Patent application title: Novel Centromeric Protein Shugoshin
Inventors:
Yoshinori Watanabe (Tokyo, JP)
Assignees:
Japan Science & Technology Agency
IPC8 Class: AC07K1600FI
USPC Class:
5303879
Class name: Globulins immunoglobulin, antibody, or fragment thereof, other than immunoglobulin antibody, or fragment thereof that is conjugated or adsorbed binds specifically-identified amino acid sequence
Publication date: 2009-08-13
Patent application number: 20090203887
Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
Patent application title: Novel Centromeric Protein Shugoshin
Inventors:
Yoshinori Watanabe
Agents:
Locke Lord Bissell & Liddell LLP;Attn: IP Docketing
Assignees:
Japan Science & Technology Agency
Origin: NEW YORK, NY US
IPC8 Class: AC07K1600FI
USPC Class:
5303879
Abstract:
The present invention is to provide meiosis-specific novel kinetochore
protein Sgo1 (shugoshin) derived from fission yeast Schizosaccharomyces
pombe, and a homologue or paralogue thereof having a regulatory activity
of chromosome segregation; and DNAs encoding them; as a factor ensuring
the retention of unidirection and cohesion in sister centromere at
meiosis I in cooperation with cohesin. To elucidate the proteins
protecting Rec8 during anaphase, the present inventor screened in fission
yeast genes for a gene that inhibits mitotic growth and prevents sister
chromatid from the separation at anaphase, when co-expressed with Rec8.
In this approach, meiosis-specific protein Sgo1 that protects (Shugo)
centromeric Rec8 from the degradation at anaphase I was indentified.
Further, a budding yeast Sgo1 homologue and a fission yeast mitotic
paralogue Sgo2 were identified.Claims:
1. An isolated DNA encoding a protein consisting of an amino acid sequence
shown in SEQ ID NO: 18.
2. An isolated DNA encoding a protein comprising an amino acid sequence where one or several amino acids are deleted, replaced or added in an amino acid sequence shown in SEQ ID NO: 18, and having a regulatory activity of chromosome segregation.
3. An isolated DNA consisting of a base sequence shown in SEQ ID NO: 17 or a complementary sequence thereof.
4. An isolated DNA containing part or whole of a base sequence shown in SEQ ID NO: 17 or a complementary sequence thereof, and encoding a protein that has a regulatory activity of chromosome segregation.
5. An isolated DNA hybridizing with the DNA according to claim 3 under stringent conditions and encoding a protein that has a regulatory activity of chromosome segregation.
6. An isolated protein consisting of an amino acid sequence shown in SEQ ID NO: 18.
7. An isolated protein consisting of an amino acid sequence where one or several amino acids are deleted, replaced or added in an amino acid sequence shown in SEQ ID NO: 18, and having a regulatory activity of chromosome segregation.
8. An isolated protein encoded by a DNA hybridizing with the antisense strand of a DNA encoding the protein according to claim 6 under stringent conditions.
9. A fusion protein in which the protein according to claim 6 is bound with a marker protein and/or a peptide tag.
10. A fusion protein in which the protein according to claim 7 is bound with a marker protein and/or a peptide tag.
11. A fusion protein in which the protein according to claim 8 is bound with a marker protein and/or a peptide tag.
12. An antibody specifically binding to the protein according to claim 6.
13. An antibody specifically binding to the protein according to claim 7.
14. An antibody specifically binding to the protein according to claim 8.
15. The antibody according to claim 12, which is a monoclonal antibody.
16. The antibody according to claim 13, which is a monoclonal antibody.
17. The antibody according to claim 14, which is a monoclonal antibody.
Description:
[0001]This application is a divisional application of U.S. patent
application Ser. No. 10/581,158 filed Jan. 30, 2007, which is national
phase entry of International Application No. PCT/JP2004/017428 filed on
Nov. 24, 2004, which claims priority benefit of Japanese Application No.
JP 2003-401943 filed Dec. 1, 2003 and Japanese Application No. JP
2004-279450 filed Sep. 27, 2004, the contents of each of which are
incorporated in their entireties.
TECHNICAL FIELD
[0002]The present invention relates to a protector protein Sgo1 (shugoshin) of cohesin Rec8 derived from fission yeast Schizosaccharomyces pombe, its homologue and paralogue having a regulatory activity of chromosome segregation, and DNAs encoding them.
BACKGROUND OF THE INVENTION
[0003]In eukaryotes, sister chromatid cohesion is established during S phase of cell cycle and. maintained throughout G2 until M phase. During mitosis, this cohesion is destroyed along the entire length of chromosome, allowing sister chromatid to segregate to the opposite sides of cell (equational division) and ensuring that each daughter cell receives one copy of each chromosome. In contrast, meiosis consists of two rounds of chromosome segregation following a single round of DNA replication, leading to the formation of four haploid gametes from one diploid germ cell. During meiosis I, homologous chromosomes (homologues) pair to recombine, forming chiasmata in which one sister chromatid from one homologue is covalently attached to a sister chromatid from the other homologue. Hence, in order for homologues to segregate at meiosis I, cohesion of sister chromatid is necessary to be dissociated along the chromosome arms to resolve chiasmata. However, sister chromatid cohesion is retained at centromere until meiosis II, and utilizes the residual centromeric cohesion when sister chromatid segregates, in the same manner as it does in mitosis. Thus, meiotic division requires sister chromatid cohesion to be dissociated in two steps. However, the molecular mechanism for protection of centromeric cohesion only during meiosis I and only at the centromere has remained to be elucidated (e.g., see Annu Rev Genet 35, 673-745 (2001)).
[0004]There are important clues as to the molecular nature of sister chromatid cohesion, and the mechanism dissociating sister chromatid cohesion at the onset of anaphase (e.g., see Annu Rev Genet 35, 673-745 (2001); Curr Opin Cell Biol 12, 297-301 (2000); Curr Biol 13, R104-14 (2003); Annu Rev Cell Dev Biol 17, 753-77 (2001); Genes Dev 16, 399-414 (2002>>. In various eukaryotes, sister chromatid cohesion depends on a multisubunit cohesin complex including Scc1 (Rad21 in fission yeast Schizosaccharomyces pombe). Anaphase promoting complex (APC)-dependent degradation of the securin, Cut2/Pds1, allows to dissociate the Cut1/Esp1 endopeptidase (separase), which in turn cleaves Rad21/Scc1, dissociating sister chromatid cohesion. During meiosis, the cohesion subunit Rad21/Scc1 is replaced with a meiotic counterpart, Rec8 (e.g., see Cell 98, 91-103 (1999); Mol. Cell. Biol. 19, 3515-3528 (1999); Nature 400, 461-4 (1999); Genes Dev 15, 1349-60 (2001); J Cell Biol 160, 657-70 (2003)). As Rec8 complex resides only at centromere after meiosis I and the depletion of Rec8 destroys centromeric cohesion, the presence of Rec8 at centromere has been thought to confer the persistence of cohesion throughout meiosis I (e.g., see Nat Cell Biol 1, E125-7 (1999)). Several lines of evidence suggest that Rec8 along chromosome arms is cleaved by separase at anaphase I while centromeric Rec8 is specifically protected until metaphase II (e.g., see Cell 103, 387-98 (2000); Embo J 22, 5643-53 (2003)). Budding yeast SP013 has been implicated in the protection of centromeric Rec8 (e.g., see Genes Dev 16, 1659-71 (2002); Genes Dev 16, 1672-81 (2002)), but SP013 is not centromeric and may function indirectly. Drosophila MEI-S332 is a protein residing at centromere, is required for the persistence of centromeric cohesion during meiosis I, and has features of a candidate protector of meiotic centromeric cohesion, although the details of such protection have so far not been elucidated (e.g., see Annu Rev Cell Dev Biol 17, 753-77 (2001); Cell 83, 247-256 (1995)). Despite the completion of genome sequencing projects on several organisms, no homologue of these proteins has emerged, preventing the formulation of .a generalized view of the protection. Concurrently, studies in fission yeast have illuminated the importance of pericentromeric heterochromatin for recruiting centromeric Rec8 complexes and ensuring centromeric cohesion during meiosis I (e.g., see Science 300, 1152-5 (2003)). However, pericentromeric heterochromatin cannot alone confer the specific protection of Rec8 at meiosis I toward meiosis II.
DISCLOSURE OF THE INVENTION
[0005]Almost all the eukaryotes including human propagate offsprings by sexual reproduction evolutionarily predominant with a mixture of genome. Meiosis that reduces chromosome number in half is a core part of the sexual reproduction mechanism. In somatic mitosis, two kinetochores of sister chromatid are caught by spindle microtuble extended from the opposite poles, and sister chromatid is evenly segregated to the both poles by concurrently dissolving the cohesion of arms and centromeres (equational division). In contrast, in meiosis I kinetochores of sister chromatids are caught by spindle microtuble extended from the same pole, and segregated to the same pole while retaining the cohesion at centromere (meiotic division). Next, for the first time in meiosis II the cohesion of centromere site of sister chromatid is dissolved, and separated toward one pole or the other of the two poles respectively, which culminates in the generation of accurate four haploid gametes. Meiosis-specific meiotic division is a modality of chromosome segregation conserved in almost all the eukaryotes, from yeast to human, however regulatory mechanism at the molecular level has remained enigmatic for a long time. The present inventor has demonstrated that meiosis-specific chromosome cohesion factor, cohesin plays an essential role in this regulation by using fission yeast (Nature 400, 461-4 (1999); Science 300, 1152-5 (2003); Nature 409, 359-363 (2001)). An object of the present invention is to provide meiosis-specific novel kinetochore protein Sgo1 (shugoshin) derived from fission yeast Schizosaccharomyces pombe, and a homologue or paralogue thereof having a regulatory activity of chromosome segregation; and DNAs encoding them; as a factor ensuring the retention of unidirection and cohesion in sister centromere at meiosis I in cooperation with cohesin.
[0006]Meiosis comprises two steps of specialized nuclear divisions for producing haploid gametes. To accomplish this, sister chromatid cohesion is necessary to be dissociated in a stepwise manner, first from chromosome arms at anaphase I and second from centromeres at anaphase II. In particular, the factors that protect centromeric cohesion during meiosis I have heretofore remained undissolved. To elucidate the proteins protecting Rec8 during anaphase, the present inventor screened in fission yeast genes for a gene that inhibits mitotic growth and prevents sister chromatid from the separation at anaphase, when co-expressed with Rec8. In this approach, meiosis-specific protein that is a protector of Rec8 in fission yeast and protects (Shugo) centromeric Rec8 from the degradation at anaphase I was indentified, and named Sgo1 (Shugoshin, a Japanese for "guardian spirit"). It was also identified that shugoshin plays an important role in mitotic chromosome segregation. and then identified a budding yeast Sg01 homologue and a fission yeast mitotic paralogue Sgo2. A marginal similarity between Sgo1 and Drosophila MEI-S332 was identified. and Sgo1 homologue in other eukaryotes was also identified. Shugoshin-like proteins in animal cells, which were predicted from the sequence, also have functional conservation with yeast shugoshin. The present invention has been thus completed based on this knowledge.
[0007]That is, the present invention relates to (1) a DNA encoding a following protein (a) or (b): (a) a protein consisting of an amino acid sequence shown in SEQ ID NO: 2, (b) a protein comprising an amino acid sequence where one or several amino acids are deleted. replaced or added in an amino acid sequence shown in SEQ ID NO: 2, and having a regulatory activity of chromosome segregation; (2) a DNA consisting of a base sequence shown in SEQ ID NO: 1 or a complementary sequence thereof; (3) a DNA containing part or whole of a base sequence shown in SEQ ID NO: 1 or a complementary sequence thereof, and encoding a protein that has a regulatory activity of chromosome segregation; (4) a DNA hybridizing with the DNA according to "2" under stringent conditions and encoding a protein that has a regulatory activity of chromosome segregation; (5) a protein consisting of an amino acid sequence shown in SEQ ID NO: 2; and (6) a protein consisting of an amino acid sequence where one or several amino acids are deleted, replaced or added in an amino acid sequence shown in SEQ ID NO: 2, and having a regulatory activity of chromosome segregation.
[0008]The present invention also relates to (7) a DNA encoding a following protein (a) or (b): (a) a protein consisting of an amino acid sequence shown in SEQ ID NO: 4, (b) a protein consisting of an amino acid sequence where one or several amino acids are deleted, replaced or added in an amino acid sequence shown in SEQ ID NO: 4, and having a regulatory activity of chromosome segregation; (8) a DNA consisting of a base sequence shown in SEQ ID NO: 3 or a complementary sequence thereof; (9) a DNA containing part or whole of a base sequence shown in SEQ ID NO: 3 or a complementary sequence thereof, and encoding a protein that has a regulatory activity of chromosome segregation; (10) a DNA hybridizing with the DNA according to "8" under stringent conditions and encoding a protein that has a regulatory activity of chromosome segregation; (11) a protein consisting of an amino acid sequence shown in SEQ ID NO: 4; and (12) a protein consisting of an amino acid sequence where one or several amino acids are deleted, replaced or added in an amino acid sequence shown in SEQ 10 NO: 4, and having a regulatory activity of chromosome segregation.
[0009]The present invention further relates to (13) a DNA encoding a following protein (a) or (b): (a) a protein consisting of an amino acid sequence shown in SEQ ID NO: 6, (b) a protein consisting of an amino acid sequence where one or several amino acids are deleted, replaced or added in an amino acid sequence shown in SEQ ID NO: 6, and having a regulatory activity of chromosome segregation; (14) a DNA consisting of a base sequence shown in SEQ ID NO: 5 or a complementary sequence thereof; (15) a DNA containing part or whole of a base sequence shown in SEQ ID NO: 5 or a complementary sequence thereof, and encoding a protein that has a regulatory activity of chromosome segregation; (16) a DNA hybridizing with the DNA according to "14" under stringent conditions and encoding a protein that has a regulatory activity of chromosome segregation; (17) a protein consisting of an amino acid sequence shown in SEQ ID NO: 6; and (18) a protein consisting of an amino acid sequence where one or several amino acids are deleted, replaced or added in an amino acid sequence shown in SEQ ID NO: 6, and having a regulatory activity of chromosome segregation.
[0010]The present invention still further relates to (19) a DNA encoding a following protein (a) or (b) that has a regulatory activity of chromosome segregation: (a) a protein consisting of an amino acid sequence shown in SEQ ID NO: 8, 10, 12, 14, 16, 18 or 20, (b) a protein consisting of an amino acid sequence where one or several amino acids are deleted, replaced or added in an amino acid sequence shown in SEQ ID NO: 8, 10, 12, 14, 16, 18 or 20; (20) a DNA consisting of a base sequence shown in SEQ ID NO: 7, 9, 11, 13, 15, 17 or 19 or a complementary sequence thereof, and encoding a protein that has a regulatory activity of chromosome segregation; (21) a DNA containing part or whole of a base sequence shown in SEQ ID NO: 7, 9, 11, 13, 15, 17 or 19 or a complementary sequence thereof, and encoding a protein that has a regulatory activity of chromosome segregation; (22) a DNA hybridizing with the DNA according to "7", "9", "11", "13", "15", "17" or "19" under stringent conditions and encoding a protein that has a regulatory activity of chromosome segregation; (23) a protein consisting of an amino acid sequence shown in SEQ ID NO: 8, 10, 12, 14, 16, 18 or 20, and having a regulatory activity of chromosome segregation; and (24) a protein consisting of an amino acid sequence where one or several amino acids are deleted, replaced or added in an amino amino acid sequence shown in SEQ ID NO: 8, 10, 12, 14, 16, 18 or 20, and having a regulatory activity of chromosome segregation.
[0011]Furthermore, the present invention relates to (25) a fusion protein in which the protein according to "5", "6", "11", "12", "23" or "24" is bound with a marker protein and/or a peptide tag; (26) an antibody specifically binding to the protein according to "5". "6", "11", "12", "23" or "24"; and (27) the antibody according to "26", which is a monoclonal antibody.
BRIEF DESCRIPTION OF DRAWINGS
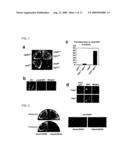
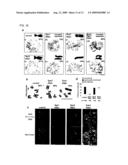
[0012]FIG. 1 is a set of pictures showing that sister chromatids are not segregated during mitosis by co-expression of Sgo1 and Rec8 in the present invention. a.) The cen2-GFP strains expressing the genes indicated by endogenous promoters (a constitutive chromatin promoter for rad21+ or rec8+, and a thiamine-repressible promoter Pnmt1 for Sgo1+) were streaked on a thiamine-depleted plate. b.) Samples of Padh1-rec8+Pnmt1-sgo1+ cells cultured for 15 hours at 30° C. after thiamine depletion. The non-segregation of cen2-GFP (asterisk) was identified in the septate junction cells. c.) The non-segregations of cen2-GFP were counted (n>100). d.) The Padh1-rec8+-GFP strains were cultured with or without the use of Pnmt1-sgo1+ in the same manner as (b). Samples of cells at interphase and anaphase are shown.
[0013]FIG. 2 is a set of pictures showing that sister chromatid segregation was undergone in mitosis by expression of non-cleavable Rec8. The plasmid pREP41-rec8-RDRD (expressing non-cleavable Rec8 (Embo J 22, 5643-53 (2003))) was integrated into the chromosome of cen2-GFP cell strains (+Rec8-RDRD), and the cells were streaked on plates with or without the presence of thiamine. The host strain cells (-Rec8-RDRD) were similarly cultured as a control. Note that Rec8-RDRD is expressed only on the thiamine-free plate. Samples of cells cultured in culture medium for 15 hours at 30° C. after the depletion of thiamine.
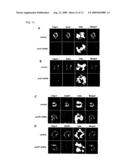
[0014]FIG. 3 is a set of pictures showing that sgo1 of the present invention is required to protect Rec8 and thereby cohesion at centromeres arises during anaphase of meiosis I. a.) As for one of the homologues marked with cen2-GFP, segregation during meiosis was observed in wild-type and sgo1Δ cells (n>170). A normal segregation pattern of cen2-GFP is illustrated (left). Samples of sgo1Δ cells are shown (right). b.) Separation of sister cen2-GFP dots after meiosis I (mes1Δarrest) is evident in sgo1Δ cells. c.) The Rec8-GFP signal was observed in the indicated cells at late anaphase I (n>30) and at prometaphase II (n>100), and the frequency of centromeric Rec8-GFP displayed in the cells was counted. The spindles were visualized by expressing CFP-Atb2 (a2-tubulin) (Curr Biol 11, 836-45 (2001)). d.) Rec8-GFP levels throughout the indicated chromosome sites in the arrested cells were measured prior to meiosis I (mei4Δ arrest) by ChIP assay with the use of anti-GFP antibodies. The bottom panel shows Schizosaccharomyces pombe chromosome I schematically, and the primers (cnt, imr, dg, dh, lys1, mes1) were used there.
[0015]FIG. 4 is a set of pictures showing that Sgo1 of the present invention localizes at pericentromeric regions during meiosis I. a.) Synchronous meiosis of diploid pat1-114/pat1-114 cell strains (Embo J 22, 5643-53 (2003)) was sampled, meiotic nuclear division was monitored by DAPI staining, and the protein level of Sgo1 was detected by Western blotting with the use of anti-Sgo1 antibodies. b.) Sgo1 (green) was counterstained with tubulin (red) and DAPI (4'6'-diamidino-2-phenylindole) (blue) at the indicated stages in meiotic cells. c.) A sgo1+-GFP cell co-expressing mis6+-CFP was examined under fluorescence microscopy. Sgo1-GFP (green) and Mis6-CFP (red) are merged. d.) Sgo1-GFP levels throughout the indicated chromosome sites in cells arrested at metaphase I were measured by ChIP assay with the use of anti-GFP antibodies. The same primers as for FIG. 2d in synchronism with additional primers at mat (heterochromatin region at the mating type locus) and TAS (telomere associated sequence) were used. e.) Sgo1-GFP (green) was detected at metaphase I in the indicated cells that express CFP Atb2 to visualize spindles (red). f.) Rec8-HA was expressed with or without Sgo1-FLAG in proliferating cells, and the extracts were immunoprecipitated with anti-FLAG antibody. g.) A model for the action of shugoshin in meiosis. Shugoshin protects centromeric Rec8 complexes from cleaving by separase at the onset of anaphase I, thereby preserves the centromeric cohesion until meiosis II. Shugoshin is degraded depending on APC during anaphase II.
[0016]FIG. 5 is a set of pictures showing the time-dependent change of the expression levels of Sgo1 and Rec8 in synchronous culture of haploid pat1-114 cell strains (wt), and of cut1-206 or Prad21-slp1 cells. The expression of slp1+ (a fission yeast CDC20 homologue required for APC activation (Mol Cell Biol 17, 742-50 (1997))) was repressed during meiosis in Prad21-slp1 cells where slp1 promoter was replaced with rad21. Meiotic nuclear division was monitored by DAPI staining, and the protein levels of Sgo1, Rec8, and tubulin (control) were measured by western blotting with the use of anti-Sgo1, anti-Rec8 and anti-tubulin antibodies, respectively. Although cut1-206 cells together with normal kinetics led to Sgo1 degradation, Rec8 degradation was delayed. Prad21-slp1 cells showed delayed degradation of Sgo1 as well as Rec8. Arrowheads indicate a cleavage product of Rec8 by separase Cut1.
[0017]FIG. 6 is a set of pictures showing that ectopic expression of sgo1+ inhibits the growth of the cut1-206 mutant. Chromosomal sgo1+ promoter was replaced with Pnmt1 or Pnmt41 (a weaker version of Pnmt1), and the effect on the mitotic growth in cut1-206 temperature-sensitive cells was examined. The indicated cells were streaked on a plate without thiamine and cultured for 3 days at 28° C. The cut1-206 cells moderately expressing Sgo1 by Pnmt1, arrested mitotic growth even at the permissive temperature, whereas cut1+ cells grew normally.
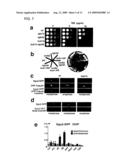
[0018]FIG. 7 is a set of pictures showing that Sgo2 of the present invention plays an important role in mitotic at centromere. a.) Serial dilutions of the indicated cultures were spotted onto YEA plates containing 0, 5 or 10 μg/ml of TBZ, and cultured for 3 days at 30° C. b.) The indicated strains were streaked on YEA plates and cultured for 3 days at 30° C. c.) Sgo2-GFP (green) was detected at anaphase I in wild-types and in bub1Δ cells that express CFP-Atb2 to visualize spindles (red). DNA was stained with Hoechst (blue). Wild-type cells at anaphase are also shown. d.) The sgo2+-GFP mis6+-HA cells were fixed and stained with anti-GFP and anti-HA antibodies. e.) Sgo2-GFP levels were measured throughout the indicated chromosome sites in cells arrested at prometaphase or in asynchronous cells by ChIP assay.
[0019]FIG. 8 is a set of pictures showing the results of analysis of budding yeast shugoshin ScSgo1 of the present invention. a.) Budding yeast ScSGO1-GFP diploids in proliferation were fixed with methanol and counterstained with DAPI. b.) ScSGO1-Myc NDC10-HA cells were fixed, and stained with DAPI and antibodies against Myc and HA. c.) ScSGO1-GFP diploids causing meiosis in culture medium were fixed with methanol and counterstained with DAPI. d.) Serial dilutions of the indicated cultures were spotted onto YPD plates containing 0 or 15 μg/ml of benomyl. e.) Chromosome loss was analyzed in wild-types (wt) and Scsgo1Δ mutants by a colony sectoring assay. The loss of nonessential chromosome fragments resulted in a red sector in a white colony. As a positive control, ubr1Δ mutant was used (Nature 410, 955-9 (2001)). The frequency of sectoring colonies is shown at the bottom (n>120). f.) Samples of segregation of cenV-GFP in Scsgo1Δ tetrads. The segregation patterns in tetrads were mostly classified as one of the three shown at the bottom. The each population (n=200) is also shown. g.) ScSGO1-Myc diploids were induced by synchronous meiosis and were examined the segregation of cenV-GFP marked on one of two homologues at meiosis I and meiosis II. Although most of the cells caused reductional segregation pattern at meiosis I (96%, n=207), the incidence of non-segregation was high at meiosis II (34%, n=322). h.) The cells marked with cenV-GFP on both homologues were induced to meiosis, and counterstained with anti-tubulin antibody and DAPI. Cells at late anaphase I were examined for cenV-GFP dots. ScSGO1-Myc cells frequently showed split cenV-GFP dots at either pair of sister chromatids (72%, n=138), while control wild-type cells did not (<2%, n=106).
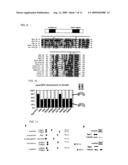
[0020]FIG. 9 is a set of pictures showing sequences of the amino terminal coiled-coil regions and carboxyl terminal basic regions of shugoshin-like proteins in various organisms. The primary sequences of the amino terminal regions of Sgo1 are conserved in Schizosaccharomyces pombe (Sgo1 and Sgo2), budding yeast (ScSgo1) and Neurospora crassa (B23G1.060), while the sequences containing ME1-S332 in other species are not conserved, all presumably carry coiled-coil motif (predicted by COILS program (Science 252, 1162-4 (1991))). See the arrowheads, asterisks and circles in the pictures. The sequences in FIG. 9 respectively correspond to the following SEQ ID NOs: Sg01_Sp18: SEQ ID NO: 21; Sg02_Sp10: SEQ ID NO: 22; Sg01_Sc40: SEQ ID NO:23; B23GI.060_Nc19: SEQ ID NO: 24; Mei-S332_Dm2: SEQ ID NO: 25; Sg01_Sp277: SEQ ID NO: 26; Sg01_Sp569: SEQ ID NO: 27; Sg01_Sc364: SEQ ID NO: 28; B23GI.060_Nc464: SEQ ID NO: 29; Mei-S332_Dm367: SEQ ID NO: 30; C33H5.15_Ce: SEQ ID NO: 31; AT3G10440.1_At: SEQ ID NO: 32; AT5G04320.1_At: SEQ ID NO: 33; BAB29295.1_Mm: SEQ ID NO: 34; Tripin_Mm: SEQ ID NO:35; Q9BVA8_Hs: SEQ ID NO: 36; Tripin_Hs: SEQ ID NO: 37.
[0021]FIG. 10 is a picture showing the results of examination of sgo1 mutations that were generated within conserved regions. Both h+sgo1Δ and h-sgo1Δcen2-GFP cells transformed with the indicated plasmid, were mixed on SPA plates and monitored for segregation of cen2-GFP at miosis II. A plasmid pREP81 bearing a weak version of the thiamine-repressible nmt1 promoter was used to express sgo1. Control cells carrying plasmid pREP81-sgo1 (wt) showed nearly 80% the segregation at meiosis II, whereas cells expressing non-segregation sgo1 allele showed random segregation (50% segregation). Any of the mutations tested, except a non-conserved site mutation 297TA, did not complement sgo1Δ in this assay. The means of two independent experiments are shown (n>100).
[0022]FIG. 11 (a) is a picture showing schematic representation of the shugoshin family proteins. A predicted coiled-coil (red) and a conserved basic region (blue) exist in the N-terminal and C-terminal regions respectively. Further, FIG. 11 (b) is a picture showing the result of analysis in HeLa cell extracts by western blotting after transfection with siRNA.
[0023]FIG. 12 is a set of pictures showing the results that HeLa cells were stained (green) with antibody against hSgo1 or hSgo2 prepared from rabbit, concurrently stained with tubulin antibody and DAPI, and then respectively co-stained with spindle (red) and chromosome DNA (blue). Meanwhile, the cells were fixed with paraformaldehyde.
[0024]FIG. 13 is a set of pictures showing the results that HeLa cells at prometaphase and metaphase were stained with antibodies against hsgo1 or hSgo2 (green), and concurrently co-stained with antibodies against centromere protein CENP-A (a, c; red), antibodies against passenger protein Aurora B of chromosome localized within kinetochore from prophase to metaphase (b, d; red), and DAPI (blue). Both signals of hSgo1 and hSgo2 showed signals at the sites close to CENP-A dots on chromosome. From the above, it was revealed that both hsgo1 and hSgo2 are centromere proteins. Furthermore, both sites of Sgo1 and Aurora B were practically the same at prometaphase and metaphase, whereas Sgo2 was placed just outside Aurora B. From the above, it was revealed that both hsgo1 and hSgo2 are placed within kinetochore from prometaphase to metaphase.
[0025]FIG. 14 is a picture showing the results of RNAi experiments that targeted hsgo1 and hSgo2 respectively. The expressions in any proteins were significantly suppressed after 48 hours, thereby the cells arrested in mitosis (total in the figure) were accumulated. As the accumulation was dissolved by suppressing a spindle checkpoint factor BubR1 by RNAi, it was suggested that hSgo1 and hSgo2 directly or indirectly function during the process where spindle take kinetochore properly at centromeres.
[0026]FIG. 15 is a set of pictures showing the results, where RNAi experiments targeting hsgo1 was performed by using HeLa cells, and then the cells were mounted on a slide glass and stained with Giemsa. It was revealed that sister chromatid strongly adhered at centromere site in control cells; but in cells suppressed hsgo1, the adhesion at centromere site was weak, and easily detached by the experiment operation.
[0027]FIG. 16 is a set of pictures showing that Sgo1 and Bub1 are required for condensation at centromeres in mitosis. (a) By treatments with siRNA, chromosome spread was performed in mitotic HeLa cells stained with Giemsa. Representative spread is shown together with the occurrence rates. More than one hundred of the prophases and prometaphases were observed for each RNAi. An example of sister chromatid pair is magnified at the top. (b) After treatment with nocodazole for 4 hours, chromosome spread was observed in cells interfered with RNAi. Examples of the spread are shown with the frequency (n>100). (c) HeLa cells expressing Scc1-myc were fixed at 36 hours after the treatment with siRNAs. The cells were immunostained with anti-myc-antibody (green) and anti-centromere-antibody (ACA) (red). DNA was stained with DAPI (blue). (d) Rates of the cells showing Scc1-myc staining are shown. Cells expressing Scc1-myc in this cell line were less than 25%. Scale bar shows 10 μm.
[0028]FIG. 17 is a set of pictures showing the results of RNAi experiments targeting Bub1, respectively. (A, B) RNAi experiments targeting Bub1 were performed respectively, and resulted in disappearance of the localization of both proteins, hSgo1 and hSgo2 at centromere. (C, D) As the localization of both proteins, hSgo1 and hSgo2 at centromere was normal in RNAi experiments targeting a control, BubR1; the significance of the results of Bub1 was ensured. It is shown that Bub1 and BubR1 are similar but different proteins, and the localization of hSgo1 and hSgo2 at centromere depends on Bub1 (A, B), but not on BubR1 (C, D).
[0029]FIG. 18 is a set of pictures showing the results that a clone in which cDNA of mouse shugoshin homologous gene (SEQ ID NOs: 21 and 23) is fused with GFP gene was generated by using retroviral vector, and expressed in human HeLa cells. It was revealed that any of the GFP fusion proteins is co-localized with human kinetochore protein Bub1 in mitosis. The appended drawings of the figures are presented to further describe the invention and to assist in its understanding through clarification of its various aspects.
BEST MODE OF CARRYING OUT THE INVENTION
[0030]As for a protein of the present invention, a protein Sgo1 (shugoshin) comprising an amino acid sequence shown in SEQ ID NO: 2 and having a regulatory activity of chromosome segregation; a protein comprising the amino acid sequence shown in SEQ ID NO: 2 where one or several amino acids are deleted, replaced or added, and having a regulatory activity of chromosome segregation; a paralogue Sgo2 of protein Sgo1 comprising an amino acid sequence shown in SEQ ID NO: 4 and having a regulatory activity of chromosome segregation; a protein comprising the amino acid sequence shown in SEQ ID NO: 4 where one or several amino acids are deleted, replaced or added, and having a regulatory activity of chromosome segregation; a Saccharomyces cerevisiae homologue ScSgo1 of protein Sgo1 comprising an amino acid sequence shown in SEQ ID NO: 6 and having a regulatory activity of chromosome segregation; a protein comprising the amino acid sequence shown in SEQ ID NO: 6 where one or several amino acids are deleted, replaced or added, and having a regulatory activity of chromosome segregation; a protein (NC) comprising an amino acid sequence shown in SEQ ID NO: 8 and having a Neurospora crassa-derived regulatory activity of chromosome segregation; a protein comprising the amino acid sequence shown in SEQ ID NO: 8 where one or several amino acids are deleted, replaced or added, and having a regulatory activity of chromosome segregation; a protein (At) comprising an amino acid sequence shown in SEQ ID NO: 10 or 12 and having a Arabidopsis-derived regulatory activity of chromosome segregation; a protein comprising the amino acid sequence shown in SEQ ID NO: 10 or 12 where one or several amino acids are deleted, replaced or added, and having a regulatory activity of chromosome segregation; a protein (Mm) comprising an amino acid sequence shown in SEQ ID NO: 14 or 16 and having a mouse-derived regulatory activity of chromosome segregation; a protein comprising the amino acid sequence shown in SEQ ID NO: 14 or 16 where one or several amino acids are deleted, replaced or added, and having a regulatory activity of chromosome segregation; a protein (Hs) comprising an amino acid sequence shown in SEQ ID NO: 18 or 20 and having a human-derived regulatory activity of chromosome segregation; and a protein comprising the amino acid sequence shown in SEQ ID NO: 18 or 20 where one or several amino acids are deleted, replaced or added, and having a regulatory activity of chromosome segregation; can be exemplified. Further, as for the regulatory activity of chromosome segregation described in the above, although it is not especially limited as long as the activities regulate chromosome segregation, for example, activities correctly regulating chromosome segregation of germ cells and/or of somatic cell division are preferable, and activities protecting (Shugo) the centromere of sister chromatid from the separation in meiosis I is more preferable. In addition, proteins of the present invention can be prepared by known methods based on DNA-sequence information and the like, and the derivations are not limited to yeast, mouse, human and the like. Furthermore, for example, Sgo1 (shugoshin) mutant that is a protein comprising an amino acid sequence shown in SEQ ID NO: 2 where one or several amino acids are deleted, replaced or added, and having a regulatory activity of chromosome segregation, can be prepared by ordinary methods such as known gene manipulation, point mutation and the like.
[0031]As for a DNA of the present invention, a DNA encoding a protein of the present invention that has a regulatory activity of chromosome segregation: a DNA derived from fission yeast Schizosaccharomyces pombe, comprising a base sequence shown in SEQ ID NO: 1 or 3 or a complementary sequence thereof; and a DNA containing part or whole of these sequences, encoding a protein that has a regulatory activity of chromosome segregation: a DNA derived from Saccharomyces cerevisiae, comprising a base sequence shown in SEQ ID NO: 5 or a complementary sequence thereof; and a DNA containing part or whole of these sequences, encoding a protein that has a regulatory activity of chromosome segregation: a DNA derived from Neurospora crassa, comprising a base sequence shown in SEQ ID NO: 7 or a complementary sequence thereof, and encoding a protein that has a regulatory activity of chromosome segregation; and a DNA containing part or whole of these sequences, encoding a protein that has a regulatory activity of chromosome segregation: a DNA derived from Arabidopsis, comprising a base sequence shown in SEQ ID NO: 9 or 11 or a complementary sequence thereof, and encoding a protein that has a regulatory activity of chromosome segregation; and a DNA containing part or whole of these sequences, encoding a protein that has a regulatory activity of chromosome segregation: a DNA derived from mouse, comprising a base sequence shown in SEQ ID NO: 13 or 15 or a complementary sequence thereof, and encoding a protein that has a regulatory activity of chromosome segregation; and a DNA containing part or whole of these sequences, encoding a protein that has a regulatory activity of chromosome segregation: a DNA derived from human, comprising a base sequence shown in SEQ ID NO: 17 or 19 or a complementary sequence thereof, and encoding a protein that has a regulatory activity of chromosome segregation; and a DNA containing part or whole of these sequences, encoding a protein that has a regulatory activity of chromosome segregation: a DNA hybridizing with the above DNA under stringent conditions, encoding a protein that has a regulatory activity of chromosome segregation: and the like, can be exemplified.
[0032]These DNAs can be prepared by known methods based on DNA-sequence information, such as a gene or cDNA library of yeast, mouse, human and the like. Further, using a base sequence shown in SEQ ID NO: 1, 3, 5, 7, 9, 11, 13, 15, 17, 19, or others or a complementary sequence thereof, or part or whole of these sequences as a probe, DNA libraries of yeast, mouse, human and the like are hybridized under stringent conditions, and the intended DNA encoding a protein that has a regulatory activity of chromosome segregation can be obtained by isolating the DNAs that hybridized with the probes. As for a condition of hybridization to obtain the DNA; hybridization at 42° C., and washing treatment by a buffer containing 1×SSC and 0.1% SDS at 42° C.; preferably hybridization at 65° C., and washing treatment by a buffer containing 0.1×SSC and 0.1% SDS at 65° C.; can be exemplified. Moreover, as for an element affecting the stringency of hybridization, there are various elements other than the above described temperature conditions, those skilled in the art can actualize the stringency equivalent to that of hybridization as exemplified in the above with an appropriate combination of various elements.
[0033]As for a fusion protein of the present invention, any protein can be used as long as the protein of the present invention is bound to a marker protein and/or a peptide tag, as for a marker protein, it is not especially limited but a conventionally known marker protein, for example, alkaline phosphatase, Fc region of antibody, HRP, GFP and the like can be exemplified. Further, as for a peptide tag of the present invention, conventionally known peptide tags such as Myc, His, FLAG and GST tags can be specifically exemplified. The fusion protein can be produced by ordinary methods; and is useful for purification of protein Sgo1 and the like by using the affinity of Ni-NTA and His tag, and for a reagent for study in the art.
[0034]As for an antibody specifically binding to a protein of the present invention, immunospecific antibodies such as monoclonal antibody, polyclonal antibody, chimeric antibody, single-stranded antibody, humanized antibody and the like, can be specifically exemplified. These antibodies can be produced by ordinary methods with the use of proteins such as the above-mentioned Sgo1 or part thereof as an antigen, and among them a monoclonal antibody is preferable in terms of specificity. Antibodies such as a monoclonal antibody are useful for elucidating the localization of Sgo1 and others in vivo.
[0035]The above-mentioned antibodies of the present invention can be generated with the use of common protocol by administering proteins of the present invention or fragments containing epitope thereof, or cells expressing the protein on their membrane surfaces, to animals (preferably non-human). For example, for preparation of a monoclonal antibody any method such as hybridoma (Nature 256, 495-497, 1975), trioma, human B cell hybridoma (Immunology Today 4, 72, 1983) and EBV-hybridoma (MONOCLONAL ANTIBODIES AND CANCER THERAPY, pp. 77-96, Alan R. Liss, Inc., 1985), by which antibodies are generated from cultures of continuous cell lines, can be used.
[0036]To generate a single-stranded antibody against a protein of the present invention, a method for preparation of single-stranded antibody (U.S. Pat. No. 4,946,778) can be applied. Further, to express a humanized antibody, transgenic mouse or other mammals can be used, clones that express a protein of the present invention with the use of the above-mentioned antibody can be isolated/identified, and its polypeptide can be purified by affinity chromatography. Antibodies against peptide containing proteins of the present invention or antigen epitopes thereof can be possibly used for diagnosis and treatment of cancer, or of chromosome segregation diseases such as infertility or Down's syndrome using a regulatory factor of chromosome segregation as an index.
[0037]Functional analysis of a protein of the present invention can be performed by using fusion proteins fused with, for example; fluorescent substances such as FITC (fluorescein isocyanate) or tetramethyl rhodamine isocyanate; radioisotopes such as 125I, 32P, 14C, 35S or 3H; labelings with enzymes such as alkaline phosphatase, peroxidase, β-galactosidase or phycoerythrin; fluorescence emission proteins such as green fluorescent protein (GFP); or the like, to antibodies such as the above-mentioned monoclonal antibodies. As an immunological assay method with the use of antibody of the present invention, methods such as RIA, ELISA, Fluorescent antibody method, Plaque forming cell assay, Spotting method, Hemagglutination testing, Ouchterlony method can be exemplified.
[0038]The present invention will be explained in detail in the following by referring to the examples, but the technical scope of the present invention will not be limited to these.
Example 1
Method
(Screening of Rec8 Protector)
[0039]The present inventor examined a gene that is toxic only when co-expressed with Rec8 in vegetative cells. The Rec8 encoding sequence that was fused with GFP was cloned into pREP82 (ura4+ marker) under the thiamine-repressible nmt1+ promoter, to construct pREP82-rec8+-GFP. A Schizosaccharomyces pombe cDNA library constructed by mRNA that was prepared from meiotic cells, and a pREP3 vector (nmt1+ promoter, LEU2+ marker) (Y. Akiyoshi and Y. W., unpublished) were used. The leu1 ura4-D18 cells carrying pREP82-rec8+-GFP were transformed with the cDNA library, spread on agar plates containing thiamine (promoter-off) and incubated for 3 days at 30° C. The colonies were then replicated on two thiamine-free agar plates: one that contains uracil and 5'-fluoroortoic acid (5'-FOA) where only cells lacked the plasmid pREP82-rec8+-CFP can grow (thereby expresses a library clone alone), and the other that does not contain 5'-FOA (allows co-expression of rec8+-GFP and a library clone). The present inventor added Phloxine B, a drug that stains dead cells red, onto the both agar plates, thereby illuminated sick colonies. After incubation for two days, the colonies exhibiting sickness only on the co-expression agar plate were picked up, and the library-derived plasmids were recovered and analyzed.
(Schizosaccharomyces pombe Strains)
[0040]Deletion and tagging of GFP or FLAG to endogenous sgo1+ and sgo2+ were performed by a PCR-based gene targeting method (Yeast 14, 943-951 (1998)). By inserting GFP into the C-terminus of the PCR-amplified sgo1+-FLAG, sgo1+-FLAG-GFP was generated and integrated into the endogenous sgo1 locus. Further, a endogenous promoter of the sgo1+ was replaced with a nmt promoter to generate Pnmt-sgo1+ or Pnmt-sgo1+-FLAG-GFP by the PCR-based gene targeting method. The proteins tagged to Sgo1-GFP or Sgo1-FLAG was deleted depending on the purpose. A mei4Δ mutation was used to arrest meiotic cells prior to meiosis I (close to late prophase in meiosis I), and a mes1 Δ mutation was used to arrest after meiosis I, as described previously (Nature 400, 461-4 (1999)).
(Observation of Chromosomes Marked with GFP)
[0041]To observe the segregation patterns of homologues at meiosis I, h90 cells retaining cen2-GFP (Embo J 22, 2284-96 (2003)) were spotted on meiosis-inducing medium, SPA. To examine the segregation patterns of sister chromatids, opposite mating type cells, one marked with cen2-GFP and the other not marked, were mixed and spotted on SPA. After incubation for one day, the zygotes were monitored for GFP. Images were obtained under a microscope (Axioplan2, Zeiss) equipped with a cooled CCD camera (Quantix, Photometrics) and by using Metamorph software (Universal Imaging Corporation). Seven Z-sections for GFP signals were converted to single two-dimensional images by taking the maximum signal at each pixel position in the images.
(Chromatin Immunoprecipitation; ChIp)
[0042]Diploid sgo1+-FLAG-GFP was used for ChIP with Sgo1. To achieve a highly synchronous culture, the endogenous slp1+ promoter was replaced with the rad21+ promoter that is not active during meiosis, and the cells were arrested at metaphase I. The cells were incubated in nitrogen-depleted medium for 17 hours at 30° C., and 60% the cells or less were arrested at metaphase I. For ChIP with Sgo2, nda3-KM311 sgo2+-GFP cells were proliferated at 30° C., and then shifted to 18° C. After incubation for 8 hours, most of the cells were arrested at metaphase. The cells were fixed with 3% para-formaldehyde for 30 minutes at 18° C., and extracts were prepared. The DNA was broken to an average size of 400 bp, and the extracts were immunoprecipitated with rabbit anti-GFP antibodies (Clontech). DNAs prepared from the whole cell crude extracts, or immunoprecipitated chromatin fractions were analyzed by quantitative PCR, with a LightCycler or a Lightcycler-DNA Master SYBR Green I kit (Roche Molecular Biochemicals). Antibody-minus samples were used as controls in each experiment to explain the nonspecific binding in the ChIP fractions.
(Preparation of Anti-Sgo1 Antibodies)
[0043]Sgo1+ ORF was PCR-amplified from an Schizosaccharomyces pombe cDNA library, and inserted into plasmids pGEX4T-2 (Pharmacia Biotech) and pET-19b (Novagen) respectively to prepare recombinant proteins GST-Sgo1 and His-Sgo1. GST-Sgo1 was used to immunize rabbit, and the raised antibodies were purified by His-Sgo1 as described previously (Embo J 22, 5643-53 (2003)). Furthermore, for the purpose of analyzing proteins (SEQ ID NOs: 18 and 20; hSgo1 and hSgo2 respectively) encoding human shugoshin homologous gene (SEQ ID NOs: 17 and 19), part of hSgo1 and hSgo2 was expressed in E coli, and antibodies against hSgo1 and hSgo2 were produced by injecting the protein into rabbit.
(Immunostaining)
[0044]To stain endogenous Sgo1, wild-type diploid cells cultured for 5 hours in MM-N were fixed with 3% formaldehyde for 40 min at 30° C., and stained by the method described previously (Embo J 22, 5643-53 (2003)). To stain Sgo2-GFP and Mis6-HA, logarithmically growing cells were used. Sgo1 was detected by using rabbit anti-Sgo1 antibody at 1:50 and Alexa488-conjugated anti-rabbit antibody (Molecular Probes) at 1:100. Tubulin was detected by using mouse anti-tubulin antibody TAT-1 (provided by Keith Gull) at 1:200 and Cy3-tagged anti-mouse antibody (Chemicon) at 1:2000. Cells were counterstained with DAPI to visualize DNA. The Sgo2-GFP was detected by using mouse anti-GFP antibody (Roche) at 1:50 and BODIPY FL-conjugated anti-mouse antibody (Molecular Probes) at 1:100. The Mis6-HA was detected by using rabbit anti-HA antibody Y-11 (Santa Cruz) at 1:50 and Alexa488-conjugated anti-rabbit antibody at 1:100. Cells were counterstained with DAPI to visualize DNA. Further, immunostaining was performed by using rabbit anti-hSgo1 antibody and rabbit anti-hSgo2 antibody in the same manner as the above.
(Communoprecipitation)
[0045]Padh-rec8+-3HA Pnmt41-sgo1+-FLAG-GFP strain cells and control Padh-rec8+-3HA strain cells were cultured without thiamine for 15 hours at 30° C., collected, and the extracts were prepared. To liberate chromatin-bound proteins, the extracts were treated with DNase I. After clarifying the extracts by centrifugation, the Sgo1-FLAG-GFP protein was immunoprecipitated with anti-FLAG antibody M2 (Sigma). The Rec8-3HA and Sgo1-FLAG-GFP were detected by anti-HA antibody Y-11 and anti-FLAG antibody M2, respectively.
(Analysis of Budding Yeast)
[0046]All sample strains except those for chromosome loss assay are derivative of SK1 (Cell 98, 91-103 (1999)). The chromosome loss assay was performed as described previously (Nature 410, 955-9 (2001)). The ScSGO1 gene was deleted or epitope-tagged by using PCR generated cassettes (Yeast 14, 953-961 (1998)). Accurate gene targeting was checked by PCR. URA3-GFP dots marking chromosome V (cenV-GFP) were described previously (Cell 98, 91-103 (1999)). Sporulation was induced by culturing diploid cells at 30° C. as described previously (Dev Cell 4, 535-48 (2003)). In situ immunofluorescence was performed as described previously (Dev Cell 4, 535-48 (2003)).
(Cell Culture)
[0047]HeLa cells were cultured in DMEM supplemented with 10% fetal bovine serum and 0.03% L-Glutamine. The HeLa cell strain expressing Scc1-myc was cultured with 200 μg/ml of G418 (Invitrogen) and 100 μg/ml of Hygromycin B (Wako). Expression of Scc1-myc was induced by incubation with 2 μg/ml of Doxycyclin (Sigma) for 48 hours.
(Preparation of Anti-Human Sgo Antibody)
[0048]As the information for N-terminal amino acid sequence of human Sgo1 was not obtained from the databases, the present inventor cloned a cDNA fragment that was amplified from a cDNA library (BD Biosciences) with the use of primers recognizing the cloning site of λTriplEx: CTCGGGAAGCGCGCCATTGTG (SEQ ID NO: 38) and the DNA sequence corresponding to the numbers 237-242 in amino acid sequence of Q9BVA8: CCTGGCTGAATCAGCTTTGGTG (SEQ ID NO: 39). The Sequencing revealed that the Sgo1 mRNA encodes a protein having 527 amino acids. To obtain polyclonal antibodies against Sgo1, a cDNA fragment encoding the numbers 109-491 in amino acid sequence of Sgo1 was amplified and inserted into the reading frames of plasmids pGEX4T-2 (Amersham) and pET19b (Novagen) to produce GST-Sgo1 and His-Sgo1 respectively, and followed by immunization of a rabbit (QIAGEN) (performed according to the manufacturer's instructions). His-Sgo1 was affinity-purified on CNBr-activated sepharose (Amersham). Antibodies against Sgo2 were raised with GST-Sgo2 (amino acid numbers 331-631) and purified with His-Sgo2 in the same manner as the above.
(Immunofluorescence Microscopy and Chromosome Spreading)
[0049]Immunofluorescent staining was performed as described in the above, by using anti-human Sgo1 (1:1000), anti-human Sgo2 antiserum (1:10000), anti-Bub1 (1:1000, MBL), anti-BubR1 (1:1000, MBL), anti-CENP-A (1:1000, MBL), anti-Aurora B AIM-1 (1:1000, BD Biosciences) and anti-tubulin DM1A (1:1000, Sigma). Immunostaining of Scc1-myc was performed as described in the above, by using anti-myc CM-100 (1:1000, Gramsch Laboratories) and ACA (1:1000, provided from Dr. Yoshinari Takasaki). As a secondary antibody, Alexa Fluor 488 goat anti-rabbit antibody (1:1000, Molecular Probes), Cy3 conjugated anti-mouse antibody (1:1000, CHEMICON), and Cy3 conjugated donkey anti-human antibody (1:1000, Jackson ImmunoResearch Laboratories. Inc) were used. 3 μg/ml of Hoechst 33342 or 0.5 μg/ml of DAPI were used for counter staining. Images were taken by using SlideBook or MetaMorph software.
(Chromosome Spreading)
[0050]HeLa cells during mitosis were collected by mitotic shake-off and incubated with 330 nM of nocodazole for 0 up to 4 hours. Chromosome spreading was performed as described in the above.
(Immunoblotting)
[0051]HeLa cells were boiled with the sample buffer and resolved by SDS-polyacrylamide gel electrophoresis. Proteins were transferred to Immobilon-P membrane (Millipore), followed by blocking with 5% Skim milk (Nacalai) in TBST (150 mM of NaCl, 20 mM of Tris-HCl pH7.4, 0.05% Tween-20). Antibody incubations were performed in 0.1% skim milk TBST supplemented with anti-Sgo1 antibody (1:1000), anti-Sgo2 antibody (1:1000), anti-Bub1 antibody (1:500) and anti-tubulin antibody (1:3000). Blots were produced by ECL (Amersham).
(RNAi)
[0052]As a siRNA target sequence, hSgo1: AAGUCUACUGAUAAUGUCUUATT (SEQ ID NO:40) and hSgo2: AAGCACUACCACUUUGAAUAATT (SEQ ID NO:41), and human Sgo1: GUGAGCCUCUGUGAAUCAATT (SEQ ID NO:42) and human Sgo2: GCUCUCAUGAACAAUAACUTT (SEQ ID NO:43) were respectively selected on hSgo1RNA or hSgo2RNA. Furthermore, as a siRNA target sequence, GAGUGAUCACGAUUUCUAATT (SEQ ID NO: 44) was selected on other siRNA target sequence, Bub1 RNA; siRNA target sequence, AACGGGCAUUUGAAUAUGAAA (SEQ ID NO: 45, see JCS, 117, 1577-1589 (2004)) was selected at 2 sites on a spindle checkpoint factor BubR1 RNA. These sequences were synthesized as double strand, and introduced into cells by using oligofectamine (Invitrogen). Furthermore similarly, when producing HIV vector, HeLa cells were transfected with HIV plasmid vector, pMD.G (VSV-G env expressing plasmid), pMDLg/p.RRE (the third generation packaging plasmid) and pRSV Rev (Rev expressing plasmid) by calcium phosphate method, collected the culture supernatant 48 hours after the transfection, and condensed to use as a virus vector.
Example 2
Results
(Identification of Shugoshin Sgo1 in Fission Yeast)
[0053]The replacement of the mitotic cohesin, Rad21/Scc1, with the meiotic version, Rec8, is a prerequisite for protecting centromeric sister chromatid cohesion through anaphase of meiosis I ((Cell 103, 1155-68 (2000), Mol Cell Biol 23, 3965-73 (2003)). However, when Rec8 was expressed ectopically during mitosis, Rec8 was localized largely at centromeres but disappeared at anaphase, with sister chromatids segregating to opposite sides (FIGS. 1c and d). Moreover, the ectopic expression of non-cleavable Rec8 during mitosis (note that Rec8 is cleaved by separase Cut1 during meiosis (Embo J 22, 5643-53 (2003))) resulted in an inability to separate sister chromatids (see FIG. 2). Thus, in contrast to the situation during meiosis I, centromeric Rec8 is cleaved by separase during mitosis, and results in separation of sister chromatids. The present inventor thus postulated a meiosis I specific centromeric protector of Rec8 from these observations. To identify this factor, the present inventor searched for a gene that generates toxicity during mitotic growth only when co-expressed with Rec8. This screening identified a novel gene, sgo1+ (ORF: SPBP35G2.03C). The hindrance of growth by Sgo1 was significantly dependent on Rec8, as Sgo1 had little effect on growth when co-expressed with Rad21 (FIG. 1a). Co-expression of rec8+ and sgo1+ resulted in high frequency of the blocked nuclear division, as centromere-associated green fluorescent protein markers (cen2-GFP) segregated to the same side of a septated cell highly frequently (see Figs. b and c). To test the possibility that Sgo1 protects Rec8 from degradation at anaphase, the localization of Rec8 was examined in associated with Sgo1 expression, Rec8 tagged with GFP at its carboxyl terminus was expressed under the control of a constitutive adh1 promoter and induced Sgo1 by using a thiamine-repressible nmt1 promoter. Consequently it was found that the Rec8-GFP signal persisted through anaphase only when Sgo1 was co-expressed (FIG. 1d). As Sgo1 is expressed exclusively in meiosis (DNA micro array data (Nat Genet. 32, 143-7 (2002)), see below), it was found from the above-mentioned results, that Sgo1 is a protector of Rec8 during meiosis.
(Sgo1 Protects Centromeric Cohesion at Meiosis I)
[0054]To examine whether Sgo1 is actually required for the protection of Rec8 during meiosis, the entire ORF encoding sgo1+ was deleted, and the phenotype was examined. Sgo1Δ cells are viable and showed normal vegetative growth, consistent with the concept that sgo1+ is a meiosis-specific gene. To examine the meiotic chromosome segregation of sgo1Δ cells, centromere-linked sequences were marked with GFP (cen2-GFP) on only one of the two homologues in a zygote, and the segregation of the GFP dots were monitored during meiosis I. It was revealed that meiosis I emerged normally in sgo1Δ cells, as sister chromatid pairs generally moved together to the same side of each zygote. Therefore, monopolar attachment was intact (FIG. 3a). Moreover, by marking cen2-GFP on both chromosomes, it was determined that accurate segregation was undergone with homologues at meiosis I (data not shown). However, sister chromatid pairs failed to segregate properly at meiosis II, non-segregation was caused in 50% of the cells or less (FIG. 3a). This value is consistent with random chromosome segregation at meiosis II.
[0055]To examine centromeric cohesion, cen2-GFP marked on both homologues was monitored in zygotes arrested prior to meiosis II via a mes1Δ mutation. Supporting the above results, sgo1Δ cells frequently showed precocious division of centromeres as split cen2-GFP signals prevailed in the dyad nuclei (FIG. 3b). Finally, it was examined whether protection of Rec8 at centromeres is dependent on Sgo1 by monitoring Rec8-GFP at late anaphase I and prometaphase II. While it is significant that Rec8 signals were centromeric in wild-type cells, the Rec8 signals had largely disappeared from the centromeres at these stages in sgo1Δ cells (FIG. 3c). Although all phenotypes of sgo1Δ cells are reminiscent of heterochromatin-deficient Schizosaccharomyces pombe, in which Rec8 localization to the pericentromeric regions is decreased and centromeric cohesion is lost during meiosis I, leading to random division at meiosis II (Science 300, 1152-5 (2003)). Chromatin binding by Rec8 was examined in cells arrested prior to meiosis I by using a chromatin immunoprecipitation (ChIP) assay. In marked contrast to heterochromatin-deficient cells, Rec8 localization was intact in sgo1Δ cells at the pericentromeric regions as well as all other regions tested. These results suggest that the loss of centromeric Rec8 after meiosis I is caused not by an initial defect in Rec8 localization to centromeres but rather by a defect in the preservation of centromeric Rec8 during meiosis I. The above results indicated that the Cut1 separase becomes active at the onset of anaphase I and cleaves most chromosomal Rec8, leaving only centromeric Rec8 intact (Embo J 22, 5643-53 (2003)). These results indicated that Sgo1 plays an essential role in protecting centromeric cohesion throughout meiosis I by protecting cohesin Rec8 from separase cleavage.
(Sgo1 Localizes at Centromeres During Meiosis I)
[0056]To detect the Sgo1 protein, Sgo1-specific antibodies were produced, and the results of Western blotting indicated that Sgo1 is expressed only around at meiosis I (FIG. 4a). The results of immunofluorescence microscopy on cells at various stages of meiosis revealed that Sgo1 appears at late prophase of meiosis I and is fully localized as several punctuate dots by the point of metaphase I (FIG. 4b). These dots were co-localized with the Mis6 kinetochore protein (Cell 90, 131-143 (1997)), and indicated that Sgo1 is a centromere-associating protein (FIG. 4c). At the onset of anaphase I, Sgo1 signals decrease dramatically. It was found that Sgo1 remains undegraded at centromeres in APC-depleted cells arrested at metaphase I but undergoes normal degradation in separase-defective cells (FIG. 5), and indicated that Sgo1 degradation at anaphase I is regulated more directly by the APC rather than through separase. Although residual Sgo1 signals were detectable at the centromeres in early anaphase I, they disappeared completely by the end of anaphase I (FIG. 4b). This indicates that a substantial amount of Sgo1 is required at the onset of anaphase I when separase is fully activated. However, it is considered that the amounts of Sgo1 required are smaller and smaller as anaphase I progressed. This idea is tenable when the separase, activity is quickly down-regulated or when the access to chromosomes is prevented during anaphase I. Sgo1 never reappears during meiosis II (FIG. 4b), and which is consistent with the idea that Sgo1 is required for the protection of Red8 only during meiosis I.
[0057]The present inventor has already reported that Rec8 localization at pericentromeric regions is especially important for the persistence of centromeric cohesion throughout meiosis I (Science 300, 1152-5 (2003)). If Sgo1 is a centromeric protector of Rec8, then it might be expected to localize there as well. To test this possibility, Rec8 localization was delineated more precisely by using the ChIP assay. Sgo1 actually associated with pericentrmeric heterochromatin regions rather than with central core regions along the centromere sequences (FIG. 4d). As the results of immunoprecipitation experiments indicated that Sgo1 interacts with Rec8 complexes in vivo (FIG. 4f), the protection was carried out through close interaction. Concurrently, these results indicate that Sgo1 resides at pericentromeric regions and acts to protect centromeric Rec8 from the cleavage of separase at anaphase I (FIG. 4d). It was found that the localization of Rec8 does not depend on Sgo1, and vice versa (FIG. 3d, figure not shown). Actually, the Rec8 and the Sgo1 are in fact independently generated at pericentromeric regions, as for the localization, the Rec8 and the Sgo1 depend on heterochromatin and Bub1 kinase respectively (FIG. 4e). In contrast, Rec8 and Sgo1 are localized at centromeres in swi6Δ (heterochromatin deficient) and bub1Δ cells respectively (FIG. 4e). Thus by localizing independently, it can be ensured that Rec8 is protected only at centromeres not along the chromosomal arm regions.
[0058]Further, it is indicated that shugoshin shields Rec8 physically from the action of separase and counteracts the effects. On this point, even when the strong expression of Sgo1 dose not express Rec8, the mitotic growth was moderately disturbed (figure not shown); and even when the temperature is tolerated for cut1 allele, it was found that cut1 mutant was killed by moderate expression of Sgo1 (FIG. 6).
(Sgo2 is a Mitotic Sgo1 Paralogue in Fission Yeast)
[0059]By a conventional BLAST search of genome databases, the present inventor identified Sgo1-like proteins from Saccharomyces cerevisiae and Neurospora crassa, and indicated that Sgo1 is a conserved protein (see below). In the same search, a Schizosaccharomyces pombe Sgo1 paralogue which the present inventor named Sgo2, was also identified (ORF: SPAC15A10.15). The sgo2+ gene was disrupted, and it was identified that sgo2Δ cells are viable but show sensitivity to the spindle destabilizing drug thiabendazole (TBZ) (FIG. 7a). As sgo1Δ cells never show such a defect, this phenotype is remarkable. To investigate its cellular distribution, the endogenous sgo2+ gene was tagged with GFP. In proliferating cells, Sgo2-GFP was observed as two or three dots in the nucleus (FIG. 7d). However, Sgo2-GFP co-localized with the centromere protein Mis6 at metaphase and disappeared during anaphase (FIGS. 7c and d). The results of ChIP assays showed that Sgo2 chromatin association is detectable only on synchronous populations of mitotic cells, and that chromatin association is localized to the pericentromeric regions (FIG. 7e). By enhancing this localization, sgo2 deletion confers a dramatic defect to chromosome segregation when the heterochromatin-deficient swi6Δ mutation was bound thereto, however which by itself impairs centromeric function slightly (Science 269, 1429-31 (1995)) (FIG. 7b). These results indicate that Sgo2 cooperates with centromeric heterochromatin factors to ensure chromosome segregation at mitosis. Moreover, it was found that sgo2Δ cells have a modest increase (up to 15%) in non-segregation of homologues at meiosis I, and indicated that Sgo2 is also important for promoting proper meiosis I. However, the role of Sgo2 does not overlap with that of Sgo1, as sgo1Δ neither causes an apparent defect at meiosis I (FIG. 3a) nor enhances a defect of sgo2 in meiosis.
(Shugoshin Localization Controlled by Bub1)
[0060]As centromeric Rec8 cannot be detected after meiosis I in fission yeast bub1 mutants, a conserved centromere-associated kinase Bub1 is considered to function in protecting Rec8 during meiosis, (Nat Cell Biol 3, 522-6 (2001)) (FIG. 3c). Although bub1 mutation has pleiotropic effects in meiotic chromosome segregation, it is considered that Sgo1 function can be targeted by Bub1 activity. To elucidate this problem, Sgo1-GFP signals were examined in bub1Δ cells undergoing meiosis. Obviously, Bub1Δ cells were almost completely devoid of accurate centromeric Sgo1-GFP signals, instead showed a diffuse fluorescence in the nucleus (FIG. 4e). Similar results were obtained by using the bub1-K762R point mutation that abolishes the kinase activity (Embo J 22, 1075-87 (2003)). Although substantial levels of Sgo1 protein were detected in meiotic bub1Δ cells by Western blot analysis (figure not shown), Bub1 dose not influence protein stability of Sgo1. Thus, the kinase activity of Bub1 is required for incorporating Sgo1 to centromeres, and the observed defects in centromeric protection in bub1Δ cells can be explained by impaired localization of Sgo1.
[0061]In parallel experiments, it was identified that mitotic Sgo2 localization at centromeres was similarly disturbed in bub1 mutants (FIG. 7c). It has been indicated that loss of Bub1 function causes centromeric function to be weakened (J Cell Biol 143, 1775-87 (1998)). In this regard, the bub1-K762R mutation shows co-lethality with swi6Δ, a mutation that also slightly impairs centromeric function via its role in pericentromeric heterochromatin formation. It was found that sgo2Δ similarly shows co-lethality with swi6Δ (FIG. 7b), and exhibits severe miss-segregation of chromosomes at mitosis (figure not shown). As the sgo2Δ bub1Δ double mutant showed no cumulative defects at all in growth or TBZ sensitivity (FIG. 7a), Sgo2 and Bub1 tandem function was confirmed to ensure chromosome segregation in mitosis by these genetic analyses. Taken all together, the above results revealed that the incorporation of Sgo1 and Sgo2 to centromeres is a crucial function of Bub1 kinase in meiosis and mitosis, respectively.
(Characteristics of a Budding Yeast Sgo1 Homologue)
[0062]The present inventor identified a single Sgo1 homologue, ScSgo1 in budding yeast (ORF: YOR073W), which has so far not been analyzed. The cellular localization of ScSgo1 was examined by tagging endogenous ScSGO1 with GFP. ScSgo1-GFP was detected mainly as a single dot in proliferating cells, but only in a limited subset of the population (FIG. 8a). Scsgo1-GFP was not detected during the G1/S period (i.e. in cells with no bud or a small bud) but appeared as a dot in G2/M (cells with a large bud and a single nucleus) and disappeared at anaphase (cells with a large bud and a stretched nucleus) (FIG. 8a). The dot is co-localized with Ndc10 kinetochore protein (FIG. 8b). During meiosis, ScSgo1-GFP was detected at the kinetochore only at metaphase I, but never during anaphase I or meiosis II (FIG. 8c). Thus, the pattern of ScSgo1 localization closely resembles that of SpSgo2 in mitosis and SpSgo1 in meiosis.
[0063]The ScSGO1 gene was disrupted to examine the function of ScSgo1. Although the Scsgo1Δ cells were viable, they grew slowly and showed sensitivity to the spindle destabilizing drug benomyl (FIG. 8d), and indicated that centromeric function might be impaired. And then the chromosome loss rates in Scsgo1Δ cells were compared with those in wild-type cells by a colony sectoring assay. Whereas 40% of the Scsgo1Δ colonies contained red sectors (which indicate chromosome loss), less than 2% wild-type colonies contained such sectors (FIG. 8e). It was concluded that ScSgo1 plays a crucial role at centromeres to ensure mitotic chromosome segregation. At the onset of meiosis, Scsgo1Δ cells showed significant defects that many cells are arrested with a single nucleus in the meiotic condition. However, among the leaked tetranucleate products of meiosis, the distribution pattern of cenV-GFP was consistent with proper segregation at meiosis I with the exception of random segregation at meiosis II (FIG. 8f). It was also found that tagging chromosomal ScSGO1 with 13Myc at its carboxyl terminus, which by itself causes no detectable defects in mitotic growth or meiosis I, resulted in impaired segregation at meiosis II (34% non-segregation indicates 68% random segregation)(FIG. 8g). Moreover, the ScSGO1-Myc cells showed frequent separation of sister centromeres at late meiotic anaphase I (FIG. 8h), indicated that centromeric cohesion was not properly protected. Concurrently, these results support the idea that ScSgo1 plays a crucial role in protecting centromeric cohesion throughout meiosis I, and meiosis II was ensured thereby as is the case with fission yeast Sgo1.
(Conservation of Shugoshin Among Eukaryotes)
[0064]BLAST searches identified only three Sgo1-like proteins, which were all in fungi: Schizosaccharomyces pombe Sgo2, Saccharomyces cerevisiae ScSgo1, and Neurospora crassa B23G1.060. As the two conserved regions were found in these proteins, the related proteins are searched under conditions of two block sequences by the BLOCK MAKER and MAST programs (Nucleic Acids Res 26, 309-12 (1998), Bioinformatics 14, 48-54 (1998)). This approach extracted several candidate proteins from various eukaryotes including fly, worm, plant, mouse and human (see SEQ ID Nos: 21-37; drosophila Dm, Ce, Arabidopsis At, mouse Mm and human Hs, respectively, in FIG. 9). Especially, this list includes Drosophila ME1-S332, which is previously characterized as a protein essential for preserving centromeric cohesion in meiosis (Cell 83, 247-256 (1995)), although the similarity score is marginal (E-value=10). All other proteins in the list show a short stretch of similarity in the carboxyl terminal basic regions, while the primary sequences in the first block are not conserved except that they all contain a putative coiled-coil. The space and sequences between these two blocks diverge among the proteins. As these blocks were previously identified to be important for ME1-S332 function (Genes Dev. 12, 3843-3856 (1998)), the importance of the conserved regions in Sgo1 was investigated. Several amino acids were changed individually to alanines in these similarity blocks and the function of the mutant proteins in vivo was examined (FIG. 10). It was found that three conserved amino acids known to be important for ME1-S332 function were also required for Sgo1 function (13N, 34V and 368S in ME1-S332; 29N, 501 and 294S in Sgo1) (marked as arrowheads in FIG. 9). Further, other conserved amino acids in the second block (293P, 296R, 298K, 299L and 300R in Sgo1) were also all required for Sgo1 function (asterisks in FIG. 9), and non-conserved residue 297T could be changed to alanine without impairing function (circle in FIG. 9). These results indicated that the marginal structural similarity observed among Schizosaccharomyces pombe Sgo1 and other proteins in various eukaryotes is important. Plants and mammals carry two shugoshin-like proteins, suggesting the possibility that the function of shugoshin diverges to complete mitosis and meiosis as in fission yeast.
(Proteins Encoding Human Shugoshin Homologous Gene are Specifically Localized at Centromeres in Mitosis)
[0065]The present inventor previously identified two putative human Sgo proteins, Sgo1 and Sgo2 in the database, although their overall sequence homology to known Sgo proteins in any species other than human is marginal (FIG. 11a). To examine whether these proteins identified in the database are actually human Sgo homologs, the present inventor examined the localization of the proteins. For this end, the present inventor cultured rabbit polyclonal antibodies against recombinant proteins that were produced in bacteria. The obtained Sgo1 antibodies detected an up to 70 kD band (predicted molecular weight is 60 kD) in the HeLa cell extracts, and the signal was significantly reduced when cells were treated with siRNA that targets Sgo1 mRNA (FIG. 11b). Similarly, Sgo2 antibodies detected an up to 120 kD band (predicted molecular weight is 145 kD), the signal was reduced in extracts obtained from cells treated with Sgo2 siRNA (FIG. 11b). These data indicate that both Sgo1 and Sgo2 are expressed at least in proliferating HeLa cells. Next, for the purpose of analyzing proteins (SEQ ID NOs: 18 and 20, respectively hSgo1 and hSgo2) encoding human shugoshin homologous gene (SEQ ID NOs: 17 and 19) that was presumed to be human Sgo homologues, part of hSgo1 and hSgo2 was expressed in E. coli, and antibodies against hSgo1 and hSgo2 were produced by injecting the protein into rabbit, HeLa cells were stained with the antibodies and concurrently with tublin antibodies and DAPI, and co-stained with spindle and chromosome DNA respectively, and the expression of hSgo1 and hSgo2 proteins that were both endogeneous in proliferating cells was examined. The results are shown in FIG. 12. As shown in FIG. 12, both signals of hsgo1 and hSgo2 were also observed as dots on chromosomes from prometaphase to metaphase. As a result of the immunostaining, it was identified that both proteins, hsgo1 and hSgo2 are specifically localized at centromeres at mitotic phase. Further, HeLa cells at prometaphase and metaphase were stained with antibodies against hsgo1 or hSgo2; concurrently co-stained with antibodies against centromere protein CENP-A, and DAPI; and examined the expression of hsgo1 and hSgo2 proteins. The results are shown in FIG. 13. As shown in FIG. 13, both signals of hSgo1 and hSgo2 were observed at sites close to CENP-A dots on chromosomes. As a result of the above, it was revealed that both hsgo1 and hSgo2 are centromere proteins. Further, to examine this possibility, Aurora B, which is a passenger protein of chromosome known to be localized within kinetochore from prophase to metaphase, was stained. The sites of Sgo1 and Aurora B were practically the same at prometaphase and metaphase, whereas Sgo2 was placed just outside Aurora B (see FIG. 13). As a result of the above, it was revealed that both hsgo1 and hSgo2 are placed within kinetochores from prometaphase to metaphase. Representative views of sister kinetochore are magnified on the right. Scale bar is 10 μm.
(Proteins Encoding Human Shugoshin Homologous Gene are Specifically Localized at Centromeres in Mitosis and Play an Important Role to Progress Chromosome Segregation)
[0066]RNAi experiments targeting hsgo1 and hSgo2 were performed respectively. The results are shown in FIG. 14. As a result, the expressions in any proteins were significantly suppressed 48 hours later, the cells arrested in mitosis (total, in figure) were accumulated as indicated in FIG. 14. As described above, it was strongly suggested that any protein localized at centromeres in mitosis plays an important role for progressing chromosome segregation. As the accumulation was dissolved by suppressing a spindle checkpoint factor BubR1 by RNAi, it was suggested that hsgo1 and hSgo2 are directly or indirectly function during the process where spindle properly takes the kinetochore at centromeres as described below.
[0067]Further, the cells for which RNAi experiments targeting hsgo1 was performed by using HeLa cells were mounted on a slide glass and stained with Giemsa. The results are shown in FIG. 15. It was revealed that sister chromatid at prophase strongly adhered at centromere site in control cells where RNAi was not performed; while in cells suppressing hsgo1 expression, where RNAi was performed, the adhesion was weak at centromere site, and easily detached. Consequently, it was demonstrated that hsgo1 has an important role to maintain the strong cohesion at centromere site in mitosis in proliferating cells. Mitotic cells where Sgo1 protein knockdown was performed by RNAi experiments were collected, and the chromosomes were spread to observe chromosome structure directly. In control cells, sister chromatids were resolved along the arm regions but showed the primary constriction at centromeres (FIG. 16a i). Amazingly, in Sgo1-depleted cells, sister chromatids were often separated along the whole chromosome length (FIG. 16a iii). In samples where sister chromatids stayed densely close, although sister chromatids did not indicate the primary constriction (FIG. 16a iv), this suggests that centromeric cohesion was lost selectively. Nocodazole treatment activates the spindle checkpoint; thereby the cell cycle is arrested at prometaphase. Such prolonged arrest in M phase causes the complete separation of the connectivity from the chromosomal arm regions. For this reason, sister chromatids are only connected at centromeres, and form `Xshaped` chromosome (FIG. 16b, control). As expected, nocodazole-treatment caused the complete separation of sister chromatids along the chromosome length in Sgo1 RNAi cells (up to 97%) (FIGS. 16c and d). Consequently, it was demonstrated that hSgo1 plays an important role to maintain the strong cohesion at chromosomal centromere site in mitosis in proliferating cells.
[0068]RNAi experiments targeting Bub1 were performed respectively. The results are shown in FIG. 17. Consequently, the localization of either protein of the hsgo1 and hSgo2 to centromere was disappeared. This result means that the conclusion, "localization of shugoshin to centromere depends on Bub1 kinase", which was found in yeast by the present inventor, is also conserved in higher organisms.
[0069]Next, clone where cDNA of mouse shugoshin homologous genes (SEQ ID NOs: 21 and 23) was fused with GFP gene was produced by using retroviral vector and expressed in human HeLa cells. The results are shown in FIG. 18. Consequently, it was revealed that any of the GFP fusion proteins are also co-localized with human kinetochore protein Bub1 in mitosis.
[0070]The analysis of the above hsgo1 and hSgo2 and the analysis results obtained with the use of mouse shugoshin homologous genes were strongly suggested that shugoshin-like protein in animal cells, which were predicted from the sequence, also have functional conservation with yeast shugoshin.
INDUSTRIAL APPLICABILITY
[0071]Shugoshin of the present invention that is a regulatory factor of chromosome segregation widely conserved in eukaryotic cells, can be advantageously used for studies on the induction mechanism of cancer in somatic division, the chromosome segregation diseases such as infertility or Down's syndrome in meiotic division, and the like besides on the elucidation of mechanism in chromosome segregation.
Sequence CWU
1
451960DNAyeast 1atgaactttc aatttataaa ttcaaatata aacaatgaag ataaattgcc
gatggagtcg 60ttgaaaaaga aatttttaaa acaaaatcgt gaaattataa aaataaatac
tcagctttct 120ataaaaatta gagaatctga aaacgaaatt caagatttga tacaagaaaa
tttcactttg 180aaaagttatt tggttaaact tgaagctcga tttcgcaatc aatctcaaac
tgaggacttg 240ttaaaaaact tctttcctga gatacaaacc attcacaaaa agatttcaca
agtgcaaagt 300ttactgaaga ttatagagaa aaagtgttca tcagatttcc tcgaagcgaa
tgtaaaaagt 360caatttacaa cctgtgaaaa taaagattcg aaagaagatt atcagatttt
gcataataaa 420cgcttggagt atgtatcatt taatgatgaa cttaaaagtc tcgaaacagg
gcaaccattg 480tattgttttc aagatttcca aaaaaaagtc catggtcctc cggctctatc
tgaaaaacct 540ggaaaatgta tattaaaaga taaaaccaat gcccacgtaa acaaaatacc
acaagatgag 600gtgaattact cattgccgca aaaaaatatc accatctttt caaaggaatt
aaaagaaaac 660gaatttgaat ccatcaacga gggcgaaact gaagaagaaa aggctaaaac
atcaaatgtt 720tgtgtttgta ttccttgtaa aagtgctgaa cagataactg accttaaagg
acaagcaacc 780ggagacagct ccccatgtga ttttgaagaa tctcaaccaa ggattaatgg
acgtgaaaaa 840ctaagacgat cagtcaaagt gataaactat gcaataccca gtttgcgaac
taaactacga 900cgagactttg acttaccatc tgatagaaaa cgcaaacgac atcccagagg
caaagcataa 9602319PRTyeast 2Met Asn Phe Gln Phe Ile Asn Ser Asn Ile
Asn Asn Glu Asp Lys Leu1 5 10
15Pro Met Glu Ser Leu Lys Lys Lys Phe Leu Lys Gln Asn Arg Glu Ile
20 25 30Ile Lys Ile Asn Thr Gln
Leu Ser Ile Lys Ile Arg Glu Ser Glu Asn 35 40
45Glu Ile Gln Asp Leu Ile Gln Glu Asn Phe Thr Leu Lys Ser
Tyr Leu 50 55 60Val Lys Leu Glu Ala
Arg Phe Arg Asn Gln Ser Gln Thr Glu Asp Leu65 70
75 80Leu Lys Asn Phe Phe Pro Glu Ile Gln Thr
Ile His Lys Lys Ile Ser 85 90
95Gln Val Gln Ser Leu Leu Lys Ile Ile Glu Lys Lys Cys Ser Ser Asp
100 105 110Phe Leu Glu Ala Asn
Val Lys Ser Gln Phe Thr Thr Cys Glu Asn Lys 115
120 125Asp Ser Lys Glu Asp Tyr Gln Ile Leu His Asn Lys
Arg Leu Glu Tyr 130 135 140Val Ser Phe
Asn Asp Glu Leu Lys Ser Leu Glu Thr Gly Gln Pro Leu145
150 155 160Tyr Cys Phe Gln Asp Phe Gln
Lys Lys Val His Gly Pro Pro Ala Leu 165
170 175Ser Glu Lys Pro Gly Lys Cys Ile Leu Lys Asp Lys
Thr Asn Ala His 180 185 190Val
Asn Lys Ile Pro Gln Asp Glu Val Asn Tyr Ser Leu Pro Gln Lys 195
200 205Asn Ile Thr Ile Phe Ser Lys Glu Leu
Lys Glu Asn Glu Phe Glu Ser 210 215
220Ile Asn Glu Gly Glu Thr Glu Glu Glu Lys Ala Lys Thr Ser Asn Val225
230 235 240Cys Val Cys Ile
Pro Cys Lys Ser Ala Glu Gln Ile Thr Asp Leu Lys 245
250 255Gly Gln Ala Thr Gly Asp Ser Ser Pro Cys
Asp Phe Glu Glu Ser Gln 260 265
270Pro Arg Ile Asn Gly Arg Glu Lys Leu Arg Arg Ser Val Lys Val Ile
275 280 285Asn Tyr Ala Ile Pro Ser Leu
Arg Thr Lys Leu Arg Arg Asp Phe Asp 290 295
300Leu Pro Ser Asp Arg Lys Arg Lys Arg His Pro Arg Gly Lys Ala305
310 31531944DNAyeast 3atgtcgaaag catctctttc
cccgaacgta gaagacttga aaaaaaagca aattcgacag 60tataaggaaa ttatacgaat
aagcaaggca caatcaatta gaattaaaga attgcagtta 120gaaaatgaac ggttgctttc
ggaaaatatc gatttgagga ctacagcgat aaacttggaa 180gagcaactcg aaaccgtgca
aaacgaaaac gaagaaaaca aaacaaagtt agctgcatta 240cttaatcgat ttcatgaaga
aacagataat tttttatcaa aattaagtct ttgtcagcaa 300gaaatacaag acaccttcaa
accagtggag gctaacttag cttacgatgt cgatacggat 360tctgaagacc ttgacgagga
atccgtcgtg aaagataccg aagaaataat tgagcaagct 420cagcatgatg tttccttacg
aaatttaagt ggaatagagg atgaaaatat aattgatgac 480ggagaaactg ctataaatga
acaaaaaaaa agagaagcta atgttttttc cgacacgcaa 540tcagcacctc agctaaaatc
cggcaaagcc ctcccagctg attttgaaaa tccttacaat 600ctatccaatt cgaaacctgt
aaataataat aatgaagata gagttgaagc ggttacttct 660gaaaataaat ctatcgattc
tgctcctcag gaaaaaaatc atgaatacga aatcgttagt 720ccaaaatcat tatccaacaa
aattaataat caagcagctg cacaaagaag aaccgaagaa 780gataatgcaa atggagttgc
tcaagaagaa aatgagggtt cacaagaagc tcattttcat 840agcagaatac aatctgatac
agtaatacaa agtacaccca ctaaacggaa atgggacgtt 900gacattcaaa ataaacaaat
taatctggct tctgcagcta ccaatgttac cggttatgta 960tcggagaccg atagtcgccc
caatcgcgca aactctttgg attctgctgt ccttcttgtg 1020caatcttcaa ataaaagtaa
ccgaaatggg catcatattt cagatcctaa tttaaatagc 1080tccatatcgt tgaagtttgc
gcctgaagat actgcgcata attcattaac ttcacaagag 1140aatgttgggc ctcaggttac
gacgacttct ctgtcaaata tgactgttgc tgaatctcct 1200cgtacagaca ctccaaggga
aataaacggg ttagtagact cttctgtcac taatgggaac 1260gaaaaatttt ctgtagaaat
aatgaatgac tctaacaaaa ttggactgaa tcctaaatct 1320tttaccgacg aagagcggga
aattttaaca ctttttcgaa atcctcccat gagactgtca 1380agtgaacctc catcttcaaa
tggattttca atagcccatc ccaataattc tccgttacgt 1440ccgccatcgc tacaaggaat
attgaatgct gaagatcgac cttacgaaat tgagccgtca 1500cgtagctcct ttgctaccaa
cgatacgggc tcctataata atttggaact tctgtcatct 1560gtaacgaatt tgaaatcccc
taatgagaac gatcgtgtga cgaaaactca gtcgcgaaga 1620gaaacaaaag tgaaaaggcg
aagaaaagct cggattcaag aaacttctga agaaagtaca 1680gtagtcaatg agccaaatga
aaaacctgat ggaaggagcc gaagggaacg gaaaaaggtt 1740aattacgctt tgcctggatt
aaggacgaaa ttaagacgga atttcgattt accttcagat 1800catgtaaaag ctaaaaaaac
gagacgtgct cctaagaact ctgagaatga ttcagctacc 1860aaaacagaaa ccgcaaacat
tacttctgaa gcacccacta cttcagaagt aacccttgaa 1920aactccgaaa cccttaattt
gtaa 19444647PRTyeast 4Met Ser
Lys Ala Ser Leu Ser Pro Asn Val Glu Asp Leu Lys Lys Lys1 5
10 15Gln Ile Arg Gln Tyr Lys Glu Ile
Ile Arg Ile Ser Lys Ala Gln Ser 20 25
30Ile Arg Ile Lys Glu Leu Gln Leu Glu Asn Glu Arg Leu Leu Ser
Glu 35 40 45Asn Ile Asp Leu Arg
Thr Thr Ala Ile Asn Leu Glu Glu Gln Leu Glu 50 55
60Thr Val Gln Asn Glu Asn Glu Glu Asn Lys Thr Lys Leu Ala
Ala Leu65 70 75 80Leu
Asn Arg Phe His Glu Glu Thr Asp Asn Phe Leu Ser Lys Leu Ser
85 90 95Leu Cys Gln Gln Glu Ile Gln
Asp Thr Phe Lys Pro Val Glu Ala Asn 100 105
110Leu Ala Tyr Asp Val Asp Thr Asp Ser Glu Asp Leu Asp Glu
Glu Ser 115 120 125Val Val Lys Asp
Thr Glu Glu Ile Ile Glu Gln Ala Gln His Asp Val 130
135 140Ser Leu Arg Asn Leu Ser Gly Ile Glu Asp Glu Asn
Ile Ile Asp Asp145 150 155
160Gly Glu Thr Ala Ile Asn Glu Gln Lys Lys Arg Glu Ala Asn Val Phe
165 170 175Ser Asp Thr Gln Ser
Ala Pro Gln Leu Lys Ser Gly Lys Ala Leu Pro 180
185 190Ala Asp Phe Glu Asn Pro Tyr Asn Leu Ser Asn Ser
Lys Pro Val Asn 195 200 205Asn Asn
Asn Glu Asp Arg Val Glu Ala Val Thr Ser Glu Asn Lys Ser 210
215 220Ile Asp Ser Ala Pro Gln Glu Lys Asn His Glu
Tyr Glu Ile Val Ser225 230 235
240Pro Lys Ser Leu Ser Asn Lys Ile Asn Asn Gln Ala Ala Ala Gln Arg
245 250 255Arg Thr Glu Glu
Asp Asn Ala Asn Gly Val Ala Gln Glu Glu Asn Glu 260
265 270Gly Ser Gln Glu Ala His Phe His Ser Arg Ile
Gln Ser Asp Thr Val 275 280 285Ile
Gln Ser Thr Pro Thr Lys Arg Lys Trp Asp Val Asp Ile Gln Asn 290
295 300Lys Gln Ile Asn Leu Ala Ser Ala Ala Thr
Asn Val Thr Gly Tyr Val305 310 315
320Ser Glu Thr Asp Ser Arg Pro Asn Arg Ala Asn Ser Leu Asp Ser
Ala 325 330 335Val Leu Leu
Val Gln Ser Ser Asn Lys Ser Asn Arg Asn Gly His His 340
345 350Ile Ser Asp Pro Asn Leu Asn Ser Ser Ile
Ser Leu Lys Phe Ala Pro 355 360
365Glu Asp Thr Ala His Asn Ser Leu Thr Ser Gln Glu Asn Val Gly Pro 370
375 380Gln Val Thr Thr Thr Ser Leu Ser
Asn Met Thr Val Ala Glu Ser Pro385 390
395 400Arg Thr Asp Thr Pro Arg Glu Ile Asn Gly Leu Val
Asp Ser Ser Val 405 410
415Thr Asn Gly Asn Glu Lys Phe Ser Val Glu Ile Met Asn Asp Ser Asn
420 425 430Lys Ile Gly Leu Asn Pro
Lys Ser Phe Thr Asp Glu Glu Arg Glu Ile 435 440
445Leu Thr Leu Phe Arg Asn Pro Pro Met Arg Leu Ser Ser Glu
Pro Pro 450 455 460Ser Ser Asn Gly Phe
Ser Ile Ala His Pro Asn Asn Ser Pro Leu Arg465 470
475 480Pro Pro Ser Leu Gln Gly Ile Leu Asn Ala
Glu Asp Arg Pro Tyr Glu 485 490
495Ile Glu Pro Ser Arg Ser Ser Phe Ala Thr Asn Asp Thr Gly Ser Tyr
500 505 510Asn Asn Leu Glu Leu
Leu Ser Ser Val Thr Asn Leu Lys Ser Pro Asn 515
520 525Glu Asn Asp Arg Val Thr Lys Thr Gln Ser Arg Arg
Glu Thr Lys Val 530 535 540Lys Arg Arg
Arg Lys Ala Arg Ile Gln Glu Thr Ser Glu Glu Ser Thr545
550 555 560Val Val Asn Glu Pro Asn Glu
Lys Pro Asp Gly Arg Ser Arg Arg Glu 565
570 575Arg Lys Lys Val Asn Tyr Ala Leu Pro Gly Leu Arg
Thr Lys Leu Arg 580 585 590Arg
Asn Phe Asp Leu Pro Ser Asp His Val Lys Ala Lys Lys Thr Arg 595
600 605Arg Ala Pro Lys Asn Ser Glu Asn Asp
Ser Ala Thr Lys Thr Glu Thr 610 615
620Ala Asn Ile Thr Ser Glu Ala Pro Thr Thr Ser Glu Val Thr Leu Glu625
630 635 640Asn Ser Glu Thr
Leu Asn Leu 64551773DNAyeast 5atgccgaaga gaaaaattgc
tcctaacaag gaaagcagca ggcgtacggt ctcccacgat 60gatttaaccc cacaaataca
agaatttcaa aacctaatgg atctcgaatc gcaaaaagtg 120gaaaacatca gacagtcgta
ttcgaggcaa aactccctgc tggccaagga taactccata 180ttaaaaatta aagttaatag
cttggaaaaa aaaataagcc agctggtaca agaaaacgtg 240actctacgat ctaaaacctc
tataagcgaa gctatctaca gggaacggtt aagtaatcaa 300ctacaagtca ttgaaaacgg
tattattcaa agatttgacg aaatttttta tatgtttgag 360aacgtacgta aaaacgaaaa
tttgcccagt tcgagcttaa gaacaatgtt gaagagaacg 420agttccaggt caagatcatg
ctcattgtca tcacccacat actcaaaaag ttacactagg 480ttatcaaatc acgagaataa
cctgtcgcat gaatcaagtt ttaacaagga cgatggtcca 540gatcttgagc ctaaggctaa
aaaaaggaag agttctaggc ggcaatctat gtttgtatcc 600acgagtttag aacctgaaga
cgaaaccggt gaaaacgaac ccatgatgga aaattcctct 660gtagaggtac cggcagaatc
acacgagtct gcgcaagtgg aggaaacaat agatgcctta 720aaccctgaag aggaaaatag
cgattctgtc agtaatttta ccaattcaat tatagaatac 780tccataccag aggagaatcc
gacagaaccc gagcattcat cttctaaact agaaatattc 840aatgacagta caaatatgct
aagtacagtg ccgtcaaatc ctttgccgtt gcctttacca 900ggcccatccg caactttacc
tactaccact agcgatgctt caacggtcta tccttcatca 960agttcttcta ctaattctca
tccaaagacc aaaattaagc attccatgaa gccgcctagg 1020atagaactga agaaaaaggt
tattgacgaa gtcatgcccg taagtaacat gagcagcaac 1080agcgaaatat catttacgag
aactagaaga actcgtggta aagctgtaga ttacactttg 1140ccttctttaa gagccaaaat
gaggaggcct tcagaaaaac ttgtggatgc tactactgtg 1200attgatatac atgatctaca
ggtttccaag agaaatcggg aaacttcaca taaaaggaaa 1260agtttatccc aagattcaat
acccgacgaa ccgcaattga gagaagtcgt cgtctcaaag 1320gattatggaa ctccaaaagg
gaaaaaaacg gaagatgaaa tacacgagga taccgctcat 1380ctaatgacca cttccaacaa
caacagcaac aacaaaaacg aaaaaaaact aactagcaac 1440aatagcccta aaaaatcgtc
gcctttactt gacattacaa ataaatcgga gaataagaaa 1500aagtcaacaa gaactaaaaa
attgttcaaa aatgcaattg tcaataattt atctgatgaa 1560aattctacta cgcgaccctc
caagtcgtca aagggaacca gtaataataa caacaattac 1620aacaatttcg acaataacaa
ttcaaacatt aataatgtta ataataaatc tgttagcttt 1680agactaaatg aagatgattt
agcagtattt gatttatttg gaaatggtaa ggcagtgaaa 1740catcaaccaa aaacatatcg
caccaaaaaa tga 17736590PRTyeast 6Met Pro
Lys Arg Lys Ile Ala Pro Asn Lys Glu Ser Ser Arg Arg Thr1 5
10 15Val Ser His Asp Asp Leu Thr Pro
Gln Ile Gln Glu Phe Gln Asn Leu 20 25
30Met Asp Leu Glu Ser Gln Lys Val Glu Asn Ile Arg Gln Ser Tyr
Ser 35 40 45Arg Gln Asn Ser Leu
Leu Ala Lys Asp Asn Ser Ile Leu Lys Ile Lys 50 55
60Val Asn Ser Leu Glu Lys Lys Ile Ser Gln Leu Val Gln Glu
Asn Val65 70 75 80Thr
Leu Arg Ser Lys Thr Ser Ile Ser Glu Ala Ile Tyr Arg Glu Arg
85 90 95Leu Ser Asn Gln Leu Gln Val
Ile Glu Asn Gly Ile Ile Gln Arg Phe 100 105
110Asp Glu Ile Phe Tyr Met Phe Glu Asn Val Arg Lys Asn Glu
Asn Leu 115 120 125Pro Ser Ser Ser
Leu Arg Thr Met Leu Lys Arg Thr Ser Ser Arg Ser 130
135 140Arg Ser Cys Ser Leu Ser Ser Pro Thr Tyr Ser Lys
Ser Tyr Thr Arg145 150 155
160Leu Ser Asn His Glu Asn Asn Leu Ser His Glu Ser Ser Phe Asn Lys
165 170 175Asp Asp Gly Pro Asp
Leu Glu Pro Lys Ala Lys Lys Arg Lys Ser Ser 180
185 190Arg Arg Gln Ser Met Phe Val Ser Thr Ser Leu Glu
Pro Glu Asp Glu 195 200 205Thr Gly
Glu Asn Glu Pro Met Met Glu Asn Ser Ser Val Glu Val Pro 210
215 220Ala Glu Ser His Glu Ser Ala Gln Val Glu Glu
Thr Ile Asp Ala Leu225 230 235
240Asn Pro Glu Glu Glu Asn Ser Asp Ser Val Ser Asn Phe Thr Asn Ser
245 250 255Ile Ile Glu Tyr
Ser Ile Pro Glu Glu Asn Pro Thr Glu Pro Glu His 260
265 270Ser Ser Ser Lys Leu Glu Ile Phe Asn Asp Ser
Thr Asn Met Leu Ser 275 280 285Thr
Val Pro Ser Asn Pro Leu Pro Leu Pro Leu Pro Gly Pro Ser Ala 290
295 300Thr Leu Pro Thr Thr Thr Ser Asp Ala Ser
Thr Val Tyr Pro Ser Ser305 310 315
320Ser Ser Ser Thr Asn Ser His Pro Lys Thr Lys Ile Lys His Ser
Met 325 330 335Lys Pro Pro
Arg Ile Glu Leu Lys Lys Lys Val Ile Asp Glu Val Met 340
345 350Pro Val Ser Asn Met Ser Ser Asn Ser Glu
Ile Ser Phe Thr Arg Thr 355 360
365Arg Arg Thr Arg Gly Lys Ala Val Asp Tyr Thr Leu Pro Ser Leu Arg 370
375 380Ala Lys Met Arg Arg Pro Ser Glu
Lys Leu Val Asp Ala Thr Thr Val385 390
395 400Ile Asp Ile His Asp Leu Gln Val Ser Lys Arg Asn
Arg Glu Thr Ser 405 410
415His Lys Arg Lys Ser Leu Ser Gln Asp Ser Ile Pro Asp Glu Pro Gln
420 425 430Leu Arg Glu Val Val Val
Ser Lys Asp Tyr Gly Thr Pro Lys Gly Lys 435 440
445Lys Thr Glu Asp Glu Ile His Glu Asp Thr Ala His Leu Met
Thr Thr 450 455 460Ser Asn Asn Asn Ser
Asn Asn Lys Asn Glu Lys Lys Leu Thr Ser Asn465 470
475 480Asn Ser Pro Lys Lys Ser Ser Pro Leu Leu
Asp Ile Thr Asn Lys Ser 485 490
495Glu Asn Lys Lys Lys Ser Thr Arg Thr Lys Lys Leu Phe Lys Asn Ala
500 505 510Ile Val Asn Asn Leu
Ser Asp Glu Asn Ser Thr Thr Arg Pro Ser Lys 515
520 525Ser Ser Lys Gly Thr Ser Asn Asn Asn Asn Asn Tyr
Asn Asn Phe Asp 530 535 540Asn Asn Asn
Ser Asn Ile Asn Asn Val Asn Asn Lys Ser Val Ser Phe545
550 555 560Arg Leu Asn Glu Asp Asp Leu
Ala Val Phe Asp Leu Phe Gly Asn Gly 565
570 575Lys Ala Val Lys His Gln Pro Lys Thr Tyr Arg Thr
Lys Lys 580 585
59072325DNANeurospora crassa 7atggcccgcc tcaacgaaca agccatgtcg tctgtcgcgt
tgtcaacaga caatctcgag 60ctcctgcgta ggaagttcct cagacaaaac agagatattg
ctcgagtcaa ttccacacag 120tcactccgta tccgtgggtt ggagaatgaa tgcgctcgtt
tgctgtcgga aaacctcgaa 180ctccgtggtc aggtcttgcg cctcgaaaag gagctccaag
acaacgctgc gcgaagggtg 240gccgatcatg cgctcgaggt caaggccaag atggagacgc
agttggcgga actcagttcg 300ctgctggcaa gcttagggga gccgccctcg aagcggcgcc
tttcagaaga gaggcgatac 360gcgcagcctc gaccgagcgt tcaccggagc cctcccttac
gaagagcacg ccaggaggcc 420gaccaggaac tactggctga gcaggaagga aggctaccgc
cgatatacga gaacaagacg 480tatgcgcgag ccacaatgaa cagtgaagaa atcctggcgc
tgtgcatgca ggcagacgat 540tcgaatgact cgccagatat cggaccgccg ccagtatcta
ggtttgtcga ggatgatatg 600gtcatacctt gttcaccatc gccaaacaag aacgccgagg
ctgaagaaac ggaaactacc 660gagcaagtgg aagagagccc tagggctctt caagtaccgc
cgtcattatc gccgcctaaa 720ctggactacg acaggagacc aaacatgatc ctattcagcc
cacccaaaga atcgagagtg 780gcagaaccct ccaaaatgtt cagtccccct ccgatggaac
caccgaaaca gtccacatcg 840gctgtaccga gtgagacaat acgagcaggc ctcaagcgaa
agttgaacgg cgacaaccaa 900aacgaaccca acaaggcaac caagcttcaa caaggaaagg
agaatggcaa tgagactggg 960atcaagaaag gactctctgc ccgcgacccg cacaagagga
aaagcatcaa agagaccgca 1020acgaaaccga gagccccgct gtcagcaaag agcacgaacg
agcacattgt ctctccgaag 1080aagccggcga agccccacca agtggccgac gattttaagc
cggtgaaggt gcacaaggcg 1140tcaaagggta aagagaaagt cgacctgccc gctccggaca
agaagtcagc agtagaagaa 1200acgcaaggaa attctacgtc ggcattcacg aaagtcgaga
tcctcccgcc ggctctggaa 1260cctactcctg aagttgcaga gattcctgaa accgatattc
tgatcacacc tggaacacca 1320gagcgcgcct ctgaaagcac tgttgtgacc cacgataccc
cgccgccagc ccacatttca 1380tccaatggag agacgtcgcg gcctagcagg cgtgctagag
cggctatcag ctatacagag 1440cccaatctgc gcgacaagat gcgacgaccg accaaagagc
tctttgatgc cgtttctggg 1500gagggcaagt tcctacacag gccgacatcg caacagcaac
agcagcaacg caagggcgac 1560gagtcagcac cgacgtcagt tagcaaggtc aaggtcgagc
catcgccggc ggtggatata 1620agtagtctga ccagcagtgc gctgtttgaa aaagagaagg
agaaggaacc acagccggat 1680gaaggaatat tatctccaaa cggcatcctc ccaagctcag
tagacctggg aaggagaaga 1740cgcgcctcat ccttctctac tgcagcccct gcaatgacaa
ttccttcggt ccaagaacaa 1800tcaactctaa acctcccagc cgcggacgag accgatgaaa
acgccgcggt cgaggctcag 1860attcagaagg agctgagtaa tagtattaca acacggccca
ggggtggaaa ggggaggcaa 1920tcaatgagcc gttccgtacc cacgatccca acagaaaatt
acgagcacga ggacgcacaa 1980ctctcgacga actcagcctc ggtggatctt tacgactttg
ctagttgtgc gtctccggat 2040agcgcagcac cccagctaga agcgactacc ggcgatgttc
ctgttaataa gaaggcaccc 2100aaaggttcaa gaagagcgtc ctcagctgct tcgaccgaga
caacagcaac agcatccgca 2160aagccaagat cttcccgaaa aagggcttcg atgctggtgc
cgaagaaaag cttgtgggct 2220gaagagttag cgcaggagga agaggatgag gaagatgtcg
gcaatgacag tggcgggtcc 2280ttgtccaagg ggagggcctc gaggaggaga agcatgatgc
tttga 23258774PRTNeurospora crassa 8Met Ala Arg Leu Asn
Glu Gln Ala Met Ser Ser Val Ala Leu Ser Thr1 5
10 15Asp Asn Leu Glu Leu Leu Arg Arg Lys Phe Leu
Arg Gln Asn Arg Asp 20 25
30Ile Ala Arg Val Asn Ser Thr Gln Ser Leu Arg Ile Arg Gly Leu Glu
35 40 45Asn Glu Cys Ala Arg Leu Leu Ser
Glu Asn Leu Glu Leu Arg Gly Gln 50 55
60Val Leu Arg Leu Glu Lys Glu Leu Gln Asp Asn Ala Ala Arg Arg Val65
70 75 80Ala Asp His Ala Leu
Glu Val Lys Ala Lys Met Glu Thr Gln Leu Ala 85
90 95Glu Leu Ser Ser Leu Leu Ala Ser Leu Gly Glu
Pro Pro Ser Lys Arg 100 105
110Arg Leu Ser Glu Glu Arg Arg Tyr Ala Gln Pro Arg Pro Ser Val His
115 120 125Arg Ser Pro Pro Leu Arg Arg
Ala Arg Gln Glu Ala Asp Gln Glu Leu 130 135
140Leu Ala Glu Gln Glu Gly Arg Leu Pro Pro Ile Tyr Glu Asn Lys
Thr145 150 155 160Tyr Ala
Arg Ala Thr Met Asn Ser Glu Glu Ile Leu Ala Leu Cys Met
165 170 175Gln Ala Asp Asp Ser Asn Asp
Ser Pro Asp Ile Gly Pro Pro Pro Val 180 185
190Ser Arg Phe Val Glu Asp Asp Met Val Ile Pro Cys Ser Pro
Ser Pro 195 200 205Asn Lys Asn Ala
Glu Ala Glu Glu Thr Glu Thr Thr Glu Gln Val Glu 210
215 220Glu Ser Pro Arg Ala Leu Gln Val Pro Pro Ser Leu
Ser Pro Pro Lys225 230 235
240Leu Asp Tyr Asp Arg Arg Pro Asn Met Ile Leu Phe Ser Pro Pro Lys
245 250 255Glu Ser Arg Val Ala
Glu Pro Ser Lys Met Phe Ser Pro Pro Pro Met 260
265 270Glu Pro Pro Lys Gln Ser Thr Ser Ala Val Pro Ser
Glu Thr Ile Arg 275 280 285Ala Gly
Leu Lys Arg Lys Leu Asn Gly Asp Asn Gln Asn Glu Pro Asn 290
295 300Lys Ala Thr Lys Leu Gln Gln Gly Lys Glu Asn
Gly Asn Glu Thr Gly305 310 315
320Ile Lys Lys Gly Leu Ser Ala Arg Asp Pro His Lys Arg Lys Ser Ile
325 330 335Lys Glu Thr Ala
Thr Lys Pro Arg Ala Pro Leu Ser Ala Lys Ser Thr 340
345 350Asn Glu His Ile Val Ser Pro Lys Lys Pro Ala
Lys Pro His Gln Val 355 360 365Ala
Asp Asp Phe Lys Pro Val Lys Val His Lys Ala Ser Lys Gly Lys 370
375 380Glu Lys Val Asp Leu Pro Ala Pro Asp Lys
Lys Ser Ala Val Glu Glu385 390 395
400Thr Gln Gly Asn Ser Thr Ser Ala Phe Thr Lys Val Glu Ile Leu
Pro 405 410 415Pro Ala Leu
Glu Pro Thr Pro Glu Val Ala Glu Ile Pro Glu Thr Asp 420
425 430Ile Leu Ile Thr Pro Gly Thr Pro Glu Arg
Ala Ser Glu Ser Thr Val 435 440
445Val Thr His Asp Thr Pro Pro Pro Ala His Ile Ser Ser Asn Gly Glu 450
455 460Thr Ser Arg Pro Ser Arg Arg Ala
Arg Ala Ala Ile Ser Tyr Thr Glu465 470
475 480Pro Asn Leu Arg Asp Lys Met Arg Arg Pro Thr Lys
Glu Leu Phe Asp 485 490
495Ala Val Ser Gly Glu Gly Lys Phe Leu His Arg Pro Thr Ser Gln Gln
500 505 510Gln Gln Gln Gln Arg Lys
Gly Asp Glu Ser Ala Pro Thr Ser Val Ser 515 520
525Lys Val Lys Val Glu Pro Ser Pro Ala Val Asp Ile Ser Ser
Leu Thr 530 535 540Ser Ser Ala Leu Phe
Glu Lys Glu Lys Glu Lys Glu Pro Gln Pro Asp545 550
555 560Glu Gly Ile Leu Ser Pro Asn Gly Ile Leu
Pro Ser Ser Val Asp Leu 565 570
575Gly Arg Arg Arg Arg Ala Ser Ser Phe Ser Thr Ala Ala Pro Ala Met
580 585 590Thr Ile Pro Ser Val
Gln Glu Gln Ser Thr Leu Asn Leu Pro Ala Ala 595
600 605Asp Glu Thr Asp Glu Asn Ala Ala Val Glu Ala Gln
Ile Gln Lys Glu 610 615 620Leu Ser Asn
Ser Ile Thr Thr Arg Pro Arg Gly Gly Lys Gly Arg Gln625
630 635 640Ser Met Ser Arg Ser Val Pro
Thr Ile Pro Thr Glu Asn Tyr Glu His 645
650 655Glu Asp Ala Gln Leu Ser Thr Asn Ser Ala Ser Val
Asp Leu Tyr Asp 660 665 670Phe
Ala Ser Cys Ala Ser Pro Asp Ser Ala Ala Pro Gln Leu Glu Ala 675
680 685Thr Thr Gly Asp Val Pro Val Asn Lys
Lys Ala Pro Lys Gly Ser Arg 690 695
700Arg Ala Ser Ser Ala Ala Ser Thr Glu Thr Thr Ala Thr Ala Ser Ala705
710 715 720Lys Pro Arg Ser
Ser Arg Lys Arg Ala Ser Met Leu Val Pro Lys Lys 725
730 735Ser Leu Trp Ala Glu Glu Leu Ala Gln Glu
Glu Glu Asp Glu Glu Asp 740 745
750Val Gly Asn Asp Ser Gly Gly Ser Leu Ser Lys Gly Arg Ala Ser Arg
755 760 765Arg Arg Ser Met Met Leu
77091671DNAArabidopsis thaliana 9atggttcgag cgacggttct gaatgtcggt
gatcacgcca gtgaaggtgt gcgtactaac 60aaagctaaag gagagaaaat ggttctggaa
cctccgatga acagtgcaca aagacgaaag 120ttgggggata ttactaattt gcagaatcag
aagaatctaa tgaatcaggg agcgaagcat 180cagcaacaag ctatattaat ctcttctaaa
gaaaacgctg aaaatcttca aaaggcactg 240agaaattctt ctgaaaacac aaagctgatg
aaagtcgtca tggagagaga tggaatcaaa 300agtgatctga agaaacttag gattgaattt
cagaaggttc aagaacagaa tttgctactt 360gcccaggcta acactcgtat cttggcgctg
aaggtacttc agcacgaact tggttgcaag 420aatgggttag tcatggccag gaaaatgctg
cttaaggctc aagcaaatgc ttgtggtggg 480gcttgcaaaa cctttcagcc aaatgatgca
gatcatgagc atgcttccgg gagctccaac 540gctaactcat tgcaaagaaa tgagaaagcc
aacagtaaaa ggagagtttc tggaaggaag 600aatcccgcca attccgaggt attagatata
attggcagat cgggagagac atgtcagatg 660gaagacaaca ttgacaacaa gaagttggtc
tctgatagtg acaatgatgc tgaaaaccat 720ataaatgaca atgtccaaag caaaagatat
tgtgcaggaa gacagagtag cagttctaag 780actcgagaag ccagccaaac agaaaccttg
caaaaggtgg ttgacgccaa agaaattaag 840ggggatgcaa ggttttcttt gacaaagcat
tctgactggt taaaatctca agaacctgag 900ccatctgaaa gcctatacga gtcaaggttc
cctttgagaa ggcgttctgc ccggttaaaa 960tctcaagaac ctgagccatc tgaaagcttc
catgactcaa tagagacaac caagaggagg 1020aggtcggcaa taaggtctgc tatgtttaat
atccaagagc tgggcgttat tcaaaacttg 1080aacggtttac ctgatgatca agagattgct
gcaaaggcca gatgctctgc acgtgaacag 1140tctaccgggt ctaaacccga agcagtagaa
ccacatgaca caaaagagat aatcgggaaa 1200agcaggatat ctttgagaag acagtctgcg
aggtttaatt tccaagagct gggcgtgact 1260gaaaacttga atggtccaca tgatgatcaa
acgattgctg caaatgccag atgctgtgca 1320agtgaacagt ctatcgggtc taaacccgaa
gcagtagaac cacatgacat tgaagagaga 1380atcgggaaaa tcagagtctc ttcaagaaga
caatctgcaa acattgaaac tccgagagcc 1440atcaaagaac ctgcaaatcc gcctttgcat
gatgacaatg ttgaggagtc tagtcagata 1500tcatgttcag tttcaatgga gcttaaaaga
gaatcaaaga agaaaccaac aggcgacgaa 1560tcagaggaaa tgagaaaaac aactgttgga
agaccttcaa ggcaagctgc tgaaaaaatc 1620aaatcgtaca aggaaccttc acttaaggag
aagatgcgag ggggcttctg a 167110556PRTArabidopsis thaliana 10Met
Val Arg Ala Thr Val Leu Asn Val Gly Asp His Ala Ser Glu Gly1
5 10 15Val Arg Thr Asn Lys Ala Lys
Gly Glu Lys Met Val Leu Glu Pro Pro 20 25
30Met Asn Ser Ala Gln Arg Arg Lys Leu Gly Asp Ile Thr Asn
Leu Gln 35 40 45Asn Gln Lys Asn
Leu Met Asn Gln Gly Ala Lys His Gln Gln Gln Ala 50 55
60Ile Leu Ile Ser Ser Lys Glu Asn Ala Glu Asn Leu Gln
Lys Ala Leu65 70 75
80Arg Asn Ser Ser Glu Asn Thr Lys Leu Met Lys Val Val Met Glu Arg
85 90 95Asp Gly Ile Lys Ser Asp
Leu Lys Lys Leu Arg Ile Glu Phe Gln Lys 100
105 110Val Gln Glu Gln Asn Leu Leu Leu Ala Gln Ala Asn
Thr Arg Ile Leu 115 120 125Ala Leu
Lys Val Leu Gln His Glu Leu Gly Cys Lys Asn Gly Leu Val 130
135 140Met Ala Arg Lys Met Leu Leu Lys Ala Gln Ala
Asn Ala Cys Gly Gly145 150 155
160Ala Cys Lys Thr Phe Gln Pro Asn Asp Ala Asp His Glu His Ala Ser
165 170 175Gly Ser Ser Asn
Ala Asn Ser Leu Gln Arg Asn Glu Lys Ala Asn Ser 180
185 190Lys Arg Arg Val Ser Gly Arg Lys Asn Pro Ala
Asn Ser Glu Val Leu 195 200 205Asp
Ile Ile Gly Arg Ser Gly Glu Thr Cys Gln Met Glu Asp Asn Ile 210
215 220Asp Asn Lys Lys Leu Val Ser Asp Ser Asp
Asn Asp Ala Glu Asn His225 230 235
240Ile Asn Asp Asn Val Gln Ser Lys Arg Tyr Cys Ala Gly Arg Gln
Ser 245 250 255Ser Ser Ser
Lys Thr Arg Glu Ala Ser Gln Thr Glu Thr Leu Gln Lys 260
265 270Val Val Asp Ala Lys Glu Ile Lys Gly Asp
Ala Arg Phe Ser Leu Thr 275 280
285Lys His Ser Asp Trp Leu Lys Ser Gln Glu Pro Glu Pro Ser Glu Ser 290
295 300Leu Tyr Glu Ser Arg Phe Pro Leu
Arg Arg Arg Ser Ala Arg Leu Lys305 310
315 320Ser Gln Glu Pro Glu Pro Ser Glu Ser Phe His Asp
Ser Ile Glu Thr 325 330
335Thr Lys Arg Arg Arg Ser Ala Ile Arg Ser Ala Met Phe Asn Ile Gln
340 345 350Glu Leu Gly Val Ile Gln
Asn Leu Asn Gly Leu Pro Asp Asp Gln Glu 355 360
365Ile Ala Ala Lys Ala Arg Cys Ser Ala Arg Glu Gln Ser Thr
Gly Ser 370 375 380Lys Pro Glu Ala Val
Glu Pro His Asp Thr Lys Glu Ile Ile Gly Lys385 390
395 400Ser Arg Ile Ser Leu Arg Arg Gln Ser Ala
Arg Phe Asn Phe Gln Glu 405 410
415Leu Gly Val Thr Glu Asn Leu Asn Gly Pro His Asp Asp Gln Thr Ile
420 425 430Ala Ala Asn Ala Arg
Cys Cys Ala Ser Glu Gln Ser Ile Gly Ser Lys 435
440 445Pro Glu Ala Val Glu Pro His Asp Ile Glu Glu Arg
Ile Gly Lys Ile 450 455 460Arg Val Ser
Ser Arg Arg Gln Ser Ala Asn Ile Glu Thr Pro Arg Ala465
470 475 480Ile Lys Glu Pro Ala Asn Pro
Pro Leu His Asp Asp Asn Val Glu Glu 485
490 495Ser Ser Gln Ile Ser Cys Ser Val Ser Met Glu Leu
Lys Arg Glu Ser 500 505 510Lys
Lys Lys Pro Thr Gly Asp Glu Ser Glu Glu Met Arg Lys Thr Thr 515
520 525Val Gly Arg Pro Ser Arg Gln Ala Ala
Glu Lys Ile Lys Ser Tyr Lys 530 535
540Glu Pro Ser Leu Lys Glu Lys Met Arg Gly Gly Phe545 550
555111341DNAArabidopsis thaliana 11atggataaag aagagacgca
gcagaaggaa aatatgctat tctcttccca ggaatatgct 60gcaaagcttc aaaaggcatt
tcctcttcac tttaatcttg aaaacatgac actgatgaaa 120gctctagcac accgaaataa
actcgtcgag ttgagcggta ttgagattca gaaactgagg 180attaacttac ggagtgtgca
ggaaaagaat ttgcagcttg ctcaggcaaa cagtcagatg 240ttagcgctca aggatctcca
gcatgaactt ggctgcaaga atgctttact taaagtcaag 300aaacatcttg aggagcaagt
acttccacgt acacatcatg aatcgaaaga caaggtttca 360gcaagcgctt ctgatgggga
ttgcaaatcc tttcaggtgc atgacataaa acataaagat 420accaagagaa agcgaacaac
aaggataaaa tcttcagtaa gtgccgacgt caagccaata 480cctgtgaatg attctaacag
taaagctaac cgtaaaagaa gagtttctgg agtaatagat 540actactggta ttcccgaaga
gatctgtcag actgaagatg acattgataa gggggttgtc 600tctcgagggg taaaccaaga
tattgacaat gttgtcaaca agaagtttgt tcctgatgca 660gcaaacccgg taaaagagag
tgtgcatcgc aagaggcaat gtacacgaag gcaatctacc 720agatttgatg ttcaagaaac
taaacaaacg gaaaagttgc ttgagatgga tggtgccaaa 780gaaagtaaag aaaccgcaag
cttctctttg agaagacggt ctgctcggtt aaggcacgaa 840gaagctgaac catgtaaaag
cttacatgag ggagacgaag tcagggagac aatcaagagg 900agaagagtct ctttaagact
gtctgcaagg tttgatatac aagaaccgca tgtgactgaa 960acctcgaatg ctgacgatgc
aagaagcata gtaatcgaag aatctgctgg atcaagatcg 1020gaatctgtag aaccatccga
aagcaggcat gaaacaaaag agataacccg gaaacgcagt 1080ttctcaacga gaagacaatc
aacaaagggt aaatctcaaa ccgatgaagc cattaaagaa 1140atagcgacag acccatcttt
ggtcaacacc atagttcaag agtgtgatca ggaaacagaa 1200tcaaaggata agcctaaagc
tgatgaaaac gaagggatga caagaagatc atctgtggga 1260agaccatcga gacatgccgc
agagaaagtc caatcataca gagaagtctc acttagagta 1320aagatgcgac gaaaatgcta a
134112446PRTArabidopsis
thaliana 12Met Asp Lys Glu Glu Thr Gln Gln Lys Glu Asn Met Leu Phe Ser
Ser1 5 10 15Gln Glu Tyr
Ala Ala Lys Leu Gln Lys Ala Phe Pro Leu His Phe Asn 20
25 30Leu Glu Asn Met Thr Leu Met Lys Ala Leu
Ala His Arg Asn Lys Leu 35 40
45Val Glu Leu Ser Gly Ile Glu Ile Gln Lys Leu Arg Ile Asn Leu Arg 50
55 60Ser Val Gln Glu Lys Asn Leu Gln Leu
Ala Gln Ala Asn Ser Gln Met65 70 75
80Leu Ala Leu Lys Asp Leu Gln His Glu Leu Gly Cys Lys Asn
Ala Leu 85 90 95Leu Lys
Val Lys Lys His Leu Glu Glu Gln Val Leu Pro Arg Thr His 100
105 110His Glu Ser Lys Asp Lys Val Ser Ala
Ser Ala Ser Asp Gly Asp Cys 115 120
125Lys Ser Phe Gln Val His Asp Ile Lys His Lys Asp Thr Lys Arg Lys
130 135 140Arg Thr Thr Arg Ile Lys Ser
Ser Val Ser Ala Asp Val Lys Pro Ile145 150
155 160Pro Val Asn Asp Ser Asn Ser Lys Ala Asn Arg Lys
Arg Arg Val Ser 165 170
175Gly Val Ile Asp Thr Thr Gly Ile Pro Glu Glu Ile Cys Gln Thr Glu
180 185 190Asp Asp Ile Asp Lys Gly
Val Val Ser Arg Gly Val Asn Gln Asp Ile 195 200
205Asp Asn Val Val Asn Lys Lys Phe Val Pro Asp Ala Ala Asn
Pro Val 210 215 220Lys Glu Ser Val His
Arg Lys Arg Gln Cys Thr Arg Arg Gln Ser Thr225 230
235 240Arg Phe Asp Val Gln Glu Thr Lys Gln Thr
Glu Lys Leu Leu Glu Met 245 250
255Asp Gly Ala Lys Glu Ser Lys Glu Thr Ala Ser Phe Ser Leu Arg Arg
260 265 270Arg Ser Ala Arg Leu
Arg His Glu Glu Ala Glu Pro Cys Lys Ser Leu 275
280 285His Glu Gly Asp Glu Val Arg Glu Thr Ile Lys Arg
Arg Arg Val Ser 290 295 300Leu Arg Leu
Ser Ala Arg Phe Asp Ile Gln Glu Pro His Val Thr Glu305
310 315 320Thr Ser Asn Ala Asp Asp Ala
Arg Ser Ile Val Ile Glu Glu Ser Ala 325
330 335Gly Ser Arg Ser Glu Ser Val Glu Pro Ser Glu Ser
Arg His Glu Thr 340 345 350Lys
Glu Ile Thr Arg Lys Arg Ser Phe Ser Thr Arg Arg Gln Ser Thr 355
360 365Lys Gly Lys Ser Gln Thr Asp Glu Ala
Ile Lys Glu Ile Ala Thr Asp 370 375
380Pro Ser Leu Val Asn Thr Ile Val Gln Glu Cys Asp Gln Glu Thr Glu385
390 395 400Ser Lys Asp Lys
Pro Lys Ala Asp Glu Asn Glu Gly Met Thr Arg Arg 405
410 415Ser Ser Val Gly Arg Pro Ser Arg His Ala
Ala Glu Lys Val Gln Ser 420 425
430Tyr Arg Glu Val Ser Leu Arg Val Lys Met Arg Arg Lys Cys 435
440 445131554DNAmouse 13atggctaagg
aaaggtgtca gaaaaggtcc tttcaagata cccttgaaga cattaagaat 60cgaatgaaag
aaaaaaggaa taaaaatttg gcggggattg ggaaacgcaa gtcctttatt 120gttgcaccgg
gccaagtacc cactaacact gctacactac tgagatatta ccaagataac 180aacaggttgt
tagtcttggc tttggaaaat gagaaatcca aagtgagaga agcacaggat 240gtcatcctgc
aactgagaaa agaatgctac taccttactt gtcagctgta tgcattgaaa 300gagaagctaa
cttcccgaca aagtgaagaa actactcaga actggaaagg acgtccctca 360gacgtggtct
ccagcattga caatacgacc agggacttgt cagggaagtc cttacagcaa 420attgctgttg
aagaaactga ttgtccttac caaaccacag aaccaagtcc tgctgttact 480ccagagacac
agggttgcga ttttgattca ggtaaagttg agtctactga tgaagtctta 540cccagaacta
tatctatccg tcgccattta aggaaagatt ttagtaatat aagccactcc 600acgactttgg
aggattgtaa agccagtcca agagtggcac agtctctgga agttaaagga 660agtagatgta
gagaagtaac cgtaaccctg cacagacttg aaaatgtttg tctgtggaac 720aaagaccaaa
ttagcttatg ttctagactg attaacccag caaagattac tgaaacagaa 780gtcattttat
catctaaacc tgaacaaata gaaagcaagc ataaacgtgc acgaaaaaga 840agagcagagc
aaagaagaac caagcagaga tgcaaatcaa aatcctcatt gaggagtaag 900gggaacaaaa
acaaagataa gcagggttta ccccctacta cactggatgg aggtattggt 960tcctgtgatg
cttacgattt taatctaaaa gggacggtcc accccacccc tttccgacaa 1020aaaatgaaca
atggctgcaa caaagaaacg gatagcagca actcagaagt gagtgacctc 1080gaatgcagta
cctctgagga tgagtctgat gacctctacc tgcctccctc caagcgcttg 1140cgagactaca
gagagtcaga gagagcagtt accaggcctc ggtctaaaag aggacttcag 1200tacccagatg
ggaaagagag gaaggaggtg ctgccatcta cagctcctac tggtatccca 1260cctgagactc
aagagtcacc tcgttgtagc ctaaaggatg tcaccaatat cctgcagtgt 1320cctagagtga
agatcaggaa gccttctctg cctccaaagc ggcgtgaaga cagcccagca 1380gtggctctga
ctaaacgcag gtgtagcacc atcaaaagct ataaagagcc aacactcgct 1440tcaaagctaa
gaagagggga ccctttcacg gacttgtgtt tcttgaattc tcctattttc 1500aagcagaaaa
ggggtatgag atgtcctaaa agaagaacca agcaaacaca gtaa
155414517PRTmosue 14Met Ala Lys Glu Arg Cys Gln Lys Arg Ser Phe Gln Asp
Thr Leu Glu1 5 10 15Asp
Ile Lys Asn Arg Met Lys Glu Lys Arg Asn Lys Asn Leu Ala Gly 20
25 30Ile Gly Lys Arg Lys Ser Phe Ile
Val Ala Pro Gly Gln Val Pro Thr 35 40
45Asn Thr Ala Thr Leu Leu Arg Tyr Tyr Gln Asp Asn Asn Arg Leu Leu
50 55 60Val Leu Ala Leu Glu Asn Glu Lys
Ser Lys Val Arg Glu Ala Gln Asp65 70 75
80Val Ile Leu Gln Leu Arg Lys Glu Cys Tyr Tyr Leu Thr
Cys Gln Leu 85 90 95Tyr
Ala Leu Lys Glu Lys Leu Thr Ser Arg Gln Ser Glu Glu Thr Thr
100 105 110Gln Asn Trp Lys Gly Arg Pro
Ser Asp Val Val Ser Ser Ile Asp Asn 115 120
125Thr Thr Arg Asp Leu Ser Gly Lys Ser Leu Gln Gln Ile Ala Val
Glu 130 135 140Glu Thr Asp Cys Pro Tyr
Gln Thr Thr Glu Pro Ser Pro Ala Val Thr145 150
155 160Pro Glu Thr Gln Gly Cys Asp Phe Asp Ser Gly
Lys Val Glu Ser Thr 165 170
175Asp Glu Val Leu Pro Arg Thr Ile Ser Ile Arg Arg His Leu Arg Lys
180 185 190Asp Phe Ser Asn Ile Ser
His Ser Thr Thr Leu Glu Asp Cys Lys Ala 195 200
205Ser Pro Arg Val Ala Gln Ser Leu Glu Val Lys Gly Ser Arg
Cys Arg 210 215 220Glu Val Thr Val Thr
Leu His Arg Leu Glu Asn Val Cys Leu Trp Asn225 230
235 240Lys Asp Gln Ile Ser Leu Cys Ser Arg Leu
Ile Asn Pro Ala Lys Ile 245 250
255Thr Glu Thr Glu Val Ile Leu Ser Ser Lys Pro Glu Gln Ile Glu Ser
260 265 270Lys His Lys Arg Ala
Arg Lys Arg Arg Ala Glu Gln Arg Arg Thr Lys 275
280 285Gln Arg Cys Lys Ser Lys Ser Ser Leu Arg Ser Lys
Gly Asn Lys Asn 290 295 300Lys Asp Lys
Gln Gly Leu Pro Pro Thr Thr Leu Asp Gly Gly Ile Gly305
310 315 320Ser Cys Asp Ala Tyr Asp Phe
Asn Leu Lys Gly Thr Val His Pro Thr 325
330 335Pro Phe Arg Gln Lys Met Asn Asn Gly Cys Asn Lys
Glu Thr Asp Ser 340 345 350Ser
Asn Ser Glu Val Ser Asp Leu Glu Cys Ser Thr Ser Glu Asp Glu 355
360 365Ser Asp Asp Leu Tyr Leu Pro Pro Ser
Lys Arg Leu Arg Asp Tyr Arg 370 375
380Glu Ser Glu Arg Ala Val Thr Arg Pro Arg Ser Lys Arg Gly Leu Gln385
390 395 400Tyr Pro Asp Gly
Lys Glu Arg Lys Glu Val Leu Pro Ser Thr Ala Pro 405
410 415Thr Gly Ile Pro Pro Glu Thr Gln Glu Ser
Pro Arg Cys Ser Leu Lys 420 425
430Asp Val Thr Asn Ile Leu Gln Cys Pro Arg Val Lys Ile Arg Lys Pro
435 440 445Ser Leu Pro Pro Lys Arg Arg
Glu Asp Ser Pro Ala Val Ala Leu Thr 450 455
460Lys Arg Arg Cys Ser Thr Ile Lys Ser Tyr Lys Glu Pro Thr Leu
Ala465 470 475 480Ser Lys
Leu Arg Arg Gly Asp Pro Phe Thr Asp Leu Cys Phe Leu Asn
485 490 495Ser Pro Ile Phe Lys Gln Lys
Arg Gly Met Arg Cys Pro Lys Arg Arg 500 505
510Thr Lys Gln Thr Gln 515153495DNAmouse 15atggagtacc
cagggataaa agttgacact gttacctctg gaattcagag acgagtgaag 60ggcagaattg
caaagacaaa tttgaatgtt tctcttgctt caaagatcaa agcaaaaata 120ttaaacaatt
cttctatttt caagatctct ctaaagcaca acaacagagc attagcgcgg 180gcccttagta
aagagaaaga gaattctcga agaattacta ccgaaaagat gcaattacag 240aaagaagtag
agaaactgaa ttttgagaat acctttcttc gcttaaagtt aaataccttg 300aataagaagc
ttgtagaaat agaatcgcat gtgagcaatg atttgttaac tgcaattgaa 360ataagcagtc
tttctgagtt ccaccaaggt tcttttctcc tgtcagctac caagaaacaa 420aggaacagta
agcagtgcaa gcctgcgcat cttccatatg caagagttct gttaacttca 480gaaaatgatg
atgatgatgg tgctgatgat aaatggcaga caaagtgtaa caacagaact 540atatcaaaga
cctcacctga tagtacctct tcagtatcaa gacaaccttc atccttacat 600cagtgcaatt
tgaaagcatt ccctcctaaa gaagataatc agaagacatg tgggtcaggt 660catttagaac
atacttcaag tgttgatata cttcctaatg agagccactc agatcaaagt 720cctaagagtt
ctctgagtga gatgaaaact gctccatctc ccagcctcag aagggaaaaa 780ttatcacatg
gtaatgtgac tatgaggaag aagtgtgtgt cttcaactcc agacattctg 840tatgtgacag
atttagatca ccaaccaact tcaagtccag gatcaaattg gaataatgag 900atacatggtc
atactaatga aaccagcaat aacacgcaaa gaaatgccga gtgttttctt 960gacttacctt
ctgagtcttc cagtgagcct gacgcaaagc gcatggagct agtgcagaag 1020aacaccgata
gctttcactt ccagaaaact gtatatgatg ccgctgatat ggagttaact 1080gctactgaca
taggcaagat tgtagcagtt tcaaaaagca agaaaaatca aaataagaaa 1140aaggcagact
gtagaaagga gactttcaga aaagtgaaag gtgcaagctc tgataaaaag 1200agagaaagct
caaagagaga atgtaaagat ggttcagaag taggtgctga ggaagaggct 1260gatgcagcca
gagcagaaag aggcgctggt gtcctggatg gcagagggga ttcagaagag 1320ccaaactgca
tttccagtac tgagcagcca tctcaggtaa acacgcaaaa gaaaagaacc 1380ctccagaaca
gctcagatca ggagaacatt caaaatacga agaggaggca aacatatacg 1440acagatgagc
aagaggaaac aaaccctttc tccagacatt cagtcaaatt tcttcaagat 1500ggtaaatttg
atctgtgtca gaaaacccta catcataatt taagtaagcc ttctcgacag 1560acatttgtga
ttcgtaagtc agaaaaagat aacttatttc caaatcaaga agataaagac 1620accatttctg
aaaacctaga agttacaaat gaatttcata tagatgatct ttccatcgaa 1680gctaatgaaa
atgtatgtga ccatgagact cagacaatgt tggacttgaa aaagtctgtc 1740agtgctcaac
aaaatcaaac aaaaataaat aagactaagc agaaaataaa tcgaaggaca 1800aaaataattt
ctgtcatgag ccaagtatat gaggacaatg ataaagatat tcacgtccta 1860gaaaaagaca
actttccctt tcatacccaa gcaaataaag aaaccaccag tggaaaccta 1920gaaagttcaa
aagaatttga atcacctctt cttttcacaa gagacaacgg aagcttacgt 1980gactgtaaga
cccagaatgt tctggatctg cacaagcaaa ttcctgatct ataccctgat 2040cggaatgagt
cccagattag caaaatccct aggcaaaaag taaatcgcaa gacagaagta 2100atttctggag
tgaaatgttt tagtaatgac caaggtgttc attgctcaga aaaggataag 2160tctttgttac
tacaaaagga taaagacttc ccaggaactt taaaagactt aagtgagttt 2220gatacgcctg
ctttttgtaa caaagatagt gcaaagtcgt gtgattataa gtctgaaatg 2280ctcttggggt
tgaaaaaaca tgaccctaat atgcaacctg cttgtcaaga tgattcaaaa 2340gcaggtaaga
aacttagaca aaaggtaaat cgaaaaacag aaataatttc taaaatcacc 2400caaatacatg
aaaatgatag aggaagtaca catgactcat taaataagaa gctctgtcag 2460aaggttaata
tatcaaaaat catttctcaa atgaaccaaa tatatgagac tattaatgaa 2520gatggaaatg
gctttaaaag ctctatcaaa gattgcgaag atattaaaag ttgtgacttt 2580ggggaaatca
acagtaataa aaaggaaaat tatgatccaa ttcaagatcc ttgcacactg 2640gttaaaaaaa
caaagagaaa gggatcatgt aaagcaggga gcagtttggc aggagctaag 2700aacaggtgtg
gtttgcagtt aacagactct tcccaggtac agtctgtccc cttagactct 2760ggcttaagac
accatccaaa cgaagcagat tctggtcctg gagagcagac taacctgcca 2820aagatgcaga
aacaaagcgc tgggaggtca ctgggagatg ctttctctgt gagtctggga 2880aaagaaggaa
gccgcccagc caaagcagtt agtaaaatga cacccaaatc aaagaagaga 2940aagctccctc
tcggttgttc tcctgaaacc cacgggacgg tggagataac acccaacact 3000gacctcgcta
aggctgttga ctcccaacag actgagaagg agaactattt ggagaaggag 3060aaaattgcca
agaggaagcc agatttttgt acaaaggtgt tgaaaccttt atctgagaca 3120tgttcatcta
acataaagaa ttcttccttg gacagtatgt gtaagagttc gctacctttg 3180agtatttctt
ctagaaaaac cctgatgctg gaagaaagtt cttccctgga gagtacatgc 3240atctttcaag
taggtgatgc cgctcatgag aagataacga caggcacacg taatccccac 3300cacaggacac
agaagtcgac accgggtagc agaacgtccc tggtcttggt ggataccagt 3360tctgtttcag
ataccaaccc tgctaacccc gagaatgagt cagaagggca gtcttcacac 3420ccaatgagaa
ggaaaagaca gtgcgtccct ctcaacctga cagagccaag ccttagaagc 3480aagatgagga
gataa
3495161164PRTmouse 16Met Glu Tyr Pro Gly Ile Lys Val Asp Thr Val Thr Ser
Gly Ile Gln1 5 10 15Arg
Arg Val Lys Gly Arg Ile Ala Lys Thr Asn Leu Asn Val Ser Leu 20
25 30Ala Ser Lys Ile Lys Ala Lys Ile
Leu Asn Asn Ser Ser Ile Phe Lys 35 40
45Ile Ser Leu Lys His Asn Asn Arg Ala Leu Ala Arg Ala Leu Ser Lys
50 55 60Glu Lys Glu Asn Ser Arg Arg Ile
Thr Thr Glu Lys Met Gln Leu Gln65 70 75
80Lys Glu Val Glu Lys Leu Asn Phe Glu Asn Thr Phe Leu
Arg Leu Lys 85 90 95Leu
Asn Thr Leu Asn Lys Lys Leu Val Glu Ile Glu Ser His Val Ser
100 105 110Asn Asp Leu Leu Thr Ala Ile
Glu Ile Ser Ser Leu Ser Glu Phe His 115 120
125Gln Gly Ser Phe Leu Leu Ser Ala Thr Lys Lys Gln Arg Asn Ser
Lys 130 135 140Gln Cys Lys Pro Ala His
Leu Pro Tyr Ala Arg Val Leu Leu Thr Ser145 150
155 160Glu Asn Asp Asp Asp Asp Gly Ala Asp Asp Lys
Trp Gln Thr Lys Cys 165 170
175Asn Asn Arg Thr Ile Ser Lys Thr Ser Pro Asp Ser Thr Ser Ser Val
180 185 190Ser Arg Gln Pro Ser Ser
Leu His Gln Cys Asn Leu Lys Ala Phe Pro 195 200
205Pro Lys Glu Asp Asn Gln Lys Thr Cys Gly Ser Gly His Leu
Glu His 210 215 220Thr Ser Ser Val Asp
Ile Leu Pro Asn Glu Ser His Ser Asp Gln Ser225 230
235 240Pro Lys Ser Ser Leu Ser Glu Met Lys Thr
Ala Pro Ser Pro Ser Leu 245 250
255Arg Arg Glu Lys Leu Ser His Gly Asn Val Thr Met Arg Lys Lys Cys
260 265 270Val Ser Ser Thr Pro
Asp Ile Leu Tyr Val Thr Asp Leu Asp His Gln 275
280 285Pro Thr Ser Ser Pro Gly Ser Asn Trp Asn Asn Glu
Ile His Gly His 290 295 300Thr Asn Glu
Thr Ser Asn Asn Thr Gln Arg Asn Ala Glu Cys Phe Leu305
310 315 320Asp Leu Pro Ser Glu Ser Ser
Ser Glu Pro Asp Ala Lys Arg Met Glu 325
330 335Leu Val Gln Lys Asn Thr Asp Ser Phe His Phe Gln
Lys Thr Val Tyr 340 345 350Asp
Ala Ala Asp Met Glu Leu Thr Ala Thr Asp Ile Gly Lys Ile Val 355
360 365Ala Val Ser Lys Ser Lys Lys Asn Gln
Asn Lys Lys Lys Ala Asp Cys 370 375
380Arg Lys Glu Thr Phe Arg Lys Val Lys Gly Ala Ser Ser Asp Lys Lys385
390 395 400Arg Glu Ser Ser
Lys Arg Glu Cys Lys Asp Gly Ser Glu Val Gly Ala 405
410 415Glu Glu Glu Ala Asp Ala Ala Arg Ala Glu
Arg Gly Ala Gly Val Leu 420 425
430Asp Gly Arg Gly Asp Ser Glu Glu Pro Asn Cys Ile Ser Ser Thr Glu
435 440 445Gln Pro Ser Gln Val Asn Thr
Gln Lys Lys Arg Thr Leu Gln Asn Ser 450 455
460Ser Asp Gln Glu Asn Ile Gln Asn Thr Lys Arg Arg Gln Thr Tyr
Thr465 470 475 480Thr Asp
Glu Gln Glu Glu Thr Asn Pro Phe Ser Arg His Ser Val Lys
485 490 495Phe Leu Gln Asp Gly Lys Phe
Asp Leu Cys Gln Lys Thr Leu His His 500 505
510Asn Leu Ser Lys Pro Ser Arg Gln Thr Phe Val Ile Arg Lys
Ser Glu 515 520 525Lys Asp Asn Leu
Phe Pro Asn Gln Glu Asp Lys Asp Thr Ile Ser Glu 530
535 540Asn Leu Glu Val Thr Asn Glu Phe His Ile Asp Asp
Leu Ser Ile Glu545 550 555
560Ala Asn Glu Asn Val Cys Asp His Glu Thr Gln Thr Met Leu Asp Leu
565 570 575Lys Lys Ser Val Ser
Ala Gln Gln Asn Gln Thr Lys Ile Asn Lys Thr 580
585 590Lys Gln Lys Ile Asn Arg Arg Thr Lys Ile Ile Ser
Val Met Ser Gln 595 600 605Val Tyr
Glu Asp Asn Asp Lys Asp Ile His Val Leu Glu Lys Asp Asn 610
615 620Phe Pro Phe His Thr Gln Ala Asn Lys Glu Thr
Thr Ser Gly Asn Leu625 630 635
640Glu Ser Ser Lys Glu Phe Glu Ser Pro Leu Leu Phe Thr Arg Asp Asn
645 650 655Gly Ser Leu Arg
Asp Cys Lys Thr Gln Asn Val Leu Asp Leu His Lys 660
665 670Gln Ile Pro Asp Leu Tyr Pro Asp Arg Asn Glu
Ser Gln Ile Ser Lys 675 680 685Ile
Pro Arg Gln Lys Val Asn Arg Lys Thr Glu Val Ile Ser Gly Val 690
695 700Lys Cys Phe Ser Asn Asp Gln Gly Val His
Cys Ser Glu Lys Asp Lys705 710 715
720Ser Leu Leu Leu Gln Lys Asp Lys Asp Phe Pro Gly Thr Leu Lys
Asp 725 730 735Leu Ser Glu
Phe Asp Thr Pro Ala Phe Cys Asn Lys Asp Ser Ala Lys 740
745 750Ser Cys Asp Tyr Lys Ser Glu Met Leu Leu
Gly Leu Lys Lys His Asp 755 760
765Pro Asn Met Gln Pro Ala Cys Gln Asp Asp Ser Lys Ala Gly Lys Lys 770
775 780Leu Arg Gln Lys Val Asn Arg Lys
Thr Glu Ile Ile Ser Lys Ile Thr785 790
795 800Gln Ile His Glu Asn Asp Arg Gly Ser Thr His Asp
Ser Leu Asn Lys 805 810
815Lys Leu Cys Gln Lys Val Asn Ile Ser Lys Ile Ile Ser Gln Met Asn
820 825 830Gln Ile Tyr Glu Thr Ile
Asn Glu Asp Gly Asn Gly Phe Lys Ser Ser 835 840
845Ile Lys Asp Cys Glu Asp Ile Lys Ser Cys Asp Phe Gly Glu
Ile Asn 850 855 860Ser Asn Lys Lys Glu
Asn Tyr Asp Pro Ile Gln Asp Pro Cys Thr Leu865 870
875 880Val Lys Lys Thr Lys Arg Lys Gly Ser Cys
Lys Ala Gly Ser Ser Leu 885 890
895Ala Gly Ala Lys Asn Arg Cys Gly Leu Gln Leu Thr Asp Ser Ser Gln
900 905 910Val Gln Ser Val Pro
Leu Asp Ser Gly Leu Arg His His Pro Asn Glu 915
920 925Ala Asp Ser Gly Pro Gly Glu Gln Thr Asn Leu Pro
Lys Met Gln Lys 930 935 940Gln Ser Ala
Gly Arg Ser Leu Gly Asp Ala Phe Ser Val Ser Leu Gly945
950 955 960Lys Glu Gly Ser Arg Pro Ala
Lys Ala Val Ser Lys Met Thr Pro Lys 965
970 975Ser Lys Lys Arg Lys Leu Pro Leu Gly Cys Ser Pro
Glu Thr His Gly 980 985 990Thr
Val Glu Ile Thr Pro Asn Thr Asp Leu Ala Lys Ala Val Asp Ser 995
1000 1005Gln Gln Thr Glu Lys Glu Asn Tyr
Leu Glu Lys Glu Lys Ile Ala 1010 1015
1020Lys Arg Lys Pro Asp Phe Cys Thr Lys Val Leu Lys Pro Leu Ser
1025 1030 1035Glu Thr Cys Ser Ser Asn
Ile Lys Asn Ser Ser Leu Asp Ser Met 1040 1045
1050Cys Lys Ser Ser Leu Pro Leu Ser Ile Ser Ser Arg Lys Thr
Leu 1055 1060 1065Met Leu Glu Glu Ser
Ser Ser Leu Glu Ser Thr Cys Ile Phe Gln 1070 1075
1080Val Gly Asp Ala Ala His Glu Lys Ile Thr Thr Gly Thr
Arg Asn 1085 1090 1095Pro His His Arg
Thr Gln Lys Ser Thr Pro Gly Ser Arg Thr Ser 1100
1105 1110Leu Val Leu Val Asp Thr Ser Ser Val Ser Asp
Thr Asn Pro Ala 1115 1120 1125Asn Pro
Glu Asn Glu Ser Glu Gly Gln Ser Ser His Pro Met Arg 1130
1135 1140Arg Lys Arg Gln Cys Val Pro Leu Asn Leu
Thr Glu Pro Ser Leu 1145 1150 1155Arg
Ser Lys Met Arg Arg 1160171584DNAHomo sapiens 17atggccaagg aaagatgcct
gaaaaagtcc tttcaagata gtcttgaaga cataaagaag 60cgaatgaaag agaaaaggaa
taaaaacttg gcagagattg gcaaacgcag gtcttttata 120gctgcaccat gccaaataat
caccaacact tctacactgc tgaaaaatta ccaagacaac 180aacaaaatgt tagttttagc
tttggaaaat gaaaaatcca aagtgaaaga agcccaagat 240atcatcctac agctgagaaa
agaatgttac tatctcacat gtcagctata tgcattgaaa 300ggaaaactta catcacaaca
aacagtagaa cctgctcaga accaggaaat atgttcctct 360ggaatggacc ccaatagtga
tgacagctcc agaaatttat ttgtgaagga tttaccgcaa 420attcctcttg aagaaactga
acttccagga caaggagaat catttcaaat agaagatcag 480atacctacta ttcctcaaga
cacactggga gttgattttg attcaggtga agctaagtct 540actgataatg tcttacctag
aactgtatct gttcgtagca gtttaaagaa acattgtaac 600agtatatgtc agtttgatag
cttggatgat tttgaaacca gtcatttggc agggaagtct 660tttgaattcg aaagagttgg
atttttagac ccactagtaa acatgcacat acctgaaaat 720gtacaacaca atgcttgtca
atggagcaag gaccaagtta acttatcacc aaagctgatt 780cagccaggaa cgtttactaa
aacaaaagaa gacattttag aatctaaatc tgaacaaact 840aaaagtaagc aaagagatac
acaagaaaga aaaagagaag agaaaagaaa agctaacagg 900agaaaatcaa aacgtatgtc
aaaatataaa gagaataaaa gcgaaaataa aaaaactgtt 960ccccaaaaaa aaatgcacaa
atctgtcagt tccaatgatg cttacaattt taatttggaa 1020gagggtgttc atcttactcc
tttccgacaa aaagtgagca atgactctaa tagagaagaa 1080aacaacgagt ctgaagtgag
cctctgtgaa tcaagtggtt caggagatga ttccgatgac 1140ctctatttgc ccacttgcaa
gtacattcag aatcccacga gcaattcaga tagaccagtc 1200accaggcctc tagctaaaag
agcactgaaa tacacagatg aaaaagagac ggagggttct 1260aagccaacaa aaactcctac
cactacacca cctgaaactc agcagtcacc tcatcttagc 1320ctgaaggata tcaccaatgt
ctccttgtat cctgttgtga aaatcagaag actttctctt 1380tctccaaaaa agaataaagc
aagcccagca gtggctctgc ctaaacgtag gtgcacagcc 1440agcgtgaact ataaggagcc
caccctcgct tcgaaactga gaagagggga cccttttaca 1500gatttgtgtt ttttgaattc
tcctattttc aagcagaaaa aggatttgag acgttctaaa 1560aaaagtatga aacaaataca
atga 158418527PRTHomo sapiens
18Met Ala Lys Glu Arg Cys Leu Lys Lys Ser Phe Gln Asp Ser Leu Glu1
5 10 15Asp Ile Lys Lys Arg Met
Lys Glu Lys Arg Asn Lys Asn Leu Ala Glu 20 25
30Ile Gly Lys Arg Arg Ser Phe Ile Ala Ala Pro Cys Gln
Ile Ile Thr 35 40 45Asn Thr Ser
Thr Leu Leu Lys Asn Tyr Gln Asp Asn Asn Lys Met Leu 50
55 60Val Leu Ala Leu Glu Asn Glu Lys Ser Lys Val Lys
Glu Ala Gln Asp65 70 75
80Ile Ile Leu Gln Leu Arg Lys Glu Cys Tyr Tyr Leu Thr Cys Gln Leu
85 90 95Tyr Ala Leu Lys Gly Lys
Leu Thr Ser Gln Gln Thr Val Glu Pro Ala 100
105 110Gln Asn Gln Glu Ile Cys Ser Ser Gly Met Asp Pro
Asn Ser Asp Asp 115 120 125Ser Ser
Arg Asn Leu Phe Val Lys Asp Leu Pro Gln Ile Pro Leu Glu 130
135 140Glu Thr Glu Leu Pro Gly Gln Gly Glu Ser Phe
Gln Ile Glu Asp Gln145 150 155
160Ile Pro Thr Ile Pro Gln Asp Thr Leu Gly Val Asp Phe Asp Ser Gly
165 170 175Glu Ala Lys Ser
Thr Asp Asn Val Leu Pro Arg Thr Val Ser Val Arg 180
185 190Ser Ser Leu Lys Lys His Cys Asn Ser Ile Cys
Gln Phe Asp Ser Leu 195 200 205Asp
Asp Phe Glu Thr Ser His Leu Ala Gly Lys Ser Phe Glu Phe Glu 210
215 220Arg Val Gly Phe Leu Asp Pro Leu Val Asn
Met His Ile Pro Glu Asn225 230 235
240Val Gln His Asn Ala Cys Gln Trp Ser Lys Asp Gln Val Asn Leu
Ser 245 250 255Pro Lys Leu
Ile Gln Pro Gly Thr Phe Thr Lys Thr Lys Glu Asp Ile 260
265 270Leu Glu Ser Lys Ser Glu Gln Thr Lys Ser
Lys Gln Arg Asp Thr Gln 275 280
285Glu Arg Lys Arg Glu Glu Lys Arg Lys Ala Asn Arg Arg Lys Ser Lys 290
295 300Arg Met Ser Lys Tyr Lys Glu Asn
Lys Ser Glu Asn Lys Lys Thr Val305 310
315 320Pro Gln Lys Lys Met His Lys Ser Val Ser Ser Asn
Asp Ala Tyr Asn 325 330
335Phe Asn Leu Glu Glu Gly Val His Leu Thr Pro Phe Arg Gln Lys Val
340 345 350Ser Asn Asp Ser Asn Arg
Glu Glu Asn Asn Glu Ser Glu Val Ser Leu 355 360
365Cys Glu Ser Ser Gly Ser Gly Asp Asp Ser Asp Asp Leu Tyr
Leu Pro 370 375 380Thr Cys Lys Tyr Ile
Gln Asn Pro Thr Ser Asn Ser Asp Arg Pro Val385 390
395 400Thr Arg Pro Leu Ala Lys Arg Ala Leu Lys
Tyr Thr Asp Glu Lys Glu 405 410
415Thr Glu Gly Ser Lys Pro Thr Lys Thr Pro Thr Thr Thr Pro Pro Glu
420 425 430Thr Gln Gln Ser Pro
His Leu Ser Leu Lys Asp Ile Thr Asn Val Ser 435
440 445Leu Tyr Pro Val Val Lys Ile Arg Arg Leu Ser Leu
Ser Pro Lys Lys 450 455 460Asn Lys Ala
Ser Pro Ala Val Ala Leu Pro Lys Arg Arg Cys Thr Ala465
470 475 480Ser Val Asn Tyr Lys Glu Pro
Thr Leu Ala Ser Lys Leu Arg Arg Gly 485
490 495Asp Pro Phe Thr Asp Leu Cys Phe Leu Asn Ser Pro
Ile Phe Lys Gln 500 505 510Lys
Lys Asp Leu Arg Arg Ser Lys Lys Ser Met Lys Gln Ile Gln 515
520 525193798DNAHomo sapiens 19atggagtgcc
cagtgatgga aactggctca ctttttacct caggaattaa gagacatttg 60aaagacaaaa
gaatttcaaa gactactaag ttgaatgttt ctcttgcttc aaaaataaaa 120acaaaaatac
taaataattc ttctattttc aaaatatctt taaagcacaa caacagggca 180ttagctcagg
ctcttagtag agaaaaagag aattctcgaa gaattacaac tgaaaagatg 240ctattgcaaa
aagaagtaga gaaactgaat tttgagaaca catttcttcg cctaaagcta 300aataacttga
ataagaagct tatagacata gaagctctca tgaacaataa cttgataact 360gcaactgaaa
tgagcagtct ttctgagttc catcagagtt cctttctact gtcagctagc 420aagaagaaac
gagttagtaa acagtgcaag ttgatgcgtc ttccatttgc aagggttcca 480ttaacttcaa
atgatgatga agatgaagat aaagagaaaa tgcagtgtga caacaatatt 540aaatcaaaga
cattacctga tattccctct tcaggatcaa caacacaacc tttatcaact 600caggataatt
cggaagtgtt atttcttaaa gaaaataatc aaaatgtata tggtttagat 660gattcagaac
atatttcttc tatagttgat gtacctccca gagaaagcca ttcccactca 720gaccaaagtt
ctaagacttc tctaatgagt gagatgagaa acgcccagtc tattggccgc 780agatgggaga
aaccatctcc tagtaatgtg actgaaagga agaagcgtgg gtcatcttgg 840gaatcaaata
atctttctgc agacactccc tgtgcaacag ttttagataa acaacacatt 900tcaagtccag
aattaaattg caataatgag ataaatggtc atactaatga aacaaatact 960gaaatgcaaa
gaaataaaca ggatcttcct ggcttatctt ctgagtctgc cagagaacct 1020aatgcagagt
gcatgaatca aattgaggat aatgatgact ttcaattgca gaaaactgtg 1080tatgatgctg
acatggattt aactgctagt gaagtcagca aaattgtcac agtctcaaca 1140ggcattaaaa
agaaaagtaa taaaaaaaca aatgaacatg gaatgaaaac tttcagaaaa 1200gtgaaagatt
ccagctctga aaaaaagaga gaaagatcaa agagacagtt taaaaatagt 1260tcagatgtcg
atattgggga aaagattgaa aacaggacag aaagatctga tgtcctggat 1320ggcaaaaggg
gtgcagaaga tcccggtttt attttcaata atgaacagct ggctcagatg 1380aatgaacagc
tggctcaggt gaatgaacta aagaaaatga cccttcaaac tggctttgaa 1440caaggtgaca
gagaaaatgt actgtgtaat aaaaaggaga aaagaataac aaatgagcaa 1500gaggaaacat
actctttatc ccaaagttca ggtaaatttc accaggagag taaatttgat 1560aagggtcaga
attccctaac ttgtaataaa agtaaagctt ctagacagac atttgtgatt 1620cacaaattag
aaaaagataa cttactccca aaccaaaagg ataaagtaac catttatgaa 1680aacctagacg
tcacaaatga atttcacaca gccaatcttt ccaccaaaga taatggaaat 1740ttatgtgatt
atgggaccca caatatattg gatttgaaaa agtatgtcac tgatattcaa 1800ccctcagagc
aaaatgaatc aaacattaat aagcttagaa agaaagtaaa ccggaagaca 1860gaaataattt
ctggaatgaa ccacatgtat gaagataatg ataaagatgt ggtgcatggc 1920ctaaaaaaag
gtaatttttt tttcaaaacc caagaggata aagaacctat ctctgaaaac 1980atagaagttt
ccaaagagct tcaaatccca gctctttcta ctagagataa tgaaaatcaa 2040tgtgactata
ggacccagaa tgtgttgggt ttgcaaaagc agatcaccaa tatgtacccc 2100gttcagcaaa
atgaatcaaa agttaataag aagcttaggc agaaagtaaa tcggaagaca 2160gaaataattt
ctgaagtgaa tcatttagat aatgacaaaa gtatagaata cacagttaaa 2220agtcactcac
tctttttaac gcaaaaagat aaggaaataa tccccggaaa cctagaagac 2280ccaagtgagt
ttgaaacacc tgctctttct accaaagata gtggaaacct gtatgattct 2340gagattcaaa
atgttttggg ggtgaaacat ggccatgata tgcaacctgc ttgtcaaaat 2400gattcaaaaa
taggtaagaa gcctagacta aatgtatgtc aaaagtcaga aataattcct 2460gaaaccaacc
aaatatatga gaatgataac aaaggtgtac atgacctaga aaaagataac 2520ttcttctctc
taaccccaaa ggataaagaa acaatttctg aaaatctaca agtcacaaat 2580gaatttcaaa
cagttgatct tctcatcaaa gataatggaa atttatgtga ttatgacacc 2640cagaatatat
tggagttgaa aaagtatgtt actgatagga aatctgctga gcaaaatgaa 2700tcaaaaataa
ataagctcag gaataaagtg aattggaaga cagaaataat ttctgaaatg 2760aaccagatat
atgaggataa tgataaagat gcacatgtcc aagaaagcta tacaaaagat 2820cttgatttta
aagtaaataa atctaaacaa aaacttgaat gccaagacat tatcaataaa 2880cactatatgg
aagtcaacag taatgaaaag gaaagttgtg atcaaatttt agattcctac 2940aaagtagtta
aaaaacgtaa gaaagaatca tcatgcaagg caaagaacat tttgacaaaa 3000gctaagaaca
aacttgcttc acagttaaca gaatcttcac agacatctat ctccttagaa 3060tctgatttaa
aacatattac tagtgaagca gattctgatc caggaaaccc agttgaacta 3120tgtaagactc
agaagcaaag cactaccact ttgaataaaa aagatctccc ttttgtggaa 3180gaaataaaag
aaggagagtg tcaggttaaa aaggtaaata aaatgacatc taagtcaaag 3240aaaaggaaga
cctccataga tccttctcca gagagccatg aagtaatgga aagaatactt 3300gacagcgttc
agggaaagtc tactgtatct gaacaagctg ataaggaaaa caatttggag 3360aatgagaaaa
tggtcaaaaa taagccagac ttttacacaa aggcatttag atctttgtct 3420gagatacatt
cacctaacat acaagattct tcctttgaca gtgttcgtga aggtttagta 3480cctttgagcg
tttcttctgg taaaaatgtg ataataaaag aaaattttgc cttggagtgc 3540tccccagcct
ttcaagtaag tgatgatgag catgagaaga tgaacaagat gaaatttaaa 3600gtcaaccgga
gaacccaaaa atcaggaata ggtgatagac cattacagga cttgtcaaat 3660accagttttg
tttcaaataa cactgctgaa tctgaaaata agtcagaaga tctatcttca 3720gaacggacaa
gcagaagaag aaggtgtact cctttctatt ttaaagagcc aagcctcaga 3780gacaagatga
gaagatga
3798201265PRTHomo sapiens 20Met Glu Cys Pro Val Met Glu Thr Gly Ser Leu
Phe Thr Ser Gly Ile1 5 10
15Lys Arg His Leu Lys Asp Lys Arg Ile Ser Lys Thr Thr Lys Leu Asn
20 25 30Val Ser Leu Ala Ser Lys Ile
Lys Thr Lys Ile Leu Asn Asn Ser Ser 35 40
45Ile Phe Lys Ile Ser Leu Lys His Asn Asn Arg Ala Leu Ala Gln
Ala 50 55 60Leu Ser Arg Glu Lys Glu
Asn Ser Arg Arg Ile Thr Thr Glu Lys Met65 70
75 80Leu Leu Gln Lys Glu Val Glu Lys Leu Asn Phe
Glu Asn Thr Phe Leu 85 90
95Arg Leu Lys Leu Asn Asn Leu Asn Lys Lys Leu Ile Asp Ile Glu Ala
100 105 110Leu Met Asn Asn Asn Leu
Ile Thr Ala Ile Glu Met Ser Ser Leu Ser 115 120
125Glu Phe His Gln Ser Ser Phe Leu Leu Ser Ala Ser Lys Lys
Lys Arg 130 135 140Ile Ser Lys Gln Cys
Lys Leu Met Arg Leu Pro Phe Ala Arg Val Pro145 150
155 160Leu Thr Ser Asn Asp Asp Glu Asp Glu Asp
Lys Glu Lys Met Gln Cys 165 170
175Asp Asn Asn Ile Lys Ser Lys Thr Leu Pro Asp Ile Pro Ser Ser Gly
180 185 190Arg Thr Thr Gln Pro
Leu Ser Thr Gln Asp Asn Ser Gly Val Leu Phe 195
200 205Leu Lys Glu Asn Asn Gln His Val Tyr Gly Leu Asp
Asp Ser Glu His 210 215 220Ile Ser Ser
Ile Val Asp Val Pro Pro Arg Glu Ser His Ser His Ser225
230 235 240Asp Gln Ser Ser Lys Thr Ser
Leu Met Ser Glu Met Arg Asn Ala Gln 245
250 255Ser Ile Gly Arg Arg Trp Glu Lys Pro Ser Pro Ser
Asn Val Thr Glu 260 265 270Arg
Lys Lys Arg Gly Ser Ser Trp Glu Ser Asn Asn Leu Ser Ala Asp 275
280 285Thr Pro Cys Ala Thr Val Leu Asp Lys
Gln His Ile Ser Ser Pro Glu 290 295
300Leu Asn Cys Asn Asn Glu Ile Asn Gly His Thr Asn Glu Thr Asn Thr305
310 315 320Glu Met Gln Arg
Asn Lys Gln Asp Leu Pro Gly Leu Ser Ser Glu Ser 325
330 335Ala Arg Glu Pro Asn Ala Glu Cys Met Asn
Gln Ile Glu Asp Asn Asp 340 345
350Asp Phe Gln Leu Gln Lys Thr Val Tyr Asp Ala Asp Met Asp Leu Thr
355 360 365Ala Ser Glu Val Ser Lys Ile
Val Thr Val Ser Thr Gly Ile Lys Lys 370 375
380Lys Ser Asn Lys Lys Thr Asn Glu His Gly Met Lys Thr Phe Arg
Lys385 390 395 400Val Lys
Asp Ser Ser Ser Glu Lys Lys Arg Glu Arg Ser Lys Arg Gln
405 410 415Phe Lys Asn Ser Ser Asp Val
Asp Ile Gly Glu Lys Ile Glu Asn Arg 420 425
430Thr Glu Arg Ser Asp Val Leu Asp Gly Lys Arg Gly Ala Glu
Asp Pro 435 440 445Gly Leu Phe Phe
Asn Asn Glu Gln Leu Ala Gln Met Asn Glu Gln Leu 450
455 460Ala Gln Val Asn Glu Leu Lys Lys Met Thr Leu Gln
Thr Gly Phe Glu465 470 475
480Gln Gly Asp Arg Glu Asn Val Leu Cys Asn Lys Lys Glu Lys Arg Val
485 490 495Thr Asn Glu Gln Glu
Glu Thr Tyr Ser Leu Ser Gln Ser Ser Gly Lys 500
505 510Phe His Gln Glu Ser Lys Phe Asp Lys Gly Gln Asn
Ser Leu Thr Cys 515 520 525Asn Lys
Ser Lys Ala Ser Arg Gln Thr Phe Val Ile His Lys Leu Glu 530
535 540Lys Asp Asn Leu Leu Pro Asn Gln Lys Asp Lys
Val Thr Ile Tyr Glu545 550 555
560Asn Leu Asp Val Thr Asn Glu Phe His Thr Ala Asn Leu Ser Thr Lys
565 570 575Asp Asn Gly Asn
Leu Cys Asp Tyr Gly Thr His Asn Ile Leu Asp Leu 580
585 590Lys Lys Tyr Val Thr Asp Ile Gln Pro Ser Glu
Gln Asn Glu Ser Asn 595 600 605Ile
Asn Lys Leu Arg Lys Lys Val Asn Arg Lys Thr Glu Ile Ile Ser 610
615 620Gly Met Asn His Met Tyr Glu Asp Asn Asp
Lys Asp Val Val His Gly625 630 635
640Leu Lys Lys Gly Asn Phe Phe Phe Lys Thr Gln Glu Asp Lys Glu
Pro 645 650 655Ile Ser Glu
Ser Ile Glu Val Ser Lys Glu Leu Gln Ile Pro Ala Leu 660
665 670Ser Thr Arg Asp Asn Glu Asn Gln Cys Asp
Tyr Arg Thr Gln Asn Val 675 680
685Leu Gly Leu Gln Lys Gln Ile Thr Asn Met Tyr Pro Val Gln Gln Asn 690
695 700Glu Ser Lys Val Asn Lys Lys Leu
Arg Gln Lys Val Asn Arg Lys Thr705 710
715 720Glu Ile Ile Ser Glu Val Asn His Leu Asp Asn Asp
Lys Ser Ile Glu 725 730
735Tyr Thr Val Lys Ser His Ser Leu Phe Leu Thr Gln Lys Asp Lys Glu
740 745 750Ile Ile Pro Gly Asn Leu
Glu Asp Pro Ser Glu Phe Glu Thr Pro Ala 755 760
765Leu Ser Thr Lys Asp Ser Gly Asn Leu Tyr Asp Ser Glu Ile
Gln Asn 770 775 780Val Leu Gly Val Lys
His Gly His Asp Met Gln Pro Ala Cys Gln Asn785 790
795 800Asp Ser Lys Ile Gly Lys Lys Pro Arg Leu
Asn Val Cys Gln Lys Ser 805 810
815Glu Ile Ile Pro Glu Thr Asn Gln Ile Tyr Glu Asn Asp Asn Lys Gly
820 825 830Val His Asp Leu Glu
Lys Asp Asn Phe Phe Ser Leu Thr Pro Lys Asp 835
840 845Lys Glu Thr Ile Ser Glu Asn Leu Gln Val Thr Asn
Glu Phe Gln Thr 850 855 860Val Asp Leu
Leu Ile Lys Asp Asn Gly Asn Leu Cys Asp Tyr Asp Thr865
870 875 880Gln Asn Ile Leu Glu Leu Lys
Lys Tyr Val Thr Asp Arg Lys Ser Ala 885
890 895Glu Gln Asn Glu Ser Lys Ile Asn Lys Leu Arg Asn
Lys Val Asn Trp 900 905 910Lys
Thr Glu Ile Ile Ser Glu Met Asn Gln Ile Tyr Glu Asp Asn Asp 915
920 925Lys Asp Ala His Val Gln Glu Ser Tyr
Thr Lys Asp Leu Asp Phe Lys 930 935
940Val Asn Lys Ser Lys Gln Lys Leu Glu Cys Gln Asp Ile Ile Asn Lys945
950 955 960His Tyr Met Glu
Val Asn Ser Asn Glu Lys Glu Ser Cys Asp Gln Ile 965
970 975Leu Asp Ser Tyr Lys Val Val Lys Lys Arg
Lys Lys Glu Ser Ser Cys 980 985
990Lys Ala Lys Asn Ile Leu Thr Lys Ala Lys Asn Lys Leu Ala Ser Gln
995 1000 1005Leu Thr Glu Ser Ser Gln
Thr Ser Ile Ser Leu Glu Ser Asp Leu 1010 1015
1020Lys His Ile Thr Ser Glu Ala Asp Ser Asp Pro Gly Asn Pro
Val 1025 1030 1035Glu Leu Cys Lys Thr
Gln Lys Gln Ser Thr Thr Thr Leu Asn Lys 1040 1045
1050Lys Asp Leu Pro Phe Val Glu Glu Ile Lys Glu Gly Glu
Cys Gln 1055 1060 1065Val Lys Lys Val
Asn Lys Met Thr Ser Lys Ser Lys Lys Arg Lys 1070
1075 1080Thr Ser Ile Asp Pro Ser Pro Glu Ser His Glu
Val Met Glu Arg 1085 1090 1095Ile Leu
Asp Ser Val Gln Gly Lys Ser Thr Val Ser Glu Gln Ala 1100
1105 1110Asp Lys Glu Asn Asn Leu Glu Asn Glu Lys
Met Val Lys Asn Lys 1115 1120 1125Pro
Asp Phe Tyr Thr Lys Ala Phe Arg Ser Leu Ser Glu Ile His 1130
1135 1140Ser Pro Asn Ile Gln Asp Ser Ser Phe
Asp Ser Val Arg Glu Gly 1145 1150
1155Leu Val Pro Leu Ser Val Ser Ser Gly Lys Asn Val Ile Ile Lys
1160 1165 1170Glu Asn Phe Ala Leu Glu
Cys Ser Pro Ala Phe Gln Val Ser Asp 1175 1180
1185Asp Glu His Glu Lys Met Asn Lys Met Lys Phe Lys Val Asn
Arg 1190 1195 1200Arg Thr Gln Lys Ser
Gly Ile Gly Asp Arg Pro Leu Gln Asp Leu 1205 1210
1215Ser Asn Thr Ser Phe Val Ser Asn Asn Thr Ala Glu Ser
Glu Asn 1220 1225 1230Lys Ser Glu Asp
Leu Ser Ser Glu Arg Thr Ser Arg Arg Arg Arg 1235
1240 1245Cys Thr Pro Phe Tyr Phe Lys Glu Pro Ser Leu
Arg Asp Lys Met 1250 1255 1260Arg Arg
12652145PRTyeast 21Met Glu Ser Leu Lys Lys Lys Phe Leu Lys Gln Asn Arg
Glu Ile Ile1 5 10 15Lys
Ile Asn Thr Gln Leu Ser Ile Lys Ile Arg Glu Ser Glu Asn Glu 20
25 30Ile Gln Asp Leu Ile Gln Glu Asn
Phe Thr Leu Lys Ser 35 40
452245PRTyeast 22Val Glu Asp Leu Lys Lys Lys Gln Ile Arg Gln Tyr Lys Glu
Ile Ile1 5 10 15Arg Ile
Ser Lys Ala Gln Ser Ile Arg Ile Lys Glu Leu Gln Leu Glu 20
25 30Asn Glu Arg Leu Leu Ser Glu Asn Ile
Asp Leu Arg Thr 35 40
452345PRTyeast 23Val Glu Asn Ile Arg Gln Ser Tyr Ser Arg Gln Asn Ser Leu
Leu Ala1 5 10 15Lys Asp
Asn Ser Ile Leu Lys Ile Lys Val Asn Ser Leu Glu Lys Lys 20
25 30Ile Ser Gln Leu Val Gln Glu Asn Val
Thr Leu Arg Ser 35 40
452445PRTNeurospora crassa 24Leu Glu Leu Leu Arg Arg Lys Phe Leu Arg Gln
Asn Arg Asp Ile Ala1 5 10
15Arg Val Asn Ser Thr Gln Ser Leu Arg Ile Arg Gly Leu Glu Asn Glu
20 25 30Cys Ala Arg Leu Leu Ser Glu
Asn Leu Glu Leu Arg Gly 35 40
452545PRTDactylicapnos macrocapnos 25Gly Ser Lys Val Glu Gln Gln Tyr Lys
Leu Leu Asn Ala Glu Leu Met1 5 10
15Asp Gln Val Gln Lys Gln Arg Leu Glu Ile Gly Glu Tyr Arg Lys
Arg 20 25 30Val Ile Ser Leu
Glu Arg Glu Ile Met Asp Ile Arg Glu 35 40
452627PRTyeast 26Gly Arg Glu Lys Leu Arg Arg Ser Val Lys Val Ile
Asn Tyr Ala Ile1 5 10
15Pro Ser Leu Arg Thr Lys Leu Arg Arg Asp Phe 20
252727PRTyeast 27Pro Asp Gly Arg Ser Arg Arg Glu Arg Lys Lys Val Asn
Tyr Ala Leu1 5 10 15Pro
Gly Leu Arg Thr Lys Leu Arg Arg Asn Phe 20
252828PRTyeast 28Ser Phe Thr Arg Thr Arg Arg Thr Arg Gly Lys Ala Val Asp
Tyr Thr1 5 10 15Leu Pro
Ser Leu Arg Ala Lys Met Arg Arg Pro Ser 20
252928PRTNeurospora crassa 29Glu Thr Ser Arg Pro Ser Arg Arg Ala Arg Ala
Ala Ile Ser Tyr Thr1 5 10
15Glu Pro Asn Leu Arg Asp Lys Met Arg Arg Pro Thr 20
253027PRTDactylicapnos macrocapnos 30Asn Ser Ala Arg Pro Ser Arg
Ser Cys Arg Pro Thr Ser Leu Val Glu1 5 10
15Pro Ser Leu Lys Asn Lys Leu Arg Asn Gly Ser
20 253128PRTCaenorhabditis elegans 31Thr Val Arg Arg Gln
Arg Ser Ala Lys Met Asn Ile Lys Ser Leu Lys1 5
10 15Glu Pro Ser Gly Lys Asp Lys Leu Arg Arg Pro
Gly 20 253229PRTArabidopsis thaliana 32Thr
Val Gly Arg Pro Ser Arg Gln Ala Ala Glu Lys Ile Lys Ser Tyr1
5 10 15Lys Glu Pro Ser Leu Lys Glu
Lys Met Arg Gly Gly Phe 20
253329PRTArabidopsis thaliana 33Ser Val Gly Arg Pro Ser Arg His Ala Ala
Glu Lys Val Gln Ser Tyr1 5 10
15Arg Glu Val Ser Leu Arg Val Lys Met Arg Arg Lys Cys 20
253428PRTmouse 34Ala Val Ala Leu Thr Lys Arg Arg Cys Ser
Thr Ile Lys Ser Tyr Lys1 5 10
15Glu Pro Thr Leu Ala Ser Lys Leu Arg Arg Gly Asp 20
253525PRTmouse 35His Pro Met Arg Arg Lys Arg Gln Cys Val Pro
Leu Asn Leu Thr Glu1 5 10
15Pro Ser Leu Arg Ser Lys Met Arg Arg 20
253628PRTHomo sapiens 36Ala Val Ala Leu Pro Lys Arg Arg Cys Thr Ala Ser
Val Asn Tyr Lys1 5 10
15Glu Pro Thr Leu Ala Ser Lys Leu Arg Arg Gly Asp 20
253726PRTHomo sapiens 37Ser Glu Arg Thr Ser Arg Arg Arg Arg Cys
Thr Pro Phe Tyr Phe Lys1 5 10
15Glu Pro Ser Leu Arg Asp Lys Met Arg Arg 20
253821DNAArtificial Sequence?TriplEx 38ctcgggaagc gcgccattgt g
213922DNAHomo sapiens 39cctggctgaa
tcagctttgg tg
224023DNAArtificialhSgo1 40aagucuacug auaaugucuu att
234123DNAArtificial SequencehSgo2 41aagcacuacc
acuuugaaua att
234221DNAArtificial SequencehSgo1 42gugagccucu gugaaucaat t
21 4321DNAArtificial SequencehSgo2
43gcucucauga acaauaacut t
21 4421DNAArtificial SequencesiRNA,Target1 44gagugaucac gauuucuaat t
21 4521DNAArtificial
SequencesiRNA,Target2 45aacgggcauu ugaauaugaa a
21
User Contributions:
comments("1"); ?> comment_form("1"); ?>Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
User Contributions:
Comment about this patent or add new information about this topic: