Patent application title: Vectors and cells for preparing immunoprotective compositions derived from transgenic plants
Inventors:
Guy A. Cardineau (Tempe, AZ, US)
Hugh Stanley Mason (Phoenix, AZ, US)
Joyce M. Vaneck (Ithaca, NY, US)
Dwayne D. Kirk (Mesa, AZ, US)
Amanda Maree Walmsley (Mesa, AZ, US)
IPC8 Class: AA61K39145FI
USPC Class:
4242061
Class name: Virus or component thereof reassortant or deletion mutant virus influenza virus
Publication date: 2009-01-15
Patent application number: 20090017065
Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
Patent application title: Vectors and cells for preparing immunoprotective compositions derived from transgenic plants
Inventors:
Guy A. Cardineau
Hugh Stanley Mason
Joyce M. VanEck
Dwayne D. Kirk
Amanda Maree Walmsley
Agents:
Edwards Angell Palmer & Dodge LLP
Assignees:
Origin: BOSTON, MA US
IPC8 Class: AA61K39145FI
USPC Class:
4242061
Abstract:
The inventions is drawn towards vectors and methods useful for preparing
genetically transformed plant cells that express immunogens from
pathogenic organisms which are used to produce immunoprotective particles
useful in vaccine preparations. The invention includes plant optimized
genes that encode the HN protein of Newcastle Disease Virus. The
invention also relates to methods of producing an antigen in a transgenic
plant.Claims:
1. A transgenic plant comprising a vector, wherein said vector comprises
SEQ ID NO: 1.
2. A transgenic plant comprising a vector, wherein said vector is selected from the group consisting of pCHN, pGHN, pGHN151, pGHN153, pMHN, pUHN.
3. The transgenic plant of claim 1, wherein a plant-functional promoter is operably linked to SEQ ID NO:1.
4. A vaccine comprising a recombinant viral antigenic protein and a pharmaceutically acceptable carrier, wherein the viral antigenic protein is the HN antigen of Newcastle Disease virus produced by a vector comprising SEQ ID NO: 1 or a vector selected from the group consisting of pCHN, pGHN, pGHN151, pGHN153, pMHN, pUHN, and wherein said vaccine is capable of eliciting an immune response upon administration to an animal.
5. The vaccine of claim 4, wherein said HN protein comprises SEQ ID NO:2.
6. The vaccine of claim 4, wherein said HN protein is produced in a plant cell.
7. A method for protecting an animal against NewCastle Disease Virus comprising administering an effective amount of the vaccine of claim 4 to an animal.
8. A vaccine comprising a recombinant viral antigenic protein and a pharmaceutically acceptable carrier, wherein the viral antigenic protein is the HA antigen of Avian Influenza Virus produced by a vector for expressing an immunoprotective antigen in a plant cell comprising a DNA sequence encoding the HA antigen of Avian Influenza Virus, wherein the vector is pCHA, and wherein said vaccine is capable of eliciting an immune response upon administration to an animal.
9. The vaccine of claim 8, wherein said HA antigen is produced in a plant cell.
10. A method for protecting an animal against Avian Influenza Virus comprising administering an effective amount of the vaccine of claim 8 to an animal.
11. The method of claim 7 or 10, wherein the vaccine is administered orally, intranasaly, intraperitonealy, intramuscularly, intravenously or subcutaneously.
12. The method of claim 7 or 10, wherein the effective amount is at a range of 1 μg to 50 μg per kilogram of bodyweight.
13. A method of producing an antigen in a transgenic plant comprising the steps of:a) producing a transgenic plant comprising a vector encoding said antigen;b) incubating said plant under conditions wherein said plant expresses said antigen; andwherein said plant is incubated prior to the onset of ripening.
14. The method of claim 13, wherein said plant comprises a fruit that ripens.
15. The method of claim 13, wherein said plant is a tomato plant.
16. The method of claim 14, wherein the fruit of said plant is harvested prior to the onset of ripening.
17. The method of claim 16, wherein said antigen is isolated from said harvested fruit.
18. The method of claim 13, wherein said antigen is selected from the group consisting of HN antigen of Newcastle Disease Virus, HA antigen of Avian Influenza Virus, LTB, NVCP, zona pellucida glycoprotein and HBsAg.
Description:
[0001]This application is a divisional of Ser. No. 11/450,102, filed Jun.
8, 2006, which is a divisional of U.S. Pat. No. 7,132,291, filed May 4,
2004, which claims the benefit of U.S. Provisional Application No.
60/467,998, filed on May 5, 2003. The entire teachings of the above
applications are incorporated herein by reference.
FIELD OF INVENTION
[0002]The present invention generally relates to the field of plant molecular biology as it applies to the recombinant production of plant-made vaccines.
BACKGROUND OF THE INVENTION
[0003]Recombinant DNA technology has provided substantial improvements in the safety, quality, efficacy and cost of pharmaceutical and veterinary medicaments including vaccines. Plant produced mucosal vaccines were invented by Curtiss & Cardineau. See U.S. Pat. Nos. 5,654,184; 5,679,880 and 5,686,079 herein incorporated by reference. Others have described transgenic plants expressing immunoprotective antigens and methods for production including Arntzen, Mason and Lam. See U.S. Pat. Nos. 5,484,717; 5,914,123; 6,034,298; 6,136,320; 6,194,560; and 6,395,964 herein incorporated by reference.
[0004]Vaccines produced in plant systems offer a number of advantages over conventional production systems. Conventionally produced vaccines strains (live and vectored) may revert towards virulence or carry biological contaminants from the production process. Subunit vaccines may be difficult to produce and purify due to protein instability issues and will not be glycosylated when produced in prokaryotes.
[0005]Plant cell production avoids the need for animal-sourced components in growth media essentially eliminating the risk of transmitting pathogenic contaminants from the production process. Plant cells are capable of post translational glycosylation, and plant cell growth media is generally less expensive and easier to handle and prepare compared to conventional growth media presently used in the manufacture of vaccines.
[0006]Systemic immunity to a particular pathogen results from activation of the immune system in response to antigen presented by a particular pathogenic organism or via a vaccine designed to protect against a particular pathogenic agent. Exposure to a pathogen is often through mucosal surfaces that are constantly exposed and challenged by pathogenic organisms.
[0007]Mucosal and oral immunity results from the production of sIgA (secretory IgA) antibodies in secretions that bathe all mucosal surfaces of the respiratory tract, gastrointestinal tract and the genitourinary tract and in secretions from all secretory glands. McGhee, J. R. et al., Annals N Y. Acad. Sci. 409, (1983). These sIgA antibodies act to prevent colonization of pathogens on a mucosal surface (Williams, R. C. et al., Science 177, 697 (1972); McNabb, P. C. et al., Ann. Rev. Microbiol. 35, 477 (1981) and thus act as a first line of defense to prevent colonization or invasion through a mucosal surface. The production of sIgA can be stimulated either by local immunization of the secretory gland or tissue or by presentation of an antigen to either the GALT (gut-associated lymphoid tissue or Peyer's patches) or the BALT (bronchial-associated lymphoid tissue). Cebra, J. J. et al., Cold Spring Harbor Symp. Quant. Biol. 41, 210 (1976); Bienenstock, J. M., Adv. Exp. Med. Biol. 107, 53 (1978); Weisz-Carrington, P. Et al., J. Immunol. 123, 1705 (1979); McCaughan, G. et al., Internal Rev. Physiol 28, 131 (1983). Membranous microfold cells, otherwise known as M Cells, cover the surface of the GALT and BALT and may be associated with other secretory mucosal surfaces. M cells act to sample antigens from the luminal space adjacent to the mucosal surface and transfer such antigens to antigen-presenting cells (dendritic cells and macrophages), which in turn present the antigen to a T lymphocyte (in the case of T-dependent antigens), which process the antigen for presentation to a committed B cell. B cells are then stimulated to proliferate, migrate and ultimately be transformed into an antibody-secreting plasma cell producing IgA against the presented antigen. When the antigen is taken up by M cells overlying the GALT and BALT, a generalized mucosal immunity results with sIgA against the antigen being produced by all secretory tissues in the body. Cebra et al., supra; Bienenstock et al., supra; Weinz-Carrington et al., supra; McCaughan et al., supra. Oral immunization is therefore a most important route to stimulate a generalized mucosal immune response and, in addition, leads to local stimulation of a secretory immune response in the oral cavity and in the gastrointestinal tract.
[0008]Mucosal immunity can also be advantageously transferred to offspring. Immunity in neonates may be passively acquired through colostrum and/or milk. This has been referred to as lactogenic immunity and is an efficient way to protect animals during early life. sIgA is the major immunoglobulin in milk and is most efficiently induced by mucosal immunization.
[0009]The M cells overlying the Peyer's patches of the gut-associated lymphoid tissue are capable of taking up a diversity of antigenic material and particles (Sneller, M. C. and Strober, W., J. Inf. Dis. 154, 737 (1986). Because of their abilities to take up latex and polystyrene spheres, charcoal, microcapsules and other soluble and particulate matter, it is possible to deliver a diversity of materials to the GALT independent of any specific adhesive-type property of the material to be delivered.
[0010]Vectors and cells useful for producing transgenic plant-derived immunoprotective antigens, and improved methods of antigen production would greatly facilitate the development, manufacture and efficacy plant-produced vaccines.
SUMMARY OF THE INVENTION
[0011]The invention is based on plant optimized sequences encoding an immunoprotective antigen of interest. In particular, the invention is based on a plant optimized DNA sequence encoding the HN antigen of Newcastle Disease Virus or a DNA sequence encoding the HA antigen of Avian Influenza Virus. The invention also includes a recombinant expression vector for effecting expression of an immunoprotective antigen gene in a plant cell, as well as plant cells and transgenic plants comprising the expression vector, as well as vaccines comprising a protein product of the expression vector. The invention also relates to methods of protecting against the effects of a pathogen utilizing the vaccines of the invention. The invention further relates to methods of producing an antigen in a transgenic plant.
[0012]The invention provides for an isolated plant optimized nucleotide sequence encoding the HN antigen of Newcastle Disease Virus comprising the sequence of SEQ ID NO: 1, as well as a recombinant expression vector comprising SEQ ID NO: 1.
[0013]In one embodiment, the vector is selected from the group consisting of pCHN, pGHN, pGHN151, pGHN153, pMHN, pUHN.
[0014]In another embodiment, the vector comprises a plant-functional promoter is operably linked to SEQ ID NO: 1.
[0015]The invention also provides for a recombinant expression vector for expressing an immunoprotective antigen in a plant cell comprising a DNA sequence encoding the HA antigen of Avian Influenza Virus, wherein the vector is pCHA
[0016]The invention further provides for a transgenic plant cell for expression of an immunogenic antigen comprising a vector of the invention. The plant cell includes a tomato plant cell or a tobacco plant cell, as well as a cell from any of the plant species described hereinbelow.
[0017]The invention further provides for a transgenic plant comprising a vector of the invention.
[0018]The invention also provides for a vaccine comprising a recombinant viral antigenic protein and a pharmaceutically acceptable carrier, wherein the viral antigenic protein is the HN antigen of Newcastle Disease virus produced by a vector of the invention, and wherein the vaccine is capable of eliciting an immune response upon administration to an animal.
[0019]In one embodiment, the HN protein of the vaccine comprises SEQ ID NO:2. The HN protein of the vaccine can be produced in a plant cell.
[0020]The invention also provides for a vaccine comprising a recombinant viral antigenic protein and a pharmaceutically acceptable carrier, wherein the viral antigenic protein is the HA antigen of Avian Influenza Virus produced by a vector of the invention, and wherein the vaccine is capable of eliciting an immune response upon administration to an animal. In one embodiment, the HA antigen of the vaccine is produced in a plant cell.
[0021]The invention also provides for a method for protecting an animal against NewCastle Disease Virus or Avian Influenza Virus comprising administering an effective amount of the appropriate vaccine of the invention to an animal. According to one embodiment of the method, wherein the vaccine is administered orally, intranasaly, intraperitonealy, intramuscularly, intravenously or subcutaneously. In one embodiment of the method, the effective amount of the vaccine is at a range of 1 μg to 50 μg per kilogram of body weight.
[0022]The invention also provides for a method of producing an antigen in a transgenic plant comprising the steps of: a) producing a transgenic plant comprising a vector encoding the antigen; b) incubating the plant under conditions wherein the plant expresses the antigen; and wherein the plant is incubated prior to the onset of ripening.
[0023]In one embodiment, the plant comprises a fruit that ripens.
[0024]In another embodiment, the plant is a tomato plant.
[0025]In another embodiment, the fruit of the plant is harvested prior to the onset of ripening. According to one embodiment of this method, the antigen is isolated from the harvested fruit.
[0026]In another embodiment, the antigen is selected from the group consisting of HN antigen of Newcastle Disease Virus, HA antigen of Avian Influenza Virus, LTB, NVCP, zona pellucida glycoprotein and HBsAg.
BRIEF DESCRIPTION OF THE FIGURES
[0027]FIGS. 1a and 1b. The plant optimized coding sequence (SEQ ID NO: 1) and protein sequence (SEQ ID NO: 2) of the HN gene of NDV strain "Lasota"
[0028]FIG. 2. Map of pBBV-PHAS-iaaH that contains the plant selectable marker PAT (phosphinothricin acetyl transferase), includes the constitutive CsVMV (cassaya vein mosaic virus) promoter and is terminated by the MAS 3' (mannopine synthase) element. LB and RB (left and right T-DNA border) elements from Agrobacterium delineate the boundaries of the DNA that is integrated into the plant genome.
[0029]FIG. 3. Map of pCP!H which is a "template vector" used as a starting plasmid for a variety of plant expression vectors for expressing immunoprotective antigens.
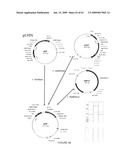
[0030]FIG. 4. Map of pCHN expression vector for NDV HN protein. This vector comprising the HN expression cassette includes the constitutive CsVMV promoter and is terminated by the soybean vspB 3' element.
[0031]FIG. 5. Map of pgHN expression vector for NDV HN protein. This vector comprising the HN expression cassette includes the tuber-specific GBSS promoter with TEV 5' UTRand is terminated by the soybean vspB 3' element.
[0032]FIG. 6. Map of pgHN151 expression vector for NDV HN protein. The HN expression vector or cassette includes the tuber-specific GBSS promoter with its native 5' UTR and intron, and is terminated by the soybean vspB 3' element. The vector is derived from pBBV-PHAS-iaaH, which contains the plant selectable marker PAT, includes the CsVMV promoter and is terminated by the MAS 3' element. LB and RB, left and right T-DNA border elements delineate the boundaries of the DNA that is integrated into the plant genome.
[0033]FIG. 7. Map of pgHN153 expression vector for NDV HN protein. The HN expression vector includes the tuber-specific GBSS promoter with its native 5' UTR and intron, and is terminated by the bean phaseolin 3' element. The vector is derived from pBBV-PHAS-iaaH, which contains the plant selectable marker PAT, includes the CsVMV promoter and is terminated by the MAS 3' element. LB and RB, left and right T-DNA border elements delineate the boundaries of the DNA that is integrated into the plant genome.
[0034]FIG. 8. Map of pMHN expression vector for NDV HN protein. The HN expression vector includes the constitutive 40CSAMAS promoter (P2 direction) and is terminated by the soybean vspB 3' element. The vector is derived from pBBV-PHAS-iaaH, which contains the plant selectable marker PAT, includes the CsVMV promoter and is terminated by the MAS 3' element. LB and RB, left and right T-DNA border elements delineate the boundaries of the DNA that is integrated into the plant genome.
[0035]FIG. 9. Map of pCHA expression vector for the HA gene of the AIV A/turkey/Wisconsin/68 (H5N9).
[0036]FIG. 10. The DNA (SEQ ID NO: 3) and protein (SEQ ID NO: 4) sequences of the HA gene of AIV A/turkey/Wisconsin/68 (H5N9).
[0037]FIG. 11. Map of pGLTB intermediate vector.
[0038]FIG. 12. Map of pCLT105 intermediate vector.
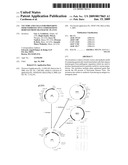
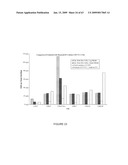
[0039]FIG. 13. HA expression in transgenic NT1 cell lines using pGPTV-HAO or pCHA. Callus cells growing on solid media were extracted and assayed for HA by ELISA and for total protein by the Bradford method. Data are presented as ng HA per μg total protein. A01/12, a high-expressing line selected from several pGPTV-HAO-transformed lines. CHA, lines transformed with pCHA. CVMV/13, a vector-only transformed line. Separate samples were extracted and assayed Jul. 12, 2001 or Jul. 27, 2001.
[0040]FIG. 14. Repeated assays of pCHA-transformed NT1 cell lines.
[0041]FIG. 15. Western blot for AIV HA expression in pCHA-transformed NT1 cell lines. NT1 cell lines were grown in liquid suspension culture, and extracts were resolved by SDS-PAGE, electro-transferred to PVDF membrane, and probed with chicken anti-AIV-H5 from USDA/SEPRL, which is also used as the detector antibody in the HA quantitation ELISA. Lanes 1 and 10, molecular size standards; lane 2, HN Reference Antigen at 1:800, 31.25 ng/well; lane 3, CHA-13 (1:2); lane 4, CHA-42 (1:2); lane 5, CHA-43 (1:2); lane 6, CHA-44 (1:2); lane 7, CHA-61 (1:2); lane 8, GPTV-HAO grown with Kanamycin (1:2); lane 9, GPTV-HAO grown without Kanamycin (1:2).
[0042]FIG. 16. HA expression in microtubers of pCHA-transformed potato plantlets. Microtubers were generated in vitro from stem nodes of tissue culture plantlets. Samples were extracted and assayed for HA by ELISA. Data are presented as ng HA per g fresh microtuber weight. Line numbers indicate independent transgenic lines. Desiree, a non-transformed line. Standard error bars represent standard deviation of multiple determinations.
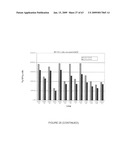
[0043]FIG. 17. HA expression in leaves of pCHA-transformed potato plants. Leaf samples from greenhouse-grown plants were extracted and assayed for HA by ELISA and for total protein by the Bradford method. Data are presented as ng HA per μg total protein. Line numbers indicate independent transgenic lines. Standard deviations of multiple determinations are shown.
[0044]FIG. 18. HA expression in tubers of soil-grown pCHA-transformed potato plants. Greenhouse-grown plants were harvested and tubers sampled for extraction and ELISA for HA expression. Four replicate samples were analyzed, and the standard deviations are shown. Data are presented as ng HA per g fresh tuber weight.
[0045]FIG. 19. Expression of NDV-HN in NT1 cells transformed with pCHN. Extracts f cells growing on solid media were assayed by ELISA for NDV-HN, using inactivated NDV as a reference standard. Total soluble protein (TSP) was assayed by the Bradford method (BioRad) using bovine serum albumin as a standard.
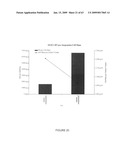
[0046]FIG. 20. Expression of HN per cell mass in pCHN-transformed NT1 lines.
[0047]FIG. 21. Stability of expression of HN in pCHN-transformed NT1 cell lines.
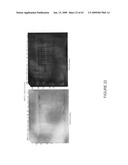
[0048]FIG. 22. Western blot of pCHN-transformed NT1 cells using HN-specific antibodies. NT1 cells cultured on solid media (5P, 7P) or in liquid suspension (5-6, 5-13, 7-6, 7-13) were extracted and subjected to SDS-PAGE, followed by Western blot probed with either monoclonal (left) or polyclonal (right) antibodies. MW, molecular weight markers, indicated by numbers at left in kDa. Lanes 11.9+, 5.85+, 2.99+, 1.45+ indicate amount (ng) of reference standard inactivated NDV run in those lanes. C, nontransgenic NT1 cell extract.
[0049]FIG. 23. HN antigen is maintained in freeze-dried pCHN-transformed NT1 cells and on storage of extracts at 4° C. NT1 cells were freeze-dried, extracted, and subjected to ELISA. The results for freeze-dried cells were analyzed by either a log or linear regression model, as indicated in the inset. The values are corrected to indicate HN content per mass of fresh weight cells using estimates of water loss on drying. Fresh cells were extracted and analyzed immediately (2/19/2) or stored at 4° C. for 1 week prior to assay.
[0050]FIG. 24. Sucrose gradient analysis of HN antigen shows particulate character. Extracts of pCHN-transformed NT1 cell lines CHN-7 or CHN-18, or reference standard inactivated NDV were sedimented in 10-50% sucrose/PBS gradients at 350,000 g for 5 h. Fractions were collected and assayed by ELISA for HN. Fraction 1 is the top of gradient.
[0051]FIG. 25. Expression of HN in pMHN- and pCHN-transformed NT1 cell lines.
[0052]FIG. 26. HN expression in pCHN-transformed potato.
[0053]FIG. 27. Particle behavior of HN antigen extracted from pCHN-transformed potato tubers.
[0054]FIG. 28. HN expression in microtubers of pGHN-transformed potato plants.
[0055]FIG. 29. Expression of HN in tubers of pGHN- and pGHN151-transformed potato plants.
[0056]FIG. 30. T-DNA region from the construct pCHN.
[0057]FIG. 31. Effect of ripening on wild type TA234 tomato fruit pH.
[0058]FIG. 32. Effect of ripening on wild type TA234 tomato fruit total soluble protein.
[0059]FIG. 33. Southern analysis of T0 CHN tomato lines. The positive control is an EcoRI digest of the plasmid pCHN loaded to indicate the intensity of 2 copies of the HN gene per genome while the negative control is DNA from wild type TA234 tomato.
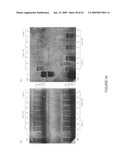
[0060]FIG. 34. Total RNA from wild type and transgenic tomato fruit. (a) Methylene blue stain of membrane. (b) Northern analysis. NC represents the negative control, total RNA from wild type fruit; L, MBI Fermentas (Hanover, Md.) high range RNA ladder; 1-6, corresponding ripening stage of fruit.
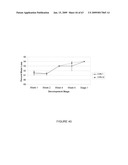
[0061]FIG. 35. ELISA analysis of HN concentration in ripening CHN tomato fruit. (a) Tomato line CHN-1. (b) Tomato line CHN-10. (c) Tomato line CHN-12. (d) Tomato line CHN-32. Except in line CHN-12 where only stage 1 fruit had three reps, bars represent the mean of 3 samples from 3 different fruit, error bars indicate the standard error of the mean.
[0062]FIG. 36. Western analysis of crude protein extracts from wild type and transgenic tomato fruit and leaves and NT1 cell extracts. NF, represents tomato fruit negative control--wild type fruit; NL tomato leaf negative control--wild type leaf; NNT NT1 cell negative control--non-transformed cell lines; 119, transgenic NT1 cell line 119; L10, leaf from transgenic tomato line 10; L32, leaf from tomato line 32; HN, animal derived Lasota NDV virus; M, Bio-Rad's precision plus protein all blue standard; 1-1, fruit from line CHN-1, stage 1 of ripening; 1-3, fruit from line CHN-1, stage 3 of ripening; 1-6, fruit from line CHN-1, stage 6 of ripening; 32-1, fruit from line CHN-32, stage 1 of ripening; 32-3, fruit from line CHN-32, stage 3 of ripening; 32-6, fruit from line CHN-32, stage 6 of ripening; 10-1, fruit from CHN-10, stage 1 of ripening. Protein size is give in kDa.
[0063]FIG. 37. Haemagglutination activity in the fruit and leaves of CHN tomatoes.
[0064]FIG. 38. Change in maturing fruit diameter. "Week" indicates the amount of time post pollination. Points indicate the mean of three measurements while the bars indicate the standard errors of the means.
[0065]FIG. 39. Change in fruit mass of maturing tomato fruit. "Week" indicates the amount of time post pollination. Each point represents the mean of the three measurements while the bars indicate the standard errors of the means.
[0066]FIG. 40. Water loss from maturing tomato fruit upon lyophilization. "Week" indicates the amount of time post pollination. Points represent the mean of three measurements while the bars indicate the standard errors of the means.
[0067]FIG. 41. Concentration of HN per gram of fresh tomato fruit. "Week" indicates the amount of time post pollination. Bars represent the average of three samples. Bars labeled with the same letter are not significantly different (α=0.05). Error bars indicate the standard error of the means.
[0068]FIG. 42. Amount of HN in maturing tomato fruit. "Week" indicates the amount of time post pollination. Bars represent the average of three replicate HN contents multiplied by the masses. Bars labeled with the same letter are not significantly different (α=0.05). Error bars indicate the standard error of the mean.
[0069]FIG. 43. The regulated biological agent (PCHN) insert in CHN-18 master seed.
[0070]FIG. 44. DNA sequence of the whole gene insert in CHN-18 master seed (SEQ ID NO: 12).
[0071]FIG. 45. pCHA vector sequence (SEQ ID NO: 24).
[0072]FIG. 46. pMHN vector sequence (SEQ ID NO: 25).
[0073]FIG. 47. pCHN vector sequence (SEQ ID NO: 26).
[0074]FIG. 48. Construction of pUHN.
SUMMARY OF THE SEQUENCES
[0075]SEQ ID NOS: 1 and 2, shown in FIG. 1, are the plant optimized coding sequence and protein sequence of the HN gene of NDV strain "Lasota".
[0076]SEQ ID NOS: 3 and 4, shown in FIG. 10, are the DNA and protein sequences of the HA gene of AIV A/turkey/Wisconsin/68 (H5N9).
[0077]SEQ ID NO: 5 is a PCR primer used to end-tailor the CsVMV promoter on pCP!H.
[0078]SEQ ID NO: 6 is a PCR primer used to end-tailor the CsVMV promoter on pCP!H.
[0079]SEQ ID NO: 7 is a mutagenic primer used to create a Nco I site.
[0080]SEQ ID NO: 8 is a forward primer complimentary to the 5' region.
[0081]SEQ ID NO: 9 is a mutagenic primer used to create a XhoI I site.
[0082]SEQ ID NO: 10 is a PCR labeled probe made by using the primer HNa.
[0083]SEQ ID NO: 11 is a PCR labeled probe made by using the primer HNb.
[0084]SEQ ID NO: 12 is the DNA sequence of the whole gene insert in CHN-18 master seed.
[0085]SEQ ID NO: 13 is the DNA sequence encoding Hepatitis B virus Strain Gly D surface antigen, complete cds. (GenBank accession AF134148).
[0086]SEQ ID NO: 14 is the protein sequence of Hepatitis B virus Strain Gly D surface antigen. (GenBank accession AAD31865).
[0087]SEQ ID NO: 15 is the DNA sequence encoding Homo sapiens zona pellucida glycoprotein 3 (sperm receptor) (ZP3), mRNA. (GenBank accession NM--007155).
[0088]SEQ ID NO: 16 is the protein sequence of Homo sapiens zona pellucida glycoprotein 3 preproprotein (sperm receptor) (ZP3). (GenBank accession NP--009086).
[0089]SEQ ID NO: 17 is the DNA sequence encoding Avian influenza virus hemagglutinin (HA) mRNA, complete cds. (GenBank accession U67783).
[0090]SEQ ID NO: 18 is the protein sequence of Avian influenza virus hemagglutinin (HA). (GenBank accession AAC58999).
[0091]SEQ ID NO: 19 is the DNA sequence encoding Newcastle disease virus hemagglutinin-neuraminidase (HN), mRNA, complete cds. (GenBank accession AY510092).
[0092]SEQ ID NO: 20 is the protein sequence of Newcastle disease virus hemagglutinin-neuraminidase (HN). (GenBank accession AAS10195).
[0093]SEQ ID NO: 21 is the DNA sequence encoding Gallus gallus zona pellucida glycoprotein 3 (sperm receptor) (ZP3), mRNA. (GenBank accession NM--204389).
[0094]SEQ ID NO: 22 is the protein sequence of Gallus gallus zona pellucida glycoprotein 3 (sperm receptor) (ZP3). (GenBank accession NP--989720).
[0095]SEQ ID NO: 23 is the DNA sequence of Duck hepatitis B virus. (GenBank accession X58569).
[0096]SEQ ID NO: 24 is the DNA sequence of vector pCHA.
[0097]SEQ ID NO: 25 is the DNA sequence of vector pMHN.
[0098]SEQ ID NO: 26 is the DNA sequence of vector pCHN.
DEFINITIONS
[0099]As used herein, "an immunogen or immunoprotective antigen" is a non-self substance that elicits a humoral and/or cellular immune response in healthy animals such that the animal is protected against future exposure to a pathogen bearing the immunogen. The pathogens are typically agents such as viruses, bacteria, fungi and protozoa. Immunogens may also be antigenic portions of pathogens including cell wall components and viral coat proteins.
[0100]As used herein, "an immunoprotective particle" is a particle or vesicle derived from a transgenic plant cell that expresses an immunogen that, when appropriately administered to an animal, provides protection against future exposure to a pathogen bearing the immunogen.
[0101]As used herein, "vaccination or vaccinating" is defined as a means for providing protection against a pathogen by inoculating a host with an immunogenic preparation of a pathogenic agent, or a non-virulent form or part thereof, such that the host immune system is stimulated and prevents or attenuates subsequent host reactions to later exposures of the pathogen. "Providing protection" refers to stimulating an immune response as defined hereinbelow.
[0102]As used herein, "a vaccine" is a composition used to vaccinate an animal that contains at least one immunoprotective antigenic substances.
[0103]As used herein, "a pathogenic organism" is a bacterium, virus, fungus, or protozoan that causes a disease or medical condition in an animal which it has infected.
[0104]As used herein, "an adjuvant" is a substance that accentuates, increases, or enhances the immune response to an immunogen or antigen. As used herein, an increase, or accentuation or enhancement means a 2-fold or more, for example, 2, 3, 4, 5, 10, 20, 30, 40, 50, 60, 70, 80, 90, 100, or 1000-fold or more increase in the amount of antibody produced, for example, in the response to an antigen administered in the presence of an adjuvant as compared to in the absence of an adjuvant. An increase, accentuation or enhancement also means at least 5% or more antibody production, for example, 5, 6, 10, 20, 30, 40, 50, 60 70, 80, 90 or 100% or more, for example, in response to an antigen administered in the presence versus the absence of an adjuvant. Adjuvants typically enhance both the humoral and cellular immune response but an increased response to either in the absence of the other qualifies to define an adjuvant. Moreover, adjuvants and their uses are well known to immunologists and are typically employed to enhance the immune response when doses of immunogen are limited or when the immunogen is poorly immunogenic or when the route of administration is sub-optimal. Thus the term `adjuvanting amount` is that quantity of adjuvant capable of enhancing the immune response to a given immunogen or antigen. The mass that equals an adjuvanting amount will vary and is dependant on a variety of factors including but not limited to the characteristics of the immunogen, the quantity of immunogen administered, the host species, the route of administration, and the protocol for administering the immunogen. The adjuvanting amount can readily be quantified by routine experimentation given a particular set of circumstances. This is well within the ordinarily skilled artisan's purview and typically employs the use of routine dose response determinations to varying amounts of administered immunogen and adjuvant. Responses are measured by determining serum antibody titers raised in response to the immunogen using enzyme linked immunosorbant assays, radio immune assays, hemagglutination assays and the like.
[0105]As used herein, a "transgenic plant cell" refers to a plant cell which stably expresses a foreign gene, wherein the foreign gene is integrated into the plant cell chromosome and does not carry with it a viral vector sequence unique to a virus, where the foreign gene is passed onto the next cell generation and is capable of being expressed from the host plant cell chromosome. In addition, "transgenic plant material" refers to a "transgenic cell suspension" comprising one or a plurality of "transgenic plant cells" obtained by well-known cell culture techniques (Street, HE. 1973, Plant tissue and cell culture: botanical monographs. Vol II, University of California, Berkeley).
[0106]As used herein, a "trangenic plant" refers to a plant, the cells of which stably express a "heterologous" foreign gene, wherein the foreign gene is integrated into the plant cell chromosome and does not carry with it a viral vector sequence unique to a virus, where the foreign gene is passed onto the next plant generation and is capable of being expressed from the host plant cell chromosome. A "transgenic plant" comprises a "plurality of transgenic plant cells". A "transgenic plant" refers to the whole plant, or a part thereof including, but not limited to roots, stems, leaves, stalks, seeds, fruit, tubers, flowers, pollen, and the like. Examples of heterologous foreign genes include, but are not limited to, Norwalk virus capsid protein (NVCP), Avian Influenza hemagglutination antigen (AIV-HA), Newcastle Disease Virus neuraminidase (NDV-HN), zona pellucida glycoprotein 3 (ZP3), and Hepatitis B surface Antigen (HBsAg).
[0107]Transgenic plant is herein defined as a plant cell culture, plant cell line, plant, or progeny thereof derived from a transformed plant cell or protoplast, wherein the genome of the transformed plant contains foreign DNA, introduced by laboratory techniques, not originally present in a native, non-transgenic plant cell of the same species. The terms "transgenic plant" and "transformed plant" have sometimes been used in the art as synonymous terms to define a plant whose DNA contains an exogenous DNA molecule.
[0108]As used herein, an "edible plant" refers to a plant which may be consumed by an animal, has nutritional value and is not toxic. An "edible plant" may be a "food" which is a plant or a material obtained from a plant which is ingested by humans or other animals. The term "food" is intended to include plant material which may be fed to humans and other animals or a processed plant material which is fed to humans and other animals. Materials obtained from a plant are intended to include a component of a plant which is eventually ingested by a human or other animal. Examples of "edible plant" include, but are not limited to, tomato plants, rice plants, wheat plants, corn plants, carrot plants, potato plants, apple plants, soybean plants, alfalfa plants, medicago plants, vegetable plants, and fruit plants or any of the edible plants described herein.
[0109]In some cases an "edible plant" is "capable of being ingested for its nutritional value", which refers to a plant or portion thereof that provides a source of metabolizable energy, supplementary or necessary vitamins or co-factors, roughage or otherwise beneficial effect upon ingestion by an animal. Thus, where the animal to be treated by the methods of the present invention is an herbivore capable of bacterial-aided digestion of cellulose, such a food might be represented by a transgenic grass plant. Other edible plants include vegetables and fruits. Similarly, although transgenic lettuce plants, for example, do not substantially contribute energy sources, building block molecules such as proteins, carbohydrates or fats, nor other necessary or supplemental vitamins or cofactors, a lettuce plant transgenic for the nucleic acid molecules described herein used as food for an animal would fall under the definition of a food as used herein if the ingestion of the lettuce contributed roughage to the benefit of the animal, even if the animal could not digest the cellulosic content of lettuce. An "edible plant" therefore excludes tobacco.
[0110]As used herein, "immune response" refers to a response made by the immune system of an organism to a substance, which includes but is not limited to foreign or self proteins. There are three general types of "immune response" including, but not limited to mucosal, humoral and cellular "immune responses." A "mucosal immune response" results from the production of secretory IgA (sIga) antibodies in secretions that bathe all mucosal surfaces of the respiratory tract, gastrointestinal tract and the genitourinary tract and in secretions from all secretory glands (McGhee, J. R. et al., 1983, Annals NY Acad. Sci. 409). These sIgA antibodies act to prevent colonization of pathogens on a mucosal surface (Williams, R. C. et al., Science 177, 697 (1972); McNabb, P. C. et al., Ann. Rev. Microbiol. 35, 477 (1981)) and thus act as a first line of defense to prevent colonization or invasion through a mucosal surface. The production of sIgA can be stimulated either by local immunization of the secretory gland or tissue or by presentation of an antigen to either the gut-associated lymphoid tissue (GALT or Peyer's patches) or the bronchial-associated lymphoid tissue (BALT; Cebra, J. J. et al., Cold Spring Harbor Symp. Quant. Biol. 41, 210 (1976); Bienenstock, J. M., Adv. Exp. Med. Biol. 107, 53 (1978); Weisz-Carrington, P. et al., J. Immunol 123, 1705 (1979); McCaughan, G. et al., Internal Rev. Physiol 28, 131 (1983)). Membranous microfold cells, otherwise known as M cells, cover the surface of the GALT and BALT and may be associated with other secretory mucosal surfaces. M cells act to sample antigens from the luminal space adjacent to the mucosal surface and transfer such antigens to antigen-presenting cells (dendritic cells and macrophages), which in turn present the antigen to a T lymphocyte (in the case of T-dependent antigens), which process the antigen for presentation to a committed B cell. B cells are then stimulated to proliferate, migrate and ultimately be transformed into an antibody-secreting plasma cell producing IgA against the presented antigen. When the antigen is taken up by M cells overlying the GALT and BALT, a generalized mucosal immunity results with sIgA against the antigen being produced by all secretory tissues in the body (Cebra et al., supra; Bienenstock et al., supra; Weinz-Carrington et al., supra; McCaughan et al., supra). Oral immunization is therefore an important route to stimulate a generalized mucosal immune response and, in addition, leads to local stimulation of a secretory immune response in the oral cavity and in the gastrointestinal tract.
[0111]An "immune response" may be measured using techniques known to those of skill in the art. For example, serum, blood or other secretions may be obtained from an organism for which an "immune response" is suspected to be present, and assayed for the presence of the above mentioned immunoglobulins using an enzyme-linked immuno-absorbant assay (ELISA; U.S. Pat. No. 5,951,988; Ausubel et al., Short Protocols in Molecular Biology 3d Ed. John Wiley & Sons, Inc. 1995). According to the present invention, a protein of the present invention can be said to stimulate an "immune response" if the quantitative measure of immunoglobulins in an animal treated with a protein of interest detected by ELISA is statistically different (for example, is increased or decreased by 2-fold or more, for example, 2, 3, 4, 5, 10, 20, 30, 40, 50, 60, 70, 80, 90, 100, or 1000-fold or more increase or decrease in the amount of antibody produced. An increase or decrease also means at least 5% or more antibody production, for example, 5, 6, 10, 20, 30, 40, 50, 60 70, 80, 90 or 100% or more, or at least 5% or more of a decrease in antibody production) from the measure of immunoglobulins detected in an animal not treated with a protein of interest, wherein said immunoglobulins are specific for the protein of interest. A statistical test known in the art and useful to determining the difference in measured immunoglobulin levels includes, but is not limited to ANOVA, Student's T-test, and the like, wherein the P value is at least <0.1, <0.05, <0.01, <0.005, <0.001, and even <0.0001.
[0112]An "immune response" may be measured using other techniques such as immunohistochemistry using labeled antibodies which are specific for portions of the immunoglobulins raised during the "immune response". Tissue (e.g., ovarian tissue) from an animal to which a protein of interest has been administered according to the invention may be obtained and processed for immunohistochemistry using techniques well known in the art (Ausubel et al., Short Protocols in Molecular Biology 3d Ed. John Wiley & Sons, Inc. 1995). Microscopic data obtained by immunohistochemistry may be quantitated by scanning the immunohistochemically stained tissue sample and quantitating the level of staining using a computer software program known to those of skill in the art including, but not limited to NIH Image (National Institutes of Health, Bethesda, Md.). According to the present invention, a protein of the present invention can be said to stimulate an "immune response" if the quantitative measure of immunohistochemical staining in an animal treated with a protein of interest is statistically different (as defined by an increase or decrease discussed hereinabove) from the measure of immunohistochemical staining detected in an animal not treated with the protein of interest, wherein said histochemical staining requires binding specific for that protein. A statistical test known in the art may be used to determine the difference in measured immunohistochemical staining levels including, but not limited to ANOVA, Student's T-test, and the like, wherein the P value is at least <0.1, <0.05, <0.01, <0.005, <0.001, and even <0.0001.
[0113]A "mucosal immune response" may be "detected" using any of the above referenced techniques. For example, an ELISA assay may be employed using anti-IgA antibodies to detect and measure the mucosal-specific immunoglobulins (Dickinson, B. L. & Clements, J. D. Dissociation of Escherichia coli heat-labile enterotoxin adjuvanticity from ADP-ribosyltransferase activity. Infect Immun 63, 1617-1623 (1995)).
[0114]A "humoral immune response" comprises the production of antibodies in response to an antigen or antigens. A cellular immune response includes responses such as a helper T-cell (CD4.sup.+) response and a cytotoxic T-cell lymphocyte (CD8.sup.+) response. A mucosal immune response (or secretory immune response) comprises the production of secretory (sIga) antibodies. An immune response can comprise one or a combination of these responses.
[0115]As used herein, "animal" refers to an organism classified within the phylogenetic kingdom Animalia. As used herein, an "animal" also refers to a mammal. Animals, useful in the present invention, include, but are not limited to mammals, marsupials, mice, dogs, cats, cows, humans, deer, horses, sheep, livestock, poultry, chickens, turkeys, ostrich, fish, fin fish, shell fish, and the like.
[0116]As used herein, "monocotyledonous" refers to a type of plant whose embryos have one cotyledon or seed leaf. Examples of "monocots" include, but are not limited to lilies; grasses; corn; grains, including oats, wheat and barley; orchids; irises; onions and palms.
[0117]As used herein, "dicotyledonous" refers to a type of plant whose embryos have two seed halves or cotyledons. Examples of "dicots" include, but are not limited to tobacco; tomato; the legumes including alfalfa; oaks; maples; roses; mints; squashes; daisies; walnuts; cacti; violets and buttercups.
[0118]As used herein, "vector" refers to a nucleic acid molecule capable of transporting another nucleic acid to which it has been linked. One type of vector is a "plasmid", which refers to a circular double stranded nucleic acid loop into which additional nucleic acid segments can be ligated. Certain vectors are capable of autonomous replication in a host cell into which they are introduced (e.g., bacterial vectors having a bacterial origin of replication and episomal mammalian vectors). Other vectors (e.g., non-episomal mammalian vectors) are integrated into the genome of a host cell upon introduction into the host cell, and thereby are replicated along with the host genome. Moreover, certain vectors are capable of directing the expression of genes to which they are operatively linked. Such vectors are referred to herein as "expression vectors". In general, expression vectors of utility in recombinant nucleic acid techniques are often in the form of plasmids. In the present specification, "plasmid" and "vector" can be used interchangeably as the plasmid is the most commonly used form of vector.
[0119]As used herein, "promoter" refers to a sequence of DNA, usually upstream (5') of the coding region of a structural gene, which controls the expression of the coding region by providing recognition and binding sites for RNA polymerase and other factors which may be required for initiation of transcription. The selection of the promoter will depend upon the nucleic acid sequence of interest. A "plant-functional promoter" refers to a "promoter" which is capable of supporting the initiation of transcription in plant cells. "Plant-functional promoters" useful in the present invention include, but are not limited to the 35S promoter of the cauliflower mosaic virus (CaMV); promoters of seed storage protein genes such as Zma10Kz or Zmag12, light inducible genes such as ribulose bisphosphate carboxylase small subunit (rbcS), stress induced genes such as alcohol dehydrogenase (Adh1), or "housekeeping genes" that express in all cells (such as Zmact, a maize actin gene); the tomato E8 promoter; ubiquitin; mannopine synthetase (mas); rice actin 1; soybean seed protein glycinin (Gy1); soybean vegetative storage protein (vsp); and granule-bound starch synthase (gbss). Other "plant-functional promoters" include promoters for genes which are known to give high expression in edible plant parts, such as the patatin gene promoter from potato.
[0120]As used herein, "operably linked" refers to a juxtaposition wherein the components described are in a relationship permitting them to function in their intended manner. A control sequence "operably linked" to a coding sequence is ligated in such a way that expression of the coding sequence is achieved under conditions compatible with the control sequences. A promoter sequence is "operably-linked" to a gene when it is in sufficient proximity to the transcription start site of a gene to regulate transcription of the gene.
[0121]As used herein, "administered" refers to the delivery of the transgenic plant material, cells, compositions, and pharmaceutical formulations of the present invention to an animal in such a manner so to guarantee that the "delivered" material contacts a mucosal surface of the animal to which it was administered. Routes of "delivery" useful in the present invention include, but are not limited to oral delivery, nasal delivery, intraperitoneal delivery, intramuscular, intravenous or subcutaneous delivery rectal or vaginal delivery (e.g., by suppository, or topical administration), or a route of delivery wherein the delivered material directly contacts a mucosal surface (i.e., "mucosal delivery"). As used herein, "pharmaceutically acceptable" means a non-toxic material that does not interfere with the effectiveness of the biological activity of the active ingredient(s). The characteristics of the carrier will depend on the route of administration.
[0122]As used herein, a "mucosal surface", "mucosal membrane", or "mucosa" refers to the well known medical definition of these structures, which is the surface or lining of a structure comprising an epithelium, lamina propria, and, in the digestive tract, a layer of smooth muscle. Examples of "mucosal surfaces" include, but are not limited to the inner coat of the bronchi, the mucous layer of the tympanic cavity, the inner mucous coat of the colon, the inner layer of the ductus deferens, the inner coat of the esophagus, the mucous coat of the small intestine, the mucous coat of the larynx, the mucous membrane of the tongue, the pituitary membrane, the mucous membrane of the oral cavity, the mucous membrane of the pharynx, the inner mucous layer of the trachea, the lining of the auditory tube, the mucous layer of the uterine tube, the inner layer of the ureter, the inner layer of the urethra, the endometrium, the mucous membrane of the vagina, the mucous layer of the stomach, the inner coat of the urinary bladder, and the mucous membrane of the seminal vesicle.
[0123]As used herein, a "carrier" refers to an inert and non-toxic material suitable for accomplishing or enhancing delivery of the vaccine of the present invention into an animal. Examples of a carrier include, but are not limited to water, phosphate buffered saline, or saline, and further may include an adjuvant. Adjuvants such as incomplete Freund's adjuvant, aluminum phosphate, aluminum hydroxide, or alum are materials well known in the art.
[0124]The present invention also provides pharmaceutical and veterinary compositions comprising an immunoprotective particle of the present invention in combination with one or more pharmaceutically acceptable adjuvants carriers, diluents, and excipients. Such pharmaceutical compositions may also be referred to as vaccines and are formulated in a manner well known in the pharmaceutical vaccine arts.
[0125]"Administering" or "administer" is defined as the introduction of a substance into the body of an animal and includes oral, nasal, rectal, vaginal and parenteral routes. The claimed compositions may be administered individually or in combination with other therapeutic agents via any route of administration, including but not limited to subcutaneous (SQ) intramuscular (IM), intravenous (IV), mucosal, nasal or oral. The compositions may be administered via the SQ or IM route. Especially preferred is the mucosal route, and most preferred is the oral route.
[0126]As used herein, "an effective amount or dosage of the vaccine" is an amount necessary to stimulate an innate immune response as defined herein and as detected by the assays described herein as in a human or animal sufficient for the human or animal to effectively resist a challenge mounted by a pathogen. For example, in one embodiment, "an effective amount or dosage of the vaccine" causes an increase in the amount of antibody that binds to the immunoprotective antigen of the vaccine. As used herein, an increase means a 2-fold or more, for example, 2, 3, 4, 5, 10, 20, 30, 40, 50, 60, 70, 80, 90, 100, or 1000-fold or more increase in the amount of antibody produced by the vaccinated subject as compared to an unvaccinated subject. An increase also means at least 5% or more antibody production, for example, 5, 6, 10, 20, 30, 40, 50, 60 70, 80, 90 or 100% or more, by a vaccinated subject as compared to an unvaccinated subject. The dosages administered to such human or animal will be determined by a physician or veterinarian in light of the relevant circumstances including the particular immunoprotective particle or combination of particles, the condition of the human or animal, and the chosen route of administration. The dosage ranges presented herein are not intended to limit the scope of the invention in any way and are presented as general guidance for the skilled practitioner. The effective dosage can be estimated initially either in cell culture assays, or in animal models, usually mice, rabbits, dogs, or pigs. The animal model is also used to achieve a desirable concentration range and route of administration. Such information can then be used to determine useful dosages and routes for administration in humans.
[0127]The exact dosage is chosen by the individual physician in view of the patient to be treated. Dosage and administration are adjusted to provide sufficient levels of the active moiety or to maintain the desired effect. Additional factors which may be taken into account include the severity of the disease state; age, weight and gender of the subject; diet, time and frequency of administration, drug combination(s), reaction sensitivities, and tolerance/response to therapy. Long acting pharmaceutical compositions might be administered every 3 to 4 days, every week, or once every two weeks depending on half-life and clearance rate of the particular formulation.
[0128]The particular dosages of an antigenic composition of the invention will depend on many factors including, but not limited to the species, age, and general condition of the human or animal to which the composition is administered, and the mode of administration of the composition. An effective amount of the composition of the invention can be readily determined using only routine experimentation. In vitro and in vivo models (for example poultry) can be employed to identify appropriate doses. Generally, 0.1, 1.0, 1.5, 2.0, 5, 10, or 100 mg/kg of an antigen will be administered to a large mammal, such as a baboon, chimpanzee, or human. If desired, co-stimulatory molecules or adjuvants can also be provided before, after, or together with the antigenic compositions. Preferably, the dosage of antigen is administered in the range of 1 ng to 0.5 mg/kg bodyweight, more preferably, 1 mg to 50 mg/kg of body weight.
[0129]The efficacy of an edible vaccine according to the invention is determined by demonstrating that the administration of the vaccine prevents or ameliorates the symptoms of the disease being treated or caused by the pathogen of interest, by at least 5%, preferably 10-20% and more preferably, 25-100%.
[0130]"Bird" is herein defined as any warm-blooded vertebrate member of the class Aves having forelimbs modified into wings, scaly legs, a beak, and bearing young in hard-shelled eggs. For purposes of this specification, preferred groups of birds are domesticated chickens, turkeys, ostriches, ducks, geese, and cornish game hens. A more preferred group is domesticated chickens and turkeys. The most preferred bird for purposes of this invention is the domesticated chicken, including broilers and layers.
[0131]The methods and compositions of the present invention are directed toward immunizing and protecting humans and animals, preferably domestic animals, such as birds (poultry), cows, sheep, goats, pigs, horses, cats, dogs and llamas, and most preferably birds. Certain of these animal species can have multiple stomachs and digestive enzymes specific for the decomposition of plant matter, and may otherwise readily inactivate other types of oral vaccines. While not meant to be a limitation of the invention, ingestion of transgenic plant cells, and compositions derived therefrom, can result in immunization of the animals at the site of the oral mucosa including the tonsils.
[0132]As used herein, "fruit" refers to the ovary of an angiosperm flower and the associated structures (e.g. the receptacle or parts of the floral tube) that enlarge and develop to form a mass of tissue surrounding the seeds. According to the invention, the particular tissues that are involved in fruit development vary with the species, but tissues involved in fruit development according to the invention, are always derived from the maternal parent of the progeny seeds.
[0133]As used herein, "ripe" refers to a stage of fruit development that is characterized by changes in pigmentation, the conversion of acids and starches to free sugars, and breakdown of cell walls that results in softening of the fruit.
[0134]As used herein, "fruit ripening conditions" refer to conditions under which the developmental processes involved in fruit ripening can occur, including cell division and expansion of maternal tissues that occurs after fertilization of ovaries. As used herein, for example, production of ethylene is a chemical signal that stimulates the genetic program for ripening in climacteric fruits such as tomato.
[0135]As used herein, "prior to the onset of fruit ripening" refers to a stage in fruit development wherein less than 10% (for example, 9.9, 9, 8, 7, 6, 5, 4, 3, 2, 1, 0.5%) of the fruit has undergone a change in pigmentation. "Prior to the onset of fruit ripening" also refers to a stage in fruit development wherein less than 10% (for example, 9.9, 9, 8, 7, 6, 5, 4, 3, 2, 1, 0.5%) of the acids and starches of a fruit are converted to sugar. "Prior to the onset of fruit ripening" also refers to a stage in fruit development wherein less than 10% (for example, 9.9, 9, 8, 7, 6, 5, 4, 3, 2, 1, 0.5%) of the cell wall material of a fruit is degraded.
[0136]As used herein, "incubating" includes growing a plant either in the field or in a controlled or uncontrolled laboratory or indoor setting. In one embodiment of the invention, an antigen is produced in a plant by "incubating", as defined herein, the plant under conditions wherein said plant expresses the antigen prior to the onset of fruit ripening.
DETAILED DESCRIPTION OF THE INVENTION
[0137]The invention relates to sequences encoding an antigen of interest, for example a plant optimized sequence encoding HN antigen of Newcastle Disease Virus or HA antigen of Avian Influenza Virus. The invention also relates to vectors, plant cells, transgenic plants and vaccines comprising the plant optimized sequences of the invention. The invention further relates to methods of protecting against viral infection, for example infection by Newcastle Disease Virus of Avian Influenza Virus. The invention also relates to methods of antigen production in transgenic plants.
Immunoprotective Antigens Useful According to the Invention
[0138]The invention provides for plant cells and transgenic plants expressing a heterologous foreign gene. A heterologous foreign gene of the invention can be any gene of interest including but not limited to Norwalk virus capsid protein (NVCP) (Genbank Accession Number: M87661, GenBank #AF093797, Genome for Norwalk Virus, Genbank Accession Number AAB50466, for NV capsid protein), Avian Influenza hemagluttination antigen (AIV-HA) (Genbank Accession Number U67783 and AAC58999), Newcastle Disease Virus neuraminidase (NDV-HN) (Genbank Accession Numbers NM-204389, NP-989720, NP-009086, and NM-007155) (Genbank Accession Number: AY510092 and AAS10195), zona pellucida glycoprotein 3(ZP3), Hepatitis B surface Antigen (HBsAg) (Genbank Accession Numbers AF134148, AAD31865, X58569, GenBank #AF090842), shigatoxin B (StxB) (Genbank #AJ132761), staphylococcus enterotoxin B (SEB)(GenBank #M11118), E. coli labile toxin B (LT-B)(GenBank#AB01677), and E. coli labile toxin A subunit (LT-A) (GenBank #AB011677).
[0139]Newcastle's disease virus (NDV) is a member of the Paramyxovirus genus of the Paramyxoviridae. Viruses in this genus are enveloped negative-strand RNA viruses that also include parainfluenza viruses like Sendai, respiratory syncytial, mumps and measles viruses (Kingsbury et al., 1978, Intervirology, 10:137-152). Virions are characterized by the presence of two surface glycoproteins including hemagglutinin neuraminidase (HN) a 74 kDA protein and a smaller fusion (F) protein. HN is involved in two important functions including cell attachment by recognition of sialic acid containing cell receptors, and neuraminidase activity cleaving sialic acid from progeny virus particles to prevent self-agglutination. The F protein mediates virus-to-cell and cell-to-cell fusion and hemolysis. See Scheid, A., and Choppin, P. W. (1973) J. Virology. 11, 263-271; Scheid, A, and Choppin, P. W. (1974) Virology 57, 470-490; Lamb, R. A., and Kolakofsky, D. (1996). Paramyxoviridae: the viruses and their replication, p. 577-604. In B. N. Fields, D. M. Knipe, and P. M. Howley (ed.), Fields virology, 3rd ed. Lippincott-Raven Publishers, Philadelphia, Pa. Polyvalent sera prepared against either protein are capable of neutralizing the infectivity of the virus. See Mertz, D. C., Scheid, A., and Choppin, P. W. (1980) J. Exp. Med. 151, 275-288.
[0140]Avian influenza virus is described in Suarez et al., Virus Res. 1997, 51:115 and Sockett, Can. Med. Assoc. J., 1998, 158:369, incorporated herein by reference in their entirety. The hemagglutinin gene of avian influenza virus is described in Barun et al., 1998, Nuc. Acids. Res., 16:4181, incorporated herein by reference in its entirety.
Preparation of the Constructs of the Invention
[0141]An expression cassette according to the invention comprises a DNA sequence encoding at least one immunoprotective antigen operably linked to transcriptional and translational control regions functional in a plant cell. Preferably the invention provides plant expression cassettes that are useful for expressing immunoprotective antigen transgenes in plants. These cassettes comprise the following elements that are operably linked from 5' to 3':
[0142]A) a plant gene promoter sequence that naturally expresses in plants;
[0143]B) a nucleic acid sequence encoding an immunoprotective antigen of interest; and
[0144]C) a 3'UTR.
[0145]Promoters useful in this embodiment are any known promoters that are functional in a plant. Many such promoters are well known to the ordinarily skilled artisan. Such promoters include promoters normally associated with other genes, and/or promoters isolated from any bacterial, viral, eukaryotic, or plant cell. It may be advantageous to employ a promoter that effectively directs the expression of the foreign coding sequence in the cell or tissue type chosen for expression. The use of promoter and cell type combinations for protein expression is generally known to those of skill in the art of molecular biology, for example, see Sambrook et al., In: Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y., 1989. The promoters employed may be constitutive, or inducible, and can be used under the appropriate conditions to direct high level expression of the introduced DNA segment, such as is advantageous in the large-scale production of recombinant proteins or peptides. The term "constitutive" used in the context of a promoter means that the promoter is capable of directing transcription of an operably linked nucleic acid sequence in the absence of a stimulus (e.g., heat shock, chemicals, etc.). In contrast, an "inducible" promoter is one which is capable of directing a level of transcription of an operably linked nucleic acid sequence in the presence of a stimulus (e.g., heat shock, chemicals, etc.), wherein the level of the transcription is different from that in the absence of the stimulus. As used herein, "inducible" also refers to expressed in the presence of an exogenous or endogenous chemical (for example an alcohol, a hormone, or a growth factor), in the presence of light and/or in response to developmental changes. As used herein, "inducible" also refers to expressed in any tissue in the presence of a chemical inducer". As used herein, "chemical induction" according to the invention refers to the physical application of a exogenous or endogenous substance (including macromolecules e.g. proteins, or nucleic acids) to a plant or a plant organ (e.g. by spraying a liquid solution comprising a chemical inducer on leaves, application of a liquid solution to roots or exposing plants or plant organs to gas or vapor) which has the effect of causing the target promoter present in the cells of the plant or plant organ to increase the rate of transcription.
[0146]Some exemplary plant functional promoters, which can be used to express a structural gene of the present invention, are among the following: CaMV 35S and 19S promoters (U.S. Pat. No. 5,352,605 and U.S. Pat. No. 5,530,196); patatin promoter (U.S. Pat. No. 5,436,393); a B33 promoter sequence of a patatin gene derived from Solanum tuberosum, and which leads to a tuber specific expression of sequences fused to the B33 promoter (U.S. Pat. No. 5,436,393); tomato E8 promoter (WO 94/24298); tomato fruit promoters (U.S. Pat. No. 5,556,653); -a plant ubiquitin promoter system (U.S. Pat. Nos. 5,614,399 and 5,510,474); 5 cis-regulatory elements of abscisic acid-responsive gene expression (U.S. Pat. No. 5,824,865); promoter from a badnavirus, rice tungro bacilliform virus (RTBV) (U.S. Pat. No. 5,824,857); a chemically inducible promoter fragment from the 5' flanking region adjacent the coding-region of a tobacco PR-1a gene (U.S. Pat. No. 5,789,214); a raspberry drul promoter (U.S. Pat. No. 5,783,394); strawberry promoters and genes (WO 98/31812); promoter is the napin promoter, the phaseolin promoter, and the DC3 promoter (U.S. Pat. No. 5,773,697); a LEA promoter (U.S. Pat. No. 5,723,765); 5' transcriptional regulatory region for sink organ specific expression (U.S. Pat. No. 5,723,757); G-box related sequence motifs, specifically Iwt and PA motifs, which function as cis-elements of promoters, to regulate the expression of heterologous genes in transgenic plants (U.S. Pat. No. 5,723,751); P119 promoters and their use (U.S. Pat. No. 5,633,440); Group 2 (Gp2) plant promoter sequences (U.S. Pat. No. 5,608,144); nucleic acid promoter fragments derived from several genes from corn, petunia and tobacco (U.S. Pat. No. 5,608,143); promoter sequences isolated from the nuclear gene for chloroplast GS2 glutamine synthetase and from two nuclear genes for cytosolic GS3 glutamine synthetase in the pea plant, Pisum sativum (U.S. Pat. No. 5,391,725); full-length transcript promoter from figwort mosaic virus (FMV) (U.S. Pat. No. 5,378,619); an isocitrate lyase promoter (U.S. Pat. No. 5,689,040); a microspore-specific regulatory element (U.S. Pat. No. 5,633,438); expression of heterologous genes in transgenic plants and plant cells using plant asparagine synthetase promoters (U.S. Pat. No. 5,595,896); a promoter region that drives expression of a 1450 base TR transcript in octopine-type crown gall tumors (U.S. Pat. No. 4,771,002); promoter sequences from the gene from the small subunit of ribulose-1,5-bisphosphate carboxylase (U.S. Pat. No. 4,962,028); the Arabidopsis histone H4 promoter (U.S. Pat. No. 5,491,288); a seed-specific plant promoter (U.S. Pat. No. 5,767,363); a 21 bp promoter element which is capable of imparting root expression capability to a rbcS-3A promoter, normally a green tissue specific promoter (U.S. Pat. No. 5,023,179); promoters of tissue-preferential transcription of associated DNA sequences in plants, particularly in the roots (U.S. Pat. No. 5,792,925); Brassica sp. polygalacturonase promoter (U.S. Pat. No. 5,689,053); a seed coat-specific cryptic promoter region (U.S. Pat. No. 5,824,863); a chemically inducible nucleic acid promoter fragment isolated from the tobacco PR-1a gene inducible by application of a benzo-1,2,3-thiadiazole, an isonicotinic acid compound, or a salicylic acid compound (U.S. Pat. No. 5,689,044); promoter fragment isolated from a cucumber chitinase/lysozyme gene that is inducible by application of benzo-1,2,3-thiadiazole (U.S. Pat. No. 5,654,414); a constitutive promoter from tobacco that directs expression in at least ovary, flower, immature embryo, mature embryo, seed, stem, leaf and root tissues (U.S. Pat. No. 5,824,872); alteration of gene expression in plants (U.S. Pat. No. 5,223,419); a recombinant promoter for gene expression in monocotyledenous plants (U.S. Pat. No. 5,290,924); method for using TMV to overproduce peptides and proteins (WO 95/21248); nucleic acid comprising shoot meristem-specific promoter and regulated sequence (WO 98/05199); phaseolin promoter and structural gene (EP-B-0122791); plant promoters [sub domain of CaMV 35S] (U.S. Pat. No. 5,097,025); use of tomato E8-derived promoters to express heterologous genes, e.g. 5-adenosylmethionine hydrolase in ripening fruit (WO 94/24294); method of using transactivation proteins to control gene expression in transgenic plants (U.S. Pat. No. 5,801,027); DNA molecules encoding inducible plant promoters and tomato Adh2 enzyme (U.S. Pat. No. 5,821,398); synthetic plant core promoter and upstream regulatory element (WO 97/47756); monocot having dicot wound inducible promoter (U.S. Pat. No. 5,684,239); selective gene expression in plants (U.S. Pat. No. 5,110,732); CaMV 35S enhanced mannopine synthase promoter and method for using the same (U.S. Pat. No. 5,106,739); seed specific transcription regulation (U.S. Pat. No. 5,420,034); seed specific promoter region (U.S. Pat. No. 5,623,067); DNA promoter fragments from wheat (U.S. Pat. No. 5,139,954); chimeric regulatory regions and gene cassettes for use in plants (WO 95/14098); production of gene products to high levels (WO 90/13658); HMG promoter expression system and post harvest production of gene products in plants and plant cell cultures (U.S. Pat. No. 5,670,349); gene expression system comprising the promoter region of the alpha amylase genes in plants (U.S. Pat. No. 5,712,112).
[0147]A preferred group of promoters is the cassaya vein mosaic virus promoters described in U.S. patent application Ser. No. 09/202,838, herein incorporated by reference in its entirety; the phaseolin promoters described in U.S. Pat. No. 5,591,605, herein incorporated by reference in its entirety; rice actin promoters described in U.S. Pat. No. 5,641,876, herein incorporated by reference in its entirety; the per5 promoter described in WO 98/56921, herein incorporated by reference in its entirety; and the gamma zein promoters described in WO 00/12681.
[0148]A promoter DNA sequence is said to be "operably linked" to a coding DNA sequence if the two are situated such that the promoter DNA sequence influences the transcription of the coding DNA sequence. For example, if the coding DNA sequence codes for the production of a protein, the promoter DNA sequence would be operably linked to the coding DNA sequence if the promoter DNA sequence affects the expression of the protein product from the coding DNA sequence.
[0149]Construction of gene cassettes is readily accomplished utilizing well known methods, such as those disclosed in Sambrook et al. (1989); and Ausubel et al. (1987) Current Protocols in Molecular Biology, John Wiley and Sons, New York, N.Y. The present invention also includes DNA sequences having substantial sequence homology with the disclosed sequences encoding immunoprotective antigens such that they are able to have the disclosed effect on expression. As used in the present application, the term "substantial sequence homology" is used to indicate that a nucleotide sequence (in the case of DNA or RNA) or an amino acid sequence (in the case of a protein or polypeptide) exhibits substantial, functional or structural equivalence with another nucleotide or amino acid sequence. Any functional or structural differences between sequences having substantial sequence homology will be de minimis; that is they will not affect the ability of the sequence to function as indicated in the present application. Sequences that have substantial sequence homology with the sequences disclosed herein are usually variants of the disclosed sequence, such as mutations, but may also be synthetic sequences.
[0150]In most cases, sequences having 95% homology to the sequences specifically disclosed herein will function as equivalents, and in many cases considerably less homology, for example 75% or 80%, will be acceptable. Locating the parts of these sequences that are not critical may be time consuming, but is routine and well within the skill in the art. Exemplary techniques for modifying oligonucleotide sequences include using polynucleotide-mediated, site-directed mutagenesis. See Zoller et al. (1984); Higuchi et al. (1988); Ho et al. (1989); Horton et al. (1989); and PCR Technology: Principles and Applications for DNA Amplification, (ed.) Erlich (1989).
[0151]The invention provides for a plant optimized sequence encoding an immunoprotective antigen of interest. A plant-optimized coding sequence is designed with hybrid codon preference reflecting tomato and potato codon usage (Ausubel F., et al., eds. (1994) Current Protocols in Molecular Biology, vol. 3, p. A.1C.3 Haq TA, Mason H S, Clements J D, Arntzen C J (1995).
[0152]A plant optimized sequence of the invention can be prepared as described in U.S. Pat. No. 5,380,831, incorporated by reference herein in its entirety. In general, the frequency of codon usage for a target plant of interest is used to adjust the codon usage frequency of a target gene of interest, for example, NDV HN.
[0153]The native sequence is scanned for sequence motifs that might result in interference with expression in the target plant, such as poly-A addition sites, Shaw/Kamen degradation sites, splice junction sites, and anything related to RNA termination or potential hairpin formation, etc. Runs of A/T sequences are often avoided. In one embodiment, it is preferable to keep strings of A/T to four or fewer in a row, if possible, since most regulatory sites tend to contain runs of A's and T's (e.g. AATAAA consensus poly-A or ATTTA Shaw/Kamen). In general it is useful to scan for about 16 putative poly-A addition sequences based on identified sites from plants found in the literature. In certain embodiments, it is useful to search for C/G runs since they can stabilize hairpin stem formation. Since monocots tend to favor third position C's and G's somewhat more than dicots, the relevance of identification of C/G runs may depend on the host plant target for expression.
[0154]In one embodiment wherein a gene is expressed in both dicots and monocots, an overall plant codon usage frequency is used as a basis for sequence optimization.
[0155]For the purposes of the present invention the term membrane anchor sequence contemplates that which the ordinarily skilled artisan understands about the term. Membrane anchor sequences include transmembrane protein sequences and are found in many naturally occurring proteins. Such membrane anchor sequences vary in size but always are comprised of a series of amino acids with lipophilic or aliphatic side chains that favor the hydrophobic environment within the membrane. During RNA translation and post translational processing, the anchor sequences integrate and become embedded in the cell membrane and function to anchor, or loosely attach the protein to a cellular membrane component allowing hydrophilic portions of the protein to be exposed to, and interact with, the aqueous milieu inside or outside of the cell.
[0156]In preparing the constructs of this invention, the various DNA fragments may be manipulated, so as to provide for the DNA sequences in the proper orientation and, as appropriate, in the proper reading frame. Adapters or linkers may be employed for joining the DNA fragments or other manipulations may be involved to provide for convenient restriction sites, removal of superfluous DNA, removal of restriction sites, or the like.
[0157]In carrying out the various steps, cloning is employed, so as to amplify a vector containing the promoter/gene of interest for subsequent introduction into the desired host cells. A wide variety of cloning vectors are available, where the cloning vector includes a replication system functional in E. coli and a marker which allows for selection of the transformed cells. Illustrative vectors include pBR322, pUC series, pACYC184, Bluescript series (Stratagene) etc. Thus, the sequence may be inserted into the vector at an appropriate restriction site(s), the resulting plasmid used to transform the E. coli host (e.g., E. coli strains HB101, JM101 and DH5α), the E. coli grown in an appropriate nutrient medium and the cells harvested and lysed and the plasmid recovered. Analysis may involve sequence analysis, restriction analysis, electrophoresis, or the like. After each manipulation the DNA sequence to be used in the final construct may be restricted and joined to the next sequence, where each of the partial constructs may be cloned in the same or different plasmids.
[0158]Vectors are available or can be readily prepared for transformation of plant cells. In general, plasmid or viral vectors should contain all the DNA control sequences necessary for both maintenance and expression of a heterologous DNA sequence in a given host. Such control sequences generally include a leader sequence and a DNA sequence coding for translation start-signal codon, a translation terminator codon, and a DNA sequence coding for a 3' UTR signal controlling messenger RNA processing. Selection of appropriate elements to optimize expression in any particular species is a matter of ordinary skill in the art utilizing the teachings of this disclosure. Finally, the vectors should desirably have a marker gene that is capable of providing a phenotypical property which allows for identification of host cells containing the vector.
[0159]The activity of the foreign coding sequence inserted into plant cells is dependent upon the influence of endogenous plant DNA adjacent to the insert. Generally, the insertion of heterologous genes appears to be random using any transformation technique; however, technology currently exists for producing plants with site specific recombination of DNA into plant cells (see WO 91/09957). Any method or combination of methods resulting in the expression of the desired sequence or sequences under the control of the promoter is acceptable.
[0160]The present invention is not limited to any particular method for transforming plant cells. Technology for introducing DNA into plant cells is well-known to those of skill in the art. Four basic methods for delivering foreign DNA into plant cells have been described. Chemical methods (Graham and van der Eb, Virology, 54(02):536-539, 1973; Zatloukal, Wagner, Cotten, Phillips, Plank, Steinlein, Curiel, Bimstiel, Ann. N.Y. Acad. Sci., 660:136-153, 1992); Physical methods including microinjection (Capecchi, Cell, 22(2):479-488, 1980), electroporation (Wong and Neumann, Biochim. Biophys. Res. Conmmun. 107(2):584-587, 1982; Fromm, Taylor, Walbot, Proc. Natl. Acad. Sci. USA, 82(17):5824-5828, 1985; U.S. Pat. No. 5,384,253) and the gene gun (Johnston and Tang, Methods Cell. Biol., 43(A):353-365, 1994; Fynan, Webster, Fuller, Haynes, Santoro, Robinson, Proc. Natl. Acad. Sci. USA 90(24):11478-11482, 1993); Viral methods (Clapp, Clin. Perinatol., 20(1):155-168, 1993; Lu, Xiao, Clapp, Li, Broxmeyer, J. Exp. Med. 178(6):2089-2096, 1993; Eglitis and Anderson, Biotechniques, 6(7):608-614, 1988; Eglitis, Kantoff, Kohn, Karson, Moen, Lothrop, Blaese, Anderson, Avd. Exp. Med. Biol., 241:19-27, 1988); and Receptor-mediated methods (Curiel, Agarwal, Wagner, Cotten, Proc. Natl. Acad. Sci. USA, 88(19):8850-8854, 1991; Curiel, Wagner, Cotten, Bimstiel, Agarwal, Li, Loechel, Hu, Hum. Gen. Ther., 3(2):147-154, 1992; Wagner et al., Proc. Natl. Acad. Sci. USA, 89, (13):6099-6103, 1992).
[0161]The introduction of DNA into plant cells by means of electroporation is well-known to those of skill in the art. Plant cell wall-degrading enzymes, such as pectin-degrading enzymes, are used to render the recipient cells more susceptible to transformation by electroporation than untreated cells. To effect transformation by electroporation one may employ either friable tissues such as a suspension culture of cells, or embryogenic callus, or immature embryos or other organized tissues directly. It is generally necessary to partially degrade the cell walls of the target plant material to pectin-degrading enzymes or mechanically wounding in a controlled manner. Such treated plant material is ready to receive foreign DNA by electroporation.
[0162]Another method for delivering foreign transforming DNA to plant cells is by microprojectile bombardment. In this method, microparticles are coated with foreign DNA and delivered into cells by a propelling force. Such micro particles are typically made of tungsten, gold, platinum, and similar metals. An advantage of microprojectile bombardment is that neither the isolation of protoplasts (Cristou et al., 1988, Plant Physiol., 87:671-674) nor the susceptibility to Agrobacterium infection is required. An illustrative embodiment of a method for delivering DNA into maize cells by acceleration is a Biolistics Particle Delivery System, which can be used to propel particles coated with DNA or cells through a screen onto a filter surface covered with corn cells cultured in suspension. The screen disperses the particles so that they are not delivered to the recipient cells in large aggregates. For the bombardment, cells in suspension are preferably concentrated on filters or solid culture medium. Alternatively, immature embryos or other target cells may be arranged on solid culture medium. The cells to be bombarded are positioned at an appropriate distance below the macroprojectile stopping plate. In bombardment transformation, one may optimize the prebombardment culturing conditions and the bombardment parameters to yield the maximum numbers of stable transformants. Both the physical and biological parameters for bombardment are important in this technology. Physical factors are those that involve manipulating the DNA/microprojectile precipitate or those that affect the flight and velocity of the microprojectiles. Biological factors include all steps involved in manipulation of cells before and immediately after bombardment, the osmotic adjustment of target cells to help alleviate the trauma associated with bombardment, and also the nature of the transforming DNA, such as linearized DNA or intact supercoiled plasmids.
[0163]Agrobacterium-mediated transfer is a widely applicable system for introducing foreign DNA into plant cells because the DNA can be introduced into whole plant tissues, eliminating the need to regenerate an intact plant from a protoplast. The use of Agrobacterium-mediated plant integrating vectors to introduce DNA into plant cells is well known in the art. See, for example, the methods described in Fraley et al., 1985, Biotechnology, 3:629; Rogers et al., 1987, Meth. in Enzymol., 153:253-277. Further, the integration of the Ti-DNA is a relatively precise process resulting in few rearrangements. The region of DNA to be transferred is defined by the border sequences, and intervening DNA is usually inserted into the plant genome as described in Spielmann et al., 1986, Mol. Gen. Genet., 205:34; Jorgensen et al., 1987, Mol. Gen. Genet., 207:471.
[0164]Modern Agrobacterium transformation vectors are capable of replication in E. coli as well as Agrobacterium, allowing for convenient manipulations. Moreover, recent technological advances in vectors for Agrobacterium-mediated gene transfer have improved the arrangement of genes and restriction sites in the vectors to facilitate construction of vectors capable of expressing various proteins or polypeptides. Convenient multi-linker regions flanked by a promoter and a polyadenylation site for direct expression of inserted polypeptide coding genes are suitable for present purposes. In addition, Agrobacterium containing both armed and disarmed Ti genes can be used for the transformations.
[0165]Transformation of plant protoplasts can be achieved using methods based on calcium phosphate precipitation, polyethylene glycol treatment, electroporation, and combinations of these treatments (see, e.g., Potrykus et al., 1985, Mol. Gen. Genet., 199:183; Marcotte et al., Nature, 335:454, 1988). Application of these systems to different plant species depends on the ability to regenerate the particular species from protoplasts.
[0166]Once the plant cells have been transformed, selected and checked for antigen expression, it is possible in some cases to regenerate whole fertile plants. This will greatly depend on the plant species chosen. Methods for regenerating numerous plant species have been reported in the literature and are well known to the skilled artisan. For practice of the present invention, it is preferable to transform plant cell lines that can be cultured and scaled-up rapidly by avoiding the generally lengthy regeneration step. In addition the use of plant cell cultures avoids open field production and greatly reduces the chances of gene escape and food contamination. Tobacco suspension cell cultures such as NT-1 and BY-2 (An, G., 1985 Plant Physiol. 79, 568-570) are preferred because these lines are particularly susceptible to handling in culture, are readily transformed, produce stably integrated events and are amenable to cryopreservation.
[0167]The tobacco suspension cell line, NT-1, is suitable for the practice of the present invention. NT-1 cells were originally developed from Nicotiana tabacum L.cv. bright yellow 2. The NT-1 cell line is widely used and readily available; though, any tobacco suspension cell line is consistent with the practice of the invention. It is worth noting that the origins of the NT-1 cell line are unclear. Moreover, the cell line appears variable and is prone to change in response to culture conditions. NT-1 cells suitable for use in the examples below are available from the American Type Culture Collection under accession number ATCC No. 74840. See also U.S. Pat. No. 6,140,075, herein incorporated by reference.
[0168]Many plant cell culture techniques and systems ranging from laboratory-scale shaker flasks to multi-thousand liter bioreactor vessels have been described and are well know in the art of plant cell culture. See for example Fischer, R. et al, 1999 Biotechnol. Appl. Biochem. 30, 109-112 and Doran, P., 2000 Current Opinions in Biotechnology 11, 199-204. After the transformed plant cells have been cultured to the mass desired, they are harvested, gently washed and placed in a suitable buffer for sonication. Many different buffers are compatible with the present invention. In general the buffer is an aqueous isotonic buffered salt solution at or near a neutral pH value that does not contain any detergent. Preferred buffers include Dulbeccos Phosphate Buffered Saline and PBS containing 1 mM EDTA.
[0169]For sonication, the washed cells are placed in buffer in a range of about 0.01 gm/ml to about 5.0 gm/ml, preferably in a range of about 0.1 gm/ml to about 0.5 gm/ml (washed wet weight cells per volume of buffer). Many commercially available sonication instruments are consistent with the invention and sonication times range from about 5 to about 20 seconds, preferably about 15 to about 20 seconds. The resulting particles are membrane vesicles that may range in size from a few microns to several hundred microns and expose the recombinant, immunoprotective, anchored proteins.
[0170]An immunoprotective agent or antigen of interest is expressed and isolated according to methods well known in the art and described in the examples herein below.
[0171]In one embodiment, a method of producing an antigen of interest comprises preparing a transgenic plant comprising a vector encoding the antigen. The plant is incubated under conditions wherein the plant expresses the antigen prior to the onset of ripening of the plant. According to this embodiment, the plant has a fruit that ripens (including but not limited to tomato, banana, citrus, melon, strawberry, pineapple, stonefruit, mango, pumpkin, squash etc.) The antigen produced according to this method can be isolated from the plant, or from the fruit of the plant prior to administration. Alternatively, the antigen is not isolated from the plant but is administed in a crude, food-processed or raw form. The details of this method are described in the Examples below.
Plants Useful According to the Invention
[0172]The present invention also provides for a transgenic plant transformed with the constructs of the invention. Plants that can be used for practice of the present invention include any dicotyledon and monocotyledon. These include, but are not limited to, tobacco, tomato, potato, eggplant, pepino, yam, soybean, pea, sugar beet, lettuce, bell pepper, celery, carrot, asparagus, onion, grapevine, muskmelon, strawberry, rice, sunflower, rapeseed/canola, wheat, oats, maize, cotton, walnut, spruce/conifer, poplar and apple, berries such as strawberries, raspberries, alfalfa and banana. Since many edible plants used by humans for food or as components of animal feed are dicotyledenous plants, dicotyledons are typically employed, although monocotyledon transformation is also applicable especially in the production of certain grains useful for animal feed. It is particularly advantageous in certain disease prevention for human infants to produce a vaccine in a juice for ease of administration to humans such as juice of tomato, soybean, and carrot, or milk. Cells and seeds derived from these plant vaccines are also useful according to the invention.
[0173]Representative plants that have been transformed with this system and representative references are listed in Table A. Other plants having edible parts, or which can be processed to afford isolated protein, can be transformed by the same methods or routine modifications thereof.
TABLE-US-00001 TABLE A Plant Reference Tobacco Barton, K. et al., (1983) Cell 32, 1033 Tomato Fillatti, J. et al., (1987) Bio/Technology 5, 726-730 Potato Hoekema, A. et al., (1989) Bio/Technology 7: 273-278 Eggplant Filipponee, E. et al., (1989) Plant Cell Rep. 8: 370-373 Pepino Atkinson, R. et al., (1991) Plant Cell Rep. 10: 208-212 Yam Shafer, W. et al., (1987) Nature. 327: 529-532 Soybean Delzer, B., et al., (1990) Crop Sci. 30: 320-322 Pea Hobbs, S. et al., (1989) Plant Cell Rep. 8: 274-277 Sugar beet Kallerhoff, J. et al., (1990) Plant Cell Rep. 9: 224-228 Lettuce Michelmore, R., et al., (1987) Plant Cell Rep. 6: 439-442 Bell pepper Liu, W. et al., (1990) Plant Cell Rep. 9: 360-364 Celery Liu, C-N. et al., (1992) Plant Mol. Biol. 1071-1087 Carrot Liu, C-N. et al, (1992) Plant Mol Biol. 1071-1087 Asparagus Delbriel, B. et al., (1993) Plant Cell Rep. 12: 129-132 Onion Dommisse, E. et al.; (1990) Plant Sci. 69: 249-257 Grapevine Baribault, T., et al., (1989) Plant Cell Rep. 8: 137-140 Muskmelon Fang, G., et al., (1990) Plant Cell Rep. 9: 160-164 Strawberry Nehra, N. et al., (1990) Plant Cell Rep. 9: 10-13 Rice Raineri, D. et al., (1990) Bio/Technology. 8: 33-38 Sunflower Schrammeijer, B. et al., (1990) Plant Cell Rep. 9: 55-60 Rapeseed/ Pua, E. et al., (1987) Bio/Technology 5. 815 Canola Wheat Mooney, P. et al., (1991) Plant Cell Tiss. Organ Cult. 25: 209-218 Oats Donson, J. et al., (1988) Virology. 162: 248-250 Maize Gould, J. et al., (1991) Plant Physiol. 95: 426-434 Alfalfa Chabaud, M. et al., (1988) Plant Cell Rep. 7: 512-516 Cotton Umbeck, P. et al., (1987) Bio/Technology. 5: 263-266 Walnut McGranahan, G. et al., (1990) Plant Cell Rep. 8: 512-516 Spruce/Conifer Ellis, D. et al., (1989) Plant Cell Rep. 8: 16-20 Poplar Python, F. et al., (1987) Bio/Technology 5: 1323 Apple James, P. et al., (1989) Plant Cell Rep. 7: 658-661
[0174]A transgenic plant transformed with a vector described hereinabove is another aspect of the present invention.
[0175]Potato varieties FL 1607 ("Frito Lay 1607") and Desiree, and tomato variety Tanksley TA234TM2R are particularly preferred varieties, which have been transformed with binary vectors using the methods described herein. Of these transformed varieties, Desiree is the only commercial variety; the other varieties can be obtained from Frito-Lay (Rhinelander, Wis.) and Steve Tanksley (Dept. of Plant Breeding, Cornell Univ.). Potato variety FL1607 allows rapid transformation but is not a good agronomic variety as it suffers from hollow heart.
[0176]Tomato is preferred as a model system for expression of foreign proteins because of its ease of genetic transformation, and because fruit-specific, ripening dependent promoters are available for regulated expression (Giovannoni et al., 1989).
[0177]The invention includes whole plants, plant cells, plant organs, plant tissues, plant seeds, protoplasts, callus, cell cultures, and any group of plant cells organized into structural and/or functional units capable of expressing at least a polynucleotide of the invention. Preferably, whole plants, plant cells, plant organs, plant tissues, plant seeds, protoplasts, callus, cell cultures, and any group of plant cells produce 0.001, 0.01, 1, 5, 10, 25, 50, 100, 500, or 11000 μg of polypeptide of the invention per gram of total soluble plant material.
Use, Dosage and Administration of a Vaccine According to the Invention
[0178]Food plant produced antigens provide a less expensive source of antigen, that does not require animal-sourced components, for the preparation of vaccines.
[0179]The vaccines according to the invention are useful for protection against a pathogen of interest and against viral infection.
[0180]1. Administration
[0181]The invention provides for methods of administering a vaccine according to the invention to a mammal to prevent viral infection.
[0182]In one embodiment, a vaccine is administered orally (either by feeding or by oral gavage) to ensure inducing a mucosal immune response as well as to take advantage of cost and convenience. Conveniently, an oral administration step entails consuming a transgenic plant or plant part according to the invention. An edible vaccine according to the invention can be in the form of a plant part, an extract, a juice, a liquid, a powder or a tablet.
[0183]An vaccine according to the invention may also be administered by via an intranasal route in the form of a nasal spray. Alternatively, a vaccine according to the invention may be administered orally, intraperitoneally, intramuscularly, intravenously, or subcutaneously.
[0184]The invention provides for compositions comprising an edible vaccine admixed with a physiologically compatible carrier. As used herein, "physiologically compatible carrier" refers to a physiologically acceptable diluent such as water, phosphate buffered saline, or saline, and further may include an adjuvant. Adjuvants such as incomplete Freund's adjuvant, aluminum phosphate, aluminum hydroxide, or alum are materials well known in the art.
[0185]The invention also provides for pharmaceutical compositions. In addition to the active ingredients, these pharmaceutical compositions may contain suitable pharmaceutically acceptable carrier preparations which can be used pharmaceutically.
[0186]Pharmaceutical compositions for oral administration can be formulated using pharmaceutically acceptable carriers well known in the art in dosages suitable for oral administration. Such carriers enable the pharmaceutical compositions to be formulated as tablets, pills, dragees, capsules, liquids, gels, syrups, slurries, suspensions and the like, for ingestion by the patient.
[0187]Pharmaceutical preparations for oral use can be obtained through combination of active compounds with solid excipient, optionally grinding a resulting mixture, and processing the mixture of granules, after adding suitable auxiliaries, if desired, to obtain tablets or dragee cores. Suitable excipients are carbohydrate or protein fillers such as sugars, including lactose, sucrose, mannitol, or sorbitol; starch from corn, wheat, rice, potato, or other plants; cellulose such as methyl cellulose, hydroxypropylmethyl-cellulose, or sodium carboxymethyl cellulose; and gums including arabic and tragacanth; and proteins such as gelatin and collagen. If desired, disintegrating or solubilizing agents may be added, such as the cross-linked polyvinyl pyrrolidone, agar, alginic acid, or a salt thereof, such as sodium alginate.
[0188]Dragee cores are provided with suitable coatings such as concentrated sugar solutions, which may also contain gum arabic, talc, polyvinylpyrrolidone, carbopol gel, polyethylene glycol, and/or titanium dioxide, lacquer solutions, and suitable organic solvents or solvent mixtures. Dyestuffs or pigments may be added to the tablets or dragee coatings for product identification or to characterize the quantity of active compound, i.e., dosage.
[0189]Pharmaceutical preparations which can be used orally include push-fit capsules made of gelatin, as well as soft, sealed capsules made of gelatin and a coating such as glycerol or sorbitol. Push-fit capsules can contain active ingredients mixed with a filler or binders such as lactose or starches, lubricants such as talc or magnesium stearate, and, optionally, stabilizers. In soft capsules, the active compounds may be dissolved or suspended in suitable liquids, such as fatty oils, liquid paraffin, or liquid polyethylene glycol with or without stabilizers.
[0190]Pharmaceutical formulations for parenteral administration include aqueous solutions of active compounds. For injection, the pharmaceutical compositions of the invention may be formulated in aqueous solutions, preferably in physiologically compatible buffers such as Hank's solution, Ringer's solution, or physiologically buffered saline. Aqueous injection suspensions may contain substances which increase the viscosity of the suspension, such as sodium carboxymethyl cellulose, sorbitol, or dextran. Additionally, suspensions of the active solvents or vehicles include fatty oils such as sesame oil, or synthetic fatty acid esters, such as ethyl oleate or triglycerides, or liposomes. Optionally, the suspension may also contain suitable stabilizers or agents which increase the solubility of the compounds to allow for the preparation of highly concentrated solutions.
[0191]For nasal administration, penetrants appropriate to the particular barrier to be permeated are used in the formulation. Such penetrants are generally known in the art.
[0192]2. Manufacture and Storage
[0193]The pharmaceutical compositions of the present invention may be manufactured in a manner known in the art, e.g. by means of conventional mixing, dissolving, granulating, dragee-making, levitating, emulsifying, encapsulating, entrapping or lyophilizing processes.
[0194]The pharmaceutical composition may be provided as a salt and can be formed with many acids, including but not limited to hydrochloric, sulfuric, acetic, lactic, tartaric, malic, succinic, etc . . . Salts tend to be more soluble in aqueous or other protonic solvents that are the corresponding free base forms. In other cases, the preferred preparation may be a lyophilized powder in 1 mM-50 mM histidine, 0.1%-2% sucrose, 2%-7% mannitol at a pH range of 4.5 to 5.5 that is combined with buffer prior to use.
[0195]After pharmaceutical compositions comprising a compound of the invention formulated in an acceptable carrier have been prepared, they can be placed in an appropriate container and labeled for treatment of an indicated condition with information including amount, frequency and method of administration.
[0196]3. Therapeutically Effective Dose
[0197]Pharmaceutical compositions suitable for use in the present invention include compositions wherein the active ingredients are contained in an effective amount to achieve the intended purpose. The determination of an effective dose is well within the capability of those skilled in the art.
[0198]For any compound, the therapeutically effective dose can be estimated initially either in cell culture assays, or in animal models, usually birds, mice, rabbits, dogs, or pigs. The animal model is also used to achieve a desirable concentration range and route of administration. Such information can then be use to determine useful doses and routes for administration in humans.
[0199]A therapeutically effective dose refers to that amount of protein or its antibodies, antagonists, or inhibitors which prevent or ameliorate the symptoms or conditions, for example caused by viral infection. Therapeutic efficacy and toxicity of such compounds can be determined by standard pharmaceutical procedures in cell cultures or experimental animals, eg, ED50 (the dose therapeutically effective in 50% of the population) and LD50 (the dose lethal to 50% of the population). The dose ratio between therapeutic and toxic effects is the therapeutic index, and it can be expressed as the ratio, LD50/ED50. Pharmaceutical compositions which exhibit large therapeutic indices are preferred. The data obtained from cell culture assays and animals studies is used in formulating a range of dosage for human use. The dosage of such compounds lies preferably within a range of circulating concentrations that include the ED50 with little or no toxicity. The dosage varies within this range depending upon the dosage from employed, sensitivity of the patient, and the route of administration.
[0200]The exact dosage is chosen by the individual physician or veterinarian in view of the patient to be treated. Dosage and administration are adjusted to provide sufficient levels of the active moiety or to maintain the desired effect. Additional factors which may be taken into account include the severity of the disease state; age, weight and gender of the subject; diet, time and frequency of administration, drug combination(s), reaction sensitivities, and tolerance/response to therapy. Long acting pharmaceutical compositions might be administered every 3 to 4 days, every week, or once every two weeks depending on a half-life and clearance rate of the particular formulation.
[0201]In general, compositions contain from about 0.5% to about 50% of the compounds in total, depending on the desired doses and the type of composition to be used. The amount of the compounds, however, is best defined as the effective amount, that is, the amount of each compound which provides the desired dose to the subject in need of such treatment. The activity of the adjunctive combinations does not depend on the nature of the composition, so the composition is chosen and formulated solely for convenience and economy. Any of the combinations may be formulated in any desired form of composition.
[0202]Dosage amounts may vary from 0.1 to 100,000 micrograms of recombinant protein; transformed plant cell, or transformed transgenic plant per subject per day, for example, 1 ug, 10 ug, 100 ug, 500 ug, 1 mg, 10 mg, and even up to a total dose of about 1 g per subject per day, depending upon the route of administration. In one embodiment, the dosage is in the range of 1 ng to) 0.5 mg per kilogram bodyweight. In another embodiment, the dosage is in the range of 1 μg to 50 μg per kilogram bodyweight. In another embodiment, the dosage is in the range of 1 to 25 μg per kilogram bodyweight. In another embodiment, the dosage is in the range of 2 to 25 μg per kg body weight. In another embodiment, the dosage is in the range of 2 to 15 μg per kg bodyweight. For example, in one embodiment HN antigen is administered subcutaneously in a range of 2.5 to 5 μg, and IN/ocularly in a range of 0.5 to 12 μg; HA antigen is administered subcutaneously at a dose of 1 to 5 mg, IN/ocuraly in a range of 24 to 26 μg; VP2 antigen is administered subcutaneously in a range of 8 to 17 μg, and LT antigen is administered orally in a range of 50 to 100 ng, subcutaneously in a range of 2-10 μg and IN/ocularly in a range of 2 to 10 μg; Guidance as to particular dosages and methods of delivery is provided in the literature. See U.S. Pat. Nos. 4,657,760; 5,206,344; or 5,225,212, hereby incorporated by reference. Those skilled in the art will employ different formulations for nucleotides than for proteins or their inhibitors. Similarly, delivery of polynucleotide or polypeptides will be specific to particular cells, conditions, locations, etc.
Testing the Efficacy of a Vaccine of the Invention
[0203]The efficacy of a vaccine according to the invention is determined by demonstrating that the administration of the vaccine prevents or ameliorates the symptoms of the viral infection being treated or prevented or the symptoms induced by the pathogen of interest, by at least 5%, preferably 10-20% and more preferably, 25-100%.
[0204]The efficacy of a vaccine according to the invention is determined by measuring antibody production in response to vaccination with a plant derived protein of interest, detection of the production of antibody in response to vaccination with a plant derived protein of interest, wherein the antibody inhibits hemagluttination, and assessing the mortality of a subject that has been inoculated and then challenged with a vaccine comprising an immunoprotective antigen of the invention (all as described hereinbelow).
[0205]Having now generally described the invention, the same will be more readily understood through reference to the following Examples which are provided by way of illustration, and are not intended to be limiting of the present invention, unless specified.
EXAMPLE 1
Vectors
[0206]Gene Construction The coding sequence of the HN gene of NDV strain "Lasota" (GenBank accession AF077761) was analyzed for codon use and the presence of undesired sequence motifs that could mediate spurious mRNA processing and instability, or methylation of genomic DNA. See Adang M J, Brody M S, Cardineau G, Eagan N, Roush R T, Shewmaker C K, Jones A, Oakes J V, McBride K E (1993) The construction and expression of Bacillus thuringiensis cryIIIA gene in protoplasts and potato plants. Plant Mol Biol 21:1131-1145. A plant-optimized coding sequence was designed with hybrid codon preference reflecting tomato and potato codon usage (Ausubel F., et al., eds. (1994) Current Protocols in Molecular Biology, vol. 3, p. A.1C.3 Haq TA, Mason H S, Clements J D, Arntzen C J (1995) Oral immunization with a recombinant bacterial antigen produced in transgenic plants. Science 268:714-716). The designed sequence is shown in FIG. 1. The synthetic HN gene was assembled by a commercial supplier (Retrogen) and was received in two separate plasmids containing either the 5' (p4187-4203-1) or 3' (p42111-4235-ic-1) half of the gene cloned into pCR-Blunt.
[0207]Plasmid construction: Binary vectors for Agrobacterium-mediated plant transformations were constructed based on vector pBBV-PHAS-iaaH shown in FIG. 2, which uses the plant selection marker phosphinothricin acetyl transferase (PAT), described in U.S. Pat. Nos. 5,879,903; 5,637,489; 5,276,268; and 5,273,894 herein incorporated by reference, driven by the constitutive cassaya vein mosaic virus promoter (CsVMV) described in WO 97/48819. The iaaH gene and the phaseolin promoter sequence were deleted by digestion of pBBV-PHAS-iaaH with PacI and religated to form pCVMV-PAT; then the single HindIII site was deleted by filling it with Klenow enzyme and religating to form pCP!H. The CsVMV promoter was end-tailored by PCR using primers CVM-Asc (5'-ATGGCGCGCCAGAAGGTAATTATCCAAG SEQ ID NO:5) and CVM-Xho (5'-ATCTCGAGCCATGGTTTGGATCCA SEQ ID NO:6) on template pCP!H, and the product was cloned in EcoRV-digested, T-tailed pBluescriptKS to make pKS-CVM7. A map of pCP!H is shown in FIG. 3. The HN expression cassette pKS-CHN was constructed by ligating the vector pKS-CVM7/NcoI-EcoRI with 3 insert fragments: the HN 5' half on NcoI/PstI, the HN 3' half on PstI/KpnI, and the soybean vspB 3' element on KpnI-EcoRI (Haq 1995). The binary T-DNA vector pCHN was then assembled by ligation of the vector pCP!H/AscI-EcoRI and the AscI-EcoRI fragment of pKS-CHN. A map of pCHN is shown in FIG. 4.
[0208]The granule bound starch synthase (GBSS) promoter, described in U.S. Pat. No. 5,824,798 herein incorporated by reference, was used to make other vectors. These constructs were made using a promoter fragment amplified from genomic DNA of Solanum tuberosum L. cv. "Desiree" using primers designed from the sequence in GenBank accession X83220 for the Chinese potato cultivar "Dongnong". A mutagenic primer "GSS-Nco" (5'-tgccatggtgatgtgtggtctacaa SEQ ID NO:7) was used to create a Nco I site overlapping the translation initiation codon, along with forward primer "GSS-1.8F" (5'-gatctgacaagtcaagaaaattg SEQ ID NO:8) complimentary to the 5' region at -1800 bp; the 1825 bp PCR product was cloned in T-tailed pBluescriptKS to make pKS-GBN, and sequenced. A mutagenic primer "GSS-Xho" (5'-agctcGAGCTGTGTGAGTGAGTG SEQ ID NO:9) was used to create a XhoI site just 3' of the transcription start site along with primer "GSS-1.8F"; the 1550 bp PCR product was cloned in T-tailed pBluescriptKS to make pKS-GBX, and sequenced.
[0209]A GBSS promoter expression cassette containing the TEV 5'UTR (untranslated region), described in U.S. Pat. No. 5,891,665 herein incorporated by reference, was assembled by ligation of vector pTH210 digested with HindIII/XhoI with the HindIII/XhoI fragment of pKS-GBX, which effected a substitution of the CaMV 35S promoter with the 811 bp GBSS promoter, to make pTH252A. See Haq TA, Mason H S, Clements J D, Arntzen C J (1995) Oral immunization with a recombinant bacterial antigen produced in transgenic plants. Science 268:714-716. The HN gene was inserted into pTH252A/NcoI-KpnI by ligation with the HN 5' half on NcoI/PstI and the HN 3' half on PstI/KpnI to make pHN252A. The binary T-DNA vector pgHN was made by ligation of the vector pGLTB (shown in FIG. 11) digested with NsiI and EcoRI with the fragments pHN252A/NsiI-KpnI and pTH210/KpnI-EcoRI. A map of pgHN is shown in FIG. 5.
[0210]A GBSS promoter expression cassette containing the GBSS 5'UTR, described in U.S. Pat. No. 5,824,798 herein incorporated by reference, with its intron, was assembled by ligation of vector pTH210 (Haq 1995) digested with HindIII/NcoI with the HindIII/NcoI fragment of pKS-GBN, which effected a substitution of the (cauliflower mosaic virus) CaMV 35S promoter/TEV 5'UTR with the 1084 bp GBSS promoter/5'-UTR, to make pTH251 A. The binary T-DNA vectorpgHN151 was madeby ligation of the vectorpCLT105 (shown in FIG. 12) with fragments pTH251A/HindIII-NcoI and pHN252A/NcoI-KpnI. A map of pgHN151 is shown in FIG. 6.
[0211]A GBSS promoter expression cassette containing the GBSS 5'UTR with its intron and the bean phaseolin 3' element (described in U.S. Pat. Nos. 5,270,200; 6,184,437; 6,320,101, herein incorporated by reference) was constructed. First, pCP!H was digested at the unique KpnI site, blunted with T4 DNA polymerase, and religated to make pCP!HK, which has the KpnI site removed. pCP!HK was digested with NsiI, followed by blunting with T4 DNA polymerase, and then digestion with PacI. The resulting vector was ligated with a 2848 bp fragment from pgHN151 digested with SacI, followed by blunting with T4 DNA polymerase, and then digestion with PacI, to make pgHN153. A map of pgHN153 is shown in FIG. 7.
[0212]A chimeric constitutive promoter (4OCSΔMAS U.S. Pat. Nos. 5,001,060; 5,573,932 and 5,290,924 herein incorporated by reference) was used to construct another expression vector for HN. Plasmid, pAGM149, was digested with EcoRV and partial digestion with BamHI. This fragment was ligated with pCHN digested with PmeI/PstI and the 5' half of the synthetic HN gene obtained by digestion of pKS-CHN with BamHI/PstI. The resulting pMHN is shown in FIG. 8.
[0213]A plasmid containing the HA gene of AIV A/turkey/Wisconsin/68 (H5N9) was obtained from David Suarez (SEPRL, Athens, Ga.). It was end-tailored by PCR to add restriction sites NcoI at the 5' and KpnI at the 3' end, and inserted into the vector pIBT210.1 (Haq et al., 1995), containing the 35S promoter, TEV 5'-UTR, and vspB 3' end. The expression cassette was transferred to the binary vector pGPTV-Kan (Becker et al., Plant Mol Biol 1992; 20: 1195-7) by digestion with HindIII and EcoRI (partial), to make pIBT-HAO. The HA gene/vspB3' end fragment from pIBT-HAO was obtained by digestion with NcoI and EcoRI (partial), and inserted into pKS-CVM7 to make pKS-CHA. The cassette containing the CsVMV promoter, HA gene, and vspB3' end was obtained from pKS-CHA by digestion with AscI and EcoRI (partial), and ligated with pCP!H to make pCHA, shown in FIG. 9.
[0214]A dicot expression vector containing the plant-optimized nucleotide sequence of NDV-HN was constructed. The completed construct contained the gene cassette; Arabidopsis thaliana (At) Ubiquitin 3 (Ubi3) promoter v2/Newcastle Disease Virus Hemagluttin Neuraminidase (NDV-HN)/vspb 3'UTR:: Cassaya Vein Mosaic Virus (CsVMV) promoter/PAT selectable marker/Arabidopsis thaliana (At) ORF 25 3'UTR in a binary expression vector.
[0215]The expression cassette was assembled by completing a 3-way ligation (FIG. 48). The binary vector pCGUS was modified by removing the CsVMV promoter and GUS gene. A restriction enzyme digest with the enzymes HinDIII and KpnI (New England Biolabs) liberated a DNA fragment of 8310 bp. The NDV-HN gene was isolated from the plasmid pCHN as an NcoI/KpnI (New England Biolabs) restriction enzyme digestion DNA fragment of 1731 bp. Finally, the AtUbi3 promoter v2 was isolated from pDAB7121 as an NcoI/HindIII (New England Biolabs) restriction enzyme digestion. The resulting reaction produced a DNA fragment of 1732 bp. The DNA of all three enzyme digestions was excised from agarose gel via the "QiaexII Gel Extraction Kit" (Qiagen). A 3-way ligation was completed using equimolar concentrations of all three DNA fragments. The ligation was catalyzed by the "T4 DNA Ligase" (New England Biolabs). The resulting ligation product was transformed into "One Shot Top10 Chemically Competent E. coli." (Invitrogen). Two colonies were isolated from this transformation. Initial screening via restriction enzyme digestion indicated that both clones produced the expected DNA banding pattern. The restriction enzyme reactions that were completed used the following enzymes; EcoRV, FspI, HinDIII, NcoI, SacI, ScaI (New England Biolabs). Further confirmation of the correct construct involved a sequencing reaction over the AtUbi 3' promoter v2/NDV-HN border. A sequencing reaction with the primer pUHN2 (tgg ttg gag cct agg gta ct) was completed using the "Beckman CEQ Quick Start Kit" (Beckman Coulter). The results of this sequencing reaction indicated that the AtUbi3' promoter v2 DNA fragment did ligate with the NDV-HN DNA fragment at the intended NcoI restriction site. Sequencing across the NDV-HN/pCGUS border and pCGUS/AtUbi3' promoter v2 border required additional steps. A PCR reaction of both borders was completed. The NDV-HN/pCGUS border and pCGUS/AtUbi3' promoter v2 border were PCR amplified using the "FailSafe PCR Kit" (Epicenter). Two reactions for the NDV-HN/pCGUS border were completed using the FailSafe buffer's B and C with the PCR primers KpnI 5' (act aat act taa tga taa ca) and KpnI 3' (ata cac tac ctc cac atg tt). The PCR reactions for the pCGUS/AtUbi3' promoter v2 border were completed using FailSafe buffer's B and C with the PCR primers HinDIII 5' (tgccggttttcaggtaac ata) and HinDIII 3' (agt tag gcc cga ata gtt tga a). All of the PCR reactions produced DNA fragments of the expected length (˜600 bp). The PCR amplifications of the border regions were cloned into the "TOPO TA cloning kit with pCR2.1-TOPO" (Invitrogen). Clones containing the amplified border region were identified via an EcoRI restriction enzyme digestion (New England Biolabs). To confirm that the intended ligation at these border junctions did occur, a sequencing reaction was completed using the "Beckman CEQ Quick Start Kit" (Beckman Coulter) with the M13 reverse sequencing primer (aac agc tat gac cat g). The results of these sequencing reactions indicated that the correct ligation reaction did occur at the pCGUS/NDV-HN and the pCGUS/At Ubi 3 promoter v2 borders.
EXAMPLE 2
Preparation of Transgenic Nicotiana tabacum
[0216]Three to 4 days prior to transformation, a 1 week old NT-1 culture was sub-cultured to fresh medium by adding 2 ml of the NT-1 culture into 40 ml NT-1 media. The sub-culture was maintained in the dark at 25±1° C. on a shaker at 100 rpm.
NT-1 Medium
TABLE-US-00002 [0217] Reagent Per liter MS salts 4.3 g MES stock (20X) 50 ml B1 inositol stock (100X) 10 ml Miller's I stock 3 ml 2,4-D (1 mg/ml) 2.21 ml Sucrose 30 g pH to 5.7 ± 0.03
[0218]B1 Inositol Stock (100×)(1 liter) [0219]Thiamine HCl (Vit B1)-0.1 g [0220]MES (20×) (1 liter) [0221]MES (2-N-morpholinoethanesulfonic acid)-10 g [0222]Myoinositol-10 g [0223]Miller's I (1 liter) [0224]KH2PO4-60 g
[0225]Agrobacterium tumefaciens containing the expression vector of interest was streaked from a glycerol stock onto a plate of LB medium containing 50 mg/l spectinomycin. The bacterial culture was incubated in the dark at 30° C. for 24 to 48 hours. One well-formed colony was selected, and transferred to 3 ml of YM medium containing 50 mg/L spectinomycin. The liquid culture was incubated in the dark at 30° C. in an incubator shaker at 250 rpm until the OD600 was 0.5-0.6. This took approximately 24 hrs.
LB Medium
TABLE-US-00003 [0226] Reagent Per liter Bacto-tryptone 10 g Yeast extract 5 g NaCl 10 g Difco Bacto Agar 15 g
YM Medium
TABLE-US-00004 [0227] Reagent Per liter Yeast extract 400 mg Mannitol 10 g NaCl 100 mg MgSO4•7H20 200 mg KH2PO4 500 mg
[0228](Alternatively, YM in powder form can be purchased (Gibco BRL; catalog #10090-011). To make liquid culture medium, add 11.1 g to 1 liter water.)
[0229]On the day of transformation, 1 μl of 20 mM acetosyringone was added per ml of NT-1 culture. The acetosyringone stock was made in ethanol the day of the transformation. The NT-1 cells were wounded to increase the transformation efficiency. For wounding, the suspension culture was drawn up and down repeatedly (20 times) through a 10 ml wide-bore sterile pipette. Four milliliters of the suspension was transferred into each of 10, 60×15 mm Petri plates. One plate was set aside to be used as a non-transformed control. Approximately, 50 to 100 μl of Agrobacterium suspension was added to each of the remaining 9 plates. The plates were wrapped with parafilm then incubated in the dark on a shaker at 100 rpm at 25±1° C. for 3 days.
[0230]Cells were transferred to a sterile, 50 ml conical centrifuge tube, and brought up to a final volume of 45 ml with NTC medium (NT-1 medium containing 500 mg/L carbenicillin, added after autoclaving). They were mixed, then centrifuged at 1000 rpm for 10 min in a centrifuge equipped with a swinging bucket rotor. The supernatant was removed, and the resultant pellet was resuspended in 45 ml of NTC. The wash was repeated. The suspension was centrifuged, the supernatant was discarded, and the pellet was resuspended in 40 ml NTC. Aliquots of 5 ml were plated onto each Petri plate (150×15 mm) containing NTCB10 medium (NTC medium solidified with 8 g/l Agar/Agar; supplemented with 10 mg/l bialaphos, added after autoclaving). Plates were wrapped with parafilm then maintained in the dark at 25°±1° C. Before transferring to the culture room, plates were left open in the laminar flow hood to allow excess liquid to evaporate. After 6 to 8 weeks, putative transformants appeared. They were selected and transferred to fresh NTCB5 (NTC medium solidified with 8 g/l Agar/Agar; supplemented with 5 mg/l bialaphos, added after autoclaving). The plates were wrapped with parafilm and cultured in the dark at 25°±1° C.
[0231]Putative transformants appeared as small clusters of callus on a background of dead, non-transformed cells. These calli were transferred to NTCB5 medium and allowed to grow for several weeks. Portions of each putative transformant were selected for ELISA analysis. After at least 2 series of analysis by ELISA, lines with the highest antigen levels were selected. The amount of callus material for each of the elite lines was then multiplied in plate cultures and occasionally in liquid cultures. The resulting transformed NT-1 cell lines expressed and accumulated the HN protein from Newcastle Disease Virus (Lasota strain), or transformed cell line CHA expressed the HA protein from Avian Influenza Virus. These lines contain an undetermined number of copies of the T-DNA region of the plasmids stably integrated into the nuclear chromosomal DNA. The transgenic CHN NT-1 cells accumulate HN at levels up to 1% of total soluble protein as determined by HN-specific ELISA.
[0232]Transgenic NT1 cell and potato lines selected for Bialaphos® resistance were propagated and evaluated for HN expression by ELISA. High-expressing NT1 cell lines were established in liquid suspension culture. Potato lines containing a constitutive promoter construct (pCHN, pMHN) were screened for expression in leaf tissue, and selected lines were transferred to soil and cultured in a greenhouse to obtain tubers for evaluation. Potato lines containing a tuber-specific GBSS promoter construct (PGHN, pGHN151, pGHN153) were screened for expression using microtubers developed in vitro.
[0233]HN expression in pCHN-transformed NT1 lines using the CsVMV promoter. Expression of HN in NT1 cell lines assayed from callus growing on solid media is shown in FIG. 19. The highest expressing lines were CHN-5 (8.5 ng/μg TSP) and CHN-18 (6.2 ng/μg TSP). Lines CHN-1 and CHN-5 were established in liquid suspension culture. The expression of HN per unit cell mass in these cultures is shown in FIG. 20. Line CHN-5 showed expression of HN at 6.7 μg per g cell mass. The same cell lines shown in FIG. 20 were evaluated multiple times, and some new lines assayed at the last time point, stability of expression of HN in the NT1 lines (FIG. 21). Western blotting of extracts from lines CHN-5 and CHN-7 showed a single reactive band co-migrating with the reference standard when probed with monoclonal antibody, and showed an additional smaller band when probed with a polyclonal antiserum (FIG. 22).
[0234]Effects of freeze-drying of fresh cells and of storage of extracts at 4° C. In order to examine the effect of drying on antigen stability, freeze-dried NT1 cells were extracted and assayed by ELISA. Extracts of freeze-dried cells showed no loss or apparent increase in HN content per fresh cell mass (FIG. 23). Furthermore, extracts of fresh cells stored at 4° C. for one week showed an increase in HN content assayed by ELISA (FIG. 23). We have observed a similar effect with another membrane-bound viral protein, the hepatitis B surface antigen. It may result from oxidation of cysteine residues to form correct disulfide bonds, which results in display of the appropriate antigenic epitopes.
[0235]Particle behavior of plant-expressed HN antigen. In order to evaluate assembly of NT1 cell-expressed antigen to form particulate structures, sucrose gradient sedimentation was performed on crude cell extracts. The profiles shown in FIG. 24 indicate that the NT1 cell-derived HN showed 2 peaks of ELISA reactive material. One peak co-sedimented with inactivated virus particles, while the other peak sedimented more slowly but still showed particulate character. These data provide evidence that the HN protein is correctly inserted into the ER membrane.
[0236]HN expression in pMHN-transformed NT1 lines using the (4-ocs)ΔMAS promoter. HN expression in several NT1 cell lines transformed with pMHN, compared to pCHN-transformed NT1 cell lines, is shown in FIG. 25. Expression in pMHN-transformed lines was at least as high as in pCHN-transformed lines, with the highest accumulation of HN observed at approximately 30 μg per gram cell mass. Maximal HN expression in pCHN-transformed NT1 cells was less than 20 μg per gram cell mass. Bialaphos® resistant NT1 cell lines were also generated and assayed for HA expression by ELISA as described previously. In the first set of assays, only one pCHA-transformed line accumulated HA to a similar extent as the pGPTV-HAO line #12 that was previously generated (FIG. 13). In this experiment, the expression range was up to 2.5 ng/μg TSP. In repeated experiments with these and new pCHA-transformed lines, accumulation of HA ranged up to 18 ng/μg TSP in line CHA-13 (FIG. 14). Selected lines from this group were analyzed by Western blot. In all pCHA-lines tested, a reactive band at the expected size of ˜68 kDa was observed (FIG. 15). These data show that the HA protein was correctly processed to remove the signal peptide, and accumulated in a stable form. Previous studies on pGPTV-HAO-transformed NT1 cells by non-denaturing Western blot (unpublished studies), showed that the HA assembled oligomeric structures, probably the native trimer that occurs on the surface of the AIV virion.
EXAMPLE 3
Cryopreservation
[0237]Cell Culture: NT-1 tobacco suspension cultures (non-transgenic and transgenic lines) were maintained in 250 ml Erlenmeyer flasks. Initially, the cells were cultured in a modified liquid Linsmaier and Skoog medium (LS) (1965). The medium, designated LSg, contained LS salts and vitamins, 30 g 1-1 glucose, and 0.05 mg 1-1 2,4-dichlorophenoxyacetic acid (2,4-D). The pH of the medium was adjusted to 5.8. The cultures were transferred to fresh medium weekly by transferring 6 ml of 7-day-old cultures into 50 ml of LSg medium.
[0238]Based on poor growth of the cells in LSg medium, two additional media were investigated which were designated KCMS and NT-1. KCMS contained Murashige and Skoog (MS) (1962) salts, 1.3 mg 1-1 thiamine, 200 mg 1-1 KH2PO4, 30 g 1-1 sucrose, 0.2 mg 1-1 and 0.1 mg 1-1 kinetin. NT-1 medium contained MS salts, 180 mg 1-1 KH2PO4, 0.5 mg 1-1 2-N-morpholinoethanesulfonic acid, 1 mg 1-1 thiamine, 100 mg 1-1 myoinositol, 30 g 1-1 sucrose, and 2.21 mg 1-1 2,4-D. The pH of both media was adjusted to 5.7. For transfers to fresh medium, 2 ml of a 7-day-old culture were transferred into 48 ml of either KCMS or NT-1. All suspension cultures were maintained in the dark at 25° C. on an orbital shaker at 100 rpm
[0239]Preculture: Three days after subculture (late exponential growth period), the cells were precultured in their respective medium (LSg, KCMS, or NT-1) by replacing one third of the medium with 1M mannitol (for final concentration of 0.3M), for 24 to 72 h.
[0240]Heat Shock Treatment: After preculture, cultures were placed on an orbital shaker at 100 rpm at 37° C. for 2 h. They were transferred back to the shaker at 25° C. for 4 h before vitrification.
[0241]Vitrification: The vitrification solution designated PVS2/100% contained 30% glycerol, 15% ethylene glycol, and 15% DMSO in a 0.4 M sucrose solution. A PVS2/20% solution was made by diluting PVS2/100%. Both solutions were adjusted to a pH of 5.8, autoclaved, then stored at 4° C.
[0242]To start the vitrification process, 4 ml of ice cold PVS2/20% was added to a 1 ml settled cell volume of cells. The preparation was incubated on ice for 5 min, then 1 ml of cold PVS2/100% was added at 1-minute intervals until a total volume of 9 ml was achieved. The preparations were centrifuged for 1 min at 7500-8000 rpm. The supernatant was discarded, then 0.5 ml of PVS2/100% was added at 2,1-min intervals while the cells remained on ice. Then 1 ml of PVS2/100% was added 3 times at 1-min intervals. Half a milliliter of this mixture was then transferred into each of 6 cryogenic straws (Continental Plastic Corporation, Delavan, Wis.). The straws were heat sealed at each end with hot forceps, and immersed immediately into liquid nitrogen. The remaining 2 ml were not frozen and served as controls.
[0243]Recovery: After 1 h in liquid nitrogen, the vitrified cells were thawed in a 40° C. water bath for 3-5 sec, then immediately diluted with 7 ml of cold 1.2 M sucrose. The cells were kept on wet ice for 20 min, then centrifuged for 3 min at 7500-8000 rpm. Cells were transferred to 2 layers of filter paper (42.5 mm Whatman) on LSg or NT-1 solidified medium containing 0.75% agarose (Invitrogen Life Technologies, Carlsbad, Calif.) in 60×15 mm tissue culture plates. The cultures were maintained at 25° C. in the dark. Two days after plating, the cells were transferred to fresh NT-1 solidified medium. Transgenic lines were initially plated on medium without a selection agent until cell growth was well established and covered a small plate. They were then transferred to medium containing the appropriate selection.
[0244]Thereafter, the cells were transferred to fresh medium approximately every 2 wk. When cell growth nearly covered the surface of the medium in the plate, the cells were removed from the filter paper and transferred to larger plates (100×15 mm). When the callus again covered the plate, the cells were transferred to NT-1 medium solidified with 8 g 1-1 Agar (Sigma, St. Louis, Mo.) for maintenance.
EXAMPLE 4
Antigen Preparation
[0245]Whole wet NT-1 cells expressing either HN, HA or null control were harvested directly from cell culture and filtered to remove excess media by placing a Spectramesh 30 filter in a Buchner funnel and pouring cells and media through the filter using a slight vacuum. 0.5 grams of cells were placed in 2 mls of buffer (Dulbeccos Phosphate Buffered Saline and 1 mM EDTA), and then sonicated for 15 to 20 seconds on ice. Sonication was performed using a Branson 450 sonifier with a replaceable microtip at output control of 8, duty cycle 60 for varying amounts of time. Sonicates were then placed on ice until use.
EXAMPLE 5
Antigen Extraction
[0246]To examine whether non-detergent treatments could release ELISA signal from transformed NT-1 cells and allow retention of biological activity, a series of treatments were set up that involved comparison of treatments without detergent and various levels of sonication. The results were striking in that periods of sonication greater than 20 seconds in extraction buffer completely destroyed hemagglutination activity of HN from a pCHN bearing NT-1 cell line, but not ELISA signal. In contrast, sonication for only 20 seconds in DPBS not only released antigen detectable by ELISA signal, but the soluble protein extracts demonstrated excellent hemagglutination activity (see Table 1).
TABLE-US-00005 TABLE 1 Comparison of extraction methods on hemagglutination activity of plant-derived HN Ext. buffer DPBS DPBS Ext. buffer DPBS F/T F/T Sonic. Sonic. Sonic. Sonic. Sonic. Sample 1.5 min 1.5 min 15 sec 15 sec 15 sec pCHN-18-NT-1 ≦2 256 4096 1024 1024 pCHA-47-NT-1 ≦2 -- 64 16 16 NT-1 ≦2 ≦2 ≦2 ≦2 ≦2 Native NDV1 256 512 128 nd nd 1Native NDV was sonicated for 2 minutes. Ext. buffer - 50 mM sodium ascorbate, 1 mM EDTA, 1 mM PMSF, and 0.1% Triton X-100 pH 7.2; DPBS--Dulbeccos phosphate buffered saline; sonic.--sonication; F/T--freeze-thaw; nd--not done for this experiment.
[0247]Plant-derived HN extracted without detergent was used as the antigen in hemagglutination inhibition assays to determine if polyclonal antibody produced to native virus could recognize and inhibit agglutination of RBC's by the plant-derived HN. The results indicate that native antibody will recognize the hemagglutination epitope of the plant-derived HN in a similar manner as native virus' (Table 2). The data from Table 2 also demonstrates that control NT-1 cells or NT-1 cells expressing a non-hemagglutinating protein do not agglutinate red blood cells nor are affected by NDV specific serum. In this experiment, extracts of plant-derived protein were diluted to 4 HA units, and then treated with NDV specific polyclonal antisera. Four HA units is the standard amount of virus used for titration of serum.
TABLE-US-00006 TABLE 2 Comparison of hemagglutination inhibition (HAI) activity of plant-derived HN and native virus Hemagglutination HN Inhibition Titer Concentration Hemagglutination (chicken anti-NDV Sample ELISA Titer polyclonal antibody) NDV allantoic 20 ug/ml 4* 4096 fluid (native) NT control cell None ≦2 ≦8 pCHN-7-NT-1 1.5 ug/g fresh >64 512 weight CHN-18-NT-1 12 ug/g fresh ≧4096 1024 weight CLT-101-14- None ≦2 ≦8 NT-1 *Stock virus is diluted such that 4HA units, a 1:4 dilution of the stock will generate a positive HA but a dilution of 1:8 will not hemagglutinate. This is the concentration of virus used to titer antibody, the endpoint dilution of antibody that will interfere with 4 HA units of virus is considered to be the HAI titer of the antibody preparation.
[0248]The above data demonstrates that using an extraction method that does not utilize detergent and reduces the amount of cell disruption produces an extracellular fraction that retains hemagglutination activity for transformed NT-1 cell lines expressing HN or HA. To determine if HN protein from non-detergent extracted NT-1 cells had additional biological activity that may be relevant to vaccine efficacy, the HN extracts were examined for ability to bind to chicken cell receptors. immuno-fluorescence staining indicated that CEF cells treated with native virus or pCHN-18 extracts were indistinguishable. Thus, plant-derived HN retains virus-like ability to bind to receptors on target cell surfaces.
[0249]The combined data from Tables 1 and 2, together with the hemagglutination and immunofluorescence assays discussed above, suggest that the HN protein derived transgenic NT-1 cells will retain both immunological and biological features if processed and formulated correctly. Most significant of the data provided above is that antisera to native virus will recognize plant-derived HN in HAI tests. Chickens that contain at least 4 fold higher titer of HAI activity above background are almost always certain of protection against challenge from virulent virus. To test whether the plant derived protein extracted in non-detergent as described above would generate antibody in target animals species both HA and HN protein were prepared and inoculated into chickens and rabbits.
EXAMPLE 6
Quantitative ELISA
[0250]HN
[0251]Quantitative ELISA for HN is performed by coating plates on the day prior to running the assay. 50 μl per well of Capture Antibody (Rabbit anti-HN in 50% glycerol, diluted (1:500) in 0.01M Borate Buffer) is added to each well of each flat bottom 96-well microtiter plate. The plate is covered and incubated at 2° C.-7° C. overnight, (12-18 hours). The coated ELISA plate(s) are allowed to equilibrate to room temperature (approximately 20-30 minutes) and then washed three times with 200-300 μl per well per wash with PBS-T. The entire plate is blocked to prevent non-specific reactions by adding 200 μl per well of 3% Skim Milk Blocking Solution. The plate(s) is(are) then incubated for 2 hours (+10 minutes) at 37° C.±2° C. (covered with a plate cover or equivalent). HN Reference antigen (Ag) in 1% Skim Milk Blocker is added to a concentration of 250 ng HN/ml; experimental antigens are diluted in 1% Blocker. The HN ELISA plate(s) are washed one time with PBS-T and 100 μl per well of diluted HN Reference Antigen and HN Test Samples are added to Row B. 50 μl per well of 1% Blocker is added to all remaining wells. The samples are serially diluted down the plate by transferring 50 μl per well from row B to row G, mixing 4-5 times with the pipette before each transfer. The plate(s) are covered and incubated 1 hour (+10 minutes) at 37° C.±2° C.; and the ELISA plate(s) are washed three times with PBS-T. Fifty μl of NDV HN 4A Ascites Fluid in 50% glycerol (1:2000) in 3% Blocker is added to each well and the plates are covered and incubated 1 hour (+10 minutes) at 37° C.±2° C. The ELISA plate(s) are washed three times with PBS-T and 50 μl of rabbit anti-Mouse IgG in 50% glycerol (1:3000) in 3% Blocker is added to each well. The plates are covered and incubated 1 hour (+10 minutes) at 37° C.±2° C. ELISA plate(s) are washed three times with PBS-T and 50 μl of ABTS Peroxidase Substrate Solution (equilibrated at RT (room temperature) for at least 30 minutes) is added to each well. The plates are covered and incubated at RT in the dark for 15-20 minutes. The Optical Density (OD) of the wells are read at a wavelength of 405 nm (with a 492 nm Reference Filter). The initial dilution of the HN Reference Antigen should be within 0.7-1.0 OD, this serves as the positive control for the ELISA.
[0252]HA
[0253]For quantitative ELISA of HA, the plates are coated on the day prior to running the assay. Fifty μl per well of Capture Antibody (goat anti-Hav5 in 50% glycerol, diluted (1:1000) in 0.01M Borate Buffer) is added to each well of flat bottom 96-well microtiter plate(s)). The plate(s) are covered and incubate at 2° C.-7° C. overnight, (12-18 hours). The coated ELISA plate(s) is(are) allowed to equilibrate to room temperature (approximately 20-30 minutes) and is(are) then washed three times with 200-300 μl per well per wash with PBS-T. The entire plate is blocked to prevent non-specific reactions by adding 200 μl per well of 3% Skim Milk Blocking Solution. The plate(s) is(are) then incubated for 2 hours (+10 minutes) at 37° C.±2° C. (covered with a plate cover or equivalent). AIV-HA (allanotoic fluid) reference Antigen is added in 1% Skim Milk Blocker to a concentration of 1000 ng HA/ml and experimental antigens are diluted in 1% Blocker. The HA ELISA plate(s) are washed one time with PBS-T and 100 μl per well of diluted HA reference antigen and HA Test Samples are added to Row B. 50 μl per well of 1% Blocker is added to all remaining wells. The samples are serially diluted down the plate by transferring 50 μl per well from row B to row G, mixing 4-5 times with the pipette before each transfer. The plate(s) are covered and incubated 1 hour (+10 minutes) at 37° C.±2° C. The ELISA plate(s) are washed three times with PBS-T. Fifty μl of chicken anti-AIV polyclonal antisera in 50% glycerol (1:2000) in 3% Blocker is added to each well and the plates are covered and incubated 1 hour (+10 minutes) at 37° C.±2° C. The ELISA plate(s) are washed three times with PBS-T and then 50 μl of goat anti-chicken IgG in 50% glycerol (1:3000) in 3% Blocker is added to each well. The plates are covered and incubated 1 hour (+10 minutes) at 37° C.±2° C. The ELISA plate(s) are washed three times with PBS-T and 50 μl of ABTS Peroxidase Substrate Solution (equilibrated at RT for at least 30 minutes) is added to each well. The plate(s) are covered and incubated at RT in the dark for 15-20 minutes. The Optical Density (OD) of the wells read at a wavelength of 405 nm (with a 492 nm Reference Filter). The initial dilution of the HA Reference Antigen should be within 0.7-1.0 OD, this serves as the positive control for the ELISA.
EXAMPLE 7
Serum ELISA
[0254]NDV-HN
[0255]Plates are coated with rabbit a-NDV pooled antiserum (Mixed 1:2 with 50% glycerol in water) diluted (1:2000) in 0.01 M borate buffer (100 μl/well). Plates are incubated overnight at 2-7° C., covered and then equilibrated for approximately 20-30 minutes at room temperature. Plates are washed 3× with PBS-T (1×PBS+0.05% Tween-20) at 300 μl/well with the Titertek M96 plate washer or equivalent. Plates are blocked with 5% skim milk in PBS-T (Blocking Buffer) (200 μl/well) and incubated for 2 hours at 37° C. Plates are washed 1× with PBS-T at 300 μl/well with the Titertek M96 plate washer or equivalent. NDV allantoic fluid is diluted 1:200 in Blocking Buffer. 100 μl/well of the diluted antigen is added to the plate, and plates are incubated for 1 hour at 37° C. Plates are washed 3× with PBS-T at 300 μl/well with the Titertek M96 plate washer or equivalent. Test chicken serum samples are diluted (1:50). Negative control serum is diluted (1:50) (Neg. Control 27NOV00). Positive control serum is diluted (1:10,000 or 1:20,000) (SPAFAS Chicken α-NDV serum). All serum samples are diluted in Blocking Buffer. 100 μl/well of Negative Control Serum is added to Column 1 Rows B-G; 200 μI/well of Positive Control Serum is added to Columns 2-3 Row A; 200 μl/well of Test Serum Samples is added to Rows A appropriate columns. This allows 4 samples per plate with 8 dilutions per sample. 100 μl/well of Blocking Buffer is added to all remaining wells The Positive Control Serum and the Test Serum Samples are serially two-fold diluted down the plate. The samples are diluted down the plate from Row A to Row H, discarding the remaining 100 μl/well. Plates are incubated for 1 hour at 37° C. and then washed 3× with PBS-T at 300 μl/well with the Titertek M96 plate washer or equivalent. The Goat α-Chicken IgG (H&L)-HRP is diluted (1:3000) in Blocking Buffer. 100 μl/well of the diluted conjugate is added to each plate. Once the conjugate is added to the plates, the ABTS substrate is equilibrated at RT in the dark. Plates are incubated for 1 hour at 37° C. and then washed 3× with PBS-T at 300 μl/well using the Titertek M96 plate washer or equivalent. 100 μl/well of pre-warmed ABTS substrate is added to each plate, waiting 2-3 minutes between plates. Plates are read at dual wavelength 405/490 nm on the Tecan Sunrise plate reader or equivalent when the first dilution of the positive control reaches an absorbance of between 0.7 and 1.0.
[0256]AIV-HA
[0257]Plates are coated with Rabbit α-HA pooled antiserum diluted (1:1000) in 0.01 M borate buffer and incubated overnight at 2-7° C., covered. Plates are equilibrated for approximately 20-30 minutes at room temperature and then washed 3× with PBS-T (PBS Stock+0.05% Tween-20) at 300 μl/well using the Titertek M96 plate washer or equivalent. The plates are blocked with 5% skim milk in PBS-T (Blocking Buffer) (200 μl/well) and incubated for 1 hour at 37° C. The plates are washed 1× with PBS-T at 300 μl/well using the Titertek M96 plate washer or equivalent. Inactivated TIW/68 AIV Allantoic Fluid is diluted (1:100) in Blocking Buffer and 100 μl/well of the diluted antigen is added to the plate. Plates are incubated for 1 hour at 37° C. The plates are washed 3× with PBS-T at 300 μl/well using the Titertek M96 plate washer or equivalent. Test chicken serum samples are diluted (1:50). Negative control serum is diluted (1:50). Positive control serum is diluted (1:25600) (USDA/SEPRL Chicken α-AIV (TIW/68 antiserum) All serum is diluted in Blocking Buffer. 100 μl/well of Negative Control Serum is added to Column 1 Rows B-G; 200 μl/well of Positive Control Serum is added to Columns 2-3 Row A; 200 μl/well of Test Serum Samples is added to Row A in appropriate columns; 100 μl/well of Blocking Buffer is added to all remaining wells. The Positive Control Serum and the Test Serum Samples are serially diluted two-fold down the plate, discarding the remaining 100 μl/well. The plates are incubated for 1 hour at 37° C. The plates are washed 3× with PBS-T (300 μl/well) using the Titertek M96 plate washer or equivalent. Goat a-Chicken IgG (H&L)-HRP is diluted (1:3000) in Blocking Buffer and 100 μl/well of the diluted conjugate is added to each plate. Once the conjugate is added to the plates, the ABTS substrate is equilibrated at RT in the dark. The plates are incubated for 1 hour at 37° C. and then washed 3× with PBS-T at 300 μl/well using the Titertek M96 plate washer or equivalent. 100 μl/well of equilibrated ABTS substrate is added to each plate, leaving 2-3 minutes between plates. Plates are read at dual wavelength 405/490 nm on the Tecan Sunrise plate reader or equivalent when the first dilution of the positive control reaches an absorbance of between 0.7 and 1.0.
EXAMPLE 8
Antigenicity in Rabbits
[0258]To test whether the plant derived protein extracted in non-detergent as described above would generate antibody in target animals species both HA and HN protein were prepared and inoculated into rabbits. New Zealand White rabbits 3 months of age were inoculated with HA-AW or HN-NDV according to the dose schedule provided in Table 3. For the primary inoculation the antigen was mixed with Complete Freund's Adjuvant (CFA) and Incomplete Freund's Adjuvant was used for all booster inoculations. The antibody titers induced by both proteins are provided in Table 4. The results indicate that after two inoculations HAI antibody titers were induced by both proteins. However, the titers of the AIV-HA inoculated rabbits were higher than those induced by the NDV-HN protein. This may be significant since the AW/HA protein had lower overall biological activity (hemagglutination) per unit of AIV-HA protein than NDV-HN (Table 3 column 4). This may indicate that the AIV-HA protein derived from plants is more potent per unit of protein than the NDV-HN protein considering that the quantitation methods from both proteins are accurate. It also suggests that the AIV-HA protein formulated in this manner would be immunogenic in chickens.
TABLE-US-00007 TABLE 3 Dose levels for inoculation into rabbits ELISA Hemagglu- Results BCA TSP1 tination Hemagglutination (ug Results units per Sample Endpoint Titers protein/ml) (mg/ml) ug protein NT ≦2 0.00 1.44 0 Control CHN-7 2048 13.53 2.48 3027 CHN-18 1024 9.21 4.10 2223 CHA-13 32 3.80 6.15 168 CHA-47 16 2.88 5.25 111 1All NT-1 samples were provided from non freeze-dried material. BCA--bicinchoninic acid, primary component of Pierce Chemical BCA protein assay kit; TSP--total soluble protein.
TABLE-US-00008 TABLE 4 Serology Results from NT-1 derived AIV-HA and NDV-HN inoculated rabbits Treatment with Sample NDV HAI Titers NDV ELISA Titers NDV-HN Number Pre-Bleed 6 week 8 week 10 week Pre-Bleed 6 week 8 week 10 week Sup from NT Control Cells 2723 ≦8 ≦8 ≦8 ≦8 0 0 0 0 Sup from HN - 18 Cells 2724 ≦8 23 23 23 0 815 554 888 Sup from HN - 18 Cells 2725 ≦8 11 23 23 0 0 585 591 Sup from HN - 7 Cells 2726 ≦8 45 23 23 0 607 461 0 Sup from HN - 7 Cells 2727 ≦8 45 45 45 0 1396 2008 1270 Treatment with Sample AIV HAI Titers AIV ELISA Titers AIV-HA Number Pre-Bleed 6 week 8 week 10 week Pre-Bleed 6 week 8 week 10 week Sup from NT Control Cells 2723 ≦8 ≦8 ≦8 ≦8 <25 <25 <25 <25 Sup from HA - 13 Cells 2728 ≦8 362 362 362 <25 25600 25600 25600 Sup from HA - 13 Cells 2729 ≦8 181 181 11 <25 3200 3200 <25 Sup from HA - 47 Cells 2730 ≦8 181 362 724 <25 12800 25600 25600 Sup from HA - 47 Cells 2731 ≦8 ≦8 362 362 <25 50 12800 25600
[0259]To examine the efficacy of the plant derived antigens in chickens the HN protein was inoculated into chickens that were 2 days of age and 10 days of age. The dose concentrations used for these studies are provided in Table 5. All vaccine inoculum were formulated with the soluble fraction of NT-1 cells grown 15-20 days in shaker flasks at 25° C. Adjuvant used in both trials was MPL-TDM from Corixa, Inc. Intranasal groups were given MPL alone as the adjuvant.
EXAMPLE 9
Challenge in Poultry
[0260]Two-day old SPF chicks were inoculated by various routes using biologically active (hemagglutination positive) NDV-HN protein derived from NT-1 with the amount of HN protein per inoculation shown in Table 5. The serological and challenge results of this trial are provided in Table 6. All control groups responded as expected in that birds not receiving NDV-HN antigen in the inoculum had 100% mortality, whereas, control birds receiving 20 ug of native NDV by SQ had 100% survival. In the experimental treatment groups there was 75% protection in group #3 (SQ inoculation without adjuvant) and 80% protection in group #4 (SQ inoculation with adjuvant). The remaining treatment groups, which were inoculated by IN and oral routes, had 100% mortality. However, in group 6 two birds had a delay in mortality, indicating that these birds may have been sensitized to vaccination (see Table 9)
TABLE-US-00009 TABLE 5 Dose levels used per inoculation for poultry trial ug HN/Bird Group Day 0 Day 14 Day 21 NT Control (SQ) 0 0 0 NDV All. Fluid (SQ) 20 20 20 HN Tobacco (SQ) 150 230 180 HN Tobacco (IN) 6 14 14 HN Tobacco (OG) 114 282 136 HN Tobacco 114og + 700of* 282og/1400of* 136 + 2366* (OG + OF) Average 3590 5810 6025 hemagglutination units per ug HN *Dose based on wet weight expression per mass cells mixed with feed; IN--intranasal; SQ--subcutaneous; OG--oral gavage; OF--on feed mixtures.
TABLE-US-00010 TABLE 6 Serology and challenge results from poultry trial NDV HAI titers NDV ELISA titers Sample Day Day Pre Post Day Pre Post Surv. Treatment Number 14 21 Day 28 Chall. Chall. 28 Chall. Chall. Chall. 1. Control NT Cells 924 ≦8 ≦8 ≦8 ≦8 na 0 0 0 No SQ 1061 ≦8 ≦8 ≦8 ≦8 na 0 0 0 No 1073 ≦8 ≦8 ≦8 ≦8 na 0 0 0 No 1077 ≦8 ≦8 ≦8 ≦8 na 0 0 0 No 1081 ≦8 ≦8 ≦8 ≦8 na 0 0 0 No 2. NDV HN allantoic 1063 11 1448 2896 724 724 11956 9245 7294 Yes fluid SQ 1068 11 1448 1024 724 362 9216 7639 6122 Yes 1072 45 1448 1448 724 362 11592 7500 5937 Yes 1083 23 724 724 181 91 5697 4919 3011 Yes 1089 45 1024 1448 362 181 15181 7449 6085 Yes 3. NDV HN tobacco 797 23 45 45 ≦8 2896 0 0 19036 Yes SQ 1066 ≦8 16 45 ≦8 724 450 0 15087 Yes 1085 ≦8 ≦8 23 ≦8 na 0 0 na No 1095 ≦8 23 45 ≦8 724 436 0 10043 Yes 4. ND V HN tobacco 1067 11 45 45 ≦8 181 592 0 4912 Yes MPL/TDM adjuvant 1080 ≦8 181 181 45 181 1911 871 5048 Yes SQ 1093 11 45 45 ≦8 ≦8 0 0 0 Yes 1094 11 91 91 11 11 747 199 0 Yes 1098 ≦8 23 45 ≦8 na 0 0 na No 5. NDV HN tobacco 796 ≦8 ≦8 ≦8 ≦8 na 0 0 na No IN 925 ≦8 ≦8 ≦8 ≦8 na 0 0 na No 1065 ≦8 ≦8 ≦8 ≦8 na 0 0 na No 1084 ≦8 ≦8 ≦8 ≦8 na 0 0 na No 1092 ≦8 ≦8 ≦8 ≦8 na 0 0 na No 6. NDV HN tobacco + 921 ≦8 ≦8 ≦8 ≦8 na 0 0 na No MPL adjuvant 923 ≦8 11 ≦8 ≦8 na 0 0 na No IN 1069 ≦8 ≦8 ≦8 ≦8 na 0 0 na No 1074 ≦8 11 ≦8 ≦8 na 0 0 na No 1088 ≦8 11 8 ≦8 na 0 0 na No 7. NDV HN tobacco 723 ≦8 ≦8 ≦8 ≦8 na 0 0 na No Oral gavage 1062 ≦8 ≦8 ≦8 ≦8 na 0 0 na No 1075 ≦8 8 ≦8 ≦8 na 0 0 na No 1079 ≦8 8 ≦8 ≦8 na 0 0 na No 1086 ≦8 ≦8 ≦8 ≦8 na 0 0 na No 8. NDV HN tobacco + 1070 ≦8 ≦8 ≦8 ≦8 na 0 0 na No MPL/TDM adjuvant 1082 ≦8 ≦8 ≦8 ≦8 na 0 0 na No Oral gavage + On feed 1091 ≦8 ≦8 ≦8 ≦8 na 0 0 na No 1097 ≦8 ≦8 ≦8 ≦8 na 0 0 na No 1100 ≦8 ≦8 ≦8 ≦8 na 0 0 na No All birds receive 102 EID50 Texas GB strain of NDV. Birds were challenged 24 days post last vaccination. Bird numbers bolded had a delayed onset to mortality see Table 9.
[0261]In a subsequent trial, eighteen 10-day old SPF birds were inoculated according to the schedule and dose amounts described in Table 7. Results of this trial are shown in Table 8. One control group (#3), a non-vaccinated non-challenged treatment was used to show that the housing and facility had no adverse affects on general health of the chickens. Control groups in this trial also responded as expected. Since birds from both trials were challenged at the same facility, treatment group #2 served as a positive control for both poultry trials. In the remaining groups, all of which were inoculated SQ with HN derived from NT-1 cells, there was 100% survival in group #7, 80% survival in each of groups 5 and 6, and 60% survival in group 4 (see Table 8).
TABLE-US-00011 TABLE 7 Dose levels of antigen used per inoculation ug HN/Bird (Subcutaneous) Group Day 0 Day 14 Day 21 NT Control 0 0 0 NDV All. Fluid 20 20 20 HN Tobacco 20 20 20 (Low Dose) HN Tobacco 150 100 100 (High Dose) Average hemagglutination 3590 2625 2625 units per ug HN
[0262]Conclusions: Using a procedure that provides recovery of hemagglutination of HA and HN protein, preparations for these plant-derived proteins were inoculated into two separate animal species by subcutaneous (SQ) route to determine if antibody could be induced that will inhibit hemagglutination (HAI antibody). In one trial, HN protein was formulated to have high hemagglutination activity to total soluble protein ratios. These materials were then inoculated by SQ, intranasal (IN) and by oral routes. The results indicate that both HA-AIV and HN-NDV protein derived from NT-1 cells will induce hemagglutination inhibition (HAI) antibody in rabbits when formulated in this manner. In addition, HN-NDV derived from NT-1 inoculated by SQ route in chickens will induce (HAI) antibodies and protect against virulent NDV challenge.
[0263]The results from these trials indicate that the HN-NDV protein derived from NT-1 cells is immunogenic in birds when inoculated by SQ. In most cases birds protected from challenge had a detectable HAI titer post challenge, however, there were exceptions to this observation. Two birds (bird #1093 and #1047, respectively) did not have a detectable HAI or ELISA titer after challenge but survived challenge (Tables 6 and 8).
TABLE-US-00012 TABLE 8 Serology and challenge results NDV HAI titer NDV ELISA titers Sample Pre Post Pre Post Surv. Treatment Number Day 21 Day 28 Chall. Chall. Day 21 Day 28 Chall. Chall. Chall. 1. Control allantoic 1026 ≦8 ≦8 ≦8 na 0 0 0 na No fluid 1027 ≦8 ≦8 ≦8 na 0 0 0 na No 1028 ≦8 ≦8 ≦8 na 0 0 0 na No 1029 ≦8 ≦8 ≦8 na 0 0 0 na No 1030 ≦8 ≦8 ≦8 na 0 0 0 na No 2. NDV HN allantoic 1031 362 91 181 91 9177 6937 5533 3551 Yes fluid 1032 362 724 181 91 12393 16533 7909 6080 Yes 20 ug/dose 1033 724 362 181 181 8622 15291 6766 6362 Yes 1034 362 181 181 181 7875 10071 6487 5822 Yes 1035 724 724 181 181 9681 16133 7537 6539 Yes 3. Control tobacco 1036 ≦8 23 ≦8 ≦8 0 0 0 0 n/c 1037 ≦8 11 ≦8 ≦8 0 0 0 0 n/c 1038 ≦8 ≦8 ≦8 ≦8 0 0 0 0 n/c 1039 ≦8 ≦8 ≦8 ≦8 0 0 0 0 n/c 1040 ≦8 23 ≦8 ≦8 0 0 0 0 n/c 4. NDV HN tobacco 1041 11 <8 ≦8 1448 0 0 0 14042 Yes 20 ug/dose 1042 23 11 ≦8 2896 0 0 0 19263 Yes 1043 32 11 ≦8 na 0 0 0 na No 1044 ≦8 11 ≦8 na 0 0 0 na No 1045 23 11 ≦8 1024 0 0 0 11770 Yes 5. NDV HN tobacco 1046 23 23 ≦8 1448 0 674 0 11243 Yes 20 ug/dose + 1047 23 23 ≦8 ≦8 0 963 0 0 Yes MPL/TDM 1048 23 23 ≦8 na 396 757 0 na No emulsion adjuvant 1049 45 23 ≦8 362 0 804 0 6239 Yes 1050 11 11 ≦8 1448 0 398 0 15948 Yes 6. NDV HN tobacco 1051 91 45 23 181 1096 1137 565 4547 Yes 250 ug/dose 1052 45 45 ≦8 181 1166 998 0 7376 Yes 1053 23 23 ≦8 2896 0 0 0 16712 Yes 1054 23 23 ≦8 na 646 838 0 na No 1055 45 45 23 91 705 563 448 4902 Yes 7. NDV HN tobacco 1056 45 45 11 23 746 948 174 926 Yes 250 ug/dose + 1057 45 23 11 724 556 892 0 11542 Yes MPL/TDM 1058 45 23 23 724 780 1588 630 9915 Yes emulsion adjuvant 1059 91 64 23 91 2004 3090 1016 4690 Yes 1060 45 45 11 181 916 1522 448 6620 Yes All birds received 102 EID50 Texas GB strain of NDV, except group 1, which received 104 EID50 Texas GB strain of NDV and group 3, which was the non-challenge control. Birds were challenged 31 days post last vaccination. Bird numbers bolded had a delayed onset to mortality see Table 9. n/c--nonchallenged.
[0264]Typically, a high titer response after challenge is indicative of a good sensitizing immunization, yet, it is not unprecedented with native or recombinant derived NDV antigen that birds will be protected without a detectable HAI or ELISA antibody titer (Winterfield, R. W., Dhillon, A. S., and L. J. Alby, 1979. Vaccination of Chickens against Newcastle Disease with Live and Inactivated Newcastle Disease Virus. Poultry. Sci. 59: 240-246). It is proposed that either a cellular immune response or a local immune response is involved with providing immune protection in a vaccinated bird where no humoral antibody response can be detected (Agrawal, P. K. and D. L. Reynolds. 1991. Evaluation of the cell mediated immune response of chickens vaccinated with Newcastle disease virus as determined by the under-agarose leukocyte-migration inhibition technique. Avian Dis. 35: 360-364).
[0265]In some cases, although there was a detectable titer at the end of the vaccination schedule at day 28, there was no protection to challenge. However, in all cases when this situation occurred there was no detectable antibody titer before (on the day of challenge) or after challenge indicating that these birds were not sensitized effectively (compare Tables 4 and 6). Differences observed between the two poultry trials may be attributed to the fact that antigen used in the first trial had a much higher biological activity per microgram of HN. Antigen used in this trial had at least a 2 fold higher level of hemagglutination activity per microgram of protein than antigen in the latter trial (compare Tables 5 and 7). This may be one reason why birds treated with non-adjuvanted antigen (group #3, Table 6) developed detectable ELISA and HAI titers by day 28, whereas, birds treated with non-adjuvanted antigen (group #4, Table 8) did not show ELISA titers by day 28. Another difference which may have contributed to the results is the age of the birds. Birds in the first trial were vaccinated at day 2 of age, whereas, birds in the second trial were vaccinated at day 10 of age.
TABLE-US-00013 TABLE 9 Death on days after challenge D D % Group # Treatment Route Chall. D 1 D 2 D 3 D 4 D 5 D 6 D 7 D 8 D 9 10 11-14 Surv. 1-018 Control allantoic fluid SQ 104 0 0 3 2 -- -- -- -- -- -- -- 0 EID50 2-018 NDV HN allantoic fluid - 20 ug/ SQ 102 0 0 0 0 0 0 0 0 0 0 0 100 dose EID50 3-018 Control tobacco SQ None 0 0 0 0 0 0 0 0 0 0 0 100 4-018 NDV HN tobacco derived - 20 SQ 102 0 0 0 0 1 0 0 0 1 -- -- 60 ug/dose EID50 5-018 NDV HN tobacco derived - 20 SQ 102 0 0 0 0 0 1 0 0 0 0 0 80 ug/dose + MPL/TDM EID50 Emulsion adjuvant 6-018 NDV HN tobacco derived 250 ug/ SQ 102 0 0 0 0 0 0 1 0 0 0 0 80 dose EID50 7-018 NDV HN tobacco derived 250 ug/ SQ 102 0 0 0 0 0 0 0 0 0 0 0 100 dose + MPL/TDM Emulsion EID50 adjuvant 1-016 Control tobacco SQ 102 0 0 0 4 1 -- -- -- -- -- -- 0 EID50 2-016 NDV HN allantoic fluid SQ 102 0 0 0 0 0 0 0 0 0 0 0 100 EID50 3-016 NDV HN tobacco derived SQ 102 0 0 0 0 0 0 0 0 0 0 1 80 EID50 4-016 NDV HN tobacco derived + SQ 102 0 0 0 0 0 0 1 0 0 0 0 80 MPL/TDM Emulsion adjuvant EID50 5-016 NDV HN tobacco derived IN 102 0 0 0 2 3 -- -- -- -- -- -- 0 EID50 6-016 NDV HN tobacco derived + MPL IN 102 0 0 0 3 0 0 0 2 -- -- -- 0 adjuvant EID50 7-016 NDV HN tobacco derived oral gav. 102 0 0 0 3 2 -- -- -- -- -- -- 0 EID50 8-016 NDV HN tobacco derived + MPL/ oral 102 0 0 0 2 3 -- -- -- -- -- -- 0 TDM adj EID50
[0266]Adjuvant also seems to be an important feature in formulating the antigen in these trials. Although the adjuvant effect was not evident when using higher doses of NT-1 derived NDV-HN (compare groups 3 and 4, Table 6 with groups 6 and 7, Table 8), there was a clear adjuvant effect when using a low dose of NDV-HN (compare groups 4 and 5, Table 8). In addition, although there was 100% mortality in groups inoculated by intranasal route, the group that received adjuvant had 2 birds (#923 and #1088) that did not die until day 8, which was 3 days after all negative control birds had succumbed to challenge (Table 9). The significance of the delay to mortality in the intranasal group receiving adjuvanted antigen is not significant, however, it is interesting that birds #923 and #1088 were two birds with detectable HAI antibody titers at day 21 of the trial and were treated with less antigen per dose than the other treatment groups (see Table 5).
[0267]It is clear from the data provided here that HN-NDV derived from transformed NT-1 cells is efficacious against virulent challenge to NDV. The immune response to the plant derived antigen has several similarities to immune response to native antigen. 1) Although the antibody titers are higher for native antigen pre-challenge, a 20 ug dose inoculated SQ will provide protection against challenge for both native and plant derived antigen. 2) Antibody titers must have similar duration of response in that in both studies challenge was performed 24 days and 31 days post vaccination. 3) All birds producing a positive HAI antibody response (above background) at the end of the vaccination schedule and post challenge were protected from NDV associated pathology.
[0268]No birds were protected from challenge when inoculated by oral or intranasal route. In the case of the oral administered birds, inoculums of 100 to 300 ug of HN-NDV soluble protein from NT-1 cells along with 700 to 2400 ug of HN-NDV feed as whole wet cells per inoculation did not elicit a detectable antibody titer after three doses nor were birds protected from challenge. Because the HN-NDV from NT-1 has definite binding capability to red blood cells and to (CEF) chick embryo fibroblasts, the ability to bind to native receptors did not supplement binding or delivery to antigen sampling sites on the bird mucosal surface in this study.
[0269]The data provided here show that HN-NDV antigen derived from transgenic NT-1 cell culture will protect against virulent challenge when administered SQ. Furthermore, despite feeding several milligrams of antigen in the oral treatment groups, no HAI antibody was induced in the systemic compartment and no protection against challenge was observed. Thus, as previous data have shown, antigens that do not have a natural affinity or invasiveness for the antigen presenting sites on the mucosal surface need to be targeted to those sites with the aid of a protein that does sensitize the mucosal surface.
EXAMPLE 10
Preparation and Analysis of Transgenic Potato
[0270]Binary vectors pCHN, pgHN, pMHN and pCHA were used to transform potato (cv. Desiree) and transgenic tubers analysed for expression of the recombinant HN antigen from NDV, or the recombinant HA antigen from Avian Influenza Virus.
[0271]Plant material. In vitro plants of Solanum tuberosum cv Desiree were provided by Dr. Steven Slack, Department of Plant Pathology, Cornell University. For propagation, nodal segments were transferred to a shoot propagation medium designated CM which contained MS salts (Murashige and Skoog, 1962), 100 mg/l myoinositol, 0.4 mg/l thiamine, 20 g/l sucrose, and 8 g/l Agar/Agar (Sigma Chemical Co., St. Louis, Mo.; catalog #A-1296). The pH of the medium was adjusted to pH 5.7 before the addition of the Agar/Agar. One nodal explant was placed in each test tube and maintained at 24°±1° C. under a photoperiod of 16 h (light)/8 h (dark) at 74 μE m-2 s-1. The source of light for these cultures and those described throughout this report was from a mixture of cool and warm fluorescent bulbs (F40CW and F40WW) (Philips Lighting Co., www.lighting.philips.com/index.htm). Nodal explants were harvested and transferred to fresh medium every 6 weeks.
[0272]Agrobacterium preparation. Agrobacterium tumefaciens containing the gene construct of interest was streaked from a glycerol stock maintained at -80° C. onto Petri plates of LB medium which contained 10 g/l bacto tryptone, 5 g/l yeast extract, 10 g/l NaCl, 50 mg/l spectinomycin, and 15 g/l Difco Bacto Agar (Difco Laboratories, Detroit, Mich.; catalog #DF 0140-01). Four well-formed colonies were picked with a sterile pipet tip, then added to 50 ml of YM medium (Gibco BRL cat.# 10090-011) containing 50 mg/l spectinomycin. Cultures were grown in a shaking incubator at 28° C. and 100 rpm for 24 hrs, or until the culture reached an OD600 of 0.5-0.7. It takes approximately 24 hrs to reach this OD. When the desired OD was reached, the cells were centrifuged at 8000 rpm for 10 min at 20° C. The pellet was resuspended in MS liquid medium (MS salts, 2 mg/l glycine, 0.5 mg/l nicotinic acid, 0.5 mg/l pyridoxine, 0.4 mg/l thiamine, 0.25 mg/l folic acid, 0.05 mg/l d-biotin, 100 mg/l myoinositol, 30 g/l sucrose, pH 5.6) at the same original volume as the YM selective medium.
[0273]Infection. Stem internode segments 0.5-1 cm in length were excised from six-week-old in vitro plants and inoculated the same day. Approximately 100 internode explants were incubated per 50 ml of inoculum for 10 min, agitating occasionally. After the incubation, they were blotted onto sterile filter paper, then transferred to medium designated CIM which contained MS salts, 2 mg/l glycine, 0.5 mg/l nicotinic acid, 0.5 mg/l pyridoxine, 0.4 mg/l thiamine, 0.25 mg/l folic acid, 0.05 mg/l D-biotin, 100 mg/l myoinositol, 30 g/l sucrose (grade II; PhytoTechnology Laboratories, Shawnee Mission, Kans.), 1 mg/l benzyladenine (BA), 2 mg/l naphthaleneacetic acid (NAA) (added after autoclaving), and 6 g/l Agar/Agar (PhytoTechnology Laboratories, Shawnee Mission, Kans.). The pH of the medium was adjusted to 5.6 before the addition of the Agar/Agar. One hundred explants were cultured per 100×20 mm Petri plates. All cultures were maintained at 24°±1° C. under a photoperiod of 16 h (light)/8 h (dark) at 74 μE m-2s-1.
[0274]Plant regeneration. After 48 hrs of cocultivation, the explants were transferred to 3C5ZR bialaphos selective medium which contained MS salts, 0.1 mg/l thiamine, 0.5 mg/l nicotinic acid, 0.5 mg/l pyridoxine, 100 mg/l myoinositol, 30 g/l sucrose, 0.5 mg/l indole-3-acetic acid (IAA) (added after autoclaving), 3 mg/l zeatin riboside (added after autoclaving), 500 mg/l carbenicillin (added after autoclaving) (Agri-Bio, Miami, Fla.), 5 mg/l bialaphos (added after autoclaving) (Duchefa, http://www.duchefa.coml/), and 8 g/l Agar/Agar. The pH of the medium was adjusted to 5.9 before the addition of the Agar/Agar. Twenty-five internode segments were cultured per 100×20 mm Petri plate and the plates were sealed with Nesco film (Karlan Research Products, Santa Rosa, CA). Explants were transferred weekly for 1 month, then every 10-14 days. All cultures were maintained at 24°±1° C. under a photoperiod of 16 h (light)/8 h (dark) at 74 μM-2S-1.
[0275]When regenerants were approximately 0.5-1 cm in length, they were excised and transferred to bialaphos selective rooting medium which contained the same components as CM with the addition of 500 mg/l carbenicillin (added after autoclaving) and 5 mg/l bialaphos (added after autoclaving). Five regenerants were cultured per GA7 Magenta box. Once the shoots rooted, the shoot tip from each plant was transferred to CM in test tubes for maintenance.
[0276]Microtubers. Microtubers were induced on plant material for an early indication of expression in tubers. This was especially applicable for transgenic lines containing genes driven by the tuber-specific promoter, GBSS. Nodal segments were placed on microtuber medium which contained 1/2 strength MS salts, 5 mg/l kinetin, 80 g/l sucrose, 0.25 mM ancymidol (added after autoclaving), 9 g/l Agar/Agar. The pH of the medium was adjusted to 5.85 prior to the addition of the Agar/Agar. The cultures were maintained in the dark at 18° C. The microtubers were analyzed by ELISA for antigen expression levels.
[0277]PCR analysis. Genomic DNA was isolated from leaves from 3-4-week-old putative transformants. Leaf samples were homogenized at room temperature in 500 μl of an extraction buffer containing 200 mM Tris HCl (pH 7.5), 250 mM NaCl, 25 mM EDTA, and 0.5% SDS. They were allowed to remain at room temperature for 1 hr, then centrifuged at 12,000 rpm for 5 min. The supernatant was removed to a new tube, 500 μl of isopropanol was added, then the samples either remained at room temperature for 5-10 min, or were placed at -20° C. overnight. They were then centrifuged at 13,000 rpm for 5 min, and the supernatant was discarded. The resultant pellet was washed with 70% ethanol, dried, then resuspended in 100 μl of TE buffer.
[0278]The primer set was designed such that the forward primer was in the CVMV promoter and the reverse primer (PAT R2) was in the PAT gene which resulted in a product size of approximately 500 bp. Amplified DNA fragments were run on a 1% agarose gel, stained with ethidium bromide, and visualized under a UV light.
[0279]ELISA analysis of leaves. Leaf material was harvested into tubes then placed on ice. Lines with the highest antigen levels were selected for propagation, then transferred to the greenhouse.
[0280]Greenhouse acclimation. Plants with well-formed root systems were transferred to Jiffy 7 pots. The pots were placed in trays and covered with plastic domes. After approximately 2 weeks, the domes were removed. The plants were transferred to 3 gallon pots containing Cornell soil mix when the roots systems had grown through the mesh on the Jiffy 7 pots.
[0281]HN expression in pCHN-transformed potato plants. Potato (Solanum tuberosum L. cv. Desiree) plants were transformed with pCHN, and regenerated Bialaphos® resistant plants were screened for expression of HN in leaves by ELISA. Several lines were selected based on leaf expression (FIG. 26), propagated, and transplanted to soil for greenhouse culture. At maturity, tubers were harvested, extracted, and assayed for HN content by ELISA (FIG. 26). HN accumulation varied among individual tubers from the same line, but in general expression was correlated with the HN content of leaves within each line (FIG. 26). The best expression was observed in tubers of lines 6, 21, 27, and 34, with the highest accumulation observed at ˜11 μg HN per g fresh tuber mass.
[0282]Particle behavior of potato tuber-expressed HN antigen. In order to evaluate assembly of NT1 cell-expressed antigen to form particulate structures, sucrose gradient sedimentation was performed on pCHN-transformed potato tuber extracts. The profiles shown in FIG. 27 indicate that the tuber-derived HN showed 2 peaks of ELISA reactive material, similar to the NT1 cell-derived HN shown in FIG. 24.
[0283]HN expression in potato plants transformed with pGHN and pGHN151. Potato (Solanum tuberosum L. cv. Desiree) plants were transformed with pGHN or pGHN151, and regenerated Bialaphos® resistant plants were screened for expression of HN in microtubers by ELISA. Several pGHN-transformed lines were selected based on microtuber expression (FIG. 28), propagated, and transplanted to soil for greenhouse culture. Transformation with pGHN151 was relatively inefficient, resulting in only one line that showed expression in microtubers (GHN151-6), which was transplanted to soil for greenhouse culture. At maturity, tubers were harvested, extracted, and assayed for HN content by ELISA (FIG. 28). HN accumulation varied among individual tubers from the same line, but in general expression was correlated with the HN content of microtubers within each line (FIG. 29). The best expression was observed in tubers of lines GHN-1, 30, 47, and 54, with the highest accumulation observed at 40 μg HN per g fresh tuber mass. Expression in tubers of line GHN151-6 varied between 6 and 12 μg HN per g fresh tuber mass. It is possible that the intron-containing GBSS promoter construct pGHN151 was unstable in transgenic plants or in Agrobacterium, resulting in poor expression with this construct.
[0284]ELISA analysis of tubers. Approximately 3-4 months after plants were transferred to soil, assorted tissue were harvested for analysis. FIG. 16 shows expression of HA in microtubers of pCHA transformed microtubers, which ranged up to 700 ng/g fresh weight. This is similar to the accumulation observed in pGPTV-HAO-transformed tubers (HA gene driven by CaMV 35S promoter), which was maximal at 1 ug/g fresh weight. Selected lines were transplanted to soil and grown in the greenhouse. Leaves of soil-grown plants were sampled and assayed by ELISA (FIG. 17). Expression of HA in leaves was very poor (<0.025 ng/μg TSP), which is consistent with the earlier assays with leaves of tissue culture plants. Tubers of mature plants were harvested, extracted, and evaluated for HA expression by ELISA. Accumulation of HA in tubers was maximally 500 ng/g fresh weight (FIG. 18). The expression observed in microtubers produced in vitro was well correlated with the expression in soil-grown tubers (FIGS. 16 and 18), thus the microtuber is a good model for expression of HA with pCHA.
EXAMPLE 11
Preparation and Analysis of Transgenic Tomato
[0285]Binary vectors pCHN, pMHN, and pUHN were used to transform tomato (variety TA234) and transgenic fruit and leaves analysed for expression of the recombinant HN protein from Newcastle Disease Virus.
[0286]Plant material. Seeds from a tomato line designated TA234 were used for transformations. TA234, originally known as Momor, is a verticillium and tobacco mosaic virus resistant line derived from the variety Moneymaker. Seeds were surface sterilized in 20% Clorox, for 20 min, rinsed 3 times with sterile distilled water, then cultured on 1/2 MSO medium (See below) in Magenta boxes. They were maintained at 24°±1° C., under a photoperiod of 16 h (light)/8 h (dark) at 74 μm-2S1. The source of light for these cultures and those described throughout this report was a mixture of cool and warm fluorescent bulbs (F40CW and F40WW) (Philips Lighting Co., www.ligting.philips.com/index.htm). The seed cultures were maintained for 6-8 days depending upon the stage of cotyledon growth. Cotyledons were excised before the first true leaves emerged. If cotyledon sections were longer than 1 cm, they were cut into two 0.5 cm segments. Cotyledon sections were placed on feeder layer plates which were prepared one day prior to transformation. The feeder layer consisted of NT1 suspension cultured cells plated on KCMS medium (See below) which had been subcultured (2 mls of cells:48 ml of liquid KCMS) 7 days prior to plating. The plated suspension culture was covered with a sterile 7 cm Whatman filter paper. Cotyledon sections were placed on top of the filter paper.
[0287]Agrobacterium preparation. Agrobacterium tumefaciens containing the gene construct of interest was streaked from glycerol stocks maintained at -80° C. onto fresh plates of LB medium (See Appendix) containing the appropriate antibiotic. For DAS constructs, 50 mg/l spectinomycin was added to the LB medium. The cultures were incubated for 24-48 hrs at 28° C. The duration of the incubation time was dependent upon colony size. If pin-point colonies developed after 24 hrs, the cultures were incubated for an additional day.
[0288]When the colonies were of a well-formed size, liquid cultures were prepared. Four colonies were picked with a sterile pipette tip, then added to 50 ml of YM selective medium containing 50 mg/l spectinomycin for DAS constructs (See below). Cultures were grown in a shaking incubator at 28° C. and 100 rpm for 24 hrs, or until the culture reached an OD600 of 0.5-0.6. It takes approximately 24 hrs to reach this OD. When the desired OD was reached, the cells were centrifuged at 8000 rpm for 10 min at 20° C. The pellet was resuspended in MS liquid medium at the same original volume as the YM selective medium.
[0289]Infection. Cotyledon explants were cultured on the feeder layer plates 1 day prior to infection with Agrobacterium. For infection, they were incubated in the Agrobacterium suspension for 10 min, then the suspension was removed. The explants were blotted on sterile paper towels, then placed with the adaxial sides down on the original feeder plate cultures. They were maintained at 19° C. in the dark for 48 hrs of cocultivation.
[0290]Plant regeneration: After cocultivation, cotyledon explants were cultured with the adaxial sides up on selective 2Z medium containing 3 mg/l bialaphos. The cultures were maintained at 24±2° C. under a 16-hr photoperiod of cool white fluorescent lights. Three weeks later, the cultures were transferred to 1Z medium containing 3 mg/l bialaphos (See below), then to fresh medium at 3 week intervals. When shoots began to regenerate, the cultures were transferred to the same 1Z medium with bialaphos in Magenta boxes. When shoots were 2 cm tall, they were transferred to selective rooting medium containing 2 mg/l bialaphos (See below) in Magenta boxes. Plants were maintained at 24±1° C. under a 16-hr photoperiod of cool white fluorescent lights. After approximately 3 weeks, cuttings from these plants were transferred again to selective rooting medium containing bialaphos, however, timentin was not included to determine if there was Agrobacterium contamination present.
[0291]Analysis. Plants that rooted on selective rooting medium were selected for ELISA analysis. Leaf material was harvested, transferred to 2 ml conical screw cap tubes, and placed on ice. ELISA was performed at least twice before selecting the lines containing the highest antigen level. The elite lines were propagated and transferred to the greenhouse.
[0292]Greenhouse acclimation. Plants were transferred to the greenhouse when they had a well-developed root system. The agar medium was washed off the roots before transferring the plants to 6-inch pots containing Cornell mix. They were covered with plastic domes. During the next week, the domes were gradually lifted to acclimate the plants. After approximately 5 weeks, the plants are transferred to 3-gallon pots containing Cornell mix.
Media Ingredients
1/2 MSO
TABLE-US-00014 [0293] Per liter MS salts 2.15 g Myoinositol 100 mg Thiamine HCl stock (0.4 mg/ml) 5 ml Pyridoxine HCl stock (0.5 mg/ml) 1 ml Nicotinic acid stock (0.5 mg/ml) 1 ml Sucrose 10 g pH to 5.8 ± 0.03 Agar/Agar 8 g
KCMS
TABLE-US-00015 [0294] Per liter MS salts 4.3 g Thiamine HCl stock (1 mg/ml) 1.3 ml Myoinositol 100 mg 2,4-D stock (1 mg/ml) 200 μl KH2PO4 200 mg Kinetin stock (1 mg/ml) 100 μl Sucrose 30 g pH to 5.5 ± 0.03 Agargel 5.2 g
LB
TABLE-US-00016 [0295] Per liter Bacto-tryptone 10 g Yeast extract 5 g NaCl 10 g Difco Bacto Agar 15 g
YM
TABLE-US-00017 [0296] Per liter Yeast extract 400 mg Mannitol 10 g NaCl 100 mg MgSO4•7H20 200 mg KH2PO4 500 mg
Alternatively, YM in powder form can be purchased (Gibco BRL, catalog #10090-011). To make liquid culture medium, add 11.1 g to 1 liter water.
MS Liquid Medium
TABLE-US-00018 [0297] Per liter MS salts 4.3 g Myoinositol 100 mg Glycine 2 mg Nicotinic acid 0.5 mg Pyridoxine HCl 0.5 mg Thiamine HCl 0.4 mg Folic acid 0.25 mg D-biotin 0.05 mg Sucrose 30 g pH 5.6
2Z
TABLE-US-00019 [0298] Per liter MS salts 4.3 g Myoinositol 100 mg Nitsch vitamins stock (1000X)* 1 ml Sucrose 20 g pH to 6.0 ± 0.3 Agargel 5.2 g
Selective Rooting Medium
TABLE-US-00020 [0299] Per liter MS salts 4.3 g Nitsch vitamins stock (1000x)* 1 ml Sucrose 30 g pH to 6.0 + 0.03 Difco Bacto Agar 8 g
Add the following filter-sterilized components per liter after autoclaving:Bialaphos: 2 ml of a 1 mg/ml stock solutionTimentin: 3 ml of a 100 mg/ml stock solution
Nitsch Vitamins Stock (1000×)
TABLE-US-00021 [0300] Per 50 ml Glycine 0.1 g Nicotinic acid 0.5 g Pyridoxine HCl 0.025 g Thiamine HCl 0.025 g Folic acid 0.025 g d-biotin 0.002 g
[0301]Adjust pH to 7.0 to clear solution.
EXAMPLE 12
Tomato as a Production System of Edible Vaccines
[0302]Assembly of a Synthetic HN Gene. A HN expression cassette that includes the promoter of the Casava vein mosaic virus (CsVMV) and terminated by the 3' element of the Soybean Vegetative Storage Protein (VSP) was assembled and inserted into binary vectors by the Mason Laboratory (The Boyce Thompson Institute for Plant Research (BTI)) for Agrobacterium-mediated plant transformations. The vector carries the gene encoding the plant selection marker phosphinothricin acetyl transferase (PAT, described in U.S. Pat. Nos. 5,879,903; 5,637,489; 5,276,268; and 5,273,894) (FIG. 30).
[0303]Tomato Transformation and Regeneration. Agrobacterium-mediated transformation of tomato cotyledons (variety Tanksley TA234TM2R) was performed according to Frary and Earle [12] by the Van Eck Laboratory (BTI). Regenerating explants were transferred to fresh medium every 3 weeks. Green shoots were transferred to GA-7 boxes (Magenta Corporation, Chicago, Ill.) containing rooting media (MS salts 4.3 g, 1 ml/l Nitsch vitamins 1000×, 30 g/l sucrose, 8 g/l difco bacto agar, pH 6.0, supplemented with 50 mg/l kanamycin, 300 mg/l timentin and 4 mg/l IBA) when they were approximately 1 cm tall. Rooted plantlets were transferred to soil and maintained at 28° C. under a 12 hours photoperiod.
[0304]Protein Extraction and Elisa Analysis. Crude protein extracts were made by homogenizing 1 mg of fresh leaf, fruit or NT1 cell material per 5 μL of PBS or 1 mg of dried leaf, fruit or NT1 cell material per 10 μL of PBS in a QBiogene (Carlsbad, Calif., USA) Fast Prep machine. Insoluble material was removed by centrifugation at 14,000 rpm in an Eppendorf 5415C microcentrifuge at 4° C. for 5 minutes. The resulting sample supernatants were kept on ice during analysis and subsequently stored at -80° C.
[0305]Ninety-six well, microtiter plates (Costar 3590, Fisher Scientific, PA, USA check) were coated with 100 μl per well of a 1 in 1,000 dilution of SPAFAS chicken anti-NDV polyclonal antibody (Benchmark Biolabs, N E) in 0.01 M borate buffer. The plates were covered and incubated overnight at 4° C. The plates were equilibrated to room temperature for 30 minutes then washed three times with 300 μl per well phosphate buffered saline with 0.05% Tween-20 (PBST). The plates were blocked with 200 μl per well, 3% skim milk in PBST at 37° C. for two hours then washed three times with PBST before 50 μl per well of protein extracts were added. ELISAs were performed on two replicates on a series of two-fold dilutions of the crude extracts in 5% skim milk+PBS+0.05% Tween-20. The plates were incubated for one hour at 37° C. before washing three times with PBST. One hundred microliters of the primary antibody, HN Mab 4A (Benchmark Biolabs), diluted 1 in 250 in 1% skim milk in PBST, was added to each well and incubated for one hour at 37° C. The plates were then washed three times with PBST before 100 μl per well of goat, anti-mouse IgG horse radish peroxidase (HRP) conjugate (Sigma, St Louis, Mo., USA) diluted 1 in 3,000 in 1% skim milk in PBST was added and left to incubate at 37° C. for one hour. The plates were washed four times with PBST before 50 μl per well of TMB Peroxidase EIA Substrate kit (BioRad) was added and incubated for five minutes at room temperature. Absorbance at 450 nm was measured in a ThermoMax Micropla reader. ELISA data obtained by anti-HN ELISA was converted to microgram per gram of fresh weight by reference to a standard curve constructed using purified HN (Benchmark Biolabs).
[0306]The top four lines based on HN expression in the leaves were used for fruit analysis.
[0307]Nucleic Acid Extraction. Ten milliliters of extraction buffer (4% p-amino salicylic acid, 1% 1,5 naphthalenedisulfonic acid, disodium salt hydrate), 3 ml CTAB buffer, and 10 ml buffer-saturated phenol (pH 4.3) were added to a 50 ml falcon tube and heated at 70° C. in a water bath for 10 minutes. About 3.5 g of individual tomato fruit were ground in liquid nitrogen then added to the heated tube and vortexed vigorously for 30 seconds. Ten milliliters of chloroform:isoarnylalcohol (24:1) was added. The resulting slurry was vortexed for 30 seconds before centrifuging for 20 minutes at 10,000 rpm at 4° C. The aqueous phase was transferred to a 50 mL falcon tube, mixed with 2 volumes of ethanol, and precipitated for 15 minutes at room temperature. The extract was then centrifuged for 15 minutes at 10,000 rpm at 4° C. before the supernatant was discarded. The resulting nucleic acid pellet was resuspended in 2 ml DEPC treated water, mixed with an equal volume of 4 M LiCl, and precipitated at -20° C. overnight. The extract was centrifuged at 10,000 rpm at 4° C. for 20 minutes. The supernatant containing genomic DNA was removed to a different tube, precipitated with 2 volumes of ethanol and stored at -20° C. overnight. Meanwhile, the RNA pellet was resuspended in DEPC treated water and stored at -20° C. The following day, the DNA pellet was centrifuged down at 10,000 rpm at 4° C. for 20 minutes and resuspended in water containing 1 μg/ml Rnase A.
[0308]Southern Analysis. Fifteen micrograms of tomato genomic or 330 ng of pCHN plasmid DNA were digested with 5 units of restriction enzyme EcoRI per μg DNA at 37° C. for 20 to 24 hours. Uncut and digested samples were run overnight in a 1.0% TAE agarose gel. The gel was prepared for transfer by one 20 minute depurination wash (0.25 M HCl), two 30 minute denaturation washes (1.5 MNaCl, 0.5 MNaOH) and two 30 minute neutralization washes (0.5 M Tris HCl, pH 7, 3 MNaCl). DNA was then transferred to a nylon membrane (Zeta-Probe blotting membranes, Bio-Rad, Hercules, Calif., USA) by capillary transfer and fixed by UV cross-linkage using a Bio-Rad GS Gene Linker. A PCR labeled probe was made by using the primer set HNa (CCG AGC AGT TTC ACA AGT GG, SEQ ID NO: 10) and HNb (CCT GAT CTT GCT TCA CGT ACA, SEQ ID NO: 11) on a pCHN template. DIG labeled dCTP was incorporated into the 1734 bp amplicon using the Roche Molecular Biochemical DIG PCR Probe Synthesis kit according to manufacturer's instructions. The amplification was performed over 37 cycles using an iCycler Gradient Thermo Cycler (BioRad, Hercules, Calif., USA). The template was initially melted at 94° C. for 5 minutes followed by 35 cycles of 94° C. for 30 seconds, 56° C. for 30 seconds, and 72° C. for 90 seconds. A final extension step was performed at 72° C. for 5 minutes before soaking at 4° C. Hybridization bottles and 10 ml DIG Easy Hyb (Roche Scientific, Mannheim, Germany) per membrane were pre-warmed to 45° C. in a hybridization oven. Membranes were prehybridized for 60 minutes at 45° C. and hybridized overnight at 45° C. with a probe concentration of 5 μl/ml DIG Easy Hyb.
[0309]Post hybridization washes and detection were performed as per manufacturer's instructions (Roche--DIG wash and block buffer set and DIG Luminescent Detection Kit). Labeled membranes were visualized after exposure to film.
[0310]Northern Analysis. Thirty micrograms of total RNA from tomato transformants and wild type plants and 1.25 μg of ladder (high range RNA ladder, MBI Fermentas, Hanover, Md.) were denatured with formaldehyde/formamide and run for two hours in a 1% agarose 3-(N-morpholino) propanesulfonic acid (MOPS) formaldehyde gel at 80V. The RNA was transferred to zeta probe membrane (BioRad, Hercules, Calif., USA) by upward capillary action and fixed by UV cross-linkage. The membrane was then stained with 0.04% methylene blue in 0.5M sodium acetate to determine if RNA transfer was successful and to confirm that RNA concentrations in all samples were similar. A PCR labeled, DNA probe was made using the primer set HNa and HNb on a pCHN template. DIG labeled dCTP was incorporated into the 1734 bp amplicon as per manufacturer's instructions (PCR DIG Probe Synthesis kit, Roche Scientific, Mannheim, Germany). The amplifications were performed as described for Southern analysis. Hybridization bottles and 10 ml DIG Easy Hyb (Roche) per membrane were pre-warmed to 45° C. in a hybridization oven. Membranes were pre-hybridized for at least 90 minutes at 45° C. and hybridized overnight with a probe concentration of 7.5 μl/ml DIG Easy Hyb. Post hybridization washes and detection were performed as per manufacturer's instructions (Roche--DIG wash and block buffer set and DIG Luminescent Detection Kit). Labeled membranes were visualized after exposure to film.
[0311]Western Analysis. Purified HN supplied by Benchmark Biolabs and NT1 cell line 119 transformed with pCHN (supplied by BTI) were used as positive controls. Twenty microliters of protein extracts were added to 4 μl 6×SDS gel loading buffer (300 mM Tris-HCl, pH 6.8, 600 mMDTT, 12% SDS, 0.6% bromophenol blue, 60% glycerol), boiled for 10 minutes and loaded into a 15% sodium dodecyl sulfate polyacrylamide gel. The gel was run in tris-glycine buffer (25 mM Tris, 250 mM Glycine, pH 8.3, 0.1% SDS) at 30 milliamps per gel until the dye front ran about 5 mm from the gel bottom. The separated proteins were transferred from the gel to a PVDF membrane using a BioRad Trans Blot Cell (50 V for 2 hours or overnight at 7 V). All membrane washes were performed in PBST (PBS+0.1% Tween-20) at room temperature unless otherwise stated. The membrane was blocked with 5% skim milk+PBS+0.1% Tween-20 overnight at 4° C. or for two hours at room temperature using slow rotation in a hybridization incubator (Fisher Scientific, Tustin, Calif., USA). The membrane was washed twice briefly before incubating for one hour at 37° C. with a 1 in 50,000 dilution of the primary antibody, mouse anti-HN Mab14F antiserum (Benchmark Biolabs) in 1% skim milk+PBS+0.1% Tween-20. The membrane was briefly rinsed in PBST before a 15 minute wash and three 5 minute washes then incubated in a 1 in 30,000 dilution of an anti-rabbit IgG horseradish peroxidase (HRP) conjugate (Sigma) for an hour at 37° C. with slow rotation. The membrane was rinsed, then subjected to a 15 minute wash and three 5 minute washes. Detection was performed using the Amersham ECL+kit as per manufacturer's instructions.
[0312]Haemagglutination Activity. To make a 1% chicken red blood cell (cRBC) standardized solution, cRBCs in Alsevers solution (Colorado Serum, Co) were transferred into a 15 ml conical tube and centrifuged at 250 g for 10 minutes. The supernatant was aspirated and the pellet resuspended in 10 ml Dulbecco's phosphate-buffered saline without calcium and magnesium (DPBS) (Cellgro, Mediatech, Inc, Kansas City, Mo.). The suspension was centrifuged at 250 g for 10 minutes. The washes and resuspensions were repeated until the supernatant was clear. Once this was achieved the cells were pelleted and the supernatant aspirated to leave the packed RBC pellet. The pellet was then diluted in 1% DPBS-(volume to volume). Four hundred microliters of the 1% RBC solution was transferred to a small tube, 1.6 ml of deionized water was added before being vortexed at high speed for 20 seconds to lyse the cells. A cRBC solution was not used unless the absorbance at 540 nm of the lysed cells was 0.4-0.5. A 96-well, U bottom plate (Falcon) was sprayed with antistatic spray before 50 μl per well of DPBS was added. Fifty microliters of the samples, including the positive control of NDV HN and negative control of DPBS, was added to the first row, mixed through repeated pipetting then serially diluted by transferring 50 μl to the next row. Fifty microliters of the standardized cRBC was added to each well before the dilutions were incubated on a plate shaker at 600 rpm for 20-30 seconds. The plates were then incubated at 5° C. for an hour before the control NDV HN wells were checked for haemagglutination. Once this was achieved the final read was made. The HA titer was taken as the reciprocal of the highest dilution that was positive for agglutination.
[0313]Analysis. Fruit ripening is a developmentally and genetically regulated process that is characterized by many biochemical and physiological changes, including increases in the rate of ethylene biosynthesis and respiration, chlorophyll degradation, pigment accumulation, textural modifications such as fruit softening, changes in the levels of sugars and organic acids, and production of volatile aromatic compounds (Brady CJ. Annu Rev. Plant Physiol. 1987; 38: 155-178). There is a distinct relationship between fruit pH and solids content (mainly sugars, Benton Jones J. Tomato plant culture: in the field, greenhouse, and home garden. New York: CRC Press, 1999). The degree of ripeness is also a factor that affects pH. Ripening of wild type TA234 caused the fruit pH to decrease significantly (α=0.05) (FIG. 31) and the total soluble protein to generally decrease (FIG. 32). Although the extent of decrease varies, this has been found in other studies (Benton Jones J. Tomato plant culture: in the field, greenhouse, and home garden. New York: CRC Press, 1999).
[0314]The four lines expressing the highest level of HN in leaf tissue had varying phenotypes. The phenotypes of lines CHN-1, CHN-12 and CHN-32 were indistinguishable from control plants while line CHN-10 showed traits indicative of polyploidy such as thick, wrinkly leaves and late flower set and fruit development. Comparing the intensity of signals between the plasmid controls and the genomic samples in Southern analysis, the lines had between 1 and 4 copies of the transgene (FIG. 33) with CHN-1 having two copies, CHN-10 four copies, CHN-12 having one copy, and CHN-32 having 2 copies. Since CHN-10 is likely a polyploid line, it will not be used in further studies or for production of vaccine batches.
[0315]Methylene blue staining of the northern membrane before the pre-hybridization and hybridization steps revealed the transfer of RNA from the gel to the membrane was successful and that there were similar concentrations of RNA in each of the samples (FIG. 34a). Northern analysis of total RNA specific for the HN gene demonstrated no band in the wild type negative control however a band of about 5000 nucleotides was seen in the transgenic tomato lines (FIG. 34b). Since the HN gene expression cassette with promoter and terminator is 2,832 bp and the expression cassettes within the T-DNA combined are 4323 bp, the transcript is larger than expected and was thought due to read through of the VSP 3' termination signal during transcription of the HN gene. Transcript prevalence varied at different stages of fruit ripening within a line and between lines. Line CHN-10 displayed a distinct decrease in HN-specific mRNA content as the fruit ripened (FIG. 34b). However since only one fruit was sampled for northern analysis no pattern could be discerned between RNA prevalence and stage of ripening in the other CHN tomato lines.
[0316]Anti-HN ELISA of transgenic lines revealed a trend of decreasing concentration as the fruit ripened (FIG. 35). Stages 4 and 5 had significantly less HN per gram fresh weight than stage 1 fruit in lines CHN-1 (FIG. 35a) and CHN-10 (FIG. 35b) and stage 2 fruit in CHN-32 (FIG. 35c) (α=0.05). HN concentration was not significantly different at the same stage between different varieties (α=0.05). HN concentration varied between 71.1 and 3.5 μg/g fresh weight with the highest concentration in each line being 67.2 μg/g fresh weight in stage 1 fruit of CHN-1, 63.3 μg/g fresh weight in stage 1 fruit of CHN-10, 65.4 μg/g fresh weight in stage 1 fruit of CHN-12 and 71.1 μg/g fresh weight in stage 2 fruit of CHN-32. The best stage to harvest a HN tomato crop would therefore be at the green or early breaker stage.
[0317]Western analysis of the Lasota NDV HN positive control while overloaded, revealed bands around 49, 74, 78, 90, 120, 200, 250 and larger than 250 kDa (FIG. 36). The 78 kDa band was designated HN monomers, the smaller bands degradation products. The 90 kDa band was thought a different glycosylation form and the 120 kDa and larger bands as HN polymers. Western analysis of transgenic fruit at different stages was difficult due to the low expression of HN, however two bands around 78 and 70 kDa were visible. These bands were also present in the leaves of transgenic tomato plants and in the NT1 cell line 119 but were not present in the tomato or NT1 cell line negative controls. It was thought the 78 and 70 kDa bands represented different glycosylation forms of the HN antigen or the 70 kDa band a truncated version of the HN monomer. Additional bands thought to be degradation products were visible around 20 and 48 kDa in leaf samples of transgenic tomatoes and 48 kDa in the NT1119 line.
[0318]NF, represents tomato fruit negative control--wild type fruit; NL tomato leaf negative control--wild type leaf, NNT NT1 cell negative control--non-transformed cell lines; 119, transgenic NT1 cell line 119; L10, leaf from transgenic tomato line 10; L32, leaf from tomato line 32; HN, animal derived Lasota NDV virus; M, Bio-Rad's precision plus protein all blue standard; 1-1, fruit from line CHN-1, stage 1 of ripening; 1-3, fruit from line CHN-1, stage 3 of ripening; 1-6, fruit from line CHN-1, stage 6 of ripening; 32-1, fruit from line CHN-32, stage 1 of ripening; 32-3, fruit from line CHN-32, stage 3 of ripening; 32-6, fruit from line CHN-32, stage 6 of ripening; 10-1, fruit from CHN-10, stage 1 of ripening. Protein size is give in kDa.
[0319]Haemagglutination assays of the freeze-dried green fruit and leaves of the transgenic tomato revealed haemagglutination activity in all lines (FIG. 37). Activity was highest in the leaves in lines CHN-1, CHN-12 and CHN-32 with CHN-10 being the only line to have higher activity in the fruit. Line CHN-32 displayed the highest haemagglutination activities of 512 and 2,048 in the fruit and leaves (FIG. 37a) as well as the highest haemagglutination activity of 10,928 units per microgram HN (FIG. 37b). The CHN-1 line had the second highest haemagglutination activities of 128 and 256 in the fruit and leaves as well as the second highest activity of 3,994 units per μg HN (FIGS. 37a and b). Despite being mis-processed during transcription, the synthetic HN gene was translated into a functional protein.
[0320]Although CHN-10 had higher HN activity in the fruit, the target organ for animal trials and vaccine delivery, the probable polyploid status of this line in addition to its slowness to flower and fruit made it an unlikely candidate for future studies. Taking into account HN expression levels and HN activity in the four lines, the CHN-1 and CHN-32 lines were chosen for future analysis.
[0321]Western analysis, ELISA and haemagglutinin activity assays show that tomato is capable of expressing a HN protein of the correct size (78 kDa) that is antigenic in ELISAs and retains haemagglutination activity. The optimal time to harvest tomato fruit expressing HN under the control of the CsVMV promoter was the early stage of fruit ripening. This decreased the time to harvest by 2 weeks and increased HN expression approximately 15-fold. Despite the protein being correctly processed, northern analysis reveals that the gene is not processed correctly at the DNA level. The 5,000 nucleotide transcript is likely due to read through of the HN gene terminator. Lines CHN-1 and CHN-32 were deemed the best lines for progression to additional studies.
EXAMPLE 13
HN Expression During Maturation of Tomato Fruit
[0322]HN levels in maturing tomato fruit to determine if immature fruit are capable of expressing higher levels of HN than stage one tomatoes.
[0323]The fruit of red-fleshy tomato varieties is said to be mature when it has completed growth but is still completely green. This stage of ripeness is known as "green" or "stage one". Tomatoes are usually picked at this stage then ripened with ethylene. The stages following include "breakers" or "stage two" when there is a definite break in color from green to tannish-yellow or pink (vine ripened tomatoes are picked at this stage); "turning" or "stage three" when more than 10% but less than 30% (for example, 11, 12, 13, 14, 15, 20, 25, 29%) of the surface of the tomato shows a definite change in color from green to tannish-yellow, pink or red; "pink" or "stage four" when more than 30% but less than 60% (for example, 31, 32, 33, 34, 35, 40, 45, 50, 55, 59%) of the surface of the fruit is pink or red in color; "light red" or "stage five" when more than 60% but less than 90% (for example, 61, 62, 63, 64, 65, 70. 75, 80, 85, 89%) of the surface of the fruit is pinkish-red or red; "red" or "stage six" when more than 90% (for example, 91, 92, 93, 94, 95, 99, 100%) of the surface of the fruit is red.
[0324]Expression of the synthetic, plant optimized, gene for the Newcastle disease virus haemagglutinin neuraminidase (H) protein driven by the Casava Vein Mosaic Virus promoter (CsVMV), decreased as the tomato fruit ripened. It was determined that in stage one green tomato fruit, HN expression was approximately 12 μg/g fresh weight (FW) and that this steadily decreased as the tomato ripened to stage six red tomato fruit to an approximate value of 2.5 μg/g FW.
[0325]One T1 plant from each of the elite lines in the T0 CHN generation (CHN-1, CHN-32) was germinated and allowed to grow. When flowering began, cross-pollination was prevented by each flower being self-pollinated by hand and enclosed in a paper towel. The individual flower was dated and allowed to fruit. Three fruit from each plant line were harvested at one week post-pollination, two weeks post-pollination, four weeks post-pollination, six weeks post-pollination, and finally at stage one green fruit (about eight weeks post pollination). Fruit diameter (mm) and fresh mass (g) were recorded before ELISA analysis of the HN content and lyophilization. To measure the diameter of the tomato fruit, a vernier caliper was applied to the widest part of the tomato fruit perpendicular to the stem and the measurement in millimeters recorded. Mass was determined using a gram balance.
[0326]HNELISA. SPAFAS chicken A-NDV polyclonal antibody diluted 1:1500 in 0.01 M borate buffer was used to coat a 96 well ELISA plate. One hundred microliters of the dilution was pipetted into each well before the plate was covered, and left overnight at 4° C. The next morning the plate was allowed to equilibrate for 30 minutes at 24° C., before being washed three times with PBST (0.05% tween). A solution of 3% skim milk was made and 200 μl added to each well. The plate was placed at 37° C. and allowed to block for two hours.
[0327]To collect a sample, a coring tool (size 1) was pushed through the center of the tomato along the horizontal axis. Any gelatinous material or seeds were excluded from the sample. Using a scalpel, approximately 1 cm of the tomato was collected and placed into the sample tube that was then placed on ice. The actual sample weight was calculated by subtracting the individual tube weight from the total mass then 20× tomato extraction buffer (4M Nacl (final concentration 100 mM), 0.5M EDTA (final concentration1mM), 20% Triton-x 100 (final concentration 10%), Leupeptin (final concentration 10 ug/ml), 0.5M NaPi, pH 7.0 (final concentration 50 mM), brought to volume with milliQ water) (mass by volume) was added to the tube. A ceramic bead was added to the sample tube and the sample homogenized using a fast prep machine at speed 4.0 for 30 seconds. The homogenized samples were centrifuged and then set aside on ice while the standard curve was prepared.
[0328]After the plate had blocked for two hours it was washed three times with PBST (PBS plus 0.05% tween). For the NDV HN standard curve, 100 μl of 232 ng/ml NDV purified stock (1:80) was pipetted into the second well of the second row. Fifty microliters of 1% skim milk was pipetted into the remaining wells of the second row and each of the wells in rows 3-8 of the microtiter plate. Fifty microliters of each plant sample was then added to the 1% skim milk in wells 3-11 of the second row. The purified NDV as well as the plant samples were then serially diluted down the plate by pipetting 50 μl out of row two and into row three and so on down the plate. The samples were mixed in the wells by pipetting up and down after each dilution step. The initial concentration of the samples was a 40-fold dilution. The plate was placed back into the 37° C. incubator for 1 hour.
[0329]After incubating for one hour the plate was washed three times with PBST and the primary antibody added. The primary antibody NDV HN Mab 4A was diluted to a concentration of 1:250 in 1% skim milk and 100 μl added to each well. This was allowed to incubate for one hour at 37° C. Next the plate was washed three times with PBST and the secondary antibody goat anti-mouse IgG, was added to each well at a concentration of 1:3000 in 1% skim milk. This was allowed to incubate for one hour at 37° C.
[0330]The plate was washed four times with PBST and 50 μl TMB substrate was added to each well. After five minutes had elapsed the TMB was neutralized with 1N H2SO4. The plate was then read on a spectrophotometer at a wavelength of 450 nm.
[0331]The percent water loss was determine by removing the seeds from each tomato fruit, measuring the mass, freezing at -20° C., lyophilizing, then reweighing the tomato fruit.
[0332]To take into account the increase in fruit size in addition to HN concentration, the HN content of a tomato fruit at each of the maturation stages selected was calculated by multiplying the fruit HN concentration by the fruit mass. In addition, the data generated from this study was used to calculate the possible number of doses produced from the CHN elite lines if fruit were harvested at stage one or at four weeks post pollination. In this model it was assumed that the same number of fruit would be produced from a plant harvested when fruit were at stage three and a plant harvested when fruit were four weeks post pollination and that one dose would be 50 μg of antigen.
Results
Fruit Physiology
[0333]As expected, fruit size, mass and percent water loss increased with time (FIGS. 38, 39, and 40). Only small variations were observed between repetitions of measurements taken at the same maturation stage within the same plant line. No significant difference was seen between line 1 and 32 at the same stage in maturation (α=0.05).
HN Content of Maturing Fruit
[0334]Concentration of HN in maturing fruit peaked at two weeks post pollination then decreased as the fruit matured (FIG. 41). There was no significant difference between the two tomato lines at the same stage of maturation nor between the HN content in the first four weeks after pollination (α=0.05). There was significantly more HN in fruit two weeks post pollination than in fruit six weeks after pollination and when mature and in the stage 1 of ripening (α=0.05).
[0335]To take into account the increase in fruit size in addition to HN content, the amount of HN of a tomato fruit at each of the maturation stages selected was calculated. The HN content within a fruit peaked at four weeks post pollination (723 μg for line CHN-1 and 630.9 μg for line CHN-32) before decreasing with further fruit maturation (FIG. 42). There was no significant difference between lines at the same stage of maturation (α=0.05).
[0336]Calculations were made for the number of doses produced by the HN elite lines if fruit were harvested at stage 1 or four weeks post pollination (Table 10). It was assumed the same number of fruit would be produced from each harvest and that one dose would be 50 μg of antigen. These data indicate that if 33 fruit are harvested at stage one, 126 doses would be produced; and if fruit were harvested four weeks post pollination, 486 doses would be produced. Thus harvest time is reduced by four weeks and there is a 286% increase in doses yielded.
TABLE-US-00022 TABLE 10 Effect of harvest time on number of HN doses produced. Four Weeks Post Characteristic Stage 1 Pollination Antigen Concentration 2.5 18.4 (μg/g) Average Weight 76.2 40 (g/fruit) Number of Fruit Harvested 33 33 Total Mass of Fruit 2514.6 1320 Produced (g) Total Mass of Antigen 6.3 24.3 (mg) Number of 50 μg Doses 126 486
[0337]Although HN concentration peaked at 38.8-42 μg/g fresh weight in tomato fruit two weeks post pollination, no significant difference was found in HN concentration in the first four weeks after pollination (α=0.05). When mass was taken into account however, tomato fruit that were four weeks post pollination averaged between 631-723 μg HN, which was a significantly higher amount of HN than the other maturation stages tested (a=0.05). Since percentage of water loss between fruit at stage one of ripening and two and four weeks post pollination did not varying greatly (total difference of 2.6%) the significantly higher antigen content at four weeks pollination was not a factor of varying water content or dilution of antigen. These data suggest that the best time to harvest tomato fruit expressing HN under the control of the CsVMV promoter is four weeks post pollination.
[0338]To identify easily when fruit are four weeks post pollination, fruit mass and diameter were recorded throughout their maturation. Fruit four weeks post pollination averaged a mass of 40 g and a diameter between 42 and 45 mm. Fruit size is affected by genetics, temperature, day length and plant age. The small standard errors of our means indicated that genetics is not presently an issue with regards to plant-by-plant variation in fruit size. However the effect of stress (due to change in temperature, day length and plant age) means that fruit size may not always prove an accurate indication of time post pollination. Large temperature fluctuations, day length and plant age should therefore be kept in mind, when approximating time of pollination using fruit size.
[0339]To calculate the benefit of harvesting earlier in the maturation of tomato fruit we constructed a conservative model that assumed 33 fruit are harvested from one plant over an average production period, and one dose of HN would be 50%g. The model was deemed conservative since one dose is likely to be less than 50 μg and plants that have fruit harvested four weeks post pollination would have reduced metabolic burden and would likely produce more fruit than plants that have fruit harvested at a later stage. These data suggest that by harvesting the fruit at four weeks post pollination, the time required for fruit to be ready for harvest is reduced by four weeks and there is a 286% increase in doses yielded.
[0340]Tomato therefore is capable of expressing large quantities of HN when harvested at an optimal time in fruit maturation.
EXAMPLE 14
Preparation of CHN-18 Master Seed
[0341]Master Seed Passage: Master Seed passage 2 was used for DNA extraction.DNA Extraction and PCR Amplification: DNA extraction was performed as described herein. PCR amplification for the HN and PAT gene expression cassettes were conducted separately. There was a 24 bp overlap between the PCR products of HN and PAT expression cassettes.
[0342]For HN gene cassette amplification, 50 μL PCR reaction contained 2.5 units of Takara Ex Tag DNA polymerase (Takara Shuzo Co, Shiga, Japan, catalog #RR001A), 0.2 μM of each primers (CHN01/CHN03), 5 μL of 10× reaction buffer containing MgCl2, 0.2 mM of each dNTP, and 200 ng of genomic DNA. The PCR was performed with a Gen Amp PCR 9700 system, manufactured by Applied Biosystem (Foster City, Calif.) at the following condition: 94° C. for 5 min for 1 cycle, 94° C. for 30 sec, 60° C. 30 sec and 72° C. for 3 min 30 sec for 40 cycles, 72° C. for 7 min.
[0343]For PAT gene cassette amplification, a 50 μL PCR reaction contained 2.5 units of Takara Ex Tag DNA polymerase (Takara Shuzo Co, Shiga, Japan, catalog #RR001A), 0.2 μM of each primers (CHN02/CHN04), 5 μL of 10× reaction buffer containing MgCl2, 0.2 mM of each dNTP, and 200 ng of genomic DNA were used. The PCR was performed with a Gen Amp PCR 9700 system, manufactured by Applied Biosystem (Foster City, Calif.) at the following condition: 94° C. for 5 min for 1 cycle, 94° C. for 30 sec, 56° C. 30 sec and 72° C. for 2 min 30 sec fro 40 cycles, 72° C. for 7 min.
Cloning of PCR Products: After agarose gel electrophoresis and visual observation, PCR products were purified using MiniElute PCR Purification Kit (Qiagen, Valencia, Calif. Catalog #28004) according to the manufacturer's protocol. Purified PCR products were cloned into pCR® II-TOPO vector using TOPO TA Cloning® kit (Invitrogen, Carlsbad, Calif., Catalog #051302) according to the manufacturer's protocol.Plasmid DNA Extraction Plasmid DNA was extracted using Qiaprep Spin Minprep kit (Qiagen, Valencia, Calif., catalog #27106) according to the manufacturer's protocol.DNA Sequencing and Analysis: Plasmid DNAs containing cloned PCR products were sent to Lark Technologies Inc (Houston, Tex.) for sequencing using ABI PRISM® BigDye® Primer Cycle Sequencing Kits (Applied Biosystem, Foster City, Calif.). DNA sequences were analyzed using Vector NT1 program (InforMax, Frederick, Md.).
Results
[0344]DNA sequences from the HN and PAT cassettes were assembled and compared with the sequences from a virtual plasmid map of pCHN. DNA sequences of all the genetic elements including the CsVMV promoter, HN and PAT coding sequences, vspB 3' UTR, and MAS 3' UTR were identical to the ones in the virtual plasmid map pCHN. Based on the virtual plasmid map pCHN, the PCR product including the whole gene insert in CHN-18 Master Seed using primer CHN01/CHN02 was 4757 bp. However, the actual cloned and sequenced PCR product including the whole gene insert in CHN-18 Master Seed was 4768 bp. By sequence comparison, 7 additional DNA bases were located in the junction region (poly cloning site) between the PAT coding sequences and MAS 3' UTR, another additional 4 DNA bases were located outside of the 3' end of MAS3' UTR (FIG. 43). The inconsistenct DNA base number between the virtual plasmid map and the actual sequencing data most likely occurred during the virtual creation of plasmid map pCHN. Open reading frame analysis of the entire insert sequence indicated there were only the expected HN and PAT open reading frames, and the 11 DNA bases did not result in any changes in the existing HN and PAT open reading frame and did not create any new opening reading frames.
CONCLUSIONS
[0345]By comparison with the virtual sequence from plasmid map pCHN, the actual DNA sequence of the whole gene insert in CHN-18 Master Seed was identical to what was expected except for extra 11 DNA bases outside all the genetic elements in the gene insert. These 11 DNA bases do not have any effect on the existing HN and PAT open reading frames.
[0346]The principles, preferred embodiments and modes of operation of the present invention have been described in the foregoing specification and examples. The invention that is intended to be protected herein, however, is not to be construed as limited to the particular forms specifically disclosed, since they are to be regarded as illustrative rather than restrictive. Variations and changes may be made by those skilled in the art without departing from the spirit and scope of the invention.
[0347]The practice of the present invention will employ, unless otherwise indicated, conventional techniques of molecular biology, cell biology, microbiology and recombinant DNA techniques, which are within the skill of the art. Such techniques are explained fully in the literature. See, e.g., Sambrook, Fritsch & Maniatis, 1989, Molecular Cloning: A Laboratory Manual, Second Edition; Oligonucleotide Synthesis (M. J. Gait, ed., 1984); Nucleic Acid Hybridization (B. D. Harnes & S. J. Higgins, eds., 1984); A Practical Guide to Molecular Cloning (B. Perbal, 1984); (Harlow, E. and Lane, D.) Using Antibodies: A Laboratory Manual (1999) Cold Spring Harbor Laboratory Press; and a series, Methods in Enzymology (Academic Press, Inc.); Short Protocols In Molecular Biology, (Ausubel et al., ed., 1995).
[0348]All patents, patent applications, and published references cited herein are hereby incorporated by reference in their entirety. While this invention has been particularly shown and described with references to preferred embodiments thereof, it will be understood by those skilled in the art that various changes in form and details may be made therein without departing from the scope of the invention encompassed by the appended claims.
Sequence CWU
1
2611760DNANewcastle disease virus 1ccatggaccg agcagtttca caagtggctc
ttgagaatga tgagagggaa gccaagaaca 60cttggaggct tatctttcgg atagccattc
tctttcttac tgttgtcacc ctagcaatct 120ctgttgcatc attactctat tctatgggag
caagcacccc ctcagactta gttggcatac 180ccacacgaat ctctagggct gaagagaaga
ttaccagtac cctaggctcc aaccaggatg 240ttgtggaccg aatctacaaa caagttgcac
ttgaaagtca acttgcatta ctcaacacag 300aaactaccat catgaatgca atcaccagcc
tatcctatca gatcaatggg gctgccaaca 360attcaggttg gggagcccca attcatgatc
cagactacat tggaggtatt ggcaaagaac 420tcattgtaga tgatgcttca gatgttacat
ctttctatcc ttcagctttc caggaacatc 480tgaacttcat tcctgcaccc acaactggga
gtgggtgcac tcggataccc tcatttgaca 540tgagtgctac acactattgc tatacacaca
atgtcattct atctggctgt cgtgaccatt 600ctcactctta tcagtactta gcacttggag
ttcttcgtac atctgctact ggtagagtgt 660tcttctcaac tcttcgcagt atcaatcttg
atgatacaca gaatcgcaaa agttgctctg 720tatctgctac acctttgggc tgtgatatgc
tatgcagtaa agtaacagaa actgaagaag 780aggactacaa ttctgcagtc cctacaagga
tggtgcatgg cagattgggt tttgatggtc 840aataccatga gaaagatttg gatgtcacta
cattgtttgg ggattgggta gctaactatc 900caggagttgg aggtggtagc ttcattgact
ccagagtctg gttctctgtc tatggtggtt 960tgaaacctaa cagtcctagt gatactgtgc
aagagggaaa gtatgttatc tacaagaggt 1020acaatgatac ttgtcctgat gagcaagact
atcagattcg aatggctaag tcatcataca 1080aaccaggaag atttggaggt aagaggatac
aacaagctat tctcagtatc aaggttagca 1140catcattggg agaagatcca gtccttactg
ttccaccaaa cactgtaaca ttgatgggag 1200ctgagggaag gattcttact gttggtacta
gccactttct ctatcaacgt ggaagttcct 1260actttagccc agcgttactg tatccaatga
ctgtgagcaa caagacagct acattacatt 1320caccatatac tttcaatgcc tttacaagac
ctggatcgat tccttgccaa gcttcagcta 1380gatgtccgaa ttcgtgtgtg actggagttt
acactgatcc ttaccctttg atcttctacc 1440gtaatcatac cttgagaggg gtgtttggaa
caatgttaga tggtgttcaa gctaggttga 1500atcctgcctc tgctgtgttt gattctacat
ccagatcaag gataaccaga gtttcctcta 1560gttctactaa ggcagcatac actacctcca
catgtttcaa agttgtaaag acgaacaaga 1620cctattgtct gagcatagct gagatttcta
acactctctt tggggaattc agaattgttc 1680cacttttggt ggagattctg aaagatgatg
gtgtacgtga agcaagatca ggttaagtct 1740tcggatccgg taccgagctc
17602577PRTNewcastle disease virus 2Met
Asp Arg Ala Val Ser Gln Val Ala Leu Glu Asn Asp Glu Arg Glu1
5 10 15Ala Lys Asn Thr Trp Arg Leu
Ile Phe Arg Ile Ala Ile Leu Phe Leu 20 25
30Thr Val Val Thr Leu Ala Ile Ser Val Ala Ser Leu Leu Tyr
Ser Met 35 40 45Gly Ala Ser Thr
Pro Ser Asp Leu Val Gly Ile Pro Thr Arg Ile Ser 50 55
60Arg Ala Glu Glu Lys Ile Thr Ser Thr Leu Gly Ser Asn
Gln Asp Val65 70 75
80Val Asp Arg Ile Tyr Lys Gln Val Ala Leu Glu Ser Pro Leu Ala Leu
85 90 95Leu Asn Thr Glu Thr Thr
Ile Met Asn Ala Ile Thr Ser Leu Ser Tyr 100
105 110Gln Ile Asn Gly Ala Ala Asn Asn Ser Gly Trp Gly
Ala Pro Ile His 115 120 125Asp Pro
Asp Tyr Ile Gly Gly Ile Gly Lys Glu Leu Ile Val Asp Asp 130
135 140Ala Ser Asp Val Thr Ser Phe Tyr Pro Ser Ala
Phe Gln Glu His Leu145 150 155
160Asn Phe Ile Pro Ala Pro Thr Thr Gly Ser Gly Cys Thr Arg Ile Pro
165 170 175Ser Phe Asp Met
Ser Ala Thr His Tyr Cys Tyr Thr His Asn Val Ile 180
185 190Leu Ser Gly Cys Arg Asp His Ser His Ser Tyr
Gln Tyr Leu Ala Leu 195 200 205Gly
Val Leu Arg Thr Ser Ala Thr Gly Arg Val Phe Phe Ser Thr Leu 210
215 220Arg Ser Ile Asn Leu Asp Asp Thr Gln Asn
Arg Lys Ser Cys Ser Val225 230 235
240Ser Ala Thr Pro Leu Gly Cys Asp Met Leu Cys Ser Lys Val Thr
Glu 245 250 255Thr Glu Glu
Glu Asp Tyr Asn Ser Ala Val Pro Thr Arg Met Val His 260
265 270Gly Arg Leu Gly Phe Asp Gly Gln Tyr His
Glu Lys Asp Leu Asp Val 275 280
285Thr Thr Leu Phe Gly Asp Trp Val Ala Asn Tyr Pro Gly Val Gly Gly 290
295 300Gly Ser Phe Ile Asp Ser Arg Val
Trp Phe Ser Val Tyr Gly Gly Leu305 310
315 320Lys Pro Asn Ser Pro Ser Asp Thr Val Gln Glu Gly
Lys Tyr Val Ile 325 330
335Tyr Lys Arg Tyr Asn Asp Thr Cys Pro Asp Glu Gln Asp Tyr Gln Ile
340 345 350Arg Met Ala Lys Ser Ser
Tyr Lys Pro Gly Arg Phe Gly Gly Lys Arg 355 360
365Ile Gln Gln Ala Ile Leu Ser Ile Lys Val Ser Thr Ser Leu
Gly Glu 370 375 380Asp Pro Val Leu Thr
Val Pro Pro Asn Thr Val Thr Leu Met Gly Ala385 390
395 400Glu Gly Arg Ile Leu Thr Val Gly Thr Ser
His Phe Leu Tyr Gln Arg 405 410
415Gly Ser Ser Tyr Phe Ser Pro Ala Leu Leu Tyr Pro Met Thr Val Ser
420 425 430Asn Lys Thr Ala Thr
Leu His Ser Pro Tyr Thr Phe Asn Ala Phe Thr 435
440 445Arg Pro Gly Ser Ile Pro Cys Gln Ala Ser Ala Arg
Cys Pro Asn Ser 450 455 460Cys Val Thr
Gly Val Tyr Thr Asp Pro Tyr Pro Leu Ile Phe Tyr Arg465
470 475 480Asn His Thr Leu Arg Gly Val
Phe Gly Thr Met Leu Asp Gly Val Gln 485
490 495Ala Arg Leu Asn Pro Ala Ser Ala Val Phe Asp Ser
Thr Ser Arg Ser 500 505 510Arg
Ile Thr Arg Val Ser Ser Ser Ser Thr Lys Ala Ala Tyr Thr Thr 515
520 525Ser Thr Cys Phe Lys Val Val Lys Thr
Asn Lys Thr Tyr Cys Leu Ser 530 535
540Ile Ala Glu Ile Ser Asn Thr Leu Phe Gly Glu Phe Arg Ile Val Pro545
550 555 560Leu Leu Val Glu
Ile Leu Lys Asp Asp Gly Val Arg Glu Ala Arg Ser 565
570 575Gly31647DNAAvian influenza virus
3gaccaaatct gcatcggtta tcatgcaaac aattcaacaa aacaagttga cacaatcatg
60gagaagaatg tgacggtcac acatgctcaa gatatactgg aaaaagagca caacgggaaa
120ctctgcagtc tcaaaggagt gaggcccctc attctgaagg attgcagtgt ggctggatgg
180cttcttggga acccaatgtg tgatgagttc ctaaatgtac cggaatggtc atatattgta
240gagaaggaca atccaaccaa tggcttatgt tatccgggag acttcaatga ttatgaagaa
300ctgaagtatt taatgagcaa cacaaaccat tttgagaaaa ttcaaataat ccctaggaac
360tcttggtcca atcatgatgc ctcatcagga gtgagctcag catgcccata caatggtagg
420tcttcctttt tcaggagtgt ggtgtggttg atcaagaaga gtaatgtata cccaacaata
480aagaggacct acaataacac caatgtagag gaccttctga tattgtgggg aatccatcac
540cctaatgatg cagcggaaca aacggaactc tatcagaact cgaacactta tgtgtctgta
600ggaacatcaa cactaaatca gaggtcaatt ccagaaatag ctaccaggcc caaagtgaat
660ggacaaagtg gaagaataga atttttctgg acaatactaa ggccgaacga tgcaatcagc
720tttgaaagta atgggaactt tatagctcct gaatatgcat acaagatagt taaaaaggga
780gattcagcaa tcatgagaag cgaactggag tatggcaact gtgataccaa atgtcagacc
840ccagtgggtg ctataaattc cagtatgcct tttcacaatg ttcatcccct taccattgga
900gagtgtccca aatatgtcaa atcagataaa ctggtccttg caacaggact gaggaacgtg
960cctcagagag aaacaagagg tctgtttgga gcaatagcag gattcataga aggggggtgg
1020caaggaatgg tagatggatg gtatggttac catcatagca acgagcaggg aagtggatat
1080gctgcagaca aagagtccac tcagaaagca atcgacggga tcaccaataa agtcaactca
1140atcattgaca aaatgaacac tcaattcgaa gccgttggga aagaattcaa caacttagaa
1200aggagaatag aaaatttgaa taagaaaatg gaagatggat ttctagatgt atggacttac
1260aatgcagaac ttctggtgct catggaaaat gaaagaactc tggatttcca tgattcatat
1320gtcaagaacc tatacgataa ggtccgactc cagctgagag ataatgcaaa agaattgggc
1380aatgggtgtt tggagttctc ccacaaatgt gacaatgaat gcatggaaag tgtgagaaac
1440ggaacgtatg actatccaca atactcagaa gaatcaaggc tgaacagaga ggaaatagat
1500ggagtcaaat tggagtcaat gggcacctat cagatactat caatttactc aacagtggcg
1560agttccctag cactggcaat catggtagct ggtctgtctt tttggatgtg ctccaatgga
1620tcattgcaat gcagaatttg catctag
16474548PRTAvian influenza virus 4Asp Gln Ile Cys Ile Gly Tyr His Ala Asn
Asn Ser Thr Lys Gln Val1 5 10
15Asp Thr Ile Met Glu Lys Asn Val Thr Val Thr His Ala Gln Asp Ile
20 25 30Leu Glu Lys Glu His Asn
Gly Lys Leu Cys Ser Leu Lys Gly Val Arg 35 40
45Pro Leu Ile Leu Lys Asp Cys Ser Val Ala Gly Trp Leu Leu
Gly Asn 50 55 60Pro Met Cys Asp Glu
Phe Leu Asn Val Pro Glu Trp Ser Tyr Ile Val65 70
75 80Glu Lys Asp Asn Pro Thr Asn Gly Leu Cys
Tyr Pro Gly Asp Phe Asn 85 90
95Asp Tyr Glu Glu Leu Lys Tyr Leu Met Ser Asn Thr Asn His Phe Glu
100 105 110Lys Ile Gln Ile Ile
Pro Arg Asn Ser Trp Ser Asn His Asp Ala Ser 115
120 125Ser Gly Val Ser Ser Ala Cys Pro Tyr Asn Gly Arg
Ser Ser Phe Phe 130 135 140Arg Ser Val
Val Trp Leu Ile Lys Lys Ser Asn Val Tyr Pro Thr Ile145
150 155 160Lys Arg Thr Tyr Asn Asn Thr
Asn Val Glu Asp Leu Leu Ile Leu Trp 165
170 175Gly Ile His His Pro Asn Asp Ala Ala Glu Gln Thr
Glu Leu Tyr Gln 180 185 190Asn
Ser Asn Thr Tyr Val Ser Val Gly Thr Ser Thr Leu Asn Gln Arg 195
200 205Ser Ile Pro Glu Ile Ala Thr Arg Pro
Lys Val Asn Gly Gln Ser Gly 210 215
220Arg Ile Glu Phe Phe Trp Thr Ile Leu Arg Pro Asn Asp Ala Ile Ser225
230 235 240Phe Glu Ser Asn
Gly Asn Phe Ile Ala Pro Glu Tyr Ala Tyr Lys Ile 245
250 255Val Lys Lys Gly Asp Ser Ala Ile Met Arg
Ser Glu Leu Glu Tyr Gly 260 265
270Asn Cys Asp Thr Lys Cys Gln Thr Pro Val Gly Ala Ile Asn Ser Ser
275 280 285Met Pro Phe His Asn Val His
Pro Leu Thr Ile Gly Glu Cys Pro Lys 290 295
300Tyr Val Lys Ser Asp Lys Leu Val Leu Ala Thr Gly Leu Arg Asn
Val305 310 315 320Pro Gln
Arg Glu Thr Arg Gly Leu Phe Gly Ala Ile Ala Gly Phe Ile
325 330 335Glu Gly Gly Trp Gln Gly Met
Val Asp Gly Trp Tyr Gly Tyr His His 340 345
350Ser Asn Glu Gln Gly Ser Gly Tyr Ala Ala Asp Lys Glu Ser
Thr Gln 355 360 365Lys Ala Ile Asp
Gly Ile Thr Asn Lys Val Asn Ser Ile Ile Asp Lys 370
375 380Met Asn Thr Gln Phe Glu Ala Val Gly Lys Glu Phe
Asn Asn Leu Glu385 390 395
400Arg Arg Ile Glu Asn Leu Asn Lys Lys Met Glu Asp Gly Phe Leu Asp
405 410 415Val Trp Thr Tyr Asn
Ala Glu Leu Leu Val Leu Met Glu Asn Glu Arg 420
425 430Thr Leu Asp Phe His Asp Ser Tyr Val Lys Asn Leu
Tyr Asp Lys Val 435 440 445Arg Leu
Gln Leu Arg Asp Asn Ala Lys Glu Leu Gly Asn Gly Cys Leu 450
455 460Glu Phe Ser His Lys Cys Asp Asn Glu Cys Met
Glu Ser Val Arg Asn465 470 475
480Gly Thr Tyr Asp Tyr Pro Gln Tyr Ser Glu Glu Ser Arg Leu Asn Arg
485 490 495Glu Glu Ile Asp
Gly Val Lys Leu Glu Ser Met Gly Thr Tyr Gln Ile 500
505 510Leu Ser Ile Tyr Ser Thr Val Ala Ser Ser Leu
Ala Leu Ala Ile Met 515 520 525Val
Ala Gly Leu Ser Phe Trp Met Cys Ser Asn Gly Ser Leu Gln Cys 530
535 540Arg Ile Cys Ile545528DNAArtificial
SequencePCR primer CVM-Asc 5atggcgcgcc agaaggtaat tatccaag
28624DNAArtificial SequencePCR primer CVM-Xho
6atctcgagcc atggtttgga tcca
24725DNAArtificial SequenceMutagenic primer GSS-Nco 7tgccatggtg
atgtgtggtc tacaa
25823DNAArtificial SequenceForward primer GSS-1.8F 8gatctgacaa gtcaagaaaa
ttg 23923DNAArtificial
SequenceMutagenic primer GSS-Xho 9agctcgagct gtgtgagtga gtg
231020DNAArtificial SequencePrimer HNa
10ccgagcagtt tcacaagtgg
201121DNAArtificial SequencePrimer HNb 11cctgatcttg cttcacgtac a
21124767DNAArtificial SequenceDNA
Sequence of the whole gene insert in CHN-18 Master Seed 12gaggtctaca
ggccaaattc gctcttagcc gtacaatatt actcaccgga tcggccgctt 60aattaagttt
aaaccctgca ggaaagcccg ggcaaaggcg cgccagaagg taattatcca 120gatgtagcat
caagaatcca atgtttacgg gaaaaactat ggaagtatta tgtgagctca 180gcaagaagca
gatcaatatg cggcacatat gcaacctatg ttcaaaaatg aagaatgtac 240agatacaaga
tcctatactg ccagaatacg aagaagaata cgtagaaatt gaaaaagaag 300aaccaggcga
agaaaagaat cttgaagacg taagcactga cgacaacaat gaaaagaaga 360agataaggtc
ggtgattgtg aaagagacat agaggacaca tgtaaggtgg aaaatgtaag 420ggcggaaagt
aaccttatca caaaggaatc ttatccccca ctacttatcc ttttatattt 480ttccgtgtca
tttttgccct tgagttttcc tatataagga accaagttcg gcatttgtga 540aaacaagaaa
aaatttggtg taagctattt tctttgaagt actgaggata caacttcaga 600gaaatttgta
agtttgtgga tccaaaccat ggaccgagca gtttcacaag tggctcttga 660gaatgatgag
agggaagcca agaacacttg gaggcttatc tttcggatag ccattctctt 720tcttactgtt
gtcaccctag caatctctgt tgcatcatta ctctattcta tgggagcaag 780caccccctca
gacttagttg gcatacccac acgaatctct agggctgaag agaagattac 840cagtacccta
ggctccaacc aggatgttgt ggaccgaatc tacaaacaag ttgcacttga 900aagtccactt
gcattactca acacagaaac taccatcatg aatgcaatca ccagcctatc 960ctatcagatc
aatggggctg ccaacaattc aggttgggga gccccaattc atgatccaga 1020ctacattgga
ggtattggca aagaactcat tgtagatgat gcttcagatg ttacatcttt 1080ctatccttca
gctttccagg aacatctgaa cttcattcct gcacccacaa ctgggagtgg 1140gtgcactcgg
ataccctcat ttgacatgag tgctacacac tattgctata cacacaatgt 1200cattctatct
ggctgtcgtg accattctca ctcttatcag tacttagcac ttggagttct 1260tcgtacatct
gctactggta gagtgttctt ctcaactctt cgcagtatca atcttgatga 1320tacacagaat
cgcaaaagtt gctctgtatc tgctacacct ttgggctgtg atatgctatg 1380cagtaaagta
acagaaactg aagaagagga ctacaattct gcagtcccta caaggatggt 1440gcatggcaga
ttgggttttg atggtcaata ccatgagaaa gatttggatg tcactacatt 1500gtttggggat
tgggtagcta actatccagg agttggaggt ggtagcttca ttgactccag 1560agtctggttc
tctgtctatg gtggtttgaa acctaacagt cctagtgata ctgtgcaaga 1620gggaaagtat
gttatctaca agaggtacaa tgatacttgt cctgatgagc aagactatca 1680gattcgaatg
gctaagtcat catacaaacc aggaagattt ggaggtaaga ggatacaaca 1740agctattctc
agtatcaagg ttagcacatc attgggagaa gatccagtcc ttactgttcc 1800accaaacact
gtaacattga tgggagctga gggaaggatt cttactgttg gtactagcca 1860ctttctctat
caacgtggaa gttcctactt tagcccagcg ttactgtatc caatgactgt 1920gagcaacaag
acagctacat tacattcacc atatactttc aatgccttta caagacctgg 1980atcgattcct
tgccaagctt cagctagatg tccgaattcg tgtgtgactg gagtttacac 2040tgatccttac
cctttgatct tctaccgtaa tcataccttg agaggggtgt ttggaacaat 2100gttagatggt
gttcaagcta ggttgaatcc tgcctctgct gtgtttgatt ctacatccag 2160atcaaggata
accagagttt cctctagttc tactaaggca gcatacacta cctccacatg 2220tttcaaagtt
gtaaagacga acaagaccta ttgtctgagc atagctgaga tttctaacac 2280tctctttggg
gaattcagaa ttgttccact tttggtggag attctgaaag atgatggtgt 2340acgtgaagca
agatcaggtt aagtcttcgg atccggtacc gagctctctc aacaatctag 2400ctagagtttg
ctcctatcta tatgtaataa ggtatgctga tatgcactat tcaaatagga 2460gcattagcta
tgtttgttaa tgtcacttta tgttatgtgg gtaagtcacc taagacactc 2520cacgtaccta
cttgttgtct cttacgcggc tttaataaat cttctgccct tgttccatat 2580ttactaatta
tccctttctt cactaaaaga aaattgttat cattaagtat tagtctttag 2640aacatatgag
gtctttaatt gggtaggttt tacaaattaa ctaatataaa atgtcataaa 2700atccacgtgg
ttaaacaaat gcagaaaatc gacgtcgtct attggaccga cagttgctat 2760taatataatg
ggccaccata gtagactgac aaataaatta cctgacaaca tcgtttcaca 2820aaaaaacaaa
cacaaaaagg gagtgcattt tccagggcat ttttgtaata aaaaacagat 2880aaaagggagt
gcaatagaaa tataggggtg tggaaatagt gatttgagca cgtcttgaag 2940cgaattcgcg
gccggccaga aggtaattat ccaagatgta gcatcaagaa tccaatgttt 3000acgggaaaaa
ctatggaagt attatgtgag ctcagcaaga agcagatcaa tatgcggcac 3060atatgcaacc
tatgttcaaa aatgaagaat gtacagatac aagatcctat actgccagaa 3120tacgaagaag
aatacgtaga aattgaaaaa gaagaaccag gcgaagaaaa gaatcttgaa 3180gacgtaagca
ctgacgacaa caatgaaaag aagaagataa ggtcggtgat tgtgaaagag 3240acatagagga
cacatgtaag gtggaaaatg taagggcgga aagtaacctt atcacaaagg 3300aatcttatcc
cccactactt atccttttat atttttccgt gtcatttttg cccttgagtt 3360ttcctatata
aggaaccaag ttcggcattt gtgaaaacaa gaaaaaattt ggtgtaagct 3420attttctttg
aagtactgag gatacaactt cagagaaatt tgtaagtttg tggatccaaa 3480ccatggcttc
tccggagagg agaccagttg agattaggcc agctacagca gctgatatgg 3540ccgcggtttg
tgatatcgtt aaccattaca ttgagacgtc tacagtgaac tttaggacag 3600agccacaaac
accacaagag tggattgatg atctagagag gttgcaagat agataccctt 3660ggttggttgc
tgaggttgag ggtgttgtgg ctggtattgc ttacgctggg ccctggaagg 3720ctaggaacgc
ttacgattgg acagttgaga gtactgttta cgtgtcacat aggcatcaaa 3780ggttgggcct
aggatccaca ttgtacacac atttgcttaa gtctatggag gcgcaaggtt 3840ttaagtctgt
ggttgctgtt ataggccttc caaacgatcc atctgttagg ttgcatgagg 3900ctttgggata
cacagcccgg ggtacattgc gcgcagctgg atacaagcat ggtggatggc 3960atgatgttgg
tttttggcaa agggattttg agttgccagc tcctccaagg ccagctaggc 4020cagttaccca
gatctgaggt accctgagct cggtcacctg tccaacagtc tcagggttaa 4080tgtctatgta
tcttaaataa tgttgtcggt attttgtaat ctcatataga ttttcactgt 4140gcgacgcaaa
aatattaaat aaatattatt attatctacg ttttgattga gatatcatca 4200atattataat
aaaaatatcc attaaacacg atttgataca aatgacagtc aataatctga 4260tttgaatatt
tattaattgt aacgaattac ataaagatcg aatagaaaat actgcactgc 4320aaatgaaaat
taacacatac taataaatgc gtcaaatatc tttgccaaga tcaagcggag 4380tgagggcctc
atatccggtc tcagttacaa gcacggtatc cccgaagcgc gctccaccaa 4440tgccctcgac
atagatgccg ggctcgacgc tgaggacatt gcctaccttg agcatggtct 4500cagcgccggc
tttaagctca atcccatccc aatctgaata tcctatcccg cgcccagtcc 4560ggtgtaagaa
cgggtctgtc catccacctc tgttgcggcc aattctgatc tggcccccat 4620ttggacgtga
atgtagacac gtcgatataa agatttccga attagaataa tttgtttatt 4680gctttcgcct
ataaatacga cggatcgtaa tttgtcgttt tatcaaaatg tactttcatt 4740ttataataac
gctgcggaca tctacat
476713681DNAHepatitis B virus 13atggagaaca tcacatcagg attcctagga
cccctgctcg tgttacaggc ggggtttttc 60ttgttgacaa gaatcctcac aataccgcag
agtctagact cgtggtggac ttctctcaat 120tttctagggg gatcacccgt gtgtcttggc
caaaattcgc agtccccaac ctccaatcac 180tcaccaacct cctgtcctcc aatctgtcct
ggttatcgct ggatgtgtct gcggcgtttt 240atcatattcc tcttcatcct gctgctatgc
ctcatcttct tattggttct tctggattat 300caaggtatgt tgcccgtttg tcctctaatt
ccaggatcaa caacaaccag tacgggacca 360tgcaaaacct gcacgactcc tgctcaaggg
aactctatgt ttccctcatg ttgctgtaca 420aaacctacgg atgggaattg cacctgtatt
cccatcccat cgtcctgggc tttcgcaaaa 480tacctatggg agtgggcctc agtccgtttc
tcttggctca gtttactagt gccatttgtt 540cagtggttcg tagggctttc ccccactgtt
tggctttcag ctatatggat gatgtggtat 600tgggggccaa gtctgtacag catcgtgagt
ccctttatac cgctgttacc aattttcttt 660tgtctctggg tatacattta a
68114226PRTHepatitis B virus 14Met Glu
Asn Ile Thr Ser Gly Phe Leu Gly Pro Leu Leu Val Leu Gln1 5
10 15Ala Gly Phe Phe Leu Leu Thr Arg
Ile Leu Thr Ile Pro Gln Ser Leu 20 25
30Asp Ser Trp Trp Thr Ser Leu Asn Phe Leu Gly Gly Ser Pro Val
Cys 35 40 45Leu Gly Gln Asn Ser
Gln Ser Pro Thr Ser Asn His Ser Pro Thr Ser 50 55
60Cys Pro Pro Ile Cys Pro Gly Tyr Arg Trp Met Cys Leu Arg
Arg Phe65 70 75 80Ile
Ile Phe Leu Phe Ile Leu Leu Leu Cys Leu Ile Phe Leu Leu Val
85 90 95Leu Leu Asp Tyr Gln Gly Met
Leu Pro Val Cys Pro Leu Ile Pro Gly 100 105
110Ser Thr Thr Thr Ser Thr Gly Pro Cys Lys Thr Cys Thr Thr
Pro Ala 115 120 125Gln Gly Asn Ser
Met Phe Pro Ser Cys Cys Cys Thr Lys Pro Thr Asp 130
135 140Gly Asn Cys Thr Cys Ile Pro Ile Pro Ser Ser Trp
Ala Phe Ala Lys145 150 155
160Tyr Leu Trp Glu Trp Ala Ser Val Arg Phe Ser Trp Leu Ser Leu Leu
165 170 175Val Pro Phe Val Gln
Trp Phe Val Gly Leu Ser Pro Thr Val Trp Leu 180
185 190Ser Ala Ile Trp Met Met Trp Tyr Trp Gly Pro Ser
Leu Tyr Ser Ile 195 200 205Val Ser
Pro Phe Ile Pro Leu Leu Pro Ile Phe Phe Cys Leu Trp Val 210
215 220Tyr Ile225151467DNAHomo sapiens 15cgcagttgtg
cacaccccag cccaggctct ggcaggctgc caggcggggc tcgccgcctt 60cgcagtgcac
gttgtccagc aggatgtgtc cggtgccata gccgaagaag gcgttggtgg 120tggcggccat
ggccgccccg cagcccagct gacgacagac cacagcggca tccggcagcc 180cccagtcgtc
gtcacacacg gtgccccaca ggccactgtg caggatctcc actcggccct 240gacacagggt
ggagccagcc atcctgagac gtccgtacag cccgtactgg tggagtgtca 300ggaggccact
ctgatggtca tggtcagcaa agaccttttt ggcaccggga agctcatcag 360ggctgctgac
ctcaccttgg gcccagaggc ctgtgagcct ctggtctcca tggacacaga 420agatgtggtc
aggtttgagg ttggactcca cgagtgtggc aacagcatgc aggtaactga 480cgatgccctg
gtgtacagca ccttcctgct ccatgacccc cgccccgtgg gaaacctgtc 540catcgtgagg
actaaccgcg cagagattcc catcgagtgc cgctacccca ggcagggcaa 600tgtgagcagc
caggccatcc tgcccacctg gttgcccttc aggaccacgg tgttctcaga 660ggagaagctg
actttctctc tgcgtctgat ggaggagaac tggaacgctg agaagaggtc 720ccccaccttc
cacctgggag atgcagccca cctccaggca gaaatccaca ctggcagcca 780cgtgccactg
cggttgtttg tggaccactg cgtggccaca ccgacaccag accagaatgc 840ctccccttat
cacaccatcg tggacttcca tggctgtctt gtcgacggtc tcactgatgc 900ctcttctgca
ttcaaagttc ctcgacccgg gccagataca ctccagttca cagtggatgt 960cttccacttt
gctaatgact ccagaaacat gatatacatc acctgccacc tgaaggtcac 1020cctagctgag
caggacccag atgaactcaa caaggcctgt tccttcagca agccttccaa 1080cagctggttc
ccagtggaag gctcggctga catctgtcaa tgctgtaaca aaggtgactg 1140tggcactcca
agccattcca ggaggcagcc tcatgtcatg agccagtggt ccaggtctgc 1200ttcccgtaac
cgcaggcatg tgacagaaga agcagatgtc accgtggggc cactgatctt 1260cctggacagg
aggggtgacc atgaagtaga gcagtgggct ttgccttctg acacctcagt 1320ggtgctgctg
ggcgtaggcc tggctgtggt ggtgtccctg actctgactg ctgttatcct 1380ggttctcacc
aggaggtgtc gcactgcctc ccaccctgtg tctgcttccg aataaaagaa 1440gaaagcaata
aaaaaaaaaa aaaaaaa 146716373PRTHomo
sapiens 16Met Val Met Val Ser Lys Asp Leu Phe Gly Thr Gly Lys Leu Ile
Arg1 5 10 15Ala Ala Asp
Leu Thr Leu Gly Pro Glu Ala Cys Glu Pro Leu Val Ser 20
25 30Met Asp Thr Glu Asp Val Val Arg Phe Glu
Val Gly Leu His Glu Cys 35 40
45Gly Asn Ser Met Gln Val Thr Asp Asp Ala Leu Val Tyr Ser Thr Phe 50
55 60Leu Leu His Asp Pro Arg Pro Val Gly
Asn Leu Ser Ile Val Arg Thr65 70 75
80Asn Arg Ala Glu Ile Pro Ile Glu Cys Arg Tyr Pro Arg Gln
Gly Asn 85 90 95Val Ser
Ser Gln Ala Ile Leu Pro Thr Trp Leu Pro Phe Arg Thr Thr 100
105 110Val Phe Ser Glu Glu Lys Leu Thr Phe
Ser Leu Arg Leu Met Glu Glu 115 120
125Asn Trp Asn Ala Glu Lys Arg Ser Pro Thr Phe His Leu Gly Asp Ala
130 135 140Ala His Leu Gln Ala Glu Ile
His Thr Gly Ser His Val Pro Leu Arg145 150
155 160Leu Phe Val Asp His Cys Val Ala Thr Pro Thr Pro
Asp Gln Asn Ala 165 170
175Ser Pro Tyr His Thr Ile Val Asp Phe His Gly Cys Leu Val Asp Gly
180 185 190Leu Thr Asp Ala Ser Ser
Ala Phe Lys Val Pro Arg Pro Gly Pro Asp 195 200
205Thr Leu Gln Phe Thr Val Asp Val Phe His Phe Ala Asn Asp
Ser Arg 210 215 220Asn Met Ile Tyr Ile
Thr Cys His Leu Lys Val Thr Leu Ala Glu Gln225 230
235 240Asp Pro Asp Glu Leu Asn Lys Ala Cys Ser
Phe Ser Lys Pro Ser Asn 245 250
255Ser Trp Phe Pro Val Glu Gly Ser Ala Asp Ile Cys Gln Cys Cys Asn
260 265 270Lys Gly Asp Cys Gly
Thr Pro Ser His Ser Arg Arg Gln Pro His Val 275
280 285Met Ser Gln Trp Ser Arg Ser Ala Ser Arg Asn Arg
Arg His Val Thr 290 295 300Glu Glu Ala
Asp Val Thr Val Gly Pro Leu Ile Phe Leu Asp Arg Arg305
310 315 320Gly Asp His Glu Val Glu Gln
Trp Ala Leu Pro Ser Asp Thr Ser Val 325
330 335Val Leu Leu Gly Val Gly Leu Ala Val Val Val Ser
Leu Thr Leu Thr 340 345 350Ala
Val Ile Leu Val Leu Thr Arg Arg Cys Arg Thr Ala Ser His Pro 355
360 365Val Ser Ala Ser Glu
370171740DNAAvian influenza virus 17atggaaagaa tagtgattgc ccttgcaata
atcaacattg tcaaaggtga ccaaatctgc 60attggttatc atgcaaacaa ttcaacagag
caggttgaca caatcatgga gaagaatgtg 120acggtcacac atgctcagga catactggaa
aaagagcaca atgggaaact ctgcagtctt 180aaaggagtga ggcccctcat tctgaaggat
tgcagtgtcg ctgggtggct tcttggaaac 240ccaatgtgtg atgaattcct gaatgtaccg
gaatggtcat acattgtgga aaaagataat 300ccagtcaatg gcctgtgcta tccaggagac
ttcaacgatt atgaagaact gaagcattta 360atgagcagca caaaccattt tgagaaaatt
cagataattc ctaggaactc ttggtccacc 420catgatgcct catcaggagt gagctcagca
tgcccataca atggtaggtc ttcctttttc 480aggaatgtag tgtggttgat caagaagaat
aatgcgtacc caacaataaa gaggacctac 540aacaacacca atgtagaaga ccttttaata
ttatggggaa tccaccaccc taatgatgca 600gcagaacaaa caaaactcta ccagaactcg
aacacttatg tgtctgtagg aacatcaaca 660ctgaatcaga ggtcaatccc agaaatagcc
accagaccca aagtgaacgg acaaagtgga 720agaatggaat ttttttggac aatactaagg
ccgaacgatg caatcagctt tgaaagtaat 780gggaacttta tagctcctga atatgcgtac
aagattgtta aaaaaggaga ttcagcaatc 840atgaaaagtg aactggagta tggtaactgt
gataccaaat gtcagacccc agtgggtgct 900ataaattcca gtatgccttt ccacaatgtt
catcccctta ccattgggga gtgccccaag 960tatgtcaaat cggacaaact ggtccttgca
acaggactaa gaaacgtacc ccaaagagaa 1020acaagaggcc tatttggagc aatagcagga
ttcatagaag gaggatggca aggaatggta 1080gatggatggt atggatacca tcatagcaat
gagcagggaa gtggatatgc tgcagacaaa 1140gaatctaccc agaaagcaat cgatgggatc
accaataaag taaactcaat cattgacaaa 1200atgaacactc aattcgaagc cgttgggaaa
gaattcaaca acctagaaag gagaatagaa 1260aatttgaata agaaaatgga agatgggttt
ttagatgtat ggacttacaa tgcagaactt 1320ctagtgctca tggaaaacga aagaactctg
gatttccatg attcaaatgt caagaactta 1380tacgataagg tccgactcca gctgagagac
aatgcaaaag aattaggcaa cgggtgcttt 1440gaattctacc acaagtgtga caatgaatgc
atggaaagtg tgagaaatgg aacgtatgac 1500tatccacaat actcagaaga atcaagactg
aacagggagg aaatagacgg agtcaaattg 1560gaatcaatgg gcacttatca gatactatca
atctactcaa cagtggcgag ttccctagca 1620ctggcaatca tggtagctgg tctatctttt
tggatgtgct ccaatggatc attgcagtgc 1680agaatttgca tctagaattg tgagttcaga
ttataattaa aaacacccta gtttctactg 174018564PRTAvian influenza virus
18Met Glu Arg Ile Val Ile Ala Leu Ala Ile Ile Asn Ile Val Lys Gly1
5 10 15Asp Gln Ile Cys Ile Gly
Tyr His Ala Asn Asn Ser Thr Glu Gln Val 20 25
30Asp Thr Ile Met Glu Lys Asn Val Thr Val Thr His Ala
Gln Asp Ile 35 40 45Leu Glu Lys
Glu His Asn Gly Lys Leu Cys Ser Leu Lys Gly Val Arg 50
55 60Pro Leu Ile Leu Lys Asp Cys Ser Val Ala Gly Trp
Leu Leu Gly Asn65 70 75
80Pro Met Cys Asp Glu Phe Leu Asn Val Pro Glu Trp Ser Tyr Ile Val
85 90 95Glu Lys Asp Asn Pro Val
Asn Gly Leu Cys Tyr Pro Gly Asp Phe Asn 100
105 110Asp Tyr Glu Glu Leu Lys His Leu Met Ser Ser Thr
Asn His Phe Glu 115 120 125Lys Ile
Gln Ile Ile Pro Arg Asn Ser Trp Ser Thr His Asp Ala Ser 130
135 140Ser Gly Val Ser Ser Ala Cys Pro Tyr Asn Gly
Arg Ser Ser Phe Phe145 150 155
160Arg Asn Val Val Trp Leu Ile Lys Lys Asn Asn Ala Tyr Pro Thr Ile
165 170 175Lys Arg Thr Tyr
Asn Asn Thr Asn Val Glu Asp Leu Leu Ile Leu Trp 180
185 190Gly Ile His His Pro Asn Asp Ala Ala Glu Gln
Thr Lys Leu Tyr Gln 195 200 205Asn
Ser Asn Thr Tyr Val Ser Val Gly Thr Ser Thr Leu Asn Gln Arg 210
215 220Ser Ile Pro Glu Ile Ala Thr Arg Pro Lys
Val Asn Gly Gln Ser Gly225 230 235
240Arg Met Glu Phe Phe Trp Thr Ile Leu Arg Pro Asn Asp Ala Ile
Ser 245 250 255Phe Glu Ser
Asn Gly Asn Phe Ile Ala Pro Glu Tyr Ala Tyr Lys Ile 260
265 270Val Lys Lys Gly Asp Ser Ala Ile Met Lys
Ser Glu Leu Glu Tyr Gly 275 280
285Asn Cys Asp Thr Lys Cys Gln Thr Pro Val Gly Ala Ile Asn Ser Ser 290
295 300Met Pro Phe His Asn Val His Pro
Leu Thr Ile Gly Glu Cys Pro Lys305 310
315 320Tyr Val Lys Ser Asp Lys Leu Val Leu Ala Thr Gly
Leu Arg Asn Val 325 330
335Pro Gln Arg Glu Thr Arg Gly Leu Phe Gly Ala Ile Ala Gly Phe Ile
340 345 350Glu Gly Gly Trp Gln Gly
Met Val Asp Gly Trp Tyr Gly Tyr His His 355 360
365Ser Asn Glu Gln Gly Ser Gly Tyr Ala Ala Asp Lys Glu Ser
Thr Gln 370 375 380Lys Ala Ile Asp Gly
Ile Thr Asn Lys Val Asn Ser Ile Ile Asp Lys385 390
395 400Met Asn Thr Gln Phe Glu Ala Val Gly Lys
Glu Phe Asn Asn Leu Glu 405 410
415Arg Arg Ile Glu Asn Leu Asn Lys Lys Met Glu Asp Gly Phe Leu Asp
420 425 430Val Trp Thr Tyr Asn
Ala Glu Leu Leu Val Leu Met Glu Asn Glu Arg 435
440 445Thr Leu Asp Phe His Asp Ser Asn Val Lys Asn Leu
Tyr Asp Lys Val 450 455 460Arg Leu Gln
Leu Arg Asp Asn Ala Lys Glu Leu Gly Asn Gly Cys Phe465
470 475 480Glu Phe Tyr His Lys Cys Asp
Asn Glu Cys Met Glu Ser Val Arg Asn 485
490 495Gly Thr Tyr Asp Tyr Pro Gln Tyr Ser Glu Glu Ser
Arg Leu Asn Arg 500 505 510Glu
Glu Ile Asp Gly Val Lys Leu Glu Ser Met Gly Thr Tyr Gln Ile 515
520 525Leu Ser Ile Tyr Ser Thr Val Ala Ser
Ser Leu Ala Leu Ala Ile Met 530 535
540Val Ala Gly Leu Ser Phe Trp Met Cys Ser Asn Gly Ser Leu Gln Cys545
550 555 560Arg Ile Cys
Ile191734DNANewcastle disease virus 19atggaccgcg ccgttagcca agttgcgtta
gagaatgatg aaagagaggc aaaaaataca 60tggcgcttga tattccggat tgcaatctta
ttcttaacag tagtgacctt ggctatatct 120gtagcctccc ttttatatag catgggggct
agcacaccta gcgatcttgt aggcataccg 180actaggattt ccagggcaga agaaaagatt
acatctacac ttggttccaa tcaagatgta 240gtagatagga tatataagca agtggccctt
gagtctccgt tggcattgtt aaaaactgag 300accacaatta tgaacgcaat aacatctctc
tcttatcaga ttaatggagc tgcaaacaac 360agcgggtggg gggcacctat ccatgaccca
gattatatag gggggatagg caaagaactc 420attgtagatg atgctagtga tgtcacatca
ttctatccct ctgcatttca agaacatctg 480aattttatcc cggcgcctac tacaggatca
ggttgcactc gaataccctc atttgacatg 540agtgctaccc attactgcta cacccataat
gtaatattgt ctggatgcag agatcactca 600cattcatatc agtatttagc acttggtgtg
ctccggacat ctgcaacagg gggggtattc 660ttttctactc tgcgttccat caacctggac
gacacccaaa atcggaagtc ttgcagtgtg 720agtgcaactc ccctgggttg tgatatgctg
tgctcgaaag tcacggagac agaggaagaa 780gattataact cagctgtccc tacgcggatg
gtacatggga ggttagggtt cgacggccag 840taccacgaaa aggacctaga tgtcacaaca
ttattcgggg actgggtggc caactaccca 900ggagtagggg gtggatcttt tattgacagc
cgcgtatggt tctcagttta cggagggtta 960aaacccaatt cacccagtga cactgtacag
gaagggaaat atgtgatata caagcaatac 1020aatgacacat gcccagatga gcaagactac
cagattcgaa tggccaagtc ttcgtataag 1080cctggacggt ttggtgggaa acgcatacag
caggctatct tatctatcaa ggtgtcaaca 1140tccttaggcg aagacccggt actgactgta
ccgcccaaca cagtcacact catgggggcc 1200gaaggcagaa ttctcacagt agggacatct
catttcttgt atcaacgagg gtcatcatac 1260ttctctcccg cgttattata tcctatgaca
gtcagcaaca aaacagccac tcttcatagt 1320ccttatacat tcaatgcctt cactcggcca
ggtagtatcc cttgccaggc ttcagcaaga 1380tgccccaacc cgtgtgttac tggagtctat
acagatccat atcccctaat cttctataga 1440aaccacacct tgcgaggggt attcgggaca
atgcttgatg gtgtacaagc aagacttaac 1500cctgcgtctg cagtattcga tagcacatcc
cgcagtcgca ttactcgagt gagttcaagc 1560agtaccaaag cagcatacac aacatcaact
tgttttaaag tggtcaagac taataagacc 1620tattgtctca gcattgctga aatatctaat
actctcttcg gagaattcag aatcgtcccg 1680ttactagttg agatcctcaa agatgacggg
gttagagaag ccaggtctgg ctag 173420577PRTNewcastle disease virus
20Met Asp Arg Ala Val Ser Gln Val Ala Leu Glu Asn Asp Glu Arg Glu1
5 10 15Ala Lys Asn Thr Trp Arg
Leu Ile Phe Arg Ile Ala Ile Leu Phe Leu 20 25
30Thr Val Val Thr Leu Ala Ile Ser Val Ala Ser Leu Leu
Tyr Ser Met 35 40 45Gly Ala Ser
Thr Pro Ser Asp Leu Val Gly Ile Pro Thr Arg Ile Ser 50
55 60Arg Ala Glu Glu Lys Ile Thr Ser Thr Leu Gly Ser
Asn Gln Asp Val65 70 75
80Val Asp Arg Ile Tyr Lys Gln Val Ala Leu Glu Ser Pro Leu Ala Leu
85 90 95Leu Lys Thr Glu Thr Thr
Ile Met Asn Ala Ile Thr Ser Leu Ser Tyr 100
105 110Gln Ile Asn Gly Ala Ala Asn Asn Ser Gly Trp Gly
Ala Pro Ile His 115 120 125Asp Pro
Asp Tyr Ile Gly Gly Ile Gly Lys Glu Leu Ile Val Asp Asp 130
135 140Ala Ser Asp Val Thr Ser Phe Tyr Pro Ser Ala
Phe Gln Glu His Leu145 150 155
160Asn Phe Ile Pro Ala Pro Thr Thr Gly Ser Gly Cys Thr Arg Ile Pro
165 170 175Ser Phe Asp Met
Ser Ala Thr His Tyr Cys Tyr Thr His Asn Val Ile 180
185 190Leu Ser Gly Cys Arg Asp His Ser His Ser Tyr
Gln Tyr Leu Ala Leu 195 200 205Gly
Val Leu Arg Thr Ser Ala Thr Gly Gly Val Phe Phe Ser Thr Leu 210
215 220Arg Ser Ile Asn Leu Asp Asp Thr Gln Asn
Arg Lys Ser Cys Ser Val225 230 235
240Ser Ala Thr Pro Leu Gly Cys Asp Met Leu Cys Ser Lys Val Thr
Glu 245 250 255Thr Glu Glu
Glu Asp Tyr Asn Ser Ala Val Pro Thr Arg Met Val His 260
265 270Gly Arg Leu Gly Phe Asp Gly Gln Tyr His
Glu Lys Asp Leu Asp Val 275 280
285Thr Thr Leu Phe Gly Asp Trp Val Ala Asn Tyr Pro Gly Val Gly Gly 290
295 300Gly Ser Phe Ile Asp Ser Arg Val
Trp Phe Ser Val Tyr Gly Gly Leu305 310
315 320Lys Pro Asn Ser Pro Ser Asp Thr Val Gln Glu Gly
Lys Tyr Val Ile 325 330
335Tyr Lys Gln Tyr Asn Asp Thr Cys Pro Asp Glu Gln Asp Tyr Gln Ile
340 345 350Arg Met Ala Lys Ser Ser
Tyr Lys Pro Gly Arg Phe Gly Gly Lys Arg 355 360
365Ile Gln Gln Ala Ile Leu Ser Ile Lys Val Ser Thr Ser Leu
Gly Glu 370 375 380Asp Pro Val Leu Thr
Val Pro Pro Asn Thr Val Thr Leu Met Gly Ala385 390
395 400Glu Gly Arg Ile Leu Thr Val Gly Thr Ser
His Phe Leu Tyr Gln Arg 405 410
415Gly Ser Ser Tyr Phe Ser Pro Ala Leu Leu Tyr Pro Met Thr Val Ser
420 425 430Asn Lys Thr Ala Thr
Leu His Ser Pro Tyr Thr Phe Asn Ala Phe Thr 435
440 445Arg Pro Gly Ser Ile Pro Cys Gln Ala Ser Ala Arg
Cys Pro Asn Pro 450 455 460Cys Val Thr
Gly Val Tyr Thr Asp Pro Tyr Pro Leu Ile Phe Tyr Arg465
470 475 480Asn His Thr Leu Arg Gly Val
Phe Gly Thr Met Leu Asp Gly Val Gln 485
490 495Ala Arg Leu Asn Pro Ala Ser Ala Val Phe Asp Ser
Thr Ser Arg Ser 500 505 510Arg
Ile Thr Arg Val Ser Ser Ser Ser Thr Lys Ala Ala Tyr Thr Thr 515
520 525Ser Thr Cys Phe Lys Val Val Lys Thr
Asn Lys Thr Tyr Cys Leu Ser 530 535
540Ile Ala Glu Ile Ser Asn Thr Leu Phe Gly Glu Phe Arg Ile Val Pro545
550 555 560Leu Leu Val Glu
Ile Leu Lys Asp Asp Gly Val Arg Glu Ala Arg Ser 565
570 575Gly211377DNAGallus gallus 21atgctgggtg
agctggaagc aggcgggatg caaggcggcc gcgtggtgct ggggctcctg 60tgctgcttgg
tggccggggt gggctcctac acgccctggg acatctcctg ggcagcacga 120ggggacccct
ctgcctggtc ctggggggcc gaggcgcact cacgggccgt ggccggctcg 180cacccggtgg
ccgtgcagtg ccaagaggcg cagctggtgg tgacggtgca cagggacctc 240ttcgggaccg
ggcgtctcat caacgctgct gacctgactc tgggcccggc tgcctgcaag 300cactcctcgc
tcaacgccgc acacaacacc gtcaccttcg ccgccggcct ccacgagtgc 360ggcagcgtcg
tgcaggtgac gccagacacc ctcatctacc gcacgctcat caactacgac 420cccagccctg
ctagcaaccc cgtcatcatc cgcaccaacc ctgctgtcat ccccatcgag 480tgccactacc
ccaggaggga gaacgtgagc agcaatgcca tccggcccac ctggtccccc 540ttcaactccg
cactgtcagc cgaggagagg ctggtgttct ccctgcgcct catgagtgat 600gactggagca
cagagagacc cttcaccggc ttccagctgg gcgacatcct caacatccag 660gccgaggtca
gcactgagaa ccatgtgccc ctgcggctct ttgtggacag ctgtgtggct 720gccctgagcc
ctgacggtga ctcctcgccc cactacgcca tcattgactt caacgggtgc 780ttagtggatg
ggagagtgga tgatactagc tctgccttca tcacaccccg gccacgggag 840gatgtgctga
ggttcaggat cgatgtcttc aggtttgcgg gggacaacag gaacctgatc 900tacatcacct
gccacctgaa ggtgacccca gcagaccaag gcccagaccc tcagaacaag 960gcttgctcct
tcaataaagc cagaaacacc tgggtgccag tggaaggcag ccgggatgtc 1020tgcaactgct
gtgagacagg caactgcgag ccgcctgcgc tctcccggag gctcaacccc 1080atggagagat
ggcagagccg ccgcttccgt cgtgatgccg ggaaagaggt tgcagctgat 1140gtggtcattg
gccccgtgtt gctctcggcg gacccgggag ctgtgggaca gcaggaggag 1200ggtggtgacg
gtgcggcggt gatggtgccc agcgtgggga cggggctggt gtgcgtggcc 1260gtggctgtag
ctctggctgc cgttggggtg gctgtatgta ttgcacgcaa gggatgcacc 1320cgaacctcaa
ctgcggtgtg agtgcagggc gagccgtgaa taaagcctgg aaaggcc
137722446PRTGallus gallus 22Met Leu Gly Glu Leu Glu Ala Gly Gly Met Gln
Gly Gly Arg Val Val1 5 10
15Leu Gly Leu Leu Cys Cys Leu Val Ala Gly Val Gly Ser Tyr Thr Pro
20 25 30Trp Asp Ile Ser Trp Ala Ala
Arg Gly Asp Pro Ser Ala Trp Ser Trp 35 40
45Gly Ala Glu Ala His Ser Arg Ala Val Ala Gly Ser His Pro Val
Ala 50 55 60Val Gln Cys Gln Glu Ala
Gln Leu Val Val Thr Val His Arg Asp Leu65 70
75 80Phe Gly Thr Gly Arg Leu Ile Asn Ala Ala Asp
Leu Thr Leu Gly Pro 85 90
95Ala Ala Cys Lys His Ser Ser Leu Asn Ala Ala His Asn Thr Val Thr
100 105 110Phe Ala Ala Gly Leu His
Glu Cys Gly Ser Val Val Gln Val Thr Pro 115 120
125Asp Thr Leu Ile Tyr Arg Thr Leu Ile Asn Tyr Asp Pro Ser
Pro Ala 130 135 140Ser Asn Pro Val Ile
Ile Arg Thr Asn Pro Ala Val Ile Pro Ile Glu145 150
155 160Cys His Tyr Pro Arg Arg Glu Asn Val Ser
Ser Asn Ala Ile Arg Pro 165 170
175Thr Trp Ser Pro Phe Asn Ser Ala Leu Ser Ala Glu Glu Arg Leu Val
180 185 190Phe Ser Leu Arg Leu
Met Ser Asp Asp Trp Ser Thr Glu Arg Pro Phe 195
200 205Thr Gly Phe Gln Leu Gly Asp Ile Leu Asn Ile Gln
Ala Glu Val Ser 210 215 220Thr Glu Asn
His Val Pro Leu Arg Leu Phe Val Asp Ser Cys Val Ala225
230 235 240Ala Leu Ser Pro Asp Gly Asp
Ser Ser Pro His Tyr Ala Ile Ile Asp 245
250 255Phe Asn Gly Cys Leu Val Asp Gly Arg Val Asp Asp
Thr Ser Ser Ala 260 265 270Phe
Ile Thr Pro Arg Pro Arg Glu Asp Val Leu Arg Phe Arg Ile Asp 275
280 285Val Phe Arg Phe Ala Gly Asp Asn Arg
Asn Leu Ile Tyr Ile Thr Cys 290 295
300His Leu Lys Val Thr Pro Ala Asp Gln Gly Pro Asp Pro Gln Asn Lys305
310 315 320Ala Cys Ser Phe
Asn Lys Ala Arg Asn Thr Trp Val Pro Val Glu Gly 325
330 335Ser Arg Asp Val Cys Asn Cys Cys Glu Thr
Gly Asn Cys Glu Pro Pro 340 345
350Ala Leu Ser Arg Arg Leu Asn Pro Met Glu Arg Trp Gln Ser Arg Arg
355 360 365Phe Arg Arg Asp Ala Gly Lys
Glu Val Ala Ala Asp Val Val Ile Gly 370 375
380Pro Val Leu Leu Ser Ala Asp Pro Gly Ala Val Gly Gln Gln Glu
Glu385 390 395 400Gly Gly
Asp Gly Ala Ala Val Met Val Pro Ser Val Gly Thr Gly Leu
405 410 415Val Cys Val Ala Val Ala Val
Ala Leu Ala Ala Val Gly Val Ala Val 420 425
430Cys Ile Ala Arg Lys Gly Cys Thr Arg Thr Ser Thr Ala Val
435 440 445233024DNAAnas sp.
23catgcacacc tgaaagctta tgcaaagatt aacgaggaat cactggatag ggctaggaga
60ttgctttggt ggcattacaa ttgtttactg tggggagaag ctaacgttac taattatatt
120tctcggcttc gcacttggct atcaacacct gagagataca gaggccgaga tgccccaacc
180attgaagcaa tcactagacc aatccaagtg gctcagggag gcagaaaaac atcttcgggt
240actagaaaac ctcgtggact cgaacctaga agaagaaaag ttaaaaccac agttgtctat
300gggagaagac gttcaaagtc cagggatagg agagcccctt caccccaacg tgcgggctcc
360cctctcccgc gtagttcgag cagccacaga agatctccct cgcctaggaa atagattacc
420tgctaggcat caccttggta aattgtcagg attatatcaa atgaagggat gtacatttaa
480ccctgaatgg aaagtacctg atatttcgga tactcatttt gatatgcaaa tagtaaatga
540gtgcccttcc cgaaattgga aatatctgac tccagccaaa ttctggccca agagcatttc
600ctactttcct gtacaggcag gggttaaagc taagtaccct gacaatgtga tgcaacatga
660atcaatagta ggtaaatatt taaccaggct ctatgaagca ggaatccttt ataagcggat
720atctaaacat ttggtcacat ttaaaggtca gccttataat tgggaacttc aataccttgt
780caagcaacat caagttcctg atgggtcaac aacctgcaaa atcaatggac gtgcggagaa
840tcgaaggagg agaactcctg ctaaatcaat tagcaggccg catgatccca aaagggacag
900tcacatggtc gggcaaattt ccaacaatag atcacatatt agaccatgtg caaacaatgg
960aggaaataaa cactcttcaa aaccaagggg cttggcctgc tggggcggga aggagagcag
1020gattaaccaa tcctgctcct caagagattc ctcagcccca gtggactccc gaagaagatc
1080agaaagctcg cgaagctttt cgtcgttatc aagaagagag accaccagag accaccacaa
1140ttcctcccac atctccaacg cagtggaaac tgcaacccgg ggacgatcca ctcctgggaa
1200acaagtctct gctcgagact cacccgcttt accagaatac cgagccagcc gtgtctgtaa
1260taaagactcc tccactgaga aagaaaatgt ctggtacctt cgggggaata ctagctggcc
1320taatcggatt actggtaagc tttttcttgt tgataaaaat tctagaaata cttcggaggc
1380tagattggtg gtggatttct ctcagttctc caaagggaaa aatgcaatgc gctttccaag
1440atactggagc ccaaacctct ccacattacg tcggatcttg cccgtgggga tgcccaggat
1500ttctctggac ttatctcagg ctttttatca tcttcctctt aatcctgcta gtagcagcag
1560gcttgctgta tctgacggac aacgggtcta ctattttagg aaagctccga tgggagtcgg
1620tcttagccct ttcctcctcc atctcttcac tactgccctc ggatccgaaa tcgctcgtcg
1680ctttaatgtt tggactttta cttatatgga tgacttcctc ctctgccacc caaacgctcg
1740tcaccttaac tcaattagcc acgctgtctg ctctttttta caagagctag gaataagaat
1800aaactttgac aaaactactc catcaccagt caacgaaatt agattcctcg gttatcaaat
1860tgatcaacga ttcatgaaga ttgaagaaag cagatggaaa gaattacgga ctgtaattaa
1920aaagataaaa attggagaat ggtatgactg gaaatgtatt cagagatttg tcgggcattt
1980aaactttgtg ttgccattta ccaaaggtaa catagaaatg ttaaaaccaa tgtatgctgc
2040tataactcat aaagtcaatt ttagcttctc ttctgcctat aggactttgc tgtacaaatt
2100aactatgggt gtttgtaaat tatcaatcaa accaaagtcc tctgtacctt tgccacgtgt
2160agctacggat gctaccccaa cacatggcgc aatatcccat atcaccggcg ggagcgcagt
2220gtttgctttt tcaaaggtca gagatataca tatacaggaa ttgctgatgg tatgtttagc
2280taaattaatg attaagccta gatgcatact aaccgattct acctttgttt gtcacaaacg
2340ttatcagacg ttaccatgga attttgcagt gtttgccaaa caattgttat cttctatacc
2400attgtacttt gtaccgagca aatataatcc tgctgacggc ccatccaggc acaaaccgcc
2460tgattggacg gctgttacat acacccctct ctcgaaagca atatatattc cacataggct
2520atgtggaact taagaattac acccctctcc ttcggagctg cctgccaagg tatctttacg
2580tctacattgc tgttgtcgtg tttgactgta cctttggtat gtaccattgt ttatgattct
2640tgcttatata tggatatcaa tgcttctaga gccttagcca atgtttatga tttgccagat
2700gatttcttcc caaaaattga tgatcttgta agggatgcga aggatgcttt agaaccttat
2760tggagatcag attcaataaa gaaacatgtt ttaattgcaa ctcactttgt ggatcttatt
2820gaagacttct ggcaaactac tcagggtatg catgaaatag ctgaagcctt aagagcagtt
2880ataccaccta ctacaacacc agttcccgca ggatatctga ttcagcacga agaggctgag
2940gagattcctc tgggagattt atttaaacat caggaagaaa ggatagttag tttccaaccg
3000gattatccta ttactgcacg aatt
30242410575DNAArtificial SequencepCHA vector sequence 24ggccgcaaca
gaggtggatg gacagacccg ttcttacacc ggactgggcg cgggatagga 60tattcagatt
gggatgggat tgagcttaaa gccggcgctg agaccatgct caaggtaggc 120aatgtcctca
gcgtcgagcc cggcatctat gtcgagggca ttggtggagc gcgcttcggg 180gataccgtgc
ttgtaactga gaccggatat gaggccctca ctccgcttga tcttggcaaa 240gatatttgac
gcatttatta gtatgtgtta attttcattt gcagtgcagt attttctatt 300cgatctttat
gtaattcgtt acaattaata aatattcaaa tcagattatt gactgtcatt 360tgtatcaaat
cgtgtttaat ggatattttt attataatat tgatgatatc tcaatcaaaa 420cgtagataat
aataatattt atttaatatt tttgcgtcgc acagtgaaaa tctatatgag 480attacaaaat
accgacaaca ttatttaaga tacatagaca ttaaccctga gactgttgga 540cagagctcat
tggtacctca gatctgggta actggcctaa ctggccttgg aggagctggc 600aactcaaaat
ccctttgcca aaaaccaaca tcatgccatc caccatgctt gtatccagct 660gcgcgcaatg
taccccgggc tgtgtatccc aaagcctcat gcaacctaac agatggatcg 720tttggaaggc
ctataacagc aaccacagac ttaaaacctt gcgcctccat agacttaagc 780aaatgtgtgt
acaatgtgga tcctaggccc aacctttgat gcctatgtga cacgtaaaca 840gtactctcaa
ctgtccaatc gtaagcgttc ctagccttcc agggcccagc gtaagcaata 900ccagccacaa
caccctcaac ctcagcaacc aaccaagggt atctatcttg caacctctct 960agatcatcaa
tccactcttg tggtgtttgt ggctctgtcc taaagttcac tgtagacgtc 1020tcaatgtaat
ggttaacgat atcacaaacc gcggccatat cagctgctgt agctggccta 1080atctcaactg
gtctcctctc cggagaagcc atggtttgga tccacaaact tacaaatttc 1140tctgaagttg
tatcctcagt acttcaaaga aaatagctta caccaatttt ttcttgtttt 1200cacaaatgcc
gaacttggtt ccttatatag gaaaactcaa gggcaaaaat gacacggaaa 1260aatataaaag
gataagtagt gggggataag attcctttgt gataaggtta ctttccgccc 1320ttacattttc
caccttacat gtgtcctcta tgtctctttc acaatcaccg accttatctt 1380cttcttttca
ttgttgtcgt cagtgcttac gtcttcaaga ttcttttctt cgcctggttc 1440ttctttttca
atttctacgt attcttcttc gtattctggc agtataggat cttgtatctg 1500tacattcttc
atttttgaac ataggttgca tatgtgccgc atattgatct gcttcttgct 1560gagctcacat
aatacttcca tagtttttcc cgtaaacatt ggattcttga tgctacatct 1620tggataatta
ccttctggcc ggccgcgaat tcgcttcaag acgtgctcaa atcactattt 1680ccacacccct
atatttctat tgcactccct tttaactgtt ttttattaca aaaatgccct 1740ggaaaatgca
ctcccttttt gtgtttgtta tttagtgaaa cgatgttgtc aggtaattta 1800tttgtcagtc
tactatggtg gcccattata ttaatagcaa ctgtcggtcc aatagacgac 1860gtcgattttc
tgcatttgtt taaccacgtg gattttatga cattttatat tagttaattt 1920gtaaaaccta
cccaattaaa gacctcatat gttctaaaga ctaatactta atgataacaa 1980ttttctttta
gtgaagaaag ggataattag taaatatgga acaagggcag aagatttatt 2040aaagccggta
agagacaaca acgtaggtac gtggagtgtc ttaggtgact tacccacata 2100acataaagtg
acattaacaa acatagctaa tgctcctatt tgaatagtgc atatcagcat 2160accttattac
atatagatag gagcaaactc tagctagatt gttgagagag ctcggtacct 2220taaaatctga
actcacaatc ctagatgcaa attctgcact gcaatgatcc attggagcac 2280atccaaaaag
acagaccagc taccatgatt gccagtgcta gggaactcgc cactgttgag 2340tagattgata
gtatctgata ggtgcccatt gactccaatt tgactccatc tatttcctct 2400ctgttcagcc
ttgattcttc tgagtattgt ggatagtcat acgttccgtt tctcacactt 2460tccatgcatt
cattgtcaca tttgtggtag aactcaaaac acccattgcc caattctttt 2520gcattatctc
tcagctggag tcggacctta tcgtataggt tcttgacata tgaatcatgg 2580aaatccagag
ttctttcatt ttccatgagc accagaagtt ctgcattgta agtccataca 2640tctagaaatc
catcttccat tttcttattc aaattttcta ttctcctttc taagttgttg 2700aattctttcc
caacggcttc gaattgagtg ttcattttgt caatgattga gttgacttta 2760ttggtgatcc
cgtcgattgc tttctgagtg gactctttgt ctgcagcata tccacttccc 2820tgctcgttgc
tatgatggta accataccat ccatctacca ttccttgcca ccccccttct 2880atgaatcctg
ctattgctcc aaacagacct cttgtttctc tctgaggcac gttcctcagt 2940cctgttgcaa
ggaccagttt atctgatttg acatatttgg gacactctcc aatggtaagg 3000ggatgaacat
tgtgaaaagg catactggaa tttatagcac ccactggggt ctgacatttg 3060gtatcacagt
tgccatactc cagttcgctt ctcatgattg ctgaatctcc ctttttaact 3120atcttgtatg
catattcagg agctataaag ttcccattac tttcaaagct gattgcatcg 3180ttcggcctta
gtattgtcca gaaaaattct attcttccac tttgtccatt cactttgggc 3240ctggtagcta
tttctggaat tgacctctga tttagtgttg atgttcctac agacacataa 3300gtgttcgagt
tctgatagag ttccgtttgt tccgctgcat cattagggtg atggattccc 3360cacaatatca
gaaggtcctc tacattggtg ttattgtagg tcctctttat tgttgggtat 3420gcattactct
tcttgatcaa ccacaccaca ttcctgaaaa aggaagatct accattgtat 3480gggcatgctg
agctcactcc tgatgaggca tcatgattgg accaagagtt cctagggatt 3540atttgaattt
tctcaaaatg gtttgtgttg ctcattaaat acttcagttc ttcataatca 3600ttgaagtctc
ccggataaca taagccattg gttggattgt ccttctctac aatatatgac 3660cattccggta
catttaggaa ctcatcacac attgggttcc caagaagcca tccagccaca 3720ctgcaatcct
tcagaatgag gggcctcact cctttgagac tgcagagttt cccgttgtgc 3780tctttttcca
gtatatcttg agcatgtgtg accgtaacat tcttctccat gattgtgtca 3840acttgttttg
ttgaattgtt tgcatgataa ccgatgcaga tttggtcacc tttgacaacg 3900ctgattattg
caagggcaat cactattctt tccatccatg gtttggatcc acaaacttac 3960aaatttctct
gaagttgtat cctcagtact tcaaagaaaa tagcttacac caattttttc 4020ttgttttcac
aaatgccgaa cttggttcct tatataggaa aactcaaggg caaaaatgac 4080acggaaaaat
ataaaaggat aagtagtggg ggataagatt cctttgtgat aaggttactt 4140tccgccctta
cattttccac cttacatgtg tcctctatgt ctctttcaca atcaccgacc 4200ttatcttctt
cttttcattg ttgtcgtcag tgcttacgtc ttcaagattc ttttcttcgc 4260ctggttcttc
tttttcaatt tctacgtatt cttcttcgta ttctggcagt ataggatctt 4320gtatctgtac
attcttcatt tttgaacata ggttgcatat gtgccgcata ttgatctgct 4380tcttgctgag
ctcacataat acttccatag tttttcccgt aaacattgga ttcttgatgc 4440tacatcttgg
ataattacct tctggcgcgc ctttgcccgg gctttcctgc agggtttaaa 4500cttaattaag
cggccgatcc ggtgagtaat attgtacggc taagagcgaa tttggcctgt 4560agacctcaat
tgcgagcttt ctaatttcaa actattcggg cctaactttt ggtgtgatga 4620tgctgactgg
caggatatat accgttgtaa tttgagctcg tgtgaataag tcgctgtgta 4680tgtttgtttg
attgtttctg ttggagtgca gcccatttca ccggacaagt cggctagatt 4740gatttagccc
tgatgaactg ccgaggggaa gccatcttga gcgcggaatg ggaatggatc 4800gaaccgggag
cacaggatga cgcctaacaa ttcattcaag ccgacaccgc ttcgcggcgc 4860ggcttaattc
aggagttaaa catcatgagg gaagcggtga tcgccgaagt atcgactcaa 4920ctatcagagg
tagttggcgt catcgagcgc catctcgaac cgacgttgct ggccgtacat 4980ttgtacggct
ccgcagtgga tggcggcctg aagccacaca gtgatattga tttgctggtt 5040acggtgaccg
taaggcttga tgaaacaacg cggcgagctt tgatcaacga ccttttggaa 5100acttcggctt
cccctggaga gagcgagatt ctccgcgctg tagaagtcac cattgttgtg 5160cacgacgaca
tcattccgtg gcgttatcca gctaagcgcg aactgcaatt tggagaatgg 5220cagcgcaatg
acattcttgc aggtatcttc gagccagcca cgatcgacat tgatctggct 5280atcttgctga
caaaagcaag agaacatagc gttgccttgg taggtccagc ggcggaggaa 5340ctctttgatc
cggttcctga acaggatcta tttgaggcgc taaatgaaac cttaacgcta 5400tggaactcgc
cgcccgactg ggctggcgat gagcgaaatg tagtgcttac gttgtcccgc 5460atttggtaca
gcgcagtaac cggcaaaatc gcgccgaagg atgtcgctgc cgactgggca 5520atggagcgcc
tgccggccca gtatcagccc gtcatacttg aagctaggca ggcttatctt 5580ggacaagaag
atcgcttggc ctcgcgcgca gatcagttgg aagaatttgt tcactacgtg 5640aaaggcgaga
tcaccaaggt agtcggcaaa taatgtctaa caattcgttc aagccgacgc 5700cgcttcgcgg
cgcggcttaa ctcaagcgtt agagagctgg ggaagactat gcgcgatctg 5760ttgaaggtgg
ttctaagcct cgtacttgcg atggcatttc gatcgaaagg ggtacaaatt 5820cccactaagc
gctcgggggc tgagaaagcc cagtaaggaa acaactgtag gttcgagtcg 5880cgagatcccc
cggaaccaaa ggaagtaggt taaacccgct ccgatcaggc cgagccacgc 5940caggccgaga
acattggttc ctgtaggcat cgggattggc ggatcaaaca ctaaagctac 6000tggaacgagc
agaagtcctc cggccgccag ttgccaggcc gtaaaggtga gcagaggcac 6060gggaggttgc
cacttgcggg tcagcacggt tccgaacgcc atggaaaccg cccccgccag 6120gcccgctgcg
acgccgacag gatctagcgc tgcgtttggt gtcaacacca acagcgccac 6180gcccgcagtt
ccgcaaatag cccccaggac cgccatcaat cgtatcgggc tacctagcag 6240agcggcagag
atgaacacga ccatcagcgg ctgcacagcg cctaccgtcg ccgcgacccg 6300cccggcaggc
ggtagaccga aataaacaac aagctccaga atagcgaaat attaagtgcg 6360ccgaggatga
agatgcgcat ccaccagatt cccgttggaa tctgtcggac gatcatcacg 6420agcaataaac
ccgccggcaa cgcccgcagc agcataccgg cgacccctcg gcctcgctgt 6480tcgggctcca
cgaaaacgcc ggacagatgc gccttgtgag cgtccttggg gccgtcctcc 6540tgtttgaaga
ccgacagccc aatgatctcg ccgtcgatgt aggcgccgaa tgccacggca 6600tctcgcaacc
gttcagcgaa cgcctccatg ggctttttct cctcgtgctc gtaaacggac 6660ccgaacatct
ctggagcttt cttcagggcc gacaatcgga tctcgcggaa atcctgcacg 6720tcggccgctc
caagccgtcg aatctgagcc ttaatcacaa ttgtcaattt taatcctctg 6780tttatcggca
gttcgtagag cgcgccgtgc gcccgagcga tactgagcga agcaagtgcg 6840tcgagcagtg
cccgcttgtt cctgaaatgc cagtaaagcg ctggctgctg aacccccagc 6900cggaactgac
cccacaaggc cctagcgttt gcaatgcacc aggtcatcat tgacccaggc 6960gtgttccacc
aggccgctgc ctcgcaactc ttcgcaggct tcgccgacct gctcgcgcca 7020cttcttcacg
cgggtggaat ccgatccgca catgaggcgg aaggtttcca gcttgagcgg 7080gtacggctcc
cggtgcgagc tgaaatagtc gaacatccgt cgggccgtcg gcgacagctt 7140gcggtacttc
tcccatatga atttcgtgta gtggtcgcca gcaaacagca cgacgatttc 7200ctcgtcgatc
aggacctggc aacgggacgt tttcttgcca cggtccagga cgcggaagcg 7260gtgcagcagc
gacaccgatt ccaggtgccc aacgcggtcg gacgtgaagc ccatcgccgt 7320cgcctgtagg
cgcgacaggc attcctcggc cttcgtgtaa taccggccat tgatcgacca 7380gcccaggtcc
tggcaaagct cgtagaacgt gaaggtgatc ggctcgccga taggggtgcg 7440cttcgcgtac
tccaacacct gctgccacac cagttcgtca tcgtcggccc gcagctcgac 7500gccggtgtag
gtgatcttca cgtccttgtt gacgtggaaa atgaccttgt tttgcagcgc 7560ctcgcgcggg
attttcttgt tgcgcgtggt gaacagggca gagcgggccg tgtcgtttgg 7620catcgctcgc
atcgtgtccg gccacggcgc aatatcgaac aaggaaagct gcatttcctt 7680gatctgctgc
ttcgtgtgtt tcagcaacgc ggcctgcttg gcctcgctga cctgttttgc 7740caggtcctcg
ccggcggttt ttcgcttctt ggtcgtcata gttcctcgcg tgtcgatggt 7800catcgacttc
gccaaacctg ccgcctcctg ttcgagacga cgcgaacgct ccacggcggc 7860cgatggcgcg
ggcagggcag ggggagccag ttgcacgctg tcgcgctcga tcttggccgt 7920agcttgctgg
accatcgagc cgacggactg gaaggtttcg cggggcgcac gcatgacggt 7980gcggcttgcg
atggtttcgg catcctcggc ggaaaacccc gcgtcgatca gttcttgcct 8040gtatgccttc
cggtcaaacg tccgattcat tcaccctcct tgcgggattg ccccgactca 8100cgccggggca
atgtgccctt attcctgatt tgacccgcct ggtgccttgg tgtccagata 8160atccacctta
tcggcaatga agtcggtccc gtagaccgtc tggccgtcct tctcgtactt 8220ggtattccga
atcttgccct gcacgaatac cagcgacccc ttgcccaaat acttgccgtg 8280ggcctcggcc
tgagagccaa aacacttgat gcggaagaag tcggtgcgct cctgcttgtc 8340gccggcatcg
ttgcgccact cttcattaac cgctatatcg aaaattgctt gcggcttgtt 8400agaattgcca
tgacgtacct cggtgtcacg ggtaagatta ccgataaact ggaactgatt 8460atggcnnctc
gaaattccct cggtcttgcc ttgctcgtcg gtgatgtact tcaccagctc 8520cgcgaagtcg
ctcttcttga tggagcgcat ggggacgtgc ttggcaatca cgcgcacccc 8580ccggccgttt
tagcggctaa aaaagtcatg gctctgccct cgggcggacc acgcccatca 8640tgaccttgcc
aagctcgtcc tgcttctctt cgatcttcgc cagcagggcg aggatcgtgg 8700catcaccgaa
ccgcgccgtg cgcgggtcgt cggtgagcca gagtttcagc aggccgccca 8760ggcggcccag
gtcgccattg atgcgggcca gctcgcggac gtgctcatag tccacgacgc 8820ccgtgatttt
gtagccctgg ccgacggcca gcaggtaggc cgacaggctc atgccggccg 8880ccgccgcctt
ttcctcaatc gctcttcgtt cgtctggaag gcagtacacc ttgataggtg 8940ggctgccctt
cctggttggc ttggtttcat cagccatccg cttgccctca tctgttacgc 9000cggcggtagc
cggccagcct cgcagagcag gattcccgtt gagcaccgcc aggtgcgaat 9060aagggacagt
gaagaaggaa cacccgctcg cgggtgggcc tacttcacct atcctgcccg 9120gctgacgccg
ttggatacac caaggaaagt ctacacgaac cctttggcaa aatcctgtat 9180atcgtgcgaa
aaaggatgga tataccgaaa aaatcgctat aatgaccccg aagcagggtt 9240atgcagcgga
aaagatccgt cgaccctttc cgacgctcac cgggctggtt gccctcgccg 9300ctgggctggc
ggccgtctat ggccctgcaa acgcgccaga aacgccgtcg aagccgtgtg 9360cgagacaccg
cggccgccgg cgttgtggat accacgcgga aaacttggcc ctcactgaca 9420gatgaggggc
ggacgttgac acttgagggg ccgactcacc cggcgcggcg ttgacagatg 9480aggggcaggc
tcgatttcgg ccggcgacgt ggagctggcc agcctcgcaa atcggcgaaa 9540acgcctgatt
ttacgcgagt ttcccacaga tgatgtggac aagcctgggg ataagtgccc 9600tgcggtattg
acacttgagg ggcgcgacta ctgacagatg aggggcgcga tccttgacac 9660ttgaggggca
gagtgatgac agatgagggg cgcacctatt gacatttgag gggctgtcca 9720caggcagaaa
atccagcatt tgcaagggtt tccgcccgtt tttcggccac cgctaacctg 9780tcttttaacc
tgcttttaaa ccaatattta taaaccttgt ttttaaccag ggctgcgccc 9840tggcgcgtga
ccgcgcacgc cgaagggggg tgccccccct tctcgaaccc tcccggcccg 9900ctaacgcggg
cctcccatcc ccccaggggc tgcgcccctc ggccgcgaac ggcctcaccc 9960caaaaatggc
aggccaagct agcttgcttg gtcgttccgg tacgtaccgt gaacgtcggc 10020tcgattgtac
ctgcgttcaa atactttgcg atcgtgttgc gcgcctgccc ggtgcgtcgg 10080ctgatctcac
ggatcgactg cttctctcgc aacgccatcc gacggatgat gtttaaaagt 10140cccatgtgga
tcactccgtt gccccgtcgc tcaccgtgtt ggggggaagg tgcacatggc 10200tcagttctca
atggaaatta tctgcctaac cggctcagtt ctgcgtagaa accaacatgc 10260aagctccacc
gggtgcaaag cggcagcggc ggcaggatat attcaattgt aaatggcttc 10320atgtccggga
aatctacatg gatcagcaat gagtatgatg gtcaatatgg agaaaaagaa 10380agagtaatta
ccaatttttt ttcaattcaa aaatgtagat gtccgcagcg ttattataaa 10440atgaaagtac
attttgataa aacgacaaat tacgatccgt cgtatttata ggcgaaagca 10500ataaacaaat
tattctaatt cggaaatctt tatttcgacg tgtctacatt cacgtccaaa 10560tgggggcggc
gaatt
105752510677DNAArtificial SequencepMHN vector sequence 25ggccgcaaca
gaggtggatg gacagacccg ttcttacacc ggactgggcg cgggatagga 60tattcagatt
gggatgggat tgagcttaaa gccggcgctg agaccatgct caaggtaggc 120aatgtcctca
gcgtcgagcc cggcatctat gtcgagggca ttggtggagc gcgcttcggg 180gataccgtgc
ttgtaactga gaccggatat gaggccctca ctccgcttga tcttggcaaa 240gatatttgac
gcatttatta gtatgtgtta attttcattt gcagtgcagt attttctatt 300cgatctttat
gtaattcgtt acaattaata aatattcaaa tcagattatt gactgtcatt 360tgtatcaaat
cgtgtttaat ggatattttt attataatat tgatgatatc tcaatcaaaa 420cgtagataat
aataatattt atttaatatt tttgcgtcgc acagtgaaaa tctatatgag 480attacaaaat
accgacaaca ttatttaaga tacatagaca ttaaccctga gactgttgga 540cagagctcat
tggtacctca gatctgggta actggcctaa ctggccttgg aggagctggc 600aactcaaaat
ccctttgcca aaaaccaaca tcatgccatc caccatgctt gtatccagct 660gcgcgcaatg
taccccgggc tgtgtatccc aaagcctcat gcaacctaac agatggatcg 720tttggaaggc
ctataacagc aaccacagac ttaaaacctt gcgcctccat agacttaagc 780aaatgtgtgt
acaatgtgga tcctaggccc aacctttgat gcctatgtga cacgtaaaca 840gtactctcaa
ctgtccaatc gtaagcgttc ctagccttcc agggcccagc gtaagcaata 900ccagccacaa
caccctcaac ctcagcaacc aaccaagggt atctatcttg caacctctct 960agatcatcaa
tccactcttg tggtgtttgt ggctctgtcc taaagttcac tgtagacgtc 1020tcaatgtaat
ggttaacgat atcacaaacc gcggccatat cagctgctgt agctggccta 1080atctcaactg
gtctcctctc cggagaagcc atggtttgga tccacaaact tacaaatttc 1140tctgaagttg
tatcctcagt acttcaaaga aaatagctta caccaatttt ttcttgtttt 1200cacaaatgcc
gaacttggtt ccttatatag gaaaactcaa gggcaaaaat gacacggaaa 1260aatataaaag
gataagtagt gggggataag attcctttgt gataaggtta ctttccgccc 1320ttacattttc
caccttacat gtgtcctcta tgtctctttc acaatcaccg accttatctt 1380cttcttttca
ttgttgtcgt cagtgcttac gtcttcaaga ttcttttctt cgcctggttc 1440ttctttttca
atttctacgt attcttcttc gtattctggc agtataggat cttgtatctg 1500tacattcttc
atttttgaac ataggttgca tatgtgccgc atattgatct gcttcttgct 1560gagctcacat
aatacttcca tagtttttcc cgtaaacatt ggattcttga tgctacatct 1620tggataatta
ccttctggcc ggccgcgaat tcgcttcaag acgtgctcaa atcactattt 1680ccacacccct
atatttctat tgcactccct tttaactgtt ttttattaca aaaatgccct 1740ggaaaatgca
ctcccttttt gtgtttgtta tttagtgaaa cgatgttgtc aggtaattta 1800tttgtcagtc
tactatggtg gcccattata ttaatagcaa ctgtcggtcc aatagacgac 1860gtcgattttc
tgcatttgtt taaccacgtg gattttatga cattttatat tagttaattt 1920gtaaaaccta
cccaattaaa gacctcatat gttctaaaga ctaatactta atgataacaa 1980ttttctttta
gtgaagaaag ggataattag taaatatgga acaagggcag aagatttatt 2040aaagccggta
agagacaaca acgtaggtac gtggagtgtc ttaggtgact tacccacata 2100acataaagtg
acattaacaa acatagctaa tgctcctatt tgaatagtgc atatcagcat 2160accttattac
atatagatag gagcaaactc tagctagatt gttgagagag ctcggtaccg 2220gatccgaaga
cttaacctga tcttgcttca cgtacaccat catctttcag aatctccacc 2280aaaagtggaa
caattctgaa ttccccaaag agagtgttag aaatctcagc tatgctcaga 2340caataggtct
tgttcgtctt tacaactttg aaacatgtgg aggtagtgta tgctgcctta 2400gtagaactag
aggaaactct ggttatcctt gatctggatg tagaatcaaa cacagcagag 2460gcaggattca
acctagcttg aacaccatct aacattgttc caaacacccc tctcaaggta 2520tgattacggt
agaagatcaa agggtaagga tcagtgtaaa ctccagtcac acacgaattc 2580ggacatctag
ctgaagcttg gcaaggaatc gatccaggtc ttgtaaaggc attgaaagta 2640tatggtgaat
gtaatgtagc tgtcttgttg ctcacagtca ttggatacag taacgctggg 2700ctaaagtagg
aacttccacg ttgatagaga aagtggctag taccaacagt aagaatcctt 2760ccctcagctc
ccatcaatgt tacagtgttt ggtggaacag taaggactgg atcttctccc 2820aatgatgtgc
taaccttgat actgagaata gcttgttgta tcctcttacc tccaaatctt 2880cctggtttgt
atgatgactt agccattcga atctgatagt cttgctcatc aggacaagta 2940tcattgtacc
tcttgtagat aacatacttt ccctcttgca cagtatcact aggactgtta 3000ggtttcaaac
caccatagac agagaaccag actctggagt caatgaagct accacctcca 3060actcctggat
agttagctac ccaatcccca aacaatgtag tgacatccaa atctttctca 3120tggtattgac
catcaaaacc caatctgcca tgcaccatcc ttgtagggac tgcagaattg 3180tagtcctctt
cttcagtttc tgttacttta ctgcatagca tatcacagcc caaaggtgta 3240gcagatacag
agcaactttt gcgattctgt gtatcatcaa gattgatact gcgaagagtt 3300gagaagaaca
ctctaccagt agcagatgta cgaagaactc caagtgctaa gtactgataa 3360gagtgagaat
ggtcacgaca gccagataga atgacattgt gtgtatagca atagtgtgta 3420gcactcatgt
caaatgaggg tatccgagtg cacccactcc cagttgtggg tgcaggaatg 3480aagttcagat
gttcctggaa agctgaagga tagaaagatg taacatctga agcatcatct 3540acaatgagtt
ctttgccaat acctccaatg tagtctggat catgaattgg ggctccccaa 3600cctgaattgt
tggcagcccc attgatctga taggataggc tggtgattgc attcatgatg 3660gtagtttctg
tgttgagtaa tgcaagtgga ctttcaagtg caacttgttt gtagattcgg 3720tccacaacat
cctggttgga gcctagggta ctggtaatct tctcttcagc cctagagatt 3780cgtgtgggta
tgccaactaa gtctgagggg gtgcttgctc ccatagaata gagtaatgat 3840gcaacagaga
ttgctagggt gacaacagta agaaagagaa tggctatccg aaagataagc 3900ctccaagtgt
tcttggcttc cctctcatca ttctcaagag ccacttgtga aactgctcgg 3960tccatggttt
ggatccgcga tttggtgtat cgagattggt tatgaaattc agatgctagt 4020gtaatgtatt
ggtaatttgg gaagatataa taggaagcaa ggctatttat ccatttctga 4080aaaggcgaaa
tggcgtcacc gcgagcgtca cgctctagtc gaccatgtac gtaagcgctt 4140acgtttttgg
tggaccccct cgaccatgta cgtaagcgct tacgtttttg gtggaccccc 4200tcgaccatgt
acgtaagcgc ttacgttttt ggtggacccc ctcgaccatg tacgtaagcg 4260cttacgtttt
tggtggaccc cctcgacgga tcccccctcg accctagacg tatctattca 4320aaagtcgtta
atggctgcgg atcaagaaaa agttggaata gaaacagaat acccgcgaaa 4380ttcaggcccg
gttgccatgt cctacacgcc gaaataaacg accaaattag tagaaaaata 4440aaaactagct
cagatactta cgtcacgtct tgcgcactga tttgaaaaat ctcaatataa 4500acaaagacgg
ccacaagaaa aaaccaaaac accgatattc attaatctta tctagtttct 4560caaaaaaatt
catatcttcc acaccctcga gatctagata aacttaatta agcggccgat 4620ccggtgagta
atattgtacg gctaagagcg aatttggcct gtagacctca attgcgagct 4680ttctaatttc
aaactattcg ggcctaactt ttggtgtgat gatgctgact ggcaggatat 4740ataccgttgt
aatttgagct cgtgtgaata agtcgctgtg tatgtttgtt tgattgtttc 4800tgttggagtg
cagcccattt caccggacaa gtcggctaga ttgatttagc cctgatgaac 4860tgccgagggg
aagccatctt gagcgcggaa tgggaatgga tcgaaccggg agcacaggat 4920gacgcctaac
aattcattca agccgacacc gcttcgcggc gcggcttaat tcaggagtta 4980aacatcatga
gggaagcggt gatcgccgaa gtatcgactc aactatcaga ggtagttggc 5040gtcatcgagc
gccatctcga accgacgttg ctggccgtac atttgtacgg ctccgcagtg 5100gatggcggcc
tgaagccaca cagtgatatt gatttgctgg ttacggtgac cgtaaggctt 5160gatgaaacaa
cgcggcgagc tttgatcaac gaccttttgg aaacttcggc ttcccctgga 5220gagagcgaga
ttctccgcgc tgtagaagtc accattgttg tgcacgacga catcattccg 5280tggcgttatc
cagctaagcg cgaactgcaa tttggagaat ggcagcgcaa tgacattctt 5340gcaggtatct
tcgagccagc cacgatcgac attgatctgg ctatcttgct gacaaaagca 5400agagaacata
gcgttgcctt ggtaggtcca gcggcggagg aactctttga tccggttcct 5460gaacaggatc
tatttgaggc gctaaatgaa accttaacgc tatggaactc gccgcccgac 5520tgggctggcg
atgagcgaaa tgtagtgctt acgttgtccc gcatttggta cagcgcagta 5580accggcaaaa
tcgcgccgaa ggatgtcgct gccgactggg caatggagcg cctgccggcc 5640cagtatcagc
ccgtcatact tgaagctagg caggcttatc ttggacaaga agatcgcttg 5700gcctcgcgcg
cagatcagtt ggaagaattt gttcactacg tgaaaggcga gatcaccaag 5760gtagtcggca
aataatgtct aacaattcgt tcaagccgac gccgcttcgc ggcgcggctt 5820aactcaagcg
ttagagagct ggggaagact atgcgcgatc tgttgaaggt ggttctaagc 5880ctcgtacttg
cgatggcatt tcgatcgaaa ggggtacaaa ttcccactaa gcgctcgggg 5940gctgagaaag
cccagtaagg aaacaactgt aggttcgagt cgcgagatcc cccggaacca 6000aaggaagtag
gttaaacccg ctccgatcag gccgagccac gccaggccga gaacattggt 6060tcctgtaggc
atcgggattg gcggatcaaa cactaaagct actggaacga gcagaagtcc 6120tccggccgcc
agttgccagg ccgtaaaggt gagcagaggc acgggaggtt gccacttgcg 6180ggtcagcacg
gttccgaacg ccatggaaac cgcccccgcc aggcccgctg cgacgccgac 6240aggatctagc
gctgcgtttg gtgtcaacac caacagcgcc acgcccgcag ttccgcaaat 6300agcccccagg
accgccatca atcgtatcgg gctacctagc agagcggcag agatgaacac 6360gaccatcagc
ggctgcacag cgcctaccgt cgccgcgacc cgcccggcag gcggtagacc 6420gaaataaaca
acaagctcca gaatagcgaa atattaagtg cgccgaggat gaagatgcgc 6480atccaccaga
ttcccgttgg aatctgtcgg acgatcatca cgagcaataa acccgccggc 6540aacgcccgca
gcagcatacc ggcgacccct cggcctcgct gttcgggctc cacgaaaacg 6600ccggacagat
gcgccttgtg agcgtccttg gggccgtcct cctgtttgaa gaccgacagc 6660ccaatgatct
cgccgtcgat gtaggcgccg aatgccacgg catctcgcaa ccgttcagcg 6720aacgcctcca
tgggcttttt ctcctcgtgc tcgtaaacgg acccgaacat ctctggagct 6780ttcttcaggg
ccgacaatcg gatctcgcgg aaatcctgca cgtcggccgc tccaagccgt 6840cgaatctgag
ccttaatcac aattgtcaat tttaatcctc tgtttatcgg cagttcgtag 6900agcgcgccgt
gcgcccgagc gatactgagc gaagcaagtg cgtcgagcag tgcccgcttg 6960ttcctgaaat
gccagtaaag cgctggctgc tgaaccccca gccggaactg accccacaag 7020gccctagcgt
ttgcaatgca ccaggtcatc attgacccag gcgtgttcca ccaggccgct 7080gcctcgcaac
tcttcgcagg cttcgccgac ctgctcgcgc cacttcttca cgcgggtgga 7140atccgatccg
cacatgaggc ggaaggtttc cagcttgagc gggtacggct cccggtgcga 7200gctgaaatag
tcgaacatcc gtcgggccgt cggcgacagc ttgcggtact tctcccatat 7260gaatttcgtg
tagtggtcgc cagcaaacag cacgacgatt tcctcgtcga tcaggacctg 7320gcaacgggac
gttttcttgc cacggtccag gacgcggaag cggtgcagca gcgacaccga 7380ttccaggtgc
ccaacgcggt cggacgtgaa gcccatcgcc gtcgcctgta ggcgcgacag 7440gcattcctcg
gccttcgtgt aataccggcc attgatcgac cagcccaggt cctggcaaag 7500ctcgtagaac
gtgaaggtga tcggctcgcc gataggggtg cgcttcgcgt actccaacac 7560ctgctgccac
accagttcgt catcgtcggc ccgcagctcg acgccggtgt aggtgatctt 7620cacgtccttg
ttgacgtgga aaatgacctt gttttgcagc gcctcgcgcg ggattttctt 7680gttgcgcgtg
gtgaacaggg cagagcgggc cgtgtcgttt ggcatcgctc gcatcgtgtc 7740cggccacggc
gcaatatcga acaaggaaag ctgcatttcc ttgatctgct gcttcgtgtg 7800tttcagcaac
gcggcctgct tggcctcgct gacctgtttt gccaggtcct cgccggcggt 7860ttttcgcttc
ttggtcgtca tagttcctcg cgtgtcgatg gtcatcgact tcgccaaacc 7920tgccgcctcc
tgttcgagac gacgcgaacg ctccacggcg gccgatggcg cgggcagggc 7980agggggagcc
agttgcacgc tgtcgcgctc gatcttggcc gtagcttgct ggaccatcga 8040gccgacggac
tggaaggttt cgcggggcgc acgcatgacg gtgcggcttg cgatggtttc 8100ggcatcctcg
gcggaaaacc ccgcgtcgat cagttcttgc ctgtatgcct tccggtcaaa 8160cgtccgattc
attcaccctc cttgcgggat tgccccgact cacgccgggg caatgtgccc 8220ttattcctga
tttgacccgc ctggtgcctt ggtgtccaga taatccacct tatcggcaat 8280gaagtcggtc
ccgtagaccg tctggccgtc cttctcgtac ttggtattcc gaatcttgcc 8340ctgcacgaat
accagcgacc ccttgcccaa atacttgccg tgggcctcgg cctgagagcc 8400aaaacacttg
atgcggaaga agtcggtgcg ctcctgcttg tcgccggcat cgttgcgcca 8460ctcttcatta
accgctatat cgaaaattgc ttgcggcttg ttagaattgc catgacgtac 8520ctcggtgtca
cgggtaagat taccgataaa ctggaactga ttatggcnnc tcgaaattcc 8580ctcggtcttg
ccttgctcgt cggtgatgta cttcaccagc tccgcgaagt cgctcttctt 8640gatggagcgc
atggggacgt gcttggcaat cacgcgcacc ccccggccgt tttagcggct 8700aaaaaagtca
tggctctgcc ctcgggcgga ccacgcccat catgaccttg ccaagctcgt 8760cctgcttctc
ttcgatcttc gccagcaggg cgaggatcgt ggcatcaccg aaccgcgccg 8820tgcgcgggtc
gtcggtgagc cagagtttca gcaggccgcc caggcggccc aggtcgccat 8880tgatgcgggc
cagctcgcgg acgtgctcat agtccacgac gcccgtgatt ttgtagccct 8940ggccgacggc
cagcaggtag gccgacaggc tcatgccggc cgccgccgcc ttttcctcaa 9000tcgctcttcg
ttcgtctgga aggcagtaca ccttgatagg tgggctgccc ttcctggttg 9060gcttggtttc
atcagccatc cgcttgccct catctgttac gccggcggta gccggccagc 9120ctcgcagagc
aggattcccg ttgagcaccg ccaggtgcga ataagggaca gtgaagaagg 9180aacacccgct
cgcgggtggg cctacttcac ctatcctgcc cggctgacgc cgttggatac 9240accaaggaaa
gtctacacga accctttggc aaaatcctgt atatcgtgcg aaaaaggatg 9300gatataccga
aaaaatcgct ataatgaccc cgaagcaggg ttatgcagcg gaaaagatcc 9360gtcgaccctt
tccgacgctc accgggctgg ttgccctcgc cgctgggctg gcggccgtct 9420atggccctgc
aaacgcgcca gaaacgccgt cgaagccgtg tgcgagacac cgcggccgcc 9480ggcgttgtgg
ataccacgcg gaaaacttgg ccctcactga cagatgaggg gcggacgttg 9540acacttgagg
ggccgactca cccggcgcgg cgttgacaga tgaggggcag gctcgatttc 9600ggccggcgac
gtggagctgg ccagcctcgc aaatcggcga aaacgcctga ttttacgcga 9660gtttcccaca
gatgatgtgg acaagcctgg ggataagtgc cctgcggtat tgacacttga 9720ggggcgcgac
tactgacaga tgaggggcgc gatccttgac acttgagggg cagagtgatg 9780acagatgagg
ggcgcaccta ttgacatttg aggggctgtc cacaggcaga aaatccagca 9840tttgcaaggg
tttccgcccg tttttcggcc accgctaacc tgtcttttaa cctgctttta 9900aaccaatatt
tataaacctt gtttttaacc agggctgcgc cctggcgcgt gaccgcgcac 9960gccgaagggg
ggtgcccccc cttctcgaac cctcccggcc cgctaacgcg ggcctcccat 10020ccccccaggg
gctgcgcccc tcggccgcga acggcctcac cccaaaaatg gcaggccaag 10080ctagcttgct
tggtcgttcc ggtacgtacc gtgaacgtcg gctcgattgt acctgcgttc 10140aaatactttg
cgatcgtgtt gcgcgcctgc ccggtgcgtc ggctgatctc acggatcgac 10200tgcttctctc
gcaacgccat ccgacggatg atgtttaaaa gtcccatgtg gatcactccg 10260ttgccccgtc
gctcaccgtg ttggggggaa ggtgcacatg gctcagttct caatggaaat 10320tatctgccta
accggctcag ttctgcgtag aaaccaacat gcaagctcca ccgggtgcaa 10380agcggcagcg
gcggcaggat atattcaatt gtaaatggct tcatgtccgg gaaatctaca 10440tggatcagca
atgagtatga tggtcaatat ggagaaaaag aaagagtaat taccaatttt 10500ttttcaattc
aaaaatgtag atgtccgcag cgttattata aaatgaaagt acattttgat 10560aaaacgacaa
attacgatcc gtcgtattta taggcgaaag caataaacaa attattctaa 10620ttcggaaatc
tttatttcga cgtgtctaca ttcacgtcca aatgggggcg gcgaatt
106772610603DNAArtificial SequencepCHN vector sequence 26tgcgtagaaa
ccaacatgca agctccaccg ggtgcaaagc ggcagcggcg gcaggatata 60ttcaattgta
aatggcttca tgtccgggaa atctacatgg atcagcaatg agtatgatgg 120tcaatatgga
gaaaaagaaa gagtaattac caattttttt tcaattcaaa aatgtagatg 180tccgcagcgt
tattataaaa tgaaagtaca ttttgataaa acgacaaatt acgatccgtc 240gtatttatag
gcgaaagcaa taaacaaatt attctaattc ggaaatcttt atttcgacgt 300gtctacattc
acgtccaaat gggggcggcg aattggccgc aacagaggtg gatggacaga 360cccgttctta
caccggactg ggcgcgggat aggatattca gattgggatg ggattgagct 420taaagccggc
gctgagacca tgctcaaggt aggcaatgtc ctcagcgtcg agcccggcat 480ctatgtcgag
ggcattggtg gagcgcgctt cggggatacc gtgcttgtaa ctgagaccgg 540atatgaggcc
ctcactccgc ttgatcttgg caaagatatt tgacgcattt attagtatgt 600gttaattttc
atttgcagtg cagtattttc tattcgatct ttatgtaatt cgttacaatt 660aataaatatt
caaatcagat tattgactgt catttgtatc aaatcgtgtt taatggatat 720ttttattata
atattgatga tatctcaatc aaaacgtaga taataataat atttatttaa 780tatttttgcg
tcgcacagtg aaaatctata tgagattaca aaataccgac aacattattt 840aagatacata
gacattaacc ctgagactgt tggacagagc tcattggtac ctcagatctg 900ggtaactggc
ctaactggcc ttggaggagc tggcaactca aaatcccttt gccaaaaacc 960aacatcatgc
catccaccat gcttgtatcc agctgcgcgc aatgtacccc gggctgtgta 1020tcccaaagcc
tcatgcaacc taacagatgg atcgtttgga aggcctataa cagcaaccac 1080agacttaaaa
ccttgcgcct ccatagactt aagcaaatgt gtgtacaatg tggatcctag 1140gcccaacctt
tgatgcctat gtgacacgta aacagtactc tcaactgtcc aatcgtaagc 1200gttcctagcc
ttccagggcc cagcgtaagc aataccagcc acaacaccct caacctcagc 1260aaccaaccaa
gggtatctat cttgcaacct ctctagatca tcaatccact cttgtggtgt 1320ttgtggctct
gtcctaaagt tcactgtaga cgtctcaatg taatggttaa cgatatcaca 1380aaccgcggcc
atatcagctg ctgtagctgg cctaatctca actggtctcc tctccggaga 1440agccatggtt
tggatccaca aacttacaaa tttctctgaa gttgtatcct cagtacttca 1500aagaaaatag
cttacaccaa attttttctt gttttcacaa atgccgaact tggttcctta 1560tataggaaaa
ctcaagggca aaaatgacac ggaaaaatat aaaaggataa gtagtggggg 1620ataagattcc
tttgtgataa ggttactttc cgcccttaca ttttccacct tacatgtgtc 1680ctctatgtct
ctttcacaat caccgacctt atcttcttct tttcattgtt gtcgtcagtg 1740cttacgtctt
caagattctt ttcttcgcct ggttcttctt tttcaatttc tacgtattct 1800tcttcgtatt
ctggcagtat aggatcttgt atctgtacat tcttcatttt tgaacatagg 1860ttgcatatgt
gccgcatatt gatctgcttc ttgctgagct cacataatac ttccatagtt 1920tttcccgtaa
acattggatt cttgatgcta catcttggat aattaccttc tggccggccg 1980cgaattcgct
tcaagacgtg ctcaaatcac tatttccaca cccctatatt tctattgcac 2040tcccttttaa
ctgtttttta ttacaaaaat gccctggaaa atgcactccc tttttgtgtt 2100tgttatttag
tgaaacgatg ttgtcaggta atttatttgt cagtctacta tggtggccca 2160ttatattaat
agcaactgtc ggtccaatag acgacgtcga ttttctgcat ttgtttaacc 2220acgtggattt
tatgacattt tatattagtt aatttgtaaa acctacccaa ttaaagacct 2280catatgttct
aaagactaat acttaatgat aacaattttc ttttagtgaa gaaagggata 2340attagtaaat
atggaacaag ggcagaagat ttattaaagc cggtaagaga caacaacgta 2400ggtacgtgga
gtgtcttagg tgacttaccc acataacata aagtgacatt aacaaacata 2460gctaatgctc
ctatttgaat agtgcatatc agcatacctt attacatata gataggagca 2520aactctagct
agattgttga gagagctcgg taccggatcc gaagacttaa cctgatcttg 2580cttcacgtac
accatcatct ttcagaatct ccaccaaaag tggaacaatt ctgaattccc 2640caaagagagt
gttagaaatc tcagctatgc tcagacaata ggtcttgttc gtctttacaa 2700ctttgaaaca
tgtggaggta gtgtatgctg ccttagtaga actagaggaa actctggtta 2760tccttgatct
ggatgtagaa tcaaacacag cagaggcagg attcaaccta gcttgaacac 2820catctaacat
tgttccaaac acccctctca aggtatgatt acggtagaag atcaaagggt 2880aaggatcagt
gtaaactcca gtcacacacg aattcggaca tctagctgaa gcttggcaag 2940gaatcgatcc
aggtcttgta aaggcattga aagtatatgg tgaatgtaat gtagctgtct 3000tgttgctcac
agtcattgga tacagtaacg ctgggctaaa gtaggaactt ccacgttgat 3060agagaaagtg
gctagtacca acagtaagaa tccttccctc agctcccatc aatgttacag 3120tgtttggtgg
aacagtaagg actggatctt ctcccaatga tgtgctaacc ttgatactga 3180gaatagcttg
ttgtatcctc ttacctccaa atcttcctgg tttgtatgat gacttagcca 3240ttcgaatctg
atagtcttgc tcatcaggac aagtatcatt gtacctcttg tagataacat 3300actttccctc
ttgcacagta tcactaggac tgttaggttt caaaccacca tagacagaga 3360accagactct
ggagtcaatg aagctaccac ctccaactcc tggatagtta gctacccaat 3420ccccaaacaa
tgtagtgaca tccaaatctt tctcatggta ttgaccatca aaacccaatc 3480tgccatgcac
catccttgta gggactgcag aattgtagtc ctcttcttca gtttctgtta 3540ctttactgca
tagcatatca cagcccaaag gtgtagcaga tacagagcaa cttttgcgat 3600tctgtgtatc
atcaagattg atactgcgaa gagttgagaa gaacactcta ccagtagcag 3660atgtacgaag
aactccaagt gctaagtact gataagagtg agaatggtca cgacagccag 3720atagaatgac
attgtgtgta tagcaatagt gtgtagcact catgtcaaat gagggtatcc 3780gagtgcaccc
actcccagtt gtgggtgcag gaatgaagtt cagatgttcc tggaaagctg 3840aaggatagaa
agatgtaaca tctgaagcat catctacaat gagttctttg ccaatacctc 3900caatgtagtc
tggatcatga attggggctc cccaacctga attgttggca gccccattga 3960tctgatagga
taggctggtg attgcattca tgatggtagt ttctgtgttg agtaatgcaa 4020gtggactttc
aagtgcaact tgtttgtaga ttcggtccac aacatcctgg ttggagccta 4080gggtactggt
aatcttctct tcagccctag agattcgtgt gggtatgcca actaagtctg 4140agggggtgct
tgctcccata gaatagagta atgatgcaac agagattgct agggtgacaa 4200cagtaagaaa
gagaatggct atccgaaaga taagcctcca agtgttcttg gcttccctct 4260catcattctc
aagagccact tgtgaaactg ctcggtccat ggtttggatc cacaaactta 4320caaatttctc
tgaagttgta tcctcagtac ttcaaagaaa atagcttaca ccaaattttt 4380tcttgttttc
acaaatgccg aacttggttc cttatatagg aaaactcaag ggcaaaaatg 4440acacggaaaa
atataaaagg ataagtagtg ggggataaga ttcctttgtg ataaggttac 4500tttccgccct
tacattttcc accttacatg tgtcctctat gtctctttca caatcaccga 4560ccttatcttc
ttcttttcat tgttgtcgtc agtgcttacg tcttcaagat tcttttcttc 4620gcctggttct
tctttttcaa tttctacgta ttcttcttcg tattctggca gtataggatc 4680ttgtatctgt
acattcttca tttttgaaca taggttgcat atgtgccgca tattgatctg 4740cttcttgctg
agctcacata atacttccat agtttttccc gtaaacattg gattcttgat 4800gctacatctt
ggataattac cttctggcgc gcctttgccc gggctttcct gcagggttta 4860aacttaatta
agcggccgat ccggtgagta atattgtacg gctaagagcg aatttggcct 4920gtagacctca
attgcgagct ttctaatttc aaactattcg ggcctaactt ttggtgtgat 4980gatgctgact
ggcaggatat ataccgttgt aatttgagct cgtgtgaata agtcgctgtg 5040tatgtttgtt
tgattgtttc tgttggagtg cagcccattt caccggacaa gtcggctaga 5100ttgatttagc
cctgatgaac tgccgagggg aagccatctt gagcgcggaa tgggaatgga 5160tcgaaccggg
agcacaggat gacgcctaac aattcattca agccgacacc gcttcgcggc 5220gcggcttaat
tcaggagtta aacatcatga gggaagcggt gatcgccgaa gtatcgactc 5280aactatcaga
ggtagttggc gtcatcgagc gccatctcga accgacgttg ctggccgtac 5340atttgtacgg
ctccgcagtg gatggcggcc tgaagccaca cagtgatatt gatttgctgg 5400ttacggtgac
cgtaaggctt gatgaaacaa cgcggcgagc tttgatcaac gaccttttgg 5460aaacttcggc
ttcccctgga gagagcgaga ttctccgcgc tgtagaagtc accattgttg 5520tgcacgacga
catcattccg tggcgttatc cagctaagcg cgaactgcaa tttggagaat 5580ggcagcgcaa
tgacattctt gcaggtatct tcgagccagc cacgatcgac attgatctgg 5640ctatcttgct
gacaaaagca agagaacata gcgttgcctt ggtaggtcca gcggcggagg 5700aactctttga
tccggttcct gaacaggatc tatttgaggc gctaaatgaa accttaacgc 5760tatggaactc
gccgcccgac tgggctggcg atgagcgaaa tgtagtgctt acgttgtccc 5820gcatttggta
cagcgcagta accggcaaaa tcgcgccgaa ggatgtcgct gccgactggg 5880caatggagcg
cctgccggcc cagtatcagc ccgtcatact tgaagctagg caggcttatc 5940ttggacaaga
agatcgcttg gcctcgcgcg cagatcagtt ggaagaattt gttcactacg 6000tgaaaggcga
gatcaccaag gtagtcggca aataatgtct aacaattcgt tcaagccgac 6060gccgcttcgc
ggcgcggctt aactcaagcg ttagagagct ggggaagact atgcgcgatc 6120tgttgaaggt
ggttctaagc ctcgtacttg cgatggcatt tcgatcgaaa ggggtacaaa 6180ttcccactaa
gcgctcgggg gctgagaaag cccagtaagg aaacaactgt aggttcgagt 6240cgcgagatcc
cccggaacca aaggaagtag gttaaacccg ctccgatcag gccgagccac 6300gccaggccga
gaacattggt tcctgtaggc atcgggattg gcggatcaaa cactaaagct 6360actggaacga
gcagaagtcc tccggccgcc agttgccagg ccgtaaaggt gagcagaggc 6420acgggaggtt
gccacttgcg ggtcagcacg gttccgaacg ccatggaaac cgcccccgcc 6480aggcccgctg
cgacgccgac aggatctagc gctgcgtttg gtgtcaacac caacagcgcc 6540acgcccgcag
ttccgcaaat agcccccagg accgccatca atcgtatcgg gctacctagc 6600agagcggcag
agatgaacac gaccatcagc ggctgcacag cgcctaccgt cgccgcgacc 6660cgcccggcag
gcggtagacc gaaataaaca acaagctcca gaatagcgaa atattaagtg 6720cgccgaggat
gaagatgcgc atccaccaga ttcccgttgg aatctgtcgg acgatcatca 6780cgagcaataa
acccgccggc aacgcccgca gcagcatacc ggcgacccct cggcctcgct 6840gttcgggctc
cacgaaaacg ccggacagat gcgccttgtg agcgtccttg gggccgtcct 6900cctgtttgaa
gaccgacagc ccaatgatct cgccgtcgat gtaggcgccg aatgccacgg 6960catctcgcaa
ccgttcagcg aacgcctcca tgggcttttt ctcctcgtgc tcgtaaacgg 7020acccgaacat
ctctggagct ttcttcaggg ccgacaatcg gatctcgcgg aaatcctgca 7080cgtcggccgc
tccaagccgt cgaatctgag ccttaatcac aattgtcaat tttaatcctc 7140tgtttatcgg
cagttcgtag agcgcgccgt gcgcccgagc gatactgagc gaagcaagtg 7200cgtcgagcag
tgcccgcttg ttcctgaaat gccagtaaag cgctggctgc tgaaccccca 7260gccggaactg
accccacaag gccctagcgt ttgcaatgca ccaggtcatc attgacccag 7320gcgtgttcca
ccaggccgct gcctcgcaac tcttcgcagg cttcgccgac ctgctcgcgc 7380cacttcttca
cgcgggtgga atccgatccg cacatgaggc ggaaggtttc cagcttgagc 7440gggtacggct
cccggtgcga gctgaaatag tcgaacatcc gtcgggccgt cggcgacagc 7500ttgcggtact
tctcccatat gaatttcgtg tagtggtcgc cagcaaacag cacgacgatt 7560tcctcgtcga
tcaggacctg gcaacgggac gttttcttgc cacggtccag gacgcggaag 7620cggtgcagca
gcgacaccga ttccaggtgc ccaacgcggt cggacgtgaa gcccatcgcc 7680gtcgcctgta
ggcgcgacag gcattcctcg gccttcgtgt aataccggcc attgatcgac 7740cagcccaggt
cctggcaaag ctcgtagaac gtgaaggtga tcggctcgcc gataggggtg 7800cgcttcgcgt
actccaacac ctgctgccac accagttcgt catcgtcggc ccgcagctcg 7860acgccggtgt
aggtgatctt cacgtccttg ttgacgtgga aaatgacctt gttttgcagc 7920gcctcgcgcg
ggattttctt gttgcgcgtg gtgaacaggg cagagcgggc cgtgtcgttt 7980ggcatcgctc
gcatcgtgtc cggccacggc gcaatatcga acaaggaaag ctgcatttcc 8040ttgatctgct
gcttcgtgtg tttcagcaac gcggcctgct tggcctcgct gacctgtttt 8100gccaggtcct
cgccggcggt ttttcgcttc ttggtcgtca tagttcctcg cgtgtcgatg 8160gtcatcgact
tcgccaaacc tgccgcctcc tgttcgagac gacgcgaacg ctccacggcg 8220gccgatggcg
cgggcagggc agggggagcc agttgcacgc tgtcgcgctc gatcttggcc 8280gtagcttgct
ggaccatcga gccgacggac tggaaggttt cgcggggcgc acgcatgacg 8340gtgcggcttg
cgatggtttc ggcatcctcg gcggaaaacc ccgcgtcgat cagttcttgc 8400ctgtatgcct
tccggtcaaa cgtccgattc attcaccctc cttgcgggat tgccccgact 8460cacgccgggg
caatgtgccc ttattcctga tttgacccgc ctggtgcctt ggtgtccaga 8520taatccacct
tatcggcaat gaagtcggtc ccgtagaccg tctggccgtc cttctcgtac 8580ttggtattcc
gaatcttgcc ctgcacgaat accagcgacc ccttgcccaa atacttgccg 8640tgggcctcgg
cctgagagcc aaaacacttg atgcggaaga agtcggtgcg ctcctgcttg 8700tcgccggcat
cgttgcgcca ctcttcatta accgctatat cgaaaattgc ttgcggcttg 8760ttagaattgc
catgacgtac ctcggtgtca cgggtaagat taccgataaa ctggaactga 8820ttatggcnnc
tcgaaattcc ctcggtcttg ccttgctcgt cggtgatgta cttcaccagc 8880tccgcgaagt
cgctcttctt gatggagcgc atggggacgt gcttggcaat cacgcgcacc 8940ccccggccgt
tttagcggct aaaaaagtca tggctctgcc ctcgggcgga ccacgcccat 9000catgaccttg
ccaagctcgt cctgcttctc ttcgatcttc gccagcaggg cgaggatcgt 9060ggcatcaccg
aaccgcgccg tgcgcgggtc gtcggtgagc cagagtttca gcaggccgcc 9120caggcggccc
aggtcgccat tgatgcgggc cagctcgcgg acgtgctcat agtccacgac 9180gcccgtgatt
ttgtagccct ggccgacggc cagcaggtag gccgacaggc tcatgccggc 9240cgccgccgcc
ttttcctcaa tcgctcttcg ttcgtctgga aggcagtaca ccttgatagg 9300tgggctgccc
ttcctggttg gcttggtttc atcagccatc cgcttgccct catctgttac 9360gccggcggta
gccggccagc ctcgcagagc aggattcccg ttgagcaccg ccaggtgcga 9420ataagggaca
gtgaagaagg aacacccgct cgcgggtggg cctacttcac ctatcctgcc 9480cggctgacgc
cgttggatac accaaggaaa gtctacacga accctttggc aaaatcctgt 9540atatcgtgcg
aaaaaggatg gatataccga aaaaatcgct ataatgaccc cgaagcaggg 9600ttatgcagcg
gaaaagatcc gtcgaccctt tccgacgctc accgggctgg ttgccctcgc 9660cgctgggctg
gcggccgtct atggccctgc aaacgcgcca gaaacgccgt cgaagccgtg 9720tgcgagacac
cgcggccgcc ggcgttgtgg ataccacgcg gaaaacttgg ccctcactga 9780cagatgaggg
gcggacgttg acacttgagg ggccgactca cccggcgcgg cgttgacaga 9840tgaggggcag
gctcgatttc ggccggcgac gtggagctgg ccagcctcgc aaatcggcga 9900aaacgcctga
ttttacgcga gtttcccaca gatgatgtgg acaagcctgg ggataagtgc 9960cctgcggtat
tgacacttga ggggcgcgac tactgacaga tgaggggcgc gatccttgac 10020acttgagggg
cagagtgatg acagatgagg ggcgcaccta ttgacatttg aggggctgtc 10080cacaggcaga
aaatccagca tttgcaaggg tttccgcccg tttttcggcc accgctaacc 10140tgtcttttaa
cctgctttta aaccaatatt tataaacctt gtttttaacc agggctgcgc 10200cctggcgcgt
gaccgcgcac gccgaagggg ggtgcccccc cttctcgaac cctcccggcc 10260cgctaacgcg
ggcctcccat ccccccaggg gctgcgcccc tcggccgcga acggcctcac 10320cccaaaaatg
gcaggccaag ctagcttgct tggtcgttcc ggtacgtacc gtgaacgtcg 10380gctcgattgt
acctgcgttc aaatactttg cgatcgtgtt gcgcgcctgc ccggtgcgtc 10440ggctgatctc
acggatcgac tgcttctctc gcaacgccat ccgacggatg atgtttaaaa 10500gtcccatgtg
gatcactccg ttgccccgtc gctcaccgtg ttggggggaa ggtgcacatg 10560gctcagttct
caatggaaat tatctgccta accggctcag ttc 10603
User Contributions:
comments("1"); ?> comment_form("1"); ?>Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
User Contributions:
Comment about this patent or add new information about this topic: