glu Thr Thr Thr Ile Leu Tyr Arg
Gly Gly Val Phe Met Arg 1 5 10
15 Ala Ala Arg Phe Tyr Asp Arg Gly Asp Ile Arg Ile Asp Glu Ile
Asn 20 25 30 Glu
Pro Ile Val Lys Ala Gly Gln Val Gly Ile Asp Val Ala Trp Cys 35
40 45 Gly Ile Cys Gly Thr Asp
Leu His Glu Phe Leu Asp Gly Pro Ile Phe 50 55
60 Cys Pro Ser Ala Glu His Pro Asn Pro Ile Thr
Gly Glu Val Pro Pro 65 70 75
80 Val Thr Leu Gly His Glu Met Ser Gly Val Val Asn Phe Ile Gly Glu
85 90 95 Gly Val
Ser Gly Leu Lys Val Gly Asp His Val Val Val Glu Pro Tyr 100
105 110 Ile Val Pro Glu Gly Thr Asp
Thr Ser Glu Thr Gly His Tyr Asn Leu 115 120
125 Ser Glu Gly Ser Asn Phe Ile Gly Leu Gly Gly Asn
Gly Gly Gly Leu 130 135 140
Ala Glu Lys Ile Ser Val Asp Glu Arg Trp Val His Lys Ile Pro Asp 145
150 155 160 Asn Leu Pro
Leu Asp Glu Ala Ala Leu Ile Glu Pro Leu Ser Val Gly 165
170 175 Tyr His Ala Val Glu Arg Ala Asn
Leu Ser Glu Lys Ser Thr Val Leu 180 185
190 Val Val Gly Ala Gly Pro Ile Gly Leu Leu Thr Ala Ala
Val Ala Lys 195 200 205
Ala Gln Gly His Thr Val Ile Ile Ser Glu Pro Ser Gly Leu Arg Arg 210
215 220 Lys Lys Ala Gln
Glu Ala Gln Val Ala Asp Tyr Phe Phe Asn Pro Ile 225 230
235 240 Glu Asp Asp Ile Gln Ala Lys Val His
Glu Ile Asn Glu Lys Gly Val 245 250
255 Asp Ala Ala Phe Glu Cys Thr Ser Val Gln Pro Gly Phe Asp
Ala Cys 260 265 270
Leu Asp Ala Ile Arg Met Gly Gly Thr Val Val Ile Val Ala Ile Trp
275 280 285 Gly Lys Pro Ala
Ser Val Asp Met Ala Lys Leu Val Ile Lys Glu Ala 290
295 300 Asn Leu Leu Gly Thr Ile Ala Tyr
Asn Asn Thr His Pro Lys Thr Ile 305 310
315 320 Asp Leu Val Ser Thr Gly Lys Ile Lys Leu Asp Gln
Phe Ile Thr Ala 325 330
335 Lys Ile Gly Leu Asp Asp Leu Ile Asp Lys Gly Phe Asp Thr Leu Ile
340 345 350 His His Asn
Glu Thr Ala Val Lys Ile Leu Val Ser Pro Thr Gly Lys 355
360 365 Gly Leu 370
90705DNAPyrococcus furiosus 90atgaaggttg ccgtaattac tggggcatcc cgtggaatcg
gggaagctat agcaaaggcc 60cttgctgaag atggatattc ccttgcctta ggggctagaa
gtgttgatag gttagagaag 120attgccaagg aactcagcga aaaacatggg gtggaggtat
tttacgacta cctcgatgta 180tcaaaaccag aaagcgttga agagtttgca aggaaaacgc
tagctcactt tggagatgtg 240gacgttgttg tggccaatgc ggggcttggt tactttggta
ggcttgaaga gcttacagaa 300gagcagttcc acgaaatgat tgaagtaaac cttttgggag
tttggagaac aataaaagct 360ttcttaaact ccttaaagcg gactggagga gtggctattg
ttgttacttc agatgtttct 420gcaaggctac ttccatacgg tggaggttat gtggcaacta
aatgggctgc aagagcattg 480gtaaggacct tccagattga gaatccagat gtgaggttct
tcgagctaag acctggagca 540gtagatacat attttggagg gagcaaagct gggaagccaa
aggagcaagg gtatttaaaa 600cctgaggaag ttgctgaggc agtaaaatac ctcctaagac
ttccaaagga tgttagggtt 660gaggaattaa tgttgcgctc aatttatcaa aaacctgagt
attga 70591234PRTPyrococcus furiosus 91Met Lys Val Ala
Val Ile Thr Gly Ala Ser Arg Gly Ile Gly Glu Ala 1 5
10 15 Ile Ala Lys Ala Leu Ala Glu Asp Gly
Tyr Ser Leu Ala Leu Gly Ala 20 25
30 Arg Ser Val Asp Arg Leu Glu Lys Ile Ala Lys Glu Leu Ser
Glu Lys 35 40 45
His Gly Val Glu Val Phe Tyr Asp Tyr Leu Asp Val Ser Lys Pro Glu 50
55 60 Ser Val Glu Glu Phe
Ala Arg Lys Thr Leu Ala His Phe Gly Asp Val 65 70
75 80 Asp Val Val Val Ala Asn Ala Gly Leu Gly
Tyr Phe Gly Arg Leu Glu 85 90
95 Glu Leu Thr Glu Glu Gln Phe His Glu Met Ile Glu Val Asn Leu
Leu 100 105 110 Gly
Val Trp Arg Thr Ile Lys Ala Phe Leu Asn Ser Leu Lys Arg Thr 115
120 125 Gly Gly Val Ala Ile Val
Val Thr Ser Asp Val Ser Ala Arg Leu Leu 130 135
140 Pro Tyr Gly Gly Gly Tyr Val Ala Thr Lys Trp
Ala Ala Arg Ala Leu 145 150 155
160 Val Arg Thr Phe Gln Ile Glu Asn Pro Asp Val Arg Phe Phe Glu Leu
165 170 175 Arg Pro
Gly Ala Val Asp Thr Tyr Phe Gly Gly Ser Lys Ala Gly Lys 180
185 190 Pro Lys Glu Gln Gly Tyr Leu
Lys Pro Glu Glu Val Ala Glu Ala Val 195 200
205 Lys Tyr Leu Leu Arg Leu Pro Lys Asp Val Arg Val
Glu Glu Leu Met 210 215 220
Leu Arg Ser Ile Tyr Gln Lys Pro Glu Tyr 225 230
921665DNASalmonella typhimurium 92atgagatcga aaagatttga
agcactggcg aaacgccctg tgaatcagga cggctttgtt 60aaggagtgga tcgaagaagg
ctttatcgcg atggaaagcc cgaacgaccc aaaaccgtcg 120ataaaaatcg ttaacggcgc
ggtaaccgag ctggacggaa aaccggttag cgaattcgac 180ctgatcgacc actttatcgc
ccgctacggc atcaacctga accgcgccga agaagtgatg 240gcgatggatt cggtcaagct
ggctaacatg ctgtgcgatc cgaacgtcaa gcgcagcgaa 300atcgttccgc taaccaccgc
gatgacccca gcgaaaattg tcgaagtggt ttcgcatatg 360aacgtggttg agatgatgat
ggcgatgcag aaaatgcgcg cccgccgtac tccatctcaa 420caggcgcacg tcaccaacgt
taaagacaac ccggtgcaaa ttgccgccga tgccgccgaa 480ggcgcatggc gcgggtttga
cgaacaagag acgacggttg cggtagcgcg ctatgcgccg 540ttcaacgcca tcgcgctgct
ggttggttct caggtaggtc gtccgggggt actgactcaa 600tgctcgctgg aagaagccac
cgagctgaag ctcggcatgc tgggccacac ctgctacgcc 660gaaaccatct ccgtttacgg
caccgagccg gtcttcaccg acggtgacga taccccatgg 720tcgaagggct tcttagcctc
ttcctacgcc tctcgcggcc tgaaaatgcg cttcacctcc 780ggctccggct ccgaagtgca
gatgggctac gccgaaggca aatccatgct gtatctggaa 840gcgcgctgca tctatatcac
caaagccgcg ggcgttcagg ggctgcaaaa cggctccgta 900agcagcatcg gcgtaccgtc
tgccgtgccg tcaggcattc gtgccgtgct ggcggaaaac 960ctgatctgct cttcgctgga
tctggaatgc gcctccagta acgaccagac cttcacccac 1020tccgatatgc gtcgtaccgc
tcgcctgctg atgcagttcc tgccgggtac cgactttatc 1080tcctccggtt attccgcggt
gccgaactac gacaacatgt tcgccggttc caacgaagat 1140gcggaagact ttgacgacta
caacgttatc cagcgtgacc tgaaagtgga cggcggtctg 1200cgcccggttc gcgaagagga
cgttatcgcc atccgtaaca aagccgcccg cgcgctgcag 1260gccgtgtttg ccggaatggg
actgccgccg attaccgatg aagaagttga agccgcgacc 1320tatgcccacg gttcgaaaga
tatgccggag cgcaacatcg tcgaagacat caagttcgcc 1380caggaaatca tcaataaaaa
ccgcaacggt ctggaagttg tgaaagcgct ggctcagggc 1440gggtttaccg acgtggccca
ggacatgctc aacatccaga aagccaagct aaccggcgac 1500tatttgcaca cctccgccat
tatcgtcggc gacggacaag tgctctctgc ggttaatgac 1560gtcaatgact atgccggtcc
ggcaacaggt tatcgcctgc agggagaacg ctgggaagag 1620attaaaaaca tccctggcgc
tcttgatccc aacgagattg attaa 166593554PRTSalmonella
typhimurium 93Met Arg Ser Lys Arg Phe Glu Ala Leu Ala Lys Arg Pro Val Asn
Gln 1 5 10 15 Asp
Gly Phe Val Lys Glu Trp Ile Glu Glu Gly Phe Ile Ala Met Glu
20 25 30 Ser Pro Asn Asp Pro
Lys Pro Ser Ile Lys Ile Val Asn Gly Ala Val 35
40 45 Thr Glu Leu Asp Gly Lys Pro Val Ser
Glu Phe Asp Leu Ile Asp His 50 55
60 Phe Ile Ala Arg Tyr Gly Ile Asn Leu Asn Arg Ala Glu
Glu Val Met 65 70 75
80 Ala Met Asp Ser Val Lys Leu Ala Asn Met Leu Cys Asp Pro Asn Val
85 90 95 Lys Arg Ser Glu
Ile Val Pro Leu Thr Thr Ala Met Thr Pro Ala Lys 100
105 110 Ile Val Glu Val Val Ser His Met Asn
Val Val Glu Met Met Met Ala 115 120
125 Met Gln Lys Met Arg Ala Arg Arg Thr Pro Ser Gln Gln Ala
His Val 130 135 140
Thr Asn Val Lys Asp Asn Pro Val Gln Ile Ala Ala Asp Ala Ala Glu 145
150 155 160 Gly Ala Trp Arg Gly
Phe Asp Glu Gln Glu Thr Thr Val Ala Val Ala 165
170 175 Arg Tyr Ala Pro Phe Asn Ala Ile Ala Leu
Leu Val Gly Ser Gln Val 180 185
190 Gly Arg Pro Gly Val Leu Thr Gln Cys Ser Leu Glu Glu Ala Thr
Glu 195 200 205 Leu
Lys Leu Gly Met Leu Gly His Thr Cys Tyr Ala Glu Thr Ile Ser 210
215 220 Val Tyr Gly Thr Glu Pro
Val Phe Thr Asp Gly Asp Asp Thr Pro Trp 225 230
235 240 Ser Lys Gly Phe Leu Ala Ser Ser Tyr Ala Ser
Arg Gly Leu Lys Met 245 250
255 Arg Phe Thr Ser Gly Ser Gly Ser Glu Val Gln Met Gly Tyr Ala Glu
260 265 270 Gly Lys
Ser Met Leu Tyr Leu Glu Ala Arg Cys Ile Tyr Ile Thr Lys 275
280 285 Ala Ala Gly Val Gln Gly Leu
Gln Asn Gly Ser Val Ser Ser Ile Gly 290 295
300 Val Pro Ser Ala Val Pro Ser Gly Ile Arg Ala Val
Leu Ala Glu Asn 305 310 315
320 Leu Ile Cys Ser Ser Leu Asp Leu Glu Cys Ala Ser Ser Asn Asp Gln
325 330 335 Thr Phe Thr
His Ser Asp Met Arg Arg Thr Ala Arg Leu Leu Met Gln 340
345 350 Phe Leu Pro Gly Thr Asp Phe Ile
Ser Ser Gly Tyr Ser Ala Val Pro 355 360
365 Asn Tyr Asp Asn Met Phe Ala Gly Ser Asn Glu Asp Ala
Glu Asp Phe 370 375 380
Asp Asp Tyr Asn Val Ile Gln Arg Asp Leu Lys Val Asp Gly Gly Leu 385
390 395 400 Arg Pro Val Arg
Glu Glu Asp Val Ile Ala Ile Arg Asn Lys Ala Ala 405
410 415 Arg Ala Leu Gln Ala Val Phe Ala Gly
Met Gly Leu Pro Pro Ile Thr 420 425
430 Asp Glu Glu Val Glu Ala Ala Thr Tyr Ala His Gly Ser Lys
Asp Met 435 440 445
Pro Glu Arg Asn Ile Val Glu Asp Ile Lys Phe Ala Gln Glu Ile Ile 450
455 460 Asn Lys Asn Arg Asn
Gly Leu Glu Val Val Lys Ala Leu Ala Gln Gly 465 470
475 480 Gly Phe Thr Asp Val Ala Gln Asp Met Leu
Asn Ile Gln Lys Ala Lys 485 490
495 Leu Thr Gly Asp Tyr Leu His Thr Ser Ala Ile Ile Val Gly Asp
Gly 500 505 510 Gln
Val Leu Ser Ala Val Asn Asp Val Asn Asp Tyr Ala Gly Pro Ala 515
520 525 Thr Gly Tyr Arg Leu Gln
Gly Glu Arg Trp Glu Glu Ile Lys Asn Ile 530 535
540 Pro Gly Ala Leu Asp Pro Asn Glu Ile Asp 545
550 94675DNASalmonella typhimurium
94atggaaatta atgaaaaatt gctgcgccag ataattgaag acgtactccg cgatatgaag
60ggcagcgata aacccgtctc gtttaatgcg cctgcggcat ccacagcacc acagaccgct
120gcgcctgcgg gcgacggctt tctgaccgaa gtgggcgaag cgcgccaggg cactcagcag
180gacgaagtca ttatcgccgt cggcccggca tttggcctgg cgcaaaccgt caatatcgtc
240ggcttaccgc ataagagcat tctgcgcgaa gtcattgccg gtattgaaga agaaggcatc
300aaggcgcgcg tgattcgctg ctttaaatct tccgacgtgg cgttcgtcgc cgttgaaggt
360aaccgcctga gcggatccgg catctccatc ggcatccagt cgaaaggtac tacggttatc
420caccagcagg ggctaccgcc gctctccaac ctggagctgt tcccgcaggc accgctgctg
480acgctggaaa cctaccgtca gattggtaaa aacgccgccc gctatgcgaa acgagaatca
540ccgcagccgg tccctacgct caatgaccag atggcacgcc cgaagtacca ggcaaagtcg
600gccattttgc atattaaaga gaccaagtac gtcgtgacgg gcaaaaaccc gcaggaactg
660cgcgtggcgc tttga
67595224PRTSalmonella typhimurium 95Met Glu Ile Asn Glu Lys Leu Leu Arg
Gln Ile Ile Glu Asp Val Leu 1 5 10
15 Arg Asp Met Lys Gly Ser Asp Lys Pro Val Ser Phe Asn Ala
Pro Ala 20 25 30
Ala Ser Thr Ala Pro Gln Thr Ala Ala Pro Ala Gly Asp Gly Phe Leu
35 40 45 Thr Glu Val Gly
Glu Ala Arg Gln Gly Thr Gln Gln Asp Glu Val Ile 50
55 60 Ile Ala Val Gly Pro Ala Phe Gly
Leu Ala Gln Thr Val Asn Ile Val 65 70
75 80 Gly Leu Pro His Lys Ser Ile Leu Arg Glu Val Ile
Ala Gly Ile Glu 85 90
95 Glu Glu Gly Ile Lys Ala Arg Val Ile Arg Cys Phe Lys Ser Ser Asp
100 105 110 Val Ala Phe
Val Ala Val Glu Gly Asn Arg Leu Ser Gly Ser Gly Ile 115
120 125 Ser Ile Gly Ile Gln Ser Lys Gly
Thr Thr Val Ile His Gln Gln Gly 130 135
140 Leu Pro Pro Leu Ser Asn Leu Glu Leu Phe Pro Gln Ala
Pro Leu Leu 145 150 155
160 Thr Leu Glu Thr Tyr Arg Gln Ile Gly Lys Asn Ala Ala Arg Tyr Ala
165 170 175 Lys Arg Glu Ser
Pro Gln Pro Val Pro Thr Leu Asn Asp Gln Met Ala 180
185 190 Arg Pro Lys Tyr Gln Ala Lys Ser Ala
Ile Leu His Ile Lys Glu Thr 195 200
205 Lys Tyr Val Val Thr Gly Lys Asn Pro Gln Glu Leu Arg Val
Ala Leu 210 215 220
96522DNASalmonella typhimurium 96atgaataccg acgcaattga atcgatggtc
cgggacgtat tgagccgcat gaacagcctg 60cagggcgatg cgccagcagc ggctcctgcg
gcaggcggca cgtcccgcag cgcaaaggtc 120agcgactacc cgctggcgaa caaacacccg
gaatgggtga aaaccgccac caataaaacg 180ctggacgact ttacgctgga aaacgtgctg
agcaataaag tcaccgctca ggatatgcgt 240attaccccgg aaaccctgcg cttacaggcc
tctatcgcca aagatgcggg tcgcgaccgg 300ctggcgatga acttcgaacg cgccgccgaa
ctgaccgcgg taccggacga tcgcattctt 360gaaatctaca acgcccttcg tccgtatcgt
tcaacgaaag aagagctgct cgctatcgcc 420gacgatctcg aaaaccgtta tcaggcaaag
atttgcgcag ctttcgttcg tgaagcggca 480gggctgtacg ttgagcgtaa aaaactcaaa
ggcgacgatt aa 52297173PRTSalmonella typhimurium
97Met Asn Thr Asp Ala Ile Glu Ser Met Val Arg Asp Val Leu Ser Arg 1
5 10 15 Met Asn Ser Leu
Gln Gly Asp Ala Pro Ala Ala Ala Pro Ala Ala Gly 20
25 30 Gly Thr Ser Arg Ser Ala Lys Val Ser
Asp Tyr Pro Leu Ala Asn Lys 35 40
45 His Pro Glu Trp Val Lys Thr Ala Thr Asn Lys Thr Leu Asp
Asp Phe 50 55 60
Thr Leu Glu Asn Val Leu Ser Asn Lys Val Thr Ala Gln Asp Met Arg 65
70 75 80 Ile Thr Pro Glu Thr
Leu Arg Leu Gln Ala Ser Ile Ala Lys Asp Ala 85
90 95 Gly Arg Asp Arg Leu Ala Met Asn Phe Glu
Arg Ala Ala Glu Leu Thr 100 105
110 Ala Val Pro Asp Asp Arg Ile Leu Glu Ile Tyr Asn Ala Leu Arg
Pro 115 120 125 Tyr
Arg Ser Thr Lys Glu Glu Leu Leu Ala Ile Ala Asp Asp Leu Glu 130
135 140 Asn Arg Tyr Gln Ala Lys
Ile Cys Ala Ala Phe Val Arg Glu Ala Ala 145 150
155 160 Gly Leu Tyr Val Glu Arg Lys Lys Leu Lys Gly
Asp Asp 165 170
981677DNALactobacillus collinoides 98ttggaacgtc aaaaaagatt tgaaaaatta
gagaaacgtc cagtgcattt agatgggttc 60gttaagaact gggacgacga aggtttagtt
gcccttaacg gtaagaacga tccaaagcca 120agcattacga tcgaaaacgg tgttgttact
gaaatggatg gtaagaagaa ggcagacttc 180gaccttatcg acaagtacat cgctgaatac
gggatcaact tggacaatgc tgaaaagact 240ttaaacacag attcagttaa gatcgccaac
atgatgtgtg atcctaacgt ctcccgtgct 300gaaattattg aatatacaac tgctatgaca
ccagccaagg ctgctgaagt tatcagccag 360ttaaacttcg ctgaaatgat catggcaact
caaaagatgc ggccacgtcg gacccctatg 420actcaagtcc acgctaccaa cactttggat
aacccagttg aaatcgctgc tgatgctgcc 480gaagctgcat tacgtggggt tcctgaagaa
gaaaccacca ctgccattgc tcggtatgcg 540ccaatgaacg ctatttcaat catggttggg
gcccaagcag gccgtcctgg tgttatcacc 600caatgttcag ttgaagaagc tgacgaattg
agtttgggga tgcgtgggtt tactgcctat 660gctgaaacca tttcagttta tgggactgac
cgggtcttca ctgatggtga tgatacccct 720tggtcaaaag gtttcttagc ttcttgctac
gcttcacgtg gtttgaagat gcggtttact 780tcaggtgccg gttcagaagc tatgatgggc
tacactgaag gtaaatcaat gctttacctt 840gaagctcgtt gtatctacat taccaaggcg
tcaggtgttc aaggtctgca aaacggtggt 900gttagttgta tcgggatgcc aggtgccgtc
gttggtggta tccgtgaagt cttaggtgaa 960aacttactat gtatgtcact tgatgttgaa
tgtgcttctg gttgtgacca agccttctct 1020cactctgaca ttcgtcggac tggccggatg
attggccaat tcatcgctgg tactgattac 1080ctgtcatcag gttacgctgc cgaagaaaac
atggataaca ccttcgctgg ttcaaacatg 1140gatgttctgg actacgatga ttacatcact
ttggaacgtg atatggctat taacggtggt 1200atcatgccaa ttaccgaaga ggaatctatt
aagattcgtc acaaggctgc ggttgctatc 1260caagctgtct ttgatggctt aggcctacca
cagatcactg atgaagaagt tgaagccgca 1320acttatggca gcaattcaaa cgacatgcca
aaacgtgaca tggttcaaga tatgaaagct 1380gctcaaggtc tgatgactcg tggcattact
gttgttgacg ttatcaaggc cttatatgac 1440catgatatta aagacgtcgc tgaggctgtg
cttaagttag cgcaacaaaa ggtttgtggt 1500gattacctgc aaacatctgc tgtcttcttg
gatggttgga agtgtacttc agctattaac 1560aacgctaacg attacaaagg cccaggtact
ggttaccgtc tatgggaaga caaagacaaa 1620tgggatcgtc tagaaaacgt tccgtgggct
ttggatcctc agaagttgga attctaa 167799558PRTLactobacillus collinoides
99Met Glu Arg Gln Lys Arg Phe Glu Lys Leu Glu Lys Arg Pro Val His 1
5 10 15 Leu Asp Gly Phe
Val Lys Asn Trp Asp Asp Glu Gly Leu Val Ala Leu 20
25 30 Asn Gly Lys Asn Asp Pro Lys Pro Ser
Ile Thr Ile Glu Asn Gly Val 35 40
45 Val Thr Glu Met Asp Gly Lys Lys Lys Ala Asp Phe Asp Leu
Ile Asp 50 55 60
Lys Tyr Ile Ala Glu Tyr Gly Ile Asn Leu Asp Asn Ala Glu Lys Thr 65
70 75 80 Leu Asn Thr Asp Ser
Val Lys Ile Ala Asn Met Met Cys Asp Pro Asn 85
90 95 Val Ser Arg Ala Glu Ile Ile Glu Tyr Thr
Thr Ala Met Thr Pro Ala 100 105
110 Lys Ala Ala Glu Val Ile Ser Gln Leu Asn Phe Ala Glu Met Ile
Met 115 120 125 Ala
Thr Gln Lys Met Arg Pro Arg Arg Thr Pro Met Thr Gln Val His 130
135 140 Ala Thr Asn Thr Leu Asp
Asn Pro Val Glu Ile Ala Ala Asp Ala Ala 145 150
155 160 Glu Ala Ala Leu Arg Gly Val Pro Glu Glu Glu
Thr Thr Thr Ala Ile 165 170
175 Ala Arg Tyr Ala Pro Met Asn Ala Ile Ser Ile Met Val Gly Ala Gln
180 185 190 Ala Gly
Arg Pro Gly Val Ile Thr Gln Cys Ser Val Glu Glu Ala Asp 195
200 205 Glu Leu Ser Leu Gly Met Arg
Gly Phe Thr Ala Tyr Ala Glu Thr Ile 210 215
220 Ser Val Tyr Gly Thr Asp Arg Val Phe Thr Asp Gly
Asp Asp Thr Pro 225 230 235
240 Trp Ser Lys Gly Phe Leu Ala Ser Cys Tyr Ala Ser Arg Gly Leu Lys
245 250 255 Met Arg Phe
Thr Ser Gly Ala Gly Ser Glu Ala Met Met Gly Tyr Thr 260
265 270 Glu Gly Lys Ser Met Leu Tyr Leu
Glu Ala Arg Cys Ile Tyr Ile Thr 275 280
285 Lys Ala Ser Gly Val Gln Gly Leu Gln Asn Gly Gly Val
Ser Cys Ile 290 295 300
Gly Met Pro Gly Ala Val Val Gly Gly Ile Arg Glu Val Leu Gly Glu 305
310 315 320 Asn Leu Leu Cys
Met Ser Leu Asp Val Glu Cys Ala Ser Gly Cys Asp 325
330 335 Gln Ala Phe Ser His Ser Asp Ile Arg
Arg Thr Gly Arg Met Ile Gly 340 345
350 Gln Phe Ile Ala Gly Thr Asp Tyr Leu Ser Ser Gly Tyr Ala
Ala Glu 355 360 365
Glu Asn Met Asp Asn Thr Phe Ala Gly Ser Asn Met Asp Val Leu Asp 370
375 380 Tyr Asp Asp Tyr Ile
Thr Leu Glu Arg Asp Met Ala Ile Asn Gly Gly 385 390
395 400 Ile Met Pro Ile Thr Glu Glu Glu Ser Ile
Lys Ile Arg His Lys Ala 405 410
415 Ala Val Ala Ile Gln Ala Val Phe Asp Gly Leu Gly Leu Pro Gln
Ile 420 425 430 Thr
Asp Glu Glu Val Glu Ala Ala Thr Tyr Gly Ser Asn Ser Asn Asp 435
440 445 Met Pro Lys Arg Asp Met
Val Gln Asp Met Lys Ala Ala Gln Gly Leu 450 455
460 Met Thr Arg Gly Ile Thr Val Val Asp Val Ile
Lys Ala Leu Tyr Asp 465 470 475
480 His Asp Ile Lys Asp Val Ala Glu Ala Val Leu Lys Leu Ala Gln Gln
485 490 495 Lys Val
Cys Gly Asp Tyr Leu Gln Thr Ser Ala Val Phe Leu Asp Gly 500
505 510 Trp Lys Cys Thr Ser Ala Ile
Asn Asn Ala Asn Asp Tyr Lys Gly Pro 515 520
525 Gly Thr Gly Tyr Arg Leu Trp Glu Asp Lys Asp Lys
Trp Asp Arg Leu 530 535 540
Glu Asn Val Pro Trp Ala Leu Asp Pro Gln Lys Leu Glu Phe 545
550 555 100693DNALactobacillus
collinoides 100gtgagttcag aaatcgatga aacattgctt agaaatatca ttaaaggcgt
tttaaatgaa 60gttcaaaact ctgatacgcc aatttccttt ggtggccaag atgcagcccc
agttgccggt 120gccaaggaag gtgccgcacc agaaaagaag ttggattggt tccaacacgt
tggaatcgcc 180aaaccaggtt tgtcaaagga tgaagttgta attggtgttg ccccagcatt
tgctgaagtg 240ttgacgcaaa ctatgacgaa gatccaacac aaagacatcc tgcgtcaaat
cattgccgga 300gttgaagaag aaggtctcaa ggcccgtgtc gttaaggttt atcggacttc
agacgtttcc 360ttcgtttccg ctgatgttga caagttgtca ggttcaggaa tttcagttgc
cgttcaatca 420aaggggacaa cgattattca ccaaaaggat caagcaccgt tgtcaaacct
tgaattgttc 480ccacaggctc cagttttgac attggacgct taccgtcaaa tcggtaagaa
cgctgcccag 540tatgctaagg gtatgtcacc aaccccagtg ccaacaatta acgaccagat
ggcacgtgtg 600caatatcaag cactttctgc tttgatgcac atcaaggaaa caaaacaggt
tgttgttggg 660aagcctgctg aagaaattaa ggtaaccttt tag
693101230PRTLactobacillus collinoides 101Met Ser Ser Glu Ile
Asp Glu Thr Leu Leu Arg Asn Ile Ile Lys Gly 1 5
10 15 Val Leu Asn Glu Val Gln Asn Ser Asp Thr
Pro Ile Ser Phe Gly Gly 20 25
30 Gln Asp Ala Ala Pro Val Ala Gly Ala Lys Glu Gly Ala Ala Pro
Glu 35 40 45 Lys
Lys Leu Asp Trp Phe Gln His Val Gly Ile Ala Lys Pro Gly Leu 50
55 60 Ser Lys Asp Glu Val Val
Ile Gly Val Ala Pro Ala Phe Ala Glu Val 65 70
75 80 Leu Thr Gln Thr Met Thr Lys Ile Gln His Lys
Asp Ile Leu Arg Gln 85 90
95 Ile Ile Ala Gly Val Glu Glu Glu Gly Leu Lys Ala Arg Val Val Lys
100 105 110 Val Tyr
Arg Thr Ser Asp Val Ser Phe Val Ser Ala Asp Val Asp Lys 115
120 125 Leu Ser Gly Ser Gly Ile Ser
Val Ala Val Gln Ser Lys Gly Thr Thr 130 135
140 Ile Ile His Gln Lys Asp Gln Ala Pro Leu Ser Asn
Leu Glu Leu Phe 145 150 155
160 Pro Gln Ala Pro Val Leu Thr Leu Asp Ala Tyr Arg Gln Ile Gly Lys
165 170 175 Asn Ala Ala
Gln Tyr Ala Lys Gly Met Ser Pro Thr Pro Val Pro Thr 180
185 190 Ile Asn Asp Gln Met Ala Arg Val
Gln Tyr Gln Ala Leu Ser Ala Leu 195 200
205 Met His Ile Lys Glu Thr Lys Gln Val Val Val Gly Lys
Pro Ala Glu 210 215 220
Glu Ile Lys Val Thr Phe 225 230 102522DNALactobacillus
collinoides 102atgagtgaag tagatgactt agtagctaga attgctgctc agctacaaca
aagtggaaac 60gcttctagtg cctcaactag tgccggtact tctgctggtt ccgagaaaga
attaggcgca 120gcagattacc cactatttga aaagcaccca gatcaaatca agacgccatc
aggtaaaaat 180gttgaagaaa tcaccttgga aaatgttatt aacggcaagg tagacgcaaa
ggatatgcgg 240attacgcccg caaccctgaa gttacaaggt gaaattgctg ccaacgcagg
tcggccagca 300atccaacgga acttccagcg ggcttctgaa ttaacttcag ttcccgatga
tgttgttttg 360gacttatata attcattacg gccattccgt tcaaccaagc aagaattatt
ggataccgcc 420aaggagcttc gtgacaagta tcacgcacct atctgtgccg gctggttcga
agaagcagcc 480gaaaactacg aagtcaacaa gaagttgaag ggcgataact ag
522103173PRTLactobacillus collinoides 103Met Ser Glu Val Asp
Asp Leu Val Ala Arg Ile Ala Ala Gln Leu Gln 1 5
10 15 Gln Ser Gly Asn Ala Ser Ser Ala Ser Thr
Ser Ala Gly Thr Ser Ala 20 25
30 Gly Ser Glu Lys Glu Leu Gly Ala Ala Asp Tyr Pro Leu Phe Glu
Lys 35 40 45 His
Pro Asp Gln Ile Lys Thr Pro Ser Gly Lys Asn Val Glu Glu Ile 50
55 60 Thr Leu Glu Asn Val Ile
Asn Gly Lys Val Asp Ala Lys Asp Met Arg 65 70
75 80 Ile Thr Pro Ala Thr Leu Lys Leu Gln Gly Glu
Ile Ala Ala Asn Ala 85 90
95 Gly Arg Pro Ala Ile Gln Arg Asn Phe Gln Arg Ala Ser Glu Leu Thr
100 105 110 Ser Val
Pro Asp Asp Val Val Leu Asp Leu Tyr Asn Ser Leu Arg Pro 115
120 125 Phe Arg Ser Thr Lys Gln Glu
Leu Leu Asp Thr Ala Lys Glu Leu Arg 130 135
140 Asp Lys Tyr His Ala Pro Ile Cys Ala Gly Trp Phe
Glu Glu Ala Ala 145 150 155
160 Glu Asn Tyr Glu Val Asn Lys Lys Leu Lys Gly Asp Asn
165 170 1041665DNAKlebsiella pneumoniae
104atgagatcga aaagatttga agcactggcg aaacgccctg tgaatcagga tggtttcgtt
60aaggagtgga ttgaagaggg ctttatcgcg atggaaagtc ctaacgatcc caaaccttct
120atccgcatcg tcaacggcgc ggtgaccgaa ctcgacggta aaccggttga cgagttcgac
180ctgattgacc actttatcgc gcgctacggc attaatctcg cccgggccga agaagtgatg
240gccatggatt cggttaagct cgccaacatg ctctgcgacc cgaacgttaa acgcagcgac
300atcgtgccgc tcactaccgc gatgaccccg gcgaaaatcg tggaagtggt gtcgcatatg
360aacgtggtcg agatgatgat ggcgatgcaa aaaatgcgcg cccgccgcac gccgtcccag
420caggcgcatg tcactaatat caaagataat ccggtacaga ttgccgccga cgccgctgaa
480ggcgcatggc gcggctttga cgaacaggag accaccgtcg ccgtggcgcg ctacgcgcgg
540ttcaacgcca tcgccctgct ggtgggttca caggttggcc gccccggcgt cctcacccag
600tgttcgctgg aagaagccac cgagctgaaa ctgggcatgc tgggccacac ctgctatgcc
660gaaaccattt cggtatacgg tacggaaccg gtgtttaccg atggcgatga cactccatgg
720tcgaaaggct tcctcgcctc ctcctacgcc tcgcgcggcc tgaaaatgcg ctttacctcc
780ggttccggtt ctgaagtaca gatgggctat gccgaaggca aatcgatgct ttatctcgaa
840gcgcgctgca tctacatcac caaagccgcc ggggtgcaag gcctgcagaa tggctccgtc
900agctgtatcg gcgtaccgtc cgccgtgccg tccgggatcc gcgccgtact ggcggaaaac
960ctgatctgct cagcgctgga tctggagtgc gcctccagca acgatcaaac ctttacccac
1020tcggatatgc ggcgtaccgc gcgtctgctg atgcagttcc tgccaggcac cgacttcatc
1080tcctccggtt actcggcggt gcccaactac gacaacatgt tcgccggttc caacgaagat
1140gccgaagact tcgatgacta caacgtgatc cagcgcgacc tgaaggtcga tgggggtctg
1200cggccggtgc gtgaagagga cgtgatcgcc attcgcaaca aagccgcccg cgcgctgcag
1260gcggtatttg ccggcatggg tttgccgcct attacggatg aagaggtaga agccgccacc
1320tacgcccacg gttcaaaaga tatgcctgag cgcaatatcg tcgaggacat caagtttgct
1380caggagatca tcaacaagaa ccgcaacggc ctggaggtgg tgaaagccct ggcgaaaggc
1440ggcttccccg atgtcgccca ggacatgctc aatattcaga aagccaagct caccggcgac
1500tacctgcata cctccgccat cattgttggc gagggccagg tgctctcggc cgtgaatgac
1560gtgaacgatt atgccggtcc ggcaacaggc taccgcctgc aaggcgagcg ctgggaagag
1620attaaaaata tcccgggcgc gctcgatccc aatgaacttg gctaa
1665105554PRTKlebsiella pneumoniae 105Met Arg Ser Lys Arg Phe Glu Ala Leu
Ala Lys Arg Pro Val Asn Gln 1 5 10
15 Asp Gly Phe Val Lys Glu Trp Ile Glu Glu Gly Phe Ile Ala
Met Glu 20 25 30
Ser Pro Asn Asp Pro Lys Pro Ser Ile Arg Ile Val Asn Gly Ala Val
35 40 45 Thr Glu Leu Asp
Gly Lys Pro Val Asp Glu Phe Asp Leu Ile Asp His 50
55 60 Phe Ile Ala Arg Tyr Gly Ile Asn
Leu Ala Arg Ala Glu Glu Val Met 65 70
75 80 Ala Met Asp Ser Val Lys Leu Ala Asn Met Leu Cys
Asp Pro Asn Val 85 90
95 Lys Arg Ser Asp Ile Val Pro Leu Thr Thr Ala Met Thr Pro Ala Lys
100 105 110 Ile Val Glu
Val Val Ser His Met Asn Val Val Glu Met Met Met Ala 115
120 125 Met Gln Lys Met Arg Ala Arg Arg
Thr Pro Ser Gln Gln Ala His Val 130 135
140 Thr Asn Ile Lys Asp Asn Pro Val Gln Ile Ala Ala Asp
Ala Ala Glu 145 150 155
160 Gly Ala Trp Arg Gly Phe Asp Glu Gln Glu Thr Thr Val Ala Val Ala
165 170 175 Arg Tyr Ala Arg
Phe Asn Ala Ile Ala Leu Leu Val Gly Ser Gln Val 180
185 190 Gly Arg Pro Gly Val Leu Thr Gln Cys
Ser Leu Glu Glu Ala Thr Glu 195 200
205 Leu Lys Leu Gly Met Leu Gly His Thr Cys Tyr Ala Glu Thr
Ile Ser 210 215 220
Val Tyr Gly Thr Glu Pro Val Phe Thr Asp Gly Asp Asp Thr Pro Trp 225
230 235 240 Ser Lys Gly Phe Leu
Ala Ser Ser Tyr Ala Ser Arg Gly Leu Lys Met 245
250 255 Arg Phe Thr Ser Gly Ser Gly Ser Glu Val
Gln Met Gly Tyr Ala Glu 260 265
270 Gly Lys Ser Met Leu Tyr Leu Glu Ala Arg Cys Ile Tyr Ile Thr
Lys 275 280 285 Ala
Ala Gly Val Gln Gly Leu Gln Asn Gly Ser Val Ser Cys Ile Gly 290
295 300 Val Pro Ser Ala Val Pro
Ser Gly Ile Arg Ala Val Leu Ala Glu Asn 305 310
315 320 Leu Ile Cys Ser Ala Leu Asp Leu Glu Cys Ala
Ser Ser Asn Asp Gln 325 330
335 Thr Phe Thr His Ser Asp Met Arg Arg Thr Ala Arg Leu Leu Met Gln
340 345 350 Phe Leu
Pro Gly Thr Asp Phe Ile Ser Ser Gly Tyr Ser Ala Val Pro 355
360 365 Asn Tyr Asp Asn Met Phe Ala
Gly Ser Asn Glu Asp Ala Glu Asp Phe 370 375
380 Asp Asp Tyr Asn Val Ile Gln Arg Asp Leu Lys Val
Asp Gly Gly Leu 385 390 395
400 Arg Pro Val Arg Glu Glu Asp Val Ile Ala Ile Arg Asn Lys Ala Ala
405 410 415 Arg Ala Leu
Gln Ala Val Phe Ala Gly Met Gly Leu Pro Pro Ile Thr 420
425 430 Asp Glu Glu Val Glu Ala Ala Thr
Tyr Ala His Gly Ser Lys Asp Met 435 440
445 Pro Glu Arg Asn Ile Val Glu Asp Ile Lys Phe Ala Gln
Glu Ile Ile 450 455 460
Asn Lys Asn Arg Asn Gly Leu Glu Val Val Lys Ala Leu Ala Lys Gly 465
470 475 480 Gly Phe Pro Asp
Val Ala Gln Asp Met Leu Asn Ile Gln Lys Ala Lys 485
490 495 Leu Thr Gly Asp Tyr Leu His Thr Ser
Ala Ile Ile Val Gly Glu Gly 500 505
510 Gln Val Leu Ser Ala Val Asn Asp Val Asn Asp Tyr Ala Gly
Pro Ala 515 520 525
Thr Gly Tyr Arg Leu Gln Gly Glu Arg Trp Glu Glu Ile Lys Asn Ile 530
535 540 Pro Gly Ala Leu Asp
Pro Asn Glu Leu Gly 545 550
106687DNAKlebsiella pneumoniae 106atggaaatta acgaaacgct gctgcgccag
attatcgaag aggtgctgtc ggagatgaaa 60tcaggcgcag ataagccggt ctcctttagc
gcgccggcgt ctgtcgcctc tgccgcgccg 120gtcgccgttg cgcctgtgtc cggcgacagc
ttcctgacgg aaatcggcga agccaaaccc 180ggcacgcagc aggatgaagt cattattgcc
gtcgggccag cgtttggtct ggcgcaaacc 240gccaatatcg tcggcattcc gcataaaaat
attctgcgcg aagtgatcgc cggcattgag 300gaagaaggca tcaaagcccg ggtgatccgc
tgctttaagt catctgacgt cgccttcgtg 360gcagtggaag gcaaccgcct gagcggctcc
ggcatctcga tcggtattca gtcgaaaggc 420accaccgtca tccaccagcg cggcctgccg
ccgctttcca atctggaact cttcccgcag 480gcgccgctgt taacgctgga aacctaccgt
cagattggca aaaacgccgc gcgctacgcc 540aaacgcgagt cgccgcagcc ggtgccgacg
cttaacgatc agatggctcg tcccaaatac 600caggcgaagt cggccatttt gcacattaaa
gagaccaaat acgtggtgac gggcaaaaac 660ccgcaggaac tgcgcgtggc gctttaa
687107228PRTKlebsiella pneumoniae
107Met Glu Ile Asn Glu Thr Leu Leu Arg Gln Ile Ile Glu Glu Val Leu 1
5 10 15 Ser Glu Met Lys
Ser Gly Ala Asp Lys Pro Val Ser Phe Ser Ala Pro 20
25 30 Ala Ser Val Ala Ser Ala Ala Pro Val
Ala Val Ala Pro Val Ser Gly 35 40
45 Asp Ser Phe Leu Thr Glu Ile Gly Glu Ala Lys Pro Gly Thr
Gln Gln 50 55 60
Asp Glu Val Ile Ile Ala Val Gly Pro Ala Phe Gly Leu Ala Gln Thr 65
70 75 80 Ala Asn Ile Val Gly
Ile Pro His Lys Asn Ile Leu Arg Glu Val Ile 85
90 95 Ala Gly Ile Glu Glu Glu Gly Ile Lys Ala
Arg Val Ile Arg Cys Phe 100 105
110 Lys Ser Ser Asp Val Ala Phe Val Ala Val Glu Gly Asn Arg Leu
Ser 115 120 125 Gly
Ser Gly Ile Ser Ile Gly Ile Gln Ser Lys Gly Thr Thr Val Ile 130
135 140 His Gln Arg Gly Leu Pro
Pro Leu Ser Asn Leu Glu Leu Phe Pro Gln 145 150
155 160 Ala Pro Leu Leu Thr Leu Glu Thr Tyr Arg Gln
Ile Gly Lys Asn Ala 165 170
175 Ala Arg Tyr Ala Lys Arg Glu Ser Pro Gln Pro Val Pro Thr Leu Asn
180 185 190 Asp Gln
Met Ala Arg Pro Lys Tyr Gln Ala Lys Ser Ala Ile Leu His 195
200 205 Ile Lys Glu Thr Lys Tyr Val
Val Thr Gly Lys Asn Pro Gln Glu Leu 210 215
220 Arg Val Ala Leu 225
108525DNAKlebsiella pneumoniae 108atgaataccg acgcaattga atccatggta
cgcgacgtgc tgagccggat gaacagccta 60caggacgggg taacgcccgc gccagccgcg
ccgacaaacg acaccgttcg ccagccaaaa 120gttagcgact acccgttagc gacctgccat
ccggagtggg tcaaaaccgc taccaataaa 180acgctcgatg acctgacgct ggagaacgta
ttaagcgatc gcgttacggc gcaggacatg 240cgcatcactc cggaaacgct gcgtatgcag
gcggcgatcg cccaggatgc cggacgcgat 300cggctggcga tgaactttga gcgggccgca
gagctcaccg cggttcccga cgaccgaatc 360cttgagatct acaacgccct gcgcccatac
cgttccaccc aggcggagct actggcgatc 420gctgatgacc tcgagcatcg ctaccaggca
cgactctgtg ccgcctttgt tcgggaagcg 480gccgggctgt acatcgagcg taagaagctg
aaaggcgacg attaa 525109174PRTKlebsiella pneumoniae
109Met Asn Thr Asp Ala Ile Glu Ser Met Val Arg Asp Val Leu Ser Arg 1
5 10 15 Met Asn Ser Leu
Gln Asp Gly Val Thr Pro Ala Pro Ala Ala Pro Thr 20
25 30 Asn Asp Thr Val Arg Gln Pro Lys Val
Ser Asp Tyr Pro Leu Ala Thr 35 40
45 Cys His Pro Glu Trp Val Lys Thr Ala Thr Asn Lys Thr Leu
Asp Asp 50 55 60
Leu Thr Leu Glu Asn Val Leu Ser Asp Arg Val Thr Ala Gln Asp Met 65
70 75 80 Arg Ile Thr Pro Glu
Thr Leu Arg Met Gln Ala Ala Ile Ala Gln Asp 85
90 95 Ala Gly Arg Asp Arg Leu Ala Met Asn Phe
Glu Arg Ala Ala Glu Leu 100 105
110 Thr Ala Val Pro Asp Asp Arg Ile Leu Glu Ile Tyr Asn Ala Leu
Arg 115 120 125 Pro
Tyr Arg Ser Thr Gln Ala Glu Leu Leu Ala Ile Ala Asp Asp Leu 130
135 140 Glu His Arg Tyr Gln Ala
Arg Leu Cys Ala Ala Phe Val Arg Glu Ala 145 150
155 160 Ala Gly Leu Tyr Ile Glu Arg Lys Lys Leu Lys
Gly Asp Asp 165 170
1101833DNAKlebsiella oxytoca 110atgcgatata tagctggcat tgatatcggc
aactcatcga cggaagtcgc cctggcgacc 60ctggatgagg ctggcgcgct gacgatcacc
cacagcgcgc tggcggaaac caccggaatc 120aaaggcacgt tgcgtaacgt gttcgggatt
caggaggcgc tcgccctcgt cgccagaggc 180gccgggatcg ccgtcagcga tatttcgctc
atccgcatca acgaagcgac gccggtgatt 240ggcgatgtgg cgatggaaac cattaccgaa
accatcatca ccgaatcgac catgatcggc 300cataacccga aaacgcccgg cggcgcgggg
cttggcacag gcatcaccat tacgccgcag 360gagctgctaa cccgcccggc ggacgcgccc
tatatcctgg tggtgtcgtc ggcgttcgat 420tttgccgata tcgccagcgt gattaacgct
tccctgcgcg ccgggtatca gattaccggc 480gtcattttac agcgcgacga tggcgtgctg
gtcagcaacc ggctggaaaa accgctgccg 540atcgttgacg aagtgctgta catcgaccgc
attccgctgg ggatgctggc ggcgattgag 600gtcgccgttc cggggaaggt catcgaaacc
ctctctaacc cttacggcat cgccaccgtc 660tttaacctca gccccgagga gacgaagaac
atcgtcccga tggcccgggc gctgattggc 720aaccgttccg ccgtggtggt caaaacgcca
tccggcgacg tcaaagcgcg cgcgataccc 780gccggtaatc ttgagctgct ggcccagggc
cgtagcgtgc gcgtggatgt ggccgccggc 840gccgaagcca tcatgaaagc ggtcgacggc
tgcggcaggc tcgataacgt caccggcgaa 900tccggcacca atatcggcgg catgctggaa
cacgtgcgcc agaccatggc cgagctgacc 960aacaagccga gcagcgaaat atttattcag
gacctgctgg ccgttgatac ctcggtaccg 1020gtgagcgtta ccggcggtct ggccggggag
ttctcgctgg agcaggccgt gggcatcgcc 1080tcgatggtga aatcggatcg cctgcagatg
gcaatgatcg cccgcgaaat cgagcagaag 1140ctcaatatcg acgtgcagat cggcggcgca
gaggccgaag ccgccatcct gggggcgctg 1200accacgccgg gcaccacccg accgctggcg
atcctcgacc tcggcgcggg ctccaccgat 1260gcctccatca tcaaccccaa aggcgacatc
atcgccaccc atctcgccgg cgcaggcgac 1320atggtgacga tgattattgc ccgcgagctg
gggctggaag accgctatct ggcggaagag 1380atcaagaagt acccgctggc taaggtggaa
agcctgttcc atttacgcca cgaggacggc 1440agcgtgcagt tcttctccac gccgctgccg
cccgccgtgt tcgcccgcgt ctgcgtggtg 1500aaagcggacg aactggtgcc gctgcccggc
gatttagcgc tggaaaaagt gcgcgccatt 1560cgccgcagcg ccaaagagcg ggtctttgtc
accaacgccc tgcgcgcgct gcgtcaggtc 1620agccccaccg gcaacattcg cgatattccg
ttcgtggtgc tggtcggcgg ttcgtcgctg 1680gatttcgaag tcccgcagct ggtcaccgat
gcgctggcgc actaccgcct ggttgccgga 1740cggggaaata ttcgcggcag cgagggcccc
cgaaacgcgg tggccaccgg cctgattctc 1800tcctggcata aggagtttgc gcatgaacgg
taa 1833111610PRTKlebsiella oxytoca 111Met
Arg Tyr Ile Ala Gly Ile Asp Ile Gly Asn Ser Ser Thr Glu Val 1
5 10 15 Ala Leu Ala Thr Leu Asp
Glu Ala Gly Ala Leu Thr Ile Thr His Ser 20
25 30 Ala Leu Ala Glu Thr Thr Gly Ile Lys Gly
Thr Leu Arg Asn Val Phe 35 40
45 Gly Ile Gln Glu Ala Leu Ala Leu Val Ala Arg Gly Ala Gly
Ile Ala 50 55 60
Val Ser Asp Ile Ser Leu Ile Arg Ile Asn Glu Ala Thr Pro Val Ile 65
70 75 80 Gly Asp Val Ala Met
Glu Thr Ile Thr Glu Thr Ile Ile Thr Glu Ser 85
90 95 Thr Met Ile Gly His Asn Pro Lys Thr Pro
Gly Gly Ala Gly Leu Gly 100 105
110 Thr Gly Ile Thr Ile Thr Pro Gln Glu Leu Leu Thr Arg Pro Ala
Asp 115 120 125 Ala
Pro Tyr Ile Leu Val Val Ser Ser Ala Phe Asp Phe Ala Asp Ile 130
135 140 Ala Ser Val Ile Asn Ala
Ser Leu Arg Ala Gly Tyr Gln Ile Thr Gly 145 150
155 160 Val Ile Leu Gln Arg Asp Asp Gly Val Leu Val
Ser Asn Arg Leu Glu 165 170
175 Lys Pro Leu Pro Ile Val Asp Glu Val Leu Tyr Ile Asp Arg Ile Pro
180 185 190 Leu Gly
Met Leu Ala Ala Ile Glu Val Ala Val Pro Gly Lys Val Ile 195
200 205 Glu Thr Leu Ser Asn Pro Tyr
Gly Ile Ala Thr Val Phe Asn Leu Ser 210 215
220 Pro Glu Glu Thr Lys Asn Ile Val Pro Met Ala Arg
Ala Leu Ile Gly 225 230 235
240 Asn Arg Ser Ala Val Val Val Lys Thr Pro Ser Gly Asp Val Lys Ala
245 250 255 Arg Ala Ile
Pro Ala Gly Asn Leu Glu Leu Leu Ala Gln Gly Arg Ser 260
265 270 Val Arg Val Asp Val Ala Ala Gly
Ala Glu Ala Ile Met Lys Ala Val 275 280
285 Asp Gly Cys Gly Arg Leu Asp Asn Val Thr Gly Glu Ser
Gly Thr Asn 290 295 300
Ile Gly Gly Met Leu Glu His Val Arg Gln Thr Met Ala Glu Leu Thr 305
310 315 320 Asn Lys Pro Ser
Ser Glu Ile Phe Ile Gln Asp Leu Leu Ala Val Asp 325
330 335 Thr Ser Val Pro Val Ser Val Thr Gly
Gly Leu Ala Gly Glu Phe Ser 340 345
350 Leu Glu Gln Ala Val Gly Ile Ala Ser Met Val Lys Ser Asp
Arg Leu 355 360 365
Gln Met Ala Met Ile Ala Arg Glu Ile Glu Gln Lys Leu Asn Ile Asp 370
375 380 Val Gln Ile Gly Gly
Ala Glu Ala Glu Ala Ala Ile Leu Gly Ala Leu 385 390
395 400 Thr Thr Pro Gly Thr Thr Arg Pro Leu Ala
Ile Leu Asp Leu Gly Ala 405 410
415 Gly Ser Thr Asp Ala Ser Ile Ile Asn Pro Lys Gly Asp Ile Ile
Ala 420 425 430 Thr
His Leu Ala Gly Ala Gly Asp Met Val Thr Met Ile Ile Ala Arg 435
440 445 Glu Leu Gly Leu Glu Asp
Arg Tyr Leu Ala Glu Glu Ile Lys Lys Tyr 450 455
460 Pro Leu Ala Lys Val Glu Ser Leu Phe His Leu
Arg His Glu Asp Gly 465 470 475
480 Ser Val Gln Phe Phe Ser Thr Pro Leu Pro Pro Ala Val Phe Ala Arg
485 490 495 Val Cys
Val Val Lys Ala Asp Glu Leu Val Pro Leu Pro Gly Asp Leu 500
505 510 Ala Leu Glu Lys Val Arg Ala
Ile Arg Arg Ser Ala Lys Glu Arg Val 515 520
525 Phe Val Thr Asn Ala Leu Arg Ala Leu Arg Gln Val
Ser Pro Thr Gly 530 535 540
Asn Ile Arg Asp Ile Pro Phe Val Val Leu Val Gly Gly Ser Ser Leu 545
550 555 560 Asp Phe Glu
Val Pro Gln Leu Val Thr Asp Ala Leu Ala His Tyr Arg 565
570 575 Leu Val Ala Gly Arg Gly Asn Ile
Arg Gly Ser Glu Gly Pro Arg Asn 580 585
590 Ala Val Ala Thr Gly Leu Ile Leu Ser Trp His Lys Glu
Phe Ala His 595 600 605
Glu Arg 610 112378DNAKlebsiella oxytoca 112atgaacggta atcacagcgc
cccggccatc gcgatcgccg tcatcgacgg ctgcgacggc 60ctgtggcgcg aagtgctgct
gggtatcgaa gaggaaggta tccctttccg gctccagcat 120cacccggccg gagaggtcgt
ggacagcgcc tggcaggcgg cgcgcagctc gccgctgctg 180gtgggcatcg cctgcgaccg
ccatatgctg gtcgtgcact acaagaattt acccgcatcg 240gcgccgcttt ttacgctgat
gcatcatcag gacagtcagg cccatcgcaa caccggtaat 300aacgcggcac ggctggtcaa
ggggatccct ttccgggatc tgaatagcga agcaacagga 360gaacagcagg atgaataa
378113125PRTKlebsiella
oxytoca 113Met Asn Gly Asn His Ser Ala Pro Ala Ile Ala Ile Ala Val Ile
Asp 1 5 10 15 Gly
Cys Asp Gly Leu Trp Arg Glu Val Leu Leu Gly Ile Glu Glu Glu
20 25 30 Gly Ile Pro Phe Arg
Leu Gln His His Pro Ala Gly Glu Val Val Asp 35
40 45 Ser Ala Trp Gln Ala Ala Arg Ser Ser
Pro Leu Leu Val Gly Ile Ala 50 55
60 Cys Asp Arg His Met Leu Val Val His Tyr Lys Asn Leu
Pro Ala Ser 65 70 75
80 Ala Pro Leu Phe Thr Leu Met His His Gln Asp Ser Gln Ala His Arg
85 90 95 Asn Thr Gly Asn
Asn Ala Ala Arg Leu Val Lys Gly Ile Pro Phe Arg 100
105 110 Asp Leu Asn Ser Glu Ala Thr Gly Glu
Gln Gln Asp Glu 115 120 125
1141833DNASalmonella typhimurium 114atgcgatata tagctggcat tgacatcggt
aactcatcaa cggaagtcgc actggcgcgg 60caagatgaga ctggcgcact gacgattaca
cacagcgcgc tggcggaaac caccgggatc 120aaaggcacgt tgcgtaacgt gttcggcatt
caggaagcgc tcgccctcgt cgcaaagcgc 180gcggggatca atgtcagaga tatttcgctc
atccgcatta acgaagccac gccggtgatt 240ggcgatgtgg cgatggaaac cattaccgaa
accatcatca ccgaatcgac aatgatcggc 300cataacccaa aaacgccggg cggagcaggc
cttggtgtgg gtatcacgat tacgccggag 360gagctgttaa cccgcccggc ggactcgtcc
tatattctgg tggtatcgtc agcctttgat 420tttgctgata tcgccaatgt tatcaacgcc
tcaatgcgcg ccggatacca gattaccggc 480gtcattttgc agcgcgacga tggcgtactg
gtcagcaacc ggctggaaaa atcgctaccg 540attgtcgatg aagttctgta catcgaccgc
attccgctgg ggatgctggc ggcgattgaa 600gtcgccgtgc cgggaaaggt tatcgaaacc
ctctctaacc cttacggcat cgccaccgta 660tttaatctca acgccgatga gacaaaaaac
atcgtcccga tggcgcgcgc gctgattggc 720aaccgttccg ccgtggtggt taaaacgcca
tccggcgacg tcaaagcgcg cgcaataccc 780gccggtaacc tggagctgca ggctcagggt
cgtaccgtgc gcgtggatgt tgccgccggt 840gccgaagcca tcatgaaagc ggtggacggt
tgcggcaagc tcgacaacgt caccggcgag 900gccgggacca atatcggcgg catgctggag
cacgtgcgcc agaccatggc cgaactgacc 960aacaagccga gcagtgagat tttcattcag
gatctactgg ccgttgacac ctcggttccg 1020gtgagcgtca ccggcggtct ggccggggag
ttctcgctgg agcaggccgt cggcatcgcc 1080tcgatggtga aatcagaccg tctgcagatg
gcgatgattg cccgtgaaat tgagcagaag 1140cttaatatcg acgtgcagat cggcggcgct
gaggctgaag ccgccattct gggcgcgctg 1200accacgccgg gtaccacccg accgctggcg
atcctcgacc tcggcgcggg ctccaccgat 1260gcctccatca tcaaccctaa aggcgaaatc
atcgccaccc atctcgccgg ggcaggcgac 1320atggtcacga tgattattgc ccgcgaactg
gggctggaag accgctatct ggcggaagag 1380atcaaaaaat acccgctggc taaggtcgaa
agcctgttcc acttacgcca cgaggacggc 1440agcgtccagt tcttcccgac gccgctgcct
cccgccgtgt tcgcccgcgt ctgcgtggtg 1500aaaccggacg aactggtgcc gcttcccggc
gacttagcgc tggaaaaagt gcgcgccatt 1560cgccgcagcg ctaaagaacg cgtctttgtc
accaacgccc tgcgcgcgct gcgccaggtc 1620agtccaaccg gcaacattcg cgatattccg
ttcgtggtgc tggtcggcgg ctcgtcgctg 1680gatttcgaag ttccgcagct ggtcaccgat
gcgctggcgc actaccgcct ggtcgccggg 1740cgaggaaata ttcgcggcag cgaaggccca
agaaacgcgg tggccaccgg cctgattctc 1800tcctggcata aggagtttgc gcatggacag
taa 1833115610PRTSalmonella typhimurium
115Met Arg Tyr Ile Ala Gly Ile Asp Ile Gly Asn Ser Ser Thr Glu Val 1
5 10 15 Ala Leu Ala Arg
Gln Asp Glu Thr Gly Ala Leu Thr Ile Thr His Ser 20
25 30 Ala Leu Ala Glu Thr Thr Gly Ile Lys
Gly Thr Leu Arg Asn Val Phe 35 40
45 Gly Ile Gln Glu Ala Leu Ala Leu Val Ala Lys Arg Ala Gly
Ile Asn 50 55 60
Val Arg Asp Ile Ser Leu Ile Arg Ile Asn Glu Ala Thr Pro Val Ile 65
70 75 80 Gly Asp Val Ala Met
Glu Thr Ile Thr Glu Thr Ile Ile Thr Glu Ser 85
90 95 Thr Met Ile Gly His Asn Pro Lys Thr Pro
Gly Gly Ala Gly Leu Gly 100 105
110 Val Gly Ile Thr Ile Thr Pro Glu Glu Leu Leu Thr Arg Pro Ala
Asp 115 120 125 Ser
Ser Tyr Ile Leu Val Val Ser Ser Ala Phe Asp Phe Ala Asp Ile 130
135 140 Ala Asn Val Ile Asn Ala
Ser Met Arg Ala Gly Tyr Gln Ile Thr Gly 145 150
155 160 Val Ile Leu Gln Arg Asp Asp Gly Val Leu Val
Ser Asn Arg Leu Glu 165 170
175 Lys Ser Leu Pro Ile Val Asp Glu Val Leu Tyr Ile Asp Arg Ile Pro
180 185 190 Leu Gly
Met Leu Ala Ala Ile Glu Val Ala Val Pro Gly Lys Val Ile 195
200 205 Glu Thr Leu Ser Asn Pro Tyr
Gly Ile Ala Thr Val Phe Asn Leu Asn 210 215
220 Ala Asp Glu Thr Lys Asn Ile Val Pro Met Ala Arg
Ala Leu Ile Gly 225 230 235
240 Asn Arg Ser Ala Val Val Val Lys Thr Pro Ser Gly Asp Val Lys Ala
245 250 255 Arg Ala Ile
Pro Ala Gly Asn Leu Glu Leu Gln Ala Gln Gly Arg Thr 260
265 270 Val Arg Val Asp Val Ala Ala Gly
Ala Glu Ala Ile Met Lys Ala Val 275 280
285 Asp Gly Cys Gly Lys Leu Asp Asn Val Thr Gly Glu Ala
Gly Thr Asn 290 295 300
Ile Gly Gly Met Leu Glu His Val Arg Gln Thr Met Ala Glu Leu Thr 305
310 315 320 Asn Lys Pro Ser
Ser Glu Ile Phe Ile Gln Asp Leu Leu Ala Val Asp 325
330 335 Thr Ser Val Pro Val Ser Val Thr Gly
Gly Leu Ala Gly Glu Phe Ser 340 345
350 Leu Glu Gln Ala Val Gly Ile Ala Ser Met Val Lys Ser Asp
Arg Leu 355 360 365
Gln Met Ala Met Ile Ala Arg Glu Ile Glu Gln Lys Leu Asn Ile Asp 370
375 380 Val Gln Ile Gly Gly
Ala Glu Ala Glu Ala Ala Ile Leu Gly Ala Leu 385 390
395 400 Thr Thr Pro Gly Thr Thr Arg Pro Leu Ala
Ile Leu Asp Leu Gly Ala 405 410
415 Gly Ser Thr Asp Ala Ser Ile Ile Asn Pro Lys Gly Glu Ile Ile
Ala 420 425 430 Thr
His Leu Ala Gly Ala Gly Asp Met Val Thr Met Ile Ile Ala Arg 435
440 445 Glu Leu Gly Leu Glu Asp
Arg Tyr Leu Ala Glu Glu Ile Lys Lys Tyr 450 455
460 Pro Leu Ala Lys Val Glu Ser Leu Phe His Leu
Arg His Glu Asp Gly 465 470 475
480 Ser Val Gln Phe Phe Pro Thr Pro Leu Pro Pro Ala Val Phe Ala Arg
485 490 495 Val Cys
Val Val Lys Pro Asp Glu Leu Val Pro Leu Pro Gly Asp Leu 500
505 510 Ala Leu Glu Lys Val Arg Ala
Ile Arg Arg Ser Ala Lys Glu Arg Val 515 520
525 Phe Val Thr Asn Ala Leu Arg Ala Leu Arg Gln Val
Ser Pro Thr Gly 530 535 540
Asn Ile Arg Asp Ile Pro Phe Val Val Leu Val Gly Gly Ser Ser Leu 545
550 555 560 Asp Phe Glu
Val Pro Gln Leu Val Thr Asp Ala Leu Ala His Tyr Arg 565
570 575 Leu Val Ala Gly Arg Gly Asn Ile
Arg Gly Ser Glu Gly Pro Arg Asn 580 585
590 Ala Val Ala Thr Gly Leu Ile Leu Ser Trp His Lys Glu
Phe Ala His 595 600 605
Gly Gln 610 116372DNASalmonella typhimurium 116atggacagta
atcacagcgc cccggctatc gtcattaccg ttatcaacga ctgcgccagc 60ctctggcacg
aagtgctgct gggcattgaa gaggaaggca tccctttcct gcttcagcat 120cacccggctg
gagatatcgt tgacagcgcc tggcaggcgg cgcgcagctc gccgctgctg 180gtcggcattg
cctgcgatcg acactcgctg gtcgtgcatt acaagaattt acccgcatcg 240gcgccgcttt
ttacgctgat gcatcatcag gacagtcagg cccaacgcaa caccggtaat 300aacgcggcac
ggctggtcaa agggatccct ttcgggatct ccatgcttaa tcacaggaga 360acggcagtat
ga
372117123PRTSalmonella typhimurium 117Met Asp Ser Asn His Ser Ala Pro Ala
Ile Val Ile Thr Val Ile Asn 1 5 10
15 Asp Cys Ala Ser Leu Trp His Glu Val Leu Leu Gly Ile Glu
Glu Glu 20 25 30
Gly Ile Pro Phe Leu Leu Gln His His Pro Ala Gly Asp Ile Val Asp
35 40 45 Ser Ala Trp Gln
Ala Ala Arg Ser Ser Pro Leu Leu Val Gly Ile Ala 50
55 60 Cys Asp Arg His Ser Leu Val Val
His Tyr Lys Asn Leu Pro Ala Ser 65 70
75 80 Ala Pro Leu Phe Thr Leu Met His His Gln Asp Ser
Gln Ala Gln Arg 85 90
95 Asn Thr Gly Asn Asn Ala Ala Arg Leu Val Lys Gly Ile Pro Phe Gly
100 105 110 Ile Ser Met
Leu Asn His Arg Arg Thr Ala Val 115 120
1181833DNALactobacillus collinoides 118atgacacgtg taattggtgt tgatatcggg
aattcctcta cagaagttgc gcttgctgat 60gtgtctgaca gtggtgaagt aaatttcatt
aattctggaa tttccgatac aactggcatt 120aaaggtacta aacaaaattt gatcggggtg
cgtaaatcca tccagatcgt tttgaaaaag 180tcgaatatgc aaatttccga tgttgacctg
attcggatca acgaagcaac gcccgttatc 240ggtgatgttg ccatggagac catcaccgaa
acggtgatta ctgaatcgac gatgatcggc 300cacaacccag ggactcctgg gggtgtcggt
actggttctg gttacacggt gaatttgctt 360gatttgttga gccaaacgga taaggatcgt
ccttatatcg ttatcatctc gaaagaaatc 420gattttgctg acgcagctaa gctgatcaac
gcttatgtgg cttctggtta taatattacc 480gctgccattc tgcaaagtga tgatggggtg
ctgatcaata atcggttgac ccataagatt 540cccatcgtgg atgaagtctc acagatcgac
aaggtaccgt tgaacatgct tgccgcagtg 600gaagttgcac cgcctggcaa agtaattgct
caactttcca acccgtatgg cattgccaca 660ctgttcgaac tttcctctga agaaaccaag
aacattgtgc cagttgcccg agccttaatc 720ggaaaccggt cagcggttgt tattaaaacc
cctgccggtg atgttaaagc tcgtgttatc 780ccagccggga aaatcttgat caatggccaa
ccgaatggtc atggtgaagt taacgttgcg 840gctggtgccg atgccatcat gaaaaaggtg
aacgagttcg atagtgtcga tgacattacc 900ggtgaatcgg gcactaacgt tggtgggatg
cttgaaaaag ttcgtcaaac aatggctgag 960ttgaccgaca agcaaaatag cgacattgcc
attcaagatt tattagctgt caatacgtcc 1020gttccagtaa cggtgcgtgg tggtctggct
ggtgaattct caatggaaca agccgttggg 1080attgctgcta tggtcaaatc tgatcacttg
caaatgcaag cgattgcaga cctgatgaaa 1140gatgaatttc acgttcaagt cgaaatcggc
ggtgctgaag ctgaatcagc catcctcggt 1200gcgctaacaa cgccagggac gacaaaacca
attgccatcc ttgatttggg ggctggttca 1260acggatgcat caattatcaa ccaaaaggac
gaaaaggtcg ctattcactt ggctggtgcc 1320ggtgatatgg ttaccatgat catcaattct
gaacttgggt tggaagaccc atatttagct 1380gaggatatta agaaatatcc gctggctaaa
gttgataatc tattccagct acggcatgaa 1440gatggtgccg ttcaattctt tgaagatcca
ttacctgctg atttatttgc cagagttgtg 1500gctgttaaac cagatggtta cgaaccactt
cctggtaatt tgagtatcga gaaagttaaa 1560atcgtccgtc aaactgctaa gaagcgggtg
ttcgtaacga acgcaattcg tgccttacac 1620cacgttagcc caacaggtaa tatccgagat
atcccatttg tggtcattgt cggcggctca 1680gccctcgatt ttgaaattcc acaattggtc
accgatgaat tatcacactt taacttagtt 1740gcaggtcgtg gtaatattcg gggaattgaa
ggtccacgga acgccgtggc aactggtttg 1800attctttcat acgcgagtga gaagagggga
tag 1833119610PRTLactobacillus collinoides
119Met Thr Arg Val Ile Gly Val Asp Ile Gly Asn Ser Ser Thr Glu Val 1
5 10 15 Ala Leu Ala Asp
Val Ser Asp Ser Gly Glu Val Asn Phe Ile Asn Ser 20
25 30 Gly Ile Ser Asp Thr Thr Gly Ile Lys
Gly Thr Lys Gln Asn Leu Ile 35 40
45 Gly Val Arg Lys Ser Ile Gln Ile Val Leu Lys Lys Ser Asn
Met Gln 50 55 60
Ile Ser Asp Val Asp Leu Ile Arg Ile Asn Glu Ala Thr Pro Val Ile 65
70 75 80 Gly Asp Val Ala Met
Glu Thr Ile Thr Glu Thr Val Ile Thr Glu Ser 85
90 95 Thr Met Ile Gly His Asn Pro Gly Thr Pro
Gly Gly Val Gly Thr Gly 100 105
110 Ser Gly Tyr Thr Val Asn Leu Leu Asp Leu Leu Ser Gln Thr Asp
Lys 115 120 125 Asp
Arg Pro Tyr Ile Val Ile Ile Ser Lys Glu Ile Asp Phe Ala Asp 130
135 140 Ala Ala Lys Leu Ile Asn
Ala Tyr Val Ala Ser Gly Tyr Asn Ile Thr 145 150
155 160 Ala Ala Ile Leu Gln Ser Asp Asp Gly Val Leu
Ile Asn Asn Arg Leu 165 170
175 Thr His Lys Ile Pro Ile Val Asp Glu Val Ser Gln Ile Asp Lys Val
180 185 190 Pro Leu
Asn Met Leu Ala Ala Val Glu Val Ala Pro Pro Gly Lys Val 195
200 205 Ile Ala Gln Leu Ser Asn Pro
Tyr Gly Ile Ala Thr Leu Phe Glu Leu 210 215
220 Ser Ser Glu Glu Thr Lys Asn Ile Val Pro Val Ala
Arg Ala Leu Ile 225 230 235
240 Gly Asn Arg Ser Ala Val Val Ile Lys Thr Pro Ala Gly Asp Val Lys
245 250 255 Ala Arg Val
Ile Pro Ala Gly Lys Ile Leu Ile Asn Gly Gln Pro Asn 260
265 270 Gly His Gly Glu Val Asn Val Ala
Ala Gly Ala Asp Ala Ile Met Lys 275 280
285 Lys Val Asn Glu Phe Asp Ser Val Asp Asp Ile Thr Gly
Glu Ser Gly 290 295 300
Thr Asn Val Gly Gly Met Leu Glu Lys Val Arg Gln Thr Met Ala Glu 305
310 315 320 Leu Thr Asp Lys
Gln Asn Ser Asp Ile Ala Ile Gln Asp Leu Leu Ala 325
330 335 Val Asn Thr Ser Val Pro Val Thr Val
Arg Gly Gly Leu Ala Gly Glu 340 345
350 Phe Ser Met Glu Gln Ala Val Gly Ile Ala Ala Met Val Lys
Ser Asp 355 360 365
His Leu Gln Met Gln Ala Ile Ala Asp Leu Met Lys Asp Glu Phe His 370
375 380 Val Gln Val Glu Ile
Gly Gly Ala Glu Ala Glu Ser Ala Ile Leu Gly 385 390
395 400 Ala Leu Thr Thr Pro Gly Thr Thr Lys Pro
Ile Ala Ile Leu Asp Leu 405 410
415 Gly Ala Gly Ser Thr Asp Ala Ser Ile Ile Asn Gln Lys Asp Glu
Lys 420 425 430 Val
Ala Ile His Leu Ala Gly Ala Gly Asp Met Val Thr Met Ile Ile 435
440 445 Asn Ser Glu Leu Gly Leu
Glu Asp Pro Tyr Leu Ala Glu Asp Ile Lys 450 455
460 Lys Tyr Pro Leu Ala Lys Val Asp Asn Leu Phe
Gln Leu Arg His Glu 465 470 475
480 Asp Gly Ala Val Gln Phe Phe Glu Asp Pro Leu Pro Ala Asp Leu Phe
485 490 495 Ala Arg
Val Val Ala Val Lys Pro Asp Gly Tyr Glu Pro Leu Pro Gly 500
505 510 Asn Leu Ser Ile Glu Lys Val
Lys Ile Val Arg Gln Thr Ala Lys Lys 515 520
525 Arg Val Phe Val Thr Asn Ala Ile Arg Ala Leu His
His Val Ser Pro 530 535 540
Thr Gly Asn Ile Arg Asp Ile Pro Phe Val Val Ile Val Gly Gly Ser 545
550 555 560 Ala Leu Asp
Phe Glu Ile Pro Gln Leu Val Thr Asp Glu Leu Ser His 565
570 575 Phe Asn Leu Val Ala Gly Arg Gly
Asn Ile Arg Gly Ile Glu Gly Pro 580 585
590 Arg Asn Ala Val Ala Thr Gly Leu Ile Leu Ser Tyr Ala
Ser Glu Lys 595 600 605
Arg Gly 610 120351DNALactobacillus collinoides 120atggcatttg
attctgaacg tccgtcaatt ctattggcga caccaacggg ttctaatggc 60caacttccag
aagttctaaa accaatgctc aatggtattg aagaagaaca gattcctttt 120cagattctcg
atatggaagg cggttcagca gttgagcggg cttataacgc gtcagttgct 180tcacgattat
cagtgggcgt tgggtttgat gatgcacata tcattgtgca ttataaaaac 240ttgaaaccag
aaaaaccgct gtttgatgtt gccatcactg atgcagcatc cattcgtaaa 300gttggcgcaa
acgccgctcg acttgtaaag ggagttccat tcaagaagta a
351121116PRTLactobacillus collinoides 121Met Ala Phe Asp Ser Glu Arg Pro
Ser Ile Leu Leu Ala Thr Pro Thr 1 5 10
15 Gly Ser Asn Gly Gln Leu Pro Glu Val Leu Lys Pro Met
Leu Asn Gly 20 25 30
Ile Glu Glu Glu Gln Ile Pro Phe Gln Ile Leu Asp Met Glu Gly Gly
35 40 45 Ser Ala Val Glu
Arg Ala Tyr Asn Ala Ser Val Ala Ser Arg Leu Ser 50
55 60 Val Gly Val Gly Phe Asp Asp Ala
His Ile Ile Val His Tyr Lys Asn 65 70
75 80 Leu Lys Pro Glu Lys Pro Leu Phe Asp Val Ala Ile
Thr Asp Ala Ala 85 90
95 Ser Ile Arg Lys Val Gly Ala Asn Ala Ala Arg Leu Val Lys Gly Val
100 105 110 Pro Phe Lys
Lys 115 122453PRTVibrio fluvialis 122Met Asn Lys Pro Gln Ser
Trp Glu Ala Arg Ala Glu Thr Tyr Ser Leu 1 5
10 15 Tyr Gly Phe Thr Asp Met Pro Ser Leu His Gln
Arg Gly Thr Val Val 20 25
30 Val Thr His Gly Glu Gly Pro Tyr Ile Val Asp Val Asn Gly Arg
Arg 35 40 45 Tyr
Leu Asp Ala Asn Ser Gly Leu Trp Asn Met Val Ala Gly Phe Asp 50
55 60 His Lys Gly Leu Ile Asp
Ala Ala Lys Ala Gln Tyr Glu Arg Phe Pro 65 70
75 80 Gly Tyr His Ala Phe Phe Gly Arg Met Ser Asp
Gln Thr Val Met Leu 85 90
95 Ser Glu Lys Leu Val Glu Val Ser Pro Phe Asp Ser Gly Arg Val Phe
100 105 110 Tyr Thr
Asn Ser Gly Ser Glu Ala Asn Asp Thr Met Val Lys Met Leu 115
120 125 Trp Phe Leu His Ala Ala Glu
Gly Lys Pro Gln Lys Arg Lys Ile Leu 130 135
140 Thr Arg Trp Asn Ala Tyr His Gly Val Thr Ala Val
Ser Ala Ser Met 145 150 155
160 Thr Gly Lys Pro Tyr Asn Ser Val Phe Gly Leu Pro Leu Pro Gly Phe
165 170 175 Val His Leu
Thr Cys Pro His Tyr Trp Arg Tyr Gly Glu Glu Gly Glu 180
185 190 Thr Glu Glu Gln Phe Val Ala Arg
Leu Ala Arg Glu Leu Glu Glu Thr 195 200
205 Ile Gln Arg Glu Gly Ala Asp Thr Ile Ala Gly Phe Phe
Ala Glu Pro 210 215 220
Val Met Gly Ala Gly Gly Val Ile Pro Pro Ala Lys Gly Tyr Phe Gln 225
230 235 240 Ala Ile Leu Pro
Ile Leu Arg Lys Tyr Asp Ile Pro Val Ile Ser Asp 245
250 255 Glu Val Ile Cys Gly Phe Gly Arg Thr
Gly Asn Thr Trp Gly Cys Val 260 265
270 Thr Tyr Asp Phe Thr Pro Asp Ala Ile Ile Ser Ser Lys Asn
Leu Thr 275 280 285
Ala Gly Phe Phe Pro Met Gly Ala Val Ile Leu Gly Pro Glu Leu Ser 290
295 300 Lys Arg Leu Glu Thr
Ala Ile Glu Ala Ile Glu Glu Phe Pro His Gly 305 310
315 320 Phe Thr Ala Ser Gly His Pro Val Gly Cys
Ala Ile Ala Leu Lys Ala 325 330
335 Ile Asp Val Val Met Asn Glu Gly Leu Ala Glu Asn Val Arg Arg
Leu 340 345 350 Ala
Pro Arg Phe Glu Glu Arg Leu Lys His Ile Ala Glu Arg Pro Asn 355
360 365 Ile Gly Glu Tyr Arg Gly
Ile Gly Phe Met Trp Ala Leu Glu Ala Val 370 375
380 Lys Asp Lys Ala Ser Lys Thr Pro Phe Asp Gly
Asn Leu Ser Val Ser 385 390 395
400 Glu Arg Ile Ala Asn Thr Cys Thr Asp Leu Gly Leu Ile Cys Arg Pro
405 410 415 Leu Gly
Gln Ser Val Val Leu Cys Pro Pro Phe Ile Leu Thr Glu Ala 420
425 430 Gln Met Asp Glu Met Phe Asp
Lys Leu Glu Lys Ala Leu Asp Lys Val 435 440
445 Phe Ala Glu Val Ala 450
1231122DNAErwinia caratovora subsp. atroseptica 123atgtctgacg gacgactcac
cgcacttttt cctgcattcc cacacccggc gtccaatcag 60cccgtatttg ccgaggcttc
accgcacgac gacgagttaa tgacgcaggc cgtaccgcag 120gtttcctgtc agcaggcgtt
ggcgattgcg cagcaagaat atggcttgtc tgggcagatg 180tcgctgcttc agggcgagcg
tgatgtgaat ttctgtctga cggtgacgcc agatgaacgc 240tacatgctga aagtcatcaa
tgcggcagaa cctgccgacg tcagcaattt ccaaaccgcg 300ctgctgctgc atcttgcccg
tcaggcacct gaactgcccg taccgcgtat caggtcgaca 360aaagcgggtc agtcggaaac
aggcgttgag atcgatggtg tactgctgcg tgtgcggctt 420gtgagctatc tggcaggaat
gccgcagtat ctggcctcac cgtcaacggc gctgatgccg 480cagttggggg gaacgctggc
gcagttggat aacgcgcttc acagctttac gcatccggcg 540gcaaaccgtg cgctgctgtg
ggatatcagc cgggcagagc aggtgcgtcc ttacctcgat 600ttcgtttctg aaccgcagca
gtatcagcat cttcagcgta tttttgaccg ttatgacagt 660aacgttgctc ctctgttgac
gacgctacgt cgtcaggtca ttcataacga tctgaatccg 720cataacgtgc tggtggatgg
atcgtcgccg acgcgggtta ctggcattat cgattttggc 780gatgccgtat ttgccccgtt
aatttgcgaa gtcgcgacgg cactggcgta tcagatcggc 840gatggaaccg atttgttgga
gcatgttgtg ccgtttgttg cggcctatca ccaacgcatt 900ccgttagcac cggaggagat
tgcgctgtta cccgatctga tagcgacccg tatggcgctg 960accctgacca ttgcgcagtg
gcgagcatcg cgttatcccg acaatcggga gtatctgctg 1020cgtaacgtgc cgcgctgttg
gcacagtttg cagcgcattg cgacctattc ccatgcgcaa 1080tttttgactc gcctacagca
ggtttgcccg gagaatgcgc ga 1122124374PRTErwinia
caratovora subsp. atroseptica 124Met Ser Asp Gly Arg Leu Thr Ala Leu Phe
Pro Ala Phe Pro His Pro 1 5 10
15 Ala Ser Asn Gln Pro Val Phe Ala Glu Ala Ser Pro His Asp Asp
Glu 20 25 30 Leu
Met Thr Gln Ala Val Pro Gln Val Ser Cys Gln Gln Ala Leu Ala 35
40 45 Ile Ala Gln Gln Glu Tyr
Gly Leu Ser Gly Gln Met Ser Leu Leu Gln 50 55
60 Gly Glu Arg Asp Val Asn Phe Cys Leu Thr Val
Thr Pro Asp Glu Arg 65 70 75
80 Tyr Met Leu Lys Val Ile Asn Ala Ala Glu Pro Ala Asp Val Ser Asn
85 90 95 Phe Gln
Thr Ala Leu Leu Leu His Leu Ala Arg Gln Ala Pro Glu Leu 100
105 110 Pro Val Pro Arg Ile Arg Ser
Thr Lys Ala Gly Gln Ser Glu Thr Gly 115 120
125 Val Glu Ile Asp Gly Val Leu Leu Arg Val Arg Leu
Val Ser Tyr Leu 130 135 140
Ala Gly Met Pro Gln Tyr Leu Ala Ser Pro Ser Thr Ala Leu Met Pro 145
150 155 160 Gln Leu Gly
Gly Thr Leu Ala Gln Leu Asp Asn Ala Leu His Ser Phe 165
170 175 Thr His Pro Ala Ala Asn Arg Ala
Leu Leu Trp Asp Ile Ser Arg Ala 180 185
190 Glu Gln Val Arg Pro Tyr Leu Asp Phe Val Ser Glu Pro
Gln Gln Tyr 195 200 205
Gln His Leu Gln Arg Ile Phe Asp Arg Tyr Asp Ser Asn Val Ala Pro 210
215 220 Leu Leu Thr Thr
Leu Arg Arg Gln Val Ile His Asn Asp Leu Asn Pro 225 230
235 240 His Asn Val Leu Val Asp Gly Ser Ser
Pro Thr Arg Val Thr Gly Ile 245 250
255 Ile Asp Phe Gly Asp Ala Val Phe Ala Pro Leu Ile Cys Glu
Val Ala 260 265 270
Thr Ala Leu Ala Tyr Gln Ile Gly Asp Gly Thr Asp Leu Leu Glu His
275 280 285 Val Val Pro Phe
Val Ala Ala Tyr His Gln Arg Ile Pro Leu Ala Pro 290
295 300 Glu Glu Ile Ala Leu Leu Pro Asp
Leu Ile Ala Thr Arg Met Ala Leu 305 310
315 320 Thr Leu Thr Ile Ala Gln Trp Arg Ala Ser Arg Tyr
Pro Asp Asn Arg 325 330
335 Glu Tyr Leu Leu Arg Asn Val Pro Arg Cys Trp His Ser Leu Gln Arg
340 345 350 Ile Ala Thr
Tyr Ser His Ala Gln Phe Leu Thr Arg Leu Gln Gln Val 355
360 365 Cys Pro Glu Asn Ala Arg 370
1251272DNAErwinia caratovora subsp. atroseptica
125atgacagcga cagaagcttt gctggcgcgc cgtcagcgag tgttgggcgg cggttatcgc
60ctgttttatg aagagccgct gcatgtcgcg cgcggcgagg gcgtgtggct gttcgatcac
120caagggaaac gttatctgga tgtctacaat aatgtggctt cggtcggaca ttgccacccc
180gcggtggttg aagccgtggc gcgacagagc gcacaactca atacccacac gcgctatttg
240caccacgcga ttgtcgattt tgcggaagat ttgctgagcg aatttcccgc cgaattgaac
300aatgtaatgc tgacctgtac cggcagtgag gctaacgatc tggcgctgcg tatcgcccga
360catgtcacgg gcgggacggg gatgttggtg acgcgctggg cgtatcacgg cgtgaccagc
420gcgctggcgg aactgtctcc gtcgctgggg gatggcgttg tgcgcggtag ccatgtgaag
480ctgatcgacg cgccagacac ttatcgtcag cccggtgcat ttcttaccag cattcgtgaa
540gcgctggcgc agatgcaacg ggaaggtatt cgtcctgcgg cgctgctggt agataccatt
600ttttccagcg atggcgtgtt ctgtgcgccg gaaggcgaaa tggcacaggc ggcggcgttg
660atccgtcagg cgggcgggct gtttattgcg gatgaagtgc agccgggctt cgggcgcacc
720ggggaatcac tgtggggctt tgcgcgccac aatgtcgtcc ctgatttggt gagtctaggg
780aaaccgatgg gcaacggaca tcccatcgct ggattggtgg ggcgttccgc tctgttcgac
840gcatttgggc gcgatgtgcg ctatttcaat acctttggcg gcaatccggt ttcctgtcag
900gcggcgcacg cggtgctgcg ggtgattcgg gaagagcagt tgcagcagaa tgcccagcgg
960gtcggtgatt atctgcggca agggttgcag caactggcgc agcatttccc gctgattggt
1020gatattcggg cttacggcct gtttattggt gcggagctgg tcagcgatcg cgaaagtaaa
1080acgccggcaa gtgaatccgc gttgcaggtg gtgaatgcga tgcgccaacg tggtgtgctc
1140atcagcgcga cggggccagc ggcgaacata ctgaaaattc gcccgccgct ggtgtttctg
1200gaagaacacg ccgatgtgtt cttaaccacg ctgagtgacg ttttagcgct catcggcact
1260cgtgcacaga ga
1272126424PRTErwinia caratovora subsp. atroseptica 126Met Thr Ala Thr Glu
Ala Leu Leu Ala Arg Arg Gln Arg Val Leu Gly 1 5
10 15 Gly Gly Tyr Arg Leu Phe Tyr Glu Glu Pro
Leu His Val Ala Arg Gly 20 25
30 Glu Gly Val Trp Leu Phe Asp His Gln Gly Lys Arg Tyr Leu Asp
Val 35 40 45 Tyr
Asn Asn Val Ala Ser Val Gly His Cys His Pro Ala Val Val Glu 50
55 60 Ala Val Ala Arg Gln Ser
Ala Gln Leu Asn Thr His Thr Arg Tyr Leu 65 70
75 80 His His Ala Ile Val Asp Phe Ala Glu Asp Leu
Leu Ser Glu Phe Pro 85 90
95 Ala Glu Leu Asn Asn Val Met Leu Thr Cys Thr Gly Ser Glu Ala Asn
100 105 110 Asp Leu
Ala Leu Arg Ile Ala Arg His Val Thr Gly Gly Thr Gly Met 115
120 125 Leu Val Thr Arg Trp Ala Tyr
His Gly Val Thr Ser Ala Leu Ala Glu 130 135
140 Leu Ser Pro Ser Leu Gly Asp Gly Val Val Arg Gly
Ser His Val Lys 145 150 155
160 Leu Ile Asp Ala Pro Asp Thr Tyr Arg Gln Pro Gly Ala Phe Leu Thr
165 170 175 Ser Ile Arg
Glu Ala Leu Ala Gln Met Gln Arg Glu Gly Ile Arg Pro 180
185 190 Ala Ala Leu Leu Val Asp Thr Ile
Phe Ser Ser Asp Gly Val Phe Cys 195 200
205 Ala Pro Glu Gly Glu Met Ala Gln Ala Ala Ala Leu Ile
Arg Gln Ala 210 215 220
Gly Gly Leu Phe Ile Ala Asp Glu Val Gln Pro Gly Phe Gly Arg Thr 225
230 235 240 Gly Glu Ser Leu
Trp Gly Phe Ala Arg His Asn Val Val Pro Asp Leu 245
250 255 Val Ser Leu Gly Lys Pro Met Gly Asn
Gly His Pro Ile Ala Gly Leu 260 265
270 Val Gly Arg Ser Ala Leu Phe Asp Ala Phe Gly Arg Asp Val
Arg Tyr 275 280 285
Phe Asn Thr Phe Gly Gly Asn Pro Val Ser Cys Gln Ala Ala His Ala 290
295 300 Val Leu Arg Val Ile
Arg Glu Glu Gln Leu Gln Gln Asn Ala Gln Arg 305 310
315 320 Val Gly Asp Tyr Leu Arg Gln Gly Leu Gln
Gln Leu Ala Gln His Phe 325 330
335 Pro Leu Ile Gly Asp Ile Arg Ala Tyr Gly Leu Phe Ile Gly Ala
Glu 340 345 350 Leu
Val Ser Asp Arg Glu Ser Lys Thr Pro Ala Ser Glu Ser Ala Leu 355
360 365 Gln Val Val Asn Ala Met
Arg Gln Arg Gly Val Leu Ile Ser Ala Thr 370 375
380 Gly Pro Ala Ala Asn Ile Leu Lys Ile Arg Pro
Pro Leu Val Phe Leu 385 390 395
400 Glu Glu His Ala Asp Val Phe Leu Thr Thr Leu Ser Asp Val Leu Ala
405 410 415 Leu Ile
Gly Thr Arg Ala Gln Arg 420
12735DNAArtificial SeqquencePrimer 127ctccggaatt catgtctgac ggacgactca
ccgca 3512846DNAArtificial SeqquencePrimer
128ttccaatgca ttggctgcag ttatctctgt gcacgagtgc cgatga
4612940DNAArtificial SeqquencePrimer 129aacagccaag cttggctgca gtcatcgcgc
attctccggg 4013040DNAArtificial SeqquencePrimer
130tctccggaat tcatgacgtc tgaaatgaca gcgacagaag
4013133DNAArtificial SequencePrimer 131gctaacagga ggaagaattc atggggggtt
ctc 3313233DNAArtificial SequencePrimer
132gagaaccccc catgaattct tcctcctgtt agc
33133723DNAKlebsiella terrigena 133atgcaaaaag tcgcacttgt caccggcgcc
ggtcagggca tcggtaaagc tatcgccctg 60cgtctggtga aggatggatt tgccgtggca
atcgccgatt acaacgacgc tacggccaca 120gcggtagccg ctgaaatcaa ccaggccggc
ggccgcgcgg tggccattaa ggtcgacgtc 180tcgcgccggg accaggtttt cgccgccgtt
gagcaggcgc gtaaagccct gggcggattc 240aacgttatcg tcaacaacgc cggcatcgcg
ccgtcaacgc cgatcgagtc catcaccgag 300gagatcgtcg accgggtcta taacatcaac
gttaagggcg tcatctgggg gatgcaggcg 360gcggtggagg ccttcaaaaa agaggggcac
ggcgggaaga tcgtcaacgc ctgctcccag 420gccggccacg tcggcaaccc ggagctggcg
gtctacagtt cgagtaaatt cgccgtgcgc 480ggcctgacgc aaaccgccgc ccgcgatctg
gcgccgctgg gcatcaccgt taacggcttc 540tgcccaggga tcgttaagac gccaatgtgg
gcggagattg accgtcagtg tcggaagcgg 600cgggcaaacc gctgggctac ggcacggctg
aatttgccaa acgcatcacc cttggccgcc 660tgtcggagcc tgaagacgtc gccgcctgcg
tgtcgttcct cgccagcccg gattccgact 720ata
723134241PRTKlebsiella terrigena 134Met
Gln Lys Val Ala Leu Val Thr Gly Ala Gly Gln Gly Ile Gly Lys 1
5 10 15 Ala Ile Ala Leu Arg Leu
Val Lys Asp Gly Phe Ala Val Ala Ile Ala 20
25 30 Asp Tyr Asn Asp Ala Thr Ala Thr Ala Val
Ala Ala Glu Ile Asn Gln 35 40
45 Ala Gly Gly Arg Ala Val Ala Ile Lys Val Asp Val Ser Arg
Arg Asp 50 55 60
Gln Val Phe Ala Ala Val Glu Gln Ala Arg Lys Ala Leu Gly Gly Phe 65
70 75 80 Asn Val Ile Val Asn
Asn Ala Gly Ile Ala Pro Ser Thr Pro Ile Glu 85
90 95 Ser Ile Thr Glu Glu Ile Val Asp Arg Val
Tyr Asn Ile Asn Val Lys 100 105
110 Gly Val Ile Trp Gly Met Gln Ala Ala Val Glu Ala Phe Lys Lys
Glu 115 120 125 Gly
His Gly Gly Lys Ile Val Asn Ala Cys Ser Gln Ala Gly His Val 130
135 140 Gly Asn Pro Glu Leu Ala
Val Tyr Ser Ser Ser Lys Phe Ala Val Arg 145 150
155 160 Gly Leu Thr Gln Thr Ala Ala Arg Asp Leu Ala
Pro Leu Gly Ile Thr 165 170
175 Val Asn Gly Phe Cys Pro Gly Ile Val Lys Thr Pro Met Trp Ala Glu
180 185 190 Ile Asp
Arg Gln Cys Arg Lys Arg Arg Ala Asn Arg Trp Ala Thr Ala 195
200 205 Arg Leu Asn Leu Pro Asn Ala
Ser Pro Leu Ala Ala Cys Arg Ser Leu 210 215
220 Lys Thr Ser Pro Pro Ala Cys Arg Ser Ser Pro Ala
Arg Ile Pro Thr 225 230 235
240 Ile 135554PRTClostridium pasteurianum 135Met Lys Ser Lys Arg Phe
Gln Val Leu Ser Glu Arg Pro Val Asn Lys 1 5
10 15 Asp Gly Phe Ile Gly Glu Trp Pro Glu Glu Gly
Leu Ile Ala Met Ser 20 25
30 Ser Pro Asn Asp Pro Lys Pro Ser Ile Lys Ile Lys Glu Gly Lys
Val 35 40 45 Ile
Glu Leu Asp Gly Lys Asn Arg Glu Asp Phe Asp Met Ile Asp Arg 50
55 60 Phe Ile Ala Asn Tyr Gly
Ile Asn Leu Asn Arg Ala Glu Asp Val Ile 65 70
75 80 Lys Met Asp Ser Val Lys Leu Ala Lys Met Leu
Val Asp Ile Asn Val 85 90
95 Asp Arg Lys Thr Ile Val Glu Leu Thr Thr Ala Met Thr Pro Ala Lys
100 105 110 Ile Val
Glu Val Val Gly Asn Met Asn Val Val Glu Met Met Met Ala 115
120 125 Leu Gln Lys Met Arg Ala Arg
Lys Thr Pro Ser Asn Gln Cys His Val 130 135
140 Thr Asn Leu Lys Asp Asn Pro Val Gln Ile Ala Ala
Asp Ala Ala Glu 145 150 155
160 Ala Ala Ile Arg Gly Phe Asp Glu Gln Glu Thr Thr Val Gly Ile Val
165 170 175 Arg Tyr Ala
Pro Phe Asn Ala Leu Ala Leu Leu Val Gly Ala Gln Val 180
185 190 Gly Arg Gly Gly Val Leu Thr Gln
Cys Ala Ile Glu Glu Ala Thr Glu 195 200
205 Leu Glu Leu Gly Met Arg Gly Leu Thr Ser Tyr Ala Glu
Thr Val Ser 210 215 220
Val Tyr Gly Thr Glu Asn Val Phe Thr Asp Gly Asp Asp Thr Pro Trp 225
230 235 240 Ser Lys Ala Phe
Leu Ala Ser Ala Tyr Ala Ser Arg Gly Leu Lys Met 245
250 255 Arg Phe Thr Ser Gly Ser Gly Ser Glu
Ala Leu Met Gly Tyr Ala Glu 260 265
270 Gly Lys Ser Met Leu Tyr Leu Glu Ala Arg Cys Ile Tyr Ile
Thr Lys 275 280 285
Ala Ala Gly Val Gln Gly Leu Gln Asn Gly Ser Val Ser Cys Ile Gly 290
295 300 Met Thr Gly Ala Leu
Pro Ser Gly Ile Arg Ala Val Leu Gly Glu Asn 305 310
315 320 Leu Ile Thr Thr Met Leu Asp Ile Glu Val
Ala Ser Ala Asn Asp Gln 325 330
335 Thr Phe Ser His Ser Asp Ile Arg Arg Thr Ala Arg Met Leu Met
Gln 340 345 350 Met
Leu Pro Gly Thr Asp Phe Ile Phe Ser Gly Tyr Ser Ser Val Pro 355
360 365 Asn Tyr Asp Asn Met Phe
Ala Gly Ser Asn Phe Asp Ala Glu Asp Phe 370 375
380 Asp Asp Tyr Asn Val Ile Gln Arg Asp Leu Met
Val Asp Gly Gly Leu 385 390 395
400 Arg Pro Val Ser Glu Glu Glu Val Ile Thr Ile Arg Asn Lys Ala Ala
405 410 415 Arg Ala
Ile Gln Ala Val Phe Glu Gly Leu Lys Leu Pro Ala Ile Thr 420
425 430 Asp Glu Glu Val Glu Ala Val
Thr Tyr Ser His Gly Ser Lys Asp Val 435 440
445 Pro Glu Arg Asn Val Val Glu Asp Leu Lys Ala Ala
Glu Glu Met Ile 450 455 460
Asn Arg Gly Ile Thr Gly Ile Asp Val Val Lys Ala Leu Ser Lys His 465
470 475 480 Gly Phe Asp
Asp Ile Ala Glu Asn Ile Leu Asn Met Leu Lys Gln Arg 485
490 495 Ile Ser Gly Asp Tyr Leu Gln Thr
Ser Ala Ile Ile Asp Lys Asn Phe 500 505
510 Asn Val Val Ser Ala Val Asn Asp Cys Asn Asp Tyr Met
Gly Pro Gly 515 520 525
Thr Gly Tyr Arg Leu Ser Lys Glu Arg Trp Asp Glu Ile Lys Asn Ile 530
535 540 Pro Asn Ala Met
Lys Pro Glu Asp Ile Lys 545 550
136179PRTClostridium pasteurianum 136Met Glu Leu Lys Glu Lys Asp Ile Ala
Leu Ser Gly Asn Gln Ser Asn 1 5 10
15 Glu Val Val Ile Gly Ile Ala Pro Ala Phe Gly Lys Tyr Gln
His Gln 20 25 30
Ser Ile Val Gly Val Pro His Asp Lys Ile Leu Arg Glu Leu Ile Ala
35 40 45 Gly Ile Glu Glu
Glu Gly Leu Lys Ser Arg Val Val Arg Ile Ile Arg 50
55 60 Thr Ser Asp Val Ser Phe Ile Ala
His Asp Ala Ala Val Leu Ser Gly 65 70
75 80 Ser Gly Ile Gly Ile Gly Ile Gln Ser Lys Gly Thr
Thr Val Ile His 85 90
95 Gln Lys Asp Leu Leu Pro Leu Asn Asn Leu Glu Leu Phe Pro Gln Ala
100 105 110 Pro Leu Leu
Asp Leu Asp Ile Phe Arg Leu Ile Gly Lys Asn Ala Ala 115
120 125 Lys Tyr Ala Lys Gly Glu Ser Pro
Asn Pro Val Pro Thr Arg Asn Asp 130 135
140 Gln Met Val Arg Pro Lys Phe Gln Ala Lys Ala Ala Leu
Leu His Ile 145 150 155
160 Lys Glu Thr Lys His Val Val Gln Asn Ala Lys Pro Ile Glu Leu Glu
165 170 175 Ile Ile Ser
137146PRTClostridium pasteurianum 137Met Ser Asp Ile Thr Asn Asn Ile Lys
Val Asp Tyr Glu Asn Asp Tyr 1 5 10
15 Pro Leu Ala Ala Lys Arg Ser Glu Trp Ile Lys Thr Pro Thr
Gly Lys 20 25 30
Asn Leu Lys Asp Ile Thr Leu Glu Ala Val Ile Asp Glu Asn Val Lys
35 40 45 Ala Glu Asp Val
Arg Ile Ser Arg Asp Thr Leu Glu Leu Gln Ala Gln 50
55 60 Val Ala Glu Gly Ser Gly Arg Cys
Ala Ile Ala Arg Asn Phe Arg Arg 65 70
75 80 Ala Ala Glu Leu Ile Ser Ile Ser Asp Glu Arg Ile
Leu Glu Ile Tyr 85 90
95 Asn Ala Leu Arg Pro Tyr Arg Ser Thr Lys Asn Glu Leu Leu Ala Ile
100 105 110 Ala Asp Glu
Leu Glu Glu Lys Tyr Asp Ala Lys Val Asn Ala Asp Phe 115
120 125 Ile Arg Glu Ala Ala Glu Val Tyr
Ser Lys Arg Asn Lys Val Arg Ile 130 135
140 Glu Asp 145 138555PRTEscherichia blattae 138Met
Arg Arg Ser Lys Arg Phe Glu Val Leu Glu Lys Arg Pro Val Asn 1
5 10 15 Gln Asp Gly Leu Ile Gly
Glu Trp Pro Glu Glu Gly Leu Ile Ala Met 20
25 30 Gly Ser Pro Trp Asp Pro Pro Ser Ser Val
Lys Val Glu Gln Gly Arg 35 40
45 Ile Val Glu Leu Asp Gly Lys Ala Arg Ala Asp Phe Asp Met
Ile Asp 50 55 60
Arg Phe Ile Ala Asp Tyr Ala Ile Asn Ile Glu Glu Thr Glu His Ala 65
70 75 80 Met Gly Leu Asp Ala
Leu Thr Ile Ala Arg Met Leu Val Asp Ile Asn 85
90 95 Val Ser Arg Ala Glu Ile Ile Lys Val Thr
Thr Ala Ile Thr Pro Ala 100 105
110 Lys Ala Val Glu Val Met Ser His Met Asn Val Val Glu Met Met
Met 115 120 125 Ala
Leu Gln Lys Met Arg Ala Arg Arg Thr Pro Ser Asn Gln Cys His 130
135 140 Val Thr Asn Leu Lys Asp
Asn Pro Val Gln Ile Ala Ala Asp Ala Ala 145 150
155 160 Glu Ala Gly Ile Arg Gly Phe Ser Glu Gln Glu
Thr Thr Val Gly Ile 165 170
175 Ala Arg Tyr Ala Pro Phe Asn Ala Leu Ala Leu Leu Ile Gly Ser Gln
180 185 190 Ser Gly
Arg Pro Gly Val Leu Thr Gln Cys Ser Val Glu Glu Ala Thr 195
200 205 Glu Leu Glu Leu Gly Met Arg
Gly Phe Thr Ser Tyr Ala Glu Thr Val 210 215
220 Ser Val Tyr Gly Thr Glu Ala Val Phe Thr Asp Gly
Asp Asp Thr Pro 225 230 235
240 Trp Ser Lys Ala Phe Leu Ala Ser Ala Tyr Ala Ser Arg Gly Leu Lys
245 250 255 Met Arg Tyr
Thr Ser Gly Thr Gly Ser Glu Ala Leu Met Gly Tyr Ala 260
265 270 Glu Ser Lys Ser Met Leu Tyr Leu
Glu Ser Arg Cys Ile Phe Ile Thr 275 280
285 Lys Gly Ala Gly Val Gln Gly Leu Gln Asn Gly Ala Val
Ser Cys Ile 290 295 300
Gly Met Thr Gly Ala Val Pro Ser Gly Ile Arg Ala Val Leu Ala Glu 305
310 315 320 Asn Leu Ile Ala
Ser Met Leu Asp Leu Glu Val Ala Ser Ala Asn Asp 325
330 335 Gln Thr Phe Ser His Ser Asp Ile Arg
Arg Thr Ala Arg Thr Leu Met 340 345
350 Gln Met Leu Pro Gly Thr Asp Phe Ile Phe Ser Gly Tyr Ser
Ala Val 355 360 365
Pro Asn Tyr Asp Asn Met Phe Ala Gly Ser Asn Phe Asp Ala Glu Asp 370
375 380 Phe Asp Asp Tyr Asn
Ile Leu Gln Arg Asp Leu Met Val Asp Gly Gly 385 390
395 400 Leu Arg Pro Val Ser Glu Glu Glu Thr Ile
Ala Ile Arg Asn Lys Ala 405 410
415 Ala Arg Ala Val Gln Ala Val Phe Arg Glu Leu Gly Leu Pro Pro
Val 420 425 430 Thr
Asp Glu Glu Val Thr Ala Ala Thr Tyr Ala His Gly Ser Lys Asp 435
440 445 Met Pro Pro Arg Asn Val
Val Glu Asp Leu Ser Ala Val Glu Glu Met 450 455
460 Met Lys Arg Asn Ile Thr Gly Leu Asp Ile Val
Arg Ala Leu Ser Val 465 470 475
480 Asn Gly Phe Asp Asp Val Ala Asn Asn Ile Leu Asn Met Leu Arg Gln
485 490 495 Arg Val
Thr Gly Asp Tyr Leu Gln Thr Ser Ala Ile Leu Asp Arg Glu 500
505 510 Phe Glu Val Val Ser Ala Val
Asn Asp Ile Asn Asp Tyr Gln Gly Pro 515 520
525 Gly Thr Gly Tyr Arg Ile Ser Pro Gln Arg Trp Glu
Glu Ile Lys Asn 530 535 540
Ile Ala Thr Val Ile Gln Pro Asp Ser Ile Glu 545 550
555 139196PRTEscherichia blattae 139Met Glu Thr Thr Gln
Lys Lys Ala Pro Val Phe Thr Leu Asn Leu Val 1 5
10 15 Glu Ser Gly Val Ala Lys Pro Gly Glu Arg
Ser Asp Glu Val Val Ile 20 25
30 Gly Val Gly Pro Ala Phe Asp Lys Tyr Gln His Lys Thr Leu Ile
Asp 35 40 45 Met
Pro His Lys Ala Ile Ile Lys Glu Leu Val Ala Gly Val Glu Glu 50
55 60 Glu Gly Leu His Ala Arg
Val Val Arg Ile Leu Arg Thr Ser Asp Val 65 70
75 80 Ser Phe Met Ala Trp Asp Ala Ala Asn Leu Ser
Gly Ser Gly Ile Gly 85 90
95 Ile Gly Ile Gln Ser Lys Gly Thr Thr Val Ile His Gln Arg Asp Leu
100 105 110 Leu Pro
Leu Ser Asn Leu Glu Leu Phe Ser Gln Ala Pro Leu Leu Thr 115
120 125 Leu Glu Thr Tyr Arg Gln Ile
Gly Lys Asn Ala Ala Arg Tyr Ala Arg 130 135
140 Lys Glu Ser Pro Ser Pro Val Pro Val Val Asn Asp
Gln Met Val Arg 145 150 155
160 Pro Lys Phe Met Ala Lys Ala Ala Leu Phe His Ile Lys Glu Thr Lys
165 170 175 His Val Val
Ala Asp Ala Lys Pro Val Thr Leu Asn Ile Glu Ile Thr 180
185 190 Arg Glu Glu Ala 195
140141PRTEscherichia blattae 140Met Thr Thr Thr Lys Met Ser Ala Ala Asp
Tyr Pro Leu Ala Ser Arg 1 5 10
15 Cys Pro Glu Arg Ile Gln Thr Pro Thr Gly Lys Pro Leu Thr Asp
Ile 20 25 30 Thr
Leu Glu Asn Val Leu Ala Gly Lys Val Gly Pro Gln Asp Val Arg 35
40 45 Ile Ser Arg Glu Thr Leu
Glu Tyr Gln Ala Gln Ile Ala Glu Gln Met 50 55
60 His Arg His Ala Ile Ala Arg Asn Leu Arg Arg
Ala Gly Glu Leu Ile 65 70 75
80 Ala Ile Pro Asp Ala Arg Ile Leu Glu Ile Tyr Asn Ala Leu Arg Pro
85 90 95 Tyr Arg
Ser Ser Val Glu Glu Leu Leu Ala Ile Ala Asp Glu Leu Glu 100
105 110 Thr Arg Tyr Gln Ala Thr Val
Asn Ala Ala Phe Ile Arg Glu Ala Ala 115 120
125 Glu Val Tyr Arg Gln Arg Asp Lys Leu Arg Lys Glu
Ala 130 135 140
141555PRTCitrobacter freundii 141Met Arg Arg Ser Lys Arg Phe Glu Val Leu
Ala Gln Arg Pro Val Asn 1 5 10
15 Gln Asp Gly Leu Ile Gly Glu Trp Pro Glu Glu Gly Leu Ile Ala
Met 20 25 30 Glu
Ser Pro Tyr Asp Pro Ala Ser Ser Val Lys Val Glu Asn Gly Arg 35
40 45 Ile Val Glu Leu Asp Gly
Lys Ser Arg Ala Glu Phe Asp Met Ile Asp 50 55
60 Arg Phe Ile Ala Asp Tyr Ala Ile Asn Val Pro
Glu Ala Glu Arg Ala 65 70 75
80 Met Gln Leu Asp Ala Leu Glu Ile Ala Arg Met Leu Val Asp Ile His
85 90 95 Val Ser
Arg Glu Glu Ile Ile Ala Ile Thr Thr Ala Ile Thr Pro Ala 100
105 110 Lys Arg Leu Glu Val Met Ala
Gln Met Asn Val Val Glu Met Met Met 115 120
125 Ala Leu Gln Lys Met Arg Ala Arg Arg Thr Pro Ser
Asn Gln Cys His 130 135 140
Val Thr Asn Leu Lys Asp Asn Pro Val Gln Ile Ala Ala Asp Ala Ala 145
150 155 160 Glu Ala Gly
Ile Arg Gly Phe Ser Glu Gln Glu Thr Thr Val Gly Ile 165
170 175 Ala Arg Tyr Ala Pro Phe Asn Ala
Leu Ala Leu Leu Val Gly Ser Gln 180 185
190 Cys Gly Ala Pro Gly Val Leu Thr Gln Cys Ser Val Glu
Glu Ala Thr 195 200 205
Glu Leu Glu Leu Gly Met Arg Gly Leu Thr Ser Tyr Ala Glu Thr Val 210
215 220 Ser Val Tyr Gly
Thr Glu Ser Val Phe Thr Asp Gly Asp Asp Thr Pro 225 230
235 240 Trp Ser Lys Ala Phe Leu Ala Ser Ala
Tyr Ala Ser Arg Gly Leu Lys 245 250
255 Met Arg Tyr Thr Ser Gly Thr Gly Ser Glu Ala Leu Met Gly
Tyr Ser 260 265 270
Glu Ser Lys Ser Met Leu Tyr Leu Glu Ser Arg Cys Ile Phe Ile Thr
275 280 285 Lys Gly Ala Gly
Val Gln Gly Leu Gln Asn Gly Ala Val Ser Cys Ile 290
295 300 Gly Met Thr Gly Ala Val Pro Ser
Gly Ile Arg Ala Val Leu Ala Glu 305 310
315 320 Asn Leu Ile Ala Ser Met Leu Asp Leu Glu Val Ala
Ser Ala Asn Asp 325 330
335 Gln Thr Phe Ser His Ser Asp Ile Arg Arg Thr Ala Arg Thr Leu Met
340 345 350 Gln Met Leu
Pro Gly Thr Asp Phe Ile Phe Ser Gly Tyr Ser Ala Val 355
360 365 Pro Asn Tyr Asp Asn Met Phe Ala
Gly Ser Asn Phe Asp Ala Glu Asp 370 375
380 Phe Asp Asp Tyr Asn Ile Leu Gln Arg Asp Leu Met Val
Asp Gly Gly 385 390 395
400 Leu Arg Pro Val Thr Glu Glu Glu Thr Ile Ala Ile Arg Asn Lys Ala
405 410 415 Ala Arg Ala Ile
Gln Ala Val Phe Arg Glu Leu Gly Leu Pro Leu Ile 420
425 430 Ser Asp Glu Glu Val Asp Ala Ala Thr
Tyr Ala His Gly Ser Lys Asp 435 440
445 Met Pro Ala Arg Asn Val Val Glu Asp Leu Ala Ala Val Glu
Glu Met 450 455 460
Met Lys Arg Asn Ile Thr Gly Leu Asp Ile Val Gly Ala Leu Ser Ser 465
470 475 480 Ser Gly Phe Glu Asp
Ile Ala Ser Asn Ile Leu Asn Met Leu Arg Gln 485
490 495 Arg Val Thr Gly Asp Tyr Leu Gln Thr Ser
Ala Ile Leu Asp Arg Gln 500 505
510 Phe Asp Val Val Ser Ala Val Asn Asp Ile Asn Asp Tyr Gln Gly
Pro 515 520 525 Gly
Thr Gly Tyr Arg Ile Ser Ala Glu Arg Trp Ala Glu Ile Lys Asn 530
535 540 Ile Ala Gly Val Val Gln
Pro Gly Ser Ile Glu 545 550 555
142194PRTCitrobacter freundii 142Met Glu Cys Thr Thr Glu Arg Lys Pro Val
Phe Thr Leu Gln Val Ser 1 5 10
15 Glu Gly Glu Ala Ala Lys Ala Asp Glu Arg Val Asp Glu Val Val
Ile 20 25 30 Gly
Val Gly Pro Ala Phe Asp Lys Tyr Gln His Lys Thr Leu Ile Asp 35
40 45 Met Pro His Lys Ala Ile
Leu Lys Glu Leu Val Ala Gly Ile Glu Glu 50 55
60 Glu Gly Leu His Ala Arg Val Val Arg Ile Leu
Arg Thr Ser Asp Val 65 70 75
80 Ser Phe Met Ala Trp Asp Ala Ala Asn Leu Ser Gly Ser Gly Ile Gly
85 90 95 Ile Gly
Ile Gln Ser Lys Gly Thr Thr Val Ile His Gln Arg Asp Leu 100
105 110 Leu Pro Leu Ser Asn Leu Glu
Leu Phe Ser Gln Ala Pro Leu Leu Thr 115 120
125 Leu Glu Thr Tyr Arg Gln Ile Gly Lys Asn Ala Ala
Arg Tyr Ala Arg 130 135 140
Lys Glu Ser Pro Ser Pro Val Pro Val Val Asn Asp Gln Met Val Arg 145
150 155 160 Pro Lys Phe
Met Ala Lys Ala Ala Leu Phe His Ile Lys Glu Thr Lys 165
170 175 His Val Val Gln Asp Arg Ala Pro
Val Thr Leu His Ile Ala Leu Val 180 185
190 Arg Glu 143142PRTCitrobacter freundii 143Met Asn
Asp Asn Ile Met Thr Ala Gln Asp Tyr Pro Leu Ala Thr Arg 1 5
10 15 Cys Pro Glu Lys Ile Gln Thr
Pro Thr Gly Lys Pro Leu Thr Glu Ile 20 25
30 Thr Leu Glu Asn Val Leu Ala Gly Arg Val Gly Pro
Gln Asp Val Arg 35 40 45
Ile Ser Gln Gln Thr Leu Glu Tyr Gln Ala Gln Ile Ala Glu Gln Met
50 55 60 Gln Arg His
Ala Val Ala Arg Asn Phe Arg Arg Ala Ala Glu Leu Ile 65
70 75 80 Ala Ile Pro Asp Ala Arg Ile
Leu Glu Ile Tyr Asn Ala Leu Arg Pro 85
90 95 Phe Arg Ser Ser Phe Ala Glu Leu Gln Ala Ile
Ala Asp Glu Leu Glu 100 105
110 His Thr Trp His Ala Thr Val Asn Ala Gly Phe Val Arg Glu Ser
Ala 115 120 125 Glu
Val Tyr Leu Gln Arg Asn Lys Leu Arg Lys Gly Ser Gln 130
135 140 1441359DNAArtificial sequenceCodon
optimized Vibrio fluvialis aminepyruvate transaminase 144atgaacaaac
cacagtcttg ggaagctcgt gcagaaacct attctctgta cggcttcact 60gacatgccgt
ccctgcacca gcgtggtact gttgttgtca cgcacggcga aggtccgtac 120attgttgacg
tcaatggtcg ccgttatctg gacgctaatt ctggcctgtg gaatatggtt 180gcaggttttg
accataaggg tctgatcgac gcagctaagg ctcagtacga gcgttttccg 240ggctaccatg
cgttcttcgg tcgtatgagc gatcagacgg tgatgctgtc cgaaaaactg 300gtagaagtct
ctccgttcga cagcggccgt gtgttctata cgaacagcgg tagcgaagca 360aacgacacta
tggttaagat gctgtggttc ctgcatgcgg cggaaggtaa gccacaaaag 420cgcaaaattc
tgacccgttg gaacgcgtat cacggcgtta ctgcagttag cgcctccatg 480accggtaaac
cgtacaacag cgttttcggt ctgccgctgc caggtttcgt tcacctgact 540tgccctcact
actggcgtta cggtgaagaa ggcgagacgg aagaacaatt cgttgcacgc 600ctggcacgcg
aactggaaga gactatccag cgtgagggtg ctgacactat cgctggcttc 660tttgctgagc
cggttatggg tgcaggtggt gttattccgc ctgctaaagg ttattttcag 720gctattctgc
caatcctgcg taaatatgac atcccggtta tctctgacga agttatctgt 780ggttttggtc
gcactggcaa cacctggggt tgcgtaactt atgattttac tccggatgct 840atcatctcta
gcaaaaacct gaccgccggt ttcttcccga tgggcgcagt gatcctgggt 900ccagaactga
gcaagcgcct ggaaaccgca attgaagcaa tcgaggaatt tccgcacggc 960tttaccgcgt
ccggccatcc ggtaggctgt gcaatcgcgc tgaaagcgat cgatgttgtt 1020atgaacgaag
gcctggcgga aaacgttcgc cgtctggcac cgcgcttcga agaacgtctg 1080aaacatatcg
cggaacgtcc gaacattggt gaatatcgtg gtatcggttt tatgtgggct 1140ctggaggcag
tcaaagacaa agcgtctaaa actccgttcg atggcaatct gagcgtgagc 1200gaacgtatcg
ccaacacttg caccgacctg ggtctgatct gccgtccact gggccaaagc 1260gtagtgctgt
gtccgccgtt tatcctgacc gaagcgcaaa tggacgaaat gttcgacaaa 1320ctggagaaag
cactggataa agtgttcgca gaggtggca
13591451668DNAKlebsiella pneumoniae 145atgaaaagat caaaacgatt tgcagtactg
gcccagcgcc ccgtcaatca ggacgggctg 60attggcgagt ggcctgaaga ggggctgatc
gccatggaca gcccctttga cccggtctct 120tcagtaaaag tggacaacgg tctgatcgtc
gaactggacg gcaaacgccg ggaccagttt 180gacatgatcg accgatttat cgccgattac
gcgatcaacg ttgagcgcac agagcaggca 240atgcgcctgg aggcggtgga aatagcccgt
atgctggtgg atattcacgt cagccgggag 300gagatcattg ccatcactac cgccatcacg
ccggccaaag cggtcgaggt gatggcgcag 360atgaacgtgg tggagatgat gatggcgctg
cagaagatgc gtgcccgccg gaccccctcc 420aaccagtgcc acgtcaccaa tctcaaagat
aatccggtgc agattgccgc tgacgccgcc 480gaggccggga tccgcggctt ctcagaacag
gagaccacgg tcggtatcgc gcgctacgcg 540ccgtttaacg ccctggcgct gttggtcggt
tcgcagtgcg gccgccccgg cgtgttgacg 600cagtgctcgg tggaagaggc caccgagctg
gagctgggca tgcgtggctt aaccagctac 660gccgagacgg tgtcggtcta cggcaccgaa
gcggtattta ccgacggcga tgatacgccg 720tggtcaaagg cgttcctcgc ctcggcctac
gcctcccgcg ggttgaaaat gcgctacacc 780tccggcaccg gatccgaagc gctgatgggc
tattcggaga gcaagtcgat gctctacctc 840gaatcgcgct gcatcttcat tactaaaggc
gccggggttc agggactgca aaacggcgcg 900gtgagctgta tcggcatgac cggcgctgtg
ccgtcgggca ttcgggcggt gctggcggaa 960aacctgatcg cctctatgct cgacctcgaa
gtggcgtccg ccaacgacca gactttctcc 1020cactcggata ttcgccgcac cgcgcgcacc
ctgatgcaga tgctgccggg caccgacttt 1080attttctccg gctacagcgc ggtgccgaac
tacgacaaca tgttcgccgg ctcgaacttc 1140gatgcggaag attttgatga ttacaacatc
ctgcagcgtg acctgatggt tgacggcggc 1200ctgcgtccgg tgaccgaggc ggaaaccatt
gccattcgcc agaaagcggc gcgggcgatc 1260caggcggttt tccgcgagct ggggctgccg
ccaatcgccg acgaggaggt ggaggccgcc 1320acctacgcgc acggcagcaa cgagatgccg
ccgcgtaacg tggtggagga tctgagtgcg 1380gtggaagaga tgatgaagcg caacatcacc
ggcctcgata ttgtcggcgc gctgagccgc 1440agcggctttg aggatatcgc cagcaatatt
ctcaatatgc tgcgccagcg ggtcaccggc 1500gattacctgc agacctcggc cattctcgat
cggcagttcg aggtggtgag tgcggtcaac 1560gacatcaatg actatcaggg gccgggcacc
ggctatcgca tctctgccga acgctgggcg 1620gagatcaaaa atattccggg cgtggttcag
cccgacacca ttgaataa 1668146555PRTKlebsiella pneumoniae
146Met Lys Arg Ser Lys Arg Phe Ala Val Leu Ala Gln Arg Pro Val Asn 1
5 10 15 Gln Asp Gly Leu
Ile Gly Glu Trp Pro Glu Glu Gly Leu Ile Ala Met 20
25 30 Asp Ser Pro Phe Asp Pro Val Ser Ser
Val Lys Val Asp Asn Gly Leu 35 40
45 Ile Val Glu Leu Asp Gly Lys Arg Arg Asp Gln Phe Asp Met
Ile Asp 50 55 60
Arg Phe Ile Ala Asp Tyr Ala Ile Asn Val Glu Arg Thr Glu Gln Ala 65
70 75 80 Met Arg Leu Glu Ala
Val Glu Ile Ala Arg Met Leu Val Asp Ile His 85
90 95 Val Ser Arg Glu Glu Ile Ile Ala Ile Thr
Thr Ala Ile Thr Pro Ala 100 105
110 Lys Ala Val Glu Val Met Ala Gln Met Asn Val Val Glu Met Met
Met 115 120 125 Ala
Leu Gln Lys Met Arg Ala Arg Arg Thr Pro Ser Asn Gln Cys His 130
135 140 Val Thr Asn Leu Lys Asp
Asn Pro Val Gln Ile Ala Ala Asp Ala Ala 145 150
155 160 Glu Ala Gly Ile Arg Gly Phe Ser Glu Gln Glu
Thr Thr Val Gly Ile 165 170
175 Ala Arg Tyr Ala Pro Phe Asn Ala Leu Ala Leu Leu Val Gly Ser Gln
180 185 190 Cys Gly
Arg Pro Gly Val Leu Thr Gln Cys Ser Val Glu Glu Ala Thr 195
200 205 Glu Leu Glu Leu Gly Met Arg
Gly Leu Thr Ser Tyr Ala Glu Thr Val 210 215
220 Ser Val Tyr Gly Thr Glu Ala Val Phe Thr Asp Gly
Asp Asp Thr Pro 225 230 235
240 Trp Ser Lys Ala Phe Leu Ala Ser Ala Tyr Ala Ser Arg Gly Leu Lys
245 250 255 Met Arg Tyr
Thr Ser Gly Thr Gly Ser Glu Ala Leu Met Gly Tyr Ser 260
265 270 Glu Ser Lys Ser Met Leu Tyr Leu
Glu Ser Arg Cys Ile Phe Ile Thr 275 280
285 Lys Gly Ala Gly Val Gln Gly Leu Gln Asn Gly Ala Val
Ser Cys Ile 290 295 300
Gly Met Thr Gly Ala Val Pro Ser Gly Ile Arg Ala Val Leu Ala Glu 305
310 315 320 Asn Leu Ile Ala
Ser Met Leu Asp Leu Glu Val Ala Ser Ala Asn Asp 325
330 335 Gln Thr Phe Ser His Ser Asp Ile Arg
Arg Thr Ala Arg Thr Leu Met 340 345
350 Gln Met Leu Pro Gly Thr Asp Phe Ile Phe Ser Gly Tyr Ser
Ala Val 355 360 365
Pro Asn Tyr Asp Asn Met Phe Ala Gly Ser Asn Phe Asp Ala Glu Asp 370
375 380 Phe Asp Asp Tyr Asn
Ile Leu Gln Arg Asp Leu Met Val Asp Gly Gly 385 390
395 400 Leu Arg Pro Val Thr Glu Ala Glu Thr Ile
Ala Ile Arg Gln Lys Ala 405 410
415 Ala Arg Ala Ile Gln Ala Val Phe Arg Glu Leu Gly Leu Pro Pro
Ile 420 425 430 Ala
Asp Glu Glu Val Glu Ala Ala Thr Tyr Ala His Gly Ser Asn Glu 435
440 445 Met Pro Pro Arg Asn Val
Val Glu Asp Leu Ser Ala Val Glu Glu Met 450 455
460 Met Lys Arg Asn Ile Thr Gly Leu Asp Ile Val
Gly Ala Leu Ser Arg 465 470 475
480 Ser Gly Phe Glu Asp Ile Ala Ser Asn Ile Leu Asn Met Leu Arg Gln
485 490 495 Arg Val
Thr Gly Asp Tyr Leu Gln Thr Ser Ala Ile Leu Asp Arg Gln 500
505 510 Phe Glu Val Val Ser Ala Val
Asn Asp Ile Asn Asp Tyr Gln Gly Pro 515 520
525 Gly Thr Gly Tyr Arg Ile Ser Ala Glu Arg Trp Ala
Glu Ile Lys Asn 530 535 540
Ile Pro Gly Val Val Gln Pro Asp Thr Ile Glu 545 550
555 147585DNAKlebsiella pneumoniae 147gtgcaacaga
caacccaaat tcagccctct tttaccctga aaacccgcga gggcggggta 60gcttctgccg
atgaacgcgc cgatgaagtg gtgatcggcg tcggccctgc cttcgataaa 120caccagcatc
acactctgat cgatatgccc catggcgcga tcctcaaaga gctgattgcc 180ggggtggaag
aagaggggct tcacgcccgg gtggtgcgca ttctgcgcac gtccgacgtc 240tcctttatgg
cctgggatgc ggccaacctg agcggctcgg ggatcggcat cggtatccag 300tcgaagggga
ccacggtcat ccatcagcgc gatctgctgc cgctcagcaa cctggagctg 360ttctcccagg
cgccgctgct gacgctggag acctaccggc agattggcaa aaacgctgcg 420cgctatgcgc
gcaaagagtc accttcgccg gtgccggtgg tgaacgatca gatggtgcgg 480ccgaaattta
tggccaaagc cgcgctattt catatcaaag agaccaaaca tgtggtgcag 540gacgccgagc
ccgtcaccct gcacatcgac ttagtaaggg agtga
585148194PRTKlebsiella pneumoniae 148Met Gln Gln Thr Thr Gln Ile Gln Pro
Ser Phe Thr Leu Lys Thr Arg 1 5 10
15 Glu Gly Gly Val Ala Ser Ala Asp Glu Arg Ala Asp Glu Val
Val Ile 20 25 30
Gly Val Gly Pro Ala Phe Asp Lys His Gln His His Thr Leu Ile Asp
35 40 45 Met Pro His Gly
Ala Ile Leu Lys Glu Leu Ile Ala Gly Val Glu Glu 50
55 60 Glu Gly Leu His Ala Arg Val Val
Arg Ile Leu Arg Thr Ser Asp Val 65 70
75 80 Ser Phe Met Ala Trp Asp Ala Ala Asn Leu Ser Gly
Ser Gly Ile Gly 85 90
95 Ile Gly Ile Gln Ser Lys Gly Thr Thr Val Ile His Gln Arg Asp Leu
100 105 110 Leu Pro Leu
Ser Asn Leu Glu Leu Phe Ser Gln Ala Pro Leu Leu Thr 115
120 125 Leu Glu Thr Tyr Arg Gln Ile Gly
Lys Asn Ala Ala Arg Tyr Ala Arg 130 135
140 Lys Glu Ser Pro Ser Pro Val Pro Val Val Asn Asp Gln
Met Val Arg 145 150 155
160 Pro Lys Phe Met Ala Lys Ala Ala Leu Phe His Ile Lys Glu Thr Lys
165 170 175 His Val Val Gln
Asp Ala Glu Pro Val Thr Leu His Ile Asp Leu Val 180
185 190 Arg Glu 149426DNAKlebsiella
pneumoniae 149atgagcgaga aaaccatgcg cgtgcaggat tatccgttag ccacccgctg
cccggagcat 60atcctgacgc ctaccggcaa accattgacc gatattaccc tcgagaaggt
gctctctggc 120gaggtgggcc cgcaggatgt gcggatctcc cgccagaccc ttgagtacca
ggcgcagatt 180gccgagcaga tgcagcgcca tgcggtggcg cgcaatttcc gccgcgcggc
ggagcttatc 240gccattcctg acgagcgcat tctggctatc tataacgcgc tgcgcccgtt
ccgctcctcg 300caggcggagc tgctggcgat cgccgacgag ctggagcaca cctggcatgc
gacagtgaat 360gccgcctttg tccgggagtc ggcggaagtg tatcagcagc ggcataagct
gcgtaaagga 420agctaa
426150141PRTKlebsiella pneumoniae 150Met Ser Glu Lys Thr Met
Arg Val Gln Asp Tyr Pro Leu Ala Thr Arg 1 5
10 15 Cys Pro Glu His Ile Leu Thr Pro Thr Gly Lys
Pro Leu Thr Asp Ile 20 25
30 Thr Leu Glu Lys Val Leu Ser Gly Glu Val Gly Pro Gln Asp Val
Arg 35 40 45 Ile
Ser Arg Gln Thr Leu Glu Tyr Gln Ala Gln Ile Ala Glu Gln Met 50
55 60 Gln Arg His Ala Val Ala
Arg Asn Phe Arg Arg Ala Ala Glu Leu Ile 65 70
75 80 Ala Ile Pro Asp Glu Arg Ile Leu Ala Ile Tyr
Asn Ala Leu Arg Pro 85 90
95 Phe Arg Ser Ser Gln Ala Glu Leu Leu Ala Ile Ala Asp Glu Leu Glu
100 105 110 His Thr
Trp His Ala Thr Val Asn Ala Ala Phe Val Arg Glu Ser Ala 115
120 125 Glu Val Tyr Gln Gln Arg His
Lys Leu Arg Lys Gly Ser 130 135 140
1511824DNAKlebsiella pneumoniae 151atgccgttaa tagccgggat tgatatcggc
aacgccacca ccgaggtggc gctggcgtcc 60gactacccgc aggcgagggc gtttgttgcc
agcgggatcg tcgcgacgac gggcatgaaa 120gggacgcggg acaatatcgc cgggaccctc
gccgcgctgg agcaggccct ggcgaaaaca 180ccgtggtcga tgagcgatgt ctctcgcatc
tatcttaacg aagccgcgcc ggtgattggc 240gatgtggcga tggagaccat caccgagacc
attatcaccg aatcgaccat gatcggtcat 300aacccgcaga cgccgggcgg ggtgggcgtt
ggcgtgggga cgactatcgc cctcgggcgg 360ctggcgacgc tgccggcggc gcagtatgcc
gaggggtgga tcgtactgat tgacgacgcc 420gtcgatttcc ttgacgccgt gtggtggctc
aatgaggcgc tcgaccgggg gatcaacgtg 480gtggcggcga tcctcaaaaa ggacgacggc
gtgctggtga acaaccgcct gcgtaaaacc 540ctgccggtgg tggatgaagt gacgctgctg
gagcaggtcc ccgagggggt aatggcggcg 600gtggaagtgg ccgcgccggg ccaggtggtg
cggatcctgt cgaatcccta cgggatcgcc 660accttcttcg ggctaagccc ggaagagacc
caggccatcg tccccatcgc ccgcgccctg 720attggcaacc gttccgcggt ggtgctcaag
accccgcagg gggatgtgca gtcgcgggtg 780atcccggcgg gcaacctcta cattagcggc
gaaaagcgcc gcggagaggc cgatgtcgcc 840gagggcgcgg aagccatcat gcaggcgatg
agcgcctgcg ctccggtacg cgacatccgc 900ggcgaaccgg gcacccacgc cggcggcatg
cttgagcggg tgcgcaaggt aatggcgtcc 960ctgaccggcc atgagatgag cgcgatatac
atccaggatc tgctggcggt ggatacgttt 1020attccgcgca aggtgcaggg cgggatggcc
ggcgagtgcg ccatggagaa tgccgtcggg 1080atggcggcga tggtgaaagc ggatcgtctg
caaatgcagg ttatcgcccg cgaactgagc 1140gcccgactgc agaccgaggt ggtggtgggc
ggcgtggagg ccaacatggc catcgccggg 1200gcgttaacca ctcccggctg tgcggcgccg
ctggcgatcc tcgacctcgg cgccggctcg 1260acggatgcgg cgatcgtcaa cgcggagggg
cagataacgg cggtccatct cgccggggcg 1320gggaatatgg tcagcctgtt gattaaaacc
gagctgggcc tcgaggatct ttcgctggcg 1380gaagcgataa aaaaataccc gctggccaaa
gtggaaagcc tgttcagtat tcgtcacgag 1440aatggcgcgg tggagttctt tcgggaagcc
ctcagcccgg cggtgttcgc caaagtggtg 1500tacatcaagg agggcgaact ggtgccgatc
gataacgcca gcccgctgga aaaaattcgt 1560ctcgtgcgcc ggcaggcgaa agagaaagtg
tttgtcacca actgcctgcg cgcgctgcgc 1620caggtctcac ccggcggttc cattcgcgat
atcgcctttg tggtgctggt gggcggctca 1680tcgctggact ttgagatccc gcagcttatc
acggaagcct tgtcgcacta tggcgtggtc 1740gccgggcagg gcaatattcg gggaacagaa
gggccgcgca atgcggtcgc caccgggctg 1800ctactggccg gtcaggcgaa ttaa
1824152607PRTKlebsiella pneumoniae
152Met Pro Leu Ile Ala Gly Ile Asp Ile Gly Asn Ala Thr Thr Glu Val 1
5 10 15 Ala Leu Ala Ser
Asp Tyr Pro Gln Ala Arg Ala Phe Val Ala Ser Gly 20
25 30 Ile Val Ala Thr Thr Gly Met Lys Gly
Thr Arg Asp Asn Ile Ala Gly 35 40
45 Thr Leu Ala Ala Leu Glu Gln Ala Leu Ala Lys Thr Pro Trp
Ser Met 50 55 60
Ser Asp Val Ser Arg Ile Tyr Leu Asn Glu Ala Ala Pro Val Ile Gly 65
70 75 80 Asp Val Ala Met Glu
Thr Ile Thr Glu Thr Ile Ile Thr Glu Ser Thr 85
90 95 Met Ile Gly His Asn Pro Gln Thr Pro Gly
Gly Val Gly Val Gly Val 100 105
110 Gly Thr Thr Ile Ala Leu Gly Arg Leu Ala Thr Leu Pro Ala Ala
Gln 115 120 125 Tyr
Ala Glu Gly Trp Ile Val Leu Ile Asp Asp Ala Val Asp Phe Leu 130
135 140 Asp Ala Val Trp Trp Leu
Asn Glu Ala Leu Asp Arg Gly Ile Asn Val 145 150
155 160 Val Ala Ala Ile Leu Lys Lys Asp Asp Gly Val
Leu Val Asn Asn Arg 165 170
175 Leu Arg Lys Thr Leu Pro Val Val Asp Glu Val Thr Leu Leu Glu Gln
180 185 190 Val Pro
Glu Gly Val Met Ala Ala Val Glu Val Ala Ala Pro Gly Gln 195
200 205 Val Val Arg Ile Leu Ser Asn
Pro Tyr Gly Ile Ala Thr Phe Phe Gly 210 215
220 Leu Ser Pro Glu Glu Thr Gln Ala Ile Val Pro Ile
Ala Arg Ala Leu 225 230 235
240 Ile Gly Asn Arg Ser Ala Val Val Leu Lys Thr Pro Gln Gly Asp Val
245 250 255 Gln Ser Arg
Val Ile Pro Ala Gly Asn Leu Tyr Ile Ser Gly Glu Lys 260
265 270 Arg Arg Gly Glu Ala Asp Val Ala
Glu Gly Ala Glu Ala Ile Met Gln 275 280
285 Ala Met Ser Ala Cys Ala Pro Val Arg Asp Ile Arg Gly
Glu Pro Gly 290 295 300
Thr His Ala Gly Gly Met Leu Glu Arg Val Arg Lys Val Met Ala Ser 305
310 315 320 Leu Thr Gly His
Glu Met Ser Ala Ile Tyr Ile Gln Asp Leu Leu Ala 325
330 335 Val Asp Thr Phe Ile Pro Arg Lys Val
Gln Gly Gly Met Ala Gly Glu 340 345
350 Cys Ala Met Glu Asn Ala Val Gly Met Ala Ala Met Val Lys
Ala Asp 355 360 365
Arg Leu Gln Met Gln Val Ile Ala Arg Glu Leu Ser Ala Arg Leu Gln 370
375 380 Thr Glu Val Val Val
Gly Gly Val Glu Ala Asn Met Ala Ile Ala Gly 385 390
395 400 Ala Leu Thr Thr Pro Gly Cys Ala Ala Pro
Leu Ala Ile Leu Asp Leu 405 410
415 Gly Ala Gly Ser Thr Asp Ala Ala Ile Val Asn Ala Glu Gly Gln
Ile 420 425 430 Thr
Ala Val His Leu Ala Gly Ala Gly Asn Met Val Ser Leu Leu Ile 435
440 445 Lys Thr Glu Leu Gly Leu
Glu Asp Leu Ser Leu Ala Glu Ala Ile Lys 450 455
460 Lys Tyr Pro Leu Ala Lys Val Glu Ser Leu Phe
Ser Ile Arg His Glu 465 470 475
480 Asn Gly Ala Val Glu Phe Phe Arg Glu Ala Leu Ser Pro Ala Val Phe
485 490 495 Ala Lys
Val Val Tyr Ile Lys Glu Gly Glu Leu Val Pro Ile Asp Asn 500
505 510 Ala Ser Pro Leu Glu Lys Ile
Arg Leu Val Arg Arg Gln Ala Lys Glu 515 520
525 Lys Val Phe Val Thr Asn Cys Leu Arg Ala Leu Arg
Gln Val Ser Pro 530 535 540
Gly Gly Ser Ile Arg Asp Ile Ala Phe Val Val Leu Val Gly Gly Ser 545
550 555 560 Ser Leu Asp
Phe Glu Ile Pro Gln Leu Ile Thr Glu Ala Leu Ser His 565
570 575 Tyr Gly Val Val Ala Gly Gln Gly
Asn Ile Arg Gly Thr Glu Gly Pro 580 585
590 Arg Asn Ala Val Ala Thr Gly Leu Leu Leu Ala Gly Gln
Ala Asn 595 600 605
153354DNAKlebsiella pneumoniae 153atgtcgcttt caccgccagg cgtacgcctg
ttttacgatc cgcgcgggca ccatgccggc 60gccatcaatg agctgtgctg ggggctggag
gagcaggggg tcccctgcca gaccataacc 120tatgacggag gcggtgacgc cgctgcgctg
ggcgccctgg cggccagaag ctcgcccctg 180cgggtgggta tcgggctcag cgcgtccggc
gagatagccc tcactcatgc ccagctgccg 240gcggacgcgc cgctggctac cggacacgtc
accgatagcg acgatcaact gcgtacgctc 300ggcgccaacg ccgggcagct ggttaaagtc
ctgccgttaa gtgagagaaa ctga 354154117PRTKlebsiella pneumoniae
154Met Ser Leu Ser Pro Pro Gly Val Arg Leu Phe Tyr Asp Pro Arg Gly 1
5 10 15 His His Ala Gly
Ala Ile Asn Glu Leu Cys Trp Gly Leu Glu Glu Gln 20
25 30 Gly Val Pro Cys Gln Thr Ile Thr Tyr
Asp Gly Gly Gly Asp Ala Ala 35 40
45 Ala Leu Gly Ala Leu Ala Ala Arg Ser Ser Pro Leu Arg Val
Gly Ile 50 55 60
Gly Leu Ser Ala Ser Gly Glu Ile Ala Leu Thr His Ala Gln Leu Pro 65
70 75 80 Ala Asp Ala Pro Leu
Ala Thr Gly His Val Thr Asp Ser Asp Asp Gln 85
90 95 Leu Arg Thr Leu Gly Ala Asn Ala Gly Gln
Leu Val Lys Val Leu Pro 100 105
110 Leu Ser Glu Arg Asn 115
1551125DNAartificial sequenceCodon optimized amino alcohol kinase from
Erwinia caratovora subsp. atroseptica 155atgagcgatg gccgtctgac
cgcactgttt cctgcatttc cacatccggc atccaaccag 60ccagtgtttg cggaggcttc
cccgcacgac gatgaactga tgacgcaggc ggtgccgcag 120gtttcctgcc agcaagccct
ggcaattgcc cagcaggaat atggcctgag cggtcagatg 180agcctgctgc agggcgaacg
tgacgttaat ttctgtctga ccgtaacgcc agatgaacgc 240tatatgctga aagtcatcaa
cgctgctgaa ccggcagatg tgagcaactt tcagactgcg 300ctgctgctgc acctggcacg
tcaggcgcca gaactgccag tccctcgtat ccgctccacg 360aaggctggtc agtctgaaac
gggcgtcgaa attgatggtg ttctgctgcg tgtgcgtctg 420gtttcctacc tggctggcat
gccgcagtac ctggcgtctc cgagcacggc actgatgcca 480cagctgggcg gtactctggc
gcagctggac aacgctctgc actctttcac ccatccggcg 540gctaaccgtg ctctgctgtg
ggacatctcc cgcgcagagc aggtccgccc gtacctggac 600ttcgttagcg agccgcagca
gtatcagcac ctgcagcgca tctttgatcg ctatgactct 660aacgtggcac cgctgctgac
gacgctgcgc cgccaggtta tccacaacga cctgaacccg 720cataacgtcc tggtcgatgg
ttccagcccg acgcgcgtca cgggtatcat cgacttcggc 780gatgcagtgt tcgcgccgct
gatctgtgag gttgcgaccg ctctggcgta ccaaattggc 840gacggcacgg atctgctgga
acatgtggta ccgtttgtcg cagcgtatca ccagcgtatt 900ccgctggcgc cggaggaaat
cgccctgctg ccagatctga tcgcgacccg catggcactg 960actctgacca tcgctcagtg
gcgtgcgtct cgctacccag ataaccgcga atacctgctg 1020cgcaacgtgc cgcgctgctg
gcactccctg cagcgtatcg caacttacag ccacgcacaa 1080tttctgacgc gcctgcagca
ggtttgccca gaaaacgctc gttga 11251561275DNAartificial
sequenceCodon optimized amino alcohol O-phosphate lyase from Erwinia
caratovora subsp. atroseptica 156atgactgcaa ctgaagctct gctggcacgt
cgtcagcgcg ttctgggcgg tggctaccgt 60ctgttctacg aagaaccgct gcatgttgca
cgcggcgaag gtgtatggct gttcgatcat 120cagggtaaac gttacctgga cgtatataac
aacgtagcta gcgtaggtca ctgtcacccg 180gccgttgtag aagcggtcgc gcgtcaatct
gcgcaactga acacccatac gcgctacctg 240catcacgcga tcgtagattt tgctgaagat
ctgctgtctg agttcccggc agaactgaac 300aacgtcatgc tgacctgtac tggctccgaa
gcgaacgacc tggccctgcg cattgcgcgt 360cacgttacgg gtggtaccgg catgctggtg
acccgttggg cctaccatgg tgttacgtcc 420gctctggcgg agctgtcccc gtccctgggc
gacggcgtag tacgcggttc ccacgtaaag 480ctgatcgatg ctccggatac ctaccgtcag
ccgggtgctt tcctgacctc tatccgcgaa 540gcgctggcac agatgcagcg tgaaggtatt
cgtccggcgg ctctgctggt tgatactatc 600ttctcctccg acggtgtatt ctgtgcgccg
gaaggtgaga tggcccaggc agccgcactg 660atccgtcagg ccggtggcct gttcattgcg
gacgaagtgc agccgggctt tggtcgtacc 720ggtgaatccc tgtggggttt cgcacgtcat
aacgtggttc cagatctggt ttctctgggc 780aaaccgatgg gtaacggcca tccgattgct
ggtctggtag gtcgctccgc actgttcgac 840gcttttggtc gtgatgttcg ctactttaat
actttcggcg gtaacccagt atcctgccag 900gcggcacatg ctgttctgcg cgttatccgt
gaagaacagc tgcagcagaa cgcgcagcgt 960gttggtgatt atctgcgcca aggtctgcag
cagctggcac aacacttccc gctgatcggt 1020gacattcgtg catatggtct gtttatcggt
gctgaactgg tttccgaccg tgaatccaaa 1080accccagcga gcgagtctgc actgcaggtt
gttaacgcga tgcgtcagcg tggtgtactg 1140atctccgcaa ccggcccggc ggcgaacatt
ctgaagatcc gtcctccgct ggtattcctg 1200gaggaacacg cggacgtgtt cctgactacc
ctgtccgacg tgctggcgct gatcggtact 1260cgtgcacagc gttaa
127515777DNAartificial sequencePrimer
157caggaggaat taaccatggg gggttctcat catcatcatc atcatggtga cgatgacgat
60aagatgagcg atggccg
7715877DNAartificial sequencePrimer 158cggccatcgc tcatcttatc gtcatcgtca
ccatgatgat gatgatgatg agaacccccc 60atggttaatt cctcctg
7715920DNAartificial sequencePrimer
159ggacctgctt cgctttatcg
2016015DNAartificial sequencePrimer 160gctagagatg atagc
1516118DNAartificial sequencePrimer
161ggaagagact atccagcg
1816250DNAartificial sequencePrimer 162gcgcgcccgg gaagaaggag ctcttcacca
tgaacaaacc acagtcttgg 5016328DNAartificial sequencePrimer
163gcgcgcccgg gttcatgcca cctctgcg
281642432DNAErwinia caratovora subsp. atroseptica 164atgtctgacg
gacgactcac cgcacttttt cctgcattcc cacacccggc gtccaatcag 60cccgtatttg
ccgaggcttc accgcacgac gacgagttaa tgacgcaggc cgtaccgcag 120gtttcctgtc
agcaggcgtt ggcgattgcg cagcaagaat atggcttgtc tgggcagatg 180tcgctgcttc
agggcgagcg tgatgtgaat ttctgtctga cggtgacgcc agatgaacgc 240tacatgctga
aagtcatcaa tgcggcagaa cctgccgacg tcagcaattt ccaaaccgcg 300ctgctgctgc
atcttgcccg tcaggcacct gaactgcccg taccgcgtat caggtcgaca 360aaagcgggtc
agtcggaaac aggcgttgag atcgatggtg tactgctgcg tgtgcggctt 420gtgagctatc
tggcaggaat gccgcagtat ctggcctcac cgtcaacggc gctgatgccg 480cagttggggg
gaacgctggc gcagttggat aacgcgcttc acagctttac gcatccggcg 540gcaaaccgtg
cgctgctgtg ggatatcagc cgggcagagc aggtgcgtcc ttacctcgat 600ttcgtttctg
aaccgcagca gtatcagcat cttcagcgta tttttgaccg ttatgacagt 660aacgttgctc
ctctgttgac gacgctacgt cgtcaggtca ttcataacga tctgaatccg 720cataacgtgc
tggtggatgg atcgtcgccg acgcgggtta ctggcattat cgattttggc 780gatgccgtat
ttgccccgtt aatttgcgaa gtcgcgacgg cactggcgta tcagatcggc 840gatggaaccg
atttgttgga gcatgttgtg ccgtttgttg cggcctatca ccaacgcatt 900ccgttagcac
cggaggagat tgcgctgtta cccgatctga tagcgacccg tatggcgctg 960accctgacca
ttgcgcagtg gcgagcatcg cgttatcccg acaatcggga gtatctgctg 1020cgtaacgtgc
cgcgctgttg gcacagtttg cagcgcattg cgacctattc ccatgcgcaa 1080tttttgactc
gcctacagca ggtttgcccg gagaatgcgc gatgaaccag aaaggaatga 1140cgtctatgac
gtctgaaatg acagcgacag aagctttgct ggcgcgccgt cagcgagtgt 1200tgggcggcgg
ttatcgcctg ttttatgaag agccgctgca tgtcgcgcgc ggcgagggcg 1260tgtggctgtt
cgatcaccaa gggaaacgtt atctggatgt ctacaataat gtggcttcgg 1320tcggacattg
ccaccccgcg gtggttgaag ccgtggcgcg acagagcgca caactcaata 1380cccacacgcg
ctatttgcac cacgcgattg tcgattttgc ggaagatttg ctgagcgaat 1440ttcccgccga
attgaacaat gtaatgctga cctgtaccgg cagtgaggct aacgatctgg 1500cgctgcgtat
cgcccgacat gtcacgggcg ggacggggat gttggtgacg cgctgggcgt 1560atcacggcgt
gaccagcgcg ctggcggaac tgtctccgtc gctgggggat ggcgttgtgc 1620gcggtagcca
tgtgaagctg atcgacgcgc cagacactta tcgtcagccc ggtgcatttc 1680ttaccagcat
tcgtgaagcg ctggcgcaga tgcaacggga aggtattcgt cctgcggcgc 1740tgctggtaga
taccattttt tccagcgatg gcgtgttctg tgcgccggaa ggcgaaatgg 1800cacaggcggc
ggcgttgatc cgtcaggcgg gcgggctgtt tattgcggat gaagtgcagc 1860cgggcttcgg
gcgcaccggg gaatcactgt ggggctttgc gcgccacaat gtcgtccctg 1920atttggtgag
tctagggaaa ccgatgggca acggacatcc catcgctgga ttggtggggc 1980gttccgctct
gttcgacgca tttgggcgcg atgtgcgcta tttcaatacc tttggcggca 2040atccggtttc
ctgtcaggcg gcgcacgcgg tgctgcgggt gattcgggaa gagcagttgc 2100agcagaatgc
ccagcgggtc ggtgattatc tgcggcaagg gttgcagcaa ctggcgcagc 2160atttcccgct
gattggtgat attcgggctt acggcctgtt tattggtgcg gagctggtca 2220gcgatcgcga
aagtaaaacg ccggcaagtg aatccgcgtt gcaggtggtg aatgcgatgc 2280gccaacgtgg
tgtgctcatc agcgcgacgg ggccagcggc gaacatactg aaaattcgcc 2340cgccgctggt
gtttctggaa gaacacgccg atgtgttctt aaccacgctg agtgacgttt 2400tagcgctcat
cggcactcgt gcacagagat aa 2432
User Contributions:
Comment about this patent or add new information about this topic: