Patent application title: METHODS AND COMPOSITIONS FOR NERVE REGENERATION
Inventors:
Yimin Zou (Chicago, IL, US)
Assignees:
THE UNIVERSITY OF CHICAGO
IPC8 Class: AA61K3817FI
USPC Class:
4241721
Class name: Drug, bio-affecting and body treating compositions immunoglobulin, antiserum, antibody, or antibody fragment, except conjugate or complex of the same with nonimmunoglobulin material binds eukaryotic cell or component thereof or substance produced by said eukaryotic cell (e.g., honey, etc.)
Publication date: 2008-12-04
Patent application number: 20080299135
Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
Patent application title: METHODS AND COMPOSITIONS FOR NERVE REGENERATION
Inventors:
Yimin Zou
Agents:
MICHAEL BEST & FRIEDRICH LLP
Assignees:
THE UNIVERSITY OF CHICAGO
Origin: MADISON, WI US
IPC8 Class: AA61K3817FI
USPC Class:
4241721
Abstract:
Methods and compositions for modulating growth of a neuron with a Wnt, a
Wnt-like substance, and/or a chemical compound affecting a Wnt signaling
pathway are disclosed. Also disclosed are methods for identifying a
substance that modulates growth of a neuron by obtaining a candidate
substance and contacting the candidate substance with the neuron are
disclosed and methods for modulating growth of a neuron in a subject
using a Wnt, a Wnt-like substance, and/or a chemical compound affecting a
Wnt signaling pathway. The Wnt, Wnt-like substance, and/or chemical
compounds affecting a Wnt signaling pathway can be delivered to the
subject using gene therapy techniques. Also disclosed are pharmaceutical
compositions for modulating growth of a neuron in a mammal that include a
Wnt or a Wnt-like substance. Methods and compositions for inhibiting
growth of a neuron are also disclosed.Claims:
1. A method for modulating the directional growth of a mammalian neuron
comprising contacting the neuron with an inhibitor of frizzled.
2. The method of claim 1, wherein the frizzled inhibitor comprises a frizzled antibody.
3. The method of claim 2, wherein the frizzled antibody inhibits a frizzled selected from the group consisting of frizzled3, frizzled8, and frizzled9.
4. The method of claim 1, wherein the neuron is contacted with the inhibitor in a spinal cord.
5. The method of claim 4, wherein the inhibitor is provided as a concentration gradient.
6. The method of claim 5, wherein the concentration gradient is provided as a decreasing anterior-posterior concentration gradient along the spinal cord.
7. The method of claim 4, wherein the directional growth of the neuron occurs along the anterior-posterior axis of the spinal cord.
8. The method of claim 4, wherein the directional growth of the neuron is along the spinothalamic pathway.
9. The method of claim 4, wherein the spinal cord has been damaged.
10. The method of claim 1, wherein the neuron is a commissural neuron.
11. The method of claim 10, wherein the commissural neuron is contacted with the inhibitor post-midline crossing.
12. The method of claim 1, wherein the frizzled inhibitor comprises a sFRP.
13. The method of claim 12, wherein the sFRP is selected from sFRP1, sFRP2 and sFRP3.
14. The method of claim 1, wherein the neuron is further contacted with a neuronal growth inhibitor.
15. The method of claim 1, wherein the neuron is further contacted with a substance that blocks activity of a neuronal growth inhibitor.
16. The method of claim 1, wherein the neuron is a motor neuron or a sensory neuron.
17. The method of claim 1, wherein the neuron is a damaged neuron.
18. The method of claim 17, wherein the directional growth of the neuron facilitates regeneration of the neuron.
19. The method of claim 1, wherein the inhibitor repels neuronal growth.
20. The method of claim 1, wherein the inhibitor is provided as a pharmaceutical composition.
Description:
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001]This application is a divisional of U.S. application Ser. No. 10/847,972 filed May 17, 2004, which claims the benefit of U.S. Provisional Application No. 60/470,913 filed May 15, 2003, and both are incorporated herein by reference in their entireties.
BACKGROUND OF THE INVENTION
[0002]1. Field of the Invention
[0003]The present invention relates generally to the fields of molecular biology, cell biology, pharmacology, developmental neuroscience, neurology, neurosurgery and regenerative biology. More particularly, it concerns methods and compositions for modulating regeneration of a nerve cell using a Wnt, a Wnt-like substance, and/or a chemical compound affecting a Wnt signaling pathway. It also concerns methods and compositions for inhibiting growth of a neuron using inhibitors of neuronal growth that act via the Wnt signaling pathways, such as a Secreted Frizzled-Related Protein (sFRP), sFRP-like substance, Ryk, or Ryk-like substance.
[0004]2. Description of Related Art
[0005]The central nervous system (CNS) is connected by ascending sensory pathways and descending motor or regulatory pathways. In the CNS, somatosensory pathways ascend to the brain centers, and motor pathways controlling body movement descend from the brain to the spinal cord (Fitzgerald, 1996). The molecular mechanisms of axonal connections along the longitudinal axis of the CNS have remained a long-standing mystery.
[0006]Unlike the peripheral nervous system, damage to the central nervous system axons, such as spinal cord axons cannot be repaired, causing permanent impairment of neural function, such as in paralysis. The spinal cord serves important functions in the central nervous system. One such function is to allow communication of the body and the brain. The nerve fibers within the spinal cord carry messages to and from the brain to other parts of the body. In general sensory information from the body travels along the spinal cord up to the brain and instruction from the brain, such as motor command, travels along the spinal cord down from the brain. Thus, the spinal cord can be compared to a telephone cable, which connects the central office (brain) to the individual homes.
[0007]The term spinal cord injury refers to any injury of the neurons within the spinal canal. Spinal cord injury can occur from either trauma or disease to the vertebral column or the spinal cord itself. Most spinal cord injuries are the result of trauma to the vertebral column causing a fracture of the bone, or tearing of the ligaments with displacement of the bony column producing a pinching of the spinal cord. The majority of broken necks and broken backs, or vertebral fractures, do not cause any spinal cord damage; however, in 10-14% of the cases where a vertebral trauma has occurred, the damage is of such severity it results in damage to the spinal cord.
[0008]Spinal cord injury primarily occurs in young men with the greatest number of injuries occurring in the 16-30 age group. Patients with a spinal cord injury are designated as having tetraplegia (preferred to quadriplegia) or paraplegia. Tetraplegia refers to injuries to the cervical spinal cord and paraplegia refers to injuries below the cervical spinal cord. Patients with tetraplegia are slightly more common (51.7%) than patients with paraplegia. The majority of spinal cord injuries, about 37.4%, are sustained during a motor vehicle accident. Acts of violence are the second most common cause at 25.9%, falls are third at 21.5% and sports injuries are fourth at 7.1%.
[0009]It is estimated that the annual incidence of spinal cord injury (SCI), not including those who die at the scene of the accident, is approximately 40 cases per million population in the U.S., or approximately 11,000 new cases each year. The number of people in the U.S. who are alive today and who have SCI has been estimated to be between 721 and 906 per million population. This corresponds to between 183,000 and 230,000 persons.
[0010]Treatment options for patients with spinal cord injuries are limited. Often, patients with SCI are left with severe, permanent disabilities. A major reason for the limited availability of treatment options is the fact that there is little known about factors that can control and modulate nerve growth and regeneration following spinal cord injury. For example, the precise molecular mechanisms that guide axons along the anterior-posterior (A-P) axis of the spinal cord are unknown.
[0011]Axonal connections are patterned along the A-P and dorsal-ventral (D-V) neuraxes, wiring a large number of neurons into an intricate network. Axon guidance along the D-V axis has been a major focus of study in a number of experimental systems in recent years (Tessier-Lavigne and Goodman, 1996; Dickson, 2002). Much work has concentrated on the question of how axons are guided towards and away from the ventral midline and how midline crossing is regulated. Guidance molecules, such as Netrin-1 and members of the Slit and Semaphorin families, play pivotal roles in the dorsal-ventral guidance of axons (Tessier-Lavigne and Goodman, 1996; Dickson, 2002). The nature of the anterior-posterior guidance cues remains an enigma. Four classes of axon guidance molecules have been described (Tessier-Lavigne and Goodman, 1996): long-range attractants, long-range repellents, contact-mediated attractants and contact-mediated repellents. It is currently unknown whether a general gradient of attractant(s) or repellent(s) along the anterior-posterior axis guides axons to grow along this axis, or whether this guidance is mediated by more regional or segmental cues. The question of axon guidance along the A-P axis is of particularly interest in the spinal cord, where multiple classes of axons project either anteriorly or posteriorly along the length of the spinal cord. For example, somatosensory pathways ascend from the spinal cord to the brain and motor pathways descend from the brain to the spinal cord, with both the ascending and descending pathways carrying topographic information (FitzGerald, 1996).
[0012]The dorsal spinal cord commissural neurons form several ascending somatosensory pathways, such as the spinothalamic tracts, which send pain and temperature sensations to the brain (Ramon y Cajal, 1893; Altman and Bayer, 1984). The cell bodies of commissural neurons are located in the dorsal spinal cord. During embryonic development, commissural neurons project axons to the ventral midline. Once they reach the floor plate, they cross the midline and enter the contralateral side of the spinal cord. After midline crossing, commissural axons make a remarkably sharp anterior turn towards the brain (Ramon y Cajal, 1893; Altman and Bayer 1984; Tessier-Lavigne, 1994). All dorsal spinal cord commissural axons along the entire anterior-posterior length of the spinal cord project anteriorly after midline crossing. The initial ventral growth of the commissural axons is controlled by a gradient of a diffusible chemoattractant, Netrin-1 (Serafini et al., 1994; Kennedy et al., 1994; Serafini et al., 1996). As the axons cross the midline, they lose responsiveness to Netrin-1 (Shirasaki et al., 1998). Interestingly, while losing responsiveness to Netrin-1 during midline crossing, commissural axons gain responsiveness to several chemorepellents, which are located in the midline and the ventral spinal cord (Zou et al., 2000). These repellents help to expel the axons from the midline and to turn axons from their dorsal-ventral trajectory into their longitudinal pathways along the anterior-posterior axis by preventing axons from overshooting into the contralateral ventral spinal cord and recrossing the floor plate; the axons thus become "squeezed" into their longitudinal pathway (Zou et al., 2000). The expression pattern of the Slits and Semaphorins identified in these studies have been examined, but no anterior-posterior gradient of these chemorepellents in the spinal cord has been identified, suggesting that these repellents do not control anterior-posterior pathfinding.
[0013]Wnt polypeptides are secreted cysteine-rich glycosylated polypeptides that play a role in the development of a wide range of organisms. The Wnt family of polypeptides bind to an extracellular domain of a family of cell surface proteins called Frizzled receptors, and may play a role in embryonic induction, generation of cell polarity, and specification of cell fate.
[0014]Wnts are encoded by a large gene family, whose members have been found in round worms, insects, cartilaginous fish and vertebrates (Sidow, 1994). Wnts are thought to function in a variety of developmental and physiological processes since many diverse species have multiple conserved Wnt genes (McMahon, 1992; Nusse and Varmus, 1992). The Wnt growth factor family includes at least 18 genes identified in the human by cDNA cloning (see, e.g., Vant Veer et al., 1984; Miller, 2001).
[0015]Wnts may play a role in local cell signaling and neurogenesis. Biochemical studies have shown that much of the secreted Wnt protein can be found associated with the cell surface or extracellular matrix rather than freely diffusible in the medium (Papkoff and Schryver, 1990; Bradley and Brown, 1990). Studies of mutations in Wnt genes have indicated a role for Wnts in growth control and tissue patterning. In Drosophila, wingless (wg) encodes a Wnt gene (Rijsenijk et al., 1987) and wg mutations alter the pattern of embryonic ectoderm, neurogenesis, and imaginal disc outgrowth (Morata and Lawrence, 1977; Baker, 1988; Klingensmith and Nusse, 1994). Knock-out mutations in mice have shown Wnts to be essential for brain development (McMahon and Bradley, 1990; Thomas and Cappechi, 1990). However, a role for Wnts in mammalian directional axonal growth regulation in the spinal cord has not previously been suggested or considered.
[0016]The identification of modulators of neuronal growth and regeneration following SCI could be applied in new forms of treatment of patients with this debilitating condition. The identification of modulators of neuronal growth and regeneration could also be applied in the treatment of patients with other disorders involving neuronal dysfunction, such as neurodegenerative diseases. Agents that can promote axonal growth along the A-P axis following injury to the spinal cord may be applied to help prevent the permanent paralysis that is often associated with SCI. Therefore, there is a need for better treatments of SCI, and a greater understanding of modulators of neuronal growth and regeneration might lead to improved methods of treatment of this devastating disorder.
SUMMARY OF THE INVENTION
[0017]The inventor has found that Wnt proteins play a general role in anterior-posterior patterns of CNS axons, which connect the brain and the spinal cord.
[0018]The invention disclosed herein is based on the discovery of a molecular regulatory system involving Wnt proteins that is involved in the normal formation of the spinal cord axon connection. A chemoattractant gradient exists inside the spinal cord, and this chemoattractant gradient guides the anterior projection of post-crossing spinal cord commissural neurons along the A-P axis towards the brain during embryogensis. In particular, it has been discovered that several Wnt proteins can stimulate the extension of post-crossing but not pre-crossing commissural axons in the spinal cord. Wnt4 was found to be expressed in a decreasing A-P gradient in the floor plate of the spinal cord. sFRPs, inhibitors of Wnts, were found to disrupt the A-P pathfinding of post-crossing spinal cord commissural neurons. However, Wnt4 protein was found to rescue the anterior turn of the misrouting axons and also reorient axons posteriorly, suggesting that Wnt4 plays an instructive role in orienting directional axonal growth. In addition, commissural axons infz3 knockout mice were found to display A-P guidance defects after midline crossing. In view of these findings, Wnt, Wnt-like substances, and/or chemical compounds affecting a Wnt signaling pathway can be used as novel agents to modulate neuronal growth, and can be used in new forms of treatment of diseases and conditions associated with neuronal dysfunction, such as SCI (Lyuksyotova et al., 2003).
[0019]The inventor has further found that a different set of Wnt proteins pattern the connections of corticospinal tract (CST) axons projecting along the opposite direction by a repulsive mechanism. CST axons project from the motor cortex of the brain to the spinal cord motor circuits and send voluntary movement signals from the brain to the body. Several Wnt genes were found to be expressed at the dorsal funiculus in an anterior-to-posterior decreasing gradient at the cervical spinal cord, where CST axons first enter the spinal cord and a anterior-to-posterior increasing gradient at the lumbar spinal cord level, forming a "half-pipe" gradient. Wnt1 and Wnt5a can repel CST axons in collagen gel assays. A repulsive Wnt receptor, Ryk (Oshikawa et al., 2003; Halford et al., 2000), is expressed in the CST axons and can be detected at the pyramidal decussation and in the dorsal funiculus. Antibodies against the ectodomain of Ryk can block the repulsion of Wnt1. Finally, intrathecal injection of a Wnt inhibitor, secreted Frizzled related protein 2 (sFRP2), at the rostral cervical level (C1 and C2), can inhibit the posterior growth of CST axons in vivo, leading to weaker grip strength.
[0020]The inventor has also found that Wnts play important roles in patterning the synaptic connections once they reach their target. This process of target selection ensures the specific neuron to neuron connection and is essential to the development of the functional circuits throughout the nervous system. Therefore, Wnts can be used to ensure specific synaptic reconnection in repair damaged neural circuits.
[0021]Certain embodiments of the present invention are generally concerned with methods for modulating growth of a neuron comprising contacting the neuron with a Wnt, a Wnt-like substance, and/or a chemical compound affecting a Wnt signaling pathway. The definitions of Wnt, Wnt-like substance, and chemical compound affecting a Wnt signaling pathway are discussed in detail in the specification below.
[0022]In the context of the invention, the terms "contact" or "contacting" are defined to mean any manner in which a compound is brought into a position where it can mediate, modulate, or inhibit the growth of a neuron. "Contacting" can comprise injecting a diffusable or non-diffusable substance into the neuron or an area adjacent a neuron. "Contacting" can comprise placing a nucleic acid encoding a compound into or close to a neuron or non-neuronal cell in a manner such that the nucleic acid is expressed to make the compound in a manner in which it can act upon the neuron. Those of skill in the art, following the teachings of this specification, will be able to contact neurons with substances in any manner.
[0023]The methods for modulating growth of a neuron may, in certain embodiments, be methods for stimulating growth of a neuron, methods for regenerating a damaged neuron, or methods for guiding growth of a neuron along the anterior-posterior axis. In other embodiments, the methods for modulating growth of a neuron are further defined as methods for directionally orienting axon growth of a neuron between the spinal cord and the brain.
[0024]The neuron to be modulated may be any neuron. However, in certain embodiments, the neuron is a neuron in the spinal cord that has been damaged. For example, the spinal cord may have been damaged by traumatic spinal cord injury. The damage may have resulted in impaired function of the neuron.
[0025]In certain embodiments, the method for modulating growth of a neuron is a method for modulating growth of a neuron in a subject. Although any subject is contemplated by the present invention, in certain embodiments the subject may be a patient with a disorder of the spinal cord. The disorder of the spinal cord may be any disorder, such as a traumatic spinal cord injury. The traumatic spinal cord injury may or may not have resulted in paralysis of the subject. In further embodiments, the patient is a patient with a neurodegenerative disease.
[0026]The neuron to be modulated can be a sensory or a motor neuron. In certain embodiments, the neuron is contacted with a Wnt, a Wnt-like substance, and/or a chemical compound affecting a Wnt signaling pathway that further involves exposing the neuron to a gradient of the Wnt, Wnt-like substance, and/or chemical compound affecting a Wnt signaling pathway. The gradient may be in the spinal cord, such as a decreasing anterior-posterior gradient within the spinal cord. In other embodiments, exposing the neuron to the gradient involves stimulating directionally-oriented axon growth of the neuron along the anterior-posterior axis. Any direction of axon growth is contemplated by the present invention. In certain embodiments, the axon growth is directed from the spinal cord to the brain, such as in the growth of neurons in ascending somatosensory pathways. In other embodiments, the axon growth is directed from the brain to the spinal cord, such as in the growth of neurons in descending motor pathways or other regulatory pathways. In further embodiments, the axon growth is directed along the spinothalamic pathway.
[0027]Any Wnt is contemplated by the present invention. A detailed discussion of Wnts is provided in the specification below. For example, the Wnt protein may be Wnt1, Wnt2, Wnt2b, Wnt3, Wnt3a, Wnt4, Wnt5a, Wnt5b, Wnt 6, Wnt7a, Wnt7b, Wnt8a, Wnt8b, Wnt9a, Wnt9b, Wnt10a, Wnt10b, Wnt11, or Wnt16 protein. One of skill in the art would be familiar with the range of Wnts available that are contemplated by the present invention. In certain embodiments, the Wnt is a Wnt1, Wnt4, Wnt5a, Wnt6, or Wnt7b protein. In certain embodiments, the Wnt protein will be a mammalian Wnt protein, for example a human or murine Wnt protein, or a homolog thereof from another vertebrate species.
[0028]In further embodiments, the Wnt-like substance is a Wnt polypeptide. Any Wnt polypeptide is contemplated by the present invention. For example, the Wnt polypeptide may be a Wnt1, Wnt2, Wnt2b, Wnt3, Wnt3a, Wnt4, Wnt5a, Wnt5b, Wnt 6, Wnt7a, Wnt7b, Wnt8a, Wnt8b, Wnt9a, Wnt9b, Wnt10a, Wnt10b, Wnt11, or Wnt16 polypeptide. One of skill in the art would be familiar with the range of Wnt polypeptides available that are contemplated by the present invention. In certain embodiments, the Wnt polypeptide is a Wnt1, Wnt4, Wnt5a, Wnt6, or Wnt7b polypeptide. Wnt polypeptides are discussed in greater detail in the specification below. In certain embodiments, the Wnt polypeptide will be a mammalian Wnt protein, for example a human or murine Wnt polypeptide, or a homolog thereof from another vertebrate species.
[0029]In further embodiments, the Wnt-like substance is a Wnt peptide. Any Wnt peptide is contemplated by the present invention. For example, the Wnt peptide may be a Wnt1, Wnt2, Wnt2b, Wnt3, Wnt3a, Wnt4, Wnt5a, Wnt5b, Wnt 6, Wnt7a, Wnt7b, Wnt8a, Wnt8b, Wnt9a, Wnt9b, Wnt10a, Wnt10b, Wnt11, or Wnt16 peptide. One of skill in the art would be familiar with the range of Wnt peptides available that are contemplated by the present invention. In certain embodiments, the Wnt peptide is a Wnt1, Wnt4, Wnt5a, Wnt6, or Wnt7b peptide. Wnt peptides are discussed in greater detail in the specification below. In certain embodiments, the Wnt protein will be a mammalian Wnt peptide, for example a human or murine Wnt peptide, or a homolog thereof from another vertebrate species.
[0030]In other embodiments, the Wnt-like substance is a mimetic of Wnt or a mutant Wnt. The definitions of mimetic Wnt and mutant Wnt are discussed in the specification below. Any Wnt mimetic is contemplated by the present invention. For example, the Wnt mimetic may be a Wnt1, Wnt2, Wnt2b, Wnt3, Wnt3a, Wnt4, Wnt5a, Wnt5b, Wnt 6, Wnt7a, Wnt7b, Wnt8a, Wnt8b, Wnt9a, Wnt9b, Wnt10a, Wnt10b, Wnt11, or Wnt16 mimetic. One of skill in the art would be familiar with the range of Wnt mimetics available that are contemplated by the present invention. In certain embodiments, the Wnt mimetic is a Wnt1, Wnt4, Wnt5a, Wnt6, or Wnt7b mimetic. In certain embodiments, the Wnt mimetic will be a mammalian Wnt mimetic, for example a human or murine Wnt mimetic, or a homolog thereof from another vertebrate species. Any Wnt mutant is contemplated by the present invention. For example, the Wnt mutant may be a Wnt1, Wnt2, Wnt2b, Wnt3, Wnt3a, Wnt4, Wnt5a, Wnt5b, Wnt 6, Wnt7a, Wnt7b, Wnt8a, Wnt8b, Wnt9a, Wnt9b, Wnt10a, Wnt10b, Wnt11, or Wnt16 mutant. One of skill in the art would be familiar with the range of Wnt mutants available that are contemplated by the present invention. In certain embodiments, the Wnt mutant is a Wnt1, Wnt4, Wnt5a, Wnt6, or Wnt7b mutant. In certain embodiments, the Wnt mutant will be a mammalian Wnt mutant, for example a human or murine Wnt mutant, or a homolog thereof from another vertebrate species. In other embodiments, the Wnt-like substance is a small molecule.
[0031]Further embodiments of the present invention involve use of chemical compounds affecting a Wnt signaling pathway to modulate growth of a neuron. The definition of such chemical compounds is described in the specification below. One of ordinary skill in the art would be familiar with the wide range of such compounds available which can modulate the Wnt signaling pathway. For example, in certain embodiments, the chemical compound affecting a Wnt signaling pathway is lithium.
[0032]The Wnt, Wnt-like substance, and/or chemical compound affecting a Wnt signaling pathway may include a fused amino acid sequence that is designed to facilitate incorporation of the polypeptide into the intracellular compartment of a cell. For example, the Wnt-like substance may include a polypeptide encoding an amino acid TAT sequence from HIV. In another example, the Wnt-like substance may include a polypeptide encoding an Antp amino acid sequence. In another example, the Wnt-like substance may include a polypeptide encoding a VP22 amino acid sequence from HSV.
[0033]In certain embodiments, the Wnt, Wnt-like substance, and/or chemical compound affecting a Wnt signaling pathway further includes an expression cassette comprising a promoter, active in a cell, operably linked to a polynucleotide encoding the Wnt, Wnt-like substance, and/or chemical compound affecting a Wnt signaling pathway. For example, the polypeptide may be a Wnt1, Wnt2, Wnt2b, Wnt3, Wnt3a, Wnt4, Wnt5a, Wnt5b, Wnt 6, Wnt7a, Wnt7b, Wnt8a, Wnt8b, Wnt9a, Wnt9b, Wnt10a, Wnt10b, Wnt11, or Wnt16 polypeptide. In certain embodiments, the Wnt polypeptide is a Wnt1, Wnt4, Wnt5a, Wnt6, or Wnt7b polypeptide. In other embodiments, the expression cassette is carried in a viral vector. Although any viral vector is contemplated by the present invention, examples include an adenoviral vector, a retroviral vector, an adeno-associated viral vector, a vaccinia viral vector, or a pox viral vector. In other embodiments, the expression cassette is carried in a nonviral vector, such as a liposome. One of skill in the art would be familiar with a wide range of viral and nonviral vectors available to be of use in the present invention.
[0034]Any promoter is contemplated for use in the present invention, as long as it facilitates expression of the polynucleotide. One of skill in the art would be familiar with the wide range of promoters available. For example, the promoter may be a constitutive promoter, an inducible promoter, or a tissue-specific promoter.
[0035]Certain embodiments of the present invention involve obtaining the Wnt, Wnt-like substance, and/or chemical compound affecting a Wnt signaling pathway from media of cultured cells. Although any cultured cells are contemplated by the present invention, in certain embodiments the cultured cells comprise an expression cassette including a promoter, active in the cultured cells, operably linked to a polynucleotide encoding Wnt, a Wnt-like substance, and/or a chemical compound affecting a Wnt signaling pathway. The characteristics of expression cassettes that have been previously discussed above apply to these embodiments of the present invention.
[0036]Further embodiments of the present invention provide for methods of inhibiting growth of a neuron. In certain embodiments, these methods involve contacting the neuron with a mutant Wnt.
[0037]Additional embodiments of the present invention include methods for identifying a substance that modulates growth of a neuron, including: (a) obtaining a candidate substance; (b) contacting said candidate substance with said neuron; and (c) measuring modulation of growth of said neuron. In certain embodiments, an explant assay is used in the methods for identifying a substance that modulates growth of a neuron. For example, the explant assay may involve use of cultured spinal cord. Any method to measure modulation of neuronal growth is contemplated by the present invention. However, in certain embodiments anterior turning of axons of the neuron is measured.
[0038]Any candidate substance is contemplated by the present invention. For example, the candidate substance may include a protein, a polypeptide, a peptide, mimetic, mutant, or a small molecule as described above. In a certain embodiments, the candidate substance is a Wnt-like substance, such as a Wnt peptide. Any Wnt peptide is contemplated by the present invention. For example, the Wnt peptide may be a Wnt1 peptide, a Wnt3 peptide, a Wnt4 peptide, a Wnt5a peptide, a Wnt6 peptide, or a Wnt7b peptide. In certain embodiments, the Wnt peptide is a mimetic of Wnt, such as a mimetic of Wnt1, a mimetic of Wnt3, a mimetic of Wnt4, a mimetic of Wnt5a, a mimetic of Wnt6, or a mimetic of Wnt7b. In a further embodiment, the Wnt-like substance is a mimetic of Wnt4. Alternatively, the Wnt-like substance may be a mutant Wnt, such as a mutant Wnt1 polypeptide, a mutant Wnt3 polypeptide, a mutant Wnt4 polypeptide, a mutant Wnt5a polypeptide, a mutant Wnt6 polypeptide, or a mutant Wnt7b polypeptide. In still further embodiments, the Wnt-like substance is a small molecule. In other embodiments, the chemical compound affecting a Wnt signaling pathway is a chemical compound that functionally or structurally resembles lithium.
[0039]Any method of measuring growth of a neuron is contemplated by the present methods for identifying modulators of nerve growth. These methods have been discussed above. For example, measuring modulation of growth of a neuron may further involve measuring stimulation of growth of the neuron, measuring regeneration of a damaged neuron, or measuring growth of said neuron along the anterior-posterior axis. In addition, these methods also involve method for directionally orienting axon growth of the neuron between the spinal cord and the brain.
[0040]The present invention also includes methods of modulating growth of a neuron in a subject, including: (a) providing a composition that includes a Wnt, a Wnt-like substance, and/or a chemical compound affecting a Wnt signaling pathway; and a pharmaceutical preparation suitable for delivery to the subject; and (b) administering the composition to the subject. The methods for modulating neuron growth of the present invention contemplate measurement of neuronal growth by any known means, as discussed above. For example, the method of modulating neuron growth may be defined as a method of promoting growth and regeneration of a neuron in a subject, a method of promoting axon growth and regeneration in a subject, or a method of promoting directionally-oriented axon growth in a subject. Directionally-oriented axon growth may be along the anterior-posterior axis such as from the spinal cord to the brain, or from the brain to the spinal cord.
[0041]The methods for modulating neuron growth in a subject contemplated by the present invention also include methods of treating a subject with a spinal cord disorder. Any spinal cord disorder is contemplated by the present invention. For example, the spinal cord disorder may be a traumatic spinal cord disorder, a disorder of motor and/or sensory neurons, a neurodegenerative disorder, or a disorder resulting in paralysis.
[0042]The methods of the present invention also contemplate exposing the neuron to a gradient of said Wnt, said Wnt-like substance, and/or said chemical compound affecting a Wnt signaling pathway. As discussed above, the gradient may be in the spinal cord, such as a decreasing gradient along the anterior-posterior axis.
[0043]Any Wnt, Wnt-like substance, and chemical compound affecting a Wnt signaling pathway, as discussed above and in the specification below, is contemplated by the present methods of modulating neuron growth in a subject. Mimetics and mutants of Wnts and Wnt-like substances are contemplated by the present invention, as are embodiments wherein the Wnt or Wnt-like substance further comprises an expression cassette comprising a promoter, active in a cell, operably linked to a polynucleotide encoding the Wnt or the Wnt-like substance. These expression cassettes have been discussed above, and are discussed in greater detail in later sections of this specification.
[0044]In certain embodiments, administering the composition of Wnt, Wnt-like substance, and/or chemical compound affecting a Wnt signaling pathway involves contacting the composition with the spinal cord of the subject. In certain embodiments, a gradient of the Wnt, Wnt-like substance, and/or chemical compound affecting a Wnt signaling pathway is created along the anterior-posterior axis. For example, the gradient may be between the spinal cord and the brain, such as a decreasing anterior-posterior gradient. In certain embodiments, the nerve cell is contacted with a modulator of neuronal growth identified by one of the previously described methods.
[0045]Certain embodiments of the present invention pertain to pharmaceutical compositions for modulating growth of a neuron in a mammal, including: (a) a Wnt, a Wnt-like substance, and/or a chemical compound affecting a Wnt signaling pathway; and (b) a pharmaceutical preparation suitable for delivery to the mammal. Neuronal growth may be modulated by any of the methods discussed above. In certain embodiments, the mammal is a human, such as a patient with a spinal cord disorder. Any Wnt, Wnt-like substance, and/or chemical compound affecting a Wnt-signaling pathway, as discussed above, is contemplated by the present invention. In certain embodiments, the composition comprises an expression cassette comprising a promoter, active in a cell, operably linked to a polynucleotide encoding the Wnt, the Wnt-like substance, and/or the chemical compound affecting a Wnt signaling pathway. Expression cassettes have been discussed above in the context of other embodiments of the present invention.
[0046]Additional embodiments of the present invention involve methods of inhibiting or controlling the growth of a neuron in a subject, by administering an inhibitor of a Wnt to the subject. In some cases, that inhibitor may be an sFRP, a Ryk protein, or an analog thereof. In general some such methods include: (a) providing a composition that includes an sFRP, an sFRP-like substance, a Ryk or a Ryk-like substance and a pharmaceutical preparation suitable for delivery to the subject; and (b) administering said composition to the subject. sFRPs are compounds that can affect a Wnt signaling pathway by binding to Wnt proteins with high affinity and blocking the interaction of Wnts with their receptors, the Frizzleds. sFRPs and sFRP-like substances are defined and discussed in detail below.
[0047]In certain embodiments, the composition comprises an sFRP protein. sFRPs are diffusable proteins that bind and modulate Wnts. Any sFRP protein from any species is contemplated by the present invention. For example, the sFRP protein may be sFRP1 protein, sFRP2 protein, or sFRP3 protein. In other embodiments, the sFRP-like substance is an sFRP polypeptide. For example, the sFRP polypeptide may be sFRP1 polypeptide, sFRP2 polypeptide, or sFRP3 polypeptide. In other embodiments, the sFRP-like substance is a peptide, such as sFRP1 peptide, sFRP2 peptide, or sFRP3 peptide. In further embodiments, the sFRP-like substance is a mutant sFRP, such as a mutant sFRP1 polypeptide, a mutant sFRP2 polypeptide, or a mutant sFRP3 polypeptide. In still further embodiments, the sFRP-like substance includes a small molecule that is functionally similar to a sFRP.
[0048]In other embodiments, the composition comprises a Ryk protein. Ryk is a receptor on neurons that binds Wnts and mediates repulsion of neurons in response to Wnts. Any Ryk protein or homolog from any species is contemplated by the present invention, for example, Drosphila Derailed protein may be employed in some embodiments. For example, the Ryk or Ryk-like substance may be a Ryk protein, polypeptide, peptide, mutant, or mimetic. In still further embodiments, the Ryk-like substance includes a small molecule that is functionally similar to a Ryk.
[0049]Other embodiments of the invention involve the contacting of a neuron with a combination of a Wnt and another substance, in order to provide a combination therapy. Such embodiments of the invention are important because, as discussed herein, the regeneration of neurons into a properly functioning spinal cord will often involve a combination of directional and other clues.
[0050]In some embodiments, one will wish to contact a neuron with a substance that blocks activity of a neuronal growth inhibitor. Such neuronal growth inhibitors include the myelin inhibitors Nogo, MAG, and Omgp, which have been shown to inhibit the growth of sensory neurons. Further, as discussed herein, Wnts can, if expressed in the adult spinal cord, inhibit the proper growth of CST motor neurons. In this regard, there are some Wnts that are expressed in normal adult spinal cords, and a variety of Wnts that may be is abnormally expressed in the neuron upon neuronal injury, as discussed below. In some embodiments of the invention, the substance that blocks the activity of the neuronal growth inhibitor is an antibody directed against a receptor for the inhibitor on the neuron or against the inhibitor itself. For example, such an antibody can be directed against a Wnt, Nogo, MAG, or OMgp. In some preferred embodiments, the antibody is directed against Wnt5a, Wnt8, or a Wnt that is expressed abnormally in the neuron due to injury, or against a receptor of any such Wnt. In other cases, the substance that blocks activity of a neuronal growth inhibitor is a Ryk, Ryk-like substance, sFRP or sFRP-like substance. In some preferred embodiments, one will want to block the activity of two or more inhibitors in the course of treating a neuron, spinal cord, and/or patient. For example, in order to allow an injured spinal cord comprising both injured sensory and injured motor neurons to regenerate in an appropriate manner, those of skill will understand that there may be a need to apply a compound to block the myelin inhibitors and prevent them from inhibiting the growth of sensory neurons, while also applying a compound to block Wnt inhibition of the growth of motor neurons.
[0051]The instant invention also involves contacting neurons with combinations of at least one Wnt and at least one other substance that attracts or repels neuronal growth. In some embodiments, the at least one other substance will be a substance attracts neuronal growth, for example, but not limited to a Wnt, Netrin, Shh, Cell adhesion molecule, Ig superfamily member, Cadherin, Integrin, EphrinB, ECM molecule, or HGF. In some embodiments, the at least one other substance will be a substance that repels neuronal growth, for example but not limited to, a Semaphorins, Netrin, Slit, Wnt, BMP, Ephrin, or member of the Ig superfamily. In many embodiments, contacting said neuron with a substance that attracts or repels neuronal growth will comprise exposing said neuron to a gradient of said substance. And, in some embodiments, the neuron will be exposed to a gradient of at least two such substances. In some cases, it will be beneficial to apply inhibitors of these substances that attract or repel neuronal growth at various portions of a regenerating spinal cord, in order to control the growth of the spinal cord, such inhibitors can be small molecules, peptides, proteins, or polypeptides that bind the substance, antibodies directed against the substance or a receptor of the substance, etc.
[0052]Some embodiments will involve the exposure of the neuron to a gradient of an attractive Wnt, some will involve exposure of the neuron to a gradient of a repulsive Wnt, some will involve exposure of the neuron to gradients of both attractive and repulsive Wnts. Attractive Wnts can include, but not be limited to, Wnt1, Wnt4, Wnt5a, Wnt 6, and Wnt7. Repulsive Wnts can include, but not be limited to Wnt5a or Wnt1. Those of skill in the art will be able to determine attractive and repulsive Wnts following the teachings herein, and will understand that the same Wnt may have an attractive property in regard to some contexts or some types of neurons and a repulsive property in regard to other contexts or types of neurons.
[0053]In some cases, it will be beneficial to apply one or more Wnt to the site of a spinal cord injury, such that the Wnt(s) will provide attractive guidance to those neurons that need to be attracted to the site of injury during regeneration and repellant guidance to those neurons that need to grow away from the site of injury during regeneration. In this regard, Wnt(s) applied at the site of an injury will provide directional guidance to axonal growth and cause sensory neurons to grow up through the site of the injury and repel motor neurons to grow down through the site of the injury. Further, in this embodiment, it may be beneficial to inhibit the Ryk pathway at the site of the injury so that motor neurons growing through the site of the injury are not inhibited by any Wnts present in the injury site, whether those Wnts are applied to the injury site, or expressed there as a result of normal adult Wnt expression or injury-induced Wnt expression. One may also apply a blocker of myelin inhibitors to the injury site, to prevent such inhibitors from impacting the growth of sensory neurons through the site.
[0054]Of course, combinations of Wnts, substances that block inhibitors of neuronal growth, and/or substances that attract or repel neuronal growth can be determined by those of skill in the art following the teaching contained herein. These various components of these combinations may be administered simultaneously, or separated by time. Individual components may be administered a single time or in a series of administrations. They may be administered in a single pharmaceutical composition, or in separate compositions. Those of skill in the art will be able to follow the teachings of this specification to determine appropriate dosage regimes and schedules of the various active agents.
[0055]Other embodiments of the invention involve pharmaceutical compositions comprising at least one Wnt, Wnt-like substance, or compound affecting a Wnt signaling pathway in combination with at least one substance that blocks an inhibitor of neuronal growth, and/or substance that attracts or repels neuronal growth. Further, kits comprising combinations of these various components, in separate or single containers are also within the scope of the invention.
[0056]As used herein the specification, "a" or "an" may mean one or more. As used herein in the claim(s), when used in conjunction with the word "comprising", the words "a" or "an" may mean one or more than one. As used herein "another" may mean at least a second or more.
[0057]The term "therapeutically effective" as used herein refers to an amount of a compound required to effect neuronal growth in the context of the manners described herein.
BRIEF DESCRIPTION OF THE DRAWINGS
[0058]The following drawings form part of the present specification and are included to further demonstrate certain aspects of the present invention. The invention may be better understood by reference to one or more of these drawings in combination with the detailed description of specific embodiments presented herein.
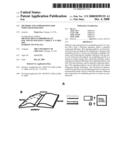
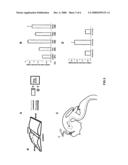
[0059]FIG. 1A, FIG. 1B, FIG. 1C, FIG. 1D, FIG. 1E. Diffusible cue(s) guides commissural axons along the anterior-posterior axis. FIG. 1A: Transverse section of an E13 rat spinal cord showing the dorsal-ventral trajectory (solid line) and the anterior-posterior trajectory (dashed line) of commissural axons. FIG. 1B: "Open-book" view of an E13 rat spinal cord showing midline-crossing and anterior turning of commissural axons. The subpopulation of commissural axons represented by the dashed line project anteriorly along a medial pathway, close to the floor plate (the ventral funiculus). The subpopulation of commissural axons represented by the solid line project along the floor plate initially, but gradually fan out to occupy more lateral positions (the lateral funiculus). Both populations project anteriorly immediately after midline crossing and were often observed in the DiI injections. FIG. 1C: A gradient of diffusible guidance cue(s) might be disrupted when the explants are cut shorter, causing misrouting of commissural axons along the A-P axis. FIG. 1D: A gradient of nondiffusible guidance cue(s) will not be affected when the explants are cut shorter and the axons should still project anteriorly. FIG. 1E: Quantification of date. Anterior turn indicates normal projection. Knotting/stalling and random A-P turns are abnormal behaviors observed in shorter explants. DiI injections usually label a cohort of axons. In the short explants, some of the axons in the cohort appeared stalling, while others turned posteriorly. These injection stiles were counted for both stalling and the random turn behavior. Therefore, the percent of all projection patterns summed up more than 100%. N=number of explants. All scale bars: 100 μm.
[0060]FIG. 2A, FIG. 2B. The anterior guidance cue(s). FIG. 2A: If the anterior guidance cue(s) is attractive, higher concentrations of the attractant(s) should be found at the anterior end of the explants. The explant tissues close to the anterior end will likely lose the gradient, whereas the posterior end will maintain the gradient. Therefore, axons close to the anterior injection sites will likely be misrouted and the axons close to the posterior end will likely project anteriorly (top panel). If the anterior guidance cue(s) is repulsive, higher concentration of the repellent(s) should be present at the posterior end. The explant tissues close to the posterior end might lose the gradient, whereas the explant tissues close to the anterior end might still maintain the A-P gradient. As a result, axons at the posterior injection sites should show abnormal behavior, whereas those at the anterior injection sites might be normal (bottom panel). FIG. 2B: Quantification of the "open-book" assays with anterior, middle and posterior injections. Note that in some of the injections sites, DiI labeled a cohort of axons. Some of the axons in the cohort appeared stalling, whereas others turned posteriorly at the anterior end of the explants. These injections sites were counted for both stalling and the random turn behavior. Therefore, the percent of all projection patterns summed up more than 100%. n=number of injection sites.
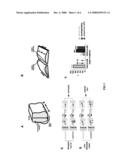
[0061]FIG. 3A, FIG. 3B, FIG. 3C, FIG. 3D. Multiple Wnt proteins stimulate the extension of post-crossing commissural axons. FIG. 3A: Diagram showing the design of "post-crossing" and "pre-crossing" assays. FIG. 3B: Quantification of post-crossing commissural axon extension stimulated by Wnts as described in Zou et al., 2000. FIG. 3C: Schematic diagram of commissural axons projecting towards their brain target, ventral-posterior-lateral region of the thalamus. Dotted square indicates the area of diencephalon dissected for the co-culture experiments. FIG. 3D: Quantification of post-crossing commissural axon growth in response to thalamic target.
[0062]FIG. 4A, FIG. 4B: sFRPs block the anterior turning of post-crossing commissural axons in "open-book" explants. FIG. 4A: Diagram showing the design of experiments. COS cells were transfected with vector only control or sFRP-expressing constructs and resuspended in collagen gel and embedded inside the bottom collagen gel pad. Long "open-book" explants were placed on top of the bottom collagen gel and embedded in the top collagen gel pad. After overnight culturing, tissues were fixed and D11 injected to reveal the projection of commissural axons. FIG. 4B: Quantification of effects of sFRP1, 2, 3 alone or combined. The method of quantification was the same as in FIG. 1 and FIG. 2. n=number of injection sites.
[0063]FIG. 5A, FIG. 5B, FIG. 5C, FIG. 5D, FIG. 5F. Wnt4 gradient rescues A-P guidance defects and can reorient post-crossing commissural axons posteriorly. FIG. 5A, FIG. 5B: Diagrams showing the design of the rescue experiments. COS cell aggregates transfected with either vector only or Wnt4 expression construct were placed to the anterior side of the short "open-book" explants. After overnight culturing, commissural axons were analyzed by DiI labeling of the fixed tissues. FIG. 5C: Quantifications of Wnt4 rescue experiments. The method of quantification was the same as in FIG. 1, FIG. 2, and FIG. 4. FIG. 5D, FIG. 5E: Diagram showing the design of the reorientation experiments. COS cell aggregates transfected with either vector only or Wnt4-expression construct were placed to the posterior side of the short "open-book" explants. After overnight culturing, commissural axons were analyzed by DiI labeling of the fixed explants. FIG. 5F: Quantification of the Wnt4 reorientation experiments. n=number of injection sites. Bars on the far right indicate the percentage of the injection sites whereby all axons turned posteriorly.
[0064]FIG. 6. Frizzled 3 is specifically required for the anterior-posterior guidance of post-crossing commissural axons. Quantification of the post-crossing A-P guidance defects in frizzled 3 knockout mice. Four litters of frizzled 3 knockout mice were analyzed (three litters were analyzed in blinded experiments). A total of 7 mutant embryos were analyzed. The A-P randomization and stalling were observed at 100% penetrance in all injections sites along the entire A-P axis of the spinal cord. n=number of injection sites.
DESCRIPTION OF ILLUSTRATIVE EMBODIMENTS
[0065]The present invention is based on the discovery that Wnts guide axon pathfinding in development and can play a role in correct spinal cord and neuronal regeneration.
[0066]The inventor has shown that a Wnt/Frizzled pathway mediates attractive effects in sensory axon guidance along the anterior-posterior axis. Additionally, the inventor shows here that vertebrate corticospinal cord axons are repelled by Wnts and the repulsion is mediated by the vertebrate homologue of Derailed, Ryk. Ryk is not expressed in the commissural neurons, consistent with the finding that commissural axons are attracted by Wnts. Interestingly, the repulsive effect of Wnt5 on fly axons appears to be independent of Frizzleds. Therefore, Wnts appear to attract axons via a Frizzled-dependent pathway and repel axons via a Ryk dependent pathway. CST axons do express Frizzleds, such as Frizzled 3. Therefore, it appears that Ryk is dominate over Frizzleds and mediates repulsion even in the presence of Frizzleds. Taken together, these studies provide evidence that Wnts, like other guidance cues, are bifunctional, capable of attracting some axons and repelling others, and suggest that Wnt proteins might have a widespread and phylogenetically conserved function in guiding axons during the wiring of the nervous system. These studies demonstrate that one continuous molecular gradient of diffusible guidance cue(s) along the entire anterior-posterior axis of the spinal cord controls the navigation decisions along the A-P axis.
[0067]The present invention seeks to exploit the inventor's discovery by providing for methods and compositions for modulating growth of a nerve cell using a Wnt, Wnt-like substances, and/or chemical compounds to stimulate the pathways of Wnt signaling to modulate nerve growth and guidance. These methods and compositions can be used in a wide variety of therapeutic contexts where nerve growth and regeneration would be beneficial. For example, a Wnt, a Wnt-like substance, and/or a chemical compound affecting a Wnt signaling pathway can be used to stimulate axonal growth of a damaged neuron along the A-P axis of a patient with SCI. Because it has also been observed that the Wnts are expressed in the several regions in the brain and the components of the Wnt signaling pathways are also present in axons of other central nervous system neurons, it is possible that Wnts and agents that stimulate or inhibit Wnt signaling can be used to modulate growth and directional guidance of axons in the central nervous system.
A. Wnt, Wnt-Like Substances, and Compounds Affecting a Wnt Signaling Pathway
[0068]1. Wnt and Wnt-Like Substances
[0069]The present invention pertains to use of Wnt and Wnt-like substances in various contexts. For example, various embodiments of the present invention pertain to methods for modulating growth of a neuron that involve contacting a neuron with a Wnt or a Wnt-like substance. Other embodiments pertain to methods for modulating growth of a neuron in a subject, that involve providing the subject with a pharmaceutical composition that includes a Wnt or a Wnt-like substance. Additional embodiments pertain to pharmaceutical compositions for modulating growth of a neuron in a mammal, that include a Wnt or a Wnt-like substance.
[0070]As discussed above, Wnts are secreted cysteine-rich glycosylated proteins that play a role in the development of a wide range of organisms. Wnts are thought to function in a variety of developmental and physiological processes since many diverse species have multiple conserved Wnt genes (McMahon, 1992; Nusse and Varmus, 1992). The Wnt growth factor family includes at least 19 genes identified in mammals, including Wnt1, Wnt2, Wnt2b, Wnt3, Wnt3a, Wnt4, Wnt5a, Wnt5b, Wnt 6, Wnt7a, Wnt7b, Wnt8a, Wnt8b, Wnt9a, Wnt9b, Wnt10a, Wnt10b, Wnt11, and Wnt16. Similar numbers of Wnt genes are present in other vertebrate species. Of course, further Wnts may be discovered and/or characterized in the future, and those of skill will be able to employ any such Wnts in the context of the invention. Further, those of skill will be able to use the teachings herein to obtain and use Wnts of any species in the context of the invention.
[0071]Throughout this application, the term "Wnt" is intended to refer to any consecutive amino acid sequence that includes the full-length amino acid sequence of a Wnt from any organism, such as a human or a mouse Wnt. Wnt can be a human Wnt protein, or a Wnt protein from any other species, such as mouse or chick. Thus, for example, Wnt can be used to refer to the full-length amino acid sequence encoded by any of the 19 genes identified in human. Alternatively, Wnt can refer to a murine Wnt protein, such as murine Wnt4. Wnt can also refer to an amino acid sequence that is longer than the full-length consecutive amino acid sequence of a Wnt, as long as it includes a full-length Wnt amino acid sequence.
[0072]Throughout this application, the term "Wnt protein" is intended to refer to the full-length amino acid sequence that is encoded by a Wnt gene. Thus, "Wnt" may refer to a Wnt protein or an amino acid sequence that is longer than a Wnt protein if additional non-Wnt amino acids are included in the sequence. Also included in the definition of "Wnt" is a truncated sequence of a Wnt protein, a mutated Wnt protein, or a Wnt amino acid sequence that is less than the full-length amino acid sequence of a Wnt, as long as the amino acid sequence retains an acceptable level of the equivalent biological activity of a full-length Wnt protein.
[0073]The human and murine full-length native amino acid sequences and the native nucleic acids encoding them are described by GenBank accession number in the Table 1. Further, summary of human and murine Wnts is provided in Miller, 2001. Specifically, Table 1 of Miller, 2001, which includes Genbank accession numbers of human and mouse Wnt genes, is herein specifically incorporated by reference.
TABLE-US-00001 TABLE 1 HUMAN MOUSE Nucleic Acid Amino Acid Nucleic Acid Amino Acid Wnt1 NM005430 NP005421 Wnt1 NM133955 NP598716 SEQ ID 1 SEQ ID 2 SEQ ID 39 SEQ ID 40 Wnt2 BC029854 AAH29854 Wnt2 BC026373 AAH26373 SEQ ID 3 SEQ ID 4 SEQ ID 41 SEQ ID 42 Wnt2B NM024494 NP078613 Wnt2B NM009520 NP033546 SEQ ID 5 SEQ ID 6 SEQ ID 43 SEQ ID 44 Wnt3 NM030753 NP110380 Wnt3 NM009521 P17553 SEQ ID 7 SEQ ID 8 SEQ ID 45 SEQ ID 46 Wnt3A NM033131 NP149122 Wnt3A NM009522 NP033548 SEQ ID 9 SEQ ID 10 SEQ ID 47 SEQ ID 48 Wnt4 NM030761 NP110388 Wnt4 NM009523 NP033549 SEQ ID 11 SEQ ID 12 SEQ ID 49 SEQ ID 50 Wnt5A NM003392 NP003383 Wnt5A NM009524 NP033550 SEQ ID 13 SEQ ID 14 SEQ ID 51 SEQ ID 52 Wnt5B BC001749 AAH01749 Wnt5B BC010775 AAH10775 SEQ ID 15 SEQ ID 16 SEQ ID 53 SEQ ID 54 Wnt6 NM006522 NP006513 Wnt6 NM009526 NP033552 SEQ ID 17 SEQ ID 18 SEQ ID 55 SEQ ID 56 Wnt7A BC008811 AAH08811 Wnt7A BC049093 AAH49093 SEQ ID 19 SEQ ID 20 SEQ ID 57 SEQ ID 58 Wnt7B NM058238 NP478679 Wnt7B NM009528 NP033554 SEQ ID 21 SEQ ID 22 SEQ ID 59 SEQ ID 60 Wnt8A NM058244 NP490645 Wnt8A NM009290 NP033316 SEQ ID 23 SEQ ID 24 SEQ ID 61 SEQ ID 62 Wnt8B NM003393 NP003384 Wnt8B NM011720 NP035850 SEQ ID 25 SEQ ID 26 SEQ ID 63 SEQ ID 64 Wnt9A NM003395 NP003386 Wnt9A NM139298 NP647459 SEQ ID 27 SEQ ID 28 SEQ ID 65 SEQ ID 66 Wnt9B NM003396 NP003387 Wnt9B NM011719 NP035849 SEQ ID 29 SEQ ID 30 SEQ ID 67 SEQ ID 68 Wnt10A BC052234 AAH52234 Wnt10A BC014737 AAH14737 SEQ ID 31 SEQ ID 32 SEQ ID 69 SEQ ID 70 Wnt10B NM003394 NP003385 Wnt10B NM011718 NP035848 SEQ ID 33 SEQ ID 34 SEQ ID 71 SEQ ID 72 Wnt11 NM004626 NP004617 Wnt11 NM009519 NP033545 SEQ ID 35 SEQ ID 36 SEQ ID 73 SEQ ID 74 Wnt16 NM057168 NP476509 Wnt16 NM053116 NP444346 SEQ ID 37 SEQ ID 38 SEQ ID 75 SEQ ID 76
[0074]Throughout this application, the term "Wnt-like substance" is intended to refer to a Wnt polypeptide, a Wnt peptide, a Wnt mimetic, or a small molecule that is functionally and/or structurally similar to a Wnt.
[0075]The term "Wnt polypeptide" includes any amino acid sequence that includes fewer consecutive amino acids of a Wnt than the full-length amino acid sequence of a Wnt. "Wnt polypeptide" includes not only consecutive amino acid sequences from a human Wnt, but from any other species, such as mouse. Thus, for example, a Wnt polypeptide can include, but is not limited to, about 11, about 12, about 13, about 14, about 15, about 16, about 17, about 18, about 19, about 20, about 21, about 22, about 23, about 24, about 25, about 26, about 27, about 28, about 29, about 30, about 31, about 32, about 33, about 34, about 35, about 36, about 37, about 38, about 39, about 40, about 41, about 42, about 43, about 44, about 45, about 46, about 47, about 48, about 49, about 50, about 51, about 52, about 53, about 54, about 55, about 56, about 57, about 58, about 59, about 60, about 61, about 62, about 63, about 64, about 65, about 66, about 67, about 68, about 69, about 70, about 71, about 72, about 73, about 74, about 75, about 76, about 77, about 78, about 79, about 80, about 81, about 82, about 83, about 84, about 85, about 86, about 87, about 88, about 89, about 90, about 91, about 92, about 93, about 94, about 95, about 96, about 97, about 98, about 99, about 100, about 110, about 120, about 130, about 140, about 150, about 160, about 170, about 180, about 190, about 200, about 210, about 220, about 230, about 240, about 250, about 275, about 300, about 325, about 350, about 375, about 400, about 425, about 450, about 475, about 500, about 525, about 550, about 575, about 600, about 625, about 650, about 675, about 700, about 725, about 750, about 775, about 800, about 825, about 850, about 875, about 900, about 925, about 950, about 975, about 1000, about 1100, about 1200, about 1300, about 1400, about 1500, about 1750, about 2000, about 2250, about 2500 or greater amino molecule residues of a Wnt, and any range derivable therein, as long as the amino acid sequence includes less than the full-length consecutive amino acid sequence of a Wnt. Included within the definition of "Wnt polypeptide" are potential amino acid sequences that include additional amino acids, other than Wnt amino acid sequences.
[0076]The term "Wnt peptide" includes any amino acid sequence that includes ten or fewer consecutive amino acid sequence of a Wnt amino acid sequence. "Wnt peptide" includes not only consecutive amino acid sequences from a human Wnt, but from any other species, such as mouse. Thus, for example, a Wnt peptide may include 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 consecutive amino acids of a Wnt. Additional amino acids can also be included, which may be other than Wnt amino acid sequences.
[0077]Included within the definition of "Wnt-like substance" is a "mimetic of Wnt." Throughout this application, "mimetic of Wnt" is intended to refer to any molecule other than the full-length sequence of a Wnt that is able to maintain an acceptable level of equivalent biological activity as a Wnt.
[0078]It is well understood by the skilled artisan that, inherent in the definition of a "mimetic of Wnt," is the concept that there is a limit to the number of changes that may be made within a defined portion of the molecule and still result in a molecule with an acceptable level of equivalent biological activity, e.g., ability of Wnt4 to modulate neuronal growth and regeneration. "Mimetic of Wnt" is thus defined herein as any Wnt polypeptide in which some, or most, of the amino acids may be substituted so long as the polypeptide retains substantially similar activity in the context of the uses set forth herein. Of course, a plurality of distinct proteins/polypeptides/peptides with different substitutions may easily be made and used in accordance with the invention. Additionally, in the context of the invention, a mimetic of Wnt can be a Wnt homologue polypeptide from any species or organism, including, but not limited to, a human polypeptide. One of ordinary skill in the art will understand that many mimetics of Wnt would likely exist and can be identified using commonly available techniques.
[0079]The present invention may utilize Wnts, Wnt polypeptides, Wnt peptides, mimetics of Wnt, or mutants of Wnt, that are purified from a natural source or from recombinantly-produced material. Those of ordinary skill in the art would know how to produce these amino acid sequences from recombinantly-produced material. This material may use the 20 common amino acids in naturally synthesized proteins, or one or more modified or unusual amino acids. Generally, "purified" will refer to an Wnt composition that has been subjected to fractionation to remove various other proteins, polypeptides, or peptides, and which composition substantially retains its activity. Purification may be substantial, in which the Wnt or Wnt-like substance is the predominant species, or to homogeneity, which purification level would permit accurate degradative sequencing.
[0080]Amino acid sequence mutants of a Wnt also are encompassed by the present invention, and are included within the definition of "Wnt-like substance." Amino acid sequence mutants of a Wnt of any species, such as human and mouse Wnt, is contemplated by the present invention. Amino acid sequence mutants of a Wnt can be substitutional mutants or insertional mutants. Insertional mutants typically involve the addition of material at a non-terminal point in the peptide. This may include the insertion of a few residues; an immunoreactive epitope; or simply a single residue. The added material may be modified, such as by methylation, acetylation, and the like. Alternatively, additional residues may be added to the N-terminal or C-terminal ends of the peptide.
[0081]Amino acid substitutions are generally based on the relative similarity of the amino acid side-chain substituents, or example, their hydrophobicity, hydrophilicity, charge, size, and the like. An analysis of the size, shape and type of the amino acid side-chain substituents reveals that arginine, lysine and histidine are all positively charged residues; that alanine, glycine and serine are all a similar size; and that phenylalanine, tryptophan and tyrosine all have a generally similar shape. Therefore, based upon these considerations, arginine, lysine and histidine; alanine, glycine and serine; and phenylalanine, tryptophan and tyrosine; are defined herein as biologically functional equivalents.
[0082]Amino acid substitutions are generally based on the relative similarity of the amino acid side-chain substituents, or example, their hydrophobicity, hydrophilicity, charge, size, and the like. An analysis of the size, shape and type of the amino acid side-chain substituents reveals that arginine, lysine and histidine are all positively charged residues; that alanine, glycine and serine are all a similar size; and that phenylalanine, tryptophan and tyrosine all have a generally similar shape. Therefore, based upon these considerations, arginine, lysine and histidine; alanine, glycine and serine; and phenylalanine, tryptophan and tyrosine; are defined herein as biologically functional equivalents.
[0083]In making changes, the hydropathic index of amino acids may be considered. Each amino acid has been assigned a hydropathic index on the basis of their hydrophobicity and charge characteristics, these are: isoleucine (+4.5); valine (+4.2); leucine (+3.8); phenylalanine (+2.8); cysteine/cystine (+2.5); methionine (+1.9); alanine (+1.8); glycine (-0.4); threonine (-0.7); serine (-0.8); tryptophan (-0.9); tyrosine (-1.3); proline (-1.6); histidine (-3.2); glutamate (-3.5); glutamine (-3.5); aspartate (-3.5); asparagine (-3.5); lysine (-3.9); and arginine (-4.5).
[0084]The importance of the hydropathic amino acid index in conferring interactive biological function on a protein is generally understood in the art (Kyte and Doolittle, 1982, incorporated by reference herein). It is known that certain amino acids may be substituted for other amino acids having a similar hydropathic index or score and still retain a similar biological activity. In making changes based upon the hydropathic index, the substitution of amino acids whose hydropathic indices are within +2 is preferred, those which are within +1 are particularly preferred, and those within +0.5 are even more particularly preferred.
[0085]It is understood that an amino acid can be substituted for another having a similar hydrophilicity value and still obtain a biologically equivalent protein. As detailed in U.S. Pat. No. 4,554,101, the following hydrophilicity values have been assigned to amino acid residues: arginine (+3.0); lysine (+3.0); aspartate (+3.0+1); glutamate (+3.0+1); serine (+0.3); asparagine (+0.2); glutamine (+0.2); glycine (0); threonine (-0.4); proline (-0.5+1); alanine (-0.5); histidine (-0.5); cysteine (-1.0); methionine (-1.3); valine (-1.5); leucine (-1.8); isoleucine (-1.8); tyrosine (-2.3); phenylalanine (-2.5); tryptophan (-3.4).
[0086]In making changes based upon similar hydrophilicity values, the substitution of amino acids whose hydrophilicity values are within +2 is preferred, those which are within +1 are particularly preferred, and those within +0.5 are even more particularly preferred.
[0087]Certain embodiments of the present invention utilize Wnt-like substances that are fusion proteins that are preferentially translocated through biological membranes. In particular, a Wnt or a Wnt-like substance such as a Wnt polypeptide may be fused to a particular protein, polypeptide, or peptide sequence that promotes facilitated intracellular delivery of the fusion protein into the targeted cell. Although any fusion protein with the property of facilitated intracellular delivery is contemplated by the present invention, specific examples include fusion proteins utilizing the HIV TAT sequence (Nagahara et al., 1998), the third helix of the Antennapedia homeodomain (Antp) (Derossi et al., 1994), and the HSV-1 structural protein VP22 (Elliott and O'Hare, 1997).
[0088]Small molecules are also included within the definition of "Wnt-like substance" in the context of the present invention. Throughout this application, the term "small molecule" is intended to refer to any small molecule not included within the definition of Wnt polypeptide, Wnt peptide, mimetic of Wnt, or mutant of Wnt, wherein the molecule is relatively small in size and wherein the molecule has an acceptable level of biological activity of a Wnt. For example, the small molecule may be a synthetic substance which is not an amino acid sequence, which is functionally able to promote axonal growth and regeneration in a manner analogous to a Wnt.
[0089]2. Polynucleotides Encoding a Wnt or a Wnt-Like Substance
[0090]Various aspects of the present invention require polynucleotides encoding an Wnt or a Wnt-like substance. For example, various embodiments include methods for modulating neuronal growth that involve contacting the neuron with an expression cassette that includes a promoter that is a cell, operably linked to a polynucleotide encoding either an Wnt or a Wnt-like substance. In other embodiments, the invention pertains to methods for modulating growth of a neuron in a subject that include administering to the subject a composition that includes an expression cassette operably inked to a polynucleotide encoding either a Wnt or a Wnt-like substance. In still other embodiments, the invention includes pharmaceutical compositions for modulating growth of a neuron in a mammal, that include a Wnt or a Wnt-like substance.
[0091]The polynucleotide encoding the full length amino acid sequences of the known human and murine Wnts are contained in Table 1. The polynucleotides according to the present invention may encode an entire Wnt sequence (e.g., the amino acid sequence of SEQ ID NO:2), or a Wnt-like substance such as a Wnt polypeptide or a Wnt peptide. The polynucleotides may be derived from genomic DNA, i.e., cloned directly from the genome of a particular organism.
[0092]In other embodiments, however, the polynucleotides may be complementary DNA (cDNA). cDNA is DNA prepared using messenger RNA (mRNA) as a template. Thus, a cDNA does not contain any interrupted coding sequences and usually contains almost exclusively the coding region(s) for the corresponding protein. In other embodiments, the polynucleotide may be produced synthetically.
[0093]It may be advantageous to combine portions of the genomic DNA with cDNA or synthetic sequences to generate specific constructs. For example, where an intron is desired in the ultimate construct, a genomic clone will need to be used. Introns may be derived from other genes in addition to a Wnt gene. The cDNA or a synthesized polynucleotide may provide more convenient restriction sites for the remaining portion of the construct and, therefore, would be used for the rest of the sequence.
[0094]The present invention is not limited to the sequences disclosed by GenBank and SEQ ID NO in Table 1, but includes polynucleotides encoding any Wnt or Wnt-like substance (discussed above). These polynucleotides encoding a Wnt or a Wnt-like substance may be naturally-occurring homologous polynucleotide sequences from other organisms. For example, polynucleotides encoding a Wnt or a Wnt-like substance include those polynucleotides encoding the human amino acid functional equivalent sequences previously described. These sequences are provided by way of example, and are not meant to be a summary of all available polynucleotide sequences encoding a Wnt or a Wnt-like substance. A person of ordinary skill in the art would understand that commonly available experimental techniques can be used to identify or synthesize polynucleotides encoding other Wnts. The present invention also encompasses chemically synthesized mutants of these sequences.
[0095]Another kind of sequence variant results from codon variation. Because there are several codons for most of the 20 normal amino acids, many different DNAs can encode a Wnt or a Wnt-like substance. Reference to the following table will allow such variants to be identified.
TABLE-US-00002 TABLE 2 Amino Acids Codons Alanine Ala A GCA GCC GCG GCU Cysteine Cys C UGC UGU Aspartic acid Asp D GAC GAU Glutamic acid Glu E GAA GAG Phenylalanine Phe F UUC UUU Glycine Gly G GGA GGC GGG GGU Histidine His H CAC CAU Isoleucine Ile I AUA AUC AUU Lysine Lys K AAA AAG Leucine Leu L UUA UUG CUA CUC CUG CUU Methionine Met M AUG Asparagine Asn N AAC AAU Proline Pro P CCA CCC CCG CCU Glutamine Gln Q CAA CAG Arginine Arg R AGA AGG CGA CGC CGG CGU Serine Ser S AGC AGU UCA UCC UCG UCU Threonine Thr T ACA ACC ACG ACU Valine Val V GUA GUC GUG GUU Tryptophan Trp W UGG Tyrosine Tyr Y UAC UAU
[0096]Allowing for the degeneracy of the genetic code, sequences that have between about 50% and about 75%, or between about 76% and about 99%, of nucleotides that are identical to the nucleotides disclosed herein will be preferred. Sequences that are within the scope of "a polynucleotide encoding a Wnt or a Wnt-like substance" are those that are capable of base-pairing with a polynucleotide segment set forth above under intracellular conditions.
[0097]As stated above, the encoding sequences may be full length genomic or cDNA copies, or large fragments thereof. The present invention also may employ shorter oligonucleotides. Sequences of 17 bases long should occur only once in the human genome and, therefore, suffice to specify a unique target sequence. Although shorter oligomers are easier to make and increase in vivo accessibility, numerous other factors are involved in determining the specificity of base-pairing. Both binding affinity and sequence specificity of an oligonucleotide to its complementary target increases with increasing length. It is contemplated that oligonucleotides of 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19 or 20 base pairs will be used, for example, in the preparation of mutants of Wnt and in PCR reactions.
[0098]Any sequence of 17 bases long should occur only once in the human genome and, therefore, suffice to specify a unique target sequence. Although shorter oligomers are easier to make and increase in vivo accessibility, numerous other factors are involved in determining the specificity of hybridization. Both binding affinity and sequence specificity of an oligonucleotide to its complementary target increases with increasing length.
[0099]In certain embodiments, one may wish to employ constructs which include other elements, for example, those which include C-5 propyne pyrimidines. Oligonucleotides which contain C-5 propyne analogues of uridine and cytidine have been shown to bind RNA with high affinity (Wagner et al., 1993).
[0100]3. Compounds that can Affect the Wnt Signaling Pathways
[0101]a. Chemical Compounds that can Affect the Wnt Signaling Pathway
[0102]As an alternative approach to using the a Wnt or a Wnt-like substance to directly modulate axon growth and guidance to promote axonal regeneration to cure spinal cord injury and other central nervous system damage, chemical compounds which affect the Wnt signaling pathways and affect axonal regeneration can also be applied to promote and guidance axon regeneration. Such chemical compound can be discovered by "chemical genetics", screening libraries of chemical compounds or testing known compounds that have an effect on Wnt signaling. For example, lithium is known to stimulate Wnt signaling and can promote axon extension (Hall et al., 2000; Klein and Melton, 1996; Lucas and Salinas, 1997). Therefore, chemical substances, such as lithium, can be used to regulate the Wnt pathway and help regenerate spinal cord axons and other central nervous system axons.
[0103]b. sFRPS can Affect the Wnt Signaling Pathways
[0104]Secreted Frizzled-related proteins (sFRPs) are soluble proteins that can bind to Wnt proteins with high affinities and can block the interaction of Wnts with their receptors, the Frizzleds (Wodarz and Nusse, 1998). Any sFRP, whether from human or any other species such as mouse, is contemplated by the present invention. In addition, the definition of sFRP-like substance is defined in a similar manner as Wnt-like substance, and includes mimetics of sFRP and mutant sFRPs.
[0105]The definition of sFRP, sFRP-like substance, sFRP protein, and sFRP polypeptide are defined in a manner analogous to the definitions provided above in reference to Wnt and Wnt-like substance, discussed supra.
[0106]The full-length amino acid sequence of human sFRP1 (Genbank accession number NP--003003) is provided herein as SEQ ID NO:77. The full-length amino acid sequence of human sFRP2 (Genbank accession number XP--050625) is provided herein as SEQ ID NO:78. The full-length amino acid sequence of human sFRP3 (Genbank accession number NP--001454) is provided herein as SEQ ID NO:79. The full-length amino acid sequence of murine sFRP1 (Genbank accession number NP--038862) is provided herein as SEQ ID NO:80. The full-length amino acid sequence of murine sFRP2 (Genbank accession number NP--033170) is provided herein as SEQ ID NO:81. The full-length amino acid sequence of murine sFRP3 (Genbank accession number AAC53147) is provided herein as SEQ ID NO:82.
[0107]c. Ryk can Affect the Wnt Signaling Pathways
[0108]Ryk is a protein that can bind to Wnt proteins with high affinities and can block the activity of at least some of Wnts. Ryk is a vertebrate homolog of the Drosphila Derailed protein, a receptor tyrosine-like protein. Any Ryk, whether from human or any other species such as mouse, is contemplated by the present invention. In addition, the definition of Ryk-like substance is defined in a similar manner as Wnt-like substance, and includes mimetics of Ryk and mutant Ryks.
[0109]The definition of Ryk, Ryk-like substance, Ryk protein, and Ryk polypeptide are defined in a manner analogous to the definitions provided above in reference to Wnt and Wnt-like substance, discussed supra.
[0110]The full-length amino acid sequence of human Ryk (Genbank accession number NM--002958) is provided herein as SEQ ID NO:83. The full-length amino acid sequence of murine Ryk (Genbank accession number BC--006963) is provided herein as SEQ ID NO:84. The full-length amino acid sequence of Derailed (Genbank accession number L47260) is provided herein as SEQ ID NO:85.
B. Inhibitors of Axonal Growth
[0111]The adult central nervous system is a largely inhibitory environment for axonal growth and regeneration. Therefore, in the context of obtaining regeneration of the CNS, it is likely that the blocking of such inhibitors will be needed.
[0112]Additionally, multiple inhibitors present in the central nervous system myelin, such as Nogo, MAG and OMgp, prevent axonal growth after injury. Other inhibitors present in glial scar, such as CSPG, also inhibit axonal outgrowth. It is not fully understood whether CSPG are the actual active components for the inhibitors of axonal regeneration or other molecules associate with CSPG are the active components.
[0113]In order to achieve effective axonal regeneration following CNS injury, it is necessary to overcome inhibition of both type of inhibitors. Those of skill in the art will understand that there are many manners in which such inhibitors can be blocked, and will, by following the teachings contained herein, be able to develop means to block these inhibitors in the context of the invention.
C. Protein Attractants and Repellants in Axonal Guidance
[0114]There are many protein attractants and repellants that play a role in axonal guidance. Further, many such axon guidance molecules are bi-functional: attractive to one type of axons and repulsive to another, depending on the receptor composition in the responding growth cones.
[0115]A number of molecules direct axonal growth during development. These compounds are play important roles in embryonic development, and may function in the same or a similar way in the adult CNS.
[0116]Attractants and repellants can be divided into two general categories, diffusable and non-diffusable. Diffusible attractants include, but are not limited to, Netrins, Shh, Wnts, and HGF. Diffusible repellents include, but are not limited to, Secreted Semaphorins, Netrins, Slits, Wnts, and BMPs. Non-diffusible attractants include, but are not limited to: cell adhesion molecules such as members of the Ig superfamily, Cadherins, and Integrins; Ephrins; and ECM molecules. Non-diffusable repellents include, but are not limited to, Ephrins, members of the Ig superfamily, and membrane-bound Semaphorins.
[0117]Those of skill in the art will be able to use these, and any other attractants or repellants in the context of the invention. For example, those of skill in the are will be able to use these attractants or repellants to create suitable gradients for guiding neuronal growth.
[0118]In the context of the invention, native attractants or repellants may be employed. Further, proteins, polypeptides, peptides, mutants, and/or mimetics of these attractants or repellants may be employed, with the definitions of these provided above in reference to Wnt and Wnt-like substance, discussed supra.
D. Targeted Diseases and Conditions
[0119]The present invention contemplates methods of treating a subject that includes administering to the subject a composition that includes a Wnt, a Wnt-like substance, and/or a chemical compound affecting a Wnt signaling pathway in a pharmaceutical preparation suitable for delivery to the subject. Other axonal guidance molecules or substances that block neuronal inhibitors can be administered in combination. The subject can be a patient with a disease wherein neuronal dysfunction plays a prominent role in the pathophysiology. For example, the patient may have a disorder of the spinal cord. Any disorder of the spinal cord is contemplated by the present invention. In certain embodiments, the disorder of the spinal cord is traumatic spinal cord injury (discussed above). The traumatic spinal cord injury may or may not have resulted in paralysis of the subject. The neuronal dysfunction can be by any mechanism. For example, cell death can be the result of acute traumatic injury or degeneration.
[0120]In certain embodiments, the Wnt, Wnt-like substance, and/or a chemical compound affecting the Wnt signaling pathway is administered to a subject for the purpose of stimulating and promoting directed axonal growth and regeneration along the anterior-posterior axis of the spinal cord.
[0121]Any disease or condition wherein there is neuronal dysfunction is contemplated by the present invention. In addition to SCI, other examples include Parkinson's disease, where dopaminergic neurons undergo degeneration and ALS where neurons in the motor systems undergo degeneration. In these cases, stem cells are being developed so that they can be transplanted to the midbrain and the spinal cord, respectively, so that they can populate and make proper connection with their targets. The establishment of new connections require the directly growth of axons from these neural stem cells. Wnt and Wnt-like substances and other chemical compounds affecting a Wnt signaling pathway can be used in growth and guidance of regenerating axons from these stem cells.
E. Nucleic Acids
[0122]1. Overview
[0123]Certain embodiments of the invention pertain to methods utilizing compositions that include an nucleic acids. In particular, the methods for modulating growth of a neuron may involve contacting the neuron with a Wnt, a Wnt-like substance, and/or a chemical compound affecting a Wnt signaling pathway that further includes an expression cassette. The methods of treating a subject may involve administering to the subject a composition of a Wnt, a Wnt-like substance, and/or a chemical compound affecting a Wnt signaling pathway that includes an expression cassette. One of skill in the art would understand the techniques relating to use of expression cassettes to deliver polynucleotide sequences to cells or subjects. Particular aspects of these techniques of these techniques are summarized in this specification. This brief summary is in no way designed to be an exhaustive overview of all available experimental techniques related to expression cassettes since one of skill in the art would already be familiar with these techniques.
[0124]Throughout this application, the term "expression cassette" is meant to include any type of genetic construct containing a nucleic acid coding for a gene product in which part or all of the nucleic acid encoding sequence is capable of being transcribed. The transcript may be translated into a protein or polypeptide, but it need not be. Thus, in certain embodiments, expression includes both transcription of a gene and translation of a mRNA into a polypeptide.
[0125]In order for the expression cassette to effect expression of a polypeptide, the polynucleotide encoding the polynucleotide will be under the transcriptional control of a promoter. A "promoter" is a control sequence that is a region of a nucleic acid sequence at which initiation and rate of transcription are controlled. It may contain genetic elements at which regulatory proteins and molecules may bind such as RNA polymerase and other transcription factors. The phrase "operatively linked" means that a promoter is in a correct functional location and/or orientation in relation to a nucleic acid sequence to control transcriptional initiation and/or expression of that sequence. A promoter may or may not be used in conjunction with an "enhancer," which refers to a cis-acting regulatory sequence involved in the transcriptional activation of a nucleic acid sequence. One of skill in the art would understand how to use a promoter or enhancer to promote expression of a Wnt, a Wnt-like substance, and/or a chemical compound affecting a Wnt signaling pathway.
[0126]In certain embodiments of the invention, the delivery of an expression cassette in a cell may be identified in vitro or in vivo by including a marker in the expression vector. The marker would result in an identifiable change to the transfected cell permitting easy identification of expression. The selectable marker employed is not believed to be important, so long as it is capable of being expressed along with the polynucleotide of the expression cassette. Examples of selectable markers are well known to one of skill in the art.
[0127]A specific initiation signal also may be required for efficient translation of coding sequences. These signals include the ATG initiation codon or adjacent sequences. Exogenous translational control signals, including the ATG initiation codon, may need to be provided. One of ordinary skill in the art would readily be capable of determining this and providing the necessary signals.
[0128]In certain embodiments of the invention, the use of internal ribosome entry sites (IRES) elements are used to create multigene, or polycistronic, messages. IRES elements are able to bypass the ribosome scanning model of 5' methylated Cap dependent translation and begin translation at internal sites (Pelletier and Sonenberg, 1988). One of skill in the art would be familiar with use of IRES in expression cassettes.
[0129]Expression cassettes can include a multiple cloning site (MCS), which is a nucleic acid region that contains multiple restriction enzyme sites, any of which can be used in conjunction with standard recombinant technology to digest the vector. See Carbonelli et al. (1999); Levenson et al. (1998); Cocea (1997). "Restriction enzyme digestion" refers to catalytic cleavage of a nucleic acid molecule with an enzyme that functions only at specific locations in a nucleic acid molecule. Techniques involving restriction enzymes and ligation reactions are well known to those of skill in the art of recombinant technology.
[0130]In expression, one will typically include a polyadenylation signal to effect proper polyadenylation of the transcript. The nature of the polyadenylation signal is not believed to be crucial to the successful practice of the invention, and/or any such sequence may be employed. One of skill in the art would understand how to use these signals to effect proper polyadenylation of the transcript.
[0131]In certain embodiments of the present invention, the expression cassette comprises a virus or engineered construct derived from a viral genome. The ability of certain viruses to enter cells via receptor-mediated endocytosis and, in some cases, integrate into the host cell chromosomes, have made them attractive candidates for gene transfer in to mammalian cells. However, because it has been demonstrated that direct uptake of naked DNA, as well as receptor-mediated uptake of DNA complexes, expression vectors need not be viral but, instead, may be any plasmid, cosmid or phage construct that is capable of supporting expression of encoded genes in mammalian cells, such as pUC or Bluescript® plasmid series. One of ordinary skill in the art would be familiar with use of viruses as tools to promote expression of the polypeptide.
[0132]In certain embodiments of the invention, a treated cell may be identified in vitro or in vivo by including a marker in the expression vector. Such markers would confer an identifiable change to the cell permitting easy identification of cells containing the expression vector. Generally, a selectable marker is one that confers a property that allows for selection. A positive selectable marker is one in which the presence of the marker allows for its selection, while a negative selectable marker is one in which its presence prevents its selection. An example of a positive selectable marker is a drug resistance marker.
[0133]Usually the inclusion of a drug selection marker aids in the cloning and identification of transformants, for example, genes that confer resistance to neomycin, puromycin, hygromycin, DHFR, GPT, zeocin and histidinol are useful selectable markers. In addition to markers conferring a phenotype that allows for the discrimination of transformants based on the implementation of conditions, other types of markers including screenable markers such as GFP, whose basis is calorimetric analysis, are also contemplated. Alternatively, screenable enzymes such as herpes simplex virus thymidine kinase (tk) or chloramphenicol acetyltransferase (CAT) may be utilized. One of skill in the art would also know how to employ immunologic markers, possibly in conjunction with FACS analysis. The marker used is not believed to be important, so long as it is capable of being expressed simultaneously with the nucleic acid encoding a gene product. Further examples of selectable and screenable markers are well known to one of skill in the art.
F. Gene Transfer
[0134]1. Viral Vectors
[0135]In certain embodiments, the methods and compositions of the invention utilize expression cassette which includes a polynucleotide encoding a Wnt, a Wnt-like substance, a chemical compound affecting a Wnt signaling pathway, another axonal guidance molecule, and/or substance that blocks a neuronal inhibitor can be administered in combination, carried in a vector. One of ordinary skill in the art would understand use of vectors since these experimental methods are well-known in the art. In particular, techniques using "viral vectors" are well-known in the art. A viral vector is meant to include those constructs containing viral sequences sufficient to (a) support packaging of the expression cassette and (b) to ultimately express a recombinant gene construct that has been cloned therein.
[0136]One method for delivery of the recombinant DNA involves the use of an adenovirus expression vector. Although adenovirus vectors are known to have a low capacity for integration into genomic DNA, this feature is counterbalanced by the high efficiency of gene transfer afforded by these vectors.
[0137]Adenoviruses are currently the most commonly used vector for gene transfer in clinical settings. Among the advantages of these viruses is that they are efficient at gene delivery to both nondividing an dividing cells and can be produced in large quantities. The vector comprises a genetically engineered form of adenovirus. Knowledge of the genetic organization or adenovirus, a 36 kb, linear, double-stranded DNA virus, allows substitution of large pieces of adenoviral DNA with foreign sequences up to 7 kb (Grunhaus et al., 1992). In contrast to retrovirus, the adenoviral infection of host cells does not result in chromosomal integration because adenoviral DNA can replicate in an episomal manner without potential genotoxicity. Also, adenoviruses are structurally stable, and no genome rearrangement has been detected after extensive amplification.
[0138]Adenovirus is particularly suitable for use as a gene transfer vector because of its mid-sized genome, ease of manipulation, high titer, wide target-cell range and high infectivity. A person of ordinary skill in the art would be familiar with experimental methods using adenoviral vectors.
[0139]The adenovirus vector may be replication defective, or at least conditionally defective, and the nature of the adenovirus vector is not believed to be crucial to the successful practice of the invention. The adenovirus may be of any of the 42 different known serotypes or subgroups A-F. Adenovirus type 5 of subgroup C is the preferred starting material in order to obtain the conditional replication-defective adenovirus vector for use in the present invention. This is because Adenovirus type 5 is a human adenovirus about which a great deal of biochemical and genetic information is known, and it has historically been used for most constructions employing adenovirus as a vector.
[0140]Adenovirus growth and manipulation is known to those of skill in the art, and exhibits broad host range in vitro and in vivo. This group of viruses can be obtained in high titers, e.g., 109-1011 plaque-forming units per ml, and they are highly infective. The life cycle of adenovirus does not require integration into the host cell genome. The foreign genes delivered by adenovirus vectors are episomal and, therefore, have low genotoxicity to host cells. No side effects have been reported in studies of vaccination with wild-type adenovirus (Couch et al., 1963; Top et al., 1971), demonstrating their safety and therapeutic potential as in vivo gene transfer vectors.
[0141]The retroviruses are a group of single-stranded RNA viruses characterized by an ability to convert their RNA to double-stranded DNA in infected cells by a process of reverse-transcription (Coffin, 1990). The resulting DNA then stably integrates into cellular chromosomes as a provirus and directs synthesis of viral proteins. The integration results in the retention of the viral gene sequences in the recipient cell and its descendants. The retroviral genome contains three genes, gag, pol, and env that code for capsid proteins, polymerase enzyme, and envelope components, respectively. A sequence found upstream from the gag gene contains a signal for packaging of the genome into virions. Two long terminal repeat (LTR) sequences are present at the 5' and 3' ends of the viral genome. These contain strong promoter and enhancer sequences and are also required for integration in the host cell genome (Coffin, 1990).
[0142]In order to construct a retroviral vector, a nucleic acid encoding a gene of interest is inserted into the viral genome in the place of certain viral sequences to produce a virus that is replication-defective. A person of ordinary skill in the art would be familiar with well-known techniques that are available to construct a retroviral vector.
[0143]Adeno-associated virus (AAV) is an attractive vector system for use in the present invention as it has a high frequency of integration and it can infect nondividing cells, thus making it useful for delivery of genes into mammalian cells in tissue culture (Muzyczka, 1992). AAV has a broad host range for infectivity (Tratschin, et al., 1984; Laughlin, et al., 1986; Lebkowski, et al., 1988; McLaughlin, et al., 1988), which means it is applicable for use with the present invention. Details concerning the generation and use of rAAV vectors are described in U.S. Pat. No. 5,139,941 and U.S. Pat. No. 4,797,368, each incorporated herein by reference.
[0144]AAV is a dependent parvovirus in that it requires coinfection with another virus (either adenovirus or a member of the herpes virus family) to undergo a productive infection in cultured cells (Muzyczka, 1992). In the absence of coinfection with helper virus, the wild-type AAV genome integrates through its ends into human chromosome 19 where it resides in a latent state as a provirus (Kotin et al., 1990; Samulski et al., 1991). rAAV, however, is not restricted to chromosome 19 for integration unless the AAV Rep protein is also expressed (Shelling and Smith, 1994). When a cell carrying an AAV provirus is superinfected with a helper virus, the AAV genome is "rescued" from the chromosome or from a recombinant plasmid, and a normal productive infection is established (Samulski et al., 1989; McLaughlin et al., 1988; Kotin et al., 1990; Muzyczka, 1992).
[0145]Typically, recombinant AAV (rAAV) virus is made by cotransfecting a plasmid containing the gene of interest flanked by the two AAV terminal repeats (McLaughlin et al., 1988; Samulski et al., 1989; each incorporated herein by reference) and an expression plasmid containing the wild-type AAV coding sequences without the terminal repeats, for example pIM45 (McCarty et al., 1991; incorporated herein by reference). A person of ordinary skill in the art would be familiar with techniques available to generate vectors using AAV virus.
[0146]Herpes simplex virus (HSV) has generated considerable interest in treating nervous system disorders due to its tropism for neuronal cells, but this vector also can be exploited for other tissues given its wide host range. Another factor that makes HSV an attractive vector is the size and organization of the genome. Because HSV is large, incorporation of multiple genes or expression cassettes is less problematic than in other smaller viral systems. In addition, the availability of different viral control sequences with varying performance (temporal, strength, etc.) makes it possible to control expression to a greater extent than in other systems. It also is an advantage that the virus has relatively few spliced messages, further easing genetic manipulations.
[0147]HSV also is relatively easy to manipulate and can be grown to high titers. Thus, delivery is less of a problem, both in terms of volumes needed to attain sufficient MOI and in a lessened need for repeat dosings. For a review of HSV as a gene therapy vector, see Glorioso et al. (1995). A person of ordinary skill in the art would be familiar with well-known techniques for use of HSV as vectors.
[0148]Vaccinia virus vectors have been used extensively because of the ease of their construction, relatively high levels of expression obtained, wide host range and large capacity for carrying DNA. Vaccinia contains a linear, double-stranded DNA genome of about 186 kb that exhibits a marked "A-T" preference. Inverted terminal repeats of about 10.5 kb flank the genome. The majority of essential genes appear to map within the central region, which is most highly conserved among poxviruses. Estimated open reading frames in vaccinia virus number from 150 to 200. Although both strands are coding, extensive overlap of reading frames is not common.
[0149]Other viral vectors may be employed as constructs in the present invention. For example, vectors derived from viruses such as poxvirus may be employed. A molecularly cloned strain of Venezuelan equine encephalitis (VEE) virus has been genetically refined as a replication competent vaccine vector for the expression of heterologous viral proteins (Davis et al., 1996). Studies have demonstrated that VEE infection stimulates potent CTL responses and has been suggested that VEE may be an extremely useful vector for immunizations (Caley et al., 1997). It is contemplated in the present invention, that VEE virus may be useful in targeting dendritic cells.
[0150]A polynucleotide may be housed within a viral vector that has been engineered to express a specific binding ligand. The virus particle will thus bind specifically to the cognate receptors of the target cell and deliver the contents to the cell. A novel approach designed to allow specific targeting of retrovirus vectors was developed based on the chemical modification of a retrovirus by the chemical addition of lactose residues to the viral envelope. This modification can permit the specific infection of hepatocytes via sialoglycoprotein receptors.
[0151]Another approach to targeting of recombinant retroviruses was designed in which biotinylated antibodies against a retroviral envelope protein and against a specific cell receptor were used. The antibodies were coupled via the biotin components by using streptavidin (Roux et al., 1989). Using antibodies against major histocompatibility complex class I and class II antigens, they demonstrated the infection of a variety of human cells that bore those surface antigens with an ecotropic virus in vitro (Roux et al., 1989).
[0152]2. Nonviral Vectors
[0153]Several non-viral methods for the transfer of expression vectors into cells also are contemplated by the present invention. These include calcium phosphate precipitation (Graham and Van Der Eb, 1973; Chen and Okayama, 1987; Rippe et al., 1990) DEAE-dextran (Gopal, 1985), electroporation (Tur-Kaspa et al., 1986; Potter et al., 1984), direct microinjection (Harland and Weintraub, 1985), DNA-loaded liposomes (Nicolau and Sene, 1982; Fraley et al., 1979) and liofectamine-DNA complex, cell sonication (Fechheimer et al., 1987), gene bombardment using high velocity microprojectiles (Yang et al., 1990), polycations (Bousssif et al., 1995) and receptor-mediated transfection (Wu and Wu, 1987; Wu and Wu, 1988). Some of these techniques may be successfully adapted for in vivo or ex vivo use. A person of ordinary skill in the art would be familiar with the techniques pertaining to use of nonviral vectors, and would understand that other types of nonviral vectors than those disclosed herein are contemplated by the present invention.
[0154]In a further embodiment of the invention, the expression cassette may be entrapped in a liposome or lipid formulation. Liposomes are vesicular structures characterized by a phospholipid bilayer membrane and an inner aqueous medium. Multilamellar liposomes have multiple lipid layers separated by aqueous medium. They form spontaneously when phospholipids are suspended in an excess of aqueous solution. The lipid components undergo self-rearrangement before the formation of closed structures and entrap water and dissolved solutes between the lipid bilayers (Ghosh and Bachhawat, 1991). Also contemplated is a gene construct complexed with Lipofectamine (Gibco BRL). One of ordinary skill in the art would be familiar with techniques utilizing liposomes and lipid formulations.
[0155]Lipid based non-viral formulations provide an alternative to adenoviral gene therapies. Although many cell culture studies have documented lipid based non-viral gene transfer, systemic gene delivery via lipid based formulations has been limited. A major limitation of non-viral lipid based gene delivery is the toxicity of the cationic lipids that comprise the non-viral delivery vehicle. The in vivo toxicity of liposomes partially explains the discrepancy between in vitro and in vivo gene transfer results. Another factor contributing to this contradictory data is the difference in liposome stability in the presence and absence of serum proteins. The interaction between liposomes and serum proteins has a dramatic impact on the stability characteristics of liposomes (Yang and Huang, 1997). Cationic liposomes attract and bind negatively charged serum proteins. Liposomes coated by serum proteins are either dissolved or taken up by macrophages leading to their removal from circulation. Current in vivo liposomal delivery methods use subcutaneous, intradermal, or intracranial injection to avoid the toxicity and stability problems associated with cationic lipids in the circulation. The interaction of liposomes and plasma proteins is responsible for the disparity between the efficiency of in vitro (Felgner et al., 1987) and in vivo gene transfer (Zhu et al., 1993; Solodin et al., 1995; Thierry et al., 1995; Tsukamoto et al., 1995; Aksentijevich et al, 1996).
[0156]The production of lipid formulations often is accomplished by sonication or serial extrusion of liposomal mixtures after (I) reverse phase evaporation (II) dehydration-rehydration (III) detergent dialysis and (IV) thin film hydration. Once manufactured, lipid structures can be used to encapsulate compounds that are toxic (chemotherapeutics) or labile (nucleic acids) when in circulation. Liposomal encapsulation has resulted in a lower toxicity and a longer serum half-life for such compounds (Gabizon et al., 1990). Numerous disease treatments are using lipid based gene transfer strategies to enhance conventional or establish novel therapies, in particular therapies for treating hyperproliferative diseases.
G. Screening Assays
[0157]The present invention also contemplates the screening of candidate substances for the ability to modulate growth of a neuron. Particularly preferred candidate substances will be those useful in stimulating directional axonal growth along the A-P axis of the spinal cord. In the screening assays of the present invention, the candidate substance may first be screened for basic biochemical activity and then tested for its ability to modulate activity, at the cellular, tissue or whole animal level. In certain embodiments, an explant assay such as an assay using cultured spinal cord sections may be used in the screening methods. Any method known to those of skill in the art may be used in the claimed invention to conduct the screening assays.
[0158]1. Modulators and Assay Formats
[0159]a. Assay Formats
[0160]The present invention provides methods of screening for modulators of growth of a neuron. In one embodiment, the present invention is directed to a method of:
[0161](a) obtaining a candidate substance;
[0162](b) contacting the candidate substance with a neuron; and
[0163](c) measuring modulation of growth of the neuron.
[0164]In an example of yet another embodiment, the assay looks at anterior turning of axons of the neuron.
[0165]b. Inhibitors and Activators
[0166]An inhibitor according to the present invention may be one which exerts an inhibitory effect on the growth of a neuron. By the same token, an activator according to the present invention may be one which exerts a stimulatory effect on the growth of a neuron.
[0167]c. Candidate Substances
[0168]As used herein, the term "candidate substance" refers to any molecule that may potentially modulate regeneration of a neuron. The candidate substance may be a protein or fragment thereof, a polypeptide, a peptide, a small molecule inhibitor, or even a nucleic acid molecule. It may prove to be the case that the most useful pharmacological compounds will be compounds that are structurally related to compounds which interact naturally with Wnts, Wnt-like substances, or chemical compounds affecting Wnt signaling pathways. Creating and examining the action of such molecules is known as "rational drug design," and include making predictions relating to the structure of target molecules.
[0169]The goal of rational drug design is to produce structural analogs of biologically active polypeptides or target compounds. By creating such analogs, it is possible to fashion drugs which are more active or stable than the natural molecules, which have different susceptibility to alteration or which may affect the function of various other molecules. In one approach, one would generate a three-dimensional structure for a Wnt, and then design a molecule for its ability to interact with the Wnt. Alternatively, one could design a partially functional fragment of a Wnt or a Wnt-like substance (binding, but no activity), thereby creating a competitive inhibitor. This could be accomplished by x-ray crystallography, computer modeling or by a combination of both approaches.
[0170]It also is possible to use antibodies to ascertain the structure of a target compound or inhibitor. In principle, this approach yields a pharmacore upon which subsequent drug design can be based. It is possible to bypass protein crystallography altogether by generating anti-idiotypic antibodies to a functional, pharmacologically active antibody. As a mirror image of a mirror image, the binding site of anti-idiotype would be expected to be an analog of the original antigen. The anti-idiotype could then be used to identify and isolate peptides from banks of chemically- or biologically-produced peptides. Selected peptides would then serve as the pharmacore. Anti-idiotypes may be generated using the methods described herein for producing antibodies, using an antibody as the antigen.
[0171]On the other hand, one may simply acquire, from various commercial sources, small molecule libraries that are believed to meet the basic criteria for useful drugs in an effort to "brute force" the identification of useful compounds. Screening of such libraries, including combinatorially generated libraries (e.g., peptide libraries), is a rapid and efficient way to screen large number of related (and unrelated) compounds for activity. Combinatorial approaches also lend themselves to rapid evolution of potential drugs by the creation of second, third and fourth generation compounds modeled of active, but otherwise undesirable compounds.
[0172]Candidate compounds may include fragments or parts of naturally-occurring compounds or may be found as active combinations of known compounds which are otherwise inactive. It is proposed that compounds isolated from natural sources, such as animals, bacteria, fungi, plant sources, including leaves and bark, and marine samples may be assayed as candidates for the presence of potentially useful pharmaceutical agents. It will be understood that the pharmaceutical agents to be screened could also be derived or synthesized from chemical compositions or man-made compounds. Thus, it is understood that the candidate substance identified by the present invention may be polypeptide, polynucleotide, small molecule inhibitors or any other compounds that may be designed through rational drug design starting from known modulators of neuronal growth.
[0173]Other suitable inhibitors include antisense molecules, ribozymes, and antibodies (including single chain antibodies).
[0174]It will, of course, be understood that all the screening methods of the present invention are useful in themselves notwithstanding the fact that effective candidates may not be found. The invention provides methods for screening for such candidates, not solely methods of finding them.
[0175]2. In Vitro Assays
[0176]A quick, inexpensive and easy assay to run is a binding assay. Binding of a molecule to a target may, in and of itself, be inhibitory, due to steric, allosteric or charge-charge interactions. This can be performed in solution or on a solid phase and can be utilized as a first round screen to rapidly eliminate certain compounds before moving into more sophisticated screening assays. In one embodiment of this kind, the screening of compounds that bind to a Wnt or fragment thereof is provided.
[0177]The target may be either free in solution, fixed to a support, expressed in or on the surface of a cell. Either the target or the compound may be labeled, thereby permitting determining of binding. In another embodiment, the assay may measure the inhibition of binding of a target to a natural or artificial substrate or binding partner (such as a Wnt). Competitive binding assays can be performed in which one of the agents (Wnt) is labeled. Usually, the target will be the labeled species, decreasing the chance that the labeling will interfere with the binding moiety's function. One may measure the amount of free label versus bound label to determine binding or inhibition of binding.
[0178]A technique for high throughput screening of compounds is described in WO 84/03564. Large numbers of small peptide test compounds are synthesized on a solid substrate, such as plastic pins or some other surface. The peptide test compounds are reacted with, for example, with a Wnt, and washed. Bound polypeptide is detected by various methods.
[0179]Purified target, such as the Wnt, can be coated directly onto plates for use in the aforementioned drug screening techniques. However, non-neutralizing antibodies to the polypeptide can be used to immobilize the polypeptide to a solid phase. Also, fusion proteins containing a reactive region (preferably a terminal region) may be used to link an active region (e.g., the C-terminus of the Wnt) to a solid phase.
[0180]Explant culture assays, such as the collagen gel assays described above, are very convenient systems to test the function of the Wnts, Wnt-like substances, and chemical compounds affecting a Wnt signaling pathway in axonal growth and guidance before applying them to animal-based tests. They can also be used as screening methods.
[0181]3. In Cyto Assays
[0182]Various cell lines that express a Wnt, a Wnt-like substance, and/or a chemical compound affecting a Wnt signaling pathway can be utilized for screening of candidate substances. For example, cells containing a Wnt or a Wnt-like substance with an engineered indicator can be used to study various functional attributes of candidate compounds. In such assays, the compound would be formulated appropriately, given its biochemical nature, and contacted with a target cell.
[0183]Depending on the assay, culture may be required. As discussed above, the cell may then be examined by virtue of a number of different physiologic assays (e.g., axon growth). Alternatively, molecular analysis may be performed in which the function of a Wnt or a Wnt-like substance and related pathways may be explored. This involves assays such as those for protein expression, enzyme function, substrate utilization, mRNA expression (including differential display of whole cell or polyA RNA) and others.
[0184]4. In Vivo Assays
[0185]The present invention particularly contemplates the use of various animal models. Transgenic animals may be created with constructs that permit Wnt expression and activity to be controlled and monitored. The generation of these animals has been described elsewhere in this document.
[0186]Treatment of these animals with test compounds will involve the administration of the compound, in an appropriate form, to the animal. Administration will be by any route the could be utilized for clinical or non-clinical purposes, including but not limited to oral, nasal, buccal, or even topical. Alternatively, administration may be by intrathecal, intratracheal instillation, bronchial instillation, intradermal, subcutaneous, intramuscular, intraperitoneal or intravenous injection. Specifically contemplated are systemic intravenous injection, regional administration via blood or lymph supply.
[0187]5. Production of Inhibitors
[0188]In an extension of any of the previously described screening assays, the present invention also provide for methods of producing inhibitors. The methods comprising any of the preceding screening steps followed by an additional step of "producing the candidate substance identified as a modulator of" the screened activity.
H. Pharmaceutical Preparations
[0189]Pharmaceutical preparations of a Wnt, a Wnt-like substance, and/or a chemical compound affecting a Wnt signaling pathway for modulation of growth of a neuron in a mammal are contemplated by the present invention.
[0190]1. Formulations
[0191]Any type of pharmaceutical preparation of a Wnt, a Wnt-like substance, a chemical compound affecting a Wnt signaling pathway, another axonal guidance molecule, and/or substance that blocks a neuronal inhibitor is contemplated by the current invention. One of skill in art would be familiar with the wide range of types of pharmaceutical preparations that are available, and would be familiar with skills needed to generate these pharmaceutical preparations.
[0192]In certain embodiments of the present invention, the pharmaceutical preparation will be an aqueous composition. Aqueous compositions of the present invention comprise an effective amount an of a Wnt, a Wnt-like substance, a chemical compound affecting a Wnt signaling pathway, another axonal guidance molecule, and/or substance that blocks a neuronal inhibitor, and the like, dissolved or dispersed in a pharmaceutically acceptable carrier or aqueous medium. Aqueous compositions of gene therapy vectors expressing any of the foregoing are also contemplated. The phrases "pharmaceutical composition" or "pharmaceutical preparation" or "pharmacologically effective" or "pharmaceutically acceptable" refer to molecular entities and compositions that do not produce an adverse, allergic or other untoward reaction when administered to an animal, or a human, as appropriate.
[0193]As used herein, "pharmaceutical preparation" includes any and all solvents, dispersion media, coatings, antibacterial and antifungal agents, isotonic and absorption delaying agents and the like. The use of such media and agents for pharmaceutical active substances is well known in the art. Except insofar as any conventional media or agent is incompatible with the active ingredient, its use in the therapeutic compositions is contemplated. Supplementary active ingredients can also be incorporated into the compositions. For human administration, preparations should meet sterility, pyrogenicity, general safety and purity standards as required by FDA Office of Biologics standards.
[0194]The biological material should be extensively dialyzed to remove undesired small molecular weight molecules and/or lyophilized for more ready formulation into a desired vehicle, where appropriate. The active compounds will then generally be formulated for administration by any known route, such as parenteral administration. The preparation of an aqueous composition containing an active agent of the invention disclosed herein as a component or active ingredient will be known to those of skill in the art in light of the present disclosure.
[0195]An agent or substance of the present invention can be formulated into a composition in a neutral or salt form. Pharmaceutically acceptable salts, include the acid addition salts (formed with the free amino groups of the protein) and which are formed with inorganic acids such as, for example, hydrochloric or phosphoric acids, or such organic acids as acetic, oxalic, tartaric, mandelic, and the like. A person of ordinary skill in the art would be familiar with techniques for generation of salt forms. The carrier can also be a solvent or dispersion medium containing, for example, water, ethanol, polyol (for example, glycerol, propylene glycol, and liquid polyethylene glycol, and the like), suitable mixtures thereof, and vegetable oils.
[0196]The present invention contemplates a Wnt, a Wnt-like substance, a chemical compound affecting a Wnt signaling pathway, another axonal guidance molecule, and/or substance that blocks a neuronal inhibitor that will be in pharmaceutical preparations that are sterile solutions for intravascular injection or for application by any other route. A person of ordinary skill in the art would be familiar with techniques for generating sterile solutions for injection or application by any other route. Sterile injectable solutions are prepared by incorporating the active compounds in the required amount in the appropriate solvent with various of the other ingredients familiar to a person of skill in the art.
[0197]Upon formulation, solutions will be administered in a manner compatible with the dosage formulation and in such amount as is therapeutically effective. The formulations are easily administered in a variety of dosage forms, such as the type of injectable solutions described above, but drug release capsules and the like can also be employed.
[0198]For parenteral administration in an aqueous solution, for example, the solution should be suitably buffered if necessary and the liquid diluent first rendered isotonic with sufficient saline or glucose. These particular aqueous solutions are especially suitable for intravenous, intramuscular, subcutaneous and intraperitoneal administration. In this connection, sterile aqueous media which can be employed will be known to those of skill in the art in light of the present disclosure. Formulations for administration via lumbar puncture into the cerebrospinal fluid are also contemplated by the present invention.
[0199]The active agents disclosed herein may be formulated within a therapeutic mixture to comprise about 0.0001 to 1.0 milligrams, or about 0.001 to 0.1 milligrams, or about 0.1 to 1.0 or even about 10 milligrams per dose or so. Multiple doses can also be administered.
[0200]In addition to the compounds formulated for parenteral administration, such as intravenous injection or via lumbar puncture, other pharmaceutically acceptable forms include, e.g., tablets or other solids for oral administration; liposomal formulations; and time release capsules.
[0201]Oral formulations include such normally employed excipients as, for example, pharmaceutical grades of mannitol, lactose, starch, magnesium stearate, sodium saccharine, cellulose, magnesium carbonate and the like. These compositions take the form of solutions, suspensions, tablets, pills, capsules, sustained release formulations or powders. A person of ordinary skill in the art would be familiar with well-known techniques for preparation of oral formulations. Such compositions and preparations should contain at least 0.1% of active compound. The percentage of the compositions and preparations may, of course, be varied and may conveniently be between about 2 to about 75% of the weight of the unit, or preferably between 25-60%. The amount of active compounds in such therapeutically useful compositions is such that a suitable dosage will be obtained.
[0202]The use of liposomes and/or nanoparticles is also contemplated for the introduction of the modulator of cell death or gene therapy vectors into host cells. The formation and use of liposomes is generally known to those of skill in the art.
[0203]2. Dosage
[0204]An effective amount of the therapeutic or preventive agent is determined based on the intended goal, for example inhibition of cell death. The quantity to be administered, both according to number of treatments and dose, depends on the subject to be treated, the state of the subject and the protection desired. Precise amounts of the therapeutic composition also depend on the judgment of the practitioner and are peculiar to each individual.
[0205]In certain embodiments, it may be desirable to provide a continuous supply of the therapeutic compositions to the patient. For example, following traumatic spinal cord injury, a continuous administration of the therapeutic agent may be administered for a defined period of time, such as direct injection into the cerebrospinal fluid. For various approaches, delayed release formulations could be used that provide limited but constant amounts of the therapeutic agent over an extended period of time. Continuous perfusion of the region of interest may be preferred.
[0206]Those of skill in the art are well aware of how to apply gene delivery to in vivo and ex vivo situations. For viral vectors, one generally will prepare a viral vector stock. Depending on the kind of virus and the titer attainable, one will deliver 1×104, 1×105, 1×106, 1×107, 1×108, 1×109, 1×1010, 1×1011 or 1×1012 infectious particles to the patient. Similar figures may be extrapolated for liposomal or other non-viral formulations by comparing relative uptake efficiencies. Formulation as a pharmaceutically acceptable composition is discussed above.
[0207]3. Tracers to Monitor Gene Expression Following Administration
[0208]Certain embodiments of the present invention employ delivery of a Wnt, a Wnt-like substance, a chemical compound affecting a Wnt signaling pathway, another axonal guidance molecule, and/or substance that blocks a neuronal inhibitor to the target area of interest using expression cassettes. It may be important to determine whether the target site has been effectively contacted with the expression cassette. This may be accomplished by identifying cells in which the expression construct is actively producing the desired polypeptide product. Tagging of the exogenous polypeptide with a tracer element would provide definitive evidence for expression of that molecule and not an endogenous version thereof. Thus, the methods and compositions of the claimed invention may involve tagging of the polypeptide encoded by the expression cassette with a tracer element. A person of ordinary skill in the art would be familiar with these methods of tagging the encoded polypeptide.
I. Combination Therapy
[0209]In order to increase the effectiveness of the compositions and methods disclosed herein, it may be desirable to combine a variety of agents into one or more pharmaceutical compositions that can be administered in a regime that is effective in the treatment of the neuronal injuries or disorders described herein. As discussed elsewhere in this specification, those of skill in the art may wish to apply a combination of neuronal attractive, repellant, inhibitory, and/or inhibition blocking substances to the neurons to facilitate appropriate neuronal growth and/or function. This may involve contacting the neuron or spinal cord with these agent(s) at the same time. This may be achieved by contacting the neuron or spinal cord with a single composition or pharmacological formulation that includes multiple agents, or by contacting the cell with two distinct compositions or formulations, at the same time.
[0210]Alternatively, the agents may be applied to the neuron or spinal cord in series or succession at intervals ranging from minutes to weeks. In embodiments where two agent are applied separately to the neuron or spinal cord, one may wish ensure that a significant period of time did not expire between the time of each delivery, such that the agents will be able to exert an advantageously combined effect on the neuron(s). In such instances, it is contemplated that one may contact the cell with both modalities within about 12-24 hours of each other and, more preferably, within about 6-12 hours of each other. In some situations, it may be desirable to extend the time period for treatment significantly, however, where several days (2, 3, 4, 5, 6 or 7) to several weeks (1, 2, 3, 4, 5, 6, 7 or 8) lapse between the respective administrations. In other embodiments, two or more agents applied separately to the neuron or spinal cord with sufficient such that the agents will be able to separately exert their beneficial therapeutic effects on the neurons. In such instances, it is contemplated that one may contact the cell with both modalities In some situations, it may be desirable to extend the time period for treatment such that several days (2, 3, 4, 5, 6 or 7) to several weeks (1, 2, 3, 4, 5, 6, 7 or 8) lapse between the respective administrations.
[0211]Various combinations, in an exemplary embodiment, may be employed. For example, any number of regimes may be employed as set forth bells where "A" is a Wnt, Wnt-like substance, or chemical compound effecting a Wnt-signaling pathway and "B" a further Wnt, Wnt-like substance, or chemical compound effecting a Wnt-signaling pathway, a compound providing attractive or repellant guidance to neuronal growth, inhibitor of neuronal growth, or blocker of an inhibitor of neuronal growth:
TABLE-US-00003 A/B/A B/A/B B/B/A A/A/B A/B/B B/A/A A/B/B/B B/A/B/B B/B/B/A B/B/A/B A/A/B/B A/B/A/B A/B/B/A B/B/A/A B/A/B/A B/A/A/B A/A/A/B B/A/A/A A/B/A/A A/A/B/A.
[0212]Administration of the agents to a patient will follow general protocols for the administration as known to those of skill in the art and set-forth herein. It is expected that the treatment cycles may be repeated as necessary. It also is contemplated that various standard therapies, as well as surgical intervention, may be applied in combination with the application of the agents.
J. EXAMPLES
[0213]The following examples are included to demonstrate preferred embodiments of the invention. It should be appreciated by those of skill in the art that the techniques disclosed in the examples which follow represent techniques discovered by the inventor to function well in the practice of the invention, and thus can be considered to constitute preferred modes for its practice. However, those of skill in the art should, in light of the present disclosure, appreciate that many changes can be made in the specific embodiments which are disclosed and still obtain a like or similar result without departing from the spirit and scope of the invention.
Example 1
Materials and Methods
[0214]Collagen gel assays. E13 rat spinal cord explants were cultured in collagen gel matrix as described previously (Tessier-Lavigne et al., 1988; Zou et al., 2000). These explants are either "open-book" or post-crossing or pre-crossing for the spinal cord commissural axons. COS7 cells were transfected with various expression constructs with FuGene6 reagent (Roche). The explants were typically cultured for 16-20 hours and fixed in 4% PFA for two hours. The "open-book" explants were analyzed by lipophilic DiI labelling using iontophoresis. The post-crossing explants were stained with a monoclonal antibody (E7) against P3 tubulin (Hybridoma Bank for Developmental Studies). The pre-crossing explants were stained with a monoclonal antibody (4D7) against TAG-1 (Hybridoma Bank for Developmental Studies). Both antibodies were detected using secondary antibodies conjugated with horseradish peroxidase and visualized with 3,3'-diaminobenzene (DAB) (Sigma). Quantification of the post-crossing assays was done as described previously (Zou et al., 2000). The relative total axon bundle length was obtained by normalizing the total length of axons in the presence of Wnt-expressing COS cell aggregates against that in the presence of vector-only transfected COS cell aggregates. The explant assays were performed in three to four sets of multiple explants for each Wnt and an average fold of increase and a standard error were obtained for each Wnt from these sets. Therefore, the relative total length of vector only was defined as 1. n indicates the total number of explants for each construct.
[0215]Axon labelling. To reveal the commissural axon projections inside the spinal cord tissue, the inventor used D11 labelling. D11 is a lipophilic dye that becomes highly fluorescent when incorporated in membrane to reveal the shape of the cells and membrane protrusions. In order to focus on relatively smaller numbers of axons and produce more consistent and reproducible injection results, the inventor uses iontophoresis (Fraser, 1996) and point the injection sites with a micromanipulator (Fine Science Tools). DiI was dissolved in MeCl2 (Sigma) at 1 mg/ml. The Dye was delivered into spinal cord tissues with a SD9 current injector (Grass Telefactor). Glass needles were pulled with Narishige PC-10 pipette puller.
[0216]In situ hybridization. Mouse E10.9-E13.5 embryos were fixed for either whole-mount or section in situ hybridization as previously described (Keino-Masu et al., 1996; Zou et al., 1997). Specific probes for Wnt1, Wnt4, Wnt6 were obtained by PCR from Wnt1, Wnt4 and Wnt6 constructs in pcDNA1 (Fan et al., 1997) and subcloned into TOPO II vector (Invitrogen). Wnt5a and Wnt7b probes were obtained by RT-PCR from mouse E11.5 embryonic mRNA and subcloned in TOPO II vector.
[0217]Immunohistochemistry. E1.5 embryos of frizzled 3 knockout embryos, wild type and heterozygous littermates were fixed for immunohistochemistry with TAG-1 (4D7) antibody as previously described (Serafini et al., 1996).
[0218]Wnt and sFRP expression constructs. Wnt1, Wnt4 and Wnt6 full-length cDNA were subcloned into pcDNA3 with Myc epitope tag from pcDNA1 (Fan et al., 1997). Wnt5a expression construct in pCS2 was a kind gift from Dr. Xi He at Children's Hospital at Harvard Medical School and was subcloned into pcDNA3 with Myc epitope tag. Wnt7b cDNA was cloned by RT-PCR from E11.5 mouse embryonic mRNA and subcloned into pcDNA3 with Myc epitope tag. Mouse sFRP1 cDNA construct was a kind gift from Dr. Xi He (Finch et al., 1997). Mouse sFRP2 and sFRP3 cDNAs were cloned by RT-PCR from E1.5 mouse embryonic mRNA and subcloned into pcDNA3 with Myc Epitope tag.
[0219]Intrathecal injection. sFRP2 was overexpressed using the bacculovirus system (Lyuksyutova et al., 2003). The overexpressed sFRP2 is tagged with 6×His epitope and can be purified with affinity columns. Purified sFRP2 protein was dialyzed into artificial cerebrospinal fluid and injected into postnatal day 1 mice and rats, followed by one more injection on postnatal day 3. At postnatal day 5, animals were sacrificed, fixed by cardiac perfusion, and dissected for obtaining the spinal cord tissue. Serial sections were obtained along the A-P axis, and the CST axons will be examined by immunohistochemistry.
[0220]Behavioral test of injected animals. The functional consequence of sFRP2 injection will be assessed by observing the movement behavior of the injected mice and measuring the strength of the hind paw. A pilot set of experiments with 12 rats and found that 50% of the injected animals displayed a reduction in CST fibers, and that approximately 50% of the injected animals showed splayed hind paws and slowed movement at two weeks after birth.
Example 2
The A-P Guidance Cue(s) is Diffusible
[0221]When a segment of E13 rat spinal cord is cultured in collagen gel for 16-18 hours, commissural axons were observed to project ventrally, cross the midline and turn anteriorly within the explant, mimicking their in vivo pathfinding. Commissural axon trajectories in these "open-book" explants can be revealed by lipophilic D11 injection into the dorsal side of the explants by iontophoresis (Fraser, 1996). Most of the commissural axons in E13 rat spinal cord "open-book" preparations fixed immediately after dissection (without culturing) are only just approaching the midline or in the process of midline crossing. Therefore, the midline crossing and anterior turning of the commissural axons observed with DiI labeling occurred during the "open-book" culture period.
[0222]FIG. 1A schematically demonstrates that during embryonic development, commissural neurons project axons to the ventral midline. Once they reach the floor plate, they cross the midline and enter the contralateral side of the spinal cord, as diagrammed in FIG. 1B. It was reasoned that if A-P guidance is controlled by a diffusible gradient of either an attractant(s) or a repellents(s), then cutting the "open-book" explants shorter might lead to the loss of the gradient within the explants and therefore lead to abnormal pathfinding along the anterior-posterior axis (FIG. 1C); if A-P guidance is controlled by a non-diffusible cue(s), commissural axons will still have the normal anterior turn in shorter explants, because the gradient will be maintained (FIG. 1D).
[0223]Open-book" explants of different anterior-posterior lengths (3 mm, 2 mm, 1 mm and 0.5 mm) were systematically cultured and commissural axon growth was analyzed using focal D11 injection by iontophoresis into the dorsal spinal cord. When the length was reduced to 0.5 mm, abnormal pathfinding behavior of the post-crossing commissural axons was consistently observed, which included knotting, stalling and randomized turning along the A-P axis. This behavior contrasted sharply with that observed in 3 mm explants, in which all axons turned anteriorly. In both short and long explants, commissural axon pathfinding from the dorsal spinal cord to the floor plate was normal. These results were quantified and are shown in FIG. 1E. Because each DiI injection labels a cohort of axons, the inventor quantified the results by categorizing axonal behavior observed for each DiI injection site, as previously described (Zou et al., 2000). If all axons turned anteriorly in one injection, it was counted as an anterior (correct) turn; if many axons appeared to stall or make knots after midline crossing, it was counted as "knotting/stalling"; if a significant number of axons projected posteriorly or all axons projected posteriorly, it was counted as "random turn (A/P)". The frequency of each category is presented as percentage of all injected sites. Some of the sites display both knotting/stalling and random turn behavior so that the total percentage can be greater than 100%. All of the post-crossing commissural axons in the long explants turned correctly. In the short explants, axons formed knots or stalled after midline crossing, or turned randomly both anteriorly and posteriorly. Only 18% of the injection sites in the short explants showed normal anterior turning, presumably due to the loss of guidance information in the short explants. Therefore, the guidance cue(s) that directs the anterior turn is likely diffusible. These results do not address the source of the diffusible cue(s) in the neural tube or how the gradient is established. The diffusible cue(s) can be either expressed at differential levels along the anterior-posterior axis of the spinal cord or secreted from an anterior or a posterior tissue source.
Example 3
The A-P Guidance Cue(s) is Attractive
[0224]To address whether the A-P guidance cue is attractive or repulsive, DiI was focally injected into the dorsal spinal cord close to the anterior end, in the middle and close to the posterior end of the long explants (3 mm or longer). The axons in the middle and close to the posterior end of the explants were found to always project anteriorly, whereas the axons close to the anterior end almost always make mistakes: they either stall after they cross the midline or they project both anteriorly and posteriorly after midline crossing, or sometimes only posteriorly. The results were quantified using the same criteria as shown in FIG. 1E. The quantification is shown in FIG. 2B. The axons close to the anterior end of the explants behave similarly to those in the short explants (0.5 mm), whereas the axons in the middle and posterior part of the explants behave normally. These results are consistent with the possibility that a gradient of an attractive cue(s) plays a role in the anterior turn of the post-crossing commissural axons. Interestingly, it was consistently found that the axons close to the anterior end of the explant have a much higher frequency (93%) of turning posteriorly than those in the shorter explants (64%). It is possible that the remaining attractant(s) in the middle and posterior parts of the longer explants creates a counter gradient after the attractant(s) diffuse out from the anterior end, turning the axons posteriorly. This abnormal behavior of the anterior injection sites is true for explants taken anywhere along the entire length of the spinal cord, suggesting that a general anterior-posterior gradient of diffusible attractant(s) controls the anterior turn of the post-crossing commissural axons along the length of the spinal cord.
[0225]It is possible that the axons located close to the anterior end of the long explants might be misrouted, because the gradient might be destroyed due to diffusion of the attractant(s) out of the explants, whereas the axons close to the posterior end will turn normally, as the tissue anterior to these turning points will still contain higher concentrations of the attractant(s) (FIG. 2A, upper panel). On the other hand, if the cue(s) were repulsive, the axons close to the posterior end of the explants might not be able to turn anteriorly correctly because the gradient might be disrupted due to the diffusion of the repellent(s) out of the explants, whereas the axons at the anterior end of the explants will not be affected, because the tissue posterior to the injection site will still contain higher amounts of the repellent(s) (FIG. 2A, bottom panel).
[0226]In order to rule out the possibility that cutting at the anterior end itself produces a repulsive signal, which repels post-crossing commissural axons, studies were conducted to determine whether a cut in the "open-book" explants can prevent axons from projecting rostrally. A cut was introduced within the explants on one side of the "open-book" spinal cords. The spinal cord explants were cultured overnight and the contralateral dorsal spinal cord explants were injected 200 μm-300 μm posterior to the cut site. Commissural axons still projected rostrally and could traverse the cut site, behaving as if they were in the middle of the long "open-book" explants.
[0227]Although the cut spinal cords sometimes appeared to be reconnected after overnight culture, they are not sealed back and can be easily separated again at the cut site. And yet, axons can grow through the cut site. This suggests that the A-P gradient of the guidance cue(s) is preserved in such a preparation and a cut (damage) to the spinal cord itself does not produce a cue(s) to repel post-crossing commissural axons. In fact, these axons were faced with two "copies" of cut edge compared to those in short explants. If cut edge produced a repellent, then axons posterior to the internal cut edge would display more severe defects than those in short explants alone. This also demonstrates that the distance between the anterior injection sites and the border of the explants (200 μm-300 μm) is sufficient for commissural axons to turn anteriorly and the failure of anterior turning in short "open-book" explants and at the anterior end of long "open-book" explants is not due to spatial or physical restrictions but rather due to the disruption of the gradient of a guidance cue(s). These results are all consistent with an interpretation that the abnormal axonal behavior at the anterior end of the "open-book" explants is caused by the disruption of a gradient of an attractive molecule(s).
Example 4
Wnt Family Proteins are Candidate A-P Guidance Cue(s)
[0228]To identify the diffusible guidance cue(s) directing the anterior turn after midline crossing, a candidate gene approach was used. It had been observed that an embryonic limb bud can stimulate the extension of commissural axons only after they have crossed the midline using the "post-crossing" explant assay (Zou et al., 2000). In this assay, commissural axons grow out of the explant after crossing the floor plate, making it possible to test the effects of secreted factors on the axons (see diagram in FIG. 3A). As axon guidance molecules are often expressed in multiple tissues during development, it was hypothesized that the factor(s) in the limb bud that stimulates extension of post-crossing commissural axons might be related to the attractant(s) that affect these same axons in vivo (Serafini et al., 1996; Ebens et al., 1996). Therefore, candidates expressed in the limb bud were tested using the post-crossing commissural axon explant assay by expressing these molecules in COS cell aggregates positioned next to post-crossing explants in collagen gels (FIG. 3A). Candidate molecules found in the limb bud include HGF (Ebens et al., 1996), FGF4 (Bueno and Heath, 1996), FGF8 (Bueno and Heath, 1996), BMP4 (Francis et al., 1994), BMP7 (Hofmann et al., 1996; Augsburger et al., 1999), Shh (Bueno and Heath, 1996), and Wnt1 (Zakany and Duboule, 1993). Wnt4 was also tested, because it is expressed in the floor plate (Ungar et al., 1995; Liu et al., 2000; Saulnier et al., 2002) and Wnt 6 (Fan et al., 1997). Of these factors, only Wnt1, Wnt4 and Wnt6 were found to stimulate the extension of the post-crossing commissural axons. Additional Wnt proteins that are expressed either in the spinal cord or in the limb bud were tested, namely Wnt5a (Dealy et al., 1993) and Wnt7b (Parr et al., 2001; Shu et al., 2002), and found that these two Wnts can also stimulate the extension of the post-crossing commissural axons. Wnt1 stimulates post-crossing axon extension relatively weakly, whereas Wnt4, Wnt5a and Wnt7b can increase the extension of post-crossing axons by 2-3 fold on average (FIG. 3B). None of these Wnts affect the outgrowth of pre-crossing commissural axons, in contrast to Netrin-1, used as a positive control (Serafini et al., 1994).
[0229]If a gradient of diffusible attractant(s) guide commissural axons anteriorly, it might be expected that the tissues anterior to commissural axons can attract post-crossing commissural axons. From previous work of the inventor, both the spinal cord and the floor plate have a potent net repulsive effect to post-crossing commissural axons (Zou et al., 2000). It is possible that the attractant(s) for post-crossing axons are not as diffusible as Semaphorins and Slit proteins precluding the possibility of revealing the function of the attractant(s) in the post-crossing collagen gel assays. Alternatively, the attractant(s) might be expressed in a more restricted fashion and cannot produce a consistently strong attractive effect in assays depending on the orientations of tissues in cultures. In order to circumvent this obstacle and test the model of anterior attractant(s), the function of a major brain target for commissural axons, the ventral-posterior-lateral nucleus of the thalamus, was examined, which is the synaptic target of the spinothalamic tracts (FitzGerald, 1996). The inventor found that the E13.5 ventral-posterior-lateral nucleus can similarly stimulate the extension of the post-crossing commissural axons by three fold (FIG. 3C). In contrast, at an earlier stage (E11.5), the diencephalon region destined to be the ventral posterior thalamus does not have any growth stimulating activity, suggesting that the E13.5 thalamus activity is specific. At E11.5, the earliest populations of commissural axons just crossed the midline and turned anteriorly inside the spinal cord and have not reached the forebrain yet.
[0230]To determine whether any of these Wnts are likely to affect commissural axon growth in vivo, the expression patterns of Wnts were examined by in situ hybridization in developing mouse embryos during the stages when commissural axons are crossing the midline and turning anteriorly into their longitudinal pathway. Expression of some of these genes in the developing spinal cord has been examined before (Kispert et al., 1996; Liu et al., 2000; Saulnier et al., 2002; Shu et al., 2002; Krylova et al., 2002). At E11.5 (equivalent to E13 rat), Wnt1 is expressed at high levels in the roof plate but diffusely and weakly throughout the spinal cord. Wnt4 is specifically enriched in the floor plate and the ventricular zone and has a decreasing anterior-to-posterior gradient along the entire length of the floor plate at E10.5 as well as E13.5, whereas the expression in the ventricular zone does not show any gradient. A similar anterior-posterior gradient of Wnt4 expression was also observed in the floor plate of E11.5 and E12.5 mouse embryos (data not shown). Wnt5a is expressed widely in the spinal cord but is particularly abundant in the ventral areas of the spinal cord next to the lateral funiculus. Wnt7b is expressed in the ventricular zone of the spinal cord and specifically on the two lateral margins of the floor plate, where the anterior turning of the post-crossing commissural axons occurs. Wnt7b appears to have a decreasing anterior-to-posterior gradient in the ventricular zone but does not display an A-P gradient in the floor plate. Wnt6 and Wnt11 (Kispert et al., 1996) are not expressed in the spinal cord. Wnt3 is expressed in the motor columns but not in the ventral midline or the ventral or lateral funiculi (Krylova et al., 2002) and therefore may not be relevant to commissural axon pathfinding along the anterior-posterior axis. Therefore, several Wnts are expressed in the right place at the right developmental stages to function as regulators of the growth of the post-crossing commissural axons. In particular, the Wnt4 expression displays a clear anterior-posterior gradient along the entire length of the floor plate throughout the time when commissural axons are turning anteriorly after midline crossing (from E10.5 to E13.5). This suggests that Wnt4 might play a role in the anterior-posterior turning decision of post-crossing commissural axons along the entire length of the spinal cord. Interestingly, a similar Wnt4b gradient in the floor plate along the anterior-posterior axis has also been found in zebrafish embryos at similar developmental stages (Liu et al., 2000). Because the ventral posterior lateral nucleus of the thalamus can stimulate the extension of the post-crossing commissural axons, the inventor tested whether any of the Wnt genes are expressed in the thalamus. The inventor found that Wnt1 and Wnt4 genes are expressed at high levels in the thalamus. At E13.5, Wnt4 is expressed in a highly restricted pattern in the thalamus, including the dorsal lateral geniculate nucleus (dLGN) and the ventral-posterior-lateral nucleus (VPL). Wnt1 is also expressed in the dLGN and the VPL at the same stage. Interestingly, Wnt4 and Wnt1 have reciprocal gradients. Wnt4 is expressed at higher level in the dLGN than in the VPL, whereas Wnt1 is expressed at higher level in the VPL than in the dLGN. However, both are expressed in the VPL and the areas used in the explant assays include the VPL. At E11.5, neither Wnt1 nor Wnt4 is expressed in the dorsal diencephalon region destined to be the VPL of the thalamus, consistent with the observation that E11.5 thalamus does not stimulate the extension of the post-crossing commissural axons. Based on the expression pattern of the Wnt genes, the Wnt protein(s) gradient is more likely formed by graded expression levels along the anterior-posterior axis rather than diffusion from the brain targets.
Example 5
SFRPs Can Disrupt Anterior-Posterior Guidance of Commissural Axons
[0231]To test directly whether Wnts are required for the proper anterior turn of the post-crossing commissural axons, potent Wnt inhibitors were used to block the function of all Wnts in the "open-book" explants. Secreted Frizzled-related proteins (sFRPs), are soluble proteins that bind to Wnt proteins with high affinities and thus can block the interaction of Wnts with their receptors, the Frizzleds (Wodarz and Nusse, 1998). sFRPs were produced in the "open-book" collagen gel assays by including sFRP-expressing COS cells in the bottom layer of collagen gel (FIG. 4A). The "open-book" of long spinal cord explants were placed on top of the bottom collagen and embedded in the top collagen gel. This system was first tested with Netrin-1 expressing cells in the bottom collagen and it was found that axons can extend from the pre-crossing spinal cord explants, suggesting that the molecules expressed in the bottom collagen can diffuse effectively into the top collagen. As a control, COS cells transfected with vector only and embedded in the bottom collagen had no growth-promoting activity.
[0232]It was found that in the presence of any of the three sFRPs (sFRP 1, sFRP2 and sFRP3) or a mixture of all three sFRPs, anterior turning of commissural axons after midline crossing are severely impaired. Instead, they either stall or turn randomly along the anterior-posterior axis, displaying behaviors similar to those observed in the short explant studies discussed above and the anterior injection sites discussed above. In contrast, in the presence of the vector-only-transfected COS cells in the bottom collagen, all commissural axons turned anteriorly after midline crossing. As shown in FIG. 4B, in the presence of sFRP1, only 11% of the injection sites displayed correct anterior turns; in the presence of sFRP2 or sFRP3, only about 25% of the injections sites turned correctly. Therefore, most of the injection sites showed abnormal projections along the A-P axis when the function of the Wnt proteins were blocked. A-P guidance of commissural axons at all anterior-posterior levels was disrupted in the presence of any of the sFRPs or a mixture of all sFRPs. No abnormal pathfinding behavior was observed in the pre-crossing segment of the commissural axons, suggesting that the Wnt signaling pathway is not required for the dorsal-ventral projection of the pre-crossing commissural axons. Similar anterior-posterior guidance defects of post-crossing commissural axons were observed when a purified Frizzled-8 ectodomain-Fc fusion protein was added to the "open-book" culture, whereas an Fc only control protein did not exert any effects.
Example 6
A Wnt4 Gradient Can Rescue A-P Guidance Defects and Reorient Axons Posteriorly
[0233]In short "open-book" explants, post-crossing axons lose A-P directionality presumably due to the disruption of a Wnt gradient. In order to further test this hypothesis, studies were conducted to determine whether applying a localized anterior source of Wnt protein(s) can rescue the anterior turn of commissural axons after midline crossing in these short explants. The inventor placed COS cell aggregates expressing Wnt4 anterior to the short explants and tested whether the post-crossing axons can turn towards the Wnt4 cell aggregates (FIG. 5A and FIG. 5B). It was found that Wnt4 expressing COS cells can attract post-crossing commissural axons and rescue A-P guidance defects found in short explants, whereas COS cells transfected with vector only had no effects (FIG. 5C). Only 25% of the explants displayed correct anterior turns in the vector only control, whereas 75% of the explants displayed clear turning towards the Wnt4-expressing COS cell aggregates. Thus, A-P pathfinding errors caused by loss of an A-P gradient of guidance cue(s) can be rescued when a Wnt4 gradient is applied.
[0234]To further test whether Wnt4 can function as an instructive cue to direct axon growth, studies were conducted to determine whether placing COS cell aggregates posterior to the short explants can reorient axons posteriorly (FIG. 5D and FIG. 5E). It was found that Wnt4 can readily redirect the growth of the post-crossing commissural axons to turn posteriorly, whereas the COS cell transfected with vector only did not affect the behavior of the post-crossing axons in the short explants, suggesting that Wnt4 is an instructive cue rather than permissive cue. Quantification of data was carried out using the same criteria throughout the these studies. For the reorientation experiments, if all axons turned posteriorly, that injection site was counted as posterior turn and shown in the bars to the far right in FIG. 5F.
[0235]In order to test whether anterior tissue contain instructive attractant(s) for commissural axons, studies were conducted to attempt to reorient post-crossing commissural axons posteriorly by putting the ventral-posterior thalamus posterior to the "open-book" explants. It was found that in contrast to the Wnt4-overexpressing COS cells, thalamus could not reproducibly reorient axons. The expression of Wnt proteins in the thalamus may not be sufficient to allow Wnt proteins to diffuse into the "open-book" explants to redirect axons. It was found that anterior spinal cord tissue could not reorient axons, either. The spinal cord contains potent repellents to post-crossing commissural axons, such as Sema3B, Sema3F and the Slit proteins, to prevent them from re-entering the grey matter and has a net repulsive effect on post-crossing commissural axons in collagen gel assays (Zou et al., 2000). The Wnt4 protein gradient in the spinal cord is only restricted to the floor plate. The rest of the ventricular zone does not have Wnt4 expression gradient. Therefore, it is very hard to recreate a Wnt4 counter gradient in the "open-book" assay by putting a piece of spinal cord posterior to the explants.
Example 7
Frizzled 3 is Required for Anterior-Posterior Guidance of the Post-Crossing Commissural Axons In Vivo
[0236]Three frizzled genes, which encode receptors for Wnts, fz3, fz8 and fz9, have been found to be expressed in the spinal cord (Borello et al., 1999). This was confirmed by in situ hybridization that fz3, fz8 and fz9 are indeed expressed in the spinal cord from E9.5 to E13.5 during the time when commissural axons are making anterior turns. Among the three frizzleds, fz3 is the most relevant, because it is expressed broadly in the spinal cord, covering the area where commissural neuron cell bodies are located. Interestingly, fz3 transcripts appear to be enriched in the ventral funiculi where post-crossing commissural axons are located at a E11.5, when a large number of commissural axons have already crossed the midline. Fz8 is expressed more weakly and is not expressed in the most dorsal portion of the spinal cord. Fz9 is only expressed in the ventricular zone where non-differentiated neurons are localized but not in the dorsal mantle zone where commissural neuron cell bodies are located. Commissural axon projections in fz3 knockout embryos (Wang et al., 2002) were examined by immunohistochemistry and DiI labeling with a monoclonal antibody against TAG-1, a commissural axonal marker that only labels the pre-crossing and the midline crossing segments of the commissural axons but not the post-crossing segment of the commissural axons. It was found that the dorsal-ventral projection of pre-crossing commissural axons were normal compared to wild type control, but post-crossing commissural axons projected randomly along the anterior-posterior axis after midline crossing with 100% penetrance. From crosses between fz3 heterozygotes, four litters among which were seven homozygous mutants were examined. For three of these litters, the dissected spinal cords were analyzed without knowledge of their genotypes. In these blinded experiments, 5/5 mutant and 11/11 wild type or heterozygous spinal cords were correctly identified; the probability of this occurring by chance is 4×10-5. It was found that in all injection sites, commissural axons either turned randomly along the anterior-posterior axis or stalled after midline crossing, whereas their pre-crossing trajectory was normal, consistent with the observations discussed above using explant assays, suggesting that the Wnt/Frizzled pathway is only required for anterior-posterior axon guidance after midline crossing in vivo. As previously reported, no spinal cord patterning defects were observed in the fz3 knockout mice at this stage of development as assessed by markers such as Nkx2.2, HNF-3β, Lim2, and Isl1 (Wang et al., 2002). Both the dorsal-ventral and anterior-posterior pathfinding of commissural axons are normal in LRP6-/- embryos although dramatic patterning defects were observed in these animals (Pinson et al., 2000), suggesting that the canonical Wnt/β-catenin signaling pathway is not involved in the differentiation, the dorsal-ventral pathfinding and the anterior-posterior guidance decision of commissural axons at the midline.
Example 8
Wnt Genes are Expressed in a "Half-Pipe" Gradient Along the Neonatal Spinal Cord
[0237]Because corticospinal tract axons project posteriorly along the dorsal funiculus of the spinal cord, the inventor examined the expression pattern of Wnt genes around the dorsal funiculus by in situ hybridization. The inventor cloned the entire family of rodent Wnt genes (including 19 members) and performed in situ hybridization at postnatal days 0 and 3 along the anterior-posterior axis. The inventor found that five Wnt genes are expressed in the dorsal midline and dorsal funiculus. Wnt1, and Wnt5a are expressed at a higher level. The other Wnts, Wnt7b, Wnt8a, and Wnt9a, are expressed at lower levels. Along the anterior-posterior axis, all of these Wnt genes have a high-to-low gradient from the cervical and thoracic level. Intriguingly, all these Wnt genes display a reverse gradient at the lumbar level: low-to-high gradient. Therefore, multiple Wnt genes are expressed in a biphasic gradient, or "half-pipe" gradient.
[0238]The biphasic gradient along the entire spinal cord suggests that Wnts first "push" CST axons posteriorly along the cervical and thoracic cord but then act as stop signal to terminate the CST axons at the lumber cord, much like the motion in a "half-pipe".
Example 9
Wnt Proteins Repel Frontal Cortical Axons
[0239]In order to test whether Wnts can guide corticospinal tract axons, the inventor performed explant assays to evaluate the function of Wnt proteins in frontal cortical axons in collagen gel. Postnatal day 0 brains were dissected out and sliced with tissue chopper. Layer 5 cortical explants were dissected from the frontal motor cortical region and culture in collagen for 60 hours. Long axons grew out in the collagen gel and are stained positively with a corticospinal tract marker, a monoclonal antibody against N-CAM, 5A5. COS cells were transfected with Wnt expression constructs and made into cell aggregates, and the inventor positioned the cell aggregates next to the cortical explants dissected out from postnatal P0 frontal cortex. The inventor found that Wnt1 protein potently inhibits the outgrowth of axons from the frontal cortex in these assays, suggesting that corticospinal tract axons might respond to Wnt proteins as they pathfind along the spinal cord in vivo. Very few axons grew out in the collagen gel, and the axon's length is much reduced as well. A slight repulsive effect can be observed. To address the possibility that the cell aggregates may be secreting too much Wnt1 protein so that axons cannot grow out of the explants, the inventor diluted the transfected COS cells with untransfected COS cells and found that Wnt1 shows robust repulsion when diluted. The inventor tested the function of Wnt1 on E18.5 cortical axons and found Wnt1 can only weakly repel frontal cortical axons. CST axons reach the spinal cord at P0. At E18.5, the CST axons are still in the midbrain and the hindbrain. The time course of Wnt1 responsiveness is consistent with it role in CST axon pathfinding once CST axons enter the spinal cord. Wnt5a also repel postnatal motor cortical axons.
Example 10
Wnt Proteins Also Regulate the A-P Pathfinding of the CST Axons
[0240]The inventor found that several Wnt genes are expressed in a high-to-low gradient in the gray matter cupping the dorsal funiculus from the cervical to the thoracic spinal cord where corticospinal tract axons first enter the spinal cord and project posteriorly at postnatal day 0. At the lumbar spinal cord, Wnt gene expression in the gray matter displays a reversed gradient (low-to-high) forming a "half-pipe" gradient along the entire length of the spinal cord. Such gradient persists from P0 to at least P5. The functional studies showed that Wnt proteins could repel axons from frontal motor cortex in a collagen gel assay. Therefore, first gradient guides CST axons to project from the cervical cord to the thoracic cord, and the second reverse gradient helps to stop CST axons at the lumbar level.
Example 11
A Repulsive Wnt Receptor, Ryk, is Expressed in the CST Axons
[0241]Along the Entire A-P trajectory Axon guidance molecules are often bi-functional, attracting some axons while repelling others, depending on the guidance receptor composition in the responding neurons. Vertebrate commissural axons are attracted by Wnts, whereas frontal cortical axons are repelled by Wnts. In Drosophila, Wnt5 was found to play a repulsive role in the pathway selection before midline crossing (Yoshikawa et al., 2003). This repulsion is mediated by a Wnt receptor called Derailed through direct binding and is independent of Frizzled (Yoshikawa et al., 2003). The inventor found that the vertebrate Derailed, Ryk (Halford et al., 2000), is not expressed in commissural axons, although Frizzled3 is, and Frizzled3 is required for mediating Wnt attraction (Lyuksyutova et al., 2003).
[0242]Further investigating why the cortical axons are repelled by Wnts, the inventor first generated an in situ probe for Ryk and found that the Ryk gene is expressed in layers 5 and 6 of the frontal cortex. The levels of Ryk expression at E18.5 are much lower than are that of P0. The inventor obtained a published antibody against the mouse Ryk protein (Kamitori et al., 2002) and performed immunohistochemistry, and the inventor found that Ryk protein is present in layer 5 neurons and is present in the internal capsule of E18.5 brain. The inventor then generated polyclonal antibodies against the extracellular domain of Ryk and further confirmed that Ryk protein is present in the CST axons forming the pyramidal decussation and the pyramidal tracts in the dorsal funiculus of the spinal cord. Therefore, Ryk is expressed in the CST axons at the right time to mediates Wnt repulsion.
Example 12
Ryk Antibodies can Block the Repulsion of CST Axons by Wnts
[0243]To demonstrate that Ryk is involved in mediating Wnt repulsion in vertebrate axons, the inventor used the polyclonal antibodies generated against the ectodomain of Ryk and tested whether the Ryk antibodies can block the repulsion by Wnts in collagen gel assays. The inventor found that addition of purified Ryk antibodies in collagen gel assays blocked the repulsive effects of Wnt proteins, suggesting that Ryk does mediate Wnt repulsion in vertebrates and may play important roles in CST axon guidance such as the anterior-posterior guidance of CST axons in vivo. The inventor found that in the presence of Wnt1 protein, frontal cortical axons tend to grow much shorter and away from the pointed source of Wnt1. When Ryk antibodies were included, frontal cortical axons were no longer repelled, and the outgrowth was increased.
Example 13
Intrathecal Injection of sFRP2 Protein at Cervical Level Caused Reduction of CST Fibers in the Dorsal Funiculus and Impaired Motor Function
[0244]To address the in vivo function of the repulsive effects of Wnt proteins on corticospinal tract axon guidance, the inventor injected purified sFRP2 protein to postnatal cervical spinal cord at P1, P3 and then analyzed the CST axon projection in P5 spinal cord. Transverse section of the vehicle and sFRP2 injected animals were collected every 800 um along the entire A-P axis of the P5 spinal cord and stained with a CST marker 5A5. The inventor found that the dorsal funiculus areas are much reduced in injected animals, suggesting that the posterior growth of CST axons was interfered. Similar results were obtained from multiple groups of mice and rats. Some animals were raised to adulthood and their motor functions were analyzed. The inventor found that the sFRP2 injected animals display consistent weakening of grip strength throughout the entire period of the tests, suggesting the posterior growth defects caused by sFPR2 injection interfered motor system development.
[0245]These studies suggest that Wnt proteins control not only the guidance of ascending sensory axons, but also that of the descending motor pathways through a Ryk-dependent signaling pathway.
Example 14
Additional Studies Involving Injection of Wnt Inhibitors into Spinal Cords
[0246]In addition to the studies described above, the sFRP2 protein has also been injected to the lumbar and sacral spinal cord on postnatal day 5 and 7 and animals were fixed on day 9. Data obtained from these studies will indicate whether inhibiting Wnt function in the posterior portion of the spinal cord will cause overshooting of corticospinal tract axons, leading to abnormal development of the motor system, and provide further information allowing one of skill to develop appropriate regimes for spinal cord regeneration.
[0247]Additionally, Anti-Ryk antibodies have also been injected to both the cervical and lumbar spinal cord regions to allow for the analysis of anatomical defects of motor axon growth and behavioral defects. These studies, have confirmed that Ryk is an inhibitor of Wnt-mediated action on neurons and a target for therapeutics.
Example 15
In Situ Hybridization Studies of Wnts Expression in Normal and Injured Spinal Cords
[0248]To study patterns of Wnt expression, the inventor cloned the entire family of Wnts and performed in situ hybridization. Most of the Wnts are no longer expressed in the adult spinal cord. One Wnt gene, Wnt5a, is expressed highly in the spinal cord. Wnt8a is weakly expressed.
[0249]Researchers have found that it is possible to regenerate sensory axons by blocking inhibitors of axon growth but it is nearly impossible to regenerate corticospinal cord (Sivasankaran et al. 2004). It is possible to that the Wnt5a is expressed in the adult spinal cord and other Wnts that become induced at injured sites in the spinal cord result in inhibition of normal cord growth. Because corticospinal tract axons are repelled by Wnts and sensory axons are attracted by Wnts, abnormal Wnt production after injury can result in selective inhibition of the motor cortical axons in the spinal cord. Any injury-induced Wnts, together with Wnt5a, may cause a repulsive environment so that the adult axons fail to regenerate.
[0250]One can use the data from in situ studies of normal and injured spinal cords to study whether various Wnt genes are induced upon spinal cord injury. To obtain data from injured spinal cords, an adult mouse spinal cord can be lesioned at cervical and thoracic levels by a hemi-section injury paradigm. The animals can be fixed at day 1, 7, 14 and one month after injury and the expression patterns of Wnt genes determined by in situ hybridization and compared to data from uninjured spinal cords.
[0251]Data from the studies described in this example can be used to determine appropriate substances to use to prevent any injury-induced Wnts from preventing proper neuronal regeneration.
Example 16
Transgenic Mice Studies
[0252]In order to further demonstrate the roles of Wnts in neuronal guidance and regeneration, a variety of transgenic mice lines were created. In these lines, generally, a dominant-negative inhibitor transgene is expressed to produce an inhibitor of a Wnt inside relevant neurons. These transgenic mice are produced by methods well-known to those of skill in the art.
[0253]For example, transgenic mice lines expressing specific dominant-negative inhibitors of Wnt intracellular signaling (dominant-negative disheveled) in a subset (Neurogenin-2 expressing) of commissural neurons (which likely give rise to pain sensory pathway and are attracted by Wnts to project to the brain) showed a kangaroo gait phenotype in the hindlimb. The gait (hopping) behavior appears to depend on the texture of the surface the mice were walking on, suggesting sensory system defects in these neuron. These data demonstrate that Wnt signaling is important for the normal wiring of the nervous system and support a cell-autonomous mechanism, meaning that the Wnt signaling pathway is required in the neurons which are responding to the Wnt gradient.
[0254]One can conduct further transgenic animal studies to show the roles in which the Wnt signaling pathway in axon sensory axon guidance and provide tools for axonal regeneration inhibitors of Wnt signaling. In this regard, dominant negative transgenic animals can be created in subsets of commissural neurons to further test the role of Wnt signaling pathway in commissural neurons. Axonal projection and mouse behavior can be analyzed in these animals.
[0255]Further once transgenic mice are created, spinal cord lesion experiments as described above can be carried out and the Wnt expression pattern in the injured spinal cord analyzed. The function of Wnts in adult spinal cord axons can be tested to see whether Wnts continue to attract sensory axons and repel motor axons. If Wnt attract sensory axons, the induction of Wnts may be helpful for axon regeneration. But if Wnts repel motor axons, the induced Wnts in the spinal cord will block regenerative growth of motor axons. In this case, anti-Ryk antibody, which blocks the repulsive function of Wnt specifically will be applied to block the inhibitory effects of Wnts on motor axon regeneration. Alternatively, interference of the Ryk signaling specifically will also block the repulsion, allowing regeneration to occur. These results will provide insights to how Wnts can be used to help spinal cord axon regeneration.
Example 17
Psychoactive Drugs in Combination with Wnt Therapy
[0256]Psychoactive drugs, such as amphetamine, improve functional recovery following stroke in experimental animals, suggesting a role in promoting nerve repair and regeneration (Long and Young, 2003). In view of the teachings of this specification, those of skill will be able to determine the effects of these drugs on Wnt signaling, axon guidance, and regeneration. Those of skill will then be able to further modify such drugs and/or their treatment regimes to enhance the drug effect on regeneration and reduce side effects without losing the effect on regeneration.
[0257]It is expected, in view of the teachings of this specification, that a combination of Wnt inhibitors or psychoactive drugs will be beneficial in promoting axonal regeneration.
Example 18
Wnts Pattern Synaptic Connections
[0258]Wnts not only are axon guidance molecules controlling pathfinding of axons toward their targets but also play important roles in patterning the synaptic connections once they reach their target. This process of target selection ensures the specific neuron to neuron connection and is essential to the development of the functional circuits throughout the nervous system. The inventor has found, at least in the somatosensory system and the visual system, Wnts play critical roles in patterning these synaptic target connections to establish topographic map. For example, when, in an animal model, Wnts are mis-expressed in the synaptic neuronal target area, the tectum, there is misconnection of the axons at the tectum and a resulting disrupted target map, causing the animals to be blind. Likewise, if Ryk is inhibited in a transgenic mouse in which a dominant-negative Ryk inhibitor is expressed in retinal ganglia cell neurons, similar results occur.
[0259]These studies suggest that Wnts play a role in patterning synapic connections and that the Wnt pathway can be modulated in manners discussed elsewhere in this specification to ensure specific synaptic reconnection in repair damaged neural circuits.
Example 19
Testing of Wnt, Wnt-Like Substances, and Compounds Affecting a Wnt Signaling Pathway
[0260]Based on the disclosure of the specification and the knowledge available to one of ordinary skill in the art, Wnts, Wnt-like substances, chemical compounds affecting a Wnt signaling pathway, sFRPs, sFRP-like substances, Ryk, Ryk-like substances, blockers of neuronal growth inhibitors, neuronal growth inhibitors, and/or repulsive and attractive neuronal guidance molecules can be identified. The candidate substances that have been identified can then be tested in accordance with the techniques disclosed in the specification, and evaluated for the ability to modulate neuronal growth. Testing can be conducted in vitro, such as by use of the previously disclosed explant assay, or in vivo in animal models of neuronal damage. One of ordinary skill in the art would be familiar with the numerous methods and techniques that can be employed to test candidate substances affecting a Wnt signaling pathway for ability to promote neuronal growth and regeneration.
Example 20
Clinical Trials of the Use of a Wnts, Wnt-Like Substances, and/or Chemical Compounds Affecting a Wnt Signaling Pathway in the Treatment of Diseases in General
[0261]This example is generally concerned with the development of human treatment protocols using Wnts, Wnt-like substances, chemical compounds affecting a Wnt signaling pathway, sFRPs, sFRP-like substances, Ryk, Ryk-like substances, blockers of neuronal growth inhibitors, neuronal growth inhibitors, and/or repulsive and attractive neuronal guidance molecules in the treatment of diseases such as those previously discussed in this specification. In particular, such drug treatment can be of use in the clinical treatment of various diseases in which neuronal dysfunction plays a role. Examples of these diseases include traumatic spinal cord injury. A more detailed example pertaining to traumatic spinal cord injury is discussed in the next example.
[0262]The various elements of conducting a clinical trial, including patient treatment and monitoring, will be known to those of skill in the art in light of the present disclosure. The following information can be used as a general guideline for use in establishing use of Wnts, Wnt-like substances, chemical compounds affecting a Wnt signaling pathway, sFRPs, sFRP-like substances, Ryk, Ryk-like substances, blockers of neuronal growth inhibitors, neuronal growth inhibitors, and/or repulsive and attractive neuronal guidance molecules in clinical trials. Patients with the targeted disease can be newly diagnosed patients or patients with existing disease. Patients with existing disease may include those who have failed to respond to at least one course of conventional therapy.
[0263]The Wnts, Wnt-like substances, chemical compounds affecting a Wnt signaling pathway, sFRPs, sFRP-like substances, Ryk, Ryk-like substances, blockers of neuronal growth inhibitors, neuronal growth inhibitors, and/or repulsive and attractive neuronal guidance molecules may be administered alone or in combination with the another therapeutic agent. The agents may be administered intravenously, directly into the cerebrospinal fluid, or by another mechanism that is specific to the disease that is being treated. The agent may also be administered intraoperatively, such as by direct application to the spinal cord during surgery.
[0264]The starting dose may, for example, be 0.5 mg/kg body weight. Three patients may be treated at each dose level in the absence of a defined level of toxicity. Dose escalation may be done by 100% increments (e.g., 0.5 mg, 1 mg, 2 mg, 4 mg) until drug related toxicity of a specific level develops. Thereafter dose escalation may proceed by 25% increments. The administered dose may be fractionated.
[0265]The therapeutic agent may be administered over a short infusion time or at a steady rate of infusion over a period of days. The infusion may be administered alone or in combination with other agents. The infusion given at any dose level will be dependent upon the toxicity achieved after each.
[0266]Physical examination, laboratory tests, and other clinical studies specific to the disease being treated may, of course, be performed before treatment and at intervals of about 3-4 weeks later. Laboratory studies can include CBC, differential and platelet count, urinalysis, SMA-12-100 (liver and renal function tests), coagulation profile, and any other appropriate chemistry studies to determine the extent of disease, or determine the cause of existing symptoms. If necessary, appropriate biological markers in serum can be monitored.
Example 21
Clinical Trials of the Use of a Wnt or a Wnt-Like Substance or Chemical Compounds affecting a Wnt Signaling Pathways in the Treatment of Spinal Cord Injury
[0267]This example is concerned with the development of human treatment protocols using a Wnts, Wnt-like substances, chemical compounds affecting a Wnt signaling pathway, sFRPs, sFRP-like substances, Ryk, Ryk-like substances, blockers of neuronal growth inhibitors, neuronal growth inhibitors, and/or repulsive and attractive neuronal guidance molecules in the treatment of spinal cord injury. The various elements of conducting a clinical trial, including patient treatment and monitoring, will be known to those of skill in the art in light of the present disclosure. The following information can be used as a general guideline for use in establishing clinical trials pertaining to spinal cord treatment.
[0268]Patients with spinal cord injury for clinical study will typically have failed to respond to at least one course of conventional therapy. Measurable disease is not required.
[0269]The therapeutic agent may be administered alone or in combination with the another chemotherapeutic agent. The administration may be intravenously, directly into or around the spinal cord, or in any other manner known to those of skill in the art. The starting dose may be 0.5 mg/kg body weight. Three patients may be treated at each dose level in the absence of grade >3 toxicity. Dose escalation may be done by 100% increments (0.5 mg, 1 mg, 2 mg, 4 mg) until toxicity is detected. Thereafter dose escalation may proceed by 25% increments.
[0270]The therapeutic agent may be administered over a short infusion time or at a steady rate of infusion over a 7 to 21 day period. The agent may be administered alone or in combination with agents for treatment of spinal cord injury. The infusion given at any dose level will be dependent upon the toxicity achieved after each. Increasing doses of the Wnts, Wnt-like substances, chemical compounds affecting a Wnt signaling pathway, sFRPs, sFRP-like substances, Ryk, Ryk-like substances, blockers of neuronal growth inhibitors, neuronal growth inhibitors, and/or repulsive and attractive neuronal guidance molecules, in combination with other therapeutic agents will be administered to groups of patients until approximately 60% of patients show unacceptable toxicity. Doses that are 2/3 of this value could be defined as the safe dose.
[0271]Physical examination, neurological function, and laboratory tests can, of course, be performed before treatment and at intervals of about 3-4 weeks later. Laboratory studies should include CBC, differential and platelet count, urinalysis, SMA-12-100 (liver and renal function tests), coagulation profile, and any other appropriate chemistry studies to determine the extent of disease, or determine the cause of existing symptoms. Also appropriate biological markers in serum can be monitored.
[0272]To monitor disease course and evaluate the response, it is contemplated that the patients may be examined for neurological function. Laboratory studies such as a CBC, differential and platelet count, coagulation profile, and/or SMA-12-100 shall be performed weekly. Appropriate clinical studies such as radiological studies should be performed and repeated every 8 weeks to evaluate response.
[0273]Clinical response may be defined by acceptable measure. For example, a response may be defined by improvement in neurological dysfunction, and can be graded using parameters known to those of skill in the art.
[0274]All of the compositions and methods disclosed and claimed herein can be made and executed without undue experimentation in light of the present disclosure. While the compositions and methods of this invention have been described in terms of preferred embodiments, it will be apparent to those of skill in the art that variations may be applied to the compositions and methods and in the steps or in the sequence of steps of the method described herein without departing from the concept, spirit and scope of the invention. More specifically, it will be apparent that certain agents which are both chemically and physiologically related may be substituted for the agents described herein while the same or similar results would be achieved. All such similar substitutes and modifications apparent to those skilled in the art are deemed to be within the spirit, scope and concept of the invention as defined by the appended claims.
REFERENCES
[0275]The following references, to the extent that they provide exemplary procedural or other details supplementary to those set forth herein, are specifically incorporated herein by reference. [0276]U.S. Pat. No. 4,554,101 [0277]U.S. Pat. No. 4,797,368 [0278]U.S. Pat. No. 5,139,941 [0279]Aksentijevich et al., Hum. Gene Ther., 7(9):1111-1122, 1996. [0280]Altman et al., Adv. Anat. Embryol Cell Biol., 85:1-164, 1984. [0281]Augsburger et al., Neuron, 24:127-141, 1999. [0282]Borello et al., Mech. Dev., 89:173-177, 1999. [0283]Boussif et al., Proc. Natl. Acad. Sci. USA, 92(16):7297-7301, 1995. [0284]Bradley and Brown, EMBO J., 9:1569-1575, 1990. [0285]Bueno et al., Int. J. Dev. Biol. Suppl. 1:79 S-80S, 1996. [0286]Caley et al., J. Virology, 71(4):3031-3038, 1997. [0287]Carbonelli et al., FEMS Microbiol. Lett., 177(1):75-82, 1999. [0288]Chen and Okayama, Mol. Cell Biol., 7(8):2745-2752, 1987. [0289]Christiansen et al., Mech. Dev., 51:341-350, 1995. [0290]Cocea, Biotechniques, 23(5):814-816, 1997. [0291]Coffin, In: Virology, Fields et al., eds., Raven Press, NY, 1437-1500, 1990. [0292]Couch et al., Am. Rev. Resp. Dis., 88:394-403, 1963. [0293]Davis et al., Curr. Biol., 6:146-148, 1996. [0294]Dealy et al., Mech. Dev., 43:175-186, 1993. [0295]Derossi et al., J. Biol. Chem., 269:10444-10450, 1994. [0296]Dickson, Science, 298:1959-1964, 2002. [0297]Ebens et al., Neuron, 17:1157-1172, 1996. [0298]Elliott and O'Hare, Cell, 88:223-233, 1997. [0299]Fan et al., Dev. Biol., 191:160-165, 1997. [0300]Fechheimer et al., Proc. Natl. Acad. Sci. USA, 84:8463-8467, 1987. [0301]Finch et al., Proc. Natl. Acad. Sci. USA, 94:6770-6775, 1997. [0302]FitzGerald, In: Neuroanatomy Basic and Clinical, W. B. Sauders Company LTD, 1996. [0303]Fraley et al., Proc. Natl. Acad. Sci. USA, 76:3348-3352, 1979. [0304]Francis et al., Development, 120:209-218, 1994. [0305]Fraser, In: Methods in Cell Biology, 147-160, Academic Press, Inc., 1996. [0306]Gabizon et al., Cancer Res., 50(19):6371-6378, 1990. [0307]Gavin et al., Genes Dev., 4:2319-2332, 1990. [0308]Glorioso et al., Mol. Biotechnol., 4(1):87-99, 1995. [0309]Gopal, Mol. Cell Biol., 5:1188-1190, 1985. [0310]Graham and Van Der Eb, Virology, 52:456-467, 1973. [0311]Grunhaus et al., Seminar in Virology, 200(2):535-546, 1992. [0312]Halford et al., Nat. Genet., 25:414-418, 2000. [0313]Hall et al., Cell, 100:525-535, 2000. [0314]Harland and Weintraub, J. Cell Biol., 101: 1094-1099, 1985. [0315]Hofmann et al., Dev. Genet., 19:43-50, 1996. [0316]Joosten et al., Brain Res. Dev. Brain Res., 94:99-105, 1996. [0317]Kamitori et al., Brain Res. Mol. Brain Res., 104:255-266, 2002. [0318]Keino-Masu et al., Cell, 87:175-185, 1996. [0319]Kennedy et al., Cell, 78:425-435, 1994. [0320]Kispert et al., Development, 122:3627-3637, 1996. [0321]Klein and Melton, Proc. Natl. Acad. Sci. USA, 93:8455-59, 1996. [0322]Klingensmith and Nusse, Dev. Biol., 166:396-414, 1994. [0323]Kotin et al., Proc. Natl. Acad. Sci. USA, 87(6):2211-5, 1990. [0324]Krylova et al., Neuron, 35:1043-1056, 2002. [0325]Laughlin et al., J. Virol., 60(2):515-524, 1986. [0326]Lebkowski et al., Mol. Cell. Biol., 8(10):3988-3996, 1988. [0327]Levenson et al., Hum Gene Ther, 9(8):1233-6, 1998. [0328]Liu et al., Mech. Dev., 91:409-413, 2000. [0329]Long and Young, Qjm 96:673-685, 2003. [0330]Lucas and Salinas, Dev. Bio., 192:31-44, 1997. [0331]Lyuksyutova et al., Science, 302:1984-1988, 2003. [0332]McCarty et al., J. Virol., 65(6):2936-2945, 1991. [0333]McLaughlin et al., J. Virol., 62(6):1963-1973, 1988. [0334]McMahon, Trends Genet., 8:236-242, 1992. [0335]McMahon and Bradley, Cell, 62:1073-1085, 1990. [0336]Miller, Genome Biology, 3(1):3001.1-3001.15, 2001. [0337]Morata and Lawrence, Dev. Biol., 56:227-240, 1977. [0338]Muzyczka, Curr Top Microbiol Immunol, 158:97-129, 1992. [0339]Nagahara et al., Nature Medicine, 4:1449-1452, 1998. [0340]Nicolau and Sene, Biochim. Biophys. Acta, 721:185-190, 1982. [0341]Nusse and Varmus, Cell, 69:1073-1087, 1992. [0342]Papkoff and Schryver, Mol. Cell Biol., 10:2723-2730, 1990. [0343]Parr et al., Dev. Biol., 237:324-332, 2001. [0344]Paxinos, In: The Rat Nervous System, 2nd Ed., Academic Press, 1995. [0345]Pelletier and Sonenberg, Nature, 334:320-325, 1988. [0346]Pinson et al., Nature, 407:535-538, 2000. [0347]Potter et al., Proc. Natl. Acad. Sci. USA, 81:7161-7165, 1984. [0348]Ramon y Cajal, La Cellule, 9:119-258, 1893. [0349]Rijsenijk et al., Cell, 50:649-657, 1987. [0350]Rippe et al., Mol. Cell Biol., 10:689-695, 1990. [0351]Roux et al., Proc. Nat'l Acad. Sci. USA, 86:9079-9083, 1989. [0352]Samulski et al., EMBO J., 10:3941-3950, 1991. [0353]Samulski et al., J. Virol, 63:3822-3828, 1989. [0354]Saulnier et al., Dev. Biol., 248:13-28, 2002. [0355]Serafini et al., Cell, 78:409-424, 1994. [0356]Serafini et al., Cell, 87:1001-1014, 1996. [0357]Shelling and Smith, Gene Therapy, 1: 165-169, 1994. [0358]Shirasaki et al., Science, 279:105-107, 1998. [0359]Shu et al., Development, 129:4831-4842, 2002. [0360]Sivasankaran et al., Nat Neurosci 7:261-268, 2004. [0361]Solodin et al., Biochemistry, 34(41):13537-13544, 1995. [0362]Tessier-Lavigne et al., Nature, 336:775-758, 1988. [0363]Tessier-Lavigne et al., Science, 274:1123-1133, 1996. [0364]Tessier-Lavigne, Curr. Opin. Genet. Dev., 4:596-601, 1994. [0365]Thierry et al., Proc. Natl. Acad. Sci. USA, 92(21):9742-9746, 1995. [0366]Thomas and Cappechi, Nature, 346:847-850, 1990. [0367]Top et al., J. Infect. Dis., 124:155-160, 1971. [0368]Tratschin et al., Mol. Cell Biol., 4:2072-2081, 1984. [0369]Tsukamoto et al., Nat. Genet., 9(3):243-248, 1995. [0370]Tur-Kaspa et al., Mol. Cell Biol., 6:716-718, 1986. [0371]Ungar et al., Mech. Dev., 52:153-164, 1995. [0372]Vant Veer et al., Mol. Cell Biol., 4:2532-2534, 1984. [0373]Wagner et al., Science, 260:1510-1513, 1993. [0374]Wang et al., J. Neurosci., 22:8563-8573, 2002. [0375]Wodarz et al., Annu. Rev. Cell Dev. Biol., 14:59-88, 1998. [0376]Wu and Wu, Biochemistry, 27:887-892, 1988. [0377]Wu and Wu, J. Biol. Chem., 262:4429-4432, 1987. [0378]Yang et al., Proc Natl. Acad Sci. USA, 87:9568-9572, 1990. [0379]Yoshikawa et al., Nature, 422(6932):583-588, 2003. [0380]Zakany et al., Nature, 362:546-549, 1993. [0381]Zhu et al., Science, 261(5118):209-211, 1993. [0382]Zou et al., Cell, 102:363-375, 2000. [0383]Zou, et al., Development, 124:793-804, 1997.
Sequence CWU
1
8512368DNAHomo sapiensCDS(199)..(1311) 1gcggtgccgc ccgccgtggc cgcctcagcc
caccagccgg gaccgcgagc catgctgtcc 60gccgcccgcc cccagggttg ttaaagccag
actgcgaact ctcgccactg ccgccaccgc 120cgcgtcccgt cccaccgtcg cgggcaacaa
ccaaagtcgc cgcaactgca gcacagagcg 180ggcaaagcca ggcaggcc atg ggg ctc
tgg gcg ctg ttg cct ggc tgg gtt 231 Met Gly Leu
Trp Ala Leu Leu Pro Gly Trp Val 1 5
10tct gct acg ctg ctg ctg gcg ctg gcc gct ctg ccc gca gcc
ctg gct 279Ser Ala Thr Leu Leu Leu Ala Leu Ala Ala Leu Pro Ala Ala
Leu Ala 15 20 25gcc aac agc
agt ggc cga tgg tgg ggt att gtg aac gta gcc tcc tcc 327Ala Asn Ser
Ser Gly Arg Trp Trp Gly Ile Val Asn Val Ala Ser Ser 30
35 40acg aac ctg ctt aca gac tcc aag agt ctg caa
ctg gta ctc gag ccc 375Thr Asn Leu Leu Thr Asp Ser Lys Ser Leu Gln
Leu Val Leu Glu Pro 45 50 55agt ctg
cag ctg ttg agc cgc aaa cag cgg cgt ctg ata cgc caa aat 423Ser Leu
Gln Leu Leu Ser Arg Lys Gln Arg Arg Leu Ile Arg Gln Asn60
65 70 75ccg ggg atc ctg cac agc gtg
agt ggg ggg ctg cag agt gcc gtg cgc 471Pro Gly Ile Leu His Ser Val
Ser Gly Gly Leu Gln Ser Ala Val Arg 80 85
90gag tgc aag tgg cag ttc cgg aat cgc cgc tgg aac tgt
ccc act gct 519Glu Cys Lys Trp Gln Phe Arg Asn Arg Arg Trp Asn Cys
Pro Thr Ala 95 100 105cca ggg
ccc cac ctc ttc ggc aag atc gtc aac cga ggc tgt cga gaa 567Pro Gly
Pro His Leu Phe Gly Lys Ile Val Asn Arg Gly Cys Arg Glu 110
115 120acg gcg ttt atc ttc gct atc acc tcc gcc
ggg gtc acc cat tcg gtg 615Thr Ala Phe Ile Phe Ala Ile Thr Ser Ala
Gly Val Thr His Ser Val 125 130 135gcg
cgc tcc tgc tca gaa ggt tcc atc gaa tcc tgc acg tgt gac tac 663Ala
Arg Ser Cys Ser Glu Gly Ser Ile Glu Ser Cys Thr Cys Asp Tyr140
145 150 155cgg cgg cgc ggc ccc ggg
ggc ccc gac tgg cac tgg ggg ggc tgc agc 711Arg Arg Arg Gly Pro Gly
Gly Pro Asp Trp His Trp Gly Gly Cys Ser 160
165 170gac aac att gac ttc ggc cgc ctc ttc ggc cgg gag
ttc gtg gac tcc 759Asp Asn Ile Asp Phe Gly Arg Leu Phe Gly Arg Glu
Phe Val Asp Ser 175 180 185ggg
gag aag ggg cgg gac ctg cgc ttc ctc atg aac ctt cac aac aac 807Gly
Glu Lys Gly Arg Asp Leu Arg Phe Leu Met Asn Leu His Asn Asn 190
195 200gag gca ggc cgt acg acc gta ttc tcc
gag atg cgc cag gag tgc aag 855Glu Ala Gly Arg Thr Thr Val Phe Ser
Glu Met Arg Gln Glu Cys Lys 205 210
215tgc cac ggg atg tcc ggc tca tgc acg gtg cgc acg tgc tgg atg cgg
903Cys His Gly Met Ser Gly Ser Cys Thr Val Arg Thr Cys Trp Met Arg220
225 230 235ctg ccc acg ctg
cgc gcc gtg ggc gat gtg ctg cgc gac cgc ttc gac 951Leu Pro Thr Leu
Arg Ala Val Gly Asp Val Leu Arg Asp Arg Phe Asp 240
245 250ggc gcc tcg cgc gtc ctg tac ggc aac cgc
ggc agc aac cgc gct tcg 999Gly Ala Ser Arg Val Leu Tyr Gly Asn Arg
Gly Ser Asn Arg Ala Ser 255 260
265cga gcg gag ctg ctg cgc ctg gag ccg gaa gac ccg gcc cac aaa ccg
1047Arg Ala Glu Leu Leu Arg Leu Glu Pro Glu Asp Pro Ala His Lys Pro
270 275 280ccc tcc ccc cac gac ctc gtc
tac ttc gag aaa tcg ccc aac ttc tgc 1095Pro Ser Pro His Asp Leu Val
Tyr Phe Glu Lys Ser Pro Asn Phe Cys 285 290
295acg tac agc gga cgc ctg ggc aca gca ggc acg gca ggg cgc gcc tgt
1143Thr Tyr Ser Gly Arg Leu Gly Thr Ala Gly Thr Ala Gly Arg Ala Cys300
305 310 315aac agc tcg tcg
ccc gcg ctg gac ggc tgc gag ctg ctc tgc tgc ggc 1191Asn Ser Ser Ser
Pro Ala Leu Asp Gly Cys Glu Leu Leu Cys Cys Gly 320
325 330agg ggc cac cgc acg cgc acg cag cgc gtc
acc gag cgc tgc aac tgc 1239Arg Gly His Arg Thr Arg Thr Gln Arg Val
Thr Glu Arg Cys Asn Cys 335 340
345acc ttc cac tgg tgc tgc cac gtc agc tgc cgc aac tgc acg cac acg
1287Thr Phe His Trp Cys Cys His Val Ser Cys Arg Asn Cys Thr His Thr
350 355 360cgc gta ctg cac gag tgt ctg
tga ggcgctgcgc ggactcgccc ccaggaaacg 1341Arg Val Leu His Glu Cys Leu
365 370ctctcctcga gccctccccc aaacagactc gctagcactc
aagacccggt tattcgccca 1401cccgagtacc tccagtcaca ctccccgcgg ttcatacgca
tcccatctct cccacttcct 1461cctacctggg gactcctcaa accacttgcc tggggcggca
tgaaccctct tgccatcctg 1521atggacctgc cccggaccta cctccctccc tctccgcggg
agaccccttg ttgcactgcc 1581ccctgcttgg ccaggaggtg agagaaggat gggtcccctc
cgccatgggg tcggctcctg 1641atggtgtcat tctgcctgct ccatcgcgcc agcgacctct
ctgcctctct tcttcccctt 1701tgtcctgcgt tttctccggg tcctcctaag tcccttccta
ttctcctgcc atgggtgcag 1761accctgaacc cacacctggg catcagggcc tttctcctcc
ccacctgtag ctgaagcagg 1821aggttacagg gcaaaagggc agctgtgatg atgtggaaat
gaggttgggg gaaccagcag 1881aaatgccccc attctcccag tctctgtcgt ggagccattg
aacagctgtg agccatgcct 1941ccctgggcca cctcctaccc cttcctgtcc tgcctcctca
tcagtgtgta aataatttgc 2001actgaaacgt ggatacagag ccacgagttt ggatgttgta
aataaaacta tttattgtgc 2061tgggtcccag cctggtttgc aaagaccacc tccaacccaa
cccaatccct ctccactctt 2121ctctcctttc tccctgcagc cttttctggt ccctcttctc
tcctcagttt ctcaaagatg 2181cgtttgcctc ctggaatcag tatttccttc cactgtagct
attagcggct cctcgccccc 2241accagtgtag catcttcctc tgcagaataa aatctctatt
tttatcgatg acttggtggc 2301ttttccttga atccagaaca caaccttgtt tgtggtgtcc
cctatcctcc ccttttacca 2361ctcccag
23682370PRTHomo sapiens 2Met Gly Leu Trp Ala Leu
Leu Pro Gly Trp Val Ser Ala Thr Leu Leu1 5
10 15Leu Ala Leu Ala Ala Leu Pro Ala Ala Leu Ala Ala
Asn Ser Ser Gly 20 25 30Arg
Trp Trp Gly Ile Val Asn Val Ala Ser Ser Thr Asn Leu Leu Thr 35
40 45Asp Ser Lys Ser Leu Gln Leu Val Leu
Glu Pro Ser Leu Gln Leu Leu 50 55
60Ser Arg Lys Gln Arg Arg Leu Ile Arg Gln Asn Pro Gly Ile Leu His65
70 75 80Ser Val Ser Gly Gly
Leu Gln Ser Ala Val Arg Glu Cys Lys Trp Gln 85
90 95Phe Arg Asn Arg Arg Trp Asn Cys Pro Thr Ala
Pro Gly Pro His Leu 100 105
110Phe Gly Lys Ile Val Asn Arg Gly Cys Arg Glu Thr Ala Phe Ile Phe
115 120 125Ala Ile Thr Ser Ala Gly Val
Thr His Ser Val Ala Arg Ser Cys Ser 130 135
140Glu Gly Ser Ile Glu Ser Cys Thr Cys Asp Tyr Arg Arg Arg Gly
Pro145 150 155 160Gly Gly
Pro Asp Trp His Trp Gly Gly Cys Ser Asp Asn Ile Asp Phe
165 170 175Gly Arg Leu Phe Gly Arg Glu
Phe Val Asp Ser Gly Glu Lys Gly Arg 180 185
190Asp Leu Arg Phe Leu Met Asn Leu His Asn Asn Glu Ala Gly
Arg Thr 195 200 205Thr Val Phe Ser
Glu Met Arg Gln Glu Cys Lys Cys His Gly Met Ser 210
215 220Gly Ser Cys Thr Val Arg Thr Cys Trp Met Arg Leu
Pro Thr Leu Arg225 230 235
240Ala Val Gly Asp Val Leu Arg Asp Arg Phe Asp Gly Ala Ser Arg Val
245 250 255Leu Tyr Gly Asn Arg
Gly Ser Asn Arg Ala Ser Arg Ala Glu Leu Leu 260
265 270Arg Leu Glu Pro Glu Asp Pro Ala His Lys Pro Pro
Ser Pro His Asp 275 280 285Leu Val
Tyr Phe Glu Lys Ser Pro Asn Phe Cys Thr Tyr Ser Gly Arg 290
295 300Leu Gly Thr Ala Gly Thr Ala Gly Arg Ala Cys
Asn Ser Ser Ser Pro305 310 315
320Ala Leu Asp Gly Cys Glu Leu Leu Cys Cys Gly Arg Gly His Arg Thr
325 330 335Arg Thr Gln Arg
Val Thr Glu Arg Cys Asn Cys Thr Phe His Trp Cys 340
345 350Cys His Val Ser Cys Arg Asn Cys Thr His Thr
Arg Val Leu His Glu 355 360 365Cys
Leu 37032102DNAHomo sapiensCDS(53)..(1135) 3gcatggcgcc cgcacacgga
gtctgacctg atgcagacgc aagggggtta at atg aac 58
Met Asn
1gcc cct ctc ggt gga atc tgg ctc tgg ctc cct ctg
ctc ttg acc tgg 106Ala Pro Leu Gly Gly Ile Trp Leu Trp Leu Pro Leu
Leu Leu Thr Trp 5 10 15ctc acc
ccc gag gtc aac tct tca tgg tgg tac atg aga gct aca ggt 154Leu Thr
Pro Glu Val Asn Ser Ser Trp Trp Tyr Met Arg Ala Thr Gly 20
25 30ggc tcc tcc agg gtg atg tgc gat aat gtg cca
ggc ctg gtg agc agc 202Gly Ser Ser Arg Val Met Cys Asp Asn Val Pro
Gly Leu Val Ser Ser35 40 45
50cag cgg cag ctg tgt cac cga cat cca gat gtg atg cgt gcc att agc
250Gln Arg Gln Leu Cys His Arg His Pro Asp Val Met Arg Ala Ile Ser
55 60 65cag ggc gtg gcc gag
tgg aca gca gaa tgc cag cac cag ttc cgc cag 298Gln Gly Val Ala Glu
Trp Thr Ala Glu Cys Gln His Gln Phe Arg Gln 70
75 80cac cgc tgg aat tgc aac acc ctg gac agg gat cac
agc ctt ttt ggc 346His Arg Trp Asn Cys Asn Thr Leu Asp Arg Asp His
Ser Leu Phe Gly 85 90 95agg gtc
cta ctc cga agt agt cgg gaa tct gcc ttt gtt tat gcc atc 394Arg Val
Leu Leu Arg Ser Ser Arg Glu Ser Ala Phe Val Tyr Ala Ile 100
105 110tcc tca gct gga gtt gta ttt gcc atc acc agg
gcc tgt agc caa gga 442Ser Ser Ala Gly Val Val Phe Ala Ile Thr Arg
Ala Cys Ser Gln Gly115 120 125
130gaa gta aaa tcc tgt tcc tgt gat cca aag aag atg gga agc gcc aag
490Glu Val Lys Ser Cys Ser Cys Asp Pro Lys Lys Met Gly Ser Ala Lys
135 140 145gac agc aaa ggc att
ttt gat tgg ggt ggc tgc agt gat aac att gac 538Asp Ser Lys Gly Ile
Phe Asp Trp Gly Gly Cys Ser Asp Asn Ile Asp 150
155 160tat ggg atc aaa ttt gcc cgc gca ttt gtg gat gca
aag gaa agg aaa 586Tyr Gly Ile Lys Phe Ala Arg Ala Phe Val Asp Ala
Lys Glu Arg Lys 165 170 175gga aag
gat gcc aga gcc ctg atg aat ctt cac aac aac aga gct ggc 634Gly Lys
Asp Ala Arg Ala Leu Met Asn Leu His Asn Asn Arg Ala Gly 180
185 190agg aag gct gta aag cgg ttc ttg aaa caa gag
tgc aag tgc cac ggg 682Arg Lys Ala Val Lys Arg Phe Leu Lys Gln Glu
Cys Lys Cys His Gly195 200 205
210gtg agc ggc tca tgt act ctc agg aca tgc tgg ctg gcc atg gcc gac
730Val Ser Gly Ser Cys Thr Leu Arg Thr Cys Trp Leu Ala Met Ala Asp
215 220 225ttc agg aaa acg ggc
gat tat ctc tgg agg aag tac aat ggg gcc atc 778Phe Arg Lys Thr Gly
Asp Tyr Leu Trp Arg Lys Tyr Asn Gly Ala Ile 230
235 240cag gtg gtc atg aac cag gat ggc aca ggt ttc act
gtg gct aac gag 826Gln Val Val Met Asn Gln Asp Gly Thr Gly Phe Thr
Val Ala Asn Glu 245 250 255agg ttt
aag aag cca acg aaa aat gac ctc gtg tat ttt gag aat tct 874Arg Phe
Lys Lys Pro Thr Lys Asn Asp Leu Val Tyr Phe Glu Asn Ser 260
265 270cca gac tac tgt atc agg gac cga gag gca ggc
tcc ctg ggt aca gca 922Pro Asp Tyr Cys Ile Arg Asp Arg Glu Ala Gly
Ser Leu Gly Thr Ala275 280 285
290ggc cgt gtg tgc aac ctg act tcc cgg ggc atg gac agc tgt gaa gtc
970Gly Arg Val Cys Asn Leu Thr Ser Arg Gly Met Asp Ser Cys Glu Val
295 300 305atg tgc tgt ggg aga
ggc tac gac acc tcc cat gtc acc cgg atg acc 1018Met Cys Cys Gly Arg
Gly Tyr Asp Thr Ser His Val Thr Arg Met Thr 310
315 320aag tgt ggg tgt aag ttc cac tgg tgc tgc gcc gtg
cgc tgt cag gac 1066Lys Cys Gly Cys Lys Phe His Trp Cys Cys Ala Val
Arg Cys Gln Asp 325 330 335tgc ctg
gaa gct ctg gat gtg cac aca tgc aag gcc ccc aag aac gct 1114Cys Leu
Glu Ala Leu Asp Val His Thr Cys Lys Ala Pro Lys Asn Ala 340
345 350gac tgg aca acc gct aca tga ccccagcagg
cgtcaccatc caccttccct 1165Asp Trp Thr Thr Ala Thr355
360tctacaagga ctccattgga tctgcaagaa cactggacct ttgggttctt tctgggggga
1225tatttcctaa ggcatgtggc ctttatctca acggaagccc cctcttcctc cctgggggcc
1285ccaggatggg ggggccacac gctgcaccta aagcctaccc tattctatcc atctcctggt
1345gttctgcagt catctcccct cctggcgagt tctctttgga aatagcatga caggctgttc
1405agccgggagg gtggtgggcc cagaccactg tctccaccca ccttgacgtt tcttctttct
1465agagcagttg gccaagcaga aaaaaaagtg tctcaaagga gctttctcaa tgtcttccca
1525caaatggtcc caattaagaa attccatact tctctcagat gggaacagta aagaaagcag
1585aatcaactgc ccctgactta actttaactt ttgaaaagac caagactttt gtctgatcaa
1645gtggttttac agctaccacc cttaggggta attggtaatt acctggagaa gaatggcttt
1705caataccctt ttaagtttaa aatgtgtatt tttcaaggca tttattgcca tattaaaatc
1765tgatgtaaca aggtggggac gtgtgtcctt tggtactatg gtgtgttgta tctttgtaag
1825agcaaaagcc tcagaaaggg attgctttgc attactgtcc ccttgatata aaaaatcttt
1885agggaatgag agttccttct cacttagaat ctgaagggaa ttaaaaagaa gatgaatggt
1945ctggcaatat tctgtaacta ttgggtgaat atggtggaaa ataatttagt ggatggaata
2005tcagaagtat atctgtacag atcaagaaaa aaagggagaa taaaattcct atctcatatt
2065atgcatgtga cccaaaaaaa aaaaaaaaaa aaaaaaa
21024360PRTHomo sapiens 4Met Asn Ala Pro Leu Gly Gly Ile Trp Leu Trp Leu
Pro Leu Leu Leu1 5 10
15Thr Trp Leu Thr Pro Glu Val Asn Ser Ser Trp Trp Tyr Met Arg Ala
20 25 30Thr Gly Gly Ser Ser Arg Val
Met Cys Asp Asn Val Pro Gly Leu Val 35 40
45Ser Ser Gln Arg Gln Leu Cys His Arg His Pro Asp Val Met Arg
Ala 50 55 60Ile Ser Gln Gly Val Ala
Glu Trp Thr Ala Glu Cys Gln His Gln Phe65 70
75 80Arg Gln His Arg Trp Asn Cys Asn Thr Leu Asp
Arg Asp His Ser Leu 85 90
95Phe Gly Arg Val Leu Leu Arg Ser Ser Arg Glu Ser Ala Phe Val Tyr
100 105 110Ala Ile Ser Ser Ala Gly
Val Val Phe Ala Ile Thr Arg Ala Cys Ser 115 120
125Gln Gly Glu Val Lys Ser Cys Ser Cys Asp Pro Lys Lys Met
Gly Ser 130 135 140Ala Lys Asp Ser Lys
Gly Ile Phe Asp Trp Gly Gly Cys Ser Asp Asn145 150
155 160Ile Asp Tyr Gly Ile Lys Phe Ala Arg Ala
Phe Val Asp Ala Lys Glu 165 170
175Arg Lys Gly Lys Asp Ala Arg Ala Leu Met Asn Leu His Asn Asn Arg
180 185 190Ala Gly Arg Lys Ala
Val Lys Arg Phe Leu Lys Gln Glu Cys Lys Cys 195
200 205His Gly Val Ser Gly Ser Cys Thr Leu Arg Thr Cys
Trp Leu Ala Met 210 215 220Ala Asp Phe
Arg Lys Thr Gly Asp Tyr Leu Trp Arg Lys Tyr Asn Gly225
230 235 240Ala Ile Gln Val Val Met Asn
Gln Asp Gly Thr Gly Phe Thr Val Ala 245
250 255Asn Glu Arg Phe Lys Lys Pro Thr Lys Asn Asp Leu
Val Tyr Phe Glu 260 265 270Asn
Ser Pro Asp Tyr Cys Ile Arg Asp Arg Glu Ala Gly Ser Leu Gly 275
280 285Thr Ala Gly Arg Val Cys Asn Leu Thr
Ser Arg Gly Met Asp Ser Cys 290 295
300Glu Val Met Cys Cys Gly Arg Gly Tyr Asp Thr Ser His Val Thr Arg305
310 315 320Met Thr Lys Cys
Gly Cys Lys Phe His Trp Cys Cys Ala Val Arg Cys 325
330 335Gln Asp Cys Leu Glu Ala Leu Asp Val His
Thr Cys Lys Ala Pro Lys 340 345
350Asn Ala Asp Trp Thr Thr Ala Thr 355
36051970DNAHomo sapiensCDS(20)..(1195) 5cgggagtctt cggggagct atg ctg aga
ccg ggt ggt gcg gag gaa gct gcg 52 Met Leu Arg
Pro Gly Gly Ala Glu Glu Ala Ala 1 5
10cag ctc ccg ctt cgg cgc gcc agc gcc ccg gtc cct gtg ccg
tcg ccc 100Gln Leu Pro Leu Arg Arg Ala Ser Ala Pro Val Pro Val Pro
Ser Pro 15 20 25gcg gcc ccc
gac ggc tcc cgg gct tcg gcc cgc cta ggt ctt gcc tgc 148Ala Ala Pro
Asp Gly Ser Arg Ala Ser Ala Arg Leu Gly Leu Ala Cys 30
35 40ctt ctg ctc ctg ctg ctg ctg acg ctg ccg gcc
cgc gta gac acg tcc 196Leu Leu Leu Leu Leu Leu Leu Thr Leu Pro Ala
Arg Val Asp Thr Ser 45 50 55tgg tgg
tac att ggg gca ctg ggg gca cga gtg atc tgt gac aat atc 244Trp Trp
Tyr Ile Gly Ala Leu Gly Ala Arg Val Ile Cys Asp Asn Ile60
65 70 75cct ggt ttg gtg agc cgg cag
cgg cag ctg tgc cag cgt tac cca gac 292Pro Gly Leu Val Ser Arg Gln
Arg Gln Leu Cys Gln Arg Tyr Pro Asp 80 85
90atc atg cgt tca gtg ggc gag ggt gcc cga gaa tgg atc
cga gag tgt 340Ile Met Arg Ser Val Gly Glu Gly Ala Arg Glu Trp Ile
Arg Glu Cys 95 100 105cag cac
caa ttc cgc cac cac cgc tgg aac tgt acc acc ctg gac cgg 388Gln His
Gln Phe Arg His His Arg Trp Asn Cys Thr Thr Leu Asp Arg 110
115 120gac cac acc gtc ttt ggc cgt gtc atg ctc
aga agt agc cga gag gca 436Asp His Thr Val Phe Gly Arg Val Met Leu
Arg Ser Ser Arg Glu Ala 125 130 135gct
ttt gta tat gcc atc tca tca gca ggg gta gtc cac gct att act 484Ala
Phe Val Tyr Ala Ile Ser Ser Ala Gly Val Val His Ala Ile Thr140
145 150 155cgc gcc tgt agc cag ggt
gaa ctg agt gtg tgc agc tgt gac ccc tac 532Arg Ala Cys Ser Gln Gly
Glu Leu Ser Val Cys Ser Cys Asp Pro Tyr 160
165 170acc cgt ggc cga cac cat gac cag cgt ggg gac ttt
gac tgg ggt ggc 580Thr Arg Gly Arg His His Asp Gln Arg Gly Asp Phe
Asp Trp Gly Gly 175 180 185tgc
agt gac aac atc cac tac ggt gtc cgt ttt gcc aag gcc ttc gtg 628Cys
Ser Asp Asn Ile His Tyr Gly Val Arg Phe Ala Lys Ala Phe Val 190
195 200gat gcc aag gag aag agg ctt aag gat
gcc cgg gcc ctc atg aac tta 676Asp Ala Lys Glu Lys Arg Leu Lys Asp
Ala Arg Ala Leu Met Asn Leu 205 210
215cat aat aac cgc tgt ggt cgc acg gct gtg cgg cgg ttt ctg aag ctg
724His Asn Asn Arg Cys Gly Arg Thr Ala Val Arg Arg Phe Leu Lys Leu220
225 230 235gag tgt aag tgc
cat ggc gtg agt ggt tcc tgt act ctg cgc acc tgc 772Glu Cys Lys Cys
His Gly Val Ser Gly Ser Cys Thr Leu Arg Thr Cys 240
245 250tgg cgt gca ctc tca gat ttc cgc cgc aca
ggt gat tac ctg cgg cga 820Trp Arg Ala Leu Ser Asp Phe Arg Arg Thr
Gly Asp Tyr Leu Arg Arg 255 260
265cgc tat gat ggg gct gtg cag gtg atg gcc acc caa gat ggt gcc aac
868Arg Tyr Asp Gly Ala Val Gln Val Met Ala Thr Gln Asp Gly Ala Asn
270 275 280ttc acc gca gcc cgc caa ggc
tat cgc cgt gcc acc cgg act gat ctt 916Phe Thr Ala Ala Arg Gln Gly
Tyr Arg Arg Ala Thr Arg Thr Asp Leu 285 290
295gtc tac ttt gac aac tct cca gat tac tgt gtc ttg gac aag gct gca
964Val Tyr Phe Asp Asn Ser Pro Asp Tyr Cys Val Leu Asp Lys Ala Ala300
305 310 315ggt tcc cta ggc
act gca ggc cgt gtc tgc agc aag aca tca aaa gga 1012Gly Ser Leu Gly
Thr Ala Gly Arg Val Cys Ser Lys Thr Ser Lys Gly 320
325 330aca gac ggt tgt gaa atc atg tgc tgt ggc
cga ggg tac gac aca act 1060Thr Asp Gly Cys Glu Ile Met Cys Cys Gly
Arg Gly Tyr Asp Thr Thr 335 340
345cga gtc acc cgt gtt acc cag tgt gag tgc aaa ttc cac tgg tgc tgt
1108Arg Val Thr Arg Val Thr Gln Cys Glu Cys Lys Phe His Trp Cys Cys
350 355 360gct gta cgg tgc aag gaa tgc
aga aat act gtg gac gtc cat act tgc 1156Ala Val Arg Cys Lys Glu Cys
Arg Asn Thr Val Asp Val His Thr Cys 365 370
375aaa gcc ccc aag aag gca gag tgg ctg gac cag acc tga acacacagat
1205Lys Ala Pro Lys Lys Ala Glu Trp Leu Asp Gln Thr380
385 390acctcactca tccctccaat tcaagcctct caactcaaaa
gcacaagatc cttgcatgca 1265caccttcctc caccctccac cctgggctgc taccgcttct
atttaaggat gtagagagta 1325atccataggg accatggtgt cctggctggt tccttagccc
tgggaaggag ttgtcagggg 1385atataagaaa ctgtgcaagc tccctgattt cccgctctgg
agatttgaag ggagagtaga 1445agagataggg ggtctttaga gtgaaatgag ttgcactaaa
gtacgtagtt gaggctcctt 1505ttttctttcc tttgcaccag cttcccgaca cttcttggtg
tgcaagagga agggtacctg 1565tagagagctt ctttttgttt ctacctggcc aaagttagat
gggacaaaga tgaatggcat 1625gtcccttctc tgaagtccgt ttgagcagaa ctacctggta
ccccgaaaga aaaatcttag 1685gctaccacat tctattattg agagcctgag atgttagcca
tagtggacaa ggttccattc 1745acatgctcat atgtttataa actgtgtttt gtagaagaaa
aagaatcata acaatacaaa 1805cacacattca ttctctcttt ttctctctac cattctcaac
ctgtattgga cagcactgcc 1865tcttttgctt acttgctgcc tgttcaaact gaggtggaat
gcagtggttc ccatgcttaa 1925cagatcatta aaacacccta gaacactcct aggatagatt
aatgt 19706391PRTHomo sapiens 6Met Leu Arg Pro Gly Gly
Ala Glu Glu Ala Ala Gln Leu Pro Leu Arg1 5
10 15Arg Ala Ser Ala Pro Val Pro Val Pro Ser Pro Ala
Ala Pro Asp Gly 20 25 30Ser
Arg Ala Ser Ala Arg Leu Gly Leu Ala Cys Leu Leu Leu Leu Leu 35
40 45Leu Leu Thr Leu Pro Ala Arg Val Asp
Thr Ser Trp Trp Tyr Ile Gly 50 55
60Ala Leu Gly Ala Arg Val Ile Cys Asp Asn Ile Pro Gly Leu Val Ser65
70 75 80Arg Gln Arg Gln Leu
Cys Gln Arg Tyr Pro Asp Ile Met Arg Ser Val 85
90 95Gly Glu Gly Ala Arg Glu Trp Ile Arg Glu Cys
Gln His Gln Phe Arg 100 105
110His His Arg Trp Asn Cys Thr Thr Leu Asp Arg Asp His Thr Val Phe
115 120 125Gly Arg Val Met Leu Arg Ser
Ser Arg Glu Ala Ala Phe Val Tyr Ala 130 135
140Ile Ser Ser Ala Gly Val Val His Ala Ile Thr Arg Ala Cys Ser
Gln145 150 155 160Gly Glu
Leu Ser Val Cys Ser Cys Asp Pro Tyr Thr Arg Gly Arg His
165 170 175His Asp Gln Arg Gly Asp Phe
Asp Trp Gly Gly Cys Ser Asp Asn Ile 180 185
190His Tyr Gly Val Arg Phe Ala Lys Ala Phe Val Asp Ala Lys
Glu Lys 195 200 205Arg Leu Lys Asp
Ala Arg Ala Leu Met Asn Leu His Asn Asn Arg Cys 210
215 220Gly Arg Thr Ala Val Arg Arg Phe Leu Lys Leu Glu
Cys Lys Cys His225 230 235
240Gly Val Ser Gly Ser Cys Thr Leu Arg Thr Cys Trp Arg Ala Leu Ser
245 250 255Asp Phe Arg Arg Thr
Gly Asp Tyr Leu Arg Arg Arg Tyr Asp Gly Ala 260
265 270Val Gln Val Met Ala Thr Gln Asp Gly Ala Asn Phe
Thr Ala Ala Arg 275 280 285Gln Gly
Tyr Arg Arg Ala Thr Arg Thr Asp Leu Val Tyr Phe Asp Asn 290
295 300Ser Pro Asp Tyr Cys Val Leu Asp Lys Ala Ala
Gly Ser Leu Gly Thr305 310 315
320Ala Gly Arg Val Cys Ser Lys Thr Ser Lys Gly Thr Asp Gly Cys Glu
325 330 335Ile Met Cys Cys
Gly Arg Gly Tyr Asp Thr Thr Arg Val Thr Arg Val 340
345 350Thr Gln Cys Glu Cys Lys Phe His Trp Cys Cys
Ala Val Arg Cys Lys 355 360 365Glu
Cys Arg Asn Thr Val Asp Val His Thr Cys Lys Ala Pro Lys Lys 370
375 380Ala Glu Trp Leu Asp Gln Thr385
39071506DNAHomo sapiensCDS(120)..(1187) 7gcgcttctga caagcccgaa
agtcatttcc aatctcaagt ggactttgtt ccaactattg 60ggggcgtcgc tccccctctt
catggtcgcg ggcaaacttc ctcctcggcg cctcttcta 119atg gag ccc cac ctg ctc
ggg ctg ctc ctc ggc ctc ctg ctc ggt ggc 167Met Glu Pro His Leu Leu
Gly Leu Leu Leu Gly Leu Leu Leu Gly Gly1 5
10 15acc agg gtc ctc gct ggc tac cca att tgg tgg tcc
ctg gcc ctg ggc 215Thr Arg Val Leu Ala Gly Tyr Pro Ile Trp Trp Ser
Leu Ala Leu Gly 20 25 30cag
cag tac aca tct ctg ggc tca cag ccc ctg ctc tgc ggc tcc atc 263Gln
Gln Tyr Thr Ser Leu Gly Ser Gln Pro Leu Leu Cys Gly Ser Ile 35
40 45cca ggc ctg gtc ccc aag caa ctg cgc
ttc tgc cgc aat tac atc gag 311Pro Gly Leu Val Pro Lys Gln Leu Arg
Phe Cys Arg Asn Tyr Ile Glu 50 55
60atc atg ccc agc gtg gcc gag ggc gtg aag ctg ggc atc cag gag tgc
359Ile Met Pro Ser Val Ala Glu Gly Val Lys Leu Gly Ile Gln Glu Cys65
70 75 80cag cac cag ttc cgg
ggc cgc cgc tgg aac tgc acc acc ata gat gac 407Gln His Gln Phe Arg
Gly Arg Arg Trp Asn Cys Thr Thr Ile Asp Asp 85
90 95agc ctg gcc atc ttt ggg ccc gtc ctc gac aaa
gcc acc cgc gag tcg 455Ser Leu Ala Ile Phe Gly Pro Val Leu Asp Lys
Ala Thr Arg Glu Ser 100 105
110gcc ttc gtt cac gcc atc gcc tcg gcc ggc gtg gcc ttc gcc gtc acc
503Ala Phe Val His Ala Ile Ala Ser Ala Gly Val Ala Phe Ala Val Thr
115 120 125cgc tcc tgc gcc gag ggc acc
tcc acc att tgc ggc tgt gac tcg cat 551Arg Ser Cys Ala Glu Gly Thr
Ser Thr Ile Cys Gly Cys Asp Ser His 130 135
140cat aag ggg ccg cct ggc gaa ggc tgg aag tgg ggc ggc tgc agc gag
599His Lys Gly Pro Pro Gly Glu Gly Trp Lys Trp Gly Gly Cys Ser Glu145
150 155 160gac gct gac ttc
ggc gtg tta gtg tcc agg gag ttc gcg gat gcg cgc 647Asp Ala Asp Phe
Gly Val Leu Val Ser Arg Glu Phe Ala Asp Ala Arg 165
170 175gag aac agg ccg gac gcg cgc tcg gcc atg
aac aag cac aac aac gag 695Glu Asn Arg Pro Asp Ala Arg Ser Ala Met
Asn Lys His Asn Asn Glu 180 185
190gcg ggc cgc acg act atc ctg gac cac atg cac ctc aaa tgc aag tgc
743Ala Gly Arg Thr Thr Ile Leu Asp His Met His Leu Lys Cys Lys Cys
195 200 205cac ggg ctg tcg ggc agc tgt
gag gtg aag acc tgc tgg tgg gcg cag 791His Gly Leu Ser Gly Ser Cys
Glu Val Lys Thr Cys Trp Trp Ala Gln 210 215
220cct gac ttc cgt gcc atc ggt gac ttc ctc aag gac aag tat gac agc
839Pro Asp Phe Arg Ala Ile Gly Asp Phe Leu Lys Asp Lys Tyr Asp Ser225
230 235 240gcc tcg gag atg
gta gta gag aag cac cgt gag tcc cga ggc tgg gtg 887Ala Ser Glu Met
Val Val Glu Lys His Arg Glu Ser Arg Gly Trp Val 245
250 255gag acc ctc cgg gcc aag tac tcg ctc ttc
aag cca ccc acg gag agg 935Glu Thr Leu Arg Ala Lys Tyr Ser Leu Phe
Lys Pro Pro Thr Glu Arg 260 265
270gac ctg gtc tac tac gag aac tcc ccc aac ttt tgt gag ccc aac cca
983Asp Leu Val Tyr Tyr Glu Asn Ser Pro Asn Phe Cys Glu Pro Asn Pro
275 280 285gag acg ggt tcc ttt ggc aca
agg gac cgg act tgc aat gtc acc tcc 1031Glu Thr Gly Ser Phe Gly Thr
Arg Asp Arg Thr Cys Asn Val Thr Ser 290 295
300cac ggc atc gat ggc tgc gat ctg ctc tgc tgt ggc cgg ggc cac aac
1079His Gly Ile Asp Gly Cys Asp Leu Leu Cys Cys Gly Arg Gly His Asn305
310 315 320acg agg acg gag
aag cgg aag gaa aaa tgc cac tgc atc ttc cac tgg 1127Thr Arg Thr Glu
Lys Arg Lys Glu Lys Cys His Cys Ile Phe His Trp 325
330 335tgc tgc tac gtc agc tgc cag gag tgt att
cgc atc tac gac gtg cac 1175Cys Cys Tyr Val Ser Cys Gln Glu Cys Ile
Arg Ile Tyr Asp Val His 340 345
350acc tgc aag tag ggcaccaggg cgctgggaag gggtgaagtg tgtggctggg
1227Thr Cys Lys 355cggattcagc gaagtctcat gggaagcagg acctagagcc
gggcacagcc ctcagcgtca 1287gacagcaagg aactgtcacc agccgcacgc gtggtaaatg
acccagaccc aactcgcctg 1347tggacgggga ggctctccct ctctctcatc ttacatttct
caccctactc tggatggtgt 1407gtggttttta aagaaggggg ctttcttttt agttctctag
ggtctgatag gaacagacct 1467gaggcttatc tttgcacatg ttaaagaaaa aaaaaaaaa
15068355PRTHomo sapiens 8Met Glu Pro His Leu Leu
Gly Leu Leu Leu Gly Leu Leu Leu Gly Gly1 5
10 15Thr Arg Val Leu Ala Gly Tyr Pro Ile Trp Trp Ser
Leu Ala Leu Gly 20 25 30Gln
Gln Tyr Thr Ser Leu Gly Ser Gln Pro Leu Leu Cys Gly Ser Ile 35
40 45Pro Gly Leu Val Pro Lys Gln Leu Arg
Phe Cys Arg Asn Tyr Ile Glu 50 55
60Ile Met Pro Ser Val Ala Glu Gly Val Lys Leu Gly Ile Gln Glu Cys65
70 75 80Gln His Gln Phe Arg
Gly Arg Arg Trp Asn Cys Thr Thr Ile Asp Asp 85
90 95Ser Leu Ala Ile Phe Gly Pro Val Leu Asp Lys
Ala Thr Arg Glu Ser 100 105
110Ala Phe Val His Ala Ile Ala Ser Ala Gly Val Ala Phe Ala Val Thr
115 120 125Arg Ser Cys Ala Glu Gly Thr
Ser Thr Ile Cys Gly Cys Asp Ser His 130 135
140His Lys Gly Pro Pro Gly Glu Gly Trp Lys Trp Gly Gly Cys Ser
Glu145 150 155 160Asp Ala
Asp Phe Gly Val Leu Val Ser Arg Glu Phe Ala Asp Ala Arg
165 170 175Glu Asn Arg Pro Asp Ala Arg
Ser Ala Met Asn Lys His Asn Asn Glu 180 185
190Ala Gly Arg Thr Thr Ile Leu Asp His Met His Leu Lys Cys
Lys Cys 195 200 205His Gly Leu Ser
Gly Ser Cys Glu Val Lys Thr Cys Trp Trp Ala Gln 210
215 220Pro Asp Phe Arg Ala Ile Gly Asp Phe Leu Lys Asp
Lys Tyr Asp Ser225 230 235
240Ala Ser Glu Met Val Val Glu Lys His Arg Glu Ser Arg Gly Trp Val
245 250 255Glu Thr Leu Arg Ala
Lys Tyr Ser Leu Phe Lys Pro Pro Thr Glu Arg 260
265 270Asp Leu Val Tyr Tyr Glu Asn Ser Pro Asn Phe Cys
Glu Pro Asn Pro 275 280 285Glu Thr
Gly Ser Phe Gly Thr Arg Asp Arg Thr Cys Asn Val Thr Ser 290
295 300His Gly Ile Asp Gly Cys Asp Leu Leu Cys Cys
Gly Arg Gly His Asn305 310 315
320Thr Arg Thr Glu Lys Arg Lys Glu Lys Cys His Cys Ile Phe His Trp
325 330 335Cys Cys Tyr Val
Ser Cys Gln Glu Cys Ile Arg Ile Tyr Asp Val His 340
345 350Thr Cys Lys 35592932DNAHomo
sapiensCDS(79)..(1137) 9agctcccagg gcccggcccc ccccggcgct cacgctctcg
gggcggactc ccggccctcc 60gcgccctctc gcgcggcg atg gcc cca ctc gga tac
ttc tta ctc ctc tgc 111 Met Ala Pro Leu Gly Tyr
Phe Leu Leu Leu Cys 1 5
10agc ctg aag cag gct ctg ggc agc tac ccg atc tgg tgg tcg ctg gct
159Ser Leu Lys Gln Ala Leu Gly Ser Tyr Pro Ile Trp Trp Ser Leu Ala
15 20 25gtt ggg cca cag tat tcc tcc
ctg ggc tcg cag ccc atc ctg tgt gcc 207Val Gly Pro Gln Tyr Ser Ser
Leu Gly Ser Gln Pro Ile Leu Cys Ala 30 35
40agc atc ccg ggc ctg gtc ccc aag cag ctc cgc ttc tgc agg aac
tac 255Ser Ile Pro Gly Leu Val Pro Lys Gln Leu Arg Phe Cys Arg Asn
Tyr 45 50 55gtg gag atc atg ccc agc
gtg gcc gag ggc atc aag att ggc atc cag 303Val Glu Ile Met Pro Ser
Val Ala Glu Gly Ile Lys Ile Gly Ile Gln60 65
70 75gag tgc cag cac cag ttc cgc ggc cgc cgg tgg
aac tgc acc acc gtc 351Glu Cys Gln His Gln Phe Arg Gly Arg Arg Trp
Asn Cys Thr Thr Val 80 85
90cac gac agc ctg gcc atc ttc ggg ccc gtg ctg gac aaa gct acc agg
399His Asp Ser Leu Ala Ile Phe Gly Pro Val Leu Asp Lys Ala Thr Arg
95 100 105gag tcg gcc ttt gtc cac
gcc att gcc tca gcc ggt gtg gcc ttt gca 447Glu Ser Ala Phe Val His
Ala Ile Ala Ser Ala Gly Val Ala Phe Ala 110 115
120gtg aca cgc tca tgt gca gaa ggc acg gcc gcc atc tgt ggc
tgc agc 495Val Thr Arg Ser Cys Ala Glu Gly Thr Ala Ala Ile Cys Gly
Cys Ser 125 130 135agc cgc cac cag ggc
tca cca ggc aag ggc tgg aag tgg ggt ggc tgt 543Ser Arg His Gln Gly
Ser Pro Gly Lys Gly Trp Lys Trp Gly Gly Cys140 145
150 155agc gag gac atc gag ttt ggt ggg atg gtg
tct cgg gag ttc gcc gac 591Ser Glu Asp Ile Glu Phe Gly Gly Met Val
Ser Arg Glu Phe Ala Asp 160 165
170gcc cgg gag aac cgg cca gat gcc cgc tca gcc atg aac cgc cac aac
639Ala Arg Glu Asn Arg Pro Asp Ala Arg Ser Ala Met Asn Arg His Asn
175 180 185aac gag gct ggg cgc cag
gcc atc gcc agc cac atg cac ctc aag tgc 687Asn Glu Ala Gly Arg Gln
Ala Ile Ala Ser His Met His Leu Lys Cys 190 195
200aag tgc cac ggg ctg tcg ggc agc tgc gag gtg aag aca tgc
tgg tgg 735Lys Cys His Gly Leu Ser Gly Ser Cys Glu Val Lys Thr Cys
Trp Trp 205 210 215tcg caa ccc gac ttc
cgc gcc atc ggt gac ttc ctc aag gac aag tac 783Ser Gln Pro Asp Phe
Arg Ala Ile Gly Asp Phe Leu Lys Asp Lys Tyr220 225
230 235gac agc gcc tcg gag atg gtg gtg gag aag
cac cgg gag tcc cgc ggc 831Asp Ser Ala Ser Glu Met Val Val Glu Lys
His Arg Glu Ser Arg Gly 240 245
250tgg gtg gag acc ctg cgg ccg cgc tac acc tac ttc aag gtg ccc acg
879Trp Val Glu Thr Leu Arg Pro Arg Tyr Thr Tyr Phe Lys Val Pro Thr
255 260 265gag cgc gac ctg gtc tac
tac gag gcc tcg ccc aac ttc tgc gag ccc 927Glu Arg Asp Leu Val Tyr
Tyr Glu Ala Ser Pro Asn Phe Cys Glu Pro 270 275
280aac cct gag acg ggc tcc ttc ggc acg cgc gac cgc acc tgc
aac gtc 975Asn Pro Glu Thr Gly Ser Phe Gly Thr Arg Asp Arg Thr Cys
Asn Val 285 290 295agc tcg cac ggc atc
gac ggc tgc gac ctg ctg tgc tgc ggc cgc ggc 1023Ser Ser His Gly Ile
Asp Gly Cys Asp Leu Leu Cys Cys Gly Arg Gly300 305
310 315cac aac gcg cga gcg gag cgg cgc cgg gag
aag tgc cgc tgc gtg ttc 1071His Asn Ala Arg Ala Glu Arg Arg Arg Glu
Lys Cys Arg Cys Val Phe 320 325
330cac tgg tgc tgc tac gtc agc tgc cag gag tgc acg cgc gtc tac gac
1119His Trp Cys Cys Tyr Val Ser Cys Gln Glu Cys Thr Arg Val Tyr Asp
335 340 345gtg cac acc tgc aag tag
gcaccggccg cggctccccc tggacggggc 1167Val His Thr Cys Lys
350gggccctgcc tgagggtggg cttttccctg ggtggagcag gactcccacc taaacggggc
1227agtactcctc cctgggggcg ggactcctcc ctgggggtgg ggctcctacc tgggggcaga
1287actcctacct gaaggcaggg ctcctccctg gagctagtgt ctcctctctg gtggctgggc
1347tgctcctgaa tgaggcggag ctccaggatg gggaggggct ctgcgttggc ttctccctgg
1407ggacggggct cccctggaca gaggcggggc tacagattgg gcggggcttc tcttgggtgg
1467gacagggctt ctcctgcggg ggcgaggccc ctcccagtaa gggcgtggct ctgggtgggc
1527ggggcactag gtaggcttct acctgcaggc ggggctcctc ctgaaggagg cggggctcta
1587ggatggggca cggctctggg gtaggctgct ccctgagggc ggagcgcctc cttaggagtg
1647gggttttatg gtggatgagg cttcttcctg gatggggcag agcttctcct gaccagggca
1707aggccccttc cacgggggct gtggctctgg gtgggcgtgg cctgcatagg ctccttcctg
1767tgggtggggc ttctctggga ccaggctcca atggggcggg gcttctctcc gcgggtggga
1827ctcttccctg ggaaccgccc tcctgattaa ggcgtggctt ctgcaggaat cccggctcca
1887gagcaggaaa ttcagcccac cagccacctc atccccaacc ccctgtaagg ttccatccac
1947ccctgcgtcg agctgggaag gttccatgaa gcgagtcggg tccccaaccc gtgcccctgg
2007gatccgaggg cccctctcca agcgcctggc tttggaatgc tccaggcgcg ccgacgcctg
2067tgccacccct tcctcagcct ggggtttgac cacccacctg accaggggcc ctacctgggg
2127aaagcctgaa gggcctccca gcccccaacc ccaagaccaa gcttagtcct gggagaggac
2187agggacttcg cagaggcaag cgaccgaggc cctcccaaag aggcccgccc tgcccgggct
2247cccacaccgt caggtactcc tgccagggaa ctggcctgct gcgccccagg ccccgcccgt
2307ctctgctctg ctcagctgcg cccccttctt tgcagctgcc cagcccctcc tccctgccct
2367cgggtctccc cacctgcact ccatccagct acaggagaga tagaagcctc tcgtcccgtc
2427cctccctttc ctccgcctgt ccacagcccc ttaagggaaa ggtaggaaga gaggtccagc
2487cccccaggct gcccagagct gctggtctca tttgggggcg ttcgggaggt ttggggggca
2547tcaacccccc gactgtgctg ctcgcgaagg tcccacagcc ctgagatggg ccggccccct
2607tcctggcccc tcatggcggg actggagaaa tggtccgctt tcctggagcc aatggcccgg
2667cccctcctga ctcatccgcc tggcccggga atgaatgggg aggccgctga acccacccgg
2727cccatatccc tggttgcctc atggccagcg cccctcagcc tctgccactg tgaaccggct
2787cccaccctca aggtgcgggg agaagaagcg gccaggcggg gcgccccaag agcccaaaag
2847agggcacacc gccatcctct gcctcaaatt ctgcgttttt ggttttaatg ttatatctga
2907tgctgctata tccactgtcc aacgg
293210352PRTHomo sapiens 10Met Ala Pro Leu Gly Tyr Phe Leu Leu Leu Cys
Ser Leu Lys Gln Ala1 5 10
15Leu Gly Ser Tyr Pro Ile Trp Trp Ser Leu Ala Val Gly Pro Gln Tyr
20 25 30Ser Ser Leu Gly Ser Gln Pro
Ile Leu Cys Ala Ser Ile Pro Gly Leu 35 40
45Val Pro Lys Gln Leu Arg Phe Cys Arg Asn Tyr Val Glu Ile Met
Pro 50 55 60Ser Val Ala Glu Gly Ile
Lys Ile Gly Ile Gln Glu Cys Gln His Gln65 70
75 80Phe Arg Gly Arg Arg Trp Asn Cys Thr Thr Val
His Asp Ser Leu Ala 85 90
95Ile Phe Gly Pro Val Leu Asp Lys Ala Thr Arg Glu Ser Ala Phe Val
100 105 110His Ala Ile Ala Ser Ala
Gly Val Ala Phe Ala Val Thr Arg Ser Cys 115 120
125Ala Glu Gly Thr Ala Ala Ile Cys Gly Cys Ser Ser Arg His
Gln Gly 130 135 140Ser Pro Gly Lys Gly
Trp Lys Trp Gly Gly Cys Ser Glu Asp Ile Glu145 150
155 160Phe Gly Gly Met Val Ser Arg Glu Phe Ala
Asp Ala Arg Glu Asn Arg 165 170
175Pro Asp Ala Arg Ser Ala Met Asn Arg His Asn Asn Glu Ala Gly Arg
180 185 190Gln Ala Ile Ala Ser
His Met His Leu Lys Cys Lys Cys His Gly Leu 195
200 205Ser Gly Ser Cys Glu Val Lys Thr Cys Trp Trp Ser
Gln Pro Asp Phe 210 215 220Arg Ala Ile
Gly Asp Phe Leu Lys Asp Lys Tyr Asp Ser Ala Ser Glu225
230 235 240Met Val Val Glu Lys His Arg
Glu Ser Arg Gly Trp Val Glu Thr Leu 245
250 255Arg Pro Arg Tyr Thr Tyr Phe Lys Val Pro Thr Glu
Arg Asp Leu Val 260 265 270Tyr
Tyr Glu Ala Ser Pro Asn Phe Cys Glu Pro Asn Pro Glu Thr Gly 275
280 285Ser Phe Gly Thr Arg Asp Arg Thr Cys
Asn Val Ser Ser His Gly Ile 290 295
300Asp Gly Cys Asp Leu Leu Cys Cys Gly Arg Gly His Asn Ala Arg Ala305
310 315 320Glu Arg Arg Arg
Glu Lys Cys Arg Cys Val Phe His Trp Cys Cys Tyr 325
330 335Val Ser Cys Gln Glu Cys Thr Arg Val Tyr
Asp Val His Thr Cys Lys 340 345
350111595DNAHomo sapiensCDS(196)..(1251) 11ggctctgggg cggcgctgac
agtctggtcc gcgccgggca gcgggcgcag cagcgggcag 60gctgccggca ggcacacgga
ggcagagccc cgccgcgcgc gccccggccc gcccgcgggc 120gcccacctgc agccccgacg
ggaggccccc cgcggccgca gccgctgccc cgggccgggc 180gcccgcggcg gcacc atg
agt ccc cgc tcg tgc ctg cgt tcg ctg cgc ctc 231 Met
Ser Pro Arg Ser Cys Leu Arg Ser Leu Arg Leu 1
5 10ctc gtc ttc gcc gtc ttc tca gcc gcc gcg agc aac
tgg ctg tac ctg 279Leu Val Phe Ala Val Phe Ser Ala Ala Ala Ser Asn
Trp Leu Tyr Leu 15 20 25gcc aag
ctg tcg tcg gtg ggg agc atc tca gag gag gag acg tgc gag 327Ala Lys
Leu Ser Ser Val Gly Ser Ile Ser Glu Glu Glu Thr Cys Glu 30
35 40aaa ctc aag ggc ctg atc cag agg cag gtg cag
atg tgc aag cgg aac 375Lys Leu Lys Gly Leu Ile Gln Arg Gln Val Gln
Met Cys Lys Arg Asn45 50 55
60ctg gaa gtc atg gac tcg gtg cgc cgc ggt gcc cag ctg gcc att gag
423Leu Glu Val Met Asp Ser Val Arg Arg Gly Ala Gln Leu Ala Ile Glu
65 70 75gag tgc cag tac cag
ttc cgg aac cgg cgc tgg aac tgc tcc aca ctc 471Glu Cys Gln Tyr Gln
Phe Arg Asn Arg Arg Trp Asn Cys Ser Thr Leu 80
85 90gac tcc ttg ccc gtc ttc ggc aag gtg gtg acg caa
ggg act cgg gag 519Asp Ser Leu Pro Val Phe Gly Lys Val Val Thr Gln
Gly Thr Arg Glu 95 100 105gcg gcc
ttc gtg tac gcc atc tct tcg gca ggt gtg gcc ttt gca gtg 567Ala Ala
Phe Val Tyr Ala Ile Ser Ser Ala Gly Val Ala Phe Ala Val 110
115 120acg cgg gcg tgc agc agt ggg gag ctg gag aag
tgc ggc tgt gac agg 615Thr Arg Ala Cys Ser Ser Gly Glu Leu Glu Lys
Cys Gly Cys Asp Arg125 130 135
140aca gtg cat ggg gtc agc cca cag ggc ttc cag tgg tca gga tgc tct
663Thr Val His Gly Val Ser Pro Gln Gly Phe Gln Trp Ser Gly Cys Ser
145 150 155gac aac atc gcc tac
ggt gtg gcc ttc tca cag tcg ttt gtg gat gtg 711Asp Asn Ile Ala Tyr
Gly Val Ala Phe Ser Gln Ser Phe Val Asp Val 160
165 170cgg gag aga agc aag ggg gcc tcg tcc agc aga gcc
ctc atg aac ctc 759Arg Glu Arg Ser Lys Gly Ala Ser Ser Ser Arg Ala
Leu Met Asn Leu 175 180 185cac aac
aat gag gcc ggc agg aag gcc atc ctg aca cac atg cgg gtg 807His Asn
Asn Glu Ala Gly Arg Lys Ala Ile Leu Thr His Met Arg Val 190
195 200gaa tgc aag tgc cac ggg gtg tca ggc tcc tgt
gag gta aag acg tgc 855Glu Cys Lys Cys His Gly Val Ser Gly Ser Cys
Glu Val Lys Thr Cys205 210 215
220tgg cga gcc gtg ccg ccc ttc cgc cag gtg ggt cac gca ctg aag gag
903Trp Arg Ala Val Pro Pro Phe Arg Gln Val Gly His Ala Leu Lys Glu
225 230 235aag ttt gat ggt gcc
act gag gtg gag cca cgc cgc gtg ggc tcc tcc 951Lys Phe Asp Gly Ala
Thr Glu Val Glu Pro Arg Arg Val Gly Ser Ser 240
245 250agg gca ctg gtg cca cgc aac gca cag ttc aag ccg
cac aca gat gag 999Arg Ala Leu Val Pro Arg Asn Ala Gln Phe Lys Pro
His Thr Asp Glu 255 260 265gac ctg
gtg tac ttg gag cct agc ccc gac ttc tgt gag cag gac atg 1047Asp Leu
Val Tyr Leu Glu Pro Ser Pro Asp Phe Cys Glu Gln Asp Met 270
275 280cgc agc ggc gtg ctg ggc acg agg ggc cgc aca
tgc aac aag acg tcc 1095Arg Ser Gly Val Leu Gly Thr Arg Gly Arg Thr
Cys Asn Lys Thr Ser285 290 295
300aag gcc atc gac ggc tgt gag ctg ctg tgc tgt ggc cgc ggc ttc cac
1143Lys Ala Ile Asp Gly Cys Glu Leu Leu Cys Cys Gly Arg Gly Phe His
305 310 315acg gcg cag gtg gag
ctg gct gaa cgc tgc agc tgc aaa ttc cac tgg 1191Thr Ala Gln Val Glu
Leu Ala Glu Arg Cys Ser Cys Lys Phe His Trp 320
325 330tgc tgc ttc gtc aag tgc cgg cag tgc cag cgg ctc
gtg gag ttg cac 1239Cys Cys Phe Val Lys Cys Arg Gln Cys Gln Arg Leu
Val Glu Leu His 335 340 345acg tgc
cga tga ccgcctgcct agccctgcgc cggcaaccac ctagtggccc 1291Thr Cys
Arg 350agggaaggcc gataatttaa acagtctccc accacctacc ccaagagata
ctggttgtat 1351tttttgttct ggtttggttt ttgggtcctc atgttattta ttgccgaaac
caggcaggca 1411accccaaggg caccaaccag ggcctcccca aagcctgggc ctttgtggct
gccactgacc 1471aaagggacct tgctcgtgcc gctggctgcc cgcatgtggc tgccactgac
cactcagttg 1531ttatctgtgt ccgtttttct acttgcagac ctaaggtgga gtaacaagga
gtattaccac 1591caca
159512351PRTHomo sapiens 12Met Ser Pro Arg Ser Cys Leu Arg Ser
Leu Arg Leu Leu Val Phe Ala1 5 10
15Val Phe Ser Ala Ala Ala Ser Asn Trp Leu Tyr Leu Ala Lys Leu
Ser 20 25 30Ser Val Gly Ser
Ile Ser Glu Glu Glu Thr Cys Glu Lys Leu Lys Gly 35
40 45Leu Ile Gln Arg Gln Val Gln Met Cys Lys Arg Asn
Leu Glu Val Met 50 55 60Asp Ser Val
Arg Arg Gly Ala Gln Leu Ala Ile Glu Glu Cys Gln Tyr65 70
75 80Gln Phe Arg Asn Arg Arg Trp Asn
Cys Ser Thr Leu Asp Ser Leu Pro 85 90
95Val Phe Gly Lys Val Val Thr Gln Gly Thr Arg Glu Ala Ala
Phe Val 100 105 110Tyr Ala Ile
Ser Ser Ala Gly Val Ala Phe Ala Val Thr Arg Ala Cys 115
120 125Ser Ser Gly Glu Leu Glu Lys Cys Gly Cys Asp
Arg Thr Val His Gly 130 135 140Val Ser
Pro Gln Gly Phe Gln Trp Ser Gly Cys Ser Asp Asn Ile Ala145
150 155 160Tyr Gly Val Ala Phe Ser Gln
Ser Phe Val Asp Val Arg Glu Arg Ser 165
170 175Lys Gly Ala Ser Ser Ser Arg Ala Leu Met Asn Leu
His Asn Asn Glu 180 185 190Ala
Gly Arg Lys Ala Ile Leu Thr His Met Arg Val Glu Cys Lys Cys 195
200 205His Gly Val Ser Gly Ser Cys Glu Val
Lys Thr Cys Trp Arg Ala Val 210 215
220Pro Pro Phe Arg Gln Val Gly His Ala Leu Lys Glu Lys Phe Asp Gly225
230 235 240Ala Thr Glu Val
Glu Pro Arg Arg Val Gly Ser Ser Arg Ala Leu Val 245
250 255Pro Arg Asn Ala Gln Phe Lys Pro His Thr
Asp Glu Asp Leu Val Tyr 260 265
270Leu Glu Pro Ser Pro Asp Phe Cys Glu Gln Asp Met Arg Ser Gly Val
275 280 285Leu Gly Thr Arg Gly Arg Thr
Cys Asn Lys Thr Ser Lys Ala Ile Asp 290 295
300Gly Cys Glu Leu Leu Cys Cys Gly Arg Gly Phe His Thr Ala Gln
Val305 310 315 320Glu Leu
Ala Glu Arg Cys Ser Cys Lys Phe His Trp Cys Cys Phe Val
325 330 335Lys Cys Arg Gln Cys Gln Arg
Leu Val Glu Leu His Thr Cys Arg 340 345
350135855DNAHomo sapiensCDS(319)..(1461) 13agttgcctgc gcgccctcgc
cggaccggcg gctccctagt tgcgccccga ccaggccctg 60cccttgctgc cggctcgcgc
gcgtccgcgc cccctccatt cctgggcgca tcccagctct 120gccccaactc gggagtccag
gcccgggcgc cagtgcccgc ttcagctccg gttcactgcg 180cccgccggac gcgcgccgga
ggactccgca gccctgctcc tgaccgtccc cccaggctta 240acccggtcgc tccgctcgga
ttcctcggct gcgctcgctc gggtggcgac ttcctccccg 300cgccccctcc ccctcgcc atg
aag aag tcc att gga ata tta agc cca gga 351 Met
Lys Lys Ser Ile Gly Ile Leu Ser Pro Gly 1
5 10gtt gct ttg ggg atg gct gga agt gca atg tct tcc
aag ttc ttc cta 399Val Ala Leu Gly Met Ala Gly Ser Ala Met Ser Ser
Lys Phe Phe Leu 15 20 25gtg
gct ttg gcc ata ttt ttc tcc ttc gcc cag gtt gta att gaa gcc 447Val
Ala Leu Ala Ile Phe Phe Ser Phe Ala Gln Val Val Ile Glu Ala 30
35 40aat tct tgg tgg tcg cta ggt atg aat
aac cct gtt cag atg tca gaa 495Asn Ser Trp Trp Ser Leu Gly Met Asn
Asn Pro Val Gln Met Ser Glu 45 50
55gta tat att ata gga gca cag cct ctc tgc agc caa ctg gca gga ctt
543Val Tyr Ile Ile Gly Ala Gln Pro Leu Cys Ser Gln Leu Ala Gly Leu60
65 70 75tct caa gga cag aag
aaa ctg tgc cac ttg tat cag gac cac atg cag 591Ser Gln Gly Gln Lys
Lys Leu Cys His Leu Tyr Gln Asp His Met Gln 80
85 90tac atc gga gaa ggc gcg aag aca ggc atc aaa
gaa tgc cag tat caa 639Tyr Ile Gly Glu Gly Ala Lys Thr Gly Ile Lys
Glu Cys Gln Tyr Gln 95 100
105ttc cga cat cga agg tgg aac tgc agc act gtg gat aac acc tct gtt
687Phe Arg His Arg Arg Trp Asn Cys Ser Thr Val Asp Asn Thr Ser Val
110 115 120ttt ggc agg gtg atg cag ata
ggc agc cgc gag acg gcc ttc aca tac 735Phe Gly Arg Val Met Gln Ile
Gly Ser Arg Glu Thr Ala Phe Thr Tyr 125 130
135gcg gtg agc gca gca ggg gtg gtg aac gcc atg agc cgg gcg tgc cgc
783Ala Val Ser Ala Ala Gly Val Val Asn Ala Met Ser Arg Ala Cys Arg140
145 150 155gag ggc gag ctg
tcc acc tgc ggc tgc agc cgc gcc gcg cgc ccc aag 831Glu Gly Glu Leu
Ser Thr Cys Gly Cys Ser Arg Ala Ala Arg Pro Lys 160
165 170gac ctg ccg cgg gac tgg ctc tgg ggc ggc
tgc ggc gac aac atc gac 879Asp Leu Pro Arg Asp Trp Leu Trp Gly Gly
Cys Gly Asp Asn Ile Asp 175 180
185tat ggc tac cgc ttt gcc aag gag ttc gtg gac gcc cgc gag cgg gag
927Tyr Gly Tyr Arg Phe Ala Lys Glu Phe Val Asp Ala Arg Glu Arg Glu
190 195 200cgc atc cac gcc aag ggc tcc
tac gag agt gct cgc atc ctc atg aac 975Arg Ile His Ala Lys Gly Ser
Tyr Glu Ser Ala Arg Ile Leu Met Asn 205 210
215ctg cac aac aac gag gcc ggc cgc agg acg gtg tac aac ctg gct gat
1023Leu His Asn Asn Glu Ala Gly Arg Arg Thr Val Tyr Asn Leu Ala Asp220
225 230 235gtg gcc tgc aag
tgc cat ggg gtg tcc ggc tca tgt agc ctg aag aca 1071Val Ala Cys Lys
Cys His Gly Val Ser Gly Ser Cys Ser Leu Lys Thr 240
245 250tgc tgg ctg cag ctg gca gac ttc cgc aag
gtg ggt gat gcc ctg aag 1119Cys Trp Leu Gln Leu Ala Asp Phe Arg Lys
Val Gly Asp Ala Leu Lys 255 260
265gag aag tac gac agc gcg gcg gcc atg cgg ctc aac agc cgg ggc aag
1167Glu Lys Tyr Asp Ser Ala Ala Ala Met Arg Leu Asn Ser Arg Gly Lys
270 275 280ttg gta cag gtc aac agc cgc
ttc aac tcg ccc acc aca caa gac ctg 1215Leu Val Gln Val Asn Ser Arg
Phe Asn Ser Pro Thr Thr Gln Asp Leu 285 290
295gtc tac atc gac ccc agc cct gac tac tgc gtg cgc aat gag agc acc
1263Val Tyr Ile Asp Pro Ser Pro Asp Tyr Cys Val Arg Asn Glu Ser Thr300
305 310 315ggc tcg ctg ggc
acg cag ggc cgc ctg tgc aac aag acg tcg gag ggc 1311Gly Ser Leu Gly
Thr Gln Gly Arg Leu Cys Asn Lys Thr Ser Glu Gly 320
325 330atg gat ggc tgc gag ctc atg tgc tgc ggc
cgt ggc tac gac cag ttc 1359Met Asp Gly Cys Glu Leu Met Cys Cys Gly
Arg Gly Tyr Asp Gln Phe 335 340
345aag acc gtg cag acg gag cgc tgc cac tgc aag ttc cac tgg tgc tgc
1407Lys Thr Val Gln Thr Glu Arg Cys His Cys Lys Phe His Trp Cys Cys
350 355 360tac gtc aag tgc aag aag tgc
acg gag atc gtg gac cag ttt gtg tgc 1455Tyr Val Lys Cys Lys Lys Cys
Thr Glu Ile Val Asp Gln Phe Val Cys 365 370
375aag tag tgggtgccac ccagcactca gccccgctcc caggacccgc ttatttatag
1511Lys380aaagtacagt gattctggtt tttggttttt agaaatattt tttatttttc
cccaagaatt 1571gcaaccggaa ccattttttt tcctgttacc atctaagaac tctgtggttt
attattaata 1631ttataattat tatttggcaa taatgggggt gggaaccaag aaaaatattt
attttgtgga 1691tctttgaaaa ggtaatacaa gacttctttt gatagtatag aatgaagggg
aaataacaca 1751taccctaact tagctgtgtg gacatggtac acatccagaa ggtaaagaaa
tacattttct 1811ttttctcaaa tatgccatca tatgggatgg gtaggttcca gttgaaagag
ggtggtagaa 1871atctattcac aattcagctt ctatgaccaa aatgagttgt aaattctctg
gtgcaagata 1931aaaggtcttg ggaaaacaaa acaaaacaaa acaaacctcc cttccccagc
agggctgcta 1991gcttgctttc tgcattttca aaatgataat ttacaatgga aggacaagaa
tgtcatattc 2051tcaaggaaaa aaggtatatc acatgtctca ttctcctcaa atattccatt
tgcagacaga 2111ccgtcatatt ctaatagctc atgaaatttg ggcagcaggg aggaaagtcc
ccagaaatta 2171aaaaatttaa aactcttatg tcaagatgtt gatttgaagc tgttataaga
attaggattc 2231cagattgtaa aaagatcccc aaatgattct ggacactaga tttttttgtt
tggggaggtt 2291ggcttgaaca taaatgaaaa tatcctgtta ttttcttagg gatacttggt
tagtaaatta 2351taatagtaaa aataatacat gaatcccatt cacaggttct cagcccaagc
aacaaggtaa 2411ttgcgtgcca ttcagcactg caccagagca gacaacctat ttgaggaaaa
acagtgaaat 2471ccaccttcct cttcacactg agccctctct gattcctccg tgttgtgatg
tgatgctggc 2531cacgtttcca aacggcagct ccactgggtc ccctttggtt gtaggacagg
aaatgaaaca 2591ttaggagctc tgcttggaaa acagttcact acttagggat ttttgtttcc
taaaactttt 2651attttgagga gcagtagttt tctatgtttt aatgacagaa cttggctaat
ggaattcaca 2711gaggtgttgc agcgtatcac tgttatgatc ctgtgtttag attatccact
catgcttctc 2771ctattgtact gcaggtgtac cttaaaactg ttcccagtgt acttgaacag
ttgcatttat 2831aaggggggaa atgtggttta atggtgcctg atatctcaaa gtcttttgta
cataacatat 2891atatatatat acatatatat aaatataaat ataaatatat ctcattgcag
ccagtgattt 2951agatttacag tttactctgg ggttatttct ctgtctagag cattgttgtc
cttcactgca 3011gtccagttgg gattattcca aaagtttttt gagtcttgag cttgggctgt
ggccctgctg 3071tgatcatacc ttgagcacga cgaagcaacc ttgtttctga ggaagcttga
gttctgactc 3131actgaaatgc gtgttgggtt gaagatatct tttttctttt ctgcctcacc
cctttgtctc 3191caacctccat ttctgttcac tttgtggaga gggcattact tgttcgttat
agacatggac 3251gttaagagat attcaaaact cagaagcatc agcaatgttt ctcttttctt
agttcattct 3311gcagaatgga aacccatgcc tattagaaat gacagtactt attaattgag
tccctaagga 3371atattcagcc cactacatag atagcttttt tttttttttt tttaataagg
acacctcttt 3431ccaaacagtg ccatcaaata tgttcttatc tcagacttac gttgttttaa
aagtttggaa 3491agatacacat ctttcatacc ccccttaggc aggttggctt tcatatcacc
tcagccaact 3551gtggctctta atttattgca taatgatatt cacatcccct cagttgcagt
gaattgtgag 3611caaaagatct tgaaagcaaa aagcactaat tagtttaaaa tgtcactttt
ttggttttta 3671ttatacaaaa accatgaagt acttttttta tttgctaaat cagattgttc
ctttttagtg 3731actcatgttt atgaagagag ttgagtttaa caatcctagc ttttaaaaga
aactatttaa 3791tgtaaaatat tctacatgtc attcagatat tatgtatatc ttctagcctt
tattctgtac 3851ttttaatgta catatttctg tcttgcgtga tttgtatatt tcactggttt
aaaaaacaaa 3911catcgaaagg cttatgccaa atggaagata gaatataaaa taaaacgtta
cttgtatatt 3971ggtaagtggt ttcaattgtc cttcagataa ttcatgtgga gatttttgga
gaaaccatga 4031cggatagttt aggatgacta catgtcaaag taataaaaga gtggtgaatt
ttaccaaaac 4091caagctattt ggaagcttca aaaggtttct atatgtaatg gaacaaaagg
ggaattctct 4151tttcctatat atgttcctta caaaaaaaaa aaaaaaagaa atcaagcaga
tggcttaaag 4211ctggttatag gattgctcac attcttttag cattatgcat gtaacttaat
tgttttagag 4271cgtgttgctg ttgtaacatc ccagagaaga atgaaaaggc acatgctttt
atccgtgacc 4331agatttttag tccaaaaaaa tgtatttttt tgtgtgttta ccactgcaac
tattgcacct 4391ctctatttga atttactgtg gaccatgtgt ggtgtctcta tgccctttga
aagcagtttt 4451tataaaaaga aagcccgggt ctgcagagaa tgaaaactgg ttggaaacta
aaggttcatt 4511gtgttaagtg caattaatac aagttattgt gcttttcaaa aatgtacacg
gaaatctgga 4571cagtgctgca cagattgata cattagcctt tgctttttct ctttccggat
aaccttgtaa 4631catattgaaa ccttttaagg atgccaagaa tgcattattc cacaaaaaaa
cagcagacca 4691acatatagag tgtttaaaat agcatttctg ggcaaattca aactcttgtg
gttctaggac 4751tcacatctgt ttcagttttt cctcagttgt atattgacca gtgttcttta
ttgcaaaaac 4811atatacccga tttagcagtg tcagcgtatt ttttcttctc atcctggagc
gtattcaaga 4871tcttcccaat acaagaaaat taataaaaaa tttatatata ggcagcagca
aaagagccat 4931gttcaaaata gtcattatgg gctcaaatag aaagaagact tttaagtttt
aatccagttt 4991atctgttgag ttctgtgagc tactgacctc ctgagactgg cactgtgtaa
gttttagttg 5051cctaccctag ctcttttctc gtacaatttt gccaatacca agtttcaatt
tgtttttaca 5111aaacattatt caagccacta gaattatcaa atatgacgct atagcagagt
aaatactctg 5171aataagagac cggtactagc taactccaag agatcgttag cagcatcagt
ccacaaacac 5231ttagtggccc acaatatata gagagataga aaaggtagtt ataacttgaa
gcatgtattt 5291aatgcaaata ggcacgaagg cacaggtcta aaatactaca ttgtcactgt
aagctatact 5351tttaaaatat ttattttttt taaagtattt tctagtcttt tctctctctg
tggaatggtg 5411aaagagagat gccgtgtttt gaaagtaaga tgatgaaatg aatttttaat
tcaagaaaca 5471ttcagaaaca taggaattaa aacttagaga aatgatctaa tttccctgtt
cacacaaact 5531ttacacttta atctgatgat tggatatttt attttagtga aacatcatct
tgttagctaa 5591ctttaaaaaa tggatgtaga atgattaaag gttggtatga ttttttttta
atgtatcagt 5651ttgaacctag aatattgaat taaaatgctg tctcagtatt ttaaaagcaa
aaaaggaatg 5711gaggaaaatt gcatcttaga ccatttttat atgcagtgta caatttgctg
ggctagaaat 5771gagataaaga ttatttattt ttgttcatat cttgtacttt tctattaaaa
tcattttatg 5831aaatccaaaa aaaaaaaaaa aaaa
585514380PRTHomo sapiens 14Met Lys Lys Ser Ile Gly Ile Leu Ser
Pro Gly Val Ala Leu Gly Met1 5 10
15Ala Gly Ser Ala Met Ser Ser Lys Phe Phe Leu Val Ala Leu Ala
Ile 20 25 30Phe Phe Ser Phe
Ala Gln Val Val Ile Glu Ala Asn Ser Trp Trp Ser 35
40 45Leu Gly Met Asn Asn Pro Val Gln Met Ser Glu Val
Tyr Ile Ile Gly 50 55 60Ala Gln Pro
Leu Cys Ser Gln Leu Ala Gly Leu Ser Gln Gly Gln Lys65 70
75 80Lys Leu Cys His Leu Tyr Gln Asp
His Met Gln Tyr Ile Gly Glu Gly 85 90
95Ala Lys Thr Gly Ile Lys Glu Cys Gln Tyr Gln Phe Arg His
Arg Arg 100 105 110Trp Asn Cys
Ser Thr Val Asp Asn Thr Ser Val Phe Gly Arg Val Met 115
120 125Gln Ile Gly Ser Arg Glu Thr Ala Phe Thr Tyr
Ala Val Ser Ala Ala 130 135 140Gly Val
Val Asn Ala Met Ser Arg Ala Cys Arg Glu Gly Glu Leu Ser145
150 155 160Thr Cys Gly Cys Ser Arg Ala
Ala Arg Pro Lys Asp Leu Pro Arg Asp 165
170 175Trp Leu Trp Gly Gly Cys Gly Asp Asn Ile Asp Tyr
Gly Tyr Arg Phe 180 185 190Ala
Lys Glu Phe Val Asp Ala Arg Glu Arg Glu Arg Ile His Ala Lys 195
200 205Gly Ser Tyr Glu Ser Ala Arg Ile Leu
Met Asn Leu His Asn Asn Glu 210 215
220Ala Gly Arg Arg Thr Val Tyr Asn Leu Ala Asp Val Ala Cys Lys Cys225
230 235 240His Gly Val Ser
Gly Ser Cys Ser Leu Lys Thr Cys Trp Leu Gln Leu 245
250 255Ala Asp Phe Arg Lys Val Gly Asp Ala Leu
Lys Glu Lys Tyr Asp Ser 260 265
270Ala Ala Ala Met Arg Leu Asn Ser Arg Gly Lys Leu Val Gln Val Asn
275 280 285Ser Arg Phe Asn Ser Pro Thr
Thr Gln Asp Leu Val Tyr Ile Asp Pro 290 295
300Ser Pro Asp Tyr Cys Val Arg Asn Glu Ser Thr Gly Ser Leu Gly
Thr305 310 315 320Gln Gly
Arg Leu Cys Asn Lys Thr Ser Glu Gly Met Asp Gly Cys Glu
325 330 335Leu Met Cys Cys Gly Arg Gly
Tyr Asp Gln Phe Lys Thr Val Gln Thr 340 345
350Glu Arg Cys His Cys Lys Phe His Trp Cys Cys Tyr Val Lys
Cys Lys 355 360 365Lys Cys Thr Glu
Ile Val Asp Gln Phe Val Cys Lys 370 375
380152252DNAHomo sapiensCDS(184)..(1263) 15gaccattagc aggcacccag
gcctgtcttt ggctcggaaa cggtggcccc caatgtagcc 60tagtttgaac ctaggaactg
caggaccaga gagattccac tggagcctga tggacgggtg 120acagagggaa ccctactctg
gaaactgtca gtcccagggc actggggagg gctgaggccg 180acc atg ccc agc ctg ctg
ctg ctg ttc acg gct gct ctg ctg tcc agc 228 Met Pro Ser Leu Leu
Leu Leu Phe Thr Ala Ala Leu Leu Ser Ser 1 5
10 15tgg gct cag ctt ctg aca gac gcc aac tcc tgg
tgg tca tta gct ttg 276Trp Ala Gln Leu Leu Thr Asp Ala Asn Ser Trp
Trp Ser Leu Ala Leu 20 25
30aac ccg gtg cag aga ccc gag atg ttt atc atc ggt gcc cag ccc gtg
324Asn Pro Val Gln Arg Pro Glu Met Phe Ile Ile Gly Ala Gln Pro Val
35 40 45tgc agt cag ctt ccc ggg ctc
tcc cct ggc cag agg aag ctg tgc caa 372Cys Ser Gln Leu Pro Gly Leu
Ser Pro Gly Gln Arg Lys Leu Cys Gln 50 55
60ttg tac cag gag cac atg gcc tac ata ggg gag gga gcc aag act
ggc 420Leu Tyr Gln Glu His Met Ala Tyr Ile Gly Glu Gly Ala Lys Thr
Gly 65 70 75atc aag gaa tgc cag cac
cag ttc cgg cag cgg cgg tgg aat tgc agc 468Ile Lys Glu Cys Gln His
Gln Phe Arg Gln Arg Arg Trp Asn Cys Ser80 85
90 95aca gcg gac aac gca tct gtc ttt ggg aga gtc
atg cag ata ggc agc 516Thr Ala Asp Asn Ala Ser Val Phe Gly Arg Val
Met Gln Ile Gly Ser 100 105
110cga gag acc gcc ttc acc cac gcg gtg agc gcc gcg ggc gtg gtc aac
564Arg Glu Thr Ala Phe Thr His Ala Val Ser Ala Ala Gly Val Val Asn
115 120 125gcc atc agc cgg gcc tgc
cgc gag ggc gag ctc tcc acc tgc ggc tgc 612Ala Ile Ser Arg Ala Cys
Arg Glu Gly Glu Leu Ser Thr Cys Gly Cys 130 135
140agc cgg acg gcg cgg ccc aag gac ctg ccc cgg gac tgg ctg
tgg ggc 660Ser Arg Thr Ala Arg Pro Lys Asp Leu Pro Arg Asp Trp Leu
Trp Gly 145 150 155ggc tgt ggg gac aac
gtg gag tac ggc tac cgc ttc gcc aag gag ttt 708Gly Cys Gly Asp Asn
Val Glu Tyr Gly Tyr Arg Phe Ala Lys Glu Phe160 165
170 175gtg gat gcc cgg gag cga gag aag aac ttt
gcc aaa gga tca gag gag 756Val Asp Ala Arg Glu Arg Glu Lys Asn Phe
Ala Lys Gly Ser Glu Glu 180 185
190cag ggc cgg gtg ctc atg aac ctg caa aac aac gag gcc ggt cgc agg
804Gln Gly Arg Val Leu Met Asn Leu Gln Asn Asn Glu Ala Gly Arg Arg
195 200 205gct gtg tat aag atg gca
gac gta gcc tgc aaa tgc cac ggc gtc tcg 852Ala Val Tyr Lys Met Ala
Asp Val Ala Cys Lys Cys His Gly Val Ser 210 215
220ggg tcc tgc agc ctc aag acc tgc tgg ctg cag ctg gcc gag
ttc cgc 900Gly Ser Cys Ser Leu Lys Thr Cys Trp Leu Gln Leu Ala Glu
Phe Arg 225 230 235aag gtc ggg gac cgg
ctg aag gag aag tac gac agc gcg gcc gcc atg 948Lys Val Gly Asp Arg
Leu Lys Glu Lys Tyr Asp Ser Ala Ala Ala Met240 245
250 255cgc gtc acc cgc aag ggc cgg ctg gag ctg
gtc aac agc cgc ttc acc 996Arg Val Thr Arg Lys Gly Arg Leu Glu Leu
Val Asn Ser Arg Phe Thr 260 265
270cag ccc acc ccg gag gac ctg gtc tat gtg gac ccc agc ccc gac tac
1044Gln Pro Thr Pro Glu Asp Leu Val Tyr Val Asp Pro Ser Pro Asp Tyr
275 280 285tgc ctg cgc aac gag agc
acg ggc tcc ctg ggc acg cag ggc cgc ctc 1092Cys Leu Arg Asn Glu Ser
Thr Gly Ser Leu Gly Thr Gln Gly Arg Leu 290 295
300tgc aac aag acc tcg gag ggc atg gat ggc tgt gag ctc atg
tgc tgc 1140Cys Asn Lys Thr Ser Glu Gly Met Asp Gly Cys Glu Leu Met
Cys Cys 305 310 315ggg cgt ggc tac aac
cag ttc aag agc gtg cag gtg gag cgc tgc cac 1188Gly Arg Gly Tyr Asn
Gln Phe Lys Ser Val Gln Val Glu Arg Cys His320 325
330 335tgc aag ttc cac tgg tgc tgc ttc gtc agg
tgt aag aag tgc acg gag 1236Cys Lys Phe His Trp Cys Cys Phe Val Arg
Cys Lys Lys Cys Thr Glu 340 345
350atc gtg gac cag tac atc tgt aaa tag cccggagggc ctgctcccgg
1283Ile Val Asp Gln Tyr Ile Cys Lys 355
360ccccccctgc actctgcctc acaaaggtct atattatata aatctatata aatctatttt
1343atatttgtat aagtaaatgg gtgggtgcta tacaatggaa agatgaaaat ggaaaggaag
1403agcttattta agagacgctg gagatctctg aggagtggac tttgctggtt ctctcctctt
1463ggtgggtggg agacagggct ttttctctcc ctctggcgag gactctcagg atgtagggac
1523ttggaaatat ttactgtctg tccaccacgg cctggaggag ggaggttgtg gttggatgga
1583ggagatgatc ttgtctggaa gtctagagtc tttgttggtt agaggactgc ctgtgatcct
1643ggccactagg ccaagaggcc ctatgaaggt ggcgggaact cagcttcaac ctcgatgtct
1703tcagggtctt gtccagaatg tagatgggtt ccgtaagagg cctggtgctc tcttactctt
1763tcatccacgt gcacttgtgc ggcatctgca gtttacagga acggctcctt ccctaaaatg
1823agaagtccaa ggtcatctct ggcccagtga ccacagagag atctgcacct cccggacttc
1883aggcctgcct ttccagcgag aattcttcat cctccacggt tcactagctc ctacctgaag
1943aggaaagggg gccatttgac ctgacatgtc aggaaagccc taaactgaat gtttgcgcct
2003gggctgcaga agccagggtg catgaccagg ctgcgtggac gttatactgt cttcccccac
2063ccccggggag gggaagcttg agctgctgct gtcactcctc caccgaggga ggcctcacaa
2123accacaggac gctgcaacgg gtcaggctgg cgggcccggc gtgctcatca tctctgcccc
2183aggtgtacgg tttctctctg acattaaatg cccttcatgg aaaaaaaaaa aagaaaaaaa
2243aaaaaaaaa
225216359PRTHomo sapiens 16Met Pro Ser Leu Leu Leu Leu Phe Thr Ala Ala
Leu Leu Ser Ser Trp1 5 10
15Ala Gln Leu Leu Thr Asp Ala Asn Ser Trp Trp Ser Leu Ala Leu Asn
20 25 30Pro Val Gln Arg Pro Glu Met
Phe Ile Ile Gly Ala Gln Pro Val Cys 35 40
45Ser Gln Leu Pro Gly Leu Ser Pro Gly Gln Arg Lys Leu Cys Gln
Leu 50 55 60Tyr Gln Glu His Met Ala
Tyr Ile Gly Glu Gly Ala Lys Thr Gly Ile65 70
75 80Lys Glu Cys Gln His Gln Phe Arg Gln Arg Arg
Trp Asn Cys Ser Thr 85 90
95Ala Asp Asn Ala Ser Val Phe Gly Arg Val Met Gln Ile Gly Ser Arg
100 105 110Glu Thr Ala Phe Thr His
Ala Val Ser Ala Ala Gly Val Val Asn Ala 115 120
125Ile Ser Arg Ala Cys Arg Glu Gly Glu Leu Ser Thr Cys Gly
Cys Ser 130 135 140Arg Thr Ala Arg Pro
Lys Asp Leu Pro Arg Asp Trp Leu Trp Gly Gly145 150
155 160Cys Gly Asp Asn Val Glu Tyr Gly Tyr Arg
Phe Ala Lys Glu Phe Val 165 170
175Asp Ala Arg Glu Arg Glu Lys Asn Phe Ala Lys Gly Ser Glu Glu Gln
180 185 190Gly Arg Val Leu Met
Asn Leu Gln Asn Asn Glu Ala Gly Arg Arg Ala 195
200 205Val Tyr Lys Met Ala Asp Val Ala Cys Lys Cys His
Gly Val Ser Gly 210 215 220Ser Cys Ser
Leu Lys Thr Cys Trp Leu Gln Leu Ala Glu Phe Arg Lys225
230 235 240Val Gly Asp Arg Leu Lys Glu
Lys Tyr Asp Ser Ala Ala Ala Met Arg 245
250 255Val Thr Arg Lys Gly Arg Leu Glu Leu Val Asn Ser
Arg Phe Thr Gln 260 265 270Pro
Thr Pro Glu Asp Leu Val Tyr Val Asp Pro Ser Pro Asp Tyr Cys 275
280 285Leu Arg Asn Glu Ser Thr Gly Ser Leu
Gly Thr Gln Gly Arg Leu Cys 290 295
300Asn Lys Thr Ser Glu Gly Met Asp Gly Cys Glu Leu Met Cys Cys Gly305
310 315 320Arg Gly Tyr Asn
Gln Phe Lys Ser Val Gln Val Glu Arg Cys His Cys 325
330 335Lys Phe His Trp Cys Cys Phe Val Arg Cys
Lys Lys Cys Thr Glu Ile 340 345
350Val Asp Gln Tyr Ile Cys Lys 355171726DNAHomo
sapiensCDS(224)..(1321) 17ggcacgagcg caggagacac aggcgctggc tgccccgtcc
gctctccgcc tccgccgcgc 60cctcctcgcc cgggatgggc ccccccgccg ccgccggatc
cctcgcctcc cggccgccgc 120cgttgcgctc gccgcgctcg cactgaagcc cgggccctcg
cgcgccgcgg ttcgccccgc 180agcctcgccc cctgcccacc cgggcggccg tagggcggtc
acg atg ctg ccg ccc 235
Met Leu Pro Pro 1tta ccc
tcc cgc ctc ggg ctg ctg ctg ctg ctg ctc ctg tgc ccg gcg 283Leu Pro
Ser Arg Leu Gly Leu Leu Leu Leu Leu Leu Leu Cys Pro Ala5
10 15 20cac gtc ggc gga ctg tgg tgg
gct gtg ggc agc ccc ttg gtt atg gac 331His Val Gly Gly Leu Trp Trp
Ala Val Gly Ser Pro Leu Val Met Asp 25 30
35cct acc agc atc tgc agg aag gca cgg cgg ctg gcc ggg
cgg cag gcc 379Pro Thr Ser Ile Cys Arg Lys Ala Arg Arg Leu Ala Gly
Arg Gln Ala 40 45 50gag ttg
tgc cag gct gag ccg gaa gtg gtg gca gag cta gct cgg ggc 427Glu Leu
Cys Gln Ala Glu Pro Glu Val Val Ala Glu Leu Ala Arg Gly 55
60 65gcc cgg ctc ggg gtg cga gag tgc cag ttc
cag ttc cgc ttc cgc cgc 475Ala Arg Leu Gly Val Arg Glu Cys Gln Phe
Gln Phe Arg Phe Arg Arg 70 75 80tgg
aat tgc tcc agc cac agc aag gcc ttt gga cgc atc ctg caa cag 523Trp
Asn Cys Ser Ser His Ser Lys Ala Phe Gly Arg Ile Leu Gln Gln85
90 95 100gac att cgg gag acg gcc
ttc gtg ttc gcc atc act gcg gcc ggc gcc 571Asp Ile Arg Glu Thr Ala
Phe Val Phe Ala Ile Thr Ala Ala Gly Ala 105
110 115agc cac gcc gtc acg cag gcc tgt tct atg ggc gag
ctg ctg cag tgc 619Ser His Ala Val Thr Gln Ala Cys Ser Met Gly Glu
Leu Leu Gln Cys 120 125 130ggc
tgc cag gcg ccc cgc ggg cgg gcc cct ccc cgg ccc tcc ggc ctg 667Gly
Cys Gln Ala Pro Arg Gly Arg Ala Pro Pro Arg Pro Ser Gly Leu 135
140 145ccc ggc acc ccc gga ccc cct ggc ccc
gcg ggc tcc ccg gaa ggc agc 715Pro Gly Thr Pro Gly Pro Pro Gly Pro
Ala Gly Ser Pro Glu Gly Ser 150 155
160gcc gcc tgg gag tgg gga ggc tgc ggc gac gac gtg gac ttc ggg gac
763Ala Ala Trp Glu Trp Gly Gly Cys Gly Asp Asp Val Asp Phe Gly Asp165
170 175 180gag aag tcg agg
ctc ttt atg gac gcg cgg cac aag cgg gga cgc gga 811Glu Lys Ser Arg
Leu Phe Met Asp Ala Arg His Lys Arg Gly Arg Gly 185
190 195gac atc cgc gcg ttg gtg caa ctg cac aac
aac gag gcg ggc agg ctg 859Asp Ile Arg Ala Leu Val Gln Leu His Asn
Asn Glu Ala Gly Arg Leu 200 205
210gcc gtg cgg agc cac acg cgc acc gag tgc aaa tgc cac ggg ctg tcg
907Ala Val Arg Ser His Thr Arg Thr Glu Cys Lys Cys His Gly Leu Ser
215 220 225gga tca tgc gcg ctg cgc acc
tgc tgg cag aag ctg cct cca ttt cgc 955Gly Ser Cys Ala Leu Arg Thr
Cys Trp Gln Lys Leu Pro Pro Phe Arg 230 235
240gag gtg ggc gcg cgg ctg ctg gag cgc ttc cac ggc gcc tca cgc gtc
1003Glu Val Gly Ala Arg Leu Leu Glu Arg Phe His Gly Ala Ser Arg Val245
250 255 260atg ggc acc aac
gac ggc aag gcc ctg ctg ccc gcc gtc cgc acg ctc 1051Met Gly Thr Asn
Asp Gly Lys Ala Leu Leu Pro Ala Val Arg Thr Leu 265
270 275aag ccg ccg ggc cga gcg gac ctc ctc tac
gcc gcc gat tcg ccc gac 1099Lys Pro Pro Gly Arg Ala Asp Leu Leu Tyr
Ala Ala Asp Ser Pro Asp 280 285
290ttt tgc gcc ccc aac cga cgc acc ggc tcc ccc ggc acg cgc ggt cgc
1147Phe Cys Ala Pro Asn Arg Arg Thr Gly Ser Pro Gly Thr Arg Gly Arg
295 300 305gcc tgc aat agc agc gcc ccg
gac ctc agc ggc tgc gac ctg ctg tgc 1195Ala Cys Asn Ser Ser Ala Pro
Asp Leu Ser Gly Cys Asp Leu Leu Cys 310 315
320tgc ggc cgc ggg cac cgc cag gag agc gtg cag ctc gaa gag aac tgc
1243Cys Gly Arg Gly His Arg Gln Glu Ser Val Gln Leu Glu Glu Asn Cys325
330 335 340ctg tgc cgc ttc
cac tgg tgc tgc gta gta cag tgc cac cgt tgc cgt 1291Leu Cys Arg Phe
His Trp Cys Cys Val Val Gln Cys His Arg Cys Arg 345
350 355gtg cgc aag gag ctc agc ctc tgc ctg tga
cccgccgccc ggccgctaga 1341Val Arg Lys Glu Leu Ser Leu Cys Leu
360 365ctgacttcgc gcagcggtgg ctcgcacctg tgggacctca
gggcaccggc accgggcgcc 1401tctcgccgct cgagcccagc ctctccctgc caaagcccaa
ctcccagggc tctggaaatg 1461gtgaggcgag gggcttgaga ggaacgccca cccacgaagg
cccagggcgc cagacggccc 1521cgaaaaggcg ctcggggagc gtttaaagga cactgtacag
gccctccctc cccttggcct 1581ctaggaggaa acagtttttt agactggaaa aaagccagtc
taaaggcctc tggatactgg 1641gctccccaga actgctggcc acaggatggt gggtgaggtt
agtatcaata aagatattta 1701aaccaaaaaa aaaaaaaaaa aaaaa
172618365PRTHomo sapiens 18Met Leu Pro Pro Leu Pro
Ser Arg Leu Gly Leu Leu Leu Leu Leu Leu1 5
10 15Leu Cys Pro Ala His Val Gly Gly Leu Trp Trp Ala
Val Gly Ser Pro 20 25 30Leu
Val Met Asp Pro Thr Ser Ile Cys Arg Lys Ala Arg Arg Leu Ala 35
40 45Gly Arg Gln Ala Glu Leu Cys Gln Ala
Glu Pro Glu Val Val Ala Glu 50 55
60Leu Ala Arg Gly Ala Arg Leu Gly Val Arg Glu Cys Gln Phe Gln Phe65
70 75 80Arg Phe Arg Arg Trp
Asn Cys Ser Ser His Ser Lys Ala Phe Gly Arg 85
90 95Ile Leu Gln Gln Asp Ile Arg Glu Thr Ala Phe
Val Phe Ala Ile Thr 100 105
110Ala Ala Gly Ala Ser His Ala Val Thr Gln Ala Cys Ser Met Gly Glu
115 120 125Leu Leu Gln Cys Gly Cys Gln
Ala Pro Arg Gly Arg Ala Pro Pro Arg 130 135
140Pro Ser Gly Leu Pro Gly Thr Pro Gly Pro Pro Gly Pro Ala Gly
Ser145 150 155 160Pro Glu
Gly Ser Ala Ala Trp Glu Trp Gly Gly Cys Gly Asp Asp Val
165 170 175Asp Phe Gly Asp Glu Lys Ser
Arg Leu Phe Met Asp Ala Arg His Lys 180 185
190Arg Gly Arg Gly Asp Ile Arg Ala Leu Val Gln Leu His Asn
Asn Glu 195 200 205Ala Gly Arg Leu
Ala Val Arg Ser His Thr Arg Thr Glu Cys Lys Cys 210
215 220His Gly Leu Ser Gly Ser Cys Ala Leu Arg Thr Cys
Trp Gln Lys Leu225 230 235
240Pro Pro Phe Arg Glu Val Gly Ala Arg Leu Leu Glu Arg Phe His Gly
245 250 255Ala Ser Arg Val Met
Gly Thr Asn Asp Gly Lys Ala Leu Leu Pro Ala 260
265 270Val Arg Thr Leu Lys Pro Pro Gly Arg Ala Asp Leu
Leu Tyr Ala Ala 275 280 285Asp Ser
Pro Asp Phe Cys Ala Pro Asn Arg Arg Thr Gly Ser Pro Gly 290
295 300Thr Arg Gly Arg Ala Cys Asn Ser Ser Ala Pro
Asp Leu Ser Gly Cys305 310 315
320Asp Leu Leu Cys Cys Gly Arg Gly His Arg Gln Glu Ser Val Gln Leu
325 330 335Glu Glu Asn Cys
Leu Cys Arg Phe His Trp Cys Cys Val Val Gln Cys 340
345 350His Arg Cys Arg Val Arg Lys Glu Leu Ser Leu
Cys Leu 355 360 365191732DNAHomo
sapiensCDS(306)..(1355) 19gaggggcggg ggctggaggc agcagcgccc ccgcactccc
cgcgtctcgc acacttgcac 60cggtcgctcg cgcgcagccc ggcgtcgccc cacgccgcgc
tcgctcctcc ctccctcctc 120ccgctccgtg gctcccgtgc tcctggcgag gctcaggcgc
ggagcgcgcg gacgggcgca 180ccgacagacg gccccgggga cgcctcggct cgcgcctccc
gggcgggcta tgttgattgc 240cccgccgggg ccggcccgcg ggatcagcac agcccggccc
gcggccccgg cggccaatcg 300ggact atg aac cgg aaa gcg cgg cgc tgc ctg ggc
cac ctc ttt ctc agc 350 Met Asn Arg Lys Ala Arg Arg Cys Leu Gly
His Leu Phe Leu Ser 1 5 10
15ctg ggc atg gtc tac ctc cgg atc ggt ggc ttc tcc tca gtg gta gct
398Leu Gly Met Val Tyr Leu Arg Ile Gly Gly Phe Ser Ser Val Val Ala
20 25 30ctg ggc gca agc atc
atc tgt aac aag atc cca ggc ctg gct ccc aga 446Leu Gly Ala Ser Ile
Ile Cys Asn Lys Ile Pro Gly Leu Ala Pro Arg 35
40 45cag cgg gcg atc tgc cag agc cgg ccc gac gcc atc
atc gtc ata gga 494Gln Arg Ala Ile Cys Gln Ser Arg Pro Asp Ala Ile
Ile Val Ile Gly 50 55 60gaa ggc
tca caa atg ggc ctg gac gag tgt cag ttt cag ttc cgc aat 542Glu Gly
Ser Gln Met Gly Leu Asp Glu Cys Gln Phe Gln Phe Arg Asn 65
70 75ggc cgc tgg aac tgc tct gca ctg gga gag cgc
acc gtc ttc ggg aag 590Gly Arg Trp Asn Cys Ser Ala Leu Gly Glu Arg
Thr Val Phe Gly Lys80 85 90
95gag ctc aaa gtg ggg agc cgg gag gct gcg ttc acc tac gcc atc att
638Glu Leu Lys Val Gly Ser Arg Glu Ala Ala Phe Thr Tyr Ala Ile Ile
100 105 110gcc gcc ggc gtg gcc
cac gcc atc aca gct gcc tgt acc cag ggc aac 686Ala Ala Gly Val Ala
His Ala Ile Thr Ala Ala Cys Thr Gln Gly Asn 115
120 125ctg agc gac tgt ggc tgc gac aaa gag aag caa ggc
cag tac cac cgg 734Leu Ser Asp Cys Gly Cys Asp Lys Glu Lys Gln Gly
Gln Tyr His Arg 130 135 140gac gag
ggc tgg aag tgg ggt ggc tgc tct gcc gac atc cgc tac ggc 782Asp Glu
Gly Trp Lys Trp Gly Gly Cys Ser Ala Asp Ile Arg Tyr Gly 145
150 155atc ggc ttc gcc aag gtc ttt gtg gat gcc cgg
gag atc aag cag aat 830Ile Gly Phe Ala Lys Val Phe Val Asp Ala Arg
Glu Ile Lys Gln Asn160 165 170
175gcc cgg act ctc atg aac ttg cac aac aac gag gca ggc cga aag atc
878Ala Arg Thr Leu Met Asn Leu His Asn Asn Glu Ala Gly Arg Lys Ile
180 185 190ctg gag gag aac atg
aag ctg gaa tgt aag tgc cac ggc gtg tca ggc 926Leu Glu Glu Asn Met
Lys Leu Glu Cys Lys Cys His Gly Val Ser Gly 195
200 205tcg tgc acc acc aag acg tgc tgg acc aca ctg cca
cag ttt cgg gag 974Ser Cys Thr Thr Lys Thr Cys Trp Thr Thr Leu Pro
Gln Phe Arg Glu 210 215 220ctg ggc
tac gtg ctc aag gac aag tac aac gag gcc gtt cac gtg gag 1022Leu Gly
Tyr Val Leu Lys Asp Lys Tyr Asn Glu Ala Val His Val Glu 225
230 235cct gtg cgt gcc agc cgc aac aag cgg ccc acc
ttc ctg aag atc aag 1070Pro Val Arg Ala Ser Arg Asn Lys Arg Pro Thr
Phe Leu Lys Ile Lys240 245 250
255aag cca ctg tcg tac cgc aag ccc atg gac acg gac ctg gtg tac atc
1118Lys Pro Leu Ser Tyr Arg Lys Pro Met Asp Thr Asp Leu Val Tyr Ile
260 265 270gag aag tcg ccc aac
tac tgc gag gag gac ccg gtg acc ggc agt gtg 1166Glu Lys Ser Pro Asn
Tyr Cys Glu Glu Asp Pro Val Thr Gly Ser Val 275
280 285ggc acc cag ggc cgc gcc tgc aac aag acg gct ccc
cag gcc agc ggc 1214Gly Thr Gln Gly Arg Ala Cys Asn Lys Thr Ala Pro
Gln Ala Ser Gly 290 295 300tgt gac
ctc atg tgc tgt ggg cgt ggc tac aac acc cac cag tac gcc 1262Cys Asp
Leu Met Cys Cys Gly Arg Gly Tyr Asn Thr His Gln Tyr Ala 305
310 315cgc gtg tgg cag tgc aac tgt aag ttc cac tgg
tgc tgc tat gtc aag 1310Arg Val Trp Gln Cys Asn Cys Lys Phe His Trp
Cys Cys Tyr Val Lys320 325 330
335tgc aac acg tgc agc gag cgc acg gag atg tac acg tgc aag tga
1355Cys Asn Thr Cys Ser Glu Arg Thr Glu Met Tyr Thr Cys Lys
340 345 350gccccgtgtg cacaccaccc
tcccgctgca agtcagattg ctgggaggac tggaccgttt 1415ccaagctgcg ggctccctgg
caggatgctg agcttgtctt ttctgctgag gagggtactt 1475ttcctgggtt tcctgcaggc
atccgtgggg gaaaaaaaat ctctcagagc cctcaactat 1535tctgttccac acccaatgct
gctccaccct cccccagaca cagcccaggt ccctccgcgg 1595ctggagcgaa gccttctgca
gcaggaactc tggacccctg ggcctcatca cagcaatatt 1655taacaattta ttctgataaa
aataatatta atttatttaa ttaaaaagaa ttcttccaca 1715aaaaaaaaaa aaaaaaa
173220349PRTHomo sapiens
20Met Asn Arg Lys Ala Arg Arg Cys Leu Gly His Leu Phe Leu Ser Leu1
5 10 15Gly Met Val Tyr Leu Arg
Ile Gly Gly Phe Ser Ser Val Val Ala Leu 20 25
30Gly Ala Ser Ile Ile Cys Asn Lys Ile Pro Gly Leu Ala
Pro Arg Gln 35 40 45Arg Ala Ile
Cys Gln Ser Arg Pro Asp Ala Ile Ile Val Ile Gly Glu 50
55 60Gly Ser Gln Met Gly Leu Asp Glu Cys Gln Phe Gln
Phe Arg Asn Gly65 70 75
80Arg Trp Asn Cys Ser Ala Leu Gly Glu Arg Thr Val Phe Gly Lys Glu
85 90 95Leu Lys Val Gly Ser Arg
Glu Ala Ala Phe Thr Tyr Ala Ile Ile Ala 100
105 110Ala Gly Val Ala His Ala Ile Thr Ala Ala Cys Thr
Gln Gly Asn Leu 115 120 125Ser Asp
Cys Gly Cys Asp Lys Glu Lys Gln Gly Gln Tyr His Arg Asp 130
135 140Glu Gly Trp Lys Trp Gly Gly Cys Ser Ala Asp
Ile Arg Tyr Gly Ile145 150 155
160Gly Phe Ala Lys Val Phe Val Asp Ala Arg Glu Ile Lys Gln Asn Ala
165 170 175Arg Thr Leu Met
Asn Leu His Asn Asn Glu Ala Gly Arg Lys Ile Leu 180
185 190Glu Glu Asn Met Lys Leu Glu Cys Lys Cys His
Gly Val Ser Gly Ser 195 200 205Cys
Thr Thr Lys Thr Cys Trp Thr Thr Leu Pro Gln Phe Arg Glu Leu 210
215 220Gly Tyr Val Leu Lys Asp Lys Tyr Asn Glu
Ala Val His Val Glu Pro225 230 235
240Val Arg Ala Ser Arg Asn Lys Arg Pro Thr Phe Leu Lys Ile Lys
Lys 245 250 255Pro Leu Ser
Tyr Arg Lys Pro Met Asp Thr Asp Leu Val Tyr Ile Glu 260
265 270Lys Ser Pro Asn Tyr Cys Glu Glu Asp Pro
Val Thr Gly Ser Val Gly 275 280
285Thr Gln Gly Arg Ala Cys Asn Lys Thr Ala Pro Gln Ala Ser Gly Cys 290
295 300Asp Leu Met Cys Cys Gly Arg Gly
Tyr Asn Thr His Gln Tyr Ala Arg305 310
315 320Val Trp Gln Cys Asn Cys Lys Phe His Trp Cys Cys
Tyr Val Lys Cys 325 330
335Asn Thr Cys Ser Glu Arg Thr Glu Met Tyr Thr Cys Lys 340
345212250DNAHomo sapiensCDS(96)..(1145) 21gagtctgccc
gcagccccct ggcccctgcc cggccctgcg tgcccgcgcg tccctccggc 60cgcgctgtct
atggcgcagc ccccctccct ggatc atg cac aga aac ttt cgc 113
Met His Arg Asn Phe Arg
1 5aag tgg att ttc tac gtg ttt ctc tgc ttt
ggc gtc ctg tac gtg aag 161Lys Trp Ile Phe Tyr Val Phe Leu Cys Phe
Gly Val Leu Tyr Val Lys 10 15
20ctc gga gca ctg tca tcc gtg gtg gcc ctg gga gcc aac atc atc tgc
209Leu Gly Ala Leu Ser Ser Val Val Ala Leu Gly Ala Asn Ile Ile Cys
25 30 35aac aag att cct ggc cta gcc ccg
cgg cag cgt gcc atc tgc cag agt 257Asn Lys Ile Pro Gly Leu Ala Pro
Arg Gln Arg Ala Ile Cys Gln Ser 40 45
50cgg ccc gat gcc atc att gtg att ggg gag ggg gcg cag atg ggc atc
305Arg Pro Asp Ala Ile Ile Val Ile Gly Glu Gly Ala Gln Met Gly Ile55
60 65 70aac gag tgc cag tac
cag ttc cgc ttc gga cgc tgg aac tgc tct gcc 353Asn Glu Cys Gln Tyr
Gln Phe Arg Phe Gly Arg Trp Asn Cys Ser Ala 75
80 85ctc ggc gag aag acc gtc ttc ggg caa gag ctc
cga gta ggg agc cgt 401Leu Gly Glu Lys Thr Val Phe Gly Gln Glu Leu
Arg Val Gly Ser Arg 90 95
100gag gct gcc ttc acg tac gcc atc acc gcg gct ggc gtg gcg cac gcc
449Glu Ala Ala Phe Thr Tyr Ala Ile Thr Ala Ala Gly Val Ala His Ala
105 110 115gtc acc gct gcc tgc agc caa
ggg aac ctg agc aac tgc ggc tgc gac 497Val Thr Ala Ala Cys Ser Gln
Gly Asn Leu Ser Asn Cys Gly Cys Asp 120 125
130cgc gag aag cag ggc tac tac aac caa gcc gag ggc tgg aag tgg ggc
545Arg Glu Lys Gln Gly Tyr Tyr Asn Gln Ala Glu Gly Trp Lys Trp Gly135
140 145 150ggc tgc tcg gcc
gac gtg cgt tac ggc atc gac ttc tcc cgg cgc ttc 593Gly Cys Ser Ala
Asp Val Arg Tyr Gly Ile Asp Phe Ser Arg Arg Phe 155
160 165gtg gac gct cgg gag atc aag aag aac gcg
cgg cgc ctc atg aac ctg 641Val Asp Ala Arg Glu Ile Lys Lys Asn Ala
Arg Arg Leu Met Asn Leu 170 175
180cat aac aat gag gcc ggc agg aag gtt cta gag gac cgg atg cag ctg
689His Asn Asn Glu Ala Gly Arg Lys Val Leu Glu Asp Arg Met Gln Leu
185 190 195gag tgc aag tgc cac ggc gtg
tct ggc tcc tgc acc acc aaa acc tgc 737Glu Cys Lys Cys His Gly Val
Ser Gly Ser Cys Thr Thr Lys Thr Cys 200 205
210tgg acc acg ctg ccc aag ttc cga gag gtg ggc cac ctg ctg aag gag
785Trp Thr Thr Leu Pro Lys Phe Arg Glu Val Gly His Leu Leu Lys Glu215
220 225 230aag tac aac gcg
gcc gtg cag gtg gag gtg gtg cgg gcc agc cgt ctg 833Lys Tyr Asn Ala
Ala Val Gln Val Glu Val Val Arg Ala Ser Arg Leu 235
240 245cgg cag ccc acc ttc ctg cgc atc aaa cag
ctg cgc agc tat cag aag 881Arg Gln Pro Thr Phe Leu Arg Ile Lys Gln
Leu Arg Ser Tyr Gln Lys 250 255
260ccc atg gag aca gac ctg gtg tac att gag aag tcg ccc aac tac tgc
929Pro Met Glu Thr Asp Leu Val Tyr Ile Glu Lys Ser Pro Asn Tyr Cys
265 270 275gag gag gac gcg gcc acg ggc
agc gtg ggc acg cag ggc cgt ctc tgc 977Glu Glu Asp Ala Ala Thr Gly
Ser Val Gly Thr Gln Gly Arg Leu Cys 280 285
290aac cgc acg tcg ccc ggc gcg gac ggc tgt gac acc atg tgc tgc ggc
1025Asn Arg Thr Ser Pro Gly Ala Asp Gly Cys Asp Thr Met Cys Cys Gly295
300 305 310cga ggc tac aac
acc cac cag tac acc aag gtg tgg cag tgc aac tgc 1073Arg Gly Tyr Asn
Thr His Gln Tyr Thr Lys Val Trp Gln Cys Asn Cys 315
320 325aaa ttc cac tgg tgc tgc ttc gtc aag tgc
aac acc tgc agc gag cgc 1121Lys Phe His Trp Cys Cys Phe Val Lys Cys
Asn Thr Cys Ser Glu Arg 330 335
340acc gag gtc ttc acc tgc aag tga ggccaggccc ggaggcggcc gcgggcaccc
1175Thr Glu Val Phe Thr Cys Lys 345 350tggaacccgg
cggcattttg cacatccact cctcaccttc cctgccttgg tgctgccagc 1235agcagacata
gacgggtgca gaagcgggga gctccaggtg caggagggca ccggccgggg 1295cccacgccct
ctgcccgcct ccctggggct ccttcctgcc acctcctccc atcacctcct 1355gcggcagaac
agcacccgtg acccacccag agagcaaggc caggggtctt ggtgctcccc 1415gacggggccc
ggcaagttct ctttcttctc tctgggaaaa tgaacgtcca ggacacacct 1475gtatcccaga
gagcaaagtg atgaggagac tgagcgtccc cagccccacc tggcggcatg 1535gacacagaaa
agctacgccg gctggcctct ccagaccagt tcccaggctg ggtctgccgc 1595tgggccctgg
ggcggtgggg acagatgttg acacaaatta tttatgtttt cttagtatca 1655gaagaggatt
ctcggcacta acacatagcc agtcctaact ccgtactctg tgtcagccca 1715tcccctagac
accctctgtt tcctttcccg gccccacctg gccggccctc tgcccctgca 1775gagctgaggc
agcctggggt tgatggggac cacgcggtgc ctgcaggtcc tagaagtgag 1835ctgggcaggg
gctcttcaga ccacacagcc ctgaccgggc cttggaggag agccatggac 1895aggctcctcc
atgccgtctt tccttctttt gaaaatccta tcaatggctg ggcgcggtgg 1955ctcacacctg
taatcccagc actttgggag accgaggcag gtggatcacc tgaggtcagg 2015agttcgagac
cagcctggcc aacgtggtga aaccctgtct ctactaaaaa tacaaaaatt 2075agctgggcgt
ggtggcgtgc acctgtaatc ccagctactc aggaggctga gacaggacac 2135ttgcttgaac
ccgggaggtg gaggttgcaa tgagccaaga ttgtgccact gtattccaac 2195ttgggtgaca
gagcacgact ctgtctcaaa aaaaaaaaaa aaaaaaaaaa aaaaa 225022349PRTHomo
sapiens 22Met His Arg Asn Phe Arg Lys Trp Ile Phe Tyr Val Phe Leu Cys
Phe1 5 10 15Gly Val Leu
Tyr Val Lys Leu Gly Ala Leu Ser Ser Val Val Ala Leu 20
25 30Gly Ala Asn Ile Ile Cys Asn Lys Ile Pro
Gly Leu Ala Pro Arg Gln 35 40
45Arg Ala Ile Cys Gln Ser Arg Pro Asp Ala Ile Ile Val Ile Gly Glu 50
55 60Gly Ala Gln Met Gly Ile Asn Glu Cys
Gln Tyr Gln Phe Arg Phe Gly65 70 75
80Arg Trp Asn Cys Ser Ala Leu Gly Glu Lys Thr Val Phe Gly
Gln Glu 85 90 95Leu Arg
Val Gly Ser Arg Glu Ala Ala Phe Thr Tyr Ala Ile Thr Ala 100
105 110Ala Gly Val Ala His Ala Val Thr Ala
Ala Cys Ser Gln Gly Asn Leu 115 120
125Ser Asn Cys Gly Cys Asp Arg Glu Lys Gln Gly Tyr Tyr Asn Gln Ala
130 135 140Glu Gly Trp Lys Trp Gly Gly
Cys Ser Ala Asp Val Arg Tyr Gly Ile145 150
155 160Asp Phe Ser Arg Arg Phe Val Asp Ala Arg Glu Ile
Lys Lys Asn Ala 165 170
175Arg Arg Leu Met Asn Leu His Asn Asn Glu Ala Gly Arg Lys Val Leu
180 185 190Glu Asp Arg Met Gln Leu
Glu Cys Lys Cys His Gly Val Ser Gly Ser 195 200
205Cys Thr Thr Lys Thr Cys Trp Thr Thr Leu Pro Lys Phe Arg
Glu Val 210 215 220Gly His Leu Leu Lys
Glu Lys Tyr Asn Ala Ala Val Gln Val Glu Val225 230
235 240Val Arg Ala Ser Arg Leu Arg Gln Pro Thr
Phe Leu Arg Ile Lys Gln 245 250
255Leu Arg Ser Tyr Gln Lys Pro Met Glu Thr Asp Leu Val Tyr Ile Glu
260 265 270Lys Ser Pro Asn Tyr
Cys Glu Glu Asp Ala Ala Thr Gly Ser Val Gly 275
280 285Thr Gln Gly Arg Leu Cys Asn Arg Thr Ser Pro Gly
Ala Asp Gly Cys 290 295 300Asp Thr Met
Cys Cys Gly Arg Gly Tyr Asn Thr His Gln Tyr Thr Lys305
310 315 320Val Trp Gln Cys Asn Cys Lys
Phe His Trp Cys Cys Phe Val Lys Cys 325
330 335Asn Thr Cys Ser Glu Arg Thr Glu Val Phe Thr Cys
Lys 340 345231899DNAHomo
sapiensCDS(101)..(1156) 23cagaattttc tcacataaat actgaggaag accctgccct
ctcctcactc ctctggactt 60ggccctgagc tggacctggt ccactggggt aggcagggcg
atg ggg aac ctg ttt 115
Met Gly Asn Leu Phe 1
5atg ctc tgg gca gct ctg ggc ata tgc tgt gct gca ttc agt gcc tct
163Met Leu Trp Ala Ala Leu Gly Ile Cys Cys Ala Ala Phe Ser Ala Ser
10 15 20gcc tgg tca gtg aac
aat ttc ctg ata aca ggt ccc aag gcc tat ctg 211Ala Trp Ser Val Asn
Asn Phe Leu Ile Thr Gly Pro Lys Ala Tyr Leu 25
30 35acc tac acg act agt gtg gcc ttg ggt gcc cag agt
ggc atc gag gag 259Thr Tyr Thr Thr Ser Val Ala Leu Gly Ala Gln Ser
Gly Ile Glu Glu 40 45 50tgc aag
ttc cag ttt gct tgg gaa cgc tgg aac tgc cct gaa aat gct 307Cys Lys
Phe Gln Phe Ala Trp Glu Arg Trp Asn Cys Pro Glu Asn Ala 55
60 65ctt cag ctc tcc acc cac aac agg ctg aga agt
gct acc aga gag act 355Leu Gln Leu Ser Thr His Asn Arg Leu Arg Ser
Ala Thr Arg Glu Thr70 75 80
85tcc ttc ata cat gct atc agc tct gct gga gtc atg tac atc atc acc
403Ser Phe Ile His Ala Ile Ser Ser Ala Gly Val Met Tyr Ile Ile Thr
90 95 100aag aac tgt agc atg
ggt gac ttc gaa aac tgt ggc tgt gat ggg tca 451Lys Asn Cys Ser Met
Gly Asp Phe Glu Asn Cys Gly Cys Asp Gly Ser 105
110 115aac aat gga aaa aca gga ggc cat ggc tgg atc tgg
gga ggc tgc agc 499Asn Asn Gly Lys Thr Gly Gly His Gly Trp Ile Trp
Gly Gly Cys Ser 120 125 130gac aat
gtg gaa ttt ggg gaa agg atc tcc aaa ctc ttt gtg gac agt 547Asp Asn
Val Glu Phe Gly Glu Arg Ile Ser Lys Leu Phe Val Asp Ser 135
140 145ttg gag aag ggg aag gat gcc aga gcc ctg atg
aat ctt cac aac aac 595Leu Glu Lys Gly Lys Asp Ala Arg Ala Leu Met
Asn Leu His Asn Asn150 155 160
165agg gcc ggc aga ctg gca gtg aga gcc acc atg aaa agg aca tgc aaa
643Arg Ala Gly Arg Leu Ala Val Arg Ala Thr Met Lys Arg Thr Cys Lys
170 175 180tgt cat ggc atc tct
ggg agc tgc agc ata cag aca tgc tgg ctg cag 691Cys His Gly Ile Ser
Gly Ser Cys Ser Ile Gln Thr Cys Trp Leu Gln 185
190 195ctg gct gaa ttc cgg gag atg gga gac tac cta aag
gcc aag tat gac 739Leu Ala Glu Phe Arg Glu Met Gly Asp Tyr Leu Lys
Ala Lys Tyr Asp 200 205 210cag gcg
ctg aaa att gaa atg gat aag cgg cag ctg aga gct ggg aac 787Gln Ala
Leu Lys Ile Glu Met Asp Lys Arg Gln Leu Arg Ala Gly Asn 215
220 225agc gcc gag ggc cac tgg gtg ccc gct gag gcc
ttc ctt cct agc gca 835Ser Ala Glu Gly His Trp Val Pro Ala Glu Ala
Phe Leu Pro Ser Ala230 235 240
245gag gcg gaa ctg atc ttt tta gag gaa tca cca gat tac tgt acc tgc
883Glu Ala Glu Leu Ile Phe Leu Glu Glu Ser Pro Asp Tyr Cys Thr Cys
250 255 260aat tcc agc ctg ggc
atc tat ggc aca gag ggt cgt gag tgc cta cag 931Asn Ser Ser Leu Gly
Ile Tyr Gly Thr Glu Gly Arg Glu Cys Leu Gln 265
270 275aac agc cac aac aca tcc agg tgg gag cga cgt agc
tgt ggg cgc ctg 979Asn Ser His Asn Thr Ser Arg Trp Glu Arg Arg Ser
Cys Gly Arg Leu 280 285 290tgc act
gag tgt ggg ctg cag gtg gaa gag agg aaa act gag gtc ata 1027Cys Thr
Glu Cys Gly Leu Gln Val Glu Glu Arg Lys Thr Glu Val Ile 295
300 305agc agc tgt aac tgc aaa ttc cag tgg tgc tgt
acg gtc aag tgt gac 1075Ser Ser Cys Asn Cys Lys Phe Gln Trp Cys Cys
Thr Val Lys Cys Asp310 315 320
325cag tgt agg cat gtg gtg agc aag tat tac tgc gca cgc tcc cca ggc
1123Gln Cys Arg His Val Val Ser Lys Tyr Tyr Cys Ala Arg Ser Pro Gly
330 335 340agt gcc cag tcc ctg
ggt aag ggc agt gcc tga taatacccca cacaagttca 1176Ser Ala Gln Ser Leu
Gly Lys Gly Ser Ala 345 350cttgattaat
tgcatcagtg gaaggggaca tagcttctct cttagagaga acagattgga 1236aagcaatcgg
aaaattgcag ttttggtctg tagtcctcat gatatctgct atcagtgggg 1296aaaatggagg
cccaagattc tacagcatat tcctggcggg gctgaaattg gaacctgggc 1356ctcctgactt
tggcagaccc ccatttcatc tttcctgcaa actactttcc catctttgtg 1416cctgtactta
tgcagctttc tacagggaga gtttggtttg gggtctatat ctagagggac 1476cttcaaagta
tttgttcctt taaatttcag accatgtcca acccagctgt gctgctggga 1536atcaggagaa
tagaagcaaa aaacgaaaga gttctgttca gacttctgaa gagcagcctg 1596tggctacaaa
tctatgctga taaatgagat tgagaactca actgtatttt gccataaatg 1656cttctaagat
atatccagct gggacttcta ttactccctt tggaaacctt aagatcaaaa 1716agggaataag
aaacccttct tctgtatccc aataatccac caggataaag gagaaactag 1776aaatatgcaa
ctcccttgat ttcagtgttt ggcaggtaac aaaaaattga gacccagaca 1836ctggtcaaca
ggaaaacaat acagactccc agaattagaa agtgttattt taatgcaacc 1896tag
189924351PRTHomo
sapiens 24Met Gly Asn Leu Phe Met Leu Trp Ala Ala Leu Gly Ile Cys Cys
Ala1 5 10 15Ala Phe Ser
Ala Ser Ala Trp Ser Val Asn Asn Phe Leu Ile Thr Gly 20
25 30Pro Lys Ala Tyr Leu Thr Tyr Thr Thr Ser
Val Ala Leu Gly Ala Gln 35 40
45Ser Gly Ile Glu Glu Cys Lys Phe Gln Phe Ala Trp Glu Arg Trp Asn 50
55 60Cys Pro Glu Asn Ala Leu Gln Leu Ser
Thr His Asn Arg Leu Arg Ser65 70 75
80Ala Thr Arg Glu Thr Ser Phe Ile His Ala Ile Ser Ser Ala
Gly Val 85 90 95Met Tyr
Ile Ile Thr Lys Asn Cys Ser Met Gly Asp Phe Glu Asn Cys 100
105 110Gly Cys Asp Gly Ser Asn Asn Gly Lys
Thr Gly Gly His Gly Trp Ile 115 120
125Trp Gly Gly Cys Ser Asp Asn Val Glu Phe Gly Glu Arg Ile Ser Lys
130 135 140Leu Phe Val Asp Ser Leu Glu
Lys Gly Lys Asp Ala Arg Ala Leu Met145 150
155 160Asn Leu His Asn Asn Arg Ala Gly Arg Leu Ala Val
Arg Ala Thr Met 165 170
175Lys Arg Thr Cys Lys Cys His Gly Ile Ser Gly Ser Cys Ser Ile Gln
180 185 190Thr Cys Trp Leu Gln Leu
Ala Glu Phe Arg Glu Met Gly Asp Tyr Leu 195 200
205Lys Ala Lys Tyr Asp Gln Ala Leu Lys Ile Glu Met Asp Lys
Arg Gln 210 215 220Leu Arg Ala Gly Asn
Ser Ala Glu Gly His Trp Val Pro Ala Glu Ala225 230
235 240Phe Leu Pro Ser Ala Glu Ala Glu Leu Ile
Phe Leu Glu Glu Ser Pro 245 250
255Asp Tyr Cys Thr Cys Asn Ser Ser Leu Gly Ile Tyr Gly Thr Glu Gly
260 265 270Arg Glu Cys Leu Gln
Asn Ser His Asn Thr Ser Arg Trp Glu Arg Arg 275
280 285Ser Cys Gly Arg Leu Cys Thr Glu Cys Gly Leu Gln
Val Glu Glu Arg 290 295 300Lys Thr Glu
Val Ile Ser Ser Cys Asn Cys Lys Phe Gln Trp Cys Cys305
310 315 320Thr Val Lys Cys Asp Gln Cys
Arg His Val Val Ser Lys Tyr Tyr Cys 325
330 335Ala Arg Ser Pro Gly Ser Ala Gln Ser Leu Gly Lys
Gly Ser Ala 340 345
350252117DNAHomo sapiensCDS(136)..(1191) 25tccgcttaca caccaaggaa
agttgggctt tgaagaattc catccccatg gccactggag 60gaagaatatt tctccgtctt
gcttacccat ctcccagttt tttggaattt tctctagctg 120ttactccaga ggatt atg
ttt ctt tca aag cct tct gtg tac atc tgt ctt 171 Met
Phe Leu Ser Lys Pro Ser Val Tyr Ile Cys Leu 1
5 10ttc acc tgt gtc ctc caa ctc agc cac agc tgg tcg
gtg aac aat ttc 219Phe Thr Cys Val Leu Gln Leu Ser His Ser Trp Ser
Val Asn Asn Phe 15 20 25ctg atg
act ggt cca aag gct tac ctg att tac tcc agc agt gtg gca 267Leu Met
Thr Gly Pro Lys Ala Tyr Leu Ile Tyr Ser Ser Ser Val Ala 30
35 40gct ggt gcc cag agt ggt att gaa gaa tgc aag
tat cag ttt gcc tgg 315Ala Gly Ala Gln Ser Gly Ile Glu Glu Cys Lys
Tyr Gln Phe Ala Trp45 50 55
60gac cgc tgg aac tgc cct gag aga gcc ctg cag ctg tcc agc cat ggt
363Asp Arg Trp Asn Cys Pro Glu Arg Ala Leu Gln Leu Ser Ser His Gly
65 70 75ggg ctt cgc agt gcc
aat cgg gag aca gca ttt gtg cat gcc atc agt 411Gly Leu Arg Ser Ala
Asn Arg Glu Thr Ala Phe Val His Ala Ile Ser 80
85 90tct gct gga gtc atg tac acc ctg act aga aac tgc
agc ctt gga gat 459Ser Ala Gly Val Met Tyr Thr Leu Thr Arg Asn Cys
Ser Leu Gly Asp 95 100 105ttt gat
aac tgt ggc tgt gat gac tcc cgc aac ggg caa ctg ggg gga 507Phe Asp
Asn Cys Gly Cys Asp Asp Ser Arg Asn Gly Gln Leu Gly Gly 110
115 120caa ggc tgg ctg tgg gga ggc tgc agt gac aat
gtg ggc ttc gga gag 555Gln Gly Trp Leu Trp Gly Gly Cys Ser Asp Asn
Val Gly Phe Gly Glu125 130 135
140gcg att tcc aag cag ttt gtc gat gcc ctg gaa aca gga cag gat gca
603Ala Ile Ser Lys Gln Phe Val Asp Ala Leu Glu Thr Gly Gln Asp Ala
145 150 155cgg gca gcc atg aac
ctg cac aac aac gag gct ggc cgc aag gcg gtg 651Arg Ala Ala Met Asn
Leu His Asn Asn Glu Ala Gly Arg Lys Ala Val 160
165 170aag ggc acc atg aaa cgc acg tgt aag tgc cat ggc
gtg tct ggc agc 699Lys Gly Thr Met Lys Arg Thr Cys Lys Cys His Gly
Val Ser Gly Ser 175 180 185tgc acc
acg cag acc tgt tgg ctg cag ctg ccc gag ttc cgc gag gtg 747Cys Thr
Thr Gln Thr Cys Trp Leu Gln Leu Pro Glu Phe Arg Glu Val 190
195 200ggc gcg cac ctg aag gag aag tac cac gca gca
ctc aag gtg gac ctg 795Gly Ala His Leu Lys Glu Lys Tyr His Ala Ala
Leu Lys Val Asp Leu205 210 215
220ctg cag ggt gct ggc aac agc gcg gcc gcc cgc ggc gcc atc gcc gac
843Leu Gln Gly Ala Gly Asn Ser Ala Ala Ala Arg Gly Ala Ile Ala Asp
225 230 235acc ttt cgc tcc atc
tct acc cgg gag ctg gtg cac ctg gag gac tcc 891Thr Phe Arg Ser Ile
Ser Thr Arg Glu Leu Val His Leu Glu Asp Ser 240
245 250ccg gac tac tgc ctg gag aac aaa acg cta ggg ctg
ctg ggc acc gaa 939Pro Asp Tyr Cys Leu Glu Asn Lys Thr Leu Gly Leu
Leu Gly Thr Glu 255 260 265ggc cga
gag tgc cta agg cgc ggg cgg gcc ctg ggt cgc tgg gaa ctc 987Gly Arg
Glu Cys Leu Arg Arg Gly Arg Ala Leu Gly Arg Trp Glu Leu 270
275 280cgc agc tgc cgc cgg ctc tgc ggg gac tgc ggg
ctg gcg gtg gag gag 1035Arg Ser Cys Arg Arg Leu Cys Gly Asp Cys Gly
Leu Ala Val Glu Glu285 290 295
300cgc cgg gcc gag acc gtg tcc agc tgc aac tgc aag ttc cac tgg tgc
1083Arg Arg Ala Glu Thr Val Ser Ser Cys Asn Cys Lys Phe His Trp Cys
305 310 315tgt gca gtc cgc tgc
gag cag tgc cgc cgg agg gtc acc aag tac ttc 1131Cys Ala Val Arg Cys
Glu Gln Cys Arg Arg Arg Val Thr Lys Tyr Phe 320
325 330tgt agc cgc gca gag cgg ccg cgg ggg ggc gct gcg
cac aaa ccc ggg 1179Cys Ser Arg Ala Glu Arg Pro Arg Gly Gly Ala Ala
His Lys Pro Gly 335 340 345aga aaa
ccc taa gggtttcctc tgccccctcc ttttcccact ggttcttggc 1231Arg Lys
Pro 350ttcctttaga gaccccggta attgtggaac ctagggaatg gggaacccgc
tctcccagac 1291ctagggatcc tgaaagggaa aaactgcaat ttctccaaag cttgccactt
tccagcctgt 1351ttccccaatt cctctgtgct ctcctaaagc tctgtctgaa tcctcgcagc
cacacctagg 1411tctgaaaact caggctttga gttactgatc ttccttggat taggaaaaca
ggtgttcctc 1471ctcccctctc ctatcagccc taatctctga cctagcctat caacccttag
gcgctggaaa 1531aaccttctca tacacgcagg acccaggtta actcaaagct ttgccctttt
gcccactgtc 1591tgctaccagg ggctcaccct ctgctgcacc tctcttctgc acagctcctc
ccctgctact 1651gctgaccaaa ttcccaggaa tcttgaatgc tttctctcct cttctccctt
tcctttccca 1711aaaaaaactg aggaaactgg ccccggaaaa gcatgtcttt ggggttggtt
cctagaggca 1771gaggttgaag atggaagagg gagctctgga gtgctaactt gaacaccaag
ggtgctactc 1831atccctatgg tatcatatca tgaatggact ttactagtgg ggcaatgact
ttcctagaca 1891ataacccgag ggactccaga tacatacccc gaaggtctag gaaatacgtt
aagggcagat 1951tacagtcatt tcctaccctt taaaggtaac ttctcccttc tcctgaccta
cttcctccta 2011gcaaccaact ttacctcttc ttctccaaag gatctttgtt cctctgagcc
aagactgagg 2071taaataaagc cactttcctc ttcagatcct ggtctgcacc tctaga
211726351PRTHomo sapiens 26Met Phe Leu Ser Lys Pro Ser Val Tyr
Ile Cys Leu Phe Thr Cys Val1 5 10
15Leu Gln Leu Ser His Ser Trp Ser Val Asn Asn Phe Leu Met Thr
Gly 20 25 30Pro Lys Ala Tyr
Leu Ile Tyr Ser Ser Ser Val Ala Ala Gly Ala Gln 35
40 45Ser Gly Ile Glu Glu Cys Lys Tyr Gln Phe Ala Trp
Asp Arg Trp Asn 50 55 60Cys Pro Glu
Arg Ala Leu Gln Leu Ser Ser His Gly Gly Leu Arg Ser65 70
75 80Ala Asn Arg Glu Thr Ala Phe Val
His Ala Ile Ser Ser Ala Gly Val 85 90
95Met Tyr Thr Leu Thr Arg Asn Cys Ser Leu Gly Asp Phe Asp
Asn Cys 100 105 110Gly Cys Asp
Asp Ser Arg Asn Gly Gln Leu Gly Gly Gln Gly Trp Leu 115
120 125Trp Gly Gly Cys Ser Asp Asn Val Gly Phe Gly
Glu Ala Ile Ser Lys 130 135 140Gln Phe
Val Asp Ala Leu Glu Thr Gly Gln Asp Ala Arg Ala Ala Met145
150 155 160Asn Leu His Asn Asn Glu Ala
Gly Arg Lys Ala Val Lys Gly Thr Met 165
170 175Lys Arg Thr Cys Lys Cys His Gly Val Ser Gly Ser
Cys Thr Thr Gln 180 185 190Thr
Cys Trp Leu Gln Leu Pro Glu Phe Arg Glu Val Gly Ala His Leu 195
200 205Lys Glu Lys Tyr His Ala Ala Leu Lys
Val Asp Leu Leu Gln Gly Ala 210 215
220Gly Asn Ser Ala Ala Ala Arg Gly Ala Ile Ala Asp Thr Phe Arg Ser225
230 235 240Ile Ser Thr Arg
Glu Leu Val His Leu Glu Asp Ser Pro Asp Tyr Cys 245
250 255Leu Glu Asn Lys Thr Leu Gly Leu Leu Gly
Thr Glu Gly Arg Glu Cys 260 265
270Leu Arg Arg Gly Arg Ala Leu Gly Arg Trp Glu Leu Arg Ser Cys Arg
275 280 285Arg Leu Cys Gly Asp Cys Gly
Leu Ala Val Glu Glu Arg Arg Ala Glu 290 295
300Thr Val Ser Ser Cys Asn Cys Lys Phe His Trp Cys Cys Ala Val
Arg305 310 315 320Cys Glu
Gln Cys Arg Arg Arg Val Thr Lys Tyr Phe Cys Ser Arg Ala
325 330 335Glu Arg Pro Arg Gly Gly Ala
Ala His Lys Pro Gly Arg Lys Pro 340 345
350271631DNAHomo sapiensCDS(12)..(1109) 27ggcgcggcaa g atg ctg
gat ggg tcc ccg ctg gcg cgc tgg ctg gcc gcg 50 Met Leu
Asp Gly Ser Pro Leu Ala Arg Trp Leu Ala Ala 1 5
10gcc ttc ggg ctg acg ctg ctg ctc gcc gcg ctg cgc cct
tcg gcc gcc 98Ala Phe Gly Leu Thr Leu Leu Leu Ala Ala Leu Arg Pro
Ser Ala Ala 15 20 25tac ttc ggg ctg
acg ggc agc gag ccc ctg acc atc ctc ccg ctg acc 146Tyr Phe Gly Leu
Thr Gly Ser Glu Pro Leu Thr Ile Leu Pro Leu Thr30 35
40 45ctg gag cca gag gcg gcc gcc cag gcg
cac tac aag gcc tgc gac cgg 194Leu Glu Pro Glu Ala Ala Ala Gln Ala
His Tyr Lys Ala Cys Asp Arg 50 55
60ctg aag ctg gag cgg aag cag cgg cgc atg tgc cgc cgg gac ccg
ggc 242Leu Lys Leu Glu Arg Lys Gln Arg Arg Met Cys Arg Arg Asp Pro
Gly 65 70 75gtg gca gag acg
ctg gtg gag gcc gtg agc atg agt gcg ctc gag tgc 290Val Ala Glu Thr
Leu Val Glu Ala Val Ser Met Ser Ala Leu Glu Cys 80
85 90cag ttc cag ttc cgc ttt gag cgc tgg aac tgc acg
ctg gag ggc cgc 338Gln Phe Gln Phe Arg Phe Glu Arg Trp Asn Cys Thr
Leu Glu Gly Arg 95 100 105tac cgg gcc
agc ctg ctc aag cga ggc ttc aag gag act gcc ttc ctc 386Tyr Arg Ala
Ser Leu Leu Lys Arg Gly Phe Lys Glu Thr Ala Phe Leu110
115 120 125tat gcc atc tcc tcg gct ggc
ctg acg cac gca ctg gcc aag gcg tgc 434Tyr Ala Ile Ser Ser Ala Gly
Leu Thr His Ala Leu Ala Lys Ala Cys 130
135 140agc gcg ggc cgc atg gag cgc tgt acc tgc gat gag
gca ccc gac ctg 482Ser Ala Gly Arg Met Glu Arg Cys Thr Cys Asp Glu
Ala Pro Asp Leu 145 150 155gag
aac cgt gag gcc tgg cag tgg ggg ggc tgc gga gac aac ctt aag 530Glu
Asn Arg Glu Ala Trp Gln Trp Gly Gly Cys Gly Asp Asn Leu Lys 160
165 170tac agc agc aag ttc gtc aag gaa ttc
ctg ggc aga cgg tca agc aag 578Tyr Ser Ser Lys Phe Val Lys Glu Phe
Leu Gly Arg Arg Ser Ser Lys 175 180
185gat ctg cga gcc cgt gtg gac ttc cac aac aac ctc gtg ggt gtg aag
626Asp Leu Arg Ala Arg Val Asp Phe His Asn Asn Leu Val Gly Val Lys190
195 200 205gtg atc aag gct
ggg gtg gag acc acc tgc aag tgc cac ggc gtg tca 674Val Ile Lys Ala
Gly Val Glu Thr Thr Cys Lys Cys His Gly Val Ser 210
215 220ggc tca tgc acg gtg cgg acc tgc tgg cgg
cag ttg gcg cct ttc cat 722Gly Ser Cys Thr Val Arg Thr Cys Trp Arg
Gln Leu Ala Pro Phe His 225 230
235gag gtg ggc aag cat ctg aag cac aag tat gag acg gca ctc aag gtg
770Glu Val Gly Lys His Leu Lys His Lys Tyr Glu Thr Ala Leu Lys Val
240 245 250ggc agc acc acc aat gaa gct
gcc ggc gag gca ggt gcc atc tcc cca 818Gly Ser Thr Thr Asn Glu Ala
Ala Gly Glu Ala Gly Ala Ile Ser Pro 255 260
265cca cgg ggc cgt gcc tcg ggg gca ggt ggc agc gac ccg ctg ccc cgc
866Pro Arg Gly Arg Ala Ser Gly Ala Gly Gly Ser Asp Pro Leu Pro Arg270
275 280 285act cca gag ctg
gtg cac ctg gat gac tcg cct agc ttc tgc ctg gct 914Thr Pro Glu Leu
Val His Leu Asp Asp Ser Pro Ser Phe Cys Leu Ala 290
295 300ggc cgc ttc tcc ccg ggc acc gct ggc cgt
agg tgc cac cgt gag aag 962Gly Arg Phe Ser Pro Gly Thr Ala Gly Arg
Arg Cys His Arg Glu Lys 305 310
315aac tgc gag agc atc tgc tgt ggc cgc ggc cat aac aca cag agc cgg
1010Asn Cys Glu Ser Ile Cys Cys Gly Arg Gly His Asn Thr Gln Ser Arg
320 325 330gtg gtg aca agg ccc tgc cag
tgc cag gtg cgt tgg tgc tgc tat gtg 1058Val Val Thr Arg Pro Cys Gln
Cys Gln Val Arg Trp Cys Cys Tyr Val 335 340
345gag tgc agg cag tgc acg cag cgt gag gag gtc tac acc tgc aag ggc
1106Glu Cys Arg Gln Cys Thr Gln Arg Glu Glu Val Tyr Thr Cys Lys Gly350
355 360 365tga gttcccaggc
cctgccagcc ctgctgcaca gggtgcaggc attgcacacg 1159gtgtgaaggg
tctacacctg cacaggctga gttcctgggc tcgaccagcc cagctgcgtg 1219gggtacaggc
attgcacaca gtgtgaatgg gtctacacct gcatgggctg agtccctggg 1279ctcagaccta
gcagcgtggg gtagtccctg ggctcagtcc tagctgcatg gggtgcaggc 1339attgcacaga
gcatgaatgg gcctacacct gccaaggctg aatccctggg cccagccagc 1399cctgctgcac
atggcacagg cattgcacac ggtgtgagga gtgtacacct gcaagggctg 1459aggccctggg
cccagtcagc cctgctgctc agagtgcagg cattgcacat ggtgtgagaa 1519ggtctacacc
tgcaagggac gagtccccgg gcctggccaa ccctgctgtg cagggtgagg 1579gccatgcatg
ctagtatgag gggtctacac ctgcaaggac tgagaggctt tt 163128365PRTHomo
sapiens 28Met Leu Asp Gly Ser Pro Leu Ala Arg Trp Leu Ala Ala Ala Phe
Gly1 5 10 15Leu Thr Leu
Leu Leu Ala Ala Leu Arg Pro Ser Ala Ala Tyr Phe Gly 20
25 30Leu Thr Gly Ser Glu Pro Leu Thr Ile Leu
Pro Leu Thr Leu Glu Pro 35 40
45Glu Ala Ala Ala Gln Ala His Tyr Lys Ala Cys Asp Arg Leu Lys Leu 50
55 60Glu Arg Lys Gln Arg Arg Met Cys Arg
Arg Asp Pro Gly Val Ala Glu65 70 75
80Thr Leu Val Glu Ala Val Ser Met Ser Ala Leu Glu Cys Gln
Phe Gln 85 90 95Phe Arg
Phe Glu Arg Trp Asn Cys Thr Leu Glu Gly Arg Tyr Arg Ala 100
105 110Ser Leu Leu Lys Arg Gly Phe Lys Glu
Thr Ala Phe Leu Tyr Ala Ile 115 120
125Ser Ser Ala Gly Leu Thr His Ala Leu Ala Lys Ala Cys Ser Ala Gly
130 135 140Arg Met Glu Arg Cys Thr Cys
Asp Glu Ala Pro Asp Leu Glu Asn Arg145 150
155 160Glu Ala Trp Gln Trp Gly Gly Cys Gly Asp Asn Leu
Lys Tyr Ser Ser 165 170
175Lys Phe Val Lys Glu Phe Leu Gly Arg Arg Ser Ser Lys Asp Leu Arg
180 185 190Ala Arg Val Asp Phe His
Asn Asn Leu Val Gly Val Lys Val Ile Lys 195 200
205Ala Gly Val Glu Thr Thr Cys Lys Cys His Gly Val Ser Gly
Ser Cys 210 215 220Thr Val Arg Thr Cys
Trp Arg Gln Leu Ala Pro Phe His Glu Val Gly225 230
235 240Lys His Leu Lys His Lys Tyr Glu Thr Ala
Leu Lys Val Gly Ser Thr 245 250
255Thr Asn Glu Ala Ala Gly Glu Ala Gly Ala Ile Ser Pro Pro Arg Gly
260 265 270Arg Ala Ser Gly Ala
Gly Gly Ser Asp Pro Leu Pro Arg Thr Pro Glu 275
280 285Leu Val His Leu Asp Asp Ser Pro Ser Phe Cys Leu
Ala Gly Arg Phe 290 295 300Ser Pro Gly
Thr Ala Gly Arg Arg Cys His Arg Glu Lys Asn Cys Glu305
310 315 320Ser Ile Cys Cys Gly Arg Gly
His Asn Thr Gln Ser Arg Val Val Thr 325
330 335Arg Pro Cys Gln Cys Gln Val Arg Trp Cys Cys Tyr
Val Glu Cys Arg 340 345 350Gln
Cys Thr Gln Arg Glu Glu Val Tyr Thr Cys Lys Gly 355
360 365291464DNAHomo sapiensCDS(38)..(1111) 29gcgaggagat
gctagagggc gcagcgccgc cagcacc atg cgc ccc ccg ccc gcg 55
Met Arg Pro Pro Pro Ala
1 5ctg gcc ctg gcc ggg ctc tgc ctg ctg
gcg ctg ccc gcc gcc gcc gcc 103Leu Ala Leu Ala Gly Leu Cys Leu Leu
Ala Leu Pro Ala Ala Ala Ala 10 15
20tcc tac ttc ggc ctg acc ggg cgg gaa gtc ctg acg ccc ttc cca gga
151Ser Tyr Phe Gly Leu Thr Gly Arg Glu Val Leu Thr Pro Phe Pro Gly
25 30 35ttg ggc act gcg gca gcc ccg
gca cag ggc ggg gcc cac ctg aag cag 199Leu Gly Thr Ala Ala Ala Pro
Ala Gln Gly Gly Ala His Leu Lys Gln 40 45
50tgt gac ctg ctg aag ctg tcc cgg cgg cag aag cag ctc tgc cgg agg
247Cys Asp Leu Leu Lys Leu Ser Arg Arg Gln Lys Gln Leu Cys Arg Arg55
60 65 70gag ccc ggc ctg
gct gag acc ctg agg gat gct gcg cac ctc ggc ctg 295Glu Pro Gly Leu
Ala Glu Thr Leu Arg Asp Ala Ala His Leu Gly Leu 75
80 85ctt gag tgc cag ttt cag ttc cgg cat gag
cgc tgg aac tgt agc ctg 343Leu Glu Cys Gln Phe Gln Phe Arg His Glu
Arg Trp Asn Cys Ser Leu 90 95
100gag ggc agg atg ggc ctg ctc aag aga ggc ttc aaa gag aca gct ttc
391Glu Gly Arg Met Gly Leu Leu Lys Arg Gly Phe Lys Glu Thr Ala Phe
105 110 115ctg tac gcg gtg tcc tct gcc
gcc ctc acc cac acc ctg gcc cgg gcc 439Leu Tyr Ala Val Ser Ser Ala
Ala Leu Thr His Thr Leu Ala Arg Ala 120 125
130tgc agc gct ggg cgc atg gag cgc tgc acc tgt gat gac tct ccg ggg
487Cys Ser Ala Gly Arg Met Glu Arg Cys Thr Cys Asp Asp Ser Pro Gly135
140 145 150ctg gag agc cgg
cag gcc tgg cag tgg ggc gtg tgc ggt gac aac ctc 535Leu Glu Ser Arg
Gln Ala Trp Gln Trp Gly Val Cys Gly Asp Asn Leu 155
160 165aag tac agc acc aag ttt ctg agc aac ttc
ctg ggg tcc aag aga gga 583Lys Tyr Ser Thr Lys Phe Leu Ser Asn Phe
Leu Gly Ser Lys Arg Gly 170 175
180aac aag gac ctg cgg gca cgg gca gac gcc cac aat acc cac gtg ggc
631Asn Lys Asp Leu Arg Ala Arg Ala Asp Ala His Asn Thr His Val Gly
185 190 195atc aag gct gtg aag agt ggc
ctc agg acc acg tgt aag tgc cat ggc 679Ile Lys Ala Val Lys Ser Gly
Leu Arg Thr Thr Cys Lys Cys His Gly 200 205
210gta tca ggc tcc tgt gcc gtg cgc acc tgc tgg aag cag ctc tcc ccg
727Val Ser Gly Ser Cys Ala Val Arg Thr Cys Trp Lys Gln Leu Ser Pro215
220 225 230ttc cgt gag acg
ggc cag gtg ctg aaa ctg cgc tat gac tcg gct gtc 775Phe Arg Glu Thr
Gly Gln Val Leu Lys Leu Arg Tyr Asp Ser Ala Val 235
240 245aag gtg tcc agt gcc acc aat gag gcc ttg
ggc cgc cta gag ctg tgg 823Lys Val Ser Ser Ala Thr Asn Glu Ala Leu
Gly Arg Leu Glu Leu Trp 250 255
260gcc cct gcc agg cag ggc agc ctc acc aaa ggc ctg gcc cca agg tct
871Ala Pro Ala Arg Gln Gly Ser Leu Thr Lys Gly Leu Ala Pro Arg Ser
265 270 275ggg gac ctg gtg tac atg gag
gac tca ccc agc ttc tgc cgg ccc agc 919Gly Asp Leu Val Tyr Met Glu
Asp Ser Pro Ser Phe Cys Arg Pro Ser 280 285
290aag tac tca cct ggc aca gca ggt agg gtg tgc tcc cgg gag gcc agc
967Lys Tyr Ser Pro Gly Thr Ala Gly Arg Val Cys Ser Arg Glu Ala Ser295
300 305 310tgc agc agc ctg
tgc tgc ggg cgg ggc tat gac acc cag agc cgc ctg 1015Cys Ser Ser Leu
Cys Cys Gly Arg Gly Tyr Asp Thr Gln Ser Arg Leu 315
320 325gtg gcc ttc tcc tgc cac tgc cag gtg cag
tgg tgc tgc tac gtg gag 1063Val Ala Phe Ser Cys His Cys Gln Val Gln
Trp Cys Cys Tyr Val Glu 330 335
340tgc cag caa tgt gtg cag gag gag ctt gtg tac acc tgc aag cac tag
1111Cys Gln Gln Cys Val Gln Glu Glu Leu Val Tyr Thr Cys Lys His
345 350 355gcctactgcc cagcaagcca
gtctggcact gccaggacct cctgtggcac ccttcaagct 1171gcccagccgg ccctctgggc
agactgtcat cacatgcatg cataaaccgg catgtgtgcc 1231aatgcacacg agtgtgccac
tcaccaccat tccttggcca gccttttgcc tccctcgata 1291ctcaacaaag agaagcaaag
cctcctccct taacccaagc atccccaacc ttgttgagga 1351cttggagagg agggcagagt
gagaaagaca tggagggaaa taagggagac caagagcaca 1411gcaggactga aattttggac
gggagagagg ggctattcca tcttgcttcc tgg 146430357PRTHomo sapiens
30Met Arg Pro Pro Pro Ala Leu Ala Leu Ala Gly Leu Cys Leu Leu Ala1
5 10 15Leu Pro Ala Ala Ala Ala
Ser Tyr Phe Gly Leu Thr Gly Arg Glu Val 20 25
30Leu Thr Pro Phe Pro Gly Leu Gly Thr Ala Ala Ala Pro
Ala Gln Gly 35 40 45Gly Ala His
Leu Lys Gln Cys Asp Leu Leu Lys Leu Ser Arg Arg Gln 50
55 60Lys Gln Leu Cys Arg Arg Glu Pro Gly Leu Ala Glu
Thr Leu Arg Asp65 70 75
80Ala Ala His Leu Gly Leu Leu Glu Cys Gln Phe Gln Phe Arg His Glu
85 90 95Arg Trp Asn Cys Ser Leu
Glu Gly Arg Met Gly Leu Leu Lys Arg Gly 100
105 110Phe Lys Glu Thr Ala Phe Leu Tyr Ala Val Ser Ser
Ala Ala Leu Thr 115 120 125His Thr
Leu Ala Arg Ala Cys Ser Ala Gly Arg Met Glu Arg Cys Thr 130
135 140Cys Asp Asp Ser Pro Gly Leu Glu Ser Arg Gln
Ala Trp Gln Trp Gly145 150 155
160Val Cys Gly Asp Asn Leu Lys Tyr Ser Thr Lys Phe Leu Ser Asn Phe
165 170 175Leu Gly Ser Lys
Arg Gly Asn Lys Asp Leu Arg Ala Arg Ala Asp Ala 180
185 190His Asn Thr His Val Gly Ile Lys Ala Val Lys
Ser Gly Leu Arg Thr 195 200 205Thr
Cys Lys Cys His Gly Val Ser Gly Ser Cys Ala Val Arg Thr Cys 210
215 220Trp Lys Gln Leu Ser Pro Phe Arg Glu Thr
Gly Gln Val Leu Lys Leu225 230 235
240Arg Tyr Asp Ser Ala Val Lys Val Ser Ser Ala Thr Asn Glu Ala
Leu 245 250 255Gly Arg Leu
Glu Leu Trp Ala Pro Ala Arg Gln Gly Ser Leu Thr Lys 260
265 270Gly Leu Ala Pro Arg Ser Gly Asp Leu Val
Tyr Met Glu Asp Ser Pro 275 280
285Ser Phe Cys Arg Pro Ser Lys Tyr Ser Pro Gly Thr Ala Gly Arg Val 290
295 300Cys Ser Arg Glu Ala Ser Cys Ser
Ser Leu Cys Cys Gly Arg Gly Tyr305 310
315 320Asp Thr Gln Ser Arg Leu Val Ala Phe Ser Cys His
Cys Gln Val Gln 325 330
335Trp Cys Cys Tyr Val Glu Cys Gln Gln Cys Val Gln Glu Glu Leu Val
340 345 350Tyr Thr Cys Lys His
355312405DNAHomo sapiensCDS(475)..(1728) 31cccgagccgg gacagtcact
tactctacag gcagtggggc ccgacacaga cagcgccgcc 60cccgccagcc agcctcgcac
gccctcggaa gcgcaggctc ccggcgctgc gctggagggt 120tccccggcac cccagcctcc
cgtccccagc ccgctgcacc tccgggcccc ccttaccctt 180gagaggcacc gggagttgtc
gcgggggggc ctcgggaaat tccccggacc cctgtgccag 240gaggtgcccg gttcgcccgc
tcttcacccc ccgccccccc cgagggcggt gcccgggggt 300gctgccccat ggagcgggga
ggcgggcgcc gtctgctccg ggagccctga cccgagtcgg 360agctgtgtgt cgcagccgcc
ccgacccccc gccgatcatg cgccggcgcc cctggctctc 420cagtcccact gggctgtgag
ccccccactc ccagcccgtc agggcctgcg cgcc atg 477
Met
1ggc agc gcc cac cct cgc ccc tgg ctg cgg ctc cga
ccc cag ccc cag 525Gly Ser Ala His Pro Arg Pro Trp Leu Arg Leu Arg
Pro Gln Pro Gln 5 10 15ccg
cgg cca gcg ctc tgg gtg ctc ctg ttc ttc cta ctg ctg ctg gct 573Pro
Arg Pro Ala Leu Trp Val Leu Leu Phe Phe Leu Leu Leu Leu Ala 20
25 30gct gcc atg ccc agg tca gca ccc aat
gac att ctg gac ctc cgc ctc 621Ala Ala Met Pro Arg Ser Ala Pro Asn
Asp Ile Leu Asp Leu Arg Leu 35 40
45ccc ccg gag ccc gtg ctc aat gcc aac aca gtg tgc cta aca ttg cca
669Pro Pro Glu Pro Val Leu Asn Ala Asn Thr Val Cys Leu Thr Leu Pro50
55 60 65ggc ctg agc cgg cgg
cag atg gag gtg tgt gtg cgt cac cct gat gtg 717Gly Leu Ser Arg Arg
Gln Met Glu Val Cys Val Arg His Pro Asp Val 70
75 80gct gcc tca gcc ata cag ggc atc cag atc gcc
atc cac gaa tgc caa 765Ala Ala Ser Ala Ile Gln Gly Ile Gln Ile Ala
Ile His Glu Cys Gln 85 90
95cac caa ttc agg gac cag cgc tgg aac tgc tca agc ctg gag act cgc
813His Gln Phe Arg Asp Gln Arg Trp Asn Cys Ser Ser Leu Glu Thr Arg
100 105 110aac aag atc ccc tat gag agt
ccc atc ttc agc aga ggt ttc cga gag 861Asn Lys Ile Pro Tyr Glu Ser
Pro Ile Phe Ser Arg Gly Phe Arg Glu 115 120
125agc gct ttt gcc tac gcc atc gca gca gct ggc gtg gtg cac gcc gtg
909Ser Ala Phe Ala Tyr Ala Ile Ala Ala Ala Gly Val Val His Ala Val130
135 140 145tcc aat gcg tgt
gcc ctg ggc aaa ctg aag gcc tgt ggc tgt gat gcg 957Ser Asn Ala Cys
Ala Leu Gly Lys Leu Lys Ala Cys Gly Cys Asp Ala 150
155 160tcc cgg cga ggg gac gag gag gcc ttc cgt
agg aag ctg cac cgc tta 1005Ser Arg Arg Gly Asp Glu Glu Ala Phe Arg
Arg Lys Leu His Arg Leu 165 170
175caa ctg gat gca ctg cag cgt ggt aag ggc ctg agc cat ggg gtc ccg
1053Gln Leu Asp Ala Leu Gln Arg Gly Lys Gly Leu Ser His Gly Val Pro
180 185 190gaa cac cca gcc ctg ccc aca
gcc agc cca ggc ctg cag gac tcc tgg 1101Glu His Pro Ala Leu Pro Thr
Ala Ser Pro Gly Leu Gln Asp Ser Trp 195 200
205gag tgg ggc ggc tgc agc ccc gac atg ggc ttc ggg gag cgc ttt tct
1149Glu Trp Gly Gly Cys Ser Pro Asp Met Gly Phe Gly Glu Arg Phe Ser210
215 220 225aag gac ttt ctg
gac tcc cgg gag cct cac aga gac atc cac gcg aga 1197Lys Asp Phe Leu
Asp Ser Arg Glu Pro His Arg Asp Ile His Ala Arg 230
235 240atg agg ctt cac aac aac cga gtt ggg agg
cag gca gtg atg gag aac 1245Met Arg Leu His Asn Asn Arg Val Gly Arg
Gln Ala Val Met Glu Asn 245 250
255atg cgg cgg aag tgc aag tgc cac ggc acg tca ggc agc tgc cag ctc
1293Met Arg Arg Lys Cys Lys Cys His Gly Thr Ser Gly Ser Cys Gln Leu
260 265 270aag acg tgc tgg cag gtg acg
ccc gag ttc cgc acc gtg ggg gcg ctg 1341Lys Thr Cys Trp Gln Val Thr
Pro Glu Phe Arg Thr Val Gly Ala Leu 275 280
285ctg cgc agc cgc ttc cac cgc gcc acg ctc atc cgg ccg cac aac cgc
1389Leu Arg Ser Arg Phe His Arg Ala Thr Leu Ile Arg Pro His Asn Arg290
295 300 305aac ggc ggc cag
ctg gag ccg ggc cca gcg ggg gca ccc tcg ccg gct 1437Asn Gly Gly Gln
Leu Glu Pro Gly Pro Ala Gly Ala Pro Ser Pro Ala 310
315 320ccg ggc gct ccc ggg ccg cgc cga cgg gcc
agc ccc gcc gac ctg gtc 1485Pro Gly Ala Pro Gly Pro Arg Arg Arg Ala
Ser Pro Ala Asp Leu Val 325 330
335tac ttc gaa aag tct ccc gac ttc tgc gag cgc gag ccg cgc ctg gac
1533Tyr Phe Glu Lys Ser Pro Asp Phe Cys Glu Arg Glu Pro Arg Leu Asp
340 345 350tcg gcg ggc acc gtg ggc cgc
ctg tgc aac aag agc agc gcc ggc tcg 1581Ser Ala Gly Thr Val Gly Arg
Leu Cys Asn Lys Ser Ser Ala Gly Ser 355 360
365gat ggc tgc ggc agc atg tgc tgc ggc cgc ggc cac aac atc ctg cgc
1629Asp Gly Cys Gly Ser Met Cys Cys Gly Arg Gly His Asn Ile Leu Arg370
375 380 385cag acg cgc agc
gag cgc tgc cac tgc cgc ttc cac tgg tgc tgt ttc 1677Gln Thr Arg Ser
Glu Arg Cys His Cys Arg Phe His Trp Cys Cys Phe 390
395 400gtg gtc tgc gaa gag tgc cgc atc acc gag
tgg gtc agc gtc tgc aag 1725Val Val Cys Glu Glu Cys Arg Ile Thr Glu
Trp Val Ser Val Cys Lys 405 410
415tga gcggcccggg gtcccctggg ccctgatcga ggtcccctcc tggagcctgg
1778ccctctgagg cttacggtct tggcaaggca gcatcgcctt ggctcttggg aagaggagat
1838tggaccacat gatcttatag gaacccctca gctctgaggt ctgtgatcgc cggacagtcc
1898aggcctgtct gaaccccacc actcacttct gtgggctcta ggactgactg ggttcttcct
1958ccctccccga agcccagaca gttcagttgg gctgggggtt gctccacacc ctaaaacaag
2018cctcagccag gcaacccgtc agtctgtctc catcctttca ccccttccct ggagatggga
2078ggtggggaat gaatggaagc tgacgggcag agagaggagg attaaaaaaa agaaatagac
2138ataactgagc tgaagtaatt ccataaaggg cccagacagc ctcctccacc attcccttca
2198tcattcattt aacaaatatt tattttgcac tctctttgcg gcactctggg ggcggtgggg
2258tgcgtggggg tggcaatgca aggcactgag gccacagatg tgagtaagcg agacacaaca
2318cttgtcctct tggaggttac attcttgctg gggggaggca tgggcaataa acaagtaaat
2378atacaaacaa aaaaaaaaaa aaaaaaa
240532417PRTHomo sapiens 32Met Gly Ser Ala His Pro Arg Pro Trp Leu Arg
Leu Arg Pro Gln Pro1 5 10
15Gln Pro Arg Pro Ala Leu Trp Val Leu Leu Phe Phe Leu Leu Leu Leu
20 25 30Ala Ala Ala Met Pro Arg Ser
Ala Pro Asn Asp Ile Leu Asp Leu Arg 35 40
45Leu Pro Pro Glu Pro Val Leu Asn Ala Asn Thr Val Cys Leu Thr
Leu 50 55 60Pro Gly Leu Ser Arg Arg
Gln Met Glu Val Cys Val Arg His Pro Asp65 70
75 80Val Ala Ala Ser Ala Ile Gln Gly Ile Gln Ile
Ala Ile His Glu Cys 85 90
95Gln His Gln Phe Arg Asp Gln Arg Trp Asn Cys Ser Ser Leu Glu Thr
100 105 110Arg Asn Lys Ile Pro Tyr
Glu Ser Pro Ile Phe Ser Arg Gly Phe Arg 115 120
125Glu Ser Ala Phe Ala Tyr Ala Ile Ala Ala Ala Gly Val Val
His Ala 130 135 140Val Ser Asn Ala Cys
Ala Leu Gly Lys Leu Lys Ala Cys Gly Cys Asp145 150
155 160Ala Ser Arg Arg Gly Asp Glu Glu Ala Phe
Arg Arg Lys Leu His Arg 165 170
175Leu Gln Leu Asp Ala Leu Gln Arg Gly Lys Gly Leu Ser His Gly Val
180 185 190Pro Glu His Pro Ala
Leu Pro Thr Ala Ser Pro Gly Leu Gln Asp Ser 195
200 205Trp Glu Trp Gly Gly Cys Ser Pro Asp Met Gly Phe
Gly Glu Arg Phe 210 215 220Ser Lys Asp
Phe Leu Asp Ser Arg Glu Pro His Arg Asp Ile His Ala225
230 235 240Arg Met Arg Leu His Asn Asn
Arg Val Gly Arg Gln Ala Val Met Glu 245
250 255Asn Met Arg Arg Lys Cys Lys Cys His Gly Thr Ser
Gly Ser Cys Gln 260 265 270Leu
Lys Thr Cys Trp Gln Val Thr Pro Glu Phe Arg Thr Val Gly Ala 275
280 285Leu Leu Arg Ser Arg Phe His Arg Ala
Thr Leu Ile Arg Pro His Asn 290 295
300Arg Asn Gly Gly Gln Leu Glu Pro Gly Pro Ala Gly Ala Pro Ser Pro305
310 315 320Ala Pro Gly Ala
Pro Gly Pro Arg Arg Arg Ala Ser Pro Ala Asp Leu 325
330 335Val Tyr Phe Glu Lys Ser Pro Asp Phe Cys
Glu Arg Glu Pro Arg Leu 340 345
350Asp Ser Ala Gly Thr Val Gly Arg Leu Cys Asn Lys Ser Ser Ala Gly
355 360 365Ser Asp Gly Cys Gly Ser Met
Cys Cys Gly Arg Gly His Asn Ile Leu 370 375
380Arg Gln Thr Arg Ser Glu Arg Cys His Cys Arg Phe His Trp Cys
Cys385 390 395 400Phe Val
Val Cys Glu Glu Cys Arg Ile Thr Glu Trp Val Ser Val Cys
405 410 415Lys332288DNAHomo
sapiensCDS(347)..(1516) 33ggggctgcag ctccgtcagc ccggcagagc caccctgagc
tcggtgagag caaagccaga 60gcccccagtc ctttgctcgc cggcttgcta tctctctcga
tcactccctc ccttcctccc 120tcccttcctc ccggcggccg cggcggcgct ggggaagcgg
tgaagaggag tggcccggcc 180ctggaagaat gcggctctga caaggggaca gaacccagcg
cagtctcccc acggtttaag 240cagcactagt gaagcccagg caacccaacc gtgcctgtct
cggaccccgc acccaaacca 300ctggaggtcc tgatcgatct gcccaccgga gcctccgggc
ttcgac atg ctg gag 355
Met Leu Glu 1gag
ccc cgg ccg cgg cct ccg ccc tcg ggc ctc gcg ggt ctc ctg ttc 403Glu
Pro Arg Pro Arg Pro Pro Pro Ser Gly Leu Ala Gly Leu Leu Phe 5
10 15ctg gcg ttg tgc agt cgg gct cta agc aat
gag att ctg ggc ctg aag 451Leu Ala Leu Cys Ser Arg Ala Leu Ser Asn
Glu Ile Leu Gly Leu Lys20 25 30
35ttg cct ggc gag ccg ccg ctg acg gcc aac acc gtg tgc ttg acg
ctg 499Leu Pro Gly Glu Pro Pro Leu Thr Ala Asn Thr Val Cys Leu Thr
Leu 40 45 50tcc ggc ctg
agc aag cgg cag cta ggc ctg tgc ctg cgc aac ccc gac 547Ser Gly Leu
Ser Lys Arg Gln Leu Gly Leu Cys Leu Arg Asn Pro Asp 55
60 65gtg acg gcg tcc gcg ctt cag ggt ctg cac
atc gcg gtc cac gag tgt 595Val Thr Ala Ser Ala Leu Gln Gly Leu His
Ile Ala Val His Glu Cys 70 75
80cag cac cag ctg cgc gac cag cgc tgg aac tgc tcc gcg ctt gag ggc
643Gln His Gln Leu Arg Asp Gln Arg Trp Asn Cys Ser Ala Leu Glu Gly 85
90 95ggc ggc cgc ctg ccg cac cac agc gcc
atc ctc aag cgc ggt ttc cga 691Gly Gly Arg Leu Pro His His Ser Ala
Ile Leu Lys Arg Gly Phe Arg100 105 110
115gaa agt gct ttt tcc ttc tcc atg ctg gct gct ggg gtc atg
cac gca 739Glu Ser Ala Phe Ser Phe Ser Met Leu Ala Ala Gly Val Met
His Ala 120 125 130gta gcc
acg gcc tgc agc ctg ggc aag ctg gtg agc tgt ggc tgt ggc 787Val Ala
Thr Ala Cys Ser Leu Gly Lys Leu Val Ser Cys Gly Cys Gly 135
140 145tgg aag ggc agt ggt gag cag gat cgg
ctg agg gcc aaa ctg ctg cag 835Trp Lys Gly Ser Gly Glu Gln Asp Arg
Leu Arg Ala Lys Leu Leu Gln 150 155
160ctg cag gca ctg tcc cga ggc aag agt ttc ccc cac tct ctg ccc agc
883Leu Gln Ala Leu Ser Arg Gly Lys Ser Phe Pro His Ser Leu Pro Ser 165
170 175cct ggc cct ggc tca agc ccc agc
cct ggc ccc cag gac aca tgg gaa 931Pro Gly Pro Gly Ser Ser Pro Ser
Pro Gly Pro Gln Asp Thr Trp Glu180 185
190 195tgg ggt ggc tgt aac cat gac atg gac ttt gga gag
aag ttc tct cgg 979Trp Gly Gly Cys Asn His Asp Met Asp Phe Gly Glu
Lys Phe Ser Arg 200 205
210gat ttc ttg gat tcc agg gaa gct ccc cgg gac atc cag gca cga atg
1027Asp Phe Leu Asp Ser Arg Glu Ala Pro Arg Asp Ile Gln Ala Arg Met
215 220 225cga atc cac aac aac agg
gtg ggg cgc cag gtg gta act gaa aac ctg 1075Arg Ile His Asn Asn Arg
Val Gly Arg Gln Val Val Thr Glu Asn Leu 230 235
240aag cgg aaa tgc aag tgt cat ggc aca tca ggc agc tgc cag
ttc aag 1123Lys Arg Lys Cys Lys Cys His Gly Thr Ser Gly Ser Cys Gln
Phe Lys 245 250 255aca tgc tgg agg gcg
gcc cca gag ttc cgg gca gtg ggg gcg gcg ttg 1171Thr Cys Trp Arg Ala
Ala Pro Glu Phe Arg Ala Val Gly Ala Ala Leu260 265
270 275agg gag cgg ctg ggc cgg gcc atc ttc att
gat acc cac aac cgc aat 1219Arg Glu Arg Leu Gly Arg Ala Ile Phe Ile
Asp Thr His Asn Arg Asn 280 285
290tct gga gcc ttc cag ccc cgt ctg cgt ccc cgt cgc ctc tca gga gag
1267Ser Gly Ala Phe Gln Pro Arg Leu Arg Pro Arg Arg Leu Ser Gly Glu
295 300 305ctg gtc tac ttt gag aag
tct cct gac ttc tgt gag cga gac ccc act 1315Leu Val Tyr Phe Glu Lys
Ser Pro Asp Phe Cys Glu Arg Asp Pro Thr 310 315
320atg ggc tcc cca ggg aca agg ggc cgg gcc tgc aac aag acc
agc cgc 1363Met Gly Ser Pro Gly Thr Arg Gly Arg Ala Cys Asn Lys Thr
Ser Arg 325 330 335ctg ttg gat ggc tgt
ggc agc ctg tgc tgt ggc cgt ggg cac aac gtg 1411Leu Leu Asp Gly Cys
Gly Ser Leu Cys Cys Gly Arg Gly His Asn Val340 345
350 355ctc cgg cag aca cga gtt gag cgc tgc cat
tgc cgc ttc cac tgg tgc 1459Leu Arg Gln Thr Arg Val Glu Arg Cys His
Cys Arg Phe His Trp Cys 360 365
370tgc tat gtg ctg tgt gat gag tgc aag gtt aca gag tgg gtg aat gtg
1507Cys Tyr Val Leu Cys Asp Glu Cys Lys Val Thr Glu Trp Val Asn Val
375 380 385tgt aag tga gggtcagcct
taccttgggg ctggggaaga ggactgtgtg 1556Cys Lys 390agaggggcgc
cttttcagcc ctttgctctg atttccttcc aaggtcactc ttggtccctg 1616gaagcttaaa
gtatctacct ggaaacagct ttaggggtgg tgggggtcag gtggactctg 1676ggatgtgtag
ccttctcccc aacaattgga gggtcttgag gggaagctgc cacccctctt 1736ctgctcctta
gacacctgaa tggactaaga tgaaatgcac tgtattgctc ctcccacttc 1796tcaactccag
agccccttta accctgattc atactccttt tggctgggga gtccctatag 1856tttcaccact
cctctccctt gagggataac cccaggcact gtttggagcc ataagatctg 1916tatctagaaa
gagatcaccc actcctatgt actatcccca aactccttta ctgcagcctg 1976ggctccctct
tgtgggataa tgggagacag tggtagagag gtttttcttg ggaaagagac 2036agagtgctga
ggggcactct cccctgaatc ctcagagagt tgtctgtcca ggcccttagg 2096gaagttgtct
ccttccattc agatgttaat ggggaccctc caaaggaagg ggttttccca 2156tgactcttgg
agcctctttt tccttcttca gcaggaaggg tgggaaggga taatttatca 2216tactgagact
tgttcttggt tcctgtttga aactaaaata aattaagtta ctggaaaaaa 2276aaaaaaaaaa
aa 228834389PRTHomo
sapiens 34Met Leu Glu Glu Pro Arg Pro Arg Pro Pro Pro Ser Gly Leu Ala
Gly1 5 10 15Leu Leu Phe
Leu Ala Leu Cys Ser Arg Ala Leu Ser Asn Glu Ile Leu 20
25 30Gly Leu Lys Leu Pro Gly Glu Pro Pro Leu
Thr Ala Asn Thr Val Cys 35 40
45Leu Thr Leu Ser Gly Leu Ser Lys Arg Gln Leu Gly Leu Cys Leu Arg 50
55 60Asn Pro Asp Val Thr Ala Ser Ala Leu
Gln Gly Leu His Ile Ala Val65 70 75
80His Glu Cys Gln His Gln Leu Arg Asp Gln Arg Trp Asn Cys
Ser Ala 85 90 95Leu Glu
Gly Gly Gly Arg Leu Pro His His Ser Ala Ile Leu Lys Arg 100
105 110Gly Phe Arg Glu Ser Ala Phe Ser Phe
Ser Met Leu Ala Ala Gly Val 115 120
125Met His Ala Val Ala Thr Ala Cys Ser Leu Gly Lys Leu Val Ser Cys
130 135 140Gly Cys Gly Trp Lys Gly Ser
Gly Glu Gln Asp Arg Leu Arg Ala Lys145 150
155 160Leu Leu Gln Leu Gln Ala Leu Ser Arg Gly Lys Ser
Phe Pro His Ser 165 170
175Leu Pro Ser Pro Gly Pro Gly Ser Ser Pro Ser Pro Gly Pro Gln Asp
180 185 190Thr Trp Glu Trp Gly Gly
Cys Asn His Asp Met Asp Phe Gly Glu Lys 195 200
205Phe Ser Arg Asp Phe Leu Asp Ser Arg Glu Ala Pro Arg Asp
Ile Gln 210 215 220Ala Arg Met Arg Ile
His Asn Asn Arg Val Gly Arg Gln Val Val Thr225 230
235 240Glu Asn Leu Lys Arg Lys Cys Lys Cys His
Gly Thr Ser Gly Ser Cys 245 250
255Gln Phe Lys Thr Cys Trp Arg Ala Ala Pro Glu Phe Arg Ala Val Gly
260 265 270Ala Ala Leu Arg Glu
Arg Leu Gly Arg Ala Ile Phe Ile Asp Thr His 275
280 285Asn Arg Asn Ser Gly Ala Phe Gln Pro Arg Leu Arg
Pro Arg Arg Leu 290 295 300Ser Gly Glu
Leu Val Tyr Phe Glu Lys Ser Pro Asp Phe Cys Glu Arg305
310 315 320Asp Pro Thr Met Gly Ser Pro
Gly Thr Arg Gly Arg Ala Cys Asn Lys 325
330 335Thr Ser Arg Leu Leu Asp Gly Cys Gly Ser Leu Cys
Cys Gly Arg Gly 340 345 350His
Asn Val Leu Arg Gln Thr Arg Val Glu Arg Cys His Cys Arg Phe 355
360 365His Trp Cys Cys Tyr Val Leu Cys Asp
Glu Cys Lys Val Thr Glu Trp 370 375
380Val Asn Val Cys Lys385351927DNAHomo sapiensCDS(124)..(1188)
35taacccgccg cctccgctct ccccggctgc aggcggcgtg caggaccagc ggcggccgtg
60caggcggagg acttcggcgc ggctcctcct gggtgtgacc ccgggcgcgc ccgccgcgcg
120acg atg agg gcg cgg ccg cag gtc tgc gag gcg ctg ctc ttc gcc ctg
168 Met Arg Ala Arg Pro Gln Val Cys Glu Ala Leu Leu Phe Ala Leu 1
5 10 15gcg ctc cag acc ggc
gtg tgc tat ggc atc aag tgg ctg gcg ctg tcc 216Ala Leu Gln Thr Gly
Val Cys Tyr Gly Ile Lys Trp Leu Ala Leu Ser 20
25 30aag aca cca tcg gcc ctg gca ctg aac cag acg
caa cac tgc aag cag 264Lys Thr Pro Ser Ala Leu Ala Leu Asn Gln Thr
Gln His Cys Lys Gln 35 40
45ctg gag ggt ctg gtg tct gca cag gtg cag ctg tgc cgc agc aac ctg
312Leu Glu Gly Leu Val Ser Ala Gln Val Gln Leu Cys Arg Ser Asn Leu
50 55 60gag ctc atg cac acg gtg gtg cac
gcc gcc cgc gag gtc atg aag gcc 360Glu Leu Met His Thr Val Val His
Ala Ala Arg Glu Val Met Lys Ala 65 70
75tgt cgc cgg gcc ttt gcc gac atg cgc tgg aac tgc tcc tcc att gag
408Cys Arg Arg Ala Phe Ala Asp Met Arg Trp Asn Cys Ser Ser Ile Glu80
85 90 95ctc gcc ccc aac tat
ttg ctt gac ctg gag aga ggg acc cgg gag tcg 456Leu Ala Pro Asn Tyr
Leu Leu Asp Leu Glu Arg Gly Thr Arg Glu Ser 100
105 110gcc ttc gtg tat gcg ctg tcg gcc gcc gcc atc
agc cac gcc atc gcc 504Ala Phe Val Tyr Ala Leu Ser Ala Ala Ala Ile
Ser His Ala Ile Ala 115 120
125cgg gcc tgc acc tcc ggc gac ctg ccc ggc tgc tcc tgc ggc ccc gtc
552Arg Ala Cys Thr Ser Gly Asp Leu Pro Gly Cys Ser Cys Gly Pro Val
130 135 140cca ggt gag cca ccc ggg ccc
ggg aac cgc tgg gga gga tgt gcg gac 600Pro Gly Glu Pro Pro Gly Pro
Gly Asn Arg Trp Gly Gly Cys Ala Asp 145 150
155aac ctc agc tac ggg ctc ctc atg ggg gcc aag ttt tcc gat gct cct
648Asn Leu Ser Tyr Gly Leu Leu Met Gly Ala Lys Phe Ser Asp Ala Pro160
165 170 175atg aag gtg aaa
aaa aca gga tcc caa gcc aat aaa ctg atg cgt cta 696Met Lys Val Lys
Lys Thr Gly Ser Gln Ala Asn Lys Leu Met Arg Leu 180
185 190cac aac agt gaa gtg ggg aga cag gct ctg
cgc gcc tct ctg gaa atg 744His Asn Ser Glu Val Gly Arg Gln Ala Leu
Arg Ala Ser Leu Glu Met 195 200
205aag tgt aag tgc cat ggg gtg tct ggc tcc tgc tcc atc cgc acc tgc
792Lys Cys Lys Cys His Gly Val Ser Gly Ser Cys Ser Ile Arg Thr Cys
210 215 220tgg aag ggg ctg cag gag ctg
cag gat gtg gct gct gac ctc aag acc 840Trp Lys Gly Leu Gln Glu Leu
Gln Asp Val Ala Ala Asp Leu Lys Thr 225 230
235cga tac ctg tcg gcc acc aag gta gtg cac cga ccc atg ggc acc cgc
888Arg Tyr Leu Ser Ala Thr Lys Val Val His Arg Pro Met Gly Thr Arg240
245 250 255aag cac ctg gtg
ccc aag gac ctg gat atc cgg cct gtg aag gac tcg 936Lys His Leu Val
Pro Lys Asp Leu Asp Ile Arg Pro Val Lys Asp Ser 260
265 270gaa ctc gtc tat ctg cag agc tca cct gac
ttc tgc atg aag aat gag 984Glu Leu Val Tyr Leu Gln Ser Ser Pro Asp
Phe Cys Met Lys Asn Glu 275 280
285aag gtg ggc tcc cac ggg aca caa gac agg cag tgc aac aag aca tcc
1032Lys Val Gly Ser His Gly Thr Gln Asp Arg Gln Cys Asn Lys Thr Ser
290 295 300aac gga agc gac agc tgc gac
ctt atg tgc tgc ggg cgt ggc tac aac 1080Asn Gly Ser Asp Ser Cys Asp
Leu Met Cys Cys Gly Arg Gly Tyr Asn 305 310
315ccc tac aca gac cgc gtg gtc gag cgg tgc cac tgt aag tac cac tgg
1128Pro Tyr Thr Asp Arg Val Val Glu Arg Cys His Cys Lys Tyr His Trp320
325 330 335tgc tgc tac gtc
acc tgc cgc agg tgt gag cgt acc gtg gag cgc tat 1176Cys Cys Tyr Val
Thr Cys Arg Arg Cys Glu Arg Thr Val Glu Arg Tyr 340
345 350gtc tgc aag tga ggccctgccc tccgccccac
gcaggagcga ggactctgct 1228Val Cys Lys 355caaggaccct
cagcaactgg ggccaggggc ctggagacac tccatggagc tctgcttgtg 1288aattccagat
gccaggcatg ggaggcggct tgtgctttgc cttcacttgg aagccaccag 1348gaacagaagg
tctggccacc ctggaaggag ggcaggacat caaaggaaac cgacaagatt 1408aaaaataact
tggcagcctg aggctctgga gtgcccacag gctggtgtaa ggagcggggc 1468ttgggatcgg
tgagactgat acagacttga cctttcaggg ccacagagac cagcctccgg 1528gaaggggtct
gcccgccttc ttcagaatgt tctgcgggac cccctggccc accctggggt 1588ctgagcctgc
tgggcccacc acatggaatc actagcttgg gttgtaaatg ttttcttttg 1648ttttttgctt
tttcttcctt tgggatgtgg aagctacaga aatatttata aaacatagct 1708ttttctttgg
ggtggcactt ctcaattcct ctttatatat tttatatata taaatatata 1768tgtatatata
taatgatctc tattttaaaa ctagcttttt aagcagctgt atgaaataaa 1828tgctgagtga
gccccagccc gcccctgcag ttcccggcct cgtcaagtga actcggcaga 1888ccctggggct
ggcagaggga gctctccagt ttccaggca 192736354PRTHomo
sapiens 36Met Arg Ala Arg Pro Gln Val Cys Glu Ala Leu Leu Phe Ala Leu
Ala1 5 10 15Leu Gln Thr
Gly Val Cys Tyr Gly Ile Lys Trp Leu Ala Leu Ser Lys 20
25 30Thr Pro Ser Ala Leu Ala Leu Asn Gln Thr
Gln His Cys Lys Gln Leu 35 40
45Glu Gly Leu Val Ser Ala Gln Val Gln Leu Cys Arg Ser Asn Leu Glu 50
55 60Leu Met His Thr Val Val His Ala Ala
Arg Glu Val Met Lys Ala Cys65 70 75
80Arg Arg Ala Phe Ala Asp Met Arg Trp Asn Cys Ser Ser Ile
Glu Leu 85 90 95Ala Pro
Asn Tyr Leu Leu Asp Leu Glu Arg Gly Thr Arg Glu Ser Ala 100
105 110Phe Val Tyr Ala Leu Ser Ala Ala Ala
Ile Ser His Ala Ile Ala Arg 115 120
125Ala Cys Thr Ser Gly Asp Leu Pro Gly Cys Ser Cys Gly Pro Val Pro
130 135 140Gly Glu Pro Pro Gly Pro Gly
Asn Arg Trp Gly Gly Cys Ala Asp Asn145 150
155 160Leu Ser Tyr Gly Leu Leu Met Gly Ala Lys Phe Ser
Asp Ala Pro Met 165 170
175Lys Val Lys Lys Thr Gly Ser Gln Ala Asn Lys Leu Met Arg Leu His
180 185 190Asn Ser Glu Val Gly Arg
Gln Ala Leu Arg Ala Ser Leu Glu Met Lys 195 200
205Cys Lys Cys His Gly Val Ser Gly Ser Cys Ser Ile Arg Thr
Cys Trp 210 215 220Lys Gly Leu Gln Glu
Leu Gln Asp Val Ala Ala Asp Leu Lys Thr Arg225 230
235 240Tyr Leu Ser Ala Thr Lys Val Val His Arg
Pro Met Gly Thr Arg Lys 245 250
255His Leu Val Pro Lys Asp Leu Asp Ile Arg Pro Val Lys Asp Ser Glu
260 265 270Leu Val Tyr Leu Gln
Ser Ser Pro Asp Phe Cys Met Lys Asn Glu Lys 275
280 285Val Gly Ser His Gly Thr Gln Asp Arg Gln Cys Asn
Lys Thr Ser Asn 290 295 300Gly Ser Asp
Ser Cys Asp Leu Met Cys Cys Gly Arg Gly Tyr Asn Pro305
310 315 320Tyr Thr Asp Arg Val Val Glu
Arg Cys His Cys Lys Tyr His Trp Cys 325
330 335Cys Tyr Val Thr Cys Arg Arg Cys Glu Arg Thr Val
Glu Arg Tyr Val 340 345 350Cys
Lys373132DNAHomo sapiensCDS(258)..(1355) 37cccgcatctc ctgcacatct
ccacccctgc gcaggaggag atccccaggc tgctctctcc 60atctctccta cagctccctg
caaacgaggg ggaagctgct gagagtccct atcactgctg 120gccttttaat gttgtatgca
aggaggaaga gggcgaggga taacttggtg ctggacaact 180gacctgcggc ccgaagggcc
tctggggagg gggtgcaaaa gaggagcggc tgggctgggg 240gactccatgc gggggcg atg
gac agg gcg gcg ctc ctg gga ctg gcc cgc 290 Met
Asp Arg Ala Ala Leu Leu Gly Leu Ala Arg 1
5 10ttg tgc gcg ctg tgg gca gcc ctg ctc gtg ctg ttc ccc
tac gga gcc 338Leu Cys Ala Leu Trp Ala Ala Leu Leu Val Leu Phe Pro
Tyr Gly Ala 15 20 25caa gga
aac tgg atg tgg ttg ggc att gcc tcc ttc ggg gtt cca gag 386Gln Gly
Asn Trp Met Trp Leu Gly Ile Ala Ser Phe Gly Val Pro Glu 30
35 40aag ctg ggc tgc gcc aat ttg ccg ctg aac
agc cgc cag aag gag ctg 434Lys Leu Gly Cys Ala Asn Leu Pro Leu Asn
Ser Arg Gln Lys Glu Leu 45 50 55tgc
aag agg aaa ccg tac ctg ctg ccg agc atc cga gag ggc gcc cgg 482Cys
Lys Arg Lys Pro Tyr Leu Leu Pro Ser Ile Arg Glu Gly Ala Arg60
65 70 75ctg ggc att cag gag tgc
ggg agc cag ttc aga cac gag aga tgg aac 530Leu Gly Ile Gln Glu Cys
Gly Ser Gln Phe Arg His Glu Arg Trp Asn 80
85 90tgc atg atc acc gcc gcc gcc act acc gcc ccg atg
ggc gcc agc ccc 578Cys Met Ile Thr Ala Ala Ala Thr Thr Ala Pro Met
Gly Ala Ser Pro 95 100 105ctc
ttt ggc tac gag ctg agc agc ggc acc aaa gag aca gca ttt att 626Leu
Phe Gly Tyr Glu Leu Ser Ser Gly Thr Lys Glu Thr Ala Phe Ile 110
115 120tat gct gtg atg gct gca ggc ctg gtg
cat tct gtg acc agg tca tgc 674Tyr Ala Val Met Ala Ala Gly Leu Val
His Ser Val Thr Arg Ser Cys 125 130
135agt gca ggc aac atg aca gag tgt tcc tgt gac acc acc ttg cag aac
722Ser Ala Gly Asn Met Thr Glu Cys Ser Cys Asp Thr Thr Leu Gln Asn140
145 150 155ggc ggc tca gca
agt gaa ggc tgg cac tgg ggg ggc tgc tcc gat gat 770Gly Gly Ser Ala
Ser Glu Gly Trp His Trp Gly Gly Cys Ser Asp Asp 160
165 170gtc cag tat ggc atg tgg ttc agc aga aag
ttc cta gat ttc ccc atc 818Val Gln Tyr Gly Met Trp Phe Ser Arg Lys
Phe Leu Asp Phe Pro Ile 175 180
185gga aac acc acg ggc aaa gaa aac aaa gta cta tta gca atg aac cta
866Gly Asn Thr Thr Gly Lys Glu Asn Lys Val Leu Leu Ala Met Asn Leu
190 195 200cat aac aat gaa gct gga agg
cag gct gtc gcc aag ttg atg tca gta 914His Asn Asn Glu Ala Gly Arg
Gln Ala Val Ala Lys Leu Met Ser Val 205 210
215gac tgc cgc tgc cac gga gtt tcc ggc tcc tgt gct gtg aaa aca tgc
962Asp Cys Arg Cys His Gly Val Ser Gly Ser Cys Ala Val Lys Thr Cys220
225 230 235tgg aaa acc atg
tct tct ttt gaa aag att ggc cat ttg ttg aag gat 1010Trp Lys Thr Met
Ser Ser Phe Glu Lys Ile Gly His Leu Leu Lys Asp 240
245 250aaa tat gaa aac agt atc cag ata tca gac
aaa aca aag agg aaa atg 1058Lys Tyr Glu Asn Ser Ile Gln Ile Ser Asp
Lys Thr Lys Arg Lys Met 255 260
265cgc agg aga gaa aaa gat cag agg aaa ata cca atc cat aag gat gat
1106Arg Arg Arg Glu Lys Asp Gln Arg Lys Ile Pro Ile His Lys Asp Asp
270 275 280ctg ctc tat gtt aat aag tct
ccc aac tac tgt gta gaa gat aag aaa 1154Leu Leu Tyr Val Asn Lys Ser
Pro Asn Tyr Cys Val Glu Asp Lys Lys 285 290
295ctg gga atc cca ggg aca caa ggc aga gaa tgc aac cgt aca tca gag
1202Leu Gly Ile Pro Gly Thr Gln Gly Arg Glu Cys Asn Arg Thr Ser Glu300
305 310 315ggt gca gat ggc
tgc aac ctc ctc tgc tgt ggc cga ggt tac aac acc 1250Gly Ala Asp Gly
Cys Asn Leu Leu Cys Cys Gly Arg Gly Tyr Asn Thr 320
325 330cat gtg gtc agg cac gtg gag agg tgt gag
tgt aag ttc atc tgg tgc 1298His Val Val Arg His Val Glu Arg Cys Glu
Cys Lys Phe Ile Trp Cys 335 340
345tgc tat gtc cgt tgc agg agg tgt gaa agc atg act gat gtc cac act
1346Cys Tyr Val Arg Cys Arg Arg Cys Glu Ser Met Thr Asp Val His Thr
350 355 360tgc aag taa ccactccatc
cagccttggg caagatgcct cagcaatata 1395Cys Lys 365caatggcatt
gcaaccagag aggtgcccat ccctgtgcag cgctagtaaa gttgactctt 1455gcagtggaat
ccctagaacc ttggacctga gagtttccct tacctgatcg acatattttc 1515ctttatctga
tcaacccatc aatcatgtgg atttcttggg attctaatgt tgaaaaggtt 1575tatattcacc
ttttgatgat ttggggaata tatattgaca tacaaggaag ataatctgtt 1635tcctaagcaa
gaaataacag gaaagatccc ttatgccagg aggcctgcca tactcaggat 1695aagatccttg
aatatggaac ttagttacag gactcaataa tggtgggtga acattagtca 1755tttttaaaag
acacctctta tagcaataag gagacattaa catgaatctc atttattctc 1815tcagtatttt
aactgaagaa attatactgt ttgtgtgtgg atagaagatg ttgaaaagtt 1875aacataagca
ttgggtgctg acttaccctt tcatgtactt ccaaagaaag gtaatcaaaa 1935agaatcttct
taagtgatat aatatcccta aaaaaatgat cattacagat gtttagtgac 1995aaagaatcaa
tatgtaaaaa gtataatgaa tgatttagat tttaagtgcc ttttcactgg 2055gagaatctgg
aaaaacctcc ataaggtata tagcaatctt tgatctttag attcatactt 2115ttatcacaga
tcagtttcaa ctgttaaaaa cccacctctg agatactggg gggaggatcc 2175tgaaacatgc
gggaaaagga gaggtaaaca gtggaggtaa aaatataatt tcatacattg 2235taaagaaaag
caccctttaa atgtgtaaag acagtgtttt gtaaagaatt ttgtttaaaa 2295agtttctatt
ttgtaaatac agtacttaag ttatatgatt tatattaaaa catttattga 2355caaagcctaa
gagctaaggc agtaaaatta tctcataaat aatattagct tatttttttt 2415catactatta
atgctatttt tttggacatc gaagagaatt taacttagca gttagttata 2475tggatgtgta
tttcttgcta aaatgacagt tttatatgtt atagattaaa atatgttgca 2535aaatatcaaa
aatttgtgtt atttcagcag taagattaat tgaattctct tttcacatta 2595gttatgctta
actcataagg ttattataat aaattatatt agtaaaagtc ttaactggaa 2655aaaagaatct
aaatcagaat agtgatcaat ttgtggattt gatatcctgg atatttatta 2715tattttatgt
aatgctgcat ttctatttga atgttaagtg gtctttcttg tttttaatat 2775tcatgcatgt
atattcatca tattttacaa ggttcctggt aaaaattaca gggctctatt 2835taaggatgta
ttttaatgta aatgcttatg ttttttatga attgttaaat atttcagtat 2895tatatagaaa
aaaatagatt tttaaaattc agaatggaca aagagaatat tcattttctt 2955attaataaga
taaagaaatg tttccctgcc ccacagtctt cattctattt ctctttaatt 3015ttattcactg
aggcagagaa acaatttttg aaaaagagca aacccatgga aaatgtctca 3075gatctaatat
taaaatcaag actaagcatt taactgtgaa aaaaaaaaaa aaaaaaa 313238365PRTHomo
sapiens 38Met Asp Arg Ala Ala Leu Leu Gly Leu Ala Arg Leu Cys Ala Leu
Trp1 5 10 15Ala Ala Leu
Leu Val Leu Phe Pro Tyr Gly Ala Gln Gly Asn Trp Met 20
25 30Trp Leu Gly Ile Ala Ser Phe Gly Val Pro
Glu Lys Leu Gly Cys Ala 35 40
45Asn Leu Pro Leu Asn Ser Arg Gln Lys Glu Leu Cys Lys Arg Lys Pro 50
55 60Tyr Leu Leu Pro Ser Ile Arg Glu Gly
Ala Arg Leu Gly Ile Gln Glu65 70 75
80Cys Gly Ser Gln Phe Arg His Glu Arg Trp Asn Cys Met Ile
Thr Ala 85 90 95Ala Ala
Thr Thr Ala Pro Met Gly Ala Ser Pro Leu Phe Gly Tyr Glu 100
105 110Leu Ser Ser Gly Thr Lys Glu Thr Ala
Phe Ile Tyr Ala Val Met Ala 115 120
125Ala Gly Leu Val His Ser Val Thr Arg Ser Cys Ser Ala Gly Asn Met
130 135 140Thr Glu Cys Ser Cys Asp Thr
Thr Leu Gln Asn Gly Gly Ser Ala Ser145 150
155 160Glu Gly Trp His Trp Gly Gly Cys Ser Asp Asp Val
Gln Tyr Gly Met 165 170
175Trp Phe Ser Arg Lys Phe Leu Asp Phe Pro Ile Gly Asn Thr Thr Gly
180 185 190Lys Glu Asn Lys Val Leu
Leu Ala Met Asn Leu His Asn Asn Glu Ala 195 200
205Gly Arg Gln Ala Val Ala Lys Leu Met Ser Val Asp Cys Arg
Cys His 210 215 220Gly Val Ser Gly Ser
Cys Ala Val Lys Thr Cys Trp Lys Thr Met Ser225 230
235 240Ser Phe Glu Lys Ile Gly His Leu Leu Lys
Asp Lys Tyr Glu Asn Ser 245 250
255Ile Gln Ile Ser Asp Lys Thr Lys Arg Lys Met Arg Arg Arg Glu Lys
260 265 270Asp Gln Arg Lys Ile
Pro Ile His Lys Asp Asp Leu Leu Tyr Val Asn 275
280 285Lys Ser Pro Asn Tyr Cys Val Glu Asp Lys Lys Leu
Gly Ile Pro Gly 290 295 300Thr Gln Gly
Arg Glu Cys Asn Arg Thr Ser Glu Gly Ala Asp Gly Cys305
310 315 320Asn Leu Leu Cys Cys Gly Arg
Gly Tyr Asn Thr His Val Val Arg His 325
330 335Val Glu Arg Cys Glu Cys Lys Phe Ile Trp Cys Cys
Tyr Val Arg Cys 340 345 350Arg
Arg Cys Glu Ser Met Thr Asp Val His Thr Cys Lys 355
360 365393364DNAMus musculusCDS(18)..(803) 39gcggccgccg
ccccgcg atg gcc ccg cag caa ggc cgg ccg gcg ctg ccc 50
Met Ala Pro Gln Gln Gly Arg Pro Ala Leu Pro 1
5 10gcc cgc tgc gag ccg ccg gcg gcg ccg ccg
gta ccg cct cgc cga gag 98Ala Arg Cys Glu Pro Pro Ala Ala Pro Pro
Val Pro Pro Arg Arg Glu 15 20
25cgc ggg ggg cgc ggg gcg cgc ggg ccc ggg gtg tcc ggg ggt cgg ggg
146Arg Gly Gly Arg Gly Ala Arg Gly Pro Gly Val Ser Gly Gly Arg Gly
30 35 40cgc gcg ggc ggc gcc gag gga cgc
ggc gtc aag tgc gtg ctg gtc ggc 194Arg Ala Gly Gly Ala Glu Gly Arg
Gly Val Lys Cys Val Leu Val Gly 45 50
55gac ggc gcg gtg ggc aag acc agc ctg gtg gtc agc tac acc act aac
242Asp Gly Ala Val Gly Lys Thr Ser Leu Val Val Ser Tyr Thr Thr Asn60
65 70 75ggc tac ccc acc gag
tac atc cct acg gcc ttc gac aac ttc tcg gcc 290Gly Tyr Pro Thr Glu
Tyr Ile Pro Thr Ala Phe Asp Asn Phe Ser Ala 80
85 90gtg gtg tct gta gat ggg cgg cct gtg aga ctc
cag ctc tgt gac act 338Val Val Ser Val Asp Gly Arg Pro Val Arg Leu
Gln Leu Cys Asp Thr 95 100
105gca gga cag gat gag ttt gac aag ctg agg ccc ctc tgc tac acc aac
386Ala Gly Gln Asp Glu Phe Asp Lys Leu Arg Pro Leu Cys Tyr Thr Asn
110 115 120aca gac atc ttc ctg ctg tgc
ttc agc gtg gtg agc ccc aca tcc ttc 434Thr Asp Ile Phe Leu Leu Cys
Phe Ser Val Val Ser Pro Thr Ser Phe 125 130
135cag aac gtg ggc gag aag tgg gtt cca gag att cga cgt cac tgc cca
482Gln Asn Val Gly Glu Lys Trp Val Pro Glu Ile Arg Arg His Cys Pro140
145 150 155aag gcc ccc atc
atc ctg gtc ggg aca cag tcg gac ctc agg gag gac 530Lys Ala Pro Ile
Ile Leu Val Gly Thr Gln Ser Asp Leu Arg Glu Asp 160
165 170gtc aaa gtg ctc ata gaa ctg gac aag tgc
aaa gag aag ccg gtg cct 578Val Lys Val Leu Ile Glu Leu Asp Lys Cys
Lys Glu Lys Pro Val Pro 175 180
185gaa gag gcg gcg aag ctg tgc gcg gag gaa gtc aaa gct gtc tcc tac
626Glu Glu Ala Ala Lys Leu Cys Ala Glu Glu Val Lys Ala Val Ser Tyr
190 195 200atc gag tgc tca gcg ttg act
cag aaa aac ctc aaa gag gtt ttc gac 674Ile Glu Cys Ser Ala Leu Thr
Gln Lys Asn Leu Lys Glu Val Phe Asp 205 210
215gcc gcc att gtt gct ggt atc cag cac tca gac tcc cag cta cag cca
722Ala Ala Ile Val Ala Gly Ile Gln His Ser Asp Ser Gln Leu Gln Pro220
225 230 235aag aag tct aaa
agc agg acc ccg gat aag gtg cgg gac ctg tcc aag 770Lys Lys Ser Lys
Ser Arg Thr Pro Asp Lys Val Arg Asp Leu Ser Lys 240
245 250tct tgg tgg agg aag tat tgc tgc ctg gcc
tga ctctcgcaaa tagcaggtgt 823Ser Trp Trp Arg Lys Tyr Cys Cys Leu Ala
255 260ttaagctgca acagctcttt atggacgagg
ctgtcatagg atgagcccca aagcaccctc 883ttctgccctt aacttcctgt gtgcgggagc
ttagggctga gattcatatg caaaatacgt 943ttttttaaaa attgaaagtt acattttttt
tctgttaagt ctggaagctt tgagctgtag 1003acctccggat taatttatat tccatatgaa
aagggctctt caaagcgggg tgtcagcatg 1063aagttctgct gtgttgtaca ggacaaagga
gaatgaatgg gaccttctcc tgattaaggg 1123ctactgaggg ctcagtgcag ggcactgtgc
accaggcttg gtgagagtga gcaagcgtga 1183gctttgaaac cacacgagcc acccccggtt
ttgtaagggc aaagatctga aaccagcaag 1243ggccttctgc ttacgaaacc tcgagcccat
cccttctgtt tactcagatt ctcttaggat 1303tttaaaacaa ccaaacatcc cacagcctac
tggcatagtg ttggcgaaca gtgcacttgc 1363ttgttacggt tttgttttgt ttttttaaat
cacgtgacca gttatattgc tatgaaaatg 1423gtggagatgc ctcgtagaag gcgagtgctg
ggtgcacatg tgacattttc ttcagggagc 1483gactcatggt gagaccagag agggctctta
gcttgcagga ctggcttctg cagggcatct 1543gtgtcctgct gttaaaagca ggaggaggtg
cttgtctggg agctttaagt gtgctgggct 1603catatcgtcc cgtttgcaag gaattgggcc
accttgagag gccatagttg atggctatgg 1663gacacacaca cactttttcc ttaagtccac
caaaatgcct gcctgtacac acacacacac 1723acacacacac acacacacac acactggctg
gtttgctgat ggaaccctta gaccaccctc 1783ccacccccac ccctccccaa gcatggctgc
aagtgtcagg gcaccacacc ttcctcttct 1843tgacatttct ttgaacagac atcattttgt
aggatcttaa tttatacatt tttttcaggt 1903cataaaatgt gggatgaaca tactttgaac
cccagtgcct tcagggtcca ttgactaggg 1963aggcactgtc ttaggggaca ggtatgtgca
aggccttacc caccagtggc ttctcgctgc 2023aggtcatgtt tgtggcactt gttctttaag
gtgagggtct tatgaccgac tgttctgaga 2083cagccctgtg tcaggcaagc tctttcacag
ggttgtaggt atttccaaga cgccatagga 2143accagacagt gaatcatagc tatcagtttg
ctgtgggcaa ggaacctctt tttggccacc 2203tggtaacaaa attttatgtc tgtaaatttt
ttcttgctat ttaaaaaaaa aaatcaatct 2263tacgtttttc tgtaggaaaa aaaaaaacaa
gtaaaagaac aggccatatt tcaggtcaaa 2323ggcttcttcc tgctggtaaa tgggactgaa
gactttctta catcattatt aaaaggctaa 2383ttgctgaacc actagagtat atgaactgtt
tgtgaatgat attagccata gtctcctgag 2443gtgtttcctt gtggcctgag tggtaacatt
gttttgctta tggagatgct gtaactgacc 2503tagtgactca gcttatccta ttgtgcatgg
ctgtctggaa agccagcgta caagtggggc 2563tttgcatgcc ctgtgtacag agggtgggtg
ggaaagagtg aattatttaa ttttaaatgt 2623tataataaag ccaatgtagt tgagaccaag
gaaatgagca ttgagaacac aaacttgaag 2683tctggtgcca gggttgttgg acctcacacc
ctgtctctga gccacccgga agtgacataa 2743aggacgctgt gtgatcaagt tctggacact
tttctgggat gcgtaccact ggactattta 2803tgtcacaaat ctagtgggtt gacgctgccc
tgcaagtttt caatgtccct gcatcctatg 2863aagtcataat gatctgactg tactggaggt
tttcctgcat tttttacttt tcgaaaatag 2923aggtttaggc tgagaattct aaacgcatgt
gcctgggtgg gacgtcaagt cagggttctc 2983atcaaagctg agaagtggct ggaatgttca
gcttggtgtc tggggaggat cctgtgagct 3043atgtagagag gtggctcttc agcctgactc
agtgtgggct gaacgaagta cctgcagaac 3103acacggtagc aggctccaaa atcgtcacct
caagcatgcg tgcaagcaaa cttccgagaa 3163ctccgttttc tgctcggcag acgtgtgagc
agctacccag aagtctcaag ccaaaagggg 3223agcctcgctc gctggctcct ctgcaggtgc
cttatcgacc tgtgctcttc tcttttcccg 3283tgtcaaagat gttggacagg atcttgtact
tgaaacatac tgcaaatgag ttactatgaa 3343ataaattctg acctgtggcc g
336440261PRTMus musculus 40Met Ala Pro
Gln Gln Gly Arg Pro Ala Leu Pro Ala Arg Cys Glu Pro1 5
10 15Pro Ala Ala Pro Pro Val Pro Pro Arg
Arg Glu Arg Gly Gly Arg Gly 20 25
30Ala Arg Gly Pro Gly Val Ser Gly Gly Arg Gly Arg Ala Gly Gly Ala
35 40 45Glu Gly Arg Gly Val Lys Cys
Val Leu Val Gly Asp Gly Ala Val Gly 50 55
60Lys Thr Ser Leu Val Val Ser Tyr Thr Thr Asn Gly Tyr Pro Thr Glu65
70 75 80Tyr Ile Pro Thr
Ala Phe Asp Asn Phe Ser Ala Val Val Ser Val Asp 85
90 95Gly Arg Pro Val Arg Leu Gln Leu Cys Asp
Thr Ala Gly Gln Asp Glu 100 105
110Phe Asp Lys Leu Arg Pro Leu Cys Tyr Thr Asn Thr Asp Ile Phe Leu
115 120 125Leu Cys Phe Ser Val Val Ser
Pro Thr Ser Phe Gln Asn Val Gly Glu 130 135
140Lys Trp Val Pro Glu Ile Arg Arg His Cys Pro Lys Ala Pro Ile
Ile145 150 155 160Leu Val
Gly Thr Gln Ser Asp Leu Arg Glu Asp Val Lys Val Leu Ile
165 170 175Glu Leu Asp Lys Cys Lys Glu
Lys Pro Val Pro Glu Glu Ala Ala Lys 180 185
190Leu Cys Ala Glu Glu Val Lys Ala Val Ser Tyr Ile Glu Cys
Ser Ala 195 200 205Leu Thr Gln Lys
Asn Leu Lys Glu Val Phe Asp Ala Ala Ile Val Ala 210
215 220Gly Ile Gln His Ser Asp Ser Gln Leu Gln Pro Lys
Lys Ser Lys Ser225 230 235
240Arg Thr Pro Asp Lys Val Arg Asp Leu Ser Lys Ser Trp Trp Arg Lys
245 250 255Tyr Cys Cys Leu Ala
260411719DNAMus musculusCDS(54)..(1136) 41cgcatggcgc ccgcacacgg
agtctgacct gatgtagacg caagggggtt aat atg 56
Met
1aac gtc cct ctc ggt gga atc tgg ctc tgg ctc cct
ctg ctc ttg acc 104Asn Val Pro Leu Gly Gly Ile Trp Leu Trp Leu Pro
Leu Leu Leu Thr 5 10 15tgg
ctc acc cct gag gtc agc tct tca tgg tgg tac atg aga gct aca 152Trp
Leu Thr Pro Glu Val Ser Ser Ser Trp Trp Tyr Met Arg Ala Thr 20
25 30ggt ggc tcc tcc agg gtg atg tgt gac
aat gtg cca ggc ctg gtg agc 200Gly Gly Ser Ser Arg Val Met Cys Asp
Asn Val Pro Gly Leu Val Ser 35 40
45cgg cag cgt cag ctg tgc cac cga cac cca gat gtg atg cgt gcc att
248Arg Gln Arg Gln Leu Cys His Arg His Pro Asp Val Met Arg Ala Ile50
55 60 65ggc ctg ggt gtg gct
gag tgg act gca gag tgc caa cac cag ttc cgc 296Gly Leu Gly Val Ala
Glu Trp Thr Ala Glu Cys Gln His Gln Phe Arg 70
75 80cag cat cgc tgg aac tgc aac acc ctg gac aga
gat cac agc ctc ttt 344Gln His Arg Trp Asn Cys Asn Thr Leu Asp Arg
Asp His Ser Leu Phe 85 90
95ggc cgg gtc ctc ctc cga agt agt cga gaa tcg gcc ttt gtt tac gcc
392Gly Arg Val Leu Leu Arg Ser Ser Arg Glu Ser Ala Phe Val Tyr Ala
100 105 110atc tct tca gct ggc gtt gta
ttt gcc atc acc agg gcc tgt agc caa 440Ile Ser Ser Ala Gly Val Val
Phe Ala Ile Thr Arg Ala Cys Ser Gln 115 120
125gga gaa tta aag tcc tgc tcc tgt gat cca aag aag aaa gga agt gcc
488Gly Glu Leu Lys Ser Cys Ser Cys Asp Pro Lys Lys Lys Gly Ser Ala130
135 140 145aag gac agc aaa
ggc acc ttc gac tgg ggt ggc tgc agt gac aat att 536Lys Asp Ser Lys
Gly Thr Phe Asp Trp Gly Gly Cys Ser Asp Asn Ile 150
155 160gac tac ggg atc aag ttt gcc cgt gcc ttt
gta gat gcc aag gag agg 584Asp Tyr Gly Ile Lys Phe Ala Arg Ala Phe
Val Asp Ala Lys Glu Arg 165 170
175aaa ggc aag gat gcc aga gcc ctg atg aac ctt cac aac aac aga gct
632Lys Gly Lys Asp Ala Arg Ala Leu Met Asn Leu His Asn Asn Arg Ala
180 185 190gga agg aag gct gta aag cgc
ttc ttg aaa caa gaa tgc aag tgt cat 680Gly Arg Lys Ala Val Lys Arg
Phe Leu Lys Gln Glu Cys Lys Cys His 195 200
205ggt gtg agt ggc tcc tgt act ctg agg aca tgc tgg ctg gcc atg gct
728Gly Val Ser Gly Ser Cys Thr Leu Arg Thr Cys Trp Leu Ala Met Ala210
215 220 225gac ttc agg aaa
aca ggc gac tat ctc tgg agg aag tac aat ggg gcc 776Asp Phe Arg Lys
Thr Gly Asp Tyr Leu Trp Arg Lys Tyr Asn Gly Ala 230
235 240atc cag gta gtc atg aac cag gat ggc act
ggc ttc act gta gcc aat 824Ile Gln Val Val Met Asn Gln Asp Gly Thr
Gly Phe Thr Val Ala Asn 245 250
255aag agg ttt aag aag cca acg aaa aat gac ctc gtg tat ttt gag aat
872Lys Arg Phe Lys Lys Pro Thr Lys Asn Asp Leu Val Tyr Phe Glu Asn
260 265 270tct cca gac tac tgt atc agg
gac cga gag gca ggc tcc ctg ggt aca 920Ser Pro Asp Tyr Cys Ile Arg
Asp Arg Glu Ala Gly Ser Leu Gly Thr 275 280
285gcg ggc cgt gtg tgc aac ttg act tcc cga ggc atg gac agc tgc gaa
968Ala Gly Arg Val Cys Asn Leu Thr Ser Arg Gly Met Asp Ser Cys Glu290
295 300 305gtt atg tgt tgt
ggg aga ggc tat gac aca tcc cac gtc acc cgg atg 1016Val Met Cys Cys
Gly Arg Gly Tyr Asp Thr Ser His Val Thr Arg Met 310
315 320acc aag tgt gag tgt aaa ttc cac tgg tgc
tgt gcc gtg cgc tgt cag 1064Thr Lys Cys Glu Cys Lys Phe His Trp Cys
Cys Ala Val Arg Cys Gln 325 330
335gac tgc ctg gag gcc ctg gac gtg cac aca tgc aag gcc ccc aag agt
1112Asp Cys Leu Glu Ala Leu Asp Val His Thr Cys Lys Ala Pro Lys Ser
340 345 350gcc gac tgg gcg acg cct aca
tga cctcagcaga ggtcatattc gccttttctt 1166Ala Asp Trp Ala Thr Pro Thr
355 360ccctcaagga ctccaattac atcttcaagg acactggacc
tctgggttgt tttcaggggc 1226tctttcttaa ggcatgaagc cttcatctca agagaaaccc
cctttcccct ctctgggggc 1286cccaggactg ggaaccacct gctgcacata agtacaccct
attctgtcta tcttgggcat 1346tctgatgtca cctctcttcc tgctgatttc tttttggaaa
tggcatgaca ggctgttaga 1406ggaggagggt catagccccc caccactgtc acctagacat
ttcctctttg gctgcgggga 1466gaaacatcac atagcgaagg aacttcctct gtgttttccc
agattccaac aacccagaaa 1526gtctgtgttt ccctggggcg cggggtaggg atggaaagca
gaatgagctg acaccaaaat 1586ttcctcggat ttttttaaaa aaagagtaag caagggcttt
aactaagtga tagctgttga 1646tagcatcctt ggtgactttc tagagaaaga tggcttccaa
taaacatcag gttaaaacaa 1706aaaaaaaaaa aaa
171942360PRTMus musculus 42Met Asn Val Pro Leu Gly
Gly Ile Trp Leu Trp Leu Pro Leu Leu Leu1 5
10 15Thr Trp Leu Thr Pro Glu Val Ser Ser Ser Trp Trp
Tyr Met Arg Ala 20 25 30Thr
Gly Gly Ser Ser Arg Val Met Cys Asp Asn Val Pro Gly Leu Val 35
40 45Ser Arg Gln Arg Gln Leu Cys His Arg
His Pro Asp Val Met Arg Ala 50 55
60Ile Gly Leu Gly Val Ala Glu Trp Thr Ala Glu Cys Gln His Gln Phe65
70 75 80Arg Gln His Arg Trp
Asn Cys Asn Thr Leu Asp Arg Asp His Ser Leu 85
90 95Phe Gly Arg Val Leu Leu Arg Ser Ser Arg Glu
Ser Ala Phe Val Tyr 100 105
110Ala Ile Ser Ser Ala Gly Val Val Phe Ala Ile Thr Arg Ala Cys Ser
115 120 125Gln Gly Glu Leu Lys Ser Cys
Ser Cys Asp Pro Lys Lys Lys Gly Ser 130 135
140Ala Lys Asp Ser Lys Gly Thr Phe Asp Trp Gly Gly Cys Ser Asp
Asn145 150 155 160Ile Asp
Tyr Gly Ile Lys Phe Ala Arg Ala Phe Val Asp Ala Lys Glu
165 170 175Arg Lys Gly Lys Asp Ala Arg
Ala Leu Met Asn Leu His Asn Asn Arg 180 185
190Ala Gly Arg Lys Ala Val Lys Arg Phe Leu Lys Gln Glu Cys
Lys Cys 195 200 205His Gly Val Ser
Gly Ser Cys Thr Leu Arg Thr Cys Trp Leu Ala Met 210
215 220Ala Asp Phe Arg Lys Thr Gly Asp Tyr Leu Trp Arg
Lys Tyr Asn Gly225 230 235
240Ala Ile Gln Val Val Met Asn Gln Asp Gly Thr Gly Phe Thr Val Ala
245 250 255Asn Lys Arg Phe Lys
Lys Pro Thr Lys Asn Asp Leu Val Tyr Phe Glu 260
265 270Asn Ser Pro Asp Tyr Cys Ile Arg Asp Arg Glu Ala
Gly Ser Leu Gly 275 280 285Thr Ala
Gly Arg Val Cys Asn Leu Thr Ser Arg Gly Met Asp Ser Cys 290
295 300Glu Val Met Cys Cys Gly Arg Gly Tyr Asp Thr
Ser His Val Thr Arg305 310 315
320Met Thr Lys Cys Glu Cys Lys Phe His Trp Cys Cys Ala Val Arg Cys
325 330 335Gln Asp Cys Leu
Glu Ala Leu Asp Val His Thr Cys Lys Ala Pro Lys 340
345 350Ser Ala Asp Trp Ala Thr Pro Thr 355
360433576DNAMus musculusCDS(239)..(1408) 43ggagccactg
acaccgcacc cgaccgccca cacccggctc agcgctcgtc ggtctcctgg 60ccctgcacgc
tcttgggaac cctgcgtctg gctcccgggc tccacgtgcc ttgaggtcct 120cggtcgccca
tggtccccat ggccactctg tggggcgatc taggagacgc ctgagcgaag 180cccagacagt
gcccgtccac ggccctgcgg gcttcggggc gggagtctgc ggggagct 238atg ctg aag
ctg cag ggt gag gat gaa gcc gcg cag ctc gcc cct cgg 286Met Leu Lys
Leu Gln Gly Glu Asp Glu Ala Ala Gln Leu Ala Pro Arg1 5
10 15cgt gcc cgc gtc ccc gtg ccc aga ccc
acg gcc ccc gac gtg tcc cca 334Arg Ala Arg Val Pro Val Pro Arg Pro
Thr Ala Pro Asp Val Ser Pro 20 25
30tct tcc gcc cgc ctg ggt ctt gcc tgc ctg ctg ctg ctg cta ctc ctg
382Ser Ser Ala Arg Leu Gly Leu Ala Cys Leu Leu Leu Leu Leu Leu Leu
35 40 45act ctg ccg gcc cgt gta gac
acg tcc tgg tgg tac ata ggg gct ctg 430Thr Leu Pro Ala Arg Val Asp
Thr Ser Trp Trp Tyr Ile Gly Ala Leu 50 55
60gga gcc cga gtg atc tgt gac aac atc ccc ggt ctg gtg agc cgg cag
478Gly Ala Arg Val Ile Cys Asp Asn Ile Pro Gly Leu Val Ser Arg Gln65
70 75 80cgg cag ttg tgt
caa cgc tac cca gac atc atg cgc tca gta ggt gag 526Arg Gln Leu Cys
Gln Arg Tyr Pro Asp Ile Met Arg Ser Val Gly Glu 85
90 95ggt gcc cgg gaa tgg atc cga gag tgc cag
cac cag ttc cgt cac cac 574Gly Ala Arg Glu Trp Ile Arg Glu Cys Gln
His Gln Phe Arg His His 100 105
110cgc tgg aat tgc acc aca ctg gac cgg gac cac act gtc ttt ggc cgc
622Arg Trp Asn Cys Thr Thr Leu Asp Arg Asp His Thr Val Phe Gly Arg
115 120 125gcc atg ctc aga agc agc cgg
gag gca gcg ttc gtc tat gct atc tcg 670Ala Met Leu Arg Ser Ser Arg
Glu Ala Ala Phe Val Tyr Ala Ile Ser 130 135
140tca gca gga gtg gtc cac gct atc act cgg gcc tgc agc cag ggt gag
718Ser Ala Gly Val Val His Ala Ile Thr Arg Ala Cys Ser Gln Gly Glu145
150 155 160ctg agc gtg tgc
agc tgt gac cca tat acc cgc ggt cgg cac cat gat 766Leu Ser Val Cys
Ser Cys Asp Pro Tyr Thr Arg Gly Arg His His Asp 165
170 175caa cga ggg gac ttt gac tgg ggt ggc tgt
agt gac aac atc cat tac 814Gln Arg Gly Asp Phe Asp Trp Gly Gly Cys
Ser Asp Asn Ile His Tyr 180 185
190ggt gtt cgc ttt gcc aag gct ttt gtg gat gcc aaa gag aag agg ctt
862Gly Val Arg Phe Ala Lys Ala Phe Val Asp Ala Lys Glu Lys Arg Leu
195 200 205aag gat gcc cgg gcc ctc atg
aac tta cac aac aac cgc tgt ggt cgc 910Lys Asp Ala Arg Ala Leu Met
Asn Leu His Asn Asn Arg Cys Gly Arg 210 215
220acg gct gtt cgg aga ttc ctg aag ctg gag tgc aag tgt cac ggt gtg
958Thr Ala Val Arg Arg Phe Leu Lys Leu Glu Cys Lys Cys His Gly Val225
230 235 240agt ggc tcc tgt
act ctg cgc acc tgc tgg aga gca ctc tca gac ttc 1006Ser Gly Ser Cys
Thr Leu Arg Thr Cys Trp Arg Ala Leu Ser Asp Phe 245
250 255cga cgc aca ggt gac tac ctg agg agg cga
tat gat ggg gct gtg cag 1054Arg Arg Thr Gly Asp Tyr Leu Arg Arg Arg
Tyr Asp Gly Ala Val Gln 260 265
270gtg acg gcc aca cag gat ggg gcc aat ttc aca gca gcg cgc cag ggc
1102Val Thr Ala Thr Gln Asp Gly Ala Asn Phe Thr Ala Ala Arg Gln Gly
275 280 285tat cgc cac gcc acc cgg act
gat ctt gtc tac ttt gac aac tcc cct 1150Tyr Arg His Ala Thr Arg Thr
Asp Leu Val Tyr Phe Asp Asn Ser Pro 290 295
300gac tac tgt gtc ttg gac aag gct gca ggt tcc cta ggt acc gca ggc
1198Asp Tyr Cys Val Leu Asp Lys Ala Ala Gly Ser Leu Gly Thr Ala Gly305
310 315 320cgc gtc tgc agc
aag act tct aaa gga aca gat ggg tgt gaa atc atg 1246Arg Val Cys Ser
Lys Thr Ser Lys Gly Thr Asp Gly Cys Glu Ile Met 325
330 335tgt tgt ggc cga ggg tat gac aca act cgg
gtc acc cgc gtc acc cag 1294Cys Cys Gly Arg Gly Tyr Asp Thr Thr Arg
Val Thr Arg Val Thr Gln 340 345
350tgt gag tgc aaa ttc cac tgg tgc tgt gct gtg cgg tgc aag gag tgc
1342Cys Glu Cys Lys Phe His Trp Cys Cys Ala Val Arg Cys Lys Glu Cys
355 360 365aga aac act gtg gat gtc cac
aca tgc aag gcc cct aag aag gca gag 1390Arg Asn Thr Val Asp Val His
Thr Cys Lys Ala Pro Lys Lys Ala Glu 370 375
380tgg ctg gac cag acc tga acacacagaa acctcattct tccctccact
1438Trp Leu Asp Gln Thr385 390tcaagcctct gactcaaaag
cacaagaccc ttgcatgcgc accttcctct accctcaatc 1498ctgggctgct atggcttctg
tcacggacct ggagagtgat ccggagggac cccaatgtcc 1558cggccgcctg gttccttagc
cctagggacg tgttgatagg ggatggattt aggaggctga 1618gtgactccct gatggtccat
ctggaggttt gaagggagag taggagaggt ctgtcttcag 1678agtgatttga gttgcactaa
gtcaaggctc atcctcccct ttgcttgcac tgacttctga 1738tcctctttgg gtatgcaaca
ggaagggaac ctggaggtag cttccgtgtt tgatgctact 1798ctgcctgagg ataggacaga
gataaaactg cctgtccctt tgctggagac agtacgggca 1858gactatctta ggccatagta
ttctgctgag accctgagat agctagatgg gttagccaca 1918ttgaacaagg ctccacatca
tgcttctacg cagcttataa agtagtggtt tggtgaggag 1978gaaaatcaca atgctctaca
gatacacatt ctctgtgcct ccctttccac ctacatcaca 2038cagcagcagc ctgctcactg
gctgcctgtt cagagtgagg cagcttgcag tgggtcaaat 2098tcttaccagg ccattagagg
cccggaacag gattgtgaga gaatgacata gaaagcctgg 2158ctaggccttg ggacttcccc
cacatccact attccggaga ttcggtagga agggaggtaa 2218ctcatgggaa gggtgagcgc
acctgatctc aggggttcca tgaggatcag tgtatactag 2278gaaggcagag atctcgcatt
ttgctagttc ttgaggatct tcagctttga agtaggaaca 2338aaaggcagca gctatagaga
gagagctggt gctggagccg aggtggcaaa catcctataa 2398ggcctttctc atttacccag
caaatcttta ttttgtgatt caccaggtcc aactgttaac 2458tactgcacgt tccacgatcg
acttaaacag ggaaggttct ctctgtgcta ctgaccgttg 2518cctaacgagg gtacacagga
gtggagcctt caaagagagc aggcacagtg acatgggggt 2578tccaaacctt gatggtctag
ttttatgtga cctcgacaat ggtcatcttc ttccctattg 2638ataaacagaa atagtataga
aatccacagt tagacttagg tctaatccca gctatttact 2698ctctattttt tattttcagc
agggtcttta aattctcctc tcccattttc ttatctgtaa 2758agtgagggtg aaactgagat
ctaactgtgc cccaaactgt agccgactga tagacgtcat 2818caacactctc actggtcaag
tacttcctgc ttctctggga ccttctgatt tagggctgtc 2878tgggcagaca acagagtaga
ttcaaagggc tttcacaatg aattctggat atagctcctc 2938tctctcttct cagggttcct
cttcatccaa tcgtactctc agatgtttgt ggagcaacct 2998ctttctgccc aggcagcagg
aggctggggt ggggtggggt gggggggcac agctctggcc 3058acagaggcag atttatttgg
atgataggac taatatttgt gtaacctgct gagacctgtg 3118tgggagagtt tagtatggtt
tttcttttgg tgaggggatt tgctccggtt tcacatccat 3178taacacaaaa catgagctag
tcagggccct tgtggtctgt ggtgagggga tgactggaga 3238aacgggactg agtgagtcag
gcggagggaa tgtcttcctc gcagagtaga gtcaacggga 3298taactgatga gccagtggtg
gggtcacgga gggggcggag gggaagaggg acttctcttg 3358gaagagagga gttttggggg
caggggcgag aacatccaag ttacggtatc agtgatggca 3418ttggccttca ctggggagcc
agcctgaggt aaatctactt gtgctgtatt ctctttgagt 3478ttgggttctt agctgtggca
gacatctgtg acatctcata ttactccatg cctttgcctg 3538ggctccaaat tctagctgat
aaagatatac aaccactt 357644389PRTMus musculus
44Met Leu Lys Leu Gln Gly Glu Asp Glu Ala Ala Gln Leu Ala Pro Arg1
5 10 15Arg Ala Arg Val Pro Val
Pro Arg Pro Thr Ala Pro Asp Val Ser Pro 20 25
30Ser Ser Ala Arg Leu Gly Leu Ala Cys Leu Leu Leu Leu
Leu Leu Leu 35 40 45Thr Leu Pro
Ala Arg Val Asp Thr Ser Trp Trp Tyr Ile Gly Ala Leu 50
55 60Gly Ala Arg Val Ile Cys Asp Asn Ile Pro Gly Leu
Val Ser Arg Gln65 70 75
80Arg Gln Leu Cys Gln Arg Tyr Pro Asp Ile Met Arg Ser Val Gly Glu
85 90 95Gly Ala Arg Glu Trp Ile
Arg Glu Cys Gln His Gln Phe Arg His His 100
105 110Arg Trp Asn Cys Thr Thr Leu Asp Arg Asp His Thr
Val Phe Gly Arg 115 120 125Ala Met
Leu Arg Ser Ser Arg Glu Ala Ala Phe Val Tyr Ala Ile Ser 130
135 140Ser Ala Gly Val Val His Ala Ile Thr Arg Ala
Cys Ser Gln Gly Glu145 150 155
160Leu Ser Val Cys Ser Cys Asp Pro Tyr Thr Arg Gly Arg His His Asp
165 170 175Gln Arg Gly Asp
Phe Asp Trp Gly Gly Cys Ser Asp Asn Ile His Tyr 180
185 190Gly Val Arg Phe Ala Lys Ala Phe Val Asp Ala
Lys Glu Lys Arg Leu 195 200 205Lys
Asp Ala Arg Ala Leu Met Asn Leu His Asn Asn Arg Cys Gly Arg 210
215 220Thr Ala Val Arg Arg Phe Leu Lys Leu Glu
Cys Lys Cys His Gly Val225 230 235
240Ser Gly Ser Cys Thr Leu Arg Thr Cys Trp Arg Ala Leu Ser Asp
Phe 245 250 255Arg Arg Thr
Gly Asp Tyr Leu Arg Arg Arg Tyr Asp Gly Ala Val Gln 260
265 270Val Thr Ala Thr Gln Asp Gly Ala Asn Phe
Thr Ala Ala Arg Gln Gly 275 280
285Tyr Arg His Ala Thr Arg Thr Asp Leu Val Tyr Phe Asp Asn Ser Pro 290
295 300Asp Tyr Cys Val Leu Asp Lys Ala
Ala Gly Ser Leu Gly Thr Ala Gly305 310
315 320Arg Val Cys Ser Lys Thr Ser Lys Gly Thr Asp Gly
Cys Glu Ile Met 325 330
335Cys Cys Gly Arg Gly Tyr Asp Thr Thr Arg Val Thr Arg Val Thr Gln
340 345 350Cys Glu Cys Lys Phe His
Trp Cys Cys Ala Val Arg Cys Lys Glu Cys 355 360
365Arg Asn Thr Val Asp Val His Thr Cys Lys Ala Pro Lys Lys
Ala Glu 370 375 380Trp Leu Asp Gln
Thr385453000DNAMus musculusCDS(46)..(1113) 45cctcttcatg atcgccggca
aacttcctcc tcggcgctgc ttcta atg gag ccc cac 57
Met Glu Pro His
1ctg ctc ggg ctg cta ctc ggc ctc ctg ctc agt ggc acc agg
gtc ctc 105Leu Leu Gly Leu Leu Leu Gly Leu Leu Leu Ser Gly Thr Arg
Val Leu5 10 15 20gct
ggc tac cca att tgg tgg tcc ctg gcc ctg ggc cag cag tac aca 153Ala
Gly Tyr Pro Ile Trp Trp Ser Leu Ala Leu Gly Gln Gln Tyr Thr
25 30 35tct ctg gcc tcc cag cct ctg
ctc tgc ggc tcc atc cca ggc ctg gtc 201Ser Leu Ala Ser Gln Pro Leu
Leu Cys Gly Ser Ile Pro Gly Leu Val 40 45
50ccc aag caa ctg cgc ttc tgc cgc aat tac atc gag atc atg
ccc agc 249Pro Lys Gln Leu Arg Phe Cys Arg Asn Tyr Ile Glu Ile Met
Pro Ser 55 60 65gta gca gaa ggt
gtg aag ctg ggc atc cag gag tgc cag cat cag ttc 297Val Ala Glu Gly
Val Lys Leu Gly Ile Gln Glu Cys Gln His Gln Phe 70 75
80cgg ggc cgc cgg tgg aac tgt acc acc ata gat gac agc
ctg gcc atc 345Arg Gly Arg Arg Trp Asn Cys Thr Thr Ile Asp Asp Ser
Leu Ala Ile85 90 95
100ttt ggg cct gtc ttg gac aaa gcc acc cgt gaa tcg gcc ttc gtg cat
393Phe Gly Pro Val Leu Asp Lys Ala Thr Arg Glu Ser Ala Phe Val His
105 110 115gcc atc gcc tcg gct
ggt gtc gcc ttc gca gtc aca cgc tcc tgc gct 441Ala Ile Ala Ser Ala
Gly Val Ala Phe Ala Val Thr Arg Ser Cys Ala 120
125 130gag gga acc tcc acc atc tgc ggc tgt gac tca cat
cat aag ggg cca 489Glu Gly Thr Ser Thr Ile Cys Gly Cys Asp Ser His
His Lys Gly Pro 135 140 145cct gga
gaa ggc tgg aag tgg ggc ggc tgc agc gag gac gcc gac ttc 537Pro Gly
Glu Gly Trp Lys Trp Gly Gly Cys Ser Glu Asp Ala Asp Phe 150
155 160ggg gtg ctg gtg tcc cgg gaa ttt gcg gat gcg
cgg gag aac agg cca 585Gly Val Leu Val Ser Arg Glu Phe Ala Asp Ala
Arg Glu Asn Arg Pro165 170 175
180gat gcc cgc tca gct atg aac aag cac aac aat gaa gca ggc cga acg
633Asp Ala Arg Ser Ala Met Asn Lys His Asn Asn Glu Ala Gly Arg Thr
185 190 195acc atc ctg gac cac
atg cac cta aag tgt aaa tgc cac ggg ttg tcc 681Thr Ile Leu Asp His
Met His Leu Lys Cys Lys Cys His Gly Leu Ser 200
205 210ggc agc tgc gag gtg aag acc tgc tgg tgg gcc cag
ccc gac ttc cgt 729Gly Ser Cys Glu Val Lys Thr Cys Trp Trp Ala Gln
Pro Asp Phe Arg 215 220 225gcc att
ggc gac ttc ctc aag gac aag tac gac agt gcc tcc gag atg 777Ala Ile
Gly Asp Phe Leu Lys Asp Lys Tyr Asp Ser Ala Ser Glu Met 230
235 240gtg gtg gag aaa cac cgt gag tcc cga ggc tgg
gtg gag acc ctg cgg 825Val Val Glu Lys His Arg Glu Ser Arg Gly Trp
Val Glu Thr Leu Arg245 250 255
260gct aag tac gcg ctc ttc aag cca ccc acc gag agg gac ctg gtc tac
873Ala Lys Tyr Ala Leu Phe Lys Pro Pro Thr Glu Arg Asp Leu Val Tyr
265 270 275tac gag aac tcc ccc
aac ttt tgt gag ccc aac cca gag acg ggc tcc 921Tyr Glu Asn Ser Pro
Asn Phe Cys Glu Pro Asn Pro Glu Thr Gly Ser 280
285 290ttt ggt acc agg gac cgg act tgc aat gtc acc tcc
cac ggc atc gat 969Phe Gly Thr Arg Asp Arg Thr Cys Asn Val Thr Ser
His Gly Ile Asp 295 300 305ggc tgc
gat ctg ctg tgc tgt ggc cgg ggc cac aac acg agg acg gag 1017Gly Cys
Asp Leu Leu Cys Cys Gly Arg Gly His Asn Thr Arg Thr Glu 310
315 320aaa cgg aag gag aaa tgc cat tgc gtc ttc cac
tgg tgc tgc tat gtc 1065Lys Arg Lys Glu Lys Cys His Cys Val Phe His
Trp Cys Cys Tyr Val325 330 335
340agc tgc caa gag tgt att cgc atc tac gat gtg cac acc tgc aag tag
1113Ser Cys Gln Glu Cys Ile Arg Ile Tyr Asp Val His Thr Cys Lys
345 350 355tgagccaggg cactgggaag
gggtagattg tgcggctgga tccattcatc gaagtcccat 1173gagaagcagg atctagatcc
aggccagcct tcggcactgg ccagcaagga gcatggactg 1233ttgccagctg catgtgataa
acgacctgga cccagccggc ctcggacgga cgggcggctt 1293ctttctcaac taacgtctct
ccccctgctc tggatggtgt acggctttac agaggggctt 1353tctttatggt tttaccaggg
tctgctgggg acagactcga ggcttacctt tgcacatgtt 1413aaagaaaata aaaatgaaaa
aaaaaaatct accgcaacag aacaggctgg gctagtgtga 1473gctcttggcc tggtgggaag
gacaagacca tggcgagatt ctgtgtccaa gctgcctcta 1533ctcgtgacat tccaagatgc
ctctgaggtg ggaactgtga agtaggacag agccccgcag 1593tcccctcttg tccgtcgact
cccatttaaa ttggacatac cttgtcgttc tgagaaaagc 1653catagatagg tgtagctggg
atgtagtgat ggggaggccc ctggccaaca gtgggagcaa 1713gatcttgagt tttgaagacc
tcagagttct gggcggcctg ggaagccatc tgcagaacag 1773agttccttgt gggctcctgt
tttcgctagc cctgttctgc cctggagcga cagtcagatc 1833tccacgcccc tttctgttgt
tctacagtgt ccacctttac tacgcgtttt tttttttttt 1893ttcatgatga ccttgtaaat
aggtcagatg tggaggcagg tctcttctgg ctccatccac 1953cacacccaga aagaatgggc
tgctctgccc ttctcagcct tgctaaccag cagacaccga 2013ggagagcagc ggggcacctt
agagagcaat ctaaacatgg ttggcaggtg gggagggtaa 2073agagtcccac ttcctttgtg
ttagaaggca gactaccctg cgtccttttc tcccattggc 2133tgaagtaacc agaaagacaa
gagatcctta acaagccctt cttcccactt gtaaaaggga 2193tagcctatct cagttcccaa
ggatctggat tagatagata ttcaaaagag gcaagcagcg 2253aatggaggca gctcccagct
ctgttcccga cgcatgatgg tactggctgg gtttagtaag 2313gtgggtgggg ctgcacggat
caatccatca actccgtctt aaggagaatc agaaagagga 2373gataaaatgg gggaatgggg
cagaacaaag aatttgtcct ttcccgcttc tgtctagggt 2433ctgctaatgc tggcttgacg
aggggtcagc cacttctttc ctgttgtgca gttggcttgc 2493caagcaggct ccagtaggcc
cttgcctgca ctctctacca tgtgaccatg agcactgctc 2553tagggacacc tcccatccct
tcctagcacc ccaaatgccc cttcccatct ctccttccag 2613aagttggaaa tcaagtcaac
tggataacgc ttgtgtgaga cacttgagca gaacggatac 2673aacaatttac aagtctcttc
atatctatgt attctatatt aaaagtgata aagtcatgtt 2733tccggggcgt attcaagtag
ctgacaagta attatttaat aatagtacat gagcgcattg 2793taattatcct cgccatagtc
aggtaatagc atccaatggg aggtccctac caacctgctg 2853tatccaaagt tttgtaaaaa
gttgtagaag ttgttgatct ttttgatttt atattcaaaa 2913agtctctttt tataaatatt
atttattata caatgtatat acctttgagt taactaagat 2973tatatattat ataaatatat
atatatt 300046355PRTMus musculus
46Met Glu Pro His Leu Leu Gly Leu Leu Leu Gly Leu Leu Leu Ser Gly1
5 10 15Thr Arg Val Leu Ala Gly
Tyr Pro Ile Trp Trp Ser Leu Ala Leu Gly 20 25
30Gln Gln Tyr Thr Ser Leu Ala Ser Gln Pro Leu Leu Cys
Gly Ser Ile 35 40 45Pro Gly Leu
Val Pro Lys Gln Leu Arg Phe Cys Arg Asn Tyr Ile Glu 50
55 60Ile Met Pro Ser Val Ala Glu Gly Val Lys Leu Gly
Ile Gln Glu Cys65 70 75
80Gln His Gln Phe Arg Gly Arg Arg Trp Asn Cys Thr Thr Ile Asp Asp
85 90 95Ser Leu Ala Ile Phe Gly
Pro Val Leu Asp Lys Ala Thr Arg Glu Ser 100
105 110Ala Phe Val His Ala Ile Ala Ser Ala Gly Val Ala
Phe Ala Val Thr 115 120 125Arg Ser
Cys Ala Glu Gly Thr Ser Thr Ile Cys Gly Cys Asp Ser His 130
135 140His Lys Gly Pro Pro Gly Glu Gly Trp Lys Trp
Gly Gly Cys Ser Glu145 150 155
160Asp Ala Asp Phe Gly Val Leu Val Ser Arg Glu Phe Ala Asp Ala Arg
165 170 175Glu Asn Arg Pro
Asp Ala Arg Ser Ala Met Asn Lys His Asn Asn Glu 180
185 190Ala Gly Arg Thr Thr Ile Leu Asp His Met His
Leu Lys Cys Lys Cys 195 200 205His
Gly Leu Ser Gly Ser Cys Glu Val Lys Thr Cys Trp Trp Ala Gln 210
215 220Pro Asp Phe Arg Ala Ile Gly Asp Phe Leu
Lys Asp Lys Tyr Asp Ser225 230 235
240Ala Ser Glu Met Val Val Glu Lys His Arg Glu Ser Arg Gly Trp
Val 245 250 255Glu Thr Leu
Arg Ala Lys Tyr Ala Leu Phe Lys Pro Pro Thr Glu Arg 260
265 270Asp Leu Val Tyr Tyr Glu Asn Ser Pro Asn
Phe Cys Glu Pro Asn Pro 275 280
285Glu Thr Gly Ser Phe Gly Thr Arg Asp Arg Thr Cys Asn Val Thr Ser 290
295 300His Gly Ile Asp Gly Cys Asp Leu
Leu Cys Cys Gly Arg Gly His Asn305 310
315 320Thr Arg Thr Glu Lys Arg Lys Glu Lys Cys His Cys
Val Phe His Trp 325 330
335Cys Cys Tyr Val Ser Cys Gln Glu Cys Ile Arg Ile Tyr Asp Val His
340 345 350Thr Cys Lys
355472814DNAMus musculusCDS(129)..(1187) 47gaattcatgt cttacggtca
aggcagaggg cccagcgcca ctgcagccgc gccacctccc 60agggccgggc cagcccaggc
gtccgcgctc tcggggtgga ctccccccgc tgcgcgctca 120agccggcg atg gct cct
ctc gga tac ctc tta gtg ctc tgc agc ctg aag 170 Met Ala Pro
Leu Gly Tyr Leu Leu Val Leu Cys Ser Leu Lys 1 5
10cag gct ctg ggc agc tac ccg atc tgg tgg tcc ttg gct gtg
gga ccc 218Gln Ala Leu Gly Ser Tyr Pro Ile Trp Trp Ser Leu Ala Val
Gly Pro15 20 25 30cag
tac tcc tct ctg agc act cag ccc att ctc tgt gcc agc atc cca 266Gln
Tyr Ser Ser Leu Ser Thr Gln Pro Ile Leu Cys Ala Ser Ile Pro
35 40 45ggc ctg gta ccg aag cag ctg
cgc ttc tgc agg aac tac gtg gag atc 314Gly Leu Val Pro Lys Gln Leu
Arg Phe Cys Arg Asn Tyr Val Glu Ile 50 55
60atg ccc agc gtg gct gag ggt gtc aaa gcg ggc atc cag gag
tgc cag 362Met Pro Ser Val Ala Glu Gly Val Lys Ala Gly Ile Gln Glu
Cys Gln 65 70 75cac cag ttc cga
ggc cgg cgt tgg aac tgc acc acc gtc agc aac agc 410His Gln Phe Arg
Gly Arg Arg Trp Asn Cys Thr Thr Val Ser Asn Ser 80 85
90ctg gcc atc ttt ggc cct gtt ctg gac aaa gcc acc cgg
gag tca gcc 458Leu Ala Ile Phe Gly Pro Val Leu Asp Lys Ala Thr Arg
Glu Ser Ala95 100 105
110ttt gtc cat gcc atc gcc tcc gct gga gta gct ttc gca gtg aca cgc
506Phe Val His Ala Ile Ala Ser Ala Gly Val Ala Phe Ala Val Thr Arg
115 120 125tcc tgt gca gag gga
tca gct gct atc tgt ggg tgc agc agc cgc ctc 554Ser Cys Ala Glu Gly
Ser Ala Ala Ile Cys Gly Cys Ser Ser Arg Leu 130
135 140cag ggc tcc cca ggc gag ggc tgg aag tgg ggc ggc
tgt agt gag gac 602Gln Gly Ser Pro Gly Glu Gly Trp Lys Trp Gly Gly
Cys Ser Glu Asp 145 150 155att gaa
ttt gga gga atg gtc tct cgg gag ttt gcc gat gcc agg gag 650Ile Glu
Phe Gly Gly Met Val Ser Arg Glu Phe Ala Asp Ala Arg Glu 160
165 170aac cgg ccg gat gcc cgc tct gcc atg aac cgt
cac aac aat gag gct 698Asn Arg Pro Asp Ala Arg Ser Ala Met Asn Arg
His Asn Asn Glu Ala175 180 185
190ggg cgc cag gcc atc gcc agt cac atg cac ctc aag tgc aaa tgc cac
746Gly Arg Gln Ala Ile Ala Ser His Met His Leu Lys Cys Lys Cys His
195 200 205ggg cta tct ggc agc
tgt gaa gtg aag acc tgc tgg tgg tcg cag ccg 794Gly Leu Ser Gly Ser
Cys Glu Val Lys Thr Cys Trp Trp Ser Gln Pro 210
215 220gac ttc cgc acc atc ggg gat ttc ctc aag gac aag
tat gac agt gcc 842Asp Phe Arg Thr Ile Gly Asp Phe Leu Lys Asp Lys
Tyr Asp Ser Ala 225 230 235tcg gag
atg gtg gta gag aaa cac cga gag tct cgt ggc tgg gtg gag 890Ser Glu
Met Val Val Glu Lys His Arg Glu Ser Arg Gly Trp Val Glu 240
245 250acc ctg agg cca cgt tac acg tac ttc aag gtg
ccg aca gaa cgc gac 938Thr Leu Arg Pro Arg Tyr Thr Tyr Phe Lys Val
Pro Thr Glu Arg Asp255 260 265
270ctg gtc tac tac gag gcc tca ccc aac ttc tgc gaa cct aac ccc gaa
986Leu Val Tyr Tyr Glu Ala Ser Pro Asn Phe Cys Glu Pro Asn Pro Glu
275 280 285acc ggc tcc ttc ggg
acg cgt gac cgc acc tgc aat gtg agc tcg cat 1034Thr Gly Ser Phe Gly
Thr Arg Asp Arg Thr Cys Asn Val Ser Ser His 290
295 300ggc ata gat ggg tgc gac ctg ttg tgc tgc ggg cgc
ggg cat aac gcg 1082Gly Ile Asp Gly Cys Asp Leu Leu Cys Cys Gly Arg
Gly His Asn Ala 305 310 315cgc act
gag cga cgg agg gag aaa tgc cac tgt gtt ttc cat tgg tgc 1130Arg Thr
Glu Arg Arg Arg Glu Lys Cys His Cys Val Phe His Trp Cys 320
325 330tgc tac gtc agc tgc cag gag tgc aca cgt gtc
tat gac gtg cac acc 1178Cys Tyr Val Ser Cys Gln Glu Cys Thr Arg Val
Tyr Asp Val His Thr335 340 345
350tgc aag tag gagagctcct aacacgggag cagggttcat tccgaggggc
1227Cys Lysaaggttccta cctgggggcg gggttcctac ttggaggggt ctcttacttg
gggactcggt 1287tcttacttga gggcggagat cctacctgtg agggtctcat acctaaggac
ccggtttctg 1347ccttcagcct gggctcctat ttgggatctg ggttcctttt taggggagaa
gctcctgtct 1407gggatacggg tttctgcccg agggtggggc tccacttggg gatggaattc
caatttgggc 1467cggaagtcct acctcaatgg cttggactcc tctcttgacc cgacagggct
caaatggaga 1527caggtaagct actccctcaa ctaggtgggg ttcgtgcgga tgggtgggag
gggagagatt 1587agggtccctc ctcccagagg cactgctcta tctagataca tgagagggtg
cttcagggtg 1647ggccctattt gggcttgagg atcccgtggg ggcggggctt caccccgact
gggtggaact 1707tttggagacc cccttccact ggggcaaggc ttcactgaag actcatggga
tggagctcca 1767cggaaggagg agttcctgag cgagcctggg ctctgagcag gccatccagc
tcccatctgg 1827cccctttcca gtcctggtgt aaggttcaac ctgcaagcct catctgcgca
gagcaggatc 1887tcctggcaga atgaggcatg gagaagaact caggggtgat accaagacct
aacaaacccc 1947gtgcctgggt acctctttta aagctctgca ccccttcttc aagggctttc
ctagtctcct 2007tggcagagct ttcctgagga agatttgcag tcccccagag ttcaagtgaa
cacccataga 2067acagaacaga ctctatcctg agtagagagg gttctctagg aatctctatg
gggactgcta 2127ggaaggatcc tgggcatgac agcctcgtat gatagcctgc atccgctctg
acacttaata 2187ctcagatctc ccgggaaacc cagctcatcc ggtccgtgat gtccatgccc
caaatgcctc 2247agagatgttg cctcactttg agttgtatga acttcggaga catggggaca
cagtcaagcc 2307gcagagccag ggttgtttca ggacccatct gattccccag agcctgctgt
tgaggcaatg 2367gtcaccagat ccgttggcca ccaccctgtc ccgagcttct ctagtgtctg
tctggcctgg 2427aagtgaggtg ctacatacag cccatctgcc acaagagctt cctgattggt
accactgtga 2487accgtccctc cccctccaga caggggaggg gatgtggcca tacaggagtg
tgcccggaga 2547gcgcggaaag aggaagagag gctgcacacg cgtggtgact gactgtcttc
tgcctggaac 2607tttgcgttcg cgcttgtaac tttattttca atgctgctat atccacccac
cactggattt 2667agacaaaagt gattttcttt tttttttttt cttttctttc tatgaaagaa
attattttag 2727tttatagtat gtttgtttca aataatgggg aaagtaaaaa gagagaaaaa
aaaaaaaaaa 2787aaaaaaaaaa aaaaaaaaaa aaaaaaa
281448352PRTMus musculus 48Met Ala Pro Leu Gly Tyr Leu Leu Val
Leu Cys Ser Leu Lys Gln Ala1 5 10
15Leu Gly Ser Tyr Pro Ile Trp Trp Ser Leu Ala Val Gly Pro Gln
Tyr 20 25 30Ser Ser Leu Ser
Thr Gln Pro Ile Leu Cys Ala Ser Ile Pro Gly Leu 35
40 45Val Pro Lys Gln Leu Arg Phe Cys Arg Asn Tyr Val
Glu Ile Met Pro 50 55 60Ser Val Ala
Glu Gly Val Lys Ala Gly Ile Gln Glu Cys Gln His Gln65 70
75 80Phe Arg Gly Arg Arg Trp Asn Cys
Thr Thr Val Ser Asn Ser Leu Ala 85 90
95Ile Phe Gly Pro Val Leu Asp Lys Ala Thr Arg Glu Ser Ala
Phe Val 100 105 110His Ala Ile
Ala Ser Ala Gly Val Ala Phe Ala Val Thr Arg Ser Cys 115
120 125Ala Glu Gly Ser Ala Ala Ile Cys Gly Cys Ser
Ser Arg Leu Gln Gly 130 135 140Ser Pro
Gly Glu Gly Trp Lys Trp Gly Gly Cys Ser Glu Asp Ile Glu145
150 155 160Phe Gly Gly Met Val Ser Arg
Glu Phe Ala Asp Ala Arg Glu Asn Arg 165
170 175Pro Asp Ala Arg Ser Ala Met Asn Arg His Asn Asn
Glu Ala Gly Arg 180 185 190Gln
Ala Ile Ala Ser His Met His Leu Lys Cys Lys Cys His Gly Leu 195
200 205Ser Gly Ser Cys Glu Val Lys Thr Cys
Trp Trp Ser Gln Pro Asp Phe 210 215
220Arg Thr Ile Gly Asp Phe Leu Lys Asp Lys Tyr Asp Ser Ala Ser Glu225
230 235 240Met Val Val Glu
Lys His Arg Glu Ser Arg Gly Trp Val Glu Thr Leu 245
250 255Arg Pro Arg Tyr Thr Tyr Phe Lys Val Pro
Thr Glu Arg Asp Leu Val 260 265
270Tyr Tyr Glu Ala Ser Pro Asn Phe Cys Glu Pro Asn Pro Glu Thr Gly
275 280 285Ser Phe Gly Thr Arg Asp Arg
Thr Cys Asn Val Ser Ser His Gly Ile 290 295
300Asp Gly Cys Asp Leu Leu Cys Cys Gly Arg Gly His Asn Ala Arg
Thr305 310 315 320Glu Arg
Arg Arg Glu Lys Cys His Cys Val Phe His Trp Cys Cys Tyr
325 330 335Val Ser Cys Gln Glu Cys Thr
Arg Val Tyr Asp Val His Thr Cys Lys 340 345
350491101DNAMus musculusCDS(46)..(1101) 49cgggagcctt
gcggccgctg ccccgggctg ggcgcgcacg gcacc atg agc ccc cgt 57
Met Ser Pro Arg
1tcg tgc ctg cgg tcc ctg cga ctc ctc gtc ttc
gcc gtg ttc tcg gcc 105Ser Cys Leu Arg Ser Leu Arg Leu Leu Val Phe
Ala Val Phe Ser Ala5 10 15
20gcc gcg agc aat tgg ctg tac ctg gcc aag ctg tca tcg gtg ggc agc
153Ala Ala Ser Asn Trp Leu Tyr Leu Ala Lys Leu Ser Ser Val Gly Ser
25 30 35atc tcc gaa gag gag
acg tgc gag aaa ctc aaa ggc ctg atc cag agg 201Ile Ser Glu Glu Glu
Thr Cys Glu Lys Leu Lys Gly Leu Ile Gln Arg 40
45 50cag gtg cag atg tgc aaa cgg aac ctt gag gtg atg
gac tca gtg cgc 249Gln Val Gln Met Cys Lys Arg Asn Leu Glu Val Met
Asp Ser Val Arg 55 60 65cgt ggt
gcc cag ctg gcc atc gag gag tgc caa tac cag ttc cgg aac 297Arg Gly
Ala Gln Leu Ala Ile Glu Glu Cys Gln Tyr Gln Phe Arg Asn 70
75 80cgg cgc tgg aac tgt tcc aca ctg gac tcc ctc
cct gtc ttt ggg aag 345Arg Arg Trp Asn Cys Ser Thr Leu Asp Ser Leu
Pro Val Phe Gly Lys85 90 95
100gtg gtg aca caa ggg acc cgg gag gcg gcc ttt gta tac gcc atc tct
393Val Val Thr Gln Gly Thr Arg Glu Ala Ala Phe Val Tyr Ala Ile Ser
105 110 115tca gca ggt gtg gcc
ttt gca gtg aca agg gca tgc agc agt gga gaa 441Ser Ala Gly Val Ala
Phe Ala Val Thr Arg Ala Cys Ser Ser Gly Glu 120
125 130ctg gag aag tgt ggc tgt gac cgg aca gtg cac ggg
gtc agc cca cag 489Leu Glu Lys Cys Gly Cys Asp Arg Thr Val His Gly
Val Ser Pro Gln 135 140 145ggc ttc
cag tgg tca gga tgc tcg gac aac atc gcc tat ggc gta gcc 537Gly Phe
Gln Trp Ser Gly Cys Ser Asp Asn Ile Ala Tyr Gly Val Ala 150
155 160ttc tca cag tcc ttt gtg gac gtc cgg gag agg
agc aag ggg gcc tcc 585Phe Ser Gln Ser Phe Val Asp Val Arg Glu Arg
Ser Lys Gly Ala Ser165 170 175
180tcc agc cgg gca ctc atg aat ctt cac aac aac gag gct ggc agg aag
633Ser Ser Arg Ala Leu Met Asn Leu His Asn Asn Glu Ala Gly Arg Lys
185 190 195gcc atc ttg aca cac
atg cgg gtg gag tgc aag tgt cac ggg gtg tcg 681Ala Ile Leu Thr His
Met Arg Val Glu Cys Lys Cys His Gly Val Ser 200
205 210ggc tcc tgc gag gta aag acg tgc tgg cgt gct gta
ccg ccc ttc cgc 729Gly Ser Cys Glu Val Lys Thr Cys Trp Arg Ala Val
Pro Pro Phe Arg 215 220 225cag gtt
ggc cac gcg cta aag gag aag ttt gac ggt gcc acg gag gtg 777Gln Val
Gly His Ala Leu Lys Glu Lys Phe Asp Gly Ala Thr Glu Val 230
235 240gag cca cga cgc gta ggc tcc tcc cgg gcg ctg
gtg cct cgg aat gca 825Glu Pro Arg Arg Val Gly Ser Ser Arg Ala Leu
Val Pro Arg Asn Ala245 250 255
260cag ttc aag cca cat aca gat gag gac ctg gta tac ctg gag cct agc
873Gln Phe Lys Pro His Thr Asp Glu Asp Leu Val Tyr Leu Glu Pro Ser
265 270 275ccg gac ttc tgt gag
cag gac atc cgc agt ggc gtg cta ggc acg agg 921Pro Asp Phe Cys Glu
Gln Asp Ile Arg Ser Gly Val Leu Gly Thr Arg 280
285 290ggc cgc acg tgc aac aag aca tct aaa gcc att gac
ggc tgc gag cta 969Gly Arg Thr Cys Asn Lys Thr Ser Lys Ala Ile Asp
Gly Cys Glu Leu 295 300 305ctg tgc
tgt ggc cgc ggc ttc cac aca gcg caa gtg gag ctg gcc gag 1017Leu Cys
Cys Gly Arg Gly Phe His Thr Ala Gln Val Glu Leu Ala Glu 310
315 320cgc tgt ggc tgc agg ttc cac tgg tgc tgc ttc
gtc aag tgc cgg cag 1065Arg Cys Gly Cys Arg Phe His Trp Cys Cys Phe
Val Lys Cys Arg Gln325 330 335
340tgc cag cgg ctc gtg gag atg cac acg tgc cgg tga
1101Cys Gln Arg Leu Val Glu Met His Thr Cys Arg 345
35050351PRTMus musculus 50Met Ser Pro Arg Ser Cys Leu Arg Ser
Leu Arg Leu Leu Val Phe Ala1 5 10
15Val Phe Ser Ala Ala Ala Ser Asn Trp Leu Tyr Leu Ala Lys Leu
Ser 20 25 30Ser Val Gly Ser
Ile Ser Glu Glu Glu Thr Cys Glu Lys Leu Lys Gly 35
40 45Leu Ile Gln Arg Gln Val Gln Met Cys Lys Arg Asn
Leu Glu Val Met 50 55 60Asp Ser Val
Arg Arg Gly Ala Gln Leu Ala Ile Glu Glu Cys Gln Tyr65 70
75 80Gln Phe Arg Asn Arg Arg Trp Asn
Cys Ser Thr Leu Asp Ser Leu Pro 85 90
95Val Phe Gly Lys Val Val Thr Gln Gly Thr Arg Glu Ala Ala
Phe Val 100 105 110Tyr Ala Ile
Ser Ser Ala Gly Val Ala Phe Ala Val Thr Arg Ala Cys 115
120 125Ser Ser Gly Glu Leu Glu Lys Cys Gly Cys Asp
Arg Thr Val His Gly 130 135 140Val Ser
Pro Gln Gly Phe Gln Trp Ser Gly Cys Ser Asp Asn Ile Ala145
150 155 160Tyr Gly Val Ala Phe Ser Gln
Ser Phe Val Asp Val Arg Glu Arg Ser 165
170 175Lys Gly Ala Ser Ser Ser Arg Ala Leu Met Asn Leu
His Asn Asn Glu 180 185 190Ala
Gly Arg Lys Ala Ile Leu Thr His Met Arg Val Glu Cys Lys Cys 195
200 205His Gly Val Ser Gly Ser Cys Glu Val
Lys Thr Cys Trp Arg Ala Val 210 215
220Pro Pro Phe Arg Gln Val Gly His Ala Leu Lys Glu Lys Phe Asp Gly225
230 235 240Ala Thr Glu Val
Glu Pro Arg Arg Val Gly Ser Ser Arg Ala Leu Val 245
250 255Pro Arg Asn Ala Gln Phe Lys Pro His Thr
Asp Glu Asp Leu Val Tyr 260 265
270Leu Glu Pro Ser Pro Asp Phe Cys Glu Gln Asp Ile Arg Ser Gly Val
275 280 285Leu Gly Thr Arg Gly Arg Thr
Cys Asn Lys Thr Ser Lys Ala Ile Asp 290 295
300Gly Cys Glu Leu Leu Cys Cys Gly Arg Gly Phe His Thr Ala Gln
Val305 310 315 320Glu Leu
Ala Glu Arg Cys Gly Cys Arg Phe His Trp Cys Cys Phe Val
325 330 335Lys Cys Arg Gln Cys Gln Arg
Leu Val Glu Met His Thr Cys Arg 340 345
350514273DNAMus musculusCDS(637)..(1779) 51agtccctgga agcagacgtt
tcggccacag acccagagag gaggagctga caatcaggag 60gcgtgagccg cctggagtct
gcagaattcg tggtgtgaat gaactggggg catcttgggc 120acagggattg ccccccctcc
ttccccgcct cgggccacag ttgagtagtg gggcattttt 180tttcaccttc ttgtgaagaa
ttttttttat tatttgttgt aaagtctttt gcacaatcac 240gcccacattt ggggttggaa
agccctaatt accgccgtcg ctgatggacg ttagagaggg 300agcgcctcgc cgcggaacag
tcgcctgcgc gccctcgtcg gacccgcggc tcctgcactg 360tgtccccgct cggccctgcg
cttgctgctc gcccgcgcgc gccggcgccc tctcggttcc 420tgggcacatt tccacgctat
accaactcct ctgcccgagt ccgggcgcca gtgctcgctt 480ccgctccggg tcgctgcgcc
cacccgacgc gcccaggagg actccgcagc cctgctttgg 540attgtccccc aaggcttaac
cccgacgctt cgcttgaatt cctcggccgc cttcgctcgg 600gtggcgactt cctctccgtg
ccccctcccc ctcgcc atg aag aag ccc att gga 654
Met Lys Lys Pro Ile Gly
1 5ata tta agc ccg gga gtg gct ttg ggg acc gct gga ggt
gcc atg tct 702Ile Leu Ser Pro Gly Val Ala Leu Gly Thr Ala Gly Gly
Ala Met Ser 10 15 20tcc aag
ttc ttc cta atg gct ttg gcc acg ttt ttc tcc ttc gcc cag 750Ser Lys
Phe Phe Leu Met Ala Leu Ala Thr Phe Phe Ser Phe Ala Gln 25
30 35gtt gtt ata gaa gct aat tct tgg tgg tct
cta ggt atg aat aac cct 798Val Val Ile Glu Ala Asn Ser Trp Trp Ser
Leu Gly Met Asn Asn Pro 40 45 50gtt
cag atg tca gaa gta tat atc ata ggt gca cag cct ctc tgc agc 846Val
Gln Met Ser Glu Val Tyr Ile Ile Gly Ala Gln Pro Leu Cys Ser55
60 65 70caa ctg gca gga ctt tct
caa gga cag aag aaa ctc tgc cac ttg tat 894Gln Leu Ala Gly Leu Ser
Gln Gly Gln Lys Lys Leu Cys His Leu Tyr 75
80 85cag gac cac atg cag tac att gga gaa ggt gcg aag
aca ggc atc aag 942Gln Asp His Met Gln Tyr Ile Gly Glu Gly Ala Lys
Thr Gly Ile Lys 90 95 100gaa
tgc cag tac cag ttc cgg cat cgg aga tgg aac tgc agc aca gtg 990Glu
Cys Gln Tyr Gln Phe Arg His Arg Arg Trp Asn Cys Ser Thr Val 105
110 115gac aat act tct gtc ttt ggc agg gtg
atg caa ata ggc agc cga gag 1038Asp Asn Thr Ser Val Phe Gly Arg Val
Met Gln Ile Gly Ser Arg Glu 120 125
130acg gcc ttc acg tac gcg gtg agc gca gct ggg gtg gtg aac gcc atg
1086Thr Ala Phe Thr Tyr Ala Val Ser Ala Ala Gly Val Val Asn Ala Met135
140 145 150agc cga gca tgc
cgg gag ggc gag ctg tct acc tgt ggc tgc agc cgc 1134Ser Arg Ala Cys
Arg Glu Gly Glu Leu Ser Thr Cys Gly Cys Ser Arg 155
160 165gct gcg cgc ccc aag gac ctg cct cgg gac
tgg ttg tgg ggc ggc tgc 1182Ala Ala Arg Pro Lys Asp Leu Pro Arg Asp
Trp Leu Trp Gly Gly Cys 170 175
180gga gac aac atc gac tat ggc tac cgc ttc gcc aag gag ttc gtg gac
1230Gly Asp Asn Ile Asp Tyr Gly Tyr Arg Phe Ala Lys Glu Phe Val Asp
185 190 195gct aga gaa agg gaa cga atc
cac gct aag ggt tcc tat gag agc gca 1278Ala Arg Glu Arg Glu Arg Ile
His Ala Lys Gly Ser Tyr Glu Ser Ala 200 205
210cgc atc ctc atg aac tta cac aac aat gaa gca ggc cgt agg aca gta
1326Arg Ile Leu Met Asn Leu His Asn Asn Glu Ala Gly Arg Arg Thr Val215
220 225 230tac aac ctg gca
gat gta gcc tgt aag tgt cat gga gtg tct ggc tcc 1374Tyr Asn Leu Ala
Asp Val Ala Cys Lys Cys His Gly Val Ser Gly Ser 235
240 245tgt agc ctc aag acg tgc tgg ctg cag ctg
gcg gac ttc cgg aag gtg 1422Cys Ser Leu Lys Thr Cys Trp Leu Gln Leu
Ala Asp Phe Arg Lys Val 250 255
260ggc gat gcc ctc aag gag aag tat gat agc gcg gcg gcc atg agg ctc
1470Gly Asp Ala Leu Lys Glu Lys Tyr Asp Ser Ala Ala Ala Met Arg Leu
265 270 275aac agc cgg ggc aag ctg gtg
cag gtc aac agc cgc ttc aac tcc ccg 1518Asn Ser Arg Gly Lys Leu Val
Gln Val Asn Ser Arg Phe Asn Ser Pro 280 285
290acc acg cag gac ctg gtc tac atc gac ccc agt ccg gac tac tgt gtg
1566Thr Thr Gln Asp Leu Val Tyr Ile Asp Pro Ser Pro Asp Tyr Cys Val295
300 305 310cgc aac gag agc
act ggc tcg ctg ggc acg cag gga cgc ctg tgc aac 1614Arg Asn Glu Ser
Thr Gly Ser Leu Gly Thr Gln Gly Arg Leu Cys Asn 315
320 325aag acc tca gag ggg atg gac ggc tgc gag
ctc atg tgc tgt ggg cgt 1662Lys Thr Ser Glu Gly Met Asp Gly Cys Glu
Leu Met Cys Cys Gly Arg 330 335
340ggc tat gac cag ttt aag aca gtg cag acc gaa cgc tgt cat tgc aag
1710Gly Tyr Asp Gln Phe Lys Thr Val Gln Thr Glu Arg Cys His Cys Lys
345 350 355ttt cac tgg tgc tgc tat gtc
aaa tgc aag aag tgc acg gag att gtg 1758Phe His Trp Cys Cys Tyr Val
Lys Cys Lys Lys Cys Thr Glu Ile Val 360 365
370gat cag ttc gtg tgc aaa tag tggtgtgcct gcccttcacc cagtcccact
1809Asp Gln Phe Val Cys Lys375 380cccaggaccc acttatttat
agaaagtaca gtgcttctgg ttctttttat ttctccccca 1869agaattgcag ctggaaccat
gtgttttgtt ttgttttatt ttgttttttc ttttctgtta 1929ccatctaaga actctgtggt
ttattattaa tattataatt aatatttggc aatagtgggg 1989gaaactaaga aaaatattta
ttttgaggat ctttgcaaag ttagtacaaa atttctttct 2049tctgatgcta caggataaag
gggaaaaact atgtattcga acttagctgt gcagttgggg 2109gttcacatct agaaggtgta
ggagccattt tcttctcaaa cagagagtcc tttgagatgg 2169gtggtatcca ggtgaaggag
gaggtacaga cccatgaata acagttcctg tgaccaaaat 2229gaattgcagg tgctctggta
caaaagatct taaatataga tatattaaat atacatatat 2289gccaaaaata cagaatatga
gacactccct aacccagagg ttaccagcct ggttttgtgg 2349gttttttgtt ttgttttgtt
ttttcttttt ttgggttttg tttgtttgtt tgtttgtttg 2409tatttttggt gtgtgtgtgt
gtgtatttct agaatgatct tttagaaggt acaagcaaga 2469atctcatatc ttcagaagca
ggcatatcat gtatgttact gtgtcccacc tacagatact 2529ccattcatga atgggccttt
ttctaacagt tcatgaatat tggggagccg gtgggctggg 2589ggagggaggt ccccagaaat
tagaaaactt gaagtttcct acattgaggc cataatcttg 2649tgttagccca gctgattctt
aataccagac ttttagatcc ataaaggaat ttttgactaa 2709aaaaaaaaaa tcttgttttg
aaagccatct tattttctta aaaatgaaaa attacccatg 2769aatcccattt gcaacccctc
acccccacag gcaacaagaa agtcccatgt agttgagcac 2829tgcgaacacc tctgtgagga
gatgatggca gccatcttcc tgcatgatcc catgcccttt 2889ctggactctc tgctggccat
gcttccgaat ggcagccctg gtggacactc actgctggta 2949gggcagaaaa tgtacacgag
gagccatgtt cagaaccagc cacttagggg ttgttctctg 3009aggcttttct ttggaggtac
ggtaacttga tgtgttttga tgatatctct tggcccaggg 3069agtccacaga ggtgttgcag
ctgtttggtt gttatcctcc tgcgtttaga ctttccattt 3129gtgcttttcc tattaccctg
caggtgtacc ctaaaactgt tcctagtgta cttgaacagt 3189tgcatttata aggggggatg
tggtttaatg gtgcctgata tctcagtttt tttgtatata 3249acatatatat aaatatacat
atataaatat agatataatt atatctcagt gcagtctggg 3309atttagacct acagttttct
ctgggcttgc tctctgcctg gagtatcgtc cttcattgca 3369gtccaattgg gatttctttt
tttccaaaaa ttttgagtct taacattgac ctgtgacagg 3429atcctaccac gaataccagg
aagcaagcta agactcggag gaagctctca gggctcatgt 3489cctgaatgta tgttggttag
aaagtagcct ttctgcttcc tgcccatggc cagttctcca 3549ccctctcttt ggtgttcttt
gtggggaggg cactgtggtt tgtcgcagcc ctggacttcg 3609agaggctccc agaacccagg
atcaccagcc tcctgtctgt ttgcttcact cctttcccag 3669ggaggacttg ggactgtcct
gtctgacagg acggatctga gttcccgaag caaaccagct 3729caccacatag atagctagtt
taaacaatgt tttaaaataa gggcacctct gtttcaaaag 3789tgacatctgc tgtgttgttt
tcgaggcctg atactcttac aaggtttgaa aaaaaatgtg 3849tgtatccatt catgggcttg
gtagccttct ggtcacctca gtcctgtggc tcttaactta 3909ttgcccaaca atattcattt
cccctcagct acaatgaatt gcaagcaaaa gatgttgaaa 3969aaaagcacta atttagttta
aaatgtcact ttttggtttt tattctacaa aaaccatgaa 4029gttctctctc tctctctctc
tctctcttat ttgttaaatc agattatgtt ctttttttgt 4089ttttgttttt agtgattcat
gtttatgagc agagtggagt ttaacaatcc tagctttaaa 4149aaaaacctat ttaatgtaag
atattctacg catccttcag atattttgta tatcccctat 4209ggcctttatt ctgtactttt
aatgtacata tttctgtctt gtgtgatttg tatatttcac 4269tggt
427352380PRTMus musculus
52Met Lys Lys Pro Ile Gly Ile Leu Ser Pro Gly Val Ala Leu Gly Thr1
5 10 15Ala Gly Gly Ala Met Ser
Ser Lys Phe Phe Leu Met Ala Leu Ala Thr 20 25
30Phe Phe Ser Phe Ala Gln Val Val Ile Glu Ala Asn Ser
Trp Trp Ser 35 40 45Leu Gly Met
Asn Asn Pro Val Gln Met Ser Glu Val Tyr Ile Ile Gly 50
55 60Ala Gln Pro Leu Cys Ser Gln Leu Ala Gly Leu Ser
Gln Gly Gln Lys65 70 75
80Lys Leu Cys His Leu Tyr Gln Asp His Met Gln Tyr Ile Gly Glu Gly
85 90 95Ala Lys Thr Gly Ile Lys
Glu Cys Gln Tyr Gln Phe Arg His Arg Arg 100
105 110Trp Asn Cys Ser Thr Val Asp Asn Thr Ser Val Phe
Gly Arg Val Met 115 120 125Gln Ile
Gly Ser Arg Glu Thr Ala Phe Thr Tyr Ala Val Ser Ala Ala 130
135 140Gly Val Val Asn Ala Met Ser Arg Ala Cys Arg
Glu Gly Glu Leu Ser145 150 155
160Thr Cys Gly Cys Ser Arg Ala Ala Arg Pro Lys Asp Leu Pro Arg Asp
165 170 175Trp Leu Trp Gly
Gly Cys Gly Asp Asn Ile Asp Tyr Gly Tyr Arg Phe 180
185 190Ala Lys Glu Phe Val Asp Ala Arg Glu Arg Glu
Arg Ile His Ala Lys 195 200 205Gly
Ser Tyr Glu Ser Ala Arg Ile Leu Met Asn Leu His Asn Asn Glu 210
215 220Ala Gly Arg Arg Thr Val Tyr Asn Leu Ala
Asp Val Ala Cys Lys Cys225 230 235
240His Gly Val Ser Gly Ser Cys Ser Leu Lys Thr Cys Trp Leu Gln
Leu 245 250 255Ala Asp Phe
Arg Lys Val Gly Asp Ala Leu Lys Glu Lys Tyr Asp Ser 260
265 270Ala Ala Ala Met Arg Leu Asn Ser Arg Gly
Lys Leu Val Gln Val Asn 275 280
285Ser Arg Phe Asn Ser Pro Thr Thr Gln Asp Leu Val Tyr Ile Asp Pro 290
295 300Ser Pro Asp Tyr Cys Val Arg Asn
Glu Ser Thr Gly Ser Leu Gly Thr305 310
315 320Gln Gly Arg Leu Cys Asn Lys Thr Ser Glu Gly Met
Asp Gly Cys Glu 325 330
335Leu Met Cys Cys Gly Arg Gly Tyr Asp Gln Phe Lys Thr Val Gln Thr
340 345 350Glu Arg Cys His Cys Lys
Phe His Trp Cys Cys Tyr Val Lys Cys Lys 355 360
365Lys Cys Thr Glu Ile Val Asp Gln Phe Val Cys Lys 370
375 380532129DNAMus musculusCDS(132)..(1250)
53ccttgctgct tctcattcca tgagctgggg agagacagtg tggaagtcaa accatgtgtt
60tcttgagagc aggtgctgct ggggctccct gaatggcggc taggtgccaa gagggagctc
120cgctttggaa g atg ttg gtc cca ggg cat tgg gat ggg ttg agg ccg gcc
170 Met Leu Val Pro Gly His Trp Asp Gly Leu Arg Pro Ala
1 5 10atg ccc agc ctg ctg ctg gtg
gtc gtg gca gct ctg ctc tcc agc tgg 218Met Pro Ser Leu Leu Leu Val
Val Val Ala Ala Leu Leu Ser Ser Trp 15 20
25gca cag ctg ctg act gac gcc aac tcc tgg tgg tca cta gct ctg aac
266Ala Gln Leu Leu Thr Asp Ala Asn Ser Trp Trp Ser Leu Ala Leu Asn30
35 40 45cca gtg cag aga
ccg gag atg ttc atc att ggc gct cag ccc gtg tgc 314Pro Val Gln Arg
Pro Glu Met Phe Ile Ile Gly Ala Gln Pro Val Cys 50
55 60agc caa ctt cct ggg ctt tcc cca ggc cag
aga aag ctg tgt cag ttg 362Ser Gln Leu Pro Gly Leu Ser Pro Gly Gln
Arg Lys Leu Cys Gln Leu 65 70
75tat cag gag cac atg tcc tac atc ggg gag gga gcc aag acg ggc atc
410Tyr Gln Glu His Met Ser Tyr Ile Gly Glu Gly Ala Lys Thr Gly Ile
80 85 90aga gag tgc caa cac cag ttt cga
cag agg cgc tgg aac tgc agc acc 458Arg Glu Cys Gln His Gln Phe Arg
Gln Arg Arg Trp Asn Cys Ser Thr 95 100
105gtg gac aac aca tct gtc ttt ggc aga gtt atg cag ata ggt agc cga
506Val Asp Asn Thr Ser Val Phe Gly Arg Val Met Gln Ile Gly Ser Arg110
115 120 125gag act gcc ttc
acg tat gca gtg agc gcc gct ggc gtg gtg aat gcc 554Glu Thr Ala Phe
Thr Tyr Ala Val Ser Ala Ala Gly Val Val Asn Ala 130
135 140atc agc cga gcc tgc aga gag ggt gag ctg
tcc acc tgt ggc tgc agc 602Ile Ser Arg Ala Cys Arg Glu Gly Glu Leu
Ser Thr Cys Gly Cys Ser 145 150
155cgt gct gcg agg ccc aag gac ctg cct cgg gac tgg ctg tgg ggt ggc
650Arg Ala Ala Arg Pro Lys Asp Leu Pro Arg Asp Trp Leu Trp Gly Gly
160 165 170tgt gga gac aac gtg gag tac
ggc tac cgc ttt gcc aag gag ttt gtg 698Cys Gly Asp Asn Val Glu Tyr
Gly Tyr Arg Phe Ala Lys Glu Phe Val 175 180
185gat gcc cga gag cgt gag aag aac ttt gcc aag gga tcg gag gag cag
746Asp Ala Arg Glu Arg Glu Lys Asn Phe Ala Lys Gly Ser Glu Glu Gln190
195 200 205ggc cga gct ctc
atg aac cta cag aac aac gag gct ggc cgc cgg gcc 794Gly Arg Ala Leu
Met Asn Leu Gln Asn Asn Glu Ala Gly Arg Arg Ala 210
215 220gtg tat aag atg gct gat gtc gcc tgc aaa
tgt cac gga gtc tcc ggg 842Val Tyr Lys Met Ala Asp Val Ala Cys Lys
Cys His Gly Val Ser Gly 225 230
235tcc tgc agc ctc aag acc tgc tgg ctc cag ctg gcc gag ttc cgc aag
890Ser Cys Ser Leu Lys Thr Cys Trp Leu Gln Leu Ala Glu Phe Arg Lys
240 245 250gtt ggg gac cgt ttg aag gag
aag tac gac agc gcc gcg gcc atg cgc 938Val Gly Asp Arg Leu Lys Glu
Lys Tyr Asp Ser Ala Ala Ala Met Arg 255 260
265atc acc cgc cag ggc aag ctg gag ctg gcc aac agc cgc ttc aac cag
986Ile Thr Arg Gln Gly Lys Leu Glu Leu Ala Asn Ser Arg Phe Asn Gln270
275 280 285ccc acc cca gag
gac ctg gtc tac gtg gac ccc agt cct gac tac tgc 1034Pro Thr Pro Glu
Asp Leu Val Tyr Val Asp Pro Ser Pro Asp Tyr Cys 290
295 300ttg cgt aat gag acc aca ggc tcc ctg ggc
acc cag ggt cgc ctc tgc 1082Leu Arg Asn Glu Thr Thr Gly Ser Leu Gly
Thr Gln Gly Arg Leu Cys 305 310
315aac aag acc tca gag ggc atg gac ggc tgc gag ctc atg tgc tgt ggc
1130Asn Lys Thr Ser Glu Gly Met Asp Gly Cys Glu Leu Met Cys Cys Gly
320 325 330cgc ggc tat gac cgc ttc aag
agc gtt cag gtg gaa cgc tgc cac tgc 1178Arg Gly Tyr Asp Arg Phe Lys
Ser Val Gln Val Glu Arg Cys His Cys 335 340
345agg ttc cac tgg tgt tgc ttt gtc aga tgc aaa aaa tgc acc gag gtt
1226Arg Phe His Trp Cys Cys Phe Val Arg Cys Lys Lys Cys Thr Glu Val350
355 360 365gtg gac cag tat
gtc tgt aag tga ctgcaccaca cgggccttca ggccgctcct 1280Val Asp Gln Tyr
Val Cys Lys 370ctccgcctta caaaagtcta tattatataa atctatctaa
atatatttta tatttgtaca 1340aatggatgga tggatggatg atagataatc aagagaagaa
agtggagagg aagagcttag 1400gagatgctgg ccctctgtga ggactggatt ttgctggaaa
tccacaacca gtgggagaga 1460aacgggcttt tccccatttt ctggccagga cttttgggac
atgggcttga gagtgtctgt 1520gtgccatagc ctccaggagt caggtgggga ttagatgaag
gaactggact tattccacat 1580ctacagtcct gtggggaaga tgagtgtctg tgaccctggc
caggagaccc agaggccctg 1640tggaaagacc tgataactgg gatggtagcc taggtcttcc
tgaaaatgga gccagctttg 1700ggaaggggct ctgtacttcc ttcttttctc atctgagtac
acactgcagg aaagtcccct 1760gccccaatat gggggagtgg tctcaagtca ctccaaccgg
tgaccgtaag agatctgggc 1820ctccctggac cctggctctg ccttctgatg agaatgtcac
tagctcctgc ctcaagctct 1880tgtgccaaga gaaagactgt tccgtcacct gctacagcca
ggaagacgtg gagcaaacct 1940gggttttgac tggggaccaa gtgcctgttg cacaggacag
gaatctgctg tcactctgtc 2000aagggaggct ttgagaatga cagggcatgc tagcaggtca
ggtcaactgc ctgtgagact 2060gtcatctctg cccacatgta cagcgtccct ctgacattaa
atatcttttt actgaaaaaa 2120aaaaaaaaa
212954372PRTMus musculus 54Met Leu Val Pro Gly His
Trp Asp Gly Leu Arg Pro Ala Met Pro Ser1 5
10 15Leu Leu Leu Val Val Val Ala Ala Leu Leu Ser Ser
Trp Ala Gln Leu 20 25 30Leu
Thr Asp Ala Asn Ser Trp Trp Ser Leu Ala Leu Asn Pro Val Gln 35
40 45Arg Pro Glu Met Phe Ile Ile Gly Ala
Gln Pro Val Cys Ser Gln Leu 50 55
60Pro Gly Leu Ser Pro Gly Gln Arg Lys Leu Cys Gln Leu Tyr Gln Glu65
70 75 80His Met Ser Tyr Ile
Gly Glu Gly Ala Lys Thr Gly Ile Arg Glu Cys 85
90 95Gln His Gln Phe Arg Gln Arg Arg Trp Asn Cys
Ser Thr Val Asp Asn 100 105
110Thr Ser Val Phe Gly Arg Val Met Gln Ile Gly Ser Arg Glu Thr Ala
115 120 125Phe Thr Tyr Ala Val Ser Ala
Ala Gly Val Val Asn Ala Ile Ser Arg 130 135
140Ala Cys Arg Glu Gly Glu Leu Ser Thr Cys Gly Cys Ser Arg Ala
Ala145 150 155 160Arg Pro
Lys Asp Leu Pro Arg Asp Trp Leu Trp Gly Gly Cys Gly Asp
165 170 175Asn Val Glu Tyr Gly Tyr Arg
Phe Ala Lys Glu Phe Val Asp Ala Arg 180 185
190Glu Arg Glu Lys Asn Phe Ala Lys Gly Ser Glu Glu Gln Gly
Arg Ala 195 200 205Leu Met Asn Leu
Gln Asn Asn Glu Ala Gly Arg Arg Ala Val Tyr Lys 210
215 220Met Ala Asp Val Ala Cys Lys Cys His Gly Val Ser
Gly Ser Cys Ser225 230 235
240Leu Lys Thr Cys Trp Leu Gln Leu Ala Glu Phe Arg Lys Val Gly Asp
245 250 255Arg Leu Lys Glu Lys
Tyr Asp Ser Ala Ala Ala Met Arg Ile Thr Arg 260
265 270Gln Gly Lys Leu Glu Leu Ala Asn Ser Arg Phe Asn
Gln Pro Thr Pro 275 280 285Glu Asp
Leu Val Tyr Val Asp Pro Ser Pro Asp Tyr Cys Leu Arg Asn 290
295 300Glu Thr Thr Gly Ser Leu Gly Thr Gln Gly Arg
Leu Cys Asn Lys Thr305 310 315
320Ser Glu Gly Met Asp Gly Cys Glu Leu Met Cys Cys Gly Arg Gly Tyr
325 330 335Asp Arg Phe Lys
Ser Val Gln Val Glu Arg Cys His Cys Arg Phe His 340
345 350Trp Cys Cys Phe Val Arg Cys Lys Lys Cys Thr
Glu Val Val Asp Gln 355 360 365Tyr
Val Cys Lys 370551669DNAMus musculusCDS(189)..(1283) 55ccgccgcgcc
ctcctcgccc gggatgggcc cccccgccgc caccgccgcc ggagccctag 60tctccgggcc
gccgcctcgg tcgccgcgtt tgccctgaag cccggtgccc gcgcgccccg 120gctcaccccg
cagcttcact ccccaccccc agccgcctcc ccggccagac tgcggtagag 180ctctcagg atg
ctg ccg ccg gtg ccc tcc cgc ctc gga ctg ctg ctg ctg 230 Met
Leu Pro Pro Val Pro Ser Arg Leu Gly Leu Leu Leu Leu 1
5 10ctc ttg tgc ccc gcg cac gtc gat gga ctg tgg tgg
gcc gtg ggc agc 278Leu Leu Cys Pro Ala His Val Asp Gly Leu Trp Trp
Ala Val Gly Ser15 20 25
30ccc ttg gtc atg gat cct acc agc atc tgc agg aag gcc agg cgg ctg
326Pro Leu Val Met Asp Pro Thr Ser Ile Cys Arg Lys Ala Arg Arg Leu
35 40 45gca gga aga cag gcc gag
ctg tgc cag gcg gag ccg gaa gta gtg gca 374Ala Gly Arg Gln Ala Glu
Leu Cys Gln Ala Glu Pro Glu Val Val Ala 50 55
60gag ctt gcc cga ggc gca aga ctg ggg gtt cga gaa tgt
cag ttc cag 422Glu Leu Ala Arg Gly Ala Arg Leu Gly Val Arg Glu Cys
Gln Phe Gln 65 70 75ttc cgt ttc
cga cgc tgg aac tgc tcc agc cac agc aag gcc ttt ggg 470Phe Arg Phe
Arg Arg Trp Asn Cys Ser Ser His Ser Lys Ala Phe Gly 80
85 90cgc gtc ctg cag cag gac atc cga gag aca gct ttc
gtg ttt gca atc 518Arg Val Leu Gln Gln Asp Ile Arg Glu Thr Ala Phe
Val Phe Ala Ile95 100 105
110acc gca gct ggt gcc agc cac gcg gtc act caa gcc tgt tcc atg gga
566Thr Ala Ala Gly Ala Ser His Ala Val Thr Gln Ala Cys Ser Met Gly
115 120 125gag ctc cta cag tgt
ggt tgt cag gca ccc cgc ggg cgg gca ccg cct 614Glu Leu Leu Gln Cys
Gly Cys Gln Ala Pro Arg Gly Arg Ala Pro Pro 130
135 140agg ccc tcc ggc ctt ctg ggc act cct gga cct cca
gga cca act ggc 662Arg Pro Ser Gly Leu Leu Gly Thr Pro Gly Pro Pro
Gly Pro Thr Gly 145 150 155tct cca
gat gct agc gca gcc tgg gag tgg gga ggc tgc gga gac gat 710Ser Pro
Asp Ala Ser Ala Ala Trp Glu Trp Gly Gly Cys Gly Asp Asp 160
165 170gtg gac ttc ggg gat gag aag tca aga ctc ttt
atg gat gcg cag cac 758Val Asp Phe Gly Asp Glu Lys Ser Arg Leu Phe
Met Asp Ala Gln His175 180 185
190aag cgg ggc cgt gga gat atc cgt gca ttg gtg caa ctg cac aac aac
806Lys Arg Gly Arg Gly Asp Ile Arg Ala Leu Val Gln Leu His Asn Asn
195 200 205gag gcg ggc agg ctg
gcg gtg cgg agt cac acg cgc acc gag tgt aag 854Glu Ala Gly Arg Leu
Ala Val Arg Ser His Thr Arg Thr Glu Cys Lys 210
215 220tgc cat ggg ctt tcg ggt tcc tgc gct ctg cgc acc
tgc tgg cag aag 902Cys His Gly Leu Ser Gly Ser Cys Ala Leu Arg Thr
Cys Trp Gln Lys 225 230 235ctg cct
ccg ttc cgc gag gtg ggc gca cgg ctg ctg gag cgc ttc cac 950Leu Pro
Pro Phe Arg Glu Val Gly Ala Arg Leu Leu Glu Arg Phe His 240
245 250ggc gcc tcg cgc gtc atg ggc acc aac gac ggc
aaa gct ctg ctg cct 998Gly Ala Ser Arg Val Met Gly Thr Asn Asp Gly
Lys Ala Leu Leu Pro255 260 265
270gcg gtc cgc aca ctc aag cct ccc gga cga gcg gat ctc ctc tac gca
1046Ala Val Arg Thr Leu Lys Pro Pro Gly Arg Ala Asp Leu Leu Tyr Ala
275 280 285gcc gat tca ccc gac
ttc tgc gcc ccc aac cgg cgc acg ggt tcg ccg 1094Ala Asp Ser Pro Asp
Phe Cys Ala Pro Asn Arg Arg Thr Gly Ser Pro 290
295 300ggc acg cgc gga cgc gcc tgc aac agc agt gcc ccg
gac ctc agc ggc 1142Gly Thr Arg Gly Arg Ala Cys Asn Ser Ser Ala Pro
Asp Leu Ser Gly 305 310 315tgc gac
ctg ttg tgc tgc ggt cgc ggg cac cgc cag gag agc gta cag 1190Cys Asp
Leu Leu Cys Cys Gly Arg Gly His Arg Gln Glu Ser Val Gln 320
325 330ctc gag gag aac tgt ctg tgc cgc ttc cac tgg
tgc tgc gtg gtg caa 1238Leu Glu Glu Asn Cys Leu Cys Arg Phe His Trp
Cys Cys Val Val Gln335 340 345
350tgc cac cgc tgc cgg gtg cgc aag gaa ctc agc ctg tgc ctc tga
1283Cys His Arg Cys Arg Val Arg Lys Glu Leu Ser Leu Cys Leu
355 360 365cccgtcgcct gcctcggaac
tgctggcagc cacctctggg ccatctacag gactattaga 1343ttccagcagg gggcgctgtc
tgagtccagc agctccctag gaaaagtacc tatccaggcc 1403ttgggaaatt acaggggcca
gccaggaact tggggtttac accagcccac gaaagcccgg 1463gggaacatac ccctccagca
ttcccctgaa aggccctttg ctagttcctg caggagatca 1523ctccccttgg ccccccagat
ggaaataaga aagccagact ctgccctctg gaaataatat 1583tcctcagaat tactgggatg
gatgggtgag tttagtatca ataaagacat ttaaatccac 1643aaaaaaaaaa aaaaaaaaaa
aaaaaa 166956364PRTMus musculus
56Met Leu Pro Pro Val Pro Ser Arg Leu Gly Leu Leu Leu Leu Leu Leu1
5 10 15Cys Pro Ala His Val Asp
Gly Leu Trp Trp Ala Val Gly Ser Pro Leu 20 25
30Val Met Asp Pro Thr Ser Ile Cys Arg Lys Ala Arg Arg
Leu Ala Gly 35 40 45Arg Gln Ala
Glu Leu Cys Gln Ala Glu Pro Glu Val Val Ala Glu Leu 50
55 60Ala Arg Gly Ala Arg Leu Gly Val Arg Glu Cys Gln
Phe Gln Phe Arg65 70 75
80Phe Arg Arg Trp Asn Cys Ser Ser His Ser Lys Ala Phe Gly Arg Val
85 90 95Leu Gln Gln Asp Ile Arg
Glu Thr Ala Phe Val Phe Ala Ile Thr Ala 100
105 110Ala Gly Ala Ser His Ala Val Thr Gln Ala Cys Ser
Met Gly Glu Leu 115 120 125Leu Gln
Cys Gly Cys Gln Ala Pro Arg Gly Arg Ala Pro Pro Arg Pro 130
135 140Ser Gly Leu Leu Gly Thr Pro Gly Pro Pro Gly
Pro Thr Gly Ser Pro145 150 155
160Asp Ala Ser Ala Ala Trp Glu Trp Gly Gly Cys Gly Asp Asp Val Asp
165 170 175Phe Gly Asp Glu
Lys Ser Arg Leu Phe Met Asp Ala Gln His Lys Arg 180
185 190Gly Arg Gly Asp Ile Arg Ala Leu Val Gln Leu
His Asn Asn Glu Ala 195 200 205Gly
Arg Leu Ala Val Arg Ser His Thr Arg Thr Glu Cys Lys Cys His 210
215 220Gly Leu Ser Gly Ser Cys Ala Leu Arg Thr
Cys Trp Gln Lys Leu Pro225 230 235
240Pro Phe Arg Glu Val Gly Ala Arg Leu Leu Glu Arg Phe His Gly
Ala 245 250 255Ser Arg Val
Met Gly Thr Asn Asp Gly Lys Ala Leu Leu Pro Ala Val 260
265 270Arg Thr Leu Lys Pro Pro Gly Arg Ala Asp
Leu Leu Tyr Ala Ala Asp 275 280
285Ser Pro Asp Phe Cys Ala Pro Asn Arg Arg Thr Gly Ser Pro Gly Thr 290
295 300Arg Gly Arg Ala Cys Asn Ser Ser
Ala Pro Asp Leu Ser Gly Cys Asp305 310
315 320Leu Leu Cys Cys Gly Arg Gly His Arg Gln Glu Ser
Val Gln Leu Glu 325 330
335Glu Asn Cys Leu Cys Arg Phe His Trp Cys Cys Val Val Gln Cys His
340 345 350Arg Cys Arg Val Arg Lys
Glu Leu Ser Leu Cys Leu 355 360573189DNAMus
musculusCDS(255)..(1304) 57cccgcgcctc aaacacttgc cgcgatcgct ggcgcgcagc
ggcgcccctt gttgcgcttg 60ttctcccctc ctctggctcc gcggctcccg cgctctggga
cagtctccag tgcctagcgc 120ggaccgacgc accgacggac cgcccaggga gcctcggccc
gcgccccctg cgcaggctat 180gtggattgcc ccgccgggcc cggctggcgg gatcagcaca
gcccggcccg tggcacccgc 240caccagcggg gact atg acc cgg aaa gcg cgg cgc
tgc ctg ggc cac ctc 290 Met Thr Arg Lys Ala Arg Arg
Cys Leu Gly His Leu 1 5
10ttt ctc agc ctg ggc ata gtc tac ctc cgg atc ggt ggc ttc tct tcg
338Phe Leu Ser Leu Gly Ile Val Tyr Leu Arg Ile Gly Gly Phe Ser Ser
15 20 25gtg gta gct ctg ggt gcg agc atc
atc tgt aac aag atc cca ggc ctg 386Val Val Ala Leu Gly Ala Ser Ile
Ile Cys Asn Lys Ile Pro Gly Leu 30 35
40gct ccc aga cag cgg gca atc tgc cag agc cgg ccg gac gcc atc atc
434Ala Pro Arg Gln Arg Ala Ile Cys Gln Ser Arg Pro Asp Ala Ile Ile45
50 55 60gtc ata gga gaa ggc
tcc caa atg ggc ctg gac gag tgt cag ttt cag 482Val Ile Gly Glu Gly
Ser Gln Met Gly Leu Asp Glu Cys Gln Phe Gln 65
70 75ttc cga aat ggc cgt tgg aac tgc tca gcg ctg
gga gag cgt act gtc 530Phe Arg Asn Gly Arg Trp Asn Cys Ser Ala Leu
Gly Glu Arg Thr Val 80 85
90ttc ggg aag gag ctc aaa gtg ggg agt cgg gag gct gcc ttc acc tat
578Phe Gly Lys Glu Leu Lys Val Gly Ser Arg Glu Ala Ala Phe Thr Tyr
95 100 105gcg att atc gct gcg ggc gtg
gcc cat gcc atc act gct gcc tgc acc 626Ala Ile Ile Ala Ala Gly Val
Ala His Ala Ile Thr Ala Ala Cys Thr 110 115
120cag ggc aac ctg agc gac tgt ggc tgc gac aag gag aag caa ggc cag
674Gln Gly Asn Leu Ser Asp Cys Gly Cys Asp Lys Glu Lys Gln Gly Gln125
130 135 140tac cac cgg gac
gag ggc tgg aag tgg ggt ggc tgc tct gcc gac atc 722Tyr His Arg Asp
Glu Gly Trp Lys Trp Gly Gly Cys Ser Ala Asp Ile 145
150 155cgc tac ggc atc ggc ttc gcc aag gtc ttc
gtg gat gcc cgg gag atc 770Arg Tyr Gly Ile Gly Phe Ala Lys Val Phe
Val Asp Ala Arg Glu Ile 160 165
170aag cag aat gcc cgg acg ctc atg aac tta cac aat aac gag gcg ggt
818Lys Gln Asn Ala Arg Thr Leu Met Asn Leu His Asn Asn Glu Ala Gly
175 180 185cgg aag atc ctg gag gag aac
atg aag ctg gag tgt aag tgc cat ggt 866Arg Lys Ile Leu Glu Glu Asn
Met Lys Leu Glu Cys Lys Cys His Gly 190 195
200gtg tca ggc tcc tgt acc act aag acg tgc tgg acc aca ctg cca cag
914Val Ser Gly Ser Cys Thr Thr Lys Thr Cys Trp Thr Thr Leu Pro Gln205
210 215 220ttc cga gag cta
ggc tac gtg ctc aag gac aaa tac aac gag gcc gtc 962Phe Arg Glu Leu
Gly Tyr Val Leu Lys Asp Lys Tyr Asn Glu Ala Val 225
230 235cac gtg gag cct gtg cgt gcc agt cga aac
aag cgg ccc acc ttt ctg 1010His Val Glu Pro Val Arg Ala Ser Arg Asn
Lys Arg Pro Thr Phe Leu 240 245
250aag atc aag aag ccc ctg tcc tac cgc aag ccc atg gac act gac ctg
1058Lys Ile Lys Lys Pro Leu Ser Tyr Arg Lys Pro Met Asp Thr Asp Leu
255 260 265gtg tat atc gag aag tca ccc
aat tac tgt gaa gag gac cca gtg aca 1106Val Tyr Ile Glu Lys Ser Pro
Asn Tyr Cys Glu Glu Asp Pro Val Thr 270 275
280ggc agc gtg ggt acc cag ggc cga gcc tgc aat aag aca gcc cct cag
1154Gly Ser Val Gly Thr Gln Gly Arg Ala Cys Asn Lys Thr Ala Pro Gln285
290 295 300gcc agt ggc tgt
gac ctc atg tgc tgt ggc cgt ggc tac aac aca cac 1202Ala Ser Gly Cys
Asp Leu Met Cys Cys Gly Arg Gly Tyr Asn Thr His 305
310 315cag tac gcc cgg gtg tgg cag tgc aac tgc
aaa ttc cac tgg tgc tgc 1250Gln Tyr Ala Arg Val Trp Gln Cys Asn Cys
Lys Phe His Trp Cys Cys 320 325
330tac gtc aag tgt aac acg tgc agc gag cgc acg gag atg tat acg tgc
1298Tyr Val Lys Cys Asn Thr Cys Ser Glu Arg Thr Glu Met Tyr Thr Cys
335 340 345aag tga atgcggtcac aggtcagatc
acaggcagga tacagtttcc ctgcaggcca 1354Lys350ctgcctggat gctcacaggg
aaagaaccac agaagcactg tccttgtctt ttctgctgag 1414gggggagggg tattctgggt
ttcctgcaga ctcccgtggg aagcatctct cagaggcccg 1474cccattcttc tccacatgga
tgctgctcag ccaccctccc ccagacaccg cccgagcctc 1534tccagggctg gaacaaagtt
ttctacggca ggagctctgg agcctcgggc ctcgtcatag 1594caatatttaa cagtttattc
tgatatgaga taatattaat ttatttaatt aaagagaatt 1654cttccacttc gtcgggatcc
gtcttctgca atcaaagtgg actgcttgag gtcctggtgg 1714gatgacttgc taggactggg
agctgagaac agctgtacat aattattctt tatgcagatg 1774tttctactag ttgatttcac
aagtaccctt ctgcagcgct aggtgttaag tacaaagaga 1834agacggtctt tatacacata
tagatatata tatgcataca catttgtaac tttgttttgt 1894tttgtttttg ctgtttgctg
ctacctatcc agactctaag ctggtccaga tctggaattg 1954tttttctcca ggacgtgctc
ctatcctttt gccctttaca gttcaaacct ctccgttaga 2014aaagttccat tgggaatggc
gtgtgtgtga tggggacgag gatcacaaat tcccagcagt 2074ttccatcctg aaacgtgaac
cactggataa gaggctttct aagagactat ttttctatgg 2134atattttatt tatatggagt
ctgcctgcgg tgccccatgg cccatgcctc ttcttaacac 2194tggtactcac tcaggggcag
aaggacaagg ccaggtgtgt gggcaggtcc cccggggacc 2254ctcacacagc tggagcctgg
agttctattt gccaaggggg ccatagcagt taccagatgc 2314ctgggttggg tatcttctgt
gttaaacaag agggaaccat cccctggctt tagcctgcta 2374agctcagggc ttggaatggg
gtcactggat ggttatcttg ggagatgacc tctggatgag 2434cctcagcggt gggtcagtca
gtgtctcaca cactttgaga agcatgggac ctggcattca 2494tcatcaggca gaggccagct
cagggatgcc gctatcccat caggacagcc caggcactgc 2554ctctaggtga ggtgtagtcc
taagagaagg ggtcaaggag ggggaaggag gaagccaagg 2614agtgttggcc atcctcagtg
aaagcgatgg gagcgttctc tcagcagcag agacacagct 2674gtacctgtat ctctccaatg
ggaaacccct ccagaaggct ggggatattt tttatgtgtt 2734tccacatgca tttccacctg
tgtgcatgta agcacatgcg cacactcctg tgccagcact 2794ctgcggcacc tccagggtgc
tcacgggtac atgtgcttac atgtatctct ctgtgcttgg 2854gagatcagac catgtgcatg
gagctgtatg cctgagcact tgtggtctca ggggttattt 2914ccaggtatct gcatttgtgg
gtggggtgca aggtagacag cagggaactg atttgattgt 2974gttgagccac agtgagactg
caactctgaa ctctgtctcc acagctgctg gtgaaactca 3034gatgcctgtg agacaacagc
cctgagcctc atggcccaca tgctgggagc ccctcagtgt 3094ctaggtcatg tccagtcccc
cacctgggtt acatcacgac caataaacat ggctgtatgg 3154ctgatttctt cccttgaaaa
aaaaaaaaaa aaaaa 318958349PRTMus musculus
58Met Thr Arg Lys Ala Arg Arg Cys Leu Gly His Leu Phe Leu Ser Leu1
5 10 15Gly Ile Val Tyr Leu Arg
Ile Gly Gly Phe Ser Ser Val Val Ala Leu 20 25
30Gly Ala Ser Ile Ile Cys Asn Lys Ile Pro Gly Leu Ala
Pro Arg Gln 35 40 45Arg Ala Ile
Cys Gln Ser Arg Pro Asp Ala Ile Ile Val Ile Gly Glu 50
55 60Gly Ser Gln Met Gly Leu Asp Glu Cys Gln Phe Gln
Phe Arg Asn Gly65 70 75
80Arg Trp Asn Cys Ser Ala Leu Gly Glu Arg Thr Val Phe Gly Lys Glu
85 90 95Leu Lys Val Gly Ser Arg
Glu Ala Ala Phe Thr Tyr Ala Ile Ile Ala 100
105 110Ala Gly Val Ala His Ala Ile Thr Ala Ala Cys Thr
Gln Gly Asn Leu 115 120 125Ser Asp
Cys Gly Cys Asp Lys Glu Lys Gln Gly Gln Tyr His Arg Asp 130
135 140Glu Gly Trp Lys Trp Gly Gly Cys Ser Ala Asp
Ile Arg Tyr Gly Ile145 150 155
160Gly Phe Ala Lys Val Phe Val Asp Ala Arg Glu Ile Lys Gln Asn Ala
165 170 175Arg Thr Leu Met
Asn Leu His Asn Asn Glu Ala Gly Arg Lys Ile Leu 180
185 190Glu Glu Asn Met Lys Leu Glu Cys Lys Cys His
Gly Val Ser Gly Ser 195 200 205Cys
Thr Thr Lys Thr Cys Trp Thr Thr Leu Pro Gln Phe Arg Glu Leu 210
215 220Gly Tyr Val Leu Lys Asp Lys Tyr Asn Glu
Ala Val His Val Glu Pro225 230 235
240Val Arg Ala Ser Arg Asn Lys Arg Pro Thr Phe Leu Lys Ile Lys
Lys 245 250 255Pro Leu Ser
Tyr Arg Lys Pro Met Asp Thr Asp Leu Val Tyr Ile Glu 260
265 270Lys Ser Pro Asn Tyr Cys Glu Glu Asp Pro
Val Thr Gly Ser Val Gly 275 280
285Thr Gln Gly Arg Ala Cys Asn Lys Thr Ala Pro Gln Ala Ser Gly Cys 290
295 300Asp Leu Met Cys Cys Gly Arg Gly
Tyr Asn Thr His Gln Tyr Ala Arg305 310
315 320Val Trp Gln Cys Asn Cys Lys Phe His Trp Cys Cys
Tyr Val Lys Cys 325 330
335Asn Thr Cys Ser Glu Arg Thr Glu Met Tyr Thr Cys Lys 340
345593154DNAMus musculusCDS(133)..(1182) 59cgcccgcctc
ccgagccgaa gcgccggctg agcgtggtcc taccgcagct ccctggctcc 60tgcccggccc
ctgcccaccc gcgcgtcccc tccggccgca gctgtctatg gcgcagcccc 120cctccctgga
tc atg cac aga aac ttt cga aag tgg atc ttt tac gtg ttt 171
Met His Arg Asn Phe Arg Lys Trp Ile Phe Tyr Val Phe 1
5 10ctc tgc ttt ggc gtc ctc tac gtg aag ctc gga
gca ttg tca tcc gtg 219Leu Cys Phe Gly Val Leu Tyr Val Lys Leu Gly
Ala Leu Ser Ser Val 15 20 25gtg gcc
ctg gta gcc aac atc atc tgc aac aag att cct ggc ctg gcc 267Val Ala
Leu Val Ala Asn Ile Ile Cys Asn Lys Ile Pro Gly Leu Ala30
35 40 45cca cgg cag cgt gcc atc tgc
cag agc cga ccc gat gcc atc att gtg 315Pro Arg Gln Arg Ala Ile Cys
Gln Ser Arg Pro Asp Ala Ile Ile Val 50 55
60atc ggg gag ggg gcg cag atg ggc atc gac gag tgc cag
cac cag ttc 363Ile Gly Glu Gly Ala Gln Met Gly Ile Asp Glu Cys Gln
His Gln Phe 65 70 75cga ttc
ggc cgc tgg aac tgc tcc gcc ctg ggc gag aag acc gtc ttc 411Arg Phe
Gly Arg Trp Asn Cys Ser Ala Leu Gly Glu Lys Thr Val Phe 80
85 90ggg caa gaa ctc cga gta ggg agt cga gag
gct gcc ttc acc tat gcc 459Gly Gln Glu Leu Arg Val Gly Ser Arg Glu
Ala Ala Phe Thr Tyr Ala 95 100 105atc
acg gcg gcg ggc gtg gcg cat gct gtc acc gct gcc tgc agc cag 507Ile
Thr Ala Ala Gly Val Ala His Ala Val Thr Ala Ala Cys Ser Gln110
115 120 125ggc aat ctg agc aat tgt
ggc tgt gac cgg gag aag caa ggc tac tac 555Gly Asn Leu Ser Asn Cys
Gly Cys Asp Arg Glu Lys Gln Gly Tyr Tyr 130
135 140aac cag gcg gaa ggc tgg aag tgg ggg ggc tgc tca
gcg gac gtc cgc 603Asn Gln Ala Glu Gly Trp Lys Trp Gly Gly Cys Ser
Ala Asp Val Arg 145 150 155tac
ggc atc gac ttt tct cgt cgc ttt gtg gat gcc cgt gag atc aaa 651Tyr
Gly Ile Asp Phe Ser Arg Arg Phe Val Asp Ala Arg Glu Ile Lys 160
165 170aag aac gcc agg cgc ctc atg aac ctt
cac aac aat gag gcg ggc aga 699Lys Asn Ala Arg Arg Leu Met Asn Leu
His Asn Asn Glu Ala Gly Arg 175 180
185aag gtt ctg gag gac cgc atg aag ctg gaa tgt aag tgt cac ggt gtg
747Lys Val Leu Glu Asp Arg Met Lys Leu Glu Cys Lys Cys His Gly Val190
195 200 205tca ggc tcc tgt
acc acc aaa act tgc tgg acc acg cta cct aag ttc 795Ser Gly Ser Cys
Thr Thr Lys Thr Cys Trp Thr Thr Leu Pro Lys Phe 210
215 220cgc gag gtg ggc cac ctg ctc aag gag aag
tac aac gca gcg gtg cag 843Arg Glu Val Gly His Leu Leu Lys Glu Lys
Tyr Asn Ala Ala Val Gln 225 230
235gtg gag gtg gtg cga gcc agc cgc ctg cgc cag ccc acc ttc ctg cgc
891Val Glu Val Val Arg Ala Ser Arg Leu Arg Gln Pro Thr Phe Leu Arg
240 245 250atc aag cag cta cgc agc tac
cag aag cct atg gag acg gac ctg gtg 939Ile Lys Gln Leu Arg Ser Tyr
Gln Lys Pro Met Glu Thr Asp Leu Val 255 260
265tac atc gag aag tcg ccc aac tac tgc gag gag gac gcg gcc acg ggc
987Tyr Ile Glu Lys Ser Pro Asn Tyr Cys Glu Glu Asp Ala Ala Thr Gly270
275 280 285agc gtg ggc acg
cag ggc cgt ctg tgc aac cgc acc tcg ccg ggg gcc 1035Ser Val Gly Thr
Gln Gly Arg Leu Cys Asn Arg Thr Ser Pro Gly Ala 290
295 300gac ggc tgt gac acc atg tgc tgc ggc cgc
ggc tac aac acg cac cag 1083Asp Gly Cys Asp Thr Met Cys Cys Gly Arg
Gly Tyr Asn Thr His Gln 305 310
315tac acc aag gtg tgg cag tgt aac tgc aaa ttc cac tgg tgt tgc ttc
1131Tyr Thr Lys Val Trp Gln Cys Asn Cys Lys Phe His Trp Cys Cys Phe
320 325 330gtc aag tgc aac acg tgc agc
gag cgc acc gag gtc ttc acc tgc aag 1179Val Lys Cys Asn Thr Cys Ser
Glu Arg Thr Glu Val Phe Thr Cys Lys 335 340
345tga ggctcccgcg caggcgcgct cggcccctgc cgaccctgcg gccctcgcca
1232350ttattttgca catccttctt tgcttctgga gctgccagct gcaggcacag
gagggtgggg 1292atagaggtgg ggagctcgag atactccagg ctccttccta ctcgctctgt
ccccgcccag 1352catccaaggt caacgcaatg gtggtctggt acccaatgga gacaaatccc
tttacttctc 1412tttgggaaag tgaaccacaa agggaccatg agactctgag ggtcacctcc
ctgcctgtga 1472ctggacacag aaaggccaca cccaccagtc acactcaaaa cggtttcctg
ggctgtttcc 1532tgccggccct gggcagtgtg gatggatgtt gacaaaatta tttatgtttt
cttagcatca 1592gatgaggact cagtactaac gactgggtag ccagacctaa ccctatttga
ggacaccctt 1652ccctcactcc tcccggcccc tccctgcagg gtcctctgct ccttgcagaa
ctcgaggatg 1712tcagaattgg cacggaagct ggctggtggg gggactcctt atcagcacct
tgggaggggc 1772ttggtggccc tacaaggcct gagatggccg cagaggacag ccaatcttcc
attccatttg 1832gagactgtca tgcaaatcaa atgtcccttg tgtcaggctc caggcatgcc
tcgtcctctc 1892cctggtcctt caccctccca gcctgctgcc aacctccacc tccagtttac
aaattctctt 1952ctcctctgga gccaacctga cacccaggac tgccccacag gttcaggaga
ggtcagggac 2012agttgcccca catgacagat ggacagaggg caatctgaag atttactgga
gaccccacgg 2072ctctgtgaaa taaatatact gacacagccc catccagccc aactctggaa
gttgccaggg 2132tgatgggagg ctgcaccccc ttttcagtac cttgggtttt gtccttcttc
tgtgatcctg 2192atgccagaga actgacatcc agaatttagg gatgtattgg tcaggccccc
tgcctagtgt 2252ccactgatac ctgcttcagg gtccttatat tatgaggaca tgggaccctc
aaacaggggt 2312ccgtgggaag cttaatgtcc catttcctca ggcccttcca gatggggaca
gaagaactca 2372ggcctgggca tatcccaccc tttcctccac aacacatggc agggtaagaa
actgccaggg 2432ctgataatac aactgcccac agcctacccc acactaaggt gtttcatagc
agaagtccat 2492ggaaatgtgg ggtttggtgg ccaccaagcc aggtggcctg gacattgacc
tggggaaggt 2552gacccttgtt tgcccttgcc ttgcatccag ctgtgtgtcc ctatcatgtc
aggatgttcc 2612aagcctctgg gccactggaa atgtcccacc ctgatcctgg ccccatctcc
tcaccccaag 2672tcctgggata cccacgtccg tcgcccagtg tcccctgtga ggagcctggt
taacttatat 2732tgttatatag cgtcccctgt ctgtcatgtc tcttaagtta ttgtgaccta
cactgggtac 2792cggaggggat gggggatggc ttcagctgct gtcccccaag ccaggctcct
ccttctgctt 2852gaaacagacc ctcgggggcc cctgatgcca ccgaggcaat tcgcactgtc
cctgggctgc 2912caggcacctg cgcctgcact cggtcagccg cagaccttgc cttgggggag
agaggtggtt 2972agtggaccca ggcagggcac tggctgtccc aatgctgtgt gctggggtgg
aggtggccgg 3032gcaccacatg tccttgaagt gccctacttc tgatgggctg tgttcctgcc
tcctctggag 3092gggagcactt agccccaata aaagctggaa tcagaaaaaa aaaaaaaaaa
aaaaaaaaaa 3152aa
315460349PRTMus musculus 60Met His Arg Asn Phe Arg Lys Trp Ile
Phe Tyr Val Phe Leu Cys Phe1 5 10
15Gly Val Leu Tyr Val Lys Leu Gly Ala Leu Ser Ser Val Val Ala
Leu 20 25 30Val Ala Asn Ile
Ile Cys Asn Lys Ile Pro Gly Leu Ala Pro Arg Gln 35
40 45Arg Ala Ile Cys Gln Ser Arg Pro Asp Ala Ile Ile
Val Ile Gly Glu 50 55 60Gly Ala Gln
Met Gly Ile Asp Glu Cys Gln His Gln Phe Arg Phe Gly65 70
75 80Arg Trp Asn Cys Ser Ala Leu Gly
Glu Lys Thr Val Phe Gly Gln Glu 85 90
95Leu Arg Val Gly Ser Arg Glu Ala Ala Phe Thr Tyr Ala Ile
Thr Ala 100 105 110Ala Gly Val
Ala His Ala Val Thr Ala Ala Cys Ser Gln Gly Asn Leu 115
120 125Ser Asn Cys Gly Cys Asp Arg Glu Lys Gln Gly
Tyr Tyr Asn Gln Ala 130 135 140Glu Gly
Trp Lys Trp Gly Gly Cys Ser Ala Asp Val Arg Tyr Gly Ile145
150 155 160Asp Phe Ser Arg Arg Phe Val
Asp Ala Arg Glu Ile Lys Lys Asn Ala 165
170 175Arg Arg Leu Met Asn Leu His Asn Asn Glu Ala Gly
Arg Lys Val Leu 180 185 190Glu
Asp Arg Met Lys Leu Glu Cys Lys Cys His Gly Val Ser Gly Ser 195
200 205Cys Thr Thr Lys Thr Cys Trp Thr Thr
Leu Pro Lys Phe Arg Glu Val 210 215
220Gly His Leu Leu Lys Glu Lys Tyr Asn Ala Ala Val Gln Val Glu Val225
230 235 240Val Arg Ala Ser
Arg Leu Arg Gln Pro Thr Phe Leu Arg Ile Lys Gln 245
250 255Leu Arg Ser Tyr Gln Lys Pro Met Glu Thr
Asp Leu Val Tyr Ile Glu 260 265
270Lys Ser Pro Asn Tyr Cys Glu Glu Asp Ala Ala Thr Gly Ser Val Gly
275 280 285Thr Gln Gly Arg Leu Cys Asn
Arg Thr Ser Pro Gly Ala Asp Gly Cys 290 295
300Asp Thr Met Cys Cys Gly Arg Gly Tyr Asn Thr His Gln Tyr Thr
Lys305 310 315 320Val Trp
Gln Cys Asn Cys Lys Phe His Trp Cys Cys Phe Val Lys Cys
325 330 335Asn Thr Cys Ser Glu Arg Thr
Glu Val Phe Thr Cys Lys 340 345611747DNAMus
musculusCDS(57)..(1121) 61gctctgccga ccttacttct ctgcgcttgg tcctggttcc
cggactgggc aggacc atg 59
Met
1gga cac ttg tta atg ctg tgg gtg gct gcg ggc atg tgc tat cca gcc
107Gly His Leu Leu Met Leu Trp Val Ala Ala Gly Met Cys Tyr Pro Ala
5 10 15ctg ggt gct tct gcc tgg
tca gtg aac aac ttc ctg ata acc ggt ccc 155Leu Gly Ala Ser Ala Trp
Ser Val Asn Asn Phe Leu Ile Thr Gly Pro 20 25
30aag gcc tat ctg acc tac acc gcc agt gtg gcc ttg gga gct
cag att 203Lys Ala Tyr Leu Thr Tyr Thr Ala Ser Val Ala Leu Gly Ala
Gln Ile 35 40 45ggc atc gaa gag tgt
aag ttc cag ttt gcc tgg gaa cgg tgg aat tgt 251Gly Ile Glu Glu Cys
Lys Phe Gln Phe Ala Trp Glu Arg Trp Asn Cys50 55
60 65cct gag cat gct ttt cag ttt tca acc cac
aac agg ctg cga gct gcc 299Pro Glu His Ala Phe Gln Phe Ser Thr His
Asn Arg Leu Arg Ala Ala 70 75
80acg aga gag aca tcc ttc att cat gcc atc cgc tct gct gcc atc atg
347Thr Arg Glu Thr Ser Phe Ile His Ala Ile Arg Ser Ala Ala Ile Met
85 90 95tac gca gtc acc aag aac
tgc agc atg ggt gac ttg gaa aac tgc ggc 395Tyr Ala Val Thr Lys Asn
Cys Ser Met Gly Asp Leu Glu Asn Cys Gly 100 105
110tgt gac gag tca caa aat gga aaa aca ggt ggc cat ggc tgg
atc tgg 443Cys Asp Glu Ser Gln Asn Gly Lys Thr Gly Gly His Gly Trp
Ile Trp 115 120 125gga ggc tgc agc gac
aac gtg gag ttc ggg gaa aaa atc tcc aga ctc 491Gly Gly Cys Ser Asp
Asn Val Glu Phe Gly Glu Lys Ile Ser Arg Leu130 135
140 145ttc gtg gac agt ttg gag aaa ggg aag gat
gcc aga gcc ctg gtg aac 539Phe Val Asp Ser Leu Glu Lys Gly Lys Asp
Ala Arg Ala Leu Val Asn 150 155
160ctt cac aac aac agg gcc ggc aga ctg gca gtg agg gcc tcc acg aaa
587Leu His Asn Asn Arg Ala Gly Arg Leu Ala Val Arg Ala Ser Thr Lys
165 170 175agg acc tgc aag tgt cat
ggc atc tca gga agc tgc agc atc cag acg 635Arg Thr Cys Lys Cys His
Gly Ile Ser Gly Ser Cys Ser Ile Gln Thr 180 185
190tgt tgg ctg cag ctg gct gac ttc cgg cag atg gga aat tac
cta aag 683Cys Trp Leu Gln Leu Ala Asp Phe Arg Gln Met Gly Asn Tyr
Leu Lys 195 200 205gcc aag tat gac cgc
gcg ctg aaa att gag atg gac aag cgc cag cta 731Ala Lys Tyr Asp Arg
Ala Leu Lys Ile Glu Met Asp Lys Arg Gln Leu210 215
220 225agg gct ggc aac aga gcc gag ggc cgc tgg
gct ctc acg gag gcc ttc 779Arg Ala Gly Asn Arg Ala Glu Gly Arg Trp
Ala Leu Thr Glu Ala Phe 230 235
240ctt ccc agc aca gag gct gag ctg atc ttc tta gag ggg tct cct gac
827Leu Pro Ser Thr Glu Ala Glu Leu Ile Phe Leu Glu Gly Ser Pro Asp
245 250 255tac tgc aac cgc aac gcc
agc ctg agc atc cag ggc aca gag ggg agg 875Tyr Cys Asn Arg Asn Ala
Ser Leu Ser Ile Gln Gly Thr Glu Gly Arg 260 265
270gag tgc ctg cag aat gcc cgc agt gct tcc cgg cgg gag cag
cgc agc 923Glu Cys Leu Gln Asn Ala Arg Ser Ala Ser Arg Arg Glu Gln
Arg Ser 275 280 285tgt ggg cgc ctg tgc
acg gag tgc ggg ctg cag gtg gag gag agg aga 971Cys Gly Arg Leu Cys
Thr Glu Cys Gly Leu Gln Val Glu Glu Arg Arg290 295
300 305gca gag gcc gtg agc agc tgt gac tgc aac
ttt cag tgg tgt tgc act 1019Ala Glu Ala Val Ser Ser Cys Asp Cys Asn
Phe Gln Trp Cys Cys Thr 310 315
320gtc aag tgt ggc cag tgc agg cgt gtg gtg agc aga tac tac tgc aca
1067Val Lys Cys Gly Gln Cys Arg Arg Val Val Ser Arg Tyr Tyr Cys Thr
325 330 335cgc cct gta ggt agt gcc
agg ccc cgg ggc agg ggc aag gac agt gcc 1115Arg Pro Val Gly Ser Ala
Arg Pro Arg Gly Arg Gly Lys Asp Ser Ala 340 345
350tgg taa caccaccacc aaattcacgt gctgcctagt tgcaggacag
tggagataga 1171Trp 355gcctgaactt ctggcctagg ggacacagac tggaaaacaa
ttgggacatc acagggttgg 1231cctgtagacc ttccacgata ggtgggttag cctgtagacc
ttccacgata ggcggggtag 1291atggatgatc tttaagcatc ttcttcgcag gagtgaaatc
ggaaccttgt tctcctggct 1351tgtggaccca gcctttcctg cgcagttact cttggactta
agcagcttgt taaagaggga 1411gtttgatttg ggtgcacatc cagaggagcc tggaagaacc
gtattccatt aagtttcaga 1471taccgttcca cccagctgtg ctgctgggag tgcgagggaa
gagaagttaa aggaaaggaa 1531ttctgggggc gggagagatg gctcagtggt taagggccct
ggctggccct ccagaggact 1591ggctcacttc acagcaccca cttgatggct gtgaaccatc
tgtacttcta gttccagggg 1651atccaatgtc cttgcctggt ctctgtgacc accaggcaca
aatgtgcaca gacagacatt 1711tatacatata aaataataaa gtaaaaactt acattt
174762354PRTMus musculus 62Met Gly His Leu Leu Met
Leu Trp Val Ala Ala Gly Met Cys Tyr Pro1 5
10 15Ala Leu Gly Ala Ser Ala Trp Ser Val Asn Asn Phe
Leu Ile Thr Gly 20 25 30Pro
Lys Ala Tyr Leu Thr Tyr Thr Ala Ser Val Ala Leu Gly Ala Gln 35
40 45Ile Gly Ile Glu Glu Cys Lys Phe Gln
Phe Ala Trp Glu Arg Trp Asn 50 55
60Cys Pro Glu His Ala Phe Gln Phe Ser Thr His Asn Arg Leu Arg Ala65
70 75 80Ala Thr Arg Glu Thr
Ser Phe Ile His Ala Ile Arg Ser Ala Ala Ile 85
90 95Met Tyr Ala Val Thr Lys Asn Cys Ser Met Gly
Asp Leu Glu Asn Cys 100 105
110Gly Cys Asp Glu Ser Gln Asn Gly Lys Thr Gly Gly His Gly Trp Ile
115 120 125Trp Gly Gly Cys Ser Asp Asn
Val Glu Phe Gly Glu Lys Ile Ser Arg 130 135
140Leu Phe Val Asp Ser Leu Glu Lys Gly Lys Asp Ala Arg Ala Leu
Val145 150 155 160Asn Leu
His Asn Asn Arg Ala Gly Arg Leu Ala Val Arg Ala Ser Thr
165 170 175Lys Arg Thr Cys Lys Cys His
Gly Ile Ser Gly Ser Cys Ser Ile Gln 180 185
190Thr Cys Trp Leu Gln Leu Ala Asp Phe Arg Gln Met Gly Asn
Tyr Leu 195 200 205Lys Ala Lys Tyr
Asp Arg Ala Leu Lys Ile Glu Met Asp Lys Arg Gln 210
215 220Leu Arg Ala Gly Asn Arg Ala Glu Gly Arg Trp Ala
Leu Thr Glu Ala225 230 235
240Phe Leu Pro Ser Thr Glu Ala Glu Leu Ile Phe Leu Glu Gly Ser Pro
245 250 255Asp Tyr Cys Asn Arg
Asn Ala Ser Leu Ser Ile Gln Gly Thr Glu Gly 260
265 270Arg Glu Cys Leu Gln Asn Ala Arg Ser Ala Ser Arg
Arg Glu Gln Arg 275 280 285Ser Cys
Gly Arg Leu Cys Thr Glu Cys Gly Leu Gln Val Glu Glu Arg 290
295 300Arg Ala Glu Ala Val Ser Ser Cys Asp Cys Asn
Phe Gln Trp Cys Cys305 310 315
320Thr Val Lys Cys Gly Gln Cys Arg Arg Val Val Ser Arg Tyr Tyr Cys
325 330 335Thr Arg Pro Val
Gly Ser Ala Arg Pro Arg Gly Arg Gly Lys Asp Ser 340
345 350Ala Trp631634DNAMus musculusCDS(135)..(1187)
63tccgcttcat ttcaccaccc cttaacactg tttgggatcg cttacacacc aaggtagcca
60cccctctgcc tccgaggaga atgcttccca tctctcaatg tttgagtcgc tcaccctgcc
120tttctccgaa gacc atg ttt ctt atg aag ccc gtg tgc gtt ctt cta gtc
170 Met Phe Leu Met Lys Pro Val Cys Val Leu Leu Val
1 5 10act tgt gtc ctt cac cgc agc
cac gcc tgg tca gtg aac aat ttt ctg 218Thr Cys Val Leu His Arg Ser
His Ala Trp Ser Val Asn Asn Phe Leu 15 20
25atg acc ggt cca aag gct tac ctg gtc tac tcc agc agc gtg gcc
gct 266Met Thr Gly Pro Lys Ala Tyr Leu Val Tyr Ser Ser Ser Val Ala
Ala 30 35 40ggc gcc cag agt ggt att
gaa gaa tgt aaa tac cag ttt gct tgg gac 314Gly Ala Gln Ser Gly Ile
Glu Glu Cys Lys Tyr Gln Phe Ala Trp Asp45 50
55 60cgt tgg aat tgc ccc gag aga gct tta cag ctg
tcc agc cat ggt gga 362Arg Trp Asn Cys Pro Glu Arg Ala Leu Gln Leu
Ser Ser His Gly Gly 65 70
75ctt cga agc gct aac cgg gag aca gca ttt gtg cac gcc atc agc tct
410Leu Arg Ser Ala Asn Arg Glu Thr Ala Phe Val His Ala Ile Ser Ser
80 85 90gct ggg gtt atg tac acc ctg
act aga aac tgc agc ctc gga gac ttt 458Ala Gly Val Met Tyr Thr Leu
Thr Arg Asn Cys Ser Leu Gly Asp Phe 95 100
105gac aac tgt ggc tgt gat gac tcc cga aat gga caa ctg ggg ggc
caa 506Asp Asn Cys Gly Cys Asp Asp Ser Arg Asn Gly Gln Leu Gly Gly
Gln 110 115 120ggt tgg ctc tgg gga ggc
tgc agt gac aac gtg ggc ttc gga gag gca 554Gly Trp Leu Trp Gly Gly
Cys Ser Asp Asn Val Gly Phe Gly Glu Ala125 130
135 140att tcc aag cag ttt gtg gat gcc ctc gag aca
gga caa gat gcc cgg 602Ile Ser Lys Gln Phe Val Asp Ala Leu Glu Thr
Gly Gln Asp Ala Arg 145 150
155gca gcc atg aat ctg cac aac aat gag gct ggc cgc aag gcg gtc aag
650Ala Ala Met Asn Leu His Asn Asn Glu Ala Gly Arg Lys Ala Val Lys
160 165 170ggc acc atg aaa cgc acg
tgt aag tgc cac ggt gtg tcc ggc agc tgc 698Gly Thr Met Lys Arg Thr
Cys Lys Cys His Gly Val Ser Gly Ser Cys 175 180
185acc acg cag acc tgc tgg ttg caa ctg cca gag ttc cgg gag
gta ggc 746Thr Thr Gln Thr Cys Trp Leu Gln Leu Pro Glu Phe Arg Glu
Val Gly 190 195 200gcg cac ttg aag gag
aag tat cat gcg gcg ctc aag gtg gac ctg ctg 794Ala His Leu Lys Glu
Lys Tyr His Ala Ala Leu Lys Val Asp Leu Leu205 210
215 220caa ggc gcg ggc aac agc gcg gcg ggc cgc
gga gcc atc gcc gac acc 842Gln Gly Ala Gly Asn Ser Ala Ala Gly Arg
Gly Ala Ile Ala Asp Thr 225 230
235ttc cgc tcc atc tcc acc cgc gag ctg gtg cat ctg gag gac tcc cca
890Phe Arg Ser Ile Ser Thr Arg Glu Leu Val His Leu Glu Asp Ser Pro
240 245 250gac tac tgc ctg gag aac
aag acc ctg ggg ctg ctg ggc acc gag ggc 938Asp Tyr Cys Leu Glu Asn
Lys Thr Leu Gly Leu Leu Gly Thr Glu Gly 255 260
265cga gag tgt ctg cgg cgc ggg cgc gcc ctg ggt cgc tgg gag
cgc cgc 986Arg Glu Cys Leu Arg Arg Gly Arg Ala Leu Gly Arg Trp Glu
Arg Arg 270 275 280agt tgt cgc cgg ctg
tgc ggg gac tgc ggg cta gcg gtg gag gag cgc 1034Ser Cys Arg Arg Leu
Cys Gly Asp Cys Gly Leu Ala Val Glu Glu Arg285 290
295 300cgc gcc gag aca gtg tcc agc tgc aac tgc
aag ttt cac tgg tgc tgc 1082Arg Ala Glu Thr Val Ser Ser Cys Asn Cys
Lys Phe His Trp Cys Cys 305 310
315gcg gtc cgc tgc gag cag tgc cgc cgg cgg gtc acc aag tac ttc tgc
1130Ala Val Arg Cys Glu Gln Cys Arg Arg Arg Val Thr Lys Tyr Phe Cys
320 325 330agc cgc gca gag cgg ccg
ccc aga ggc gct gcg cac aaa ccg gga aag 1178Ser Arg Ala Glu Arg Pro
Pro Arg Gly Ala Ala His Lys Pro Gly Lys 335 340
345aac tcc taa gggtatctat ccctcccgcc tccacccctg ttcgtcctcg
1227Asn Ser 350gcttccttta gagacccccg gaaatagagg aacccagaat
gggggacctc gcactcccta 1287gcccagagat tctgacagga ggaggctgca gtctctaccg
agtgacactt tgtagctcac 1347tcgtaggtct caaaactgtt ataaaattct gcaagttgtt
cctgaaaaga ggatgagaac 1407aggcgagtct cctcacccca ctttacctac ttcggacccc
aatggtcgct caatgctgga 1467cctagcttat caggcctagg aagggcccct ctcagatatt
cagggtccag ggaaagacgt 1527ggcccttctc ttgctcgcca tagcttcacc tccctcctgt
gagccagagc ttctaggcct 1587agactccccg ctgttgatta ttcaagaatc taaaaacctt
gaccgta 163464350PRTMus musculus 64Met Phe Leu Met Lys
Pro Val Cys Val Leu Leu Val Thr Cys Val Leu1 5
10 15His Arg Ser His Ala Trp Ser Val Asn Asn Phe
Leu Met Thr Gly Pro 20 25
30Lys Ala Tyr Leu Val Tyr Ser Ser Ser Val Ala Ala Gly Ala Gln Ser
35 40 45Gly Ile Glu Glu Cys Lys Tyr Gln
Phe Ala Trp Asp Arg Trp Asn Cys 50 55
60Pro Glu Arg Ala Leu Gln Leu Ser Ser His Gly Gly Leu Arg Ser Ala65
70 75 80Asn Arg Glu Thr Ala
Phe Val His Ala Ile Ser Ser Ala Gly Val Met 85
90 95Tyr Thr Leu Thr Arg Asn Cys Ser Leu Gly Asp
Phe Asp Asn Cys Gly 100 105
110Cys Asp Asp Ser Arg Asn Gly Gln Leu Gly Gly Gln Gly Trp Leu Trp
115 120 125Gly Gly Cys Ser Asp Asn Val
Gly Phe Gly Glu Ala Ile Ser Lys Gln 130 135
140Phe Val Asp Ala Leu Glu Thr Gly Gln Asp Ala Arg Ala Ala Met
Asn145 150 155 160Leu His
Asn Asn Glu Ala Gly Arg Lys Ala Val Lys Gly Thr Met Lys
165 170 175Arg Thr Cys Lys Cys His Gly
Val Ser Gly Ser Cys Thr Thr Gln Thr 180 185
190Cys Trp Leu Gln Leu Pro Glu Phe Arg Glu Val Gly Ala His
Leu Lys 195 200 205Glu Lys Tyr His
Ala Ala Leu Lys Val Asp Leu Leu Gln Gly Ala Gly 210
215 220Asn Ser Ala Ala Gly Arg Gly Ala Ile Ala Asp Thr
Phe Arg Ser Ile225 230 235
240Ser Thr Arg Glu Leu Val His Leu Glu Asp Ser Pro Asp Tyr Cys Leu
245 250 255Glu Asn Lys Thr Leu
Gly Leu Leu Gly Thr Glu Gly Arg Glu Cys Leu 260
265 270Arg Arg Gly Arg Ala Leu Gly Arg Trp Glu Arg Arg
Ser Cys Arg Arg 275 280 285Leu Cys
Gly Asp Cys Gly Leu Ala Val Glu Glu Arg Arg Ala Glu Thr 290
295 300Val Ser Ser Cys Asn Cys Lys Phe His Trp Cys
Cys Ala Val Arg Cys305 310 315
320Glu Gln Cys Arg Arg Arg Val Thr Lys Tyr Phe Cys Ser Arg Ala Glu
325 330 335Arg Pro Pro Arg
Gly Ala Ala His Lys Pro Gly Lys Asn Ser 340
345 350651106DNAMus musculusCDS(8)..(1105) 65cggcaag atg
ctg gat ggg tcc ctt ctg gcg cgc tgg ctg gcc gcg gcc 49 Met
Leu Asp Gly Ser Leu Leu Ala Arg Trp Leu Ala Ala Ala 1
5 10ttc ggg ctg acg ctg ctg ctc gcc gcg ctg cgc cct
tcg gcc gcc tac 97Phe Gly Leu Thr Leu Leu Leu Ala Ala Leu Arg Pro
Ser Ala Ala Tyr15 20 25
30ttc ggg cta aca ggc agt gaa ccc ctg act atc ctc cct ctg acc ctg
145Phe Gly Leu Thr Gly Ser Glu Pro Leu Thr Ile Leu Pro Leu Thr Leu
35 40 45gag acc gag gct gcg gcc
caa gca cac tac aag gcc tgc gac agg ctg 193Glu Thr Glu Ala Ala Ala
Gln Ala His Tyr Lys Ala Cys Asp Arg Leu 50 55
60aag ctg gag cgc aag cag cgc cgc atg tgc cgc agg gac
ccg ggt gtg 241Lys Leu Glu Arg Lys Gln Arg Arg Met Cys Arg Arg Asp
Pro Gly Val 65 70 75gcc gag aca
ctg gtg gag gcc gta agc atg agt gcc ctg gag tgc cag 289Ala Glu Thr
Leu Val Glu Ala Val Ser Met Ser Ala Leu Glu Cys Gln 80
85 90tac cag ttc cgc ttt gag cgc tgg aac tgc acc ctg
gag ggc cgc tac 337Tyr Gln Phe Arg Phe Glu Arg Trp Asn Cys Thr Leu
Glu Gly Arg Tyr95 100 105
110cga gcc agc ctg ctc aag cga ggc ttc aag gag act gct ttc ctc tac
385Arg Ala Ser Leu Leu Lys Arg Gly Phe Lys Glu Thr Ala Phe Leu Tyr
115 120 125gcc atc tct tct gcc
ggc ctg acg cat gca ctg gcc aag gcc tgc agt 433Ala Ile Ser Ser Ala
Gly Leu Thr His Ala Leu Ala Lys Ala Cys Ser 130
135 140gca ggc cgc atg gag cgc tgc acg tgt gat gag gca
ccc gac ctg gaa 481Ala Gly Arg Met Glu Arg Cys Thr Cys Asp Glu Ala
Pro Asp Leu Glu 145 150 155aac cgc
gag gcc tgg cag tgg ggc ggc tgc ggg gac aac ctc aag tac 529Asn Arg
Glu Ala Trp Gln Trp Gly Gly Cys Gly Asp Asn Leu Lys Tyr 160
165 170agc agc aag ttt gtc aag gag ttc ctg ggc cgg
cgc tct agc aag gat 577Ser Ser Lys Phe Val Lys Glu Phe Leu Gly Arg
Arg Ser Ser Lys Asp175 180 185
190ttg cga gcc cga gtg gac ttc cac aac aac ctc gtg ggt gtg aag gtg
625Leu Arg Ala Arg Val Asp Phe His Asn Asn Leu Val Gly Val Lys Val
195 200 205ata aag gct gga gtg
gaa acc act tgc aaa tgc cat ggt gtg tct ggc 673Ile Lys Ala Gly Val
Glu Thr Thr Cys Lys Cys His Gly Val Ser Gly 210
215 220tcc tgc acc gtg cgg acc tgc tgg cgg cag cta gca
ccc ttc cac gag 721Ser Cys Thr Val Arg Thr Cys Trp Arg Gln Leu Ala
Pro Phe His Glu 225 230 235gtg ggc
aag cac cta aaa cac aaa tat gag acc tcg ctc aag gtg ggc 769Val Gly
Lys His Leu Lys His Lys Tyr Glu Thr Ser Leu Lys Val Gly 240
245 250agc act acc aat gaa gcc act gga gag gca ggt
gcc atc tcc cca ccg 817Ser Thr Thr Asn Glu Ala Thr Gly Glu Ala Gly
Ala Ile Ser Pro Pro255 260 265
270cgg ggc cgg gct tct ggg tca gga ggt ggc gac cca ctg ccc cga aca
865Arg Gly Arg Ala Ser Gly Ser Gly Gly Gly Asp Pro Leu Pro Arg Thr
275 280 285cca gag ctt gta cac
ctg gac gac tct ccc agc ttc tgc ctg gct ggc 913Pro Glu Leu Val His
Leu Asp Asp Ser Pro Ser Phe Cys Leu Ala Gly 290
295 300cgc ttt tcc cct ggc acg gca ggc cgc agg tgt cac
cgg gag aag aac 961Arg Phe Ser Pro Gly Thr Ala Gly Arg Arg Cys His
Arg Glu Lys Asn 305 310 315tgt gag
agt att tgt tgt ggc cga ggc cac aac aca cag agt cgt gtg 1009Cys Glu
Ser Ile Cys Cys Gly Arg Gly His Asn Thr Gln Ser Arg Val 320
325 330gtg aca agg ccc tgc caa tgc cag gtc cgc tgg
tgc tgc tac gtg gag 1057Val Thr Arg Pro Cys Gln Cys Gln Val Arg Trp
Cys Cys Tyr Val Glu335 340 345
350tgc agg cag tgt aca cag aga gag gag gtc tat acc tgc aag ggc tga c
1106Cys Arg Gln Cys Thr Gln Arg Glu Glu Val Tyr Thr Cys Lys Gly
355 360 36566365PRTMus musculus
66Met Leu Asp Gly Ser Leu Leu Ala Arg Trp Leu Ala Ala Ala Phe Gly1
5 10 15Leu Thr Leu Leu Leu Ala
Ala Leu Arg Pro Ser Ala Ala Tyr Phe Gly 20 25
30Leu Thr Gly Ser Glu Pro Leu Thr Ile Leu Pro Leu Thr
Leu Glu Thr 35 40 45Glu Ala Ala
Ala Gln Ala His Tyr Lys Ala Cys Asp Arg Leu Lys Leu 50
55 60Glu Arg Lys Gln Arg Arg Met Cys Arg Arg Asp Pro
Gly Val Ala Glu65 70 75
80Thr Leu Val Glu Ala Val Ser Met Ser Ala Leu Glu Cys Gln Tyr Gln
85 90 95Phe Arg Phe Glu Arg Trp
Asn Cys Thr Leu Glu Gly Arg Tyr Arg Ala 100
105 110Ser Leu Leu Lys Arg Gly Phe Lys Glu Thr Ala Phe
Leu Tyr Ala Ile 115 120 125Ser Ser
Ala Gly Leu Thr His Ala Leu Ala Lys Ala Cys Ser Ala Gly 130
135 140Arg Met Glu Arg Cys Thr Cys Asp Glu Ala Pro
Asp Leu Glu Asn Arg145 150 155
160Glu Ala Trp Gln Trp Gly Gly Cys Gly Asp Asn Leu Lys Tyr Ser Ser
165 170 175Lys Phe Val Lys
Glu Phe Leu Gly Arg Arg Ser Ser Lys Asp Leu Arg 180
185 190Ala Arg Val Asp Phe His Asn Asn Leu Val Gly
Val Lys Val Ile Lys 195 200 205Ala
Gly Val Glu Thr Thr Cys Lys Cys His Gly Val Ser Gly Ser Cys 210
215 220Thr Val Arg Thr Cys Trp Arg Gln Leu Ala
Pro Phe His Glu Val Gly225 230 235
240Lys His Leu Lys His Lys Tyr Glu Thr Ser Leu Lys Val Gly Ser
Thr 245 250 255Thr Asn Glu
Ala Thr Gly Glu Ala Gly Ala Ile Ser Pro Pro Arg Gly 260
265 270Arg Ala Ser Gly Ser Gly Gly Gly Asp Pro
Leu Pro Arg Thr Pro Glu 275 280
285Leu Val His Leu Asp Asp Ser Pro Ser Phe Cys Leu Ala Gly Arg Phe 290
295 300Ser Pro Gly Thr Ala Gly Arg Arg
Cys His Arg Glu Lys Asn Cys Glu305 310
315 320Ser Ile Cys Cys Gly Arg Gly His Asn Thr Gln Ser
Arg Val Val Thr 325 330
335Arg Pro Cys Gln Cys Gln Val Arg Trp Cys Cys Tyr Val Glu Cys Arg
340 345 350Gln Cys Thr Gln Arg Glu
Glu Val Tyr Thr Cys Lys Gly 355 360
365674522DNAMus musculusCDS(54)..(1133) 67gacgagcgcc tagtggcgcg
aggagatgcg agagtgcacc ggccgcctgc acc atg 56
Met
1cgc ccc gcg ccc gcg ctg gcc ctg gct gcg ctc tgc
ctg ctg gtg ctg 104Arg Pro Ala Pro Ala Leu Ala Leu Ala Ala Leu Cys
Leu Leu Val Leu 5 10 15cct
gcc gct gcc gcc gcc gcc gcc tac ttc ggc ctg acc ggt cgt gag 152Pro
Ala Ala Ala Ala Ala Ala Ala Tyr Phe Gly Leu Thr Gly Arg Glu 20
25 30gtc ctg aca ccc ttc cca ggc ctg ggt
acg gca gca gcc ccg gca cag 200Val Leu Thr Pro Phe Pro Gly Leu Gly
Thr Ala Ala Ala Pro Ala Gln 35 40
45gct ggt gct cac ctg aag cag tgt gac cta ctg aag ctg tcc agg cgg
248Ala Gly Ala His Leu Lys Gln Cys Asp Leu Leu Lys Leu Ser Arg Arg50
55 60 65cag aag cag ctc tgc
agg cgg gag ccc ggc ctg gct gag acc ctg agg 296Gln Lys Gln Leu Cys
Arg Arg Glu Pro Gly Leu Ala Glu Thr Leu Arg 70
75 80gat gct gca cac ctg ggg ctg ctg gaa tgt cag
ttc cag ttc agg cag 344Asp Ala Ala His Leu Gly Leu Leu Glu Cys Gln
Phe Gln Phe Arg Gln 85 90
95gag cgc tgg aac tgc agc ctg gag ggg agg act ggc ctg ctc cag aga
392Glu Arg Trp Asn Cys Ser Leu Glu Gly Arg Thr Gly Leu Leu Gln Arg
100 105 110ggc ttt aag gag acg gcc ttc
ctg tat gca gtg tct gca gct gcc ctc 440Gly Phe Lys Glu Thr Ala Phe
Leu Tyr Ala Val Ser Ala Ala Ala Leu 115 120
125acg cat gca ctg gcc agg gcc tgc agt gct ggg cgc atg gag cgc tgt
488Thr His Ala Leu Ala Arg Ala Cys Ser Ala Gly Arg Met Glu Arg Cys130
135 140 145act tgt gac gac
tcc cca ggc ctg gag agc cgg cag gcc tgg cag tgg 536Thr Cys Asp Asp
Ser Pro Gly Leu Glu Ser Arg Gln Ala Trp Gln Trp 150
155 160ggt gtg tgt ggt gac aat ctg aag tac agc
acc aag ttc ctc agc aac 584Gly Val Cys Gly Asp Asn Leu Lys Tyr Ser
Thr Lys Phe Leu Ser Asn 165 170
175ttc ctg ggg ccc aag aga gga agc aag gac ctg agg gcg agg gct gac
632Phe Leu Gly Pro Lys Arg Gly Ser Lys Asp Leu Arg Ala Arg Ala Asp
180 185 190gcc cac aac acc cac gtg ggc
atc aag gct gtg aag agc ggc ctg aga 680Ala His Asn Thr His Val Gly
Ile Lys Ala Val Lys Ser Gly Leu Arg 195 200
205aca acc tgc aag tgc cat ggt gtg tca ggc tcc tgt gct gtt cgt acc
728Thr Thr Cys Lys Cys His Gly Val Ser Gly Ser Cys Ala Val Arg Thr210
215 220 225tgt tgg aag cag
ctc tcc ccg ttt cgc gag acc ggc cag gtg ctg aag 776Cys Trp Lys Gln
Leu Ser Pro Phe Arg Glu Thr Gly Gln Val Leu Lys 230
235 240cta cgc tat gac acg gct gtc aag gtg tcc
agt gcc acc aac gag gcc 824Leu Arg Tyr Asp Thr Ala Val Lys Val Ser
Ser Ala Thr Asn Glu Ala 245 250
255ttg ggt cgt ctg gag cta tgg gcc ccc gct aag cca ggt ggt acc gcc
872Leu Gly Arg Leu Glu Leu Trp Ala Pro Ala Lys Pro Gly Gly Thr Ala
260 265 270aag ggc cta gcc cct cgt ccc
ggg gac ctg gtc tac atg gaa gat tct 920Lys Gly Leu Ala Pro Arg Pro
Gly Asp Leu Val Tyr Met Glu Asp Ser 275 280
285ccc agc ttc tgc cgg ccc agc aag tac tct ccg ggc acg gca ggc agg
968Pro Ser Phe Cys Arg Pro Ser Lys Tyr Ser Pro Gly Thr Ala Gly Arg290
295 300 305gtg tgt tct cga
gac tcc agt tgc agc agc cta tgc tgt ggg cga ggc 1016Val Cys Ser Arg
Asp Ser Ser Cys Ser Ser Leu Cys Cys Gly Arg Gly 310
315 320tac gac acc cag agc cgc atg gtg gtt ttc
tcc tgc cac tgt cag gtg 1064Tyr Asp Thr Gln Ser Arg Met Val Val Phe
Ser Cys His Cys Gln Val 325 330
335cag tgg tgc tgc tac gtg gag tgc cag cag tgt gca cag cag gag ctc
1112Gln Trp Cys Cys Tyr Val Glu Cys Gln Gln Cys Ala Gln Gln Glu Leu
340 345 350gtg tat acc tgc aag cgc tag
gcctccacag cgaatcccgc ggaacagcgc 1163Val Tyr Thr Cys Lys Arg
355 360gcaagcgcgc acctgtcgac gcacctgccg tgcacaagag
tgtgcgactc atctctcttc 1223cccaacagat ggttggccag cccttctgcc ttccccgaca
ctcagcaaag agaaagaaag 1283ccctgcctcc tagtcccagg atcaccaacc tgctggagga
cttggggccg gagaacagac 1343tgagaagggg aatctttgag gaccagggta gggcaggaat
gatgctgtgc gggaagagag 1403aaacatcctc ctatctcaag gccaaaaact gggaggatgg
ggaagaggga ggcggagcca 1463gctggagtgt ggggtcaggg catccatctg ggcgtggccg
atctcttgtg gtcccactct 1523aatagcagag cgctctgggt gctgcatgcc taccctgctc
ttgtggcttc gtgcactgga 1583gacttcgaaa tgtttattag gagcaaggga agcactttag
gcttgggtgg attgagtcgc 1643agagcccatg ccctgaagtc ttacgtcctg gcactcaggg
ctgccacctt gtctccttgt 1703cttgagatcc cctgtccccc aaagccattg agctctgctc
aacgagaccc ctaatatgta 1763taagaagggt gcaggagcca gtctcctcgg tgagactcag
ataaacataa ctagggttga 1823gcggggagac agtgaccctt tctctttcct ttggtccaag
gaacctttaa tcacagccca 1883gaggtggaga gaggcagggt ccaaatgcct ggaagagata
tgacaggctc tgtattgaga 1943taccactctg gagtgtgtcc taccaattcc tgtgaccagg
gacccccaag aaccgagggg 2003cccccatcca tgttagtgat acataagaac gagtgactca
tgggccacac gtctgcttcc 2063accccctgct ctcaaagatg cttgtgcagg cttttttgcc
attgctaagt ctttgccaag 2123tctgcctcct caatggtctt actcatttac taacgacctg
tcacttgggc tcccaccaga 2183ggaacaaaat gactgctggt gaatcctttg gtcattttta
atgcccccat caaggccctc 2243tgtgagagga gaggaagtag tgtacaggta caggctcaca
cgtgcacaca ctcagcctag 2303ccaggcacag acatcccaag gagcagtgcg gcgtctctcc
agcccagggc aaagacctca 2363ctggggtcac ttctggaggc tgtgagctac tccagggcag
ggcccaaggc caaccaggag 2423gaagtgacct cctttgggaa gcctttggcc atgtggctgg
ctgtgctgca ccctcctgtg 2483agcttccttc caccctgaaa tctgttgggg ttactgtctc
tctaagggag caggaagctt 2543cggaatcagc cggtactcag cactactggc cctgccagct
ccaggaaaga gacactgtgg 2603cggagaggtc cgtggggcag aaggggctac cctttcttca
gtgcctccgg gcagcatgct 2663gggaagatct ttgatggtgg aaagccccga ggcggagcca
ccgtgacctg agacccttct 2723ctgggacgac tttgccaccc acccgcagct tggcaggagg
ggtaaacaga ttgggagctg 2783ctttctactt ccctgatgaa gacagatgtg ttccttggca
acccaaggca tccttctcta 2843tgaccctaat cctgctctgg ctcgagggta caaggcaaga
atggagcctg gcaaaacttg 2903gggactagaa cacctggacc tacagccaaa tcacctgtac
cctgactcta tggccaggag 2963ggccaggggt ggaggagggt taaagatgaa cttgaagttg
agggctgagg ctgaccaacc 3023attaagactg gtgccttaag gcaccctcag tcaggtcctc
tccctccctt ctccattctt 3083tctccaaggc cccgttcccc ctaaaatccc accatagcca
tgctgggtcc ccccttcccc 3143cacactggga actttaagga agatattcac agggtatttc
tgcctacctc atacatgtaa 3203ttttcaaaaa aaattaattt atatagttaa gatatatggg
aaagtattta tgttatttat 3263atatcttctc tatttcctgg gcaccatatg gggggttgtg
tgtttaccca gaagcctctg 3323aggaaacatg gctgggtctg tctggggcct cgcagagctg
gatgcgcata gctgagaggt 3383cacagctcct gtgtctcact gtcttggagc tcgggaagca
catgtacctc ctgagataaa 3443ccccgtgaca ccaagcaggg ccttccttgt gaagtctgtg
gattctctgc ctctggcccc 3503agaggccttt ctgctctggc ccaagggttt tgctcataaa
ggacaaaaag ggtgagcagc 3563tctggatttg taaagcactt tccatcttca gaaacactcc
tctcttctct ctccctcggt 3623tacccccggt tccctatgag gtcatgccac tgttaccacg
ttccaggccc agagacggag 3683gcaggttggt caaagccagt cactctctga acccagaggt
tgaggaagag tgcatgctgc 3743gtggaacgct ggtcttcccc catggatggc atgctagttt
ctccagcaag ctgagtctca 3803tgtccccaaa gacggggact tcctgagaag cctggagaga
caagggctcc gtggatgtca 3863ctcttaggga gggtgtcctg cagccctcat tgacctccac
gactaggcta tggtctccag 3923cccctcacag ctcgtggata atttgtgttt cttcgctttt
gttttttgtc ttttcaaagt 3983gactttttcc ccactggatt tctaagtttc tctttgaaaa
tcagttcact ggcaaatggg 4043acctgcatcc tgacctggct gcctgcatca ggagcgacac
caaacagagt gcgtggggat 4103ccccaattgg cccagtgtcc cccggccctt ccttaagtca
cacaagctcc cgtgtggctt 4163tcgtgagcat ggagaacctg tcccctggtc ttagagaaag
ccagccattc tgccaccctc 4223tgtttgtctg gcagacagat taccacaccg tggctgtctt
tctagccaaa gcttcctctc 4283tcaacaccca tgaacgtcca tgcttcctgt ctgagcactg
aggagaaccc cagcggagct 4343cattgttcag tgctggaata cccatccccc ctcccgttga
ttatttaggg agtgtctgat 4403aatgccaggg gatactctgg gtgctagggc gcagaagtac
ttaagagcaa gtcccagcct 4463caggggactt atatgccggc gaggagaaag ccaacaaacc
aataaactat gcactggtt 452268359PRTMus musculus 68Met Arg Pro Ala Pro
Ala Leu Ala Leu Ala Ala Leu Cys Leu Leu Val1 5
10 15Leu Pro Ala Ala Ala Ala Ala Ala Ala Tyr Phe
Gly Leu Thr Gly Arg 20 25
30Glu Val Leu Thr Pro Phe Pro Gly Leu Gly Thr Ala Ala Ala Pro Ala
35 40 45Gln Ala Gly Ala His Leu Lys Gln
Cys Asp Leu Leu Lys Leu Ser Arg 50 55
60Arg Gln Lys Gln Leu Cys Arg Arg Glu Pro Gly Leu Ala Glu Thr Leu65
70 75 80Arg Asp Ala Ala His
Leu Gly Leu Leu Glu Cys Gln Phe Gln Phe Arg 85
90 95Gln Glu Arg Trp Asn Cys Ser Leu Glu Gly Arg
Thr Gly Leu Leu Gln 100 105
110Arg Gly Phe Lys Glu Thr Ala Phe Leu Tyr Ala Val Ser Ala Ala Ala
115 120 125Leu Thr His Ala Leu Ala Arg
Ala Cys Ser Ala Gly Arg Met Glu Arg 130 135
140Cys Thr Cys Asp Asp Ser Pro Gly Leu Glu Ser Arg Gln Ala Trp
Gln145 150 155 160Trp Gly
Val Cys Gly Asp Asn Leu Lys Tyr Ser Thr Lys Phe Leu Ser
165 170 175Asn Phe Leu Gly Pro Lys Arg
Gly Ser Lys Asp Leu Arg Ala Arg Ala 180 185
190Asp Ala His Asn Thr His Val Gly Ile Lys Ala Val Lys Ser
Gly Leu 195 200 205Arg Thr Thr Cys
Lys Cys His Gly Val Ser Gly Ser Cys Ala Val Arg 210
215 220Thr Cys Trp Lys Gln Leu Ser Pro Phe Arg Glu Thr
Gly Gln Val Leu225 230 235
240Lys Leu Arg Tyr Asp Thr Ala Val Lys Val Ser Ser Ala Thr Asn Glu
245 250 255Ala Leu Gly Arg Leu
Glu Leu Trp Ala Pro Ala Lys Pro Gly Gly Thr 260
265 270Ala Lys Gly Leu Ala Pro Arg Pro Gly Asp Leu Val
Tyr Met Glu Asp 275 280 285Ser Pro
Ser Phe Cys Arg Pro Ser Lys Tyr Ser Pro Gly Thr Ala Gly 290
295 300Arg Val Cys Ser Arg Asp Ser Ser Cys Ser Ser
Leu Cys Cys Gly Arg305 310 315
320Gly Tyr Asp Thr Gln Ser Arg Met Val Val Phe Ser Cys His Cys Gln
325 330 335Val Gln Trp Cys
Cys Tyr Val Glu Cys Gln Gln Cys Ala Gln Gln Glu 340
345 350Leu Val Tyr Thr Cys Lys Arg
355691974DNAMus musculusCDS(153)..(1406) 69ggcgggcgcc gtctgctgcg
ggagctgtga cctgagtagg agctgtgtgt cgcagccgcc 60ccacccctgc cgatcatgcg
ccggcgaccc tggttcgcca gtcccactgg gctgtgagcc 120ccccactcct ggcctgtcac
ggcccgcgcg cc atg ggc agc gcc cac cct cgc 173
Met Gly Ser Ala His Pro Arg
1 5ccc tgg ctg cgg ctc cca caa ggg ccc cag ccg cgg cct gag
ttc tgg 221Pro Trp Leu Arg Leu Pro Gln Gly Pro Gln Pro Arg Pro Glu
Phe Trp 10 15 20gcg ctc ctg ttc
ttc cta ctg ctg ctg gct gcc gct gtg ccc agg tca 269Ala Leu Leu Phe
Phe Leu Leu Leu Leu Ala Ala Ala Val Pro Arg Ser 25 30
35gca ccc aac gac atc ctg ggc ctc cgc cta ccc cca gag
ccc gtg ctc 317Ala Pro Asn Asp Ile Leu Gly Leu Arg Leu Pro Pro Glu
Pro Val Leu40 45 50
55aac gcc aac aca gtg tgc ctg aca ttg ccc ggc ctg agc cgg cgg cag
365Asn Ala Asn Thr Val Cys Leu Thr Leu Pro Gly Leu Ser Arg Arg Gln
60 65 70atg gag gtg tgt gtg cgt
cac cct gac gtg gcc gcc tct gct atc cag 413Met Glu Val Cys Val Arg
His Pro Asp Val Ala Ala Ser Ala Ile Gln 75 80
85ggc atc cag atc gcc atc cat gag tgc cag cat cag ttc
cgg gac cag 461Gly Ile Gln Ile Ala Ile His Glu Cys Gln His Gln Phe
Arg Asp Gln 90 95 100cgc tgg aac
tgc tcc agc ctg gag act cgg aac aaa gtc ccc tac gag 509Arg Trp Asn
Cys Ser Ser Leu Glu Thr Arg Asn Lys Val Pro Tyr Glu 105
110 115agc ccc atc ttc agc cga ggt ttt cga gag agt gct
ttc gcc tac gcc 557Ser Pro Ile Phe Ser Arg Gly Phe Arg Glu Ser Ala
Phe Ala Tyr Ala120 125 130
135ata gca gct gcc ggg gtg gtg cac gca gtg tcc aac gcg tgc gct ctg
605Ile Ala Ala Ala Gly Val Val His Ala Val Ser Asn Ala Cys Ala Leu
140 145 150ggt aaa ctg aag gct
tgc ggt tgc gac gcc tcc aga cgt ggg gac gaa 653Gly Lys Leu Lys Ala
Cys Gly Cys Asp Ala Ser Arg Arg Gly Asp Glu 155
160 165gaa gct ttc cgt cgg aag ctg cac cgc ttg cag ctg
gac gcg ctg cag 701Glu Ala Phe Arg Arg Lys Leu His Arg Leu Gln Leu
Asp Ala Leu Gln 170 175 180cgc gga
aag ggc ttg agc cac ggg gtc cct gaa cac ccg gcc ata ctt 749Arg Gly
Lys Gly Leu Ser His Gly Val Pro Glu His Pro Ala Ile Leu 185
190 195cct gcc agc cca ggt ctg cag gac tcc tgg gag
tgg ggt ggc tgc agt 797Pro Ala Ser Pro Gly Leu Gln Asp Ser Trp Glu
Trp Gly Gly Cys Ser200 205 210
215ccg gat gtg ggc ttc gga gaa cgc ttc tct aag gac ttt ctg gac tcc
845Pro Asp Val Gly Phe Gly Glu Arg Phe Ser Lys Asp Phe Leu Asp Ser
220 225 230cga gag cct cac aga
gac atc cat gct cga atg aga ctc cac aac aac 893Arg Glu Pro His Arg
Asp Ile His Ala Arg Met Arg Leu His Asn Asn 235
240 245cgt gtg ggc cgg cag gcg gtg atg gag aac atg cgg
cgt aag tgc aaa 941Arg Val Gly Arg Gln Ala Val Met Glu Asn Met Arg
Arg Lys Cys Lys 250 255 260tgc cac
ggc acc tca ggc agc tgc cag ctc aag acc tgc tgg cag gtg 989Cys His
Gly Thr Ser Gly Ser Cys Gln Leu Lys Thr Cys Trp Gln Val 265
270 275acg cct gag ttc cgc aca gta ggg gcg ctg ctg
cgc aac cgc ttc cac 1037Thr Pro Glu Phe Arg Thr Val Gly Ala Leu Leu
Arg Asn Arg Phe His280 285 290
295cgc gcc acg ctc atc cgg ccg cac aac cgc aac ggt ggc cag ctg gag
1085Arg Ala Thr Leu Ile Arg Pro His Asn Arg Asn Gly Gly Gln Leu Glu
300 305 310ccc ggc ccc gcg gga
gca ccc tcg cca gca ccg ggc act cca ggg ctg 1133Pro Gly Pro Ala Gly
Ala Pro Ser Pro Ala Pro Gly Thr Pro Gly Leu 315
320 325cgc cgc agg gcc agc cac tcc gac ctg gtc tac ttt
gag aaa tct ccc 1181Arg Arg Arg Ala Ser His Ser Asp Leu Val Tyr Phe
Glu Lys Ser Pro 330 335 340gac ttc
tgt gag cgc gag ccg cgc ctg gac tcg gca ggc act gtg ggc 1229Asp Phe
Cys Glu Arg Glu Pro Arg Leu Asp Ser Ala Gly Thr Val Gly 345
350 355cgc ctg tgc aat aag agc agc acg ggt ccc gat
ggc tgc ggc agc atg 1277Arg Leu Cys Asn Lys Ser Ser Thr Gly Pro Asp
Gly Cys Gly Ser Met360 365 370
375tgc tgt ggc cgc ggc cac aac att ctg cgc cag acg cgc agc gag cgc
1325Cys Cys Gly Arg Gly His Asn Ile Leu Arg Gln Thr Arg Ser Glu Arg
380 385 390tgc cac tgc cgg ttc
cac tgg tgc tgc ttc gtg gtc tgc gaa gaa tgc 1373Cys His Cys Arg Phe
His Trp Cys Cys Phe Val Val Cys Glu Glu Cys 395
400 405cgc atc acc gag tgg gtc agc gtc tgc aag tga
gcagacccaa gctcctctgg 1426Arg Ile Thr Glu Trp Val Ser Val Cys Lys
410 415gtctcaagaa tggttgtcct cttggtgcct ggcttctgcc
gctagcggat ctgagccagg 1486cagcaagcag cagccttggc tcctgagaga ggtggttggc
tcttacagcc ccgagggtct 1546acaatcacca gacagtccag atctgattga cattcctccg
ctcacctctg taggttcccc 1606tctttctgtt cctagctcag acagctgggg gtgatagtgg
agactgttcc acaccctagg 1666acaggtcacc aaagcagccc agcctggcat gcctacctcc
tgtcatctct tcttcccttc 1726cccaggagtg ataggcaatg cactgaagct gatgggcacc
ggggaagaaa actaaaaggc 1786agaaatggcc gtcatcgggc tgaagtgact ctaagggctc
cagacctctg ctcctgtctt 1846tcacttaaca gatatttatt tttgcgctct ctttgagaca
ctctctgggg aaaaagaagc 1906tccggagtct acaggctgat taagggacat ggacaataaa
ccagtaaaca cacaaaaaaa 1966aaaaaaaa
197470417PRTMus musculus 70Met Gly Ser Ala His Pro
Arg Pro Trp Leu Arg Leu Pro Gln Gly Pro1 5
10 15Gln Pro Arg Pro Glu Phe Trp Ala Leu Leu Phe Phe
Leu Leu Leu Leu 20 25 30Ala
Ala Ala Val Pro Arg Ser Ala Pro Asn Asp Ile Leu Gly Leu Arg 35
40 45Leu Pro Pro Glu Pro Val Leu Asn Ala
Asn Thr Val Cys Leu Thr Leu 50 55
60Pro Gly Leu Ser Arg Arg Gln Met Glu Val Cys Val Arg His Pro Asp65
70 75 80Val Ala Ala Ser Ala
Ile Gln Gly Ile Gln Ile Ala Ile His Glu Cys 85
90 95Gln His Gln Phe Arg Asp Gln Arg Trp Asn Cys
Ser Ser Leu Glu Thr 100 105
110Arg Asn Lys Val Pro Tyr Glu Ser Pro Ile Phe Ser Arg Gly Phe Arg
115 120 125Glu Ser Ala Phe Ala Tyr Ala
Ile Ala Ala Ala Gly Val Val His Ala 130 135
140Val Ser Asn Ala Cys Ala Leu Gly Lys Leu Lys Ala Cys Gly Cys
Asp145 150 155 160Ala Ser
Arg Arg Gly Asp Glu Glu Ala Phe Arg Arg Lys Leu His Arg
165 170 175Leu Gln Leu Asp Ala Leu Gln
Arg Gly Lys Gly Leu Ser His Gly Val 180 185
190Pro Glu His Pro Ala Ile Leu Pro Ala Ser Pro Gly Leu Gln
Asp Ser 195 200 205Trp Glu Trp Gly
Gly Cys Ser Pro Asp Val Gly Phe Gly Glu Arg Phe 210
215 220Ser Lys Asp Phe Leu Asp Ser Arg Glu Pro His Arg
Asp Ile His Ala225 230 235
240Arg Met Arg Leu His Asn Asn Arg Val Gly Arg Gln Ala Val Met Glu
245 250 255Asn Met Arg Arg Lys
Cys Lys Cys His Gly Thr Ser Gly Ser Cys Gln 260
265 270Leu Lys Thr Cys Trp Gln Val Thr Pro Glu Phe Arg
Thr Val Gly Ala 275 280 285Leu Leu
Arg Asn Arg Phe His Arg Ala Thr Leu Ile Arg Pro His Asn 290
295 300Arg Asn Gly Gly Gln Leu Glu Pro Gly Pro Ala
Gly Ala Pro Ser Pro305 310 315
320Ala Pro Gly Thr Pro Gly Leu Arg Arg Arg Ala Ser His Ser Asp Leu
325 330 335Val Tyr Phe Glu
Lys Ser Pro Asp Phe Cys Glu Arg Glu Pro Arg Leu 340
345 350Asp Ser Ala Gly Thr Val Gly Arg Leu Cys Asn
Lys Ser Ser Thr Gly 355 360 365Pro
Asp Gly Cys Gly Ser Met Cys Cys Gly Arg Gly His Asn Ile Leu 370
375 380Arg Gln Thr Arg Ser Glu Arg Cys His Cys
Arg Phe His Trp Cys Cys385 390 395
400Phe Val Val Cys Glu Glu Cys Arg Ile Thr Glu Trp Val Ser Val
Cys 405 410
415Lys712215DNAMus musculusCDS(332)..(1501) 71ctcgagcaga accacccgtg
agttaggtcg agcagagcca aagcccccgg tgcttcgtcg 60cgggttcgct cgctagctat
ctggatcact ccctcccttt taccctccct tcctcccggc 120gggcggccgc ggcgacgccg
gggaagcggc agagaggagt ggctgggcgc tgggagaatg 180ctgctccgcc gagggggctg
aacccgacag tttccccacg gtttaagccc caagagccgg 240gcccgagtga ctcaaccgcg
agccttgtgg atcctgcacc tgaaccgctg gaggctgact 300gactcgccca ccggagcctc
cgggcttcga c atg ctg gag gag ccc cgg tct 352
Met Leu Glu Glu Pro Arg Ser
1 5cgg cct ccg ccc tta ggc ctc gcg ggt ctc ctg ttc ttg gct
ttg ttc 400Arg Pro Pro Pro Leu Gly Leu Ala Gly Leu Leu Phe Leu Ala
Leu Phe 10 15 20agt cgg gct cta
agc aat gag att ctg ggc ctt aaa ctt ccc ggt gag 448Ser Arg Ala Leu
Ser Asn Glu Ile Leu Gly Leu Lys Leu Pro Gly Glu 25 30
35ccg ccg ctg acg gcc aac acc gtg tgc ttg acc ctg tcc
gga ctg agt 496Pro Pro Leu Thr Ala Asn Thr Val Cys Leu Thr Leu Ser
Gly Leu Ser40 45 50
55aag cga cag ctg ggg ctg tgc ctg cgc agc ccc gac gtg acg gcg tcg
544Lys Arg Gln Leu Gly Leu Cys Leu Arg Ser Pro Asp Val Thr Ala Ser
60 65 70gcg ctc cag ggg ctg cac
atc gcc gtt cac gag tgt cag cac cag ctg 592Ala Leu Gln Gly Leu His
Ile Ala Val His Glu Cys Gln His Gln Leu 75 80
85cgc gac cag cgc tgg aac tgc tcg gca ctg gag ggc ggc
ggc cgg ctg 640Arg Asp Gln Arg Trp Asn Cys Ser Ala Leu Glu Gly Gly
Gly Arg Leu 90 95 100ccg cac cac
agc gcc atc ctc aag cgc ggt ttc cgt gag agt gct ttc 688Pro His His
Ser Ala Ile Leu Lys Arg Gly Phe Arg Glu Ser Ala Phe 105
110 115tcc ttc tcc atg ctg gct gct ggg gtc atg cat gct
gtt gcc aca gcc 736Ser Phe Ser Met Leu Ala Ala Gly Val Met His Ala
Val Ala Thr Ala120 125 130
135tgc agc ctg ggc aag ctg gtg agc tgc ggc tgc gga tgg aag ggt agt
784Cys Ser Leu Gly Lys Leu Val Ser Cys Gly Cys Gly Trp Lys Gly Ser
140 145 150ggt gag caa gac cgg
ctt aga gcc aag ctg ctg cag ctt cag gca ctg 832Gly Glu Gln Asp Arg
Leu Arg Ala Lys Leu Leu Gln Leu Gln Ala Leu 155
160 165tct cgg ggc aag act ttc ccc atc tcc cag ccc agc
cct gtt cct ggc 880Ser Arg Gly Lys Thr Phe Pro Ile Ser Gln Pro Ser
Pro Val Pro Gly 170 175 180tca gtc
ccc agc ccc ggc ccc cag gac acg tgg gaa tgg ggt ggc tgt 928Ser Val
Pro Ser Pro Gly Pro Gln Asp Thr Trp Glu Trp Gly Gly Cys 185
190 195aac cac gac atg gac ttc gga gag aag ttc tct
cgg gat ttc ttg gat 976Asn His Asp Met Asp Phe Gly Glu Lys Phe Ser
Arg Asp Phe Leu Asp200 205 210
215tcc agg gag gct ccc cgg gac atc cag gcg aga atg cgg atc cac aac
1024Ser Arg Glu Ala Pro Arg Asp Ile Gln Ala Arg Met Arg Ile His Asn
220 225 230aac agg gtg gga cgc
cag gtg gta acg gaa aac ctg aag cgg aag tgc 1072Asn Arg Val Gly Arg
Gln Val Val Thr Glu Asn Leu Lys Arg Lys Cys 235
240 245aaa tgc cat gga acg tca ggc agc tgc caa ttc aag
acc tgt tgg agg 1120Lys Cys His Gly Thr Ser Gly Ser Cys Gln Phe Lys
Thr Cys Trp Arg 250 255 260gca gcg
cca gag ttc cgg gcc atc ggg gca gca ctg agg gag cgg ctg 1168Ala Ala
Pro Glu Phe Arg Ala Ile Gly Ala Ala Leu Arg Glu Arg Leu 265
270 275agc aga gcc atc ttt atc gat acc cac aac cgc
aac tct gga gcg ttc 1216Ser Arg Ala Ile Phe Ile Asp Thr His Asn Arg
Asn Ser Gly Ala Phe280 285 290
295cag ccc cgc cta cgt ccg cgg cgc ctc tct gga gag ctg gtt tac ttt
1264Gln Pro Arg Leu Arg Pro Arg Arg Leu Ser Gly Glu Leu Val Tyr Phe
300 305 310gag aag tct cct gac
ttc tgc gag cga gac cct act ctg ggc tcc cca 1312Glu Lys Ser Pro Asp
Phe Cys Glu Arg Asp Pro Thr Leu Gly Ser Pro 315
320 325ggc acg aga ggc cgg gct tgc aac aag acc agc cgc
ctc ttg gat ggc 1360Gly Thr Arg Gly Arg Ala Cys Asn Lys Thr Ser Arg
Leu Leu Asp Gly 330 335 340tgt ggc
agc ctg tgc tgt ggc cgt ggg cac aac gtg ctc cgg cag acg 1408Cys Gly
Ser Leu Cys Cys Gly Arg Gly His Asn Val Leu Arg Gln Thr 345
350 355cga gtg gag cgc tgc cac tgt cgt ttc cac tgg
tgc tgt tat gtg ctg 1456Arg Val Glu Arg Cys His Cys Arg Phe His Trp
Cys Cys Tyr Val Leu360 365 370
375tgt gat gag tgt aaa gtc aca gag tgg gtc aat gtg tgt aaa tga
1501Cys Asp Glu Cys Lys Val Thr Glu Trp Val Asn Val Cys Lys
380 385 390aggtgagcct cgcctaggca
cgacgaggag gagaagcact gtgtgagggc tgctctcttt 1561cagccctttg ctcggatttc
tgtctagggt ttatcgtggc tcccggaagc tcagagcatc 1621tgcctgagaa cagctctggg
ggtgtagggt caggtgaaat ctgtaacgag cagccttttg 1681tgggggaagt ggccccacac
tctgttctta aacactcgaa tagactaaga tgaaatgcac 1741tgtactgtta gcgtcttctc
tacctacagc tccctcgggc tcaggttcct acttcctttg 1801gatagggagt ctatcttttg
gccactcctc ttcctcgaag gataatagca ggcattgtgt 1861ggagtcaata agacccgtat
atatagcaag agaccacctc ttcctatttg tggttctcaa 1921actcctccac tacagcccag
aacctcctct tatgggacct cgggtgacaa taatgagagg 1981ttttcggttg gaaaaggaca
gagggcaggg aagcctcaga cagctgtctt gtcaggctct 2041tgggaggctt ctccttccgt
tcagttgttg aaagggtctc tccaaaggaa aggttttagc 2101cataactctt ggaggccctt
ttccttcttc agcaggaagg gtgggaatgg ataatttatt 2161ttactgagat gtgttcttgg
ttcctgtttg aaactaaaat aaattaagtt actg 221572389PRTMus musculus
72Met Leu Glu Glu Pro Arg Ser Arg Pro Pro Pro Leu Gly Leu Ala Gly1
5 10 15Leu Leu Phe Leu Ala Leu
Phe Ser Arg Ala Leu Ser Asn Glu Ile Leu 20 25
30Gly Leu Lys Leu Pro Gly Glu Pro Pro Leu Thr Ala Asn
Thr Val Cys 35 40 45Leu Thr Leu
Ser Gly Leu Ser Lys Arg Gln Leu Gly Leu Cys Leu Arg 50
55 60Ser Pro Asp Val Thr Ala Ser Ala Leu Gln Gly Leu
His Ile Ala Val65 70 75
80His Glu Cys Gln His Gln Leu Arg Asp Gln Arg Trp Asn Cys Ser Ala
85 90 95Leu Glu Gly Gly Gly Arg
Leu Pro His His Ser Ala Ile Leu Lys Arg 100
105 110Gly Phe Arg Glu Ser Ala Phe Ser Phe Ser Met Leu
Ala Ala Gly Val 115 120 125Met His
Ala Val Ala Thr Ala Cys Ser Leu Gly Lys Leu Val Ser Cys 130
135 140Gly Cys Gly Trp Lys Gly Ser Gly Glu Gln Asp
Arg Leu Arg Ala Lys145 150 155
160Leu Leu Gln Leu Gln Ala Leu Ser Arg Gly Lys Thr Phe Pro Ile Ser
165 170 175Gln Pro Ser Pro
Val Pro Gly Ser Val Pro Ser Pro Gly Pro Gln Asp 180
185 190Thr Trp Glu Trp Gly Gly Cys Asn His Asp Met
Asp Phe Gly Glu Lys 195 200 205Phe
Ser Arg Asp Phe Leu Asp Ser Arg Glu Ala Pro Arg Asp Ile Gln 210
215 220Ala Arg Met Arg Ile His Asn Asn Arg Val
Gly Arg Gln Val Val Thr225 230 235
240Glu Asn Leu Lys Arg Lys Cys Lys Cys His Gly Thr Ser Gly Ser
Cys 245 250 255Gln Phe Lys
Thr Cys Trp Arg Ala Ala Pro Glu Phe Arg Ala Ile Gly 260
265 270Ala Ala Leu Arg Glu Arg Leu Ser Arg Ala
Ile Phe Ile Asp Thr His 275 280
285Asn Arg Asn Ser Gly Ala Phe Gln Pro Arg Leu Arg Pro Arg Arg Leu 290
295 300Ser Gly Glu Leu Val Tyr Phe Glu
Lys Ser Pro Asp Phe Cys Glu Arg305 310
315 320Asp Pro Thr Leu Gly Ser Pro Gly Thr Arg Gly Arg
Ala Cys Asn Lys 325 330
335Thr Ser Arg Leu Leu Asp Gly Cys Gly Ser Leu Cys Cys Gly Arg Gly
340 345 350His Asn Val Leu Arg Gln
Thr Arg Val Glu Arg Cys His Cys Arg Phe 355 360
365His Trp Cys Cys Tyr Val Leu Cys Asp Glu Cys Lys Val Thr
Glu Trp 370 375 380Val Asn Val Cys
Lys385731821DNAMus musculusCDS(271)..(1335) 73gaattcgggc ctaatccgag
cctgacgccg gcgggtctcg ggcggttcgg ggagagagcg 60gactccttcc tcgctcagcc
tccccggccc gacccctcct ttgtaatttg aataaaacgc 120ctcccagccc gcgcgccgcc
ttaacccgcc gccctgttct ccgtgattgc aggcggcgtg 180cgcgcaggaa cagcagcggt
ggcctgcagg cggcggagtt cggtgcggct cctgcagggt 240gcgacccccg ggacgccggg
ccgcgcgacg atg agg gcg cgg ccg cag gtc tgc 294
Met Arg Ala Arg Pro Gln Val Cys
1 5gag gct ctg ctc ttt gcc ttg gcg ctc cac acc ggc gtg tgc
tat ggc 342Glu Ala Leu Leu Phe Ala Leu Ala Leu His Thr Gly Val Cys
Tyr Gly 10 15 20atc aag tgg ctg gca
ctg tcc aag act ccg gca gcc ttg gca ctg aat 390Ile Lys Trp Leu Ala
Leu Ser Lys Thr Pro Ala Ala Leu Ala Leu Asn25 30
35 40cag acg caa cac tgt aaa cag ctg gag ggc
ctg gtg tct gcg cag gtg 438Gln Thr Gln His Cys Lys Gln Leu Glu Gly
Leu Val Ser Ala Gln Val 45 50
55cag ctc tgc cgc agc aac ctg gag ctc atg cgc acc atc gtg cac gcc
486Gln Leu Cys Arg Ser Asn Leu Glu Leu Met Arg Thr Ile Val His Ala
60 65 70gcc cgg ggg gcc atg aag
gcc tgc cgt agg gcc ttc gct gac atg cgc 534Ala Arg Gly Ala Met Lys
Ala Cys Arg Arg Ala Phe Ala Asp Met Arg 75 80
85tgg aac tgc tcc tcc atc gag ctc gcc ccc aac tac ctg ctt
gac ctg 582Trp Asn Cys Ser Ser Ile Glu Leu Ala Pro Asn Tyr Leu Leu
Asp Leu 90 95 100gag aga ggt aca cgg
gag tca gcc ttc gtg tat gcc ctg tcg gcc gcc 630Glu Arg Gly Thr Arg
Glu Ser Ala Phe Val Tyr Ala Leu Ser Ala Ala105 110
115 120acc atc agt cac acc atc gcc cgg gcc tgc
acc tct ggc gac ctg ccc 678Thr Ile Ser His Thr Ile Ala Arg Ala Cys
Thr Ser Gly Asp Leu Pro 125 130
135ggc tgc tcc tgc ggc ccc gtc cca ggt gag cca ccc ggg ccc ggg aac
726Gly Cys Ser Cys Gly Pro Val Pro Gly Glu Pro Pro Gly Pro Gly Asn
140 145 150cgc tgg gga gga tgt gcg
gac aac ctc agc tac ggg ctc ctc atg ggg 774Arg Trp Gly Gly Cys Ala
Asp Asn Leu Ser Tyr Gly Leu Leu Met Gly 155 160
165gcc aag ttt tcc gat gct cct atg aag gtg aaa aaa aca gga
tcc caa 822Ala Lys Phe Ser Asp Ala Pro Met Lys Val Lys Lys Thr Gly
Ser Gln 170 175 180gcc aat aaa ctg atg
cgt cta cac aac agt gaa gtg ggg aga cag gct 870Ala Asn Lys Leu Met
Arg Leu His Asn Ser Glu Val Gly Arg Gln Ala185 190
195 200cta cgt gcc tcc ctg gaa acg aag tgt aaa
tgc cat ggg gtg tct ggc 918Leu Arg Ala Ser Leu Glu Thr Lys Cys Lys
Cys His Gly Val Ser Gly 205 210
215tcc tgc tcc atc cgc acc tgt tgg aag ggg ctg caa gag ctc cag gac
966Ser Cys Ser Ile Arg Thr Cys Trp Lys Gly Leu Gln Glu Leu Gln Asp
220 225 230gtg gct gct gac ctc aag
acc cgc tac ctg tca gcc acg aag gtg gta 1014Val Ala Ala Asp Leu Lys
Thr Arg Tyr Leu Ser Ala Thr Lys Val Val 235 240
245cac cgg cct atg ggc acc cgc aaa cac ttg gtg ccc aag gac
ctg gat 1062His Arg Pro Met Gly Thr Arg Lys His Leu Val Pro Lys Asp
Leu Asp 250 255 260atc cgg cct gtg aag
gac tca gaa ctt gtg tat cta cag agc tcc cct 1110Ile Arg Pro Val Lys
Asp Ser Glu Leu Val Tyr Leu Gln Ser Ser Pro265 270
275 280gac ttc tgc atg aag aat gag aag gtg gga
tcc cat ggg acc caa gac 1158Asp Phe Cys Met Lys Asn Glu Lys Val Gly
Ser His Gly Thr Gln Asp 285 290
295agg cag tgc aac aag act tcc aac ggc agt gac agc tgc gac ctc atg
1206Arg Gln Cys Asn Lys Thr Ser Asn Gly Ser Asp Ser Cys Asp Leu Met
300 305 310tgc tgt ggg cgc ggc tac
aac ccc tac acg gac aga gtg gtg gag cga 1254Cys Cys Gly Arg Gly Tyr
Asn Pro Tyr Thr Asp Arg Val Val Glu Arg 315 320
325tgt cac tgc aag tac cac tgg tgc tgc tac gtc acc tgc cgc
agg tgt 1302Cys His Cys Lys Tyr His Trp Cys Cys Tyr Val Thr Cys Arg
Arg Cys 330 335 340gag cgc acg gtg gag
cgc tac gtc tgc aag tga gaccatatgc cccacccctg 1355Glu Arg Thr Val Glu
Arg Tyr Val Cys Lys345 350 355aggaggggtg
ctgctcctct gaggacccac tcaagggcct agagaccttg gtggacttcc 1415ctgcagatgc
cagatgccag gcgtgggagg cggcttgtgc tgtgcctcca cttggaagac 1475accacaccag
gaggcctggt cgccctggga gagccggggc ttcaaaggaa actgatagga 1535ttaaaaataa
cctggcagcc tggggcctga gtgccacatg ttgccttcca ggctgctcca 1595agaagtcagg
gcagggatgg gtaagactgt gcatttgacc tttcaaggcc agaaagaccg 1655gctttctgga
atgttctttg ggaccctgtg cccaccacat ggaaccacta acttgggttg 1715taaattttta
ttttccttcc cctctccgtg ggatgtggga gttacagaaa tatttataaa 1775aatacagctt
tttcctttgg gggtgaaaaa aaaaaaaaaa gaattc 182174354PRTMus
musculus 74Met Arg Ala Arg Pro Gln Val Cys Glu Ala Leu Leu Phe Ala Leu
Ala1 5 10 15Leu His Thr
Gly Val Cys Tyr Gly Ile Lys Trp Leu Ala Leu Ser Lys 20
25 30Thr Pro Ala Ala Leu Ala Leu Asn Gln Thr
Gln His Cys Lys Gln Leu 35 40
45Glu Gly Leu Val Ser Ala Gln Val Gln Leu Cys Arg Ser Asn Leu Glu 50
55 60Leu Met Arg Thr Ile Val His Ala Ala
Arg Gly Ala Met Lys Ala Cys65 70 75
80Arg Arg Ala Phe Ala Asp Met Arg Trp Asn Cys Ser Ser Ile
Glu Leu 85 90 95Ala Pro
Asn Tyr Leu Leu Asp Leu Glu Arg Gly Thr Arg Glu Ser Ala 100
105 110Phe Val Tyr Ala Leu Ser Ala Ala Thr
Ile Ser His Thr Ile Ala Arg 115 120
125Ala Cys Thr Ser Gly Asp Leu Pro Gly Cys Ser Cys Gly Pro Val Pro
130 135 140Gly Glu Pro Pro Gly Pro Gly
Asn Arg Trp Gly Gly Cys Ala Asp Asn145 150
155 160Leu Ser Tyr Gly Leu Leu Met Gly Ala Lys Phe Ser
Asp Ala Pro Met 165 170
175Lys Val Lys Lys Thr Gly Ser Gln Ala Asn Lys Leu Met Arg Leu His
180 185 190Asn Ser Glu Val Gly Arg
Gln Ala Leu Arg Ala Ser Leu Glu Thr Lys 195 200
205Cys Lys Cys His Gly Val Ser Gly Ser Cys Ser Ile Arg Thr
Cys Trp 210 215 220Lys Gly Leu Gln Glu
Leu Gln Asp Val Ala Ala Asp Leu Lys Thr Arg225 230
235 240Tyr Leu Ser Ala Thr Lys Val Val His Arg
Pro Met Gly Thr Arg Lys 245 250
255His Leu Val Pro Lys Asp Leu Asp Ile Arg Pro Val Lys Asp Ser Glu
260 265 270Leu Val Tyr Leu Gln
Ser Ser Pro Asp Phe Cys Met Lys Asn Glu Lys 275
280 285Val Gly Ser His Gly Thr Gln Asp Arg Gln Cys Asn
Lys Thr Ser Asn 290 295 300Gly Ser Asp
Ser Cys Asp Leu Met Cys Cys Gly Arg Gly Tyr Asn Pro305
310 315 320Tyr Thr Asp Arg Val Val Glu
Arg Cys His Cys Lys Tyr His Trp Cys 325
330 335Cys Tyr Val Thr Cys Arg Arg Cys Glu Arg Thr Val
Glu Arg Tyr Val 340 345 350Cys
Lys751664DNAMus musculusCDS(279)..(1373) 75gagcagaagg ttctcacctt
ggaaagtgag ggaagctccc gcatctccag ctcatcctca 60cctctgcgcc agaggacctt
aggctacttt ctccgcctta tcttgcctag gggactgctg 120atagtctctg tccttgctgc
cctgtttaat gttaccttcc aggggaaaga gagcaaggaa 180caactgggtg ctaagaaact
gaccccaggc cctgcgggcc tctggagaga ggagacagag 240gaggagtggc tggggctggg
ggtctccatg cgtgggcc atg gac aga gcg gcg ctc 296
Met Asp Arg Ala Ala Leu
1 5ctg gcc ctg ccc agc ttg tgt gcg ctg tgg gca gcc
gtg ctg tcg ctg 344Leu Ala Leu Pro Ser Leu Cys Ala Leu Trp Ala Ala
Val Leu Ser Leu 10 15 20ctc
ccc tgc gga acc cag ggc aac tgg atg tgg ttg ggc atc gcc tct 392Leu
Pro Cys Gly Thr Gln Gly Asn Trp Met Trp Leu Gly Ile Ala Ser 25
30 35ttc ggg gta ccg gag aag ctg ggc tgc
gcc gac ttg ccg ctg aac agc 440Phe Gly Val Pro Glu Lys Leu Gly Cys
Ala Asp Leu Pro Leu Asn Ser 40 45
50cgc cag aag gag ctg tgc aag agg aaa ccg tac ctg ctg cct agc atc
488Arg Gln Lys Glu Leu Cys Lys Arg Lys Pro Tyr Leu Leu Pro Ser Ile55
60 65 70cgc gag ggc gcc agg
ctg ggc att cag gag tgc aga agc cag ttc cga 536Arg Glu Gly Ala Arg
Leu Gly Ile Gln Glu Cys Arg Ser Gln Phe Arg 75
80 85cac gag agg tgg aac tgt atg gtc gcc act acc
act tcc acc cag ctc 584His Glu Arg Trp Asn Cys Met Val Ala Thr Thr
Thr Ser Thr Gln Leu 90 95
100gcc aca gcc ccc ctc ttt ggc tat gag ctg agt agc ggc acc aag gag
632Ala Thr Ala Pro Leu Phe Gly Tyr Glu Leu Ser Ser Gly Thr Lys Glu
105 110 115aca gca ttc att tat gcc atc
atg gca gcg ggc ctg gtg cac tct gtc 680Thr Ala Phe Ile Tyr Ala Ile
Met Ala Ala Gly Leu Val His Ser Val 120 125
130acc agg tca tgc agt gca ggc aac atg acc gaa tgt tcc tgt gaa acc
728Thr Arg Ser Cys Ser Ala Gly Asn Met Thr Glu Cys Ser Cys Glu Thr135
140 145 150acc ttg cag aat
ggt ggc tca cca agt gaa ggc tgg cac tgg gga gga 776Thr Leu Gln Asn
Gly Gly Ser Pro Ser Glu Gly Trp His Trp Gly Gly 155
160 165tgc tcg gat gat gtc cag tac ggc atg tgg
ttc agc aga aag ttt cta 824Cys Ser Asp Asp Val Gln Tyr Gly Met Trp
Phe Ser Arg Lys Phe Leu 170 175
180gat ctt ccc atc aga aac acc aca gga aaa gaa agc aga gtc ctg cta
872Asp Leu Pro Ile Arg Asn Thr Thr Gly Lys Glu Ser Arg Val Leu Leu
185 190 195gcc atg aat cta cac aac aac
gaa gcg ggg cgg cag gct gtc gcc aag 920Ala Met Asn Leu His Asn Asn
Glu Ala Gly Arg Gln Ala Val Ala Lys 200 205
210tta atg tct gtg gac tgc cgc tgc cac gga gtt tcc ggc tcc tgt gct
968Leu Met Ser Val Asp Cys Arg Cys His Gly Val Ser Gly Ser Cys Ala215
220 225 230gtg aaa acc tgc
tgg aaa act atg tct tct ttt gaa aag att ggg cat 1016Val Lys Thr Cys
Trp Lys Thr Met Ser Ser Phe Glu Lys Ile Gly His 235
240 245ttt tta aag gat aaa tat gaa aac agc atc
cag atc tca gac aaa acc 1064Phe Leu Lys Asp Lys Tyr Glu Asn Ser Ile
Gln Ile Ser Asp Lys Thr 250 255
260aag agg aaa atg cgc agg aga gaa aaa gac cag agg cag acc ccc att
1112Lys Arg Lys Met Arg Arg Arg Glu Lys Asp Gln Arg Gln Thr Pro Ile
265 270 275ctc aag gat gac ttg ctg tac
gtt cat aag tct ccc aac tac tgc gtg 1160Leu Lys Asp Asp Leu Leu Tyr
Val His Lys Ser Pro Asn Tyr Cys Val 280 285
290gag aac aag aaa ctg ggg att cct ggg acc cag ggc aga gag tgc aac
1208Glu Asn Lys Lys Leu Gly Ile Pro Gly Thr Gln Gly Arg Glu Cys Asn295
300 305 310cgg aca tca gga
ggc gca gat ggc tgt aac ctc ctc tgc tgt ggc cga 1256Arg Thr Ser Gly
Gly Ala Asp Gly Cys Asn Leu Leu Cys Cys Gly Arg 315
320 325ggc tac aac acc cat gta gtc agg cac gtg
gag agg tgt gag tgt aag 1304Gly Tyr Asn Thr His Val Val Arg His Val
Glu Arg Cys Glu Cys Lys 330 335
340ttt atc tgg tgc tgc tac gtc cgc tgc agg agg tgt gaa agt atg acc
1352Phe Ile Trp Cys Cys Tyr Val Arg Cys Arg Arg Cys Glu Ser Met Thr
345 350 355gat gtc cac acg tgt aag taa
cctctccgtc cagcctagca tgagacgcct 1403Asp Val His Thr Cys Lys
360 365ctgtagtaac caaggtgtgg tgttggcatc tggagggcgc
ccctactgtg cactgatggg 1463gaagtcgctg cctgtaagag tgttcccaga cccctgggct
agtctacgat ttctttcttt 1523ctggcaggct tcaaatcaca agctgatcca gaggattgct
tgggattctg aagttgaaaa 1583ggttggcagt cgcctttgga tgatttggga aatatacatt
gatatacagg aaacatcaaa 1643tctgtttctg aagcaatgtg g
166476364PRTMus musculus 76Met Asp Arg Ala Ala Leu
Leu Ala Leu Pro Ser Leu Cys Ala Leu Trp1 5
10 15Ala Ala Val Leu Ser Leu Leu Pro Cys Gly Thr Gln
Gly Asn Trp Met 20 25 30Trp
Leu Gly Ile Ala Ser Phe Gly Val Pro Glu Lys Leu Gly Cys Ala 35
40 45Asp Leu Pro Leu Asn Ser Arg Gln Lys
Glu Leu Cys Lys Arg Lys Pro 50 55
60Tyr Leu Leu Pro Ser Ile Arg Glu Gly Ala Arg Leu Gly Ile Gln Glu65
70 75 80Cys Arg Ser Gln Phe
Arg His Glu Arg Trp Asn Cys Met Val Ala Thr 85
90 95Thr Thr Ser Thr Gln Leu Ala Thr Ala Pro Leu
Phe Gly Tyr Glu Leu 100 105
110Ser Ser Gly Thr Lys Glu Thr Ala Phe Ile Tyr Ala Ile Met Ala Ala
115 120 125Gly Leu Val His Ser Val Thr
Arg Ser Cys Ser Ala Gly Asn Met Thr 130 135
140Glu Cys Ser Cys Glu Thr Thr Leu Gln Asn Gly Gly Ser Pro Ser
Glu145 150 155 160Gly Trp
His Trp Gly Gly Cys Ser Asp Asp Val Gln Tyr Gly Met Trp
165 170 175Phe Ser Arg Lys Phe Leu Asp
Leu Pro Ile Arg Asn Thr Thr Gly Lys 180 185
190Glu Ser Arg Val Leu Leu Ala Met Asn Leu His Asn Asn Glu
Ala Gly 195 200 205Arg Gln Ala Val
Ala Lys Leu Met Ser Val Asp Cys Arg Cys His Gly 210
215 220Val Ser Gly Ser Cys Ala Val Lys Thr Cys Trp Lys
Thr Met Ser Ser225 230 235
240Phe Glu Lys Ile Gly His Phe Leu Lys Asp Lys Tyr Glu Asn Ser Ile
245 250 255Gln Ile Ser Asp Lys
Thr Lys Arg Lys Met Arg Arg Arg Glu Lys Asp 260
265 270Gln Arg Gln Thr Pro Ile Leu Lys Asp Asp Leu Leu
Tyr Val His Lys 275 280 285Ser Pro
Asn Tyr Cys Val Glu Asn Lys Lys Leu Gly Ile Pro Gly Thr 290
295 300Gln Gly Arg Glu Cys Asn Arg Thr Ser Gly Gly
Ala Asp Gly Cys Asn305 310 315
320Leu Leu Cys Cys Gly Arg Gly Tyr Asn Thr His Val Val Arg His Val
325 330 335Glu Arg Cys Glu
Cys Lys Phe Ile Trp Cys Cys Tyr Val Arg Cys Arg 340
345 350Arg Cys Glu Ser Met Thr Asp Val His Thr Cys
Lys 355 36077313PRTHomo sapiens 77Met Gly Ile Gly
Arg Ser Glu Gly Gly Arg Arg Gly Ala Leu Gly Val1 5
10 15Leu Leu Ala Leu Gly Ala Ala Leu Leu Ala
Val Gly Ser Ala Ser Glu 20 25
30Tyr Asp Tyr Val Ser Phe Gln Ser Asp Ile Gly Pro Tyr Gln Ser Gly
35 40 45Arg Phe Tyr Thr Lys Pro Pro Gln
Cys Val Asp Ile Pro Ala Asp Leu 50 55
60Arg Leu Cys His Asn Val Gly Tyr Lys Lys Met Val Leu Pro Asn Leu65
70 75 80Leu Glu His Glu Thr
Met Ala Glu Val Lys Gln Gln Ala Ser Ser Trp 85
90 95Val Pro Leu Leu Asn Lys Asn Cys His Ala Gly
Thr Gln Val Phe Leu 100 105
110Cys Ser Leu Phe Ala Pro Val Cys Leu Asp Arg Pro Ile Tyr Pro Cys
115 120 125Arg Trp Leu Cys Glu Ala Val
Arg Asp Ser Cys Glu Pro Val Met Gln 130 135
140Phe Phe Gly Phe Tyr Trp Pro Glu Met Leu Lys Cys Asp Lys Phe
Pro145 150 155 160Glu Gly
Asp Val Cys Ile Ala Met Thr Pro Pro Asn Ala Thr Glu Ala
165 170 175Ser Lys Pro Gln Gly Thr Thr
Val Cys Pro Pro Cys Asp Asn Glu Leu 180 185
190Lys Ser Glu Ala Ile Ile Glu His Leu Cys Ala Ser Glu Phe
Ala Leu 195 200 205Arg Met Lys Ile
Lys Glu Val Lys Lys Glu Asn Gly Asp Lys Lys Ile 210
215 220Val Pro Lys Lys Lys Lys Pro Leu Lys Leu Gly Pro
Ile Lys Lys Lys225 230 235
240Asp Leu Lys Lys Leu Val Leu Tyr Leu Lys Asn Gly Ala Asp Cys Pro
245 250 255Cys His Gln Leu Asp
Asn Leu Ser His His Phe Leu Ile Met Gly Arg 260
265 270Lys Val Lys Ser Gln Tyr Leu Leu Thr Ala Ile His
Lys Trp Asp Lys 275 280 285Lys Asn
Lys Glu Phe Lys Asn Phe Met Lys Lys Met Lys Asn His Glu 290
295 300Cys Pro Thr Phe Gln Ser Val Phe Lys305
31078295PRTHomo sapiens 78Met Leu Gln Gly Pro Gly Ser Leu Leu
Leu Leu Phe Leu Ala Ser His1 5 10
15Cys Cys Leu Gly Ser Ala Arg Gly Leu Phe Leu Phe Gly Gln Pro
Asp 20 25 30Phe Ser Tyr Lys
Arg Ser Asn Cys Lys Pro Ile Pro Ala Asn Leu Gln 35
40 45Leu Cys His Gly Ile Glu Tyr Gln Asn Met Arg Leu
Pro Asn Leu Leu 50 55 60Gly His Glu
Thr Met Lys Glu Val Leu Glu Gln Ala Gly Ala Trp Ile65 70
75 80Pro Leu Val Met Lys Gln Cys His
Pro Asp Thr Lys Lys Phe Leu Cys 85 90
95Ser Leu Phe Ala Pro Val Cys Leu Asp Asp Leu Asp Glu Thr
Ile Gln 100 105 110Pro Cys His
Ser Leu Cys Val Gln Val Lys Asp Arg Cys Ala Pro Val 115
120 125Met Ser Ala Phe Gly Phe Pro Trp Pro Asp Met
Leu Glu Cys Asp Arg 130 135 140Phe Pro
Gln Asp Asn Asp Leu Cys Ile Pro Leu Ala Ser Ser Asp His145
150 155 160Leu Leu Pro Ala Thr Glu Glu
Ala Pro Lys Val Cys Glu Ala Cys Lys 165
170 175Asn Lys Asn Asp Asp Asp Asn Asp Ile Met Glu Thr
Leu Cys Lys Asn 180 185 190Asp
Phe Ala Leu Lys Ile Lys Val Lys Glu Ile Thr Tyr Ile Asn Arg 195
200 205Asp Thr Lys Ile Ile Leu Glu Thr Lys
Ser Lys Thr Ile Tyr Lys Leu 210 215
220Asn Gly Val Ser Glu Arg Asp Leu Lys Lys Ser Val Leu Trp Leu Lys225
230 235 240Asp Ser Leu Gln
Cys Thr Cys Glu Glu Met Asn Asp Ile Asn Ala Pro 245
250 255Tyr Leu Val Met Gly Gln Lys Gln Gly Gly
Glu Leu Val Ile Thr Ser 260 265
270Val Lys Arg Trp Gln Lys Gly Gln Arg Glu Phe Lys Arg Ile Ser Arg
275 280 285Ser Ile Arg Lys Leu Gln Cys
290 29579325PRTHomo sapiens 79Met Val Cys Gly Ser Pro
Gly Gly Met Leu Leu Leu Arg Ala Gly Leu1 5
10 15Leu Ala Leu Ala Ala Leu Cys Leu Leu Arg Val Pro
Gly Ala Arg Ala 20 25 30Ala
Ala Cys Glu Pro Val Arg Ile Pro Leu Cys Lys Ser Leu Pro Trp 35
40 45Asn Met Thr Lys Met Pro Asn His Leu
His His Ser Thr Gln Ala Asn 50 55
60Ala Ile Leu Ala Ile Glu Gln Phe Glu Gly Leu Leu Gly Thr His Cys65
70 75 80Ser Pro Asp Leu Leu
Phe Phe Leu Cys Ala Met Tyr Ala Pro Ile Cys 85
90 95Thr Ile Asp Phe Gln His Glu Pro Ile Lys Pro
Cys Lys Ser Val Cys 100 105
110Glu Arg Ala Arg Gln Gly Cys Glu Pro Ile Leu Ile Lys Tyr Arg His
115 120 125Ser Trp Pro Glu Asn Leu Ala
Cys Glu Glu Leu Pro Val Tyr Asp Arg 130 135
140Gly Val Cys Ile Ser Pro Glu Ala Ile Val Thr Ala Asp Gly Ala
Asp145 150 155 160Phe Pro
Met Asp Ser Ser Asn Gly Asn Cys Arg Gly Ala Ser Ser Glu
165 170 175Arg Cys Lys Cys Lys Pro Ile
Arg Ala Thr Gln Lys Thr Tyr Phe Arg 180 185
190Asn Asn Tyr Asn Tyr Val Ile Arg Ala Lys Val Lys Glu Ile
Lys Thr 195 200 205Lys Cys His Asp
Val Thr Ala Val Val Glu Val Lys Glu Ile Leu Lys 210
215 220Ser Ser Leu Val Asn Ile Pro Arg Asp Thr Val Asn
Leu Tyr Thr Ser225 230 235
240Ser Gly Cys Leu Cys Pro Pro Leu Asn Val Asn Glu Glu Tyr Ile Ile
245 250 255Met Gly Tyr Glu Asp
Glu Glu Arg Ser Arg Leu Leu Leu Val Glu Gly 260
265 270Ser Ile Ala Glu Lys Trp Lys Asp Arg Leu Gly Lys
Lys Val Lys Arg 275 280 285Trp Asp
Met Lys Leu Arg His Leu Gly Leu Ser Lys Ser Asp Ser Ser 290
295 300Asn Ser Asp Ser Thr Gln Ser Gln Lys Ser Gly
Arg Asn Ser Asn Pro305 310 315
320Arg Gln Ala Arg Asn 32580314PRTMus musculus 80Met
Gly Val Gly Arg Ser Ala Arg Gly Arg Gly Gly Ala Ala Ser Gly1
5 10 15Val Leu Leu Ala Leu Ala Ala
Ala Leu Leu Ala Ala Gly Ser Ala Ser 20 25
30Glu Tyr Asp Tyr Val Ser Phe Gln Ser Asp Ile Gly Ser Tyr
Gln Ser 35 40 45Gly Arg Phe Tyr
Thr Lys Pro Pro Gln Cys Val Asp Ile Pro Val Asp 50 55
60Leu Arg Leu Cys His Asn Val Gly Tyr Lys Lys Met Val
Leu Pro Asn65 70 75
80Leu Leu Glu His Glu Thr Met Ala Glu Val Lys Gln Gln Ala Ser Ser
85 90 95Trp Val Pro Leu Leu Asn
Lys Asn Cys His Met Gly Thr Gln Val Phe 100
105 110Leu Cys Ser Leu Phe Ala Pro Val Cys Leu Asp Arg
Pro Ile Tyr Pro 115 120 125Cys Arg
Trp Leu Cys Glu Ala Val Arg Asp Ser Cys Glu Pro Val Met 130
135 140Gln Phe Phe Gly Phe Tyr Trp Pro Glu Met Leu
Lys Cys Asp Lys Phe145 150 155
160Pro Glu Gly Asp Val Cys Ile Ala Met Thr Pro Pro Asn Thr Thr Glu
165 170 175Ala Ser Lys Pro
Gln Gly Thr Thr Val Cys Pro Pro Cys Asp Asn Glu 180
185 190Leu Lys Ser Glu Ala Ile Ile Glu His Leu Cys
Ala Ser Glu Phe Ala 195 200 205Leu
Arg Met Lys Ile Lys Glu Val Lys Lys Glu Asn Gly Asp Lys Lys 210
215 220Ile Val Pro Lys Lys Lys Lys Pro Leu Lys
Leu Gly Pro Ile Lys Lys225 230 235
240Lys Glu Leu Lys Ala Leu Val Leu Phe Leu Lys Asn Gly Ala Asp
Cys 245 250 255Pro Cys His
Gln Leu Asp Asn Leu Ser His Asn Phe Leu Ile Met Gly 260
265 270Arg Lys Val Lys Ser Gln Tyr Leu Leu Thr
Ala Ile His Lys Trp Asp 275 280
285Lys Lys Asn Lys Glu Phe Lys Asn Phe Met Lys Arg Met Lys Asn His 290
295 300Glu Cys Pro Thr Phe Gln Ser Val
Phe Lys305 31081295PRTMus musculus 81Met Pro Arg Gly Pro
Ala Ser Leu Leu Leu Leu Val Leu Ala Ser His1 5
10 15Cys Cys Leu Gly Ser Ala Arg Gly Leu Phe Leu
Phe Gly Gln Pro Asp 20 25
30Phe Ser Tyr Lys Arg Ser Asn Cys Lys Pro Ile Pro Ala Asn Leu Gln
35 40 45Leu Cys His Gly Ile Glu Tyr Gln
Asn Met Arg Leu Pro Asn Leu Leu 50 55
60Gly His Glu Thr Met Lys Glu Val Leu Glu Gln Ala Gly Ala Trp Ile65
70 75 80Pro Leu Val Met Lys
Gln Cys His Pro Asp Thr Lys Lys Phe Leu Cys 85
90 95Ser Leu Phe Ala Pro Val Cys Leu Asp Asp Leu
Asp Glu Thr Ile Gln 100 105
110Pro Cys His Ser Leu Cys Val Gln Val Lys Asp Arg Cys Ala Pro Val
115 120 125Met Ser Ala Phe Gly Phe Pro
Trp Pro Asp Met Leu Glu Cys Asp Arg 130 135
140Phe Pro Gln Asp Asn Asp Leu Cys Ile Pro Leu Ala Ser Ser Asp
His145 150 155 160Leu Leu
Pro Ala Thr Glu Glu Ala Pro Lys Val Cys Glu Ala Cys Lys
165 170 175Thr Lys Asn Glu Asp Asp Asn
Asp Ile Met Glu Thr Leu Cys Lys Asn 180 185
190Asp Phe Ala Leu Lys Ile Lys Val Lys Glu Ile Thr Tyr Ile
Asn Arg 195 200 205Asp Thr Lys Ile
Ile Leu Glu Thr Lys Ser Lys Thr Ile Tyr Lys Leu 210
215 220Asn Gly Val Ser Glu Arg Asp Leu Lys Lys Ser Val
Leu Trp Leu Lys225 230 235
240Asp Ser Leu Gln Cys Thr Cys Glu Glu Met Asn Asp Ile Asn Ala Pro
245 250 255Tyr Leu Val Met Gly
Gln Lys Gln Gly Gly Glu Leu Val Ile Thr Ser 260
265 270Val Lys Arg Trp Gln Lys Gly Gln Arg Glu Phe Lys
Arg Ile Ser Arg 275 280 285Ser Ile
Arg Lys Leu Gln Cys 290 29582323PRTMus musculus 82Met
Val Cys Cys Gly Pro Gly Arg Met Leu Leu Gly Trp Ala Gly Leu1
5 10 15Leu Val Leu Ala Ala Leu Cys
Leu Leu Gln Val Pro Gly Ala Gln Ala 20 25
30Ala Ala Cys Glu Pro Val Arg Ile Pro Leu Cys Lys Ser Leu
Pro Trp 35 40 45Asn Met Thr Lys
Met Pro Asn His Leu His His Ser Thr Gln Ala Asn 50 55
60Ala Ile Leu Ala Met Glu Gln Phe Glu Gly Leu Leu Gly
Thr His Cys65 70 75
80Ser Pro Asp Leu Leu Phe Phe Leu Cys Ala Met Tyr Ala Pro Ile Cys
85 90 95Thr Ile Asp Phe Gln His
Glu Pro Ile Lys Pro Cys Lys Ser Val Cys 100
105 110Glu Arg Ala Arg Gln Gly Cys Glu Pro Ile Leu Ile
Lys Tyr Arg His 115 120 125Ser Trp
Pro Glu Ser Leu Ala Cys Asp Glu Leu Pro Val Tyr Asp Arg 130
135 140Gly Val Cys Ile Ser Pro Glu Ala Ile Val Thr
Ala Asp Gly Ala Asp145 150 155
160Phe Pro Met Asp Ser Ser Thr Gly His Cys Arg Gly Ala Ser Ser Glu
165 170 175Arg Cys Lys Cys
Lys Pro Val Arg Ala Thr Gln Lys Thr Tyr Phe Arg 180
185 190Asn Asn Tyr Asn Tyr Val Ile Arg Ala Lys Val
Lys Glu Val Lys Met 195 200 205Lys
Cys His Asp Val Thr Ala Val Val Glu Val Lys Glu Ile Leu Lys 210
215 220Ala Ser Leu Val Asn Ile Pro Arg Asp Thr
Val Asn Leu Tyr Thr Thr225 230 235
240Ser Gly Cys Leu Cys Pro Pro Leu Thr Val Asn Glu Glu Tyr Val
Ile 245 250 255Met Gly Tyr
Glu Asp Glu Glu Arg Ser Arg Leu Leu Leu Val Glu Gly 260
265 270Ser Ile Ala Glu Lys Trp Lys Asp Arg Leu
Gly Lys Lys Val Lys Arg 275 280
285Trp Asp Met Lys Leu Arg His Leu Gly Leu Gly Lys Thr Asp Ala Ser 290
295 300Asp Ser Thr Gln Asn Gln Lys Ser
Gly Arg Asn Ser Asn Pro Arg Pro305 310
315 320Ala Arg Ser83604PRTHomo sapiens 83Met Arg Gly Ala
Ala Arg Leu Gly Arg Pro Gly Arg Ser Cys Leu Pro1 5
10 15Gly Pro Ala Leu Arg Ala Ala Ala Ala Pro
Ala Leu Leu Leu Ala Arg 20 25
30Cys Ala Val Ala Ala Ala Ala Gly Leu Arg Ala Ala Ala Arg Pro Arg
35 40 45Pro Pro Glu Leu Gln Ser Ala Ser
Ala Gly Pro Ser Val Ser Leu Tyr 50 55
60Leu Ser Glu Asp Glu Val Arg Arg Leu Ile Gly Leu Asp Ala Glu Leu65
70 75 80Tyr Tyr Val Arg Asn
Asp Leu Ile Ser His Tyr Ala Leu Ser Phe Asn 85
90 95Leu Leu Val Pro Ser Glu Thr Asn Phe Leu His
Phe Thr Trp His Ala 100 105
110Lys Ser Lys Val Glu Tyr Lys Leu Gly Phe Gln Val Asp Asn Val Leu
115 120 125Ala Met Asp Met Pro Gln Val
Asn Ile Ser Val Gln Gly Glu Val Pro 130 135
140Arg Thr Leu Ser Val Phe Arg Val Glu Leu Ser Cys Thr Gly Lys
Val145 150 155 160Asp Ser
Glu Val Met Ile Leu Met Gln Leu Asn Leu Thr Val Asn Ser
165 170 175Ser Lys Asn Phe Thr Val Leu
Asn Phe Lys Arg Arg Lys Met Cys Tyr 180 185
190Lys Lys Leu Glu Glu Val Lys Thr Ser Ala Leu Asp Lys Asn
Thr Ser 195 200 205Arg Thr Ile Tyr
Asp Pro Val His Ala Ala Pro Thr Thr Ser Thr Arg 210
215 220Val Phe Tyr Ile Ser Val Gly Val Cys Cys Ala Val
Ile Phe Leu Val225 230 235
240Ala Ile Ile Leu Ala Val Leu His Leu His Asn Met Lys Arg Ile Glu
245 250 255Leu Asp Asp Ser Ile
Ser Ala Ser Ser Ser Ser Gln Gly Leu Ser Gln 260
265 270Pro Ser Thr Gln Thr Thr Gln Tyr Leu Arg Ala Asp
Thr Pro Asn Asn 275 280 285Ala Thr
Pro Ile Thr Ser Tyr Pro Thr Leu Arg Ile Glu Lys Asn Asp 290
295 300Leu Arg Ser Val Thr Leu Leu Glu Ala Lys Gly
Lys Val Lys Asp Ile305 310 315
320Ala Ile Ser Arg Glu Arg Ile Thr Leu Lys Asp Val Leu Gln Glu Gly
325 330 335Thr Phe Gly Arg
Ile Phe His Gly Ile Leu Ile Asp Glu Lys Asp Pro 340
345 350Asn Lys Glu Lys Gln Ala Phe Val Lys Thr Val
Lys Asp Gln Ala Ser 355 360 365Glu
Ile Gln Val Thr Met Met Leu Thr Glu Ser Cys Lys Leu Arg Gly 370
375 380Leu His His Arg Asn Leu Leu Pro Ile Thr
His Val Cys Ile Glu Glu385 390 395
400Gly Glu Lys Pro Met Val Ile Leu Pro Tyr Met Asn Trp Gly Asn
Leu 405 410 415Lys Leu Phe
Leu Arg Gln Cys Lys Leu Val Glu Ala Asn Asn Pro Gln 420
425 430Ala Ile Ser Gln Gln Asp Leu Val His Met
Ala Ile Gln Ile Ala Cys 435 440
445Gly Met Ser Tyr Leu Ala Arg Arg Glu Val Ile His Lys Asp Leu Ala 450
455 460Ala Arg Asn Cys Val Ile Asp Asp
Thr Leu Gln Val Lys Ile Thr Asp465 470
475 480Asn Ala Leu Ser Arg Asp Leu Phe Pro Met Asp Tyr
His Cys Leu Gly 485 490
495Asp Asn Glu Asn Arg Pro Val Arg Trp Met Ala Leu Glu Ser Leu Val
500 505 510Asn Asn Glu Phe Ser Ser
Ala Ser Asp Val Trp Ala Phe Gly Val Thr 515 520
525Leu Trp Glu Leu Met Thr Leu Gly Gln Thr Pro Tyr Val Asp
Ile Asp 530 535 540Pro Phe Glu Met Ala
Ala Tyr Leu Lys Asp Gly Tyr Arg Ile Ala Gln545 550
555 560Pro Ile Asn Cys Pro Asp Glu Leu Phe Ala
Val Met Ala Cys Cys Trp 565 570
575Ala Leu Asp Pro Glu Glu Arg Pro Lys Phe Gln Gln Leu Val Gln Cys
580 585 590Leu Thr Glu Phe His
Ala Ala Leu Gly Ala Tyr Val 595 60084405PRTMus
musculus 84Leu Asp Lys Asn Thr Ser Arg Thr Ile Tyr Asp Pro Val His Ala
Ala1 5 10 15Pro Thr Thr
Ser Thr Arg Val Phe Tyr Ile Ser Val Gly Val Cys Cys 20
25 30Ala Val Ile Phe Leu Val Ala Ile Ile Leu
Ala Val Leu His Leu His 35 40
45Ser Met Lys Arg Ile Glu Leu Asp Asp Ser Ile Ser Ala Ser Ser Ser 50
55 60Ser Gln Gly Leu Ser Gln Pro Ser Thr
Gln Thr Thr Gln Tyr Leu Arg65 70 75
80Ala Asp Thr Pro Asn Asn Ala Thr Pro Ile Thr Ser Ser Ser
Gly Tyr 85 90 95Pro Thr
Leu Arg Ile Glu Lys Asn Asp Leu Arg Ser Val Thr Leu Leu 100
105 110Glu Ala Lys Ala Lys Val Lys Asp Ile
Ala Ile Ser Arg Glu Arg Ile 115 120
125Thr Leu Lys Asp Val Leu Gln Glu Gly Thr Phe Gly Arg Ile Phe His
130 135 140Gly Ile Leu Val Asp Glu Lys
Asp Pro Asn Lys Glu Lys Gln Thr Phe145 150
155 160Val Lys Thr Val Lys Asp Gln Ala Ser Glu Val Gln
Val Thr Met Met 165 170
175Leu Thr Glu Ser Cys Lys Leu Arg Gly Leu His His Arg Asn Leu Leu
180 185 190Pro Ile Thr His Val Cys
Ile Glu Glu Gly Glu Lys Pro Met Val Val 195 200
205Leu Pro Tyr Met Asn Trp Gly Asn Leu Lys Leu Phe Leu Arg
Gln Cys 210 215 220Lys Leu Val Glu Ala
Asn Asn Pro Gln Ala Ile Ser Gln Gln Asp Leu225 230
235 240Val His Met Ala Ile Gln Ile Ala Cys Gly
Met Ser Tyr Leu Ala Arg 245 250
255Arg Glu Val Ile His Arg Asp Leu Ala Ala Arg Asn Cys Val Ile Asp
260 265 270Asp Thr Leu Gln Val
Lys Ile Thr Asp Asn Ala Leu Ser Arg Asp Leu 275
280 285Phe Pro Met Asp Tyr His Cys Leu Gly Asp Asn Glu
Asn Arg Pro Val 290 295 300Arg Trp Met
Ala Leu Glu Ser Leu Val Asn Asn Glu Phe Ser Ser Ala305
310 315 320Ser Asp Val Trp Ala Phe Gly
Val Thr Leu Trp Glu Leu Met Thr Leu 325
330 335Gly Gln Thr Pro Tyr Val Asp Ile Asp Pro Phe Glu
Met Ala Ala Tyr 340 345 350Leu
Lys Asp Gly Tyr Arg Ile Ala Gln Pro Ile Asn Cys Pro Asp Glu 355
360 365Leu Phe Ala Val Met Ala Cys Cys Trp
Ala Leu Asp Pro Glu Glu Arg 370 375
380Pro Lys Phe Gln Gln Leu Val Gln Cys Leu Thr Glu Phe His Ala Ala385
390 395 400Leu Gly Ala Tyr
Val 40585610PRTDrosophila melanogaster 85Met Ala Pro Asn
Leu Leu Thr Ile Gly Leu Leu Leu Thr Leu Ile Ala1 5
10 15Ser Gly Gln Ala His Leu Asn Ile Phe Leu
Asn Leu His Glu Val Leu 20 25
30Arg Leu Ile Gly Val Ser Ala Glu Leu Tyr Tyr Val Arg Glu Gly Ala
35 40 45Ile Asn Asp Tyr Ala Leu Asn Phe
Ala Val Pro Val Pro Ala Asn Ile 50 55
60Ser Asp Val Thr Phe Thr Trp Gln Ser Leu Val Asp His Pro Leu Pro65
70 75 80Tyr Ser Ile Asn Ile
Ala Thr Ser Asp Thr Glu Val Leu Pro Arg Pro 85
90 95Ile Leu Asn Ile Ser Arg Ile Gly Asp Val Pro
Val Glu Pro Gln Thr 100 105
110Trp Gly Ile Ala Leu Lys Cys Ser Gly Thr Arg Asn Ala Glu Val Thr
115 120 125Val Thr Ile Asn Val Glu Val
Ile Leu Asp Arg Ala Thr Asn Asn Asn 130 135
140Thr Asn Leu Ile Phe Lys Arg Lys Lys Ile Cys Leu Arg Glu Glu
Gln145 150 155 160Asp Ser
Ala His Glu Glu Tyr Asp Asp Asp Asp Leu Asp Leu Leu Gln
165 170 175Thr Ala Arg Lys Gly His Gly
Gly Asp Ile His Tyr Val Asp Arg Asn 180 185
190Asp Glu His Val Val Ala Asn Gly His Gln Ala Pro Glu Lys
Gln Arg 195 200 205Pro Val Val Thr
Glu Ser Pro Val Gly Arg Gly Asn Ser Gly Gly Ser 210
215 220Lys Arg Asp Phe Asp Pro Met Leu Arg Glu Asn Leu
Val Pro Pro Ala225 230 235
240Ser Gly Leu Val Thr Leu Ile Val Gly Gly Ile Leu Ala Leu Val Leu
245 250 255Val Ser Thr Leu Ile
Leu Ile Ala Tyr Cys Ala Lys Gly Pro Ser Lys 260
265 270Arg His Pro Ser Asn Gly Val His Leu Ile Lys Thr
Ser Ser Phe Gln 275 280 285Arg Leu
Pro Thr Ile Ser Ser Thr Ala His Asn Ser Ile Tyr Val Cys 290
295 300Pro Ser Thr Ile Thr Pro Thr Tyr Ala Thr Leu
Thr Arg Pro Phe Arg305 310 315
320Glu Tyr Glu His Glu Pro Glu Glu Phe Asn Arg Arg Leu Gln Glu Leu
325 330 335Thr Val Gln Lys
Cys Arg Val Arg Leu Ser Cys Leu Val Gln Glu Gly 340
345 350Asn Phe Gly Arg Ile Tyr Arg Gly Thr Tyr Asn
Asp Cys Gln Glu Val 355 360 365Leu
Val Lys Thr Val Ala Gln His Ala Ser Gln Leu Gln Val Asn Leu 370
375 380Leu Leu Gln Glu Ser Met Met Leu Tyr Glu
Ala Ser His Pro Asn Val385 390 395
400Leu Ser Val Leu Gly Ile Ser Ile Glu Asp Tyr Ala Thr Pro Phe
Val 405 410 415Leu Tyr Ala
Ala Thr Gly Ser Val Arg Asn Leu Lys Ser Phe Leu Gln 420
425 430Asp Pro Ser Tyr Ala Arg Ser Val Thr Thr
Ile Gln Thr Val Leu Met 435 440
445Gly Ser Gln Leu Ala Met Ala Met Glu His Leu His Asn His Gly Val 450
455 460Ile His Lys Asp Ile Ala Ala Arg
Asn Cys Val Ile Asp Asp Gln Leu465 470
475 480Arg Val Lys Leu Thr Asp Ser Ala Leu Ser Arg Asp
Leu Phe Pro Gly 485 490
495Asp Tyr Asn Ser Leu Gly Asp Gly Glu Tyr Arg Pro Ile Lys Trp Leu
500 505 510Ser Leu Glu Ala Leu Gln
Lys Ser His Tyr Asn Glu Gly Ser Asp Val 515 520
525Trp Ser Phe Gly Val Leu Met Trp Glu Met Cys Thr Leu Gly
Lys Leu 530 535 540Pro Tyr Ala Glu Ile
Asp Pro Tyr Glu Met Glu His Tyr Leu Lys Asp545 550
555 560Gly Tyr Arg Leu Ala Gln Pro Phe Asn Cys
Pro Asp Glu Leu Phe Thr 565 570
575Ile Met Ala Tyr Cys Trp Ala Ser Met Pro Ala Glu Arg Pro Ser Phe
580 585 590Ser Gln Leu Gln Ile
Cys Leu Ser Glu Phe His Thr Gln Ile Thr Arg 595
600 605Tyr Val 610
User Contributions:
comments("1"); ?> comment_form("1"); ?>Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
User Contributions:
Comment about this patent or add new information about this topic:
| People who visited this patent also read: | |
| Patent application number | Title |
|---|---|
| 20100206890 | BEVERAGE CONTAINER WITH COOLING MEANS |
| 20100206889 | SELF-HEATABLE CONTAINER |
| 20100206884 | GAS BURNER BOILING POT |
| 20100206883 | Fluid Container |
| 20100206882 | MULTI CHAMBER COOLANT TANK |