Patent application title: Hybrid immunoglobulins with moving parts
Inventors:
Daniel J. Capon (Hillsborough, CA, US)
Daniel J. Capon (Hillsborough, CA, US)
IPC8 Class: AC12P2104FI
USPC Class:
435 696
Class name: Micro-organism, tissue cell culture or enzyme using process to synthesize a desired chemical compound or composition recombinant dna technique included in method of making a protein or polypeptide blood proteins
Publication date: 2008-10-16
Patent application number: 20080254512
Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
Patent application title: Hybrid immunoglobulins with moving parts
Inventors:
Daniel J. Capon
Agents:
COOPER & DUNHAM, LLP
Assignees:
Origin: NEW YORK, NY US
IPC8 Class: AC12P2104FI
USPC Class:
435 696
Abstract:
Hybrid immunoglobulins containing moving parts are provided as well as
related compositions and methods of use and methods of production. In
addition, analogous genetic devices are provided as well as related
compositions and methods of use and methods of production.Claims:
1. A compound comprising a first stretch of consecutive amino acids, each
of which is joined to the preceding amino acid by a peptide bond and the
sequence of which comprises a binding site for a target; anda second
stretch of consecutive amino acids, each of which is joined to the
preceding amino acid by a peptide bond and the sequence of which is
identical to the sequence of the first stretch of consecutive amino acids
and which comprises an identical binding site for the target;wherein each
of the first stretch of amino acids and the second stretch of amino acids
has at a predefined end thereof a cysteine residue or a selenocysteine
residue and such cysteine residues or such selenocysteine residues are
joined by a bond having the structure:wherein each X is the same and
represents a sulfur (S) or a selenium (Se) and each C represents a
beta-carbon of one of such cysteine or selenocysteine residues.
2-7. (canceled)
8. A compound comprising a first stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which comprises a binding site for a target; anda second stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which is different from the sequence of the first stretch of consecutive amino acids and which comprises a binding site for a different moiety;wherein each of the first stretch of amino acids and the second stretch of amino acids has at a predefined end thereof a cysteine residue or a selenocysteine residue and such residues are joined by a bond having the structure:wherein each X may be the same or different and represents a sulfur (S) or a selenium (Se) and each C represents a beta-carbon of one of such cysteine or selenocysteine residues.
9-17. (canceled)
18. A multimer comprising two or more identical compounds according to any one of claims 1 or 8 joined together by at least one bond.
19-50. (canceled)
51. A method of affecting the activity of a target comprising contacting the target with a composition comprising the compound of claim 1 under conditions such that the compound binds to and affects the activity of the target.
52-62. (canceled)
63. A process of making the compound of claim 1, comprising:(a) transfecting a cell with a recombinant nucleic acid which comprises (i) a first portion, the sequence of which is a N-terminal signal sequence, contiguous with (ii) a second portion, the sequence of which encodes a stretch of consecutive amino acids contiguous with (iii) a third portion, the sequence of which encodes a C-terminal intein-containing binding domain, under conditions permitting synthesis of a chimeric polypeptide comprising the stretch of consecutive amino acids contiguous with the C-terminal intein-containing binding domain;(b) isolating the chimeric polypeptide produced in step (a);(c) treating the chimeric polypeptide so as to cause thio-mediated cleavage of the C-terminal intein-containing binding domain from the stretch of consecutive amino acids and its replacement with a C-terminal thioester;(d) treating the product of step (c) to permit the attachment of a cysteine residue to the product so as to form product with a C-terminal cysteine; and(e) oxidizing the product of step (e) in the presence of another product of step (e) under conditions permitting formation of the compound.
64-69. (canceled)
70. A compound comprising an independently folding protein domain fused to a second independently folding protein domain by non-peptide bond.
71-74. (canceled)
75. A method of making a stretch of consecutive amino acids comprising an N-terminal cysteine comprising:(a) transfecting a cell with a recombinant nucleic acid which comprises (i) a first portion, the sequence of which encodes a N-terminal signal sequence contiguous with (ii) a second portion, the sequence of which encodes a stretch of consecutive amino acids comprising a N-terminal cysteine residue, under conditions permitting (i) synthesis of a chimeric polypeptide which comprises the N-terminal signal sequence joined by a peptide bond at its C-terminus to the N-terminal cysteine of the stretch of consecutive amino acids and (ii) cleavage of the N-terminal signal sequence from the chimeric polypeptide within the cell so as to produce a stretch of consecutive amino acids comprising an N-terminal cysteine;(b) recovering the stretch of consecutive amino acids produced in step (a).
76-154. (canceled)
155. A process for making a compound comprising contacting a stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which comprises a binding site for a target with a second stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which is identical to the sequence of the first stretch of consecutive amino acids and which comprises an identical binding site for the target, wherein each of the first stretch of amino acids and the second stretch of amino acids has at a predefined end thereof a cysteine residue or a selenocysteine residue, under reducing conditions so as to make the compound.
156. A process for making a compound comprising contacting a stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which comprises a binding site for a target with a second stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which is different to the sequence of the first stretch of consecutive amino acids and which comprises an identical binding site for the target, wherein each of the first stretch of amino acids and the second stretch of amino acids has at a predefined end thereof a cysteine residue or a selenocysteine residue, under reducing conditions so as to make the compound.
157-164. (canceled)
165. A compound comprising a first stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which comprises a binding site for a target; anda second stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which is identical to the sequence of the first stretch of consecutive amino acids and which comprises an identical binding site for the target;wherein each of the first stretch of amino acids and the second stretch of amino acids has at a predefined end thereof, independently, a natural amino acid or non-natural amino having a linear aliphatic side-chain acid comprising a sulfur (S) or a selenium (Se) and wherein such sulfur (S) or a selenium (Se) are joined by a bond having the structure:wherein each X is a sulfur (S) or a selenium (Se) and each (C) represents a carbon of the linear aliphatic side-chain of one of such natural or non-natural amino acid and wherein n and m are, independently, 2, 3, 4, 5, 6, 7, 8, 9, or 10.
166-170. (canceled)
171. A compound comprising a first stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which comprises a binding site for a target; anda second stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which is different from the sequence of the first stretch of consecutive amino acids and which comprises a binding site for a different moiety;wherein each of the first stretch of amino acids and the second stretch of amino acids has at a predefined end thereof, independently, a natural amino acid or non-natural amino having a linear aliphatic side-chain acid comprising a sulfur (S) or a selenium (Se) and wherein such sulfur (S) or a selenium (Se) are joined by a bond having the structure:wherein each X may be the same or different and represents a sulfur (S) or a selenium (Se) and each (C) represents a carbon of the linear aliphatic side-chain of one of such natural or non-natural amino acid and wherein n and m are, independently, 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10.
172-176. (canceled)
177. A method of producing a protein which comprises a first polypeptide contiguous with an intein, which intein is contiguous with a second polypeptide comprising a binding domain, the method comprising transfecting an animal cell with a nucleic acid, which nucleic acid comprises (i) a first portion which encodes the polypeptide contiguous with (ii) a second portion which encodes the intein, contiguous with a and the third portion of which encodes the binding domain, under conditions such that the animal cell expresses and secretes the protein.
178-181. (canceled)
182. A compound comprising:a first stretch of consecutive amino acids each of which is joined to the preceding amino acid by a peptide bond and which first stretch of consecutive amino acids comprises an amino acid residue having a chalcogen functional group-containing side chain; anda second stretch of consecutive amino acids, comprising at least 100 amino acids, each of which is joined to the preceding amino acid by a peptide bond, wherein at least 90 consecutive amino acids thereof of the second stretch of consecutive amino acids have a sequence identical to portion of a human immunoglobulin constant region polypeptide, and wherein the second stretch of consecutive amino acids comprises an amino acid residue having a chalcogen functional group-containing side chain at a predefined terminus thereof,wherein said amino acid residue having a chalcogen functional group-containing side chain of the first stretch of consecutive amino acids and said amino acid residue having a chalcogen functional group-containing side chain of the second stretch of consecutive amino acids are joined by a bond having the structure:wherein each X represents, independently, a chalcogen, and wherein C1 represents a side chain carbon of the amino acid residue having a chalcogen functional group-containing side chain of the first stretch of consecutive amino acids and C2 represents a side chain carbon of the second stretch of consecutive amino acids.
183-194. (canceled)
195. The compound of claim 182, wherein the human immunoglobulin constant region polypeptide is a human IgG1, human IgG2, human IgG3, or human IgG4.
196. The compound of claim 195, wherein the side chain of at least one of amino acid residues having a chalcogen functional group-containing side chain comprises a C1-C10 alkylene.
197. A composition comprising two of the compounds of claim 182 bonded together via at least one disulfide bond between the second stretch of consecutive amino acids of each of the compounds.
198. A polypeptide consisting of consecutive amino acids having the sequence set forth in one of SEQ ID NOS:35 through 46, or having the sequence set forth in one of SEQ ID NOS:53 through 67, or having the sequence set forth in one of SEQ ID NOS:74 through 82, or having the sequence set forth in one of SEQ ID NOS:89 through 97.
199-201. (canceled)
202. A polypeptide consisting of consecutive amino acids having a sequence identical to a portion of the sequence set forth in SEQ ID NO:44, SEQ ID NO:64, SEQ ID NO:81 or SEQ ID NO:96, wherein at least one of the terminal residues of the polypeptide has a chalcogen functional group-containing side chain.
203. The polypeptide of claim 202, wherein the terminal residue having a chalcogen functional group-containing side chain is a cysteine or analog thereof
Description:
[0001]This application claims benefit of U.S. Provisional Application No.
60/856,864, filed Nov. 2, 2006, the contents of which are hereby
incorporated by reference into this application.
[0002]Throughout this application, various publications are referenced. The disclosures of these publications in their entireties are hereby incorporated by reference into this application in order to more fully describe the state of the art as known to those skilled therein as of the date of the invention described and claimed herein.
BACKGROUND OF THE INVENTION
[0003]All machines and devices have moving parts. The function of the moving parts is to perform work, by transforming a source of energy, in order to carry out a useful task. Moving parts cover a spectrum of sizes and shapes. At one end of the spectrum is a visible world evident in machines that perform mechanical tasks. At the other end of the spectrum is an invisible world of charge carriers utilized by devices that carry out electrical work.
[0004]This spectrum is so vast that certain of its regions have only begun to be technologically exploited. Among these are devices with moving parts of several nano-meters to several hundred nano-meters. This size range holds considerable interest to many scientists and engineers because it is comparable to the very size of molecules, the fundamental units of chemical matter. Nano-machines have the potential to exploit the unique properties of molecules, such as intermolecular binding or catalysis.
[0005]The ability to make molecules of any imaginable size and shape is one activity crucial in building nano-machines. As such it has been widely anticipated in medicine, electronics, optics, and many other fields. Tremendous commercial activity has been focused on the synthesis very large numbers of chemically distinct molecules. However, molecular configuration (differences in bonding) is just one practical means of generating diversity. Molecular conformation (differences in bond rotation) offers another important avenue for generating a universe of continuous size and shape.
[0006]Molecular conformation has certain unique advantages in strategies for creating molecules with moving parts. While atoms and chemical bonds have precise linear and angular dimensions, conformational change can provide limitless variation in the size and shape of molecules. Covalent and non-covalent chemical bonds both afford rotational degrees of freedom. Dihedral rotation around each of a series of bonds connecting distinct parts (domains) of a molecule is capable of providing the essential dynamic ingredient of nano-machines.
[0007]In general, any two given atoms interconnected by a single bond (i.e., a single electron pair) can rotate fully 360 degrees with respect to each other and with respect to the other atoms that each is bonded to. A series of consecutive single bonds is like a series of interconnected ball joints. Although limited to rotary motions, a series of consecutive single bonds, like a series of consecutive ball joints, can recapitulate the movement of other types of interconnected moving parts (e.g., a series of consecutive hinges).
[0008]One challenging aspect of creating useful nano-machines is striking a balance in the number of moving parts and the number of interconnections. Above a certain threshold, increasing the number of parts or connections in any machine is counterproductive. Thus automobile engines employ an optimal number of pistons, valves, camshafts, pulleys, and so forth.
[0009]The analogous challenge in the chemical field is illustrated by two related, but very different types of molecules, namely organic and biological polymers. A good comparison is provided by polyethylene and proteins. Polyethylenes are stretches of consecutive ethylenes, (CH2)n, interconnected by consecutive single bonds (--C--)n, while proteins are stretches of consecutive amino acids, (NHCHRCO)n, interconnected by consecutive peptide bonds (═N--C--C═)n. Unbranched polyethylenes are repeating chains of single bonds, while proteins are repeating chains of one double bond followed by two single bonds. The most important difference between these two types of chains is that polyethylene can adopt almost any conformation and thus has no definite size or shape (only a statistically averaged one), while proteins are extremely rigid and thus have very definite (and unchanging) size and shape.
[0010]A simple but reasonable comparison to a mechanical device would represent polyethylene as a machine with a high ratio of moving parts to connections, and a protein as a machine with an low ratio of moving parts to connections. Neither molecule is very suited to a machine-like task unless one takes advantage of higher order structures that each can form. For example, polyethylene is useful when its ability to form intermolecular fibers is exploited. Interesting, the ability of polyethylene to display such tertiary structure depends upon its inherent flexibility. Although some proteins can form fibers of commercial value (e.g., silk, wool and collagen), most proteins are globular and do not.
[0011]Globular proteins are, for nearly all practical purposes, machines with few if any moving parts, like a crowbar that must exert its leverage in combination with other objects, such as the human that wields it and the objects against which it is wedged. Notwithstanding, there are many instances in which there would be great value to protein-like molecules having distinct regions (e.g., binding domains) that are joined together in some manner permitting relative, yet coordinated movement. One example would be protein-like molecules capable of cooperatively binding a disease target having two or more identical binding sites. This would take full advantage of the unique properties of globular protein binding domains, namely their great specificity for targets, particularly other proteins associated with disease.
[0012]The potential commercial value of protein-like molecules that are able to cooperatively bind a disease target may be estimated quantitatively. A starting assumption is that most therapeutics in current use, whether small molecules or biopharmaceuticals, typically bind their targets non-cooperatively with affinity constants on the order of nano-molar (10-9 M). Remarkably, a cooperative therapeutic could conceivably bind the same target, with an affinity of nano-molar×nano-molar (10-9 M×10-9 M) [i.e., atto-molar (10-18 M)].
[0013]Because therapeutics are typically required in great molar excess over their targets (about one million-fold), a cooperative therapeutic would thus be equivalent to a non-cooperative therapeutic at a 10-6 smaller dose. For many current biopharmaceuticals (e.g. antibodies and immunoadhesins) this difference amounts to 1 microgram per single dose instead of 1 gram per single dose. With patient costs exceeding $1,000 per gram, this factor has great significance in new drug discovery and development as well as for existing biopharmaceuticals.
[0014]One irony associated with antibodies and immunoadhesins is that while they are symmetric proteins having two identical binding domains, they do not generally bind symmetrically to symmetric targets. The inflexible connections between the two binding domains do not provide the machine-like motion that would permit cooperative binding. Numerous attempts to engineer antibodies and immunoadhesins that bind symmetrically have failed because of the difficulty in achieving the precise geometry needed for complementary symmetries between the binding sites and target sites. Unlike materials used to make conventional machines, such as wood, metals, plastics, ceramics, and the like, molecules cannot simply be cut, wrought, cast, machined or joined to an exact size and shape.
[0015]While cooperative binding is thus not readily achieved with any single fixed size and shape, conformational flexibility between binding domains does provides a potential solution. A "one size fits all" strategy is based upon the proposition that a protein-like molecule with binding domains that move symmetrically will also be capable of binding symmetrically (i.e., cooperativity) The binding domains are driven thermodynamically into a conformation most compatible with simultaneous binding of both target sites because it represents the energetically favored conformational minima.
SUMMARY OF THE INVENTION
[0016]In an embodiment this invention provides a compound comprising a first stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which comprises a binding site for a target; and a second stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which is identical to the sequence of the first stretch of consecutive amino acids and which comprises an identical binding site for the target; wherein each of the first stretch of amino acids and the second stretch of amino acids has at a predefined end thereof a cysteine residue or a selenocysteine residue and such cysteine residues or such selenocysteine residues are joined by a bond having the structure:
wherein each X is the same and represents a sulfur (S) or a selenium (Se) and each C represents a beta-carbon of one of such cysteine or selenocysteine residues.
[0017]In an embodiment this invention also provides a compound comprising a first stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which comprises a binding site for a target; and a second stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which is different from the sequence of the first stretch of consecutive amino acids and which comprises a binding site for a different moiety; wherein each of the first stretch of amino acids and the second stretch of amino acids has at a predefined end thereof a cysteine residue or a selenocysteine residue and such residues are joined by a bond having the structure:
wherein each X may be the same or different and represents a sulfur (S) or a selenium (Se) and each C represents a beta-carbon of one of such cysteine or selenocysteine residues.
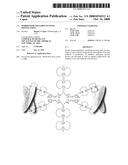
[0018]Genetic devices disclosed herein comprise two or more stretches of consecutive amino acids that are connected at a predefined terminus by a non-peptide bond. Such genetic devices are both symmetric and symmetrically binding with respect to one or more important targets (i.e., cooperative). The genetic devices herein are protein-like molecules may be described by a number of related terms that include symmetroadhesins, immuno-symmetroadhesins, hemi-symmetroadhesins, and bi-symmetroadhesins [meaning "stick to proportionately," from the Gk. symmetros "having a common measure, even, proportionate," and the L. adhaerentem, prp. of adhaerere "stick to"].
[0019]Disclosed herein is a compound comprising two or more independently-folding protein domains linked to one another through one or more non-peptide bonds, around which bond(s) dihedral rotation may occur.
BRIEF DESCRIPTION OF THE FIGURES
[0020]FIG. 1: Stretches of consecutive amino acids, with one X-terminus, depicting positions of N-, C-, S-, and Se-termini. N-terminal and C-terminal amino acid residues, drawn as a Newman-style projection, are shown above and below the projection plane, respectively: (i) A stretch of consecutive amino acids (generalized structure), having, an N-terminal amino acid residue (sidechain=R1) having a free α-amino (NH2) group, and a C-terminal amino acid residue (sidechain=Rn) having a free α-carboxyl (COOH) group. (ii) A stretch of consecutive amino acids, with N-terminal S-terminus, having, an N-terminal cysteine having free α-amino (NH2) and β-sulfhydryl (SH) groups, and a C-terminal amino acid residue having a free α-carboxyl (COOH) group. (iii) A stretch of consecutive amino acids, with C-terminal S-terminus, having, an N-terminal amino acid residue having a free α-amino (NH2) group, and a C-terminal cysteine having free α-carboxyl (COOH) and β-sulfhydryl (SH) groups. (iv) A stretch of consecutive amino acids, with N-terminal Se-terminus, having, an N-terminal selenocysteine having free α-amino (NH2) and β-selenohydryl (SeH) groups, and a C-terminal amino acid residue having a free α-carboxyl (COOH) group. (v) A stretch of consecutive amino acids, with C-terminal Se-terminus, having, a N-terminal amino acid residue having a free α-amino (NH2) group, and a C-terminal selenocysteine having free α-carboxyl (COOH) and β-selenohydryl (SeH) groups.
[0021]FIG. 2: Stretches of consecutive amino acids, with two X-termini, depicting positions of N-, C-, S-, and Se-termini. N-terminal and C-terminal amino acid residues, drawn as a Newman-style projection, are shown above and below the projection plane, respectively. (i) A stretch of consecutive amino acids, with two X-termini (generalized structure), having, an N-terminal amino acid residue (sidechain=X1) having a free α-amino (NH2) group, and a C-terminal amino acid residue (sidechain=Xn) having a free α-carboxyl (COOH) group. (ii) A stretch of consecutive amino acids, with N-terminal S-terminus and C-terminal S-terminus, having, an N-terminal cysteine having free α-amino (NH2) and β-sulfhydryl (SH) groups, and a C-terminal cysteine having free α-carboxyl (COOH) and β-sulfhydryl (SH) groups. (iii) A stretch of consecutive amino acids, with N-terminal S-terminus and C-terminal Se-terminus, having, an N-terminal cysteine having free α-amino (NH2) and β-sulfhydryl (SH) groups, and a C-terminal selenocysteine having free α-carboxyl (COOH) and β-selenohydryl (SeH) groups. (iv) A stretch of consecutive amino acids, with N-terminal Se-terminus and C-terminal S-terminus, having, an N-terminal selenocysteine having free α-amino (NH2) and β-selenohydryl (SeH) groups, and a C-terminal cysteine having free α-carboxyl (COOH) and β-sulfhydryl (SH) groups. (v) A stretch of consecutive amino acids, with N-terminal Se-terminus and C-terminal Se-terminus, having, an N-terminal selenocysteine having free α-amino (NH2) and β-selenohydryl (SeH) groups, and a C-terminal selenocysteine having free α-carboxyl (COOH) and β-selenohydryl (SeH) groups.
[0022]FIG. 3: General structure of a chimeric polypeptide consisting of a first stretch of consecutive amino acids joined at its C-terminus by a peptide bond to the N-terminus of a second stretch of consecutive amino acids. Chimeric polypeptides, like proteins found in nature, are continuous stretches of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond. Like other proteins, chimeric polypeptides have limited conformational flexibility because the peptide bond by itself provides no more than two consecutive single bonds capable of dihedral rotation along the polypeptide chain. Amino acid residues in the figure are numbered as follows: The first stretch of consecutive amino acids has length=n residues, and numbering=1, 2, 3, . . . , (n-2), (n-1), n. The second stretch of consecutive amino acids has length=p residues, and numbering=1', 2', 3', . . . , (p-a2), (p-a1), p. The chimeric polypeptide has length=(n+p) residues, and numbering=1, 2, 3, . . . , (n-2), (n-1), n, (n+1), (n+2), (n+3), (n+p-2), (n+p-1), (n+p). The major and minor tautomeric forms and the resonance structure are shown on the left, center and right, respectively.
[0023]FIG. 4A: General structure of a "symmetroadhesin" with a head-to-tail configuration consisting of a first stretch of consecutive amino acids joined at its C-terminal-X-terminus by an -X-X- bond to the N-terminal-X-terminus of a second stretch of consecutive amino acids. The -X-X- bond is not a peptide bond. Non-limiting examples of the bonds envisaged here include any combination wherein each X is a S or a Se atom. The overall polarity of head-to-tail symmetroadhesins is N- to C-terminal. Symmetroadhesins, like proteins, are stretches of consecutive amino acids each of which is joined to the preceding amino acid, but differ from proteins by substituting one or more -X-X- bonds for peptide bonds. Symmetroadhesins have greater conformational flexibility than polypeptides because each -X-X- bond provides seven adjacent single bonds capable of dihedral rotation. Amino acid residues are numbered as follows: The first stretch of consecutive amino acids has length=n residues, and numbering=1, 2, 3, . . . , (n-2), (n-1), n. The second stretch of consecutive amino acids has length=p residues, and numbering=1', 2', 3', (p-a2), (p-a1), p. The head-to-tail symmetroadhesin has length=(n+p) residues, and numbering=1, 2, 3, . . . , (n-2), (n-1), n, (n+1), (n+2), (n+3), . . . , (n+p-2), (n+p-1), (n+p). The major and minor tautomeric forms and the resonance structure are shown on the left, center and right, respectively.
[0024]FIG. 4B: The symmetroadhesin of FIG. 4A (left) compared with a symmetroadhesin (right) consisting of a first stretch of consecutive amino acids joined at the X-terminus of its penultimate C-terminal residue by an -X-X- bond to the X-terminus of the penultimate N-terminal residue of a second stretch of consecutive amino acids. Resonance structures are shown for each.
[0025]FIG. 4C: The symmetroadhesin of FIG. 4A (left) compared with a symmetroadhesin (right) consisting of a first stretch of consecutive amino acids joined at the X-terminus of its antepenultimate C-terminal residue by an -X-X- bond to the X-terminus of the antepenultimate N-terminal residue of a second stretch of consecutive amino acids. Resonance structures are shown for each.
[0026]FIG. 4D: The symmetroadhesin of FIG. 4A (left) compared with a symmetroadhesin (right) consisting of a first stretch of consecutive amino acids joined at the X-terminus of its preantepenultimate C-terminal residue by an -X-X- bond to the X-terminus of the preantepenultimate N-terminal residue of a second stretch of consecutive amino acids. Resonance structures are shown for each.
[0027]FIG. 5: General structure of a symmetroadhesin with a head-to-head configuration consisting of a first stretch of consecutive amino acids, joined at its N-terminal-X-terminus by an -X-X- bond to the N-terminal-X-terminus of a second stretch of consecutive amino acids. The overall polarity of head-to-head symmetroadhesins changes at the position of the -X-X- bond, going from C- to N-terminal to N- to C-terminal. Amino acid residues are numbered as follows: The first stretch of consecutive amino acids has length=n residues, and numbering=1, 2, 3, . . . , (n-2), (n-1), n. The second stretch of consecutive amino acids has length=p residues, and numbering=1', 2', 3', (p-a2), (p-a1), p. The head-to-head symmetroadhesin has length=(n+p) residues, and numbering=n, (n-1), (n-2), . . . , 3, 2, 1, (inversion), 1', 2', 3', . . . , (p-a2), (p-a1), p. The major and minor tautomeric forms and the resonance structure are shown on the left, center and right, respectively.
[0028]FIG. 6: General structure of a symmetroadhesin with a tail-to-tail configuration consisting of a first stretch of consecutive amino acids joined at its C-terminal-X-terminus by an -X-X- bond to the C-terminal-X-terminus of a second stretch of consecutive amino acids. The overall polarity of tail-to-tail symmetroadhesins changes at the position of the -X-X- bond, going from N- to C-terminal to C- to N-terminal. Amino acid residues are numbered as follows: The first stretch of consecutive amino acids has length=n residues, and numbering=1, 2, 3, . . . , (n-2), (n-1), n. The second stretch of consecutive amino acids has length=p residues, and numbering=1', 2', 3', (p-a2), (p-a1), p. The tail-to-tail symmetroadhesin has length=n+p residues, and numbering=1, 2, 3, . . . , (n-2), (n-1), n, (inversion), p, (p-a1), (p-a2), . . . , 3', 2', 1'. The major and minor tautomeric forms and the resonance structure are shown on the left, center and right, respectively.
[0029]FIG. 7: A schematic representation of a head-to-tail hemi-symmetroadhesin showing the all-trans conformation. All of the seven consecutive single bonds joining the C-terminal-X-terminus with the N-terminal-X-terminus are trans (N--C--C--X--X--C--C--C). The two binding domains are pointed away from one another in this conformation; a rotation of 180 degrees around any one of the seven consecutive single bonds will point the two binding domains towards one another. Hemi-symmetroadhesins with the head-to-tail configuration are asymmetric molecules regardless of conformation (compare FIGS. 7 and 8); however, two or more head-to-tail hemi-symmetroadhesins can together form a symmetric molecule.
[0030]FIG. 8: A schematic representation of a head-to-tail hemi-symmetroadhesin showing the X-cis-X conformation. All but one of seven consecutive single bonds joining the C-terminal-X-terminus with the N-terminal-X-terminus are trans (N--C--C--X-cis-X--C--C--C). The two binding domains are pointed towards one another in this conformation; further rotations of 180 degrees around any one of the seven consecutive single bonds will point the two binding domains away from one another. Hemi-symmetroadhesins with the head-to-tail configuration are asymmetric molecules regardless of conformation (compare FIGS. 7 and 8); however, two or more head-to-tail hemi-symmetroadhesins can together form a symmetric molecule.
[0031]FIG. 9: A schematic representation of a head-to-head hemi-symmetroadhesin showing the all-trans conformation. All of the seven consecutive single bonds joining the 1st N-terminal-X-terminus and 2nd N-terminal-X-terminus are trans (C--C--C--X--X--C--C--C). The two binding domains are pointed away from one another in this conformation; a rotation of 180 degrees around any one of the seven consecutive single bonds will point the two binding domains towards one another. Hemi-symmetroadhesins with the head-to-head configuration are symmetric molecules in only two of their possible conformations: the all-trans and the X-cis-X (compare FIGS. 9 and 10); however, two or more head-to-head hemi-symmetroadhesins subunits can form a molecule that has an unlimited number of symmetric conformations.
[0032]FIG. 10: A schematic representation of a head-to-head hemi-symmetroadhesin showing the X-cis-X conformation. All but one of the seven consecutive single bonds joining the 1st N-terminal-X-terminus and 2nd N-terminal-X-terminus are trans (C--C--C--X-cis-X--C--C--C). The two binding domains are pointed towards one another in this conformation; further rotations of 180 degrees around any one of the seven consecutive single bonds will point the two binding domains away from one another. Hemi-symmetroadhesins with the head-to-head configuration are symmetric molecules in only two of their possible conformations: the all-trans and the X-cis-X (compare FIGS. 9 and 10); however, two or more head-to-head hemi-symmetroadhesins subunits can form a molecule that has an unlimited number of symmetric conformations.
[0033]FIG. 11: A schematic representation of a tail-to-tail hemi-symmetroadhesin showing the all-trans conformation. All of the seven consecutive single bonds joining the 1st C-terminal-X-terminus and 2nd C-terminal-X-terminus are trans (N--C--C--X--X--C--C--N). The two binding domains are pointed away from one another in this conformation; a rotation of 180 degrees around any one of the seven consecutive single bonds will point the two binding domains towards one another. Hemi-symmetroadhesins with the tail-to-tail configuration are symmetric molecules in only two of their possible conformations: the all-trans and the X-cis-X (compare FIGS. 11 and 12); however, two or more tail-to-tail hemi-symmetroadhesins subunits can form a molecule that has an unlimited number of symmetric conformations.
[0034]FIG. 12: A schematic representation of a tail-to-tail hemi-symmetroadhesin showing the X-cis-X conformation. All but one of the seven consecutive single bonds joining the 1st C-terminal-X-terminus and 2nd C-terminal-X-terminus are trans (N--C--C--X-cis-X--C--C--N). The two binding domains are pointed towards one another in this conformation; further rotations of 180 degrees around any one of the seven consecutive single bonds will point the two binding domains away from one another. Hemi-symmetroadhesins with the tail-to-tail configuration are symmetric molecules in only two of their possible conformations: the all-trans and the X-cis-X (compare FIGS. 9 and 10); however, two or more tail-to-tail hemi-symmetroadhesins subunits can form a molecule that has an unlimited number of symmetric conformations.
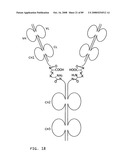
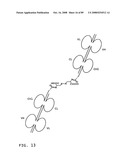
[0035]FIG. 13: A schematic representation of a tail-to-tail hemi-symmetroadhesin consisting of two immunoglobulin Fab binding domains. The all-trans conformation is shown here. All of the seven consecutive single bonds joining the 1st C-terminal-X-terminus and 2nd C-terminal-X-terminus are trans (N--C--C--X--X--C--C--N). The two Fab binding domains are pointed away from one another in this conformation; a rotation of 180 degrees around any one of the seven consecutive single bonds will point the Fab binding domains towards one another (compare FIGS. 13 and 14). The heavy chain regions are joined together by the X-X bond; the light chain regions are joined to the heavy chain regions by internal disulfide bonds. Abbreviations: VL, light chain variable region; CL, light chain constant region; VH, heavy chain variable region; CH1, heavy chain constant region 1.
[0036]FIG. 14: A schematic representation of a tail-to-tail hemi-symmetroadhesin consisting of two immunoglobulin Fab binding domains. The X-cis-X conformation is shown here. All but one of seven consecutive single bonds joining the 1st C-terminal-X-terminus and 2nd C-terminal-X-terminus are trans (N--C--C--X-cis-X--C--C--N). The two Fab binding domains are pointed towards one another in this conformation; further rotation of 180 degrees around any one of the seven consecutive single bonds will point the Fab binding domains away from one another (compare FIGS. 13 and 14). The heavy chain regions are joined together by the X-X bond; the light chain regions are joined to the heavy chain regions by internal disulfide bonds. Abbreviations: VL, light chain variable region; CL, light chain constant region; VH, heavy chain variable region; CH1, heavy chain constant region 1.
[0037]FIG. 15: Schematic representation of the immunoadhesin molecule (Capon et al. (1989) Nature 337, 525-530). Immunoadhesins are chimeric polypeptides that form disulfide-linked dimers. Each chimeric polypeptide consists of a binding domain joined at its C-terminus by a peptide bond to the N-terminus of an immunoglobulin Fc domain. Although immunoadhesins are structurally symmetric, they do not generally bind cooperatively to dimeric or multimeric target molecules. Abbreviations: CH2, heavy chain constant region 2; CH3, heavy chain constant region 3.
[0038]FIG. 16: Schematic representation of the immunoglobulin (antibody) molecule. Immunoglobulins are heterotetramers consisting of two heavy chains and two light chains. Although immunoglobulins are structurally symmetric, they do not generally bind cooperatively to dimeric or multimeric target molecules. Abbreviations: VL, light chain variable region; CL, light chain constant region; VH, heavy chain variable region; CH1, heavy chain constant region 1; CH2, heavy chain constant region 2; CH3, heavy chain constant region 3.
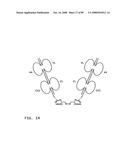
[0039]FIG. 17: A schematic representation of a head-to-tail immunosymmetroadhesin showing the all-trans conformation. Head-to-tail immunosymmetroadhesins are head-to-tail hemi-symmetroadhesins that form disulfide-linked dimers. Each hemi-symmetroadhesin consists of an immunoglobulin Fab binding domain having a C-terminal-X-terminus joined by an -X-X- bond to a immunoglobulin Fc subunit having an N-terminal-X-terminus. The dimer contains two functional Fab binding domains and one functional Fc binding domain. The seven consecutive single bonds which join each Fab domain to an Fc subunit are all trans (N--C--C--X--X--C--C--C). Symmetric rotations of 180 degrees around the first (N--C), third (C--X), fifth (X--C), or seventh (C--C) pairs of consecutive single bonds will move the two Fab domains in a first general direction (compare FIGS. 17 and 18). Symmetric rotations of 180 degrees around the second (C--C), fourth (X--X), or sixth (C--C) single bond pairs will move the two Fab domains in a second general direction (compare FIGS. 17 and 19).
[0040]FIG. 18: A schematic representation of a head-to-tail immunosymmetroadhesin showing the X-cis-C conformation. The X-cis-C conformation is obtained from the all-trans conformation by the symmetric rotation of the fifth pair of seven consecutive single bonds (N--C--C--X--X-cis-C--C--C) (compare FIGS. 17 and 18). Other conformations that are similar to the X-cis-C conformation shown here are obtained from the all-trans conformation following the symmetric rotation of the first (N-cis-C--C--X--X--C--C--C), the third (N--C--C-cis-X--X--C--C--C), or the seventh pairs (N--C--C--X--X--C--C-cis-C) of consecutive single bonds.
[0041]FIG. 19: A schematic representation of a head-to-tail immunosymmetroadhesin showing the X-cis-X conformation. The X-cis-X conformation is obtained from the all-trans conformation by the symmetric rotation of the fourth pair of seven consecutive single bonds (N--C--C--X-cis-X--C--C--C) (compare FIGS. 17 and 19). Other conformations that are similar to the X-cis-X conformation shown here are obtained from the all-trans conformation following the symmetric rotation of the second (N--C-cis-C--X--X--C-C--C), or the sixth pairs (N--C-C--X--X--C-cis-C--C) of consecutive single bonds.
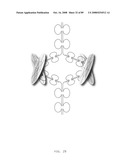
[0042]FIG. 20: A schematic representation of a tail-to-tail immunosymmetroadhesin showing the all-trans conformation. Tail-to-tail immunosymmetroadhesins are tail-to-tail hemi-symmetroadhesins that form disulfide-linked dimers. Each hemi-symmetroadhesin consists of an immunoglobulin Fab binding domain having a C-terminal-X-terminus joined by an X-X bond to an immunoglobulin Fc subunit having a C-terminal-X-terminus. The dimer contains two functional Fab binding domains and one functional Fc binding domain. The seven consecutive single bonds which join each Fab domain to an Fc subunit are all trans (N--C--C--X--X--C--C--N). Symmetric rotations of 180 degrees around the first (N--C), third (C--X), fifth (X--C), or seventh (C--N) pairs of consecutive single bonds will move the two Fab domains in a first general direction (compare FIGS. 20 and 21). Symmetric rotations of 180 degrees around the second (C--C), fourth (X--X), or sixth (C--C) single bond pairs will move the two Fab domains in a second general direction (compare FIGS. 20 and 22).
[0043]FIG. 21: A schematic representation of a tail-to-tail immunosymmetroadhesin showing the X-cis-C conformation. The X-cis-C conformation is obtained from the all-trans conformation by the symmetric rotation of the fifth pair of seven consecutive single bonds (N--C--C--X--X-cis-C--C--N) (compare FIGS. 20 and 21). Other conformations that are similar to the X-cis-C conformation shown here are obtained from the all-trans conformation following the symmetric rotation of the first (N-cis-C--C--X--X--C--C--N), the third (N--C--C-cis-X--X--C--C--N), or the seventh pairs (N--C--C--X--X--C--C-cis-N) of consecutive single bonds.
[0044]FIG. 22: A schematic representation of a tail-to-tail immunosymmetroadhesin showing the X-cis-X conformation. The X-cis-X conformation is obtained from the all-trans conformation by the symmetric rotation of the fourth pair of seven consecutive single bonds (N--C--C--X-cis-X--C--C--N) (compare FIGS. 20 and 22). Other conformations that are similar to the X-cis-X conformation shown here are obtained from the all-trans conformation following the symmetric rotation of the second (N--C-cis-C--X--X--C--C--N), or the sixth pairs (N--C--C--X--X--C-cis-C--N) of consecutive single bonds.
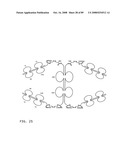
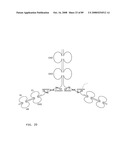
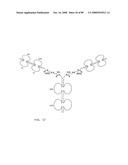
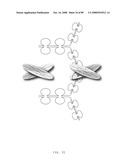
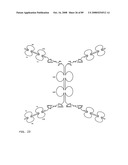
[0045]FIG. 23: Schematic representation of bi-symmetroadhesin with four Fab binding domains, showing the all-trans conformation. The molecule is a dimer of two hemi-symmetroadhesins each consisting of three stretches of consecutive amino acids. The bi-symmetroadhesin shown here is a head-to-tail, tail-to-tail hemi-symmetroadhesin that forms disulfide-linked dimers. Each hemi-symmetroadhesin consists of two immunoglobulin Fab domains having a C-terminal-X-terminus that is joined by an -X-X- bond to a immunoglobulin Fc subunit; the first Fab domain is joined to the Fc N-terminal-X-terminus, and the second Fab is joined to the Fc C-terminal-X-terminus. The dimer has four functional Fab binding domains, and one functional Fc binding domain. The seven consecutive single bonds which join all four Fab domains to an Fc subunit are all trans (N--C--C--X--X--C--C--C/N). Symmetric rotations of 180 degrees around the first (N--C), third (C--X), fifth (X--C), or seventh (C--C/N) pairs of consecutive single bonds will move the four Fab domains in a first general direction (compare FIGS. 23 and 24). Symmetric rotations of 180 degrees around the second (C--C), fourth (X--X), or sixth (C--C) single bond pairs will move the four Fab domains in a second general direction (compare FIGS. 23 and 25).
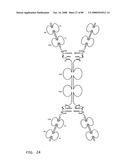
[0046]FIG. 24: Schematic representation of the X-cis-C conformation of a bi-immunosymmetroadhesin consisting of four Fab binding domains and one Fc domain. The molecule is a dimer of two hemi-symmetroadhesins each consisting of three stretches of consecutive amino acids.
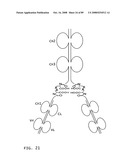
[0047]FIG. 25: Schematic representation of the X-cis-X conformation of a bi-immunosymmetroadhesin consisting of four Fab binding domains and one Fc domain. The molecule is a dimer of two hemi-symmetroadhesins each consisting of three stretches of consecutive amino acids.
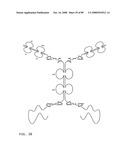
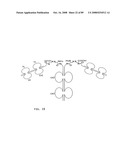
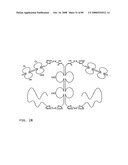
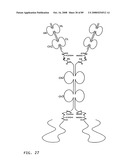
[0048]FIG. 26: Schematic representation of the all-trans conformation of a bi-immunosymmetroadhesin consisting of two Fab binding domains, one Fc domain, and two non-Fab binding domains. The molecule is a dimer of two hemi-symmetroadhesins each consisting of three stretches of consecutive amino acids. The bi-symmetroadhesin shown here is a head-to-tail, tail-to-tail hemi-symmetroadhesin that forms disulfide-linked dimers. Each hemi-symmetroadhesin consists of one immunoglobulin Fab domain having a C-terminal-X-terminus that is joined by an -X-X- bond to the N-terminal-X-terminus of an immunoglobulin Fc subunit, and one non-immunoglobulin binding domain having a C-terminal-X-terminus that is joined by an -X-X- bond to the C-terminal-X-terminus of an immunoglobulin Fc subunit. The dimer has two functional Fab binding domains, two functional non-immunoglobulin binding domains, and one functional Fc binding domain. The seven consecutive single bonds which join all four binding domain to an Fc subunit are all trans (N--C--C--X--X--C--C--C/N). Symmetric rotations of 180 degrees around the first (N--C), third (C--X), fifth (X--C), or seventh (C--C/N) pairs of consecutive single bonds will move the four binding domains in a first general direction (compare FIGS. 26 and 27). Symmetric rotations of 180 degrees around the second (C--C), fourth (X--X), or sixth (C--C) single bond pairs will move the four binding domains in a second general direction (compare FIGS. 26 and 28).
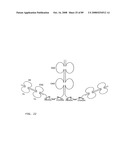
[0049]FIG. 27: Schematic representation of the X-cis-C conformation of a bi-immunosymmetroadhesin consisting of two Fab binding domains, one Fc domain, and two non-Fab binding domains. The molecule is a dimer of two hemi-symmetroadhesins each consisting of three stretches of consecutive amino acids.
[0050]FIG. 28: Schematic representation of the X-cis-X conformation of a bi-immunosymmetroadhesin consisting of two Fab binding domains, one Fc domain, and two non-Fab binding domains. The molecule is a dimer of two hemi-symmetroadhesins each consisting of three stretches of consecutive amino acids.
[0051]FIG. 29: Schematic representation of an immunoglobulin binding to a first symmetric target. The interaction is symmetric and cooperative. Both targets are bound by both immunoglobulins.
[0052]FIG. 30: Schematic representation of a symmetroadhesin binding to a first symmetric target. The interaction is symmetric and cooperative. Both targets are bound by both symmetroadhesins in a first conformation (all-trans).
[0053]FIG. 31: Schematic representation of an immunoglobulin binding to a second symmetric target. The interaction is neither symmetric and nor cooperative. Only one target is bound by each immunoglobulin.
[0054]FIG. 32: Schematic representation of a symmetroadhesin binding to a second symmetric target. The interaction is symmetric and cooperative. Both targets are bound by both symmetroadhesins in a second conformation (X-cis-C).
[0055]FIG. 33: Schematic representation of an immunoglobulin binding to a third symmetric target. The interaction is neither symmetric and nor cooperative. Only one target is bound by each immunoglobulin.
[0056]FIG. 34: Schematic representation of a symmetroadhesin binding to a third symmetric target. The interaction is symmetric and cooperative. Both targets are bound by both symmetroadhesins in a third conformation (X-cis-X).
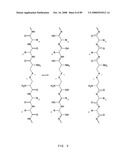
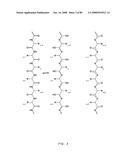
[0057]FIG. 35A: Amino acid sequences of various polypeptide synthetic intermediates of a human IgG1 Fc symmetroadhesin precursor subunit with an N-terminal-S-terminus. Part (i) shows three distinct pre-Fc polypeptides comprising, alternatively, the human sonic hedgehog (SHH), human interferon alpha-2 (IFN), or human cholesterol ester transferase (CETP) signal sequences (residues -23 to -1, -23 to -1, or -17 to -1, respectively), and the human IGHG1 Fc domain (residues 1 to 228) beginning at the fifth amino acid encoded by the hinge exon, CDKTHTCPPCP (Ellison et al. (1982) Nuc. Acids Res. 10, 4071-4079). The three distinct pre-Fc polypeptides have lengths of 251, 251 and 245 residues, respectively. Part (ii) shows the mature Fc domain (length=228) with an N-terminal-S-terminus. The N-terminal cysteine residue is underlined. (IGHG1, UniProtKB/Swiss-Prot entry P01857, Ig gamma-1 chain C region, Homo sapiens).
[0058]FIG. 35B: Amino acid sequences of various polypeptide synthetic intermediates of a human IgG2 Fc symmetroadhesin precursor subunit with an N-terminal-S-terminus. Part (i) shows three distinct pre-Fc polypeptides comprising, alternatively, the human sonic hedgehog (SHH), human interferon alpha-2 (IFN), or human cholesterol ester transferase (CETP) signal sequences (residues -23 to -1, -23 to -1, or -17 to -1, respectively), and the human IGHG2 Fc domain (residues 1 to 225) beginning at the fourth amino acid encoded by the hinge exon, CCVECPPCP (Ellison et al. (1982) Nuc. Acids Res. 10, 4071-4079). The three distinct pre-Fc polypeptides have lengths of 248, 248 and 242 residues, respectively. Part (ii) shows the mature Fc domain (length=225) with an N-terminal-S-terminus. The N-terminal cysteine residue is underlined. (IGHG2, UniProtKB/Swiss-Prot entry P01859, Ig gamma-2 chain C region, Homo sapiens).
[0059]FIG. 35C: Amino acid sequences of various polypeptide synthetic intermediates of a human IgG3 Fc symmetroadhesin precursor subunit with an N-terminal-S-terminus. Part (i) shows three distinct pre-Fc polypeptides comprising, alternatively, the human sonic hedgehog (SHH), human interferon alpha-2 (IFN), or human cholesterol ester transferase (CETP) signal sequences (residues -23 to -1, -23 to -1, or -17 to -1, respectively), and the human IGHG3 Fc domain (residues 1 to 267) beginning at the thirteenth amino acid encoded by the first hinge exon, CPRCP (Strausberg et al. (2002) Proc. Natl. Acad. Sci. 99, 16899-1690). The three distinct pre-Fc polypeptides have lengths of 290, 290 and 284 residues, respectively. Part (ii) shows the mature Fc domain (length=267) with an N-terminal-S-terminus. The N-terminal cysteine residue is underlined. (IGHG3, UniProtKB/Swiss-Prot entry Q8N4Y9, Ig gamma-3 chain C region, Homo sapiens).
[0060]FIG. 35D: Amino acid sequences of various polypeptide synthetic intermediates of a human IgG4 Fc symmetroadhesin precursor subunit with an N-terminal-S-terminus. Part (i) shows three distinct pre-Fc polypeptides comprising, alternatively, the human sonic hedgehog (SHH), human interferon alpha-2 (IFN), or human cholesterol ester transferase (CETP) signal sequences (residues -23 to -1, -23 to -1, or -17 to -1, respectively), and the human IGHG4 Fc domain (residues 1 to 222) beginning at the eighth amino acid encoded by the hinge exon, CPSCP (Strausberg et al. (2002) Proc. Natl. Acad. Sci. 99, 16899-1690). The three distinct pre-Fc polypeptides have lengths of 245, 245 and 239 residues, respectively. Part (ii) shows the mature Fc domain (length=222) with an N-terminal-S-terminus. The N-terminal cysteine residue is underlined. (IGHG4, UniProtKB/Swiss-Prot entry Q8TC63, Ig gamma-4 chain C region, Homo sapiens).
[0061]FIG. 36A: Amino acid sequences of various polypeptide synthetic intermediates of a human Fc symmetroadhesin precursor subunit with an N-terminal-X-terminus. Part (i) shows three distinct pre-Fc polypeptides comprising, alternatively, the human sonic hedgehog (SHH), human interferon alpha-2 (IFN), or human cholesterol ester transferase (CETP) signal sequences (residues -23 to -1, -23 to -1, or -17 to -1, respectively), and the human IGHG1 Fc domain (residues 1 to 222) beginning at the eleventh amino acid encoded by the hinge exon, CPPCP (Ellison et al. (1982) Nuc. Acids Res. 10, 4071-4079). The three distinct pre-Fc polypeptides have lengths of 245, 245 and 239 residues, respectively. Part (ii) shows the mature Fc domain (length=222) having an N-terminal-S-terminus. The N-terminal cysteine residue is underlined. Part (iii) shows the mature Fc domain extended by native chemical ligation (length=226) to have an N-terminal-X-terminus. The N-terminal X amino acid (e.g., cysteine, selenocysteine) is underlined; it is followed by the sixth amino acid encoded by the hinge exon, XDKTHTCPPCP. (IGHG1, UniProtKB/Swiss-Prot entry P01857, Ig gamma-1 chain C region, Homo sapiens).
[0062]FIG. 36B: Amino acid sequences of various polypeptide synthetic intermediates of a human Fc symmetroadhesin precursor subunit with an N-terminal-X-terminus. Part (i) shows three distinct pre-Fc polypeptides comprising, alternatively, the human sonic hedgehog (SHH), human interferon alpha-2 (IFN), or human cholesterol ester transferase (CETP) signal sequences (residues -23 to -1, -23 to -1, or -17 to -1, respectively), and the human IGHG1 Fc domain (residues 1 to 219) beginning at the fourteenth amino acid encoded by the hinge exon, CP (Ellison et al. (1982) Nuc. Acids Res. 10, 4071-4079). The three distinct pre-Fc polypeptides have lengths of 242, 242 and 236 residues, respectively. Part (ii) shows the mature Fc domain (length=219) having an N-terminal-S-terminus. The N-terminal cysteine residue is underlined. Part (iii) shows the mature Fc domain extended by native chemical ligation (length=222) to have an N-terminal-X-terminus. The N-terminal X amino acid (e.g., cysteine, selenocysteine) is underlined; it is followed by the twelfth amino acid encoded by the hinge exon, XPPCP. (IGHG1, UniProtKB/Swiss-Prot entry P01857, Ig gamma-1 chain C region, Homo sapiens).
[0063]FIG. 37A: Amino acid sequences of various polypeptide intermediates in an intein-based synthesis of a human Fc symmetroadhesin precursor subunit with a C-terminal-X-terminus. Part (i) shows two distinct pre-Fc-intein polypeptides comprising, alternatively, the human CD2 or CD4 signal sequences (residues -24 to -1, or -25 to -1, respectively), the human IGHG1 Fc domain (residues 1 to 224) beginning at the seventh amino acid encoded by the hinge exon (KTHTCPPCP), the human IGHG3 M1 domain (residues 225 to 241), and an Mth RIR1 intein-chitin binding domain (residues 242 to 441). The two distinct pre-Fc-intein chimeric polypeptides have lengths of 465 and 466 residues, respectively. Part (ii) shows the mature Fc-intein chimeric polypeptide (length=441) comprising the human Fc/M1 domain and the Mth RIR1 intein-chitin binding domain. The intein autocleavage site is underlined. Part (iii) shows the thioester-terminated human Fc/M1 domain (length=242). The C-terminal thio-glycine residue (Z) is underlined. Part (iv) shows the human Fc/M1 domain (length=243) with a C-terminal-X-terminus. The C-terminal X amino acid residue (e.g., cysteine, selenocysteine) is underlined. (IGHG1, UniProtKB/Swiss-Prot entry P01857, Ig gamma-1 chain C region, Homo sapiens; IGHG3, NCBI/GenBank accession BAA11363, membrane-bound-type Ig gamma-chain, Homo sapiens).
[0064]FIG. 37B: Amino acid sequences of various polypeptide synthetic intermediates of a human Fc symmetroadhesin precursor subunit with an C-terminal-S-terminus. Part (i) shows two distinct pre-Fc polypeptides comprising, alternatively, the human CD2 or CD4 signal sequences (residues -24 to -1, or -25 to -1, respectively), the human IGHG1 Fc domain (residues 1 to 224) beginning at the seventh amino acid encoded by the hinge exon (KTHTCPPCP), and a portion of the human IGHG3 M1 domain (residues 225 to 232). The two distinct pre-Fc polypeptides have lengths of 256 and 257 residues, respectively. Part (ii) shows the mature Fc domain (length=232) with an N-terminal-S-terminus. The N-terminal cysteine residue is underlined. (IGHG1, UniProtKB/Swiss-Prot entry P01857, Ig gamma-1 chain C region, Homo sapiens).
[0065]FIGS. 38A-38B: Amino acid sequences of various polypeptide intermediates in an intein-based synthesis of a human Fc symmetroadhesin precursor subunit, with an N-terminal-S-terminus and a C-terminal-X-terminus. (A) Part (i) shows three distinct pre-Fc-intein polypeptides comprising, alternatively, the human sonic hedgehog (SHH), human interferon alpha-2 (IFN), or human cholesterol ester transferase (CETP) signal sequences (residues -23 to -1, -23 to -1, or -17 to -1, respectively), the human IGHG1 Fc domain (residues 1 to 226) beginning at the fifth amino acid encoded by the hinge exon (CDKTHTCPPCP), the human IGHG3 M1 domain (residues 227 to 243), and an Mth RIR1 intein-chitin binding domain (residues 244 to 443). The three distinct pre-Fc-intein chimeric polypeptides have lengths of 466, 466 and 460 residues, respectively. Part (ii) shows the mature Fc-intein chimeric polypeptide (length=443) comprising the human Fc/M1 domain and the Mth RIR1 intein-chitin binding domain with an N-terminal-S-terminus. The N-terminal cysteine residue and intein autocleavage site are underlined. (B) Part (iii) shows the thioester-terminated human Fc/M1 domain (length=244). The N-terminal cysteine residue and C-terminal thio-glycine residue (Z) are underlined. Part (iv) shows the Fc/M1 domain (length=245) with an N-terminal-S-terminus and a C-terminal-X-terminus. The N-terminal cysteine residue and C-terminal X amino acid residue (e.g., cysteine, selenocysteine) are underlined. (IGHG1, UniProtKB/Swiss-Prot entry P01857, Ig gamma-1 chain C region, Homo sapiens; IGHG3, NCBI/GenBank accession BAA11363, membrane-bound-type Ig gamma-chain, Homo sapiens).
[0066]FIGS. 39A-39B: Amino acid sequences of various polypeptide intermediates in an intein-based synthesis of a human Fc symmetroadhesin precursor subunit, with an N-terminal-X-terminus and a C-terminal-X-terminus. (A) Part (i) shows three distinct pre-Fc-intein polypeptides comprising, alternatively, the human sonic hedgehog (SHH), human interferon alpha-2 (IFN), or human cholesterol ester transferase (CETP) signal sequences (residues -23 to -1, -23 to -1, or -17 to -1, respectively), the human IGHG1 Fc domain (residues 1 to 220) beginning at the eleventh amino acid encoded by the hinge exon (CPPCP), the human IGHG3 M1 domain (residues 221 to 237), and an Mth RIR1 intein-chitin binding domain (residues 238 to 437). The three distinct pre-Fc-intein chimeric polypeptides have lengths of 460, 460 and 454 residues, respectively. Part (ii) shows the mature Fc-intein chimeric polypeptide (length=437) comprising the human Fc/M1 domain and the Mth RIR1 intein-chitin binding domain with an N-terminal-S-terminus. The N-terminal cysteine residue and intein autocleavage site are underlined. (B) Part (iii) shows the mature Fc-intein chimeric polypeptide extended by native chemical ligation (length=443) to have an N-terminal-X-terminus. The N-terminal X amino acid (e.g., cysteine, selenocysteine) is underlined. Part (iv) shows the thioester-terminated human Fc/M1 domain (length=244). The N-terminal X amino acid residue and C-terminal thio-glycine residue (Z) are underlined. Part (v) shows the Fc/M1 domain (length=245) with an N-terminal-X-terminus and a C-terminal-X-terminus. The N-terminal X amino acid residue and C-terminal X amino acid residue are underlined. (IGHG1, UniProtKB/Swiss-Prot entry P01857, Ig gamma-1 chain C region, Homo sapiens; IGHG3, NCBI/GenBank accession BAA11363, membrane-bound-type Ig gamma-chain, Homo sapiens).
[0067]FIGS. 40A-40B: Amino acid sequences of various polypeptide intermediates in an intein-based synthesis of a human CD4 symmetroadhesin precursor subunit with a C-terminal-X-terminus. (A) Part (i) shows the pre-CD4-intein polypeptide (length=596) comprising the human CD4 signal sequence (residues -25 to -1) and extracellular domain (residues 1 to 371), and an Mth RIR1 intein-chitin binding domain (residues 372 to 571). Part (ii) shows the mature CD4-intein chimeric polypeptide (length=571) comprising the human CD4 extracellular domain and the Mth RIR1 intein-chitin binding domain. The position of the intein autocleavage site is underlined. (B) Part (iii) shows the thioester-terminated human CD4 extracellular domain (length=372). The C-terminal thio-glycine residue (Z) is underlined. Part (iv) shows the human CD4 extracellular domain (length=373) with a C-terminal-X-terminus. The C-terminal X amino acid residue (e.g., cysteine, selenocysteine) is underlined. (CD4, UniProtKB/Swiss-Prot entry P01730, T-cell surface glycoprotein CD4).
[0068]FIGS. 41A-41B: Amino acid sequences of various polypeptide intermediates in an intein-based synthesis of a Di62-VH symmetroadhesin precursor subunit with a C-terminal-X-terminus. (A) Part (i) shows the pre-Di62-VH-intein polypeptide (length=444) comprising the mouse Di62-VH signal sequence (residues -19 to -1) and variable domain (residues 1 to 117), the human CH1 contant domain (residues 118 to 225), and an Mth RIR1 intein-chitin binding domain (residues 226 to 425). Part (ii) shows the mature Di62-VH-intein chimeric polypeptide (length=425) comprising the mouse Di62-VH variable domain, the human CH1 contant domain, and the Mth RIR1 intein-chitin binding domain. The position of the intein autocleavage site is underlined. (B) Part (iii) shows the thioester-terminated mouse Di62-VH variable domain/human CH1 contant domain (length=226). The C-terminal thio-glycine residue (Z) is underlined. Part (iv) shows the mouse Di62-VH variable domain/human CH1 contant domain (length=227) with a C-terminal-X-terminus. The C-terminal X amino acid residue (e.g., cysteine, selenocysteine) is underlined. (Di62-VH, NCBI/GenBank accession CAA05416, IgG heavy chain, antigen binding of human TNF alpha subunit, Mus musculus).
[0069]FIG. 42: Amino acid sequences of various polypeptide synthetic intermediates of a Di62-Vk symmetroadhesin precursor subunit. Part (i) shows the pre-Di62-Vk polypeptide (length=234) comprising the mouse Di62-Vk signal sequence (residues -20 to -1) and variable domain (residues 1 to 107), and the human Ck contant domain (residues 108 to 214). Part (ii) shows the mature Di62-Vk chimeric polypeptide (length=214) comprising the mouse Di62-Vk variable domain, and the human Ck contant domain. (Di62-Vk, NCBI/GenBank accession CAA05417, IgG light chain, antigen binding of human TNF alpha subunit, Mus musculus).
[0070]FIG. 43: Amino acid sequences of various polypeptide intermediates in an intein-based synthesis of a human TNR1A symmetroadhesin precursor subunit with a C-terminal-X-terminus. Part (i) shows the pre-TNR1A-intein polypeptide (length=411) comprising the human TNR1A signal sequence (residues -21 to -1) and extracellular domain (residues 1 to 190), and an Mth RIR1 intein-chitin binding domain (residues 191 to 390). Part (ii) shows the mature TNR1A-intein chimeric polypeptide (length=390) comprising the human TNR1A extracellular domain and the Mth RIR1 intein-chitin binding domain. The position of the intein autocleavage site is underlined. Part (iii) shows the thioester-terminated human TNR1A extracellular domain (length=191). The C-terminal thio-glycine residue (Z) is underlined. Part (iv) shows the human TNR1A extracellular domain (length=192) with a C-terminal-X-terminus. The C-terminal X amino acid residue (e.g., cysteine, selenocysteine) is underlined. (TNR1A, UniProtKB/Swiss-Prot entry P19438, Tumor necrosis factor receptor superfamily member 1A).
[0071]FIG. 44A: Amino acid sequences of various polypeptide intermediates in an intein-based synthesis of a human TNR1B symmetroadhesin precursor subunit with a C-terminal-X-terminus. Part (i) shows the pre-TNR1B-intein polypeptide (length=457) comprising the human TNR1B signal sequence (residues -22 to -1) and extracellular domain (residues 1 to 235), and an Mth RIR1 intein-chitin binding domain (residues 236 to 435). Part (ii) shows the mature TNR1B-intein chimeric polypeptide (length=435) comprising the human TNR1B extracellular domain and the Mth RIR1 intein-chitin binding domain. The position of the intein autocleavage site is underlined. Part (iii) shows the thioester-terminated human TNR1B extracellular domain (length=236). The C-terminal thio-glycine residue (Z) is underlined. Part (iv) shows the human TNR1B extracellular domain (length=237) with a C-terminal-X-terminus. The C-terminal X amino acid residue (e.g., cysteine, selenocysteine) is underlined. (TNR1B, UniProtKB/Swiss-Prot entry 20333, Tumor necrosis factor receptor superfamily member 1B).
[0072]FIG. 44B: Amino acid sequences of various polypeptide synthetic intermediates of a TNR1B immunoadhesin precursor subunit. Part (i) shows the pre-TNR1B-immunoadhesin polypeptide (length=489) comprising the TNR1B signal sequence (residues -22 to -1) and extracellular domain (residues 1 to 235), and the human heavy contant domain (residues 236 to 467). Part (ii) shows the mature TNR1B-immunoadhesin (length=467).
[0073]FIGS. 45A-45C: Amino acid sequences of various polypeptide intermediates in an intein-based synthesis of a human VGFR1 symmetroadhesin precursor subunit with a C-terminal-X-terminus. (A) Part (i) shows the pre-VGFR1-intein polypeptide (length=958) comprising the human VGFR1 signal sequence (residues -26 to -1) and extracellular domain (residues 1 to 732), and an Mth RIR1 intein-chitin binding domain (residues 733 to 932). (B) Part (ii) shows the mature VGFR1-intein chimeric polypeptide (length=932) comprising the human VGFR1 extracellular domain and the Mth RIR1 intein-chitin binding domain. The position of the intein autocleavage site is underlined. (C) Part (iii) shows the thioester-terminated human VGFR1 extracellular domain (length=733). The C-terminal thio-glycine residue (Z) is underlined. Part (iv) shows the human VGFR1 extracellular domain (length=734) with a C-terminal-X-terminus. The C-terminal X amino acid residue (e.g., cysteine, selenocysteine) is underlined. (VGFR1, UniProtKB/Swiss-Prot entry P17948, Vascular endothelial growth factor receptor 1).
[0074]FIGS. 46A-46C: Amino acid sequences of various polypeptide intermediates in an intein-based synthesis of a human VGFR2 symmetroadhesin precursor subunit with a C-terminal-X-terminus. (A) Part (i) shows the pre-VGFR2-intein polypeptide (length=964) comprising the human VGFR2 signal sequence (residues -19 to -1) and extracellular domain (residues 1 to 745), and an Mth RIR1 intein-chitin binding domain (residues 746 to 945). (B) Part (ii) shows the mature VGFR2-intein chimeric polypeptide (length=945) comprising the human VGFR2 extracellular domain and the Mth RIR1 intein-chitin binding domain. The position of the intein autocleavage site is underlined. (C) Part (iii) shows the thioester-terminated human VGFR2 extracellular domain (length=746). The C-terminal thio-glycine residue (Z) is underlined. Part (iv) shows the human VGFR2 extracellular domain (length=747) with a C-terminal-X-terminus. The C-terminal X amino acid residue (e.g., cysteine, selenocysteine) is underlined. (VGFR2, UniProtKB/Swiss-Prot entry P35968, Vascular endothelial growth factor receptor 2).
[0075]FIGS. 47A-47C: Amino acid sequences of various polypeptide intermediates in an intein-based synthesis of a human VGFR3 symmetroadhesin precursor subunit with a C-terminal-X-terminus. (A) Part (i) shows the pre-VGFR3-intein polypeptide (length=975) comprising the human VGFR3 signal sequence (residues -24 to -1) and extracellular domain (residues 1 to 751), and an Mth RIR1 intein-chitin binding domain (residues 752 to 951). (B) Part (ii) shows the mature VGFR3-intein chimeric polypeptide (length=951) comprising the human VGFR3 extracellular domain and the Mth RIR1 intein-chitin binding domain. The position of the intein autocleavage site is underlined. (C) Part (iii) shows the thioester-terminated human VGFR3 extracellular domain (length=752). The C-terminal thio-glycine residue (Z) is underlined. Part (iv) shows the human VGFR3 extracellular domain (length=753) with a C-terminal-X-terminus. The C-terminal X amino acid residue (e.g., cysteine, selenocysteine) is underlined. (VGFR3, UniProtKB/Swiss-Prot entry P35916, Vascular endothelial growth factor receptor 3).
[0076]FIGS. 48A-48B: Amino acid sequences of various polypeptide intermediates in an intein-based synthesis of a human ERBB1 symmetroadhesin precursor subunit with a C-terminal-X-terminus. (A) Part (i) shows the pre-ERBB1-intein polypeptide (length=845) comprising the human ERBB1 signal sequence (residues -24 to -1) and extracellular domain (residues 1 to 621), and an Mth RIR1 intein-chitin binding domain (residues 622 to 821). Part (ii) shows the mature ERBB1-intein chimeric polypeptide (length=821) comprising the human ERBB1 extracellular domain and the Mth RIR1 intein-chitin binding domain. The position of the intein autocleavage site is underlined. (B) Part (iii) shows the thioester-terminated human ERBB1 extracellular domain (length=622). The C-terminal thio-glycine residue (Z) is underlined. Part (iv) shows the human ERBB1 extracellular domain (length=623) with a C-terminal-X-terminus. The C-terminal X amino acid residue (e.g., cysteine, selenocysteine) is underlined. (ERBB1, UniProtKB/Swiss-Prot entry P00533, Epidermal growth factor receptor).
[0077]FIGS. 49A-49B: Amino acid sequences of various polypeptide intermediates in an intein-based synthesis of a human ERBB2 symmetroadhesin precursor subunit with a C-terminal-X-terminus. (A) Part (i) shows the pre-ERBB2-intein polypeptide (length=852) comprising the human ERBB2 signal sequence (residues -22 to -1) and extracellular domain (residues 1 to 630), and an Mth RIR1 intein-chitin binding domain (residues 631 to 830). Part (ii) shows the mature ERBB2-intein chimeric polypeptide (length=830) comprising the human ERBB2 extracellular domain and the Mth RIR1 intein-chitin binding domain. The position of the intein autocleavage site is underlined. (B) Part (iii) shows the thioester-terminated human ERBB2 extracellular domain (length=631). The C-terminal thio-glycine residue (Z) is underlined. Part (iv) shows the human ERBB2 extracellular domain (length=632) with a C-terminal-X-terminus. The C-terminal X amino acid residue (e.g., cysteine, selenocysteine) is underlined. (ERBB2, UniProtKB/Swiss-Prot entry P04626, Receptor tyrosine-protein kinase erbB-2).
[0078]FIGS. 50A-50B: Amino acid sequences of various polypeptide intermediates in an intein-based synthesis of a human ERBB3 symmetroadhesin precursor subunit with a C-terminal-X-terminus. (A) Part (i) shows the pre-ERBB3-intein polypeptide (length=843) comprising the human ERBB3 signal sequence (residues -19 to -1) and extracellular domain (residues 1 to 624), and an Mth RIR1 intein-chitin binding domain (residues 625 to 824). Part (ii) shows the mature ERBB3-intein chimeric polypeptide (length=824) comprising the human ERBB3 extracellular domain and the Mth RIR1 intein-chitin binding domain. The position of the intein autocleavage site is underlined. (B) Part (iii) shows the thioester-terminated human ERBB3 extracellular domain (length=625). The C-terminal thio-glycine residue (Z) is underlined. Part (iv) shows the human ERBB3 extracellular domain (length=626) with a C-terminal-X-terminus. The C-terminal X amino acid residue (e.g., cysteine, selenocysteine) is underlined. (ERBB3, UniProtKB/Swiss-Prot entry P21860, Receptor tyrosine-protein kinase erbB-3).
[0079]FIGS. 51A-51B: Amino acid sequences of various polypeptide intermediates in an intein-based synthesis of a human ERBB4 symmetroadhesin precursor subunit with a C-terminal-X-terminus. (A) Part (i) shows the pre-ERBB4-intein polypeptide (length=851) comprising the human ERBB4 signal sequence (residues -25 to -1) and extracellular domain (residues 1 to 626), and an Mth RIR1 intein-chitin binding domain (residues 627 to 826). Part (ii) shows the mature ERBB4-intein chimeric polypeptide (length=826) comprising the human ERBB4 extracellular domain and the Mth RIR1 intein-chitin binding domain. The position of the intein autocleavage site is underlined. (B) Part (iii) shows the thioester-terminated human ERBB4 extracellular domain (length=627). The C-terminal thio-glycine residue (Z) is underlined. Part (iv) shows the human ERBB4 extracellular domain (length=628) with a C-terminal-X-terminus. The C-terminal X amino acid residue (e.g., cysteine, selenocysteine) is underlined. (ERBB4, UniProtKB/Swiss-Prot entry Q15303, Receptor tyrosine-protein kinase erbB-4).
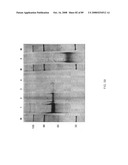
[0080]FIG. 52: Expression in 293 kidney cells of human IgG1 Fc symmetroadhesin subunits with N-terminal-S-termini. Lanes 1-6 and lanes 7-12 show the IgG1 Fc polypeptides of FIG. 35A (ii) and FIG. 36A (ii), respectively. Cell supernatants: lanes 1, 3, 5, 7, 9 and 11; cell lysates: lanes 2, 4, 6, 8, 10 and 12. Signal sequences used: SHH (lanes 1, 2, 7 and 8); IFNα (lanes 3, 4, 9, 10); CETP (lanes 5, 6, 11 and 12).
[0081]FIG. 53: Expression in 293 kidney cells of human IgG1 Fc symmetroadhesin subunits. Lanes 1-2, 3-4 and 5-6 show the IgG1 Fc polypeptides of FIG. 35A (ii), FIG. 36A (ii) and FIG. 37B (ii), respectively. Cell supernatants: (lanes 1-6). Signal sequences used: SHH (lanes 1-6).
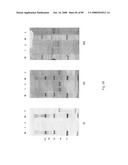
[0082]FIG. 54: Protein A purification of human IgG1 Fc symmetroadhesin subunits expressed in 293 kidney cells. Lane 2 and 8 show the IgG1 Fc polypeptides of FIG. 36A and FIG. 35A, respectively. Lanes 1-7: proteinA-sepharose column fractions for the IgG1 Fc polypeptide of FIG. 36A.
[0083]FIG. 55: Thiol-sepharose binding of proteinA-purified human IgG1 Fc symmetroadhesin subunits shown in FIG. 54. Lanes 1-3 and lanes 4-6 show the human IgG1 Fc polypeptides of FIG. 35A and FIG. 36A, respectively. Lanes 1 and 4: starting material; lanes 2 and 5: thiol-sepharose flow-thru fraction; lanes 3 and 6: thiol-sepharose bound fraction.
[0084]FIG. 56: Expression in human 293 kidney cells of human CD4-intein fusion proteins. Lanes 1-4 show the CD4-intein fusion polypeptide of FIG. 40A (ii). Cell supernatants: lanes 1 and 3; cell lysates: lanes 2 and 4.
[0085]FIG. 57: Expression in human 293 kidney cells of human TNR1B fusion proteins. Lanes 2 and 5 show the TNR1B-intein fusion protein of FIG. 44A (ii). Lanes 1 and 3 show the TNR1B-immunoadhesin fusion protein of FIG. 44B (ii). Lanes 3 and 6 show proteins from mock-transfected cells. Cell supernatants: lanes 1-3; cell lysates: lanes 4-7. Lane 7: control TNR1B-immunoadhesin (R&D Systems).
[0086]FIG. 58: TNR1B symmetroadhesin subunits with C-terminal-S-termini. Lanes 1-2 show the TNR1B polypeptide of FIG. 44A (iii) following purification by chitin affinity chromatography and cleavage/elution with MESNA. Lanes 3 shows the native ligation product between the TNR1B polypeptide of FIG. 44A (iii) with a fluorescent-labeled peptide (New England Biolabs). Panel (i): direct fluorescence; panel (ii): western blot with anti-TNR1B antibody (R&D Systems); panel (iii): SYPRO Ruby staining (Sigma-Aldrich).
[0087]FIG. 59: TNR1B symmetroadhesin subunits with C-terminal-S-termini. Lane 5 shows the TNR1B polypeptide of FIG. 44A (iv) following purification by chitin affinity chromatography and cleavage/elution with cysteine. Lanes 1-4 show TNR1B-immunoadhesin.
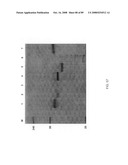
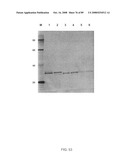
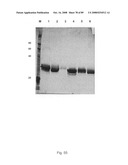
[0088]FIG. 60: TNR1B symmetroadhesin. Lanes 1-4 show the TNR1B symmetroadhesin of FIG. 44A (iv) before oxidation (lanes 1 and 4) and after oxidation in the presence of 10 mM CuSO4. Lanes 3 and 6 show a TNR1B-immunoadhesin control. Lanes 1-3: reducing conditions; lanes 4-6: non-reducing conditions. The TNR1B symmetroadhesin monomer (42 kd) and dimer (84 kd) are apparent in lanes 2 and 5, and lane 5, respectively.
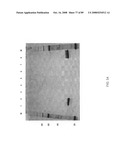
[0089]FIG. 61A-61C: TNF-alpha saturation binding analysis with various TNR1B polypeptides on the Biacore T-100. (A) The TNR1B symmetroadhesin of FIG. 44A (iv) was covalently coupled to a Biacore CM-5 chip using standard Biacore amine chemistry. (B) TNR1B immunoadhesin (R&D Systems) was covalently coupled to a Biacore CM-5 chip using standard Biacore amine chemistry. (C) The TNR1B symmetroadhesin of FIG. 44A (iv) was covalently coupled to a Biacore CM-5 chip using standard Biacore thiol chemistry. Following coupling, saturation binding analysis was carried out using TNF-alpha (R&D Systems) at the indicated concentations.
[0090]FIG. 62A-62C: Scatchard analysis of the TNF-alpha saturation binding analysis shown in FIG. 61A-61C. (A) TNR1B symmetroadhesin of FIG. 44A (iv) covalently coupled using amine chemistry; Kd=Kd=4.697×10-9 M. (B) TNR1B-immunoadhesin (R&D Systems) covalently coupled using amine chemistry; Kd=4.089×10-9 M. (C) TNR1B symmetroadhesin of FIG. 44A (iv) covalently coupled using thiol chemistry; Kd=0.8476×10-9 M.
DETAILED DESCRIPTION
[0091]This invention provides a compound comprising a first stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which comprises a binding site for a target; and a second stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which is identical to the sequence of the first stretch of consecutive amino acids and which comprises an identical binding site for the target; wherein each of the first stretch of amino acids and the second stretch of amino acids has at a predefined end thereof a cysteine residue or a selenocysteine residue and such cysteine residues or such selenocysteine residues are joined by a bond having the structure:
wherein each X is the same and represents a sulfur (S) or a selenium (Se) and each C represents a beta-carbon of one of such cysteine or selenocysteine residues.
[0092]In an embodiment, the bond has the structure:
[0093]In an embodiment, the residue at the predefined end of each of the first stretch and the second stretch of consecutive amino acids is a cysteine residue. In an embodiment the residue at the predefined end of each of the first stretch and the second stretch of consecutive amino acids is a selenocysteine residue.
[0094]This invention also provides a compound comprising a first stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which comprises a binding site for a target; and a second stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which is different from the sequence of the first stretch of consecutive amino acids and which comprises a binding site for a different moiety; wherein each of the first stretch of amino acids and the second stretch of amino acids has at a predefined end thereof a cysteine residue or a selenocysteine residue and such residues are joined by a bond having the structure:
wherein each X may be the same or different and represents a sulfur (S) or a selenium (Se) and each C represents a beta-carbon of one of such cysteine or selenocysteine residues.
[0095]In an embodiment the bond has the structure:
[0096]In an embodiment both of the residues at the predefined ends of each of the first stretch of amino acids and the second stretch of amino acids are cysteine residues. In an embodiment both of the residues at the predefined ends of each of the first stretch of amino acids and the second stretch of amino acids are selenocysteine residues. In an embodiment the residue at one predefined end of one of the first stretch or second stretch of consecutive amino acids is a cysteine residue and the residue at the other predefined end is a selenocysteine residue.
[0097]This invention provides a multimer comprising two or more identical instant compounds joined together by at least one bond. In an embodiment the multimer is a dimer. In an embodiment the multimer is a trimer. In an embodiment the multimer is a tetramer. In an embodiment of the multimer, the one or more bonds comprises a disulfide bond.
[0098]In an embodiment of the compounds the predefined end of both the first stretch of amino acids and the second stretch of amino acids is a N-terminal end thereof. In an embodiment the predefined end of both the first stretch of amino acids and the second stretch of amino acids is a C-terminal end thereof. In an embodiment the predefined end of one of the first stretch of amino acids and the second stretch of amino acids is a C-terminal end and the other predefined end is a N-terminal end.
[0099]In an embodiment of the compounds, the first stretch of amino acids comprises L-amino acids. In an embodiment, the first stretch of amino acids comprises D-amino acids. In an embodiment, the first stretch of amino acids comprises L-amino acids and D-amino acids
[0100]In an embodiment, the second stretch of amino acids comprises L-amino acids. In an embodiment, the second stretch of amino acids comprises D-amino acids. In an embodiment, the second stretch of amino acids comprises L-amino acids and D-amino acids.
[0101]In an embodiment, the first stretch of amino acids comprises at least 50 consecutive amino acids. In an embodiment the second stretch of amino acids comprises at least 50 consecutive amino acids. In an embodiment, the first and/or second stretch of amino acids is between 1 and 100, 100 and 200 or 200 and 300 amino acids in length. In an embodiment the first stretch or second stretch of amino acids comprises at least 20, 25, 30, 35, 40, or 45 consecutive amino acids.
[0102]In an embodiment, the first stretch of amino acids comprises more than one type of amino acid. In an embodiment, the second stretch of amino acids comprises more than one type of amino acid.
[0103]In an embodiment of the instant compounds, the sequence of the first and/or second stretch of amino acids corresponds to the sequence of a constant region of an immunoglobulin. In an embodiment, the immunoglobulin is a human immunoglobulin. In an embodiment, the constant region of the immunoglobulin is a constant region of an IgG, an IgA, an IgE, an IgD, or an IgM immunoglobulin. In an embodiment, the constant region of the immunoglobulin is a constant region of an IgG-1, IgG-2, IgG-3 or IgG-4 immunoglobulin. In an embodiment, the constant region of the immunoglobulin which is a constant region of an IgG-1, IgG-2, IgG-3 or IgG-4 immunoglobulin has one of the sequences set forth herein. In an embodiment, the constant region of the immunoglobulin is a constant region of an IgG immunoglobulin and comprises a hinge region, a CH6 region and a CH3 region. In an embodiment, the different moiety is an immunoeffector or immunoregulator.
[0104]In an embodiment, the target is protein. In an embodiment, the target is an EGF receptor, a HER2, a VEGF receptor, a CD20 antigen, a CD11a, an IgE immunoglobulin, a glycoprotein IIa receptor, a glycoprotein IIIa receptor, a TNF alpha, or a TNF receptor, a gp120. In an embodiment, each of the first and second stretch of consecutive amino acids comprises the amino acid sequence of any one of TNFRSF1a, TNFRSF1b, VEGFR1, VEGFR6, VEGFR3, human Erb1, human Erb2, human Erb6, human Erb3, or human Erb4. In an embodiment, the first or second stretch of consecutive amino acids comprises the amino acid sequence set forth of any one of TNFRSF1a, TNFRSF1b, VEGFR1, VEGFR6, VEGFR3, human Erb1, human Erb2, human Erb6, human Erb3, or human Erb4).
[0105]This invention provides a composition comprising any of the instant compounds in an amount effective to bind the target, and a carrier. In an embodiment, the compound is in an amount effective to bind the target, and a carrier. In an embodiment, the compound is a multimer and is in an amount effective to bind the target, and a carrier. In an embodiment, the multimer is also present in an amount effective to bind the different moiety. In an embodiment, the carrier is a pharmaceutically acceptable carrier. In an embodiment the carrier is a phosphate-buffered saline. Such a composition may be lyophilized.
[0106]This invention provides a method of affecting the activity of a target comprising contacting the target with the composition of one or more of the instant compounds, under conditions such that the compound binds to and affects the activity of the target. In one embodiment, the binding of the composition to the target increases the activity of the target. In one embodiment, the binding of the composition to the target decreases the activity of the target. In one embodiment, the target is an EGF receptor, a HER2 protein, a VEGF receptor, a CD20 antigen, a CD11a, an IgE immunoglobulin, a glycoprotein IIa receptor, a glycoprotein IIIa receptor, a gp40, a gp120, a TNF alpha, or a TNF receptor.
[0107]This invention provides a complex comprising the composition of any of the instant compounds and a third stretch of consecutive amino acids, wherein the third stretch of consecutive amino acids is bound to the one of the first or second stretch of consecutive amino acids by one or more bonds. In an embodiment the one or more bonds comprise van der Waals forces. In one embodiment the one or more bonds comprise a hydrogen bond. In one embodiment the one or bonds comprise a covalent bond. In one embodiment the one or bonds comprise a disulfide bond. In an embodiment the at least one bond is a disulfide bond. In an embodiment the disulfide bond is between two non-terminal amino acid residues. In an embodiment the disulfide bond is between two amino acid residues, at least one of which is a non-terminal amino acid residue.
[0108]This invention provides a process of making one of the instant compounds comprising: [0109](a) transfecting a cell with a recombinant nucleic acid which comprises (i) a first portion, the sequence of which is a N-terminal signal sequence, contiguous with (ii) a second portion, the sequence of which encodes a stretch of consecutive amino acids contiguous with (iii) a third portion, the sequence of which encodes a C-terminal intein-containing binding domain, under conditions permitting synthesis of a chimeric polypeptide comprising the stretch of consecutive amino acids contiguous with the C-terminal intein-containing binding domain; [0110](b) isolating the chimeric polypeptide produced in step (a); [0111](c) treating the chimeric polypeptide so as to cause thio-mediated cleavage of the C-terminal intein-containing binding domain from the stretch of consecutive amino acids and its replacement with a C-terminal thioester; [0112](d) treating the product of step (c) to permit the attachment of a cysteine residue to the product so as to form product with a C-terminal cysteine; and [0113](e) oxidizing the product of step (e) in the presence of another product of step (e) under conditions permitting formation of the compound.
[0114]In one embodiment, the recombinant nucleic acid has the sequence set forth in any one of SEQ ID NOs. 1-8. In one embodiment, the C-terminal intein-containing binding domain is an intein-chitin binding domain. In one embodiment, the C-terminal intein-containing binding domain is an Mth RIR1 intein-chitin binding domain. In one embodiment, the chimeric polypeptide is isolated in step b) by affinity chromatography. In one embodiment, the chimeric polypeptide is isolated in step b) by exposure of the product to a chitin-derivatized resin. In one embodiment, the oxidizing conditions permit formation of a disulfide bond between the C-terminal cysteine of each of the products.
[0115]This invention provides a compound comprising an independently folding protein domain fused to a second independently folding protein domain by non-peptide bond. This invention provides a compound comprising a first polypeptide chain, comprising a terminal cysteine residue, fused at its S-terminus to a S-terminus of a second polypeptide chain comprising a terminal cysteine residue. This invention provides a compound comprising a first polypeptide chain, comprising a terminal selenocysteine residue, fused at its Se-terminus to a S-terminus of a second polypeptide chain comprising a terminal cysteine residue. This invention provides a compound comprising a first polypeptide chain, comprising a terminal selenocysteine residue, fused at its Se-terminus to a Se-terminus of a second polypeptide chain comprising a terminal selenocysteine residue. This invention provides a multimer comprising two or more identical compounds according to any one of claims 66-69 joined together by at least one bond.
[0116]This invention provides a method of making a stretch of consecutive amino acids comprising an N-terminal cysteine comprising: [0117](a) transfecting a cell with a recombinant nucleic acid which comprises (i) a first portion, the sequence of which encodes a N-terminal signal sequence contiguous with (ii) a second portion, the sequence of which encodes a stretch of consecutive amino acids comprising a N-terminal cysteine residue, under conditions permitting (i) synthesis of a chimeric polypeptide which comprises the N-terminal signal sequence joined by a peptide bond at its C-terminus to the N-terminal cysteine of the stretch of consecutive amino acids and (ii) cleavage of the N-terminal signal sequence from the chimeric polypeptide within the cell so as to produce a stretch of consecutive amino acids comprising an N-terminal cysteine; [0118](b) recovering the stretch of consecutive amino acids produced in step (a).
[0119]In one embodiment of the methods disclosed herein, the stretch of consecutive amino acids is isolated in step (b).
[0120]In one embodiment, the stretch of consecutive amino acids comprises an immunoglobulin Fc polypeptide. In one embodiment, the immunoglobulin Fc polypeptide is a human immunoglobulin Fc polypeptide. In one embodiment, the N-terminal cysteine residue is a cys-5 residue of the human immunoglobulin Fc polypeptide. In one embodiment, the cell is a 293 human embryonic cell or a CHO-K1 hamster ovary cell. In one embodiment, the transfection is performed with a plasmid pSA. In one embodiment, N-terminal signal sequence is selected from a protein having a N-terminal cysteine. In one embodiment, the signal peptide is sonic hedgehog, interferon alpha-2 or cholesterol ester transferase. In one embodiment, the stretch of consecutive amino acids is recovered by affinity chromatography. In one embodiment, the cleavage of the chimeric polypeptide within the cell is effected by a cellular signal peptidase.
[0121]This invention provides a method of making a stretch of consecutive amino acids comprising an N-terminal cysteine or selenocysteine comprising: [0122](a) transfecting a cell with a recombinant nucleic acid which comprises (i) a first portion, the sequence of which encodes a N-terminal signal sequence contiguous with (ii) a second portion, the sequence of which encodes a stretch of consecutive amino acids comprising a N-terminal cysteine residue, under conditions permitting (i) synthesis of a chimeric polypeptide comprising the N-terminal signal sequence joined at its C-terminus to the N-terminal cysteine of the Fc polypeptide and (ii) cleavage of the N-terminal signal sequence from the chimeric polypeptide within the cell so as to produce a stretch of consecutive amino acids comprising an N-terminal cysteine; [0123](b) ligating the N-terminal of the stretch of consecutive amino acids produced in step (a) with a C-terminal of a peptide comprising the amino acid sequence cys-asp-lys-thr-his-thr or with a peptide comprising the amino acid sequence sec-asp-lys-thr-his-thr so as to thereby produce the stretch of consecutive amino acids comprising an N-terminal cysteine or selenocysteine; and [0124](c) recovering the stretch of consecutive amino acids produced in step (b).
[0125]In an embodiment, the stretch of consecutive amino acids comprises an immunoglobulin Fc polypeptide. In an embodiment, the immunoglobulin Fc polypeptide is a human immunoglobulin Fc polypeptide. In an embodiment, the N-terminal cysteine residue is a cys-5 residue of the human immunoglobulin Fc polypeptide. In one embodiment, the N-terminal cysteine residue is a cys-11 residue of the human immunoglobulin Fc polypeptide. In an embodiment, the peptide in step (b) comprises the amino acid sequence cys-asp-lys-thr-his-thr and the stretch of consecutive amino acids produced comprises an N-terminal cysteine. In an embodiment, peptide in step (b) comprises the amino acid sequence sec-asp-lys-thr-his-thr and the stretch of consecutive amino acids produced comprises an N-terminal selenocysteine. In an embodiment, the peptide in step (b) is protected with a N-terminal Msc protecting group prior to ligation. In an embodiment, the cell is a 293 human embryonic cell or a CHO-K1 hamster ovary cell. In an embodiment, the transfection is performed with a plasmid pSA. In an embodiment, the N-terminal signal sequence is selected from a protein having a N-terminal cysteine. In an embodiment, the signal peptide is sonic hedgehog, interferon alpha-2 or cholesterol ester transferase. In an embodiment, the stretch of consecutive amino acids is recovered by affinity chromatography. In an embodiment, the cleavage of the chimeric polypeptide within the cell is effected by a cellular signal peptidase. In embodiments other short peptide sequences with a N-terminal cysteine or selenocysteine are employed in place of those set forth above.
[0126]This invention provides process of making a stretch of consecutive amino acids comprising a C-terminal cysteine or a C-terminal selenocysteine, comprising: [0127](a) transfecting a cell with a recombinant nucleic acid which comprises (i) a first portion, the sequence of which encodes a N-terminal signal sequence, contiguous with (ii) a second portion, the sequence of which encodes a stretch of consecutive amino acids contiguous with (iii) a third portion, the sequence of which encodes a C-terminal intein-containing binding domain, under conditions permitting (i) synthesis of a chimeric polypeptide comprising the N-terminal signal sequence contiguous with the stretch of consecutive amino acids contiguous with the C-terminal intein-containing binding domain and (ii) cleavage of the N-terminal signal sequence from the chimeric polypeptide so as to produce a second chimeric polypeptide having a N-terminal lysine residue and comprising the stretch of consecutive amino acids contiguous with the C-terminal intein-containing binding domain; [0128](b) isolating the second chimeric polypeptide produced in step (a); [0129](c) treating the second chimeric polypeptide so as to cause thio-mediated cleavage of the C-terminal intein-containing binding domain from the stretch of consecutive amino acids so as to form a C-terminal thioester; [0130](d) ligating the product of step (c) with a cysteine residue or selenocysteine residue at its C-terminal so as to form product with a C-terminal cysteine or a C-terminal selenocysteine; and [0131](e) recovering the product of step (d).
[0132]In an embodiment, the stretch of consecutive amino acids contiguous comprises an IgG immunoglobulin Fc polypeptide and an IgG M1 exon. In an embodiment, the IgG immunoglobulin is a human IgG immunoglobulin. In an embodiment, the stretch of consecutive amino acids contiguous comprises a CD4 extracellular domain. In an embodiment, the N-terminal signal sequence is selected from a protein having a N-terminal lysine. In an embodiment, the N-terminal signal sequence is a CD2 T-cell surface glycoprotein or a CD4 T-cell surface glycoprotein. In an embodiment, the cell is a 293 human embryonic cell or a CHO-K1 hamster ovary cell. In an embodiment, the transfection is performed with a plasmid pSA. In an embodiment, the C-terminal intein-containing binding domain is an intein-chitin binding domain. In an embodiment, the C-terminal intein-containing binding domain is an Mth RIR1 intein-chitin binding domain. In an embodiment, the C-terminal intein-containing binding domain is a self-splicing intein-containing binding domain. In an embodiment, the chimeric polypeptide is isolated in step b) by exposure of the product to a chitin-derivatized resin. In an embodiment, the cleavage of the second chimeric polypeptide within the cell is effected by a cellular signal peptidase. In an embodiment, the product of step (c) is ligated with a cysteine residue. In an embodiment, the product of step (c) is ligated with a selenocysteine residue.
[0133]This invention provides a process of making a stretch of consecutive amino acids comprising a N-terminal cysteine and a C-terminal cysteine or selenocysteine comprising: [0134](a) transfecting a cell with a recombinant nucleic acid which comprises (i) a first portion, the sequence of which encodes a N-terminal signal sequence, contiguous with (ii) a second portion, the sequence of which encodes a stretch of consecutive amino acids contiguous with (iii) a third portion, the sequence of which encodes a C-terminal intein-containing binding domain, under conditions permitting (i) synthesis of a chimeric polypeptide comprising the N-terminal signal sequence contiguous with the stretch of consecutive amino acids contiguous with the C-terminal intein-containing binding domain and (ii) cleavage of the N-terminal signal sequence from the chimeric polypeptide so as to produce a second chimeric polypeptide having a N-terminal cysteine residue and comprising the stretch of consecutive amino acids contiguous with the C-terminal intein-containing binding domain; [0135](b) isolating the second chimeric polypeptide produced in step (a); [0136](c) treating the second chimeric polypeptide so as to cause thio-mediated cleavage of the C-terminal intein-containing binding domain from the stretch of consecutive amino acids so as to form a C-terminal thioester; [0137](d) ligating the product of step (c) with a cysteine residue or selenocysteine residue at its C-terminal so as to form product with a C-terminal cysteine or a C-terminal selenocysteine; and [0138](e) recovering the product of step (d).
[0139]In an embodiment, the stretch of consecutive amino acids contiguous comprises an IgG immunoglobulin Fc polypeptide and an IgG M1 exon. In an embodiment, the IgG immunoglobulin is a human IgG immunoglobulin. In an embodiment, the N-terminal signal sequence is selected from a protein having a N-terminal cysteine. In an embodiment, the N-terminal cysteine residue is a cys-5 residue of the human immunoglobulin Fc polypeptide. In an embodiment, the N-terminal cysteine residue is a cys-11 residue of the human immunoglobulin Fc polypeptide. In an embodiment, the N-terminal signal is a sonic hedgehog, interferon alpha-2 or cholesterol ester transferase. In an embodiment, the cell is a 293 human embryonic cell or a CHO-K1 hamster ovary cell. In an embodiment, the transfection is performed with a plasmid pSA. In an embodiment, the C-terminal intein-containing binding domain is an intein-chitin binding domain. In an embodiment, the C-terminal intein-containing binding domain is an Mth RIR1 intein-chitin binding domain. In an embodiment, the C-terminal intein-containing binding domain is a self-splicing intein-containing binding domain In an embodiment, the second chimeric polypeptide is isolated in step b) by exposure of the product to a chitin-derivatized resin. In an embodiment, the product is recovered in step e) by affinity chromatography. In an embodiment, the cleavage of the chimeric polypeptide within the cell is effected by a cellular signal peptidase. In an embodiment, the product of step (c) is ligated with a cysteine residue. In an embodiment, the product of step (c) is ligated with a selenocysteine residue.
[0140]This invention provides a process of making a stretch of consecutive amino acids comprising a N-terminal cysteine or selenocysteine and a C-terminal cysteine or selenocysteine comprising: [0141](a) transfecting a cell with a recombinant nucleic acid which comprises (i) a first portion, the sequence of which encodes a N-terminal signal sequence, contiguous with (ii) a second portion, the sequence of which encodes a stretch of consecutive amino acids contiguous with (iii) a third portion, the sequence of which encodes a C-terminal intein-containing binding domain, under conditions permitting (i) synthesis of a chimeric polypeptide comprising the N-terminal signal sequence contiguous with the stretch of consecutive amino acids contiguous with the C-terminal intein-containing binding domain and (ii) cleavage of the N-terminal signal sequence from the chimeric polypeptide so as to produce a second chimeric polypeptide having a N-terminal cysteine residue and comprising the stretch of consecutive amino acids contiguous with the C-terminal intein-containing binding domain; [0142](b) isolating the second chimeric polypeptide produced in step (a); [0143](c) (i) ligating the N-terminal of the stretch of consecutive amino acids produced in step (a) with a C-terminal of a peptide comprising the amino acid sequence cys-asp-lys-thr-his-thr or with a peptide comprising the amino acid sequence sec-asp-lys-thr-his-thr so as to thereby produce the stretch of consecutive amino acids comprising a N-terminal cysteine or a N-terminal selenocysteine, respectively; [0144](ii) treating the chimeric polypeptide so as to cause thio-mediated cleavage of the C-terminal intein-containing binding domain from the stretch of consecutive amino acids and its replacement with a C-terminal thioester; [0145](iii) ligating the product of step (c) with a cysteine residue or selenocysteine residue at its C-terminal so as to form product with a C-terminal cysteine pr with a C-terminal selenocysteine; and [0146](d) recovering the product of step (c)(iii).
[0147]In an embodiment, the stretch of consecutive amino acids contiguous comprises an IgG immunoglobulin Fc polypeptide and an IgG M1 exon. In an embodiment, the IgG immunoglobulin is a human IgG immunoglobulin. In an embodiment, the N-terminal signal sequence is selected from a protein having a N-terminal cysteine. In an embodiment, the N-terminal cysteine residue is a cys-11 residue of the human immunoglobulin Fc polypeptide. In an embodiment, N-terminal signal sequence is selected from a protein having a N-terminal cysteine. In an embodiment, the N-terminal signal is a sonic hedgehog, interferon alpha-2 or cholesterol ester transferase. In an embodiment, the cell is a 293 human embryonic cell or a CHO-K1 hamster ovary cell. In an embodiment, the transfection is performed with a plasmid pSA. In an embodiment, the C-terminal intein-containing binding domain is an intein-chitin binding domain. In an embodiment, the C-terminal intein-containing binding domain is an Mth RIR1 intein-chitin binding domain. In an embodiment, the chimeric polypeptide is isolated in step (b) by exposure of the product to a chitin-derivatized resin.
[0148]In an embodiment, the immunoglobulin Fc polypeptide is a human immunoglobulin Fc polypeptide. In an embodiment, the N-terminal cysteine residue of the Fc polypeptide is a cys-11 residue. In an embodiment, the peptide in step (c) (i) comprises the amino acid sequence cys-asp-lys-thr-his-thr and the stretch of consecutive amino In an embodiment, the peptide in step (c) (i) comprises the amino acid sequence sec-asp-lys-thr-his-thr and the stretch of consecutive amino acids produced comprises an N-terminal selenocysteine. In an embodiment, the peptide in step (c) is protected with a N-terminal Msc protecting group prior to ligation. In an embodiment, in the cell is a 293 human embryonic cell or a CHO-K1 hamster ovary cell. In an embodiment, the transfection is performed with a plasmid pSA. In an embodiment, the N-terminal signal sequence is selected from a protein having a N-terminal cysteine. In an embodiment, the signal peptide is sonic hedgehog, interferon alpha-2 or cholesterol ester transferase. In an embodiment, in the stretch of consecutive amino acids is recovered by affinity chromatography.
[0149]In an embodiment, step (c) of this instant method is performed in the order step (c) (i); step (c) (ii); step (c) (iii). In an embodiment, step (c) is performed in the order step (c)(ii); step (c)(iii); step (c)(i).
[0150]This invention provides a process for making a compound comprising contacting a stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which comprises a binding site for a target with a second stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which is identical to the sequence of the first stretch of consecutive amino acids and which comprises an identical binding site for the target, wherein each of the first stretch of amino acids and the second stretch of amino acids has at a predefined end thereof a cysteine residue or a selenocysteine residue, under reducing conditions so as to make the compound.
[0151]This invention provides a process for making a compound comprising contacting a stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which comprises a binding site for a target with a second stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which is different to the sequence of the first stretch of consecutive amino acids and which comprises an identical binding site for the target, wherein each of the first stretch of amino acids and the second stretch of amino acids has at a predefined end thereof a cysteine residue or a selenocysteine residue, under reducing conditions so as to make the compound.
[0152]In an embodiment of the instant processes, the reducing conditions do not denature the stretches of consecutive amino acids. In an embodiment, the reducing conditions comprise exposing the stretches of consecutive amino acids to a buffer comprising Tris-HCL and mercaptoethanol. In an embodiment, the buffer is between pH 7.6 and 8.4. In an embodiment, the buffer is pH 8. In an embodiment, the method further comprises exchanging the product into oxidation buffer. In an embodiment, the stretches of consecutive amino acids comprises a CD4 extracellular domain. In an embodiment, the stretches of consecutive amino acids comprises a sequence of an immunoglobulin Fc polypeptide. In an embodiment, the immunoglobulin is a human immunoglobulin.
[0153]In embodiments, the transfected cells of the instant methods are grown under conditions suitable to permit expression of the polypeptide.
[0154]In an embodiment a compound is provided comprising a first stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which comprises a binding site for a target; and
a second stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which is identical to the sequence of the first stretch of consecutive amino acids and which comprises an identical binding site for the target;wherein each of the first stretch of amino acids and the second stretch of amino acids has at a predefined end thereof, independently, a natural amino acid or non-natural amino having a linear aliphatic side-chain acid comprising a sulfur (S) or a selenium (Se) and wherein such sulfur (S) or a selenium (Se) are joined by a bond having the structure:
wherein each X is a sulfur (S) or a selenium (Se) and each (C) represents a carbon of the linear aliphatic side-chain of one of such natural or non-natural amino acid and wherein n and m are, independently, 2, 3, 4, 5, 6, 7, 8, 9, or 10.
[0155]In an embodiment, the natural amino acid is homocysteine of homoselenocysteine. In an embodiment, the first stretch and second stretch of amino acids have a homocysteine at the predefined end thereof. In an embodiment, the first stretch and second stretch of amino acids have a homoselenocysteine at the predefined end thereof. In an embodiment, the first stretch and second stretch of amino acids have a homocysteine at the predefined end thereof. In an embodiment, the predefined end is a C-terminus.
[0156]A compound is provided which comprises a first stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which comprises a binding site for a target; and
a second stretch of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond and the sequence of which is different from the sequence of the first stretch of consecutive amino acids and which comprises a binding site for a different moiety;wherein each of the first stretch of amino acids and the second stretch of amino acids has at a predefined end thereof, independently, a natural amino acid or non-natural amino having a linear aliphatic side-chain acid comprising a sulfur (S) or a selenium (Se) and wherein such sulfur (S) or a selenium (Se) are joined by a bond having the structure:
wherein each X may be the same or different and represents a sulfur (S) or a selenium (Se) and each (C) represents a carbon of the linear aliphatic side-chain of one of such natural or non-natural amino acid and wherein n and m are, independently, 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10.
[0157]In an embodiment, the natural amino acid is homocysteine of homoselenocysteine. In an embodiment, the first stretch and second stretch of amino acids have a homocysteine at the predefined end thereof. In an embodiment, the first stretch and second stretch of amino acids have a homoselenocysteine at the predefined end thereof. In an embodiment, the first stretch and second stretch of amino acids have a homocysteine at the predefined end thereof. In an embodiment, the predefined end is a C-terminus.
[0158]A method is provided of producing a protein which comprises a first polypeptide contiguous with an intein, which intein is contiguous with a second polypeptide comprising a binding domain, the method comprising transfecting an animal cell with a nucleic acid, which nucleic acid comprises (i) a first portion which encodes the polypeptide contiguous with (ii) a second portion which encodes the intein, contiguous with a and the third portion of which encodes the binding domain, under conditions such that the animal cell expresses and secretes the protein. In an embodiment, the animal cell is a derived from mammal. In an embodiment, the binding domain is a chitin-binding domain.
[0159]A composition is provided comprising a polypeptide attached to a solid surface through a terminal disulfide bond. In an embodiment, the solid surface is a chip or a bead.
[0160]A compound is provided comprising:
a first stretch of consecutive amino acids each of which is joined to the preceding amino acid by a peptide bond and which first stretch of consecutive amino acids comprises an amino acid residue having a chalcogen functional group-containing side chain; anda second stretch of consecutive amino acids, comprising at least 100 amino acids, each of which is joined to the preceding amino acid by a peptide bond, wherein at least 90 consecutive amino acids of the second stretch of consecutive amino acids have a sequence identical to portion of a human immunoglobulin constant region polypeptide, and wherein the second stretch of consecutive amino acids comprises an amino acid residue having a chalcogen functional group-containing side chain at a predefined terminus thereof,wherein said amino acid residue having a chalcogen functional group-containing side chain of the first stretch of consecutive amino acids and said amino acid residue having a chalcogen functional group-containing side chain of the second stretch of consecutive amino acids are joined by a bond having the structure:
wherein each X represents, independently, a chalcogen, and wherein C1 represents a side chain carbon of the amino acid residue having a chalcogen functional group-containing side chain of the first stretch of consecutive amino acids and C2 represents a side chain carbon of the second stretch of consecutive amino acids.
[0161]In embodiments at least wherein at least 91, 92, 93, 94, 95, 96, 97, 98, 99 or 100 consecutive amino acids of the second stretch of consecutive amino acids have a sequence identical to portion of a human immunoglobulin constant region polypeptide.
[0162]In an embodiment of the compound, at least one of C1 and C2 is a beta carbon of amino acid. In an embodiment of the compound the bond has the structure:
wherein S is sulfur. In an embodiment of the compound the bond has the structure:
[0163]wherein S is sulfur and Se is selenium.
[0164]In an embodiment of the compound the amino acid residue having a chalcogen functional group-containing side chain at the predefined terminus of the second stretch of amino acids is a cysteine. In an embodiment of the compound the amino acid residue having a chalcogen functional group-containing side chain at the predefined terminus of the second stretch of amino acids is a selenocysteine, homocysteine or homoseleneocysteine. In an embodiment of the compound the amino acid residue having a chalcogen functional group-containing side chain of the first stretch of consecutive amino acids is a cysteine. In an embodiment of the compound the amino acid residue having a chalcogen functional group-containing side chain of the first stretch of amino acids is a selenocysteine, homocysteine or homoseleneocysteine. In an embodiment of the compound the amino acid residue having a chalcogen functional group-containing side chain of the first stretch of consecutive amino acids is a terminal residue.
[0165]In an embodiment, the amino acid residue having a chalcogen functional group-containing side chain of the first stretch of consecutive amino acids is a penultimate, antepenultimate, or pre-antepenultimate terminal residue.
[0166]In an embodiment, the second stretch of consecutive amino acids has a sequence identical to a human immunoglobulin constant region. In an embodiment, the second stretch of consecutive amino acids is a portion of a human immunoglobulin constant region. In an embodiment, the first stretch of consecutive amino acids has a sequence identical to a human immunoglobulin constant region.
[0167]In an embodiment, the human immunoglobulin constant region polypeptide is a human IgG1, human IgG2, human IgG3, or human IgG4. A compound is provided wherein the side chain of at least one of amino acid residues having a chalcogen functional group-containing side chain comprises a C1-C10 alkylene.
[0168]A composition is provided comprising two of the instant compounds bonded together via at least one disulfide bond between the second stretch of consecutive amino acids of each of the compounds.
[0169]A composition comprising a polypeptide is provided comprising consecutive amino acids having the sequence set forth in one of SEQ ID NOS:35 through 46, or having the sequence set forth in one of SEQ ID NOS:53 through 67, or having the sequence set forth in one of SEQ ID NOS:74 through 82, or having the sequence set forth in one of SEQ ID NOS:89 through 97, wherein the polypeptide does not consist of a naturally-occurring immunoglobulin polypeptide, including enzyme-cleaved fragments thereof.
[0170]A composition comprising a polypeptide is provided consisting of consecutive amino acids having the sequence set forth in one of SEQ ID NOS:35 through 46, or having the sequence set forth in one of SEQ ID NOS:53 through 67, or having the sequence set forth in one of SEQ ID NOS:74 through 82, or having the sequence set forth in one of SEQ ID NOS:89 through 97.
[0171]A composition comprising the instant polypeptide is provided and a carrier. In an embodiment of the compound, the carrier is phosphate-buffered saline.
[0172]A composition is provided comprising two of the instant independently chosen polypeptides, joined by a non-peptide bond. In one embodiment the bond is a di-chalcogenide bond. In one embodiment the bond is a disulfide bond.
[0173]A composition comprising two of the instant polypeptides bonded together via at least one disulfide bond between the two polypeptides is provided.
[0174]A composition is provided comprising a polypeptide is provided consisting of consecutive amino acids having a sequence identical to a portion of the sequence set forth in SEQ ID NO:44, SEQ ID NO:64, SEQ ID NO:81 or SEQ ID NO:96, wherein at least one of the terminal residues of the polypeptide has a chalcogen functional group-containing side chain.
[0175]In an embodiment of the compound the terminal residue having a chalcogen functional group-containing side chain is a cysteine or analog thereof.
[0176]The various N-terminal signal sequences, plasmids, expression vecots, recombinant nucleic acids, stretches of consecutive amino acids, intein binding domains, cell types, recovery/isolation methods, etc set forth hereinabove are non-limiting examples, further of which are set forth in the Examples below.
DEFINITION OF TERMS
[0177]Stretch of consecutive amino acids: a plurality of amino acids arranged in a chain, each of which is joined to a preceding amino acid by a peptide bond, excepting that the first amino acid in the chain is not joined to a preceding amino acid. The amino acids of the chain may be naturally or non-naturally occurring, or may comprise a mixture thereof. The amino acids, unless otherwise indicated, may be genetically encoded, naturally-occurring but not genetically encoded, or non-naturally occurring, and any selection thereof.
[0178]In an embodiment, a stretch of consecutive amino acids has biological activity, including, but not limited to, target-binding activity or an immunoeffector activity, which biological activity is retained on the bonding of the stretch of consecutive amino acids to another stretch of consecutive amino acids by an --X--X-- bond (e.g. --S--S--, --S--Se--, --Se--Se--, or --Se--S-- bond). A "segment of consecutive amino acids" is an alternative description of "a stretch of amino acids".
[0179]N-terminal amino acid residue: the terminal residue of a stretch of two or more consecutive amino acids having a free α-amino (NH2) functional group, or a derivative of an α-amino (NH2) functional group.
[0180]N-terminus: the free α-amino (NH2) group (or derivative thereof) of a N-terminal amino acid residue.
[0181]C-terminal amino acid residue: the terminal residue of a stretch of two or more consecutive amino acids having a free α-carboxyl (COOH) functional group, or a derivative of a α-carboxyl (COOH) functional group.
[0182]C-terminus: the free α-carboxyl (COOH) group (or derivative thereof) of a C-terminal amino acid residue.
[0183]S-terminal cysteine residue: a cysteine which is the N- and/or C-terminal residue(s) of a stretch of consecutive amino acids, and which has a free β-sulfhydryl (SH) functional group, or a derivative of a β-sulfhydryl (SH) functional group.
[0184]S-terminus: the free β-sulfhydryl (SH) group (or derivative thereof) of a S-terminal cysteine residue.
[0185]Se-terminal selenocysteine residue: a selenocysteine which is the N- and/or C-terminal residue(s) of a stretch of consecutive amino acids, and which has a free β-selenohydryl (SeH) functional group, or a derivative of a β-selenohydryl (SeH) functional group.
[0186]Se-terminus: the free β-selenohydryl (SeH) group (or derivative thereof) of a Se-terminal selenocysteine residue.
[0187]X-terminal amino acid residue: a cysteine (or cysteine derivative) or homocysteine (or homocysteine derivative), selenocysteine (or selenocysteine derivative) or homoselenocysteine (or homoselenocysteine derivative), which is the N- and/or C-terminal residue(s) of a stretch of consecutive amino acids, and which has a free β-sulfhydryl (SH) or β-selenohydryl (SeH) functional group, respectively, or a sulfur-containing or selenium-containing derivative thereof.
[0188]X-terminus: the free β-sulfhydryl (SH) or β-selenohydryl (SeH) group of a S-terminal cysteine/cysteine derivative residue or of a Se-terminal selenocysteine/selenocysteine derivative residue, respectively. In addition, an X-terminus can be the free β-sulfhydryl (SH) or β-selenohydryl (SeH) group of a S-terminal homocysteine residue or Se-terminal homoselenocysteine residue.
[0189]Target: an entity which binds to a discrete selection of a stretch of consecutive amino acids or a portion of a tertiary structure thereof and includes, but is not limited to, receptors, carrier proteins, hormones, cellular adhesive proteins, tissue-specific adhesion factors, growth factors, and enzymes. Specific examples of targets include a human EGF receptor, a HER2 protein, a VEGF receptor, a human CD20 antigen, a human CD11a, a human IgE immunoglobulin, a human glycoprotein IIa receptor, a human glycoprotein IIIa receptor, a human TNF alpha, and a TNF receptor.
[0190]A "bond", unless otherwise specified, or contrary to context, is understood to include a covalent bond, a dipole-dipole interaction such as a hydrogen bond, and intermolecular interactions such as van der Waals forces.
[0191]A "Signal Sequence" is a short (3-60 amino acids long) peptide chain that directs the post-translational transport of a polypeptide.
[0192]Amino acid" as used herein, in one embodiment, means a L or D isomer of the genetically encoded amino acids, i.e. isoleucine, alanine, leucine, asparagine, lysine, aspartate, methionine, cysteine, phenylalanine, glutamate, threonine, glutamine, tryptophan, glycine, valine, proline, arginine, serine, histidine, tyrosine, selenocysteine, pyrolysine and also includes homocysteine and homoselenocysteine.
[0193]Other examples of amino acids include an L or D isomer of taurine, gaba, dopamine, lanthionine, 2-aminoisobutyric acid, dehydroalanine, ornithine and citrulline, as well as non-natural homologues and synthetically modified forms thereof including amino acids having alkylene chains shortened or lengthened by up to two carbon atoms, amino acids comprising optionally substituted aryl groups, and amino acids comprising halogenated groups, including halogenated alkyl and aryl groups as well as beta or gamma amino acids, and cyclic analogs.
[0194]Due to the presence of ionizable amino and carboxyl groups, the amino acids in these embodiments may be in the form of acidic or basic salts, or may be in neutral forms. Individual amino acid residues may also be modified by oxidation or reduction. Other contemplated modifications include hydroxylation of proline and lysine, phosphorylation of hydroxyl groups of seryl or threonyl residues, and methylation of the alpha-amino groups of lysine, arginine, and histidine side chains.
[0195]Covalent derivatives may be prepared by linking particular functional groups to the amino acid side chains or at the N- or C-termini.
[0196]Chalcogen" as used herein is limited to sulfur, selenium, tellurium and polonium only, i.e. as used herein, "chalcogen" excludes oxygen and ununhexium.
[0197]Chalcogen functional group containing side chain" as used herein is an amino acid residue side chain containing a terminally reactive non-oxygen, non-ununhexium chalcogen atom. By way of non-limiting example, an amino acid having a chalcogen functional group containing side chain would be cysteine, selenocysteine, homocysteine etc., but would not include methionine, for example, which contains a chalcogen atom (S), but not a terminally reactive chalcogen atom.
[0198]Compounds comprising amino acids with R-group substitutions are within the scope of the invention. It is understood that substituents and substitution patterns on the compounds of the instant invention can be selected by one of ordinary skill in the art to provide compounds that are chemically stable from readily available starting materials.
[0199]Natural amino acid" as used herein means a L or D isomer of the genetically encoded amino acids, i.e. isoleucine, alanine, leucine, asparagine, lysine, aspartate, methionine, cysteine, phenylalanine, glutamate, threonine, glutamine, tryptophan, glycine, valine, proline, arginine, serine, histidine, tyrosine, selenocysteine, pyrolysine and homocysteine and homoselenocysteine.
[0200]Non-natural amino acid" as used herein means a chemically modified L or D isomer of isoleucine, alanine, leucine, asparagine, lysine, aspartate, methionine, cysteine, phenylalanine, glutamate, threonine, glutamine, tryptophan, glycine, valine, proline, arginine, serine, histidine, tyrosine, selenocysteine, pyrolysine, homocysteine, homoselenocysteine, taurine, gaba, dopamine, lanthionine, 2-aminoisobutyric acid, dehydroalanine, ornithine or citrulline, including cysteine and selenocysteine derivatives having C3-C10 aliphatic side chains between the alpha carbon and the S or Se. In one embodiment the aliphatic side chain is an alkylene. In another embodiment, the aliphatic side chain is an alkenylene or alkynylene.
[0201]In addition to the stretches of consecutive amino acid sequences described herein, it is contemplated that variants thereof can be prepared by introducing appropriate nucleotide changes into the encoding DNA, and/or by synthesis of the desired consecutive amino acid sequences. Those skilled in the art will appreciate that amino acid changes may alter post-translational processes of the stretches of consecutive amino acids described herein when expression is the chosen method of synthesis (rather than chemical synthesis for example), such as changing the number or position of glycosylation sites or altering the membrane anchoring characteristics.
[0202]Variations in the sequences described herein, can be made, for example, using any of the techniques and guidelines for conservative and non-conservative mutations set forth, for instance, in U.S. Pat. No. 5,364,934. Variations may be a substitution, deletion or insertion of one or more codons encoding the consecutive amino acid sequence of interest that results in a change in the amino acid sequence as compared with the native sequence. Optionally the variation is by substitution of at least one amino acid with any other amino acid in one or more of the domains. Guidance in determining which amino acid residue may be inserted, substituted or deleted without adversely affecting the desired activity may be found by comparing the sequence with that of homologous known protein molecules and minimizing the number of amino acid sequence changes made in regions of high homology. Amino acid substitutions can be the result of replacing one amino acid with another amino acid having similar structural and/or chemical properties, such as the replacement of a leucine with a serine, i.e., conservative amino acid replacements. Insertions or deletions may optionally be in the range of about 1 to 5 amino acids. The variation allowed may be determined by systematically making insertions, deletions or substitutions of amino acids in the sequence and testing the resulting variants for activity exhibited by the full-length or mature native sequence. It is understood that any terminal variations are made within the context of the invention disclosed herein.
[0203]Amino acid sequence variants of the binding partner are prepared with various objectives in mind, including increasing the affinity of the binding partner for its ligand, facilitating the stability, purification and preparation of the binding partner, modifying its plasma half life, improving therapeutic efficacy, and lessening the severity or occurrence of side effects during therapeutic use of the binding partner.
[0204]Amino acid sequence variants of these sequences are also contemplated herein including insertional, substitutional, or deletional variants. Such variants ordinarily can prepared by site-specific mutagenesis of nucleotides in the DNA encoding the target-binding monomer, by which DNA encoding the variant is obtained, and thereafter expressing the DNA in recombinant cell culture. Fragments having up to about 100-150 amino acid residues can also be prepared conveniently by in vitro synthesis. Such amino acid sequence variants are predetermined variants and are not found in nature. The variants exhibit the qualitative biological activity (including target-binding) of the nonvariant form, though not necessarily of the same quantative value. While the site for introducing an amino acid sequence variation is predetermined, the mutation per se need not be predetermined. For example, in order to optimize the performance of a mutation at a given site, random or saturation mutagenesis (where all 20 possible residues are inserted) is conducted at the target codon and the expressed variant is screened for the optimal combination of desired activities. Such screening is within the ordinary skill in the art.
[0205]Amino acid insertions usually will be on the order of about from 1 to 10 amino acid residues; substitutions are typically introduced for single residues; and deletions will range about from 1 to 30 residues. Deletions or insertions preferably are made in adjacent pairs, i.e. a deletion of 2 residues or insertion of 2 residues. It will be amply apparent from the following discussion that substitutions, deletions, insertions or any combination thereof are introduced or combined to arrive at a final construct.
[0206]In an aspect, the invention concerns a compound comprising a stretch of consecutive amino acids having at least about 80% sequence identity, preferably at least about 81% sequence identity, more preferably at least about 82% sequence identity, yet more preferably at least about 83% sequence identity, yet more preferably at least about 84% sequence identity, yet more preferably at least about 85% sequence identity, yet more preferably at least about 86% sequence identity, yet more preferably at least about 87% sequence identity, yet more preferably at least about 88% sequence identity, yet more preferably at least about 89% sequence identity, yet more preferably at least about 90% sequence identity, yet more preferably at least about 91% sequence identity, yet more preferably at least about 92% sequence identity, yet more preferably at least about 93% sequence identity, yet more preferably at least about 94% sequence identity, yet more preferably at least about 95% sequence identity, yet more preferably at least about 96% sequence identity, yet more preferably at least about 97% sequence identity, yet more preferably at least about 98% sequence identity and yet more preferably at least about 99% sequence identity to an amino acid sequence disclosed in the specification, a figure, a SEQ ID NO. or a sequence listing of the present application.
[0207]The % amino acid sequence identity values can be readily obtained using, for example, the WU-BLAST-2 computer program (Altschul et al., Methods in Enzymology 266:460-480 (1996)).
[0208]Fragments of native sequences are provided herein. Such fragments may be truncated at the N-terminus or C-terminus, or may lack internal residues, for example, when compared with a full length native protein. Again, it is understood that any terminal variations are made within the context of the invention disclosed herein. Certain fragments lack amino acid residues that are not essential for a desired biological activity of the sequence of interest.
[0209]Any of a number of conventional techniques may be used. Desired peptide fragments or fragments of stretches of consecutive amino acids may be chemically synthesized. An alternative approach involves generating fragments by enzymatic digestion, e.g. by treating the protein with an enzyme known to cleave proteins at sites defined by particular amino acid residues, or by digesting the DNA with suitable restriction enzymes and isolating the desired fragment. Yet another suitable technique involves isolating and amplifying a DNA fragment encoding a desired polypeptide/sequence fragment, by polymerase chain reaction (PCR). Oligonucleotides that define the desired termini of the DNA fragment are employed at the 5' and 3' primers in the PCR.
[0210]In particular embodiments, conservative substitutions of interest are shown in Table 1 under the heading of preferred substitutions. If such substitutions result in a change in biological activity, then more substantial changes, denominated exemplary substitutions in Table 1, or as further described below in reference to amino acid classes, are introduced and the products screened.
TABLE-US-00001 TABLE 1 Original Exemplary Preferred Ala (A) val; leu; ile val Arg (R) lys; gln; asn lys Asn (N) gln; his; lys; arg gln Asp (D) glu glu Cys (C) ser ser Gln (Q) asn asn Glu (E) asp asp Gly (G) pro; ala ala His (H) asn; gln; lys; arg arg Ile (I) leu; val; met; ala; phe; norleucine leu Leu (L) norleucine; ile; val; met; ala; phe ile Lys (K) arg; gln; asn arg Met (M) leu; phe; ile leu Phe (F) leu; val; ile; ala; tyr leu Pro (P) ala ala Ser (S) thr thr Thr (T) ser ser Trp (W) tyr; phe tyr Tyr (Y) trp; phe; thr; ser phe Val (V) ile; leu; met; phe; ala; norleucine leu
[0211]Substantial modifications in function or immunological identity of the sequence are accomplished by selecting substitutions that differ significantly in their effect on maintaining (a) the structure of the polypeptide backbone in the area of the substitution, for example, as a sheet or helical conformation, (b) the charge or hydrophobicity of the molecule at the target site, or (c) the bulk of the side chain. Naturally occurring residues are divided into groups based on common side-chain properties:
(1) hydrophobic: norleucine, met, ala, val, leu, ile;(2) neutral hydrophilic: cys, ser, thr;(3) acidic: asp, glu;(4) basic: asn, gln, his, lys, arg;(5) residues that influence chain orientation: gly, pro;(6) aromatic: trp, tyr, phe.
[0212]Non-conservative substitutions will entail exchanging a member of one of these classes for another class. Such substituted residues also may be introduced into the conservative substitution sites or, more preferably, into the remaining (non-conserved) sites.
[0213]The variations can be made using methods known in the art such as oligonucleotide-mediated (site-directed) mutagenesis, alanine scanning, and PCR mutagenesis. Site-directed mutagenesis (Carter et al., Nucl. Acids Res., 13:4331 (1986); Zoller et al., Nucl. Acids Res., 10:6487 (1987)), cassette mutagenesis (Wells et al., Gene, 34:315 (1985)), restriction selection mutagenesis (Wells et al., Philos. Trans. R. Soc. London SerA, 317:415 (1986)) or other known techniques can be performed on the cloned DNA to produce the variant DNA.
[0214]Scanning amino acid analysis can also be employed to identify one or more amino acids along a contiguous sequence. Among the preferred scanning amino acids are relatively small, neutral amino acids. Such amino acids include alanine, glycine, serine, and cysteine. Alanine is typically a preferred scanning amino acid among this group because it eliminates the side-chain beyond the beta-carbon and is less likely to alter the main-chain conformation of the variant (Cunningham and Wells, Science, 244:1081-1085 (1989)). Alanine is also typically preferred because it is the most common amino acid. Further, it is frequently found in both buried and exposed positions (Creighton, The Proteins, (W.H. Freeman & Co., N.Y.); Chothia, J. Mol. Biol., 150:1 (1976)). If alanine substitution does not yield adequate amounts of variant, an isoteric amino acid can be used.
[0215]Covalent modifications: The stretches of consecutive amino acids may be covalently modified. One type of covalent modification includes reacting targeted amino acid residues with an organic derivatizing agent that is capable of reacting with selected side chains or the N- or C-terminal residues that are not involved in an -x-x- bond. Derivatization with bifunctional agents is useful, for instance, for crosslinking to a water-insoluble support matrix or surface for use in the method for purifying anti-sequence of interest antibodies, and vice-versa. Commonly used crosslinking agents include, e.g., 1,1-bis(diazoacetyl)-2-phenylethane, glutaraldehyde, N-hydroxysuccinimide esters, for example, esters with 4-azidosalicylic acid, homobifunctional imidoesters, including disuccinimidyl esters such as 3,3'-dithiobis(succinimidylpropionate), bifunctional maleimides such as bis-N-maleimido-1,8-octane and agents such as methyl-3-((p-azidophenyl)dithio)propioimidate.
[0216]Other modifications include deamidation of glutaminyl and asparaginyl residues to the corresponding glutamyl and aspartyl residues, respectively, hydroxylation of proline and lysine, phosphorylation of hydroxyl groups of seryl or threonyl residues, methylation of the α-amino groups of lysine, arginine, and histidine side chains (T. E. Creighton, Proteins: Structure and Molecular Properties, W.H. Freeman & Co., San Francisco, pp. 79-86 (1983)), acetylation of the N-terminal amine, and amidation of any C-terminal carboxyl group.
[0217]Another type of covalent modification comprises altering the native glycosylation pattern of the consecutive stretch of amino acids or of a polypeptide. "Altering the native glycosylation pattern" is intended for purposes herein to mean deleting one or more carbohydrate moieties found in amino acid sequences (either by removing the underlying glycosylation site or by deleting the glycosylation by chemical and/or enzymatic means), and/or adding one or more glycosylation sites that are not present in the native sequence. In addition, the phrase includes qualitative changes in the glycosylation of the native proteins, involving a change in the nature and proportions of the various carbohydrate moieties present.
[0218]Addition of glycosylation sites to the amino acid sequence may be accomplished by altering the amino acid sequence. The alteration may be made, for example, by the addition of, or substitution by, one or more serine or threonine residues to the native sequence (for O-linked glycosylation sites). The amino acid sequence may optionally be altered through changes at the DNA level, particularly by mutating the DNA encoding the amino acid sequence at preselected bases such that codons are generated that will translate into the desired amino acids.
[0219]Another means of increasing the number of carbohydrate moieties on the amino acid sequence is by chemical or enzymatic coupling of glycosides to the polypeptide. Such methods are described in the art, e.g., in WO 87/05330 published Sep. 11, 1987, and in Aplin and Wriston, CRC Crit. Rev. Biochem., pp. 259-306 (1981).
[0220]Removal of carbohydrate moieties present on the amino acid sequence may be accomplished chemically or enzymatically or by mutational substitution of codons encoding for amino acid residues that serve as targets for glycosylation. Chemical deglycosylation techniques are known in the art and described, for instance, by Hakimuddin, et al., Arch. Biochem. Biophys., 259:52 (1987) and by Edge et al., Anal. Biochem., 118:131 (1981). Enzymatic cleavage of carbohydrate moieties on polypeptides can be achieved by the use of a variety of endo- and exo-glycosidases as described by Thotakura et al., Meth. Enzymol., 138:350 (1987).
[0221]Another type of covalent modification comprises linking the amino acid sequence to one of a variety of nonproteinaceous polymers, e.g., polyethylene glycol (PEG), polypropylene glycol, or polyoxyalkylenes, in the manner set forth in U.S. Pat. No. 4,640,835; 4,496,689; 4,301,144; 4,670,417; 4,791,192 or 4,179,337, or to a tag polypeptide which provides an epitope to which an anti-tag antibody can selectively bind. Various tag polypeptides and their respective antibodies are well known in the art. Examples include poly-histidine (poly-his) or poly-histidine-glycine (poly-his-gly) tags; the flu HA tag polypeptide and its antibody 12CA5 (Field et al., Mol. Cell. Biol., 8:2159-2165 (1988)); the c-myc tag and the 8F9, 3C7, 6E10, G4, B7 and 9E10 antibodies thereto (Evan et al., Molecular and Cellular Biology, 5:3610-3616 (1985); and the Herpes Simplex virus glycoprotein D (gD) tag and its antibody (Paborsky et al., Protein Engineering, 3(6):547-553 (1990)). Other tag polypeptides include the Flag-peptide (Hopp et al., BioTechnology, 6:1204-1210 (1988)); the KT3 epitope peptide (Martin et al., Science, 255:192-194 (1992)); an alpha-tubulin epitope peptide (Skinner et al., J. Biol. Chem., 266:15163-15166 (1991)); and the T7 gene 10 protein peptide tag (Lutz-Freyermuth et al., Proc. Natl. Acad. Sci. USA, 87:6393-6397 (1990)).
Salts
[0222]Salts of the compounds disclosed herein are within the scope of the invention. As used herein, a "salt" is salt of the instant compounds which has been modified by making acid or base salts of the compounds.
Pharmaceuticals
[0223]The salt can be pharmaceutically acceptable. Examples of pharmaceutically acceptable salts include, but are not limited to, mineral or organic acid salts of basic residues such as amines; alkali or organic salts of acidic residues such as carboxcylic acids. The salts can be made using an organic or inorganic acid. Such acid salts are chlorides, bromides, sulfates, nitrates, phosphates, sulfonates, formates, tartrates, maleates, malates, citrates, benzoates, salicylates, ascorbates, and the like. Carboxylate salts are the alkaline earth metal salts, sodium, potassium or lithium.
[0224]Pharmaceutically acceptable salts of the compounds disclosed here can be prepared in any conventional manner, for example by treating a solution or suspension of the corresponding free base or acid with one chemical equivalent of a pharmaceutically acceptable acid or base. Conventional concentration or crystallization techniques can be employed to isolate the salts. Illustrative of suitable acids are acetic, lactic, succinic, maleic, tartaric, citric, gluconic, ascorbic, benzoic, cinnamic, fumaric, sulfuric, phosphoric, hydrochloric, hydrobromic, hydroiodic, sulfamic, sulfonic acids such as methanesulfonic, benzene sulfonic, p-toluenesulfonic, and related acids. Illustrative bases are sodium, potassium, and calcium.
[0225]The term "pharmaceutically acceptable carrier" is understood to include excipients, carriers or diluents. The particular carrier, diluent or excipient used will depend upon the means and purpose for which the active ingredient is being applied.
[0226]The compounds of this invention may be administered alone or in combination with one or more pharmaceutically acceptable carriers, in either single or multiple doses. Suitable pharmaceutical carriers include inert solid diluents or fillers, sterile aqueous solutions and various organic solvents. The pharmaceutical compositions disclosed herein can be readily administered in a variety of dosage forms such as injectable solutions, tablets, powders, lozenges, syrups, and the like. These pharmaceutical compositions can, if desired, contain additional ingredients such as flavorings, binders, excipients and the like. Additionally, lubricating agents such as magnesium stearate, sodium lauryl sulfate and talc may be used for tabletting purposes. Solid compositions of a similar type may also be employed as fillers in soft and hard filled gelatin capsules. Preferred materials for this include lactose or milk sugar and high molecular weight polyethylene glycols. When aqueous suspensions or elixirs are desired for oral administration, the essential active ingredient therein may be combined with various sweetening or flavoring agents, coloring matter or dyes and, if desired, emulsifying or suspending agents, together with diluents such as water, ethanol, propylene glycol, glycerin and combinations thereof.
[0227]For parenteral administration, solutions containing a compound of this invention or a pharmaceutically acceptable salt thereof in sterile aqueous solution may be employed. Such aqueous solutions should be suitably buffered if necessary and the liquid diluent first rendered isotonic with sufficient saline or glucose. These particular aqueous solutions are especially suitable for intravenous, intramuscular, subcutaneous and intraperitoneal administration. The sterile aqueous media employed are all readily available by standard techniques known to those skilled in the art.
[0228]The final pharmaceutical composition can be processed into a unit dosage form (e.g., powdered or lyophilized in a vial, a solution in a vial, tablet, capsule or sachet) and then packaged for distribution. The processing step will vary depending upon the particular unit dosage form. For example, a tablet is generally compressed under pressure into a desired shape and a capsule or sachet employs a simple fill operation. Those skilled in the art are well aware of the procedures used for manufacturing the various unit dosage forms.
[0229]The compositions of this invention may be in a variety of forms. These include, for example, liquid, semi-solid and solid dosage forms, such as liquid solutions (e.g., injectable and infusible solutions), dispersions or suspensions, tablets, pills, powders, liposomes and suppositories. The preferred form depends on the intended mode of administration and therapeutic application. Some compositions are in the form of injectable or infusible solutions. A mode of administration is parenteral (e.g., intravenous, subcutaneous, intraperitoneal, intramuscular). In an embodiment, the compound is administered by intravenous infusion or injection. In another embodiment, the compound is administered by intramuscular or subcutaneous injection.
[0230]Therapeutic compositions as contemplated herein typically must be sterile and stable under the conditions of manufacture and storage. The composition can be formulated as a solution, microemulsion, dispersion, liposome, or other ordered structure suitable to high drug concentration. Sterile injectable solutions can be prepared by incorporating the compound in the required amount in an appropriate solvent with one or a combination of ingredients as required, followed by filtered sterilization. Generally, dispersions are prepared by incorporating the active compound into a sterile vehicle that contains a basic dispersion medium and the required other ingredients from those enumerated above. In the case of sterile powders for the preparation of sterile injectable solutions, the preferred methods of preparation are vacuum drying and freeze-drying that yields a powder of the active ingredient plus any additional desired ingredient from a previously sterile-filtered solution thereof. The proper fluidity of a solution can be maintained, for example, by the use of a coating such as lecithin, by the maintenance of the required particle size in the case of dispersion and by the use of surfactants. Prolonged absorption of injectable compositions can be brought about by including in the composition an agent that delays absorption, for example, monostearate salts and gelatin.
[0231]In certain embodiments, the active compound may be prepared with a carrier that will protect the compound against rapid release, such as a controlled release formulation, including implants, transdermal patches, and microencapsulated delivery systems. Biodegradable, biocompatible polymers can be used, such as ethylene vinyl acetate, polyanhydrides, polyglycolic acid, collagen, polyorthoesters, and polylactic acid. Many methods for the preparation of such formulations are patented or generally known to those skilled in the art. See, e.g., Sustained and Controlled Release Drug Delivery Systems, J. R. Robinson, ed., Marcel Dekker, Inc., New York, 1978. In certain embodiments, the compounds of the invention may be orally administered, for example, with an inert diluent or an assimilable edible carrier. The compound (and other ingredients, if desired) may also be enclosed in a hard or soft shell gelatin capsule, compressed into tablets, or incorporated directly into the subject's diet. For oral therapeutic administration, the compounds may be incorporated with excipients and used in the form of ingestible tablets, buccal tablets, troches, capsules, elixirs, suspensions, syrups, wafers, and the like. In administering a compound of the invention by other than parenteral administration the compound may be coated with, or co-administered with, a material to prevent its inactivation.
[0232]Supplementary active compounds can also be incorporated into the compositions, for example a chemotherapeutic agent, an antineoplastic agent or an anti-tumor agent. IN addition, the compounds of the invention may be coformulated and/or coadministered with one or more additional therapeutic agents. These agents include, without limitation, antibodies that bind other targets (e.g., antibodies that bind one or more growth factors or cytokines, their cell surface receptors), binding proteins, antineoplastic agents, chemotherapeutic agents, anti-tumor agents, antisense oligonucleotides, growth factors. In one embodiment, a pharmaceutical composition of the compounds disclosed herein may comprise one or more additional therapeutic agent.
[0233]For therapeutic use, the compositions disclosed here can be administered in various manners, including soluble form by bolus injection, continuous infusion, sustained release from implants, oral ingestion, local injection (e.g. intracrdiac, intramuscular), systemic injection, or other suitable techniques well known in the pharmaceutical arts. Other methods of pharmaceutical administration include, but are not limited to oral, subcutaneously, transdermal, intravenous, intramuscular and parenteral methods of administration. Typically, a soluble composition will comprise a purified compound in conjunction with physiologically acceptable carriers, excipients or diluents. Such carriers will be nontoxic to recipients at the dosages and concentrations employed. The preparation of such compositions can entail combining a compound with buffers, antioxidants, carbohydrates including glucose, sucrose or dextrins, chelating agents such as EDTA, glutathione and other stabilizers and excipients. Neutral buffered saline or saline mixed with conspecific serum albumin are exemplary appropriate diluents. The product can be formulated as a lyophilizate using appropriate excipient solutions (e.g., sucrose) as diluents.
[0234]Other derivatives comprise the compounds/compositions of this invention covalently bonded to a nonproteinaceous polymer. The bonding to the polymer is generally conducted so as not to interfere with the preferred biological activity of the compound, e.g. the binding activity of the compound to a target. The nonproteinaceous polymer ordinarily is a hydrophilic synthetic polymer, i.e., a polymer not otherwise found in nature. However, polymers which exist in nature and are produced by recombinant or in vitro methods are useful, as are polymers which are isolated from nature. Hydrophilic polyvinyl polymers fall within the scope of this invention, e.g. polyvinylalcohol and polyvinylpyrrolidone. Particularly useful are polyalkylene ethers such as polyethylene glycol, polypropylene glycol, polyoxyethylene esters or methoxy polyethylene glycol; polyoxyalkylenes such as polyoxyethylene, polyoxypropylene, and block copolymers of polyoxyethylene and polyoxypropylene (Pluronics); polymethacrylates; carbomers; branched or unbranched polysaccharides which comprise the saccharide monomers D-mannose, D- and L-galactose, fucose, fructose, D-xylose, L-arabinose, D-glucuronic acid, sialic acid, D-galacturontc acid, D-mannuronic acid (e.g. polymannuronic acid, or alginic acid), D-glucosamine, D-galactosamine, D-glucose and neuraminic acid including homopolysaccharides and heteropolysaccharides such as lactose, amylopectin, starch, hydroxyethyl starch, amylose, dextran sulfate, dextran, dextrins, glycogen, or the polysaccharide subunit of acid mucopolysaccharides, e.g. hyaluronic acid; polymers of sugar alcohols such as polysorbitol and polymannitol; as well as heparin or heparon.
[0235]The pharmaceutical compositions of the invention may include a "therapeutically effective amount" or a "prophylactically effective amount" of a compound of the invention. A "therapeutically effective amount" refers to an amount effective, at dosages and for periods of time necessary, to achieve the desired therapeutic result. A therapeutically effective amount of the compound may vary according to factors such as the disease state, age, sex, and weight of the individual. A therapeutically effective amount is also one in which any toxic or detrimental effects of the compound are outweighed by the therapeutically beneficial effects. A "prophylactically effective amount" refers to an amount effective, at dosages and for periods of time necessary, to achieve the desired prophylactic result. Typically, since a prophylactic dose is used in subjects prior to or at an earlier stage of disease, the prophylactically effective amount will be less than the therapeutically effective amount.
Examples of Pharmaceutical Compositions
[0236]Non-limiting examples of such compositions and dosages are set forth as follows:
[0237]Compositions comprising a compound comprising a stretch of consecutive amino acids which comprises consecutive amino acids having the sequence of bevacizumab (e.g. Avastin) may comprise trehalose dihydrate, sodium phosphate (monobasic, monohydrate), sodium phosphate (dibasic, anhydrous), polysorbate 20, and Water for Injection, USP. The composition may also be lyophilized, to which the water can be added for reconstitution. In an embodiment the composition has a pH of 6.2 or about 6.2. In one embodiment the compound can be administered combination with a chemotherapeutic such as intravenous 5-fluorouracil for treatment of patients with metastatic carcinoma of the colon or rectum. In one embodiment the compound is administered in a dose of between 0.1 and 10 mg/kg given once every 14 days as an IV infusion. In a further embodiment the dose is 5 mg/kg given once every 14 days. In an embodiment the dose is between 1.0 and 2.0 mg/kg given once every 14 days. In an embodiment the dose is between 0.01 and 1.5 mg/kg given once every 14 days. In an embodiment the dose is between 0.001 and 10 mg/kg given once every 1-21 days.
[0238]Compositions comprising a compound comprising a stretch of consecutive amino acids which comprises consecutive amino acids having the sequence of trastuzumab (e.g. Herceptin) may comprise trehalose dihydrate, L-histidine HCl, L-histidine, and polysorbate 20, USP. This can be reconstituted with Bacteriostatic Water for Injection (BWFI), USP, or equivalents thereof, containing 1.1% benzyl alcohol as a preservative or equivalents thereof. In an embodiment the composition has a pH of about 6.0. The composition may also be lyophilized, to which the water can be added for reconstitution. In one embodiment the compound can be administered to subjects with metatastic breast cancer whose tumor overexpresses HER2 protein. In an embodiment the subject has received/is receiving chemotherapy. In another embodiment the compound is administered in combination with paclitaxel to subjects with metastatic breast cancer whose tumors overexpress the HER2 protein and who have not received chemotherapy for their metastatic disease. In one embodiment the compound is administered in an initial dose of between 0.1 and 10 mg/kg in a continuous 45-120 minute infusion IV infusion. In a further embodiment the compound is administered in a dose of 4 mg/kg in a 90 minute infusion. In an embodiment a weekly maintenance dose is administered to the subject at a dose of 2 mg/kg in a 30 minute infusion. In an embodiment the compound is administered in a dose of 0.5 to 1.5 mg/kg in a 90 minute infusion. In an embodiment a weekly maintenance dose is administered to the subject at a dose of 0.5-1.0 mg/kg in a 30 minute infusion. In an embodiment the compound is administered in a dose of 0.04 to 0.5 mg/kg in a 90 minute infusion. In an embodiment the compound is administered in a dose of between 0.001 to 10 mg/kg in a 90 minute infusion.
[0239]Compositions comprising a compound comprising a stretch of consecutive amino acids which comprises consecutive amino acids having the sequence of rituximab (e.g. Rituxin) may comprise sodium chloride, sodium citrate dihydrate, polysorbate 80, and Water for Injection (USP), or equivalents thereof. In an embodiment the pH of the composition is adjusted to 6.5. The composition may also be lyophilized, to which the water can be added for reconstitution. In one embodiment the compound is administered to a subject for the treatment of relapsed or refractory, low-grade or follicular, CD20-positive, B-cell non-Hodgkin's lymphoma. In one embodiment the compound is administered at a dose of 250-500 mg/m2 IV infusion once weekly for 4 or 8 doses. In a further embodiment the compound is administered at 375 mg/m2 IV infusion once weekly for 4 or 8 doses. In one embodiment the compound is administered at a dose of 150-250 mg/m2 IV infusion once weekly for 4 or 8 doses. In one embodiment the compound is administered at a dose of 1.5 to 5 mg/m2 IV infusion once weekly for 4 or 8 doses. In one embodiment the compound is administered at a dose of between 1.0 and 500 mg/m2 IV infusion once weekly for 4 or 8 doses
[0240]Compositions comprising a compound comprising a stretch of consecutive amino acids which comprises consecutive amino acids having the sequence of efalizumab (e.g. Raptiva) may comprise sucrose, L-histidine hydrochloride monohydrate, L-histidine and polysorbate 20. Such a composition may be diluted to an approporiate dosage form with sterile non-USP water, or Sterile Water for Injection, USP, or equivalents thereof. The composition may also be lyophilized, to which the water can be added for reconstitution. In one embodiment the compound is administered to a subject for the treatment of chronic moderate to severe plaque psoriasis. Such a subject may be a candidate for systemic therapy or phototherapy. In an embodiment the compound is administered in a single 0.1-1.1 mg/kg subcutaneous (SC) conditioning dose followed by weekly SC doses of 0.8-1.5 mg/kg (maximum single dose not to exceed a total of 250 mg). In a further embodiment the compound is administered in a single 0.7 mg/kg SC conditioning dose followed by weekly SC doses of 1 mg/kg (maximum single dose not to exceed a total of 200 mg). In an embodiment the compound is administered in a single 0.001-1.0 mg/kg subcutaneous (SC) conditioning dose followed by weekly SC doses of 0.008-0.015 mg/kg.
[0241]Compositions comprising a compound comprising a stretch of consecutive amino acids which comprises consecutive amino acids having the sequence of omalizumab (e.g. Xolair) may comprise sucrose, L-histidine hydrochloride monohydrate, L-histidine, and polysorbate 20. Such a composition may lyophilized. In an embodiment the composition is diluted to an appropriate dosage form with Sterile Water for Injection, USP, or equivalents thereof. In one embodiment the compound is administered to a subject for the treatment of moderate to severe persistent asthma. In another embodiment the compound is administered to a subject who has a positive skin test or in vitro reactivity to a perennial aeroallergen and whose symptoms are inadequately controlled with inhaled corticosteroids in order to decrease the incidence of asthma exacerbations. In an embodiment the compound is administered at a dose of 100 to 400 mg subcutaneously every 2 or 4 weeks. In a further embodiment the compound is administered at a dose of 150 to 375 mg SC every 2 or 4 weeks. In an embodiment the compound is administered at a dose of 25 to 150 mg subcutaneously every 2 or 4 weeks. In an embodiment the compound is administered at a dose of 1 to 4 mg subcutaneously every 2 or 4 weeks.
[0242]Compositions comprising a compound comprising a stretch of consecutive amino acids which comprises consecutive amino acids having the sequence of etanercept (e.g. Enbrel) may comprise mannitol, sucrose, and tromethamine. In an embodiment, the composition is in the form of a lyophilizate. In an embodiment, the composition is reconstituted with, for example, Sterile Bacteriostatic Water for Injection (BWFI), USP (containing 0.9% benzyl alcohol). In an embodiment the compound is administered to a subject for reducing signs and symptoms, inducing major clinical response, inhibiting the progression of structural damage, and improving physical function in subjects with moderately to severely active rheumatoid arthritis. The compound may be initiated in combination with methotrexate (MTX) or used alone. In an embodiment the compound is administered to a subject for reducing signs and symptoms of moderately to severely active polyarticular-course juvenile rheumatoid arthritis in subjects who have had an inadequate response to one or more DMARDs. In an embodiment the compound is administered to a subject for reducing signs and symptoms, inhibiting the progression of structural damage of active arthritis, and improving physical function in subjects with psoriatic arthritis. In an embodiment the compound is administered to a subject for reducing signs and symptoms in subjects with active ankylosing spondylitis. In an embodiment the compound is administered to a subject for the treatment of chronic moderate to severe plaque psoriasis. In an embodiment wherein the subject has rheumatoid arthritis, psoriatic arthritis, or ankylosing spondylitis the compound is administered at 25-75 mg per week given as one or more subcutaneous (SC) injections. In a further embodiment the compound is administered at 50 mg per week in a single SC injection. In an embodiment wherein the subject has plaque psoriasis the compound is administered at 25-75 mg twice weekly or 4 days apart for 3 months followed by a reduction to a maintenance dose of 25-75 mg per week. In a further embodiment the compound is administered at a dose of at 50 mg twice weekly or 4 days apart for 3 months followed by a reduction to a maintenance dose of 50 mg per week. In an embodiment the dose is between 2× and 10× less than the doses set forth herein. In an embodiment wherein the subject has active polyarticular-course JRA the compound may be administered at a dose of 0.2-1.2 mg/kg per week (up to a maximum of 75 mg per week). In a further embodiment the compound is administered at a dose of 0.8 mg/kg per week (up to a maximum of 50 mg per week). In some embodiments the dose is between 2× and 100× less than the doses set forth hereinabove.
[0243]Compositions comprising a compound comprising a stretch of consecutive amino acids which comprises consecutive amino acids having the sequence of infliximab (e.g. Remicade) may comprise sucrose, polysorbate 80, monobasic sodium phosphate, monohydrate, and dibasic sodium phosphate, dihydrate. Preservatives are not present in one embodiment. In an embodiment, the composition is in the form of a lyophilizate. In an embodiment, the composition is reconstituted with, for example, Water for Injection (BWFI), USP. In an embodiment the pH of the composition is 7.2 or is about 7.2. In one embodiment the compound is administered is administered to a subject with rheumatoid arthritis in a dose of 2-4 mg/kg given as an intravenous infusion followed with additional similar doses at 2 and 6 weeks after the first infusion then every 8 weeks thereafter. In a further embodiment the compound is administered in a dose of 3 mg/kg given as an intravenous infusion followed with additional similar doses at 2 and 6 weeks after the first infusion then every 8 weeks thereafter. In an embodiment the dose is adjusted up to 10 mg/kg or treating as often as every 4 weeks. In an embodiment the compound is administered in combination with methotrexate. In one embodiment the compound is administered is administered to a subject with Crohn's disease or fistulizing Crohn's disease at dose of 2-7 mg/kg given as an induction regimen at 0, 2 and 6 weeks followed by a maintenance regimen of 4-6 mg/kg every 8 weeks thereafter for the treatment of moderately to severely active Crohn's disease or fistulizing disease. In a further embodiment the compound is administered at a dose of 5 mg/kg given as an induction regimen at 0, 2 and 6 weeks followed by a maintenance regimen of 5 mg/kg every 8 weeks thereafter for the treatment of moderately to severely active Crohn's disease or fistulizing disease. In an embodiment the dose is adjusted up to 10 mg/kg. In one embodiment the compound is administered to a subject with ankylosing spondylitis at a dose of 2-7 mg/kg given as an intravenous infusion followed with additional similar doses at 2 and 6 weeks after the first infusion, then every 6 weeks thereafter. In a further embodiment the compound is administered at a dose of 5 mg/kg given as an intravenous infusion followed with additional similar doses at 2 and 6 weeks after the first infusion, then every 6 weeks thereafter. In one embodiment the compound is administered to a subject with psoriatic arthritis at a dose of 2-7 mg/kg given as an intravenous infusion followed with additional similar doses at 2 and 6 weeks after the first infusion then every 8 weeks thereafter. In a further embodiment the compound is administered at a dose of 5 mg/kg given as an intravenous infusion followed with additional similar doses at 2 and 6 weeks after the first infusion then every 8 weeks thereafter. In an embodiment the compound is administered with methotrexate. In one embodiment the compound is administered to a subject with ulcerative colitis at a dose of 2-7 mg/kg given as an induction regimen at 0, 2 and 6 weeks followed by a maintenance regimen of 2-7 mg/kg every 8 weeks thereafter for the treatment of moderately to severely active ulcerative colitis. In a further embodiment the compound is administered to a subject with ulcerative colitis at a dose of 5 mg/kg given as an induction regimen at 0, 2 and 6 weeks followed by a maintenance regimen of 5 mg/kg every 8 weeks thereafter. In some embodiments the dose is between 2× and 100× less than the doses set forth hereinabove for treating the indivisual diseases.
[0244]Compositions comprising a compound comprising a stretch of consecutive amino acids which comprises consecutive amino acids having the sequence of cetuximab (e.g. Erbitux) may comprise a preservative-free composition including sodium chloride, sodium phosphate dibasic heptahydrate, sodium phosphate monobasic monohydrate. In an embodiment, the composition is in the form of a lyophilizate. In an embodiment, the composition is reconstituted with, for example, Water for Injection, USP. In an embodiment the pH of the composition is in the range of about 7.0 to about 7.4. In one embodiment the compound is administered to a subject for the treatment of EGFR-expressing, metastatic colorectal carcinoma. In another embodiment the compound is used in combination with irinotecan for the treatment in patients who are refractory to irinotecan-based chemotherapy. In an embodiment the compound is administered at a dose of 300-500 mg/m2 as an initial loading dose (first infusion) administered as a 120-minute IV infusion (maximum infusion rate 5 mL/min). In a further embodiment the compound is administered at a dose of 400 mg/m2 as an initial loading dose (first infusion) administered as a 120-minute IV infusion (maximum infusion rate 5 mL/min). In an embodiment the weekly maintenance dose (all other infusions) is 200-300 mg/m2 infused over 60 minutes (maximum infusion rate 5 mL/min). In a further embodiment the weekly maintenance dose (all other infusions) is 250 mg/m2 infused over 60 minutes (maximum infusion rate 5 mL/min). In some embodiments the dose is between 2× and 100× less than the doses set forth hereinabove.
[0245]Compositions comprising a compound comprising a stretch of consecutive amino acids which comprises consecutive amino acids having the sequence of abciximab (e.g. Reopro) may comprise, a preservative-free composition including sodium phosphate, sodium chloride and polysorbate 80. In an embodiment, the composition is in the form of a lyophilizate. In an embodiment, the composition is diluted in, or reconstituted with, for example, Water for Injection, USP. In an embodiment the pH of the composition is 7.2 or is about 7.2. In one embodiment, the composition is used as an adjunct to percutaneous coronary intervention (PCI). In such a use the compositions may be administered as an intravenous bolus of 0.15 to 0.35 mg/kg at 10-60 minutes before the start of PCI. In a further embodiment the dose is 0.2 mg/kg. In an embodiment, the bolus is followed by continuous intravenous infusion of 0.1 to 0.15 g/kg/min for up to 12 hours. In a further embodiment the dose is 0.125 g/kg/min for up to 12 hours. In one embodiment, the composition is used as an adjunct to percutaneous coronary intervention (PCI) in subjects suffering from unstable angina. In one embodiment of such a use the compositions may be administered as an intravenous bolus at 0.1 to 0.4 mg/kg before the start of PCI followed by continuous intravenous infusion of 5-15 g/min for up to 24 hours, concluding 1 hour after the PCI. In a further embodiment the composition is administered as an intravenous bolus at 0.25 mg/kg before the start of PCI followed by continuous intravenous infusion of 10 g/min for up to 24 hours, concluding 1 hour after the PCI. In some embodiments the dose is between 2× and 100× less than the doses set forth hereinabove.
[0246]In each of the embodiments of the compositions described herein, the compositions, when in the form of a lyophilizate, may be reconstituted with, for example, sterile aqueous solutions, sterile water, Sterile Water for Injections (USP), Sterile Bacteriostatic Water for Injections (USP), and equivalents thereof known to those skilled in the art.
[0247]It is understood that in administration of any of the instant compounds, the compound may be administered in isolation, in a carrier, as part of a pharmaceutical composition, or in any appropriate vehicle.
Dosage
[0248]It is understood that where a dosage range is stated herein, e.g. 1-10 mg/kg per week, the invention disclosed herein also contemplates each integer dose, and tenth thereof, between the upper and lower limits. In the case of the example given, therefore, the invention contemplates 1.0, 1.1, 1.2, 1.3, 1.4, 1.5, 1.6, 1.7, 1.8, 1.9, 2.0, 2.1, 2.2, 2.3, 2.4 etc. mg/kg up to 10 mg/kg.
[0249]In embodiments, the compounds of the present invention can be administered as a single dose or may be administered as multiple doses.
[0250]In general, the daily dosage for treating a disorder or condition according to the methods described above will generally range from about 0.01 to about 10.0 mg/kg body weight of the subject to be treated.
[0251]Variations based on the aforementioned dosage ranges may be made by a physician of ordinary skill taking into account known considerations such as the weight, age, and condition of the person being treated, the severity of the affliction, and the particular route of administration chosen.
[0252]It is also expected that the compounds disclosed will effect cooperative binding with attendant consequences on effective dosages required.
Kits
[0253]Another aspect of the present invention provides kits comprising the compounds disclosed herein and the pharmaceutical compositions comprising these compounds. A kit may include, in addition to the compound or pharmaceutical composition, diagnostic or therapeutic agents. A kit may also include instructions for use in a diagnostic or therapeutic method. In a diagnostic embodiment, the kit includes the compound or a pharmaceutical composition thereof and a diagnostic agent. In a therapeutic embodiment, the kit includes the antibody or a pharmaceutical composition thereof and one or more therapeutic agents, such as an additional antineoplastic agent, anti-tumor agent or chemotherapeutic agent.
Subjects
[0254]In some embodiments the subject is a human. In an embodiment the subject is 18 years or older. In another embodiment the subject is less than 18 years old.
[0255]All combinations of the various elements disclosed herein are within the scope of the invention.
[0256]This invention will be better understood by reference to the Experimental Details which follow, but those skilled in the art will readily appreciate that the specific examples detailed are only illustrative of the invention as described more fully in the claims which follow thereafter.
SymmetroAdhesins
[0257]This patent specification describes genetic devices which are devices comprising functional stretches of consecutive amino acids joined in a novel manner. Until now, proteins have been genetically engineered to create new functions by virtue of their adopting a novel fixed structure or conformation. In contrast to such previously genetically engineered proteins, the genetic devices disclosed herein comprise two or more distinguishable protein domains that are connected by novel chemical bonds in a manner permitting relative motion to occur between the domains. The relative motion between protein domains constitutes the moving parts of the genetic devices, and the useful work which they are able to thereby carry out. Input energy for this work is provided by the kinetic and rotational energies intrinsic to the protein domains themselves, as well as the energy of mechanical interactions between the protein domains and solvent molecules, and the like.
[0258]This disclosure describes "symmetroadhesins", a specific class of genetic devices. Symmetroadhesins comprise two or more chemically-bonded adhesins (independently folding polypeptide binding domains or stretches of consecutive amino acids). These chemically-bonded adhesins are capable of relative motion with respect to one another, resulting in the formation of two or more symmetrically-oriented binding domains useful in binding dimeric ligands, trimeric ligands, tetrameric ligands, and the like, with greatly increased affinity. Symmetroadhesin-Fc hybrid proteins are a particularly useful embodiment of the genetic devices.
[0259]The importance of symmetry in the effectiveness of therapeutic proteins is due to the fact that most protein disease targets themselves display such a higher-order symmetric structure. For example, if the disease target consists of two proteins it is said to be a dimer, if it consists of three proteins it is said to be a trimer, and so forth. If a protein therapeutic binds to the protein disease target only on a one-at-a-time basis, then the strength of the binding is typically on the order of the nano-molar range. At this ordinary level of binding, which is typical of present-day therapeutic proteins, it becomes necessary to flood the body with an enormous and wasteful excess of the therapeutic protein relative to the protein disease target (one million-to-one basis).
[0260]Each symmetroadhesin is engineered using a simple set of rules that makes proteins into more useful therapeutics by making them more symmetric. For a symmetroadhesin designed to bind to the protein disease target on a two-at-a-time basis, the strength of binding will be on the order of the nano H nano-molar range. Cooperative binding permits such an extraordinary level of binding. At this extraordinary level of binding, it is only necessary to administer much less of the therapeutic protein relative to the protein disease target (one to one basis).
Peptide Bond Chemistry
[0261]All proteins consist of one or more polypeptide chains, stretches of consecutive amino acids, each of which is joined to the preceding amino acid by a peptide bond. Due to resonance between the single bond form (--C--N--) and double bond form (--C═N--) which occur in amides, the peptide bond has a significant degree of double bond character (--C═N--C.sub.α--C═N--). As a consequence, peptide bonds in proteins are approximately planar. Although the adjoining N--C.sub.α and C.sub.α--C bonds are relatively free to rotate, the rigidly of the peptide bond reduces the degrees of freedom of the folded polypeptide chain to the point where it behaves like a single, static object.
[0262]Consider a first stretch of N consecutive amino acids and a second stretch of P consecutive amino acids, AA1-[ ]-AAn and AA1-[ ]-AAp, with the peptide bonding shown in 1:
C.sub.α(1)--C═N--[ ]--C═N--C.sub.α(N) & C.sub.α(1)--C═N--[ ]--C═N--C.sub.α(P) 1
[0263]Joining of the first stretch of consecutive amino acids at its N-terminus by a new peptide bond to the C-terminus of the second stretch of consecutive amino acids will form a chimeric polypeptide, AA1-[ ]-AAn-AAn+1-[ ]-AAn+p, with the peptide bonding shown in 2:
C.sub.α(1)--C═N--[ ]-C═N--C.sub.α(N)--C═N--C.sub.α(N+1)--C═N--[ ]--C═N--C.sub.α(N+P) 2
[0264]The chimeric polypeptide, like either of its progenitors, is a single stretch of consecutive amino acids, each of which is also joined to the preceding amino acid by a peptide bond. The folded chimeric polypeptide chain will thus also generally behave like a single, static object.
[0265]This invention provides for novel protein-like molecules, herein termed genetic devices, which like mechanical devices are dynamic objects with two or more moving parts that are interconnected in a manner to permit relative movement. Each part, or domain, in a genetic device is a stretch of consecutive amino acids, and each interconnection is made by a non-peptide bond joining a predefined end of each stretch of consecutive amino acids. Preferably, the domains in a genetic device are binding domains. Three distinct types of genetic devices are distinguished by the topology of their interconnections as shown in 3, 4, and 5:
C.sub.α(1)--C═N--[ ]--C═N--C.sub.α(N)--Xc--Xn--C.sub.α(N+1)--C.dbd- .N--[ ]--C═N--C.sub.α(N+P) 3
C.sub.α(1)--C═N--[ ]--C═N--C.sub.α(N)--Xc--Xc--C.sub.α(P)--N═C- --[ ]--N═C--C.sub.α(1) 4
C.sub.α(N)--N═C--[ ]N═C--C.sub.α(1)--Xn--Xn--C.sub.α(1)--C═N--- [ ]--C═N--C.sub.α(P) 5
[0266]Genetic device with two identical binding domains that are connected at distinct termini (6) have an asymmetric configuration,
C.sub.α(1)--C═N--[ ]--C═N--C.sub.α(N)--Xc--Xn--C.sub.α(N+1)--C.dbd- .N--[ ]--C═N--C.sub.α(2N) 6
[0267]Genetic device with two identical binding domains that are connected at identical termini (7 and 8) have a configuration with point symmetry,
C.sub.α(1)--C═N--[ ]--C═N--C.sub.α(N)--Xc--Xc--C.sub.α(N)--N═C- --[ ]--N═C--C.sub.α(1) 7
C.sub.α(N)--N═C--[ ]--N═C--C.sub.α(1)--Xn--Xn--C.sub.α(1)--C═N- --[ ]--C═N--C.sub.α(N) 8
[0268]The genetic devices shown in 7 and 8 are herein termed hemi-symmetroadhesins. Although hemi-symmetroadhesins are have structures with point symmetry they are not capable of rotating both their binding domains independently, and thus, they are generally not able to bind symmetrically to more than one binding site in a symmetric target.
[0269]This invention also provides protein-like molecules capable of binding symmetrically to two or more binding sites in a symmetric target (i.e., cooperatively). Genetic devices that are capable of binding symmetric targets symmetrically are termed herein symmetroadhesins. Like a human body grasping a oversized, yet symmetric object (e.g., a medicine ball) with two hands instead of one, the ability of symmetroadhesins to bind symmetric targets is generally much greater than proteins.
Symmetroadhesin Subtypes
[0270]Tables 2-11 set forth various non-limiting embodiments of different symmetroadhesins. For example, Table 2, describing CD4-symmetroadhesins, shows in the top line the configuration of a CD4 hemi-symmetroadhesin, i.e. a CD4 domain with a C-terminal X-terminus, e.g. a stretch of consecutive amino acids which is a CD4 domain with a C-terminal cysteine or selenosyteine residue bonded through a non-peptide link (for example a cysteine-cysteine disulfide bond or a selenocysteine-selenocysteine diselenide bond) to a second stretch of consecutive amino acids which is a CD4 domain with a C-terminal cysteine or selenosyteine residue bonded through a non-peptide link, generically described as [CD4-Xc-Xc-CD4]. A dimer of the CD4 hemi-symmetroadhesin and a Fc hemisymmetroadhesin to form an immuno-symmetroadhesin is set forth in the second row of Table 2, for example, described as [CD4-Xc-Sn-Fc]2. In each of the tables Xc represents a C-terminal X-terminus; Xn a N-terminal X-terminus, Sn a N-terminal cysteine residue
TABLE-US-00002 TABLE 2 CD4 Symmetroadhesins Stretches of Consecutive Amino Acids Class Symmetroadhesin Configuration CD4-Xc hemi- [CD4-Xc-Xc-CD4] CD4-Xc + Sn-Fc immuno [CD4-Xc-Sn-Fc]2 CD4-Xc + Xn-Fc immuno [CD4-Xc-Xn-Fc]2 CD4-Xc + Fc-Xc immuno [Fc-Xc-Xc-CD4]2 CD4-Xc + Sn-Fc-Xc bi- [CD4-Xc-Sn-Fc-Xc-Xc-CD4]2 CD4-Xc + Xn-Fc-Xc bi- [CD4-Xc-Xn-Fc-Xc-Xc-CD4]2
TABLE-US-00003 TABLE 3 TNR Hemi-Symmetroadhesins Stretches of Consecutive Amino Acids # Symmetroadhesin Configurations TNR1-Xc 1 [TNR1-Xc-Xc-TNR1] TNR2-Xc 1 [TNR2-Xc-Xc-TNR2] TNRFab-Xc 1 [TNRFab-Xc-Xc-TNRFab] TNR1-Xc + TNR2-Xc 3 [(TNR1/2)-Xc-Xc-(TNR1/2)] TNR1-Xc + TNRFab-Xc 3 [(TNR1/Fab)-Xc-Xc-(TNR1/Fab)] TNR2-Xc + TNRFab-Xc 3 [(TNR2/Fab)-Xc-Xc-(TNR2/Fab)] TNR1-Xc + TNR2-Xc + 6 [(TNR1/2/Fab)-Xc-Xc-(TNR1/2/Fab)] TNRFab-Xc
TABLE-US-00004 TABLE 4 TNR ImmunoSymmetroadhesins Stretches of Consecutive Amino Acids # Symmetroadhesin Configurations TNR1-Xc + Sn-Fc 1 [TNR1-Xc-Sn-Fc]2 TNR1-Xc + Xn-Fc 1 [TNR1-Xc-Xn-Fc]2 TNR1-Xc + Fc-Xc 1 [Fc-Xc-Xc-TNR1]2 TNR2-Xc + Sn-Fc 1 [TNR2-Xc-Sn-Fc]2 TNR2-Xc + Xn-Fc 1 [TNR2-Xc-Xn-Fc]2 TNR2-Xc + Fc-Xc 1 [Fc-Xc-Xc-TNR2]2 TNRFab-Xc + Sn-Fc 1 [TNRFab-Xc-Sn-Fc]2 TNRFab-Xc + Xn-Fc 1 [TNRFab-Xc-Xn-Fc]2 TNRFab-Xc + Fc-Xc 1 [Fc-Xc-Xc-TNRFab]2 TNR1-Xc + TNR2-Xc + Sn-Fc 3 [(TNR1/2)-Xc-Sn-Fc]2 TNR1-Xc + TNR2-Xc + Xn-Fc 3 [(TNR1/2)-Xc-Xn-Fc]2 TNR1-Xc + TNR2-Xc + Fc-Xc 3 [Fc-Xc-Xc-(TNR1/2)]2 TNR1-Xc + TNRFab-Xc + Sn-Fc 3 [(TNR1/Fab)-Xc-Sn-Fc]2 TNR1-Xc + TNRFab-Xc + Xn-Fc 3 [(TNR1/Fab)-Xc-Xn-Fc]2 TNR1-Xc + TNRFab-Xc + Fc-Xc 3 [Fc-Xc-Xc-(TNR1/Fab)]2 TNR2-Xc + TNRFab-Xc + Sn-Fc 3 [(TNR2/Fab)-Xc-Sn-Fc]2 TNR2-Xc + TNRFab-Xc + Xn-Fc 3 [(TNR2/Fab)-Xc-Xn-Fc]2 TNR2-Xc + TNRFab-Xc + Fc-Xc 3 [Fc-Xc-Xc-(TNR2/Fab)]2 TNR1-Xc + TNR2-Xc + 6 [(TNR1/2/Fab)-Xc-Sn-Fc]2 TNRFab-Xc + Sn-Fc TNR1-Xc + TNR2-Xc + 6 [(TNR1/2/Fab)-Xc-Xn-Fc]2 TNRFab-Xc + Xn-Fc TNR1-Xc + TNR2-Xc + 6 [Fc-Xc-Xc-(TNR1/2/Fab)]2 TNRFab-Xc + Fc-Xc
TABLE-US-00005 TABLE 5 TNR Bi-Symmetroadhesins Stretches of Consecutive Amino Acids # Symmetroadhesin Configurations TNR1-Xc + Sn-Fc-Xc 1 [TNR1-Xc-Sn-Fc-Xc-Xc-TNR1]2 TNR1-Xc + Xn-Fc-Xc 1 [TNR1-Xc-Xn-Fc-Xc-Xc-TNR1]2 TNR2-Xc + Sn-Fc-Xc 1 [TNR2-Xc-Sn-Fc-Xc-Xc-TNR2]2 TNR2-Xc + Xn-Fc-Xc 1 [TNR2-Xc-Xn-Fc-Xc-Xc-TNR2]2 TNRFab-Xc + Sn-Fc-Xc 1 [TNRFab-Xc-Sn-Fc-Xc-Xc-TNRFab]2 TNRFab-Xc + Xn-Fc-Xc 1 [TNRFab-Xc-Xn-Fc-Xc-Xc-TNRFab]2 TNR1-Xc + TNR2-Xc + Sn-Fc-Xc 12 [(TNR1/2)-Xc-Sn-Fc-Xc-Xc-(TNR1/2)]2 TNR1-Xc + TNR2-Xc + Xn-Fc-Xc 12 [(TNR1/2)-Xc-Xn-Fc-Xc-Xc-(TNR1/2)]2 TNR1-Xc + TNRFab-Xc + Sn-Fc-Xc 12 [(TNR1/Fab)-Xc-Sn-Fc-Xc-Xc-(TNR1/Fab)]2 TNR1-Xc + TNRFab-Xc + Xn-Fc-Xc 12 [(TNR1/Fab)-Xc-Xn-Fc-Xc-Xc-(TNR1/Fab)]2 TNR2-Xc + TNRFab-Xc + Sn-Fc-Xc 12 [(TNR2/Fab)-Xc-Sn-Fc-Xc-Xc-(TNR2/Fab)]2 TNR2-Xc + TNRFab-Xc + Xn-Fc-Xc 12 [(TNR2/Fab)-Xc-Xn-Fc-Xc-Xc-(TNR2/Fab)]2 TNR1-Xc + TNR2-Xc + TNRFab-Xc + Sn-Fc-Xc 63 [(TNR1/2/Fab)-Xc-Xn-Fc-Xc-Xc-(TNR1/2/Fab)].- sub.2 TNR1-Xc + TNR2-Xc + TNRFab-Xc + Sn-Fc-Xc 63 [(TNR1/2/Fab)-Xc-Xn-Fc-Xc-Xc-(TNR1/2/Fab)].- sub.2
TABLE-US-00006 TABLE 6 VGFR Hemi-Symmetroadhesins Stretches of Consecutive Amino Acids # Symmetroadhesin Configurations VGFR1-Xc 1 [VGFR1-Xc-Xc-VGFR1] VGFR2-Xc 1 [VGFR2-Xc-Xc-VGFR2] VGFR3-Xc 1 [VGFR3-Xc-Xc-VGFR3] VGFR1-Xc + VGFR2-Xc 3 [(VGFR1/2)-Xc-Xc-(VGFR1/2)] VGFR1-Xc + VGFR3-Xc 3 [(VGFR1/3)-Xc-Xc-(VGFR1/3)] VGFR2-Xc + VGFR3-Xc 3 [(VGFR2/3)-Xc-Xc-(VGFR2/3)] VGFR1-Xc + VGFR2-Xc + 6 [(VGFR1/2/3)-Xc-Xc-(VGFR1/2/3)] VGFR3-Xc
TABLE-US-00007 TABLE 7 VGFR ImmunoSymmetroadhesins Stretches of Consecutive Amino Acids # Symmetroadhesin Configurations VGFR1-Xc + Sn-Fc 1 [VGFR1-Xc-Sn-Fc]2 VGFR1-Xc + Xn-Fc 1 [VGFR1-Xc-Xn-Fc]2 VGFR1-Xc + Fc-Xc 1 [Fc-Xc-Xc-VGFR1]2 VGFR2-Xc + Sn-Fc 1 [VGFR2-Xc-Sn-Fc]2 VGFR2-Xc + Xn-Fc 1 [VGFR2-Xc-Xn-Fc]2 VGFR2-Xc + Fc-Xc 1 [Fc-Xc-Xc-VGFR2]2 VGFR3-Xc + Sn-Fc 1 [VGFR3-Xc-Sn-Fc]2 VGFR3-Xc + Xn-Fc 1 [VGFR3-Xc-Xn-Fc]2 VGFR3-Xc + Fc-Xc 1 [Fc-Xc-Xc-VGFR3]2 VGFR1-Xc + VGFR2-Xc + Sn-Fc 3 [(VGFR1/2)-Xc-Sn-Fc]2 VGFR1-Xc + VGFR2-Xc + Xn-Fc 3 [(VGFR1/2)-Xc-Xn-Fc]2 VGFR1-Xc + VGFR2-Xc + Fc-Xc 3 [Fc-Xc-Xc-(VGFR1/2)]2 VGFR1-Xc + VGFR3-Xc + Sn-Fc 3 [(VGFR1/3)-Xc-Sn-Fc]2 VGFR1-Xc + VGFR3-Xc + Xn-Fc 3 [(VGFR1/3)-Xc-Xn-Fc]2 VGFR1-Xc + VGFR3-Xc + Fc-Xc 3 [Fc-Xc-Xc-(VGFR1/3)]2 VGFR2-Xc + VGFR3-Xc + Sn-Fc 3 [(VGFR2/3)-Xc-Sn-Fc]2 VGFR2-Xc + VGFR3-Xc + Xn-Fc 3 [(VGFR2/3)-Xc-Xn-Fc]2 VGFR2-Xc + VGFR3-Xc + Fc-Xc 3 [Fc-Xc-Xc-(VGFR2/3)]2 VGFR1-Xc + VGFR2-Xc + VGFR3-Xc + Sn-Fc 6 [(VGFR1/2/3)-Xc-Sn-Fc]2 VGFR1-Xc + VGFR2-Xc + VGFR3-Xc + Xn-Fc 6 [(VGFR1/2/3)-Xc-Xn-Fc]2 VGFR1-Xc + VGFR2-Xc + VGFR3-Xc + Fc-Xc 6 [Fc-Xc-Xc-(VGFR1/2/3)]2
TABLE-US-00008 TABLE 8 VGFR Bi-Symmetroadhesins Stretches of Consecutive Amino Acids # Symmetroadhesin Configurations VGFR1-Xc + Sn-Fc-Xc 1 [VGFR1-Xc-Sn-Fc-Xc-Xc-VGFR1]2 VGFR1-Xc + Xn-Fc-Xc 1 [VGFR1-Xc-Xn-Fc-Xc-Xc-VGFR1]2 VGFR2-Xc + Sn-Fc-Xc 1 [VGFR2-Xc-Sn-Fc-Xc-Xc-VGFR2]2 VGFR2-Xc + Xn-Fc-Xc 1 [VGFR2-Xc-Xn-Fc-Xc-Xc-VGFR2]2 VGFR3-Xc + Sn-Fc-Xc 1 [VGFR3-Xc-Sn-Fc-Xc-Xc-VGFR3]2 VGFR3-Xc + Xn-Fc-Xc 1 [VGFR3-Xc-Xn-Fc-Xc-Xc-VGFR3]2 VGFR1-Xc + VGFR2-Xc + Sn-Fc-Xc 12 [(VGFR1/2)-Xc-Sn-Fc-Xc-Xc-(VGFR1/2)]2 VGFR1-Xc + VGFR2-Xc + Xn-Fc-Xc 12 [(VGFR1/2)-Xc-Xn-Fc-Xc-Xc-(VGFR1/2)]2 VGFR1-Xc + VGFR3-Xc + Sn-Fc-Xc 12 [(VGFR1/3)-Xc-Sn-Fc-Xc-Xc-(VGFR1/3)]2 VGFR1-Xc + VGFR3-Xc + Xn-Fc-Xc 12 [(VGFR1/3)-Xc-Xn-Fc-Xc-Xc-(VGFR1/3)]2 VGFR2-Xc + VGFR3-Xc + Sn-Fc-Xc 12 [(VGFR2/3)-Xc-Sn-Fc-Xc-Xc-(VGFR2/3)]2 VGFR2-Xc + VGFR3-Xc + Xn-Fc-Xc 12 [(VGFR2/3)-Xc-Xn-Fc-Xc-Xc-(VGFR2/3)]2 VGFR1-Xc + VGFR2-Xc + VGFR3-Xc + Sn-Fc-Xc 63 [(VGFR1/2/3)-Xc-Xn-Fc-Xc-Xc-(VGFR1/2/3)].su- b.2 VGFR1-Xc + VGFR2-Xc + VGFR3-Xc + Sn-Fc-Xc 63 [(VGFR1/2/3)-Xc-Xn-Fc-Xc-Xc-(VGFR1/2/3)].su- b.2
TABLE-US-00009 TABLE 9 ErbB Hemi-Symmetroadhesins Stretches of Consecutive Amino Acids # Symmetroadhesin Configurations ErbB1-Xc 1 [ErbB1-Xc-Xc-ErbB1] ErbB2-Xc 1 [ErbB2-Xc-Xc-ErbB2] ErbB3-Xc 1 [ErbB3-Xc-Xc-ErbB3] ErbB4-Xc 1 [ErbB4-Xc-Xc-ErbB4] ErbB1-Xc + ErbB2-Xc 3 [(ErbB1/2)-Xc-Xc-(ErbB1/2)] ErbB1-Xc + ErbB3-Xc 3 [(ErbB1/3)-Xc-Xc-(ErbB1/3)] ErbB1-Xc + ErbB4-Xc 3 [(ErbB1/4)-Xc-Xc-(ErbB1/4)] ErbB2-Xc + ErbB3-Xc 3 [(ErbB2/3)-Xc-Xc-(ErbB2/3)] ErbB2-Xc + ErbB4-Xc 3 [(ErbB2/4)-Xc-Xc-(ErbB2/4)] ErbB3-Xc + ErbB4-Xc 3 [(ErbB3/4)-Xc-Xc-(ErbB3/4)] ErbB1-Xc + ErbB2-Xc + ErbB3-Xc 6 [(ErbB1/2/3)-Xc-Xc-(ErbB1/2/3)] ErbB1-Xc + ErbB2-Xc + ErbB4-Xc 6 [(ErbB1/2/4)-Xc-Xc-(ErbB1/2/4)] ErbB1-Xc + ErbB3-Xc + ErbB4-Xc 6 [(ErbB1/3/4)-Xc-Xc-(ErbB1/3/4)] ErbB2-Xc + ErbB3-Xc + ErbB4-Xc 6 [(ErbB2/3/4)-Xc-Xc-(ErbB2/3/4)] ErbB1-Xc + ErbB2-Xc + ErbB3-Xc + ErbB4-Xc 10 [(ErbB1/2/3/4)-Xc-Xc-(ErbB1/2/3/4)]
TABLE-US-00010 TABLE 10 ErbB ImmunoSymmetroadhesins Stretches of Consecutive Amino Acids # Symmetroadhesin Configurations ErbB1-Xc + Xn-Fc 1 [ErbB1-Xc-Xn-Fc]2 ErbB1-Xc + Fc-Xc 1 [Fc-Xc-Xc-ErbB1]2 ErbB2-Xc + Xn-Fc 1 [ErbB2-Xc-Xn-Fc]2 ErbB2-Xc + Fc-Xc 1 [Fc-Xc-Xc-ErbB2]2 ErbB3-Xc + Xn-Fc 1 [ErbB3-Xc-Xn-Fc]2 ErbB3-Xc + Fc-Xc 1 [Fc-Xc-Xc-ErbB3]2 ErbB4-Xc + Xn-Fc 1 [ErbB4-Xc-Xn-Fc]2 ErbB4-Xc + Fc-Xc 1 [Fc-Xc-Xc-ErbB4]2 ErbB1-Xc + ErbB2-Xc + Xn-Fc 3 [(ErbB1/2)-Xc-Xn-Fc]2 ErbB1-Xc + ErbB2-Xc + Fc-Xc 3 [Fc-Xc-Xc-(ErbB1/2)]2 ErbB1-Xc + ErbB3-Xc + Xn-Fc 3 [(ErbB1/3)-Xc-Xn-Fc]2 ErbB1-Xc + ErbB3-Xc + Fc-Xc 3 [Fc-Xc-Xc-(ErbB1/3)]2 ErbB1-Xc + ErbB4-Xc + Xn-Fc 3 [(ErbB1/4)-Xc-Xn-Fc]2 ErbB1-Xc + ErbB4-Xc + Fc-Xc 3 [Fc-Xc-Xc-(ErbB1/4)]2 ErbB2-Xc + ErbB3-Xc + Xn-Fc 3 [(ErbB2/3)-Xc-Xn-Fc]2 ErbB2-Xc + ErbB3-Xc + Fc-Xc 3 [Fc-Xc-Xc-(ErbB2/3)]2 ErbB2-Xc + ErbB4-Xc + Xn-Fc 3 [(ErbB2/4)-Xc-Xn-Fc]2 ErbB2-Xc + ErbB4-Xc + Fc-Xc 3 [Fc-Xc-Xc-(ErbB2/4)]2 ErbB3-Xc + ErbB4-Xc + Xn-Fc 3 [(ErbB3/4)-Xc-Xn-Fc]2 ErbB3-Xc + ErbB4-Xc + Fc-Xc 3 [Fc-Xc-Xc-(ErbB3/4)]2 ErbB1-Xc + ErbB2-Xc + ErbB3-Xc + Xn-Fc 6 [(ErbB1/2/3)-Xc-Xn-Fc]2 ErbB1-Xc + ErbB2-Xc + ErbB3-Xc + Fc-Xc 6 [Fc-Xc-Xc-(ErbB1/2/3)]2 ErbB1-Xc + ErbB2-Xc + ErbB4-Xc + Xn-Fc 6 [(ErbB1/2/4)-Xc-Xn-Fc]2 ErbB1-Xc + ErbB2-Xc + ErbB4-Xc + Fc-Xc 6 [Fc-Xc-Xc-(ErbB1/2/4)]2 ErbB1-Xc + ErbB3-Xc + ErbB4-Xc + Xn-Fc 6 [(ErbB1/3/4)-Xc-Xn-Fc]2 ErbB1-Xc + ErbB3-Xc + ErbB4-Xc + Fc-Xc 6 [Fc-Xc-Xc-(ErbB1/3/4)]2 ErbB2-Xc + ErbB3-Xc + ErbB4-Xc + Xn-Fc 6 [(ErbB2/3/4)-Xc-Xn-Fc]2 ErbB2-Xc + ErbB3-Xc + ErbB4-Xc + Fc-Xc 6 [Fc-Xc-Xc-(ErbB2/3/4)]2 ErbB1-Xc + ErbB2-Xc + ErbB3-Xc + ErbB4-Xc + Xn-Fc 10 [(ErbB1/2/3/4)-Xc-Xn-Fc]2 ErbB1-Xc + ErbB2-Xc + ErbB3-Xc + ErbB4-Xc + Fc-Xc 10 [Fc-Xc-Xc-(ErbB1/2/3/4)]2
TABLE-US-00011 TABLE 11 ErbB Bi-Symmetroadhesins Stretches of Consecutive Amino Acids # Symmetroadhesin Configurations ErbB1-Xc + Xn-Fc-Xc 1 [ErbB1-Xc-Xn-Fc-Xc-Xc-ErbB1]2 ErbB2-Xc + Xn-Fc-Xc 1 [ErbB2-Xc-Xn-Fc-Xc-Xc-ErbB2]2 ErbB3-Xc + Xn-Fc-Xc 1 [ErbB3-Xc-Xn-Fc-Xc-Xc-ErbB3]2 ErbB4-Xc + Xn-Fc-Xc 1 [ErbB4-Xc-Xn-Fc-Xc-Xc-ErbB4]2 ErbB1-Xc + ErbB2-Xc + Xn-Fc-Xc 12 [(ErbB1/2)-Xc-Xn-Fc-Xc-Xc-(ErbB1/2)]2 ErbB1-Xc + ErbB3-Xc + Xn-Fc-Xc 12 [(ErbB1/3)-Xc-Xn-Fc-Xc-Xc-(ErbB1/3)]2 ErbB1-Xc + ErbB4-Xc + Xn-Fc-Xc 12 [(ErbB1/4)-Xc-Xn-Fc-Xc-Xc-(ErbB1/4)]2 ErbB2-Xc + ErbB3-Xc + Xn-Fc-Xc 12 [(ErbB2/3)-Xc-Xn-Fc-Xc-Xc-(ErbB2/3)]2 ErbB2-Xc + ErbB4-Xc + Xn-Fc-Xc 12 [(ErbB2/4)-Xc-Xn-Fc-Xc-Xc-(ErbB2/4)]2 ErbB3-Xc + ErbB4-Xc + Xn-Fc-Xc 12 [(ErbB3/4)-Xc-Xn-Fc-Xc-Xc-(ErbB3/4)]2 ErbB1-Xc + ErbB2-Xc + ErbB3-Xc + Xn-Fc-Xc 63 [(ErbB1/2/3)-Xc-Xn-Fc-Xc-Xc-(ErbB1/2/3)].su- b.2 ErbB1-Xc + ErbB2-Xc + ErbB4-Xc + Xn-Fc-Xc 63 [(ErbB1/2/4)-Xc-Xn-Fc-Xc-Xc-(ErbB1/2/4)].su- b.2 ErbB1-Xc + ErbB3-Xc + ErbB4-Xc + Xn-Fc-Xc 63 [(ErbB1/3/4)-Xc-Xn-Fc-Xc-Xc-(ErbB1/3/4)].su- b.2 ErbB2-Xc + ErbB3-Xc + ErbB4-Xc + Xn-Fc-Xc 63 [(ErbB2/3/4)-Xc-Xn-Fc-Xc-Xc-(ErbB2/3/4)].su- b.2 ErbB1-Xc + ErbB2-Xc + ErbB3-Xc + ErbB4-Xc + Xn-Fc-Xc [(ErbB1/2/3/4)-Xc-Xn-Fc-Xc-Xc-(ErbB1/2/3/4)- ]2
Stretches of Consecutive Amino Acids
[0271]Examples of stretches of consecutive amino acids as referred to herein include, but are not limited to, consecutive amino acids including binding domains such as secreted or transmembrane proteins, intracellular binding domains and antibodies (whole or portions thereof) and modified versions thereof. The following are some non-limiting examples:
1) Immunoglobulins
[0272]Immunoglobulins are molecules containing polypeptide chains held together by intra-chain disulfide bonds, wherein at least one of the bonded amino acids is not a terminus residue, typically having two light chains and two heavy chains. In each chain, one domain (V) has a variable amino acid sequence depending on the antibody specificity of the molecule. The other domains (C) have a rather constant sequence common among molecules of the same class. The domains are numbered in sequence from the amino-terminal end.
[0273]The immunoglobulin gene superfamily consists of molecules with immunoglobulin-like domains. Members of this family include class I and class II major histocompatibility antigens, immunoglobulins, T-cell receptor alpha, beta, gamma and delta chains, CD1, CD2, CD4, CD8, CD28, the gamma, delta and epsilon chains of CD3, OX-2, Thy-1, the intercellular or neural cell adhesion molecules (1-CAM or N-CAM), lymphocyte function associated antigen-3 (LFA-3), neurocytoplasmic protein (NCP-3), poly-Ig receptor, myelin-associated glycoprotein (MAG), high affinity IgE receptor, the major glycoprotein of peripheral myelin (Po), platelet derived growth factor receptor, colony stimulating factor-1 receptor, macrophage Fc receptor, Fc gamma receptors and carcinoembryonic antigen.
[0274]It is known that one can substitute variable domains (including hypervariable regions) of one immunoglobulin for another, and from one species to another. See, for example, EP 0 173 494; EP 0 125 023; Munro, Nature 312 (13 Dec. 1984); Neuberger et al., Nature 312: (13 Dec. 1984); Sharon et al., Nature 309 (May 24, 1984); Morrison et al., Proc. Natl. Acad. Sci. USA 81:6851-6855 (1984); Morrison et al. Science 229:1202-1207 (1985); and Boulianne et al., Nature 312:643-646 (Dec. 13, 1984).
[0275]Morrison et al., Science 229:1202-1207 (1985) teaches the preparation of an immunoglobulin chimera having a variable region from one species fused to an immunoglobulin constant region from another species.
[0276]It has also been shown that it is possible to substitute immunoglobulin variable-like domains from two members of the immunoglobulin gene superfamily-CD4 and the T cell receptor--for a variable domain in an immunoglobulin; see e.g. Capon et al., Nature 337:525-531, 1989, Traunecker et al., Nature 339:68-70, 1989, Gascoigne et al., Proc. Nat. Acad. Sci. 84:2936-2940, 1987, and published European application EPO 0 325 224 A2.
[0277]U.S. Pat. No. 5,116,964 (Capon et al., May 26, 1992) hereby incorporated by reference, describes hybrid immunoglobulins commonly referred to as immunoadhesins, which combine, for example, the adhesive and targeting properties of a ligand binding partner with immunoglobulin effector functions. U.S. Pat. No. 5,336,603 (Capon et al., Aug. 9, 1994), hereby incorporated by reference, describes a heterofunctional immunoadhesin comprising a fusion protein in which a polypeptide comprising a human CD4 antigen variable (V) region is fused at its C-terminus to the N-terminus of a polypeptide comprising a constant region of an immunoglobulin chain disulfide bonded to a companion immunoglobulin heavy chain-light chain pair bearing a antibody combining site capable of binding a predetermined antigen.
[0278]Components" of immunoglobulins include antibody fragments comprise a portion of an intact antibody, preferably the antigen binding or variable region of the intact antibody. Examples of antibody fragments include Fab, Fab', F(ab')2, and Fv fragments; diabodies; linear antibodies (Zapata et al., Protein Eng. 8(10): 1057-1062 (1995)); single-chain antibody molecules; and multispecific antibodies formed from antibody fragments.
[0279]Papain digestion of antibodies produces two identical antigen-binding fragments, called "Fab" fragments, each with a single antigen-binding site, and a residual "Fc" fragment, a designation reflecting the ability to crystallize readily. Pepsin treatment yields an F(ab')2 fragment that has two antigen-combining sites and is still capable of cross-linking antigen.
[0280]Fv" is the minimum antibody fragment which contains a complete antigen-recognition and -binding site. This region consists of a dimer of one heavy- and one light-chain variable domain in tight, non-covalent association. It is in this configuration that the three CDRs of each variable domain interact to define an antigen-binding site on the surface of the VH-VL dimer. Collectively, the six CDRs confer antigen-binding specificity to the antibody. However, even a single variable domain (or half of an Fv comprising only three CDRs specific for an antigen) has the ability to recognize and bind antigen, although at a lower affinity than the entire binding site.
[0281]The Fab fragment also contains the constant domain of the light chain and the first constant domain (CH1) of the heavy chain. Fab fragments differ from Fab' fragments by the addition of a few residues at the carboxy terminus of the heavy chain CH1 domain including one or more cysteines from the antibody hinge region. Fab'-SH is the designation herein for Fab' in which the cysteine residue(s) of the constant domains bear a free thiol group. F(ab')2 antibody fragments originally were produced as pairs of Fab' fragments which have hinge cysteines between them. Other chemical couplings of antibody fragments are also known.
[0282]The "light chains" of antibodies (immunoglobulins) from any vertebrate species can be assigned to one of two clearly distinct types, called kappa and lambda, based on the amino acid sequences of their constant domains.
[0283]Depending on the amino acid sequence of the constant domain of their heavy chains, immunoglobulins can be assigned to different classes. There are five major classes of immunoglobulins: IgA, IgD, IgE, IgG, and IgM, and several of these may be further divided into subclasses (isotypes), e.g., IgG1, IgG2, IgG3, IgG4, IgA, and IgA2.
[0284]Single-chain Fv" or "sFv" antibody fragments comprise the VH and VL domains of antibody, wherein these domains are present in a single polypeptide chain. Preferably, the Fv polypeptide further comprises a polypeptide linker between the VH and VL domains which enables the sFv to form the desired structure for antigen binding. For a review of sFv, see Pluckthun in The Pharmacology of Monoclonal Antibodies, vol. 113, Rosenburg and Moore eds., Springer-Verlag, New York, pp. 269-315 (1994).
[0285]Thus, linking of the various components of immunoglobulins and other biologically active molecules has allowed production of hybrid molecules which retain the functionalities of the individual components.
[0286]In an embodiment, the invention described herein provides novel hybrid molecules which include one or more immunoglobulin components.
2) Extracellular Proteins
[0287]Extracellular proteins play important roles in, among other things, the formation, differentiation and maintenance of multicellular organisms. A discussion of various intracellular proteins of interest is set forth in U.S. Pat. No. 6,723,535, Ashkenazi et al., issued Apr. 20, 2004, hereby incorporated by reference.
[0288]The fate of many individual cells, e.g., proliferation, migration, differentiation, or interaction with other cells, is typically governed by information received from other cells and/or the immediate environment. This information is often transmitted by secreted polypeptides (for instance, mitogenic factors, survival factors, cytotoxic factors, differentiation factors, neuropeptides, and hormones) which are, in turn, received and interpreted by diverse cell receptors or membrane-bound proteins. These secreted polypeptides or signaling molecules normally pass through the cellular secretory pathway to reach their site of action in the extracellular environment.
[0289]Secreted proteins have various industrial applications, including as pharmaceuticals, diagnostics, biosensors and bioreactors. Most protein drugs available at present, such as thrombolytic agents, interferons, interleukins, erythropoietines, colony stimulating factors, and various other cytokines, are secretory proteins. Their receptors, which are membrane proteins, also have potential as therapeutic or diagnostic agents. Efforts are being undertaken by both industry and academia to identify new, native secreted proteins. Many efforts are focused on the screening of mammalian recombinant DNA libraries to identify the coding sequences for novel secreted proteins. Examples of screening methods and techniques are described in the literature (see, for example, Klein et al., Proc. Natl. Acad. Sci. 93:7108-7113 (1996); U.S. Pat. No. 5,536,637)).
[0290]Membrane-bound proteins and receptors can play important roles in, among other things, the formation, differentiation and maintenance of multicellular organisms. The fate of many individual cells, e.g., proliferation, migration, differentiation, or interaction with other cells, is typically governed by information received from other cells and/or the immediate environment. This information is often transmitted by secreted polypeptides (for instance, mitogenic factors, survival factors, cytotoxic factors, differentiation factors, neuropeptides, and hormones) which are, in turn, received and interpreted by diverse cell receptors or membrane-bound proteins. Such membrane-bound proteins and cell receptors include, but are not limited to, cytokine receptors, receptor kinases, receptor phosphatases, receptors involved in cell-cell interactions, and cellular adhesin molecules like selectins and integrins. For instance, transduction of signals that regulate cell growth and differentiation is regulated in part by phosphorylation of various cellular proteins. Protein tyrosine kinases, enzymes that catalyze that process, can also act as growth factor receptors. Examples include fibroblast growth factor receptor and nerve growth factor receptor.
[0291]Membrane-bound proteins and receptor molecules have various industrial applications, including as pharmaceutical and diagnostic agents. Receptor immunoadhesins, for instance, can be employed as therapeutic agents to block receptor-ligand interactions. The membrane-bound proteins can also be employed for screening of potential peptide or small molecule inhibitors of the relevant receptor/ligand interaction.
[0292]Examples of such proteins include EGF and growth factors.
[0293]Epidermal growth factor (EGF) is a conventional mitogenic factor that stimulates the proliferation of various types of cells including epithelial cells and fibroblasts. EGF binds to and activates the EGF receptor (EGFR), which initiates intracellular signaling and subsequent effects. The EGFR is expressed in neurons of the cerebral cortex, cerebellum, and hippocampus in addition to other regions of the central nervous system (CNS). In addition, EGF is also expressed in various regions of the CNS. Therefore, EGF acts not only on mitotic cells, but also on postmitotic neurons. In fact, many studies have indicated that EGF has neurotrophic or neuromodulatory effects on various types of neurons in the CNS. For example, EGF acts directly on cultured cerebral cortical and cerebellar neurons, enhancing neurite outgrowth and survival. On the other hand, EGF also acts on other cell types, including septal cholinergic and mesencephalic dopaminergic neurons, indirectly through glial cells. Evidence of the effects of EGF on neurons in the CNS is accumulating, but the mechanisms of action remain essentially unknown. EGF-induced signaling in mitotic cells is better understood than in postmitotic neurons. Studies of cloned pheochromocytoma PC12 cells and cultured cerebral cortical neurons have suggested that the EGF-induced neurotrophic actions are mediated by sustained activation of the EGFR and mitogen-activated protein kinase (MAPK) in response to EGF. The sustained intracellular signaling correlates with the decreased rate of EGFR down-regulation, which might determine the response of neuronal cells to EGF. It is likely that EGF is a multi-potent growth factor that acts upon various types of cells including mitotic cells and postmitotic neurons.
[0294]EGF is produced by the salivary and Brunner's glands of the gastrointestinal system, kidney, pancreas, thyroid gland, pituitary gland, and the nervous system, and is found in body fluids such as saliva, blood, cerebrospinal fluid (CSF), urine, amniotic fluid, prostatic fluid, pancreatic juice, and breast milk, Plata-Salaman, Peptides 12:653-663 (1991).
[0295]EGF is mediated by its membrane specific receptor, which contains an intrinsic tyrosine kinase. Stoscheck et al., J. Cell Biochem. 31:135-152 (1986). EGF is believed to function by binding to the extracellular portion of its receptor which induces a transmembrane signal that activates the intrinsic tyrosine kinase.
[0296]Purification and sequence analysis of the EGF-like domain has revealed the presence of six conserved cysteine residues which cross-bind to create three peptide loops, Savage et al., J. Biol. Chem. 248:7669-7672 (1979). It is now generally known that several other peptides can react with the EGF receptor which share the same generalized motif Xn CX7 CX4/5 CX10 CXCX5 GX2 CXn, where X represents any non-cysteine amino acid, and n is a variable repeat number. Non isolated peptides having this motif include TGF-alpha, amphiregulin, schwannoma-derived growth factor (SDGF), heparin-binding EGF-like growth factors and certain virally encoded peptides (e.g., Vaccinia virus, Reisner, Nature 313:801-803 (1985), Shope fibroma virus, Chang et al., Mol Cell Biol. 7:535-540 (1987), Molluscum contagiosum, Porter and Archard, J. Gen. Virol. 68:673-682 (1987), and Myxoma virus, Upton et al., J. Virol. 61:1271-1275 (1987), Prigent and Lemoine, Prog. Growth Factor Res. 4:1-24 (1992).
[0297]EGF-like domains are not confined to growth factors but have been observed in a variety of cell-surface and extracellular proteins which have interesting properties in cell adhesion, protein-protein interaction and development, Laurence and Gusterson, Tumor Biol. 11:229-261 (1990). These proteins include blood coagulation factors (factors VI, IX, X, XII, protein C, protein S, protein Z, tissue plasminogen activator, urokinase), extracellular matrix components (laminin, cytotactin, entactin), cell surface receptors (LDL receptor, thrombomodulin receptor) and immunity-related proteins (complement C1r, uromodulin).
[0298]Even more interesting, the general structure pattern of EGF-like precursors is preserved through lower organisms as well as in mammalian cells. A number of genes with developmental significance have been identified in invertebrates with EGF-like repeats. For example, the notch gene of Drosophila encodes 36 tandemly arranged 40 amino acid repeats which show homology to EGF, Wharton et al., Cell 43:557-581 (1985). Hydropathy plots indicate a putative membrane spanning domain, with the EGF-related sequences being located on the extracellular side of the membrane. Other homeotic genes with EGF-like repeats include Delta, 95F and 5ZD which were identified using probes based on Notch, and the nematode gene Lin-12 which encodes a putative receptor for a developmental signal transmitted between two specified cells.
[0299]Specifically, EGF has been shown to have potential in the preservation and maintenance of gastrointestinal mucosa and the repair of acute and chronic mucosal lesions, Konturek et al., Eur. J. Gastroenterol Hepatol. 7 (10), 933-37 (1995), including the treatment of necrotizing enterocolitis, Zollinger-Ellison syndrome, gastrointestinal ulceration gastrointestinal ulcerations and congenital microvillus atrophy, Guglietta and Sullivan, Eur. J. Gastroenterol Hepatol, 7(10), 945-50 (1995). Additionally, EGF has been implicated in hair follicle differentiation; du Cros, J. Invest. Dermatol. 101 (1 Suppl.), 106S-113S (1993), Hillier, Clin. Endocrinol. 33(4), 427-28 (1990); kidney function, Hamm et al., Semin. Nephrol. 13(1): 109-15 (1993), Harris, Am. J. Kidney Dis. 17(6): 627-30 (1991); tear fluid, van Setten et al., Int. Opthalmol 15(6); 359-62 (1991); vitamin K mediated blood coagulation, Stenflo et al., Blood 78(7): 1637-51 (1991). EGF is also implicated various skin disease characterized by abnormal keratinocyte differentiation, e.g., psoriasis, epithelial cancers such as squamous cell carcinomas of the lung, epidermoid carcinoma of the vulva and gliomas. King et al., Am. J. Med. Sci. 296:154-158 (1988).
[0300]Of great interest is mounting evidence that genetic alterations in growth factors signaling pathways are closely linked to developmental abnormalities and to chronic diseases including cancer. Aaronson, Science 254: 1146-1153 (1991). For example, c-erb-2 (also known as HER-2), a proto-oncogene with close structural similarity to EGF receptor protein, is overexpressed in human breast cancer. King et al., Science 229:974-976 (1985); Gullick, Hormones and their actions, Cooke et al., eds, Amsterdam, Elsevier, pp 349-360 (1986).
[0301]Growth factors are molecular signals or mediators that enhance cell growth or proliferation, alone or in concert, by binding to specific cell surface receptors. However, there are other cellular reactions than only growth upon expression to growth factors. As a result, growth factors are better characterized as multifunctional and potent cellular regulators. Their biological effects include proliferation, chemotaxis and stimulation of extracellular matrix production. Growth factors can have both stimulatory and inhibitory effects. For example, transforming growth factor (TGF-beta) is highly pleiotropic and can stimulate proliferation in some cells, especially connective tissue, while being a potent inhibitor of proliferation in others, such as lymphocytes and epithelial cells.
[0302]The physiological effect of growth stimulation or inhibition by growth factors depends upon the state of development and differentiation of the target tissue. The mechanism of local cellular regulation by classical endocrine molecules involves comprehends autocrine (same cell), juxtacrine (neighbor cell), and paracrine (adjacent cells) pathways. Peptide growth factors are elements of a complex biological language, providing the basis for intercellular communication. They permit cells to convey information between each other, mediate interaction between cells and change gene expression. The effect of these multifunctional and pluripotent factors is dependent on the presence or absence of other peptides.
[0303]FGF-8 is a member of the fibroblast growth factors (FGFs) which are a family of heparin-binding, potent mitogens for both normal diploid fibroblasts and established cell lines, Gospodarowicz et al. (1984), Proc. Nat. Acad. Sci. USA 81:6963. The FGF family comprises acidic FGF (FGF-1), basic FGF (FGF-2), INT-2 (FGF-3), K-FGF/HST (FGF-4), FGF-5, FGF-6, KGF (FGF-7), AIGF (FGF-8) among others. All FGFs have two conserved cysteine residues and share 30-50% sequence homology at the amino acid level. These factors are mitogenic for a wide variety of normal diploid mesoderm-derived and neural crest-derived cells, including granulosa cells, adrenal cortical cells, chondrocytes, myoblasts, corneal and vascular endothelial cells (bovine or human), vascular smooth muscle cells, lens, retina and prostatic epithelial cells, oligodendrocytes, astrocytes, chrondocytes, myoblasts and osteoblasts.
[0304]Fibroblast growth factors can also stimulate a large number of cell types in a non-mitogenic manner. These activities include promotion of cell migration into wound area (chemotaxis), initiation of new blood vessel formulation (angiogenesis), modulation of nerve regeneration and survival (neurotrophism), modulation of endocrine functions, and stimulation or suppression of specific cellular protein expression, extracellular matrix production and cell survival. Baird & Bohlen, Handbook of Exp. Pharmacol. 95(1): 369418, Springer, (1990). These properties provide a basis for using fibroblast growth factors in therapeutic approaches to accelerate wound healing, nerve repair, collateral blood vessel formation, and the like. For example, fibroblast growth factors have been suggested to minimize myocardium damage in heart disease and surgery (U.S. Pat. No. 4,378,347).
[0305]FGF-8, also known as androgen-induced growth factor (AIGF), is a 215 amino acid protein which shares 30-40% sequence homology with the other members of the FGF family. FGF-8 has been proposed to be under androgenic regulation and induction in the mouse mammary carcinoma cell line SC3. Tanaka et al., Proc. Natl. Acad. Sci. USA 89:8928-8932 (1992); Sato et al., J. Steroid Biochem. Molec. Biol. 47:91-98 (1993). As a result, FGF-8 may have a local role in the prostate, which is known to be an androgen-responsive organ. FGF-8 can also be oncogenic, as it displays transforming activity when transfected into NIH-3T3 fibroblasts. Kouhara et al., Oncogene 9 455-462 (1994). While FGF-8 has been detected in heart, brain, lung, kidney, testis, prostate and ovary, expression was also detected in the absence of exogenous androgens. Schmitt et al., J. Steroid Biochem. Mol. Biol. 57 (34): 173-78 (1996).
[0306]FGF-8 shares the property with several other FGFs of being expressed at a variety of stages of murine embryogenesis, which supports the theory that the various FGFs have multiple and perhaps coordinated roles in differentiation and embryogenesis. Moreover, FGF-8 has also been identified as a protooncogene that cooperates with Wnt-1 in the process of mammary tumorigenesis (Shackleford et al., Proc. Natl. Acad. Sci. USA 90, 740-744 (1993); Heikinheimo et al., Mech. Dev. 48:129-138 (1994)).
[0307]In contrast to the other FGFs, FGF-8 exists as three protein isoforms, as a result of alternative splicing of the primary transcript. Tanaka et al., supra. Normal adult expression of FGF-8 is weak and confined to gonadal tissue, however northern blot analysis has indicated that FGF-8 mRNA is present from day 10 through day 12 or murine gestation, which suggests that FGF-8 is important to normal development. Heikinheimo et al., Mech Dev. 48(2): 129-38 (1994). Further in situ hybridization assays between day 8 and 16 of gestation indicated initial expression in the surface ectoderm of the first bronchial arches, the frontonasal process, the forebrain and the midbrain-hindbrain junction. At days 10-12, FGF-8 was expressed in the surface ectoderm of the forelimb and hindlimb buds, the nasal its and nasopharynx, the infundibulum and in the telencephalon, diencephalon and metencephalon. Expression continues in the developing hindlimbs through day 13 of gestation, but is undetectable thereafter. The results suggest that FGF-8 has a unique temporal and spatial pattern in embryogenesis and suggests a role for this growth factor in multiple regions of ectodermal differentiation in the post-gastrulation embryo.
[0308]The TGF-beta supergene family, or simply TGF-beta superfamily, a group of secreted proteins, includes a large number of related growth and differentiation factors expressed in virtually all phyla. Superfamily members bind to specific cell surface receptors that activate signal transduction mechanisms to elicit their multifunctional cytokine effects. Kolodziejczyk and Hall, Biochem. Cell. Biol., 74:299-314 (1996); Attisano and Wrana, Cytokine Growth Factor Rev., 7:327-339 (1996); and Hill, Cellular Signaling, 8:533-544 (1996).
[0309]Members of this family include five distinct forms of TGF-beta (Sporn and Roberts, in Peptide Growth Factors and Their Receptors, Sporn and Roberts, eds. (Springer-Verlag: Berlin, 1990) pp. 419-472), as well as the differentiation factors vg1 (Weeks and Melton, Cell, 51:861-867 (1987)) and DPP-C polypeptide (Padgett et al., Nature, 325:81-84 (1987)), the hormones activin and inhibin (Mason et al., Nature 318-659-663 (1985); Mason et al., Growth Factors, 1:77-88 (1987)), the Mullerian-inhibiting substance (MIS) (Cate et al., Cell, 45: 685-698 (1986)), the bone morphogenetic proteins (BMPs) (Wozney et al., Science, 242:1528-1534 (1988); PCT WO 88/00205 published Jan. 14, 1988; U.S. Pat. No. 4,877,864 issued Oct. 31, 1989), the developmentally regulated proteins Vgr-1 (Lyons et al., Proc. Natl. Acad. Sci. USA. 86:45544558 (1989)) and Vgr-2 (Jones et al., Molec. Endocrinol., 6:1961-1968 (1992)), the mouse growth differentiation factor (GDF), such as GDF-3 and GDF-9 (Kingsley, Genes Dev., 8:133-146 (1994); McPherron and Lee, J. Biol. Chem., 268:3444-3449 (1993)), the mouse lefty/Stral (Meno et al., Nature, 381:151-155 (1996); Bouillet et al., Dev. Biol., 170: 420-433 (1995)), glial cell line-derived neurotrophic factor (GDNF) (Lin et al., Science, 260:1130-1132 (1993), neurturin (Kotzbauer et al., Nature, 384:467-470 (1996)), and endometrial bleeding-associated factor (EBAF) (Kothapalli et al., J. Clin. Invest., 99:2342-2350 (1997)). The subset BMP-2A and BMP-2B is approximately 75% homologous in sequence to DPP-C and may represent the mammalian equivalent of that protein.
[0310]The proteins of the TGF-beta superfamily are disulfide-linked homo- or heterodimers encoded by larger precursor polypeptide chains containing a hydrophobic signal sequence, a long and relatively poorly conserved N-terminal pro region of several hundred amino acids, a cleavage site (usually polybasic), and a shorter and more highly conserved C-terminal region. This C-terminal region corresponds to the processed mature protein and contains approximately 100 amino acids with a characteristic cysteine motif, i.e. the conservation of seven of the nine cysteine residues of TGF-beta among all known family members. Although the position of the cleavage site between the mature and pro regions varies among the family members, the C-terminus of all of the proteins is in the identical position, ending in the sequence Cys-X-Cys-X, but differing in every case from the TGF-beta consensus C-terminus of Cys-Lys-Cys-Ser. Sporn and Roberts, 1990, supra.
[0311]There are at least five forms of TGF-beta currently identified, TGF-beta1, TGF-beta2, TGF-beta3, TGF-beta4, and TGF-beta5. The activated form of TGF-beta1 is a homodimer formed by dimerization of the carboxy-terminal 112 amino acids of a 390 amino acid precursor. Recombinant TGF-beta1 has been cloned (Derynck et al., Nature, 316:701-705 (1985)) and expressed in Chinese hamster ovary cells (Gentry et al., Mol. Cell. Biol. 7:3418-3427 (1987)). Additionally, recombinant human TGF-beta2 (deMartin et al., EMBO J., 6:3673 (1987)), as well as human and porcine TGF-beta3 (Derynck et al., EMBO J., 7:3737-3743 (1988); ten Dijke et al., Proc. Natl. Acad. Sci. USA, 85:4715 (1988)) have been cloned. TGF-beta2 has a precursor form of 414 amino acids and is also processed to a homodimer from the carboxy-terminal 112 amino acids that shares approximately 70% homology with the active form of TGF-beta1 (Marquardt et al., J. Biol. Chem., 262:12127 (1987)). See also EP 200,341; 169,016; 268,561; and 267,463; U.S. Pat. No. 4,774,322; Cheifetz et al., Cell, 48:409-415 (1987); Jakowlew et al., Molecular Endocrin., 2:747-755 (1988); Derynck et al., J. Biol. Chem., 261:4377-4379 (1986); Sharples et al., DNA, 6:239-244 (1987); Derynck et al., Nucl. Acids. Res., 15:3188-3189 (1987); Derynck et al., Nucl. Acids. Res. 15:3187 (1987); Seyedin et al., J. Biol. Chem., 261:5693-5695 (1986); Madisen et al., DNA 7:1-8 (1988); and Hanks et al., Proc. Natl. Acad. Sci. (U.S.A.), 85:79-82 (1988).
[0312]TGF-beta4 and TGF-beta5 were cloned from a chicken chondrocyte cDNA library (Jakowlew et al., Molec. Endocrinol., 2:1186-1195 (1988)) and from a frog oocyte cDNA library, respectively.
[0313]The pro region of TGF-beta associates non-covalently with the mature TGF-beta dimer (Wakefield et al., J. Biol. Chem., 263:7646-7654 (1988); Wakefield et al., Growth Factors, 1:203-218 (1989)), and the pro regions are found to be necessary for proper folding and secretion of the active mature dimers of both TGF-beta and activin (Gray and Mason, Science, 247:1328-1330 (1990)). The association between the mature and pro regions of TGF-beta masks the biological activity of the mature dimer, resulting in formation of an inactive latent form. Latency is not a constant of the TGF-beta superfamily, since the presence of the pro region has no effect on activin or inhibin biological activity.
[0314]A unifying feature of the biology of the proteins from the TGF-beta superfamily is their ability to regulate developmental processes. TGF-beta has been shown to have numerous regulatory actions on a wide variety of both normal and neoplastic cells. TGF-beta is multifunctional, as it can either stimulate or inhibit cell proliferation, differentiation, and other critical processes in cell function (Sporn and Roberts, supra).
[0315]One member of the TGF-beta superfamily, EBAF, is expressed in endometrium only in the late secretory phase and during abnormal endometrial bleeding. Kothapalli et al., J. Clin. Invest., 99:2342-2350 (1997). Human endometrium is unique in that it is the only tissue in the body that bleeds at regular intervals. In addition, abnormal endometrial bleeding is one of the most common manifestations of gynecological diseases, and is a prime indication for hysterectomy. In situ hybridization showed that the mRNA of EBAF was expressed in the stroma without any significant mRNA expression in the endometrial glands or endothelial cells.
[0316]The predicted protein sequence of EBAF showed a strong homology to the protein encoded by mouse lefty/stra3 of the TGF-beta superfamily. A motif search revealed that the predicted EBAF protein contains most of the cysteine residues which are conserved among the TGF-beta-related proteins and which are necessary for the formation of the cysteine knot structure. The EBAF sequence contains an additional cysteine residue, 12 amino acids upstream from the first conserved cysteine residue. The only other family members known to contain an additional cysteine residue are TGF-betas, inhibins, and GDF-3. EBAF, similar to LEFTY, GDF-3/Vgr2, and GDF-9, lacks the cysteine residue that is known to form the intermolecular disulfide bond. Therefore, EBAF appears to be an additional member of the TGF-beta superfamily with an unpaired cysteine residue that may not exist as a dimer. However, hydrophobic contacts between the two monomer subunits may promote dimer formation. Fluorescence in situ hybridization showed that the ebaf gene is located on human chromosome 1 at band q42.1.
[0317]Further examples of such extracellular proteins are well known in the art, for example see U.S. Pat. No. 6,723,535.
Conjugates
[0318]The invention also pertains to conjugates of symmtroadhesins/immunosymmetroadhesins. Thus the instant compositions can be conjugated to a cytotoxic agent such as a chemotherapeutic agent, toxin (e.g., an enzymatically active toxin of bacterial, fungal, plant, or animal origin, or fragments thereof), or a radioactive isotope (i.e., a radioconjugate).
[0319]Chemotherapeutic agents useful in the generation of such immunoconjugates are well known in the art. Enzymatically active toxins and fragments thereof that can be used include diphtheria A chain, nonbinding active fragments of diphtheria toxin, exotoxin A chain (from Pseudomonas aeruginosa), ricin A chain, abrin A chain, modeccin A chain, alpha-sarcin, Aleurites fordii proteins, dianthin proteins, Phytolaca americana proteins (PAPI, PAPH, and PAP-S), momordica charantia inhibitor, curcin, crotin, sapaonaria officinalis inhibitor, gelonin, mitogellin, restrictocin, phenomycin, enomycin, and the tricothecenes. A variety of radionuclides are available for the production of radioconjugated antibodies. Examples include 212Bi, 131I, 113In, 90Y, and 186Re.
[0320]Conjugates of the compositions of the invention including cytotoxic agent are made using a variety of bifunctional protein-coupling agents such as N-succinimidyl-3-(2-pyridyldithiol) propionate (SPDP), iminothiolane (IT), bifunctional derivatives of imidoesters (such as dimethyl adipimidate HCL), active esters (such as disuccinimidyl suberate), aldehydes (such as glutareldehyde), bis-azido compounds (such as bis(p-azidobenzoyl)hexanediamine), bis-diazonium derivatives (such as bis-(p-diazoniumbenzoyl)-ethylenediamine), diisocyanates (such as tolyene 2,6-diisocyanate), and bis-active fluorine compounds (such as 1,5-difluoro-2,4-dinitrobenzene). For example, a ricin immunotoxin can be prepared as described in Vitetta et al., Science, 238:1098 (1987). Carbon-14-labeled 1-isothiocyanatobenzyl-3-methyldiethylene triaminepentaacetic acid (MX-DTPA) is an exemplary chelating agent for conjugation of radionucleotide to the antibody. See WO94111026.
Synthesis of Genetic Devices
[0321]The genetic devices disclosed herein may be synthesized by various routes. One particular route is to synthesize components in vivo by recombinant DNA technology and then to chemically modify the secreted or procured products under conditions so as to form the compounds. Alternative routes include solid-state synthesis.
General Techniques
[0322]The description below relates primarily to production of stretches of consecutive amino acids or polypeptides of interest by culturing cells transformed or transfected with a vector containing an encoding nucleic acid. It is, of course, contemplated that alternative methods, which are well known in the art, may be employed. For instance, the amino acid sequence, or portions thereof, may be produced by direct peptide synthesis using solid-phase techniques (see, e.g., Stewart et al., Solid-Phase Peptide Synthesis, W.H. Freeman Co., San Francisco, Calif. (1969); Merrifield, J. Am. Chem. Soc., 85:2149-2154 (1963)). In vitro protein synthesis may be performed using manual techniques or by automation. Automated synthesis may be accomplished, for instance, using an Applied Biosystems Peptide Synthesizer (Foster City, Calif.) using manufacturer's instructions. Various portions of the stretches of consecutive amino acids or polypeptides of interest may be chemically synthesized separately and combined using chemical or enzymatic methods to produce the full-length stretches of consecutive amino acids or polypeptides of interest.
1. Isolation of DNA Encoding Stretches of Consecutive Amino Acids or Polypeptides of Interest
[0323]Encoding DNA may be obtained from a cDNA library prepared from tissue believed to possess the mRNA of interest and to express it at a detectable level. Accordingly, human DNA can be conveniently obtained from a cDNA library prepared from human tissue, and so forth. An encoding gene may also be obtained from a genomic library or by known synthetic procedures (e.g., automated nucleic acid synthesis).
[0324]Libraries can be screened with probes (such as antibodies to the stretch of consecutive amino acids or oligonucleotides of at least about 20-80 bases) designed to identify the gene of interest or the protein encoded by it. Screening the cDNA or genomic library with the selected probe may be conducted using standard procedures, such as described in Sambrook et al., Molecular Cloning: A Laboratory Manual (New York: Cold Spring Harbor Laboratory Press, 1989). An alternative means to isolate the encoding gene is to use PCR methodology (Sambrook et al., supra; Dieffenbach et al., PCR Primer: A Laboratory Manual (Cold Spring Harbor Laboratory Press, 1995)).
[0325]The oligonucleotide sequences selected as probes should be of sufficient length and sufficiently unambiguous that false positives are minimized. The oligonucleotide is preferably labeled such that it can be detected upon hybridization to DNA in the library being screened. Methods of labeling are well known in the art, and include the use of radiolabels like 32P-labeled ATP, biotinylation or enzyme labeling. Hybridization conditions, including moderate stringency and high stringency, are provided in Sambrook et al., supra.
[0326]Sequences identified in such library screening methods can be compared and aligned to other known sequences deposited and available in public databases such as GenBank or other private sequence databases. Sequence identity (at either the amino acid or nucleotide level) within defined regions of the molecule or across the full-length sequence can be determined using methods known in the art and as described herein.
[0327]Nucleic acid having protein coding sequence may be obtained by screening selected cDNA or genomic libraries using the deduced amino acid sequence disclosed herein for the first time, and, if necessary, using conventional primer extension procedures as described in Sambrook et al., supra, to detect precursors and processing intermediates of mRNA that may not have been reverse-transcribed into cDNA.
2. Selection and Transformation of Host Cells
[0328]Host cells are transfected or transformed with expression or cloning vectors described herein for production and cultured in conventional nutrient media modified as appropriate for inducing promoters, selecting transformants, or amplifying the genes encoding the desired sequences. The culture conditions, such as media, temperature, pH and the like, can be selected by the skilled artisan without undue experimentation. In general, principles, protocols, and practical techniques for maximizing the productivity of cell cultures can be found in Mammalian Cell Biotechnology: a Practical Approach, M. Butler, ed. (IRL Press, 1991) and Sambrook et al., supra.
[0329]Methods of eukaryotic cell transfection and prokaryotic cell transformation are known to the ordinarily skilled artisan, for example, CaCl2, CaPO4, liposome-mediated and electroporation. Depending on the host cell used, transformation is performed using standard techniques appropriate to such cells. The calcium treatment employing calcium chloride, as described in Sambrook et al., supra, or electroporation is generally used for prokaryotes. Infection with Agrobacterium tumefaciens is used for transformation of certain plant cells, as described by Shaw et al., Gene, 23:315 (1983) and WO 89/05859 published Jun. 29, 1989. For mammalian cells without such cell walls, the calcium phosphate precipitation method of Graham and van der Eb, Virology, 52:456-457 (1978) can be employed. General aspects of mammalian cell host system transfections have been described in U.S. Pat. No. 4,399,216. Transformations into yeast are typically carried out according to the method of Van Solingen et al., J. Bact., 130:946 (1977) and Hsiao et al., Proc. Natl. Acad. Sci. (USA), 76:3829 (1979). However, other methods for introducing DNA into cells, such as by nuclear microinjection, electroporation, bacterial protoplast fusion with intact cells, or polycations, e.g., polybrene, polyornithine, may also be used. For various techniques for transforming mammalian cells, see Keown et al., Methods in Enzymology, 185:527-537 (1990) and Mansour et al., Nature, 336:348-352 (1988).
[0330]Suitable host cells for cloning or expressing the DNA in the vectors herein include prokaryote, yeast, or higher eukaryote cells. Suitable prokaryotes include but are not limited to eubacteria, such as Gram-negative or Gram-positive organisms, for example, Enterobacteriaceae such as E. coli. Various E. coli strains are publicly available, such as E. coli K12 strain MM294 (ATCC 31,446); E. coli X1776 (ATCC 31,537); E. coli strain W3110 (ATCC 27,325) and K5772 (ATCC 53,635). Other suitable prokaryotic host cells include Enterobacteriaceae such as Escherichia, e.g., E. coli, Enterobacter, Erwinia, Klebsiella, Proteus, Salmonella, e.g., Salmonella typhimurium, Serratia, e.g., Serratia marcescans, and Shigella, as well as Bacilli such as B. subtilis and B. licheniformis (e.g., B. licheniformis 41P disclosed in DD 266,710 published Apr. 12, 1989), Pseudomonas such as P. aeruginosa, and Streptomyces. These examples are illustrative rather than limiting. Strain W3110 is one particularly preferred host or parent host because it is a common host strain for recombinant DNA product fermentations. Preferably, the host cell secretes minimal amounts of proteolytic enzymes. For example, strain W3110 may be modified to effect a genetic mutation in the genes encoding proteins endogenous to the host, with examples of such hosts including E. coli W3110 strain 1A2, which has the complete genotype tonA; E. coli W3110 strain 9E4, which has the complete genotype tonA ptr3; E. coli W3110 strain 27C7 (ATCC 55,244), which has the complete genotype tonAptr3phoA E15 (argF-lac)169 degP ompT kanr; E. coli W3110 strain 37D6, which has the complete genotype tonA ptr3 phoA E15 (argF-lac)169 degP ompT rbs7 ilvG kanr, E. coli W3110 strain 40B4, which is strain 37D6 with a non-kanamycin resistant degP deletion mutation; and an E. coli strain having mutant periplasmic protease disclosed in U.S. Pat. No. 4,946,783 issued Aug. 7, 1990. Alternatively, in vitro methods of cloning, e.g., PCR or other nucleic acid polymerase reactions, are suitable.
[0331]In addition to prokaryotes, eukaryotic microbes such as filamentous fungi or yeast are suitable cloning or expression hosts for encoding vectors. Saccharomyces cerevisiae is a commonly used lower eukaryotic host microorganism. Others include Schizosaccharomyces pombe (Beach and Nurse, Nature, 290:140 (1981); EP 139,383 published May 2, 1985); Kluyveromyces hosts (U.S. Pat. No. 4,943,529; Fleer et al., Bio/Technology, 9:968-975 (1991)) such as, e.g., K. lactis (MW98-8C, CBS683, CBS4574; Louvencourt et al., J. Bacteriol., 737 (1983)), K. fragilis (ATCC 12,424), K. bulgaricus (ATCC 16,045), K. wickeramii (ATCC 24,178), K. waltii (ATCC 56,500), K. drosophilarum (ATCC 36,906; Van den Berg et al., Bio/Technology, 8:135 (1990)), K. thermotolerans, and K. marxianus; yarrowia (EP 402,226); Pichia pastoris (EP 183,070; Sreekrishna et al., J. Basic Microbiol., 28:265-278 (1988)); Candida; Trichoderma reesia (EP 244,234); Neurospora crassa (Case et al., Proc. Natl. Acad. Sci. USA, 76:5259-5263 (1979)); Schwanniomyces such as Schwanniomyces occidentalis (EP 394,538 published Oct. 31, 1990); and filamentous fungi such as, e.g., Neurospora, Penicillium, Tolypocladium (WO 91/00357 published Jan. 10, 1991), and Aspergillus hosts such as A. nidulans (Ballance et al., Biochem. Biophys. Res. Commun., 112:284-289 (1983); Tilburn et al., Gene, 26:205-221 (1983); Yelton et al., Proc. Natl. Acad. Sci. USA, 81:1470-1474 (1984)) and A. niger (Kelly and Hynes, EMBO J., 4:475479 (1985)). Methylotropic yeasts are suitable herein and include, but are not limited to, yeast capable of growth on methanol selected from the genera consisting of Hansenula, Candida, Kloeckera, Pichia, Saccharomyces, Torulopsis, and Rhodotorula. A list of specific species that are exemplary of this class of yeasts may be found in C. Anthony, The Biochemistry of Methylotrophs, 269 (1982).
[0332]Suitable host cells for the expression of glycosylated stretches of consecutive amino acids or polypeptides of interest are derived from multicellular organisms. Examples of invertebrate cells include insect cells such as Drosophila S2 and Spodoptera Sf9, as well as plant cells. Examples of useful mammalian host cell lines include Chinese hamster ovary (CHO) and COS cells. More specific examples include monkey kidney CV1 line transformed by SV40 (COS-7, ATCC CRL 1651); human embryonic kidney line (293 or 293 cells subcloned for growth in suspension culture, Graham et al., J. Gen Virol., 36:59 (1977)); Chinese hamster ovary cells/-DHFR (CHO, Urlaub and Chasin, Proc. Natl. Acad. Sci. USA, 77:4216 (1980)); mouse sertoli cells (TM4, Mather, Biol. Reprod., 23:243-251 (1980)); human lung cells (W138, ATCC CCL 75); human liver cells (Hep G2, HB 8065); and mouse mammary tumor (MMT 060562, ATCC CCL51). The selection of the appropriate host cell is deemed to be within the skill in the art.
3. Selection and Use of a Replicable Vector
[0333]The nucleic acid (e.g., cDNA or genomic DNA) encoding the stretch of consecutive amino acids or polypeptides of interest may be inserted into a replicable vector for cloning (amplification of the DNA) or for expression. Various vectors are publicly available. The vector may, for example, be in the form of a plasmid, cosmid, viral particle, or phage. The appropriate nucleic acid sequence may be inserted into the vector by a variety of procedures. In general, DNA is inserted into an appropriate restriction endonuclease site(s) using techniques known in the art. Vector components generally include, but are not limited to, one or more of a signal sequence, an origin of replication, one or more marker genes, an enhancer element, a promoter, and a transcription termination sequence. Construction of suitable vectors containing one or more of these components employs standard ligation techniques which are known to the skilled artisan.
[0334]The stretches of consecutive amino acids or polypeptides of interest may be produced recombinantly not only directly, but also as a fusion polypeptide with a heterologous polypeptide, which may be a signal sequence or other polypeptide having a specific cleavage site at the N-terminus of the mature protein or polypeptide. In general, the signal sequence may be a component of the vector, or it may be a part of the encoding DNA that is inserted into the vector. The signal sequence may be a prokaryotic signal sequence selected, for example, from the group of the alkaline phosphatase, penicillinase, lpp, or heat-stable enterotoxin II leaders. For yeast secretion the signal sequence may be, e.g., the yeast invertase leader, alpha factor leader (including Saccharomyces and Kluyveromyces alpha-factor leaders, the latter described in U.S. Pat. No. 5,010,182), or acid phosphatase leader, the C. albicans glucoamylase leader (EP 362,179 published Apr. 4, 1990), or the signal described in WO 90/13646 published Nov. 15, 1990. In mammalian cell expression, mammalian signal sequences may be used to direct secretion of the protein, such as signal sequences from secreted polypeptides of the same or related species, as well as viral secretory leaders.
[0335]Both expression and cloning vectors contain a nucleic acid sequence that enables the vector to replicate in one or more selected host cells. Such sequences are well known for a variety of bacteria, yeast, and viruses. The origin of replication from the plasmid pBR322 is suitable for most Gram-negative bacteria, the 2mu plasmid origin is suitable for yeast, and various viral origins (SV40, polyoma, adenovirus, VSV or BPV) are useful for cloning vectors in mammalian cells.
[0336]Expression and cloning vectors will typically contain a selection gene, also termed a selectable marker. Typical selection genes encode proteins that (a) confer resistance to antibiotics or other toxins, e.g., ampicillin, neomycin, methotrexate, or tetracycline, (b) complement auxotrophic deficiencies, or (c) supply critical nutrients not available from complex media, e.g., the gene encoding D-alanine racemase for Bacilli.
[0337]An example of suitable selectable markers for mammalian cells are those that enable the identification of cells competent to take up the encoding nucleic acid, such as DHFR or thymidine kinase. An appropriate host cell when wild-type DHFR is employed is the CHO cell line deficient in DHFR activity, prepared and propagated as described by Urlaub et al., Proc. Natl. Acad. Sci. USA, 77:4216 (1980). A suitable selection gene for use in yeast is the trp1 gene present in the yeast plasmid YRp7 (Stinchcomb et al., Nature, 282:39 (1979); Kingsman et al., Gene, 7:141 (1979); Tschemper et al., Gene, 10:157 (1980)). The trp1 gene provides a selection marker for a mutant strain of yeast lacking the ability to grow in tryptophan, for example, ATCC No. 44076 or PEP4-1 (Jones, Genetics, 85:12 (1977)).
[0338]Expression and cloning vectors usually contain a promoter operably linked to the encoding nucleic acid sequence to direct mRNA synthesis. Promoters recognized by a variety of potential host cells are well known. Promoters suitable for use with prokaryotic hosts include the beta-lactamase and lactose promoter systems (Chang et al., Nature, 275:615 (1978); Goeddel et al., Nature, 281:544 (1979)), alkaline phosphatase, a tryptophan (trp) promoter system (Goeddel, Nucleic Acids Res., 8:4057 (1980); EP 36,776), and hybrid promoters such as the tac promoter (deBoer et al., Proc. Natl. Acad. Sci. USA, 80:21-25 (1983)). Promoters for use in bacterial systems also will contain a Shine-Dalgarno (S.D.) sequence operably linked to the encoding DNA.
[0339]Examples of suitable promoting sequences for use with yeast hosts include the promoters for 3-phosphoglycerate kinase (Hitzeman et al., J. Biol. Chem., 255:2073 (1980)) or other glycolytic enzymes (Hess et al., J. Adv. Enzyme Re.g., 7:149 (1968); Holland, Biochemistry, 17:4900 (1978)), such as enolase, glyceraldehyde-3-phosphate dehydrogenase, hexokinase, pyruvate decarboxylase, phosphofructokinase, glucose-6-phosphate isomerase, 3-phosphoglycerate mutase, pyruvate kinase, triosephosphate isomerase, phosphoglucose isomerase, and glucokinase.
[0340]Other yeast promoters, which are inducible promoters having the additional advantage of transcription controlled by growth conditions, are the promoter regions for alcohol dehydrogenase 2, isocytochrome C, acid phosphatase, degradative enzymes associated with nitrogen metabolism, metallothionein, glyceraldehyde-3-phosphate dehydrogenase, and enzymes responsible for maltose and galactose utilization. Suitable vectors and promoters for use in yeast expression are further described in EP 73,657.
[0341]Transcription from vectors in mammalian host cells is controlled, for example, by promoters obtained from the genomes of viruses such as polyoma virus, fowlpox virus (UK 2,211,504 published Jul. 5, 1989), adenovirus (such as Adenovirus 2), bovine papilloma virus, avian sarcoma virus, cytomegalovirus, a retrovirus, hepatitis-B virus and Simian Virus 40 (SV40), from heterologous mammalian promoters, e.g., the actin promoter or an immunoglobulin promoter, and from heat-shock promoters, provided such promoters are compatible with the host cell systems.
[0342]Transcription of a DNA encoding the stretches of consecutive amino acids or polypeptides of interest by higher eukaryotes may be increased by inserting an enhancer sequence into the vector. Enhancers are cis-acting elements of DNA, usually about from 10 to 300 bp, that act on a promoter to increase its transcription. Many enhancer sequences are now known from mammalian genes (globin, elastase, albumin, alpha-fetoprotein, and, insulin). Typically, however, one will use an enhancer from a eukaryotic cell virus. Examples include the SV40 enhancer on the late side of the replication origin (bp 100-270), the cytomegalovirus early promoter enhancer, the polyoma enhancer on the late side of the replication origin, and adenovirus enhancers. The enhancer may be spliced into the vector at a position 5' or 3' to the coding sequence, but is preferably located at a site 5' from the promoter.
[0343]Expression vectors used in eukaryotic host cells (yeast, fungi, insect, plant, animal, human, or nucleated cells from other multicellular organisms) will also contain sequences necessary for the termination of transcription and for stabilizing the mRNA. Such sequences are commonly available from the 5' and, occasionally 3', untranslated regions of eukaryotic or viral DNAs or cDNAs. These regions contain nucleotide segments transcribed as polyadenylated fragments in the untranslated portion of the mRNA encoding stretches of consecutive amino acids or polypeptides of interest.
[0344]Still other methods, vectors, and host cells suitable for adaptation to the synthesis of stretches of consecutive amino acids or polypeptides in recombinant vertebrate cell culture are described in Gething et al., Nature 293:620-625 (1981); Mantei et al., Nature, 281:4046 (1979); EP 117,060; and EP 117,058.
4. Detecting Gene Amplification/Expression
[0345]Gene amplification and/or expression may be measured in a sample directly, for example, by conventional Southern blotting, Northern blotting to quantitate the transcription of mRNA (Thomas, Proc. Natl. Acad. Sci. USA, 77:5201-5205 (1980)), dot blotting (DNA analysis), or in situ hybridization, using an appropriately labeled probe, based on the sequences provided herein. Alternatively, antibodies may be employed that can recognize specific duplexes, including DNA duplexes, RNA duplexes, and DNA-RNA hybrid duplexes or DNA-protein duplexes. The antibodies in turn may be labeled and the assay may be carried out where the duplex is bound to a surface, so that upon the formation of duplex on the surface, the presence of antibody bound to the duplex can be detected.
[0346]Gene expression, alternatively, may be measured by immunological methods, such as immunohistochemical staining of cells or tissue sections and assay of cell culture or body fluids, to quantitate directly the expression of gene product. Antibodies useful for immunohistochemical staining and/or assay of sample fluids may be either monoclonal or polyclonal, and may be prepared in any mammal. Conveniently, the antibodies may be prepared against a native sequence stretches of consecutive amino acids or polypeptides of interest or against a synthetic peptide based on the DNA sequences provided herein or against exogenous sequence fused to DNA encoding a stretch of consecutive amino acids or polypeptide of interest and encoding a specific antibody epitope.
5. Purification of Polypeptide
[0347]Forms of the stretches of consecutive amino acids or polypeptides of interest may be recovered from culture medium or from host cell lysates. If membrane-bound, it can be released from the membrane using a suitable detergent solution (e.g. Triton-X 100) or by enzymatic cleavage. Cells employed in expression of the stretches of consecutive amino acids or polypeptides of interest can be disrupted by various physical or chemical means, such as freeze-thaw cycling, sonication, mechanical disruption, or cell lysing agents.
[0348]It may be desired to purify the stretches of consecutive amino acids or polypeptides of interest from recombinant cell proteins or polypeptides. The following procedures are exemplary of suitable purification procedures: by fractionation on an ion-exchange column; ethanol precipitation; reverse phase HPLC; chromatography on silica or on a cation-exchange resin such as DEAE; chromatofocusing; SDS-PAGE; ammonium sulfate precipitation; gel filtration using, for example, Sephadex G-75; protein A Sepharose columns to remove contaminants such as IgG; and metal chelating columns to bind epitope-tagged forms. Various methods of protein purification may be employed and such methods are known in the art and described for example in Deutscher, Methods in Enzymology, 182 (1990); Scopes, Protein Purification: Principles and Practice, Springer-Verlag, New York (1982). The purification step(s) selected will depend, for example, on the nature of the production process used and the particular stretches of consecutive amino acids or polypeptides of interest produced.
Example of Expression of Stretch of Consecutive Amino Acids or Polypeptide Component of Interest in E. coli
[0349]The DNA sequence encoding the desired amino acid sequence of interest or polypeptide is initially amplified using selected PCR primers. The primers should contain restriction enzyme sites which correspond to the restriction enzyme sites on the selected expression vector. A variety of expression vectors may be employed. An example of a suitable vector is pBR322 (derived from E. coli; see Bolivar et al., Gene, 2:95 (1977)) which contains genes for ampicillin and tetracycline resistance. The vector is digested with restriction enzyme and dephosphorylated. The PCR amplified sequences are then ligated into the vector. The vector will preferably include sequences which encode for an antibiotic resistance gene, a trp promoter, a polyhis leader (including the first six STII codons, polyhis sequence, and enterokinase cleavage site), the specific amino acid sequence of interest/polypeptide coding region, lambda transcriptional terminator, and an argU gene.
[0350]The ligation mixture is then used to transform a selected E. coli strain using the methods described in Sambrook et al., supra. Transformants are identified by their ability to grow on LB plates and antibiotic resistant colonies are then selected. Plasmid DNA can be isolated and confirmed by restriction analysis and DNA sequencing.
[0351]Selected clones can be grown overnight in liquid culture medium such as LB broth supplemented with antibiotics. The overnight culture may subsequently be used to inoculate a larger scale culture. The cells are then grown to a desired optical density, during which the expression promoter is turned on.
[0352]After culturing the cells for several more hours, the cells can be harvested by centrifugation. The cell pellet obtained by the centrifugation can be solubilized using various agents known in the art, and the solubilized amino acid sequence of interest or polypeptide can then be purified using a metal chelating column under conditions that allow tight binding of the protein.
[0353]The primers can contain restriction enzyme sites which correspond to the restriction enzyme sites on the selected expression vector, and other useful sequences providing for efficient and reliable translation initiation, rapid purification on a metal chelation column, and proteolytic removal with enterokinase. The PCR-amplified, poly-His tagged sequences can be ligated into an expression vector used to transform an E. coli host based on, for example, strain 52 (W3110 fuhA(tonA) Ion galE rpoHts(htpRts) clpP(lacIq). Transformants can first be grown in LB containing 50 mg/ml carbenicillin at 30° C. with shaking until an O.D.600 of 3-5 is reached. Cultures are then diluted 50-100 fold into C RAP media (prepared by mixing 3.57 g (NH4)2 SO4, 0.71 g sodium citrate-2H2O, 1.07 g KCl, 5.36 g Difco yeast extract, 5.36 g Sheffield hycase SF in 500 mL water, as well as 110 mM MPOS, pH 7.3, 0.55% (w/v) glucose and 7 mM MgSO4) and grown for approximately 20-30 hours at 30° C. with shaking. Samples were removed to verify expression by SDS-PAGE analysis, and the bulk culture is centrifuged to pellet the cells. Cell pellets were frozen until purification and refolding.
[0354]E. coli paste from 0.5 to 1 L fermentations (6-10 g pellets) was resuspended in 10 volumes (w/v) in 7 M guanidine, 20 mM Tris, pH 8 buffer. Solid sodium sulfite and sodium tetrathionate is added to make final concentrations of 0.1M and 0.02 M, respectively, and the solution was stirred overnight at 4° C. This step results in a denatured protein with all cysteine residues blocked by sulfitolization. The solution was centrifuged at 40,000 rpm in a Beckman Ultracentifuge for 30 min. The supernatant was diluted with 3-5 volumes of metal chelate column buffer (6 M guanidine, 20 mM Tris, pH 7.4) and filtered through 0.22 micron filters to clarify. Depending the clarified extract was loaded onto a 5 mil Qiagen Ni-NTA metal chelate column equilibrated in the metal chelate column buffer. The column was washed with additional buffer containing 50 mM imidazole (Calbiochem, Utrol grade), pH 7.4. The protein was eluted with buffer containing 250 mM imidazole. Fractions containing the desired protein were pooled and stored at 4° C. Protein concentration was estimated by its absorbance at 280 nm using the calculated extinction coefficient based on its amino acid sequence.
Expression of Consecutive Stretches of Amino Acids in Mammalian Cells
[0355]This general example illustrates a preparation of a glycosylated form of a desired amino acid sequence of interest or polypeptide component by recombinant expression in mammalian cells.
[0356]The vector pRK5 (see EP 307,247, published Mar. 15, 1989) can be employed as the expression vector. Optionally, the encoding DNA is ligated into pRK5 with selected restriction enzymes to allow insertion of the DNA using ligation methods such as described in Sambrook et al., supra.
[0357]In one embodiment, the selected host cells may be 293 cells. Human 293 cells (ATCC CCL 1573) are grown to confluence in tissue culture plates in medium such as DMEM supplemented with fetal calf serum and optionally, nutrient components and/or antibiotics. About 10 μg of the ligated vector DNA is mixed with about 1 μg DNA encoding the VA RNA gene [Thimmappaya et al., Cell 31:543 (1982)] and dissolved in 500 μl of 1 mM Tris-HCl, 0.1 mM EDTA, 0.227 M CaCl2 To this mixture is added, dropwise, 500 μl of 50 mM HEPES (pH 7.35), 280 mM NaCl, 1.5 mM NaPO4, and a precipitate is allowed to form for 10 minutes at 25° C. The precipitate is suspended and added to the 293 cells and allowed to settle for about four hours at 37° C. The culture medium is aspirated off and 2 ml of 20% glycerol in PBS is added for 30 seconds. The 293 cells are then washed with serum free medium, fresh medium is added and the cells are incubated for about 5 days.
[0358]Approximately 24 hours after the transfections, the culture medium is removed and replaced with culture medium (alone) or culture medium containing 200 μCi/ml 35S-cysteine and 200 μCi/ml 35S-methionine. After a 12 hour incubation, the conditioned medium is collected, concentrated on a spin filter, and loaded onto a 15% SDS gel. The processed gel may be dried and exposed to film for a selected period of time to reveal the presence of amino acid sequence of interest or polypeptide component. The cultures containing transfected cells may undergo further incubation (in serum free medium) and the medium is tested in selected bioassays.
[0359]In an alternative technique, the nucleic acid amino acid sequence of interest or polypeptide component may be introduced into 293 cells transiently using the dextran sulfate method described by Somparyrac et al., Proc. Natl. Acad. Sci., 12:7575 (1981). 293 cells are grown to maximal density in a spinner flask and 700 μg of the ligated vector is added. The cells are first concentrated from the spinner flask by centrifugation and washed with PBS. The DNA-dextran precipitate is incubated on the cell pellet for four hours. The cells are treated with 20% glycerol for 90 seconds, washed with tissue culture medium, and re-introduced into the spinner flask containing tissue culture medium, 5 μg/ml bovine insulin and 0.1 μg/ml bovine transferrin. After about four days, the conditioned media is centrifuged and filtered to remove cells and debris. The sample containing expressed amino acid sequence of interest or polypeptide component can then be concentrated and purified by any selected method, such as dialysis and/or column chromatography.
[0360]In another embodiment, the amino acid sequence of interest or polypeptide component can be expressed in CHO cells. The amino acid sequence of interest or polypeptide component can be transfected into CHO cells using known reagents such as CaPO4 or DEAE-dextran. As described above, the cell cultures can be incubated, and the medium replaced with culture medium (alone) or medium containing a radiolabel such as 35S-methionine. After determining the presence of amino acid sequence of interest or polypeptide component, the culture medium may be replaced with serum free medium. Preferably, the cultures are incubated for about 6 days, and then the conditioned medium is harvested. The medium containing the expressed amino acid sequence of interest or polypeptide component can then be concentrated and purified by any selected method.
[0361]Epitope-tagged amino acid sequence of interest or polypeptide component may also be expressed in host CHO cells. The amino acid sequence of interest or polypeptide component may be subcloned out of a pRK5 vector. The subclone insert can undergo PCR to fuse in frame with a selected epitope tag such as a poly-his tag into a Baculovirus expression vector. The poly-his tagged amino acid sequence of interest or polypeptide component insert can then be subcloned into a SV40 driven vector containing a selection marker such as DHFR for selection of stable clones. Finally, the CHO cells can be transfected (as described above) with the SV40 driven vector. Labeling may be performed, as described above, to verify expression. The culture medium containing the expressed poly-His tagged amino acid sequence of interest or polypeptide component can then be concentrated and purified by any selected method, such as by Ni2+-chelate affinity chromatography.
[0362]In an embodiment the amino acid sequence of interest or polypeptide component are expressed as an IgG construct (immunoadhesin), in which the coding sequences for the soluble forms (e.g. extracellular domains) of the respective proteins are fused to an IgG1 constant region sequence containing the hinge, CH2 and CH2 domains and/or is a poly-His tagged form.
[0363]Following PCR amplification, the respective DNAs are subcloned in a CHO expression vector using standard techniques as described in Ausubel et al., Current Protocols of Molecular Biology, Unit 3.16, John Wiley and Sons (1997). CHO expression vectors are constructed to have compatible restriction sites 5' and 3' of the DNA of interest to allow the convenient shuttling of cDNA's. The vector used in expression in CHO cells is as described in Lucas et al., Nucl. Acids Res. 24:9 (1774-1779 (1996), and uses the SV40 early promoter/enhancer to drive expression of the cDNA of interest and dihydrofolate reductase (DHFR). DHFR expression permits selection for stable maintenance of the plasmid following transfection.
Expression of Stretch of Consecutive Amino Acids in Yeast
[0364]The following method describes recombinant expression of a desired amino acid sequence of interest or polypeptide component in yeast.
[0365]First, yeast expression vectors are constructed for intracellular production or secretion of a stretch of consecutive amino acids from the ADH2/GAPDH promoter. DNA encoding a desired amino acid sequence of interest or polypeptide component, a selected signal peptide and the promoter is inserted into suitable restriction enzyme sites in the selected plasmid to direct intracellular expression of the amino acid sequence of interest or polypeptide component. For secretion, DNA encoding the stretch of consecutive amino acids can be cloned into the selected plasmid, together with DNA encoding the ADH2/GAPDH promoter, the yeast alpha-factor secretory signal/leader sequence, and linker sequences (if needed) for expression of the stretch of consecutive amino acids.
[0366]Yeast cells, such as yeast strain AB110, can then be transformed with the expression plasmids described above and cultured in selected fermentation media. The transformed yeast supernatants can be analyzed by precipitation with 10% trichloroacetic acid and separation by SDS-PAGE, followed by staining of the gels with Coomassie Blue stain.
[0367]Recombinant amino acid sequence of interest or polypeptide component can subsequently be isolated and purified by removing the yeast cells from the fermentation medium by centrifugation and then concentrating the medium using selected cartridge filters. The concentrate containing the amino acid sequence of interest or polypeptide component may further be purified using selected column chromatography resins.
Expression of Stretches of Consecutive Amino Acids in Baculovirus-Infected Insect Cells
[0368]The following method describes recombinant expression of stretches of consecutive amino acids in Baculovirus-infected insect cells.
[0369]The desired nucleic acid encoding the stretch of consecutive amino acids is fused upstream of an epitope tag contained with a baculovirus expression vector. Such epitope tags include poly-his tags and immunoglobulin tags (like Fc regions of IgG). A variety of plasmids may be employed, including plasmids derived from commercially available plasmids such as pVL1393 (Novagen). Briefly, the amino acid sequence of interest or polypeptide component or the desired portion of the amino acid sequence of interest or polypeptide component (such as the sequence encoding the extracellular domain of a transmembrane protein) is amplified by PCR with primers complementary to the 5' and 3' regions. The 5' primer may incorporate flanking (selected) restriction enzyme sites. The product is then digested with those selected restriction enzymes and subcloned into the expression vector.
[0370]Recombinant baculovirus is generated by co-transfecting the above plasmid and BaculoGold® virus DNA (Pharmingen) into Spodoptera frugiperda ("Sf9") cells (ATCC CRL 1711) using lipofectin (commercially available from GIBCO-BRL). After 4-5 days of incubation at 28° C., the released viruses are harvested and used for further amplifications. Viral infection and protein expression is performed as described by O'Reilley et al., Baculovirus expression vectors: A laboratory Manual, Oxford: Oxford University Press (1994).
[0371]Expressed poly-his tagged amino acid sequence of interest or polypeptide component can then be purified, for example, by Ni2+-chelate affinity chromatography as follows. Extracts are prepared from recombinant virus-infected Sf9 cells as described by Rupert et al., Nature, 362:175-179 (1993). Briefly, Sf9 cells are washed, resuspended in sonication buffer (25 mL Hepes, pH 7.9; 12.5 mM MgCl2; 0.1 mM EDTA; 10% Glycerol; 0.1% NP40; 0.4 M KCl), and sonicated twice for 20 seconds on ice. The sonicates are cleared by centrifugation, and the supernatant is diluted 50-fold in loading buffer (50 mM phosphate, 300 mM NaCl, 10% Glycerol, pH 7.8) and filtered through a 0.45 μm filter. A Ni2+-NTA agarose column (commercially available from Qiagen) is prepared with a bed volume of 5 mL, washed with 25 mL of water and equilibrated with 25 mL of loading buffer. The filtered cell extract is loaded onto the column at 0.5 mL per minute. The column is washed to baseline A280 with loading buffer, at which point fraction collection is started. Next, the column is washed with a secondary wash buffer (50 mM phosphate; 300 mM NaCl, 10% Glycerol, pH 6.0), which elutes nonspecifically bound protein. After reaching A280 baseline again, the column is developed with a 0 to 500 mM Imidazole gradient in the secondary wash buffer. One mL fractions are collected and analyzed by SDS-PAGE and silver staining or western blot with Ni2+-NTA-conjugated to alkaline phosphatase (Qiagen). Fractions containing the eluted His10-tagged sequence are pooled and dialyzed against loading buffer.
[0372]Alternatively, purification of the IgG tagged (or Fc tagged) amino acid sequence can be performed using known chromatography techniques, including for instance, Protein A or Protein G column chromatography.
[0373]Immunoadhesin (Fc containing) constructs of proteins can be purified from conditioned media as follows. The conditioned media is pumped onto a 5 ml Protein A column (Pharmacia) which is equilibrated in 20 mM Na phosphate buffer, pH 6.8. After loading, the column is washed extensively with equilibration buffer before elution with 100 mM citric acid, pH 3.5. The eluted protein is immediately neutralized by collecting 1 ml fractions into tubes containing 275 mL of 1 M Tris buffer, pH 9. The highly purified protein is subsequently desalted into storage buffer as described above for the poly-His tagged proteins. The homogeneity of the proteins is verified by SDS polyacrylamide gel (PEG) electrophoresis and N-terminal amino acid sequencing by Edman degradation.
Intein-Based C-Terminal Syntheses
[0374]As described, for example, in U.S. Pat. No. 6,849,428, issued Feb. 1, 2005, inteins are the protein equivalent of the self-splicing RNA introns (see Perler et al., Nucleic Acids Res. 22:1125-1127 (1994)), which catalyze their own excision from a precursor protein with the concomitant fusion of the flanking protein sequences, known as exteins (reviewed in Perler et al., Curr. Opin. Chem. Biol. 1:292-299 (1997); Perler, F. B. Cell 92(1):1-4 (1998); Xu et al., EMBO J. 15(19):5146-5153 (1996)).
[0375]Studies into the mechanism of intein splicing led to the development of a protein purification system that utilized thiol-induced cleavage of the peptide bond at the N-terminus of the Sce VMA intein (Chong et al., Gene 192(2):271-281 (1997)). Purification with this intein-mediated system generates a bacterially-expressed protein with a C-terminal thioester (Chong et al., (1997)). In one application, where it is described to isolate a cytotoxic protein, the bacterially expressed protein with the C-terminal thioester is then fused to a chemically-synthesized peptide with an N-terminal cysteine using the chemistry described for "native chemical ligation" (Evans et al., Protein Sci. 7:2256-2264 (1998); Muir et al., Proc. Natl. Acad. Sci. USA 95:6705-6710 (1998)).
[0376]This technique, referred to as "intein-mediated protein ligation" (IPL), represents an important advance in protein semi-synthetic techniques. However, because chemically-synthesized peptides of larger than about 100 residues are difficult to obtain, the general application of IPL was limited by the requirement of a chemically-synthesized peptide as a ligation partner.
[0377]IPL technology was significantly expanded when an expressed protein with a predetermined N-terminus, such as cysteine, was generated, as described for example in U.S. Pat. No. 6,849,428. This allows the fusion of one or more expressed proteins from a host cell, such as bacterial, yeast or mammalian cells. In one non-limiting example the intein a modified RIR1 Methanobacterium thermoautotrophicum is that cleaves at either the C-terminus or N-terminus is used which allows for the release of a bacterially expressed protein during a one-column purification, thus eliminating the need proteases entirely.
[0378]Intein technology is one example of one route to obtain components. In one embodiment, the subunits of the compounds of the invention are obtained by transfecting suitable cells, capable of expressing and secreting mature chimeric polypeptides, wherein such polypeptides comprise, for example, an adhesin domain contiguous with an isolatable c-terminal intein domain (see U.S. Pat. No. 6,849,428, Evans et al., issued Feb. 1, 2005, hereby incorporated by reference). The cells, such as mammalian cells or bacterial cells, are transfected using known recombinant DNA techniques. The secreted chimeric polypeptide can then be isolated, e.g. using a chitin-derivatized resin in the case of an intein-chitin binding domain (see U.S. Pat. No. 6,897,285, Xu et al., issued May 24, 2005, hereby incorporated by reference), and is then treated under conditions permitting thiol-mediated cleavage and release of the now C-terminal thioester-terminated adhesion subunit. The thioester-terminated adhesion subunit is readily converted to a C-terminal cysteine terminated subunit.
[0379]These subunits can be treated under oxidizing conditions to permit formation of, for example, a disulfide bond between the two terminal cysteine residues, thus forming a symmetroadhesin. In addition, this technique can be used to make the symmetroadhesin-Fc hybrid subunits by treating the individual adhesion-Fc heterodimers under conditions permitting formation of bonds between the Fc portions of the heterodimers.
EXAMPLE 1
Preparation of Immunoglobulin Fc with N-Terminal-S-Termini (S-Fc)
[0380]Digestion of immunoglobulin (IgG) with papain yields two Fab fragments and one Fc fragment (Porter (1959) Biochem. 73, 119-126). The site of proteolysis in human IgG is the heavy chain hinge region between the cys-5 and cys-11 residues, EPKSCDKTHTCPPCP, (Fleischman et al., Biochem J. (1963) 88, 220-227; Edelman et al. (1969) Proc. Natl. Acad. Sci. 63, 78-85). The cys-5 residue normally forms a disulfide bond with the human IgG light chain, which is easily cleaved under mild reducing conditions, making it an ideal candidate for Fc-like molecules with N-terminal-S-termini (S-Fc).
[0381]Accordingly, host cells were transfected with expression vectors encoding IgG1 pre-Fc chimeric polypeptides consisting of a signal peptide joined at its C-terminus by a peptide bond to the N-terminus of an Fc domain beginning at the cys-5 residue, CDKTHTCPPCP (FIG. 35A). The heterologous signal peptides used are selected from proteins with N-terminal cysteines (part i). Thus, cleavage by the cellular signal peptidase will provide a mature S-Fc protein having cys-5 at the N-terminus (part ii).
[0382]The sequences of the IgG1 precursor polypeptides of FIG. 35A are shown in SEQ ID NO: 32, SEQ ID NO:33, and SEQ ID NO:34. The sequence of the mature IgG1 polypeptide of FIG. 35A is shown in SEQ ID NO: 35. Other mature IgG1 polypeptides made by methods described in EXAMPLES 1 to 5 are shown in SEQ ID NO: 36 through SEQ ID NO: 46.
[0383]Host cells are transfected with expression vectors encoding IgG2 pre-Fc chimeric polypeptides consisting of a signal peptide joined at its C-terminus by a peptide bond to the N-terminus of an Fc domain beginning at the cys-4 residue, CCVECPPCP (FIG. 35B). The heterologous signal peptides used are selected from proteins with N-terminal cysteines (part i). Thus, cleavage by the cellular signal peptidase will provide a mature S-Fc protein having cys-4 at the N-terminus (part ii).
[0384]The sequences of the IgG2 precursor polypeptides of FIG. 35B are shown in SEQ ID NO:50, SEQ ID NO:51, and SEQ ID NO:52. The sequence of the mature IgG2 polypeptide of FIG. 35B is shown in SEQ ID NO:53. Other mature IgG2 polypeptides made by methods described in EXAMPLES 1 to 5 are shown in SEQ ID NO:54 through SEQ ID NO:67.
[0385]Host cells are transfected with expression vectors encoding IgG3 pre-Fc chimeric polypeptides consisting of a signal peptide joined at its C-terminus by a peptide bond to the N-terminus of an Fc domain beginning at the cys-13 residue, CPRCP (FIG. 35C). The heterologous signal peptides used are selected from proteins with N-terminal cysteines (part i). Thus, cleavage by the cellular signal peptidase will provide a mature S-Fc protein having cys-13 at the N-terminus (part ii).
[0386]The sequences of the IgG3 precursor polypeptides of FIG. 35C are shown in SEQ ID NO:71, SEQ ID NO:72, and SEQ ID NO:73. The sequence of the mature IgG3 polypeptide of FIG. 35C is shown in SEQ ID NO:74. Other mature IgG2 polypeptides made by methods described in EXAMPLES 1 to 5 are shown in SEQ ID NO:75 through SEQ ID NO:82.
[0387]Host cells are transfected with expression vectors encoding IgG4 pre-Fc chimeric polypeptides consisting of a signal peptide joined at its C-terminus by a peptide bond to the N-terminus of an Fc domain beginning at the cys-8 residue, CPSCP (FIG. 35D). The heterologous signal peptides used are selected from proteins with N-terminal cysteines (part i). Thus, cleavage by the cellular signal peptidase will provide a mature S-Fc protein having cys-8 at the N-terminus (part ii).
[0388]The sequences of the IgG4 precursor polypeptides of FIG. 35D are shown in SEQ ID NO:86, SEQ ID NO:87, and SEQ ID NO:88. The sequence of the mature IgG4 polypeptide of FIG. 35D is shown in SEQ ID NO:89. Other mature IgG4 polypeptides made by methods described in EXAMPLES 1 to 5 are shown in SEQ ID NO:90 through SEQ ID NO:97.
[0389]Suitable host cells include 293 human embryonic cells (ATCC CRL-1573) and CHO-K1 hamster ovary cells (ATCC CCL-61) obtained from the American Type Culture Collection (Rockville, Md.). Cells are grown at 37° C. in an atmosphere of air, 95%; carbon dioxide, 5%. 293 cells are maintained in Minimal essential medium (Eagle) with 2 mM L-glutamine and Earle's BSS adjusted to contain 1.5 g/L sodium bicarbonate, 0.1 mM non-essential amino acids, and 1.0 mM sodium pyruvate, 90%; fetal bovine serum, 10%. CHO-K1 cells are maintained in Ham's F12K medium with 2 mM L-glutamine adjusted to contain 1.5 g/L sodium bicarbonate, 90%; fetal bovine serum, 10%. Other suitable host cells include CV1 monkey kidney cells (ATCC CCL-70), COS-7 monkey kidney cells (ATCC CRL-1651), VERO-76 monkey kidney cells (ATCC CRL-1587), HELA human cervical cells (ATCC CCL-2), W138 human lung cells (ATCC CCL-75), MDCK canine kidney cells (ATCC CCL-34), BRL3A rat liver cells (ATCC CRL-1442), BHK hamster kidney cells (ATCC CCL-10), MMT060562 mouse mammary cells (ATCC CCL-51), and human CD8.sup.+ T lymphocytes (described in U.S. Ser. No. 08/258,152 incorporated herein in its entirety by reference).
[0390]An example of a suitable expression vector is plasmid pSA (SEQ ID NO:1). Plasmid pSA contains the following DNA sequence elements: 1) pBluescriptIIKS(+) (nucleotides 912 to 2941/1 to 619, GenBank Accession No. X52327), 2) a human cytomegalovirus promoter, enhancer, and first exon splice donor (nucleotides 63 to 912, GenBank Accession No. K03104), 3) a human alpha1-globin second exon splice acceptor (nucleotides 6808 to 6919, GenBank Accession No. J00153), 4) an SV40 T antigen polyadenylation site (nucleotides 2770 to 2533, Reddy et al. (1978) Science 200, 494-502), and 5) an SV40 origin of replication (nucleotides 5725 to 5578, Reddy et al., ibid). For expression of the polypeptide of interest, an EcoRI-BglII DNA fragment encoding the polypeptide is inserted into plasmid pSA between the EcoRI and BglII restriction sites located at positions 1,608 and 1,632, respectively. Other suitable expression vectors include plasmids pSVeCD4DHFR and pRKCD4 (U.S. Pat. No. 5,336,603), plasmid pIK.1.1 (U.S. Pat. No. 5,359,046), plasmid pVL-2 (U.S. Pat. No. 5,838,464), plasmid pRT43.2F3 (described in U.S. Ser. No. 08/258,152 incorporated herein in its entirety by reference), and plasmid pCDNA3.1(+) (Invitrogen, Inc.).
[0391]Suitable selectable markers include the Tn5 transposon neomycin phosphotransferase (NEO) gene (Southern and Berg (1982) J. Mol. Appl. Gen. 1, 327-341), and the dihydrofolate reductase (DHFR) cDNA (Lucas et al. (1996) Nucl. Acids Res. 24, 1774-1779). One example of a suitable expression vector that incorporates a NEO gene is plasmid pSA-NEO, which is constructed by ligating a first DNA fragment, prepared by digesting SEQ ID NO:2 with EcoRI and BglII, with a second DNA fragment, prepared by digesting SEQ ID NO:1 with EcoRI and BglII. SEQ ID NO:2 incorporates a NEO gene (nucleotides 1551 to 2345, Genbank Accession No. U00004) preceded by a sequence for translational initiation (Kozak (1991) J. Biol. Chem, 266, 19867-19870). Another example of a suitable expression vector that incorporates a NEO gene and a DHFR cDNA is plasmid pSVe-NEO-DHFR, which is constructed by ligating a first DNA fragment, prepared by digesting SEQ ID NO:2 with EcoRI and BglII, with a second DNA fragment, prepared by digesting pSVeCD4DHFR with EcoRI and BglII. Plasmid pSVe-NEO-DHFR uses SV40 early promoter/enhancers to drive expression of the NEO gene and the DHFR cDNA. Other suitable selectable markers include the XPGT gene (Mulligan and Berg (1980) Science 209, 1422-1427) and the hygromycin resistance gene (Sugden et al. (1985) Mol. Cell. Biol. 5, 410-413).
[0392]Human IgG1 DNA sequences are described in Ellison et al. (1982) Nuc. Acids Res. 10, 4071-4079) (Genbank Acc. No. Z17370)
[0393]Suitable examples of signal peptides are sonic hedgehog (SHH) (GenBank Acc. No. NM--000193), interferona-2 (IFN) (GenBank Acc. No. NP--000596), and cholesterol ester transferase (CETP) (GenBank Accession No. NM--000078). Other suitable examples include Indian hedgehog (Genbank Acc. No. NM--002181), desert hedgehog (Genbank Acc. No. NM--021044), IFNα-1 (Genbank Acc. No. NP--076918), IFNα-4 (Genbank Acc. No. NM--021068), IFNα-5 (Genbank Acc. No. NM--002169), IFNα-6 (Genbank Acc. No. NM--021002), IFNα-7 (Genbank Acc. No. NM--021057), IFNα-8 (Genbank Acc. No. NM--002170), IFNα-10 (Genbank Acc. No. NM--002171), IFNα-13 (Genbank Acc. No. NM--006900), IFNα-14 (Genbank Acc. No. NM--002172), IFNα-16 (Genbank Acc. No. NM--002173), IFNα-17 (Genbank Acc. No. NM--021268) and IFNα-21 (Genbank Acc. No. NM--002175).
[0394]Suitable expression vectors are constructed by ligation of an insert and vector fragment prepared by digestion with EcoRI and BglII. For unamplified expression, plasmid pSHH-Fc5 (SHH signal) is constructed using SEQ ID NO:3 and SEQ ID NO:1, plasmid pIFN-Fc5 (IFN signal) is constructed using SEQ ID NO:4 and SEQ ID NO:1 and plasmid pCETP-Fc5 (CETP signal) is constructed using SEQ ID NO:3 and SEQ ID NO:1. For amplified expression, plasmid PSHH-Fc5-DHFR is constructed using SEQ ID NO:3 and pSVeCD4DHFR (U.S. Pat. No. 5,336,603), pIFN-Fc5-DHFR is constructed using SEQ ID NO:4 and pSVeCD4DHFR, and pCETP-Fc5-DHFR is constructed using SEQ ID NO:5 and pSVeCD4DHFR.
[0395]Suitable expression vectors for human IgG1 Fc polypeptides were constructed by ligation of an insert fragment prepared by digestion with Hind III and EagI and a vector fragment prepared by digestion with Hind III and PspOM1. Plasmid pCDNA3-SHH-IgG1-Fc (SHH signal) was constructed using SEQ ID NO:29 and pCDNA3.1(+), plasmid pCDNA3-IFN-IgG1-Fc (IFN signal) was constructed using SEQ ID NO:30 and pCDNA3.1(+), and plasmid pCDNA3-IgG1-Fc (CETP signal) was constructed using SEQ ID NO:31 and pCDNA3.1(+).
[0396]Suitable expression vectors for human IgG2 Fc polypeptides are constructed by ligation of an insert fragment prepared by digestion with Hind III and EagI and a vector fragment prepared by digestion with Hind III and PspOM1. Plasmid pCDNA3-SHH-IgG2-Fc (SHH signal) is constructed using SEQ ID NO:47 and pCDNA3.1(+), plasmid pCDNA3-IFN-IgG2-Fc (IFN signal) is constructed using SEQ ID NO:48 and pCDNA3.1(+), and plasmid pCDNA3-IgG2-Fc (CETP signal) is constructed using SEQ ID NO:49 and pCDNA3.1(+).
[0397]Suitable expression vectors for human IgG3 Fc polypeptides are constructed by ligation of an insert fragment prepared by digestion with Hind III and EagI and a vector fragment prepared by digestion with Hind III and PspOM1. Plasmid pCDNA3-SHH-IgG3-Fc (SHH signal) is constructed using SEQ ID NO:68 and pCDNA3.1(+), plasmid pCDNA3-IFN-IgG3-Fc (IFN signal) is constructed using SEQ ID NO:69 and pCDNA3.1(+), and plasmid pCDNA3-IgG3-Fc (CETP signal) is constructed using SEQ ID NO:70 and pCDNA3.1(+).
[0398]Suitable expression vectors for human IgG4 Fc polypeptides are constructed by ligation of an insert fragment prepared by digestion with Hind III and EagI and a vector fragment prepared by digestion with Hind III and PspOM1. Plasmid pCDNA3-SHH-IgG4-Fc (SHH signal) is constructed using SEQ ID NO:83 and pCDNA3.1(+), plasmid pCDNA3-IFN-IgG4-Fc (IFN signal) is constructed using SEQ ID NO:84 and pCDNA3.1(+), and plasmid pCDNA3-IgG4-Fc (CETP signal) is constructed using SEQ ID NO:85 and pCDNA3.1(+).
[0399]In one embodiment, cells are transfected by the calcium phosphate method of Graham et al. (1977) J. Gen. Virol. 36, 59-74. A DNA mixture (10 micrograms) is dissolved in 0.5 ml of 1 mM Tris-HCl, 0.1 mM EDTA, and 227 mM CaCl2. The DNA mixture contains (in a ratio of 10:1:1) the expression vector DNA, the selectable marker DNA, and a DNA encoding the VA RNA gene (Thimmappaya et al. (1982) Cell 31, 543-551). To this mixture is added, dropwise, 0.5 mL of 50 mM Hepes (pH 7.35), 280 mM NaCl, and 1.5 mM NaPO4. The DNA precipitate is allowed to form for 10 minutes at 25° C., then suspended and added to cells grown to confluence on 100 mm plastic tissue culture dishes. After 4 hours at 37° C., the culture medium is aspirated and 2 ml of 20% glycerol in PBS is added for 0.5 minutes. The cells are then washed with serum-free medium, fresh culture medium is added, and the cells are incubated for 5 days.
[0400]In another embodiment, cells are transiently transfected by the dextran sulfate method of Somparyrac et al. (1981) Proc. Nat. Acad. Sci. 12, 7575-7579. Cells are grown to maximal density in spinner flasks, concentrated by centrifugation, and washed with PBS. The DNA-dextran precipitate is incubated on the cell pellet. After 4 hours at 37° C., the DEAE-dextran is aspirated and 20% glycerol in PBS is added for 1.5 minutes. The cells are then washed with serum-free medium, and re-introduced into spinner flasks containing fresh culture medium with 5 micrograms/ml bovine insulin and 0.1 micrograms/ml bovine transferring, and the cells are incubated for 4 days. Following transfection by either method, the conditioned media is centrifuged and filtered to remove the host cells and debris. The sample contained the S-Fc domain is then concentrated and purified by any selected method, such as dialysis and/or column chromatography (see below).
[0401]To identify the S-Fc in the cell culture supernatant, the culture medium is removed 24 hours after transfection and replaced with culture medium containing 200 microCi/ml each of 35S-methionine and 35S-cysteine. After a 12 hour incubation, the conditioned medium is collected by centrifugation to remove the host cells and debris, concentrated on a spin dialysis filter. The labeled supernatants is analyzed by immunoprecipitation with protein A sepharose beads in the absence of added antibodies. The precipitated proteins are analyzed on 7.5% polyacrylamide-SDS gels either with or without reduction with β-mercaptoethanol. The processed gel is dried and exposed to x-ray film to reveal the presence of the S-Fc domain.
[0402]For unamplified expression, plasmids pSHH-Fc-5, pIFN-Fc-5 and pCETP-Fc-5 are transfected into human 293 cells (Graham et al., J. Gen. Virol. 36:59 74 (1977)), using a high efficiency procedure (Gorman et al., DNA Prot. Eng. Tech. 2:3 10 (1990)). Media is changed to serum-free and harvested daily for up to five days. The S-Fc proteins are purified from the cell culture supernatant using protein A-Sepharose CL-4B (Pharmacia). The eluted S-Fc protein is buffer-exchanged into PBS using a Centricon-30 (Amicon), concentrated to 0.5 ml, sterile filtered using a Millex-GV (Millipore) and stored at 4° C.
[0403]For unamplified expression, plasmids pCDNA3-SHH-IgG1-Fc, pCDA3-IFN-IgG1-Fc and pCDA-3-CETP-IgG1-Fc were transfected into human 293 cells (Graham et al., J. Gen. Virol. 36:59 74 (1977)), using a high efficiency procedure (Gorman et al., DNA Prot. Eng. Tech. 2:3 10 (1990)). Media was changed to serum-free and harvested daily for up to five days. The S-Fc proteins were purified from the cell culture supernatant using protein A-Sepharose CL-4B (Pharmacia). The eluted S-Fc protein was buffer-exchanged into PBS using a Centricon-30 (Amicon), concentrated to 0.5 ml, sterile filtered using a Millex-GV (Millipore) and stored at 4° C.
[0404]FIG. 52 shows expression in 293 kidney cells of human IgG1 Fc symmetroadhesin subunits with N-terminal-S-termini. Lanes 1-6 and lanes 7-12 show the IgG1 Fc polypeptides of FIG. 35A (ii) and FIG. 36A (ii), respectively. Cell supernatants: lanes 1, 3, 5, 7, 9 and 11; cell lysates: lanes 2, 4, 6, 8, 10 and 12. Signal sequences used: SHH (lanes 1, 2, 7 and 8); IFNα (lanes 3, 4, 9, 10); CETP (lanes 5, 6, 11 and 12).
[0405]FIG. 53 shows expression in 293 kidney cells of human IgG1 Fc symmetroadhesin subunits. Lanes 1-2, 3-4 and 5-6 show the IgG1 Fc polypeptides of FIG. 35A (ii), FIG. 36A (ii) and FIG. 37B (ii), respectively. Cell supernatants: (lanes 1-6). Signal sequences used: SHH (lanes 1-6).
[0406]FIG. 54 shows Protein A purification of human IgG1 Fc symmetroadhesin subunits expressed in 293 kidney cells. Lane 2 and 8 show the IgG1 Fc polypeptides of FIG. 36A and FIG. 35A, respectively. Lanes 1-7: proteinA-sepharose column fractions for the IgG1 Fc polypeptide of FIG. 36A.
[0407]FIG. 55 shows Thiol-sepharose binding of proteinA-purified human IgG1 Fc symmetroadhesin subunits shown in FIG. 54. Lanes 1-3 and lanes 4-6 show the human IgG1 Fc polypeptides of FIG. 35A and FIG. 36A, respectively. Lanes 1 and 4: starting material; lanes 2 and 5: thiol-sepharose flow-thru fraction; lanes 3 and 6: thiol-sepharose bound fraction.
[0408]Analysis of the two IgG1 Fc polypeptides of FIG. 35A and FIG. 36A by mass spectrometry (MALDI) reveals average molecular masses of 54,552.85 and 53,173.43 daltons, respectively. Assuming each molecular mass represents the corresponding Fc dimers, the apparent difference in molecular mass between the two polypeptides (1,379.4 daltons) is in good agreement (0.6% deviation) with the predicted difference in molecular mass (1,371.5).
[0409]For amplified expression, Chinese hamster ovary (CHO) cells are transfected with dicistronic vectors pSHH-Fc5-DHFR, pIFN-Fc-5-DHFR and pCETP-Fc-5-DHFR which co-express a DHFR cDNA. Plasmids are introduced into CHO-K1 DUX B11 cells developed by L. Chasin (Columbia University) via lipofection and selected for growth in GHT-free medium (Chisholm (1996) High efficiency gene transfer in mammalian cells. In: Glover, D M, Hames, B D. DNA Cloning vol 4. Mammalian systems. Oxford Univ. Press, pp 1-41). Approximately 20 unamplified clones are randomly chosen and reseeded into 96 well plates. Relative specific productivity of each colony is monitored using an ELISA to quantitate the S-Fc protein accumulated in each well after 3 days and a fluorescent dye, Calcien AM, as a surrogate marker of viable cell number per well. Based on these data, several unamplified clones are chosen for further amplification in the presence of increasing concentrations of methotrexate. Individual clones surviving at 10, 50, and 100 nM methotrexate are chosen and transferred to 96 well plates for productivity screening. Suitable clones, which reproducibly exhibit high specific productivity, are expanded in T-flasks and used to inoculate spinner cultures. After several passages, the suspension-adapted cells are used to inoculate production cultures in GHT-containing, serum-free media supplemented with various hormones and protein hydrolysates. Harvested cell culture fluid containing the S-Fc protein is purified using protein A-Sepharose CL-4B.
EXAMPLE 2
Preparation of Immunoglobulin Fc with N-Terminal-X-Termini (X-Fc)
[0410]Selenocysteine (sec) is the 21st amino acid incorporated during ribosome-mediated protein synthesis (Zinoni et al. (1986) Proc. Natl. Acad. Sci. 83, 4650-4654; Chambers et al. (1986) EMBO J. 5, 1221-1227). The process is complex and distinct from cysteine incorporation, requiring an mRNA selenocysteine insertion element in order to decode a UGA stop codon. Protein semi-synthesis offers an alternative means for preparing Fc-like molecules having N-terminal-X-termini (X-Fc) that begin at cysteine (cys) and/or selenocysteine (sec).
[0411]Accordingly, host cells are transfected with constructs that encode pre-Fc chimeric polypeptides consisting of a signal peptide joined at its C-terminus by a peptide bond to the N-terminus of an Fc domain beginning at cys-11, CDKTHTCPPCP (FIG. 36A) and cys-14 CDKTHTCPPCP (FIG. 36B). The heterologous signal peptides used are selected from proteins with N-terminal cysteines (part i). Thus, cleavage by the cellular signal peptidase will provide a mature S-Fc protein having cys-11 at the N-terminus (part ii). Native chemical ligation is then employed to prepare mature X-Fc proteins having cys-5 or sec-5 at the N-terminus, XDKTHTCPPCP (part iii).
[0412]The sequences of the IgG1 precursor polypeptides of FIG. 36A are shown in SEQ ID NO: 101, SEQ ID NO:102, and SEQ ID NO:103. The sequence of the mature IgG1 polypeptide of FIG. 35A is shown in SEQ ID NO: 104. The sequences of the IgG1 precursor polypeptides of FIG. 36B are shown in SEQ ID NO: 109, SEQ ID NO:110, and SEQ ID NO:111. The sequence of the mature IgG1 polypeptide of FIG. 35B is shown in SEQ ID NO: 112.
[0413]Accordingly, host cells are transfected with constructs that encode pre-Fc chimeric polypeptides consisting of a signal peptide joined at its C-terminus by a peptide bond to the N-terminus of an Fc domain beginning at cys-14, CDKTHTCPPCP (FIG. 36B). The heterologous signal peptides used are selected from proteins with N-terminal cysteines (part i). Thus, cleavage by the cellular signal peptidase will provide a mature S-Fc protein having cys-11 at the N-terminus (part ii). Native chemical ligation is then employed to prepare mature X-Fc proteins having cys-11 or sec-11 at the N-terminus, XPPCP (part iii).
[0414]The S-Fc proteins with cys-11 and cys-14 at the N-terminus is first prepared using the procedures described in EXAMPLE 1. Native chemical ligation is carried out with S-Fc protein and peptide Fc-A (5-11: cys-asp-lys-thr-his-thr), or S-Fc protein and peptide Fc-B (5-11: sec-asp-lys-thr-his-thr). Suitable expression vectors are constructed by ligation of an insert and vector fragment prepared by digestion with EcoRI and BglII. For unamplified expression, plasmid pSHH-Fc11 (SHH signal) is constructed using SEQ ID NO:6 and SEQ ID NO:1, plasmid pIFN-Fc11 (IFN signal) is constructed using SEQ ID NO 7 and SEQ ID NO:1 and plasmid pCETP-Fc11 (CETP signal) is constructed using SEQ ID NO:8 and SEQ ID NO:1. For amplified expression, plasmid pSHH-Fc11-DHFR is constructed using SEQ ID NO:6 and pSVeCD4DHFR, plasmid pIFN-Fc11-DHFR is constructed using SEQ ID NO:7 and pSVeCD4DHFR, and plasmid pCETP-Fc11-DHFR is constructed using SEQ ID NO:8 and pSVeCD4DHFR.
[0415]Suitable expression vectors for human IgG1 Fc are constructed by ligation of an insert fragment prepared by digestion with Hind III and EagI and a vector fragment prepared by digestion with Hind III and PspOM1. Plasmid pCDNA3-SHH-IgG1-Fc11 (SHH signal) is constructed using SEQ ID NO:98 and pCDNA3.1(+), plasmid pCDNA3-IFN-IgG1-Fc11 (IFN signal) is constructed using SEQ ID NO:99 and pCDNA3.1(+), and plasmid pCDNA3-IgG1-Fc11 (CETP signal) is constructed using SEQ ID NO:100 and pCDNA3.1(+). Plasmid pCDNA3-SHH-IgG1-Fc14 (SHH signal) is constructed using SEQ ID NO:106 and pCDNA3.1(+), plasmid pCDNA3-IFN-IgG1-Fc14 (IFN signal) is constructed using SEQ ID NO:107 and pCDNA3.1(+), and plasmid pCDNA3-IgG1-Fc14 (CETP signal) is constructed using SEQ ID NO:108 and pCDNA3.1(+).
[0416]General principles of native chemical ligation are described in U.S. Pat. No. 6,184,344, the whole of which is incorporated herein by reference. Peptides Fc-A and Fc-B are synthesized using a TAMPAL resin from which any desired thioester can be readily obtained. After de-protection of side chain protecting groups, the resulting C-terminal activated peptides are used in native chemical ligation without further modification. To prevent the cyclization or polymerization of bifunctional peptides, the sulfhydryl moiety (peptide Fc-A) and the selenohydryl moiety (peptide Fc-B) are reversibly blocked with Msc.
[0417]Peptide synthesis is carried out by manual a solid-phase procedure using an in situ neutralization/HBTU activation procedure for Boc chemistry (Schnolzer et al. (1992) Int. J. Pept. Protein Res. 40, 180-193). After each coupling step, yields are determined by measuring residual free amine with the quantitative ninhydrin assay (Sarin et al. (1981) Anal. Biochem. 117, 147-157). Side chain protected amino acids are Boc-Asp(O-cyclohexyl)-OH, Boc-Cys(4-methylbenzyl)-OH, Boc-Lys(2-Cl-Z)-OH, and Boc-Thr(benzyl)-OH. After chain assembly is completed, peptides are deprotected and cleaved from the resin by treatment with anhydrous HF for 1 hour at 0° C. with 4% anisole as scavenger.
[0418]Peptides Fc-A and Fc-B are synthesized on trityl-associated mercaptopropionic acid leucine (TAMPAL) resin to yield C-terminal MPAL-activated thioesters (Hackeng et al. (1999) Proc. Natl. Acad. Sci. 96, 10068-10073). The N-terminal cys/sec residues of the thioester peptides are protected with 2-(methylsulfonyl)ethyl carbonate (Msc) groups in a 2 hour reaction (10-fold excess) of activated Msc nitrophenol ester dissolved in a minimal volume of dimethylformamide/5% diisopropylethylamine. The thioester-activated peptides are deprotected and cleaved from the TAMPAL resin, HPLC-purified, lyophilized, and stored until use at -20° C. Preparative reversed-phase HPLC is performed using a Vydac C-18 column (10 micrometer, 1.0 cm×25 cm). The bound peptides are eluted using a linear gradient of acetonitrile in H20/10% trifluoroacetic acid.
[0419]Starting with the purified C-terminal S-Fc protein, native chemical ligation is carried out with the Msc-NH-cys5-thr10-α-thioester peptide (Fc-A) or the Msc-NH-sec5-thr10-α-thioester peptide (Fc-B) under non-denaturing conditions as previously described (Evans et al. (1999) J. Biol. Chem. 274, 3923-3926). The thioester-activated peptides are mixed in molar excess with freshly prepared S-Fc protein (starting concentration, 1-200 micromolar). The solution is concentrated with a Centriprep 3/30 apparatus (Millipore, Mass.), then with a Centricon 3/10 apparatus to a final concentration of 0.15 to 1.2 mM for the S-Fc protein. Ligations are incubated overnight at 4° C. and visualized using SDS-page electrophoresis. After native chemical ligation, the N-terminal Msc protecting group is removed by a brief incubation (<5 minute) at pH 13. The X-Fc product is purified to remove unreacted peptides by affinity chromatography with protein A sepharose using the procedure in EXAMPLE 1.
[0420]The sequence of the native ligation product of FIG. 36A is shown in SEQ ID NO: 105. The sequence of the native ligation product of FIG. 36B is shown in SEQ ID NO: 113.
EXAMPLE 3
Preparation of Immuoglobulin Fc with C-Terminal-X-Termini (Fc-X)
[0421]IgG is expressed in two abundant forms, the soluble antibody molecule and the cell-bound B-cell receptor. Both forms arise from a single messenger RNA by alternative splicing with the result that two additional exons are added to the IgG heavy chain coding region (Tyler et al. (1982) Proc. Natl. Acad. Sci. 79, 2008-2012; Yamawaki-Kataoka et al. (1982) Proc. Natl. Acad. Sci. 79, 2623-2627). The first exon added (M1 exon) encodes a stretch of 18 amino acids, ELQLEESCAEAQDGELDG, which flexibly tethers IgG to the cell surface, making it a good choice for novel Fc-like molecules with a C-terminal-X-terminus (Fc-X). The C-terminal gly-18 residue of the M1 domain is also well-suited for preparing Fc-intein fusion proteins used in generating a C-terminal activated thioester. Following an intein autocleavage reaction, a thioester intermediate is generated that permits the facile addition of cysteine or selenocysteine to the C-terminus by native chemical ligation.
[0422]Accordingly, host cells are transfected with expression vectors that encode pre-Fc-intein chimeric polypeptides containing the M1 domain joined at its C-terminus, ELQLEESCAEAQDGELDG, by a peptide bond to the N-terminus of a self-splicing intein at the autocleavage site (FIG. 37A), or the Fc protein containing a portion of the M1 domain, ELQLEESC (FIG. 37B). To ensure that the Fc-X protein does not have an N-terminal-X-terminus, heterologous signal peptides are used that are cleaved by the cellular signal peptidase before lysine residues (part i). Thus, cleavage by the cellular signal peptidase will provide a Fc-intein fusion protein with lys-7, EPKSCDKTHTCPPCP, at the N-terminus (part ii). Excision of the intein domain by protein splicing provides an Fc-thioester intermediate (part iii). Finally, native chemical ligation of the Fc-thioester with free cysteine and/or selenocysteine is employed to prepare Fc-X proteins having the C-terminal-X-terminus, ELQLEESCAEAQDGELDGX (part iv).
[0423]The sequences of the IgG1 precursor polypeptides of FIG. 37A are shown in SEQ ID NO:116, and SEQ ID NO:117. The sequence of the mature and modified IgG1 polypeptides of FIG. 37A is shown in SEQ ID NO:118 to SEQ ID NO:120. The sequences of the IgG1 precursor polypeptides of FIG. 37B are shown in SEQ ID NO:123, and SEQ ID NO:124. The sequence of the mature IgG1 polypeptide of FIG. 37B is shown in SEQ ID NO:125.
[0424]The Fc-intein fusion protein with lys-7 at the N-terminus is prepared using the procedures described in EXAMPLE 1. The initial purification step is carried out using affinity chromatography with a chitin resin instead of protein A sepharose. After cleavage from the resin, the activated Fc-thioester intermediate is used directly for native ligation with cysteine and/or selenocysteine.
[0425]A suitable DNA sequence for the M1 membrane domain of human IgG1 is described in Strausberg et. al. (2002) Proc. Natl. Acad. Sci. 99, 16899-16903 (GenBank Acc. No. BC019046).
[0426]Suitable examples of signal peptides are the CD2 T-cell surface glycoprotein (CD2) (GenBank Acc. No. NM--001767), and the CD4 T-cell surface glycoprotein (CD4) (GenBank Acc. No. NP--000616).
[0427]A suitable example of a self-splicing intein is found in the Methanobacterium thermoautotrophicum ribonucleotide reductase large subunit (MthRIR1) (Genbank Acc. No. AE000845). To limit the intein autocleave reaction to the Fc-intein fusion junction, an MthRR1 intein variant with only N-terminal cleavage activity is prepared by changing the pro at position -1 to gly and the C-terminal asn at position 134 to ala (Evans et al. (1999) J. Biol. Chem. 274, 3923-3926). In addition, an Mth RRI intein sequence fused to a Bacillus circulans chitin binding domain is used to facilitate the purification of the Fc-intein chimeric polypeptide by affinity chromatography. A suitable sequence for the modified MthRIR1 intein is found in plasmid pTWIN-2 (New England BioLabs, MA).
[0428]Other suitable examples are found in the Mycobacterium xenopi gyrase subunit A (Mxe GyrA) (Genbank Acc. No. MXU67876), and the Saccharomyces cerevisiae vacuolar ATPase (Sce VMA1) (GenBank Acc. No. NC--001136). Many other suitable examples of self-splicing inteins are described in Inbase: the Intein Database (Perler (2002) Nucl. Acids Res. 30, 383-384).
[0429]Suitable expression vectors are constructed by ligation of an insert and vector fragment prepared by digestion with EcoRI and BglII. For unamplified expression, plasmid pCD2-Fc7-Mth (CD2 signal) is constructed with SEQ ID NO:9 and SEQ ID NO:1, and plasmid pCD4-Fc7-Mth (CD4 signal) is constructed with SEQ ID NO:10 and SEQ ID NO:1. For amplified expression, plasmid pCD2-Fc7-Mth-DHFR is constructed with SEQ ID NO:9 and pSVeCD4DHFR, and plasmid pCD4-Fc7-Mth-DHFR is constructed with SEQ ID NO:10 and pSVeCD4DHFR.
[0430]Suitable expression vectors are constructed by ligation of an insert fragment prepared by digestion with Hind III and EagI and a vector fragment prepared by digestion with Hind III and PspOM1. Plasmid pCDNA3-CD2-Fc7-Mth (CD2 signal) is constructed with SEQ ID NO:114 and pCDNA3.1(+), and plasmid pCDNA3-CD4-Fc7-Mth (CD4 signal) is constructed with SEQ ID NO:115 and pCDNA3.1(+). Plasmid pCDNA3-CD2-Fc7-ELQLEESC (CD2 signal) is constructed with SEQ ID NO:121 and pCDNA3.1(+), and plasmid pCDNA3-CD4-Fc7-ELQLEESC (CD4 signal) is constructed with SEQ ID NO:122 and pCDNA3.1(+).
[0431]General principles of chitin affinity purification and the self-splicing intein autocleavage reaction are described in U.S. Pat. No. 5,834,247, the whole of which is incorporated herein by reference.
[0432]Following host cell transfection, cell culture supernatant is applied to a column packed with chitin resin (New England BioLabs, MA) that is equilibrated in buffer A (20 mM Tris-HCl, pH 7.5 containing 500 mM NaCl). Unbound protein is washed from the column with 10 column volumes of buffer A. Thiol reagent-induced cleavage is initiated by rapidly equilibrating the chitin resin in buffer B (20 mM Tris-HCl, pH 8 containing 0.5 M NaCl and 0.1 M 2-mercaptoethane-sulfonic acid (MESNA)). The cleavage, which simultaneously generates a C-terminal thioester on the target protein, is carried out overnight at 4° C. after which the protein was eluted from the column.
[0433]Starting with the purified Fc-thioester intermediate, native chemical ligation is carried out with cysteine or selenocysteine using the procedure in EXAMPLE 2. The final Fc-X product is purified to remove unreacted cysteine and selenocysteine by affinity chromatography with protein A sepharose.
EXAMPLE 4
Preparation of Immunoglobulin Fc with N-Terminal-S-Termini and C-Terminal-X-Termini (S-Fc-X)
[0434]The S-Fc and Fc-X proteins are useful in preparing immunosymmetroadhesins having two binding domains joined to a single Fc domain (see below). Binding domains are added to the N-terminal-S-termini (S-Fc) or C-terminal-X-termini (Fc-X). S-Fc-X domains that are useful in preparing bi-symmetroadhesins with four binding domains joined to a single Fc domain are prepared using the procedures described in EXAMPLE 1 and EXAMPLE 3.
[0435]Accordingly, host cells are transfected with expression vectors that encode pre-Fc-intein chimeric polypeptides containing the M1 domain joined at its C-terminus, ELQLEESCAEAQDGELDG, by a peptide bond to the N-terminus of a self-splicing intein at the autocleavage site (FIG. 38). Heterologous signal peptides are selected from proteins with N-terminal cysteines (part i). Thus, cleavage by the cellular signal peptidase will provide a mature S-Fc-intein fusion protein having cys-5 at the N-terminus, CDKTHTCPPCP (part ii). Excision of the intein by protein splicing provides an Fc-thioester intermediate (part iii). Finally, native ligation of the Fc-thioester with free cysteine and/or selenocysteine is employed to prepare S-Fc-X proteins having the C-terminal-X-terminus, ELQLEESCAEAQDGELDGX (part iv).
[0436]The sequences of the IgG1 precursor polypeptides of FIGS. 38A-38B are shown in SEQ ID NO:126, SEQ ID NO:127, and SEQ ID NO:128. The sequence of the mature and modified IgG1 polypeptides of FIGS. 38A-38B is shown in SEQ ID NO:129 to SEQ ID NO:131.
[0437]Suitable expression vectors are constructed by ligation of an insert and vector fragment prepared by digestion with EcoRI and BglII. For unamplified expression, PSHH-Fc5-Mth is constructed with SEQ ID NO:11 and SEQ ID NO:1, pIFN-Fc5-Mth is constructed with SEQ ID NO:12 and SEQ ID NO:1, and pCETP-Fc5-Mth is constructed with SEQ ID NO:13 and SEQ ID NO:1. For amplified expression, pSHH-Fc5-Mth-DHFR is constructed with SEQ ID NO:11 and pSVeCD4DHFR, pIFN-Fc5-Mth-DHFR is constructed with SEQ ID NO:12 and pSVeCD4DHFR, and pCETP-Fc5-Mth-DHFR is constructed with SEQ ID NO:13 and pSVeCD4DHFR.
[0438]The S-Fc-intein fusion protein is first purified from the culture supernatant of transfected host cells using chitin affinity chromatography. After cleavage from the chitin resin, the Fc-thioester intermediate is directly applied to a protein A Sepharose column in order to prevent cyclization or polymerization side-reactions. The column bound activated S-Fc-thioester is used directly for native ligation with cysteine and/or selenocysteine. The column is then washed to remove excess amino acids, and the S-Fc-X is eluted from the protein A sepharose.
EXAMPLE 5
Preparation of Immunoglobulin Fc with N-Terminal-X-Termini and C-Terminal-X-Termini (X-Fc-X)
[0439]The X-Fc and Fc-X proteins are useful in preparing immunosymmetroadhesins having two binding domains joined to a single Fc domain (see below). Binding domains are added to the N-terminal-X-termini (X-Fc) or C-terminal-X-termini (Fc-X). X-Fc-X domains that are useful in preparing bi-symmetroadhesins with four binding domains joined to a single Fc domain are prepared using the procedures described in EXAMPLE 2 and EXAMPLE 3.
[0440]Accordingly, host cells are transfected with expression vectors that encode pre-Fc-intein chimeric polypeptides containing the M1 domain joined at its C-terminus, ELQLEESCAEAQDGELDG, by a peptide bond to the N-terminus of a self-splicing intein at the autocleavage site (FIG. 39). Heterologous signal peptides are selected from proteins with N-terminal cysteines (part i). Thus, cleavage by the cellular signal peptidase will provide a mature S-Fc-intein fusion protein having cys-11 at the N-terminus (part ii). Native chemical ligation is then employed to prepare X-Fc-intein fusion proteins having cys-5 or sec-5 at the N-terminus, XDKTHTCPPCP (part iii). Excision of the intein by protein splicing provides an Fc-thioester intermediate (part iv). Finally, in a second native chemical ligation reaction, the Fc-thioester with is carried out with free cysteine and/or selenocysteine to prepare X-Fc-X proteins having a C-terminal-X-terminus ELQLEESCAEAQDGELDGX (part v). Alternately, part iii is carried following parts iv and v.
[0441]The sequences of the IgG1 precursor polypeptides of FIGS. 39A-39B are shown in SEQ ID NO:132, SEQ ID NO:133, and SEQ ID NO:134. The sequence of the mature and modified IgG1 polypeptides of FIGS. 38A-38B is shown in SEQ ID NO:135 to SEQ ID NO:138.
[0442]Suitable expression vectors are constructed by ligation of an insert and vector fragment prepared by digestion with EcoRI and BglII. For unamplified expression, plasmid pSHH-Fc11-Mth is constructed with SEQ ID NO:14 and SEQ ID NO:1, pIFN-Fc11-Mth is constructed with SEQ ID NO:15 and SEQ ID NO:1, and pCETP-Fc11-Mth is constructed with SEQ ID NO:16 and SEQ ID NO:1. For amplified expression, pSHH-Fc11-Mth-DHFR is constructed with SEQ ID NO:14 and pSVeCD4DHFR, pIFN-Fc11-Mth-DHFR is constructed with SEQ ID NO:15 and pSVeCD4DHFR, and pCETP-Fc11-Mth-DHFR is constructed with SEQ ID NO:16 and pSVeCD4DHFR.
[0443]The S-Fc-intein fusion protein is first purified from the culture supernatant of transfected host cells using chitin affinity chromatography. Native chemical ligation is carried out on the chitin column with the Msc-NH-cys5-thr10-α-thioester peptide (Fc-A) or the Msc-NH-sec5-thr10-α-thioester peptide (Fc-B) as described (EXAMPLE 2). The chitin column is then washed thoroughly to remove unreacted peptide. The intein autocleavage reaction is then carried out. After cleavage, the X-Fc-thioester intermediate is directly applied to protein A Sepharose. The bound activated X-Fc-thioester is used directly for native ligation with cysteine and/or selenocysteine. The column is then washed to remove excess amino acids. The Msc-blocked X-Fc-X is eluted from the column, treated to remove the Msc protecting group, and repurified using protein A Sepharose to yield the final X-Fc-X product.
EXAMPLE 6
CD4 Symmetroadhesins
[0444]A therapeutic strategy for treating HIV-1 infection is based upon human CD4, a component of the HIV-1 receptor. CD4 immunoadhesins (Capon et al. (1989) Nature 337, 525-531) effectively block HIV-1 infectivity by binding to the gp120 envelope protein. The blocking activity resides in the CD4 extracellular domain (residues 1 to 371).
[0445]Accordingly, various CD4 symmetroadhesins are prepared using CD4-X protein, and analysed for their ability to bind gp120 and block HIV-1 infectively. The activity of CD4 symmetroadhesins is compared with CD4 immunoadhesin prepared as described (Capon et al., ibid).
[0446]CD4-X protein is prepared by the procedures of EXAMPLE 3. Host cells are transfected with expression vectors that encode the pre-CD4 chimeric polypeptide containing the CD4 extracellular domain joined at its C-terminus by a peptide bond to the N-terminus of a self-splicing intein at the autocleavage site (FIGS. 40A-40B).
[0447]Cleavage of the CD4 signal sequence (part i) by the cellular signal peptidase provides mature CD4-intein fusion protein (part ii). Excision of the intein domain provides a CD4-thioester intermediate (part iii). Finally, native chemical ligation of the CD4-thioester with free cysteine and/or selenocysteine is employed to prepare CD4 domains with C-terminal-X-termini (part iv).
[0448]The sequences of these polypeptides are shown in SEQ ID NO: 140, SEQ ID NO:141, SEQ ID NO:142, and SEQ ID NO:143.
[0449]Suitable expression vectors are constructed by ligation of an insert and vector fragment prepared by digestion with EcoRI and BglII. For unamplified expression, plasmid pCD4-Mth is constructed using SEQ ID NO:17 and SEQ ID NO:1. For amplified expression, plasmid pCD4-Mth-DHFR is constructed using SEQ ID NO:17 and pSVeCD4DHFR.
[0450]Suitable expression vectors were constructed by ligation of an insert fragment prepared by digestion with Hind III and EagI and a vector fragment prepared by digestion with Hind III and PspOM1. Plasmid pCDNA3-CD4-Mth was constructed using SEQ ID NO:139 and pCDNA3.1(+).
[0451]FIG. 56 shows expression in human 293 kidney cells of human CD4-intein fusion proteins. Lanes 1-4 show the CD4-intein fusion polypeptide of FIG. 40A (ii). Cell supernatants: lanes 1 and 3; cell lysates: lanes 2 and 4.
[0452]CD4-X is used to prepare various CD4 symmetroadhesins (Table 2). Hemi-symmetroadhesins are prepared using the CD4-X protein alone, immunosymmetroadhesins are prepared using the CD4-X protein with S-Fc, X-Fc, or Fc-X, and bi-symmetroadhesins are prepared using the CD4-X protein with S-Fc-X or X-Fc-X.
[0453]CD4 hemi-symmetroadhesin is prepared using CD4-X protein exposed to mildly reducing conditions that activate the X-termini, but do not denature the protein (Fleischman et al. (1962) Arch. Biochem. Biophys. 1 (Suppl.), 174-180; Edelman et al. (1963) Proc. Nat. Acad. Sci. (1963) 50, 753-761). CD4-X protein (0.5 to 2.0 mgs/ml) is dissolved in reducing buffer (0.05 M Tris-HCl buffer pH 8.0, made 0.1 M in 2-mercaptoethanol) and incubated for 1 hour at room temperature. The protein is then exchanged into oxidation buffer (0.1 M K2HPO4) by Sephadex G-100 chromatography, gently agitated in a round bottom glass test tube stoppered with a loosely packed cotton plug, and allowed to proceed at room temperature for 20 hours (Haber and Anfinsen (1961) J. Biol. Chem. 236, 422-424). CD4 immunosymmetroadhesins and CD4 bi-symmetroadhesins are prepared on protein A Sepharose beads using the S-Fc, X-Fc, Fc-X, S-Fc-X, or X-Fc-X proteins as indicated (Table 2). The bound proteins are gently agitated in reducing buffer for 1 hour at room temperature, then washed with oxidation buffer. CD4-X is treated with reducing buffer, added to the beads, and the reaction allowed to proceed at room temperature for 20 hours.
[0454]CD4-gp120 saturation binding analysis is carried out as described (Smith et al. (1987) Science 238, 1704-1707) using radio-iodinated gp120 prepared as described (Lasky et al. (1987) Cell 50, 975-985). Reactions (0.2) contain 0.25% NP-40, 0.1% sodium deoxycholate, 0.06 M NaCl, 0.01 M Tris-HCl, pH 8.0 (1× buffer A) with 125I-gp120 (3 ng to 670 ng at 2.9 nCi/ng). Binding is carried out for 1 hour at 0° C. in the presence or absence of 50 micrograms of unlabeled purified gp120. The bound 125I-gp120 is then determined by immunoprecipitation. Binding reactions are preabsorbed with 5 microliters of normal rabbit serum for 1 hour at 0° C., cleared with 40 microliters of 10% w/v Pansorbin (Calbiochem) for 30 minutes at 0° C., and incubated overnight at 0° C. with 2 microliters of normal serum or 5 microliters (0.25 microgram) of OKT4 monoclonal antibody (Ortho Biotech). Immunoprecipitates are collected with 10 microliters of Pansorbin, washed twice in 2× buffer and once in water, then eluted at 100° C. for 2 minutes in 0.12 M Tris-HCl pH 6.8, 4% SDS, 0.7 M mercaptoethanol. The fraction of bound 125I-gp120 is determined in a gamma counter and Scatchard analysis is used to determine the apparent dissociation contant.
[0455]HIV-1 blocking studies are carried out as described (Robert-Guroff et al. (1985) Nature 316, 72-74). Equal volumes of inhibitor and HIV-1 (60 microliters) are incubated at 4° C. for 1 hour, then the same volume of H9 cells (Gallo et al. (1984) Science 224, 500-503) at 5×106/ml is added and incubation continued for 1 hour at 37° C. Following absorption of virus, 2.5×105 cells in 150 microliters are transferred to 2 ml of incubation media. After 4 days at 37° C., the cultures are split 1:2 with fresh media and incubated for an additional 3 days. Cultures are harvested, reverse transcriptase activity is measured (Groopman et al., AIDS Res. Hum. Retroviruses (1987) 3, 71-85), and immunofluorescence with HIV-1 positive serum is determined as described (Poiesz et al. (1980) Proc. Acad. Nat. Sci. 77, 7415-7419). Challenge dose of virus is 100 TCID50 of HIV-1 strain HTLV-IIIB grown in H9 cells assayed in the same system. Incubation media is RPMI 1640 media containing 2 mM L-glutamine, 100 units/ml penicillin, 100 micrograms/ml streptomycin, 2 micrograms/ml polybrene and 20% fetal calf serum.
EXAMPLE 7
Tumor Necrosis Factor Receptor Symmetroadhesins
[0456]A therapeutic strategy for treating autoimmune disease is based upon tumor necrosis factor α (TNF-α), and its binding interaction with TNF-α antibodies and receptors (TNR). Both are an important therapeutic option in adult rheumatoid arthritis, juvenile rheumatoid arthritis, ankylosing spondylitis, psoriatic arthritis, Crohn's disease, and ulcerative colitis.
[0457]Accordingly, various TNR symmetroadhesins are prepared using TNR-X proteins, and analyzed for their ability to bind TNF-α and to block TNF-α biological activity. The activity of TNR symmetroadhesins is compared to that of TNR immunoadhesins (Ashkenazi et al. (1991) Proc. Natl. Acad. Sci. 88, 10535-10539).
[0458]Human TNR include TNR1A (Genbank Acc. No. NM--001065); and TNR1B (GenBank Acc. No. NM--001066). TNF-α antibody Di62 includes Di62 heavy chain (Genbank Acc. No. AJ002433); and Di62 light chain (Genbank Acc. No. AJ002434) (hereafter, TNR1A=TNR1; TNR1B=TNR2; and Di62-VH-CH+Di62-VkCk=TNRFab).
[0459]Di62-Vk is prepared according to the method of EXAMPLE 1. Di62-VH-X, TNR1A-X, and TNR1B-X, are prepared according to the method of EXAMPLE 3.
[0460]Host cells are co-transfected with two expression vectors that encode the Di62-VH-CH-intein chimeric polypeptide and Di62-VkCk protein result in co-expression of the Di62-VH-CH-intein:Di62-Vk-Ck protein that is used to prepare the TNFFab-X protein:
1) a pre-Di62-VH-intein chimeric polypeptide containing the Di62-VH-CH1 domain joined at its C-terminus by a peptide bond to the N-terminus of a self-splicing intein at the autocleavage site (FIGS. 41A-41B); and2) a pre-Di62-Vk polypeptide containing the Di62-Vk-Ck domain (FIG. 42).
[0461]Cleavage of the homologous Di62-VH-CH signal sequence (part i) by the cellular signal peptidase provides mature Di62-VH-CH-intein fusion protein (part ii). Excision of the intein by protein splicing provides the Di62-VH-CH-thioester intermediate (part iii). Finally, native chemical ligation of the Di62-VH-CH-thioester with free cysteine and/or selenocysteine is employed to prepare Di62-VH-CH-X protein with C-terminal-X-termini (part iv) (refer to FIGS. 41A-41B).
[0462]Cleavage of the homologous Di62-Vk-Ck signal sequence (part i) by the cellular signal peptidase provides the mature Di62-Vk-Ck protein (part ii) (refer to FIG. 42).
[0463]Host cells are transfected with expression vectors that encode the TNR1A-intein chimeric polypeptide and TNR1B-intein chimeric polypeptide to prepare the TNF1-X and TNF2-X proteins, respectively:
1) a pre-TNR1A-intein chimeric polypeptide containing the TNR1A extracellular domain joined at its C-terminus by a peptide bond to the N-terminus of a self-splicing intein at the autocleavage site (FIG. 43); and2) a pre-TNR1B-intein chimeric polypeptide containing the TNR1B extracellular domain joined at its C-terminus by a peptide bond to the N-terminus of a self-splicing intein at the autocleavage site (FIG. 44A).
[0464]Cleavage of the homologous TNR signal sequences (part i) by the cellular signal peptidase provides mature TNR-intein fusion proteins (part ii). Excision of the intein by protein splicing provides TNR-thioester intermediates (part iii). Finally, native chemical ligation of the TNR-thioesters with free cysteine and/or selenocysteine is employed to prepare TNR-X proteins with C-terminal-X-termini (part iv) (refer to FIG. 43, 44A).
[0465]The sequences of these polypeptides are shown in SEQ ID NO: 145, SEQ ID NO:146, SEQ ID NO:147, and SEQ ID NO:148 (Di62-VHCH); in SEQ ID NO: 150, and SEQ ID NO:151 (Di62-VkCk); in SEQ ID NO: 153, SEQ ID NO:154, SEQ ID NO:155, and SEQ ID NO:156 (TNR1A); in SEQ ID NO: 158, SEQ ID NO:159, SEQ ID NO:160, and SEQ ID NO:161 (TNR1B); and in SEQ ID NO: 163, and SEQ ID NO:164 (TNR1B immunoadhesin).
[0466]Suitable expression vectors are constructed by ligation of an insert and vector fragment prepared by digestion with EcoRI and BglII. For unamplified expression, plasmid pDi62-VHCH-Mth is constructed using SEQ ID NO:18 and SEQ ID NO:1, plasmid pDi62-VkCk is constructed using SEQ ID NO:19 and SEQ ID NO:1, plasmid pTNR1A-Mth is constructed using SEQ ID NO:20 and SEQ ID NO:1, and plasmid pTNR1A-Mth is constructed using SEQ ID NO:22 and SEQ ID NO:1. For amplified expression, plasmid pDi62-VHCH-Mth-DHFR is constructed using SEQ ID NO:18 and pSVeCD4DHFR, plasmid pDi62-VkCk-DHFR is constructed using SEQ ID NO:19 and pSVeCD4DHFR, plasmid pTNR1A-Mth-DHFR is constructed using SEQ ID NO:20 and pSVeCD4DHFR, and plasmid pTNR1A-Mth-DHFR is constructed using SEQ ID NO:22 and pSVeCD4DHFR.
[0467]Suitable expression vectors are constructed by ligation of an insert fragment prepared by digestion with Hind III and EagI and a vector fragment prepared by digestion with Hind III and PspOM1. Plasmid pCDA3-Di62-VHCH-Mth is constructed using SEQ ID NO:144 and pCDNA3.1(+). Plasmid pCDA3-Di62-VkCk is constructed using SEQ ID NO:149 and pCDNA3.1(+). Plasmid pCDNA-3-TNR1A-Mth-DHFR is constructed using SEQ ID NO:152 and pCDNA3.1(+). Plasmid pCDNA-3-TNR1B-Mth-DHFR was constructed using SEQ ID NO:157 and pCDNA3.1(+). Plasmid pCDNA-3-TNR1B-immunoadhesin was constructed using SEQ ID NO:162 and pCDNA3.1(+).
[0468]FIG. 57 shows the expression in human 293 kidney cells of human TNR1B fusion proteins. Lanes 2 and 5 show the TNR1B-intein fusion protein of FIG. 44A (ii). Lanes 1 and 3 show the TNR1B-immunoadhesin fusion protein of FIG. 44B (ii). Lanes 3 and 6 show proteins from mock-transfected cells. Cell supernatants: lanes 1-3; cell lysates: lanes 4-7. Lane 7: control TNR1B-immunoadhesin (R&D Systems).
[0469]FIG. 58 shows TNR1B symmetroadhesin subunits with C-terminal-S-termini. Lanes 1-2 show the TNR1B polypeptide of FIG. 44A (iii) following purification by chitin affinity chromatography and cleavage/elution with MESNA. Lanes 3 shows the native ligation product between the TNR1B polypeptide of FIG. 44A (iii) with a fluorescent-labeled peptide (New England Biolabs). Panel (i): direct fluorescence; panel (ii): western blot with anti-TNR1B antibody (R&D Systems); panel (iii): SYPRO Ruby staining (Sigma-Aldrich).
[0470]FIG. 59 shows TNR1B symmetroadhesin subunits with C-terminal-S-termini. Lane 5 shows the TNR1B polypeptide of FIG. 44A (iv) following purification by chitin affinity chromatography and cleavage/elution with cysteine. Lanes 1-4 show TNR1B-immunoadhesin.
[0471]FIG. 60 shows TNR1B symmetroadhesin. Lanes 1-4 show the TNR1B symmetroadhesin of FIG. 44A (iv) before oxidation (lanes 1 and 4) and after oxidation in the presence of 10 mM CuSO4. Lanes 3 and 6 show a TNR1B-immunoadhesin control. Lanes 1-3: reducing conditions; lanes 4-6: non-reducing conditions. The TNR1B symmetroadhesin monomer (42 kd) and dimer (84 kd) are apparent in lanes 2 and 5, and lane 5, respectively.
[0472]FIGS. 61A-61C show TNF-alpha saturation binding analysis with various TNR1B polypeptides on the Biacore T-100. (A) The TNR1B symmetroadhesin of FIG. 44A (iv) was covalently coupled to a Biacore CM-5 chip using standard Biacore amine chemistry. (B) TNR1B immunoadhesin (R&D Systems) was covalently coupled to a Biacore CM-5 chip using standard Biacore amine chemistry. (C) The TNR1B symmetroadhesin of FIG. 44A (iv) was covalently coupled to a Biacore CM-5 chip using standard Biacore thiol chemistry. Following coupling, saturation binding analysis was carried out using TNF-alpha (R&D Systems) at the indicated concentations.
[0473]FIGS. 62A-62C shows Scatchard analysis of the TNF-alpha saturation binding analysis shown in FIG. 61A-61C. (A) TNR1B symmetroadhesin of FIG. 44A (iv) covalently coupled using amine chemistry; Kd=Kd=4.697×10-9 M. (B) TNR1B-immunoadhesin (R&D Systems) covalently coupled using amine chemistry; Kd=4.089×10-9 M. (C) TNR1B symmetroadhesin of FIG. 44A (iv) covalently coupled using thiol chemistry; Kd=0.8476×10-9 M.
[0474]The TNR1-X, TNR2-X, and TNRFab-X proteins, individually and in various combinations, are used to prepare various VGFR symmetroadhesins (Tables 3, 4, and 5). The number of distinct configurations obtained for each combination, as well as the general structure for each, is also shown.
[0475]Hemi-symmetroadhesins are prepared using the TNR-X proteins alone, immunosymmetroadhesins are prepared using the TNR-X proteins with S-Fc, X-Fc, or Fc-X, and bi-symmetroadhesins are prepared using the TNR-X proteins with S-Fc-X or X-Fc-X.
[0476]Each TNR-X protein or TNR symmetroadhesin is determined by a quantitative immunoassay using subtype-specific, affinity-purified polyclonal antibodies to TNR1A, TNR1B, and Di62 (goat-anti-mouse), and immunoadhesins (TNR1A-Ig and TNR2A-Ig) as reference standards. TNR antibodies, TNR-Ig immunoadhesins, and TNF-α protein are obtained from R&D Systems (MN).
[0477]Binding of TNGR symmetroadhesins to TNF-α is studied as described (Ashkenazi et al. (1991) Proc. Natl. Acad. Sci. 88, 10535-10539). Individual samples (1 microgram/ml) are immobilized onto microtiter wells coated with goat-anti-human Ig Fc antibody. Reactions with recombinant human 125I-TNF-α (radiodinated by using lactoperoxidase to a specific activity of 19.1 microCi/microgram, 1 microCi=37 kBq) are done in phosphate-buffered saline (PBS) containing 1% bovine serum albumin for 1 hour at 24° C. Non-specific binding is determined by omitting the sample. In competition binding analyses, 125I-TNF-α is incubated with immobilized samples in the presence of increasing concentrations of unlabeled TNF-α. The Kd is determined from competition IC50 values according to the following equation: Kd=IC50/(1+[T]/KdT), where [T] is the concentration of the tracer (0.1 nm) and KdT is the Kd of the tracer determined by saturation binding (80 μM). TNF cytotoxity is studied as described (Kawade and Watanabe (1984) J. Interferon Res. 4, 571-584). Mouse L-M cells are plated in microtiter dishes (4×104 cells per well) and treated with actinomycin D (3 micrograms/ml) and TNF-α or TNF-β (1 nanogram/ml) in the presence or absence of the sample or other inhibitors. After 20 hours of incubation at 39° C., the cell survival is determined by a crystal violet dye exclusion test.
[0478]A mouse model for septic shock is studied by endotoxin injection of 6- to 8-week-old BALB/c mice. Animals are injected intravenously (i.v.) with an LD100 dose of Salmonella abortus-derived endotoxin (175 micrograms per mouse) in phosphate-buffered saline (PBS), and survival is followed for at least 78 hour. TNR1-immunoadhesin and CD4-immunoadhesin are used as the positive and negative controls, respectively. Each is diluted in PBS and injected i.v. prior to, or after, the administration of endotoxin.
EXAMPLE 8
Vascular Endothelial Growth Factor Receptor Symmetroadhesins
[0479]A therapeutic strategy for treating angiogenic disease is based upon vascular endothelial growth factors (VEGF) and their binding interaction with VEGF receptors (VGFR). VEGF-antibodies and VGFR-immunoadhesins are promising candidates for treatment in a number of metastatic carcinomas including colon, rectum, lung, and breast, and in age-related macular degeneration.
[0480]Accordingly, various VGFR symmetroadhesins are prepared using VGFR-X proteins, and analyzed for their ability to bind VEGF and to block VEGF biological activity. The activity of VGFR symmetroadhesins is compared to that of VGRF immunoadhesins (Park et al. (1994) J. Biol. Chem. 269, 25646-25654).
[0481]Human VGFR include VGFR1 (Genbank Acc. No. NM--002019); VGFR2 (GenBank Acc. No. NM--002253); and VGFR3 (GenBank Acc. No. NM--002020). VGFR1-X, VGFR2-X, and VGFR3-X are prepared according to the method of EXAMPLE 3. Host cells are transfected with expression vectors that encode:
1) a pre-VGFR1-intein chimeric polypeptide containing the VGFR1 extracellular domain joined at its C-terminus by a peptide bond to the N-terminus of a self-splicing intein at the autocleavage site (FIGS. 45A-45C);2) a pre-VGFR2-intein chimeric polypeptide containing the VGFR1 extracellular domain joined at its C-terminus by a peptide bond to the N-terminus of a self-splicing intein at the autocleavage site (FIGS. 46A-46C); and3) a pre-VGFR1-intein chimeric polypeptide containing the VGFR1 extracellular domain joined at its C-terminus by a peptide bond to the N-terminus of a self-splicing intein at the autocleavage site (FIGS. 47A-47C).
[0482]Cleavage of the VGFR signal sequence (part i) by the cellular signal peptidase provides mature VGFR-intein fusion proteins (part ii). Excision of the intein domain provides a VGFR-thioester intermediate (part iii). Finally, native chemical ligation of the VGFR-thioester with free cysteine and/or selenocysteine is employed to prepare VGFR domains with C-terminal-X-termini (part iv).
[0483]The sequences of these polypeptides are shown in SEQ ID NO: 166, SEQ ID NO:167, SEQ ID NO:168, and SEQ ID NO:169 (VGFR1); in SEQ ID NO: 171, SEQ ID NO:172, SEQ ID NO:173, and SEQ ID NO:174 (VGFR2); and in SEQ ID NO: 176, SEQ ID NO:177, SEQ ID NO:178, and SEQ ID NO:179 (VGFR3).
[0484]Suitable expression vectors are constructed by ligation of an insert and vector fragment prepared by digestion with EcoRI and BglII. For unamplified expression, plasmid pVGFR1-Mth is constructed using SEQ ID NO:22 and SEQ ID NO:1, plasmid pVGR1-Mth is constructed using SEQ ID NO:23 and SEQ ID NO:1, and plasmid pVGR1-Mth is constructed using SEQ ID NO:24 and SEQ ID NO:1. For unamplified expression, plasmid pVGFR1-Mth-DHFR is constructed using SEQ ID NO:22 and pSVeCD4DHFR, plasmid pVGR1-Mth-DHFR is constructed using SEQ ID NO:23 and pSVeCD4DHFR, and plasmid pVGR1-Mth-DHFR is constructed using SEQ ID NO:24 and pSVeCD4DHFR.
[0485]Suitable expression vectors are constructed by ligation of an insert fragment prepared by digestion with Hind III and EagI and a vector fragment prepared by digestion with Hind III and PspOM1. Plasmid pCDNA3-VGFR1-Mth is constructed using SEQ ID NO:165 and pCDNA3.1(+). Plasmid pCDNA3-VGFR2-Mth is constructed using SEQ ID NO:170 and pCDNA3.1(+). Plasmid pCDNA3-VGFR3-Mth is constructed using SEQ ID NO:175 and pCDNA3.1(+).
[0486]The VGFR1-X, VGFR2-X, and VEGFR3-X proteins, individually and in various combinations, are used to prepare various VGFR symmetroadhesins (Tables 6, 7, and 8). The number of distinct configurations obtained for each combination, as well as the general structure for each, is also shown.
[0487]Hemi-symmetroadhesins are prepared using the VGFR-X proteins alone, immunosymmetroadhesins are prepared using the VGFR-X proteins with S-Fc, X-Fc, or Fc-X, and bi-symmetroadhesins are prepared using the VGFR-X proteins with S-Fc-X or X-Fc-X.
[0488]Each VGFR-X protein or VGFR symmetroadhesin is determined by a quantitative immunoassay using subtype-specific, affinity-purified polyclonal antibodies to VGFR1, VGFR2, and VGFR3, and immunoadhesins (VGFR1-Ig, VGFR2-Ig, and VGFR3-Ig) as reference standards. VGFR antibodies, VGFR-Ig immunoadhesins, and VEGF165 protein are obtained from R&D Systems (MN).
[0489]Binding of VEGR symmetroadhesins to VEGF is studied using a VEGF saturation binding assay. Reactions (0.1 ml) contain 10% fetal bovine serum in PBS (buffer A) with 125I-VEGF165 (<9000 cpm/well, 5.69×107 cpm/microgram). Chloramine T is used to iodinated VEGF165 as described (Keyt et al. (1996) J. Biol Chem. 271, 5638-5646). Binding is carried out overnight at 4° C. in the presence or absence of 50 nanograms of unlabeled purified VEGF165. The bound 125I-VEGF165 is then determined by capture in 96-well breakaway immunoabsorbent assay plates (Nunc). Plates are coated overnight at 4° C. with 2 micrograms/ml affinity-purified goat anti-human Fc IgG (Organon-Teknika) in 50 mM Na2CO3, pH 9.6, and preblocked for 1 hr in buffer A. Binding reactions are then incubated in the coated wells for 4 hrs at room temperature, followed by 4 washes with buffer A. The fraction of bound 125I-VEGF165 is determined in a gamma counter. Data is analyzed using a 4-parameter non-linear curve fitting program (Kalidagraph, Abelbeck Software, Pa.).
[0490]Binding of VEGF to VEGR symmetroadhesins is also studied using a competition binding assay. ELISA plates are coated with 2 micrograms/ml rabbit F(ab')2 to human IgG Fc (Jackson ImmunoResearch, Pa.) and blocked with buffer A. KDR-IgG (3 nanograms/ml) in buffer A is added to the plate and incubated for 1 hr. Serially-diluted samples are incubated with 2 nM biotinylated VEGF for 1 h in tubes. The reactions are then transferred to the ELISA plates and incubated for 1 h. After washing, the fraction of biotinylated VEGF bound to KDR-Ig is detected using horseradish peroxidase-labeled streptavidin (Sigma, Mo.) followed by 3,3',5,5'-tetramethylbenzidine substrate. Data is analyzed using 4-parameter non-linear regression curve fitting analysis.
[0491]Endothelial cell growth inhibition studies are carried out as described (Leung et al. (1989) Science 246, 1306-1309). Bovine adrenal cortex capillary endothelial cells are cultured in the presence of low glucose Dulbecco's modified Eagle's medium (GIBCO) supplemented with 10% calf serum, and 2 mM glutamine (growth medium). Cells are seeded at a density of 6×103 cells/well in 6-well plates. Serially diluted samples are added to the cells at concentrations between 1 to 5000 nanograms/ml and incubated for 2 to 3 hr. Purified VEGF165 is added to a final concentration of 3 nanograms/ml. Cells are then incubated for 5 to 6 days, removed from plates with trypsin, and cell number determined in a Coulter counter (Coulter Electronics, FL). Data is analyzed using 4-parameter non-linear regression curve fitting analysis.
[0492]In vivo tumor studies are carried out as described (Kim et al. (1993) Nature 362, 841-844; Borgstrom et al. (1996) Cancer Res. 56, 4032-4039). Human A673 rhabdomyosarcoma cells (ATCC CRL-1598) are cultured in DMEM/F12 supplemented with 10% fetal bovine serum, and 2 mM glutamine. Female BALB/c nude mice, 6 to 10 weeks old, are injected subcutaneously with 2×106 tumor cells in the dorsal area in a volume of 200 microliters. Following tumor cell inoculation (24 hr.), animals (10 per group) are treated with serially diluted samples at dose of 0.05 mg/kg, 0.5 mg/kg, and 5 mg/kg, administered twice weekly intraperitoneally in a volume of 0.1 ml. Tumor size is determined at weekly intervals. Four weeks after tumor cell inoculation, animals are euthanized and the tumors removed and weighed. Statistical analysis is carried out by ANOVA.
EXAMPLE 9
ErbB Syetroadhesins
[0493]A therapeutic strategy for treating malignant disease is based upon epidermal growth factor-like receptors (ErbB) and their ligands, including the neuregulins/heregulins (NRG/HRG), and the family of EGF-related protein ligands. ErbB-antibodies and ErbB-immunoadhesins are under clinical investigation, and are well-proven in treating metastatic breast cancers overexpressing ErbB2.
[0494]Accordingly, various ErbB symmetroadhesins are prepared using ErbB-X proteins, and analyzed for their ability to bind heregulins and block heregulin biological activity. The activity of ErbB symmetroadhesins is compared to that of ErbB immunoadhesins (Sliwkowski et al. (1994) J. Biol. Chem. 269, 14661-14665).
[0495]Human ErbB include ErbB1 (Genbank Acc. No. NM--005228); ErbB2 (GenBank Acc. No. NM--004448); ErbB3 (GenBank Acc. No. NM--001982); and ErbB4 (GenBank Acc. No. NM--005235). ErbB1-X, ErbB2-X, ErbB3-X, and ErbB4-X are prepared according to the method of EXAMPLE 3. Host cells are transfected with expression vectors that encode:
1) a pre-ErbB1-intein chimeric polypeptide containing the ErbB1 extracellular domain joined at its C-terminus by a peptide bond to the N-terminus of a self-splicing intein at the autocleavage site (FIGS. 48A-48B);2) a pre-ErbB2-intein chimeric polypeptide containing the ErbB2 extracellular domain joined at its C-terminus by a peptide bond to the N-terminus of a self-splicing intein at the autocleavage site (FIGS. 49A-49B);3) a pre-ErbB3-intein chimeric polypeptide containing the ErbB3 extracellular domain joined at its C-terminus by a peptide bond to the N-terminus of a self-splicing intein at the autocleavage site (FIGS. 50A-50B); and4) a pre-ErbB4-intein chimeric polypeptide containing the ErbB4 extracellular domain joined at its C-terminus by a peptide bond to the N-terminus of a self-splicing intein at the autocleavage site (FIGS. 51A-50B).
[0496]Cleavage of the homologous ErbB signal sequence (part i) by the cellular signal peptidase provides mature ErbB-intein fusion proteins (part ii). Excision of the intein domain by protein splicing provides a ErbB-thioester intermediate (part iii). Finally, native chemical ligation of the ErbB-thioester with free cysteine and/or selenocysteine is employed to prepare VEGR domains with C-terminal-X-termini (part iv).
[0497]The sequences of these polypeptides are shown in SEQ ID NO: 181, SEQ ID NO:182, SEQ ID NO:183, and SEQ ID NO:184 (ERBB1); in SEQ ID NO: 186, SEQ ID NO:187, SEQ ID NO:188, and SEQ ID NO:189 (ERBB2); in SEQ ID NO: 191, SEQ ID NO:192, SEQ ID NO:193, and SEQ ID NO:194 (ERBB3); and in SEQ ID NO: 196, SEQ ID NO:197, SEQ ID NO:198, and SEQ ID NO:199 (ERBB4).
[0498]Suitable expression vectors are constructed by ligation of an insert and vector fragment prepared by digestion with EcoRI and BglII. For unamplified expression, plasmid pErbB1-Mth is constructed using SEQ ID NO:25 and SEQ ID NO:1, plasmid pErbB2-Mth is constructed using SEQ ID NO:26 and SEQ ID NO:1, plasmid pErbB3-Mth is constructed using SEQ ID NO:27 and SEQ ID NO:1, and plamid pErbB4-Mth is constructed using SEQ ID NO:28 and SEQ ID NO:1. For amplified expression, plasmid pErbB1-Mth-DHFR is constructed using SEQ ID NO:25 and pSVeCD4DHFR, plasmid pErbB2-Mth-DHFR is constructed using SEQ ID NO:26 and pSVeCD4DHFR, plasmid pErbB3-Mth-DHFR is constructed using SEQ ID NO:27 and pSVeCD4DHFR, and plamid pErbB4-Mth-DHFR is constructed using SEQ ID NO:28 and pSVeCD4DHFR.
[0499]Suitable expression vectors are constructed by ligation of an insert fragment prepared by digestion with Hind III and EagI and a vector fragment prepared by digestion with Hind III and PspOM1. Plasmid pCDNA3-ERBB1-Mth is constructed using SEQ ID NO:180 and pCDNA3.1(+). Plasmid pCDNA3-ERBB2-Mth is constructed using SEQ ID NO:185 and pCDNA3.1(+). Plasmid pCDNA3-ERBB3-Mth is constructed using SEQ ID NO:190 and pCDNA3.1(+). Plasmid pCDNA3-ERBB4-Mth is constructed using SEQ ID NO:195 and pCDNA3.1(+).
[0500]The ErbB1-X, ErbB2-X, ErbB3-X and ErbB4-X proteins, individually and in various combinations, are used to prepare various VGFR symmetroadhesins (Tables 9, 10, 11). The number of distinct configurations obtained for each combination, as well as the general structure for each, is also shown.
[0501]Hemi-symmetroadhesins are prepared using the ErbB-X proteins alone, immunosymmetroadhesins are prepared using the ErbB-X proteins with S-Fc, X-Fc, or Fc-X, and bi-symmetroadhesins are prepared using the ErbB-X proteins with S-Fc-X or X-Fc-X.
[0502]Each ErbB-X protein or ErbB symmetroadhesin is determined by a quantitative immunoassay using subtype-specific, affinity-purified polyclonal antibodies to ErbB1, ErbB2, ErbB3, and ErbB4, and immunoadhesins (ErbB1-Ig, ErbB2-Ig, ErbB3-Ig, and ErbB4-Ig) as reference standards. ErbB antibodies, ErbB-Ig immunoadhesins, NRG1-α177-241 protein, NRG1-β176-246 protein, NRG1-β1-246 protein, and NRG1-SMDF1-296 protein are obtained from R&D Systems (MN).
[0503]Binding of ErbB symmetroadhesins to neuregulins is studied using a HRG saturation binding assay (Sliwkowski et al. (1994) J. Biol. Chem. 269, 14661-14665). Reactions are performed in Nunc breakapart immuno-module plates. Plate wells are coated at 4° C. overnight with 100 microliters of 5 micrograms/ml goat-anti-human antibody (Boehringer Mannheim) in 50 mM carbonate buffer (pH 9.6). Plates are rinsed twice with 200 microliters wash buffer (PBS/0.05% Tween-20®) followed by a brief incubation with 100 microliters 1% BSA/PBS for 30 min at room temperature. Buffer is removed and each well is incubated with 100 microliters of the sample in 1% BSA/PBS under vigorous side-to-side rotation for 1 hour. Plates are rinsed three times with wash buffer and competitive binding is carried out by adding various amounts of cold competitor gamma-HRG and 125I-HRGβ1 and incubating at room temperature for 2 to 3 hours with vigorous side-to-side rotation. Wells are quickly rinsed three times with wash buffer, drained and individual wells are counted in a gamma-counter. Scatchard analysis is performed using a modified Ligand program (Munson, P. and Robard, D. (1980) Analytical Biochemistry 107:220-239).
[0504]The ability of ErbB symmetroadhesins to inhibit HRG-dependent proliferation is studied in the MCF7 breast carcinoma cell line (Lewis et al. (1996) Cancer Res. 56, 1457-1465). 3H-thymidine incorporation is are carried out in 96-well format. Serum-starved MCF7-7 cells are plated at 10,000 cells/well in 50:50 F12/DMEM (high glucose). Varying concentrations of sample are incubated with 1 nM HRG and added to the cells. After a 15 hour incubation, cells are labeled with 3H-thymidine to measure DNA synthesis (20 mL of 1/20 diluted tritiated thymidine stock: Amersham TRA 120 B363, 1 mCi/ml). Cells are then harvested onto GF/C unifilters (96 well format) using a Packard Filtermate 196 harvester. Filters are counted using a Packard Topcount apparatus.
Screening
[0505]This invention is particularly useful for screening compounds by using consecutive amino acid sequences/or compounds comprising such in any of a variety of drug screening techniques. The compound employed in such a test may either be free in solution, affixed to a solid support, borne on a cell surface, or located intracellularly. One method of drug screening utilizes eukaryotic or prokaryotic host cells which are stably transformed with recombinant nucleic acids expressing the compound. Drugs are screened against such transformed cells in competitive binding assays. Such cells, either in viable or fixed form, can be used for standard binding assays. One may measure, for example, the formation of complexes between compound or a fragment and the agent being tested. Alternatively, one can examine the diminution in complex formation between the compound and its target cell or target receptors caused by the agent being tested.
[0506]Thus, the present invention provides methods of screening for drugs or any other agents which can affect a disease or disorder associated with a stretch of consecutive amino acids of the compound. These methods comprise contacting such an agent with the compound or fragment thereof and assaying (I) for the presence of a complex between the agent and the compound or fragment thereof, or (ii) for the presence of a complex between the compound or fragment thereof and the cell, by methods well known in the art. In such competitive binding assays, the compound or fragment thereof is typically labeled. After suitable incubation, free compound or fragment thereof is separated from that present in bound form, and the amount of free or uncomplexed label is a measure of the ability of the particular agent to bind to the compound or fragment thereof or to interfere with the compound/cell complex.
[0507]Another technique for drug screening provides high throughput screening for compounds having suitable binding affinity to a polypeptide and is described in detail in WO 84/03564, published on Sep. 13, 1984. Briefly stated, large numbers of different small peptide test compounds are synthesized on a solid substrate, such as plastic pins or some other surface. As applied to the instant compound or fragment thereof, the peptide test compounds are reacted with compound or fragment thereof and washed. Bound compound or fragment thereof is detected by methods well known in the art. Purified compound or fragment thereof can also be coated directly onto plates for use in the aforementioned drug screening techniques. In addition, non-neutralizing antibodies can be used to capture the peptide and immobilize it on the solid support.
[0508]This invention also contemplates the use of competitive drug screening assays in which neutralizing antibodies capable of binding compound or fragment thereof specifically compete with a test compound for binding to compound or fragment thereof. In this manner, the antibodies can be used to detect the presence of any peptide which shares one or more antigenic determinants with the stretch(es) of consecutive amino acids of the compound.
Rational Drug Design
[0509]The goal of rational drug design is to produce structural analogs of biologically active polypeptide of interest (i.e., a compound of the invention, or an amino acid sequence of a compound of the invention) or of small molecules with which they interact, e.g., agonists, antagonists, or inhibitors. Any of these examples can be used to fashion drugs which are more active or stable forms of the a compound of the invention, or an amino acid sequence of a compound of the invention or which enhance or interfere with the function of the a compound of the invention, or an amino acid sequence of a compound of the invention in vivo (cf., Hodgson, Bio/Technology, 9:19-21 (1991)).
[0510]In one approach, the three-dimensional structure of a compound of the invention, or an amino acid sequence of a compound of the invention, or of a compound-inhibitor complex, is determined by x-ray crystallography, by computer modeling or, most typically, by a combination of the two approaches. Both the shape and charges of a compound of the invention, or an amino acid sequence of a compound of the invention, must be ascertained to elucidate the structure and to determine active site(s). Relevant structural information is used to then design analogous molecules or to identify efficient inhibitors. Useful examples of rational drug design may include molecules which have improved activity or stability as shown by Braxton and Wells, Biochemistry, 31:7796-7801 (1992) or which act as inhibitors, agonists, or antagonists of native peptides as shown by Athauda et al., J. Biochem., 113:742-746 (1993).
[0511]It is also possible to isolate a target-specific antibody, selected by functional assay, as described above, and then to solve its crystal structure. This approach, in principle, yields a pharmacore upon which subsequent drug design can be based. It is possible to bypass protein crystallography altogether by generating anti-idiotypic antibodies (anti-ids) to a functional, pharmacologically active antibody. As a mirror image of a mirror image, the binding site of the anti-ids would be expected to be an analog of the original receptor. The anti-id could then be used to identify and isolate peptides from banks of chemically or biologically produced peptides. The isolated peptides would then act as the pharmacore.
[0512]By virtue of the present invention, sufficient amounts of the compounds of the invention, or an amino acid sequence of a compound of the invention may be made available to perform such analytical studies as X-ray crystallography.
Assays of Biological Activity
[0513]The compounds disclosed herein, including components thereof such as a stretch of consecutive amino acids of a compound, are readily assayed using one or more standard assays of biological activity known to those in the art. The following are non-limiting examples of such assays:
Ability of the Compounds to Inhibit Vascular Endothelial Growth Factor (VEGF) Stimulated Proliferation of Endothelial Cell Growth
[0514]The ability of the various compounds of this invention to inhibit VEGF stimulated proliferation of endothelial cells was tested. Positive testing in this assay indicates the compound is useful for inhibiting endothelial cell growth in mammals where such an effect would be beneficial, e.g., for inhibiting tumor growth.
[0515]In a specific example of the assay, bovine adrenal cortical capillary endothelial cells (ACE) (from primary culture, maximum of 12-14 passages) are plated in 96-well plates at 500 cells/well per 100 microliter. Assay media includes low glucose DMEM, 10% calf serum, 2 mM glutamine, and 1× penicillin/streptomycin/fungizone. Control wells include the following: (1) no ACE cells added; (2) ACE cells alone; (3) ACE cells plus 5 ng/ml FGF; (4) ACE cells plus 3 ng/ml VEGF; (5) ACE cells plus 3 ng/ml VEGF plus 1 ng/ml TGF-beta; and (6) ACE cells plus 3 ng/ml VEGF plus 5 ng/ml LIF. The test samples, poly-his tagged compounds (in 100 microliter volumes), are then added to the wells (at dilutions of 1%, 0.1% and 0.01%, respectively). The cell cultures are incubated for 6-7 days at 37° C./5% CO2 After the incubation, the media in the wells is aspirated, the cells are washed 1× with PBS. An acid phosphatase reaction mixture (100 microliter; 0.1M sodium acetate, pH 5.5, 0.1% Triton X-100, 10 mM p-nitrophenyl phosphate) is then added to each well. After a 2 hour incubation at 37° C., the reaction is stopped by addition of 10 microliters 1M NaOH. Optical density (OD) is measured on a microplate reader at 405 nm.
[0516]The activity of the assayed compound is calculated as the percent inhibition of VEGF (3 ng/ml) stimulated proliferation (as determined by measuring acid phosphatase activity at OD 405 nm) relative to the cells without stimulation. TGF-beta can be employed as an activity reference at 1 ng/ml, since TGF-beta blocks 70-90% of VEGF-stimulated ACE cell proliferation. The results are indicative of the utility of the assayed compound in cancer therapy and specifically in inhibiting tumor angiogenesis. Numerical values (relative inhibition) are determined by calculating the percent inhibition of VEGF stimulated proliferation by the assayed compound relative to cells without stimulation and then dividing that percentage into the percent inhibition obtained by TGF-beta at 1 ng/ml which is known to block 70-90% of VEGF stimulated cell proliferation. The results are considered positive if the assayed compound exhibits 30% or greater inhibition of VEGF stimulation of endothelial cell growth (relative inhibition 30% or greater).
Retinal Neuron Survival
[0517]This assay can demonstrate if the compound tested has efficacy in enhancing the survival of retinal neuron cells and, therefore, is useful for the therapeutic treatment of retinal disorders or injuries including, for example, treating sight loss in mammals due to retinitis pigmentosum, AMD, etc.
[0518]Sprague Dawley rat pups at postnatal day 7 (mixed population: glia and retinal neuronal types) are killed by decapitation following CO2 anesthesia and the eyes are removed under sterile conditions. The neural retina is dissected away from the pigment epithelium and other ocular tissue and then dissociated into a single cell suspension using 0.25% trypsin in Ca2+, Mg2+-free PBS. The retinas are incubated at 37° C. for 7-10 minutes after which the trypsin is inactivated by adding 1 ml soybean trypsin inhibitor. The cells are plated at 100,000 cells per well in 96 well plates in DMEM/F12 supplemented with N2 and with or without the specific test compound. Cells for all experiments are grown at 37° C. in a water saturated atmosphere of 5% CO2. After 2-3 days in culture, cells are stained with calcein AM then fixed using 4% paraformaldehyde and stained with DAPI for determination of total cell count. The total cells (fluorescent) are quantified at 20× objective magnification using CCD camera and NIH image software for MacIntosh. Fields in the well are chosen at random.
[0519]The effect of various concentration of the tested compound are reported herein where percent survival is calculated by dividing the total number of calcein AM positive cells at 2-3 days in culture by the total number of DAPI-labeled cells at 2-3 days in culture. Anything above 30% survival is considered positive.
Rod Photoreceptor Cell Survival
[0520]This assay is used to show whether certain compounds of the invention act to enhance the survival/proliferation of rod photoreceptor cells and, therefore, are useful for the therapeutic treatment of retinal disorders or injuries including, for example, treating sight loss in mammals due to retinitis pigmentosum, AMD, etc. Sprague Dawley rat pups at 7 day postnatal (mixed population: glia and retinal neuronal cell types) are killed by decapitation following CO2 anesthesis and the eyes are removed under sterile conditions. The neural retina is dissected away form the pigment epithelium and other ocular tissue and then dissociated into a single cell suspension using 0.25% trypsin in Ca2+, Mg2+-free PBS. The retinas are incubated at 37° C. for 7-10 minutes after which the trypsin is inactivated by adding 1 ml soybean trypsin inhibitor. The cells are plated at 100,000 cells per well in 96 well plates in DMEM/F12 supplemented with N2. Cells for all experiments are grown at 37° C. in a water saturated atmosphere of 5% CO2. After 2-3 days in culture, cells are fixed using 4% paraformaldehyde, and then stained using CellTracker Green CMFDA. Rho 4D2 (ascites or IgG 1:100), a monoclonal antibody directed towards the visual pigment rhodopsin is used to detect rod photoreceptor cells by indirect immunofluorescence. The results are calculated as % survival: total number of calcein-rhodopsin positive cells at 2-3 days in culture, divided by the total number of rhodopsin positive cells at time 2-3 days in culture. The total cells (fluorescent) are quantified at 20× objective magnification using a CCD camera and NIH image software for MacIntosh. Fields in the well are chosen at random.
Induction of Endothelial Cell Apoptosis
[0521]The ability of the compounds disclosed herein to induce apoptosis in endothelial cells can be tested in human venous umbilical vein endothelial cells (HUVEC, Cell Systems). A positive test in the assay is indicative of the usefulness of the compound in therapeutically treating tumors as well as vascular disorders where inducing apoptosis of endothelial cells would be beneficial.
[0522]The cells are plated on 96-well microtiter plates (Amersham Life Science, cytostar-T scintillating microplate, RPNQ160, sterile, tissue-culture treated, individually wrapped), in 10% serum (CSG-medium, Cell Systems), at a density of 2×104 cells per well in a total volume of 100 μl. On day 2, test samples containing the tested compound are added in triplicate at dilutions of 1%, 0.33% and 0.11%. Wells without cells were used as a blank and wells with cells only are used as a negative control. As a positive control 1:3 serial dilutions of 50 μl of a 3× stock of staurosporine were used. The ability of the compound to induce apoptosis is determined by processing of the 96 well plates for detection of Annexin V, a member of the calcium and phospholipid binding proteins, to detect apoptosis.
[0523]0.2 ml Annexin V-Biotin stock solution (100 μg/ml) was diluted in 4.6 ml 2× Ca2+ binding buffer and 2.5% BSA (1:25 dilution). 50 μl of the diluted Annexin V-Biotin solution was added to each well (except controls) to a final concentration of 1.0 μg/ml. The samples were incubated for 10-15 minutes with Annexin-Biotin prior to direct addition of 35S-Streptavidin. 35S-Streptavidin was diluted in 2× Ca2+ Binding buffer, 2.5% BSA and was added to all wells at a final concentration of 3×104 cpm/well. The plates were then sealed, centrifuged at 1000 rpm for 15 minutes and placed on orbital shaker for 2 hours. The analysis was performed on a 1450 Microbeta Trilux (Wallac). Percent above background represents the percentage amount of counts per minute above the negative controls. Percents greater than or equal to 30% above background are considered positive.
PDB12 Cell Inhibition
[0524]This assay will demonstrates if the compounds disclosed herein have efficacy in inhibiting protein production by PDB12 pancreatic ductal cells and are, therefore, useful in the therapeutic treatment of disorders which involve protein secretion by the pancreas, including diabetes, and the like.
[0525]PDB12 pancreatic ductal cells are plated on fibronectin coated 96 well plates at 1.5×103 cells per well in 100 μL/180 μL of growth media. 100 μL of growth media with the compound test sample or negative control lacking the compound is then added to well, for a final volume of 200 μL. Controls contain growth medium containing a protein shown to be inactive in this assay. Cells are incubated for 4 days at 37° C. 20 μL of Alamar Blue Dye (AB) is then added to each well and the flourescent reading is measured at 4 hours post addition of AB, on a microtiter plate reader at 530 nm excitation and 590 nm emission. The standard employed is cells without Bovine Pituitary Extract (BPE) and with various concentrations of BPE. Buffer or CM controls from unknowns are run 2 times on each 96 well plate.
[0526]These assays allow one to calculate a percent decrease in protein production by comparing the Alamar Blue Dye calculated protein concentration produced by the compound-treated cells with the Alamar Blue Dye calculated protein concentration produced by the negative control cells. A percent decrease in protein production of greater than or equal to 25% as compared to the negative control cells is considered positive.
Stimulation of Adult Heart Hypertrophy
[0527]This assay is designed to measure the ability of the various compounds disclosed herein to stimulate hypertrophy of adult heart. A positive test in this assay indicates that the compound would be expected to be useful for the therapeutic treatment of various cardiac insufficiency disorders.
[0528]Ventricular myocytes freshly isolated from adult (250 g) Sprague Dawley rats are plated at 2000 cell/well in 180 μl volume. Cells are isolated and plated on day 1, the compound-containing test samples or growth medium only (negative control) (20 μl volume) is added on day 2 and the cells are then fixed and stained on day 5. After staining, cell size is visualized wherein cells showing no growth enhancement as compared to control cells are given a value of 0.0, cells showing small to moderate growth enhancement as compared to control cells are given a value of 1.0 and cells showing large growth enhancement as compared to control cells are given a value of 2.0. Any degree of growth enhancement as compared to the negative control cells is considered positive for the assay.
PDB12 Cell Proliferation
[0529]This assay demonstrates whether the various compounds disclosed herein have efficacy in inducing proliferation of PDB12 pancreatic ductal cells and are, therefore, useful in the therapeutic treatment of disorders which involve protein secretion by the pancreas, including diabetes, and the like.
[0530]PDB12 pancreatic ductal cells are plated on fibronectin coated 96 well plates at 1.5×103 cells per well in 100 μL/180 μL of growth media. 100 μL of growth media with the compound test sample or negative control lacking the compound tested is then added to well, for a final volume of 200 μL. Controls contain growth medium containing a protein shown to be inactive in this assay. Cells are incubated for 4 days at 37° C. 20 μL of Alamar Blue Dye (AB) is then added to each well and the flourescent reading is measured at 4 hours post addition of AB, on a microtiter plate reader at 530 nm excitation and 590 nm emission. The standard employed is cells without Bovine Pituitary Extract (BPE) and with various concentrations of BPE. Buffer or growth medium only controls from unknowns are run 2 times on each 96 well plate.
[0531]Percent increase in protein production is calculated by comparing the Alamar Blue Dye calculated protein concentration produced by the test compound-treated cells with the Alamar Blue Dye calculated protein concentration produced by the negative control cells. A percent increase in protein production of greater than or equal to 25% as compared to the negative control cells is considered positive.
Enhancement of Heart Neonatal Hypertrophy
[0532]This assay is designed to measure the ability of the compounds disclosed herein to stimulate hypertrophy of neonatal heart. Testing positive in this assay indicates the compounds to be useful for the therapeutic treatment of various cardiac insufficiency disorders.
[0533]Cardiac myocytes from 1-day old Harlan Sprague Dawley rats are obtained. Cells (180 μl at 7.5×104/ml, serum <0.1%, freshly isolated) are added on day 1 to 96-well plates previously coated with DMEM/F12+4% FCS. Test samples containing the test compound or growth medium only (negative control) (201/well) are added directly to the wells on day 1. PGF (20 μl/well) is then added on day 2 at final concentration of 10-6 M. The cells are then stained on day 4 and visually scored on day 5, wherein cells showing no increase in size as compared to negative controls are scored 0.0, cells showing a small to moderate increase in size as compared to negative controls are scored 1.0 and cells showing a large increase in size as compared to negative controls are scored 2.0. A positive result in the assay is a score of 1.0 or greater.
Stimulatory Activity in Mixed Lymphocyte Reaction (MLR) Assay
[0534]This assay is used to determine if the compounds disclosed herein are active as a stimulator of the proliferation of stimulated T-lymphocytes. Compounds which stimulate proliferation of lymphocytes are useful therapeutically where enhancement of an immune response is beneficial. A therapeutic agent may take the form of antagonists of the compounds of the invention, for example, murine-human chimeric, humanized or human antibodies against the compound.
[0535]The basic protocol for this assay is described in Current Protocols in Immunology, unit 3.12; edited by J E Coligan, A M Kruisbeek, D H Marglies, E M Shevach, W Strober, National Insitutes of Health, Published by John Wiley & Sons, Inc.
[0536]More specifically, in one assay variant, peripheral blood mononuclear cells (PBMC) are isolated from mammalian individuals, for example a human volunteer, by leukopheresis (one donor will supply stimulator PBMCs, the other donor will supply responder PBMCs). If desired, the cells are frozen in fetal bovine serum and DMSO after isolation. Frozen cells may be thawed overnight in assay media (37° C., 5% CO2) and then washed and resuspended to 3×106 cells/ml of assay media (RPMI; 10% fetal bovine serum, 1% penicillin/streptomycin, 1% glutamine, 1% HEPES, 1% non-essential amino acids, 1% pyruvate). The stimulator PBMCs are prepared by irradiating the cells (about 3000 Rads).
[0537]The assay is prepared by plating in triplicate wells a mixture of:
[0538]100:1 of test sample diluted to 1% or to 0.1%, 50:1 of irradiated stimulator cells, and 50:1 of responder PBMC cells.
[0539]100 microliters of cell culture media or 100 microliter of CD4-IgG is used as the control. The wells are then incubated at 37° C., 5% CO2 for 4 days. On day 5, each well is pulsed with tritiated thymidine (1.0 mC/well; Amersham). After 6 hours the cells are washed 3 times and then the uptake of the label is evaluated.
[0540]In another variant of this assay, PBMCs are isolated from the spleens of Balb/c mice and C57B6 mice. The cells are teased from freshly harvested spleens in assay media (RPMI; 10% fetal bovine serum, 1% penicillin/streptomycin, 1% glutamine, 1% HEPES, 1% non-essential amino acids, 1% pyruvate) and the PBMCs are isolated by overlaying these cells over Lympholyte M (Organon Teknika), centrifuging at 2000 rpm for 20 minutes, collecting and washing the mononuclear cell layer in assay media and resuspending the cells to 1×107 cells/ml of assay media. The assay is then conducted as described above.
[0541]Positive increases over control are considered positive with increases of greater than or equal to 180% being preferred. However, any value greater than control indicates a stimulatory effect for the test protein.
Pericyte c-Fos Induction
[0542]This assay shows the ability of the compounds disclosed herein of the invention act to induce the expression of c-fos in pericyte cells and, therefore, their use not only as diagnostic markers for particular types of pericyte-associated tumors but also for giving rise to antagonists which would be expected to be useful for the therapeutic treatment of pericyte-associated tumors. Specifically, on day 1, pericytes are received from VEC Technologies and all but 5 ml of media is removed from flask. On day 2, the pericytes are trypsinized, washed, spun and then plated onto 96 well plates. On day 7, the media is removed and the pericytes are treated with 100 μl of test compound samples and controls (positive control=DME+5% serum +/-PDGF at 500 ng/ml; negative control=protein 32). Replicates are averaged and SD/CV are determined. Fold increase over Protein 32 (buffer control) value indicated by chemiluminescence units (RLU) luminometer reading verses frequency is plotted on a histogram. Two-fold above Protein 32 value is considered positive for the assay. ASY Matrix: Growth media=low glucose DMEM=20% FBS+1× pen/strep+1× fungizone. Assay Media=low glucose DMEM+5% FBS.
Ability of the Compounds of the Invention to Stimulate the Release of Proteoglycans from Cartilage
[0543]The ability of the compounds disclosed herein to stimulate the release of proteoglycans from cartilage tissue can be tested as follows.
[0544]The metacarphophalangeal joint of 4-6 month old pigs is aseptically dissected, and articular cartilage was removed by free hand slicing being careful to avoid the underlying bone. The cartilage was minced and cultured in bulk for 24 hours in a humidified atmosphere of 95% air, 5% CO2 in serum free (SF) media (DME/F12 1:1) with 0.1% BSA and 100 U/ml penicillin and 100 μg/ml streptomycin. After washing three times, approximately 100 mg of articular cartilage is aliquoted into micronics tubes and incubated for an additional 24 hours in the above SF media. The compound is then added at 1% either alone or in combination with 18 ng/ml interleukin-1 alpha, a known stimulator of proteoglycan release from cartilage tissue. The supernatant is then harvested and assayed for the amount of proteoglycans using the 1,9-dimethyl-methylene blue (DMB) calorimetric assay (Farndale and Buttle, Biochem. Biophys. Acta 883:173-177 (1985)). A positive result in this assay indicates that the test compound will find use, for example, in the treatment of sports-related joint problems, articular cartilage defects, osteoarthritis or rheumatoid arthritis.
Skin Vascular Permeability Assay
[0545]This assay is used to test whether compounds of the invention stimulate an immune response and induce inflammation by inducing mononuclear cell, eosinophil and PMN infiltration at the site of injection of the animal. Compounds which stimulate an immune response are useful therapeutically where stimulation of an immune response is beneficial. This skin vascular permeability assay is conducted as follows. Hairless guinea pigs weighing 350 grams or more are anesthetized with ketamine (75-80 mg/Kg) and 5 mg/Kg xylazine intramuscularly (IM). A sample of purified compound of the invention or a conditioned media test sample is injected intradermally onto the backs of the test animals with 100 μl per injection site. It is possible to have about 10-30, preferably about 16-24, injection sites per animal. One μl of Evans blue dye (1% in physiologic buffered saline) is injected intracardially. Blemishes at the injection sites are then measured (mm diameter) at 1 hr and 6 hr post injection. Animals were sacrificed at 6 hrs after injection. Each skin injection site is biopsied and fixed in formalin. The skins are then prepared for histopathologic evaluation. Each site is evaluated for inflammatory cell infiltration into the skin. Sites with visible inflammatory cell inflammation are scored as positive. Inflammatory cells may be neutrophilic, eosinophilic, monocytic or lymphocytic. At least a minimal perivascular infiltrate at the injection site is scored as positive, no infiltrate at the site of injection is scored as negative.
Enhancement of Heart Neonatal Hypertrophy Induced by F2a
[0546]This assay is designed to measure the ability of compounds disclosed herein to stimulate hypertrophy of neonatal heart, a positive test indicating usefulness for the therapeutic treatment of various cardiac insufficiency disorders.
[0547]Cardiac myocytes from 1-day old Harlan Sprague Dawley rats were obtained. Cells (180 μl at 7.5×104/ml, serum <0.1%, freshly isolated) are added on day 1 to 96-well plates previously coated with DMEM/F12+4% FCS. Test samples containing the test compound (20 μl/well) are added directly to the wells on day 1. PGF (20 μl/well) is then added on day 2 at a final concentration of 10-6 M. The cells are then stain on day 4 and visually scored on day ˜5. Visual scores are based on cell size, wherein cells showing no increase in size as compared to negative controls are scored 0.0, cells showing a small to moderate increase in size as compared to negative controls are scored 1.0 and cells showing a large increase in size as compared to negative controls are scored 2.0. A score of 1.0 or greater is considered positive.
[0548]No PBS is included, since calcium concentration is critical for assay response. Plates are coated with DMEM/F12 plus 4% FCS (200 μl/well). Assay media included: DMEM/F12 (with 2.44 gm bicarbonate), μg/ml transferrin, 1 μg/ml insulin, 1 μg/ml aprotinin, 2 mmol/L glutamine, 100 U/ml penicillin G, 100 μg/ml streptomycin. Protein buffer containing mannitol (4%) gave a positive signal (score 3.5) at 1/10 (0.4%) and 1/100 (0.04%), but not at 1/1000 (0.004%). Therefore the test sample buffer containing mannitol is not run.
Inhibitory Activity in Mixed Lymphocyte Reaction (MLR) Assay
[0549]This example shows that one or more of the compound of the invention are active as inhibitors of the proliferation of stimulated T-lymphocytes. Compounds which inhibit proliferation of lymphocytes are useful therapeutically where suppression of an immune response is beneficial.
[0550]The basic protocol for this assay is described in Current Protocols in Immunology, unit 3.12; edited by J E Coligan, A M Kruisbeek, D H Marglies, E M Shevach, W Strober, National Insitutes of Health, Published by John Wiley & Sons, Inc.
[0551]More specifically, in one assay variant, peripheral blood mononuclear cells (PBMC) are isolated from mammalian individuals, for example a human volunteer, by leukopheresis (one donor will supply stimulator PBMCs, the other donor will supply responder PBMCs). If desired, the cells are frozen in fetal bovine serum and DMSO after isolation. Frozen cells may be thawed overnight in assay media (37° C., 5% CO2) and then washed and resuspended to 3×106 cells/ml of assay media (RPMI; 10% fetal bovine serum, 1% penicillin/streptomycin, 1% glutamine, 1% HEPES, 1% non-essential amino acids, 1% pyruvate). The stimulator PBMCs are prepared by irradiating the cells (about 3000 Rads).
[0552]The assay is prepared by plating in triplicate wells a mixture of:
100:1 of test sample diluted to 1% or to 0.1%, 50:1 of irradiated stimulator cells, and50:1 of responder PBMC cells.
[0553]100 microliters of cell culture media or 100 microliter of CD4-IgG is used as the control. The wells are then incubated at 37° C., 5% CO2 for 4 days. On day 5, each well is pulsed with tritiated thymidine (1.0 mC/well; Amersham). After 6 hours the cells are washed 3 times and then the uptake of the label is evaluated.
[0554]In another variant of this assay, PBMCs are isolated from the spleens of Balb/c mice and C57B6 mice. The cells are teased from freshly harvested spleens in assay media (RPMI; 10% fetal bovine serum, 1% penicillin/streptomycin, 1% glutamine, 1% HEPES, 1% non-essential amino acids, 1% pyruvate) and the PBMCs are isolated by overlaying these cells over Lympholyte M (Organon Teknika), centrifuging at 2000 rpm for 20 minutes, collecting and washing the mononuclear cell layer in assay media and resuspending the cells to 1×107 cells/ml of assay media. The assay is then conducted as described above.
[0555]Any decreases below control is considered to be a positive result for an inhibitory compound, with decreases of less than or equal to 80% being preferred. However, any value less than control indicates an inhibitory effect for the test protein.
Induction of Endothelial Cell Apoptosis (ELISA)
[0556]The ability of the compounds disclosed herein to induce apoptosis in endothelial cells can be tested in human venous umbilical vein endothelial cells (HUVEC, Cell Systems) using a 96-well format, in 0% serum media supplemented with 100 ng/ml VEGF, 0.1% BSA, 1× pen/strep.
[0557]A positive result in this assay indicates the usefulness of the compound for therapeutically treating any of a variety of conditions associated with undesired endothelial cell growth including, for example, the inhibition of tumor growth. Coating of 96 well plates can be prepared by allowing gelatinization to occur for >30 minutes with 100 μl of 0.2% gelatin in PBS solution. The gelatin mix is aspirated thoroughly before plating HUVEC cells at a final concentration of 2×104 cells/ml in 10% serum containing medium--100 μl volume per well. The cells were grown for 24 hours before adding test samples containing the compound of interest.
[0558]To all wells, 100 μl of 0% serum media (Cell Systems) complemented with 100 ng/ml VEGF, 0.1% BSA, 1× pen/strep is added. Test samples containing the test compound were added in triplicate at dilutions of 1%, 0.33% and 0.11%. Wells without cells were used as a blank and wells with cells only are used as a negative control. As a positive control, 1:3 serial dilutions of 50 μl of a 3× stock of staurosporine are used. The cells were incubated for 24 to 35 hours prior to ELISA.
[0559]ELISA is used to determine levels of apoptosis preparing solutions according to the Boehringer Manual [Boehringer, Cell Death Detection ELISA plus, Cat No. 1 920 685]. Sample preparations: 96 well plates are spun down at 1 krpm for 10 minutes (200 g); the supernatant is removed by fast inversion, placing the plate upside down on a paper towel to remove residual liquid. To each well, 200 μl of 1× Lysis buffer is added and incubation allowed at room temperature for 30 minutes without shaking. The plates were spun down for 10 minutes at 1 krpm, and 20 μl of the lysate (cytoplasmic fraction) is transferred into streptavidin coated MTP. 80 μl of immunoreagent mix was added to the 20 μl lystate in each well. The MTP was covered with adhesive foil and incubated at room temperature for 2 hours by placing it on an orbital shaker (200 rpm). After two hours, the supernatant was removed by suction and the wells rinsed three times with 250 μl of 1× incubation buffer per well (removed by suction). Substrate solution was added (100 μl) into each well and incubated on an orbital shaker at room temperature at 250 rpm until color development was sufficient for a photometric analysis (approx. after 10-20 minutes). A 96 well reader was used to read the plates at 405 nm, reference wavelength, 492 nm. The levels obtained for PIN 32 (control buffer) was set to 100%. Samples with levels >130% were considered positive for induction of apoptosis.
Human Venous Endothelial Cell Calcium Flux Assay
[0560]This assay is designed to determine whether compounds of the present invention show the ability to stimulate calcium flux in human umbilical vein endothelial cells (HUVEC, Cell Systems). Calcium influx is a well documented response upon binding of certain ligands to their receptors. A test compound that results in a positive response in the present calcium influx assay can be said to bind to a specific receptor and activate a biological signaling pathway in human endothelial cells. This could ultimately lead, for example, to endothelial cell division, inhibition of endothelial cell proliferation, endothelial tube formation, cell migration, apoptosis, etc.
[0561]Human venous umbilical vein endothelial cells (HUVEC, Cell Systems) in growth media (50:50 without glycine, 1% glutamine, 10 mM Hepes, 10% FBS, 10 ng/ml bFGF), are plated on 96-well microtiter ViewPlates-96 (Packard Instrument Company Part #6005182) microtiter plates at a cell density of 2×104 cells/well. The day after plating, the cells are washed three times with buffer (HBSS plus 10 mM Hepes), leaving 100 μl/well. Then 100 μl/well of 8 μM Fluo-3 (2× is added. The cells are incubated for 1.5 hours at 37° C./5% CO2. After incubation, the cells are then washed 3× with buffer (described above) leaving 100 μl/well. Test samples of the compound are prepared on different 96-well plates at 5× concentration in buffer. The positive control corresponded to 50 μM ionomycin (5×); the negative control corresponded to Protein 32. Cell plate and sample plates are run on a FLIPR (Molecular Devices) machine. The FLIPR machine added 25 μl of test sample to the cells, and readings are taken every second for one minute, then every 3 seconds for the next three minutes.
[0562]The fluorescence change from baseline to the maximum rise of the curve (Δ change) is calculated, and replicates averaged. The rate of fluorescence increase is monitored, and only those samples which had a Δ change greater than 1000 and a rise within 60 seconds, are considered positive.
Fibroblast (BHK-21) Proliferation
[0563]This assay will show if the compounds of the invention act to induce proliferation of mammalian fibroblast cells in culture and, therefore, function is useful growth factors in mammalian systems.
[0564]The assay is performed is follows. BHK-21 fibroblast cells plated in standard growth medium at 2500 cells/well in a total volume of 100 μl. The compound, beta-FGF (positive control) or nothing (negative control) are then added to the wells in the presence of 1 μg/ml of heparin for a total final volume of 200 μl. The cells are then incubated at 37° C. for 6 to 7 days. After incubation, the media is removed, the cells are washed with PBS and then an acid phosphatase substrate reaction mixture (100 μl/well) is added. The cells are then incubated at 37° C. for 2 hours. 10 μL per well of 1N NaOH is then added to stop the acid phosphatase reaction. The plates are then read at OD 405 nm. A positive in the assay is acid phosphatase activity which is at least 50% above the negative control.
Inhibition of Heart Adult Hypertrophy
[0565]This assay is designed to measure the inhibition of heart adult hypertrophy. Compounds testing positive in this assay may find use in the therapeutic treatment of cardiac disorders associated with cardiac hypertrophy.
[0566]Ventricular myocytes are freshly isolated from adult (250 g) Harlan Sprague Dawley rats and the cells are plated at 2000/well in 180 μl volume. On day two, test samples (20 μL) containing the test compound are added. On day five, the cells are fixed and then stained. An increase in ANP message can also be measured by PCR from cells after a few hours. Results are based on a visual score of cell size: 0=no inhibition, -1=small inhibition, -2=large inhibition. A score of less than 0 is considered positive. Activity reference corresponds to phenylephrine (PE) at 0.1 mM, is a positive control. Assay media included: M199 (modified)-glutamine free, NaHCO3, phenol red, supplemented with 100 nM insulin, 0.2% BSA, 5 mM cretine, 2 mM L-carnitine, 5 mM taurine, 100 U/ml penicillin G, 100 μg/ml streptomycin (CCT medium). Only inner 60 wells are used in 96 well plates. Of these, 6 wells are reserved for negative and positive (PE) controls.
Induction of c-Fos in Endothelial Cells
[0567]This assay is designed to determine whether compounds of the invention show the ability to induce c-fos in endothelial cells. Compounds testing positive in this assay would be expected to be useful for the therapeutic treatment of conditions or disorders where angiogenesis would be beneficial including, for example, wound healing, and the like (is would agonists of these compounds). Antagonists of the compounds testing positive in this assay would be expected to be useful for the therapeutic treatment of cancerous tumors.
[0568]Human venous umbilical vein endothelial cells (HUVEC, Cell Systems) in growth media (50% Ham's F12 w/o GHT: low glucose, and 50% DMEM without glycine: with NaHCO3, 1% glutamine, 10 mM HEPES, 10% FBS, 10 ng/ml bFGF) are plated on 96-well microtiter plates at a cell density of 1×104 cells/well. The day after plating, the cells are starved by removing the growth media and treating the cells with 100 μl/well test samples and controls (positive control=growth media; negative control=Protein 32 buffer=10 mM HEPES, 140 mM NaCl, 4% (w/v) mannitol, pH 6.8). The cells are incubated for 30 minutes at 37° C., in 5% CO2. The samples are removed, and the first part of the bDNA kit protocol (Chiron Diagnostics, cat. #6005-037) is followed, where each capitalized reagent/buffer listed below is available from the kit.
[0569]Briefly, the amounts of the TM Lysis Buffer and Probes needed for the tests are calculated based on information provided by the manufacturer. The appropriate amounts of thawed Probes are added to the TM Lysis Buffer. The Capture Hybridization Buffer is warmed to room temperature. The bDNA strips are set up in the metal strip holders, and 100 μl of Capture Hybridization Buffer is added to each b-DNA well needed, followed by incubation for at least 30 minutes. The test plates with the cells are removed from the incubator, and the media is gently removed using the vacuum manifold. 100 μl of Lysis Hybridization Buffer with Probes are quickly pipetted into each well of the microtiter plates. The plates are then incubated at 55° C. for 15 minutes. Upon removal from the incubator, the plates are placed on the vortex mixer with the microtiter adapter head and vortexed on the #2 setting for one minute. 80 μl of the lysate is removed and added to the bDNA wells containing the Capture Hybridization Buffer, and pipetted up and down to mix. The plates are incubated at 53° C. for at least 16 hours.
[0570]On the next day, the second part of the bDNA kit protocol is followed. Specifically, the plates are removed from the incubator and placed on the bench to cool for 10 minutes. The volumes of additions needed are calculated based upon information provided by the manufacturer. An Amplifier Working Solution is prepared by making a 1:100 dilution of the Amplifier Concentrate (20 fm/μl) in AL Hybridization Buffer. The hybridization mixture is removed from the plates and washed twice with Wash A. 50 μl of Amplifier Working Solution is added to each well and the wells are incubated at 53° C. for 30 minutes. The plates are then removed from the incubator and allowed to cool for 10 minutes. The Label Probe Working Solution is prepared by making a 1:100 dilution of Label Concentrate (40 pmoles/μl) in AL Hybridization Buffer. After the 10-minute cool-down period, the amplifier hybridization mixture is removed and the plates are washed twice with Wash A. 50 μl of Label Probe Working Solution is added to each well and the wells are incubated at 53° C. for 15 minutes. After cooling for 10 minutes, the Substrate is warmed to room temperature. Upon addition of 3 μl of Substrate Enhancer to each ml of Substrate needed for the assay, the plates are allowed to cool for 10 minutes, the label hybridization mixture is removed, and the plates are washed twice with Wash A and three times with Wash D. 50 μl of the Substrate Solution with Enhancer is added to each well. The plates are incubated for 30 minutes at 37° C. and RLU is read in an appropriate luminometer.
[0571]The replicates are averaged and the coefficient of variation is determined. The measure of activity of the fold increase over the negative control (Protein 32/HEPES buffer described above) value is indicated by chemiluminescence units (RLU). The results are considered positive if the compound exhibits at least a two-fold value over the negative buffer control. Negative control=1.00 RLU at 1.00% dilution. Positive control=8.39 RLU at 1.00% dilution.
Guinea Pig Vascular Leak
[0572]This assay is designed to determine whether the compounds of the present invention show the ability to induce vascular permeability. Compounds testing positive in this assay are expected to be useful for the therapeutic treatment of conditions which would benefit from enhanced vascular permeability including, for example, conditions which may benefit from enhanced local immune system cell infiltration.
[0573]Hairless guinea pigs weighing 350 grams or more are anesthetized with Ketamine (75-80 mg/kg) and 5 mg/kg Xylazine intramuscularly. Test samples containing the tested compound or a physiological buffer without the test compound are injected into skin on the back of the test animals with 100 μl per injection site intradermally. There are approximately 16-24 injection sites per animal. One ml of Evans blue dye (1% in PBS) is then injected intracardially. Skin vascular permeability responses to the compounds (i.e., blemishes at the injection sites of injection) are visually scored by measuring the diameter (in mm) of blue-colored leaks from the site of injection at 1 and 6 hours post administration of the test materials. The mm diameter of blueness at the site of injection is observed and recorded is well is the severity of the vascular leakage. Blemishes of at least 5 mm in diameter are considered positive for the assay when testing purified proteins, being indicative of the ability to induce vascular leakage or permeability. A response greater than 7 mm diameter is considered positive for conditioned media samples. Human VEGF at 0.1 μg/100 μl is used is a positive control, inducing a response of 15-23 mm diameter.
Detection of Endothelial Cell Apoptosis (FACS)
[0574]The ability of the compounds disclosed herein to induce apoptosis in endothelial cells is tested in human venous umbilical vein endothelial cells (HUVEC, Cell Systems) in gelatinized T175 flasks using HUVEC cells below passage 10. Compounds testing positive in this assay are expected to be useful for therapeutically treating conditions where apoptosis of endothelial cells would be beneficial including, for example, the therapeutic treatment of tumors.
[0575]On day one, the cells are split [420,000 cells per gelatinized 6 cm dishes (11×103 cells/cm2 Falcon, Primaria)] and grown in media containing serum (CS-C, Cell System) overnight or for 16 hours to 24 hours.
[0576]On day 2, the cells are washed 1× with 5 ml PBS; 3 ml of 0% serum medium is added with VEGF (100 ng/ml); and 30 μl of the PRO test compound (final dilution 1%) or 0% serum medium (negative control) is added. The mixtures are incubated for 48 hours before harvesting.
[0577]The cells are then harvested for FACS analysis. The medium is aspirated and the cells washed once with PBS. 5 ml of 1× trypsin is added to the cells in a T-175 flask, and the cells are allowed to stand until they are released from the plate (about 5-10 minutes). Trypsinization is stopped by adding 5 ml of growth media. The cells are spun at 1000 rpm for 5 minutes at 4° C. The media is aspirated and the cells are resuspended in 10 ml of 10% serum complemented medium (Cell Systems), 5 μl of Annexin-FITC (BioVison) added and chilled tubes are submitted for FACS. A positive result is determined to be enhanced apoptosis in the compound treated samples is compared to the negative control.
Induction of c-Fos in Cortical Neurons
[0578]This assay is designed to determine whether the compounds tested show the ability to induce c-fos in cortical neurons. Compounds testing positive in this assay would be expected to be useful for the therapeutic treatment of nervous system disorders and injuries where neuronal proliferation would be beneficial.
[0579]Cortical neurons are dissociated and plated in growth medium at 10,000 cells per well in 96 well plates. After approximately 2 cellular divisions, the cells are treated for 30 minutes with the compound or nothing (negative control). The cells are then fixed for 5 minutes with cold methanol and stained with an antibody directed against phosphorylated CREB. mRNA levels are then calculated using chemiluminescence. A positive in the assay is any factor that results in at least a 2-fold increase in c-fos message is compared to the negative controls.
Stimulation of Endothelial Tube Formation
[0580]This assay is designed to determine whether the compounds of the invention show the ability to promote endothelial vacuole and lumen formation in the absence of exogenous growth factors. Compounds testing positive in this assay would be expected to be useful for the therapeutic treatment of disorders where endothelial vacuole and/or lumen formation would be beneficial including, for example, where the stimulation of pinocytosis, ion pumping, vascular permeability and/or junctional formation would be beneficial.
[0581]HUVEC cells (passage <8 from primary) are mixed with type I rat tail collagen (final concentration 2.6 mg/ml) at a density of 6×105 cells per ml and plated at 50 μl per well of M199 culture media supplement with 1% FBS and 1 μM 6-FAM-FITC dye to stain the vacuoles while they are forming and in the presence of the test compound. The cells are then incubated at 37° C./5% CO2 for 48 hours, fixed with 3.7% formalin room temperature for 10 minutes, washed 5 times with M199 medium and then stained with Rh-Phalloidin at 4° C. overnight followed by nuclear staining with 4 μM DAPI. A positive result in the assay is when vacuoles are present in greater than 50% of the cells.
Detection of Compounds of the Invention that Affect Glucose and/or FFA Uptake in Skeletal Muscle
[0582]This assay is designed to determine whether compounds of the invention show the ability to affect glucose or FFA uptake by skeletal muscle cells. Compounds testing positive in this assay would be expected to be useful for the therapeutic treatment of disorders where either the stimulation or inhibition of glucose uptake by skeletal muscle would be beneficial including, for example, diabetes or hyper- or hypo-insulinemia.
[0583]In a 96 well format, a compound to be assayed is added to primary rat differentiated skeletal muscle, and allowed to incubate overnight. Then fresh media with the compound and +/- insulin are added to the wells. The sample media is then monitored to determine glucose and FFA uptake by the skeletal muscle cells. The insulin will stimulate glucose and FFA uptake by the skeletal muscle, and insulin in media without the compound is used is a positive control, and a limit for scoring. As the compound being tested may either stimulate or inhibit glucose and FFA uptake, results are scored is positive in the assay if greater than 1.5 times or less than 0.5 times the insulin control.
Rod Photoreceptor Cell Survival Assay
[0584]This assay tests the ability of the compounds of this invention act to enhance the survival/proliferation of rod photoreceptor cells and, therefore, are useful for the therapeutic treatment of retinal disorders or injuries including, for example, treating sight loss in mammals due to retinitis pigmentosum, AMD, etc.
[0585]Sprague Dawley rat pups (postnatal day 7, mixed population: glia and netinal neural cell types) are killed by decapitation following CO2 anesthesia and the eyes removed under sterile conditions. The neural retina is dissected away from the pigment epithelium and other ocular tissue and then dissociated into a single cell suspension using 0.25% trypsin in Ca2+, Mg2+-free PBS. The retinas are incubated at 37° C. in this solution for 7-10 minutes after which the trypsin is inactivated by adding 1 ml soybean trypsin inhibitor. The cells are plated at a density of approximately 10,000 cells/ml into 96 well plates in DMEM/F12 supplemented with N2. Cells for all experiments are grown at 37° C. in a water saturated atmosphere of 5% CO2. After 7-10 days in culture, the cells are stained using calcein AM or CellTracker Green CMFDA and then fixed using 4% paraformaldehyde. Rho 4D2 (ascities or IgG 1:100) monoclonal antibody directed towards the visual pigment rhodopsin is used to detect rod photoreceptor cells by indirect immunofluorescence. The results are calculated is % survival: total number of calcein--rhodopsin positive cells at 7-10 days in culture, divided by the total number of rhodopsin positive cells at time 7-10 days in culture. The total cells (fluorescent) are quantified at 20× objective magnification using a CCD camera and NIH image software for MacIntosh. Fields in the well are chosen at random.
In Vitro Antitumor Assay
[0586]The antiproliferative activity of the compounds disclosed herein can be determined in the investigational, disease-oriented in vitro anti-cancer drug discovery assay of the National Cancer Institute (NCl), using a sulforhodamine B (SRB) dye binding assay essentially is described by Skehan et al., J. Natl. Cancer Inst. 82:1107-1112 (1990). The 60 tumor cell lines employed in this study ("the NCl panel"), as well as conditions for their maintenance and culture in vitro, have been described by Monks et al., J. Natl. Cancer Inst. 83:757-766 (1991). The purpose of this screen is to initially evaluate the cytotoxic and/or cytostatic activity of the test compounds against different types of tumors (Monks et al., supra; Boyd, Cancer: Princ. Pract. Oncol. Update 3(10):1-12 [1989]).
[0587]Cells from approximately 60 human tumor cell lines are harvested with trypsin/EDTA (Gibco), washed once, resuspended in IMEM and their viability is determined. The cell suspensions are added by pipet (100 μL volume) into separate 96-well microtiter plates. The cell density for the 6-day incubation is less than for the 2-day incubation to prevent overgrowth. Inoculates are allowed a preincubation period of 24 hours at 37° C. for stabilization. Dilutions at twice the intended test concentration are added at time zero in 100 μL aliquots to the microtiter plate wells (1:2 dilution). Test compounds are evaluated at five half-log dilutions (1000 to 100.000-fold). Incubations took place for two days and six days in a 5% CO2 atmosphere and 100% humidity.
[0588]After incubation, the medium is removed and the cells are fixed in 0.1 ml of 10% trichloroacetic acid at 40° C. The plates are rinsed five times with deionized water, dried, stained for 30 minutes with 0.1 ml of 0.4% sulforhodamine B dye (Sigma) dissolved in 1% acetic acid, rinsed four times with 1% acetic acid to remove unbound dye, dried, and the stain is extracted for five minutes with 0.1 ml of 10 mM Tris base [tris(hydroxymethyl)aminomethane], pH 10.5. The absorbance (OD) of sulforhodamine B at 492 nm is measured using a computer-interfaced, 96-well microtiter plate reader.
[0589]A test sample is considered positive if it shows at least 50% growth inhibitory effect at one or more concentrations.
Determination of Compounds of this Invention that Affect Glucose or FFA Uptake by Primary Rat Adipocytes
[0590]This assay is designed to determine whether the compounds of this invention show the ability to affect glucose or FFA uptake by adipocyte cells. Compounds testing positive in this assay would be expected to be useful for the therapeutic treatment of disorders where either the stimulation or inhibition of glucose uptake by adipocytes would be beneficial including, for example, obesity, diabetes or hyper- or hypo-insulinemia.
[0591]In a 96 well format, compounds to be assayed are added to primary rat adipocytes, and allowed to incubate overnight. Samples are taken at 4 and 16 hours and assayed for glycerol, glucose and FFA uptake. After the 16 hour incubation, insulin is added to the media and allowed to incubate for 4 hours. At this time, a sample is taken and glycerol, glucose and FFA uptake is measured. Media containing insulin without the compound is used as a positive reference control. As the compound being tested may either stimulate or inhibit glucose and FFA uptake, results are scored as positive in the assay if greater than 1.5 times or less than 0.5 times the insulin control.
Chondrocyte Re-Differentiation Assay
[0592]This assay shows whether the compounds of the invention act to induce redifferentiation of chondrocytes, and therefore are expected to be useful for the treatment of various bone and/or cartilage disorders such as, for example, sports injuries and arthritis. The assay is performed as follows. Porcine chondrocytes are isolated by overnight collagenase digestion of articulary cartilage of metacarpophalangeal joints of 4-6 month old female pigs. The isolated cells are then seeded at 25,000 cells/cm2 in Ham F-12 containing 10% FBS and 4 μg/ml gentamycin. The culture media is changed every third day and the cells are then seeded in 96 well plates at 5,000 cells/well in 100 μl of the same media without serum and 100 μl of the test compound, 5 nM staurosporin (positive control) or medium alone (negative control) is added to give a final volume of 200 μl/well. After 5 days of incubation at 37° C., a picture of each well is taken and the differentiation state of the chondrocytes is determined. A positive result in the assay occurs when the redifferentiation of the chondrocytes is determined to be more similar to the positive control than the negative control.
Fetal Hemoglobin Induction in an Erythroblastic Cell Line
[0593]This assay is useful for screening compounds for the ability to induce the switch from adult hemoglobin to fetal hemoglobin in an erythroblastic cell line. Compounds testing positive in this assay are expected to be useful for therapeutically treating various mammalian hemoglobin-associated disorders such as the various thalassemias. The assay is performed as follows. Erythroblastic cells are plated in standard growth medium at 1000 cells/well in a 96 well format. The tested compound is added to the growth medium at a concentration of 0.2% or 2% and the cells are incubated for 5 days at 37° C. As a positive control, cells are treated with 100 μM hemin and as a negative control, the cells are untreated. After 5 days, cell lysates are prepared and analyzed for the expression of gamma globin (a fetal marker). A positive in the assay is a gamma globin level at least 2-fold above the negative control.
Mouse Kidney Mesangial Cell Proliferation Assay
[0594]This assay shows whether compounds of the invention act to induce proliferation of mammalian kidney mesangial cells and therefore are useful for treating kidney disorders associated with decreased mesangial cell function such as Berger disease or other nephropathies associated with Schonlein-Henoch purpura, celiac disease, dermatitis herpetiformis or Crohn disease. The assay is performed as follows. On day one, mouse kidney mesangial cells are plated on a 96 well plate in growth media (3:1 mixture of Dulbecco's modified Eagle's medium and Ham's F12 medium, 95% fetal bovine serum, 5% supplemented with 14 mM HEPES) and grown overnight. On day 2, the test compound is diluted at 2 concentrations (1% and 0.1%) in serum-free medium and added to the cells. Control samples are serum-free medium alone. On day 4, 20 μl of the Cell Titer 96 Aqueous one solution reagent (Progema) was added to each well and the colormetric reaction was allowed to proceed for 2 hours. The absorbance (OD) is then measured at 490 nm. A positive in the assay is an absorbance reading which is at least 15% above the control reading.
Proliferation of Rat Utricular Supporting Cells
[0595]This assay is used to determine of compounds of the invention act as potent mitogens for inner ear supporting cells which are auditory hair cell progenitors and, therefore, are useful for inducing the regeneration of auditory hair cells and treating hearing loss in mammals. The assay is performed as follows. Rat UEC-4 utricular epithelial cells are aliquoted into 96 well plates with a density of 3000 cells/well in 200 μl of serum-containing medium at 33° C. The cells are cultured overnight and are then switched to serum-free medium at 37° C. Various dilutions of the compounds (or nothing for a control) are then added to the cultures and the cells are incubated for 24 hours. After the 24 hour incubation, 3H-thymidine (1 μCi/well) is added and the cells are then cultured for an additional 24 hours. The cultures are then washed to remove unincorporated radiolabel, the cells harvested and Cpm per well determined. Cpm of at least 30% or greater in the test compound treated cultures as compared to the control cultures is considered a positive in the assay.
Chondrocyte Proliferation Assay
[0596]This assay is designed to determine whether compounds of the present invention show the ability to induce the proliferation and/or redifferentiation of chondrocytes in culture. Compounds testing positive in this assay would be expected to be useful for the therapeutic treatment of various bone and/or cartilage disorders such as, for example, sports injuries and arthritis.
[0597]Porcine chondrocytes are isolated by overnight collagenase digestion of articular cartilage of the metacarpophalangeal joint of 4-6 month old female pigs. The isolated cells are then seeded at 25,000 cells/cm2 in Ham F-12 containing 10% FBS and 4 μg/ml gentamycin. The culture media is changed every third day and the cells are reseeded to 25,000 cells/cm2 every five days. On day 12, the cells are seeded in 96 well plates at 5,000 cells/well in 100 μl of the same media without serum and 100 μl of either serum-free medium (negative control), staurosporin (final concentration of 5 nM; positive control) or the test compound are added to give a final volume of 200 μl/well. After 5 days at 37° C., 20 μl of Alamar blue is added to each well and the plates are incubated for an additional 3 hours at 37° C. The fluorescence is then measured in each well (Ex: 530 nm; Em: 590 nm). The fluorescence of a plate containing 200 μl of the serum-free medium is measured to obtain the background. A positive result in the assay is obtained when the fluorescence of the compound treated sample is more like that of the positive control than the negative control.
Inhibition of Heart Neonatal Hypertrophy Induced by LIF+ET-1
[0598]This assay is designed to determine whether the compounds of the present invention show the ability to inhibit neonatal heart hypertrophy induced by LIF and endothelin-1 (ET-1). A test compound that provides a positive response in the present assay would be useful for the therapeutic treatment of cardiac insufficiency diseases or disorders characterized or associated with an undesired hypertrophy of the cardiac muscle.
[0599]Cardiac myocytes from 1-day old Harlan Sprague Dawley rats (180 μl at 7.5×104/ml, serum <0.1, freshly isolated) are introduced on day 1 to 96-well plates previously coated with DMEM/F12+4% FCS. Test compound samples or growth medium alone (negative control) are then added directly to the wells on day 2 in 20 μl volume. LIF+ET-1 are then added to the wells on day 3. The cells are stained after an additional 2 days in culture and are then scored visually the next day. A positive in the assay occurs when the compound treated myocytes are visually smaller on the average or less numerous than the untreated myocytes.
Sequence CWU
1
19914195DNAEscherichia coli 1ctaaattgta agcgttaata ttttgttaaa attcgcgtta
aatttttgtt aaatcagctc 60attttttaac caataggccg aaatcggcaa aatcccttat
aaatcaaaag aatagaccga 120gatagggttg agtgttgttc cagtttggaa caagagtcca
ctattaaaga acgtggactc 180caacgtcaaa gggcgaaaaa ccgtctatca gggcgatggc
ccactacgtg aaccatcacc 240ctaatcaagt tttttggggt cgaggtgccg taaagcacta
aatcggaacc ctaaagggag 300cccccgattt agagcttgac ggggaaagcc ggcgaacgtg
gcgagaaagg aagggaagaa 360agcgaaagga gcgggcgcta gggcgctggc aagtgtagcg
gtcacgctgc gcgtaaccac 420cacacccgcc gcgcttaatg cgccgctaca gggcgcgtcc
cattcgccat tcaggctgcg 480caactgttgg gaagggcgat cggtgcgggc ctcttcgcta
ttacgccagc tggcgaaagg 540gggatgtgct gcaaggcgat taagttgggt aacgccaggg
ttttcccagt cacgacgttg 600taaaacgacg gccagtgagc gcgcaagcgg ccgcaacccg
ggaaaagctt ggccattgca 660tacgttgtat ccatatcata atatgtacat ttatattggc
tcatgtccaa cattaccgcc 720atgttgacat tgattattga ctagttatta atagtaatca
attacggggt cattagttca 780tagcccatat atggagttcc gcgttacata acttacggta
aatggcccgc ctggctgacc 840gcccaacgac ccccgcccat tgacgtcaat aatgacgtat
gttcccatag taacgccaat 900agggactttc cattgacgtc aatgggtgga gtatttacgg
taaactgccc acttggcagt 960acatcaagtg tatcatatgc caagtacgcc ccctattgac
gtcaatgacg gtaaatggcc 1020cgcctggcat tatgcccagt acatgacctt atgggacttt
cctacttggc agtacatcta 1080cgtattagtc atcgctatta ccatggtgat gcggttttgg
cagtacatca atgggcgtgg 1140atagcggttt gactcacggg gatttccaag tctccacccc
attgacgtca atgggagttt 1200gttttggcac caaaatcaac gggactttcc aaaatgtcgt
aacaactccg ccccattgac 1260gcaaatgggc ggtaggcgtg tacggtggga ggtctatata
agcagagctc gtttagtgaa 1320ccgtcagatc gcctggagac gccatccacg ctgttttgac
ctccatagaa gacaccggga 1380ccgatccagc ctccgcggcc gggaacggtg cattggaacg
cggattcccc gtgccaagag 1440tgacgtaagt accgcctata gagtctatag gcccaccccc
ttggcttctt atgcatgctc 1500ccctgctccg acccgggctc ctcgcccgcc cggacccaca
ggccaccctc aaccgtcctg 1560gccccggacc caaaccccac ccctcactct gcttctcccc
gcaggagaat tcaatcgcga 1620aagggcccaa agatctgcca taccacattt gtagaggttt
tacttgcttt aaaaaacctc 1680ccacacctcc ccctgaacct gaaacataaa atgaatgcaa
ttgttgttgt taacttgttt 1740attgcagctt ataatggtta caaataaagc aatagcatca
caaatttcac aaataaagca 1800tttttttcac tgcattctag ttgtggtttg tccaaactca
tcaatgtatc ttatcatgtc 1860tggagctagc atcccgcccc taactccgcc ctgttccgcc
cattctccgc cccatggctg 1920actaattttt tttatttatg cagaggccga ggccgcctcg
gcctctgagc tattccagaa 1980gtagtgagga ggcttttttg gaggcctagg cttttgcgtc
gagaagcgcg cttggcgtaa 2040tcatggtcat agctgtttcc tgtgtgaaat tgttatccgc
tcacaattcc acacaacata 2100cgagccggaa gcataaagtg taaagcctgg ggtgcctaat
gagtgagcta actcacatta 2160attgcgttgc gctcactgcc cgctttccag tcgggaaacc
tgtcgtgcca gctgcattaa 2220tgaatcggcc aacgcgcggg gagaggcggt ttgcgtattg
ggcgctcttc cgcttcctcg 2280ctcactgact cgctgcgctc ggtcgttcgg ctgcggcgag
cggtatcagc tcactcaaag 2340gcggtaatac ggttatccac agaatcaggg gataacgcag
gaaagaacat gtgagcaaaa 2400ggccagcaaa aggccaggaa ccgtaaaaag gccgcgttgc
tggcgttttt ccataggctc 2460cgcccccctg acgagcatca caaaaatcga cgctcaagtc
agaggtggcg aaacccgaca 2520ggactataaa gataccaggc gtttccccct ggaagctccc
tcgtgcgctc tcctgttccg 2580accctgccgc ttaccggata cctgtccgcc tttctccctt
cgggaagcgt ggcgctttct 2640catagctcac gctgtaggta tctcagttcg gtgtaggtcg
ttcgctccaa gctgggctgt 2700gtgcacgaac cccccgttca gcccgaccgc tgcgccttat
ccggtaacta tcgtcttgag 2760tccaacccgg taagacacga cttatcgcca ctggcagcag
ccactggtaa caggattagc 2820agagcgaggt atgtaggcgg tgctacagag ttcttgaagt
ggtggcctaa ctacggctac 2880actagaagga cagtatttgg tatctgcgct ctgctgaagc
cagttacctt cggaaaaaga 2940gttggtagct cttgatccgg caaacaaacc accgctggta
gcggtggttt ttttgtttgc 3000aagcagcaga ttacgcgcag aaaaaaagga tctcaagaag
atcctttgat cttttctacg 3060gggtctgacg ctcagtggaa cgaaaactca cgttaaggga
ttttggtcat gagattatca 3120aaaaggatct tcacctagat ccttttaaat taaaaatgaa
gttttaaatc aatctaaagt 3180atatatgagt aaacttggtc tgacagttac caatgcttaa
tcagtgaggc acctatctca 3240gcgatctgtc tatttcgttc atccatagtt gcctgactcc
ccgtcgtgta gataactacg 3300atacgggagg gcttaccatc tggccccagt gctgcaatga
taccgcgaga cccacgctca 3360ccggctccag atttatcagc aataaaccag ccagccggaa
gggccgagcg cagaagtggt 3420cctgcaactt tatccgcctc catccagtct attaattgtt
gccgggaagc tagagtaagt 3480agttcgccag ttaatagttt gcgcaacgtt gttgccattg
ctacaggcat cgtggtgtca 3540cgctcgtcgt ttggtatggc ttcattcagc tccggttccc
aacgatcaag gcgagttaca 3600tgatccccca tgttgtgcaa aaaagcggtt agctccttcg
gtcctccgat cgttgtcaga 3660agtaagttgg ccgcagtgtt atcactcatg gttatggcag
cactgcataa ttctcttact 3720gtcatgccat ccgtaagatg cttttctgtg actggtgagt
actcaaccaa gtcattctga 3780gaatagtgta tgcggcgacc gagttgctct tgcccggcgt
caatacggga taataccgcg 3840ccacatagca gaactttaaa agtgctcatc attggaaaac
gttcttcggg gcgaaaactc 3900tcaaggatct taccgctgtt gagatccagt tcgatgtaac
ccactcgtgc acccaactga 3960tcttcagcat cttttacttt caccagcgtt tctgggtgag
caaaaacagg aaggcaaaat 4020gccgcaaaaa agggaataag ggcgacacgg aaatgttgaa
tactcatact cttccttttt 4080caatattatt gaagcattta tcagggttat tgtctcatga
gcggatacat atttgaatgt 4140atttagaaaa ataaacaaat aggggttccg cgcacatttc
cccgaaaagt gccac 41952824DNAEscherichia coli 2gttaacgaat
tcccaccatg attgaacaag atggattgca cgcaggttct ccggccgctt 60gggtggagag
gctattcggc tatgactggg cacaacagac aatcggctgc tctgatgccg 120ccgtgttccg
gctgtcagcg caggggcgcc cggttctttt tgtcaagacc gacctgtccg 180gtgccctgaa
tgaactgcag gacgaggcag cgcggctatc gtggctggcc acgacgggcg 240ttccttgcgc
agctgtgctc gacgttgtca ctgaagcggg aagggactgg ctgctattgg 300gcgaagtgcc
ggggcaggat ctcctgtcat ctcaccttgc tcctgccgag aaagtatcca 360tcatggctga
tgcaatgcgg cggctgcata cgcttgatcc ggctacctgc ccattcgacc 420accaagcgaa
acatcgcatc gagcgagcac gtactcggat ggaagccggt cttgtcgatc 480aggatgatct
ggacgaagag catcaggggc tcgcgccagc cgaactgttc gccaggctca 540aggcgcgcat
gcccgacggc gaggatctcg tcgtgaccca tggcgatgcc tgcttgccga 600atatcatggt
ggaaaatggc cgcttttctg gattcatcga ctgtggccgg ctgggtgtgg 660cggaccgcta
tcaggacata gcgttggcta cccgtgatat tgctgaagag cttggcggcg 720aatgggctga
ccgcttcctc gtgctttacg gtatcgccgc tcccgattcg cagcgcatcg 780ccttctatcg
ccttcttgac gagttcttct gaagatctgt taac
8243785DNAArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 3gttaacgaat tcccaccatg ctgctgctgg cgagatgtct
gctgctagtc ctcgtctcct 60cgctgctggt atgctcggga ctggcgtgtg acaaaactca
cacatgccca ccgtgcccag 120cacctgaact cctgggggga ccgtcagtct tcctcttccc
cccaaaaccc aaggacaccc 180tcatgatctc ccggacccct gaggtcacat gcgtggtggt
ggacgtgagc cacgaagacc 240ctgaggtcaa gttcaactgg tacgtggacg gcgtggaggt
gcataatgcc aagacaaagc 300cgcgggagga gcagtacaac agcacgtacc gtgtggtcag
cgtcctcacc gtcctgcacc 360aggactggct gaatggcaag gagtacaagt gcaaggtctc
caacaaagcc ctcccagccc 420ccatcgagaa aaccatctcc aaagccaaag ggcagccccg
agaaccacag gtgtacaccc 480tgcccccatc ccgggatgag ctgaccaaga accaggtcag
cctgacctgc ctggtcaaag 540gcttctatcc cagcgacatc gccgtggagt gggagagcaa
tgggcagccg gagaacaact 600acaagaccac gcctcccgtg ctggactccg acggctcctt
cttcctctac agcaagctca 660ccgtggacaa gagcaggtgg cagcagggga acgtcttctc
atgctccgtg atgcatgagg 720ctctgcacaa ccactacacg cagaagagcc tctccctgtc
tccgggtaaa tgaagatctg 780ttaac
7854785DNAArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 4gttaacgaat tcccaccatg
gccttgacct ttgctttact ggtggccctc ctggtgctca 60gctgcaagtc aagctgctct
gtgggctgtg acaaaactca cacatgccca ccgtgcccag 120cacctgaact cctgggggga
ccgtcagtct tcctcttccc cccaaaaccc aaggacaccc 180tcatgatctc ccggacccct
gaggtcacat gcgtggtggt ggacgtgagc cacgaagacc 240ctgaggtcaa gttcaactgg
tacgtggacg gcgtggaggt gcataatgcc aagacaaagc 300cgcgggagga gcagtacaac
agcacgtacc gtgtggtcag cgtcctcacc gtcctgcacc 360aggactggct gaatggcaag
gagtacaagt gcaaggtctc caacaaagcc ctcccagccc 420ccatcgagaa aaccatctcc
aaagccaaag ggcagccccg agaaccacag gtgtacaccc 480tgcccccatc ccgggatgag
ctgaccaaga accaggtcag cctgacctgc ctggtcaaag 540gcttctatcc cagcgacatc
gccgtggagt gggagagcaa tgggcagccg gagaacaact 600acaagaccac gcctcccgtg
ctggactccg acggctcctt cttcctctac agcaagctca 660ccgtggacaa gagcaggtgg
cagcagggga acgtcttctc atgctccgtg atgcatgagg 720ctctgcacaa ccactacacg
cagaagagcc tctccctgtc tccgggtaaa tgaagatctg 780ttaac
7855767DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 5gttaacgaat tcccaccatg ctggctgcca cagtcctgac cctggccctg
ctgggcaatg 60cccatgcctg tgacaaaact cacacatgcc caccgtgccc agcacctgaa
ctcctggggg 120gaccgtcagt cttcctcttc cccccaaaac ccaaggacac cctcatgatc
tcccggaccc 180ctgaggtcac atgcgtggtg gtggacgtga gccacgaaga ccctgaggtc
aagttcaact 240ggtacgtgga cggcgtggag gtgcataatg ccaagacaaa gccgcgggag
gagcagtaca 300acagcacgta ccgtgtggtc agcgtcctca ccgtcctgca ccaggactgg
ctgaatggca 360aggagtacaa gtgcaaggtc tccaacaaag ccctcccagc ccccatcgag
aaaaccatct 420ccaaagccaa agggcagccc cgagaaccac aggtgtacac cctgccccca
tcccgggatg 480agctgaccaa gaaccaggtc agcctgacct gcctggtcaa aggcttctat
cccagcgaca 540tcgccgtgga gtgggagagc aatgggcagc cggagaacaa ctacaagacc
acgcctcccg 600tgctggactc cgacggctcc ttcttcctct acagcaagct caccgtggac
aagagcaggt 660ggcagcaggg gaacgtcttc tcatgctccg tgatgcatga ggctctgcac
aaccactaca 720cgcagaagag cctctccctg tctccgggta aatgaagatc tgttaac
7676767DNAArtificial Sequenceartificial construct relating to
Homo Sapiens immunoglobulin 6gttaacgaat tcccaccatg ctgctgctgg
cgagatgtct gctgctagtc ctcgtctcct 60cgctgctggt atgctcggga ctggcgtgcc
caccgtgccc agcacctgaa ctcctggggg 120gaccgtcagt cttcctcttc cccccaaaac
ccaaggacac cctcatgatc tcccggaccc 180ctgaggtcac atgcgtggtg gtggacgtga
gccacgaaga ccctgaggtc aagttcaact 240ggtacgtgga cggcgtggag gtgcataatg
ccaagacaaa gccgcgggag gagcagtaca 300acagcacgta ccgtgtggtc agcgtcctca
ccgtcctgca ccaggactgg ctgaatggca 360aggagtacaa gtgcaaggtc tccaacaaag
ccctcccagc ccccatcgag aaaaccatct 420ccaaagccaa agggcagccc cgagaaccac
aggtgtacac cctgccccca tcccgggatg 480agctgaccaa gaaccaggtc agcctgacct
gcctggtcaa aggcttctat cccagcgaca 540tcgccgtgga gtgggagagc aatgggcagc
cggagaacaa ctacaagacc acgcctcccg 600tgctggactc cgacggctcc ttcttcctct
acagcaagct caccgtggac aagagcaggt 660ggcagcaggg gaacgtcttc tcatgctccg
tgatgcatga ggctctgcac aaccactaca 720cgcagaagag cctctccctg tctccgggta
aatgaagatc tgttaac 7677767DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 7gttaacgaat tcccaccatg gccttgacct ttgctttact ggtggccctc
ctggtgctca 60gctgcaagtc aagctgctct gtgggctgcc caccgtgccc agcacctgaa
ctcctggggg 120gaccgtcagt cttcctcttc cccccaaaac ccaaggacac cctcatgatc
tcccggaccc 180ctgaggtcac atgcgtggtg gtggacgtga gccacgaaga ccctgaggtc
aagttcaact 240ggtacgtgga cggcgtggag gtgcataatg ccaagacaaa gccgcgggag
gagcagtaca 300acagcacgta ccgtgtggtc agcgtcctca ccgtcctgca ccaggactgg
ctgaatggca 360aggagtacaa gtgcaaggtc tccaacaaag ccctcccagc ccccatcgag
aaaaccatct 420ccaaagccaa agggcagccc cgagaaccac aggtgtacac cctgccccca
tcccgggatg 480agctgaccaa gaaccaggtc agcctgacct gcctggtcaa aggcttctat
cccagcgaca 540tcgccgtgga gtgggagagc aatgggcagc cggagaacaa ctacaagacc
acgcctcccg 600tgctggactc cgacggctcc ttcttcctct acagcaagct caccgtggac
aagagcaggt 660ggcagcaggg gaacgtcttc tcatgctccg tgatgcatga ggctctgcac
aaccactaca 720cgcagaagag cctctccctg tctccgggta aatgaagatc tgttaac
7678749DNAArtificial Sequenceartificial construct relating to
Homo Sapiens immunoglobulin 8gttaacgaat tcccaccatg ctggctgcca
cagtcctgac cctggccctg ctgggcaatg 60cccatgcctg cccaccgtgc ccagcacctg
aactcctggg gggaccgtca gtcttcctct 120tccccccaaa acccaaggac accctcatga
tctcccggac ccctgaggtc acatgcgtgg 180tggtggacgt gagccacgaa gaccctgagg
tcaagttcaa ctggtacgtg gacggcgtgg 240aggtgcataa tgccaagaca aagccgcggg
aggagcagta caacagcacg taccgtgtgg 300tcagcgtcct caccgtcctg caccaggact
ggctgaatgg caaggagtac aagtgcaagg 360tctccaacaa agccctccca gcccccatcg
agaaaaccat ctccaaagcc aaagggcagc 420cccgagaacc acaggtgtac accctgcccc
catcccggga tgagctgacc aagaaccagg 480tcagcctgac ctgcctggtc aaaggcttct
atcccagcga catcgccgtg gagtgggaga 540gcaatgggca gccggagaac aactacaaga
ccacgcctcc cgtgctggac tccgacggct 600ccttcttcct ctacagcaag ctcaccgtgg
acaagagcag gtggcagcag gggaacgtct 660tctcatgctc cgtgatgcat gaggctctgc
acaaccacta cacgcagaag agcctctccc 720tgtctccggg taaatgaaga tctgttaac
74991427DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 9gttaacgaat tcccaccatg agctttccat gtaaatttgt agccagcttc
cttctgattt 60tcaatgtttc ttccaaaggt gcagtctcca aaactcacac atgcccaccg
tgcccagcac 120ctgaactcct ggggggaccg tcagtcttcc tcttcccccc aaaacccaag
gacaccctca 180tgatctcccg gacccctgag gtcacatgcg tggtggtgga cgtgagccac
gaagaccctg 240aggtcaagtt caactggtac gtggacggcg tggaggtgca taatgccaag
acaaagccgc 300gggaggagca gtacaacagc acgtaccgtg tggtcagcgt cctcaccgtc
ctgcaccagg 360actggctgaa tggcaaggag tacaagtgca aggtctccaa caaagccctc
ccagccccca 420tcgagaaaac catctccaaa gccaaagggc agccccgaga accacaggtg
tacaccctgc 480ccccatcccg ggatgagctg accaagaacc aggtcagcct gacctgcctg
gtcaaaggct 540tctatcccag cgacatcgcc gtggagtggg agagcaatgg gcagccggag
aacaactaca 600agaccacgcc tcccgtgctg gactccgacg gctccttctt cctctacagc
aagctcaccg 660tggacaagag caggtggcag caggggaacg tcttctcatg ctccgtgatg
catgaggctc 720tgcacaacca ctacacgcag aagagcctct ccctgtctcc ggagctgcaa
ctggaggaga 780gctgtgcgga ggcgcaggac ggggagctgg acgggtgcgt atccggtgac
accattgtaa 840tgactagtgg cggtccgcgc actgtggctg aactggaggg caaaccgttc
accgcactga 900ttcgcggctc tggctaccca tgcccctcag gtttcttccg cacctgtgaa
cgtgacgtat 960atgatctgcg tacacgtgag ggtcattgct tacgtttgac ccatgatcac
cgtgttctgg 1020tgatggatgg tggcctggaa tggcgtgccg cgggtgaact ggaacgcggc
gaccgcctgg 1080tgatggatga tgcagctggc gagtttccgg cactggcaac cttccgtggc
ctgcgtggcg 1140ctggccgcca ggatgtttat gacgctactg tttacggtgc tagcgcattc
actgctaatg 1200gcttcattgt acacgcatgt ggcgagcagc ccgggaccgg tctgaactca
ggcctcacga 1260caaatcctgg tgtatccgct tggcaggtca acacagctta tactgcggga
caattggtca 1320catataacgg caagacgtat aaatgtttgc agccccacac ctccttggca
ggatgggaac 1380catccaacgt tcctgccttg tggcagcttc aatgaagatc tgttaac
1427101430DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 10gttaacgaat tcccaccatg aaccggggag
tcccttttag gcacttgctt ctggtgctgc 60aactggcgct cctcccagca gccactcagg
gaaaaactca cacatgccca ccgtgcccag 120cacctgaact cctgggggga ccgtcagtct
tcctcttccc cccaaaaccc aaggacaccc 180tcatgatctc ccggacccct gaggtcacat
gcgtggtggt ggacgtgagc cacgaagacc 240ctgaggtcaa gttcaactgg tacgtggacg
gcgtggaggt gcataatgcc aagacaaagc 300cgcgggagga gcagtacaac agcacgtacc
gtgtggtcag cgtcctcacc gtcctgcacc 360aggactggct gaatggcaag gagtacaagt
gcaaggtctc caacaaagcc ctcccagccc 420ccatcgagaa aaccatctcc aaagccaaag
ggcagccccg agaaccacag gtgtacaccc 480tgcccccatc ccgggatgag ctgaccaaga
accaggtcag cctgacctgc ctggtcaaag 540gcttctatcc cagcgacatc gccgtggagt
gggagagcaa tgggcagccg gagaacaact 600acaagaccac gcctcccgtg ctggactccg
acggctcctt cttcctctac agcaagctca 660ccgtggacaa gagcaggtgg cagcagggga
acgtcttctc atgctccgtg atgcatgagg 720ctctgcacaa ccactacacg cagaagagcc
tctccctgtc tccggagctg caactggagg 780agagctgtgc ggaggcgcag gacggggagc
tggacgggtg cgtatccggt gacaccattg 840taatgactag tggcggtccg cgcactgtgg
ctgaactgga gggcaaaccg ttcaccgcac 900tgattcgcgg ctctggctac ccatgcccct
caggtttctt ccgcacctgt gaacgtgacg 960tatatgatct gcgtacacgt gagggtcatt
gcttacgttt gacccatgat caccgtgttc 1020tggtgatgga tggtggcctg gaatggcgtg
ccgcgggtga actggaacgc ggcgaccgcc 1080tggtgatgga tgatgcagct ggcgagtttc
cggcactggc aaccttccgt ggcctgcgtg 1140gcgctggccg ccaggatgtt tatgacgcta
ctgtttacgg tgctagcgca ttcactgcta 1200atggcttcat tgtacacgca tgtggcgagc
agcccgggac cggtctgaac tcaggcctca 1260cgacaaatcc tggtgtatcc gcttggcagg
tcaacacagc ttatactgcg ggacaattgg 1320tcacatataa cggcaagacg tataaatgtt
tgcagcccca cacctccttg gcaggatggg 1380aaccatccaa cgttcctgcc ttgtggcagc
ttcaatgaag atctgttaac 1430111430DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 11gttaacgaat tcccaccatg ctgctgctgg cgagatgtct gctgctagtc
ctcgtctcct 60cgctgctggt atgctcggga ctggcgtgtg acaaaactca cacatgccca
ccgtgcccag 120cacctgaact cctgggggga ccgtcagtct tcctcttccc cccaaaaccc
aaggacaccc 180tcatgatctc ccggacccct gaggtcacat gcgtggtggt ggacgtgagc
cacgaagacc 240ctgaggtcaa gttcaactgg tacgtggacg gcgtggaggt gcataatgcc
aagacaaagc 300cgcgggagga gcagtacaac agcacgtacc gtgtggtcag cgtcctcacc
gtcctgcacc 360aggactggct gaatggcaag gagtacaagt gcaaggtctc caacaaagcc
ctcccagccc 420ccatcgagaa aaccatctcc aaagccaaag ggcagccccg agaaccacag
gtgtacaccc 480tgcccccatc ccgggatgag ctgaccaaga accaggtcag cctgacctgc
ctggtcaaag 540gcttctatcc cagcgacatc gccgtggagt gggagagcaa tgggcagccg
gagaacaact 600acaagaccac gcctcccgtg ctggactccg acggctcctt cttcctctac
agcaagctca 660ccgtggacaa gagcaggtgg cagcagggga acgtcttctc atgctccgtg
atgcatgagg 720ctctgcacaa ccactacacg cagaagagcc tctccctgtc tccggagctg
caactggagg 780agagctgtgc ggaggcgcag gacggggagc tggacgggtg cgtatccggt
gacaccattg 840taatgactag tggcggtccg cgcactgtgg ctgaactgga gggcaaaccg
ttcaccgcac 900tgattcgcgg ctctggctac ccatgcccct caggtttctt ccgcacctgt
gaacgtgacg 960tatatgatct gcgtacacgt gagggtcatt gcttacgttt gacccatgat
caccgtgttc 1020tggtgatgga tggtggcctg gaatggcgtg ccgcgggtga actggaacgc
ggcgaccgcc 1080tggtgatgga tgatgcagct ggcgagtttc cggcactggc aaccttccgt
ggcctgcgtg 1140gcgctggccg ccaggatgtt tatgacgcta ctgtttacgg tgctagcgca
ttcactgcta 1200atggcttcat tgtacacgca tgtggcgagc agcccgggac cggtctgaac
tcaggcctca 1260cgacaaatcc tggtgtatcc gcttggcagg tcaacacagc ttatactgcg
ggacaattgg 1320tcacatataa cggcaagacg tataaatgtt tgcagcccca cacctccttg
gcaggatggg 1380aaccatccaa cgttcctgcc ttgtggcagc ttcaatgaag atctgttaac
1430121430DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 12gttaacgaat tcccaccatg gccttgacct
ttgctttact ggtggccctc ctggtgctca 60gctgcaagtc aagctgctct gtgggctgtg
acaaaactca cacatgccca ccgtgcccag 120cacctgaact cctgggggga ccgtcagtct
tcctcttccc cccaaaaccc aaggacaccc 180tcatgatctc ccggacccct gaggtcacat
gcgtggtggt ggacgtgagc cacgaagacc 240ctgaggtcaa gttcaactgg tacgtggacg
gcgtggaggt gcataatgcc aagacaaagc 300cgcgggagga gcagtacaac agcacgtacc
gtgtggtcag cgtcctcacc gtcctgcacc 360aggactggct gaatggcaag gagtacaagt
gcaaggtctc caacaaagcc ctcccagccc 420ccatcgagaa aaccatctcc aaagccaaag
ggcagccccg agaaccacag gtgtacaccc 480tgcccccatc ccgggatgag ctgaccaaga
accaggtcag cctgacctgc ctggtcaaag 540gcttctatcc cagcgacatc gccgtggagt
gggagagcaa tgggcagccg gagaacaact 600acaagaccac gcctcccgtg ctggactccg
acggctcctt cttcctctac agcaagctca 660ccgtggacaa gagcaggtgg cagcagggga
acgtcttctc atgctccgtg atgcatgagg 720ctctgcacaa ccactacacg cagaagagcc
tctccctgtc tccggagctg caactggagg 780agagctgtgc ggaggcgcag gacggggagc
tggacgggtg cgtatccggt gacaccattg 840taatgactag tggcggtccg cgcactgtgg
ctgaactgga gggcaaaccg ttcaccgcac 900tgattcgcgg ctctggctac ccatgcccct
caggtttctt ccgcacctgt gaacgtgacg 960tatatgatct gcgtacacgt gagggtcatt
gcttacgttt gacccatgat caccgtgttc 1020tggtgatgga tggtggcctg gaatggcgtg
ccgcgggtga actggaacgc ggcgaccgcc 1080tggtgatgga tgatgcagct ggcgagtttc
cggcactggc aaccttccgt ggcctgcgtg 1140gcgctggccg ccaggatgtt tatgacgcta
ctgtttacgg tgctagcgca ttcactgcta 1200atggcttcat tgtacacgca tgtggcgagc
agcccgggac cggtctgaac tcaggcctca 1260cgacaaatcc tggtgtatcc gcttggcagg
tcaacacagc ttatactgcg ggacaattgg 1320tcacatataa cggcaagacg tataaatgtt
tgcagcccca cacctccttg gcaggatggg 1380aaccatccaa cgttcctgcc ttgtggcagc
ttcaatgaag atctgttaac 1430131412DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 13gttaacgaat tcccaccatg ctggctgcca cagtcctgac cctggccctg
ctgggcaatg 60cccatgcctg tgacaaaact cacacatgcc caccgtgccc agcacctgaa
ctcctggggg 120gaccgtcagt cttcctcttc cccccaaaac ccaaggacac cctcatgatc
tcccggaccc 180ctgaggtcac atgcgtggtg gtggacgtga gccacgaaga ccctgaggtc
aagttcaact 240ggtacgtgga cggcgtggag gtgcataatg ccaagacaaa gccgcgggag
gagcagtaca 300acagcacgta ccgtgtggtc agcgtcctca ccgtcctgca ccaggactgg
ctgaatggca 360aggagtacaa gtgcaaggtc tccaacaaag ccctcccagc ccccatcgag
aaaaccatct 420ccaaagccaa agggcagccc cgagaaccac aggtgtacac cctgccccca
tcccgggatg 480agctgaccaa gaaccaggtc agcctgacct gcctggtcaa aggcttctat
cccagcgaca 540tcgccgtgga gtgggagagc aatgggcagc cggagaacaa ctacaagacc
acgcctcccg 600tgctggactc cgacggctcc ttcttcctct acagcaagct caccgtggac
aagagcaggt 660ggcagcaggg gaacgtcttc tcatgctccg tgatgcatga ggctctgcac
aaccactaca 720cgcagaagag cctctccctg tctccggagc tgcaactgga ggagagctgt
gcggaggcgc 780aggacgggga gctggacggg tgcgtatccg gtgacaccat tgtaatgact
agtggcggtc 840cgcgcactgt ggctgaactg gagggcaaac cgttcaccgc actgattcgc
ggctctggct 900acccatgccc ctcaggtttc ttccgcacct gtgaacgtga cgtatatgat
ctgcgtacac 960gtgagggtca ttgcttacgt ttgacccatg atcaccgtgt tctggtgatg
gatggtggcc 1020tggaatggcg tgccgcgggt gaactggaac gcggcgaccg cctggtgatg
gatgatgcag 1080ctggcgagtt tccggcactg gcaaccttcc gtggcctgcg tggcgctggc
cgccaggatg 1140tttatgacgc tactgtttac ggtgctagcg cattcactgc taatggcttc
attgtacacg 1200catgtggcga gcagcccggg accggtctga actcaggcct cacgacaaat
cctggtgtat 1260ccgcttggca ggtcaacaca gcttatactg cgggacaatt ggtcacatat
aacggcaaga 1320cgtataaatg tttgcagccc cacacctcct tggcaggatg ggaaccatcc
aacgttcctg 1380ccttgtggca gcttcaatga agatctgtta ac
1412141412DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 14gttaacgaat tcccaccatg ctgctgctgg
cgagatgtct gctgctagtc ctcgtctcct 60cgctgctggt atgctcggga ctggcgtgcc
caccgtgccc agcacctgaa ctcctggggg 120gaccgtcagt cttcctcttc cccccaaaac
ccaaggacac cctcatgatc tcccggaccc 180ctgaggtcac atgcgtggtg gtggacgtga
gccacgaaga ccctgaggtc aagttcaact 240ggtacgtgga cggcgtggag gtgcataatg
ccaagacaaa gccgcgggag gagcagtaca 300acagcacgta ccgtgtggtc agcgtcctca
ccgtcctgca ccaggactgg ctgaatggca 360aggagtacaa gtgcaaggtc tccaacaaag
ccctcccagc ccccatcgag aaaaccatct 420ccaaagccaa agggcagccc cgagaaccac
aggtgtacac cctgccccca tcccgggatg 480agctgaccaa gaaccaggtc agcctgacct
gcctggtcaa aggcttctat cccagcgaca 540tcgccgtgga gtgggagagc aatgggcagc
cggagaacaa ctacaagacc acgcctcccg 600tgctggactc cgacggctcc ttcttcctct
acagcaagct caccgtggac aagagcaggt 660ggcagcaggg gaacgtcttc tcatgctccg
tgatgcatga ggctctgcac aaccactaca 720cgcagaagag cctctccctg tctccggagc
tgcaactgga ggagagctgt gcggaggcgc 780aggacgggga gctggacggg tgcgtatccg
gtgacaccat tgtaatgact agtggcggtc 840cgcgcactgt ggctgaactg gagggcaaac
cgttcaccgc actgattcgc ggctctggct 900acccatgccc ctcaggtttc ttccgcacct
gtgaacgtga cgtatatgat ctgcgtacac 960gtgagggtca ttgcttacgt ttgacccatg
atcaccgtgt tctggtgatg gatggtggcc 1020tggaatggcg tgccgcgggt gaactggaac
gcggcgaccg cctggtgatg gatgatgcag 1080ctggcgagtt tccggcactg gcaaccttcc
gtggcctgcg tggcgctggc cgccaggatg 1140tttatgacgc tactgtttac ggtgctagcg
cattcactgc taatggcttc attgtacacg 1200catgtggcga gcagcccggg accggtctga
actcaggcct cacgacaaat cctggtgtat 1260ccgcttggca ggtcaacaca gcttatactg
cgggacaatt ggtcacatat aacggcaaga 1320cgtataaatg tttgcagccc cacacctcct
tggcaggatg ggaaccatcc aacgttcctg 1380ccttgtggca gcttcaatga agatctgtta
ac 1412151412DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 15gttaacgaat tcccaccatg gccttgacct ttgctttact ggtggccctc
ctggtgctca 60gctgcaagtc aagctgctct gtgggctgcc caccgtgccc agcacctgaa
ctcctggggg 120gaccgtcagt cttcctcttc cccccaaaac ccaaggacac cctcatgatc
tcccggaccc 180ctgaggtcac atgcgtggtg gtggacgtga gccacgaaga ccctgaggtc
aagttcaact 240ggtacgtgga cggcgtggag gtgcataatg ccaagacaaa gccgcgggag
gagcagtaca 300acagcacgta ccgtgtggtc agcgtcctca ccgtcctgca ccaggactgg
ctgaatggca 360aggagtacaa gtgcaaggtc tccaacaaag ccctcccagc ccccatcgag
aaaaccatct 420ccaaagccaa agggcagccc cgagaaccac aggtgtacac cctgccccca
tcccgggatg 480agctgaccaa gaaccaggtc agcctgacct gcctggtcaa aggcttctat
cccagcgaca 540tcgccgtgga gtgggagagc aatgggcagc cggagaacaa ctacaagacc
acgcctcccg 600tgctggactc cgacggctcc ttcttcctct acagcaagct caccgtggac
aagagcaggt 660ggcagcaggg gaacgtcttc tcatgctccg tgatgcatga ggctctgcac
aaccactaca 720cgcagaagag cctctccctg tctccggagc tgcaactgga ggagagctgt
gcggaggcgc 780aggacgggga gctggacggg tgcgtatccg gtgacaccat tgtaatgact
agtggcggtc 840cgcgcactgt ggctgaactg gagggcaaac cgttcaccgc actgattcgc
ggctctggct 900acccatgccc ctcaggtttc ttccgcacct gtgaacgtga cgtatatgat
ctgcgtacac 960gtgagggtca ttgcttacgt ttgacccatg atcaccgtgt tctggtgatg
gatggtggcc 1020tggaatggcg tgccgcgggt gaactggaac gcggcgaccg cctggtgatg
gatgatgcag 1080ctggcgagtt tccggcactg gcaaccttcc gtggcctgcg tggcgctggc
cgccaggatg 1140tttatgacgc tactgtttac ggtgctagcg cattcactgc taatggcttc
attgtacacg 1200catgtggcga gcagcccggg accggtctga actcaggcct cacgacaaat
cctggtgtat 1260ccgcttggca ggtcaacaca gcttatactg cgggacaatt ggtcacatat
aacggcaaga 1320cgtataaatg tttgcagccc cacacctcct tggcaggatg ggaaccatcc
aacgttcctg 1380ccttgtggca gcttcaatga agatctgtta ac
1412161394DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 16gttaacgaat tcccaccatg ctggctgcca
cagtcctgac cctggccctg ctgggcaatg 60cccatgcctg cccaccgtgc ccagcacctg
aactcctggg gggaccgtca gtcttcctct 120tccccccaaa acccaaggac accctcatga
tctcccggac ccctgaggtc acatgcgtgg 180tggtggacgt gagccacgaa gaccctgagg
tcaagttcaa ctggtacgtg gacggcgtgg 240aggtgcataa tgccaagaca aagccgcggg
aggagcagta caacagcacg taccgtgtgg 300tcagcgtcct caccgtcctg caccaggact
ggctgaatgg caaggagtac aagtgcaagg 360tctccaacaa agccctccca gcccccatcg
agaaaaccat ctccaaagcc aaagggcagc 420cccgagaacc acaggtgtac accctgcccc
catcccggga tgagctgacc aagaaccagg 480tcagcctgac ctgcctggtc aaaggcttct
atcccagcga catcgccgtg gagtgggaga 540gcaatgggca gccggagaac aactacaaga
ccacgcctcc cgtgctggac tccgacggct 600ccttcttcct ctacagcaag ctcaccgtgg
acaagagcag gtggcagcag gggaacgtct 660tctcatgctc cgtgatgcat gaggctctgc
acaaccacta cacgcagaag agcctctccc 720tgtctccgga gctgcaactg gaggagagct
gtgcggaggc gcaggacggg gagctggacg 780ggtgcgtatc cggtgacacc attgtaatga
ctagtggcgg tccgcgcact gtggctgaac 840tggagggcaa accgttcacc gcactgattc
gcggctctgg ctacccatgc ccctcaggtt 900tcttccgcac ctgtgaacgt gacgtatatg
atctgcgtac acgtgagggt cattgcttac 960gtttgaccca tgatcaccgt gttctggtga
tggatggtgg cctggaatgg cgtgccgcgg 1020gtgaactgga acgcggcgac cgcctggtga
tggatgatgc agctggcgag tttccggcac 1080tggcaacctt ccgtggcctg cgtggcgctg
gccgccagga tgtttatgac gctactgttt 1140acggtgctag cgcattcact gctaatggct
tcattgtaca cgcatgtggc gagcagcccg 1200ggaccggtct gaactcaggc ctcacgacaa
atcctggtgt atccgcttgg caggtcaaca 1260cagcttatac tgcgggacaa ttggtcacat
ataacggcaa gacgtataaa tgtttgcagc 1320cccacacctc cttggcagga tgggaaccat
ccaacgttcc tgccttgtgg cagcttcaat 1380gaagatctgt taac
1394171820DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 17gttaacgaat tcccaccatg aaccggggag tcccttttag gcacttgctt
ctggtgctgc 60aactggcgct cctcccagca gccactcagg gaaagaaagt ggtgctgggc
aaaaaagggg 120atacagtgga actgacctgt acagcttccc agaagaagag catacaattc
cactggaaaa 180actccaacca gataaagatt ctgggaaatc agggctcctt cttaactaaa
ggtccatcca 240agctgaatga tcgcgctgac tcaagaagaa gcctttggga ccaaggaaac
ttccccctga 300tcatcaagaa tcttaagata gaagactcag atacttacat ctgtgaagtg
gaggaccaga 360aggaggaggt gcaattgcta gtgttcggat tgactgccaa ctctgacacc
cacctgcttc 420aggggcagag cctgaccctg accttggaga gcccccctgg tagtagcccc
tcagtgcaat 480gtaggagtcc aaggggtaaa aacatacagg gggggaagac cctctccgtg
tctcagctgg 540agctccagga tagtggcacc tggacatgca ctgtcttgca gaaccagaag
aaggtggagt 600tcaaaataga catcgtggtg ctagctttcc agaaggcctc cagcatagtc
tataagaaag 660agggggaaca ggtggagttc tccttcccac tcgcctttac agttgaaaag
ctgacgggca 720gtggcgagct gtggtggcag gcggagaggg cttcctcctc caagtcttgg
atcacctttg 780acctgaagaa caaggaagtg tctgtaaaac gggttaccca ggaccctaag
ctccagatgg 840gcaagaagct cccgctccac ctcaccctgc cccaggcctt gcctcagtat
gctggctctg 900gaaacctcac cctggccctt gaagcgaaaa caggaaagtt gcatcaggaa
gtgaacctgg 960tggtgatgag agccactcag ctccagaaaa atttgacctg tgaggtgtgg
ggacccacct 1020cccctaagct gatgctgagc ttgaaactgg agaacaagga ggcaaaggtc
tcgaagcggg 1080agaaggcggt gtgggtgctg aaccctgagg cggggatgtg gcagtgtctg
ctgagtgact 1140cgggacaggt cctgctggaa tccaacatca aggttctgcc cacatggtcc
accccggtgc 1200agccagggtg cgtatccggt gacaccattg taatgactag tggcggtccg
cgcactgtgg 1260ctgaactgga gggcaaaccg ttcaccgcac tgattcgcgg ctctggctac
ccatgcccct 1320caggtttctt ccgcacctgt gaacgtgacg tatatgatct gcgtacacgt
gagggtcatt 1380gcttacgttt gacccatgat caccgtgttc tggtgatgga tggtggcctg
gaatggcgtg 1440ccgcgggtga actggaacgc ggcgaccgcc tggtgatgga tgatgcagct
ggcgagtttc 1500cggcactggc aaccttccgt ggcctgcgtg gcgctggccg ccaggatgtt
tatgacgcta 1560ctgtttacgg tgctagcgca ttcactgcta atggcttcat tgtacacgca
tgtggcgagc 1620agcccgggac cggtctgaac tcaggcctca cgacaaatcc tggtgtatcc
gcttggcagg 1680tcaacacagc ttatactgcg ggacaattgg tcacatataa cggcaagacg
tataaatgtt 1740tgcagcccca cacctccttg gcaggatggg aaccatccaa cgttcctgcc
ttgtggcagc 1800ttcaatgaag atctgttaac
1820181364DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 18gttaacgaat tcccaccatg gcttgggtgt
ggaccttgct tttcctgatg gcagctgccc 60aaagtatcca agcacagatc cagttggtcc
agtctggacc tgagctgaag aagcctggag 120agacagtcaa gatctcctgc aaggcttcag
gatatacctt cacacactat ggaatgaact 180gggtgaagca ggctccagga aagggtttaa
agtggatggg ctggataaac acctacactg 240gagagccaac atatgctgat gacttcaagg
aacactttgc cttctctttg gaaacctctg 300ccagcactgt ctttttgcag atcaacaacc
tcaaaaatga ggacacggcc acatatttct 360gtgcaagaga acggggggat gctatggact
actggggtca gggaacctcc gtcaccgtct 420cctcagcctc caccaagggc ccatcggtct
tccccctggc accctcctcc aagagcacct 480ctgggggcac agcggccctg ggctgcctgg
tcaaggacta cttccccgaa ccggtgacgg 540tgtcgtggaa ctcaggcgcc ctgaccagcg
gcgtgcacac cttcccggct gtcctacagt 600cctcaggact ctactccctc agcagcgtgg
tgaccgtgcc ctccagcagc ttgggcaccc 660agacctacat ctgcaacgtg aatcacaagc
ccagcaacac caaggtggac aagaaagttg 720agcccaaatc ttgtgacaaa actcacacag
ggtgcgtatc cggtgacacc attgtaatga 780ctagtggcgg tccgcgcact gtggctgaac
tggagggcaa accgttcacc gcactgattc 840gcggctctgg ctacccatgc ccctcaggtt
tcttccgcac ctgtgaacgt gacgtatatg 900atctgcgtac acgtgagggt cattgcttac
gtttgaccca tgatcaccgt gttctggtga 960tggatggtgg cctggaatgg cgtgccgcgg
gtgaactgga acgcggcgac cgcctggtga 1020tggatgatgc agctggcgag tttccggcac
tggcaacctt ccgtggcctg cgtggcgctg 1080gccgccagga tgtttatgac gctactgttt
acggtgctag cgcattcact gctaatggct 1140tcattgtaca cgcatgtggc gagcagcccg
ggaccggtct gaactcaggc ctcacgacaa 1200atcctggtgt atccgcttgg caggtcaaca
cagcttatac tgcgggacaa ttggtcacat 1260ataacggcaa gacgtataaa tgtttgcagc
cccacacctc cttggcagga tgggaaccat 1320ccaacgttcc tgccttgtgg cagcttcaat
gaagatctgt taac 136419737DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 19gttaacgaat tcccaccatg aagtcacaga cccaggtctt cgtatttcta
ctgctctgtg 60tgtctggtgc tcatgggagt attgtgatga cccagactcc caaattcctg
cttgtatcag 120caggagacag ggttaccata acctgcacgg ccagtcagag tgtgagtaat
gatgtagttt 180ggtaccaaca gaagccaggg cagtctccta aaatgctgat gtattctgca
ttcaatcgct 240acactggagt ccctgatcgt ttcactggca gaggatacgg gacggatttc
actttcacca 300tcagctctgt gcaggctgaa gacctggcag tttatttctg tcagcaggat
tataactctc 360ctcggacgtt cggtggaggc accaagctgg agatcaaacg aactgtggct
gcaccatctg 420tcttcatctt cccgccatct gatgagcagt tgaaatctgg aactgcctct
gttgtgtgcc 480tgctgaataa cttctatccc agagaggcca aagtacagtg gaaggtggat
aacgccctcc 540aatcgggtaa ctcccaggag agtgtcacag agcaggacag caaggacagc
acctacagcc 600tcagcagcac cctgacgctg agcaaagcag actacgagaa acacaaagtc
tacgcctgcg 660aagtcaccca tcagggcctg agctcgcccg tcacaaagag cttcaacagg
ggagagtgtt 720agtgaagatc tgttaac
737201268DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 20gttaacgaat tcccaccatg ggcctctcca
ccgtgcctga cctgctgctg ccgctggtgc 60tcctggagct gttggtggga atatacccct
caggggttat tggactggtc cctcacctag 120gggacaggga gaagagagat agtgtgtgtc
cccaaggaaa atatatccac cctcaaaata 180attcgatttg ctgtaccaag tgccacaaag
gaacctactt gtacaatgac tgtccaggcc 240cggggcagga tacggactgc agggagtgtg
agagcggctc cttcaccgct tcagaaaacc 300acctcagaca ctgcctcagc tgctccaaat
gccgaaagga aatgggtcag gtggagatct 360cttcttgcac agtggaccgg gacaccgtgt
gtggctgcag gaagaaccag taccggcatt 420attggagtga aaaccttttc cagtgcttca
attgcagcct ctgcctcaat gggaccgtgc 480acctctcctg ccaggagaaa cagaacaccg
tgtgcacctg ccatgcaggt ttctttctaa 540gagaaaacga gtgtgtctcc tgtagtaact
gtaagaaaag cctggagtgc acgaagttgt 600gcctacccca gattgagaat gttaagggca
ctgaggactc aggcaccaca gtggggtgcg 660tatccggtga caccattgta atgactagtg
gcggtccgcg cactgtggct gaactggagg 720gcaaaccgtt caccgcactg attcgcggct
ctggctaccc atgcccctca ggtttcttcc 780gcacctgtga acgtgacgta tatgatctgc
gtacacgtga gggtcattgc ttacgtttga 840cccatgatca ccgtgttctg gtgatggatg
gtggcctgga atggcgtgcc gcgggtgaac 900tggaacgcgg cgaccgcctg gtgatggatg
atgcagctgg cgagtttccg gcactggcaa 960ccttccgtgg cctgcgtggc gctggccgcc
aggatgttta tgacgctact gtttacggtg 1020ctagcgcatt cactgctaat ggcttcattg
tacacgcatg tggcgagcag cccgggaccg 1080gtctgaactc aggcctcacg acaaatcctg
gtgtatccgc ttggcaggtc aacacagctt 1140atactgcggg acaattggtc acatataacg
gcaagacgta taaatgtttg cagccccaca 1200cctccttggc aggatgggaa ccatccaacg
ttcctgcctt gtggcagctt caatgaagat 1260ctgttaac
1268211403DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 21gttaacgaat tcccaccatg gcgcccgtcg ccgtctgggc cgcgctggcc
gtcggactgg 60agctctgggc tgcggcgcac gccttgcccg cccaggtggc atttacaccc
tacgccccgg 120agcccgggag cacatgccgg ctcagagaat actatgacca gacagctcag
atgtgctgca 180gcaaatgctc gccgggccaa catgcaaaag tcttctgtac caagacctcg
gacaccgtgt 240gtgactcctg tgaggacagc acatacaccc agctctggaa ctgggttccc
gagtgcttga 300gctgtggctc ccgctgtagc tctgaccagg tggaaactca agcctgcact
cgggaacaga 360accgcatctg cacctgcagg cccggctggt actgcgcgct gagcaagcag
gaggggtgcc 420ggctgtgcgc gccgctgcgc aagtgccgcc cgggcttcgg cgtggccaga
ccaggaactg 480aaacatcaga cgtggtgtgc aagccctgtg ccccggggac gttctccaac
acgacttcat 540ccacggatat ttgcaggccc caccagatct gtaacgtggt ggccatccct
gggaatgcaa 600gcatggatgc agtctgcacg tccacgtccc ccacccggag tatggcccca
ggggcagtac 660acttacccca gccagtgtcc acacgatccc aacacacgca gccaactcca
gaacccagca 720ctgctccaag cacctccttc ctgctcccaa tgggccccag ccccccagct
gaagggagca 780ctggcgacgg gtgcgtatcc ggtgacacca ttgtaatgac tagtggcggt
ccgcgcactg 840tggctgaact ggagggcaaa ccgttcaccg cactgattcg cggctctggc
tacccatgcc 900cctcaggttt cttccgcacc tgtgaacgtg acgtatatga tctgcgtaca
cgtgagggtc 960attgcttacg tttgacccat gatcaccgtg ttctggtgat ggatggtggc
ctggaatggc 1020gtgccgcggg tgaactggaa cgcggcgacc gcctggtgat ggatgatgca
gctggcgagt 1080ttccggcact ggcaaccttc cgtggcctgc gtggcgctgg ccgccaggat
gtttatgacg 1140ctactgttta cggtgctagc gcattcactg ctaatggctt cattgtacac
gcatgtggcg 1200agcagcccgg gaccggtctg aactcaggcc tcacgacaaa tcctggtgta
tccgcttggc 1260aggtcaacac agcttatact gcgggacaat tggtcacata taacggcaag
acgtataaat 1320gtttgcagcc ccacacctcc ttggcaggat gggaaccatc caacgttcct
gccttgtggc 1380agcttcaatg aagatctgtt aac
1403222906DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 22gttaacgaat tcccaccatg gtcagctact
gggacaccgg ggtcctgctg tgcgcgctgc 60tcagctgtct gcttctcaca ggatctagtt
caggttcaaa attaaaagat cctgaactga 120gtttaaaagg cacccagcac atcatgcaag
caggccagac actgcatctc caatgcaggg 180gggaagcagc ccataaatgg tctttgcctg
aaatggtgag taaggaaagc gaaaggctga 240gcataactaa atctgcctgt ggaagaaatg
gcaaacaatt ctgcagtact ttaaccttga 300acacagctca agcaaaccac actggcttct
acagctgcaa atatctagct gtacctactt 360caaagaagaa ggaaacagaa tctgcaatct
atatatttat tagtgataca ggtagacctt 420tcgtagagat gtacagtgaa atccccgaaa
ttatacacat gactgaagga agggagctcg 480tcattccctg ccgggttacg tcacctaaca
tcactgttac tttaaaaaag tttccacttg 540acactttgat ccctgatgga aaacgcataa
tctgggacag tagaaagggc ttcatcatat 600caaatgcaac gtacaaagaa atagggcttc
tgacctgtga agcaacagtc aatgggcatt 660tgtataagac aaactatctc acacatcgac
aaaccaatac aatcatagat gtccaaataa 720gcacaccacg cccagtcaaa ttacttagag
gccatactct tgtcctcaat tgtactgcta 780ccactccctt gaacacgaga gttcaaatga
cctggagtta ccctgatgaa aaaaataaga 840gagcttccgt aaggcgacga attgaccaaa
gcaattccca tgccaacata ttctacagtg 900ttcttactat tgacaaaatg cagaacaaag
acaaaggact ttatacttgt cgtgtaagga 960gtggaccatc attcaaatct gttaacacct
cagtgcatat atatgataaa gcattcatca 1020ctgtgaaaca tcgaaaacag caggtgcttg
aaaccgtagc tggcaagcgg tcttaccggc 1080tctctatgaa agtgaaggca tttccctcgc
cggaagttgt atggttaaaa gatgggttac 1140ctgcgactga gaaatctgct cgctatttga
ctcgtggcta ctcgttaatt atcaaggacg 1200taactgaaga ggatgcaggg aattatacaa
tcttgctgag cataaaacag tcaaatgtgt 1260ttaaaaacct cactgccact ctaattgtca
atgtgaaacc ccagatttac gaaaaggccg 1320tgtcatcgtt tccagacccg gctctctacc
cactgggcag cagacaaatc ctgacttgta 1380ccgcatatgg tatccctcaa cctacaatca
agtggttctg gcacccctgt aaccataatc 1440attccgaagc aaggtgtgac ttttgttcca
ataatgaaga gtcctttatc ctggatgctg 1500acagcaacat gggaaacaga attgagagca
tcactcagcg catggcaata atagaaggaa 1560agaataagat ggctagcacc ttggttgtgg
ctgactctag aatttctgga atctacattt 1620gcatagcttc caataaagtt gggactgtgg
gaagaaacat aagcttttat atcacagatg 1680tgccaaatgg gtttcatgtt aacttggaaa
aaatgccgac ggaaggagag gacctgaaac 1740tgtcttgcac agttaacaag ttcttataca
gagacgttac ttggatttta ctgcggacag 1800ttaataacag aacaatgcac tacagtatta
gcaagcaaaa aatggccatc actaaggagc 1860actccatcac tcttaatctt accatcatga
atgtttccct gcaagattca ggcacctatg 1920cctgcagagc caggaatgta tacacagggg
aagaaatcct ccagaagaaa gaaattacaa 1980tcagagatca ggaagcacca tacctcctgc
gaaacctcag tgatcacaca gtggccatca 2040gcagttccac cactttagac tgtcatgcta
atggtgtccc cgagcctcag atcacttggt 2100ttaaaaacaa ccacaaaata caacaagagc
ctggaattat tttaggacca ggaagcagca 2160cgctgtttat tgaaagagtc acagaagagg
atgaaggtgt ctatcactgc aaagccacca 2220accagaaggg ctctgtggaa agttcagcat
acctcactgt tcaaggaacc tcggacaagt 2280ctaatctgga ggggtgcgta tccggtgaca
ccattgtaat gactagtggc ggtccgcgca 2340ctgtggctga actggagggc aaaccgttca
ccgcactgat tcgcggctct ggctacccat 2400gcccctcagg tttcttccgc acctgtgaac
gtgacgtata tgatctgcgt acacgtgagg 2460gtcattgctt acgtttgacc catgatcacc
gtgttctggt gatggatggt ggcctggaat 2520ggcgtgccgc gggtgaactg gaacgcggcg
accgcctggt gatggatgat gcagctggcg 2580agtttccggc actggcaacc ttccgtggcc
tgcgtggcgc tggccgccag gatgtttatg 2640acgctactgt ttacggtgct agcgcattca
ctgctaatgg cttcattgta cacgcatgtg 2700gcgagcagcc cgggaccggt ctgaactcag
gcctcacgac aaatcctggt gtatccgctt 2760ggcaggtcaa cacagcttat actgcgggac
aattggtcac atataacggc aagacgtata 2820aatgtttgca gccccacacc tccttggcag
gatgggaacc atccaacgtt cctgccttgt 2880ggcagcttca atgaagatct gttaac
2906232924DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 23gttaacgaat tcccaccatg cagagcaagg tgctgctggc cgtcgccctg
tggctctgcg 60tggagacccg ggccgcctct gtgggtttgc ctagtgtttc tcttgatctg
cccaggctca 120gcatacaaaa agacatactt acaattaagg ctaatacaac tcttcaaatt
acttgcaggg 180gacagaggga cttggactgg ctttggccca ataatcagag tggcagtgag
caaagggtgg 240aggtgactga gtgcagcgat ggcctcttct gtaagacact cacaattcca
aaagtgatcg 300gaaatgacac tggagcctac aagtgcttct accgggaaac tgacttggcc
tcggtcattt 360atgtctatgt tcaagattac agatctccat ttattgcttc tgttagtgac
caacatggag 420tcgtgtacat tactgagaac aaaaacaaaa ctgtggtgat tccatgtctc
gggtccattt 480caaatctcaa cgtgtcactt tgtgcaagat acccagaaaa gagatttgtt
cctgatggta 540acagaatttc ctgggacagc aagaagggct ttactattcc cagctacatg
atcagctatg 600ctggcatggt cttctgtgaa gcaaaaatta atgatgaaag ttaccagtct
attatgtaca 660tagttgtcgt tgtagggtat aggatttatg atgtggttct gagtccgtct
catggaattg 720aactatctgt tggagaaaag cttgtcttaa attgtacagc aagaactgaa
ctaaatgtgg 780ggattgactt caactgggaa tacccttctt cgaagcatca gcataagaaa
cttgtaaacc 840gagacctaaa aacccagtct gggagtgaga tgaagaaatt tttgagcacc
ttaactatag 900atggtgtaac ccggagtgac caaggattgt acacctgtgc agcatccagt
gggctgatga 960ccaagaagaa cagcacattt gtcagggtcc atgaaaaacc ttttgttgct
tttggaagtg 1020gcatggaatc tctggtggaa gccacggtgg gggagcgtgt cagaatccct
gcgaagtacc 1080ttggttaccc acccccagaa ataaaatggt ataaaaatgg aatacccctt
gagtccaatc 1140acacaattaa agcggggcat gtactgacga ttatggaagt gagtgaaaga
gacacaggaa 1200attacactgt catccttacc aatcccattt caaaggagaa gcagagccat
gtggtctctc 1260tggttgtgta tgtcccaccc cagattggtg agaaatctct aatctctcct
gtggattcct 1320accagtacgg caccactcaa acgctgacat gtacggtcta tgccattcct
cccccgcatc 1380acatccactg gtattggcag ttggaggaag agtgcgccaa cgagcccagc
caagctgtct 1440cagtgacaaa cccataccct tgtgaagaat ggagaagtgt ggaggacttc
cagggaggaa 1500ataaaattga agttaataaa aatcaatttg ctctaattga aggaaaaaac
aaaactgtaa 1560gtacccttgt tatccaagcg gcaaatgtgt cagctttgta caaatgtgaa
gcggtcaaca 1620aagtcgggag aggagagagg gtgatctcct tccacgtgac caggggtcct
gaaattactt 1680tgcaacctga catgcagccc actgagcagg agagcgtgtc tttgtggtgc
actgcagaca 1740gatctacgtt tgagaacctc acatggtaca agcttggccc acagcctctg
ccaatccatg 1800tgggagagtt gcccacacct gtttgcaaga acttggatac tctttggaaa
ttgaatgcca 1860ccatgttctc taatagcaca aatgacattt tgatcatgga gcttaagaat
gcatccttgc 1920aggaccaagg agactatgtc tgccttgctc aagacaggaa gaccaagaaa
agacattgcg 1980tggtcaggca gctcacagtc ctagagcgtg tggcacccac gatcacagga
aacctggaga 2040atcagacgac aagtattggg gaaagcatcg aagtctcatg cacggcatct
gggaatcccc 2100ctccacagat catgtggttt aaagataatg agacccttgt agaagactca
ggcattgtat 2160tgaaggatgg gaaccggaac ctcactatcc gcagagtgag gaaggaggac
gaaggcctct 2220acacctgcca ggcatgcagt gttcttggct gtgcaaaagt ggaggcattt
ttcataatag 2280aaggtgccca ggaaaagacg aacttggaag ggtgcgtatc cggtgacacc
attgtaatga 2340ctagtggcgg tccgcgcact gtggctgaac tggagggcaa accgttcacc
gcactgattc 2400gcggctctgg ctacccatgc ccctcaggtt tcttccgcac ctgtgaacgt
gacgtatatg 2460atctgcgtac acgtgagggt cattgcttac gtttgaccca tgatcaccgt
gttctggtga 2520tggatggtgg cctggaatgg cgtgccgcgg gtgaactgga acgcggcgac
cgcctggtga 2580tggatgatgc agctggcgag tttccggcac tggcaacctt ccgtggcctg
cgtggcgctg 2640gccgccagga tgtttatgac gctactgttt acggtgctag cgcattcact
gctaatggct 2700tcattgtaca cgcatgtggc gagcagcccg ggaccggtct gaactcaggc
ctcacgacaa 2760atcctggtgt atccgcttgg caggtcaaca cagcttatac tgcgggacaa
ttggtcacat 2820ataacggcaa gacgtataaa tgtttgcagc cccacacctc cttggcagga
tgggaaccat 2880ccaacgttcc tgccttgtgg cagcttcaat gaagatctgt taac
2924242957DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 24gttaacgaat tcccaccatg cagcggggcg
ccgcgctgtg cctgcgactg tggctctgcc 60tgggactcct ggacggcctg gtgagtggct
actccatgac ccccccgacc ttgaacatca 120cggaggagtc acacgtcatc gacaccggtg
acagcctgtc catctcctgc aggggacagc 180accccctcga gtgggcttgg ccaggagctc
aggaggcgcc agccaccgga gacaaggaca 240gcgaggacac gggggtggtg cgagactgcg
agggcacaga cgccaggccc tactgcaagg 300tgttgctgct gcacgaggta catgccaacg
acacaggcag ctacgtctgc tactacaagt 360acatcaaggc acgcatcgag ggcaccacgg
ccgccagctc ctacgtgttc gtgagagact 420ttgagcagcc attcatcaac aagcctgaca
cgctcttggt caacaggaag gacgccatgt 480gggtgccctg tctggtgtcc atccccggcc
tcaatgtcac gctgcgctcg caaagctcgg 540tgctgtggcc agacgggcag gaggtggtgt
gggatgaccg gcggggcatg ctcgtgtcca 600cgccactgct gcacgatgcc ctgtacctgc
agtgcgagac cacctgggga gaccaggact 660tcctttccaa ccccttcctg gtgcacatca
caggcaacga gctctatgac atccagctgt 720tgcccaggaa gtcgctggag ctgctggtag
gggagaagct ggtcctgaac tgcaccgtgt 780gggctgagtt taactcaggt gtcacctttg
actgggacta cccagggaag caggcagagc 840ggggtaagtg ggtgcccgag cgacgctccc
agcagaccca cacagaactc tccagcatcc 900tgaccatcca caacgtcagc cagcacgacc
tgggctcgta tgtgtgcaag gccaacaacg 960gcatccagcg atttcgggag agcaccgagg
tcattgtgca tgaaaatccc ttcatcagcg 1020tcgagtggct caaaggaccc atcctggagg
ccacggcagg agacgagctg gtgaagctgc 1080ccgtgaagct ggcagcgtac cccccgcccg
agttccagtg gtacaaggat ggaaaggcac 1140tgtccgggcg ccacagtcca catgccctgg
tgctcaagga ggtgacagag gccagcacag 1200gcacctacac cctcgccctg tggaactccg
ctgctggcct gaggcgcaac atcagcctgg 1260agctggtggt gaatgtgccc ccccagatac
atgagaagga ggcctcctcc cccagcatct 1320actcgcgtca cagccgccag gccctcacct
gcacggccta cggggtgccc ctgcctctca 1380gcatccagtg gcactggcgg ccctggacac
cctgcaagat gtttgcccag cgtagtctcc 1440ggcggcggca gcagcaagac ctcatgccac
agtgccgtga ctggagggcg gtgaccacgc 1500aggatgccgt gaaccccatc gagagcctgg
acacctggac cgagtttgtg gagggaaaga 1560ataagactgt gagcaagctg gtgatccaga
atgccaacgt gtctgccatg tacaagtgtg 1620tggtctccaa caaggtgggc caggatgagc
ggctcatcta cttctatgtg accaccatcc 1680ccgacggctt caccatcgaa tccaagccat
ccgaggagct actagagggc cagccggtgc 1740tcctgagctg ccaagccgac agctacaagt
acgagcatct gcgctggtac cgcctcaacc 1800tgtccacgct gcacgatgcg cacgggaacc
cgcttctgct cgactgcaag aacgtgcatc 1860tgttcgccac ccctctggcc gccagcctgg
aggaggtggc acctggggcg cgccacgcca 1920cgctcagcct gagtatcccc cgcgtcgcgc
ccgagcacga gggccactat gtgtgcgaag 1980tgcaagaccg gcgcagccat gacaagcact
gccacaagaa gtacctgtcg gtgcaggccc 2040tggaagcccc tcggctcacg cagaacttga
ccgacctcct ggtgaacgtg agcgactcgc 2100tggagatgca gtgcttggtg gccggagcgc
acgcgcccag catcgtgtgg tacaaagacg 2160agaggctgct ggaggaaaag tctggagtcg
acttggcgga ctccaaccag aagctgagca 2220tccagcgcgt gcgcgaggag gatgcgggac
gctatctgtg cagcgtgtgc aacgccaagg 2280gctgcgtcaa ctcctccgcc agcgtggccg
tggaaggctc cgaggataag ggcagcatgg 2340aggggtgcgt atccggtgac accattgtaa
tgactagtgg cggtccgcgc actgtggctg 2400aactggaggg caaaccgttc accgcactga
ttcgcggctc tggctaccca tgcccctcag 2460gtttcttccg cacctgtgaa cgtgacgtat
atgatctgcg tacacgtgag ggtcattgct 2520tacgtttgac ccatgatcac cgtgttctgg
tgatggatgg tggcctggaa tggcgtgccg 2580cgggtgaact ggaacgcggc gaccgcctgg
tgatggatga tgcagctggc gagtttccgg 2640cactggcaac cttccgtggc ctgcgtggcg
ctggccgcca ggatgtttat gacgctactg 2700tttacggtgc tagcgcattc actgctaatg
gcttcattgt acacgcatgt ggcgagcagc 2760ccgggaccgg tctgaactca ggcctcacga
caaatcctgg tgtatccgct tggcaggtca 2820acacagctta tactgcggga caattggtca
catataacgg caagacgtat aaatgtttgc 2880agccccacac ctccttggca ggatgggaac
catccaacgt tcctgccttg tggcagcttc 2940aatgaagatc tgttaac
2957252567DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 25gttaacgaat tcccaccatg cgaccctccg ggacggccgg ggcagcgctc
ctggcgctgc 60tggctgcgct ctgcccggcg agtcgggctc tggaggaaaa gaaagtttgc
caaggcacga 120gtaacaagct cacgcagttg ggcacttttg aagatcattt tctcagcctc
cagaggatgt 180tcaataactg tgaggtggtc cttgggaatt tggaaattac ctatgtgcag
aggaattatg 240atctttcctt cttaaagacc atccaggagg tggctggtta tgtcctcatt
gccctcaaca 300cagtggagcg aattcctttg gaaaacctgc agatcatcag aggaaatatg
tactacgaaa 360attcctatgc cttagcagtc ttatctaact atgatgcaaa taaaaccgga
ctgaaggagc 420tgcccatgag aaatttacag gaaatcctgc atggcgccgt gcggttcagc
aacaaccctg 480ccctgtgcaa cgtggagagc atccagtggc gggacatagt cagcagtgac
tttctcagca 540acatgtcgat ggacttccag aaccacctgg gcagctgcca aaagtgtgat
ccaagctgtc 600ccaatgggag ctgctggggt gcaggagagg agaactgcca gaaactgacc
aaaatcatct 660gtgcccagca gtgctccggg cgctgccgtg gcaagtcccc cagtgactgc
tgccacaacc 720agtgtgctgc aggctgcaca ggcccccggg agagcgactg cctggtctgc
cgcaaattcc 780gagacgaagc cacgtgcaag gacacctgcc ccccactcat gctctacaac
cccaccacgt 840accagatgga tgtgaacccc gagggcaaat acagctttgg tgccacctgc
gtgaagaagt 900gtccccgtaa ttatgtggtg acagatcacg gctcgtgcgt ccgagcctgt
ggggccgaca 960gctatgagat ggaggaagac ggcgtccgca agtgtaagaa gtgcgaaggg
ccttgccgca 1020aagtgtgtaa cggaataggt attggtgaat ttaaagactc actctccata
aatgctacga 1080atattaaaca cttcaaaaac tgcacctcca tcagtggcga tctccacatc
ctgccggtgg 1140catttagggg tgactccttc acacatactc ctcctctgga tccacaggaa
ctggatattc 1200tgaaaaccgt aaaggaaatc acagggtttt tgctgattca ggcttggcct
gaaaacagga 1260cggacctcca tgcctttgag aacctagaaa tcatacgcgg caggaccaag
caacatggtc 1320agttttctct tgcagtcgtc agcctgaaca taacatcctt gggattacgc
tccctcaagg 1380agataagtga tggagatgtg ataatttcag gaaacaaaaa tttgtgctat
gcaaatacaa 1440taaactggaa aaaactgttt gggacctccg gtcagaaaac caaaattata
agcaacagag 1500gtgaaaacag ctgcaaggcc acaggccagg tctgccatgc cttgtgctcc
cccgagggct 1560gctggggccc ggagcccagg gactgcgtct cttgccggaa tgtcagccga
ggcagggaat 1620gcgtggacaa gtgcaagctt ctggagggtg agccaaggga gtttgtggag
aactctgagt 1680gcatacagtg ccacccagag tgcctgcctc aggccatgaa catcacctgc
acaggacggg 1740gaccagacaa ctgtatccag tgtgcccact acattgacgg cccccactgc
gtcaagacct 1800gcccggcagg agtcatggga gaaaacaaca ccctggtctg gaagtacgca
gacgccggcc 1860atgtgtgcca cctgtgccat ccaaactgca cctacggatg cactgggcca
ggtcttgaag 1920gctgtccaac gaatgggcct aagatcccgt ccgggtgcgt atccggtgac
accattgtaa 1980tgactagtgg cggtccgcgc actgtggctg aactggaggg caaaccgttc
accgcactga 2040ttcgcggctc tggctaccca tgcccctcag gtttcttccg cacctgtgaa
cgtgacgtat 2100atgatctgcg tacacgtgag ggtcattgct tacgtttgac ccatgatcac
cgtgttctgg 2160tgatggatgg tggcctggaa tggcgtgccg cgggtgaact ggaacgcggc
gaccgcctgg 2220tgatggatga tgcagctggc gagtttccgg cactggcaac cttccgtggc
ctgcgtggcg 2280ctggccgcca ggatgtttat gacgctactg tttacggtgc tagcgcattc
actgctaatg 2340gcttcattgt acacgcatgt ggcgagcagc ccgggaccgg tctgaactca
ggcctcacga 2400caaatcctgg tgtatccgct tggcaggtca acacagctta tactgcggga
caattggtca 2460catataacgg caagacgtat aaatgtttgc agccccacac ctccttggca
ggatgggaac 2520catccaacgt tcctgccttg tggcagcttc aatgaagatc tgttaac
2567262588DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 26gttaacgaat tcccaccatg gagctggcgg
ccttgtgccg ctgggggctc ctcctcgccc 60tcttgccccc cggagccgcg agcacccaag
tgtgcaccgg cacagacatg aagctgcggc 120tccctgccag tcccgagacc cacctggaca
tgctccgcca cctctaccag ggctgccagg 180tggtgcaggg aaacctggaa ctcacctacc
tgcccaccaa tgccagcctg tccttcctgc 240aggatatcca ggaggtgcag ggctacgtgc
tcatcgctca caaccaagtg aggcaggtcc 300cactgcagag gctgcggatt gtgcgaggca
cccagctctt tgaggacaac tatgccctgg 360ccgtgctaga caatggagac ccgctgaaca
ataccacccc tgtcacaggg gcctccccag 420gaggcctgcg ggagctgcag cttcgaagcc
tcacagagat cttgaaagga ggggtcttga 480tccagcggaa cccccagctc tgctaccagg
acacgatttt gtggaaggac atcttccaca 540agaacaacca gctggctctc acactgatag
acaccaaccg ctctcgggcc tgccacccct 600gttctccgat gtgtaagggc tcccgctgct
ggggagagag ttctgaggat tgtcagagcc 660tgacgcgcac tgtctgtgcc ggtggctgtg
cccgctgcaa ggggccactg cccactgact 720gctgccatga gcagtgtgct gccggctgca
cgggccccaa gcactctgac tgcctggcct 780gcctccactt caaccacagt ggcatctgtg
agctgcactg cccagccctg gtcacctaca 840acacagacac gtttgagtcc atgcccaatc
ccgagggccg gtatacattc ggcgccagct 900gtgtgactgc ctgtccctac aactaccttt
ctacggacgt gggatcctgc accctcgtct 960gccccctgca caaccaagag gtgacagcag
aggatggaac acagcggtgt gagaagtgca 1020gcaagccctg tgcccgagtg tgctatggtc
tgggcatgga gcacttgcga gaggtgaggg 1080cagttaccag tgccaatatc caggagtttg
ctggctgcaa gaagatcttt gggagcctgg 1140catttctgcc ggagagcttt gatggggacc
cagcctccaa cactgccccg ctccagccag 1200agcagctcca agtgtttgag actctggaag
agatcacagg ttacctatac atctcagcat 1260ggccggacag cctgcctgac ctcagcgtct
tccagaacct gcaagtaatc cggggacgaa 1320ttctgcacaa tggcgcctac tcgctgaccc
tgcaagggct gggcatcagc tggctggggc 1380tgcgctcact gagggaactg ggcagtggac
tggccctcat ccaccataac acccacctct 1440gcttcgtgca cacggtgccc tgggaccagc
tctttcggaa cccgcaccaa gctctgctcc 1500acactgccaa ccggccagag gacgagtgtg
tgggcgaggg cctggcctgc caccagctgt 1560gcgcccgagg gcactgctgg ggtccagggc
ccacccagtg tgtcaactgc agccagttcc 1620ttcggggcca ggagtgcgtg gaggaatgcc
gagtactgca ggggctcccc agggagtatg 1680tgaatgccag gcactgtttg ccgtgccacc
ctgagtgtca gccccagaat ggctcagtga 1740cctgttttgg accggaggct gaccagtgtg
tggcctgtgc ccactataag gaccctccct 1800tctgcgtggc ccgctgcccc agcggtgtga
aacctgacct ctcctacatg cccatctgga 1860agtttccaga tgaggagggc gcatgccagc
cttgccccat caactgcacc cactcctgtg 1920tggacctgga tgacaagggc tgccccgccg
agcagagagc cagccctctg acggggtgcg 1980tatccggtga caccattgta atgactagtg
gcggtccgcg cactgtggct gaactggagg 2040gcaaaccgtt caccgcactg attcgcggct
ctggctaccc atgcccctca ggtttcttcc 2100gcacctgtga acgtgacgta tatgatctgc
gtacacgtga gggtcattgc ttacgtttga 2160cccatgatca ccgtgttctg gtgatggatg
gtggcctgga atggcgtgcc gcgggtgaac 2220tggaacgcgg cgaccgcctg gtgatggatg
atgcagctgg cgagtttccg gcactggcaa 2280ccttccgtgg cctgcgtggc gctggccgcc
aggatgttta tgacgctact gtttacggtg 2340ctagcgcatt cactgctaat ggcttcattg
tacacgcatg tggcgagcag cccgggaccg 2400gtctgaactc aggcctcacg acaaatcctg
gtgtatccgc ttggcaggtc aacacagctt 2460atactgcggg acaattggtc acatataacg
gcaagacgta taaatgtttg cagccccaca 2520cctccttggc aggatgggaa ccatccaacg
ttcctgcctt gtggcagctt caatgaagat 2580ctgttaac
2588272561DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 27gttaacgaat tcccaccatg agggcgaacg acgctctgca ggtgctgggc
ttgcttttca 60gcctggcccg gggctccgag gtgggcaact ctcaggcagt gtgtcctggg
actctgaatg 120gcctgagtgt gaccggcgat gctgagaacc aataccagac actgtacaag
ctctacgaga 180ggtgtgaggt ggtgatgggg aaccttgaga ttgtgctcac gggacacaat
gccgacctct 240ccttcctgca gtggattcga gaagtgacag gctatgtcct cgtggccatg
aatgaattct 300ctactctacc attgcccaac ctccgcgtgg tgcgagggac ccaggtctac
gatgggaagt 360ttgccatctt cgtcatgttg aactataaca ccaactccag ccacgctctg
cgccagctcc 420gcttgactca gctcaccgag attctgtcag ggggtgttta tattgagaag
aacgataagc 480tttgtcacat ggacacaatt gactggaggg acatcgtgag ggaccgagat
gctgagatag 540tggtgaagga caatggcaga agctgtcccc cctgtcatga ggtttgcaag
gggcgatgct 600ggggtcctgg atcagaagac tgccagacat tgaccaagac catctgtgct
cctcagtgta 660atggtcactg ctttgggccc aaccccaacc agtgctgcca tgatgagtgt
gccgggggct 720gctcaggccc tcaggacaca gactgctttg cctgccggca cttcaatgac
agtggagcct 780gtgtacctcg ctgtccacag cctcttgtct acaacaagct aactttccag
ctggaaccca 840atccccacac caagtatcag tatggaggag tttgtgtagc cagctgtccc
cataactttg 900tggtggatca aacatcctgt gtcagggcct gtcctcctga caagatggaa
gtagataaaa 960atgggctcaa gatgtgtgag ccttgtgggg gactatgtcc caaagcctgt
gagggaacag 1020gctctgggag ccgcttccag actgtggact cgagcaacat tgatggattt
gtgaactgca 1080ccaagatcct gggcaacctg gactttctga tcaccggcct caatggagac
ccctggcaca 1140agatccctgc cctggaccca gagaagctca atgtcttccg gacagtacgg
gagatcacag 1200gttacctgaa catccagtcc tggccgcccc acatgcacaa cttcagtgtt
ttttccaatt 1260tgacaaccat tggaggcaga agcctctaca accggggctt ctcattgttg
atcatgaaga 1320acttgaatgt cacatctctg ggcttccgat ccctgaagga aattagtgct
gggcgtatct 1380atataagtgc caataggcag ctctgctacc accactcttt gaactggacc
aaggtgcttc 1440gggggcctac ggaagagcga ctagacatca agcataatcg gccgcgcaga
gactgcgtgg 1500cagagggcaa agtgtgtgac ccactgtgct cctctggggg atgctggggc
ccaggccctg 1560gtcagtgctt gtcctgtcga aattatagcc gaggaggtgt ctgtgtgacc
cactgcaact 1620ttctgaatgg ggagcctcga gaatttgccc atgaggccga atgcttctcc
tgccacccgg 1680aatgccaacc catggagggc actgccacat gcaatggctc gggctctgat
acttgtgctc 1740aatgtgccca ttttcgagat gggccccact gtgtgagcag ctgcccccat
ggagtcctag 1800gtgccaaggg cccaatctac aagtacccag atgttcagaa tgaatgtcgg
ccctgccatg 1860agaactgcac ccaggggtgt aaaggaccag agcttcaaga ctgtttagga
caaacactgg 1920tgctgatcgg caaaacccat ctgacagggt gcgtatccgg tgacaccatt
gtaatgacta 1980gtggcggtcc gcgcactgtg gctgaactgg agggcaaacc gttcaccgca
ctgattcgcg 2040gctctggcta cccatgcccc tcaggtttct tccgcacctg tgaacgtgac
gtatatgatc 2100tgcgtacacg tgagggtcat tgcttacgtt tgacccatga tcaccgtgtt
ctggtgatgg 2160atggtggcct ggaatggcgt gccgcgggtg aactggaacg cggcgaccgc
ctggtgatgg 2220atgatgcagc tggcgagttt ccggcactgg caaccttccg tggcctgcgt
ggcgctggcc 2280gccaggatgt ttatgacgct actgtttacg gtgctagcgc attcactgct
aatggcttca 2340ttgtacacgc atgtggcgag cagcccggga ccggtctgaa ctcaggcctc
acgacaaatc 2400ctggtgtatc cgcttggcag gtcaacacag cttatactgc gggacaattg
gtcacatata 2460acggcaagac gtataaatgt ttgcagcccc acacctcctt ggcaggatgg
gaaccatcca 2520acgttcctgc cttgtggcag cttcaatgaa gatctgttaa c
2561282585DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 28gttaacgaat tcccaccatg aagccggcga
caggactttg ggtctgggtg agccttctcg 60tggcggcggg gaccgtccag cccagcgatt
ctcagtcagt gtgtgcagga acggagaata 120aactgagctc tctctctgac ctggaacagc
agtaccgagc cttgcgcaag tactatgaaa 180actgtgaggt tgtcatgggc aacctggaga
taaccagcat tgagcacaac cgggacctct 240ccttcctgcg gtctgttcga gaagtcacag
gctacgtgtt agtggctctt aatcagtttc 300gttacctgcc tctggagaat ttacgcatta
ttcgtgggac aaaactttat gaggatcgat 360atgccttggc aatattttta aactacagaa
aagatggaaa ctttggactt caagaacttg 420gattaaagaa cttgacagaa atcctaaatg
gtggagtcta tgtagaccag aacaaattcc 480tttgttatgc agacaccatt cattggcaag
atattgttcg gaacccatgg ccttccaact 540tgactcttgt gtcaacaaat ggtagttcag
gatgtggacg ttgccataag tcctgtactg 600gccgttgctg gggacccaca gaaaatcatt
gccagacttt gacaaggacg gtgtgtgcag 660aacaatgtga cggcagatgc tacggacctt
acgtcagtga ctgctgccat cgagaatgtg 720ctggaggctg ctcaggacct aaggacacag
actgctttgc ctgcatgaat ttcaatgaca 780gtggagcatg tgttactcag tgtccccaaa
cctttgtcta caatccaacc acctttcaac 840tggagcacaa tttcaatgca aagtacacat
atggagcatt ctgtgtcaag aaatgtccac 900ataactttgt ggtagattcc agttcttgtg
tgcgtgcctg ccctagttcc aagatggaag 960tagaagaaaa tgggattaaa atgtgtaaac
cttgcactga catttgccca aaagcttgtg 1020atggcattgg cacaggatca ttgatgtcag
ctcagactgt ggattccagt aacattgaca 1080aattcataaa ctgtaccaag atcaatggga
atttgatctt tctagtcact ggtattcatg 1140gggaccctta caatgcaatt gaagccatag
acccagagaa actgaacgtc tttcggacag 1200tcagagagat aacaggtttc ctgaacatac
agtcatggcc accaaacatg actgacttca 1260gtgttttttc taacctggtg accattggtg
gaagagtact ctatagtggc ctgtccttgc 1320ttatcctcaa gcaacagggc atcacctctc
tacagttcca gtccctgaag gaaatcagcg 1380caggaaacat ctatattact gacaacagca
acctgtgtta ttatcatacc attaactgga 1440caacactctt cagcacaatc aaccagagaa
tagtaatccg ggacaacaga aaagctgaaa 1500attgtactgc tgaaggaatg gtgtgcaacc
atctgtgttc cagtgatggc tgttggggac 1560ctgggccaga ccaatgtctg tcgtgtcgcc
gcttcagtag aggaaggatc tgcatagagt 1620cttgtaacct ctatgatggt gaatttcggg
agtttgagaa tggctccatc tgtgtggagt 1680gtgaccccca gtgtgagaag atggaagatg
gcctcctcac atgccatgga ccgggtcctg 1740acaactgtac aaagtgctct cattttaaag
atggcccaaa ctgtgtggaa aaatgtccag 1800atggcttaca gggggcaaac agtttcattt
tcaagtatgc tgatccagat cgggagtgcc 1860acccatgcca tccaaactgc acccaagggt
gtaacggtcc cactagtcat gactgcattt 1920actacccatg gacgggccat tccactttac
cacaacatgc tagaactccc gggtgcgtat 1980ccggtgacac cattgtaatg actagtggcg
gtccgcgcac tgtggctgaa ctggagggca 2040aaccgttcac cgcactgatt cgcggctctg
gctacccatg cccctcaggt ttcttccgca 2100cctgtgaacg tgacgtatat gatctgcgta
cacgtgaggg tcattgctta cgtttgaccc 2160atgatcaccg tgttctggtg atggatggtg
gcctggaatg gcgtgccgcg ggtgaactgg 2220aacgcggcga ccgcctggtg atggatgatg
cagctggcga gtttccggca ctggcaacct 2280tccgtggcct gcgtggcgct ggccgccagg
atgtttatga cgctactgtt tacggtgcta 2340gcgcattcac tgctaatggc ttcattgtac
acgcatgtgg cgagcagccc gggaccggtc 2400tgaactcagg cctcacgaca aatcctggtg
tatccgcttg gcaggtcaac acagcttata 2460ctgcgggaca attggtcaca tataacggca
agacgtataa atgtttgcag ccccacacct 2520ccttggcagg atgggaacca tccaacgttc
ctgccttgtg gcagcttcaa tgaagatctg 2580ttaac
258529785DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 29aagcttgaat tcccaccatg ctgctgctgg cgagatgtct gctgctagtc
ctcgtctcct 60cgctgctggt atgctcggga ctggcgtgtg acaaaactca cacatgccca
ccgtgcccag 120cacctgaact cctggggggg ccctcagtct tcctcttccc cccaaaaccc
aaggacaccc 180tcatgatctc ccggacccct gaggtcacat gcgtggtggt ggacgtgagc
cacgaagacc 240ctgaggtcaa gttcaactgg tacgtggacg gcgtggaggt gcataatgcc
aagacaaagc 300cgcgggagga gcagtacaac agcacgtacc gtgtggtcag cgtcctcacc
gtcctgcacc 360aggactggct gaatggcaag gagtacaagt gcaaggtctc caacaaagcc
ctcccagccc 420ccatcgagaa aaccatctcc aaagccaaag ggcagccccg agaaccacag
gtgtacaccc 480tgcccccatc ccgggatgag ctgaccaaga accaggtcag cctgacctgc
ctggtcaaag 540gcttctatcc cagcgacatc gccgtggagt gggagagcaa tgggcagccg
gagaacaact 600acaagaccac gcctcccgtg ctggactccg acggctcctt cttcctctac
agcaagctca 660ccgtggacaa gagcaggtgg cagcagggga acgtcttctc atgctccgtg
atgcatgagg 720ctctgcacaa ccactacacg cagaagagcc tctccctgtc tccgggtaaa
tgactcgagc 780ggccg
78530785DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 30aagcttgaat tcccaccatg gccttgacct
ttgctttact ggtggccctc ctggtgctca 60gctgcaagtc aagctgctct gtgggctgtg
acaaaactca cacatgccca ccgtgcccag 120cacctgaact cctggggggg ccctcagtct
tcctcttccc cccaaaaccc aaggacaccc 180tcatgatctc ccggacccct gaggtcacat
gcgtggtggt ggacgtgagc cacgaagacc 240ctgaggtcaa gttcaactgg tacgtggacg
gcgtggaggt gcataatgcc aagacaaagc 300cgcgggagga gcagtacaac agcacgtacc
gtgtggtcag cgtcctcacc gtcctgcacc 360aggactggct gaatggcaag gagtacaagt
gcaaggtctc caacaaagcc ctcccagccc 420ccatcgagaa aaccatctcc aaagccaaag
ggcagccccg agaaccacag gtgtacaccc 480tgcccccatc ccgggatgag ctgaccaaga
accaggtcag cctgacctgc ctggtcaaag 540gcttctatcc cagcgacatc gccgtggagt
gggagagcaa tgggcagccg gagaacaact 600acaagaccac gcctcccgtg ctggactccg
acggctcctt cttcctctac agcaagctca 660ccgtggacaa gagcaggtgg cagcagggga
acgtcttctc atgctccgtg atgcatgagg 720ctctgcacaa ccactacacg cagaagagcc
tctccctgtc tccgggtaaa tgactcgagc 780ggccg
78531767DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 31aagcttgaat tcccaccatg ctggctgcca cagtcctgac cctggccctg
ctgggcaatg 60cccatgcctg tgacaaaact cacacatgcc caccgtgccc agcacctgaa
ctcctggggg 120ggccctcagt cttcctcttc cccccaaaac ccaaggacac cctcatgatc
tcccggaccc 180ctgaggtcac atgcgtggtg gtggacgtga gccacgaaga ccctgaggtc
aagttcaact 240ggtacgtgga cggcgtggag gtgcataatg ccaagacaaa gccgcgggag
gagcagtaca 300acagcacgta ccgtgtggtc agcgtcctca ccgtcctgca ccaggactgg
ctgaatggca 360aggagtacaa gtgcaaggtc tccaacaaag ccctcccagc ccccatcgag
aaaaccatct 420ccaaagccaa agggcagccc cgagaaccac aggtgtacac cctgccccca
tcccgggatg 480agctgaccaa gaaccaggtc agcctgacct gcctggtcaa aggcttctat
cccagcgaca 540tcgccgtgga gtgggagagc aatgggcagc cggagaacaa ctacaagacc
acgcctcccg 600tgctggactc cgacggctcc ttcttcctct acagcaagct caccgtggac
aagagcaggt 660ggcagcaggg gaacgtcttc tcatgctccg tgatgcatga ggctctgcac
aaccactaca 720cgcagaagag cctctccctg tctccgggta aatgactcga gcggccg
76732251PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 32Met Leu Leu Leu Ala Arg Cys Leu
Leu Leu Val Leu Val Ser Ser Leu1 5 10
15Leu Val Cys Ser Gly Leu Ala Cys Asp Lys Thr His Thr Cys
Pro Pro20 25 30Cys Pro Ala Pro Glu Leu
Leu Gly Gly Pro Ser Val Phe Leu Phe Pro35 40
45Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr50
55 60Cys Val Val Val Asp Val Ser His Glu
Asp Pro Glu Val Lys Phe Asn65 70 75
80Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys
Pro Arg85 90 95Glu Glu Gln Tyr Asn Ser
Thr Tyr Arg Val Val Ser Val Leu Thr Val100 105
110Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val
Ser115 120 125Asn Lys Ala Leu Pro Ala Pro
Ile Glu Lys Thr Ile Ser Lys Ala Lys130 135
140Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp145
150 155 160Glu Leu Thr Lys
Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe165 170
175Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu180 185 190Asn Asn Tyr Lys Thr Thr
Pro Pro Val Leu Asp Ser Asp Gly Ser Phe195 200
205Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln
Gly210 215 220Asn Val Phe Ser Cys Ser Val
Met His Glu Ala Leu His Asn His Tyr225 230
235 240Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys245
25033251PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 33Met Ala Leu Thr Phe Ala Leu Leu
Val Ala Leu Leu Val Leu Ser Cys1 5 10
15Lys Ser Ser Cys Ser Val Gly Cys Asp Lys Thr His Thr Cys
Pro Pro20 25 30Cys Pro Ala Pro Glu Leu
Leu Gly Gly Pro Ser Val Phe Leu Phe Pro35 40
45Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr50
55 60Cys Val Val Val Asp Val Ser His Glu
Asp Pro Glu Val Lys Phe Asn65 70 75
80Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys
Pro Arg85 90 95Glu Glu Gln Tyr Asn Ser
Thr Tyr Arg Val Val Ser Val Leu Thr Val100 105
110Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val
Ser115 120 125Asn Lys Ala Leu Pro Ala Pro
Ile Glu Lys Thr Ile Ser Lys Ala Lys130 135
140Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp145
150 155 160Glu Leu Thr Lys
Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe165 170
175Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu180 185 190Asn Asn Tyr Lys Thr Thr
Pro Pro Val Leu Asp Ser Asp Gly Ser Phe195 200
205Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln
Gly210 215 220Asn Val Phe Ser Cys Ser Val
Met His Glu Ala Leu His Asn His Tyr225 230
235 240Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys245
25034245PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 34Met Leu Ala Ala Thr Val Leu Thr
Leu Ala Leu Leu Gly Asn Ala His1 5 10
15Ala Cys Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro
Glu Leu20 25 30Leu Gly Gly Pro Ser Val
Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr35 40
45Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val50
55 60Ser His Glu Asp Pro Glu Val Lys Phe
Asn Trp Tyr Val Asp Gly Val65 70 75
80Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr
Asn Ser85 90 95Thr Tyr Arg Val Val Ser
Val Leu Thr Val Leu His Gln Asp Trp Leu100 105
110Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro
Ala115 120 125Pro Ile Glu Lys Thr Ile Ser
Lys Ala Lys Gly Gln Pro Arg Glu Pro130 135
140Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln145
150 155 160Val Ser Leu Thr
Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala165 170
175Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys
Thr Thr180 185 190Pro Pro Val Leu Asp Ser
Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu195 200
205Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys
Ser210 215 220Val Met His Glu Ala Leu His
Asn His Tyr Thr Gln Lys Ser Leu Ser225 230
235 240Leu Ser Pro Gly Lys24535228PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 35Cys Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro Glu
Leu Leu1 5 10 15Gly Gly
Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu20
25 30Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val
Val Asp Val Ser35 40 45His Glu Asp Pro
Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu50 55
60Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn
Ser Thr65 70 75 80Tyr
Arg Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn85
90 95Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys
Ala Leu Pro Ala Pro100 105 110Ile Glu Lys
Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln115
120 125Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr
Lys Asn Gln Val130 135 140Ser Leu Thr Cys
Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val145 150
155 160Glu Trp Glu Ser Asn Gly Gln Pro Glu
Asn Asn Tyr Lys Thr Thr Pro165 170 175Pro
Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr180
185 190Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val
Phe Ser Cys Ser Val195 200 205Met His Glu
Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu210
215 220Ser Pro Gly Lys22536228PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 36Xaa Asp Lys Thr His Thr Xaa Pro Pro Xaa Pro Ala Pro Glu
Leu Leu1 5 10 15Gly Gly
Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu20
25 30Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val
Val Asp Val Ser35 40 45His Glu Asp Pro
Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu50 55
60Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn
Ser Thr65 70 75 80Tyr
Arg Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn85
90 95Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys
Ala Leu Pro Ala Pro100 105 110Ile Glu Lys
Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln115
120 125Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr
Lys Asn Gln Val130 135 140Ser Leu Thr Cys
Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val145 150
155 160Glu Trp Glu Ser Asn Gly Gln Pro Glu
Asn Asn Tyr Lys Thr Thr Pro165 170 175Pro
Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr180
185 190Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val
Phe Ser Cys Ser Val195 200 205Met His Glu
Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu210
215 220Ser Pro Gly Lys22537222PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 37Xaa Pro Pro Xaa Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser
Val Phe1 5 10 15Leu Phe
Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro20
25 30Glu Val Thr Cys Val Val Val Asp Val Ser His Glu
Asp Pro Glu Val35 40 45Lys Phe Asn Trp
Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr50 55
60Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val Val
Ser Val65 70 75 80Leu
Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys85
90 95Lys Val Ser Asn Lys Ala Leu Pro Ala Pro Ile
Glu Lys Thr Ile Ser100 105 110Lys Ala Lys
Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro115
120 125Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser Leu
Thr Cys Leu Val130 135 140Lys Gly Phe Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly145 150
155 160Gln Pro Glu Asn Asn Tyr Lys Thr Thr
Pro Pro Val Leu Asp Ser Asp165 170 175Gly
Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp180
185 190Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met
His Glu Ala Leu His195 200 205Asn His Tyr
Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys210 215
22038219PRTArtificial Sequenceartificial construct relating to
Homo Sapiens immunoglobulin 38Cys Pro Ala Pro Glu Leu Leu Gly Gly
Pro Ser Val Phe Leu Phe Pro1 5 10
15Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val
Thr20 25 30Cys Val Val Val Asp Val Ser
His Glu Asp Pro Glu Val Lys Phe Asn35 40
45Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg50
55 60Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val
Val Ser Val Leu Thr Val65 70 75
80Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val
Ser85 90 95Asn Lys Ala Leu Pro Ala Pro
Ile Glu Lys Thr Ile Ser Lys Ala Lys100 105
110Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp115
120 125Glu Leu Thr Lys Asn Gln Val Ser Leu
Thr Cys Leu Val Lys Gly Phe130 135 140Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu145
150 155 160Asn Asn Tyr Lys Thr Thr
Pro Pro Val Leu Asp Ser Asp Gly Ser Phe165 170
175Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln
Gly180 185 190Asn Val Phe Ser Cys Ser Val
Met His Glu Ala Leu His Asn His Tyr195 200
205Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys210
21539224PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 39Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser
Val Phe Leu Phe Pro Pro1 5 10
15Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys20
25 30Val Val Val Asp Val Ser His Glu Asp
Pro Glu Val Lys Phe Asn Trp35 40 45Tyr
Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu50
55 60Glu Gln Tyr Asn Ser Thr Tyr Arg Val Val Ser
Val Leu Thr Val Leu65 70 75
80His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn85
90 95Lys Ala Leu Pro Ala Pro Ile Glu Lys
Thr Ile Ser Lys Ala Lys Gly100 105 110Gln
Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu115
120 125Leu Thr Lys Asn Gln Val Ser Leu Thr Cys Leu
Val Lys Gly Phe Tyr130 135 140Pro Ser Asp
Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn145
150 155 160Asn Tyr Lys Thr Thr Pro Pro
Val Leu Asp Ser Asp Gly Ser Phe Phe165 170
175Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn180
185 190Val Phe Ser Cys Ser Val Met His Glu
Ala Leu His Asn His Tyr Thr195 200 205Gln
Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln Leu Glu Glu Ser Cys210
215 22040235PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 40Pro Ala Pro Glu
Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro1 5
10 15Lys Pro Lys Asp Thr Leu Met Ile Ser Arg
Thr Pro Glu Val Thr Cys20 25 30Val Val
Val Asp Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp35
40 45Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr
Lys Pro Arg Glu50 55 60Glu Gln Tyr Asn
Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu65 70
75 80His Gln Asp Trp Leu Asn Gly Lys Glu
Tyr Lys Cys Lys Val Ser Asn85 90 95Lys
Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly100
105 110Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro
Pro Ser Arg Asp Glu115 120 125Leu Thr Lys
Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr130
135 140Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly
Gln Pro Glu Asn145 150 155
160Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe165
170 175Leu Tyr Ser Lys Leu Thr Val Asp Lys
Ser Arg Trp Gln Gln Gly Asn180 185 190Val
Phe Ser Cys Ser Val Met His Glu Ala Leu His Asn His Tyr Thr195
200 205Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln
Leu Glu Glu Ser Xaa210 215 220Ala Glu Ala
Gln Asp Gly Glu Leu Asp Gly Xaa225 230
23541234PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 41Xaa Asp Lys Thr His Thr Xaa Pro Pro Xaa
Pro Ala Pro Glu Leu Leu1 5 10
15Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu20
25 30Met Ile Ser Arg Thr Pro Glu Val Thr
Cys Val Val Val Asp Val Ser35 40 45His
Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu50
55 60Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu
Gln Tyr Asn Ser Thr65 70 75
80Tyr Arg Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn85
90 95Gly Lys Glu Tyr Lys Cys Lys Val Ser
Asn Lys Ala Leu Pro Ala Pro100 105 110Ile
Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln115
120 125Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu
Thr Lys Asn Gln Val130 135 140Ser Leu Thr
Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val145
150 155 160Glu Trp Glu Ser Asn Gly Gln
Pro Glu Asn Asn Tyr Lys Thr Thr Pro165 170
175Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr180
185 190Val Asp Lys Ser Arg Trp Gln Gln Gly
Asn Val Phe Ser Cys Ser Val195 200 205Met
His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu210
215 220Ser Pro Glu Leu Gln Leu Glu Glu Ser Cys225
23042228PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 42Xaa Pro Pro Xaa Pro Ala Pro Glu
Leu Leu Gly Gly Pro Ser Val Phe1 5 10
15Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg
Thr Pro20 25 30Glu Val Thr Cys Val Val
Val Asp Val Ser His Glu Asp Pro Glu Val35 40
45Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr50
55 60Lys Pro Arg Glu Glu Gln Tyr Asn Ser
Thr Tyr Arg Val Val Ser Val65 70 75
80Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr
Lys Cys85 90 95Lys Val Ser Asn Lys Ala
Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser100 105
110Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro
Pro115 120 125Ser Arg Asp Glu Leu Thr Lys
Asn Gln Val Ser Leu Thr Cys Leu Val130 135
140Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly145
150 155 160Gln Pro Glu Asn
Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser Asp165 170
175Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser
Arg Trp180 185 190Gln Gln Gly Asn Val Phe
Ser Cys Ser Val Met His Glu Ala Leu His195 200
205Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln
Leu210 215 220Glu Glu Ser
Cys22543225PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 43Cys Pro Ala Pro Glu Leu Leu Gly Gly Pro
Ser Val Phe Leu Phe Pro1 5 10
15Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr20
25 30Cys Val Val Val Asp Val Ser His Glu
Asp Pro Glu Val Lys Phe Asn35 40 45Trp
Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg50
55 60Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val Val
Ser Val Leu Thr Val65 70 75
80Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser85
90 95Asn Lys Ala Leu Pro Ala Pro Ile Glu
Lys Thr Ile Ser Lys Ala Lys100 105 110Gly
Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp115
120 125Glu Leu Thr Lys Asn Gln Val Ser Leu Thr Cys
Leu Val Lys Gly Phe130 135 140Tyr Pro Ser
Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu145
150 155 160Asn Asn Tyr Lys Thr Thr Pro
Pro Val Leu Asp Ser Asp Gly Ser Phe165 170
175Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly180
185 190Asn Val Phe Ser Cys Ser Val Met His
Glu Ala Leu His Asn His Tyr195 200 205Thr
Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln Leu Glu Glu Ser210
215 220Cys22544245PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 44Xaa Asp Lys Thr
His Thr Xaa Pro Pro Xaa Pro Ala Pro Glu Leu Leu1 5
10 15Gly Gly Pro Ser Val Phe Leu Phe Pro Pro
Lys Pro Lys Asp Thr Leu20 25 30Met Ile
Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser35
40 45His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val
Asp Gly Val Glu50 55 60Val His Asn Ala
Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr65 70
75 80Tyr Arg Val Val Ser Val Leu Thr Val
Leu His Gln Asp Trp Leu Asn85 90 95Gly
Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro100
105 110Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln
Pro Arg Glu Pro Gln115 120 125Val Tyr Thr
Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln Val130
135 140Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser
Asp Ile Ala Val145 150 155
160Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro165
170 175Pro Val Leu Asp Ser Asp Gly Ser Phe
Phe Leu Tyr Ser Lys Leu Thr180 185 190Val
Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val195
200 205Met His Glu Ala Leu His Asn His Tyr Thr Gln
Lys Ser Leu Ser Leu210 215 220Ser Pro Glu
Leu Gln Leu Glu Glu Ser Xaa Ala Glu Ala Gln Asp Gly225
230 235 240Glu Leu Asp Gly
Xaa24545239PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 45Xaa Pro Pro Xaa Pro Ala Pro Glu Leu Leu
Gly Gly Pro Ser Val Phe1 5 10
15Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro20
25 30Glu Val Thr Cys Val Val Val Asp Val
Ser His Glu Asp Pro Glu Val35 40 45Lys
Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr50
55 60Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr
Arg Val Val Ser Val65 70 75
80Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys85
90 95Lys Val Ser Asn Lys Ala Leu Pro Ala
Pro Ile Glu Lys Thr Ile Ser100 105 110Lys
Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro115
120 125Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu Val130 135 140Lys Gly Phe
Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly145
150 155 160Gln Pro Glu Asn Asn Tyr Lys
Thr Thr Pro Pro Val Leu Asp Ser Asp165 170
175Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp180
185 190Gln Gln Gly Asn Val Phe Ser Cys Ser
Val Met His Glu Ala Leu His195 200 205Asn
His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln Leu210
215 220Glu Glu Ser Xaa Ala Glu Ala Gln Asp Gly Glu
Leu Asp Gly Xaa225 230
23546236PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 46Cys Pro Ala Pro Glu Leu Leu Gly Gly Pro
Ser Val Phe Leu Phe Pro1 5 10
15Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr20
25 30Cys Val Val Val Asp Val Ser His Glu
Asp Pro Glu Val Lys Phe Asn35 40 45Trp
Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg50
55 60Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val Val
Ser Val Leu Thr Val65 70 75
80Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser85
90 95Asn Lys Ala Leu Pro Ala Pro Ile Glu
Lys Thr Ile Ser Lys Ala Lys100 105 110Gly
Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp115
120 125Glu Leu Thr Lys Asn Gln Val Ser Leu Thr Cys
Leu Val Lys Gly Phe130 135 140Tyr Pro Ser
Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu145
150 155 160Asn Asn Tyr Lys Thr Thr Pro
Pro Val Leu Asp Ser Asp Gly Ser Phe165 170
175Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly180
185 190Asn Val Phe Ser Cys Ser Val Met His
Glu Ala Leu His Asn His Tyr195 200 205Thr
Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln Leu Glu Glu Ser210
215 220Xaa Ala Glu Ala Gln Asp Gly Glu Leu Asp Gly
Xaa225 230 23547776DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 47aagcttgaat tcccaccatg ctgctgctgg cgagatgtct gctgctagtc
ctcgtctcct 60cgctgctggt atgctcggga ctggcgtgtt gtgtcgagtg cccaccgtgc
ccagcaccac 120ctgtggcagg accgtcagtc ttcctcttcc ccccaaaacc caaggacacc
ctcatgatct 180cccggacccc tgaggtcacg tgcgtggtgg tggacgtgag ccacgaagac
cccgaggtcc 240agttcaactg gtacgtggac ggcgtggagg tgcataatgc caagacaaag
ccacgggagg 300agcagttcaa cagcacgttc cgtgtggtca gcgtcctcac cgttgtgcac
caggactggc 360tgaacggcaa ggagtacaag tgcaaggtct ccaacaaagg cctcccagcc
cccatcgaga 420aaaccatctc caaaaccaaa gggcagcccc gagaaccaca ggtgtacacc
ctgcccccat 480cccgggagga gatgaccaag aaccaggtca gcctgacctg cctggtcaaa
ggcttctacc 540ccagcgacat cgccgtggag tgggagagca atgggcagcc ggagaacaac
tacaagacca 600cacctcccat gctggactcc gacggctcct tcttcctcta cagcaagctc
accgtggaca 660agagcaggtg gcagcagggg aacgtcttct catgctccgt gatgcatgag
gctctgcaca 720accactacac gcagaagagc ctctccctgt ctccgggtaa atgactcgag
cggccg 77648776DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 48aagcttgaat tcccaccatg gccttgacct
ttgctttact ggtggccctc ctggtgctca 60gctgcaagtc aagctgctct gtgggctgtt
gtgtcgagtg cccaccgtgc ccagcaccac 120ctgtggcagg accgtcagtc ttcctcttcc
ccccaaaacc caaggacacc ctcatgatct 180cccggacccc tgaggtcacg tgcgtggtgg
tggacgtgag ccacgaagac cccgaggtcc 240agttcaactg gtacgtggac ggcgtggagg
tgcataatgc caagacaaag ccacgggagg 300agcagttcaa cagcacgttc cgtgtggtca
gcgtcctcac cgttgtgcac caggactggc 360tgaacggcaa ggagtacaag tgcaaggtct
ccaacaaagg cctcccagcc cccatcgaga 420aaaccatctc caaaaccaaa gggcagcccc
gagaaccaca ggtgtacacc ctgcccccat 480cccgggagga gatgaccaag aaccaggtca
gcctgacctg cctggtcaaa ggcttctacc 540ccagcgacat cgccgtggag tgggagagca
atgggcagcc ggagaacaac tacaagacca 600cacctcccat gctggactcc gacggctcct
tcttcctcta cagcaagctc accgtggaca 660agagcaggtg gcagcagggg aacgtcttct
catgctccgt gatgcatgag gctctgcaca 720accactacac gcagaagagc ctctccctgt
ctccgggtaa atgactcgag cggccg 77649758DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 49aagcttgaat tcccaccatg ctggctgcca cagtcctgac cctggccctg
ctgggcaatg 60cccatgcctg ttgtgtcgag tgcccaccgt gcccagcacc acctgtggca
ggaccgtcag 120tcttcctctt ccccccaaaa cccaaggaca ccctcatgat ctcccggacc
cctgaggtca 180cgtgcgtggt ggtggacgtg agccacgaag accccgaggt ccagttcaac
tggtacgtgg 240acggcgtgga ggtgcataat gccaagacaa agccacggga ggagcagttc
aacagcacgt 300tccgtgtggt cagcgtcctc accgttgtgc accaggactg gctgaacggc
aaggagtaca 360agtgcaaggt ctccaacaaa ggcctcccag cccccatcga gaaaaccatc
tccaaaacca 420aagggcagcc ccgagaacca caggtgtaca ccctgccccc atcccgggag
gagatgacca 480agaaccaggt cagcctgacc tgcctggtca aaggcttcta ccccagcgac
atcgccgtgg 540agtgggagag caatgggcag ccggagaaca actacaagac cacacctccc
atgctggact 600ccgacggctc cttcttcctc tacagcaagc tcaccgtgga caagagcagg
tggcagcagg 660ggaacgtctt ctcatgctcc gtgatgcatg aggctctgca caaccactac
acgcagaaga 720gcctctccct gtctccgggt aaatgactcg agcggccg
75850248PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 50Met Leu Leu Leu Ala Arg Cys Leu
Leu Leu Val Leu Val Ser Ser Leu1 5 10
15Leu Val Cys Ser Gly Leu Ala Cys Cys Val Glu Cys Pro Pro
Cys Pro20 25 30Ala Pro Pro Val Ala Gly
Pro Ser Val Phe Leu Phe Pro Pro Lys Pro35 40
45Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val50
55 60Val Asp Val Ser His Glu Asp Pro Glu
Val Gln Phe Asn Trp Tyr Val65 70 75
80Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu
Glu Gln85 90 95Phe Asn Ser Thr Phe Arg
Val Val Ser Val Leu Thr Val Val His Gln100 105
110Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys
Gly115 120 125Leu Pro Ala Pro Ile Glu Lys
Thr Ile Ser Lys Thr Lys Gly Gln Pro130 135
140Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu Met Thr145
150 155 160Lys Asn Gln Val
Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser165 170
175Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn
Asn Tyr180 185 190Lys Thr Thr Pro Pro Met
Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr195 200
205Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val
Phe210 215 220Ser Cys Ser Val Met His Glu
Ala Leu His Asn His Tyr Thr Gln Lys225 230
235 240Ser Leu Ser Leu Ser Pro Gly
Lys24551248PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 51Met Ala Leu Thr Phe Ala Leu Leu Val Ala
Leu Leu Val Leu Ser Cys1 5 10
15Lys Ser Ser Cys Ser Val Gly Cys Cys Val Glu Cys Pro Pro Cys Pro20
25 30Ala Pro Pro Val Ala Gly Pro Ser Val
Phe Leu Phe Pro Pro Lys Pro35 40 45Lys
Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val50
55 60Val Asp Val Ser His Glu Asp Pro Glu Val Gln
Phe Asn Trp Tyr Val65 70 75
80Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln85
90 95Phe Asn Ser Thr Phe Arg Val Val Ser
Val Leu Thr Val Val His Gln100 105 110Asp
Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly115
120 125Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys
Thr Lys Gly Gln Pro130 135 140Arg Glu Pro
Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu Met Thr145
150 155 160Lys Asn Gln Val Ser Leu Thr
Cys Leu Val Lys Gly Phe Tyr Pro Ser165 170
175Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr180
185 190Lys Thr Thr Pro Pro Met Leu Asp Ser
Asp Gly Ser Phe Phe Leu Tyr195 200 205Ser
Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe210
215 220Ser Cys Ser Val Met His Glu Ala Leu His Asn
His Tyr Thr Gln Lys225 230 235
240Ser Leu Ser Leu Ser Pro Gly Lys24552242PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 52Met Leu Ala Ala Thr Val Leu Thr Leu Ala Leu Leu Gly Asn
Ala His1 5 10 15Ala Cys
Cys Val Glu Cys Pro Pro Cys Pro Ala Pro Pro Val Ala Gly20
25 30Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp
Thr Leu Met Ile35 40 45Ser Arg Thr Pro
Glu Val Thr Cys Val Val Val Asp Val Ser His Glu50 55
60Asp Pro Glu Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu
Val His65 70 75 80Asn
Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Phe Arg85
90 95Val Val Ser Val Leu Thr Val Val His Gln Asp
Trp Leu Asn Gly Lys100 105 110Glu Tyr Lys
Cys Lys Val Ser Asn Lys Gly Leu Pro Ala Pro Ile Glu115
120 125Lys Thr Ile Ser Lys Thr Lys Gly Gln Pro Arg Glu
Pro Gln Val Tyr130 135 140Thr Leu Pro Pro
Ser Arg Glu Glu Met Thr Lys Asn Gln Val Ser Leu145 150
155 160Thr Cys Leu Val Lys Gly Phe Tyr Pro
Ser Asp Ile Ala Val Glu Trp165 170 175Glu
Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Met180
185 190Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser
Lys Leu Thr Val Asp195 200 205Lys Ser Arg
Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met His210
215 220Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu
Ser Leu Ser Pro225 230 235
240Gly Lys53225PRTArtificial Sequenceartificial construct relating to
Homo Sapiens immunoglobulin 53Cys Cys Val Glu Cys Pro Pro Cys Pro
Ala Pro Pro Val Ala Gly Pro1 5 10
15Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile
Ser20 25 30Arg Thr Pro Glu Val Thr Cys
Val Val Val Asp Val Ser His Glu Asp35 40
45Pro Glu Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn50
55 60Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe
Asn Ser Thr Phe Arg Val65 70 75
80Val Ser Val Leu Thr Val Val His Gln Asp Trp Leu Asn Gly Lys
Glu85 90 95Tyr Lys Cys Lys Val Ser Asn
Lys Gly Leu Pro Ala Pro Ile Glu Lys100 105
110Thr Ile Ser Lys Thr Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr115
120 125Leu Pro Pro Ser Arg Glu Glu Met Thr
Lys Asn Gln Val Ser Leu Thr130 135 140Cys
Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu145
150 155 160Ser Asn Gly Gln Pro Glu
Asn Asn Tyr Lys Thr Thr Pro Pro Met Leu165 170
175Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp
Lys180 185 190Ser Arg Trp Gln Gln Gly Asn
Val Phe Ser Cys Ser Val Met His Glu195 200
205Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly210
215 220Lys22554225PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 54Xaa Xaa Val Glu Xaa Pro Pro Xaa Pro Ala Pro Pro Val Ala
Gly Pro1 5 10 15Ser Val
Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser20
25 30Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val
Ser His Glu Asp35 40 45Pro Glu Val Gln
Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn50 55
60Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Phe
Arg Val65 70 75 80Val
Ser Val Leu Thr Val Val His Gln Asp Trp Leu Asn Gly Lys Glu85
90 95Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu Pro
Ala Pro Ile Glu Lys100 105 110Thr Ile Ser
Lys Thr Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr115
120 125Leu Pro Pro Ser Arg Glu Glu Met Thr Lys Asn Gln
Val Ser Leu Thr130 135 140Cys Leu Val Lys
Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu145 150
155 160Ser Asn Gly Gln Pro Glu Asn Asn Tyr
Lys Thr Thr Pro Pro Met Leu165 170 175Asp
Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys180
185 190Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys
Ser Val Met His Glu195 200 205Ala Leu His
Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly210
215 220Lys22555224PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 55Xaa Val Glu Xaa
Pro Pro Xaa Pro Ala Pro Pro Val Ala Gly Pro Ser1 5
10 15Val Phe Leu Phe Pro Pro Lys Pro Lys Asp
Thr Leu Met Ile Ser Arg20 25 30Thr Pro
Glu Val Thr Cys Val Val Val Asp Val Ser His Glu Asp Pro35
40 45Glu Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu
Val His Asn Ala50 55 60Lys Thr Lys Pro
Arg Glu Glu Gln Phe Asn Ser Thr Phe Arg Val Val65 70
75 80Ser Val Leu Thr Val Val His Gln Asp
Trp Leu Asn Gly Lys Glu Tyr85 90 95Lys
Cys Lys Val Ser Asn Lys Gly Leu Pro Ala Pro Ile Glu Lys Thr100
105 110Ile Ser Lys Thr Lys Gly Gln Pro Arg Glu Pro
Gln Val Tyr Thr Leu115 120 125Pro Pro Ser
Arg Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys130
135 140Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val
Glu Trp Glu Ser145 150 155
160Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Met Leu Asp165
170 175Ser Asp Gly Ser Phe Phe Leu Tyr Ser
Lys Leu Thr Val Asp Lys Ser180 185 190Arg
Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala195
200 205Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser
Leu Ser Pro Gly Lys210 215
22056221PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 56Xaa Pro Pro Xaa Pro Ala Pro Pro Val Ala
Gly Pro Ser Val Phe Leu1 5 10
15Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu20
25 30Val Thr Cys Val Val Val Asp Val Ser
His Glu Asp Pro Glu Val Gln35 40 45Phe
Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys50
55 60Pro Arg Glu Glu Gln Phe Asn Ser Thr Phe Arg
Val Val Ser Val Leu65 70 75
80Thr Val Val His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys85
90 95Val Ser Asn Lys Gly Leu Pro Ala Pro
Ile Glu Lys Thr Ile Ser Lys100 105 110Thr
Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser115
120 125Arg Glu Glu Met Thr Lys Asn Gln Val Ser Leu
Thr Cys Leu Val Lys130 135 140Gly Phe Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln145
150 155 160Pro Glu Asn Asn Tyr Lys Thr
Thr Pro Pro Met Leu Asp Ser Asp Gly165 170
175Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln180
185 190Gln Gly Asn Val Phe Ser Cys Ser Val
Met His Glu Ala Leu His Asn195 200 205His
Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys210 215
22057218PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 57Cys Pro Ala Pro Pro Val Ala Gly
Pro Ser Val Phe Leu Phe Pro Pro1 5 10
15Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val
Thr Cys20 25 30Val Val Val Asp Val Ser
His Glu Asp Pro Glu Val Gln Phe Asn Trp35 40
45Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu50
55 60Glu Gln Phe Asn Ser Thr Phe Arg Val
Val Ser Val Leu Thr Val Val65 70 75
80His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val
Ser Asn85 90 95Lys Gly Leu Pro Ala Pro
Ile Glu Lys Thr Ile Ser Lys Thr Lys Gly100 105
110Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu
Glu115 120 125Met Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu Val Lys Gly Phe Tyr130 135
140Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn145
150 155 160Asn Tyr Lys Thr
Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe Phe165 170
175Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln
Gly Asn180 185 190Val Phe Ser Cys Ser Val
Met His Glu Ala Leu His Asn His Tyr Thr195 200
205Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys210
21558223PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 58Pro Ala Pro Pro Val Ala Gly Pro Ser Val
Phe Leu Phe Pro Pro Lys1 5 10
15Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val20
25 30Val Val Asp Val Ser His Glu Asp Pro
Glu Val Gln Phe Asn Trp Tyr35 40 45Val
Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu50
55 60Gln Phe Asn Ser Thr Phe Arg Val Val Ser Val
Leu Thr Val Val His65 70 75
80Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys85
90 95Gly Leu Pro Ala Pro Ile Glu Lys Thr
Ile Ser Lys Thr Lys Gly Gln100 105 110Pro
Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu Met115
120 125Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val
Lys Gly Phe Tyr Pro130 135 140Ser Asp Ile
Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn145
150 155 160Tyr Lys Thr Thr Pro Pro Met
Leu Asp Ser Asp Gly Ser Phe Phe Leu165 170
175Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val180
185 190Phe Ser Cys Ser Val Met His Glu Ala
Leu His Asn His Tyr Thr Gln195 200 205Lys
Ser Leu Ser Leu Ser Pro Glu Leu Gln Leu Glu Glu Ser Cys210
215 22059234PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 59Pro Ala Pro Pro Val Ala
Gly Pro Ser Val Phe Leu Phe Pro Pro Lys1 5
10 15Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu
Val Thr Cys Val20 25 30Val Val Asp Val
Ser His Glu Asp Pro Glu Val Gln Phe Asn Trp Tyr35 40
45Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg
Glu Glu50 55 60Gln Phe Asn Ser Thr Phe
Arg Val Val Ser Val Leu Thr Val Val His65 70
75 80Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys
Lys Val Ser Asn Lys85 90 95Gly Leu Pro
Ala Pro Ile Glu Lys Thr Ile Ser Lys Thr Lys Gly Gln100
105 110Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser
Arg Glu Glu Met115 120 125Thr Lys Asn Gln
Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro130 135
140Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu
Asn Asn145 150 155 160Tyr
Lys Thr Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe Phe Leu165
170 175Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp
Gln Gln Gly Asn Val180 185 190Phe Ser Cys
Ser Val Met His Glu Ala Leu His Asn His Tyr Thr Gln195
200 205Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln Leu Glu
Glu Ser Xaa Ala210 215 220Glu Ala Gln Asp
Gly Glu Leu Asp Gly Xaa225 23060231PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 60Xaa Xaa Val Glu Xaa Pro Pro Xaa Pro Ala Pro Pro Val Ala
Gly Pro1 5 10 15Ser Val
Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser20
25 30Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val
Ser His Glu Asp35 40 45Pro Glu Val Gln
Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn50 55
60Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Phe
Arg Val65 70 75 80Val
Ser Val Leu Thr Val Val His Gln Asp Trp Leu Asn Gly Lys Glu85
90 95Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu Pro
Ala Pro Ile Glu Lys100 105 110Thr Ile Ser
Lys Thr Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr115
120 125Leu Pro Pro Ser Arg Glu Glu Met Thr Lys Asn Gln
Val Ser Leu Thr130 135 140Cys Leu Val Lys
Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu145 150
155 160Ser Asn Gly Gln Pro Glu Asn Asn Tyr
Lys Thr Thr Pro Pro Met Leu165 170 175Asp
Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys180
185 190Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys
Ser Val Met His Glu195 200 205Ala Leu His
Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Glu210
215 220Leu Gln Leu Glu Glu Ser Cys225
23061230PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 61Xaa Val Glu Xaa Pro Pro Xaa Pro Ala Pro
Pro Val Ala Gly Pro Ser1 5 10
15Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg20
25 30Thr Pro Glu Val Thr Cys Val Val Val
Asp Val Ser His Glu Asp Pro35 40 45Glu
Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala50
55 60Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser
Thr Phe Arg Val Val65 70 75
80Ser Val Leu Thr Val Val His Gln Asp Trp Leu Asn Gly Lys Glu Tyr85
90 95Lys Cys Lys Val Ser Asn Lys Gly Leu
Pro Ala Pro Ile Glu Lys Thr100 105 110Ile
Ser Lys Thr Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu115
120 125Pro Pro Ser Arg Glu Glu Met Thr Lys Asn Gln
Val Ser Leu Thr Cys130 135 140Leu Val Lys
Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser145
150 155 160Asn Gly Gln Pro Glu Asn Asn
Tyr Lys Thr Thr Pro Pro Met Leu Asp165 170
175Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser180
185 190Arg Trp Gln Gln Gly Asn Val Phe Ser
Cys Ser Val Met His Glu Ala195 200 205Leu
His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu210
215 220Gln Leu Glu Glu Ser Cys225
23062227PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 62Xaa Pro Pro Xaa Pro Ala Pro Pro Val Ala
Gly Pro Ser Val Phe Leu1 5 10
15Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu20
25 30Val Thr Cys Val Val Val Asp Val Ser
His Glu Asp Pro Glu Val Gln35 40 45Phe
Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys50
55 60Pro Arg Glu Glu Gln Phe Asn Ser Thr Phe Arg
Val Val Ser Val Leu65 70 75
80Thr Val Val His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys85
90 95Val Ser Asn Lys Gly Leu Pro Ala Pro
Ile Glu Lys Thr Ile Ser Lys100 105 110Thr
Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser115
120 125Arg Glu Glu Met Thr Lys Asn Gln Val Ser Leu
Thr Cys Leu Val Lys130 135 140Gly Phe Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln145
150 155 160Pro Glu Asn Asn Tyr Lys Thr
Thr Pro Pro Met Leu Asp Ser Asp Gly165 170
175Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln180
185 190Gln Gly Asn Val Phe Ser Cys Ser Val
Met His Glu Ala Leu His Asn195 200 205His
Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln Leu Glu210
215 220Glu Ser Cys22563224PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 63Cys Pro Ala Pro Pro Val Ala Gly Pro Ser Val Phe Leu Phe
Pro Pro1 5 10 15Lys Pro
Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys20
25 30Val Val Val Asp Val Ser His Glu Asp Pro Glu Val
Gln Phe Asn Trp35 40 45Tyr Val Asp Gly
Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu50 55
60Glu Gln Phe Asn Ser Thr Phe Arg Val Val Ser Val Leu Thr
Val Val65 70 75 80His
Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn85
90 95Lys Gly Leu Pro Ala Pro Ile Glu Lys Thr Ile
Ser Lys Thr Lys Gly100 105 110Gln Pro Arg
Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu115
120 125Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val
Lys Gly Phe Tyr130 135 140Pro Ser Asp Ile
Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn145 150
155 160Asn Tyr Lys Thr Thr Pro Pro Met Leu
Asp Ser Asp Gly Ser Phe Phe165 170 175Leu
Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn180
185 190Val Phe Ser Cys Ser Val Met His Glu Ala Leu
His Asn His Tyr Thr195 200 205Gln Lys Ser
Leu Ser Leu Ser Pro Glu Leu Gln Leu Glu Glu Ser Cys210
215 22064242PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 64Xaa Xaa Val Glu Xaa Pro
Pro Xaa Pro Ala Pro Pro Val Ala Gly Pro1 5
10 15Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr
Leu Met Ile Ser20 25 30Arg Thr Pro Glu
Val Thr Cys Val Val Val Asp Val Ser His Glu Asp35 40
45Pro Glu Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val
His Asn50 55 60Ala Lys Thr Lys Pro Arg
Glu Glu Gln Phe Asn Ser Thr Phe Arg Val65 70
75 80Val Ser Val Leu Thr Val Val His Gln Asp Trp
Leu Asn Gly Lys Glu85 90 95Tyr Lys Cys
Lys Val Ser Asn Lys Gly Leu Pro Ala Pro Ile Glu Lys100
105 110Thr Ile Ser Lys Thr Lys Gly Gln Pro Arg Glu Pro
Gln Val Tyr Thr115 120 125Leu Pro Pro Ser
Arg Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr130 135
140Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu
Trp Glu145 150 155 160Ser
Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Met Leu165
170 175Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys
Leu Thr Val Asp Lys180 185 190Ser Arg Trp
Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met His Glu195
200 205Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser
Leu Ser Pro Glu210 215 220Leu Gln Leu Glu
Glu Ser Xaa Ala Glu Ala Gln Asp Gly Glu Leu Asp225 230
235 240Gly Xaa65241PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 65Xaa Val Glu Xaa Pro Pro Xaa Pro Ala Pro Pro Val Ala Gly
Pro Ser1 5 10 15Val Phe
Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg20
25 30Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser
His Glu Asp Pro35 40 45Glu Val Gln Phe
Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala50 55
60Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Phe Arg
Val Val65 70 75 80Ser
Val Leu Thr Val Val His Gln Asp Trp Leu Asn Gly Lys Glu Tyr85
90 95Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ala
Pro Ile Glu Lys Thr100 105 110Ile Ser Lys
Thr Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu115
120 125Pro Pro Ser Arg Glu Glu Met Thr Lys Asn Gln Val
Ser Leu Thr Cys130 135 140Leu Val Lys Gly
Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser145 150
155 160Asn Gly Gln Pro Glu Asn Asn Tyr Lys
Thr Thr Pro Pro Met Leu Asp165 170 175Ser
Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser180
185 190Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser
Val Met His Glu Ala195 200 205Leu His Asn
His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu210
215 220Gln Leu Glu Glu Ser Xaa Ala Glu Ala Gln Asp Gly
Glu Leu Asp Gly225 230 235
240Xaa66238PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 66Xaa Pro Pro Xaa Pro Ala Pro Pro Val Ala
Gly Pro Ser Val Phe Leu1 5 10
15Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu20
25 30Val Thr Cys Val Val Val Asp Val Ser
His Glu Asp Pro Glu Val Gln35 40 45Phe
Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys50
55 60Pro Arg Glu Glu Gln Phe Asn Ser Thr Phe Arg
Val Val Ser Val Leu65 70 75
80Thr Val Val His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys85
90 95Val Ser Asn Lys Gly Leu Pro Ala Pro
Ile Glu Lys Thr Ile Ser Lys100 105 110Thr
Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser115
120 125Arg Glu Glu Met Thr Lys Asn Gln Val Ser Leu
Thr Cys Leu Val Lys130 135 140Gly Phe Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln145
150 155 160Pro Glu Asn Asn Tyr Lys Thr
Thr Pro Pro Met Leu Asp Ser Asp Gly165 170
175Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln180
185 190Gln Gly Asn Val Phe Ser Cys Ser Val
Met His Glu Ala Leu His Asn195 200 205His
Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln Leu Glu210
215 220Glu Ser Xaa Ala Glu Ala Gln Asp Gly Glu Leu
Asp Gly Xaa225 230 23567235PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 67Cys Pro Ala Pro Pro Val Ala Gly Pro Ser Val Phe Leu Phe
Pro Pro1 5 10 15Lys Pro
Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys20
25 30Val Val Val Asp Val Ser His Glu Asp Pro Glu Val
Gln Phe Asn Trp35 40 45Tyr Val Asp Gly
Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu50 55
60Glu Gln Phe Asn Ser Thr Phe Arg Val Val Ser Val Leu Thr
Val Val65 70 75 80His
Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn85
90 95Lys Gly Leu Pro Ala Pro Ile Glu Lys Thr Ile
Ser Lys Thr Lys Gly100 105 110Gln Pro Arg
Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu115
120 125Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val
Lys Gly Phe Tyr130 135 140Pro Ser Asp Ile
Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn145 150
155 160Asn Tyr Lys Thr Thr Pro Pro Met Leu
Asp Ser Asp Gly Ser Phe Phe165 170 175Leu
Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn180
185 190Val Phe Ser Cys Ser Val Met His Glu Ala Leu
His Asn His Tyr Thr195 200 205Gln Lys Ser
Leu Ser Leu Ser Pro Glu Leu Gln Leu Glu Glu Ser Xaa210
215 220Ala Glu Ala Gln Asp Gly Glu Leu Asp Gly Xaa225
230 23568902DNAArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 68aagcttgaat
tcccaccatg ctgctgctgg cgagatgtct gctgctagtc ctcgtctcct 60cgctgctggt
atgctcggga ctggcgtgcc cacggtgccc agagcccaaa tcttgtgaca 120cacctccccc
gtgcccacgg tgcccagagc ccaaatcttg tgacacacct cccccatgcc 180cacggtgccc
agagcccaaa tcttgtgaca cacctccccc gtgcccaagg tgcccagcac 240ctgaactcct
gggaggaccg tcagtcttcc tcttcccccc aaaacccaag gataccctta 300tgatttcccg
gacccctgag gtcacgtgcg tggtggtgga cgtgagccac gaagaccccg 360aggtccagtt
caagtggtac gtggacggcg tggaggtgca taatgccaag acaaagccgc 420gggaggagca
gtacaacagc acgttccgtg tggtcagcgt cctcaccgtc ctgcaccagg 480actggctgaa
cggcaaggag tacaagtgca aggtctccaa caaagccctc ccagccccca 540tcgagaaaac
catctccaaa accaaaggac agccccgaga accacaggtg tacaccctgc 600ccccatcccg
ggaggagatg accaagaacc aggtcagcct gacctgcctg gtcaaaggct 660tctaccccag
cgacatcgcc gtggagtggg agagcagcgg gcagccggag aacaactaca 720acaccacgcc
tcccatgctg gactccgacg gctccttctt cctctacagc aagctcaccg 780tggacaagag
caggtggcag caggggaaca tcttctcatg ctccgtgatg catgaggctc 840tgcacaaccg
cttcacgcag aagagcctct ccctgtctcc gggtaaatga ctcgagcggc 900cg
90269902DNAArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 69aagcttgaat tcccaccatg gccttgacct
ttgctttact ggtggccctc ctggtgctca 60gctgcaagtc aagctgctct gtgggctgcc
cacggtgccc agagcccaaa tcttgtgaca 120cacctccccc gtgcccacgg tgcccagagc
ccaaatcttg tgacacacct cccccatgcc 180cacggtgccc agagcccaaa tcttgtgaca
cacctccccc gtgcccaagg tgcccagcac 240ctgaactcct gggaggaccg tcagtcttcc
tcttcccccc aaaacccaag gataccctta 300tgatttcccg gacccctgag gtcacgtgcg
tggtggtgga cgtgagccac gaagaccccg 360aggtccagtt caagtggtac gtggacggcg
tggaggtgca taatgccaag acaaagccgc 420gggaggagca gtacaacagc acgttccgtg
tggtcagcgt cctcaccgtc ctgcaccagg 480actggctgaa cggcaaggag tacaagtgca
aggtctccaa caaagccctc ccagccccca 540tcgagaaaac catctccaaa accaaaggac
agccccgaga accacaggtg tacaccctgc 600ccccatcccg ggaggagatg accaagaacc
aggtcagcct gacctgcctg gtcaaaggct 660tctaccccag cgacatcgcc gtggagtggg
agagcagcgg gcagccggag aacaactaca 720acaccacgcc tcccatgctg gactccgacg
gctccttctt cctctacagc aagctcaccg 780tggacaagag caggtggcag caggggaaca
tcttctcatg ctccgtgatg catgaggctc 840tgcacaaccg cttcacgcag aagagcctct
ccctgtctcc gggtaaatga ctcgagcggc 900cg
90270884DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 70aagcttgaat tcccaccatg ctggctgcca cagtcctgac cctggccctg
ctgggcaatg 60cccatgcctg cccacggtgc ccagagccca aatcttgtga cacacctccc
ccgtgcccac 120ggtgcccaga gcccaaatct tgtgacacac ctcccccatg cccacggtgc
ccagagccca 180aatcttgtga cacacctccc ccgtgcccaa ggtgcccagc acctgaactc
ctgggaggac 240cgtcagtctt cctcttcccc ccaaaaccca aggataccct tatgatttcc
cggacccctg 300aggtcacgtg cgtggtggtg gacgtgagcc acgaagaccc cgaggtccag
ttcaagtggt 360acgtggacgg cgtggaggtg cataatgcca agacaaagcc gcgggaggag
cagtacaaca 420gcacgttccg tgtggtcagc gtcctcaccg tcctgcacca ggactggctg
aacggcaagg 480agtacaagtg caaggtctcc aacaaagccc tcccagcccc catcgagaaa
accatctcca 540aaaccaaagg acagccccga gaaccacagg tgtacaccct gcccccatcc
cgggaggaga 600tgaccaagaa ccaggtcagc ctgacctgcc tggtcaaagg cttctacccc
agcgacatcg 660ccgtggagtg ggagagcagc gggcagccgg agaacaacta caacaccacg
cctcccatgc 720tggactccga cggctccttc ttcctctaca gcaagctcac cgtggacaag
agcaggtggc 780agcaggggaa catcttctca tgctccgtga tgcatgaggc tctgcacaac
cgcttcacgc 840agaagagcct ctccctgtct ccgggtaaat gactcgagcg gccg
88471290PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 71Met Leu Leu Leu Ala Arg Cys Leu
Leu Leu Val Leu Val Ser Ser Leu1 5 10
15Leu Val Cys Ser Gly Leu Ala Cys Pro Arg Cys Pro Glu Pro
Lys Ser20 25 30Cys Asp Thr Pro Pro Pro
Cys Pro Arg Cys Pro Glu Pro Lys Ser Cys35 40
45Asp Thr Pro Pro Pro Cys Pro Arg Cys Pro Glu Pro Lys Ser Cys Asp50
55 60Thr Pro Pro Pro Cys Pro Arg Cys Pro
Ala Pro Glu Leu Leu Gly Gly65 70 75
80Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu
Met Ile85 90 95Ser Arg Thr Pro Glu Val
Thr Cys Val Val Val Asp Val Ser His Glu100 105
110Asp Pro Glu Val Gln Phe Lys Trp Tyr Val Asp Gly Val Glu Val
His115 120 125Asn Ala Lys Thr Lys Pro Arg
Glu Glu Gln Phe Asn Ser Thr Phe Arg130 135
140Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys145
150 155 160Glu Tyr Lys Cys
Lys Val Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu165 170
175Lys Thr Ile Ser Lys Thr Lys Gly Gln Pro Arg Glu Pro Gln
Val Tyr180 185 190Thr Leu Pro Pro Ser Arg
Glu Glu Met Thr Lys Asn Gln Val Ser Leu195 200
205Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu
Trp210 215 220Glu Ser Ser Gly Gln Pro Glu
Asn Asn Tyr Asn Thr Thr Pro Pro Met225 230
235 240Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys
Leu Thr Val Asp245 250 255Lys Ser Arg Trp
Gln Gln Gly Asn Ile Phe Ser Cys Ser Val Met His260 265
270Glu Ala Leu His Asn Arg Phe Thr Gln Lys Ser Leu Ser Leu
Ser Pro275 280 285Gly
Lys29072290PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 72Met Ala Leu Thr Phe Ala Leu Leu Val Ala
Leu Leu Val Leu Ser Cys1 5 10
15Lys Ser Ser Cys Ser Val Gly Cys Pro Arg Cys Pro Glu Pro Lys Ser20
25 30Cys Asp Thr Pro Pro Pro Cys Pro Arg
Cys Pro Glu Pro Lys Ser Cys35 40 45Asp
Thr Pro Pro Pro Cys Pro Arg Cys Pro Glu Pro Lys Ser Cys Asp50
55 60Thr Pro Pro Pro Cys Pro Arg Cys Pro Ala Pro
Glu Leu Leu Gly Gly65 70 75
80Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile85
90 95Ser Arg Thr Pro Glu Val Thr Cys Val
Val Val Asp Val Ser His Glu100 105 110Asp
Pro Glu Val Gln Phe Lys Trp Tyr Val Asp Gly Val Glu Val His115
120 125Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe
Asn Ser Thr Phe Arg130 135 140Val Val Ser
Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys145
150 155 160Glu Tyr Lys Cys Lys Val Ser
Asn Lys Ala Leu Pro Ala Pro Ile Glu165 170
175Lys Thr Ile Ser Lys Thr Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr180
185 190Thr Leu Pro Pro Ser Arg Glu Glu Met
Thr Lys Asn Gln Val Ser Leu195 200 205Thr
Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp210
215 220Glu Ser Ser Gly Gln Pro Glu Asn Asn Tyr Asn
Thr Thr Pro Pro Met225 230 235
240Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val
Asp245 250 255Lys Ser Arg Trp Gln Gln Gly
Asn Ile Phe Ser Cys Ser Val Met His260 265
270Glu Ala Leu His Asn Arg Phe Thr Gln Lys Ser Leu Ser Leu Ser Pro275
280 285Gly Lys29073284PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 73Met Leu Ala Ala Thr Val Leu Thr Leu Ala Leu Leu Gly Asn
Ala His1 5 10 15Ala Cys
Pro Arg Cys Pro Glu Pro Lys Ser Cys Asp Thr Pro Pro Pro20
25 30Cys Pro Arg Cys Pro Glu Pro Lys Ser Cys Asp Thr
Pro Pro Pro Cys35 40 45Pro Arg Cys Pro
Glu Pro Lys Ser Cys Asp Thr Pro Pro Pro Cys Pro50 55
60Arg Cys Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe
Leu Phe65 70 75 80Pro
Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val85
90 95Thr Cys Val Val Val Asp Val Ser His Glu Asp
Pro Glu Val Gln Phe100 105 110Lys Trp Tyr
Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro115
120 125Arg Glu Glu Gln Phe Asn Ser Thr Phe Arg Val Val
Ser Val Leu Thr130 135 140Val Leu His Gln
Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val145 150
155 160Ser Asn Lys Ala Leu Pro Ala Pro Ile
Glu Lys Thr Ile Ser Lys Thr165 170 175Lys
Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg180
185 190Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr
Cys Leu Val Lys Gly195 200 205Phe Tyr Pro
Ser Asp Ile Ala Val Glu Trp Glu Ser Ser Gly Gln Pro210
215 220Glu Asn Asn Tyr Asn Thr Thr Pro Pro Met Leu Asp
Ser Asp Gly Ser225 230 235
240Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln245
250 255Gly Asn Ile Phe Ser Cys Ser Val Met
His Glu Ala Leu His Asn Arg260 265 270Phe
Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys275
28074267PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 74Cys Pro Arg Cys Pro Glu Pro Lys Ser Cys
Asp Thr Pro Pro Pro Cys1 5 10
15Pro Arg Cys Pro Glu Pro Lys Ser Cys Asp Thr Pro Pro Pro Cys Pro20
25 30Arg Cys Pro Glu Pro Lys Ser Cys Asp
Thr Pro Pro Pro Cys Pro Arg35 40 45Cys
Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro50
55 60Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg
Thr Pro Glu Val Thr65 70 75
80Cys Val Val Val Asp Val Ser His Glu Asp Pro Glu Val Gln Phe Lys85
90 95Trp Tyr Val Asp Gly Val Glu Val His
Asn Ala Lys Thr Lys Pro Arg100 105 110Glu
Glu Gln Phe Asn Ser Thr Phe Arg Val Val Ser Val Leu Thr Val115
120 125Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr
Lys Cys Lys Val Ser130 135 140Asn Lys Ala
Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Thr Lys145
150 155 160Gly Gln Pro Arg Glu Pro Gln
Val Tyr Thr Leu Pro Pro Ser Arg Glu165 170
175Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe180
185 190Tyr Pro Ser Asp Ile Ala Val Glu Trp
Glu Ser Ser Gly Gln Pro Glu195 200 205Asn
Asn Tyr Asn Thr Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe210
215 220Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser
Arg Trp Gln Gln Gly225 230 235
240Asn Ile Phe Ser Cys Ser Val Met His Glu Ala Leu His Asn Arg
Phe245 250 255Thr Gln Lys Ser Leu Ser Leu
Ser Pro Gly Lys260 26575267PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 75Xaa Pro Arg Xaa Pro Glu Pro Lys Ser Xaa Asp Thr Pro Pro
Pro Xaa1 5 10 15Pro Arg
Xaa Pro Glu Pro Lys Ser Xaa Asp Thr Pro Pro Pro Xaa Pro20
25 30Arg Xaa Pro Glu Pro Lys Ser Xaa Asp Thr Pro Pro
Pro Xaa Pro Arg35 40 45Xaa Pro Ala Pro
Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro50 55
60Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu
Val Thr65 70 75 80Cys
Val Val Val Asp Val Ser His Glu Asp Pro Glu Val Gln Phe Lys85
90 95Trp Tyr Val Asp Gly Val Glu Val His Asn Ala
Lys Thr Lys Pro Arg100 105 110Glu Glu Gln
Phe Asn Ser Thr Phe Arg Val Val Ser Val Leu Thr Val115
120 125Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys
Cys Lys Val Ser130 135 140Asn Lys Ala Leu
Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Thr Lys145 150
155 160Gly Gln Pro Arg Glu Pro Gln Val Tyr
Thr Leu Pro Pro Ser Arg Glu165 170 175Glu
Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe180
185 190Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser
Ser Gly Gln Pro Glu195 200 205Asn Asn Tyr
Asn Thr Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe210
215 220Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg
Trp Gln Gln Gly225 230 235
240Asn Ile Phe Ser Cys Ser Val Met His Glu Ala Leu His Asn Arg Phe245
250 255Thr Gln Lys Ser Leu Ser Leu Ser Pro
Gly Lys260 26576264PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 76Xaa Pro Glu Pro
Lys Ser Xaa Asp Thr Pro Pro Pro Xaa Pro Arg Xaa1 5
10 15Pro Glu Pro Lys Ser Xaa Asp Thr Pro Pro
Pro Xaa Pro Arg Xaa Pro20 25 30Glu Pro
Lys Ser Xaa Asp Thr Pro Pro Pro Xaa Pro Arg Xaa Pro Ala35
40 45Pro Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe
Pro Pro Lys Pro50 55 60Lys Asp Thr Leu
Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val65 70
75 80Val Asp Val Ser His Glu Asp Pro Glu
Val Gln Phe Lys Trp Tyr Val85 90 95Asp
Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln100
105 110Phe Asn Ser Thr Phe Arg Val Val Ser Val Leu
Thr Val Leu His Gln115 120 125Asp Trp Leu
Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala130
135 140Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Thr
Lys Gly Gln Pro145 150 155
160Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu Met Thr165
170 175Lys Asn Gln Val Ser Leu Thr Cys Leu
Val Lys Gly Phe Tyr Pro Ser180 185 190Asp
Ile Ala Val Glu Trp Glu Ser Ser Gly Gln Pro Glu Asn Asn Tyr195
200 205Asn Thr Thr Pro Pro Met Leu Asp Ser Asp Gly
Ser Phe Phe Leu Tyr210 215 220Ser Lys Leu
Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Ile Phe225
230 235 240Ser Cys Ser Val Met His Glu
Ala Leu His Asn Arg Phe Thr Gln Lys245 250
255Ser Leu Ser Leu Ser Pro Gly Lys26077224PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 77Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe
Pro Pro1 5 10 15Lys Pro
Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys20
25 30Val Val Val Asp Val Ser His Glu Asp Pro Glu Val
Gln Phe Lys Trp35 40 45Tyr Val Asp Gly
Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu50 55
60Glu Gln Phe Asn Ser Thr Phe Arg Val Val Ser Val Leu Thr
Val Leu65 70 75 80His
Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn85
90 95Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile
Ser Lys Thr Lys Gly100 105 110Gln Pro Arg
Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Glu Glu115
120 125Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val
Lys Gly Phe Tyr130 135 140Pro Ser Asp Ile
Ala Val Glu Trp Glu Ser Ser Gly Gln Pro Glu Asn145 150
155 160Asn Tyr Asn Thr Thr Pro Pro Met Leu
Asp Ser Asp Gly Ser Phe Phe165 170 175Leu
Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn180
185 190Ile Phe Ser Cys Ser Val Met His Glu Ala Leu
His Asn Arg Phe Thr195 200 205Gln Lys Ser
Leu Ser Leu Ser Pro Glu Leu Gln Leu Glu Glu Ser Cys210
215 22078235PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 78Pro Ala Pro Glu Leu Leu
Gly Gly Pro Ser Val Phe Leu Phe Pro Pro1 5
10 15Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro
Glu Val Thr Cys20 25 30Val Val Val Asp
Val Ser His Glu Asp Pro Glu Val Gln Phe Lys Trp35 40
45Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro
Arg Glu50 55 60Glu Gln Phe Asn Ser Thr
Phe Arg Val Val Ser Val Leu Thr Val Leu65 70
75 80His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys
Cys Lys Val Ser Asn85 90 95Lys Ala Leu
Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Thr Lys Gly100
105 110Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro
Ser Arg Glu Glu115 120 125Met Thr Lys Asn
Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr130 135
140Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Ser Gly Gln Pro
Glu Asn145 150 155 160Asn
Tyr Asn Thr Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe Phe165
170 175Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg
Trp Gln Gln Gly Asn180 185 190Ile Phe Ser
Cys Ser Val Met His Glu Ala Leu His Asn Arg Phe Thr195
200 205Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln Leu
Glu Glu Ser Xaa210 215 220Ala Glu Ala Gln
Gly Asp Glu Leu Asp Gly Xaa225 230
23579273PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 79Xaa Pro Arg Xaa Pro Glu Pro Lys Ser Xaa
Asp Thr Pro Pro Pro Xaa1 5 10
15Pro Arg Xaa Pro Glu Pro Lys Ser Xaa Asp Thr Pro Pro Pro Xaa Pro20
25 30Arg Xaa Pro Glu Pro Lys Ser Xaa Asp
Thr Pro Pro Pro Xaa Pro Arg35 40 45Xaa
Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro50
55 60Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg
Thr Pro Glu Val Thr65 70 75
80Cys Val Val Val Asp Val Ser His Glu Asp Pro Glu Val Gln Phe Lys85
90 95Trp Tyr Val Asp Gly Val Glu Val His
Asn Ala Lys Thr Lys Pro Arg100 105 110Glu
Glu Gln Phe Asn Ser Thr Phe Arg Val Val Ser Val Leu Thr Val115
120 125Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr
Lys Cys Lys Val Ser130 135 140Asn Lys Ala
Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Thr Lys145
150 155 160Gly Gln Pro Arg Glu Pro Gln
Val Tyr Thr Leu Pro Pro Ser Arg Glu165 170
175Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe180
185 190Tyr Pro Ser Asp Ile Ala Val Glu Trp
Glu Ser Ser Gly Gln Pro Glu195 200 205Asn
Asn Tyr Asn Thr Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe210
215 220Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser
Arg Trp Gln Gln Gly225 230 235
240Asn Ile Phe Ser Cys Ser Val Met His Glu Ala Leu His Asn Arg
Phe245 250 255Thr Gln Lys Ser Leu Ser Leu
Ser Pro Glu Leu Gln Leu Glu Glu Ser260 265
270Cys80270PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 80Xaa Pro Glu Pro Lys Ser Xaa Asp Thr Pro
Pro Pro Xaa Pro Arg Xaa1 5 10
15Pro Glu Pro Lys Ser Xaa Asp Thr Pro Pro Pro Xaa Pro Arg Xaa Pro20
25 30Glu Pro Lys Ser Xaa Asp Thr Pro Pro
Pro Xaa Pro Arg Xaa Pro Ala35 40 45Pro
Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro50
55 60Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu
Val Thr Cys Val Val65 70 75
80Val Asp Val Ser His Glu Asp Pro Glu Val Gln Phe Lys Trp Tyr Val85
90 95Asp Gly Val Glu Val His Asn Ala Lys
Thr Lys Pro Arg Glu Glu Gln100 105 110Phe
Asn Ser Thr Phe Arg Val Val Ser Val Leu Thr Val Leu His Gln115
120 125Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys
Val Ser Asn Lys Ala130 135 140Leu Pro Ala
Pro Ile Glu Lys Thr Ile Ser Lys Thr Lys Gly Gln Pro145
150 155 160Arg Glu Pro Gln Val Tyr Thr
Leu Pro Pro Ser Arg Glu Glu Met Thr165 170
175Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser180
185 190Asp Ile Ala Val Glu Trp Glu Ser Ser
Gly Gln Pro Glu Asn Asn Tyr195 200 205Asn
Thr Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr210
215 220Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln
Gln Gly Asn Ile Phe225 230 235
240Ser Cys Ser Val Met His Glu Ala Leu His Asn Arg Phe Thr Gln
Lys245 250 255Ser Leu Ser Leu Ser Pro Glu
Leu Gln Leu Glu Glu Ser Cys260 265
27081284PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 81Xaa Pro Arg Xaa Pro Glu Pro Lys Ser Xaa
Asp Thr Pro Pro Pro Xaa1 5 10
15Pro Arg Xaa Pro Glu Pro Lys Ser Xaa Asp Thr Pro Pro Pro Xaa Pro20
25 30Arg Xaa Pro Glu Pro Lys Ser Xaa Asp
Thr Pro Pro Pro Xaa Pro Arg35 40 45Xaa
Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro50
55 60Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg
Thr Pro Glu Val Thr65 70 75
80Cys Val Val Val Asp Val Ser His Glu Asp Pro Glu Val Gln Phe Lys85
90 95Trp Tyr Val Asp Gly Val Glu Val His
Asn Ala Lys Thr Lys Pro Arg100 105 110Glu
Glu Gln Phe Asn Ser Thr Phe Arg Val Val Ser Val Leu Thr Val115
120 125Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr
Lys Cys Lys Val Ser130 135 140Asn Lys Ala
Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Thr Lys145
150 155 160Gly Gln Pro Arg Glu Pro Gln
Val Tyr Thr Leu Pro Pro Ser Arg Glu165 170
175Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe180
185 190Tyr Pro Ser Asp Ile Ala Val Glu Trp
Glu Ser Ser Gly Gln Pro Glu195 200 205Asn
Asn Tyr Asn Thr Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe210
215 220Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser
Arg Trp Gln Gln Gly225 230 235
240Asn Ile Phe Ser Cys Ser Val Met His Glu Ala Leu His Asn Arg
Phe245 250 255Thr Gln Lys Ser Leu Ser Leu
Ser Pro Glu Leu Gln Leu Glu Glu Ser260 265
270Xaa Ala Glu Ala Gln Asp Gly Glu Leu Asp Gly Xaa275
28082281PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 82Xaa Pro Glu Pro Lys Ser Xaa Asp Thr Pro
Pro Pro Xaa Pro Arg Xaa1 5 10
15Pro Glu Pro Lys Ser Xaa Asp Thr Pro Pro Pro Xaa Pro Arg Xaa Pro20
25 30Glu Pro Lys Ser Xaa Asp Thr Pro Pro
Pro Xaa Pro Arg Xaa Pro Ala35 40 45Pro
Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro50
55 60Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu
Val Thr Cys Val Val65 70 75
80Val Asp Val Ser His Glu Asp Pro Glu Val Gln Phe Lys Trp Tyr Val85
90 95Asp Gly Val Glu Val His Asn Ala Lys
Thr Lys Pro Arg Glu Glu Gln100 105 110Phe
Asn Ser Thr Phe Arg Val Val Ser Val Leu Thr Val Leu His Gln115
120 125Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys
Val Ser Asn Lys Ala130 135 140Leu Pro Ala
Pro Ile Glu Lys Thr Ile Ser Lys Thr Lys Gly Gln Pro145
150 155 160Arg Glu Pro Gln Val Tyr Thr
Leu Pro Pro Ser Arg Glu Glu Met Thr165 170
175Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser180
185 190Asp Ile Ala Val Glu Trp Glu Ser Ser
Gly Gln Pro Glu Asn Asn Tyr195 200 205Asn
Thr Thr Pro Pro Met Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr210
215 220Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln
Gln Gly Asn Ile Phe225 230 235
240Ser Cys Ser Val Met His Glu Ala Leu His Asn Arg Phe Thr Gln
Lys245 250 255Ser Leu Ser Leu Ser Pro Glu
Leu Gln Leu Glu Glu Ser Xaa Ala Glu260 265
270Ala Gln Asp Gly Glu Leu Asp Gly Xaa275
28083767DNAArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 83aagcttgaat tcccaccatg ctgctgctgg
cgagatgtct gctgctagtc ctcgtctcct 60cgctgctggt atgctcggga ctggcgtgcc
catcatgccc agcacctgag ttcctggggg 120gaccatcagt cttcctgttc cccccaaaac
ccaaggacac tctcatgatc tcccggaccc 180ctgaggtcac gtgcgtggtg gtggacgtga
gccaggaaga ccccgaggtc cagttcaact 240ggtacgtgga tggcgtggag gtgcataatg
ccaagacaaa gccgcgggag gagcagttca 300acagcacgta ccgtgtggtc agcgtcctca
ccgtcctgca ccaggactgg ctgaacggca 360aggagtacaa gtgcaaggtc tccaacaaag
gcctcccgtc ctccatcgag aaaaccatct 420ccaaagccaa agggcagccc cgagagccac
aggtgtacac cctgccccca tcccaggagg 480agatgaccaa gaaccaggtc agcctgacct
gcctggtcaa aggcttctac cccagcgaca 540tcgccgtgga gtgggagagc aatgggcagc
cggagaacaa ctacaagacc acgcctcccg 600tgctggactc cgacggctcc ttcttcctct
acagcaggct aaccgtggac aagagcaggt 660ggcaggaggg gaatgtcttc tcatgctccg
tgatgcatga ggctctgcac aaccactaca 720cacagaagag cctctccctg tctctgggta
aatgactcga gcggccg 76784767DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 84aagcttgaat tcccaccatg gccttgacct ttgctttact ggtggccctc
ctggtgctca 60gctgcaagtc aagctgctct gtgggctgcc catcatgccc agcacctgag
ttcctggggg 120gaccatcagt cttcctgttc cccccaaaac ccaaggacac tctcatgatc
tcccggaccc 180ctgaggtcac gtgcgtggtg gtggacgtga gccaggaaga ccccgaggtc
cagttcaact 240ggtacgtgga tggcgtggag gtgcataatg ccaagacaaa gccgcgggag
gagcagttca 300acagcacgta ccgtgtggtc agcgtcctca ccgtcctgca ccaggactgg
ctgaacggca 360aggagtacaa gtgcaaggtc tccaacaaag gcctcccgtc ctccatcgag
aaaaccatct 420ccaaagccaa agggcagccc cgagagccac aggtgtacac cctgccccca
tcccaggagg 480agatgaccaa gaaccaggtc agcctgacct gcctggtcaa aggcttctac
cccagcgaca 540tcgccgtgga gtgggagagc aatgggcagc cggagaacaa ctacaagacc
acgcctcccg 600tgctggactc cgacggctcc ttcttcctct acagcaggct aaccgtggac
aagagcaggt 660ggcaggaggg gaatgtcttc tcatgctccg tgatgcatga ggctctgcac
aaccactaca 720cacagaagag cctctccctg tctctgggta aatgactcga gcggccg
76785749DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 85aagcttgaat tcccaccatg ctggctgcca
cagtcctgac cctggccctg ctgggcaatg 60cccatgcctg cccatcatgc ccagcacctg
agttcctggg gggaccatca gtcttcctgt 120tccccccaaa acccaaggac actctcatga
tctcccggac ccctgaggtc acgtgcgtgg 180tggtggacgt gagccaggaa gaccccgagg
tccagttcaa ctggtacgtg gatggcgtgg 240aggtgcataa tgccaagaca aagccgcggg
aggagcagtt caacagcacg taccgtgtgg 300tcagcgtcct caccgtcctg caccaggact
ggctgaacgg caaggagtac aagtgcaagg 360tctccaacaa aggcctcccg tcctccatcg
agaaaaccat ctccaaagcc aaagggcagc 420cccgagagcc acaggtgtac accctgcccc
catcccagga ggagatgacc aagaaccagg 480tcagcctgac ctgcctggtc aaaggcttct
accccagcga catcgccgtg gagtgggaga 540gcaatgggca gccggagaac aactacaaga
ccacgcctcc cgtgctggac tccgacggct 600ccttcttcct ctacagcagg ctaaccgtgg
acaagagcag gtggcaggag gggaatgtct 660tctcatgctc cgtgatgcat gaggctctgc
acaaccacta cacacagaag agcctctccc 720tgtctctggg taaatgactc gagcggccg
74986245PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 86Met Leu Leu Leu Ala Arg Cys Leu Leu Leu Val Leu Val Ser
Ser Leu1 5 10 15Leu Val
Cys Ser Gly Leu Ala Cys Pro Ser Cys Pro Ala Pro Glu Phe20
25 30Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys
Pro Lys Asp Thr35 40 45Leu Met Ile Ser
Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val50 55
60Ser Gln Glu Asp Pro Glu Val Gln Phe Asn Trp Tyr Val Asp
Gly Val65 70 75 80Glu
Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser85
90 95Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu
His Gln Asp Trp Leu100 105 110Asn Gly Lys
Glu Tyr Lys Cys Lys Val Ser Asn Lys Gly Leu Pro Ser115
120 125Ser Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln
Pro Arg Glu Pro130 135 140Gln Val Tyr Thr
Leu Pro Pro Ser Gln Glu Glu Met Thr Lys Asn Gln145 150
155 160Val Ser Leu Thr Cys Leu Val Lys Gly
Phe Tyr Pro Ser Asp Ile Ala165 170 175Val
Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr180
185 190Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe
Leu Tyr Ser Arg Leu195 200 205Thr Val Asp
Lys Ser Arg Trp Gln Glu Gly Asn Val Phe Ser Cys Ser210
215 220Val Met His Glu Ala Leu His Asn His Tyr Thr Gln
Lys Ser Leu Ser225 230 235
240Leu Ser Leu Gly Lys24587245PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 87Met Ala Leu Thr Phe Ala
Leu Leu Val Ala Leu Leu Val Leu Ser Cys1 5
10 15Lys Ser Ser Cys Ser Val Gly Cys Pro Ser Cys Pro
Ala Pro Glu Phe20 25 30Leu Gly Gly Pro
Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr35 40
45Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val
Asp Val50 55 60Ser Gln Glu Asp Pro Glu
Val Gln Phe Asn Trp Tyr Val Asp Gly Val65 70
75 80Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu
Glu Gln Phe Asn Ser85 90 95Thr Tyr Arg
Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu100
105 110Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys
Gly Leu Pro Ser115 120 125Ser Ile Glu Lys
Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro130 135
140Gln Val Tyr Thr Leu Pro Pro Ser Gln Glu Glu Met Thr Lys
Asn Gln145 150 155 160Val
Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala165
170 175Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn
Asn Tyr Lys Thr Thr180 185 190Pro Pro Val
Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Arg Leu195
200 205Thr Val Asp Lys Ser Arg Trp Gln Glu Gly Asn Val
Phe Ser Cys Ser210 215 220Val Met His Glu
Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser225 230
235 240Leu Ser Leu Gly
Lys24588239PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 88Met Leu Ala Ala Thr Val Leu Thr Leu Ala
Leu Leu Gly Asn Ala His1 5 10
15Ala Cys Pro Ser Cys Pro Ala Pro Glu Phe Leu Gly Gly Pro Ser Val20
25 30Phe Leu Phe Pro Pro Lys Pro Lys Asp
Thr Leu Met Ile Ser Arg Thr35 40 45Pro
Glu Val Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro Glu50
55 60Val Gln Phe Asn Trp Tyr Val Asp Gly Val Glu
Val His Asn Ala Lys65 70 75
80Thr Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val Ser85
90 95Val Leu Thr Val Leu His Gln Asp Trp
Leu Asn Gly Lys Glu Tyr Lys100 105 110Cys
Lys Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu Lys Thr Ile115
120 125Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln
Val Tyr Thr Leu Pro130 135 140Pro Ser Gln
Glu Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu145
150 155 160Val Lys Gly Phe Tyr Pro Ser
Asp Ile Ala Val Glu Trp Glu Ser Asn165 170
175Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser180
185 190Asp Gly Ser Phe Phe Leu Tyr Ser Arg
Leu Thr Val Asp Lys Ser Arg195 200 205Trp
Gln Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu210
215 220His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu
Ser Leu Gly Lys225 230
23589222PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 89Cys Pro Ser Cys Pro Ala Pro Glu Phe Leu
Gly Gly Pro Ser Val Phe1 5 10
15Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro20
25 30Glu Val Thr Cys Val Val Val Asp Val
Ser Gln Glu Asp Pro Glu Val35 40 45Gln
Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr50
55 60Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr
Arg Val Val Ser Val65 70 75
80Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys85
90 95Lys Val Ser Asn Lys Gly Leu Pro Ser
Ser Ile Glu Lys Thr Ile Ser100 105 110Lys
Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro115
120 125Ser Gln Glu Glu Met Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu Val130 135 140Lys Gly Phe
Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly145
150 155 160Gln Pro Glu Asn Asn Tyr Lys
Thr Thr Pro Pro Val Leu Asp Ser Asp165 170
175Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser Arg Trp180
185 190Gln Glu Gly Asn Val Phe Ser Cys Ser
Val Met His Glu Ala Leu His195 200 205Asn
His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu Gly Lys210
215 22090222PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 90Xaa Pro Ser Xaa Pro Ala
Pro Glu Phe Leu Gly Gly Pro Ser Val Phe1 5
10 15Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile
Ser Arg Thr Pro20 25 30Glu Val Thr Cys
Val Val Val Asp Val Ser Gln Glu Asp Pro Glu Val35 40
45Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala
Lys Thr50 55 60Lys Pro Arg Glu Glu Gln
Phe Asn Ser Thr Tyr Arg Val Val Ser Val65 70
75 80Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly
Lys Glu Tyr Lys Cys85 90 95Lys Val Ser
Asn Lys Gly Leu Pro Ser Ser Ile Glu Lys Thr Ile Ser100
105 110Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr
Thr Leu Pro Pro115 120 125Ser Gln Glu Glu
Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val130 135
140Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser
Asn Gly145 150 155 160Gln
Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser Asp165
170 175Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val
Asp Lys Ser Arg Trp180 185 190Gln Glu Gly
Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu His195
200 205Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu
Gly Lys210 215 22091219PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 91Cys Pro Ala Pro Glu Phe Leu Gly Gly Pro Ser Val Phe Leu
Phe Pro1 5 10 15Pro Lys
Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr20
25 30Cys Val Val Val Asp Val Ser Gln Glu Asp Pro Glu
Val Gln Phe Asn35 40 45Trp Tyr Val Asp
Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg50 55
60Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val Ser Val Leu
Thr Val65 70 75 80Leu
His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser85
90 95Asn Lys Gly Leu Pro Ser Ser Ile Glu Lys Thr
Ile Ser Lys Ala Lys100 105 110Gly Gln Pro
Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Gln Glu115
120 125Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu
Val Lys Gly Phe130 135 140Tyr Pro Ser Asp
Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu145 150
155 160Asn Asn Tyr Lys Thr Thr Pro Pro Val
Leu Asp Ser Asp Gly Ser Phe165 170 175Phe
Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser Arg Trp Gln Glu Gly180
185 190Asn Val Phe Ser Cys Ser Val Met His Glu Ala
Leu His Asn His Tyr195 200 205Thr Gln Lys
Ser Leu Ser Leu Ser Leu Gly Lys210 21592224PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 92Pro Ala Pro Glu Phe Leu Gly Gly Pro Ser Val Phe Leu Phe
Pro Pro1 5 10 15Lys Pro
Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys20
25 30Val Val Val Asp Val Ser Gln Glu Asp Pro Glu Val
Gln Phe Asn Trp35 40 45Tyr Val Asp Gly
Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu50 55
60Glu Gln Phe Asn Ser Thr Tyr Arg Val Val Ser Val Leu Thr
Val Leu65 70 75 80His
Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn85
90 95Lys Gly Leu Pro Ser Ser Ile Glu Lys Thr Ile
Ser Lys Ala Lys Gly100 105 110Gln Pro Arg
Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Gln Glu Glu115
120 125Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val
Lys Gly Phe Tyr130 135 140Pro Ser Asp Ile
Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn145 150
155 160Asn Tyr Lys Thr Thr Pro Pro Val Leu
Asp Ser Asp Gly Ser Phe Phe165 170 175Leu
Tyr Ser Arg Leu Thr Val Asp Lys Ser Arg Trp Gln Glu Gly Asn180
185 190Val Phe Ser Cys Ser Val Met His Glu Ala Leu
His Asn His Tyr Thr195 200 205Gln Lys Ser
Leu Ser Leu Ser Leu Glu Leu Gln Leu Glu Glu Ser Cys210
215 22093235PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 93Pro Ala Pro Glu Phe Leu
Gly Gly Pro Ser Val Phe Leu Phe Pro Pro1 5
10 15Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro
Glu Val Thr Cys20 25 30Val Val Val Asp
Val Ser Gln Glu Asp Pro Glu Val Gln Phe Asn Trp35 40
45Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro
Arg Glu50 55 60Glu Gln Phe Asn Ser Thr
Tyr Arg Val Val Ser Val Leu Thr Val Leu65 70
75 80His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys
Cys Lys Val Ser Asn85 90 95Lys Gly Leu
Pro Ser Ser Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly100
105 110Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro
Ser Gln Glu Glu115 120 125Met Thr Lys Asn
Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr130 135
140Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro
Glu Asn145 150 155 160Asn
Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe165
170 175Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser Arg
Trp Gln Glu Gly Asn180 185 190Val Phe Ser
Cys Ser Val Met His Glu Ala Leu His Asn His Tyr Thr195
200 205Gln Lys Ser Leu Ser Leu Ser Leu Glu Leu Gln Leu
Glu Glu Ser Xaa210 215 220Ala Glu Ala Gln
Asp Gly Glu Leu Asp Gly Xaa225 230
23594228PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 94Xaa Pro Ser Xaa Pro Ala Pro Glu Phe Leu
Gly Gly Pro Ser Val Phe1 5 10
15Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro20
25 30Glu Val Thr Cys Val Val Val Asp Val
Ser Gln Glu Asp Pro Glu Val35 40 45Gln
Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr50
55 60Lys Pro Arg Glu Glu Gln Phe Asn Ser Thr Tyr
Arg Val Val Ser Val65 70 75
80Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys85
90 95Lys Val Ser Asn Lys Gly Leu Pro Ser
Ser Ile Glu Lys Thr Ile Ser100 105 110Lys
Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro115
120 125Ser Gln Glu Glu Met Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu Val130 135 140Lys Gly Phe
Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly145
150 155 160Gln Pro Glu Asn Asn Tyr Lys
Thr Thr Pro Pro Val Leu Asp Ser Asp165 170
175Gly Ser Phe Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser Arg Trp180
185 190Gln Glu Gly Asn Val Phe Ser Cys Ser
Val Met His Glu Ala Leu His195 200 205Asn
His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Leu Glu Leu Gln Leu210
215 220Glu Glu Ser Cys22595225PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 95Cys Pro Ala Pro Glu Phe Leu Gly Gly Pro Ser Val Phe Leu
Phe Pro1 5 10 15Pro Lys
Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr20
25 30Cys Val Val Val Asp Val Ser Gln Glu Asp Pro Glu
Val Gln Phe Asn35 40 45Trp Tyr Val Asp
Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg50 55
60Glu Glu Gln Phe Asn Ser Thr Tyr Arg Val Val Ser Val Leu
Thr Val65 70 75 80Leu
His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser85
90 95Asn Lys Gly Leu Pro Ser Ser Ile Glu Lys Thr
Ile Ser Lys Ala Lys100 105 110Gly Gln Pro
Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Gln Glu115
120 125Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu
Val Lys Gly Phe130 135 140Tyr Pro Ser Asp
Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu145 150
155 160Asn Asn Tyr Lys Thr Thr Pro Pro Val
Leu Asp Ser Asp Gly Ser Phe165 170 175Phe
Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser Arg Trp Gln Glu Gly180
185 190Asn Val Phe Ser Cys Ser Val Met His Glu Ala
Leu His Asn His Tyr195 200 205Thr Gln Lys
Ser Leu Ser Leu Ser Leu Glu Leu Gln Leu Glu Glu Ser210
215 220Cys22596239PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 96Xaa Pro Ser Xaa
Pro Ala Pro Glu Phe Leu Gly Gly Pro Ser Val Phe1 5
10 15Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu
Met Ile Ser Arg Thr Pro20 25 30Glu Val
Thr Cys Val Val Val Asp Val Ser Gln Glu Asp Pro Glu Val35
40 45Gln Phe Asn Trp Tyr Val Asp Gly Val Glu Val His
Asn Ala Lys Thr50 55 60Lys Pro Arg Glu
Glu Gln Phe Asn Ser Thr Tyr Arg Val Val Ser Val65 70
75 80Leu Thr Val Leu His Gln Asp Trp Leu
Asn Gly Lys Glu Tyr Lys Cys85 90 95Lys
Val Ser Asn Lys Gly Leu Pro Ser Ser Ile Glu Lys Thr Ile Ser100
105 110Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val
Tyr Thr Leu Pro Pro115 120 125Ser Gln Glu
Glu Met Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val130
135 140Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp
Glu Ser Asn Gly145 150 155
160Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser Asp165
170 175Gly Ser Phe Phe Leu Tyr Ser Arg Leu
Thr Val Asp Lys Ser Arg Trp180 185 190Gln
Glu Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu His195
200 205Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser
Leu Glu Leu Gln Leu210 215 220Glu Glu Ser
Xaa Ala Glu Ala Gln Asp Gly Glu Leu Asp Gly Xaa225 230
23597236PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 97Cys Pro Ala Pro Glu Phe Leu Gly
Gly Pro Ser Val Phe Leu Phe Pro1 5 10
15Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu
Val Thr20 25 30Cys Val Val Val Asp Val
Ser Gln Glu Asp Pro Glu Val Gln Phe Asn35 40
45Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg50
55 60Glu Glu Gln Phe Asn Ser Thr Tyr Arg
Val Val Ser Val Leu Thr Val65 70 75
80Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys
Val Ser85 90 95Asn Lys Gly Leu Pro Ser
Ser Ile Glu Lys Thr Ile Ser Lys Ala Lys100 105
110Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Gln
Glu115 120 125Glu Met Thr Lys Asn Gln Val
Ser Leu Thr Cys Leu Val Lys Gly Phe130 135
140Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu145
150 155 160Asn Asn Tyr Lys
Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe165 170
175Phe Leu Tyr Ser Arg Leu Thr Val Asp Lys Ser Arg Trp Gln
Glu Gly180 185 190Asn Val Phe Ser Cys Ser
Val Met His Glu Ala Leu His Asn His Tyr195 200
205Thr Gln Lys Ser Leu Ser Leu Ser Leu Glu Leu Gln Leu Glu Glu
Ser210 215 220Xaa Ala Glu Ala Gln Asp Gly
Glu Leu Asp Gly Xaa225 230
23598767DNAArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 98aagcttgaat tcccaccatg ctgctgctgg
cgagatgtct gctgctagtc ctcgtctcct 60cgctgctggt atgctcggga ctggcgtgcc
caccgtgccc agcacctgaa ctcctggggg 120ggccctcagt cttcctcttc cccccaaaac
ccaaggacac cctcatgatc tcccggaccc 180ctgaggtcac atgcgtggtg gtggacgtga
gccacgaaga ccctgaggtc aagttcaact 240ggtacgtgga cggcgtggag gtgcataatg
ccaagacaaa gccgcgggag gagcagtaca 300acagcacgta ccgtgtggtc agcgtcctca
ccgtcctgca ccaggactgg ctgaatggca 360aggagtacaa gtgcaaggtc tccaacaaag
ccctcccagc ccccatcgag aaaaccatct 420ccaaagccaa agggcagccc cgagaaccac
aggtgtacac cctgccccca tcccgggatg 480agctgaccaa gaaccaggtc agcctgacct
gcctggtcaa aggcttctat cccagcgaca 540tcgccgtgga gtgggagagc aatgggcagc
cggagaacaa ctacaagacc acgcctcccg 600tgctggactc cgacggctcc ttcttcctct
acagcaagct caccgtggac aagagcaggt 660ggcagcaggg gaacgtcttc tcatgctccg
tgatgcatga ggctctgcac aaccactaca 720cgcagaagag cctctccctg tctccgggta
aatgactcga gcggccg 76799767DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 99aagcttgaat tcccaccatg gccttgacct ttgctttact ggtggccctc
ctggtgctca 60gctgcaagtc aagctgctct gtgggctgcc caccgtgccc agcacctgaa
ctcctggggg 120ggccctcagt cttcctcttc cccccaaaac ccaaggacac cctcatgatc
tcccggaccc 180ctgaggtcac atgcgtggtg gtggacgtga gccacgaaga ccctgaggtc
aagttcaact 240ggtacgtgga cggcgtggag gtgcataatg ccaagacaaa gccgcgggag
gagcagtaca 300acagcacgta ccgtgtggtc agcgtcctca ccgtcctgca ccaggactgg
ctgaatggca 360aggagtacaa gtgcaaggtc tccaacaaag ccctcccagc ccccatcgag
aaaaccatct 420ccaaagccaa agggcagccc cgagaaccac aggtgtacac cctgccccca
tcccgggatg 480agctgaccaa gaaccaggtc agcctgacct gcctggtcaa aggcttctat
cccagcgaca 540tcgccgtgga gtgggagagc aatgggcagc cggagaacaa ctacaagacc
acgcctcccg 600tgctggactc cgacggctcc ttcttcctct acagcaagct caccgtggac
aagagcaggt 660ggcagcaggg gaacgtcttc tcatgctccg tgatgcatga ggctctgcac
aaccactaca 720cgcagaagag cctctccctg tctccgggta aatgactcga gcggccg
767100749DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 100aagcttgaat tcccaccatg ctggctgcca
cagtcctgac cctggccctg ctgggcaatg 60cccatgcctg cccaccgtgc ccagcacctg
aactcctggg ggggccctca gtcttcctct 120tccccccaaa acccaaggac accctcatga
tctcccggac ccctgaggtc acatgcgtgg 180tggtggacgt gagccacgaa gaccctgagg
tcaagttcaa ctggtacgtg gacggcgtgg 240aggtgcataa tgccaagaca aagccgcggg
aggagcagta caacagcacg taccgtgtgg 300tcagcgtcct caccgtcctg caccaggact
ggctgaatgg caaggagtac aagtgcaagg 360tctccaacaa agccctccca gcccccatcg
agaaaaccat ctccaaagcc aaagggcagc 420cccgagaacc acaggtgtac accctgcccc
catcccggga tgagctgacc aagaaccagg 480tcagcctgac ctgcctggtc aaaggcttct
atcccagcga catcgccgtg gagtgggaga 540gcaatgggca gccggagaac aactacaaga
ccacgcctcc cgtgctggac tccgacggct 600ccttcttcct ctacagcaag ctcaccgtgg
acaagagcag gtggcagcag gggaacgtct 660tctcatgctc cgtgatgcat gaggctctgc
acaaccacta cacgcagaag agcctctccc 720tgtctccggg taaatgactc gagcggccg
749101245PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 101Met Leu Leu Leu Ala Arg Cys Leu Leu Leu Val Leu Val Ser
Ser Leu1 5 10 15Leu Val
Cys Ser Gly Leu Ala Cys Pro Pro Cys Pro Ala Pro Glu Leu20
25 30Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys
Pro Lys Asp Thr35 40 45Leu Met Ile Ser
Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val50 55
60Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp
Gly Val65 70 75 80Glu
Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser85
90 95Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu
His Gln Asp Trp Leu100 105 110Asn Gly Lys
Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala115
120 125Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln
Pro Arg Glu Pro130 135 140Gln Val Tyr Thr
Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln145 150
155 160Val Ser Leu Thr Cys Leu Val Lys Gly
Phe Tyr Pro Ser Asp Ile Ala165 170 175Val
Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr180
185 190Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe
Leu Tyr Ser Lys Leu195 200 205Thr Val Asp
Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser210
215 220Val Met His Glu Ala Leu His Asn His Tyr Thr Gln
Lys Ser Leu Ser225 230 235
240Leu Ser Pro Gly Lys245102245PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 102Met Ala Leu
Thr Phe Ala Leu Leu Val Ala Leu Leu Val Leu Ser Cys1 5
10 15Lys Ser Ser Cys Ser Val Gly Cys Pro
Pro Cys Pro Ala Pro Glu Leu20 25 30Leu
Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr35
40 45Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys
Val Val Val Asp Val50 55 60Ser His Glu
Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val65 70
75 80Glu Val His Asn Ala Lys Thr Lys
Pro Arg Glu Glu Gln Tyr Asn Ser85 90
95Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu100
105 110Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser
Asn Lys Ala Leu Pro Ala115 120 125Pro Ile
Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro130
135 140Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu
Thr Lys Asn Gln145 150 155
160Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala165
170 175Val Glu Trp Glu Ser Asn Gly Gln Pro
Glu Asn Asn Tyr Lys Thr Thr180 185 190Pro
Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu195
200 205Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn
Val Phe Ser Cys Ser210 215 220Val Met His
Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser225
230 235 240Leu Ser Pro Gly
Lys245103239PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 103Met Leu Ala Ala Thr Val Leu Thr Leu Ala
Leu Leu Gly Asn Ala His1 5 10
15Ala Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val20
25 30Phe Leu Phe Pro Pro Lys Pro Lys Asp
Thr Leu Met Ile Ser Arg Thr35 40 45Pro
Glu Val Thr Cys Val Val Val Asp Val Ser His Glu Asp Pro Glu50
55 60Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu
Val His Asn Ala Lys65 70 75
80Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val Val Ser85
90 95Val Leu Thr Val Leu His Gln Asp Trp
Leu Asn Gly Lys Glu Tyr Lys100 105 110Cys
Lys Val Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile115
120 125Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln
Val Tyr Thr Leu Pro130 135 140Pro Ser Arg
Asp Glu Leu Thr Lys Asn Gln Val Ser Leu Thr Cys Leu145
150 155 160Val Lys Gly Phe Tyr Pro Ser
Asp Ile Ala Val Glu Trp Glu Ser Asn165 170
175Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser180
185 190Asp Gly Ser Phe Phe Leu Tyr Ser Lys
Leu Thr Val Asp Lys Ser Arg195 200 205Trp
Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu210
215 220His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu
Ser Pro Gly Lys225 230
235104222PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 104Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu
Gly Gly Pro Ser Val Phe1 5 10
15Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro20
25 30Glu Val Thr Cys Val Val Val Asp Val
Ser His Glu Asp Pro Glu Val35 40 45Lys
Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr50
55 60Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr
Arg Val Val Ser Val65 70 75
80Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys85
90 95Lys Val Ser Asn Lys Ala Leu Pro Ala
Pro Ile Glu Lys Thr Ile Ser100 105 110Lys
Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro115
120 125Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu Val130 135 140Lys Gly Phe
Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly145
150 155 160Gln Pro Glu Asn Asn Tyr Lys
Thr Thr Pro Pro Val Leu Asp Ser Asp165 170
175Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp180
185 190Gln Gln Gly Asn Val Phe Ser Cys Ser
Val Met His Glu Ala Leu His195 200 205Asn
His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys210
215 220105228PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 105Xaa Asp Lys Thr His Thr
Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu1 5
10 15Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro
Lys Asp Thr Leu20 25 30Met Ile Ser Arg
Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser35 40
45His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly
Val Glu50 55 60Val His Asn Ala Lys Thr
Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr65 70
75 80Tyr Arg Val Val Ser Val Leu Thr Val Leu His
Gln Asp Trp Leu Asn85 90 95Gly Lys Glu
Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro100
105 110Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro
Arg Glu Pro Gln115 120 125Val Tyr Thr Leu
Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln Val130 135
140Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile
Ala Val145 150 155 160Glu
Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro165
170 175Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu
Tyr Ser Lys Leu Thr180 185 190Val Asp Lys
Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val195
200 205Met His Glu Ala Leu His Asn His Tyr Thr Gln Lys
Ser Leu Ser Leu210 215 220Ser Pro Gly
Lys225106758DNAArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 106aagcttgaat tcccaccatg ctgctgctgg
cgagatgtct gctgctagtc ctcgtctcct 60cgctgctggt atgctcggga ctggcgtgcc
cagcacctga actcctgggg gggccctcag 120tcttcctctt ccccccaaaa cccaaggaca
ccctcatgat ctcccggacc cctgaggtca 180catgcgtggt ggtggacgtg agccacgaag
accctgaggt caagttcaac tggtacgtgg 240acggcgtgga ggtgcataat gccaagacaa
agccgcggga ggagcagtac aacagcacgt 300accgtgtggt cagcgtcctc accgtcctgc
accaggactg gctgaatggc aaggagtaca 360agtgcaaggt ctccaacaaa gccctcccag
cccccatcga gaaaaccatc tccaaagcca 420aagggcagcc ccgagaacca caggtgtaca
ccctgccccc atcccgggat gagctgacca 480agaaccaggt cagcctgacc tgcctggtca
aaggcttcta tcccagcgac atcgccgtgg 540agtgggagag caatgggcag ccggagaaca
actacaagac cacgcctccc gtgctggact 600ccgacggctc cttcttcctc tacagcaagc
tcaccgtgga caagagcagg tggcagcagg 660ggaacgtctt ctcatgctcc gtgatgcatg
aggctctgca caaccactac acgcagaaga 720gcctctccct gtctccgggt aaatgactcg
agcggccg 758107758DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 107aagcttgaat tcccaccatg gccttgacct ttgctttact ggtggccctc
ctggtgctca 60gctgcaagtc aagctgctct gtgggctgcc cagcacctga actcctgggg
gggccctcag 120tcttcctctt ccccccaaaa cccaaggaca ccctcatgat ctcccggacc
cctgaggtca 180catgcgtggt ggtggacgtg agccacgaag accctgaggt caagttcaac
tggtacgtgg 240acggcgtgga ggtgcataat gccaagacaa agccgcggga ggagcagtac
aacagcacgt 300accgtgtggt cagcgtcctc accgtcctgc accaggactg gctgaatggc
aaggagtaca 360agtgcaaggt ctccaacaaa gccctcccag cccccatcga gaaaaccatc
tccaaagcca 420aagggcagcc ccgagaacca caggtgtaca ccctgccccc atcccgggat
gagctgacca 480agaaccaggt cagcctgacc tgcctggtca aaggcttcta tcccagcgac
atcgccgtgg 540agtgggagag caatgggcag ccggagaaca actacaagac cacgcctccc
gtgctggact 600ccgacggctc cttcttcctc tacagcaagc tcaccgtgga caagagcagg
tggcagcagg 660ggaacgtctt ctcatgctcc gtgatgcatg aggctctgca caaccactac
acgcagaaga 720gcctctccct gtctccgggt aaatgactcg agcggccg
758108740DNAArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 108aagcttgaat tcccaccatg ctggctgcca
cagtcctgac cctggccctg ctgggcaatg 60cccatgcctg cccagcacct gaactcctgg
gggggccctc agtcttcctc ttccccccaa 120aacccaagga caccctcatg atctcccgga
cccctgaggt cacatgcgtg gtggtggacg 180tgagccacga agaccctgag gtcaagttca
actggtacgt ggacggcgtg gaggtgcata 240atgccaagac aaagccgcgg gaggagcagt
acaacagcac gtaccgtgtg gtcagcgtcc 300tcaccgtcct gcaccaggac tggctgaatg
gcaaggagta caagtgcaag gtctccaaca 360aagccctccc agcccccatc gagaaaacca
tctccaaagc caaagggcag ccccgagaac 420cacaggtgta caccctgccc ccatcccggg
atgagctgac caagaaccag gtcagcctga 480cctgcctggt caaaggcttc tatcccagcg
acatcgccgt ggagtgggag agcaatgggc 540agccggagaa caactacaag accacgcctc
ccgtgctgga ctccgacggc tccttcttcc 600tctacagcaa gctcaccgtg gacaagagca
ggtggcagca ggggaacgtc ttctcatgct 660ccgtgatgca tgaggctctg cacaaccact
acacgcagaa gagcctctcc ctgtctccgg 720gtaaatgact cgagcggccg
740109242PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 109Met Leu Leu Leu Ala Arg Cys Leu Leu Leu Val Leu Val Ser
Ser Leu1 5 10 15Leu Val
Cys Ser Gly Leu Ala Cys Pro Ala Pro Glu Leu Leu Gly Gly20
25 30Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp
Thr Leu Met Ile35 40 45Ser Arg Thr Pro
Glu Val Thr Cys Val Val Val Asp Val Ser His Glu50 55
60Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu
Val His65 70 75 80Asn
Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr Arg85
90 95Val Val Ser Val Leu Thr Val Leu His Gln Asp
Trp Leu Asn Gly Lys100 105 110Glu Tyr Lys
Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu115
120 125Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu
Pro Gln Val Tyr130 135 140Thr Leu Pro Pro
Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser Leu145 150
155 160Thr Cys Leu Val Lys Gly Phe Tyr Pro
Ser Asp Ile Ala Val Glu Trp165 170 175Glu
Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val180
185 190Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser
Lys Leu Thr Val Asp195 200 205Lys Ser Arg
Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met His210
215 220Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu
Ser Leu Ser Pro225 230 235
240Gly Lys110242PRTArtificial Sequenceartificial construct relating to
Homo Sapiens immunoglobulin 110Met Ala Leu Thr Phe Ala Leu Leu Val
Ala Leu Leu Val Leu Ser Cys1 5 10
15Lys Ser Ser Cys Ser Val Gly Cys Pro Ala Pro Glu Leu Leu Gly
Gly20 25 30Pro Ser Val Phe Leu Phe Pro
Pro Lys Pro Lys Asp Thr Leu Met Ile35 40
45Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser His Glu50
55 60Asp Pro Glu Val Lys Phe Asn Trp Tyr Val
Asp Gly Val Glu Val His65 70 75
80Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr
Arg85 90 95Val Val Ser Val Leu Thr Val
Leu His Gln Asp Trp Leu Asn Gly Lys100 105
110Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu115
120 125Lys Thr Ile Ser Lys Ala Lys Gly Gln
Pro Arg Glu Pro Gln Val Tyr130 135 140Thr
Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser Leu145
150 155 160Thr Cys Leu Val Lys Gly
Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp165 170
175Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro
Val180 185 190Leu Asp Ser Asp Gly Ser Phe
Phe Leu Tyr Ser Lys Leu Thr Val Asp195 200
205Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met His210
215 220Glu Ala Leu His Asn His Tyr Thr Gln
Lys Ser Leu Ser Leu Ser Pro225 230 235
240Gly Lys111236PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 111Met Leu Ala Ala Thr Val
Leu Thr Leu Ala Leu Leu Gly Asn Ala His1 5
10 15Ala Cys Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser
Val Phe Leu Phe20 25 30Pro Pro Lys Pro
Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val35 40
45Thr Cys Val Val Val Asp Val Ser His Glu Asp Pro Glu Val
Lys Phe50 55 60Asn Trp Tyr Val Asp Gly
Val Glu Val His Asn Ala Lys Thr Lys Pro65 70
75 80Arg Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val
Val Ser Val Leu Thr85 90 95Val Leu His
Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val100
105 110Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr
Ile Ser Lys Ala115 120 125Lys Gly Gln Pro
Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg130 135
140Asp Glu Leu Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val
Lys Gly145 150 155 160Phe
Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro165
170 175Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu
Asp Ser Asp Gly Ser180 185 190Phe Phe Leu
Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln195
200 205Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala
Leu His Asn His210 215 220Tyr Thr Gln Lys
Ser Leu Ser Leu Ser Pro Gly Lys225 230
235112219PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 112Cys Pro Ala Pro Glu Leu Leu Gly Gly Pro
Ser Val Phe Leu Phe Pro1 5 10
15Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr20
25 30Cys Val Val Val Asp Val Ser His Glu
Asp Pro Glu Val Lys Phe Asn35 40 45Trp
Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg50
55 60Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val Val
Ser Val Leu Thr Val65 70 75
80Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser85
90 95Asn Lys Ala Leu Pro Ala Pro Ile Glu
Lys Thr Ile Ser Lys Ala Lys100 105 110Gly
Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp115
120 125Glu Leu Thr Lys Asn Gln Val Ser Leu Thr Cys
Leu Val Lys Gly Phe130 135 140Tyr Pro Ser
Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu145
150 155 160Asn Asn Tyr Lys Thr Thr Pro
Pro Val Leu Asp Ser Asp Gly Ser Phe165 170
175Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly180
185 190Asn Val Phe Ser Cys Ser Val Met His
Glu Ala Leu His Asn His Tyr195 200 205Thr
Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys210
215113222PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 113Xaa Pro Pro Cys Pro Ala Pro Glu Leu Leu
Gly Gly Pro Ser Val Phe1 5 10
15Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro20
25 30Glu Val Thr Cys Val Val Val Asp Val
Ser His Glu Asp Pro Glu Val35 40 45Lys
Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr50
55 60Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr
Arg Val Val Ser Val65 70 75
80Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys85
90 95Lys Val Ser Asn Lys Ala Leu Pro Ala
Pro Ile Glu Lys Thr Ile Ser100 105 110Lys
Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro115
120 125Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu Val130 135 140Lys Gly Phe
Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly145
150 155 160Gln Pro Glu Asn Asn Tyr Lys
Thr Thr Pro Pro Val Leu Asp Ser Asp165 170
175Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp180
185 190Gln Gln Gly Asn Val Phe Ser Cys Ser
Val Met His Glu Ala Leu His195 200 205Asn
His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys210
215 2201141427DNAArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 114aagcttgaat tcccaccatg
agctttccat gtaaatttgt agccagcttc cttctgattt 60tcaatgtttc ttccaaaggt
gcagtctcca aaactcacac atgcccaccg tgcccagcac 120ctgaactcct gggggggccc
tcagtcttcc tcttcccccc aaaacccaag gacaccctca 180tgatctcccg gacccctgag
gtcacatgcg tggtggtgga cgtgagccac gaagaccctg 240aggtcaagtt caactggtac
gtggacggcg tggaggtgca taatgccaag acaaagccgc 300gggaggagca gtacaacagc
acgtaccgtg tggtcagcgt cctcaccgtc ctgcaccagg 360actggctgaa tggcaaggag
tacaagtgca aggtctccaa caaagccctc ccagccccca 420tcgagaaaac catctccaaa
gccaaagggc agccccgaga accacaggtg tacaccctgc 480ccccatcccg ggatgagctg
accaagaacc aggtcagcct gacctgcctg gtcaaaggct 540tctatcccag cgacatcgcc
gtggagtggg agagcaatgg gcagccggag aacaactaca 600agaccacgcc tcccgtgctg
gactccgacg gctccttctt cctctacagc aagctcaccg 660tggacaagag caggtggcag
caggggaacg tcttctcatg ctccgtgatg catgaggctc 720tgcacaacca ctacacgcag
aagagcctct ccctgtctcc ggagctgcaa ctggaggaga 780gctgtgcgga ggcgcaggac
ggggagctgg acgggtgcgt atccggtgac accattgtaa 840tgactagtgg cggtccgcgc
actgtggctg aactggaggg caaaccgttc accgcactga 900ttcgcggctc tggctaccca
tgcccctcag gtttcttccg cacctgtgaa cgtgacgtat 960atgatctgcg tacacgtgag
ggtcattgct tacgtttgac ccatgatcac cgtgttctgg 1020tgatggatgg tggcctggaa
tggcgtgccg cgggtgaact ggaacgcggc gaccgcctgg 1080tgatggatga tgcagctggc
gagtttccgg cactggcaac cttccgtggc ctgcgtggcg 1140ctggccgcca ggatgtttat
gacgctactg tttacggtgc tagcgcattc actgctaatg 1200gcttcattgt acacgcatgt
ggcgagcagc ccgggaccgg tctgaactca ggcctcacga 1260caaatcctgg tgtatccgct
tggcaggtca acacagctta tactgcggga caattggtca 1320catataacgg caagacgtat
aaatgtttgc agccccacac ctccttggca ggatgggaac 1380catccaacgt tcctgccttg
tggcagcttc aatgactcga gcggccg 14271151430DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 115aagcttgaat tcccaccatg aaccggggag tcccttttag gcacttgctt
ctggtgctgc 60aactggcgct cctcccagca gccactcagg gaaaaactca cacatgccca
ccgtgcccag 120cacctgaact cctggggggg ccctcagtct tcctcttccc cccaaaaccc
aaggacaccc 180tcatgatctc ccggacccct gaggtcacat gcgtggtggt ggacgtgagc
cacgaagacc 240ctgaggtcaa gttcaactgg tacgtggacg gcgtggaggt gcataatgcc
aagacaaagc 300cgcgggagga gcagtacaac agcacgtacc gtgtggtcag cgtcctcacc
gtcctgcacc 360aggactggct gaatggcaag gagtacaagt gcaaggtctc caacaaagcc
ctcccagccc 420ccatcgagaa aaccatctcc aaagccaaag ggcagccccg agaaccacag
gtgtacaccc 480tgcccccatc ccgggatgag ctgaccaaga accaggtcag cctgacctgc
ctggtcaaag 540gcttctatcc cagcgacatc gccgtggagt gggagagcaa tgggcagccg
gagaacaact 600acaagaccac gcctcccgtg ctggactccg acggctcctt cttcctctac
agcaagctca 660ccgtggacaa gagcaggtgg cagcagggga acgtcttctc atgctccgtg
atgcatgagg 720ctctgcacaa ccactacacg cagaagagcc tctccctgtc tccggagctg
caactggagg 780agagctgtgc ggaggcgcag gacggggagc tggacgggtg cgtatccggt
gacaccattg 840taatgactag tggcggtccg cgcactgtgg ctgaactgga gggcaaaccg
ttcaccgcac 900tgattcgcgg ctctggctac ccatgcccct caggtttctt ccgcacctgt
gaacgtgacg 960tatatgatct gcgtacacgt gagggtcatt gcttacgttt gacccatgat
caccgtgttc 1020tggtgatgga tggtggcctg gaatggcgtg ccgcgggtga actggaacgc
ggcgaccgcc 1080tggtgatgga tgatgcagct ggcgagtttc cggcactggc aaccttccgt
ggcctgcgtg 1140gcgctggccg ccaggatgtt tatgacgcta ctgtttacgg tgctagcgca
ttcactgcta 1200atggcttcat tgtacacgca tgtggcgagc agcccgggac cggtctgaac
tcaggcctca 1260cgacaaatcc tggtgtatcc gcttggcagg tcaacacagc ttatactgcg
ggacaattgg 1320tcacatataa cggcaagacg tataaatgtt tgcagcccca cacctccttg
gcaggatggg 1380aaccatccaa cgttcctgcc ttgtggcagc ttcaatgact cgagcggccg
1430116465PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 116Met Ser Phe Pro Cys Lys Phe Val
Ala Ser Phe Leu Leu Ile Phe Asn1 5 10
15Val Ser Ser Lys Gly Ala Val Ser Lys Thr His Thr Cys Pro
Pro Cys20 25 30Pro Ala Pro Glu Leu Leu
Gly Gly Pro Ser Val Phe Leu Phe Pro Pro35 40
45Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys50
55 60Val Val Val Asp Val Ser His Glu Asp
Pro Glu Val Lys Phe Asn Trp65 70 75
80Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro
Arg Glu85 90 95Glu Gln Tyr Asn Ser Thr
Tyr Arg Val Val Ser Val Leu Thr Val Leu100 105
110His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser
Asn115 120 125Lys Ala Leu Pro Ala Pro Ile
Glu Lys Thr Ile Ser Lys Ala Lys Gly130 135
140Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu145
150 155 160Leu Thr Lys Asn
Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr165 170
175Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro
Glu Asn180 185 190Asn Tyr Lys Thr Thr Pro
Pro Val Leu Asp Ser Asp Gly Ser Phe Phe195 200
205Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly
Asn210 215 220Val Phe Ser Cys Ser Val Met
His Glu Ala Leu His Asn His Tyr Thr225 230
235 240Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln Leu
Glu Glu Ser Cys245 250 255Ala Glu Ala Gln
Asp Gly Glu Leu Asp Gly Cys Val Ser Gly Asp Thr260 265
270Ile Val Met Thr Ser Gly Gly Pro Arg Thr Val Ala Glu Leu
Glu Gly275 280 285Lys Pro Phe Thr Ala Leu
Ile Arg Gly Ser Gly Tyr Pro Cys Pro Ser290 295
300Gly Phe Phe Arg Thr Cys Glu Arg Asp Val Tyr Asp Leu Arg Thr
Arg305 310 315 320Glu Gly
His Cys Leu Arg Leu Thr His Asp His Arg Val Leu Val Met325
330 335Asp Gly Gly Leu Glu Trp Arg Ala Ala Gly Glu Leu
Glu Arg Gly Asp340 345 350Arg Leu Val Met
Asp Asp Ala Ala Gly Glu Phe Pro Ala Leu Ala Thr355 360
365Phe Arg Gly Leu Arg Gly Ala Gly Arg Gln Asp Val Tyr Asp
Ala Thr370 375 380Val Tyr Gly Ala Ser Ala
Phe Thr Ala Asn Gly Phe Ile Val His Ala385 390
395 400Cys Gly Glu Gln Pro Gly Thr Gly Leu Asn Ser
Gly Leu Thr Thr Asn405 410 415Pro Gly Val
Ser Ala Trp Gln Val Asn Thr Ala Tyr Thr Ala Gly Gln420
425 430Leu Val Thr Tyr Asn Gly Lys Thr Tyr Lys Cys Leu
Gln Pro His Thr435 440 445Ser Leu Ala Gly
Trp Glu Pro Ser Asn Val Pro Ala Leu Trp Gln Leu450 455
460Gln465117466PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 117Met Asn Arg Gly Val Pro
Phe Arg His Leu Leu Leu Val Leu Gln Leu1 5
10 15Ala Leu Leu Pro Ala Ala Thr Gln Gly Lys Thr His
Thr Cys Pro Pro20 25 30Cys Pro Ala Pro
Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro35 40
45Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu
Val Thr50 55 60Cys Val Val Val Asp Val
Ser His Glu Asp Pro Glu Val Lys Phe Asn65 70
75 80Trp Tyr Val Asp Gly Val Glu Val His Asn Ala
Lys Thr Lys Pro Arg85 90 95Glu Glu Gln
Tyr Asn Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val100
105 110Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys
Cys Lys Val Ser115 120 125Asn Lys Ala Leu
Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys130 135
140Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser
Arg Asp145 150 155 160Glu
Leu Thr Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe165
170 175Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser
Asn Gly Gln Pro Glu180 185 190Asn Asn Tyr
Lys Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe195
200 205Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg
Trp Gln Gln Gly210 215 220Asn Val Phe Ser
Cys Ser Val Met His Glu Ala Leu His Asn His Tyr225 230
235 240Thr Gln Lys Ser Leu Ser Leu Ser Pro
Glu Leu Gln Leu Glu Glu Ser245 250 255Cys
Ala Glu Ala Gln Asp Gly Glu Leu Asp Gly Cys Val Ser Gly Asp260
265 270Thr Ile Val Met Thr Ser Gly Gly Pro Arg Thr
Val Ala Glu Leu Glu275 280 285Gly Lys Pro
Phe Thr Ala Leu Ile Arg Gly Ser Gly Tyr Pro Cys Pro290
295 300Ser Gly Phe Phe Arg Thr Cys Glu Arg Asp Val Tyr
Asp Leu Arg Thr305 310 315
320Arg Glu Gly His Cys Leu Arg Leu Thr His Asp His Arg Val Leu Val325
330 335Met Asp Gly Gly Leu Glu Trp Arg Ala
Ala Gly Glu Leu Glu Arg Gly340 345 350Asp
Arg Leu Val Met Asp Asp Ala Ala Gly Glu Phe Pro Ala Leu Ala355
360 365Thr Phe Arg Gly Leu Arg Gly Ala Gly Arg Gln
Asp Val Tyr Asp Ala370 375 380Thr Val Tyr
Gly Ala Ser Ala Phe Thr Ala Asn Gly Phe Ile Val His385
390 395 400Ala Cys Gly Glu Gln Pro Gly
Thr Gly Leu Asn Ser Gly Leu Thr Thr405 410
415Asn Pro Gly Val Ser Ala Trp Gln Val Asn Thr Ala Tyr Thr Ala Gly420
425 430Gln Leu Val Thr Tyr Asn Gly Lys Thr
Tyr Lys Cys Leu Gln Pro His435 440 445Thr
Ser Leu Ala Gly Trp Glu Pro Ser Asn Val Pro Ala Leu Trp Gln450
455 460Leu Gln465118441PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 118Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu
Gly Gly1 5 10 15Pro Ser
Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile20
25 30Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp
Val Ser His Glu35 40 45Asp Pro Glu Val
Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val His50 55
60Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr
Tyr Arg65 70 75 80Val
Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys85
90 95Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu
Pro Ala Pro Ile Glu100 105 110Lys Thr Ile
Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr115
120 125Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn
Gln Val Ser Leu130 135 140Thr Cys Leu Val
Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp145 150
155 160Glu Ser Asn Gly Gln Pro Glu Asn Asn
Tyr Lys Thr Thr Pro Pro Val165 170 175Leu
Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp180
185 190Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser
Cys Ser Val Met His195 200 205Glu Ala Leu
His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro210
215 220Glu Leu Gln Leu Glu Glu Ser Cys Ala Glu Ala Gln
Asp Gly Glu Leu225 230 235
240Asp Gly Cys Val Ser Gly Asp Thr Ile Val Met Thr Ser Gly Gly Pro245
250 255Arg Thr Val Ala Glu Leu Glu Gly Lys
Pro Phe Thr Ala Leu Ile Arg260 265 270Gly
Ser Gly Tyr Pro Cys Pro Ser Gly Phe Phe Arg Thr Cys Glu Arg275
280 285Asp Val Tyr Asp Leu Arg Thr Arg Glu Gly His
Cys Leu Arg Leu Thr290 295 300His Asp His
Arg Val Leu Val Met Asp Gly Gly Leu Glu Trp Arg Ala305
310 315 320Ala Gly Glu Leu Glu Arg Gly
Asp Arg Leu Val Met Asp Asp Ala Ala325 330
335Gly Glu Phe Pro Ala Leu Ala Thr Phe Arg Gly Leu Arg Gly Ala Gly340
345 350Arg Gln Asp Val Tyr Asp Ala Thr Val
Tyr Gly Ala Ser Ala Phe Thr355 360 365Ala
Asn Gly Phe Ile Val His Ala Cys Gly Glu Gln Pro Gly Thr Gly370
375 380Leu Asn Ser Gly Leu Thr Thr Asn Pro Gly Val
Ser Ala Trp Gln Val385 390 395
400Asn Thr Ala Tyr Thr Ala Gly Gln Leu Val Thr Tyr Asn Gly Lys
Thr405 410 415Tyr Lys Cys Leu Gln Pro His
Thr Ser Leu Ala Gly Trp Glu Pro Ser420 425
430Asn Val Pro Ala Leu Trp Gln Leu Gln435
440119242PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 119Lys Thr His Thr Cys Pro Pro Cys Pro Ala
Pro Glu Leu Leu Gly Gly1 5 10
15Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile20
25 30Ser Arg Thr Pro Glu Val Thr Cys Val
Val Val Asp Val Ser His Glu35 40 45Asp
Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val His50
55 60Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr
Asn Ser Thr Tyr Arg65 70 75
80Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys85
90 95Glu Tyr Lys Cys Lys Val Ser Asn Lys
Ala Leu Pro Ala Pro Ile Glu100 105 110Lys
Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr115
120 125Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys
Asn Gln Val Ser Leu130 135 140Thr Cys Leu
Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp145
150 155 160Glu Ser Asn Gly Gln Pro Glu
Asn Asn Tyr Lys Thr Thr Pro Pro Val165 170
175Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp180
185 190Lys Ser Arg Trp Gln Gln Gly Asn Val
Phe Ser Cys Ser Val Met His195 200 205Glu
Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro210
215 220Glu Leu Gln Leu Glu Glu Ser Cys Ala Glu Ala
Gln Asp Gly Glu Leu225 230 235
240Asp Xaa120243PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 120Lys Thr His Thr Cys Pro Pro Cys
Pro Ala Pro Glu Leu Leu Gly Gly1 5 10
15Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu
Met Ile20 25 30Ser Arg Thr Pro Glu Val
Thr Cys Val Val Val Asp Val Ser His Glu35 40
45Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val His50
55 60Asn Ala Lys Thr Lys Pro Arg Glu Glu
Gln Tyr Asn Ser Thr Tyr Arg65 70 75
80Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn
Gly Lys85 90 95Glu Tyr Lys Cys Lys Val
Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu100 105
110Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val
Tyr115 120 125Thr Leu Pro Pro Ser Arg Asp
Glu Leu Thr Lys Asn Gln Val Ser Leu130 135
140Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp145
150 155 160Glu Ser Asn Gly
Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val165 170
175Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr
Val Asp180 185 190Lys Ser Arg Trp Gln Gln
Gly Asn Val Phe Ser Cys Ser Val Met His195 200
205Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser
Pro210 215 220Glu Leu Gln Leu Glu Glu Ser
Cys Ala Glu Ala Gln Asp Gly Glu Leu225 230
235 240Asp Gly Xaa121800DNAArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 121aagcttgaat
tcccaccatg agctttccat gtaaatttgt agccagcttc cttctgattt 60tcaatgtttc
ttccaaaggt gcagtctcca aaactcacac atgcccaccg tgcccagcac 120ctgaactcct
gggggggccc tcagtcttcc tcttcccccc aaaacccaag gacaccctca 180tgatctcccg
gacccctgag gtcacatgcg tggtggtgga cgtgagccac gaagaccctg 240aggtcaagtt
caactggtac gtggacggcg tggaggtgca taatgccaag acaaagccgc 300gggaggagca
gtacaacagc acgtaccgtg tggtcagcgt cctcaccgtc ctgcaccagg 360actggctgaa
tggcaaggag tacaagtgca aggtctccaa caaagccctc ccagccccca 420tcgagaaaac
catctccaaa gccaaagggc agccccgaga accacaggtg tacaccctgc 480ccccatcccg
ggatgagctg accaagaacc aggtcagcct gacctgcctg gtcaaaggct 540tctatcccag
cgacatcgcc gtggagtggg agagcaatgg gcagccggag aacaactaca 600agaccacgcc
tcccgtgctg gactccgacg gctccttctt cctctacagc aagctcaccg 660tggacaagag
caggtggcag caggggaacg tcttctcatg ctccgtgatg catgaggctc 720tgcacaacca
ctacacgcag aagagcctct ccctgtctcc ggagctgcaa ctggaggaga 780gctgttgact
cgagcggccg
800122803DNAArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 122aagcttgaat tcccaccatg aaccggggag
tcccttttag gcacttgctt ctggtgctgc 60aactggcgct cctcccagca gccactcagg
gaaaaactca cacatgccca ccgtgcccag 120cacctgaact cctggggggg ccctcagtct
tcctcttccc cccaaaaccc aaggacaccc 180tcatgatctc ccggacccct gaggtcacat
gcgtggtggt ggacgtgagc cacgaagacc 240ctgaggtcaa gttcaactgg tacgtggacg
gcgtggaggt gcataatgcc aagacaaagc 300cgcgggagga gcagtacaac agcacgtacc
gtgtggtcag cgtcctcacc gtcctgcacc 360aggactggct gaatggcaag gagtacaagt
gcaaggtctc caacaaagcc ctcccagccc 420ccatcgagaa aaccatctcc aaagccaaag
ggcagccccg agaaccacag gtgtacaccc 480tgcccccatc ccgggatgag ctgaccaaga
accaggtcag cctgacctgc ctggtcaaag 540gcttctatcc cagcgacatc gccgtggagt
gggagagcaa tgggcagccg gagaacaact 600acaagaccac gcctcccgtg ctggactccg
acggctcctt cttcctctac agcaagctca 660ccgtggacaa gagcaggtgg cagcagggga
acgtcttctc atgctccgtg atgcatgagg 720ctctgcacaa ccactacacg cagaagagcc
tctccctgtc tccggagctg caactggagg 780agagctgttg actcgagcgg ccg
803123256PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 123Met Ser Phe Pro Cys Lys Phe Val Ala Ser Phe Leu Leu Ile
Phe Asn1 5 10 15Val Ser
Ser Lys Gly Ala Val Ser Lys Thr His Thr Cys Pro Pro Cys20
25 30Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe
Leu Phe Pro Pro35 40 45Lys Pro Lys Asp
Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys50 55
60Val Val Val Asp Val Ser His Glu Asp Pro Glu Val Lys Phe
Asn Trp65 70 75 80Tyr
Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu85
90 95Glu Gln Tyr Asn Ser Thr Tyr Arg Val Val Ser
Val Leu Thr Val Leu100 105 110His Gln Asp
Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn115
120 125Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser
Lys Ala Lys Gly130 135 140Gln Pro Arg Glu
Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu145 150
155 160Leu Thr Lys Asn Gln Val Ser Leu Thr
Cys Leu Val Lys Gly Phe Tyr165 170 175Pro
Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn180
185 190Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser
Asp Gly Ser Phe Phe195 200 205Leu Tyr Ser
Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn210
215 220Val Phe Ser Cys Ser Val Met His Glu Ala Leu His
Asn His Tyr Thr225 230 235
240Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln Leu Glu Glu Ser Cys245
250 255124257PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 124Met Asn Arg Gly Val Pro Phe Arg His Leu Leu Leu Val Leu
Gln Leu1 5 10 15Ala Leu
Leu Pro Ala Ala Thr Gln Gly Lys Thr His Thr Cys Pro Pro20
25 30Cys Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val
Phe Leu Phe Pro35 40 45Pro Lys Pro Lys
Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr50 55
60Cys Val Val Val Asp Val Ser His Glu Asp Pro Glu Val Lys
Phe Asn65 70 75 80Trp
Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg85
90 95Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val Val
Ser Val Leu Thr Val100 105 110Leu His Gln
Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser115
120 125Asn Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile
Ser Lys Ala Lys130 135 140Gly Gln Pro Arg
Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp145 150
155 160Glu Leu Thr Lys Asn Gln Val Ser Leu
Thr Cys Leu Val Lys Gly Phe165 170 175Tyr
Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu180
185 190Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp
Ser Asp Gly Ser Phe195 200 205Phe Leu Tyr
Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly210
215 220Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu
His Asn His Tyr225 230 235
240Thr Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln Leu Glu Glu Ser245
250 255Cys125232PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 125Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu
Gly Gly1 5 10 15Pro Ser
Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile20
25 30Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp
Val Ser His Glu35 40 45Asp Pro Glu Val
Lys Phe Asn Trp Tyr Val Asp Gly Val Glu Val His50 55
60Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr
Tyr Arg65 70 75 80Val
Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys85
90 95Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu
Pro Ala Pro Ile Glu100 105 110Lys Thr Ile
Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr115
120 125Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn
Gln Val Ser Leu130 135 140Thr Cys Leu Val
Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val Glu Trp145 150
155 160Glu Ser Asn Gly Gln Pro Glu Asn Asn
Tyr Lys Thr Thr Pro Pro Val165 170 175Leu
Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp180
185 190Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser
Cys Ser Val Met His195 200 205Glu Ala Leu
His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro210
215 220Glu Leu Gln Leu Glu Glu Ser Cys225
230126466PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 126Met Leu Leu Leu Ala Arg Cys Leu Leu Leu
Val Leu Val Ser Ser Leu1 5 10
15Leu Val Cys Ser Gly Leu Ala Cys Asp Lys Thr His Thr Cys Pro Pro20
25 30Cys Pro Ala Pro Glu Leu Leu Gly Gly
Pro Ser Val Phe Leu Phe Pro35 40 45Pro
Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr50
55 60Cys Val Val Val Asp Val Ser His Glu Asp Pro
Glu Val Lys Phe Asn65 70 75
80Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg85
90 95Glu Glu Gln Tyr Asn Ser Thr Tyr Arg
Val Val Ser Val Leu Thr Val100 105 110Leu
His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser115
120 125Asn Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr
Ile Ser Lys Ala Lys130 135 140Gly Gln Pro
Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp145
150 155 160Glu Leu Thr Lys Asn Gln Val
Ser Leu Thr Cys Leu Val Lys Gly Phe165 170
175Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu180
185 190Asn Asn Tyr Lys Thr Thr Pro Pro Val
Leu Asp Ser Asp Gly Ser Phe195 200 205Phe
Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly210
215 220Asn Val Phe Ser Cys Ser Val Met His Glu Ala
Leu His Asn His Tyr225 230 235
240Thr Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln Leu Glu Glu
Ser245 250 255Cys Ala Glu Ala Gln Asp Gly
Glu Leu Asp Gly Cys Val Ser Gly Asp260 265
270Thr Ile Val Met Thr Ser Gly Gly Pro Arg Thr Val Ala Glu Leu Glu275
280 285Gly Lys Pro Phe Thr Ala Leu Ile Arg
Gly Ser Gly Tyr Pro Cys Pro290 295 300Ser
Gly Phe Phe Arg Thr Cys Glu Arg Asp Val Tyr Asp Leu Arg Thr305
310 315 320Arg Glu Gly His Cys Leu
Arg Leu Thr His Asp His Arg Val Leu Val325 330
335Met Asp Gly Gly Leu Glu Trp Arg Ala Ala Gly Glu Leu Glu Arg
Gly340 345 350Asp Arg Leu Val Met Asp Asp
Ala Ala Gly Glu Phe Pro Ala Leu Ala355 360
365Thr Phe Arg Gly Leu Arg Gly Ala Gly Arg Gln Asp Val Tyr Asp Ala370
375 380Thr Val Tyr Gly Ala Ser Ala Phe Thr
Ala Asn Gly Phe Ile Val His385 390 395
400Ala Cys Gly Glu Gln Pro Gly Thr Gly Leu Asn Ser Gly Leu
Thr Thr405 410 415Asn Pro Gly Val Ser Ala
Trp Gln Val Asn Thr Ala Tyr Thr Ala Gly420 425
430Gln Leu Val Thr Tyr Asn Gly Lys Thr Tyr Lys Cys Leu Gln Pro
His435 440 445Thr Ser Leu Ala Gly Trp Glu
Pro Ser Asn Val Pro Ala Leu Trp Gln450 455
460Leu Gln465127466PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 127Met Ala Leu Thr Phe Ala Leu Leu
Val Ala Leu Leu Val Leu Ser Cys1 5 10
15Lys Ser Ser Cys Ser Val Gly Cys Asp Lys Thr His Thr Cys
Pro Pro20 25 30Cys Pro Ala Pro Glu Leu
Leu Gly Gly Pro Ser Val Phe Leu Phe Pro35 40
45Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr50
55 60Cys Val Val Val Asp Val Ser His Glu
Asp Pro Glu Val Lys Phe Asn65 70 75
80Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys
Pro Arg85 90 95Glu Glu Gln Tyr Asn Ser
Thr Tyr Arg Val Val Ser Val Leu Thr Val100 105
110Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val
Ser115 120 125Asn Lys Ala Leu Pro Ala Pro
Ile Glu Lys Thr Ile Ser Lys Ala Lys130 135
140Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp145
150 155 160Glu Leu Thr Lys
Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe165 170
175Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu180 185 190Asn Asn Tyr Lys Thr Thr
Pro Pro Val Leu Asp Ser Asp Gly Ser Phe195 200
205Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln
Gly210 215 220Asn Val Phe Ser Cys Ser Val
Met His Glu Ala Leu His Asn His Tyr225 230
235 240Thr Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln
Leu Glu Glu Ser245 250 255Cys Ala Glu Ala
Gln Asp Gly Glu Leu Asp Gly Cys Val Ser Gly Asp260 265
270Thr Ile Val Met Thr Ser Gly Gly Pro Arg Thr Val Ala Glu
Leu Glu275 280 285Gly Lys Pro Phe Thr Ala
Leu Ile Arg Gly Ser Gly Tyr Pro Cys Pro290 295
300Ser Gly Phe Phe Arg Thr Cys Glu Arg Asp Val Tyr Asp Leu Arg
Thr305 310 315 320Arg Glu
Gly His Cys Leu Arg Leu Thr His Asp His Arg Val Leu Val325
330 335Met Asp Gly Gly Leu Glu Trp Arg Ala Ala Gly Glu
Leu Glu Arg Gly340 345 350Asp Arg Leu Val
Met Asp Asp Ala Ala Gly Glu Phe Pro Ala Leu Ala355 360
365Thr Phe Arg Gly Leu Arg Gly Ala Gly Arg Gln Asp Val Tyr
Asp Ala370 375 380Thr Val Tyr Gly Ala Ser
Ala Phe Thr Ala Asn Gly Phe Ile Val His385 390
395 400Ala Cys Gly Glu Gln Pro Gly Thr Gly Leu Asn
Ser Gly Leu Thr Thr405 410 415Asn Pro Gly
Val Ser Ala Trp Gln Val Asn Thr Ala Tyr Thr Ala Gly420
425 430Gln Leu Val Thr Tyr Asn Gly Lys Thr Tyr Lys Cys
Leu Gln Pro His435 440 445Thr Ser Leu Ala
Gly Trp Glu Pro Ser Asn Val Pro Ala Leu Trp Gln450 455
460Leu Gln465128460PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 128Met Leu Ala
Ala Thr Val Leu Thr Leu Ala Leu Leu Gly Asn Ala His1 5
10 15Ala Cys Asp Lys Thr His Thr Cys Pro
Pro Cys Pro Ala Pro Glu Leu20 25 30Leu
Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr35
40 45Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys
Val Val Val Asp Val50 55 60Ser His Glu
Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val65 70
75 80Glu Val His Asn Ala Lys Thr Lys
Pro Arg Glu Glu Gln Tyr Asn Ser85 90
95Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu100
105 110Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser
Asn Lys Ala Leu Pro Ala115 120 125Pro Ile
Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro130
135 140Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu
Thr Lys Asn Gln145 150 155
160Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala165
170 175Val Glu Trp Glu Ser Asn Gly Gln Pro
Glu Asn Asn Tyr Lys Thr Thr180 185 190Pro
Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu195
200 205Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn
Val Phe Ser Cys Ser210 215 220Val Met His
Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser225
230 235 240Leu Ser Pro Glu Leu Gln Leu
Glu Glu Ser Cys Ala Glu Ala Gln Asp245 250
255Gly Glu Leu Asp Gly Cys Val Ser Gly Asp Thr Ile Val Met Thr Ser260
265 270Gly Gly Pro Arg Thr Val Ala Glu Leu
Glu Gly Lys Pro Phe Thr Ala275 280 285Leu
Ile Arg Gly Ser Gly Tyr Pro Cys Pro Ser Gly Phe Phe Arg Thr290
295 300Cys Glu Arg Asp Val Tyr Asp Leu Arg Thr Arg
Glu Gly His Cys Leu305 310 315
320Arg Leu Thr His Asp His Arg Val Leu Val Met Asp Gly Gly Leu
Glu325 330 335Trp Arg Ala Ala Gly Glu Leu
Glu Arg Gly Asp Arg Leu Val Met Asp340 345
350Asp Ala Ala Gly Glu Phe Pro Ala Leu Ala Thr Phe Arg Gly Leu Arg355
360 365Gly Ala Gly Arg Gln Asp Val Tyr Asp
Ala Thr Val Tyr Gly Ala Ser370 375 380Ala
Phe Thr Ala Asn Gly Phe Ile Val His Ala Cys Gly Glu Gln Pro385
390 395 400Gly Thr Gly Leu Asn Ser
Gly Leu Thr Thr Asn Pro Gly Val Ser Ala405 410
415Trp Gln Val Asn Thr Ala Tyr Thr Ala Gly Gln Leu Val Thr Tyr
Asn420 425 430Gly Lys Thr Tyr Lys Cys Leu
Gln Pro His Thr Ser Leu Ala Gly Trp435 440
445Glu Pro Ser Asn Val Pro Ala Leu Trp Gln Leu Gln450
455 460129443PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 129Cys Asp Lys Thr His Thr
Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu1 5
10 15Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro
Lys Asp Thr Leu20 25 30Met Ile Ser Arg
Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser35 40
45His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly
Val Glu50 55 60Val His Asn Ala Lys Thr
Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr65 70
75 80Tyr Arg Val Val Ser Val Leu Thr Val Leu His
Gln Asp Trp Leu Asn85 90 95Gly Lys Glu
Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro100
105 110Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro
Arg Glu Pro Gln115 120 125Val Tyr Thr Leu
Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln Val130 135
140Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile
Ala Val145 150 155 160Glu
Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro165
170 175Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu
Tyr Ser Lys Leu Thr180 185 190Val Asp Lys
Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val195
200 205Met His Glu Ala Leu His Asn His Tyr Thr Gln Lys
Ser Leu Ser Leu210 215 220Ser Pro Glu Leu
Gln Leu Glu Glu Ser Cys Ala Glu Ala Gln Asp Gly225 230
235 240Glu Leu Asp Gly Cys Val Ser Gly Asp
Thr Ile Val Met Thr Ser Gly245 250 255Gly
Pro Arg Thr Val Ala Glu Leu Glu Gly Lys Pro Phe Thr Ala Leu260
265 270Ile Arg Gly Ser Gly Tyr Pro Cys Pro Ser Gly
Phe Phe Arg Thr Cys275 280 285Glu Arg Asp
Val Tyr Asp Leu Arg Thr Arg Glu Gly His Cys Leu Arg290
295 300Leu Thr His Asp His Arg Val Leu Val Met Asp Gly
Gly Leu Glu Trp305 310 315
320Arg Ala Ala Gly Glu Leu Glu Arg Gly Asp Arg Leu Val Met Asp Asp325
330 335Ala Ala Gly Glu Phe Pro Ala Leu Ala
Thr Phe Arg Gly Leu Arg Gly340 345 350Ala
Gly Arg Gln Asp Val Tyr Asp Ala Thr Val Tyr Gly Ala Ser Ala355
360 365Phe Thr Ala Asn Gly Phe Ile Val His Ala Cys
Gly Glu Gln Pro Gly370 375 380Thr Gly Leu
Asn Ser Gly Leu Thr Thr Asn Pro Gly Val Ser Ala Trp385
390 395 400Gln Val Asn Thr Ala Tyr Thr
Ala Gly Gln Leu Val Thr Tyr Asn Gly405 410
415Lys Thr Tyr Lys Cys Leu Gln Pro His Thr Ser Leu Ala Gly Trp Glu420
425 430Pro Ser Asn Val Pro Ala Leu Trp Gln
Leu Gln435 440130244PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 130Cys Asp Lys
Thr His Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu1 5
10 15Gly Gly Pro Ser Val Phe Leu Phe Pro
Pro Lys Pro Lys Asp Thr Leu20 25 30Met
Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser35
40 45His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr
Val Asp Gly Val Glu50 55 60Val His Asn
Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr65 70
75 80Tyr Arg Val Val Ser Val Leu Thr
Val Leu His Gln Asp Trp Leu Asn85 90
95Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro100
105 110Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly
Gln Pro Arg Glu Pro Gln115 120 125Val Tyr
Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln Val130
135 140Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser
Asp Ile Ala Val145 150 155
160Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro165
170 175Pro Val Leu Asp Ser Asp Gly Ser Phe
Phe Leu Tyr Ser Lys Leu Thr180 185 190Val
Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val195
200 205Met His Glu Ala Leu His Asn His Tyr Thr Gln
Lys Ser Leu Ser Leu210 215 220Ser Pro Glu
Leu Gln Leu Glu Glu Ser Cys Ala Glu Ala Gln Asp Gly225
230 235 240Glu Leu Asp
Xaa131245PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 131Cys Asp Lys Thr His Thr Cys Pro Pro Cys
Pro Ala Pro Glu Leu Leu1 5 10
15Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu20
25 30Met Ile Ser Arg Thr Pro Glu Val Thr
Cys Val Val Val Asp Val Ser35 40 45His
Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu50
55 60Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu
Gln Tyr Asn Ser Thr65 70 75
80Tyr Arg Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn85
90 95Gly Lys Glu Tyr Lys Cys Lys Val Ser
Asn Lys Ala Leu Pro Ala Pro100 105 110Ile
Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln115
120 125Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu
Thr Lys Asn Gln Val130 135 140Ser Leu Thr
Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val145
150 155 160Glu Trp Glu Ser Asn Gly Gln
Pro Glu Asn Asn Tyr Lys Thr Thr Pro165 170
175Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr180
185 190Val Asp Lys Ser Arg Trp Gln Gln Gly
Asn Val Phe Ser Cys Ser Val195 200 205Met
His Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu210
215 220Ser Pro Glu Leu Gln Leu Glu Glu Ser Cys Ala
Glu Ala Gln Asp Gly225 230 235
240Glu Leu Asp Gly Xaa245132460PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 132Met Leu Leu
Leu Ala Arg Cys Leu Leu Leu Val Leu Val Ser Ser Leu1 5
10 15Leu Val Cys Ser Gly Leu Ala Cys Pro
Pro Cys Pro Ala Pro Glu Leu20 25 30Leu
Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr35
40 45Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys
Val Val Val Asp Val50 55 60Ser His Glu
Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val65 70
75 80Glu Val His Asn Ala Lys Thr Lys
Pro Arg Glu Glu Gln Tyr Asn Ser85 90
95Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu100
105 110Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser
Asn Lys Ala Leu Pro Ala115 120 125Pro Ile
Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro130
135 140Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu
Thr Lys Asn Gln145 150 155
160Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala165
170 175Val Glu Trp Glu Ser Asn Gly Gln Pro
Glu Asn Asn Tyr Lys Thr Thr180 185 190Pro
Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu195
200 205Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn
Val Phe Ser Cys Ser210 215 220Val Met His
Glu Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser225
230 235 240Leu Ser Pro Glu Leu Gln Leu
Glu Glu Ser Cys Ala Glu Ala Gln Asp245 250
255Gly Glu Leu Asp Gly Cys Val Ser Gly Asp Thr Ile Val Met Thr Ser260
265 270Gly Gly Pro Arg Thr Val Ala Glu Leu
Glu Gly Lys Pro Phe Thr Ala275 280 285Leu
Ile Arg Gly Ser Gly Tyr Pro Cys Pro Ser Gly Phe Phe Arg Thr290
295 300Cys Glu Arg Asp Val Tyr Asp Leu Arg Thr Arg
Glu Gly His Cys Leu305 310 315
320Arg Leu Thr His Asp His Arg Val Leu Val Met Asp Gly Gly Leu
Glu325 330 335Trp Arg Ala Ala Gly Glu Leu
Glu Arg Gly Asp Arg Leu Val Met Asp340 345
350Asp Ala Ala Gly Glu Phe Pro Ala Leu Ala Thr Phe Arg Gly Leu Arg355
360 365Gly Ala Gly Arg Gln Asp Val Tyr Asp
Ala Thr Val Tyr Gly Ala Ser370 375 380Ala
Phe Thr Ala Asn Gly Phe Ile Val His Ala Cys Gly Glu Gln Pro385
390 395 400Gly Thr Gly Leu Asn Ser
Gly Leu Thr Thr Asn Pro Gly Val Ser Ala405 410
415Trp Gln Val Asn Thr Ala Tyr Thr Ala Gly Gln Leu Val Thr Tyr
Asn420 425 430Gly Lys Thr Tyr Lys Cys Leu
Gln Pro His Thr Ser Leu Ala Gly Trp435 440
445Glu Pro Ser Asn Val Pro Ala Leu Trp Gln Leu Gln450
455 460133460PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 133Met Ala Leu Thr Phe Ala
Leu Leu Val Ala Leu Leu Val Leu Ser Cys1 5
10 15Lys Ser Ser Cys Ser Val Gly Cys Pro Pro Cys Pro
Ala Pro Glu Leu20 25 30Leu Gly Gly Pro
Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr35 40
45Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val
Asp Val50 55 60Ser His Glu Asp Pro Glu
Val Lys Phe Asn Trp Tyr Val Asp Gly Val65 70
75 80Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu
Glu Gln Tyr Asn Ser85 90 95Thr Tyr Arg
Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu100
105 110Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys
Ala Leu Pro Ala115 120 125Pro Ile Glu Lys
Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro130 135
140Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys
Asn Gln145 150 155 160Val
Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala165
170 175Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn
Asn Tyr Lys Thr Thr180 185 190Pro Pro Val
Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu195
200 205Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val
Phe Ser Cys Ser210 215 220Val Met His Glu
Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser225 230
235 240Leu Ser Pro Glu Leu Gln Leu Glu Glu
Ser Cys Ala Glu Ala Gln Asp245 250 255Gly
Glu Leu Asp Gly Cys Val Ser Gly Asp Thr Ile Val Met Thr Ser260
265 270Gly Gly Pro Arg Thr Val Ala Glu Leu Glu Gly
Lys Pro Phe Thr Ala275 280 285Leu Ile Arg
Gly Ser Gly Tyr Pro Cys Pro Ser Gly Phe Phe Arg Thr290
295 300Cys Glu Arg Asp Val Tyr Asp Leu Arg Thr Arg Glu
Gly His Cys Leu305 310 315
320Arg Leu Thr His Asp His Arg Val Leu Val Met Asp Gly Gly Leu Glu325
330 335Trp Arg Ala Ala Gly Glu Leu Glu Arg
Gly Asp Arg Leu Val Met Asp340 345 350Asp
Ala Ala Gly Glu Phe Pro Ala Leu Ala Thr Phe Arg Gly Leu Arg355
360 365Gly Ala Gly Arg Gln Asp Val Tyr Asp Ala Thr
Val Tyr Gly Ala Ser370 375 380Ala Phe Thr
Ala Asn Gly Phe Ile Val His Ala Cys Gly Glu Gln Pro385
390 395 400Gly Thr Gly Leu Asn Ser Gly
Leu Thr Thr Asn Pro Gly Val Ser Ala405 410
415Trp Gln Val Asn Thr Ala Tyr Thr Ala Gly Gln Leu Val Thr Tyr Asn420
425 430Gly Lys Thr Tyr Lys Cys Leu Gln Pro
His Thr Ser Leu Ala Gly Trp435 440 445Glu
Pro Ser Asn Val Pro Ala Leu Trp Gln Leu Gln450 455
460134454PRTArtificial Sequenceartificial construct relating to
Homo Sapiens immunoglobulin 134Met Leu Ala Ala Thr Val Leu Thr Leu
Ala Leu Leu Gly Asn Ala His1 5 10
15Ala Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser
Val20 25 30Phe Leu Phe Pro Pro Lys Pro
Lys Asp Thr Leu Met Ile Ser Arg Thr35 40
45Pro Glu Val Thr Cys Val Val Val Asp Val Ser His Glu Asp Pro Glu50
55 60Val Lys Phe Asn Trp Tyr Val Asp Gly Val
Glu Val His Asn Ala Lys65 70 75
80Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr Arg Val Val
Ser85 90 95Val Leu Thr Val Leu His Gln
Asp Trp Leu Asn Gly Lys Glu Tyr Lys100 105
110Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile115
120 125Ser Lys Ala Lys Gly Gln Pro Arg Glu
Pro Gln Val Tyr Thr Leu Pro130 135 140Pro
Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser Leu Thr Cys Leu145
150 155 160Val Lys Gly Phe Tyr Pro
Ser Asp Ile Ala Val Glu Trp Glu Ser Asn165 170
175Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp
Ser180 185 190Asp Gly Ser Phe Phe Leu Tyr
Ser Lys Leu Thr Val Asp Lys Ser Arg195 200
205Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met His Glu Ala Leu210
215 220His Asn His Tyr Thr Gln Lys Ser Leu
Ser Leu Ser Pro Glu Leu Gln225 230 235
240Leu Glu Glu Ser Cys Ala Glu Ala Gln Asp Gly Glu Leu Asp
Gly Cys245 250 255Val Ser Gly Asp Thr Ile
Val Met Thr Ser Gly Gly Pro Arg Thr Val260 265
270Ala Glu Leu Glu Gly Lys Pro Phe Thr Ala Leu Ile Arg Gly Ser
Gly275 280 285Tyr Pro Cys Pro Ser Gly Phe
Phe Arg Thr Cys Glu Arg Asp Val Tyr290 295
300Asp Leu Arg Thr Arg Glu Gly His Cys Leu Arg Leu Thr His Asp His305
310 315 320Arg Val Leu Val
Met Asp Gly Gly Leu Glu Trp Arg Ala Ala Gly Glu325 330
335Leu Glu Arg Gly Asp Arg Leu Val Met Asp Asp Ala Ala Gly
Glu Phe340 345 350Pro Ala Leu Ala Thr Phe
Arg Gly Leu Arg Gly Ala Gly Arg Gln Asp355 360
365Val Tyr Asp Ala Thr Val Tyr Gly Ala Ser Ala Phe Thr Ala Asn
Gly370 375 380Phe Ile Val His Ala Cys Gly
Glu Gln Pro Gly Thr Gly Leu Asn Ser385 390
395 400Gly Leu Thr Thr Asn Pro Gly Val Ser Ala Trp Gln
Val Asn Thr Ala405 410 415Tyr Thr Ala Gly
Gln Leu Val Thr Tyr Asn Gly Lys Thr Tyr Lys Cys420 425
430Leu Gln Pro His Thr Ser Leu Ala Gly Trp Glu Pro Ser Asn
Val Pro435 440 445Ala Leu Trp Gln Leu
Gln450135437PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 135Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu
Gly Gly Pro Ser Val Phe1 5 10
15Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr Pro20
25 30Glu Val Thr Cys Val Val Val Asp Val
Ser His Glu Asp Pro Glu Val35 40 45Lys
Phe Asn Trp Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr50
55 60Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr
Arg Val Val Ser Val65 70 75
80Leu Thr Val Leu His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys85
90 95Lys Val Ser Asn Lys Ala Leu Pro Ala
Pro Ile Glu Lys Thr Ile Ser100 105 110Lys
Ala Lys Gly Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro115
120 125Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser
Leu Thr Cys Leu Val130 135 140Lys Gly Phe
Tyr Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly145
150 155 160Gln Pro Glu Asn Asn Tyr Lys
Thr Thr Pro Pro Val Leu Asp Ser Asp165 170
175Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp180
185 190Gln Gln Gly Asn Val Phe Ser Cys Ser
Val Met His Glu Ala Leu His195 200 205Asn
His Tyr Thr Gln Lys Ser Leu Ser Leu Ser Pro Glu Leu Gln Leu210
215 220Glu Glu Ser Cys Ala Glu Ala Gln Asp Gly Glu
Leu Asp Gly Cys Val225 230 235
240Ser Gly Asp Thr Ile Val Met Thr Ser Gly Gly Pro Arg Thr Val
Ala245 250 255Glu Leu Glu Gly Lys Pro Phe
Thr Ala Leu Ile Arg Gly Ser Gly Tyr260 265
270Pro Cys Pro Ser Gly Phe Phe Arg Thr Cys Glu Arg Asp Val Tyr Asp275
280 285Leu Arg Thr Arg Glu Gly His Cys Leu
Arg Leu Thr His Asp His Arg290 295 300Val
Leu Val Met Asp Gly Gly Leu Glu Trp Arg Ala Ala Gly Glu Leu305
310 315 320Glu Arg Gly Asp Arg Leu
Val Met Asp Asp Ala Ala Gly Glu Phe Pro325 330
335Ala Leu Ala Thr Phe Arg Gly Leu Arg Gly Ala Gly Arg Gln Asp
Val340 345 350Tyr Asp Ala Thr Val Tyr Gly
Ala Ser Ala Phe Thr Ala Asn Gly Phe355 360
365Ile Val His Ala Cys Gly Glu Gln Pro Gly Thr Gly Leu Asn Ser Gly370
375 380Leu Thr Thr Asn Pro Gly Val Ser Ala
Trp Gln Val Asn Thr Ala Tyr385 390 395
400Thr Ala Gly Gln Leu Val Thr Tyr Asn Gly Lys Thr Tyr Lys
Cys Leu405 410 415Gln Pro His Thr Ser Leu
Ala Gly Trp Glu Pro Ser Asn Val Pro Ala420 425
430Leu Trp Gln Leu Gln435136443PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 136Xaa Asp Lys
Thr His Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu1 5
10 15Gly Gly Pro Ser Val Phe Leu Phe Pro
Pro Lys Pro Lys Asp Thr Leu20 25 30Met
Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser35
40 45His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr
Val Asp Gly Val Glu50 55 60Val His Asn
Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr65 70
75 80Tyr Arg Val Val Ser Val Leu Thr
Val Leu His Gln Asp Trp Leu Asn85 90
95Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro100
105 110Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly
Gln Pro Arg Glu Pro Gln115 120 125Val Tyr
Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln Val130
135 140Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser
Asp Ile Ala Val145 150 155
160Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro165
170 175Pro Val Leu Asp Ser Asp Gly Ser Phe
Phe Leu Tyr Ser Lys Leu Thr180 185 190Val
Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val195
200 205Met His Glu Ala Leu His Asn His Tyr Thr Gln
Lys Ser Leu Ser Leu210 215 220Ser Pro Glu
Leu Gln Leu Glu Glu Ser Cys Ala Glu Ala Gln Asp Gly225
230 235 240Glu Leu Asp Gly Cys Val Ser
Gly Asp Thr Ile Val Met Thr Ser Gly245 250
255Gly Pro Arg Thr Val Ala Glu Leu Glu Gly Lys Pro Phe Thr Ala Leu260
265 270Ile Arg Gly Ser Gly Tyr Pro Cys Pro
Ser Gly Phe Phe Arg Thr Cys275 280 285Glu
Arg Asp Val Tyr Asp Leu Arg Thr Arg Glu Gly His Cys Leu Arg290
295 300Leu Thr His Asp His Arg Val Leu Val Met Asp
Gly Gly Leu Glu Trp305 310 315
320Arg Ala Ala Gly Glu Leu Glu Arg Gly Asp Arg Leu Val Met Asp
Asp325 330 335Ala Ala Gly Glu Phe Pro Ala
Leu Ala Thr Phe Arg Gly Leu Arg Gly340 345
350Ala Gly Arg Gln Asp Val Tyr Asp Ala Thr Val Tyr Gly Ala Ser Ala355
360 365Phe Thr Ala Asn Gly Phe Ile Val His
Ala Cys Gly Glu Gln Pro Gly370 375 380Thr
Gly Leu Asn Ser Gly Leu Thr Thr Asn Pro Gly Val Ser Ala Trp385
390 395 400Gln Val Asn Thr Ala Tyr
Thr Ala Gly Gln Leu Val Thr Tyr Asn Gly405 410
415Lys Thr Tyr Lys Cys Leu Gln Pro His Thr Ser Leu Ala Gly Trp
Glu420 425 430Pro Ser Asn Val Pro Ala Leu
Trp Gln Leu Gln435 440137244PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 137Xaa Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro Glu
Leu Leu1 5 10 15Gly Gly
Pro Ser Val Phe Leu Phe Pro Pro Lys Pro Lys Asp Thr Leu20
25 30Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val Val
Val Asp Val Ser35 40 45His Glu Asp Pro
Glu Val Lys Phe Asn Trp Tyr Val Asp Gly Val Glu50 55
60Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn
Ser Thr65 70 75 80Tyr
Arg Val Val Ser Val Leu Thr Val Leu His Gln Asp Trp Leu Asn85
90 95Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys
Ala Leu Pro Ala Pro100 105 110Ile Glu Lys
Thr Ile Ser Lys Ala Lys Gly Gln Pro Arg Glu Pro Gln115
120 125Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr
Lys Asn Gln Val130 135 140Ser Leu Thr Cys
Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile Ala Val145 150
155 160Glu Trp Glu Ser Asn Gly Gln Pro Glu
Asn Asn Tyr Lys Thr Thr Pro165 170 175Pro
Val Leu Asp Ser Asp Gly Ser Phe Phe Leu Tyr Ser Lys Leu Thr180
185 190Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val
Phe Ser Cys Ser Val195 200 205Met His Glu
Ala Leu His Asn His Tyr Thr Gln Lys Ser Leu Ser Leu210
215 220Ser Pro Glu Leu Gln Leu Glu Glu Ser Cys Ala Glu
Ala Gln Asp Gly225 230 235
240Glu Leu Asp Xaa138245PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 138Xaa Asp Lys Thr His Thr
Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu1 5
10 15Gly Gly Pro Ser Val Phe Leu Phe Pro Pro Lys Pro
Lys Asp Thr Leu20 25 30Met Ile Ser Arg
Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser35 40
45His Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val Asp Gly
Val Glu50 55 60Val His Asn Ala Lys Thr
Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr65 70
75 80Tyr Arg Val Val Ser Val Leu Thr Val Leu His
Gln Asp Trp Leu Asn85 90 95Gly Lys Glu
Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro100
105 110Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly Gln Pro
Arg Glu Pro Gln115 120 125Val Tyr Thr Leu
Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln Val130 135
140Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser Asp Ile
Ala Val145 150 155 160Glu
Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro165
170 175Pro Val Leu Asp Ser Asp Gly Ser Phe Phe Leu
Tyr Ser Lys Leu Thr180 185 190Val Asp Lys
Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val195
200 205Met His Glu Ala Leu His Asn His Tyr Thr Gln Lys
Ser Leu Ser Leu210 215 220Ser Pro Glu Leu
Gln Leu Glu Glu Ser Cys Ala Glu Ala Gln Asp Gly225 230
235 240Glu Leu Asp Gly
Xaa2451391820DNAArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 139aagcttgaat tcccaccatg aaccggggag
tcccttttag gcacttgctt ctggtgctgc 60aactggcgct cctcccagca gccactcagg
gaaagaaagt ggtgctgggc aaaaaagggg 120atacagtgga actgacctgt acagcttccc
agaagaagag catacaattc cactggaaaa 180actccaacca gataaagatt ctgggaaatc
agggctcctt cttaactaaa ggtccatcca 240agctgaatga tcgcgctgac tcaagaagaa
gcctttggga ccaaggaaac ttccccctga 300tcatcaagaa tcttaagata gaagactcag
atacttacat ctgtgaagtg gaggaccaga 360aggaggaggt gcaattgcta gtgttcggat
tgactgccaa ctctgacacc cacctgcttc 420aggggcagag cctgaccctg accttggaga
gcccccctgg tagtagcccc tcagtgcaat 480gtaggagtcc aaggggtaaa aacatacagg
gggggaagac cctctccgtg tctcagctgg 540agctccagga tagtggcacc tggacatgca
ctgtcttgca gaaccagaag aaggtggagt 600tcaaaataga catcgtggtg ctagctttcc
agaaggcctc cagcatagtc tataagaaag 660agggggaaca ggtggagttc tccttcccac
tcgcctttac agttgaaaag ctgacgggca 720gtggcgagct gtggtggcag gcggagaggg
cttcctcctc caagtcttgg atcacctttg 780acctgaagaa caaggaagtg tctgtaaaac
gggttaccca ggaccctaag ctccagatgg 840gcaagaagct cccgctccac ctcaccctgc
cccaggcctt gcctcagtat gctggctctg 900gaaacctcac cctggccctt gaagcgaaaa
caggaaagtt gcatcaggaa gtgaacctgg 960tggtgatgag agccactcag ctccagaaaa
atttgacctg tgaggtgtgg ggacccacct 1020cccctaagct gatgctgagc ttgaaactgg
agaacaagga ggcaaaggtc tcgaagcggg 1080agaaggcggt gtgggtgctg aaccctgagg
cggggatgtg gcagtgtctg ctgagtgact 1140cgggacaggt cctgctggaa tccaacatca
aggttctgcc cacatggtcc accccggtgc 1200agccagggtg cgtatccggt gacaccattg
taatgactag tggcgggccc cgcactgtgg 1260ctgaactgga gggcaaaccg ttcaccgcac
tgattcgcgg ctctggctac ccatgcccct 1320caggtttctt ccgcacctgt gaacgtgacg
tatatgatct gcgtacacgt gagggtcatt 1380gcttacgttt gacccatgat caccgtgttc
tggtgatgga tggtggcctg gaatggcgtg 1440ccgcgggtga actggaacgc ggcgaccgcc
tggtgatgga tgatgcagct ggcgagtttc 1500cggcactggc aaccttccgt ggcctgcgtg
gcgctggccg ccaggatgtt tatgacgcta 1560ctgtttacgg tgctagcgca ttcactgcta
atggcttcat tgtacacgca tgtggcgagc 1620agcccgggac cggtctgaac tcaggcctca
cgacaaatcc tggtgtatcc gcttggcagg 1680tcaacacagc ttatactgcg ggacaattgg
tcacatataa cggcaagacg tataaatgtt 1740tgcagcccca cacctccttg gcaggatggg
aaccatccaa cgttcctgcc ttgtggcagc 1800ttcaatgact cgagcggccg
1820140596PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 140Met Asn Arg Gly Val Pro Phe Arg His Leu Leu Leu Val Leu
Gln Leu1 5 10 15Ala Leu
Leu Pro Ala Ala Thr Gln Gly Lys Lys Val Val Leu Gly Lys20
25 30Lys Gly Asp Thr Val Glu Leu Thr Cys Thr Ala Ser
Gln Lys Lys Ser35 40 45Ile Gln Phe His
Trp Lys Asn Ser Asn Gln Ile Lys Ile Leu Gly Asn50 55
60Gln Gly Ser Phe Leu Thr Lys Gly Pro Ser Lys Leu Asn Asp
Arg Ala65 70 75 80Asp
Ser Arg Arg Ser Leu Trp Asp Gln Gly Asn Phe Pro Leu Ile Ile85
90 95Lys Asn Leu Lys Ile Glu Asp Ser Asp Thr Tyr
Ile Cys Glu Val Glu100 105 110Asp Gln Lys
Glu Glu Val Gln Leu Leu Val Phe Gly Leu Thr Ala Asn115
120 125Ser Asp Thr His Leu Leu Gln Gly Gln Ser Leu Thr
Leu Thr Leu Glu130 135 140Ser Pro Pro Gly
Ser Ser Pro Ser Val Gln Cys Arg Ser Pro Arg Gly145 150
155 160Lys Asn Ile Gln Gly Gly Lys Thr Leu
Ser Val Ser Gln Leu Glu Leu165 170 175Gln
Asp Ser Gly Thr Trp Thr Cys Thr Val Leu Gln Asn Gln Lys Lys180
185 190Val Glu Phe Lys Ile Asp Ile Val Val Leu Ala
Phe Gln Lys Ala Ser195 200 205Ser Ile Val
Tyr Lys Lys Glu Gly Glu Gln Val Glu Phe Ser Phe Pro210
215 220Leu Ala Phe Thr Val Glu Lys Leu Thr Gly Ser Gly
Glu Leu Trp Trp225 230 235
240Gln Ala Glu Arg Ala Ser Ser Ser Lys Ser Trp Ile Thr Phe Asp Leu245
250 255Lys Asn Lys Glu Val Ser Val Lys Arg
Val Thr Gln Asp Pro Lys Leu260 265 270Gln
Met Gly Lys Lys Leu Pro Leu His Leu Thr Leu Pro Gln Ala Leu275
280 285Pro Gln Tyr Ala Gly Ser Gly Asn Leu Thr Leu
Ala Leu Glu Ala Lys290 295 300Thr Gly Lys
Leu His Gln Glu Val Asn Leu Val Val Met Arg Ala Thr305
310 315 320Gln Leu Gln Lys Asn Leu Thr
Cys Glu Val Trp Gly Pro Thr Ser Pro325 330
335Lys Leu Met Leu Ser Leu Lys Leu Glu Asn Lys Glu Ala Lys Val Ser340
345 350Lys Arg Glu Lys Ala Val Trp Val Leu
Asn Pro Glu Ala Gly Met Trp355 360 365Gln
Cys Leu Leu Ser Asp Ser Gly Gln Val Leu Leu Glu Ser Asn Ile370
375 380Lys Val Leu Pro Thr Trp Ser Thr Pro Val Gln
Pro Gly Cys Val Ser385 390 395
400Gly Asp Thr Ile Val Met Thr Ser Gly Gly Pro Arg Thr Val Ala
Glu405 410 415Leu Glu Gly Lys Pro Phe Thr
Ala Leu Ile Arg Gly Ser Gly Tyr Pro420 425
430Cys Pro Ser Gly Phe Phe Arg Thr Cys Glu Arg Asp Val Tyr Asp Leu435
440 445Arg Thr Arg Glu Gly His Cys Leu Arg
Leu Thr His Asp His Arg Val450 455 460Leu
Val Met Asp Gly Gly Leu Glu Trp Arg Ala Ala Gly Glu Leu Glu465
470 475 480Arg Gly Asp Arg Leu Val
Met Asp Asp Ala Ala Gly Glu Phe Pro Ala485 490
495Leu Ala Thr Phe Arg Gly Leu Arg Gly Ala Gly Arg Gln Asp Val
Tyr500 505 510Asp Ala Thr Val Tyr Gly Ala
Ser Ala Phe Thr Ala Asn Gly Phe Ile515 520
525Val His Ala Cys Gly Glu Gln Pro Gly Thr Gly Leu Asn Ser Gly Leu530
535 540Thr Thr Asn Pro Gly Val Ser Ala Trp
Gln Val Asn Thr Ala Tyr Thr545 550 555
560Ala Gly Gln Leu Val Thr Tyr Asn Gly Lys Thr Tyr Lys Cys
Leu Gln565 570 575Pro His Thr Ser Leu Ala
Gly Trp Glu Pro Ser Asn Val Pro Ala Leu580 585
590Trp Gln Leu Gln595141571PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 141Lys Lys Val
Val Leu Gly Lys Lys Gly Asp Thr Val Glu Leu Thr Cys1 5
10 15Thr Ala Ser Gln Lys Lys Ser Ile Gln
Phe His Trp Lys Asn Ser Asn20 25 30Gln
Ile Lys Ile Leu Gly Asn Gln Gly Ser Phe Leu Thr Lys Gly Pro35
40 45Ser Lys Leu Asn Asp Arg Ala Asp Ser Arg Arg
Ser Leu Trp Asp Gln50 55 60Gly Asn Phe
Pro Leu Ile Ile Lys Asn Leu Lys Ile Glu Asp Ser Asp65 70
75 80Thr Tyr Ile Cys Glu Val Glu Asp
Gln Lys Glu Glu Val Gln Leu Leu85 90
95Val Phe Gly Leu Thr Ala Asn Ser Asp Thr His Leu Leu Gln Gly Gln100
105 110Ser Leu Thr Leu Thr Leu Glu Ser Pro Pro
Gly Ser Ser Pro Ser Val115 120 125Gln Cys
Arg Ser Pro Arg Gly Lys Asn Ile Gln Gly Gly Lys Thr Leu130
135 140Ser Val Ser Gln Leu Glu Leu Gln Asp Ser Gly Thr
Trp Thr Cys Thr145 150 155
160Val Leu Gln Asn Gln Lys Lys Val Glu Phe Lys Ile Asp Ile Val Val165
170 175Leu Ala Phe Gln Lys Ala Ser Ser Ile
Val Tyr Lys Lys Glu Gly Glu180 185 190Gln
Val Glu Phe Ser Phe Pro Leu Ala Phe Thr Val Glu Lys Leu Thr195
200 205Gly Ser Gly Glu Leu Trp Trp Gln Ala Glu Arg
Ala Ser Ser Ser Lys210 215 220Ser Trp Ile
Thr Phe Asp Leu Lys Asn Lys Glu Val Ser Val Lys Arg225
230 235 240Val Thr Gln Asp Pro Lys Leu
Gln Met Gly Lys Lys Leu Pro Leu His245 250
255Leu Thr Leu Pro Gln Ala Leu Pro Gln Tyr Ala Gly Ser Gly Asn Leu260
265 270Thr Leu Ala Leu Glu Ala Lys Thr Gly
Lys Leu His Gln Glu Val Asn275 280 285Leu
Val Val Met Arg Ala Thr Gln Leu Gln Lys Asn Leu Thr Cys Glu290
295 300Val Trp Gly Pro Thr Ser Pro Lys Leu Met Leu
Ser Leu Lys Leu Glu305 310 315
320Asn Lys Glu Ala Lys Val Ser Lys Arg Glu Lys Ala Val Trp Val
Leu325 330 335Asn Pro Glu Ala Gly Met Trp
Gln Cys Leu Leu Ser Asp Ser Gly Gln340 345
350Val Leu Leu Glu Ser Asn Ile Lys Val Leu Pro Thr Trp Ser Thr Pro355
360 365Val Gln Pro Gly Cys Val Ser Gly Asp
Thr Ile Val Met Thr Ser Gly370 375 380Gly
Pro Arg Thr Val Ala Glu Leu Glu Gly Lys Pro Phe Thr Ala Leu385
390 395 400Ile Arg Gly Ser Gly Tyr
Pro Cys Pro Ser Gly Phe Phe Arg Thr Cys405 410
415Glu Arg Asp Val Tyr Asp Leu Arg Thr Arg Glu Gly His Cys Leu
Arg420 425 430Leu Thr His Asp His Arg Val
Leu Val Met Asp Gly Gly Leu Glu Trp435 440
445Arg Ala Ala Gly Glu Leu Glu Arg Gly Asp Arg Leu Val Met Asp Asp450
455 460Ala Ala Gly Glu Phe Pro Ala Leu Ala
Thr Phe Arg Gly Leu Arg Gly465 470 475
480Ala Gly Arg Gln Asp Val Tyr Asp Ala Thr Val Tyr Gly Ala
Ser Ala485 490 495Phe Thr Ala Asn Gly Phe
Ile Val His Ala Cys Gly Glu Gln Pro Gly500 505
510Thr Gly Leu Asn Ser Gly Leu Thr Thr Asn Pro Gly Val Ser Ala
Trp515 520 525Gln Val Asn Thr Ala Tyr Thr
Ala Gly Gln Leu Val Thr Tyr Asn Gly530 535
540Lys Thr Tyr Lys Cys Leu Gln Pro His Thr Ser Leu Ala Gly Trp Glu545
550 555 560Pro Ser Asn Val
Pro Ala Leu Trp Gln Leu Gln565 570142372PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 142Lys Lys Val Val Leu Gly Lys Lys Gly Asp Thr Val Glu Leu
Thr Cys1 5 10 15Thr Ala
Ser Gln Lys Lys Ser Ile Gln Phe His Trp Lys Asn Ser Asn20
25 30Gln Ile Lys Ile Leu Gly Asn Gln Gly Ser Phe Leu
Thr Lys Gly Pro35 40 45Ser Lys Leu Asn
Asp Arg Ala Asp Ser Arg Arg Ser Leu Trp Asp Gln50 55
60Gly Asn Phe Pro Leu Ile Ile Lys Asn Leu Lys Ile Glu Asp
Ser Asp65 70 75 80Thr
Tyr Ile Cys Glu Val Glu Asp Gln Lys Glu Glu Val Gln Leu Leu85
90 95Val Phe Gly Leu Thr Ala Asn Ser Asp Thr His
Leu Leu Gln Gly Gln100 105 110Ser Leu Thr
Leu Thr Leu Glu Ser Pro Pro Gly Ser Ser Pro Ser Val115
120 125Gln Cys Arg Ser Pro Arg Gly Lys Asn Ile Gln Gly
Gly Lys Thr Leu130 135 140Ser Val Ser Gln
Leu Glu Leu Gln Asp Ser Gly Thr Trp Thr Cys Thr145 150
155 160Val Leu Gln Asn Gln Lys Lys Val Glu
Phe Lys Ile Asp Ile Val Val165 170 175Leu
Ala Phe Gln Lys Ala Ser Ser Ile Val Tyr Lys Lys Glu Gly Glu180
185 190Gln Val Glu Phe Ser Phe Pro Leu Ala Phe Thr
Val Glu Lys Leu Thr195 200 205Gly Ser Gly
Glu Leu Trp Trp Gln Ala Glu Arg Ala Ser Ser Ser Lys210
215 220Ser Trp Ile Thr Phe Asp Leu Lys Asn Lys Glu Val
Ser Val Lys Arg225 230 235
240Val Thr Gln Asp Pro Lys Leu Gln Met Gly Lys Lys Leu Pro Leu His245
250 255Leu Thr Leu Pro Gln Ala Leu Pro Gln
Tyr Ala Gly Ser Gly Asn Leu260 265 270Thr
Leu Ala Leu Glu Ala Lys Thr Gly Lys Leu His Gln Glu Val Asn275
280 285Leu Val Val Met Arg Ala Thr Gln Leu Gln Lys
Asn Leu Thr Cys Glu290 295 300Val Trp Gly
Pro Thr Ser Pro Lys Leu Met Leu Ser Leu Lys Leu Glu305
310 315 320Asn Lys Glu Ala Lys Val Ser
Lys Arg Glu Lys Ala Val Trp Val Leu325 330
335Asn Pro Glu Ala Gly Met Trp Gln Cys Leu Leu Ser Asp Ser Gly Gln340
345 350Val Leu Leu Glu Ser Asn Ile Lys Val
Leu Pro Thr Trp Ser Thr Pro355 360 365Val
Gln Pro Xaa370143373PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 143Lys Lys Val Val Leu Gly Lys Lys
Gly Asp Thr Val Glu Leu Thr Cys1 5 10
15Thr Ala Ser Gln Lys Lys Ser Ile Gln Phe His Trp Lys Asn
Ser Asn20 25 30Gln Ile Lys Ile Leu Gly
Asn Gln Gly Ser Phe Leu Thr Lys Gly Pro35 40
45Ser Lys Leu Asn Asp Arg Ala Asp Ser Arg Arg Ser Leu Trp Asp Gln50
55 60Gly Asn Phe Pro Leu Ile Ile Lys Asn
Leu Lys Ile Glu Asp Ser Asp65 70 75
80Thr Tyr Ile Cys Glu Val Glu Asp Gln Lys Glu Glu Val Gln
Leu Leu85 90 95Val Phe Gly Leu Thr Ala
Asn Ser Asp Thr His Leu Leu Gln Gly Gln100 105
110Ser Leu Thr Leu Thr Leu Glu Ser Pro Pro Gly Ser Ser Pro Ser
Val115 120 125Gln Cys Arg Ser Pro Arg Gly
Lys Asn Ile Gln Gly Gly Lys Thr Leu130 135
140Ser Val Ser Gln Leu Glu Leu Gln Asp Ser Gly Thr Trp Thr Cys Thr145
150 155 160Val Leu Gln Asn
Gln Lys Lys Val Glu Phe Lys Ile Asp Ile Val Val165 170
175Leu Ala Phe Gln Lys Ala Ser Ser Ile Val Tyr Lys Lys Glu
Gly Glu180 185 190Gln Val Glu Phe Ser Phe
Pro Leu Ala Phe Thr Val Glu Lys Leu Thr195 200
205Gly Ser Gly Glu Leu Trp Trp Gln Ala Glu Arg Ala Ser Ser Ser
Lys210 215 220Ser Trp Ile Thr Phe Asp Leu
Lys Asn Lys Glu Val Ser Val Lys Arg225 230
235 240Val Thr Gln Asp Pro Lys Leu Gln Met Gly Lys Lys
Leu Pro Leu His245 250 255Leu Thr Leu Pro
Gln Ala Leu Pro Gln Tyr Ala Gly Ser Gly Asn Leu260 265
270Thr Leu Ala Leu Glu Ala Lys Thr Gly Lys Leu His Gln Glu
Val Asn275 280 285Leu Val Val Met Arg Ala
Thr Gln Leu Gln Lys Asn Leu Thr Cys Glu290 295
300Val Trp Gly Pro Thr Ser Pro Lys Leu Met Leu Ser Leu Lys Leu
Glu305 310 315 320Asn Lys
Glu Ala Lys Val Ser Lys Arg Glu Lys Ala Val Trp Val Leu325
330 335Asn Pro Glu Ala Gly Met Trp Gln Cys Leu Leu Ser
Asp Ser Gly Gln340 345 350Val Leu Leu Glu
Ser Asn Ile Lys Val Leu Pro Thr Trp Ser Thr Pro355 360
365Val Gln Pro Gly Xaa3701441364DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 144aagcttgaat tcccaccatg gcttgggtgt ggaccttgct tttcctgatg
gcagctgccc 60aaagtatcca agcacagatc cagttggtcc agtctggacc tgagctgaag
aagcctggag 120agacagtcaa gatctcctgc aaggcttcag gatatacctt cacacactat
ggaatgaact 180gggtgaagca ggctccagga aagggtttaa agtggatggg ctggataaac
acctacactg 240gagagccaac atatgctgat gacttcaagg aacactttgc cttctctttg
gaaacctctg 300ccagcactgt ctttttgcag atcaacaacc tcaaaaatga ggacacggcc
acatatttct 360gtgcaagaga acggggggat gctatggact actggggtca gggaacctcc
gtcaccgtct 420cctcagcctc caccaagggg ccatcggtct tccccctggc accctcctcc
aagagcacct 480ctgggggcac agcggccctg ggctgcctgg tcaaggacta cttccccgaa
ccggtgacgg 540tgtcgtggaa ctcaggcgcc ctgaccagcg gcgtgcacac cttcccggct
gtcctacagt 600cctcaggact ctactccctc agcagcgtgg tgaccgtgcc ctccagcagc
ttgggcaccc 660agacctacat ctgcaacgtg aatcacaagc ccagcaacac caaggtggac
aagaaagttg 720agcccaaatc ttgtgacaaa actcacacag ggtgcgtatc cggtgacacc
attgtaatga 780ctagtggcgg gccccgcact gtggctgaac tggagggcaa accgttcacc
gcactgattc 840gcggctctgg ctacccatgc ccctcaggtt tcttccgcac ctgtgaacgt
gacgtatatg 900atctgcgtac acgtgagggt cattgcttac gtttgaccca tgatcaccgt
gttctggtga 960tggatggtgg cctggaatgg cgtgccgcgg gtgaactgga acgcggcgac
cgcctggtga 1020tggatgatgc agctggcgag tttccggcac tggcaacctt ccgtggcctg
cgtggcgctg 1080gccgccagga tgtttatgac gctactgttt acggtgctag cgcattcact
gctaatggct 1140tcattgtaca cgcatgtggc gagcagcccg ggaccggtct gaactcaggc
ctcacgacaa 1200atcctggtgt atccgcttgg caggtcaaca cagcttatac tgcgggacaa
ttggtcacat 1260ataacggcaa gacgtataaa tgtttgcagc cccacacctc cttggcagga
tgggaaccat 1320ccaacgttcc tgccttgtgg cagcttcaat gactcgagcg gccg
1364145444PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 145Met Ala Trp Val Trp Thr Leu Leu
Phe Leu Met Ala Ala Ala Gln Ser1 5 10
15Ile Gln Ala Gln Ile Gln Leu Val Gln Ser Gly Pro Glu Leu
Lys Lys20 25 30Pro Gly Glu Thr Val Lys
Ile Ser Cys Lys Ala Ser Gly Tyr Thr Phe35 40
45Thr His Tyr Gly Met Asn Trp Val Lys Gln Ala Pro Gly Lys Gly Leu50
55 60Lys Trp Met Gly Trp Ile Asn Thr Tyr
Thr Gly Glu Pro Thr Tyr Ala65 70 75
80Asp Asp Phe Lys Glu His Phe Ala Phe Ser Leu Glu Thr Ser
Ala Ser85 90 95Thr Val Phe Leu Gln Ile
Asn Asn Leu Lys Asn Glu Asp Thr Ala Thr100 105
110Tyr Phe Cys Ala Arg Glu Arg Gly Asp Ala Met Asp Tyr Trp Gly
Gln115 120 125Gly Thr Ser Val Thr Val Ser
Ser Ala Ser Thr Lys Gly Pro Ser Val130 135
140Phe Pro Leu Ala Pro Ser Ser Lys Ser Thr Ser Gly Gly Thr Ala Ala145
150 155 160Leu Gly Cys Leu
Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser165 170
175Trp Asn Ser Gly Ala Leu Thr Ser Gly Val His Thr Phe Pro
Ala Val180 185 190Leu Gln Ser Ser Gly Leu
Tyr Ser Leu Ser Ser Val Val Thr Val Pro195 200
205Ser Ser Ser Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val Asn His
Lys210 215 220Pro Ser Asn Thr Lys Val Asp
Lys Lys Val Glu Pro Lys Ser Cys Asp225 230
235 240Lys Thr His Thr Gly Cys Val Ser Gly Asp Thr Ile
Val Met Thr Ser245 250 255Gly Gly Pro Arg
Thr Val Ala Glu Leu Glu Gly Lys Pro Phe Thr Ala260 265
270Leu Ile Arg Gly Ser Gly Tyr Pro Cys Pro Ser Gly Phe Phe
Arg Thr275 280 285Cys Glu Arg Asp Val Tyr
Asp Leu Arg Thr Arg Glu Gly His Cys Leu290 295
300Arg Leu Thr His Asp His Arg Val Leu Val Met Asp Gly Gly Leu
Glu305 310 315 320Trp Arg
Ala Ala Gly Glu Leu Glu Arg Gly Asp Arg Leu Val Met Asp325
330 335Asp Ala Ala Gly Glu Phe Pro Ala Leu Ala Thr Phe
Arg Gly Leu Arg340 345 350Gly Ala Gly Arg
Gln Asp Val Tyr Asp Ala Thr Val Tyr Gly Ala Ser355 360
365Ala Phe Thr Ala Asn Gly Phe Ile Val His Ala Cys Gly Glu
Gln Pro370 375 380Gly Thr Gly Leu Asn Ser
Gly Leu Thr Thr Asn Pro Gly Val Ser Ala385 390
395 400Trp Gln Val Asn Thr Ala Tyr Thr Ala Gly Gln
Leu Val Thr Tyr Asn405 410 415Gly Lys Thr
Tyr Lys Cys Leu Gln Pro His Thr Ser Leu Ala Gly Trp420
425 430Glu Pro Ser Asn Val Pro Ala Leu Trp Gln Leu Gln435
440146425PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 146Gln Ile Gln Leu Val Gln
Ser Gly Pro Glu Leu Lys Lys Pro Gly Glu1 5
10 15Thr Val Lys Ile Ser Cys Lys Ala Ser Gly Tyr Thr
Phe Thr His Tyr20 25 30Gly Met Asn Trp
Val Lys Gln Ala Pro Gly Lys Gly Leu Lys Trp Met35 40
45Gly Trp Ile Asn Thr Tyr Thr Gly Glu Pro Thr Tyr Ala Asp
Asp Phe50 55 60Lys Glu His Phe Ala Phe
Ser Leu Glu Thr Ser Ala Ser Thr Val Phe65 70
75 80Leu Gln Ile Asn Asn Leu Lys Asn Glu Asp Thr
Ala Thr Tyr Phe Cys85 90 95Ala Arg Glu
Arg Gly Asp Ala Met Asp Tyr Trp Gly Gln Gly Thr Ser100
105 110Val Thr Val Ser Ser Ala Ser Thr Lys Gly Pro Ser
Val Phe Pro Leu115 120 125Ala Pro Ser Ser
Lys Ser Thr Ser Gly Gly Thr Ala Ala Leu Gly Cys130 135
140Leu Val Lys Asp Tyr Phe Pro Glu Pro Val Thr Val Ser Trp
Asn Ser145 150 155 160Gly
Ala Leu Thr Ser Gly Val His Thr Phe Pro Ala Val Leu Gln Ser165
170 175Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr
Val Pro Ser Ser Ser180 185 190Leu Gly Thr
Gln Thr Tyr Ile Cys Asn Val Asn His Lys Pro Ser Asn195
200 205Thr Lys Val Asp Lys Lys Val Glu Pro Lys Ser Cys
Asp Lys Thr His210 215 220Thr Gly Cys Val
Ser Gly Asp Thr Ile Val Met Thr Ser Gly Gly Pro225 230
235 240Arg Thr Val Ala Glu Leu Glu Gly Lys
Pro Phe Thr Ala Leu Ile Arg245 250 255Gly
Ser Gly Tyr Pro Cys Pro Ser Gly Phe Phe Arg Thr Cys Glu Arg260
265 270Asp Val Tyr Asp Leu Arg Thr Arg Glu Gly His
Cys Leu Arg Leu Thr275 280 285His Asp His
Arg Val Leu Val Met Asp Gly Gly Leu Glu Trp Arg Ala290
295 300Ala Gly Glu Leu Glu Arg Gly Asp Arg Leu Val Met
Asp Asp Ala Ala305 310 315
320Gly Glu Phe Pro Ala Leu Ala Thr Phe Arg Gly Leu Arg Gly Ala Gly325
330 335Arg Gln Asp Val Tyr Asp Ala Thr Val
Tyr Gly Ala Ser Ala Phe Thr340 345 350Ala
Asn Gly Phe Ile Val His Ala Cys Gly Glu Gln Pro Gly Thr Gly355
360 365Leu Asn Ser Gly Leu Thr Thr Asn Pro Gly Val
Ser Ala Trp Gln Val370 375 380Asn Thr Ala
Tyr Thr Ala Gly Gln Leu Val Thr Tyr Asn Gly Lys Thr385
390 395 400Tyr Lys Cys Leu Gln Pro His
Thr Ser Leu Ala Gly Trp Glu Pro Ser405 410
415Asn Val Pro Ala Leu Trp Gln Leu Gln420
425147226PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 147Gln Ile Gln Leu Val Gln Ser Gly Pro Glu
Leu Lys Lys Pro Gly Glu1 5 10
15Thr Val Lys Ile Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr His Tyr20
25 30Gly Met Asn Trp Val Lys Gln Ala Pro
Gly Lys Gly Leu Lys Trp Met35 40 45Gly
Trp Ile Asn Thr Tyr Thr Gly Glu Pro Thr Tyr Ala Asp Asp Phe50
55 60Lys Glu His Phe Ala Phe Ser Leu Glu Thr Ser
Ala Ser Thr Val Phe65 70 75
80Leu Gln Ile Asn Asn Leu Lys Asn Glu Asp Thr Ala Thr Tyr Phe Cys85
90 95Ala Arg Glu Arg Gly Asp Ala Met Asp
Tyr Trp Gly Gln Gly Thr Ser100 105 110Val
Thr Val Ser Ser Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu115
120 125Ala Pro Ser Ser Lys Ser Thr Ser Gly Gly Thr
Ala Ala Leu Gly Cys130 135 140Leu Val Lys
Asp Tyr Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser145
150 155 160Gly Ala Leu Thr Ser Gly Val
His Thr Phe Pro Ala Val Leu Gln Ser165 170
175Ser Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val Pro Ser Ser Ser180
185 190Leu Gly Thr Gln Thr Tyr Ile Cys Asn
Val Asn His Lys Pro Ser Asn195 200 205Thr
Lys Val Asp Lys Lys Val Glu Pro Lys Ser Cys Asp Lys Thr His210
215 220Thr Xaa225148227PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 148Gln Ile Gln Leu Val Gln Ser Gly Pro Glu Leu Lys Lys Pro
Gly Glu1 5 10 15Thr Val
Lys Ile Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr His Tyr20
25 30Gly Met Asn Trp Val Lys Gln Ala Pro Gly Lys Gly
Leu Lys Trp Met35 40 45Gly Trp Ile Asn
Thr Tyr Thr Gly Glu Pro Thr Tyr Ala Asp Asp Phe50 55
60Lys Glu His Phe Ala Phe Ser Leu Glu Thr Ser Ala Ser Thr
Val Phe65 70 75 80Leu
Gln Ile Asn Asn Leu Lys Asn Glu Asp Thr Ala Thr Tyr Phe Cys85
90 95Ala Arg Glu Arg Gly Asp Ala Met Asp Tyr Trp
Gly Gln Gly Thr Ser100 105 110Val Thr Val
Ser Ser Ala Ser Thr Lys Gly Pro Ser Val Phe Pro Leu115
120 125Ala Pro Ser Ser Lys Ser Thr Ser Gly Gly Thr Ala
Ala Leu Gly Cys130 135 140Leu Val Lys Asp
Tyr Phe Pro Glu Pro Val Thr Val Ser Trp Asn Ser145 150
155 160Gly Ala Leu Thr Ser Gly Val His Thr
Phe Pro Ala Val Leu Gln Ser165 170 175Ser
Gly Leu Tyr Ser Leu Ser Ser Val Val Thr Val Pro Ser Ser Ser180
185 190Leu Gly Thr Gln Thr Tyr Ile Cys Asn Val Asn
His Lys Pro Ser Asn195 200 205Thr Lys Val
Asp Lys Lys Val Glu Pro Lys Ser Cys Asp Lys Thr His210
215 220Thr Gly Xaa225149731DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 149aagcttgaat tcccaccatg aagtcacaga cccaggtctt cgtatttcta
ctgctctgtg 60tgtctggtgc tcatgggagt attgtgatga cccagactcc caaattcctg
cttgtatcag 120caggagacag ggttaccata acctgcacgg ccagtcagag tgtgagtaat
gatgtagttt 180ggtaccaaca gaagccaggg cagtctccta aaatgctgat gtattctgca
ttcaatcgct 240acactggagt ccctgatcgt ttcactggca gaggatacgg gacggatttc
actttcacca 300tcagctctgt gcaggctgaa gacctggcag tttatttctg tcagcaggat
tataactctc 360ctcggacgtt cggtggaggc accaagctgg agatcaaacg aactgtggct
gcaccatctg 420tcttcatctt cccgccatct gatgagcagt tgaaatctgg aactgcctct
gttgtgtgcc 480tgctgaataa cttctatccc agagaggcca aagtacagtg gaaggtggat
aacgccctcc 540aatcgggtaa ctcccaggag agtgtcacag agcaggacag caaggacagc
acctacagcc 600tcagcagcac cctgacgctg agcaaagcag actacgagaa acacaaagtc
tacgcctgcg 660aagtcaccca tcagggcctg agctcgcccg tcacaaagag cttcaacagg
ggagagtgtt 720agtgactcga g
731150234PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 150Met Lys Ser Gln Thr Gln Val Phe
Val Phe Leu Leu Leu Cys Val Ser1 5 10
15Gly Ala His Gly Ser Ile Val Met Thr Gln Thr Pro Lys Phe
Leu Leu20 25 30Val Ser Ala Gly Asp Arg
Val Thr Ile Thr Cys Thr Ala Ser Gln Ser35 40
45Val Ser Asn Asp Val Val Trp Tyr Gln Gln Lys Pro Gly Gln Ser Pro50
55 60Lys Met Leu Met Tyr Ser Ala Phe Asn
Arg Tyr Thr Gly Val Pro Asp65 70 75
80Arg Phe Thr Gly Arg Gly Tyr Gly Thr Asp Phe Thr Phe Thr
Ile Ser85 90 95Ser Val Gln Ala Glu Asp
Leu Ala Val Tyr Phe Cys Gln Gln Asp Tyr100 105
110Asn Ser Pro Arg Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys
Arg115 120 125Thr Val Ala Ala Pro Ser Val
Phe Ile Phe Pro Pro Ser Asp Glu Gln130 135
140Leu Lys Ser Gly Thr Ala Ser Val Val Cys Leu Leu Asn Asn Phe Tyr145
150 155 160Pro Arg Glu Ala
Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser165 170
175Gly Asn Ser Gln Glu Ser Val Thr Glu Gln Asp Ser Lys Asp
Ser Thr180 185 190Tyr Ser Leu Ser Ser Thr
Leu Thr Leu Ser Lys Ala Asp Tyr Glu Lys195 200
205His Lys Val Tyr Ala Cys Glu Val Thr His Gln Gly Leu Ser Ser
Pro210 215 220Val Thr Lys Ser Phe Asn Arg
Gly Glu Cys225 230151214PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 151Ser Ile Val
Met Thr Gln Thr Pro Lys Phe Leu Leu Val Ser Ala Gly1 5
10 15Asp Arg Val Thr Ile Thr Cys Thr Ala
Ser Gln Ser Val Ser Asn Asp20 25 30Val
Val Trp Tyr Gln Gln Lys Pro Gly Gln Ser Pro Lys Met Leu Met35
40 45Tyr Ser Ala Phe Asn Arg Tyr Thr Gly Val Pro
Asp Arg Phe Thr Gly50 55 60Arg Gly Tyr
Gly Thr Asp Phe Thr Phe Thr Ile Ser Ser Val Gln Ala65 70
75 80Glu Asp Leu Ala Val Tyr Phe Cys
Gln Gln Asp Tyr Asn Ser Pro Arg85 90
95Thr Phe Gly Gly Gly Thr Lys Leu Glu Ile Lys Arg Thr Val Ala Ala100
105 110Pro Ser Val Phe Ile Phe Pro Pro Ser Asp
Glu Gln Leu Lys Ser Gly115 120 125Thr Ala
Ser Val Val Cys Leu Leu Asn Asn Phe Tyr Pro Arg Glu Ala130
135 140Lys Val Gln Trp Lys Val Asp Asn Ala Leu Gln Ser
Gly Asn Ser Gln145 150 155
160Glu Ser Val Thr Glu Gln Asp Ser Lys Asp Ser Thr Tyr Ser Leu Ser165
170 175Ser Thr Leu Thr Leu Ser Lys Ala Asp
Tyr Glu Lys His Lys Val Tyr180 185 190Ala
Cys Glu Val Thr His Gln Gly Leu Ser Ser Pro Val Thr Lys Ser195
200 205Phe Asn Arg Gly Glu
Cys2101521268DNAArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 152aagcttgaat tcccaccatg ggcctctcca
ccgtgcctga cctgctgctg ccgctggtgc 60tcctggagct gttggtggga atatacccct
caggggttat tggactggtc cctcacctag 120gggacaggga gaagagagat agtgtgtgtc
cccaaggaaa atatatccac cctcaaaata 180attcgatttg ctgtaccaag tgccacaaag
gaacctactt gtacaatgac tgtccaggcc 240cggggcagga tacggactgc agggagtgtg
agagcggctc cttcaccgct tcagaaaacc 300acctcagaca ctgcctcagc tgctccaaat
gccgaaagga aatgggtcag gtggagatct 360cttcttgcac agtggaccgg gacaccgtgt
gtggctgcag gaagaaccag taccggcatt 420attggagtga aaaccttttc cagtgcttca
attgcagcct ctgcctcaat gggaccgtgc 480acctctcctg ccaggagaaa cagaacaccg
tgtgcacctg ccatgcaggt ttctttctaa 540gagaaaacga gtgtgtctcc tgtagtaact
gtaagaaaag cctggagtgc acgaagttgt 600gcctacccca gattgagaat gttaagggca
ctgaggactc aggcaccaca gtggggtgcg 660tatccggtga caccattgta atgactagtg
gcgggccccg cactgtggct gaactggagg 720gcaaaccgtt caccgcactg attcgcggct
ctggctaccc atgcccctca ggtttcttcc 780gcacctgtga acgtgacgta tatgatctgc
gtacacgtga gggtcattgc ttacgtttga 840cccatgatca ccgtgttctg gtgatggatg
gtggcctgga atggcgtgcc gcgggtgaac 900tggaacgcgg cgaccgcctg gtgatggatg
atgcagctgg cgagtttccg gcactggcaa 960ccttccgtgg cctgcgtggc gctggccgcc
aggatgttta tgacgctact gtttacggtg 1020ctagcgcatt cactgctaat ggcttcattg
tacacgcatg tggcgagcag cccgggaccg 1080gtctgaactc aggcctcacg acaaatcctg
gtgtatccgc ttggcaggtc aacacagctt 1140atactgcggg acaattggtc acatataacg
gcaagacgta taaatgtttg cagccccaca 1200cctccttggc aggatgggaa ccatccaacg
ttcctgcctt gtggcagctt caatgactcg 1260agcggccg
1268153411PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 153Met Gly Leu Ser Thr Val Pro Asp Leu Leu Leu Pro Leu Val
Leu Leu1 5 10 15Glu Leu
Leu Val Gly Ile Tyr Pro Ser Gly Val Ile Gly Leu Val Pro20
25 30His Leu Gly Asp Arg Glu Lys Arg Asp Ser Val Cys
Pro Gln Gly Lys35 40 45Tyr Ile His Pro
Gln Asn Asn Ser Ile Cys Cys Thr Lys Cys His Lys50 55
60Gly Thr Tyr Leu Tyr Asn Asp Cys Pro Gly Pro Gly Gln Asp
Thr Asp65 70 75 80Cys
Arg Glu Cys Glu Ser Gly Ser Phe Thr Ala Ser Glu Asn His Leu85
90 95Arg His Cys Leu Ser Cys Ser Lys Cys Arg Lys
Glu Met Gly Gln Val100 105 110Glu Ile Ser
Ser Cys Thr Val Asp Arg Asp Thr Val Cys Gly Cys Arg115
120 125Lys Asn Gln Tyr Arg His Tyr Trp Ser Glu Asn Leu
Phe Gln Cys Phe130 135 140Asn Cys Ser Leu
Cys Leu Asn Gly Thr Val His Leu Ser Cys Gln Glu145 150
155 160Lys Gln Asn Thr Val Cys Thr Cys His
Ala Gly Phe Phe Leu Arg Glu165 170 175Asn
Glu Cys Val Ser Cys Ser Asn Cys Lys Lys Ser Leu Glu Cys Thr180
185 190Lys Leu Cys Leu Pro Gln Ile Glu Asn Val Lys
Gly Thr Glu Asp Ser195 200 205Gly Thr Thr
Gly Cys Val Ser Gly Asp Thr Ile Val Met Thr Ser Gly210
215 220Gly Pro Arg Thr Val Ala Glu Leu Glu Gly Lys Pro
Phe Thr Ala Leu225 230 235
240Ile Arg Gly Ser Gly Tyr Pro Cys Pro Ser Gly Phe Phe Arg Thr Cys245
250 255Glu Arg Asp Val Tyr Asp Leu Arg Thr
Arg Glu Gly His Cys Leu Arg260 265 270Leu
Thr His Asp His Arg Val Leu Val Met Asp Gly Gly Leu Glu Trp275
280 285Arg Ala Ala Gly Glu Leu Glu Arg Gly Asp Arg
Leu Val Met Asp Asp290 295 300Ala Ala Gly
Glu Phe Pro Ala Leu Ala Thr Phe Arg Gly Leu Arg Gly305
310 315 320Ala Gly Arg Gln Asp Val Tyr
Asp Ala Thr Val Tyr Gly Ala Ser Ala325 330
335Phe Thr Ala Asn Gly Phe Ile Val His Ala Cys Gly Glu Gln Pro Gly340
345 350Thr Gly Leu Asn Ser Gly Leu Thr Thr
Asn Pro Gly Val Ser Ala Trp355 360 365Gln
Val Asn Thr Ala Tyr Thr Ala Gly Gln Leu Val Thr Tyr Asn Gly370
375 380Lys Thr Tyr Lys Cys Leu Gln Pro His Thr Ser
Leu Ala Gly Trp Glu385 390 395
400Pro Ser Asn Val Pro Ala Leu Trp Gln Leu Gln405
410154390PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 154Ile Tyr Pro Ser Gly Val Ile Gly Leu Val
Pro His Leu Gly Asp Arg1 5 10
15Glu Lys Arg Asp Ser Val Cys Pro Gln Gly Lys Tyr Ile His Pro Gln20
25 30Asn Asn Ser Ile Cys Cys Thr Lys Cys
His Lys Gly Thr Tyr Leu Tyr35 40 45Asn
Asp Cys Pro Gly Pro Gly Gln Asp Thr Asp Cys Arg Glu Cys Glu50
55 60Ser Gly Ser Phe Thr Ala Ser Glu Asn His Leu
Arg His Cys Leu Ser65 70 75
80Cys Ser Lys Cys Arg Lys Glu Met Gly Gln Val Glu Ile Ser Ser Cys85
90 95Thr Val Asp Arg Asp Thr Val Cys Gly
Cys Arg Lys Asn Gln Tyr Arg100 105 110His
Tyr Trp Ser Glu Asn Leu Phe Gln Cys Phe Asn Cys Ser Leu Cys115
120 125Leu Asn Gly Thr Val His Leu Ser Cys Gln Glu
Lys Gln Asn Thr Val130 135 140Cys Thr Cys
His Ala Gly Phe Phe Leu Arg Glu Asn Glu Cys Val Ser145
150 155 160Cys Ser Asn Cys Lys Lys Ser
Leu Glu Cys Thr Lys Leu Cys Leu Pro165 170
175Gln Ile Glu Asn Val Lys Gly Thr Glu Asp Ser Gly Thr Thr Gly Cys180
185 190Val Ser Gly Asp Thr Ile Val Met Thr
Ser Gly Gly Pro Arg Thr Val195 200 205Ala
Glu Leu Glu Gly Lys Pro Phe Thr Ala Leu Ile Arg Gly Ser Gly210
215 220Tyr Pro Cys Pro Ser Gly Phe Phe Arg Thr Cys
Glu Arg Asp Val Tyr225 230 235
240Asp Leu Arg Thr Arg Glu Gly His Cys Leu Arg Leu Thr His Asp
His245 250 255Arg Val Leu Val Met Asp Gly
Gly Leu Glu Trp Arg Ala Ala Gly Glu260 265
270Leu Glu Arg Gly Asp Arg Leu Val Met Asp Asp Ala Ala Gly Glu Phe275
280 285Pro Ala Leu Ala Thr Phe Arg Gly Leu
Arg Gly Ala Gly Arg Gln Asp290 295 300Val
Tyr Asp Ala Thr Val Tyr Gly Ala Ser Ala Phe Thr Ala Asn Gly305
310 315 320Phe Ile Val His Ala Cys
Gly Glu Gln Pro Gly Thr Gly Leu Asn Ser325 330
335Gly Leu Thr Thr Asn Pro Gly Val Ser Ala Trp Gln Val Asn Thr
Ala340 345 350Tyr Thr Ala Gly Gln Leu Val
Thr Tyr Asn Gly Lys Thr Tyr Lys Cys355 360
365Leu Gln Pro His Thr Ser Leu Ala Gly Trp Glu Pro Ser Asn Val Pro370
375 380Ala Leu Trp Gln Leu Gln385
390155191PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 155Ile Tyr Pro Ser Gly Val Ile Gly Leu Val
Pro His Leu Gly Asp Arg1 5 10
15Glu Lys Arg Asp Ser Val Cys Pro Gln Gly Lys Tyr Ile His Pro Gln20
25 30Asn Asn Ser Ile Cys Cys Thr Lys Cys
His Lys Gly Thr Tyr Leu Tyr35 40 45Asn
Asp Cys Pro Gly Pro Gly Gln Asp Thr Asp Cys Arg Glu Cys Glu50
55 60Ser Gly Ser Phe Thr Ala Ser Glu Asn His Leu
Arg His Cys Leu Ser65 70 75
80Cys Ser Lys Cys Arg Lys Glu Met Gly Gln Val Glu Ile Ser Ser Cys85
90 95Thr Val Asp Arg Asp Thr Val Cys Gly
Cys Arg Lys Asn Gln Tyr Arg100 105 110His
Tyr Trp Ser Glu Asn Leu Phe Gln Cys Phe Asn Cys Ser Leu Cys115
120 125Leu Asn Gly Thr Val His Leu Ser Cys Gln Glu
Lys Gln Asn Thr Val130 135 140Cys Thr Cys
His Ala Gly Phe Phe Leu Arg Glu Asn Glu Cys Val Ser145
150 155 160Cys Ser Asn Cys Lys Lys Ser
Leu Glu Cys Thr Lys Leu Cys Leu Pro165 170
175Gln Ile Glu Asn Val Lys Gly Thr Glu Asp Ser Gly Thr Thr Xaa180
185 190156192PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 156Ile Tyr Pro
Ser Gly Val Ile Gly Leu Val Pro His Leu Gly Asp Arg1 5
10 15Glu Lys Arg Asp Ser Val Cys Pro Gln
Gly Lys Tyr Ile His Pro Gln20 25 30Asn
Asn Ser Ile Cys Cys Thr Lys Cys His Lys Gly Thr Tyr Leu Tyr35
40 45Asn Asp Cys Pro Gly Pro Gly Gln Asp Thr Asp
Cys Arg Glu Cys Glu50 55 60Ser Gly Ser
Phe Thr Ala Ser Glu Asn His Leu Arg His Cys Leu Ser65 70
75 80Cys Ser Lys Cys Arg Lys Glu Met
Gly Gln Val Glu Ile Ser Ser Cys85 90
95Thr Val Asp Arg Asp Thr Val Cys Gly Cys Arg Lys Asn Gln Tyr Arg100
105 110His Tyr Trp Ser Glu Asn Leu Phe Gln Cys
Phe Asn Cys Ser Leu Cys115 120 125Leu Asn
Gly Thr Val His Leu Ser Cys Gln Glu Lys Gln Asn Thr Val130
135 140Cys Thr Cys His Ala Gly Phe Phe Leu Arg Glu Asn
Glu Cys Val Ser145 150 155
160Cys Ser Asn Cys Lys Lys Ser Leu Glu Cys Thr Lys Leu Cys Leu Pro165
170 175Gln Ile Glu Asn Val Lys Gly Thr Glu
Asp Ser Gly Thr Thr Gly Xaa180 185
1901571403DNAArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 157aagcttgaat tcccaccatg gcgcccgtcg
ccgtctgggc cgcgctggcc gtcggactgg 60agctctgggc tgcggcgcac gccttgcccg
cccaggtggc atttacaccc tacgccccgg 120agcccgggag cacatgccgg ctcagagaat
actatgacca gacagctcag atgtgctgca 180gcaaatgctc gccgggccaa catgcaaaag
tcttctgtac caagacctcg gacaccgtgt 240gtgactcctg tgaggacagc acatacaccc
agctctggaa ctgggttccc gagtgcttga 300gctgtggctc ccgctgtagc tctgaccagg
tggaaactca agcctgcact cgggaacaga 360accgcatctg cacctgcagg cccggctggt
actgcgcgct gagcaagcag gaggggtgcc 420ggctgtgcgc gccgctgcgc aagtgccgcc
cgggcttcgg cgtggccaga ccaggaactg 480aaacatcaga cgtggtgtgc aagccctgtg
ccccggggac gttctccaac acgacttcat 540ccacggatat ttgcaggccc caccagatct
gtaacgtggt ggccatccct gggaatgcaa 600gcatggatgc agtctgcacg tccacgtccc
ccacccggag tatggcccca ggggcagtac 660acttacccca gccagtgtcc acacgatccc
aacacacgca gccaactcca gaacccagca 720ctgctccaag cacctccttc ctgctcccaa
tgggccccag ccccccagct gaagggagca 780ctggcgacgg gtgcgtatcc ggtgacacca
ttgtaatgac tagtggcggg ccccgcactg 840tggctgaact ggagggcaaa ccgttcaccg
cactgattcg cggctctggc tacccatgcc 900cctcaggttt cttccgcacc tgtgaacgtg
acgtatatga tctgcgtaca cgtgagggtc 960attgcttacg tttgacccat gatcaccgtg
ttctggtgat ggatggtggc ctggaatggc 1020gtgccgcggg tgaactggaa cgcggcgacc
gcctggtgat ggatgatgca gctggcgagt 1080ttccggcact ggcaaccttc cgtggcctgc
gtggcgctgg ccgccaggat gtttatgacg 1140ctactgttta cggtgctagc gcattcactg
ctaatggctt cattgtacac gcatgtggcg 1200agcagcccgg gaccggtctg aactcaggcc
tcacgacaaa tcctggtgta tccgcttggc 1260aggtcaacac agcttatact gcgggacaat
tggtcacata taacggcaag acgtataaat 1320gtttgcagcc ccacacctcc ttggcaggat
gggaaccatc caacgttcct gccttgtggc 1380agcttcaatg actcgagcgg ccg
1403158457PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 158Met Ala Pro Val Ala Val Trp Ala Ala Leu Ala Val Gly Leu
Glu Leu1 5 10 15Trp Ala
Ala Ala His Ala Leu Pro Ala Gln Val Ala Phe Thr Pro Tyr20
25 30Ala Pro Glu Pro Gly Ser Thr Cys Arg Leu Arg Glu
Tyr Tyr Asp Gln35 40 45Thr Ala Gln Met
Cys Cys Ser Lys Cys Ser Pro Gly Gln His Ala Lys50 55
60Val Phe Cys Thr Lys Thr Ser Asp Thr Val Cys Asp Ser Cys
Glu Asp65 70 75 80Ser
Thr Tyr Thr Gln Leu Trp Asn Trp Val Pro Glu Cys Leu Ser Cys85
90 95Gly Ser Arg Cys Ser Ser Asp Gln Val Glu Thr
Gln Ala Cys Thr Arg100 105 110Glu Gln Asn
Arg Ile Cys Thr Cys Arg Pro Gly Trp Tyr Cys Ala Leu115
120 125Ser Lys Gln Glu Gly Cys Arg Leu Cys Ala Pro Leu
Arg Lys Cys Arg130 135 140Pro Gly Phe Gly
Val Ala Arg Pro Gly Thr Glu Thr Ser Asp Val Val145 150
155 160Cys Lys Pro Cys Ala Pro Gly Thr Phe
Ser Asn Thr Thr Ser Ser Thr165 170 175Asp
Ile Cys Arg Pro His Gln Ile Cys Asn Val Val Ala Ile Pro Gly180
185 190Asn Ala Ser Met Asp Ala Val Cys Thr Ser Thr
Ser Pro Thr Arg Ser195 200 205Met Ala Pro
Gly Ala Val His Leu Pro Gln Pro Val Ser Thr Arg Ser210
215 220Gln His Thr Gln Pro Thr Pro Glu Pro Ser Thr Ala
Pro Ser Thr Ser225 230 235
240Phe Leu Leu Pro Met Gly Pro Ser Pro Pro Ala Glu Gly Ser Thr Gly245
250 255Asp Gly Cys Val Ser Gly Asp Thr Ile
Val Met Thr Ser Gly Gly Pro260 265 270Arg
Thr Val Ala Glu Leu Glu Gly Lys Pro Phe Thr Ala Leu Ile Arg275
280 285Gly Ser Gly Tyr Pro Cys Pro Ser Gly Phe Phe
Arg Thr Cys Glu Arg290 295 300Asp Val Tyr
Asp Leu Arg Thr Arg Glu Gly His Cys Leu Arg Leu Thr305
310 315 320His Asp His Arg Val Leu Val
Met Asp Gly Gly Leu Glu Trp Arg Ala325 330
335Ala Gly Glu Leu Glu Arg Gly Asp Arg Leu Val Met Asp Asp Ala Ala340
345 350Gly Glu Phe Pro Ala Leu Ala Thr Phe
Arg Gly Leu Arg Gly Ala Gly355 360 365Arg
Gln Asp Val Tyr Asp Ala Thr Val Tyr Gly Ala Ser Ala Phe Thr370
375 380Ala Asn Gly Phe Ile Val His Ala Cys Gly Glu
Gln Pro Gly Thr Gly385 390 395
400Leu Asn Ser Gly Leu Thr Thr Asn Pro Gly Val Ser Ala Trp Gln
Val405 410 415Asn Thr Ala Tyr Thr Ala Gly
Gln Leu Val Thr Tyr Asn Gly Lys Thr420 425
430Tyr Lys Cys Leu Gln Pro His Thr Ser Leu Ala Gly Trp Glu Pro Ser435
440 445Asn Val Pro Ala Leu Trp Gln Leu
Gln450 455159435PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 159Leu Pro Ala
Gln Val Ala Phe Thr Pro Tyr Ala Pro Glu Pro Gly Ser1 5
10 15Thr Cys Arg Leu Arg Glu Tyr Tyr Asp
Gln Thr Ala Gln Met Cys Cys20 25 30Ser
Lys Cys Ser Pro Gly Gln His Ala Lys Val Phe Cys Thr Lys Thr35
40 45Ser Asp Thr Val Cys Asp Ser Cys Glu Asp Ser
Thr Tyr Thr Gln Leu50 55 60Trp Asn Trp
Val Pro Glu Cys Leu Ser Cys Gly Ser Arg Cys Ser Ser65 70
75 80Asp Gln Val Glu Thr Gln Ala Cys
Thr Arg Glu Gln Asn Arg Ile Cys85 90
95Thr Cys Arg Pro Gly Trp Tyr Cys Ala Leu Ser Lys Gln Glu Gly Cys100
105 110Arg Leu Cys Ala Pro Leu Arg Lys Cys Arg
Pro Gly Phe Gly Val Ala115 120 125Arg Pro
Gly Thr Glu Thr Ser Asp Val Val Cys Lys Pro Cys Ala Pro130
135 140Gly Thr Phe Ser Asn Thr Thr Ser Ser Thr Asp Ile
Cys Arg Pro His145 150 155
160Gln Ile Cys Asn Val Val Ala Ile Pro Gly Asn Ala Ser Met Asp Ala165
170 175Val Cys Thr Ser Thr Ser Pro Thr Arg
Ser Met Ala Pro Gly Ala Val180 185 190His
Leu Pro Gln Pro Val Ser Thr Arg Ser Gln His Thr Gln Pro Thr195
200 205Pro Glu Pro Ser Thr Ala Pro Ser Thr Ser Phe
Leu Leu Pro Met Gly210 215 220Pro Ser Pro
Pro Ala Glu Gly Ser Thr Gly Asp Gly Cys Val Ser Gly225
230 235 240Asp Thr Ile Val Met Thr Ser
Gly Gly Pro Arg Thr Val Ala Glu Leu245 250
255Glu Gly Lys Pro Phe Thr Ala Leu Ile Arg Gly Ser Gly Tyr Pro Cys260
265 270Pro Ser Gly Phe Phe Arg Thr Cys Glu
Arg Asp Val Tyr Asp Leu Arg275 280 285Thr
Arg Glu Gly His Cys Leu Arg Leu Thr His Asp His Arg Val Leu290
295 300Val Met Asp Gly Gly Leu Glu Trp Arg Ala Ala
Gly Glu Leu Glu Arg305 310 315
320Gly Asp Arg Leu Val Met Asp Asp Ala Ala Gly Glu Phe Pro Ala
Leu325 330 335Ala Thr Phe Arg Gly Leu Arg
Gly Ala Gly Arg Gln Asp Val Tyr Asp340 345
350Ala Thr Val Tyr Gly Ala Ser Ala Phe Thr Ala Asn Gly Phe Ile Val355
360 365His Ala Cys Gly Glu Gln Pro Gly Thr
Gly Leu Asn Ser Gly Leu Thr370 375 380Thr
Asn Pro Gly Val Ser Ala Trp Gln Val Asn Thr Ala Tyr Thr Ala385
390 395 400Gly Gln Leu Val Thr Tyr
Asn Gly Lys Thr Tyr Lys Cys Leu Gln Pro405 410
415His Thr Ser Leu Ala Gly Trp Glu Pro Ser Asn Val Pro Ala Leu
Trp420 425 430Gln Leu
Gln435160236PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 160Leu Pro Ala Gln Val Ala Phe Thr Pro Tyr
Ala Pro Glu Pro Gly Ser1 5 10
15Thr Cys Arg Leu Arg Glu Tyr Tyr Asp Gln Thr Ala Gln Met Cys Cys20
25 30Ser Lys Cys Ser Pro Gly Gln His Ala
Lys Val Phe Cys Thr Lys Thr35 40 45Ser
Asp Thr Val Cys Asp Ser Cys Glu Asp Ser Thr Tyr Thr Gln Leu50
55 60Trp Asn Trp Val Pro Glu Cys Leu Ser Cys Gly
Ser Arg Cys Ser Ser65 70 75
80Asp Gln Val Glu Thr Gln Ala Cys Thr Arg Glu Gln Asn Arg Ile Cys85
90 95Thr Cys Arg Pro Gly Trp Tyr Cys Ala
Leu Ser Lys Gln Glu Gly Cys100 105 110Arg
Leu Cys Ala Pro Leu Arg Lys Cys Arg Pro Gly Phe Gly Val Ala115
120 125Arg Pro Gly Thr Glu Thr Ser Asp Val Val Cys
Lys Pro Cys Ala Pro130 135 140Gly Thr Phe
Ser Asn Thr Thr Ser Ser Thr Asp Ile Cys Arg Pro His145
150 155 160Gln Ile Cys Asn Val Val Ala
Ile Pro Gly Asn Ala Ser Met Asp Ala165 170
175Val Cys Thr Ser Thr Ser Pro Thr Arg Ser Met Ala Pro Gly Ala Val180
185 190His Leu Pro Gln Pro Val Ser Thr Arg
Ser Gln His Thr Gln Pro Thr195 200 205Pro
Glu Pro Ser Thr Ala Pro Ser Thr Ser Phe Leu Leu Pro Met Gly210
215 220Pro Ser Pro Pro Ala Glu Gly Ser Thr Gly Asp
Xaa225 230 235161237PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 161Leu Pro Ala Gln Val Ala Phe Thr Pro Tyr Ala Pro Glu Pro
Gly Ser1 5 10 15Thr Cys
Arg Leu Arg Glu Tyr Tyr Asp Gln Thr Ala Gln Met Cys Cys20
25 30Ser Lys Cys Ser Pro Gly Gln His Ala Lys Val Phe
Cys Thr Lys Thr35 40 45Ser Asp Thr Val
Cys Asp Ser Cys Glu Asp Ser Thr Tyr Thr Gln Leu50 55
60Trp Asn Trp Val Pro Glu Cys Leu Ser Cys Gly Ser Arg Cys
Ser Ser65 70 75 80Asp
Gln Val Glu Thr Gln Ala Cys Thr Arg Glu Gln Asn Arg Ile Cys85
90 95Thr Cys Arg Pro Gly Trp Tyr Cys Ala Leu Ser
Lys Gln Glu Gly Cys100 105 110Arg Leu Cys
Ala Pro Leu Arg Lys Cys Arg Pro Gly Phe Gly Val Ala115
120 125Arg Pro Gly Thr Glu Thr Ser Asp Val Val Cys Lys
Pro Cys Ala Pro130 135 140Gly Thr Phe Ser
Asn Thr Thr Ser Ser Thr Asp Ile Cys Arg Pro His145 150
155 160Gln Ile Cys Asn Val Val Ala Ile Pro
Gly Asn Ala Ser Met Asp Ala165 170 175Val
Cys Thr Ser Thr Ser Pro Thr Arg Ser Met Ala Pro Gly Ala Val180
185 190His Leu Pro Gln Pro Val Ser Thr Arg Ser Gln
His Thr Gln Pro Thr195 200 205Pro Glu Pro
Ser Thr Ala Pro Ser Thr Ser Phe Leu Leu Pro Met Gly210
215 220Pro Ser Pro Pro Ala Glu Gly Ser Thr Gly Asp Gly
Xaa225 230 2351621499DNAArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 162aagcttgaat tcccaccatg gcgcccgtcg ccgtctgggc cgcgctggcc
gtcggactgg 60agctctgggc tgcggcgcac gccttgcccg cccaggtggc atttacaccc
tacgccccgg 120agcccgggag cacatgccgg ctcagagaat actatgacca gacagctcag
atgtgctgca 180gcaaatgctc gccgggccaa catgcaaaag tcttctgtac caagacctcg
gacaccgtgt 240gtgactcctg tgaggacagc acatacaccc agctctggaa ctgggttccc
gagtgcttga 300gctgtggctc ccgctgtagc tctgaccagg tggaaactca agcctgcact
cgggaacaga 360accgcatctg cacctgcagg cccggctggt actgcgcgct gagcaagcag
gaggggtgcc 420ggctgtgcgc gccgctgcgc aagtgccgcc cgggcttcgg cgtggccaga
ccaggaactg 480aaacatcaga cgtggtgtgc aagccctgtg ccccggggac gttctccaac
acgacttcat 540ccacggatat ttgcaggccc caccagatct gtaacgtggt ggccatccct
gggaatgcaa 600gcatggatgc agtctgcacg tccacgtccc ccacccggag tatggcccca
ggggcagtac 660acttacccca gccagtgtcc acacgatccc aacacacgca gccaactcca
gaacccagca 720ctgctccaag cacctccttc ctgctcccaa tgggccccag ccccccagct
gaagggagca 780ctggcgacga gcccaaatct tgtgacaaaa ctcacacatg cccaccgtgc
ccagcacctg 840aactcctggg ggggccctca gtcttcctct tccccccaaa acccaaggac
accctcatga 900tctcccggac ccctgaggtc acatgcgtgg tggtggacgt gagccacgaa
gaccctgagg 960tcaagttcaa ctggtacgtg gacggcgtgg aggtgcataa tgccaagaca
aagccgcggg 1020aggagcagta caacagcacg taccgtgtgg tcagcgtcct caccgtcctg
caccaggact 1080ggctgaatgg caaggagtac aagtgcaagg tctccaacaa agccctccca
gcccccatcg 1140agaaaaccat ctccaaagcc aaagggcagc cccgagaacc acaggtgtac
accctgcccc 1200catcccggga tgagctgacc aagaaccagg tcagcctgac ctgcctggtc
aaaggcttct 1260atcccagcga catcgccgtg gagtgggaga gcaatgggca gccggagaac
aactacaaga 1320ccacgcctcc cgtgctggac tccgacggct ccttcttcct ctacagcaag
ctcaccgtgg 1380acaagagcag gtggcagcag gggaacgtct tctcatgctc cgtgatgcat
gaggctctgc 1440acaaccacta cacgcagaag agcctctccc tgtctccggg taaatgactc
gagcggccg 1499163489PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 163Met Ala Pro Val Ala Val
Trp Ala Ala Leu Ala Val Gly Leu Glu Leu1 5
10 15Trp Ala Ala Ala His Ala Leu Pro Ala Gln Val Ala
Phe Thr Pro Tyr20 25 30Ala Pro Glu Pro
Gly Ser Thr Cys Arg Leu Arg Glu Tyr Tyr Asp Gln35 40
45Thr Ala Gln Met Cys Cys Ser Lys Cys Ser Pro Gly Gln His
Ala Lys50 55 60Val Phe Cys Thr Lys Thr
Ser Asp Thr Val Cys Asp Ser Cys Glu Asp65 70
75 80Ser Thr Tyr Thr Gln Leu Trp Asn Trp Val Pro
Glu Cys Leu Ser Cys85 90 95Gly Ser Arg
Cys Ser Ser Asp Gln Val Glu Thr Gln Ala Cys Thr Arg100
105 110Glu Gln Asn Arg Ile Cys Thr Cys Arg Pro Gly Trp
Tyr Cys Ala Leu115 120 125Ser Lys Gln Glu
Gly Cys Arg Leu Cys Ala Pro Leu Arg Lys Cys Arg130 135
140Pro Gly Phe Gly Val Ala Arg Pro Gly Thr Glu Thr Ser Asp
Val Val145 150 155 160Cys
Lys Pro Cys Ala Pro Gly Thr Phe Ser Asn Thr Thr Ser Ser Thr165
170 175Asp Ile Cys Arg Pro His Gln Ile Cys Asn Val
Val Ala Ile Pro Gly180 185 190Asn Ala Ser
Met Asp Ala Val Cys Thr Ser Thr Ser Pro Thr Arg Ser195
200 205Met Ala Pro Gly Ala Val His Leu Pro Gln Pro Val
Ser Thr Arg Ser210 215 220Gln His Thr Gln
Pro Thr Pro Glu Pro Ser Thr Ala Pro Ser Thr Ser225 230
235 240Phe Leu Leu Pro Met Gly Pro Ser Pro
Pro Ala Glu Gly Ser Thr Gly245 250 255Asp
Glu Pro Lys Ser Cys Asp Lys Thr His Thr Cys Pro Pro Cys Pro260
265 270Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe
Leu Phe Pro Pro Lys275 280 285Pro Lys Asp
Thr Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys Val290
295 300Val Val Asp Val Ser His Glu Asp Pro Glu Val Lys
Phe Asn Trp Tyr305 310 315
320Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu Glu325
330 335Gln Tyr Asn Ser Thr Tyr Arg Val Val
Ser Val Leu Thr Val Leu His340 345 350Gln
Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys355
360 365Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser
Lys Ala Lys Gly Gln370 375 380Pro Arg Glu
Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu Leu385
390 395 400Thr Lys Asn Gln Val Ser Leu
Thr Cys Leu Val Lys Gly Phe Tyr Pro405 410
415Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn420
425 430Tyr Lys Thr Thr Pro Pro Val Leu Asp
Ser Asp Gly Ser Phe Phe Leu435 440 445Tyr
Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn Val450
455 460Phe Ser Cys Ser Val Met His Glu Ala Leu His
Asn His Tyr Thr Gln465 470 475
480Lys Ser Leu Ser Leu Ser Pro Gly Lys485164467PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 164Leu Pro Ala Gln Val Ala Phe Thr Pro Tyr Ala Pro Glu Pro
Gly Ser1 5 10 15Thr Cys
Arg Leu Arg Glu Tyr Tyr Asp Gln Thr Ala Gln Met Cys Cys20
25 30Ser Lys Cys Ser Pro Gly Gln His Ala Lys Val Phe
Cys Thr Lys Thr35 40 45Ser Asp Thr Val
Cys Asp Ser Cys Glu Asp Ser Thr Tyr Thr Gln Leu50 55
60Trp Asn Trp Val Pro Glu Cys Leu Ser Cys Gly Ser Arg Cys
Ser Ser65 70 75 80Asp
Gln Val Glu Thr Gln Ala Cys Thr Arg Glu Gln Asn Arg Ile Cys85
90 95Thr Cys Arg Pro Gly Trp Tyr Cys Ala Leu Ser
Lys Gln Glu Gly Cys100 105 110Arg Leu Cys
Ala Pro Leu Arg Lys Cys Arg Pro Gly Phe Gly Val Ala115
120 125Arg Pro Gly Thr Glu Thr Ser Asp Val Val Cys Lys
Pro Cys Ala Pro130 135 140Gly Thr Phe Ser
Asn Thr Thr Ser Ser Thr Asp Ile Cys Arg Pro His145 150
155 160Gln Ile Cys Asn Val Val Ala Ile Pro
Gly Asn Ala Ser Met Asp Ala165 170 175Val
Cys Thr Ser Thr Ser Pro Thr Arg Ser Met Ala Pro Gly Ala Val180
185 190His Leu Pro Gln Pro Val Ser Thr Arg Ser Gln
His Thr Gln Pro Thr195 200 205Pro Glu Pro
Ser Thr Ala Pro Ser Thr Ser Phe Leu Leu Pro Met Gly210
215 220Pro Ser Pro Pro Ala Glu Gly Ser Thr Gly Asp Glu
Pro Lys Ser Cys225 230 235
240Asp Lys Thr His Thr Cys Pro Pro Cys Pro Ala Pro Glu Leu Leu Gly245
250 255Gly Pro Ser Val Phe Leu Phe Pro Pro
Lys Pro Lys Asp Thr Leu Met260 265 270Ile
Ser Arg Thr Pro Glu Val Thr Cys Val Val Val Asp Val Ser His275
280 285Glu Asp Pro Glu Val Lys Phe Asn Trp Tyr Val
Asp Gly Val Glu Val290 295 300His Asn Ala
Lys Thr Lys Pro Arg Glu Glu Gln Tyr Asn Ser Thr Tyr305
310 315 320Arg Val Val Ser Val Leu Thr
Val Leu His Gln Asp Trp Leu Asn Gly325 330
335Lys Glu Tyr Lys Cys Lys Val Ser Asn Lys Ala Leu Pro Ala Pro Ile340
345 350Glu Lys Thr Ile Ser Lys Ala Lys Gly
Gln Pro Arg Glu Pro Gln Val355 360 365Tyr
Thr Leu Pro Pro Ser Arg Asp Glu Leu Thr Lys Asn Gln Val Ser370
375 380Leu Thr Cys Leu Val Lys Gly Phe Tyr Pro Ser
Asp Ile Ala Val Glu385 390 395
400Trp Glu Ser Asn Gly Gln Pro Glu Asn Asn Tyr Lys Thr Thr Pro
Pro405 410 415Val Leu Asp Ser Asp Gly Ser
Phe Phe Leu Tyr Ser Lys Leu Thr Val420 425
430Asp Lys Ser Arg Trp Gln Gln Gly Asn Val Phe Ser Cys Ser Val Met435
440 445His Glu Ala Leu His Asn His Tyr Thr
Gln Lys Ser Leu Ser Leu Ser450 455 460Pro
Gly Lys4651652906DNAArtificial Sequenceartificial construct relating to
Homo Sapiens immunoglobulin 165aagcttgaat tcccaccatg gtcagctact
gggacaccgg ggtcctgctg tgcgcgctgc 60tcagctgtct gcttctcaca ggatctagtt
caggttcaaa attaaaagat cctgaactga 120gtttaaaagg cacccagcac atcatgcaag
caggccagac actgcatctc caatgcaggg 180gggaagcagc ccataaatgg tctttgcctg
aaatggtgag taaggaaagc gaaaggctga 240gcataactaa atctgcctgt ggaagaaatg
gcaaacaatt ctgcagtact ttaaccttga 300acacagctca agcaaaccac actggcttct
acagctgcaa atatctagct gtacctactt 360caaagaagaa ggaaacagaa tctgcaatct
atatatttat tagtgataca ggtagacctt 420tcgtagagat gtacagtgaa atccccgaaa
ttatacacat gactgaagga agggagctcg 480tcattccctg ccgggttacg tcacctaaca
tcactgttac tttaaaaaag tttccacttg 540acactttgat ccctgatgga aaacgcataa
tctgggacag tagaaagggc ttcatcatat 600caaatgcaac gtacaaagaa atagggcttc
tgacctgtga agcaacagtc aatgggcatt 660tgtataagac aaactatctc acacatcgac
aaaccaatac aatcatagat gtccaaataa 720gcacaccacg cccagtcaaa ttacttagag
gccatactct tgtcctcaat tgtactgcta 780ccactccctt gaacacgaga gttcaaatga
cctggagtta ccctgatgaa aaaaataaga 840gagcttccgt aaggcgacga attgaccaaa
gcaattccca tgccaacata ttctacagtg 900ttcttactat tgacaaaatg cagaacaaag
acaaaggact ttatacttgt cgtgtaagga 960gtggaccatc attcaaatct gttaacacct
cagtgcatat atatgataaa gcattcatca 1020ctgtgaaaca tcgaaaacag caggtgcttg
aaaccgtagc tggcaagcgg tcttaccggc 1080tctctatgaa agtgaaggca tttccctcgc
cggaagttgt atggttaaaa gatgggttac 1140ctgcgactga gaaatctgct cgctatttga
ctcgtggcta ctcgttaatt atcaaggacg 1200taactgaaga ggatgcaggg aattatacaa
tcttgctgag cataaaacag tcaaatgtgt 1260ttaaaaacct cactgccact ctaattgtca
atgtgaaacc ccagatttac gaaaaggccg 1320tgtcatcgtt tccagacccg gctctctacc
cactgggcag cagacaaatc ctgacttgta 1380ccgcatatgg tatccctcaa cctacaatca
agtggttctg gcacccctgt aaccataatc 1440attccgaagc aaggtgtgac ttttgttcca
ataatgaaga gtcctttatc ctggatgctg 1500acagcaacat gggaaacaga attgagagca
tcactcagcg catggcaata atagaaggaa 1560agaataagat ggctagcacc ttggttgtgg
ctgactctag aatttctgga atctacattt 1620gcatagcttc caataaagtt gggactgtgg
gaagaaacat tagcttttat atcacagatg 1680tgccaaatgg gtttcatgtt aacttggaaa
aaatgccgac ggaaggagag gacctgaaac 1740tgtcttgcac agttaacaag ttcttataca
gagacgttac ttggatttta ctgcggacag 1800ttaataacag aacaatgcac tacagtatta
gcaagcaaaa aatggccatc actaaggagc 1860actccatcac tcttaatctt accatcatga
atgtttccct gcaagattca ggcacctatg 1920cctgcagagc caggaatgta tacacagggg
aagaaatcct ccagaagaaa gaaattacaa 1980tcagagatca ggaagcacca tacctcctgc
gaaacctcag tgatcacaca gtggccatca 2040gcagttccac cactttagac tgtcatgcta
atggtgtccc cgagcctcag atcacttggt 2100ttaaaaacaa ccacaaaata caacaagagc
ctggaattat tttaggacca ggaagcagca 2160cgctgtttat tgaaagagtc acagaagagg
atgaaggtgt ctatcactgc aaagccacca 2220accagaaggg ctctgtggaa agttcagcat
acctcactgt tcaaggaacc tcggacaagt 2280ctaatctgga ggggtgcgta tccggtgaca
ccattgtaat gactagtggc gggccccgca 2340ctgtggctga actggagggc aaaccgttca
ccgcactgat tcgcggctct ggctacccat 2400gcccctcagg tttcttccgc acctgtgaac
gtgacgtata tgatctgcgt acacgtgagg 2460gtcattgctt acgtttgacc catgatcacc
gtgttctggt gatggatggt ggcctggaat 2520ggcgtgccgc gggtgaactg gaacgcggcg
accgcctggt gatggatgat gcagctggcg 2580agtttccggc actggcaacc ttccgtggcc
tgcgtggcgc tggccgccag gatgtttatg 2640acgctactgt ttacggtgct agcgcattca
ctgctaatgg cttcattgta cacgcatgtg 2700gcgagcagcc cgggaccggt ctgaactcag
gcctcacgac aaatcctggt gtatccgctt 2760ggcaggtcaa cacagcttat actgcgggac
aattggtcac atataacggc aagacgtata 2820aatgtttgca gccccacacc tccttggcag
gatgggaacc atccaacgtt cctgccttgt 2880ggcagcttca atgactcgag cggccg
2906166958PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 166Met Val Ser Tyr Trp Asp Thr Gly Val Leu Leu Cys Ala Leu
Leu Ser1 5 10 15Cys Leu
Leu Leu Thr Gly Ser Ser Ser Gly Ser Lys Leu Lys Asp Pro20
25 30Glu Leu Ser Leu Lys Gly Thr Gln His Ile Met Gln
Ala Gly Gln Thr35 40 45Leu His Leu Gln
Cys Arg Gly Glu Ala Ala His Lys Trp Ser Leu Pro50 55
60Glu Met Val Ser Lys Glu Ser Glu Arg Leu Ser Ile Thr Lys
Ser Ala65 70 75 80Cys
Gly Arg Asn Gly Lys Gln Phe Cys Ser Thr Leu Thr Leu Asn Thr85
90 95Ala Gln Ala Asn His Thr Gly Phe Tyr Ser Cys
Lys Tyr Leu Ala Val100 105 110Pro Thr Ser
Lys Lys Lys Glu Thr Glu Ser Ala Ile Tyr Ile Phe Ile115
120 125Ser Asp Thr Gly Arg Pro Phe Val Glu Met Tyr Ser
Glu Ile Pro Glu130 135 140Ile Ile His Met
Thr Glu Gly Arg Glu Leu Val Ile Pro Cys Arg Val145 150
155 160Thr Ser Pro Asn Ile Thr Val Thr Leu
Lys Lys Phe Pro Leu Asp Thr165 170 175Leu
Ile Pro Asp Gly Lys Arg Ile Ile Trp Asp Ser Arg Lys Gly Phe180
185 190Ile Ile Ser Asn Ala Thr Tyr Lys Glu Ile Gly
Leu Leu Thr Cys Glu195 200 205Ala Thr Val
Asn Gly His Leu Tyr Lys Thr Asn Tyr Leu Thr His Arg210
215 220Gln Thr Asn Thr Ile Ile Asp Val Gln Ile Ser Thr
Pro Arg Pro Val225 230 235
240Lys Leu Leu Arg Gly His Thr Leu Val Leu Asn Cys Thr Ala Thr Thr245
250 255Pro Leu Asn Thr Arg Val Gln Met Thr
Trp Ser Tyr Pro Asp Glu Lys260 265 270Asn
Lys Arg Ala Ser Val Arg Arg Arg Ile Asp Gln Ser Asn Ser His275
280 285Ala Asn Ile Phe Tyr Ser Val Leu Thr Ile Asp
Lys Met Gln Asn Lys290 295 300Asp Lys Gly
Leu Tyr Thr Cys Arg Val Arg Ser Gly Pro Ser Phe Lys305
310 315 320Ser Val Asn Thr Ser Val His
Ile Tyr Asp Lys Ala Phe Ile Thr Val325 330
335Lys His Arg Lys Gln Gln Val Leu Glu Thr Val Ala Gly Lys Arg Ser340
345 350Tyr Arg Leu Ser Met Lys Val Lys Ala
Phe Pro Ser Pro Glu Val Val355 360 365Trp
Leu Lys Asp Gly Leu Pro Ala Thr Glu Lys Ser Ala Arg Tyr Leu370
375 380Thr Arg Gly Tyr Ser Leu Ile Ile Lys Asp Val
Thr Glu Glu Asp Ala385 390 395
400Gly Asn Tyr Thr Ile Leu Leu Ser Ile Lys Gln Ser Asn Val Phe
Lys405 410 415Asn Leu Thr Ala Thr Leu Ile
Val Asn Val Lys Pro Gln Ile Tyr Glu420 425
430Lys Ala Val Ser Ser Phe Pro Asp Pro Ala Leu Tyr Pro Leu Gly Ser435
440 445Arg Gln Ile Leu Thr Cys Thr Ala Tyr
Gly Ile Pro Gln Pro Thr Ile450 455 460Lys
Trp Phe Trp His Pro Cys Asn His Asn His Ser Glu Ala Arg Cys465
470 475 480Asp Phe Cys Ser Asn Asn
Glu Glu Ser Phe Ile Leu Asp Ala Asp Ser485 490
495Asn Met Gly Asn Arg Ile Glu Ser Ile Thr Gln Arg Met Ala Ile
Ile500 505 510Glu Gly Lys Asn Lys Met Ala
Ser Thr Leu Val Val Ala Asp Ser Arg515 520
525Ile Ser Gly Ile Tyr Ile Cys Ile Ala Ser Asn Lys Val Gly Thr Val530
535 540Gly Arg Asn Ile Ser Phe Tyr Ile Thr
Asp Val Pro Asn Gly Phe His545 550 555
560Val Asn Leu Glu Lys Met Pro Thr Glu Gly Glu Asp Leu Lys
Leu Ser565 570 575Cys Thr Val Asn Lys Phe
Leu Tyr Arg Asp Val Thr Trp Ile Leu Leu580 585
590Arg Thr Val Asn Asn Arg Thr Met His Tyr Ser Ile Ser Lys Gln
Lys595 600 605Met Ala Ile Thr Lys Glu His
Ser Ile Thr Leu Asn Leu Thr Ile Met610 615
620Asn Val Ser Leu Gln Asp Ser Gly Thr Tyr Ala Cys Arg Ala Arg Asn625
630 635 640Val Tyr Thr Gly
Glu Glu Ile Leu Gln Lys Lys Glu Ile Thr Ile Arg645 650
655Asp Gln Glu Ala Pro Tyr Leu Leu Arg Asn Leu Ser Asp His
Thr Val660 665 670Ala Ile Ser Ser Ser Thr
Thr Leu Asp Cys His Ala Asn Gly Val Pro675 680
685Glu Pro Gln Ile Thr Trp Phe Lys Asn Asn His Lys Ile Gln Gln
Glu690 695 700Pro Gly Ile Ile Leu Gly Pro
Gly Ser Ser Thr Leu Phe Ile Glu Arg705 710
715 720Val Thr Glu Glu Asp Glu Gly Val Tyr His Cys Lys
Ala Thr Asn Gln725 730 735Lys Gly Ser Val
Glu Ser Ser Ala Tyr Leu Thr Val Gln Gly Thr Ser740 745
750Asp Lys Ser Asn Leu Glu Gly Cys Val Ser Gly Asp Thr Ile
Val Met755 760 765Thr Ser Gly Gly Pro Arg
Thr Val Ala Glu Leu Glu Gly Lys Pro Phe770 775
780Thr Ala Leu Ile Arg Gly Ser Gly Tyr Pro Cys Pro Ser Gly Phe
Phe785 790 795 800Arg Thr
Cys Glu Arg Asp Val Tyr Asp Leu Arg Thr Arg Glu Gly His805
810 815Cys Leu Arg Leu Thr His Asp His Arg Val Leu Val
Met Asp Gly Gly820 825 830Leu Glu Trp Arg
Ala Ala Gly Glu Leu Glu Arg Gly Asp Arg Leu Val835 840
845Met Asp Asp Ala Ala Gly Glu Phe Pro Ala Leu Ala Thr Phe
Arg Gly850 855 860Leu Arg Gly Ala Gly Arg
Gln Asp Val Tyr Asp Ala Thr Val Tyr Gly865 870
875 880Ala Ser Ala Phe Thr Ala Asn Gly Phe Ile Val
His Ala Cys Gly Glu885 890 895Gln Pro Gly
Thr Gly Leu Asn Ser Gly Leu Thr Thr Asn Pro Gly Val900
905 910Ser Ala Trp Gln Val Asn Thr Ala Tyr Thr Ala Gly
Gln Leu Val Thr915 920 925Tyr Asn Gly Lys
Thr Tyr Lys Cys Leu Gln Pro His Thr Ser Leu Ala930 935
940Gly Trp Glu Pro Ser Asn Val Pro Ala Leu Trp Gln Leu
Gln945 950 955167932PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 167Ser Lys Leu Lys Asp Pro Glu Leu Ser Leu Lys Gly Thr Gln
His Ile1 5 10 15Met Gln
Ala Gly Gln Thr Leu His Leu Gln Cys Arg Gly Glu Ala Ala20
25 30His Lys Trp Ser Leu Pro Glu Met Val Ser Lys Glu
Ser Glu Arg Leu35 40 45Ser Ile Thr Lys
Ser Ala Cys Gly Arg Asn Gly Lys Gln Phe Cys Ser50 55
60Thr Leu Thr Leu Asn Thr Ala Gln Ala Asn His Thr Gly Phe
Tyr Ser65 70 75 80Cys
Lys Tyr Leu Ala Val Pro Thr Ser Lys Lys Lys Glu Thr Glu Ser85
90 95Ala Ile Tyr Ile Phe Ile Ser Asp Thr Gly Arg
Pro Phe Val Glu Met100 105 110Tyr Ser Glu
Ile Pro Glu Ile Ile His Met Thr Glu Gly Arg Glu Leu115
120 125Val Ile Pro Cys Arg Val Thr Ser Pro Asn Ile Thr
Val Thr Leu Lys130 135 140Lys Phe Pro Leu
Asp Thr Leu Ile Pro Asp Gly Lys Arg Ile Ile Trp145 150
155 160Asp Ser Arg Lys Gly Phe Ile Ile Ser
Asn Ala Thr Tyr Lys Glu Ile165 170 175Gly
Leu Leu Thr Cys Glu Ala Thr Val Asn Gly His Leu Tyr Lys Thr180
185 190Asn Tyr Leu Thr His Arg Gln Thr Asn Thr Ile
Ile Asp Val Gln Ile195 200 205Ser Thr Pro
Arg Pro Val Lys Leu Leu Arg Gly His Thr Leu Val Leu210
215 220Asn Cys Thr Ala Thr Thr Pro Leu Asn Thr Arg Val
Gln Met Thr Trp225 230 235
240Ser Tyr Pro Asp Glu Lys Asn Lys Arg Ala Ser Val Arg Arg Arg Ile245
250 255Asp Gln Ser Asn Ser His Ala Asn Ile
Phe Tyr Ser Val Leu Thr Ile260 265 270Asp
Lys Met Gln Asn Lys Asp Lys Gly Leu Tyr Thr Cys Arg Val Arg275
280 285Ser Gly Pro Ser Phe Lys Ser Val Asn Thr Ser
Val His Ile Tyr Asp290 295 300Lys Ala Phe
Ile Thr Val Lys His Arg Lys Gln Gln Val Leu Glu Thr305
310 315 320Val Ala Gly Lys Arg Ser Tyr
Arg Leu Ser Met Lys Val Lys Ala Phe325 330
335Pro Ser Pro Glu Val Val Trp Leu Lys Asp Gly Leu Pro Ala Thr Glu340
345 350Lys Ser Ala Arg Tyr Leu Thr Arg Gly
Tyr Ser Leu Ile Ile Lys Asp355 360 365Val
Thr Glu Glu Asp Ala Gly Asn Tyr Thr Ile Leu Leu Ser Ile Lys370
375 380Gln Ser Asn Val Phe Lys Asn Leu Thr Ala Thr
Leu Ile Val Asn Val385 390 395
400Lys Pro Gln Ile Tyr Glu Lys Ala Val Ser Ser Phe Pro Asp Pro
Ala405 410 415Leu Tyr Pro Leu Gly Ser Arg
Gln Ile Leu Thr Cys Thr Ala Tyr Gly420 425
430Ile Pro Gln Pro Thr Ile Lys Trp Phe Trp His Pro Cys Asn His Asn435
440 445His Ser Glu Ala Arg Cys Asp Phe Cys
Ser Asn Asn Glu Glu Ser Phe450 455 460Ile
Leu Asp Ala Asp Ser Asn Met Gly Asn Arg Ile Glu Ser Ile Thr465
470 475 480Gln Arg Met Ala Ile Ile
Glu Gly Lys Asn Lys Met Ala Ser Thr Leu485 490
495Val Val Ala Asp Ser Arg Ile Ser Gly Ile Tyr Ile Cys Ile Ala
Ser500 505 510Asn Lys Val Gly Thr Val Gly
Arg Asn Ile Ser Phe Tyr Ile Thr Asp515 520
525Val Pro Asn Gly Phe His Val Asn Leu Glu Lys Met Pro Thr Glu Gly530
535 540Glu Asp Leu Lys Leu Ser Cys Thr Val
Asn Lys Phe Leu Tyr Arg Asp545 550 555
560Val Thr Trp Ile Leu Leu Arg Thr Val Asn Asn Arg Thr Met
His Tyr565 570 575Ser Ile Ser Lys Gln Lys
Met Ala Ile Thr Lys Glu His Ser Ile Thr580 585
590Leu Asn Leu Thr Ile Met Asn Val Ser Leu Gln Asp Ser Gly Thr
Tyr595 600 605Ala Cys Arg Ala Arg Asn Val
Tyr Thr Gly Glu Glu Ile Leu Gln Lys610 615
620Lys Glu Ile Thr Ile Arg Asp Gln Glu Ala Pro Tyr Leu Leu Arg Asn625
630 635 640Leu Ser Asp His
Thr Val Ala Ile Ser Ser Ser Thr Thr Leu Asp Cys645 650
655His Ala Asn Gly Val Pro Glu Pro Gln Ile Thr Trp Phe Lys
Asn Asn660 665 670His Lys Ile Gln Gln Glu
Pro Gly Ile Ile Leu Gly Pro Gly Ser Ser675 680
685Thr Leu Phe Ile Glu Arg Val Thr Glu Glu Asp Glu Gly Val Tyr
His690 695 700Cys Lys Ala Thr Asn Gln Lys
Gly Ser Val Glu Ser Ser Ala Tyr Leu705 710
715 720Thr Val Gln Gly Thr Ser Asp Lys Ser Asn Leu Glu
Gly Cys Val Ser725 730 735Gly Asp Thr Ile
Val Met Thr Ser Gly Gly Pro Arg Thr Val Ala Glu740 745
750Leu Glu Gly Lys Pro Phe Thr Ala Leu Ile Arg Gly Ser Gly
Tyr Pro755 760 765Cys Pro Ser Gly Phe Phe
Arg Thr Cys Glu Arg Asp Val Tyr Asp Leu770 775
780Arg Thr Arg Glu Gly His Cys Leu Arg Leu Thr His Asp His Arg
Val785 790 795 800Leu Val
Met Asp Gly Gly Leu Glu Trp Arg Ala Ala Gly Glu Leu Glu805
810 815Arg Gly Asp Arg Leu Val Met Asp Asp Ala Ala Gly
Glu Phe Pro Ala820 825 830Leu Ala Thr Phe
Arg Gly Leu Arg Gly Ala Gly Arg Gln Asp Val Tyr835 840
845Asp Ala Thr Val Tyr Gly Ala Ser Ala Phe Thr Ala Asn Gly
Phe Ile850 855 860Val His Ala Cys Gly Glu
Gln Pro Gly Thr Gly Leu Asn Ser Gly Leu865 870
875 880Thr Thr Asn Pro Gly Val Ser Ala Trp Gln Val
Asn Thr Ala Tyr Thr885 890 895Ala Gly Gln
Leu Val Thr Tyr Asn Gly Lys Thr Tyr Lys Cys Leu Gln900
905 910Pro His Thr Ser Leu Ala Gly Trp Glu Pro Ser Asn
Val Pro Ala Leu915 920 925Trp Gln Leu
Gln930168733PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 168Ser Lys Leu Lys Asp Pro Glu Leu Ser Leu
Lys Gly Thr Gln His Ile1 5 10
15Met Gln Ala Gly Gln Thr Leu His Leu Gln Cys Arg Gly Glu Ala Ala20
25 30His Lys Trp Ser Leu Pro Glu Met Val
Ser Lys Glu Ser Glu Arg Leu35 40 45Ser
Ile Thr Lys Ser Ala Cys Gly Arg Asn Gly Lys Gln Phe Cys Ser50
55 60Thr Leu Thr Leu Asn Thr Ala Gln Ala Asn His
Thr Gly Phe Tyr Ser65 70 75
80Cys Lys Tyr Leu Ala Val Pro Thr Ser Lys Lys Lys Glu Thr Glu Ser85
90 95Ala Ile Tyr Ile Phe Ile Ser Asp Thr
Gly Arg Pro Phe Val Glu Met100 105 110Tyr
Ser Glu Ile Pro Glu Ile Ile His Met Thr Glu Gly Arg Glu Leu115
120 125Val Ile Pro Cys Arg Val Thr Ser Pro Asn Ile
Thr Val Thr Leu Lys130 135 140Lys Phe Pro
Leu Asp Thr Leu Ile Pro Asp Gly Lys Arg Ile Ile Trp145
150 155 160Asp Ser Arg Lys Gly Phe Ile
Ile Ser Asn Ala Thr Tyr Lys Glu Ile165 170
175Gly Leu Leu Thr Cys Glu Ala Thr Val Asn Gly His Leu Tyr Lys Thr180
185 190Asn Tyr Leu Thr His Arg Gln Thr Asn
Thr Ile Ile Asp Val Gln Ile195 200 205Ser
Thr Pro Arg Pro Val Lys Leu Leu Arg Gly His Thr Leu Val Leu210
215 220Asn Cys Thr Ala Thr Thr Pro Leu Asn Thr Arg
Val Gln Met Thr Trp225 230 235
240Ser Tyr Pro Asp Glu Lys Asn Lys Arg Ala Ser Val Arg Arg Arg
Ile245 250 255Asp Gln Ser Asn Ser His Ala
Asn Ile Phe Tyr Ser Val Leu Thr Ile260 265
270Asp Lys Met Gln Asn Lys Asp Lys Gly Leu Tyr Thr Cys Arg Val Arg275
280 285Ser Gly Pro Ser Phe Lys Ser Val Asn
Thr Ser Val His Ile Tyr Asp290 295 300Lys
Ala Phe Ile Thr Val Lys His Arg Lys Gln Gln Val Leu Glu Thr305
310 315 320Val Ala Gly Lys Arg Ser
Tyr Arg Leu Ser Met Lys Val Lys Ala Phe325 330
335Pro Ser Pro Glu Val Val Trp Leu Lys Asp Gly Leu Pro Ala Thr
Glu340 345 350Lys Ser Ala Arg Tyr Leu Thr
Arg Gly Tyr Ser Leu Ile Ile Lys Asp355 360
365Val Thr Glu Glu Asp Ala Gly Asn Tyr Thr Ile Leu Leu Ser Ile Lys370
375 380Gln Ser Asn Val Phe Lys Asn Leu Thr
Ala Thr Leu Ile Val Asn Val385 390 395
400Lys Pro Gln Ile Tyr Glu Lys Ala Val Ser Ser Phe Pro Asp
Pro Ala405 410 415Leu Tyr Pro Leu Gly Ser
Arg Gln Ile Leu Thr Cys Thr Ala Tyr Gly420 425
430Ile Pro Gln Pro Thr Ile Lys Trp Phe Trp His Pro Cys Asn His
Asn435 440 445His Ser Glu Ala Arg Cys Asp
Phe Cys Ser Asn Asn Glu Glu Ser Phe450 455
460Ile Leu Asp Ala Asp Ser Asn Met Gly Asn Arg Ile Glu Ser Ile Thr465
470 475 480Gln Arg Met Ala
Ile Ile Glu Gly Lys Asn Lys Met Ala Ser Thr Leu485 490
495Val Val Ala Asp Ser Arg Ile Ser Gly Ile Tyr Ile Cys Ile
Ala Ser500 505 510Asn Lys Val Gly Thr Val
Gly Arg Asn Ile Ser Phe Tyr Ile Thr Asp515 520
525Val Pro Asn Gly Phe His Val Asn Leu Glu Lys Met Pro Thr Glu
Gly530 535 540Glu Asp Leu Lys Leu Ser Cys
Thr Val Asn Lys Phe Leu Tyr Arg Asp545 550
555 560Val Thr Trp Ile Leu Leu Arg Thr Val Asn Asn Arg
Thr Met His Tyr565 570 575Ser Ile Ser Lys
Gln Lys Met Ala Ile Thr Lys Glu His Ser Ile Thr580 585
590Leu Asn Leu Thr Ile Met Asn Val Ser Leu Gln Asp Ser Gly
Thr Tyr595 600 605Ala Cys Arg Ala Arg Asn
Val Tyr Thr Gly Glu Glu Ile Leu Gln Lys610 615
620Lys Glu Ile Thr Ile Arg Asp Gln Glu Ala Pro Tyr Leu Leu Arg
Asn625 630 635 640Leu Ser
Asp His Thr Val Ala Ile Ser Ser Ser Thr Thr Leu Asp Cys645
650 655His Ala Asn Gly Val Pro Glu Pro Gln Ile Thr Trp
Phe Lys Asn Asn660 665 670His Lys Ile Gln
Gln Glu Pro Gly Ile Ile Leu Gly Pro Gly Ser Ser675 680
685Thr Leu Phe Ile Glu Arg Val Thr Glu Glu Asp Glu Gly Val
Tyr His690 695 700Cys Lys Ala Thr Asn Gln
Lys Gly Ser Val Glu Ser Ser Ala Tyr Leu705 710
715 720Thr Val Gln Gly Thr Ser Asp Lys Ser Asn Leu
Glu Xaa725 730169734PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 169Ser Lys Leu
Lys Asp Pro Glu Leu Ser Leu Lys Gly Thr Gln His Ile1 5
10 15Met Gln Ala Gly Gln Thr Leu His Leu
Gln Cys Arg Gly Glu Ala Ala20 25 30His
Lys Trp Ser Leu Pro Glu Met Val Ser Lys Glu Ser Glu Arg Leu35
40 45Ser Ile Thr Lys Ser Ala Cys Gly Arg Asn Gly
Lys Gln Phe Cys Ser50 55 60Thr Leu Thr
Leu Asn Thr Ala Gln Ala Asn His Thr Gly Phe Tyr Ser65 70
75 80Cys Lys Tyr Leu Ala Val Pro Thr
Ser Lys Lys Lys Glu Thr Glu Ser85 90
95Ala Ile Tyr Ile Phe Ile Ser Asp Thr Gly Arg Pro Phe Val Glu Met100
105 110Tyr Ser Glu Ile Pro Glu Ile Ile His Met
Thr Glu Gly Arg Glu Leu115 120 125Val Ile
Pro Cys Arg Val Thr Ser Pro Asn Ile Thr Val Thr Leu Lys130
135 140Lys Phe Pro Leu Asp Thr Leu Ile Pro Asp Gly Lys
Arg Ile Ile Trp145 150 155
160Asp Ser Arg Lys Gly Phe Ile Ile Ser Asn Ala Thr Tyr Lys Glu Ile165
170 175Gly Leu Leu Thr Cys Glu Ala Thr Val
Asn Gly His Leu Tyr Lys Thr180 185 190Asn
Tyr Leu Thr His Arg Gln Thr Asn Thr Ile Ile Asp Val Gln Ile195
200 205Ser Thr Pro Arg Pro Val Lys Leu Leu Arg Gly
His Thr Leu Val Leu210 215 220Asn Cys Thr
Ala Thr Thr Pro Leu Asn Thr Arg Val Gln Met Thr Trp225
230 235 240Ser Tyr Pro Asp Glu Lys Asn
Lys Arg Ala Ser Val Arg Arg Arg Ile245 250
255Asp Gln Ser Asn Ser His Ala Asn Ile Phe Tyr Ser Val Leu Thr Ile260
265 270Asp Lys Met Gln Asn Lys Asp Lys Gly
Leu Tyr Thr Cys Arg Val Arg275 280 285Ser
Gly Pro Ser Phe Lys Ser Val Asn Thr Ser Val His Ile Tyr Asp290
295 300Lys Ala Phe Ile Thr Val Lys His Arg Lys Gln
Gln Val Leu Glu Thr305 310 315
320Val Ala Gly Lys Arg Ser Tyr Arg Leu Ser Met Lys Val Lys Ala
Phe325 330 335Pro Ser Pro Glu Val Val Trp
Leu Lys Asp Gly Leu Pro Ala Thr Glu340 345
350Lys Ser Ala Arg Tyr Leu Thr Arg Gly Tyr Ser Leu Ile Ile Lys Asp355
360 365Val Thr Glu Glu Asp Ala Gly Asn Tyr
Thr Ile Leu Leu Ser Ile Lys370 375 380Gln
Ser Asn Val Phe Lys Asn Leu Thr Ala Thr Leu Ile Val Asn Val385
390 395 400Lys Pro Gln Ile Tyr Glu
Lys Ala Val Ser Ser Phe Pro Asp Pro Ala405 410
415Leu Tyr Pro Leu Gly Ser Arg Gln Ile Leu Thr Cys Thr Ala Tyr
Gly420 425 430Ile Pro Gln Pro Thr Ile Lys
Trp Phe Trp His Pro Cys Asn His Asn435 440
445His Ser Glu Ala Arg Cys Asp Phe Cys Ser Asn Asn Glu Glu Ser Phe450
455 460Ile Leu Asp Ala Asp Ser Asn Met Gly
Asn Arg Ile Glu Ser Ile Thr465 470 475
480Gln Arg Met Ala Ile Ile Glu Gly Lys Asn Lys Met Ala Ser
Thr Leu485 490 495Val Val Ala Asp Ser Arg
Ile Ser Gly Ile Tyr Ile Cys Ile Ala Ser500 505
510Asn Lys Val Gly Thr Val Gly Arg Asn Ile Ser Phe Tyr Ile Thr
Asp515 520 525Val Pro Asn Gly Phe His Val
Asn Leu Glu Lys Met Pro Thr Glu Gly530 535
540Glu Asp Leu Lys Leu Ser Cys Thr Val Asn Lys Phe Leu Tyr Arg Asp545
550 555 560Val Thr Trp Ile
Leu Leu Arg Thr Val Asn Asn Arg Thr Met His Tyr565 570
575Ser Ile Ser Lys Gln Lys Met Ala Ile Thr Lys Glu His Ser
Ile Thr580 585 590Leu Asn Leu Thr Ile Met
Asn Val Ser Leu Gln Asp Ser Gly Thr Tyr595 600
605Ala Cys Arg Ala Arg Asn Val Tyr Thr Gly Glu Glu Ile Leu Gln
Lys610 615 620Lys Glu Ile Thr Ile Arg Asp
Gln Glu Ala Pro Tyr Leu Leu Arg Asn625 630
635 640Leu Ser Asp His Thr Val Ala Ile Ser Ser Ser Thr
Thr Leu Asp Cys645 650 655His Ala Asn Gly
Val Pro Glu Pro Gln Ile Thr Trp Phe Lys Asn Asn660 665
670His Lys Ile Gln Gln Glu Pro Gly Ile Ile Leu Gly Pro Gly
Ser Ser675 680 685Thr Leu Phe Ile Glu Arg
Val Thr Glu Glu Asp Glu Gly Val Tyr His690 695
700Cys Lys Ala Thr Asn Gln Lys Gly Ser Val Glu Ser Ser Ala Tyr
Leu705 710 715 720Thr Val
Gln Gly Thr Ser Asp Lys Ser Asn Leu Glu Gly Xaa725
7301702924DNAArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 170aagcttgaat tcccaccatg cagagcaagg
tgctgctggc cgtcgccctg tggctctgcg 60tggagacccg ggccgcctct gtgggtttgc
ctagtgtttc tcttgatctg cccaggctca 120gcatacaaaa agacatactt acaattaagg
ctaatacaac tcttcaaatt acttgcaggg 180gacagaggga cttggactgg ctttggccca
ataatcagag tggcagtgag caaagggtgg 240aggtgactga gtgcagcgat ggcctcttct
gtaagacact cacaattcca aaagtgatcg 300gaaatgacac tggagcctac aagtgcttct
accgggaaac tgacttggcc tcggtcattt 360atgtctatgt tcaagattac agatctccat
ttattgcttc tgttagtgac caacatggag 420tcgtgtacat tactgagaac aaaaacaaaa
ctgtggtgat tccatgtctc gggtccattt 480caaatctcaa cgtgtcactt tgtgcaagat
acccagaaaa gagatttgtt cctgatggta 540acagaatttc ctgggacagc aagaagggct
ttactattcc cagctacatg atcagctatg 600ctggcatggt cttctgtgaa gcaaaaatta
atgatgaaag ttaccagtct attatgtaca 660tagttgtcgt tgtagggtat aggatttatg
atgtggttct gagtccgtct catggaattg 720aactatctgt tggagaaaaa cttgtcttaa
attgtacagc aagaactgaa ctaaatgtgg 780ggattgactt caactgggaa tacccttctt
cgaagcatca gcataagaaa cttgtaaacc 840gagacctaaa aacccagtct gggagtgaga
tgaagaaatt tttgagcacc ttaactatag 900atggtgtaac ccggagtgac caaggattgt
acacctgtgc agcatccagt gggctgatga 960ccaagaagaa cagcacattt gtcagggtcc
atgaaaaacc ttttgttgct tttggaagtg 1020gcatggaatc tctggtggaa gccacggtgg
gggagcgtgt cagaatccct gcgaagtacc 1080ttggttaccc acccccagaa ataaaatggt
ataaaaatgg aatacccctt gagtccaatc 1140acacaattaa agcggggcat gtactgacga
ttatggaagt gagtgaaaga gacacaggaa 1200attacactgt catccttacc aatcccattt
caaaggagaa gcagagccat gtggtctctc 1260tggttgtgta tgtcccaccc cagattggtg
agaaatctct aatctctcct gtggattcct 1320accagtacgg caccactcaa acgctgacat
gtacggtcta tgccattcct cccccgcatc 1380acatccactg gtattggcag ttggaggaag
agtgcgccaa cgagcccagc caagctgtct 1440cagtgacaaa cccataccct tgtgaagaat
ggagaagtgt ggaggacttc cagggaggaa 1500ataaaattga agttaataaa aatcaatttg
ctctaattga aggaaaaaac aaaactgtaa 1560gtacccttgt tatccaagcg gcaaatgtgt
cagctttgta caaatgtgaa gcggtcaaca 1620aagtcgggag aggagagagg gtgatctcct
tccacgtgac caggggtcct gaaattactt 1680tgcaacctga catgcagccc actgagcagg
agagcgtgtc tttgtggtgc actgcagaca 1740gatctacgtt tgagaacctc acatggtaca
aacttggccc acagcctctg ccaatccatg 1800tgggagagtt gcccacacct gtttgcaaga
acttggatac tctttggaaa ttgaatgcca 1860ccatgttctc taatagcaca aatgacattt
tgatcatgga gcttaagaat gcatccttgc 1920aggaccaagg agactatgtc tgccttgctc
aagacaggaa gaccaagaaa agacattgcg 1980tggtcaggca gctcacagtc ctagagcgtg
tggcacccac gatcacagga aacctggaga 2040atcagacgac aagtattggg gaaagcatcg
aagtctcatg cacggcatct gggaatcccc 2100ctccacagat catgtggttt aaagataatg
agacccttgt agaagactca ggcattgtat 2160tgaaggatgg gaaccggaac ctcactatcc
gcagagtgag gaaggaggac gaaggcctct 2220acacctgcca ggcatgcagt gttcttggct
gtgcaaaagt ggaggcattt ttcataatag 2280aaggtgccca ggaaaagacg aacttggaag
ggtgcgtatc cggtgacacc attgtaatga 2340ctagtggcgg gccccgcact gtggctgaac
tggagggcaa accgttcacc gcactgattc 2400gcggctctgg ctacccatgc ccctcaggtt
tcttccgcac ctgtgaacgt gacgtatatg 2460atctgcgtac acgtgagggt cattgcttac
gtttgaccca tgatcaccgt gttctggtga 2520tggatggtgg cctggaatgg cgtgccgcgg
gtgaactgga acgcggcgac cgcctggtga 2580tggatgatgc agctggcgag tttccggcac
tggcaacctt ccgtggcctg cgtggcgctg 2640gccgccagga tgtttatgac gctactgttt
acggtgctag cgcattcact gctaatggct 2700tcattgtaca cgcatgtggc gagcagcccg
ggaccggtct gaactcaggc ctcacgacaa 2760atcctggtgt atccgcttgg caggtcaaca
cagcttatac tgcgggacaa ttggtcacat 2820ataacggcaa gacgtataaa tgtttgcagc
cccacacctc cttggcagga tgggaaccat 2880ccaacgttcc tgccttgtgg cagcttcaat
gactcgagcg gccg 2924171964PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 171Met Gln Ser Lys Val Leu Leu Ala Val Ala Leu Trp Leu Cys
Val Glu1 5 10 15Thr Arg
Ala Ala Ser Val Gly Leu Pro Ser Val Ser Leu Asp Leu Pro20
25 30Arg Leu Ser Ile Gln Lys Asp Ile Leu Thr Ile Lys
Ala Asn Thr Thr35 40 45Leu Gln Ile Thr
Cys Arg Gly Gln Arg Asp Leu Asp Trp Leu Trp Pro50 55
60Asn Asn Gln Ser Gly Ser Glu Gln Arg Val Glu Val Thr Glu
Cys Ser65 70 75 80Asp
Gly Leu Phe Cys Lys Thr Leu Thr Ile Pro Lys Val Ile Gly Asn85
90 95Asp Thr Gly Ala Tyr Lys Cys Phe Tyr Arg Glu
Thr Asp Leu Ala Ser100 105 110Val Ile Tyr
Val Tyr Val Gln Asp Tyr Arg Ser Pro Phe Ile Ala Ser115
120 125Val Ser Asp Gln His Gly Val Val Tyr Ile Thr Glu
Asn Lys Asn Lys130 135 140Thr Val Val Ile
Pro Cys Leu Gly Ser Ile Ser Asn Leu Asn Val Ser145 150
155 160Leu Cys Ala Arg Tyr Pro Glu Lys Arg
Phe Val Pro Asp Gly Asn Arg165 170 175Ile
Ser Trp Asp Ser Lys Lys Gly Phe Thr Ile Pro Ser Tyr Met Ile180
185 190Ser Tyr Ala Gly Met Val Phe Cys Glu Ala Lys
Ile Asn Asp Glu Ser195 200 205Tyr Gln Ser
Ile Met Tyr Ile Val Val Val Val Gly Tyr Arg Ile Tyr210
215 220Asp Val Val Leu Ser Pro Ser His Gly Ile Glu Leu
Ser Val Gly Glu225 230 235
240Lys Leu Val Leu Asn Cys Thr Ala Arg Thr Glu Leu Asn Val Gly Ile245
250 255Asp Phe Asn Trp Glu Tyr Pro Ser Ser
Lys His Gln His Lys Lys Leu260 265 270Val
Asn Arg Asp Leu Lys Thr Gln Ser Gly Ser Glu Met Lys Lys Phe275
280 285Leu Ser Thr Leu Thr Ile Asp Gly Val Thr Arg
Ser Asp Gln Gly Leu290 295 300Tyr Thr Cys
Ala Ala Ser Ser Gly Leu Met Thr Lys Lys Asn Ser Thr305
310 315 320Phe Val Arg Val His Glu Lys
Pro Phe Val Ala Phe Gly Ser Gly Met325 330
335Glu Ser Leu Val Glu Ala Thr Val Gly Glu Arg Val Arg Ile Pro Ala340
345 350Lys Tyr Leu Gly Tyr Pro Pro Pro Glu
Ile Lys Trp Tyr Lys Asn Gly355 360 365Ile
Pro Leu Glu Ser Asn His Thr Ile Lys Ala Gly His Val Leu Thr370
375 380Ile Met Glu Val Ser Glu Arg Asp Thr Gly Asn
Tyr Thr Val Ile Leu385 390 395
400Thr Asn Pro Ile Ser Lys Glu Lys Gln Ser His Val Val Ser Leu
Val405 410 415Val Tyr Val Pro Pro Gln Ile
Gly Glu Lys Ser Leu Ile Ser Pro Val420 425
430Asp Ser Tyr Gln Tyr Gly Thr Thr Gln Thr Leu Thr Cys Thr Val Tyr435
440 445Ala Ile Pro Pro Pro His His Ile His
Trp Tyr Trp Gln Leu Glu Glu450 455 460Glu
Cys Ala Asn Glu Pro Ser Gln Ala Val Ser Val Thr Asn Pro Tyr465
470 475 480Pro Cys Glu Glu Trp Arg
Ser Val Glu Asp Phe Gln Gly Gly Asn Lys485 490
495Ile Glu Val Asn Lys Asn Gln Phe Ala Leu Ile Glu Gly Lys Asn
Lys500 505 510Thr Val Ser Thr Leu Val Ile
Gln Ala Ala Asn Val Ser Ala Leu Tyr515 520
525Lys Cys Glu Ala Val Asn Lys Val Gly Arg Gly Glu Arg Val Ile Ser530
535 540Phe His Val Thr Arg Gly Pro Glu Ile
Thr Leu Gln Pro Asp Met Gln545 550 555
560Pro Thr Glu Gln Glu Ser Val Ser Leu Trp Cys Thr Ala Asp
Arg Ser565 570 575Thr Phe Glu Asn Leu Thr
Trp Tyr Lys Leu Gly Pro Gln Pro Leu Pro580 585
590Ile His Val Gly Glu Leu Pro Thr Pro Val Cys Lys Asn Leu Asp
Thr595 600 605Leu Trp Lys Leu Asn Ala Thr
Met Phe Ser Asn Ser Thr Asn Asp Ile610 615
620Leu Ile Met Glu Leu Lys Asn Ala Ser Leu Gln Asp Gln Gly Asp Tyr625
630 635 640Val Cys Leu Ala
Gln Asp Arg Lys Thr Lys Lys Arg His Cys Val Val645 650
655Arg Gln Leu Thr Val Leu Glu Arg Val Ala Pro Thr Ile Thr
Gly Asn660 665 670Leu Glu Asn Gln Thr Thr
Ser Ile Gly Glu Ser Ile Glu Val Ser Cys675 680
685Thr Ala Ser Gly Asn Pro Pro Pro Gln Ile Met Trp Phe Lys Asp
Asn690 695 700Glu Thr Leu Val Glu Asp Ser
Gly Ile Val Leu Lys Asp Gly Asn Arg705 710
715 720Asn Leu Thr Ile Arg Arg Val Arg Lys Glu Asp Glu
Gly Leu Tyr Thr725 730 735Cys Gln Ala Cys
Ser Val Leu Gly Cys Ala Lys Val Glu Ala Phe Phe740 745
750Ile Ile Glu Gly Ala Gln Glu Lys Thr Asn Leu Glu Gly Cys
Val Ser755 760 765Gly Asp Thr Ile Val Met
Thr Ser Gly Gly Pro Arg Thr Val Ala Glu770 775
780Leu Glu Gly Lys Pro Phe Thr Ala Leu Ile Arg Gly Ser Gly Tyr
Pro785 790 795 800Cys Pro
Ser Gly Phe Phe Arg Thr Cys Glu Arg Asp Val Tyr Asp Leu805
810 815Arg Thr Arg Glu Gly His Cys Leu Arg Leu Thr His
Asp His Arg Val820 825 830Leu Val Met Asp
Gly Gly Leu Glu Trp Arg Ala Ala Gly Glu Leu Glu835 840
845Arg Gly Asp Arg Leu Val Met Asp Asp Ala Ala Gly Glu Phe
Pro Ala850 855 860Leu Ala Thr Phe Arg Gly
Leu Arg Gly Ala Gly Arg Gln Asp Val Tyr865 870
875 880Asp Ala Thr Val Tyr Gly Ala Ser Ala Phe Thr
Ala Asn Gly Phe Ile885 890 895Val His Ala
Cys Gly Glu Gln Pro Gly Thr Gly Leu Asn Ser Gly Leu900
905 910Thr Thr Asn Pro Gly Val Ser Ala Trp Gln Val Asn
Thr Ala Tyr Thr915 920 925Ala Gly Gln Leu
Val Thr Tyr Asn Gly Lys Thr Tyr Lys Cys Leu Gln930 935
940Pro His Thr Ser Leu Ala Gly Trp Glu Pro Ser Asn Val Pro
Ala Leu945 950 955 960Trp
Gln Leu Gln172945PRTArtificial Sequenceartificial construct relating to
Homo Sapiens immunoglobulin 172Ala Ser Val Gly Leu Pro Ser Val Ser
Leu Asp Leu Pro Arg Leu Ser1 5 10
15Ile Gln Lys Asp Ile Leu Thr Ile Lys Ala Asn Thr Thr Leu Gln
Ile20 25 30Thr Cys Arg Gly Gln Arg Asp
Leu Asp Trp Leu Trp Pro Asn Asn Gln35 40
45Ser Gly Ser Glu Gln Arg Val Glu Val Thr Glu Cys Ser Asp Gly Leu50
55 60Phe Cys Lys Thr Leu Thr Ile Pro Lys Val
Ile Gly Asn Asp Thr Gly65 70 75
80Ala Tyr Lys Cys Phe Tyr Arg Glu Thr Asp Leu Ala Ser Val Ile
Tyr85 90 95Val Tyr Val Gln Asp Tyr Arg
Ser Pro Phe Ile Ala Ser Val Ser Asp100 105
110Gln His Gly Val Val Tyr Ile Thr Glu Asn Lys Asn Lys Thr Val Val115
120 125Ile Pro Cys Leu Gly Ser Ile Ser Asn
Leu Asn Val Ser Leu Cys Ala130 135 140Arg
Tyr Pro Glu Lys Arg Phe Val Pro Asp Gly Asn Arg Ile Ser Trp145
150 155 160Asp Ser Lys Lys Gly Phe
Thr Ile Pro Ser Tyr Met Ile Ser Tyr Ala165 170
175Gly Met Val Phe Cys Glu Ala Lys Ile Asn Asp Glu Ser Tyr Gln
Ser180 185 190Ile Met Tyr Ile Val Val Val
Val Gly Tyr Arg Ile Tyr Asp Val Val195 200
205Leu Ser Pro Ser His Gly Ile Glu Leu Ser Val Gly Glu Lys Leu Val210
215 220Leu Asn Cys Thr Ala Arg Thr Glu Leu
Asn Val Gly Ile Asp Phe Asn225 230 235
240Trp Glu Tyr Pro Ser Ser Lys His Gln His Lys Lys Leu Val
Asn Arg245 250 255Asp Leu Lys Thr Gln Ser
Gly Ser Glu Met Lys Lys Phe Leu Ser Thr260 265
270Leu Thr Ile Asp Gly Val Thr Arg Ser Asp Gln Gly Leu Tyr Thr
Cys275 280 285Ala Ala Ser Ser Gly Leu Met
Thr Lys Lys Asn Ser Thr Phe Val Arg290 295
300Val His Glu Lys Pro Phe Val Ala Phe Gly Ser Gly Met Glu Ser Leu305
310 315 320Val Glu Ala Thr
Val Gly Glu Arg Val Arg Ile Pro Ala Lys Tyr Leu325 330
335Gly Tyr Pro Pro Pro Glu Ile Lys Trp Tyr Lys Asn Gly Ile
Pro Leu340 345 350Glu Ser Asn His Thr Ile
Lys Ala Gly His Val Leu Thr Ile Met Glu355 360
365Val Ser Glu Arg Asp Thr Gly Asn Tyr Thr Val Ile Leu Thr Asn
Pro370 375 380Ile Ser Lys Glu Lys Gln Ser
His Val Val Ser Leu Val Val Tyr Val385 390
395 400Pro Pro Gln Ile Gly Glu Lys Ser Leu Ile Ser Pro
Val Asp Ser Tyr405 410 415Gln Tyr Gly Thr
Thr Gln Thr Leu Thr Cys Thr Val Tyr Ala Ile Pro420 425
430Pro Pro His His Ile His Trp Tyr Trp Gln Leu Glu Glu Glu
Cys Ala435 440 445Asn Glu Pro Ser Gln Ala
Val Ser Val Thr Asn Pro Tyr Pro Cys Glu450 455
460Glu Trp Arg Ser Val Glu Asp Phe Gln Gly Gly Asn Lys Ile Glu
Val465 470 475 480Asn Lys
Asn Gln Phe Ala Leu Ile Glu Gly Lys Asn Lys Thr Val Ser485
490 495Thr Leu Val Ile Gln Ala Ala Asn Val Ser Ala Leu
Tyr Lys Cys Glu500 505 510Ala Val Asn Lys
Val Gly Arg Gly Glu Arg Val Ile Ser Phe His Val515 520
525Thr Arg Gly Pro Glu Ile Thr Leu Gln Pro Asp Met Gln Pro
Thr Glu530 535 540Gln Glu Ser Val Ser Leu
Trp Cys Thr Ala Asp Arg Ser Thr Phe Glu545 550
555 560Asn Leu Thr Trp Tyr Lys Leu Gly Pro Gln Pro
Leu Pro Ile His Val565 570 575Gly Glu Leu
Pro Thr Pro Val Cys Lys Asn Leu Asp Thr Leu Trp Lys580
585 590Leu Asn Ala Thr Met Phe Ser Asn Ser Thr Asn Asp
Ile Leu Ile Met595 600 605Glu Leu Lys Asn
Ala Ser Leu Gln Asp Gln Gly Asp Tyr Val Cys Leu610 615
620Ala Gln Asp Arg Lys Thr Lys Lys Arg His Cys Val Val Arg
Gln Leu625 630 635 640Thr
Val Leu Glu Arg Val Ala Pro Thr Ile Thr Gly Asn Leu Glu Asn645
650 655Gln Thr Thr Ser Ile Gly Glu Ser Ile Glu Val
Ser Cys Thr Ala Ser660 665 670Gly Asn Pro
Pro Pro Gln Ile Met Trp Phe Lys Asp Asn Glu Thr Leu675
680 685Val Glu Asp Ser Gly Ile Val Leu Lys Asp Gly Asn
Arg Asn Leu Thr690 695 700Ile Arg Arg Val
Arg Lys Glu Asp Glu Gly Leu Tyr Thr Cys Gln Ala705 710
715 720Cys Ser Val Leu Gly Cys Ala Lys Val
Glu Ala Phe Phe Ile Ile Glu725 730 735Gly
Ala Gln Glu Lys Thr Asn Leu Glu Gly Cys Val Ser Gly Asp Thr740
745 750Ile Val Met Thr Ser Gly Gly Pro Arg Thr Val
Ala Glu Leu Glu Gly755 760 765Lys Pro Phe
Thr Ala Leu Ile Arg Gly Ser Gly Tyr Pro Cys Pro Ser770
775 780Gly Phe Phe Arg Thr Cys Glu Arg Asp Val Tyr Asp
Leu Arg Thr Arg785 790 795
800Glu Gly His Cys Leu Arg Leu Thr His Asp His Arg Val Leu Val Met805
810 815Asp Gly Gly Leu Glu Trp Arg Ala Ala
Gly Glu Leu Glu Arg Gly Asp820 825 830Arg
Leu Val Met Asp Asp Ala Ala Gly Glu Phe Pro Ala Leu Ala Thr835
840 845Phe Arg Gly Leu Arg Gly Ala Gly Arg Gln Asp
Val Tyr Asp Ala Thr850 855 860Val Tyr Gly
Ala Ser Ala Phe Thr Ala Asn Gly Phe Ile Val His Ala865
870 875 880Cys Gly Glu Gln Pro Gly Thr
Gly Leu Asn Ser Gly Leu Thr Thr Asn885 890
895Pro Gly Val Ser Ala Trp Gln Val Asn Thr Ala Tyr Thr Ala Gly Gln900
905 910Leu Val Thr Tyr Asn Gly Lys Thr Tyr
Lys Cys Leu Gln Pro His Thr915 920 925Ser
Leu Ala Gly Trp Glu Pro Ser Asn Val Pro Ala Leu Trp Gln Leu930
935 940Gln945173746PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 173Ala Ser Val
Gly Leu Pro Ser Val Ser Leu Asp Leu Pro Arg Leu Ser1 5
10 15Ile Gln Lys Asp Ile Leu Thr Ile Lys
Ala Asn Thr Thr Leu Gln Ile20 25 30Thr
Cys Arg Gly Gln Arg Asp Leu Asp Trp Leu Trp Pro Asn Asn Gln35
40 45Ser Gly Ser Glu Gln Arg Val Glu Val Thr Glu
Cys Ser Asp Gly Leu50 55 60Phe Cys Lys
Thr Leu Thr Ile Pro Lys Val Ile Gly Asn Asp Thr Gly65 70
75 80Ala Tyr Lys Cys Phe Tyr Arg Glu
Thr Asp Leu Ala Ser Val Ile Tyr85 90
95Val Tyr Val Gln Asp Tyr Arg Ser Pro Phe Ile Ala Ser Val Ser Asp100
105 110Gln His Gly Val Val Tyr Ile Thr Glu Asn
Lys Asn Lys Thr Val Val115 120 125Ile Pro
Cys Leu Gly Ser Ile Ser Asn Leu Asn Val Ser Leu Cys Ala130
135 140Arg Tyr Pro Glu Lys Arg Phe Val Pro Asp Gly Asn
Arg Ile Ser Trp145 150 155
160Asp Ser Lys Lys Gly Phe Thr Ile Pro Ser Tyr Met Ile Ser Tyr Ala165
170 175Gly Met Val Phe Cys Glu Ala Lys Ile
Asn Asp Glu Ser Tyr Gln Ser180 185 190Ile
Met Tyr Ile Val Val Val Val Gly Tyr Arg Ile Tyr Asp Val Val195
200 205Leu Ser Pro Ser His Gly Ile Glu Leu Ser Val
Gly Glu Lys Leu Val210 215 220Leu Asn Cys
Thr Ala Arg Thr Glu Leu Asn Val Gly Ile Asp Phe Asn225
230 235 240Trp Glu Tyr Pro Ser Ser Lys
His Gln His Lys Lys Leu Val Asn Arg245 250
255Asp Leu Lys Thr Gln Ser Gly Ser Glu Met Lys Lys Phe Leu Ser Thr260
265 270Leu Thr Ile Asp Gly Val Thr Arg Ser
Asp Gln Gly Leu Tyr Thr Cys275 280 285Ala
Ala Ser Ser Gly Leu Met Thr Lys Lys Asn Ser Thr Phe Val Arg290
295 300Val His Glu Lys Pro Phe Val Ala Phe Gly Ser
Gly Met Glu Ser Leu305 310 315
320Val Glu Ala Thr Val Gly Glu Arg Val Arg Ile Pro Ala Lys Tyr
Leu325 330 335Gly Tyr Pro Pro Pro Glu Ile
Lys Trp Tyr Lys Asn Gly Ile Pro Leu340 345
350Glu Ser Asn His Thr Ile Lys Ala Gly His Val Leu Thr Ile Met Glu355
360 365Val Ser Glu Arg Asp Thr Gly Asn Tyr
Thr Val Ile Leu Thr Asn Pro370 375 380Ile
Ser Lys Glu Lys Gln Ser His Val Val Ser Leu Val Val Tyr Val385
390 395 400Pro Pro Gln Ile Gly Glu
Lys Ser Leu Ile Ser Pro Val Asp Ser Tyr405 410
415Gln Tyr Gly Thr Thr Gln Thr Leu Thr Cys Thr Val Tyr Ala Ile
Pro420 425 430Pro Pro His His Ile His Trp
Tyr Trp Gln Leu Glu Glu Glu Cys Ala435 440
445Asn Glu Pro Ser Gln Ala Val Ser Val Thr Asn Pro Tyr Pro Cys Glu450
455 460Glu Trp Arg Ser Val Glu Asp Phe Gln
Gly Gly Asn Lys Ile Glu Val465 470 475
480Asn Lys Asn Gln Phe Ala Leu Ile Glu Gly Lys Asn Lys Thr
Val Ser485 490 495Thr Leu Val Ile Gln Ala
Ala Asn Val Ser Ala Leu Tyr Lys Cys Glu500 505
510Ala Val Asn Lys Val Gly Arg Gly Glu Arg Val Ile Ser Phe His
Val515 520 525Thr Arg Gly Pro Glu Ile Thr
Leu Gln Pro Asp Met Gln Pro Thr Glu530 535
540Gln Glu Ser Val Ser Leu Trp Cys Thr Ala Asp Arg Ser Thr Phe Glu545
550 555 560Asn Leu Thr Trp
Tyr Lys Leu Gly Pro Gln Pro Leu Pro Ile His Val565 570
575Gly Glu Leu Pro Thr Pro Val Cys Lys Asn Leu Asp Thr Leu
Trp Lys580 585 590Leu Asn Ala Thr Met Phe
Ser Asn Ser Thr Asn Asp Ile Leu Ile Met595 600
605Glu Leu Lys Asn Ala Ser Leu Gln Asp Gln Gly Asp Tyr Val Cys
Leu610 615 620Ala Gln Asp Arg Lys Thr Lys
Lys Arg His Cys Val Val Arg Gln Leu625 630
635 640Thr Val Leu Glu Arg Val Ala Pro Thr Ile Thr Gly
Asn Leu Glu Asn645 650 655Gln Thr Thr Ser
Ile Gly Glu Ser Ile Glu Val Ser Cys Thr Ala Ser660 665
670Gly Asn Pro Pro Pro Gln Ile Met Trp Phe Lys Asp Asn Glu
Thr Leu675 680 685Val Glu Asp Ser Gly Ile
Val Leu Lys Asp Gly Asn Arg Asn Leu Thr690 695
700Ile Arg Arg Val Arg Lys Glu Asp Glu Gly Leu Tyr Thr Cys Gln
Ala705 710 715 720Cys Ser
Val Leu Gly Cys Ala Lys Val Glu Ala Phe Phe Ile Ile Glu725
730 735Gly Ala Gln Glu Lys Thr Asn Leu Glu Xaa740
745174747PRTArtificial Sequenceartificial construct relating to
Homo Sapiens immunoglobulin 174Ala Ser Val Gly Leu Pro Ser Val Ser
Leu Asp Leu Pro Arg Leu Ser1 5 10
15Ile Gln Lys Asp Ile Leu Thr Ile Lys Ala Asn Thr Thr Leu Gln
Ile20 25 30Thr Cys Arg Gly Gln Arg Asp
Leu Asp Trp Leu Trp Pro Asn Asn Gln35 40
45Ser Gly Ser Glu Gln Arg Val Glu Val Thr Glu Cys Ser Asp Gly Leu50
55 60Phe Cys Lys Thr Leu Thr Ile Pro Lys Val
Ile Gly Asn Asp Thr Gly65 70 75
80Ala Tyr Lys Cys Phe Tyr Arg Glu Thr Asp Leu Ala Ser Val Ile
Tyr85 90 95Val Tyr Val Gln Asp Tyr Arg
Ser Pro Phe Ile Ala Ser Val Ser Asp100 105
110Gln His Gly Val Val Tyr Ile Thr Glu Asn Lys Asn Lys Thr Val Val115
120 125Ile Pro Cys Leu Gly Ser Ile Ser Asn
Leu Asn Val Ser Leu Cys Ala130 135 140Arg
Tyr Pro Glu Lys Arg Phe Val Pro Asp Gly Asn Arg Ile Ser Trp145
150 155 160Asp Ser Lys Lys Gly Phe
Thr Ile Pro Ser Tyr Met Ile Ser Tyr Ala165 170
175Gly Met Val Phe Cys Glu Ala Lys Ile Asn Asp Glu Ser Tyr Gln
Ser180 185 190Ile Met Tyr Ile Val Val Val
Val Gly Tyr Arg Ile Tyr Asp Val Val195 200
205Leu Ser Pro Ser His Gly Ile Glu Leu Ser Val Gly Glu Lys Leu Val210
215 220Leu Asn Cys Thr Ala Arg Thr Glu Leu
Asn Val Gly Ile Asp Phe Asn225 230 235
240Trp Glu Tyr Pro Ser Ser Lys His Gln His Lys Lys Leu Val
Asn Arg245 250 255Asp Leu Lys Thr Gln Ser
Gly Ser Glu Met Lys Lys Phe Leu Ser Thr260 265
270Leu Thr Ile Asp Gly Val Thr Arg Ser Asp Gln Gly Leu Tyr Thr
Cys275 280 285Ala Ala Ser Ser Gly Leu Met
Thr Lys Lys Asn Ser Thr Phe Val Arg290 295
300Val His Glu Lys Pro Phe Val Ala Phe Gly Ser Gly Met Glu Ser Leu305
310 315 320Val Glu Ala Thr
Val Gly Glu Arg Val Arg Ile Pro Ala Lys Tyr Leu325 330
335Gly Tyr Pro Pro Pro Glu Ile Lys Trp Tyr Lys Asn Gly Ile
Pro Leu340 345 350Glu Ser Asn His Thr Ile
Lys Ala Gly His Val Leu Thr Ile Met Glu355 360
365Val Ser Glu Arg Asp Thr Gly Asn Tyr Thr Val Ile Leu Thr Asn
Pro370 375 380Ile Ser Lys Glu Lys Gln Ser
His Val Val Ser Leu Val Val Tyr Val385 390
395 400Pro Pro Gln Ile Gly Glu Lys Ser Leu Ile Ser Pro
Val Asp Ser Tyr405 410 415Gln Tyr Gly Thr
Thr Gln Thr Leu Thr Cys Thr Val Tyr Ala Ile Pro420 425
430Pro Pro His His Ile His Trp Tyr Trp Gln Leu Glu Glu Glu
Cys Ala435 440 445Asn Glu Pro Ser Gln Ala
Val Ser Val Thr Asn Pro Tyr Pro Cys Glu450 455
460Glu Trp Arg Ser Val Glu Asp Phe Gln Gly Gly Asn Lys Ile Glu
Val465 470 475 480Asn Lys
Asn Gln Phe Ala Leu Ile Glu Gly Lys Asn Lys Thr Val Ser485
490 495Thr Leu Val Ile Gln Ala Ala Asn Val Ser Ala Leu
Tyr Lys Cys Glu500 505 510Ala Val Asn Lys
Val Gly Arg Gly Glu Arg Val Ile Ser Phe His Val515 520
525Thr Arg Gly Pro Glu Ile Thr Leu Gln Pro Asp Met Gln Pro
Thr Glu530 535 540Gln Glu Ser Val Ser Leu
Trp Cys Thr Ala Asp Arg Ser Thr Phe Glu545 550
555 560Asn Leu Thr Trp Tyr Lys Leu Gly Pro Gln Pro
Leu Pro Ile His Val565 570 575Gly Glu Leu
Pro Thr Pro Val Cys Lys Asn Leu Asp Thr Leu Trp Lys580
585 590Leu Asn Ala Thr Met Phe Ser Asn Ser Thr Asn Asp
Ile Leu Ile Met595 600 605Glu Leu Lys Asn
Ala Ser Leu Gln Asp Gln Gly Asp Tyr Val Cys Leu610 615
620Ala Gln Asp Arg Lys Thr Lys Lys Arg His Cys Val Val Arg
Gln Leu625 630 635 640Thr
Val Leu Glu Arg Val Ala Pro Thr Ile Thr Gly Asn Leu Glu Asn645
650 655Gln Thr Thr Ser Ile Gly Glu Ser Ile Glu Val
Ser Cys Thr Ala Ser660 665 670Gly Asn Pro
Pro Pro Gln Ile Met Trp Phe Lys Asp Asn Glu Thr Leu675
680 685Val Glu Asp Ser Gly Ile Val Leu Lys Asp Gly Asn
Arg Asn Leu Thr690 695 700Ile Arg Arg Val
Arg Lys Glu Asp Glu Gly Leu Tyr Thr Cys Gln Ala705 710
715 720Cys Ser Val Leu Gly Cys Ala Lys Val
Glu Ala Phe Phe Ile Ile Glu725 730 735Gly
Ala Gln Glu Lys Thr Asn Leu Glu Gly Xaa740
7451752957DNAArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 175aagcttgaat tcccaccatg cagcggggcg
ccgcgctgtg cctgcgactg tggctctgcc 60tgggactcct ggacggcctg gtgagtggct
actccatgac ccccccgacc ttgaacatca 120cggaggagtc acacgtcatc gacaccggtg
acagcctgtc catctcctgc aggggacagc 180accccctgga gtgggcttgg ccaggagctc
aggaggcgcc agccaccgga gacaaggaca 240gcgaggacac gggggtggtg cgagactgcg
agggcacaga cgccaggccc tactgcaagg 300tgttgctgct gcacgaggta catgccaacg
acacaggcag ctacgtctgc tactacaagt 360acatcaaggc acgcatcgag ggcaccacgg
ccgccagctc ctacgtgttc gtgagagact 420ttgagcagcc attcatcaac aagcctgaca
cgctcttggt caacaggaag gacgccatgt 480gggtgccctg tctggtgtcc atccccggcc
tcaatgtcac gctgcgctcg caaagctcgg 540tgctgtggcc agacgggcag gaggtggtgt
gggatgaccg gcggggcatg ctcgtgtcca 600cgccactgct gcacgatgcc ctgtacctgc
agtgcgagac cacctgggga gaccaggact 660tcctttccaa ccccttcctg gtgcacatca
caggcaacga gctctatgac atccagctgt 720tgcccaggaa gtcgctggag ctgctggtag
gggagaagct ggtcctgaac tgcaccgtgt 780gggctgagtt taactcaggt gtcacctttg
actgggacta cccagggaag caggcagagc 840ggggtaagtg ggtgcccgag cgacgctccc
agcagaccca cacagaactc tccagcatcc 900tgaccatcca caacgtcagc cagcacgacc
tgggctcgta tgtgtgcaag gccaacaacg 960gcatccagcg atttcgggag agcaccgagg
tcattgtgca tgaaaatccc ttcatcagcg 1020tcgagtggct caaaggaccc atcctggagg
ccacggcagg agacgagctg gtgaagctgc 1080ccgtgaagct ggcagcgtac cccccgcccg
agttccagtg gtacaaggat ggaaaggcac 1140tgtccgggcg ccacagtcca catgccctgg
tgctcaagga ggtgacagag gccagcacag 1200gcacctacac cctcgccctg tggaactccg
ctgctggcct gaggcgcaac atcagcctgg 1260agctggtggt gaatgtgccc ccccagatac
atgagaagga ggcctcctcc cccagcatct 1320actcgcgtca cagccgccag gccctcacct
gcacggccta cggggtgccc ctgcctctca 1380gcatccagtg gcactggcgg ccctggacac
cctgcaagat gtttgcccag cgtagtctcc 1440ggcggcggca gcagcaagac ctcatgccac
agtgccgtga ctggagggcg gtgaccacgc 1500aggatgccgt gaaccccatc gagagcctgg
acacctggac cgagtttgtg gagggaaaga 1560ataagactgt gagcaagctg gtgatccaga
atgccaacgt gtctgccatg tacaagtgtg 1620tggtctccaa caaggtgggc caggatgagc
ggctcatcta cttctatgtg accaccatcc 1680ccgacggctt caccatcgaa tccaagccat
ccgaggagct actagagggc cagccggtgc 1740tcctgagctg ccaagccgac agctacaagt
acgagcatct gcgctggtac cgcctcaacc 1800tgtccacgct gcacgatgcg cacgggaacc
cgcttctgct cgactgcaag aacgtgcatc 1860tgttcgccac ccctctggcc gccagcctgg
aggaggtggc acctggggcg cgccacgcca 1920cgctcagcct gagtatcccc cgcgtcgcgc
ccgagcacga gggccactat gtgtgcgaag 1980tgcaagaccg gcgcagccat gacaagcact
gccacaagaa gtacctgtcg gtgcaggccc 2040tggaagcccc tcggctcacg cagaacttga
ccgacctcct ggtgaacgtg agcgactcgc 2100tggagatgca gtgcttggtg gccggagcgc
acgcgcccag catcgtgtgg tacaaagacg 2160agaggctgct ggaggaaaag tctggagtcg
acttggcgga ctccaaccag aagctgagca 2220tccagcgcgt gcgcgaggag gatgcgggac
gctatctgtg cagcgtgtgc aacgccaagg 2280gctgcgtcaa ctcctccgcc agcgtggccg
tggaaggctc cgaggataag ggcagcatgg 2340aggggtgcgt atccggtgac accattgtaa
tgactagtgg cgggccccgc actgtggctg 2400aactggaggg caaaccgttc accgcactga
ttcgcggctc tggctaccca tgcccctcag 2460gtttcttccg cacctgtgaa cgtgacgtat
atgatctgcg tacacgtgag ggtcattgct 2520tacgtttgac ccatgatcac cgtgttctgg
tgatggatgg tggcctggaa tggcgtgccg 2580cgggtgaact ggaacgcggc gaccgcctgg
tgatggatga tgcagctggc gagtttccgg 2640cactggcaac cttccgtggc ctgcgtggcg
ctggccgcca ggatgtttat gacgctactg 2700tttacggtgc tagcgcattc actgctaatg
gcttcattgt acacgcatgt ggcgagcagc 2760ccgggaccgg tctgaactca ggcctcacga
caaatcctgg tgtatccgct tggcaggtca 2820acacagctta tactgcggga caattggtca
catataacgg caagacgtat aaatgtttgc 2880agccccacac ctccttggca ggatgggaac
catccaacgt tcctgccttg tggcagcttc 2940aatgactcga gcggccg
2957176975PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 176Met Gln Arg Gly Ala Ala Leu Cys Leu Arg Leu Trp Leu Cys
Leu Gly1 5 10 15Leu Leu
Asp Gly Leu Val Ser Gly Tyr Ser Met Thr Pro Pro Thr Leu20
25 30Asn Ile Thr Glu Glu Ser His Val Ile Asp Thr Gly
Asp Ser Leu Ser35 40 45Ile Ser Cys Arg
Gly Gln His Pro Leu Glu Trp Ala Trp Pro Gly Ala50 55
60Gln Glu Ala Pro Ala Thr Gly Asp Lys Asp Ser Glu Asp Thr
Gly Val65 70 75 80Val
Arg Asp Cys Glu Gly Thr Asp Ala Arg Pro Tyr Cys Lys Val Leu85
90 95Leu Leu His Glu Val His Ala Asn Asp Thr Gly
Ser Tyr Val Cys Tyr100 105 110Tyr Lys Tyr
Ile Lys Ala Arg Ile Glu Gly Thr Thr Ala Ala Ser Ser115
120 125Tyr Val Phe Val Arg Asp Phe Glu Gln Pro Phe Ile
Asn Lys Pro Asp130 135 140Thr Leu Leu Val
Asn Arg Lys Asp Ala Met Trp Val Pro Cys Leu Val145 150
155 160Ser Ile Pro Gly Leu Asn Val Thr Leu
Arg Ser Gln Ser Ser Val Leu165 170 175Trp
Pro Asp Gly Gln Glu Val Val Trp Asp Asp Arg Arg Gly Met Leu180
185 190Val Ser Thr Pro Leu Leu His Asp Ala Leu Tyr
Leu Gln Cys Glu Thr195 200 205Thr Trp Gly
Asp Gln Asp Phe Leu Ser Asn Pro Phe Leu Val His Ile210
215 220Thr Gly Asn Glu Leu Tyr Asp Ile Gln Leu Leu Pro
Arg Lys Ser Leu225 230 235
240Glu Leu Leu Val Gly Glu Lys Leu Val Leu Asn Cys Thr Val Trp Ala245
250 255Glu Phe Asn Ser Gly Val Thr Phe Asp
Trp Asp Tyr Pro Gly Lys Gln260 265 270Ala
Glu Arg Gly Lys Trp Val Pro Glu Arg Arg Ser Gln Gln Thr His275
280 285Thr Glu Leu Ser Ser Ile Leu Thr Ile His Asn
Val Ser Gln His Asp290 295 300Leu Gly Ser
Tyr Val Cys Lys Ala Asn Asn Gly Ile Gln Arg Phe Arg305
310 315 320Glu Ser Thr Glu Val Ile Val
His Glu Asn Pro Phe Ile Ser Val Glu325 330
335Trp Leu Lys Gly Pro Ile Leu Glu Ala Thr Ala Gly Asp Glu Leu Val340
345 350Lys Leu Pro Val Lys Leu Ala Ala Tyr
Pro Pro Pro Glu Phe Gln Trp355 360 365Tyr
Lys Asp Gly Lys Ala Leu Ser Gly Arg His Ser Pro His Ala Leu370
375 380Val Leu Lys Glu Val Thr Glu Ala Ser Thr Gly
Thr Tyr Thr Leu Ala385 390 395
400Leu Trp Asn Ser Ala Ala Gly Leu Arg Arg Asn Ile Ser Leu Glu
Leu405 410 415Val Val Asn Val Pro Pro Gln
Ile His Glu Lys Glu Ala Ser Ser Pro420 425
430Ser Ile Tyr Ser Arg His Ser Arg Gln Ala Leu Thr Cys Thr Ala Tyr435
440 445Gly Val Pro Leu Pro Leu Ser Ile Gln
Trp His Trp Arg Pro Trp Thr450 455 460Pro
Cys Lys Met Phe Ala Gln Arg Ser Leu Arg Arg Arg Gln Gln Gln465
470 475 480Asp Leu Met Pro Gln Cys
Arg Asp Trp Arg Ala Val Thr Thr Gln Asp485 490
495Ala Val Asn Pro Ile Glu Ser Leu Asp Thr Trp Thr Glu Phe Val
Glu500 505 510Gly Lys Asn Lys Thr Val Ser
Lys Leu Val Ile Gln Asn Ala Asn Val515 520
525Ser Ala Met Tyr Lys Cys Val Val Ser Asn Lys Val Gly Gln Asp Glu530
535 540Arg Leu Ile Tyr Phe Tyr Val Thr Thr
Ile Pro Asp Gly Phe Thr Ile545 550 555
560Glu Ser Lys Pro Ser Glu Glu Leu Leu Glu Gly Gln Pro Val
Leu Leu565 570 575Ser Cys Gln Ala Asp Ser
Tyr Lys Tyr Glu His Leu Arg Trp Tyr Arg580 585
590Leu Asn Leu Ser Thr Leu His Asp Ala His Gly Asn Pro Leu Leu
Leu595 600 605Asp Cys Lys Asn Val His Leu
Phe Ala Thr Pro Leu Ala Ala Ser Leu610 615
620Glu Glu Val Ala Pro Gly Ala Arg His Ala Thr Leu Ser Leu Ser Ile625
630 635 640Pro Arg Val Ala
Pro Glu His Glu Gly His Tyr Val Cys Glu Val Gln645 650
655Asp Arg Arg Ser His Asp Lys His Cys His Lys Lys Tyr Leu
Ser Val660 665 670Gln Ala Leu Glu Ala Pro
Arg Leu Thr Gln Asn Leu Thr Asp Leu Leu675 680
685Val Asn Val Ser Asp Ser Leu Glu Met Gln Cys Leu Val Ala Gly
Ala690 695 700His Ala Pro Ser Ile Val Trp
Tyr Lys Asp Glu Arg Leu Leu Glu Glu705 710
715 720Lys Ser Gly Val Asp Leu Ala Asp Ser Asn Gln Lys
Leu Ser Ile Gln725 730 735Arg Val Arg Glu
Glu Asp Ala Gly Arg Tyr Leu Cys Ser Val Cys Asn740 745
750Ala Lys Gly Cys Val Asn Ser Ser Ala Ser Val Ala Val Glu
Gly Ser755 760 765Glu Asp Lys Gly Ser Met
Glu Gly Cys Val Ser Gly Asp Thr Ile Val770 775
780Met Thr Ser Gly Gly Pro Arg Thr Val Ala Glu Leu Glu Gly Lys
Pro785 790 795 800Phe Thr
Ala Leu Ile Arg Gly Ser Gly Tyr Pro Cys Pro Ser Gly Phe805
810 815Phe Arg Thr Cys Glu Arg Asp Val Tyr Asp Leu Arg
Thr Arg Glu Gly820 825 830His Cys Leu Arg
Leu Thr His Asp His Arg Val Leu Val Met Asp Gly835 840
845Gly Leu Glu Trp Arg Ala Ala Gly Glu Leu Glu Arg Gly Asp
Arg Leu850 855 860Val Met Asp Asp Ala Ala
Gly Glu Phe Pro Ala Leu Ala Thr Phe Arg865 870
875 880Gly Leu Arg Gly Ala Gly Arg Gln Asp Val Tyr
Asp Ala Thr Val Tyr885 890 895Gly Ala Ser
Ala Phe Thr Ala Asn Gly Phe Ile Val His Ala Cys Gly900
905 910Glu Gln Pro Gly Thr Gly Leu Asn Ser Gly Leu Thr
Thr Asn Pro Gly915 920 925Val Ser Ala Trp
Gln Val Asn Thr Ala Tyr Thr Ala Gly Gln Leu Val930 935
940Thr Tyr Asn Gly Lys Thr Tyr Lys Cys Leu Gln Pro His Thr
Ser Leu945 950 955 960Ala
Gly Trp Glu Pro Ser Asn Val Pro Ala Leu Trp Gln Leu Gln965
970 975177951PRTArtificial Sequenceartificial construct
relating to Homo Sapiens immunoglobulin 177Tyr Ser Met Thr Pro Pro
Thr Leu Asn Ile Thr Glu Glu Ser His Val1 5
10 15Ile Asp Thr Gly Asp Ser Leu Ser Ile Ser Cys Arg
Gly Gln His Pro20 25 30Leu Glu Trp Ala
Trp Pro Gly Ala Gln Glu Ala Pro Ala Thr Gly Asp35 40
45Lys Asp Ser Glu Asp Thr Gly Val Val Arg Asp Cys Glu Gly
Thr Asp50 55 60Ala Arg Pro Tyr Cys Lys
Val Leu Leu Leu His Glu Val His Ala Asn65 70
75 80Asp Thr Gly Ser Tyr Val Cys Tyr Tyr Lys Tyr
Ile Lys Ala Arg Ile85 90 95Glu Gly Thr
Thr Ala Ala Ser Ser Tyr Val Phe Val Arg Asp Phe Glu100
105 110Gln Pro Phe Ile Asn Lys Pro Asp Thr Leu Leu Val
Asn Arg Lys Asp115 120 125Ala Met Trp Val
Pro Cys Leu Val Ser Ile Pro Gly Leu Asn Val Thr130 135
140Leu Arg Ser Gln Ser Ser Val Leu Trp Pro Asp Gly Gln Glu
Val Val145 150 155 160Trp
Asp Asp Arg Arg Gly Met Leu Val Ser Thr Pro Leu Leu His Asp165
170 175Ala Leu Tyr Leu Gln Cys Glu Thr Thr Trp Gly
Asp Gln Asp Phe Leu180 185 190Ser Asn Pro
Phe Leu Val His Ile Thr Gly Asn Glu Leu Tyr Asp Ile195
200 205Gln Leu Leu Pro Arg Lys Ser Leu Glu Leu Leu Val
Gly Glu Lys Leu210 215 220Val Leu Asn Cys
Thr Val Trp Ala Glu Phe Asn Ser Gly Val Thr Phe225 230
235 240Asp Trp Asp Tyr Pro Gly Lys Gln Ala
Glu Arg Gly Lys Trp Val Pro245 250 255Glu
Arg Arg Ser Gln Gln Thr His Thr Glu Leu Ser Ser Ile Leu Thr260
265 270Ile His Asn Val Ser Gln His Asp Leu Gly Ser
Tyr Val Cys Lys Ala275 280 285Asn Asn Gly
Ile Gln Arg Phe Arg Glu Ser Thr Glu Val Ile Val His290
295 300Glu Asn Pro Phe Ile Ser Val Glu Trp Leu Lys Gly
Pro Ile Leu Glu305 310 315
320Ala Thr Ala Gly Asp Glu Leu Val Lys Leu Pro Val Lys Leu Ala Ala325
330 335Tyr Pro Pro Pro Glu Phe Gln Trp Tyr
Lys Asp Gly Lys Ala Leu Ser340 345 350Gly
Arg His Ser Pro His Ala Leu Val Leu Lys Glu Val Thr Glu Ala355
360 365Ser Thr Gly Thr Tyr Thr Leu Ala Leu Trp Asn
Ser Ala Ala Gly Leu370 375 380Arg Arg Asn
Ile Ser Leu Glu Leu Val Val Asn Val Pro Pro Gln Ile385
390 395 400His Glu Lys Glu Ala Ser Ser
Pro Ser Ile Tyr Ser Arg His Ser Arg405 410
415Gln Ala Leu Thr Cys Thr Ala Tyr Gly Val Pro Leu Pro Leu Ser Ile420
425 430Gln Trp His Trp Arg Pro Trp Thr Pro
Cys Lys Met Phe Ala Gln Arg435 440 445Ser
Leu Arg Arg Arg Gln Gln Gln Asp Leu Met Pro Gln Cys Arg Asp450
455 460Trp Arg Ala Val Thr Thr Gln Asp Ala Val Asn
Pro Ile Glu Ser Leu465 470 475
480Asp Thr Trp Thr Glu Phe Val Glu Gly Lys Asn Lys Thr Val Ser
Lys485 490 495Leu Val Ile Gln Asn Ala Asn
Val Ser Ala Met Tyr Lys Cys Val Val500 505
510Ser Asn Lys Val Gly Gln Asp Glu Arg Leu Ile Tyr Phe Tyr Val Thr515
520 525Thr Ile Pro Asp Gly Phe Thr Ile Glu
Ser Lys Pro Ser Glu Glu Leu530 535 540Leu
Glu Gly Gln Pro Val Leu Leu Ser Cys Gln Ala Asp Ser Tyr Lys545
550 555 560Tyr Glu His Leu Arg Trp
Tyr Arg Leu Asn Leu Ser Thr Leu His Asp565 570
575Ala His Gly Asn Pro Leu Leu Leu Asp Cys Lys Asn Val His Leu
Phe580 585 590Ala Thr Pro Leu Ala Ala Ser
Leu Glu Glu Val Ala Pro Gly Ala Arg595 600
605His Ala Thr Leu Ser Leu Ser Ile Pro Arg Val Ala Pro Glu His Glu610
615 620Gly His Tyr Val Cys Glu Val Gln Asp
Arg Arg Ser His Asp Lys His625 630 635
640Cys His Lys Lys Tyr Leu Ser Val Gln Ala Leu Glu Ala Pro
Arg Leu645 650 655Thr Gln Asn Leu Thr Asp
Leu Leu Val Asn Val Ser Asp Ser Leu Glu660 665
670Met Gln Cys Leu Val Ala Gly Ala His Ala Pro Ser Ile Val Trp
Tyr675 680 685Lys Asp Glu Arg Leu Leu Glu
Glu Lys Ser Gly Val Asp Leu Ala Asp690 695
700Ser Asn Gln Lys Leu Ser Ile Gln Arg Val Arg Glu Glu Asp Ala Gly705
710 715 720Arg Tyr Leu Cys
Ser Val Cys Asn Ala Lys Gly Cys Val Asn Ser Ser725 730
735Ala Ser Val Ala Val Glu Gly Ser Glu Asp Lys Gly Ser Met
Glu Gly740 745 750Cys Val Ser Gly Asp Thr
Ile Val Met Thr Ser Gly Gly Pro Arg Thr755 760
765Val Ala Glu Leu Glu Gly Lys Pro Phe Thr Ala Leu Ile Arg Gly
Ser770 775 780Gly Tyr Pro Cys Pro Ser Gly
Phe Phe Arg Thr Cys Glu Arg Asp Val785 790
795 800Tyr Asp Leu Arg Thr Arg Glu Gly His Cys Leu Arg
Leu Thr His Asp805 810 815His Arg Val Leu
Val Met Asp Gly Gly Leu Glu Trp Arg Ala Ala Gly820 825
830Glu Leu Glu Arg Gly Asp Arg Leu Val Met Asp Asp Ala Ala
Gly Glu835 840 845Phe Pro Ala Leu Ala Thr
Phe Arg Gly Leu Arg Gly Ala Gly Arg Gln850 855
860Asp Val Tyr Asp Ala Thr Val Tyr Gly Ala Ser Ala Phe Thr Ala
Asn865 870 875 880Gly Phe
Ile Val His Ala Cys Gly Glu Gln Pro Gly Thr Gly Leu Asn885
890 895Ser Gly Leu Thr Thr Asn Pro Gly Val Ser Ala Trp
Gln Val Asn Thr900 905 910Ala Tyr Thr Ala
Gly Gln Leu Val Thr Tyr Asn Gly Lys Thr Tyr Lys915 920
925Cys Leu Gln Pro His Thr Ser Leu Ala Gly Trp Glu Pro Ser
Asn Val930 935 940Pro Ala Leu Trp Gln Leu
Gln945 950178752PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 178Tyr Ser Met
Thr Pro Pro Thr Leu Asn Ile Thr Glu Glu Ser His Val1 5
10 15Ile Asp Thr Gly Asp Ser Leu Ser Ile
Ser Cys Arg Gly Gln His Pro20 25 30Leu
Glu Trp Ala Trp Pro Gly Ala Gln Glu Ala Pro Ala Thr Gly Asp35
40 45Lys Asp Ser Glu Asp Thr Gly Val Val Arg Asp
Cys Glu Gly Thr Asp50 55 60Ala Arg Pro
Tyr Cys Lys Val Leu Leu Leu His Glu Val His Ala Asn65 70
75 80Asp Thr Gly Ser Tyr Val Cys Tyr
Tyr Lys Tyr Ile Lys Ala Arg Ile85 90
95Glu Gly Thr Thr Ala Ala Ser Ser Tyr Val Phe Val Arg Asp Phe Glu100
105 110Gln Pro Phe Ile Asn Lys Pro Asp Thr Leu
Leu Val Asn Arg Lys Asp115 120 125Ala Met
Trp Val Pro Cys Leu Val Ser Ile Pro Gly Leu Asn Val Thr130
135 140Leu Arg Ser Gln Ser Ser Val Leu Trp Pro Asp Gly
Gln Glu Val Val145 150 155
160Trp Asp Asp Arg Arg Gly Met Leu Val Ser Thr Pro Leu Leu His Asp165
170 175Ala Leu Tyr Leu Gln Cys Glu Thr Thr
Trp Gly Asp Gln Asp Phe Leu180 185 190Ser
Asn Pro Phe Leu Val His Ile Thr Gly Asn Glu Leu Tyr Asp Ile195
200 205Gln Leu Leu Pro Arg Lys Ser Leu Glu Leu Leu
Val Gly Glu Lys Leu210 215 220Val Leu Asn
Cys Thr Val Trp Ala Glu Phe Asn Ser Gly Val Thr Phe225
230 235 240Asp Trp Asp Tyr Pro Gly Lys
Gln Ala Glu Arg Gly Lys Trp Val Pro245 250
255Glu Arg Arg Ser Gln Gln Thr His Thr Glu Leu Ser Ser Ile Leu Thr260
265 270Ile His Asn Val Ser Gln His Asp Leu
Gly Ser Tyr Val Cys Lys Ala275 280 285Asn
Asn Gly Ile Gln Arg Phe Arg Glu Ser Thr Glu Val Ile Val His290
295 300Glu Asn Pro Phe Ile Ser Val Glu Trp Leu Lys
Gly Pro Ile Leu Glu305 310 315
320Ala Thr Ala Gly Asp Glu Leu Val Lys Leu Pro Val Lys Leu Ala
Ala325 330 335Tyr Pro Pro Pro Glu Phe Gln
Trp Tyr Lys Asp Gly Lys Ala Leu Ser340 345
350Gly Arg His Ser Pro His Ala Leu Val Leu Lys Glu Val Thr Glu Ala355
360 365Ser Thr Gly Thr Tyr Thr Leu Ala Leu
Trp Asn Ser Ala Ala Gly Leu370 375 380Arg
Arg Asn Ile Ser Leu Glu Leu Val Val Asn Val Pro Pro Gln Ile385
390 395 400His Glu Lys Glu Ala Ser
Ser Pro Ser Ile Tyr Ser Arg His Ser Arg405 410
415Gln Ala Leu Thr Cys Thr Ala Tyr Gly Val Pro Leu Pro Leu Ser
Ile420 425 430Gln Trp His Trp Arg Pro Trp
Thr Pro Cys Lys Met Phe Ala Gln Arg435 440
445Ser Leu Arg Arg Arg Gln Gln Gln Asp Leu Met Pro Gln Cys Arg Asp450
455 460Trp Arg Ala Val Thr Thr Gln Asp Ala
Val Asn Pro Ile Glu Ser Leu465 470 475
480Asp Thr Trp Thr Glu Phe Val Glu Gly Lys Asn Lys Thr Val
Ser Lys485 490 495Leu Val Ile Gln Asn Ala
Asn Val Ser Ala Met Tyr Lys Cys Val Val500 505
510Ser Asn Lys Val Gly Gln Asp Glu Arg Leu Ile Tyr Phe Tyr Val
Thr515 520 525Thr Ile Pro Asp Gly Phe Thr
Ile Glu Ser Lys Pro Ser Glu Glu Leu530 535
540Leu Glu Gly Gln Pro Val Leu Leu Ser Cys Gln Ala Asp Ser Tyr Lys545
550 555 560Tyr Glu His Leu
Arg Trp Tyr Arg Leu Asn Leu Ser Thr Leu His Asp565 570
575Ala His Gly Asn Pro Leu Leu Leu Asp Cys Lys Asn Val His
Leu Phe580 585 590Ala Thr Pro Leu Ala Ala
Ser Leu Glu Glu Val Ala Pro Gly Ala Arg595 600
605His Ala Thr Leu Ser Leu Ser Ile Pro Arg Val Ala Pro Glu His
Glu610 615 620Gly His Tyr Val Cys Glu Val
Gln Asp Arg Arg Ser His Asp Lys His625 630
635 640Cys His Lys Lys Tyr Leu Ser Val Gln Ala Leu Glu
Ala Pro Arg Leu645 650 655Thr Gln Asn Leu
Thr Asp Leu Leu Val Asn Val Ser Asp Ser Leu Glu660 665
670Met Gln Cys Leu Val Ala Gly Ala His Ala Pro Ser Ile Val
Trp Tyr675 680 685Lys Asp Glu Arg Leu Leu
Glu Glu Lys Ser Gly Val Asp Leu Ala Asp690 695
700Ser Asn Gln Lys Leu Ser Ile Gln Arg Val Arg Glu Glu Asp Ala
Gly705 710 715 720Arg Tyr
Leu Cys Ser Val Cys Asn Ala Lys Gly Cys Val Asn Ser Ser725
730 735Ala Ser Val Ala Val Glu Gly Ser Glu Asp Lys Gly
Ser Met Glu Xaa740 745
750179753PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 179Tyr Ser Met Thr Pro Pro Thr Leu Asn Ile
Thr Glu Glu Ser His Val1 5 10
15Ile Asp Thr Gly Asp Ser Leu Ser Ile Ser Cys Arg Gly Gln His Pro20
25 30Leu Glu Trp Ala Trp Pro Gly Ala Gln
Glu Ala Pro Ala Thr Gly Asp35 40 45Lys
Asp Ser Glu Asp Thr Gly Val Val Arg Asp Cys Glu Gly Thr Asp50
55 60Ala Arg Pro Tyr Cys Lys Val Leu Leu Leu His
Glu Val His Ala Asn65 70 75
80Asp Thr Gly Ser Tyr Val Cys Tyr Tyr Lys Tyr Ile Lys Ala Arg Ile85
90 95Glu Gly Thr Thr Ala Ala Ser Ser Tyr
Val Phe Val Arg Asp Phe Glu100 105 110Gln
Pro Phe Ile Asn Lys Pro Asp Thr Leu Leu Val Asn Arg Lys Asp115
120 125Ala Met Trp Val Pro Cys Leu Val Ser Ile Pro
Gly Leu Asn Val Thr130 135 140Leu Arg Ser
Gln Ser Ser Val Leu Trp Pro Asp Gly Gln Glu Val Val145
150 155 160Trp Asp Asp Arg Arg Gly Met
Leu Val Ser Thr Pro Leu Leu His Asp165 170
175Ala Leu Tyr Leu Gln Cys Glu Thr Thr Trp Gly Asp Gln Asp Phe Leu180
185 190Ser Asn Pro Phe Leu Val His Ile Thr
Gly Asn Glu Leu Tyr Asp Ile195 200 205Gln
Leu Leu Pro Arg Lys Ser Leu Glu Leu Leu Val Gly Glu Lys Leu210
215 220Val Leu Asn Cys Thr Val Trp Ala Glu Phe Asn
Ser Gly Val Thr Phe225 230 235
240Asp Trp Asp Tyr Pro Gly Lys Gln Ala Glu Arg Gly Lys Trp Val
Pro245 250 255Glu Arg Arg Ser Gln Gln Thr
His Thr Glu Leu Ser Ser Ile Leu Thr260 265
270Ile His Asn Val Ser Gln His Asp Leu Gly Ser Tyr Val Cys Lys Ala275
280 285Asn Asn Gly Ile Gln Arg Phe Arg Glu
Ser Thr Glu Val Ile Val His290 295 300Glu
Asn Pro Phe Ile Ser Val Glu Trp Leu Lys Gly Pro Ile Leu Glu305
310 315 320Ala Thr Ala Gly Asp Glu
Leu Val Lys Leu Pro Val Lys Leu Ala Ala325 330
335Tyr Pro Pro Pro Glu Phe Gln Trp Tyr Lys Asp Gly Lys Ala Leu
Ser340 345 350Gly Arg His Ser Pro His Ala
Leu Val Leu Lys Glu Val Thr Glu Ala355 360
365Ser Thr Gly Thr Tyr Thr Leu Ala Leu Trp Asn Ser Ala Ala Gly Leu370
375 380Arg Arg Asn Ile Ser Leu Glu Leu Val
Val Asn Val Pro Pro Gln Ile385 390 395
400His Glu Lys Glu Ala Ser Ser Pro Ser Ile Tyr Ser Arg His
Ser Arg405 410 415Gln Ala Leu Thr Cys Thr
Ala Tyr Gly Val Pro Leu Pro Leu Ser Ile420 425
430Gln Trp His Trp Arg Pro Trp Thr Pro Cys Lys Met Phe Ala Gln
Arg435 440 445Ser Leu Arg Arg Arg Gln Gln
Gln Asp Leu Met Pro Gln Cys Arg Asp450 455
460Trp Arg Ala Val Thr Thr Gln Asp Ala Val Asn Pro Ile Glu Ser Leu465
470 475 480Asp Thr Trp Thr
Glu Phe Val Glu Gly Lys Asn Lys Thr Val Ser Lys485 490
495Leu Val Ile Gln Asn Ala Asn Val Ser Ala Met Tyr Lys Cys
Val Val500 505 510Ser Asn Lys Val Gly Gln
Asp Glu Arg Leu Ile Tyr Phe Tyr Val Thr515 520
525Thr Ile Pro Asp Gly Phe Thr Ile Glu Ser Lys Pro Ser Glu Glu
Leu530 535 540Leu Glu Gly Gln Pro Val Leu
Leu Ser Cys Gln Ala Asp Ser Tyr Lys545 550
555 560Tyr Glu His Leu Arg Trp Tyr Arg Leu Asn Leu Ser
Thr Leu His Asp565 570 575Ala His Gly Asn
Pro Leu Leu Leu Asp Cys Lys Asn Val His Leu Phe580 585
590Ala Thr Pro Leu Ala Ala Ser Leu Glu Glu Val Ala Pro Gly
Ala Arg595 600 605His Ala Thr Leu Ser Leu
Ser Ile Pro Arg Val Ala Pro Glu His Glu610 615
620Gly His Tyr Val Cys Glu Val Gln Asp Arg Arg Ser His Asp Lys
His625 630 635 640Cys His
Lys Lys Tyr Leu Ser Val Gln Ala Leu Glu Ala Pro Arg Leu645
650 655Thr Gln Asn Leu Thr Asp Leu Leu Val Asn Val Ser
Asp Ser Leu Glu660 665 670Met Gln Cys Leu
Val Ala Gly Ala His Ala Pro Ser Ile Val Trp Tyr675 680
685Lys Asp Glu Arg Leu Leu Glu Glu Lys Ser Gly Val Asp Leu
Ala Asp690 695 700Ser Asn Gln Lys Leu Ser
Ile Gln Arg Val Arg Glu Glu Asp Ala Gly705 710
715 720Arg Tyr Leu Cys Ser Val Cys Asn Ala Lys Gly
Cys Val Asn Ser Ser725 730 735Ala Ser Val
Ala Val Glu Gly Ser Glu Asp Lys Gly Ser Met Glu Gly740
745 750Xaa1802567DNAArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 180aagcttgaat
tcccaccatg cgaccctccg ggacggccgg ggcagcgctc ctggcgctgc 60tggctgcgct
ctgcccggcg agtcgggctc tggaggaaaa gaaagtttgc caaggcacga 120gtaacaagct
cacgcagttg ggcacttttg aagatcattt tctcagcctc cagaggatgt 180tcaataactg
tgaggtggtc cttgggaatt tggaaattac ctatgtgcag aggaattatg 240atctttcctt
cttaaagacc atccaggagg tggctggtta tgtcctcatt gccctcaaca 300cagtggagcg
aatccctttg gaaaacctgc agatcatcag aggaaatatg tactacgaaa 360attcctatgc
cttagcagtc ttatctaact atgatgcaaa taaaaccgga ctgaaggagc 420tgcccatgag
aaatttacag gaaatcctgc atggcgccgt gcggttcagc aacaaccctg 480ccctgtgcaa
cgtggagagc atccagtggc gggacatagt cagcagtgac tttctcagca 540acatgtcgat
ggacttccag aaccacctgg gcagctgcca aaagtgtgat ccaagctgtc 600ccaatgggag
ctgctggggt gcaggagagg agaactgcca gaaactgacc aaaatcatct 660gtgcccagca
gtgctccggg cgctgccgtg gcaagtcccc cagtgactgc tgccacaacc 720agtgtgctgc
aggctgcaca ggcccccggg agagcgactg cctggtctgc cgcaaattcc 780gagacgaagc
cacgtgcaag gacacctgcc ccccactcat gctctacaac cccaccacgt 840accagatgga
tgtgaacccc gagggcaaat acagctttgg tgccacctgc gtgaagaagt 900gtccccgtaa
ttatgtggtg acagatcacg gctcgtgcgt ccgagcctgt ggggccgaca 960gctatgagat
ggaggaagac ggcgtccgca agtgtaagaa gtgcgaaggg ccttgccgca 1020aagtgtgtaa
cggaataggt attggtgaat ttaaagactc actctccata aatgctacga 1080atattaaaca
cttcaaaaac tgcacctcca tcagtggcga tctccacatc ctgccggtgg 1140catttagggg
tgactccttc acacatactc ctcctctgga tccacaggaa ctggatattc 1200tgaaaaccgt
aaaggaaatc acagggtttt tgctgattca ggcttggcct gaaaacagga 1260cggacctcca
tgcctttgag aacctagaaa tcatacgcgg caggaccaag caacatggtc 1320agttttctct
tgcagtcgtc agcctgaaca taacatcctt gggattacgc tccctcaagg 1380agataagtga
tggagatgtg ataatttcag gaaacaaaaa tttgtgctat gcaaatacaa 1440taaactggaa
aaaactgttt gggacctccg gtcagaaaac caaaattata agcaacagag 1500gtgaaaacag
ctgcaaggcc acaggccagg tctgccatgc cttgtgctcc cccgagggct 1560gctggggtcc
ggagcccagg gactgcgtct cttgccggaa tgtcagccga ggcagggaat 1620gcgtggacaa
gtgcaagctc ctggagggtg agccaaggga gtttgtggag aactctgagt 1680gcatacagtg
ccacccagag tgcctgcctc aggccatgaa catcacctgc acaggacggg 1740gaccagacaa
ctgtatccag tgtgcccact acattgacgg cccccactgc gtcaagacct 1800gcccggcagg
agtcatggga gaaaacaaca ccctggtctg gaagtacgca gacgccggcc 1860atgtgtgcca
cctgtgccat ccaaactgca cctacggatg cactgggcca ggtcttgaag 1920gctgtccaac
gaatgggcct aagatcccgt ccgggtgcgt atccggtgac accattgtaa 1980tgactagtgg
cgggccccgc actgtggctg aactggaggg caaaccgttc accgcactga 2040ttcgcggctc
tggctaccca tgcccctcag gtttcttccg cacctgtgaa cgtgacgtat 2100atgatctgcg
tacacgtgag ggtcattgct tacgtttgac ccatgatcac cgtgttctgg 2160tgatggatgg
tggcctggaa tggcgtgccg cgggtgaact ggaacgcggc gaccgcctgg 2220tgatggatga
tgcagctggc gagtttccgg cactggcaac cttccgtggc ctgcgtggcg 2280ctggccgcca
ggatgtttat gacgctactg tttacggtgc tagcgcattc actgctaatg 2340gcttcattgt
acacgcatgt ggcgagcagc ccgggaccgg tctgaactca ggcctcacga 2400caaatcctgg
tgtatccgct tggcaggtca acacagctta tactgcggga caattggtca 2460catataacgg
caagacgtat aaatgtttgc agccccacac ctccttggca ggatgggaac 2520catccaacgt
tcctgccttg tggcagcttc aatgactcga gcggccg
2567181845PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 181Met Arg Pro Ser Gly Thr Ala Gly Ala Ala
Leu Leu Ala Leu Leu Ala1 5 10
15Ala Leu Cys Pro Ala Ser Arg Ala Leu Glu Glu Lys Lys Val Cys Gln20
25 30Gly Thr Ser Asn Lys Leu Thr Gln Leu
Gly Thr Phe Glu Asp His Phe35 40 45Leu
Ser Leu Gln Arg Met Phe Asn Asn Cys Glu Val Val Leu Gly Asn50
55 60Leu Glu Ile Thr Tyr Val Gln Arg Asn Tyr Asp
Leu Ser Phe Leu Lys65 70 75
80Thr Ile Gln Glu Val Ala Gly Tyr Val Leu Ile Ala Leu Asn Thr Val85
90 95Glu Arg Ile Pro Leu Glu Asn Leu Gln
Ile Ile Arg Gly Asn Met Tyr100 105 110Tyr
Glu Asn Ser Tyr Ala Leu Ala Val Leu Ser Asn Tyr Asp Ala Asn115
120 125Lys Thr Gly Leu Lys Glu Leu Pro Met Arg Asn
Leu Gln Glu Ile Leu130 135 140His Gly Ala
Val Arg Phe Ser Asn Asn Pro Ala Leu Cys Asn Val Glu145
150 155 160Ser Ile Gln Trp Arg Asp Ile
Val Ser Ser Asp Phe Leu Ser Asn Met165 170
175Ser Met Asp Phe Gln Asn His Leu Gly Ser Cys Gln Lys Cys Asp Pro180
185 190Ser Cys Pro Asn Gly Ser Cys Trp Gly
Ala Gly Glu Glu Asn Cys Gln195 200 205Lys
Leu Thr Lys Ile Ile Cys Ala Gln Gln Cys Ser Gly Arg Cys Arg210
215 220Gly Lys Ser Pro Ser Asp Cys Cys His Asn Gln
Cys Ala Ala Gly Cys225 230 235
240Thr Gly Pro Arg Glu Ser Asp Cys Leu Val Cys Arg Lys Phe Arg
Asp245 250 255Glu Ala Thr Cys Lys Asp Thr
Cys Pro Pro Leu Met Leu Tyr Asn Pro260 265
270Thr Thr Tyr Gln Met Asp Val Asn Pro Glu Gly Lys Tyr Ser Phe Gly275
280 285Ala Thr Cys Val Lys Lys Cys Pro Arg
Asn Tyr Val Val Thr Asp His290 295 300Gly
Ser Cys Val Arg Ala Cys Gly Ala Asp Ser Tyr Glu Met Glu Glu305
310 315 320Asp Gly Val Arg Lys Cys
Lys Lys Cys Glu Gly Pro Cys Arg Lys Val325 330
335Cys Asn Gly Ile Gly Ile Gly Glu Phe Lys Asp Ser Leu Ser Ile
Asn340 345 350Ala Thr Asn Ile Lys His Phe
Lys Asn Cys Thr Ser Ile Ser Gly Asp355 360
365Leu His Ile Leu Pro Val Ala Phe Arg Gly Asp Ser Phe Thr His Thr370
375 380Pro Pro Leu Asp Pro Gln Glu Leu Asp
Ile Leu Lys Thr Val Lys Glu385 390 395
400Ile Thr Gly Phe Leu Leu Ile Gln Ala Trp Pro Glu Asn Arg
Thr Asp405 410 415Leu His Ala Phe Glu Asn
Leu Glu Ile Ile Arg Gly Arg Thr Lys Gln420 425
430His Gly Gln Phe Ser Leu Ala Val Val Ser Leu Asn Ile Thr Ser
Leu435 440 445Gly Leu Arg Ser Leu Lys Glu
Ile Ser Asp Gly Asp Val Ile Ile Ser450 455
460Gly Asn Lys Asn Leu Cys Tyr Ala Asn Thr Ile Asn Trp Lys Lys Leu465
470 475 480Phe Gly Thr Ser
Gly Gln Lys Thr Lys Ile Ile Ser Asn Arg Gly Glu485 490
495Asn Ser Cys Lys Ala Thr Gly Gln Val Cys His Ala Leu Cys
Ser Pro500 505 510Glu Gly Cys Trp Gly Pro
Glu Pro Arg Asp Cys Val Ser Cys Arg Asn515 520
525Val Ser Arg Gly Arg Glu Cys Val Asp Lys Cys Asn Leu Leu Glu
Gly530 535 540Glu Pro Arg Glu Phe Val Glu
Asn Ser Glu Cys Ile Gln Cys His Pro545 550
555 560Glu Cys Leu Pro Gln Ala Met Asn Ile Thr Cys Thr
Gly Arg Gly Pro565 570 575Asp Asn Cys Ile
Gln Cys Ala His Tyr Ile Asp Gly Pro His Cys Val580 585
590Lys Thr Cys Pro Ala Gly Val Met Gly Glu Asn Asn Thr Leu
Val Trp595 600 605Lys Tyr Ala Asp Ala Gly
His Val Cys His Leu Cys His Pro Asn Cys610 615
620Thr Tyr Gly Cys Thr Gly Pro Gly Leu Glu Gly Cys Pro Thr Asn
Gly625 630 635 640Pro Lys
Ile Pro Ser Gly Cys Val Ser Gly Asp Thr Ile Val Met Thr645
650 655Ser Gly Gly Pro Arg Thr Val Ala Glu Leu Glu Gly
Lys Pro Phe Thr660 665 670Ala Leu Ile Arg
Gly Ser Gly Tyr Pro Cys Pro Ser Gly Phe Phe Arg675 680
685Thr Cys Glu Arg Asp Val Tyr Asp Leu Arg Thr Arg Glu Gly
His Cys690 695 700Leu Arg Leu Thr His Asp
His Arg Val Leu Val Met Asp Gly Gly Leu705 710
715 720Glu Trp Arg Ala Ala Gly Glu Leu Glu Arg Gly
Asp Arg Leu Val Met725 730 735Asp Asp Ala
Ala Gly Glu Phe Pro Ala Leu Ala Thr Phe Arg Gly Leu740
745 750Arg Gly Ala Gly Arg Gln Asp Val Tyr Asp Ala Thr
Val Tyr Gly Ala755 760 765Ser Ala Phe Thr
Ala Asn Gly Phe Ile Val His Ala Cys Gly Glu Gln770 775
780Pro Gly Thr Gly Leu Asn Ser Gly Leu Thr Thr Asn Pro Gly
Val Ser785 790 795 800Ala
Trp Gln Val Asn Thr Ala Tyr Thr Ala Gly Gln Leu Val Thr Tyr805
810 815Asn Gly Lys Thr Tyr Lys Cys Leu Gln Pro His
Thr Ser Leu Ala Gly820 825 830Trp Glu Pro
Ser Asn Val Pro Ala Leu Trp Gln Leu Gln835 840
845182821PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 182Leu Glu Glu Lys Lys Val Cys Gln Gly Thr
Ser Asn Lys Leu Thr Gln1 5 10
15Leu Gly Thr Phe Glu Asp His Phe Leu Ser Leu Gln Arg Met Phe Asn20
25 30Asn Cys Glu Val Val Leu Gly Asn Leu
Glu Ile Thr Tyr Val Gln Arg35 40 45Asn
Tyr Asp Leu Ser Phe Leu Lys Thr Ile Gln Glu Val Ala Gly Tyr50
55 60Val Leu Ile Ala Leu Asn Thr Val Glu Arg Ile
Pro Leu Glu Asn Leu65 70 75
80Gln Ile Ile Arg Gly Asn Met Tyr Tyr Glu Asn Ser Tyr Ala Leu Ala85
90 95Val Leu Ser Asn Tyr Asp Ala Asn Lys
Thr Gly Leu Lys Glu Leu Pro100 105 110Met
Arg Asn Leu Gln Glu Ile Leu His Gly Ala Val Arg Phe Ser Asn115
120 125Asn Pro Ala Leu Cys Asn Val Glu Ser Ile Gln
Trp Arg Asp Ile Val130 135 140Ser Ser Asp
Phe Leu Ser Asn Met Ser Met Asp Phe Gln Asn His Leu145
150 155 160Gly Ser Cys Gln Lys Cys Asp
Pro Ser Cys Pro Asn Gly Ser Cys Trp165 170
175Gly Ala Gly Glu Glu Asn Cys Gln Lys Leu Thr Lys Ile Ile Cys Ala180
185 190Gln Gln Cys Ser Gly Arg Cys Arg Gly
Lys Ser Pro Ser Asp Cys Cys195 200 205His
Asn Gln Cys Ala Ala Gly Cys Thr Gly Pro Arg Glu Ser Asp Cys210
215 220Leu Val Cys Arg Lys Phe Arg Asp Glu Ala Thr
Cys Lys Asp Thr Cys225 230 235
240Pro Pro Leu Met Leu Tyr Asn Pro Thr Thr Tyr Gln Met Asp Val
Asn245 250 255Pro Glu Gly Lys Tyr Ser Phe
Gly Ala Thr Cys Val Lys Lys Cys Pro260 265
270Arg Asn Tyr Val Val Thr Asp His Gly Ser Cys Val Arg Ala Cys Gly275
280 285Ala Asp Ser Tyr Glu Met Glu Glu Asp
Gly Val Arg Lys Cys Lys Lys290 295 300Cys
Glu Gly Pro Cys Arg Lys Val Cys Asn Gly Ile Gly Ile Gly Glu305
310 315 320Phe Lys Asp Ser Leu Ser
Ile Asn Ala Thr Asn Ile Lys His Phe Lys325 330
335Asn Cys Thr Ser Ile Ser Gly Asp Leu His Ile Leu Pro Val Ala
Phe340 345 350Arg Gly Asp Ser Phe Thr His
Thr Pro Pro Leu Asp Pro Gln Glu Leu355 360
365Asp Ile Leu Lys Thr Val Lys Glu Ile Thr Gly Phe Leu Leu Ile Gln370
375 380Ala Trp Pro Glu Asn Arg Thr Asp Leu
His Ala Phe Glu Asn Leu Glu385 390 395
400Ile Ile Arg Gly Arg Thr Lys Gln His Gly Gln Phe Ser Leu
Ala Val405 410 415Val Ser Leu Asn Ile Thr
Ser Leu Gly Leu Arg Ser Leu Lys Glu Ile420 425
430Ser Asp Gly Asp Val Ile Ile Ser Gly Asn Lys Asn Leu Cys Tyr
Ala435 440 445Asn Thr Ile Asn Trp Lys Lys
Leu Phe Gly Thr Ser Gly Gln Lys Thr450 455
460Lys Ile Ile Ser Asn Arg Gly Glu Asn Ser Cys Lys Ala Thr Gly Gln465
470 475 480Val Cys His Ala
Leu Cys Ser Pro Glu Gly Cys Trp Gly Pro Glu Pro485 490
495Arg Asp Cys Val Ser Cys Arg Asn Val Ser Arg Gly Arg Glu
Cys Val500 505 510Asp Lys Cys Asn Leu Leu
Glu Gly Glu Pro Arg Glu Phe Val Glu Asn515 520
525Ser Glu Cys Ile Gln Cys His Pro Glu Cys Leu Pro Gln Ala Met
Asn530 535 540Ile Thr Cys Thr Gly Arg Gly
Pro Asp Asn Cys Ile Gln Cys Ala His545 550
555 560Tyr Ile Asp Gly Pro His Cys Val Lys Thr Cys Pro
Ala Gly Val Met565 570 575Gly Glu Asn Asn
Thr Leu Val Trp Lys Tyr Ala Asp Ala Gly His Val580 585
590Cys His Leu Cys His Pro Asn Cys Thr Tyr Gly Cys Thr Gly
Pro Gly595 600 605Leu Glu Gly Cys Pro Thr
Asn Gly Pro Lys Ile Pro Ser Gly Cys Val610 615
620Ser Gly Asp Thr Ile Val Met Thr Ser Gly Gly Pro Arg Thr Val
Ala625 630 635 640Glu Leu
Glu Gly Lys Pro Phe Thr Ala Leu Ile Arg Gly Ser Gly Tyr645
650 655Pro Cys Pro Ser Gly Phe Phe Arg Thr Cys Glu Arg
Asp Val Tyr Asp660 665 670Leu Arg Thr Arg
Glu Gly His Cys Leu Arg Leu Thr His Asp His Arg675 680
685Val Leu Val Met Asp Gly Gly Leu Glu Trp Arg Ala Ala Gly
Glu Leu690 695 700Glu Arg Gly Asp Arg Leu
Val Met Asp Asp Ala Ala Gly Glu Phe Pro705 710
715 720Ala Leu Ala Thr Phe Arg Gly Leu Arg Gly Ala
Gly Arg Gln Asp Val725 730 735Tyr Asp Ala
Thr Val Tyr Gly Ala Ser Ala Phe Thr Ala Asn Gly Phe740
745 750Ile Val His Ala Cys Gly Glu Gln Pro Gly Thr Gly
Leu Asn Ser Gly755 760 765Leu Thr Thr Asn
Pro Gly Val Ser Ala Trp Gln Val Asn Thr Ala Tyr770 775
780Thr Ala Gly Gln Leu Val Thr Tyr Asn Gly Lys Thr Tyr Lys
Cys Leu785 790 795 800Gln
Pro His Thr Ser Leu Ala Gly Trp Glu Pro Ser Asn Val Pro Ala805
810 815Leu Trp Gln Leu Gln820183622PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 183Leu Glu Glu Lys Lys Val Cys Gln Gly Thr Ser Asn Lys Leu
Thr Gln1 5 10 15Leu Gly
Thr Phe Glu Asp His Phe Leu Ser Leu Gln Arg Met Phe Asn20
25 30Asn Cys Glu Val Val Leu Gly Asn Leu Glu Ile Thr
Tyr Val Gln Arg35 40 45Asn Tyr Asp Leu
Ser Phe Leu Lys Thr Ile Gln Glu Val Ala Gly Tyr50 55
60Val Leu Ile Ala Leu Asn Thr Val Glu Arg Ile Pro Leu Glu
Asn Leu65 70 75 80Gln
Ile Ile Arg Gly Asn Met Tyr Tyr Glu Asn Ser Tyr Ala Leu Ala85
90 95Val Leu Ser Asn Tyr Asp Ala Asn Lys Thr Gly
Leu Lys Glu Leu Pro100 105 110Met Arg Asn
Leu Gln Glu Ile Leu His Gly Ala Val Arg Phe Ser Asn115
120 125Asn Pro Ala Leu Cys Asn Val Glu Ser Ile Gln Trp
Arg Asp Ile Val130 135 140Ser Ser Asp Phe
Leu Ser Asn Met Ser Met Asp Phe Gln Asn His Leu145 150
155 160Gly Ser Cys Gln Lys Cys Asp Pro Ser
Cys Pro Asn Gly Ser Cys Trp165 170 175Gly
Ala Gly Glu Glu Asn Cys Gln Lys Leu Thr Lys Ile Ile Cys Ala180
185 190Gln Gln Cys Ser Gly Arg Cys Arg Gly Lys Ser
Pro Ser Asp Cys Cys195 200 205His Asn Gln
Cys Ala Ala Gly Cys Thr Gly Pro Arg Glu Ser Asp Cys210
215 220Leu Val Cys Arg Lys Phe Arg Asp Glu Ala Thr Cys
Lys Asp Thr Cys225 230 235
240Pro Pro Leu Met Leu Tyr Asn Pro Thr Thr Tyr Gln Met Asp Val Asn245
250 255Pro Glu Gly Lys Tyr Ser Phe Gly Ala
Thr Cys Val Lys Lys Cys Pro260 265 270Arg
Asn Tyr Val Val Thr Asp His Gly Ser Cys Val Arg Ala Cys Gly275
280 285Ala Asp Ser Tyr Glu Met Glu Glu Asp Gly Val
Arg Lys Cys Lys Lys290 295 300Cys Glu Gly
Pro Cys Arg Lys Val Cys Asn Gly Ile Gly Ile Gly Glu305
310 315 320Phe Lys Asp Ser Leu Ser Ile
Asn Ala Thr Asn Ile Lys His Phe Lys325 330
335Asn Cys Thr Ser Ile Ser Gly Asp Leu His Ile Leu Pro Val Ala Phe340
345 350Arg Gly Asp Ser Phe Thr His Thr Pro
Pro Leu Asp Pro Gln Glu Leu355 360 365Asp
Ile Leu Lys Thr Val Lys Glu Ile Thr Gly Phe Leu Leu Ile Gln370
375 380Ala Trp Pro Glu Asn Arg Thr Asp Leu His Ala
Phe Glu Asn Leu Glu385 390 395
400Ile Ile Arg Gly Arg Thr Lys Gln His Gly Gln Phe Ser Leu Ala
Val405 410 415Val Ser Leu Asn Ile Thr Ser
Leu Gly Leu Arg Ser Leu Lys Glu Ile420 425
430Ser Asp Gly Asp Val Ile Ile Ser Gly Asn Lys Asn Leu Cys Tyr Ala435
440 445Asn Thr Ile Asn Trp Lys Lys Leu Phe
Gly Thr Ser Gly Gln Lys Thr450 455 460Lys
Ile Ile Ser Asn Arg Gly Glu Asn Ser Cys Lys Ala Thr Gly Gln465
470 475 480Val Cys His Ala Leu Cys
Ser Pro Glu Gly Cys Trp Gly Pro Glu Pro485 490
495Arg Asp Cys Val Ser Cys Arg Asn Val Ser Arg Gly Arg Glu Cys
Val500 505 510Asp Lys Cys Asn Leu Leu Glu
Gly Glu Pro Arg Glu Phe Val Glu Asn515 520
525Ser Glu Cys Ile Gln Cys His Pro Glu Cys Leu Pro Gln Ala Met Asn530
535 540Ile Thr Cys Thr Gly Arg Gly Pro Asp
Asn Cys Ile Gln Cys Ala His545 550 555
560Tyr Ile Asp Gly Pro His Cys Val Lys Thr Cys Pro Ala Gly
Val Met565 570 575Gly Glu Asn Asn Thr Leu
Val Trp Lys Tyr Ala Asp Ala Gly His Val580 585
590Cys His Leu Cys His Pro Asn Cys Thr Tyr Gly Cys Thr Gly Pro
Gly595 600 605Leu Glu Gly Cys Pro Thr Asn
Gly Pro Lys Ile Pro Ser Xaa610 615
620184623PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 184Leu Glu Glu Lys Lys Val Cys Gln Gly Thr
Ser Asn Lys Leu Thr Gln1 5 10
15Leu Gly Thr Phe Glu Asp His Phe Leu Ser Leu Gln Arg Met Phe Asn20
25 30Asn Cys Glu Val Val Leu Gly Asn Leu
Glu Ile Thr Tyr Val Gln Arg35 40 45Asn
Tyr Asp Leu Ser Phe Leu Lys Thr Ile Gln Glu Val Ala Gly Tyr50
55 60Val Leu Ile Ala Leu Asn Thr Val Glu Arg Ile
Pro Leu Glu Asn Leu65 70 75
80Gln Ile Ile Arg Gly Asn Met Tyr Tyr Glu Asn Ser Tyr Ala Leu Ala85
90 95Val Leu Ser Asn Tyr Asp Ala Asn Lys
Thr Gly Leu Lys Glu Leu Pro100 105 110Met
Arg Asn Leu Gln Glu Ile Leu His Gly Ala Val Arg Phe Ser Asn115
120 125Asn Pro Ala Leu Cys Asn Val Glu Ser Ile Gln
Trp Arg Asp Ile Val130 135 140Ser Ser Asp
Phe Leu Ser Asn Met Ser Met Asp Phe Gln Asn His Leu145
150 155 160Gly Ser Cys Gln Lys Cys Asp
Pro Ser Cys Pro Asn Gly Ser Cys Trp165 170
175Gly Ala Gly Glu Glu Asn Cys Gln Lys Leu Thr Lys Ile Ile Cys Ala180
185 190Gln Gln Cys Ser Gly Arg Cys Arg Gly
Lys Ser Pro Ser Asp Cys Cys195 200 205His
Asn Gln Cys Ala Ala Gly Cys Thr Gly Pro Arg Glu Ser Asp Cys210
215 220Leu Val Cys Arg Lys Phe Arg Asp Glu Ala Thr
Cys Lys Asp Thr Cys225 230 235
240Pro Pro Leu Met Leu Tyr Asn Pro Thr Thr Tyr Gln Met Asp Val
Asn245 250 255Pro Glu Gly Lys Tyr Ser Phe
Gly Ala Thr Cys Val Lys Lys Cys Pro260 265
270Arg Asn Tyr Val Val Thr Asp His Gly Ser Cys Val Arg Ala Cys Gly275
280 285Ala Asp Ser Tyr Glu Met Glu Glu Asp
Gly Val Arg Lys Cys Lys Lys290 295 300Cys
Glu Gly Pro Cys Arg Lys Val Cys Asn Gly Ile Gly Ile Gly Glu305
310 315 320Phe Lys Asp Ser Leu Ser
Ile Asn Ala Thr Asn Ile Lys His Phe Lys325 330
335Asn Cys Thr Ser Ile Ser Gly Asp Leu His Ile Leu Pro Val Ala
Phe340 345 350Arg Gly Asp Ser Phe Thr His
Thr Pro Pro Leu Asp Pro Gln Glu Leu355 360
365Asp Ile Leu Lys Thr Val Lys Glu Ile Thr Gly Phe Leu Leu Ile Gln370
375 380Ala Trp Pro Glu Asn Arg Thr Asp Leu
His Ala Phe Glu Asn Leu Glu385 390 395
400Ile Ile Arg Gly Arg Thr Lys Gln His Gly Gln Phe Ser Leu
Ala Val405 410 415Val Ser Leu Asn Ile Thr
Ser Leu Gly Leu Arg Ser Leu Lys Glu Ile420 425
430Ser Asp Gly Asp Val Ile Ile Ser Gly Asn Lys Asn Leu Cys Tyr
Ala435 440 445Asn Thr Ile Asn Trp Lys Lys
Leu Phe Gly Thr Ser Gly Gln Lys Thr450 455
460Lys Ile Ile Ser Asn Arg Gly Glu Asn Ser Cys Lys Ala Thr Gly Gln465
470 475 480Val Cys His Ala
Leu Cys Ser Pro Glu Gly Cys Trp Gly Pro Glu Pro485 490
495Arg Asp Cys Val Ser Cys Arg Asn Val Ser Arg Gly Arg Glu
Cys Val500 505 510Asp Lys Cys Asn Leu Leu
Glu Gly Glu Pro Arg Glu Phe Val Glu Asn515 520
525Ser Glu Cys Ile Gln Cys His Pro Glu Cys Leu Pro Gln Ala Met
Asn530 535 540Ile Thr Cys Thr Gly Arg Gly
Pro Asp Asn Cys Ile Gln Cys Ala His545 550
555 560Tyr Ile Asp Gly Pro His Cys Val Lys Thr Cys Pro
Ala Gly Val Met565 570 575Gly Glu Asn Asn
Thr Leu Val Trp Lys Tyr Ala Asp Ala Gly His Val580 585
590Cys His Leu Cys His Pro Asn Cys Thr Tyr Gly Cys Thr Gly
Pro Gly595 600 605Leu Glu Gly Cys Pro Thr
Asn Gly Pro Lys Ile Pro Ser Gly Xaa610 615
6201852588DNAArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 185aagcttgaat tcccaccatg gagctggcgg
ccttgtgccg ctgggggctc ctcctcgccc 60tcttgccccc cggagccgcg agcacccaag
tgtgcaccgg cacagacatg aagctgcggc 120tccctgccag tcccgagacc cacctggaca
tgctccgcca cctctaccag ggctgccagg 180tggtgcaggg aaacctggaa ctcacctacc
tgcccaccaa tgccagcctg tccttcctgc 240aggatatcca ggaggtgcag ggctacgtgc
tcatcgctca caaccaagtg aggcaggtcc 300cactgcagag gctgcggatt gtgcgaggca
cccagctctt tgaggacaac tatgccctgg 360ccgtgctaga caatggagac ccgctgaaca
ataccacccc tgtcacaggg gcctccccag 420gaggcctgcg ggagctgcag cttcgaagcc
tcacagagat cttgaaagga ggggtcttga 480tccagcggaa cccccagctc tgctaccagg
acacgatttt gtggaaggac atcttccaca 540agaacaacca gctggctctc acactgatag
acaccaaccg ctctcgggcc tgccacccct 600gttctccgat gtgtaagggc tcccgctgct
ggggagagag ttctgaggat tgtcagagcc 660tgacgcgcac tgtctgtgcc ggtggctgtg
cccgctgcaa ggggccactg cccactgact 720gctgccatga gcagtgtgct gccggctgca
cgggtcccaa gcactctgac tgcctggcct 780gcctccactt caaccacagt ggcatctgtg
agctgcactg cccagccctg gtcacctaca 840acacagacac gtttgagtcc atgcccaatc
ccgagggccg gtatacattc ggcgccagct 900gtgtgactgc ctgtccctac aactaccttt
ctacggacgt gggatcctgc accctcgtct 960gccccctgca caaccaagag gtgacagcag
aggatggaac acagcggtgt gagaagtgca 1020gcaagccctg tgcccgagtg tgctatggtc
tgggcatgga gcacttgcga gaggtgaggg 1080cagttaccag tgccaatatc caggagtttg
ctggctgcaa gaagatcttt gggagcctgg 1140catttctgcc ggagagcttt gatggggacc
cagcctccaa cactgccccg ctccagccag 1200agcagctcca agtgtttgag actctggaag
agatcacagg ttacctatac atctcagcat 1260ggccggacag cctgcctgac ctcagcgtct
tccagaacct gcaagtaatc cggggacgaa 1320ttctgcacaa tggcgcctac tcgctgaccc
tgcaagggct gggcatcagc tggctggggc 1380tgcgctcact gagggaactg ggcagtggac
tggccctcat ccaccataac acccacctct 1440gcttcgtgca cacggtgccc tgggaccagc
tctttcggaa cccgcaccaa gctctgctcc 1500acactgccaa ccggccagag gacgagtgtg
tgggcgaggg cctggcctgc caccagctgt 1560gcgcccgagg gcactgctgg ggtccaggtc
ccacccagtg tgtcaactgc agccagttcc 1620ttcggggcca ggagtgcgtg gaggaatgcc
gagtactgca ggggctcccc agggagtatg 1680tgaatgccag gcactgtttg ccgtgccacc
ctgagtgtca gccccagaat ggctcagtga 1740cctgttttgg accggaggct gaccagtgtg
tggcctgtgc ccactataag gaccctccct 1800tctgcgtggc ccgctgcccc agcggtgtga
aacctgacct ctcctacatg cccatctgga 1860agtttccaga tgaggagggc gcatgccagc
cttgccccat caactgcacc cactcctgtg 1920tggacctgga tgacaagggc tgccccgccg
agcagagagc cagccctctg acggggtgcg 1980tatccggtga caccattgta atgactagtg
gcgggccccg cactgtggct gaactggagg 2040gcaaaccgtt caccgcactg attcgcggct
ctggctaccc atgcccctca ggtttcttcc 2100gcacctgtga acgtgacgta tatgatctgc
gtacacgtga gggtcattgc ttacgtttga 2160cccatgatca ccgtgttctg gtgatggatg
gtggcctgga atggcgtgcc gcgggtgaac 2220tggaacgcgg cgaccgcctg gtgatggatg
atgcagctgg cgagtttccg gcactggcaa 2280ccttccgtgg cctgcgtggc gctggccgcc
aggatgttta tgacgctact gtttacggtg 2340ctagcgcatt cactgctaat ggcttcattg
tacacgcatg tggcgagcag cccgggaccg 2400gtctgaactc aggcctcacg acaaatcctg
gtgtatccgc ttggcaggtc aacacagctt 2460atactgcggg acaattggtc acatataacg
gcaagacgta taaatgtttg cagccccaca 2520cctccttggc aggatgggaa ccatccaacg
ttcctgcctt gtggcagctt caatgactcg 2580agcggccg
2588186852PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 186Met Glu Leu Ala Ala Leu Cys Arg Trp Gly Leu Leu Leu Ala
Leu Leu1 5 10 15Pro Pro
Gly Ala Ala Ser Thr Gln Val Cys Thr Gly Thr Asp Met Lys20
25 30Leu Arg Leu Pro Ala Ser Pro Glu Thr His Leu Asp
Met Leu Arg His35 40 45Leu Tyr Gln Gly
Cys Gln Val Val Gln Gly Asn Leu Glu Leu Thr Tyr50 55
60Leu Pro Thr Asn Ala Ser Leu Ser Phe Leu Gln Asp Ile Gln
Glu Val65 70 75 80Gln
Gly Tyr Val Leu Ile Ala His Asn Gln Val Arg Gln Val Pro Leu85
90 95Gln Arg Leu Arg Ile Val Arg Gly Thr Gln Leu
Phe Glu Asp Asn Tyr100 105 110Ala Leu Ala
Val Leu Asp Asn Gly Asp Pro Leu Asn Asn Thr Thr Pro115
120 125Val Thr Gly Ala Ser Pro Gly Gly Leu Arg Glu Leu
Gln Leu Arg Ser130 135 140Leu Thr Glu Ile
Leu Lys Gly Gly Val Leu Ile Gln Arg Asn Pro Gln145 150
155 160Leu Cys Tyr Gln Asp Thr Ile Leu Trp
Lys Asp Ile Phe His Lys Asn165 170 175Asn
Gln Leu Ala Leu Thr Leu Ile Asp Thr Asn Arg Ser Arg Ala Cys180
185 190His Pro Cys Ser Pro Met Cys Lys Gly Ser Arg
Cys Trp Gly Glu Ser195 200 205Ser Glu Asp
Cys Gln Ser Leu Thr Arg Thr Val Cys Ala Gly Gly Cys210
215 220Ala Arg Cys Lys Gly Pro Leu Pro Thr Asp Cys Cys
His Glu Gln Cys225 230 235
240Ala Ala Gly Cys Thr Gly Pro Lys His Ser Asp Cys Leu Ala Cys Leu245
250 255His Phe Asn His Ser Gly Ile Cys Glu
Leu His Cys Pro Ala Leu Val260 265 270Thr
Tyr Asn Thr Asp Thr Phe Glu Ser Met Pro Asn Pro Glu Gly Arg275
280 285Tyr Thr Phe Gly Ala Ser Cys Val Thr Ala Cys
Pro Tyr Asn Tyr Leu290 295 300Ser Thr Asp
Val Gly Ser Cys Thr Leu Val Cys Pro Leu His Asn Gln305
310 315 320Glu Val Thr Ala Glu Asp Gly
Thr Gln Arg Cys Glu Lys Cys Ser Lys325 330
335Pro Cys Ala Arg Val Cys Tyr Gly Leu Gly Met Glu His Leu Arg Glu340
345 350Val Arg Ala Val Thr Ser Ala Asn Ile
Gln Glu Phe Ala Gly Cys Lys355 360 365Lys
Ile Phe Gly Ser Leu Ala Phe Leu Pro Glu Ser Phe Asp Gly Asp370
375 380Pro Ala Ser Asn Thr Ala Pro Leu Gln Pro Glu
Gln Leu Gln Val Phe385 390 395
400Glu Thr Leu Glu Glu Ile Thr Gly Tyr Leu Tyr Ile Ser Ala Trp
Pro405 410 415Asp Ser Leu Pro Asp Leu Ser
Val Phe Gln Asn Leu Gln Val Ile Arg420 425
430Gly Arg Ile Leu His Asn Gly Ala Tyr Ser Leu Thr Leu Gln Gly Leu435
440 445Gly Ile Ser Trp Leu Gly Leu Arg Ser
Leu Arg Glu Leu Gly Ser Gly450 455 460Leu
Ala Leu Ile His His Asn Thr His Leu Cys Phe Val His Thr Val465
470 475 480Pro Trp Asp Gln Leu Phe
Arg Asn Pro His Gln Ala Leu Leu His Thr485 490
495Ala Asn Arg Pro Glu Asp Glu Cys Val Gly Glu Gly Leu Ala Cys
His500 505 510Gln Leu Cys Ala Arg Gly His
Cys Trp Gly Pro Gly Pro Thr Gln Cys515 520
525Val Asn Cys Ser Gln Phe Leu Arg Gly Gln Glu Cys Val Glu Glu Cys530
535 540Arg Val Leu Gln Gly Leu Pro Arg Glu
Tyr Val Asn Ala Arg His Cys545 550 555
560Leu Pro Cys His Pro Glu Cys Gln Pro Gln Asn Gly Ser Val
Thr Cys565 570 575Phe Gly Pro Glu Ala Asp
Gln Cys Val Ala Cys Ala His Tyr Lys Asp580 585
590Pro Pro Phe Cys Val Ala Arg Cys Pro Ser Gly Val Lys Pro Asp
Leu595 600 605Ser Tyr Met Pro Ile Trp Lys
Phe Pro Asp Glu Glu Gly Ala Cys Gln610 615
620Pro Cys Pro Ile Asn Cys Thr His Ser Cys Val Asp Leu Asp Asp Lys625
630 635 640Gly Cys Pro Ala
Glu Gln Arg Ala Ser Pro Leu Thr Gly Cys Val Ser645 650
655Gly Asp Thr Ile Val Met Thr Ser Gly Gly Pro Arg Thr Val
Ala Glu660 665 670Leu Glu Gly Lys Pro Phe
Thr Ala Leu Ile Arg Gly Ser Gly Tyr Pro675 680
685Cys Pro Ser Gly Phe Phe Arg Thr Cys Glu Arg Asp Val Tyr Asp
Leu690 695 700Arg Thr Arg Glu Gly His Cys
Leu Arg Leu Thr His Asp His Arg Val705 710
715 720Leu Val Met Asp Gly Gly Leu Glu Trp Arg Ala Ala
Gly Glu Leu Glu725 730 735Arg Gly Asp Arg
Leu Val Met Asp Asp Ala Ala Gly Glu Phe Pro Ala740 745
750Leu Ala Thr Phe Arg Gly Leu Arg Gly Ala Gly Arg Gln Asp
Val Tyr755 760 765Asp Ala Thr Val Tyr Gly
Ala Ser Ala Phe Thr Ala Asn Gly Phe Ile770 775
780Val His Ala Cys Gly Glu Gln Pro Gly Thr Gly Leu Asn Ser Gly
Leu785 790 795 800Thr Thr
Asn Pro Gly Val Ser Ala Trp Gln Val Asn Thr Ala Tyr Thr805
810 815Ala Gly Gln Leu Val Thr Tyr Asn Gly Lys Thr Tyr
Lys Cys Leu Gln820 825 830Pro His Thr Ser
Leu Ala Gly Trp Glu Pro Ser Asn Val Pro Ala Leu835 840
845Trp Gln Leu Gln850187830PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 187Thr Gln Val
Cys Thr Gly Thr Asp Met Lys Leu Arg Leu Pro Ala Ser1 5
10 15Pro Glu Thr His Leu Asp Met Leu Arg
His Leu Tyr Gln Gly Cys Gln20 25 30Val
Val Gln Gly Asn Leu Glu Leu Thr Tyr Leu Pro Thr Asn Ala Ser35
40 45Leu Ser Phe Leu Gln Asp Ile Gln Glu Val Gln
Gly Tyr Val Leu Ile50 55 60Ala His Asn
Gln Val Arg Gln Val Pro Leu Gln Arg Leu Arg Ile Val65 70
75 80Arg Gly Thr Gln Leu Phe Glu Asp
Asn Tyr Ala Leu Ala Val Leu Asp85 90
95Asn Gly Asp Pro Leu Asn Asn Thr Thr Pro Val Thr Gly Ala Ser Pro100
105 110Gly Gly Leu Arg Glu Leu Gln Leu Arg Ser
Leu Thr Glu Ile Leu Lys115 120 125Gly Gly
Val Leu Ile Gln Arg Asn Pro Gln Leu Cys Tyr Gln Asp Thr130
135 140Ile Leu Trp Lys Asp Ile Phe His Lys Asn Asn Gln
Leu Ala Leu Thr145 150 155
160Leu Ile Asp Thr Asn Arg Ser Arg Ala Cys His Pro Cys Ser Pro Met165
170 175Cys Lys Gly Ser Arg Cys Trp Gly Glu
Ser Ser Glu Asp Cys Gln Ser180 185 190Leu
Thr Arg Thr Val Cys Ala Gly Gly Cys Ala Arg Cys Lys Gly Pro195
200 205Leu Pro Thr Asp Cys Cys His Glu Gln Cys Ala
Ala Gly Cys Thr Gly210 215 220Pro Lys His
Ser Asp Cys Leu Ala Cys Leu His Phe Asn His Ser Gly225
230 235 240Ile Cys Glu Leu His Cys Pro
Ala Leu Val Thr Tyr Asn Thr Asp Thr245 250
255Phe Glu Ser Met Pro Asn Pro Glu Gly Arg Tyr Thr Phe Gly Ala Ser260
265 270Cys Val Thr Ala Cys Pro Tyr Asn Tyr
Leu Ser Thr Asp Val Gly Ser275 280 285Cys
Thr Leu Val Cys Pro Leu His Asn Gln Glu Val Thr Ala Glu Asp290
295 300Gly Thr Gln Arg Cys Glu Lys Cys Ser Lys Pro
Cys Ala Arg Val Cys305 310 315
320Tyr Gly Leu Gly Met Glu His Leu Arg Glu Val Arg Ala Val Thr
Ser325 330 335Ala Asn Ile Gln Glu Phe Ala
Gly Cys Lys Lys Ile Phe Gly Ser Leu340 345
350Ala Phe Leu Pro Glu Ser Phe Asp Gly Asp Pro Ala Ser Asn Thr Ala355
360 365Pro Leu Gln Pro Glu Gln Leu Gln Val
Phe Glu Thr Leu Glu Glu Ile370 375 380Thr
Gly Tyr Leu Tyr Ile Ser Ala Trp Pro Asp Ser Leu Pro Asp Leu385
390 395 400Ser Val Phe Gln Asn Leu
Gln Val Ile Arg Gly Arg Ile Leu His Asn405 410
415Gly Ala Tyr Ser Leu Thr Leu Gln Gly Leu Gly Ile Ser Trp Leu
Gly420 425 430Leu Arg Ser Leu Arg Glu Leu
Gly Ser Gly Leu Ala Leu Ile His His435 440
445Asn Thr His Leu Cys Phe Val His Thr Val Pro Trp Asp Gln Leu Phe450
455 460Arg Asn Pro His Gln Ala Leu Leu His
Thr Ala Asn Arg Pro Glu Asp465 470 475
480Glu Cys Val Gly Glu Gly Leu Ala Cys His Gln Leu Cys Ala
Arg Gly485 490 495His Cys Trp Gly Pro Gly
Pro Thr Gln Cys Val Asn Cys Ser Gln Phe500 505
510Leu Arg Gly Gln Glu Cys Val Glu Glu Cys Arg Val Leu Gln Gly
Leu515 520 525Pro Arg Glu Tyr Val Asn Ala
Arg His Cys Leu Pro Cys His Pro Glu530 535
540Cys Gln Pro Gln Asn Gly Ser Val Thr Cys Phe Gly Pro Glu Ala Asp545
550 555 560Gln Cys Val Ala
Cys Ala His Tyr Lys Asp Pro Pro Phe Cys Val Ala565 570
575Arg Cys Pro Ser Gly Val Lys Pro Asp Leu Ser Tyr Met Pro
Ile Trp580 585 590Lys Phe Pro Asp Glu Glu
Gly Ala Cys Gln Pro Cys Pro Ile Asn Cys595 600
605Thr His Ser Cys Val Asp Leu Asp Asp Lys Gly Cys Pro Ala Glu
Gln610 615 620Arg Ala Ser Pro Leu Thr Gly
Cys Val Ser Gly Asp Thr Ile Val Met625 630
635 640Thr Ser Gly Gly Pro Arg Thr Val Ala Glu Leu Glu
Gly Lys Pro Phe645 650 655Thr Ala Leu Ile
Arg Gly Ser Gly Tyr Pro Cys Pro Ser Gly Phe Phe660 665
670Arg Thr Cys Glu Arg Asp Val Tyr Asp Leu Arg Thr Arg Glu
Gly His675 680 685Cys Leu Arg Leu Thr His
Asp His Arg Val Leu Val Met Asp Gly Gly690 695
700Leu Glu Trp Arg Ala Ala Gly Glu Leu Glu Arg Gly Asp Arg Leu
Val705 710 715 720Met Asp
Asp Ala Ala Gly Glu Phe Pro Ala Leu Ala Thr Phe Arg Gly725
730 735Leu Arg Gly Ala Gly Arg Gln Asp Val Tyr Asp Ala
Thr Val Tyr Gly740 745 750Ala Ser Ala Phe
Thr Ala Asn Gly Phe Ile Val His Ala Cys Gly Glu755 760
765Gln Pro Gly Thr Gly Leu Asn Ser Gly Leu Thr Thr Asn Pro
Gly Val770 775 780Ser Ala Trp Gln Val Asn
Thr Ala Tyr Thr Ala Gly Gln Leu Val Thr785 790
795 800Tyr Asn Gly Lys Thr Tyr Lys Cys Leu Gln Pro
His Thr Ser Leu Ala805 810 815Gly Trp Glu
Pro Ser Asn Val Pro Ala Leu Trp Gln Leu Gln820 825
830188631PRTArtificial Sequenceartificial construct relating to
Homo Sapiens immunoglobulin 188Thr Gln Val Cys Thr Gly Thr Asp Met
Lys Leu Arg Leu Pro Ala Ser1 5 10
15Pro Glu Thr His Leu Asp Met Leu Arg His Leu Tyr Gln Gly Cys
Gln20 25 30Val Val Gln Gly Asn Leu Glu
Leu Thr Tyr Leu Pro Thr Asn Ala Ser35 40
45Leu Ser Phe Leu Gln Asp Ile Gln Glu Val Gln Gly Tyr Val Leu Ile50
55 60Ala His Asn Gln Val Arg Gln Val Pro Leu
Gln Arg Leu Arg Ile Val65 70 75
80Arg Gly Thr Gln Leu Phe Glu Asp Asn Tyr Ala Leu Ala Val Leu
Asp85 90 95Asn Gly Asp Pro Leu Asn Asn
Thr Thr Pro Val Thr Gly Ala Ser Pro100 105
110Gly Gly Leu Arg Glu Leu Gln Leu Arg Ser Leu Thr Glu Ile Leu Lys115
120 125Gly Gly Val Leu Ile Gln Arg Asn Pro
Gln Leu Cys Tyr Gln Asp Thr130 135 140Ile
Leu Trp Lys Asp Ile Phe His Lys Asn Asn Gln Leu Ala Leu Thr145
150 155 160Leu Ile Asp Thr Asn Arg
Ser Arg Ala Cys His Pro Cys Ser Pro Met165 170
175Cys Lys Gly Ser Arg Cys Trp Gly Glu Ser Ser Glu Asp Cys Gln
Ser180 185 190Leu Thr Arg Thr Val Cys Ala
Gly Gly Cys Ala Arg Cys Lys Gly Pro195 200
205Leu Pro Thr Asp Cys Cys His Glu Gln Cys Ala Ala Gly Cys Thr Gly210
215 220Pro Lys His Ser Asp Cys Leu Ala Cys
Leu His Phe Asn His Ser Gly225 230 235
240Ile Cys Glu Leu His Cys Pro Ala Leu Val Thr Tyr Asn Thr
Asp Thr245 250 255Phe Glu Ser Met Pro Asn
Pro Glu Gly Arg Tyr Thr Phe Gly Ala Ser260 265
270Cys Val Thr Ala Cys Pro Tyr Asn Tyr Leu Ser Thr Asp Val Gly
Ser275 280 285Cys Thr Leu Val Cys Pro Leu
His Asn Gln Glu Val Thr Ala Glu Asp290 295
300Gly Thr Gln Arg Cys Glu Lys Cys Ser Lys Pro Cys Ala Arg Val Cys305
310 315 320Tyr Gly Leu Gly
Met Glu His Leu Arg Glu Val Arg Ala Val Thr Ser325 330
335Ala Asn Ile Gln Glu Phe Ala Gly Cys Lys Lys Ile Phe Gly
Ser Leu340 345 350Ala Phe Leu Pro Glu Ser
Phe Asp Gly Asp Pro Ala Ser Asn Thr Ala355 360
365Pro Leu Gln Pro Glu Gln Leu Gln Val Phe Glu Thr Leu Glu Glu
Ile370 375 380Thr Gly Tyr Leu Tyr Ile Ser
Ala Trp Pro Asp Ser Leu Pro Asp Leu385 390
395 400Ser Val Phe Gln Asn Leu Gln Val Ile Arg Gly Arg
Ile Leu His Asn405 410 415Gly Ala Tyr Ser
Leu Thr Leu Gln Gly Leu Gly Ile Ser Trp Leu Gly420 425
430Leu Arg Ser Leu Arg Glu Leu Gly Ser Gly Leu Ala Leu Ile
His His435 440 445Asn Thr His Leu Cys Phe
Val His Thr Val Pro Trp Asp Gln Leu Phe450 455
460Arg Asn Pro His Gln Ala Leu Leu His Thr Ala Asn Arg Pro Glu
Asp465 470 475 480Glu Cys
Val Gly Glu Gly Leu Ala Cys His Gln Leu Cys Ala Arg Gly485
490 495His Cys Trp Gly Pro Gly Pro Thr Gln Cys Val Asn
Cys Ser Gln Phe500 505 510Leu Arg Gly Gln
Glu Cys Val Glu Glu Cys Arg Val Leu Gln Gly Leu515 520
525Pro Arg Glu Tyr Val Asn Ala Arg His Cys Leu Pro Cys His
Pro Glu530 535 540Cys Gln Pro Gln Asn Gly
Ser Val Thr Cys Phe Gly Pro Glu Ala Asp545 550
555 560Gln Cys Val Ala Cys Ala His Tyr Lys Asp Pro
Pro Phe Cys Val Ala565 570 575Arg Cys Pro
Ser Gly Val Lys Pro Asp Leu Ser Tyr Met Pro Ile Trp580
585 590Lys Phe Pro Asp Glu Glu Gly Ala Cys Gln Pro Cys
Pro Ile Asn Cys595 600 605Thr His Ser Cys
Val Asp Leu Asp Asp Lys Gly Cys Pro Ala Glu Gln610 615
620Arg Ala Ser Pro Leu Thr Xaa625
630189632PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 189Thr Gln Val Cys Thr Gly Thr Asp Met Lys
Leu Arg Leu Pro Ala Ser1 5 10
15Pro Glu Thr His Leu Asp Met Leu Arg His Leu Tyr Gln Gly Cys Gln20
25 30Val Val Gln Gly Asn Leu Glu Leu Thr
Tyr Leu Pro Thr Asn Ala Ser35 40 45Leu
Ser Phe Leu Gln Asp Ile Gln Glu Val Gln Gly Tyr Val Leu Ile50
55 60Ala His Asn Gln Val Arg Gln Val Pro Leu Gln
Arg Leu Arg Ile Val65 70 75
80Arg Gly Thr Gln Leu Phe Glu Asp Asn Tyr Ala Leu Ala Val Leu Asp85
90 95Asn Gly Asp Pro Leu Asn Asn Thr Thr
Pro Val Thr Gly Ala Ser Pro100 105 110Gly
Gly Leu Arg Glu Leu Gln Leu Arg Ser Leu Thr Glu Ile Leu Lys115
120 125Gly Gly Val Leu Ile Gln Arg Asn Pro Gln Leu
Cys Tyr Gln Asp Thr130 135 140Ile Leu Trp
Lys Asp Ile Phe His Lys Asn Asn Gln Leu Ala Leu Thr145
150 155 160Leu Ile Asp Thr Asn Arg Ser
Arg Ala Cys His Pro Cys Ser Pro Met165 170
175Cys Lys Gly Ser Arg Cys Trp Gly Glu Ser Ser Glu Asp Cys Gln Ser180
185 190Leu Thr Arg Thr Val Cys Ala Gly Gly
Cys Ala Arg Cys Lys Gly Pro195 200 205Leu
Pro Thr Asp Cys Cys His Glu Gln Cys Ala Ala Gly Cys Thr Gly210
215 220Pro Lys His Ser Asp Cys Leu Ala Cys Leu His
Phe Asn His Ser Gly225 230 235
240Ile Cys Glu Leu His Cys Pro Ala Leu Val Thr Tyr Asn Thr Asp
Thr245 250 255Phe Glu Ser Met Pro Asn Pro
Glu Gly Arg Tyr Thr Phe Gly Ala Ser260 265
270Cys Val Thr Ala Cys Pro Tyr Asn Tyr Leu Ser Thr Asp Val Gly Ser275
280 285Cys Thr Leu Val Cys Pro Leu His Asn
Gln Glu Val Thr Ala Glu Asp290 295 300Gly
Thr Gln Arg Cys Glu Lys Cys Ser Lys Pro Cys Ala Arg Val Cys305
310 315 320Tyr Gly Leu Gly Met Glu
His Leu Arg Glu Val Arg Ala Val Thr Ser325 330
335Ala Asn Ile Gln Glu Phe Ala Gly Cys Lys Lys Ile Phe Gly Ser
Leu340 345 350Ala Phe Leu Pro Glu Ser Phe
Asp Gly Asp Pro Ala Ser Asn Thr Ala355 360
365Pro Leu Gln Pro Glu Gln Leu Gln Val Phe Glu Thr Leu Glu Glu Ile370
375 380Thr Gly Tyr Leu Tyr Ile Ser Ala Trp
Pro Asp Ser Leu Pro Asp Leu385 390 395
400Ser Val Phe Gln Asn Leu Gln Val Ile Arg Gly Arg Ile Leu
His Asn405 410 415Gly Ala Tyr Ser Leu Thr
Leu Gln Gly Leu Gly Ile Ser Trp Leu Gly420 425
430Leu Arg Ser Leu Arg Glu Leu Gly Ser Gly Leu Ala Leu Ile His
His435 440 445Asn Thr His Leu Cys Phe Val
His Thr Val Pro Trp Asp Gln Leu Phe450 455
460Arg Asn Pro His Gln Ala Leu Leu His Thr Ala Asn Arg Pro Glu Asp465
470 475 480Glu Cys Val Gly
Glu Gly Leu Ala Cys His Gln Leu Cys Ala Arg Gly485 490
495His Cys Trp Gly Pro Gly Pro Thr Gln Cys Val Asn Cys Ser
Gln Phe500 505 510Leu Arg Gly Gln Glu Cys
Val Glu Glu Cys Arg Val Leu Gln Gly Leu515 520
525Pro Arg Glu Tyr Val Asn Ala Arg His Cys Leu Pro Cys His Pro
Glu530 535 540Cys Gln Pro Gln Asn Gly Ser
Val Thr Cys Phe Gly Pro Glu Ala Asp545 550
555 560Gln Cys Val Ala Cys Ala His Tyr Lys Asp Pro Pro
Phe Cys Val Ala565 570 575Arg Cys Pro Ser
Gly Val Lys Pro Asp Leu Ser Tyr Met Pro Ile Trp580 585
590Lys Phe Pro Asp Glu Glu Gly Ala Cys Gln Pro Cys Pro Ile
Asn Cys595 600 605Thr His Ser Cys Val Asp
Leu Asp Asp Lys Gly Cys Pro Ala Glu Gln610 615
620Arg Ala Ser Pro Leu Thr Gly Xaa625
6301902561DNAArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 190aagcttgaat tcccaccatg agggcgaacg
acgctctgca ggtgctgggc ttgcttttca 60gcctggcccg gggctccgag gtgggcaact
ctcaggcagt gtgtcctggg actctgaatg 120gcctgagtgt gaccggcgat gctgagaacc
aataccagac actgtacaag ctctacgaga 180ggtgtgaggt ggtgatgggg aaccttgaga
ttgtgctcac gggacacaat gccgacctct 240ccttcctgca gtggattcga gaagtgacag
gctatgtcct cgtggccatg aatgaatttt 300ctactctacc attgcccaac ctccgcgtgg
tgcgagggac ccaggtctac gatgggaagt 360ttgccatctt cgtcatgttg aactataaca
ccaactccag ccacgctctg cgccagctcc 420gcttgactca gctcaccgag attctgtcag
ggggtgttta tattgagaag aacgataagc 480tctgtcacat ggacacaatt gactggaggg
acatcgtgag ggaccgagat gctgagatag 540tggtgaagga caatggcaga agctgtcccc
cctgtcatga ggtttgcaag gggcgatgct 600ggggtcctgg atcagaagac tgccagacat
tgaccaagac catctgtgct cctcagtgta 660atggtcactg ctttggtccc aaccccaacc
agtgctgcca tgatgagtgt gccgggggct 720gctcaggccc tcaggacaca gactgctttg
cctgccggca cttcaatgac agtggagcct 780gtgtacctcg ctgtccacag cctcttgtct
acaacaagct aactttccag ctggaaccca 840atccccacac caagtatcag tatggaggag
tttgtgtagc cagctgtccc cataactttg 900tggtggatca aacatcctgt gtcagggcct
gtcctcctga caagatggaa gtagataaaa 960atgggctcaa gatgtgtgag ccttgtgggg
gactatgtcc caaagcctgt gagggaacag 1020gctctgggag ccgcttccag actgtggatt
cgagcaacat tgatggattt gtgaactgca 1080ccaagatcct gggcaacctg gactttctga
tcaccggcct caatggagac ccctggcaca 1140agatccctgc cctggaccca gagaagctca
atgtcttccg gacagtacgg gagatcacag 1200gttacctgaa catccagtcc tggccgcccc
acatgcacaa cttcagtgtt ttttccaatt 1260tgacaaccat tggaggcaga agcctctaca
accggggctt ctcattgttg atcatgaaga 1320acttgaatgt cacatctctg ggcttccgat
ccctgaagga aattagtgct gggcgtatct 1380atataagtgc caataggcag ctctgctacc
accactcttt gaactggacc aaggtgcttc 1440gggggcctac ggaagagcga ctagacatca
agcataatcg gccgcgcaga gactgcgtgg 1500cagagggcaa agtgtgtgac ccactgtgct
cctctggggg atgctgggtc ccaggccctg 1560gtcagtgctt gtcctgtcga aattatagcc
gaggaggtgt ctgtgtgacc cactgcaact 1620ttctgaatgg ggagccgcga gaatttgccc
atgaggccga atgcttctcc tgccacccgg 1680aatgccaacc catggagggc actgccacat
gcaatggctc gggctctgat acttgtgctc 1740aatgtgccca ttttcgagat ggtccccact
gtgtgagcag ctgcccccat ggagtcctag 1800gtgccaaggg tccaatctac aagtacccag
atgttcagaa tgaatgtcgg ccctgccatg 1860agaactgcac ccaggggtgt aaaggaccag
agcttcaaga ctgtttagga caaacactgg 1920tgctgatcgg caaaacccat ctgacagggt
gcgtatccgg tgacaccatt gtaatgacta 1980gtggcgggcc ccgcactgtg gctgaactgg
agggcaaacc gttcaccgca ctgattcgcg 2040gctctggcta cccatgcccc tcaggtttct
tccgcacctg tgaacgtgac gtatatgatc 2100tgcgtacacg tgagggtcat tgcttacgtt
tgacccatga tcaccgtgtt ctggtgatgg 2160atggtggcct ggaatggcgt gccgcgggtg
aactggaacg cggcgaccgc ctggtgatgg 2220atgatgcagc tggcgagttt ccggcactgg
caaccttccg tggcctgcgt ggcgctggcc 2280gccaggatgt ttatgacgct actgtttacg
gtgctagcgc attcactgct aatggcttca 2340ttgtacacgc atgtggcgag cagcccggga
ccggtctgaa ctcaggcctc acgacaaatc 2400ctggtgtatc cgcttggcag gtcaacacag
cttatactgc gggacaattg gtcacatata 2460acggcaagac gtataaatgt ttgcagcccc
acacctcctt ggcaggatgg gaaccatcca 2520acgttcctgc cttgtggcag cttcaatgac
tcgagcggcc g 2561191843PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 191Met Arg Ala Asn Asp Ala Leu Gln Val Leu Gly Leu Leu Phe
Ser Leu1 5 10 15Ala Arg
Gly Ser Glu Val Gly Asn Ser Gln Ala Val Cys Pro Gly Thr20
25 30Leu Asn Gly Leu Ser Val Thr Gly Asp Ala Glu Asn
Gln Tyr Gln Thr35 40 45Leu Tyr Lys Leu
Tyr Glu Arg Cys Glu Val Val Met Gly Asn Leu Glu50 55
60Ile Val Leu Thr Gly His Asn Ala Asp Leu Ser Phe Leu Gln
Trp Ile65 70 75 80Arg
Glu Val Thr Gly Tyr Val Leu Val Ala Met Asn Glu Phe Ser Thr85
90 95Leu Pro Leu Pro Asn Leu Arg Val Val Arg Gly
Thr Gln Val Tyr Asp100 105 110Gly Lys Phe
Ala Ile Phe Val Met Leu Asn Tyr Asn Thr Asn Ser Ser115
120 125His Ala Leu Arg Gln Leu Arg Leu Thr Gln Leu Thr
Glu Ile Leu Ser130 135 140Gly Gly Val Tyr
Ile Glu Lys Asn Asp Lys Leu Cys His Met Asp Thr145 150
155 160Ile Asp Trp Arg Asp Ile Val Arg Asp
Arg Asp Ala Glu Ile Val Val165 170 175Lys
Asp Asn Gly Arg Ser Cys Pro Pro Cys His Glu Val Cys Lys Gly180
185 190Arg Cys Trp Gly Pro Gly Ser Glu Asp Cys Gln
Thr Leu Thr Lys Thr195 200 205Ile Cys Ala
Pro Gln Cys Asn Gly His Cys Phe Gly Pro Asn Pro Asn210
215 220Gln Cys Cys His Asp Glu Cys Ala Gly Gly Cys Ser
Gly Pro Gln Asp225 230 235
240Thr Asp Cys Phe Ala Cys Arg His Phe Asn Asp Ser Gly Ala Cys Val245
250 255Pro Arg Cys Pro Gln Pro Leu Val Tyr
Asn Lys Leu Thr Phe Gln Leu260 265 270Glu
Pro Asn Pro His Thr Lys Tyr Gln Tyr Gly Gly Val Cys Val Ala275
280 285Ser Cys Pro His Asn Phe Val Val Asp Gln Thr
Ser Cys Val Arg Ala290 295 300Cys Pro Pro
Asp Lys Met Glu Val Asp Lys Asn Gly Leu Lys Met Cys305
310 315 320Glu Pro Cys Gly Gly Leu Cys
Pro Lys Ala Cys Glu Gly Thr Gly Ser325 330
335Gly Ser Arg Phe Gln Thr Val Asp Ser Ser Asn Ile Asp Gly Phe Val340
345 350Asn Cys Thr Lys Ile Leu Gly Asn Leu
Asp Phe Leu Ile Thr Gly Leu355 360 365Asn
Gly Asp Pro Trp His Lys Ile Pro Ala Leu Asp Pro Glu Lys Leu370
375 380Asn Val Phe Arg Thr Val Arg Glu Ile Thr Gly
Tyr Leu Asn Ile Gln385 390 395
400Ser Trp Pro Pro His Met His Asn Phe Ser Val Phe Ser Asn Leu
Thr405 410 415Thr Ile Gly Gly Arg Ser Leu
Tyr Asn Arg Gly Phe Ser Leu Leu Ile420 425
430Met Lys Asn Leu Asn Val Thr Ser Leu Gly Phe Arg Ser Leu Lys Glu435
440 445Ile Ser Ala Gly Arg Ile Tyr Ile Ser
Ala Asn Arg Gln Leu Cys Tyr450 455 460His
His Ser Leu Asn Trp Thr Lys Val Leu Arg Gly Pro Thr Glu Glu465
470 475 480Arg Leu Asp Ile Lys His
Asn Arg Pro Arg Arg Asp Cys Val Ala Glu485 490
495Gly Lys Val Cys Asp Pro Leu Cys Ser Ser Gly Gly Cys Trp Gly
Pro500 505 510Gly Pro Gly Gln Cys Leu Ser
Cys Arg Asn Tyr Ser Arg Gly Gly Val515 520
525Cys Val Thr His Cys Asn Phe Leu Asn Gly Glu Pro Arg Glu Phe Ala530
535 540His Glu Ala Glu Cys Phe Ser Cys His
Pro Glu Cys Gln Pro Met Glu545 550 555
560Gly Thr Ala Thr Cys Asn Gly Ser Gly Ser Asp Thr Cys Ala
Gln Cys565 570 575Ala His Phe Arg Asp Gly
Pro His Cys Val Ser Ser Cys Pro His Gly580 585
590Val Leu Gly Ala Lys Gly Pro Ile Tyr Lys Tyr Pro Asp Val Gln
Asn595 600 605Glu Cys Arg Pro Cys His Glu
Asn Cys Thr Gln Gly Cys Lys Gly Pro610 615
620Glu Leu Gln Asp Cys Leu Gly Gln Thr Leu Val Leu Ile Gly Lys Thr625
630 635 640His Leu Thr Gly
Cys Val Ser Gly Asp Thr Ile Val Met Thr Ser Gly645 650
655Gly Pro Arg Thr Val Ala Glu Leu Glu Gly Lys Pro Phe Thr
Ala Leu660 665 670Ile Arg Gly Ser Gly Tyr
Pro Cys Pro Ser Gly Phe Phe Arg Thr Cys675 680
685Glu Arg Asp Val Tyr Asp Leu Arg Thr Arg Glu Gly His Cys Leu
Arg690 695 700Leu Thr His Asp His Arg Val
Leu Val Met Asp Gly Gly Leu Glu Trp705 710
715 720Arg Ala Ala Gly Glu Leu Glu Arg Gly Asp Arg Leu
Val Met Asp Asp725 730 735Ala Ala Gly Glu
Phe Pro Ala Leu Ala Thr Phe Arg Gly Leu Arg Gly740 745
750Ala Gly Arg Gln Asp Val Tyr Asp Ala Thr Val Tyr Gly Ala
Ser Ala755 760 765Phe Thr Ala Asn Gly Phe
Ile Val His Ala Cys Gly Glu Gln Pro Gly770 775
780Thr Gly Leu Asn Ser Gly Leu Thr Thr Asn Pro Gly Val Ser Ala
Trp785 790 795 800Gln Val
Asn Thr Ala Tyr Thr Ala Gly Gln Leu Val Thr Tyr Asn Gly805
810 815Lys Thr Tyr Lys Cys Leu Gln Pro His Thr Ser Leu
Ala Gly Trp Glu820 825 830Pro Ser Asn Val
Pro Ala Leu Trp Gln Leu Gln835 840192824PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 192Ser Glu Val Gly Asn Ser Gln Ala Val Cys Pro Gly Thr Leu
Asn Gly1 5 10 15Leu Ser
Val Thr Gly Asp Ala Glu Asn Gln Tyr Gln Thr Leu Tyr Lys20
25 30Leu Tyr Glu Arg Cys Glu Val Val Met Gly Asn Leu
Glu Ile Val Leu35 40 45Thr Gly His Asn
Ala Asp Leu Ser Phe Leu Gln Trp Ile Arg Glu Val50 55
60Thr Gly Tyr Val Leu Val Ala Met Asn Glu Phe Ser Thr Leu
Pro Leu65 70 75 80Pro
Asn Leu Arg Val Val Arg Gly Thr Gln Val Tyr Asp Gly Lys Phe85
90 95Ala Ile Phe Val Met Leu Asn Tyr Asn Thr Asn
Ser Ser His Ala Leu100 105 110Arg Gln Leu
Arg Leu Thr Gln Leu Thr Glu Ile Leu Ser Gly Gly Val115
120 125Tyr Ile Glu Lys Asn Asp Lys Leu Cys His Met Asp
Thr Ile Asp Trp130 135 140Arg Asp Ile Val
Arg Asp Arg Asp Ala Glu Ile Val Val Lys Asp Asn145 150
155 160Gly Arg Ser Cys Pro Pro Cys His Glu
Val Cys Lys Gly Arg Cys Trp165 170 175Gly
Pro Gly Ser Glu Asp Cys Gln Thr Leu Thr Lys Thr Ile Cys Ala180
185 190Pro Gln Cys Asn Gly His Cys Phe Gly Pro Asn
Pro Asn Gln Cys Cys195 200 205His Asp Glu
Cys Ala Gly Gly Cys Ser Gly Pro Gln Asp Thr Asp Cys210
215 220Phe Ala Cys Arg His Phe Asn Asp Ser Gly Ala Cys
Val Pro Arg Cys225 230 235
240Pro Gln Pro Leu Val Tyr Asn Lys Leu Thr Phe Gln Leu Glu Pro Asn245
250 255Pro His Thr Lys Tyr Gln Tyr Gly Gly
Val Cys Val Ala Ser Cys Pro260 265 270His
Asn Phe Val Val Asp Gln Thr Ser Cys Val Arg Ala Cys Pro Pro275
280 285Asp Lys Met Glu Val Asp Lys Asn Gly Leu Lys
Met Cys Glu Pro Cys290 295 300Gly Gly Leu
Cys Pro Lys Ala Cys Glu Gly Thr Gly Ser Gly Ser Arg305
310 315 320Phe Gln Thr Val Asp Ser Ser
Asn Ile Asp Gly Phe Val Asn Cys Thr325 330
335Lys Ile Leu Gly Asn Leu Asp Phe Leu Ile Thr Gly Leu Asn Gly Asp340
345 350Pro Trp His Lys Ile Pro Ala Leu Asp
Pro Glu Lys Leu Asn Val Phe355 360 365Arg
Thr Val Arg Glu Ile Thr Gly Tyr Leu Asn Ile Gln Ser Trp Pro370
375 380Pro His Met His Asn Phe Ser Val Phe Ser Asn
Leu Thr Thr Ile Gly385 390 395
400Gly Arg Ser Leu Tyr Asn Arg Gly Phe Ser Leu Leu Ile Met Lys
Asn405 410 415Leu Asn Val Thr Ser Leu Gly
Phe Arg Ser Leu Lys Glu Ile Ser Ala420 425
430Gly Arg Ile Tyr Ile Ser Ala Asn Arg Gln Leu Cys Tyr His His Ser435
440 445Leu Asn Trp Thr Lys Val Leu Arg Gly
Pro Thr Glu Glu Arg Leu Asp450 455 460Ile
Lys His Asn Arg Pro Arg Arg Asp Cys Val Ala Glu Gly Lys Val465
470 475 480Cys Asp Pro Leu Cys Ser
Ser Gly Gly Cys Trp Gly Pro Gly Pro Gly485 490
495Gln Cys Leu Ser Cys Arg Asn Tyr Ser Arg Gly Gly Val Cys Val
Thr500 505 510His Cys Asn Phe Leu Asn Gly
Glu Pro Arg Glu Phe Ala His Glu Ala515 520
525Glu Cys Phe Ser Cys His Pro Glu Cys Gln Pro Met Glu Gly Thr Ala530
535 540Thr Cys Asn Gly Ser Gly Ser Asp Thr
Cys Ala Gln Cys Ala His Phe545 550 555
560Arg Asp Gly Pro His Cys Val Ser Ser Cys Pro His Gly Val
Leu Gly565 570 575Ala Lys Gly Pro Ile Tyr
Lys Tyr Pro Asp Val Gln Asn Glu Cys Arg580 585
590Pro Cys His Glu Asn Cys Thr Gln Gly Cys Lys Gly Pro Glu Leu
Gln595 600 605Asp Cys Leu Gly Gln Thr Leu
Val Leu Ile Gly Lys Thr His Leu Thr610 615
620Gly Cys Val Ser Gly Asp Thr Ile Val Met Thr Ser Gly Gly Pro Arg625
630 635 640Thr Val Ala Glu
Leu Glu Gly Lys Pro Phe Thr Ala Leu Ile Arg Gly645 650
655Ser Gly Tyr Pro Cys Pro Ser Gly Phe Phe Arg Thr Cys Glu
Arg Asp660 665 670Val Tyr Asp Leu Arg Thr
Arg Glu Gly His Cys Leu Arg Leu Thr His675 680
685Asp His Arg Val Leu Val Met Asp Gly Gly Leu Glu Trp Arg Ala
Ala690 695 700Gly Glu Leu Glu Arg Gly Asp
Arg Leu Val Met Asp Asp Ala Ala Gly705 710
715 720Glu Phe Pro Ala Leu Ala Thr Phe Arg Gly Leu Arg
Gly Ala Gly Arg725 730 735Gln Asp Val Tyr
Asp Ala Thr Val Tyr Gly Ala Ser Ala Phe Thr Ala740 745
750Asn Gly Phe Ile Val His Ala Cys Gly Glu Gln Pro Gly Thr
Gly Leu755 760 765Asn Ser Gly Leu Thr Thr
Asn Pro Gly Val Ser Ala Trp Gln Val Asn770 775
780Thr Ala Tyr Thr Ala Gly Gln Leu Val Thr Tyr Asn Gly Lys Thr
Tyr785 790 795 800Lys Cys
Leu Gln Pro His Thr Ser Leu Ala Gly Trp Glu Pro Ser Asn805
810 815Val Pro Ala Leu Trp Gln Leu
Gln820193625PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 193Ser Glu Val Gly Asn Ser Gln Ala Val Cys
Pro Gly Thr Leu Asn Gly1 5 10
15Leu Ser Val Thr Gly Asp Ala Glu Asn Gln Tyr Gln Thr Leu Tyr Lys20
25 30Leu Tyr Glu Arg Cys Glu Val Val Met
Gly Asn Leu Glu Ile Val Leu35 40 45Thr
Gly His Asn Ala Asp Leu Ser Phe Leu Gln Trp Ile Arg Glu Val50
55 60Thr Gly Tyr Val Leu Val Ala Met Asn Glu Phe
Ser Thr Leu Pro Leu65 70 75
80Pro Asn Leu Arg Val Val Arg Gly Thr Gln Val Tyr Asp Gly Lys Phe85
90 95Ala Ile Phe Val Met Leu Asn Tyr Asn
Thr Asn Ser Ser His Ala Leu100 105 110Arg
Gln Leu Arg Leu Thr Gln Leu Thr Glu Ile Leu Ser Gly Gly Val115
120 125Tyr Ile Glu Lys Asn Asp Lys Leu Cys His Met
Asp Thr Ile Asp Trp130 135 140Arg Asp Ile
Val Arg Asp Arg Asp Ala Glu Ile Val Val Lys Asp Asn145
150 155 160Gly Arg Ser Cys Pro Pro Cys
His Glu Val Cys Lys Gly Arg Cys Trp165 170
175Gly Pro Gly Ser Glu Asp Cys Gln Thr Leu Thr Lys Thr Ile Cys Ala180
185 190Pro Gln Cys Asn Gly His Cys Phe Gly
Pro Asn Pro Asn Gln Cys Cys195 200 205His
Asp Glu Cys Ala Gly Gly Cys Ser Gly Pro Gln Asp Thr Asp Cys210
215 220Phe Ala Cys Arg His Phe Asn Asp Ser Gly Ala
Cys Val Pro Arg Cys225 230 235
240Pro Gln Pro Leu Val Tyr Asn Lys Leu Thr Phe Gln Leu Glu Pro
Asn245 250 255Pro His Thr Lys Tyr Gln Tyr
Gly Gly Val Cys Val Ala Ser Cys Pro260 265
270His Asn Phe Val Val Asp Gln Thr Ser Cys Val Arg Ala Cys Pro Pro275
280 285Asp Lys Met Glu Val Asp Lys Asn Gly
Leu Lys Met Cys Glu Pro Cys290 295 300Gly
Gly Leu Cys Pro Lys Ala Cys Glu Gly Thr Gly Ser Gly Ser Arg305
310 315 320Phe Gln Thr Val Asp Ser
Ser Asn Ile Asp Gly Phe Val Asn Cys Thr325 330
335Lys Ile Leu Gly Asn Leu Asp Phe Leu Ile Thr Gly Leu Asn Gly
Asp340 345 350Pro Trp His Lys Ile Pro Ala
Leu Asp Pro Glu Lys Leu Asn Val Phe355 360
365Arg Thr Val Arg Glu Ile Thr Gly Tyr Leu Asn Ile Gln Ser Trp Pro370
375 380Pro His Met His Asn Phe Ser Val Phe
Ser Asn Leu Thr Thr Ile Gly385 390 395
400Gly Arg Ser Leu Tyr Asn Arg Gly Phe Ser Leu Leu Ile Met
Lys Asn405 410 415Leu Asn Val Thr Ser Leu
Gly Phe Arg Ser Leu Lys Glu Ile Ser Ala420 425
430Gly Arg Ile Tyr Ile Ser Ala Asn Arg Gln Leu Cys Tyr His His
Ser435 440 445Leu Asn Trp Thr Lys Val Leu
Arg Gly Pro Thr Glu Glu Arg Leu Asp450 455
460Ile Lys His Asn Arg Pro Arg Arg Asp Cys Val Ala Glu Gly Lys Val465
470 475 480Cys Asp Pro Leu
Cys Ser Ser Gly Gly Cys Trp Gly Pro Gly Pro Gly485 490
495Gln Cys Leu Ser Cys Arg Asn Tyr Ser Arg Gly Gly Val Cys
Val Thr500 505 510His Cys Asn Phe Leu Asn
Gly Glu Pro Arg Glu Phe Ala His Glu Ala515 520
525Glu Cys Phe Ser Cys His Pro Glu Cys Gln Pro Met Glu Gly Thr
Ala530 535 540Thr Cys Asn Gly Ser Gly Ser
Asp Thr Cys Ala Gln Cys Ala His Phe545 550
555 560Arg Asp Gly Pro His Cys Val Ser Ser Cys Pro His
Gly Val Leu Gly565 570 575Ala Lys Gly Pro
Ile Tyr Lys Tyr Pro Asp Val Gln Asn Glu Cys Arg580 585
590Pro Cys His Glu Asn Cys Thr Gln Gly Cys Lys Gly Pro Glu
Leu Gln595 600 605Asp Cys Leu Gly Gln Thr
Leu Val Leu Ile Gly Lys Thr His Leu Thr610 615
620Xaa625194626PRTArtificial Sequenceartificial construct relating
to Homo Sapiens immunoglobulin 194Ser Glu Val Gly Asn Ser Gln Ala
Val Cys Pro Gly Thr Leu Asn Gly1 5 10
15Leu Ser Val Thr Gly Asp Ala Glu Asn Gln Tyr Gln Thr Leu
Tyr Lys20 25 30Leu Tyr Glu Arg Cys Glu
Val Val Met Gly Asn Leu Glu Ile Val Leu35 40
45Thr Gly His Asn Ala Asp Leu Ser Phe Leu Gln Trp Ile Arg Glu Val50
55 60Thr Gly Tyr Val Leu Val Ala Met Asn
Glu Phe Ser Thr Leu Pro Leu65 70 75
80Pro Asn Leu Arg Val Val Arg Gly Thr Gln Val Tyr Asp Gly
Lys Phe85 90 95Ala Ile Phe Val Met Leu
Asn Tyr Asn Thr Asn Ser Ser His Ala Leu100 105
110Arg Gln Leu Arg Leu Thr Gln Leu Thr Glu Ile Leu Ser Gly Gly
Val115 120 125Tyr Ile Glu Lys Asn Asp Lys
Leu Cys His Met Asp Thr Ile Asp Trp130 135
140Arg Asp Ile Val Arg Asp Arg Asp Ala Glu Ile Val Val Lys Asp Asn145
150 155 160Gly Arg Ser Cys
Pro Pro Cys His Glu Val Cys Lys Gly Arg Cys Trp165 170
175Gly Pro Gly Ser Glu Asp Cys Gln Thr Leu Thr Lys Thr Ile
Cys Ala180 185 190Pro Gln Cys Asn Gly His
Cys Phe Gly Pro Asn Pro Asn Gln Cys Cys195 200
205His Asp Glu Cys Ala Gly Gly Cys Ser Gly Pro Gln Asp Thr Asp
Cys210 215 220Phe Ala Cys Arg His Phe Asn
Asp Ser Gly Ala Cys Val Pro Arg Cys225 230
235 240Pro Gln Pro Leu Val Tyr Asn Lys Leu Thr Phe Gln
Leu Glu Pro Asn245 250 255Pro His Thr Lys
Tyr Gln Tyr Gly Gly Val Cys Val Ala Ser Cys Pro260 265
270His Asn Phe Val Val Asp Gln Thr Ser Cys Val Arg Ala Cys
Pro Pro275 280 285Asp Lys Met Glu Val Asp
Lys Asn Gly Leu Lys Met Cys Glu Pro Cys290 295
300Gly Gly Leu Cys Pro Lys Ala Cys Glu Gly Thr Gly Ser Gly Ser
Arg305 310 315 320Phe Gln
Thr Val Asp Ser Ser Asn Ile Asp Gly Phe Val Asn Cys Thr325
330 335Lys Ile Leu Gly Asn Leu Asp Phe Leu Ile Thr Gly
Leu Asn Gly Asp340 345 350Pro Trp His Lys
Ile Pro Ala Leu Asp Pro Glu Lys Leu Asn Val Phe355 360
365Arg Thr Val Arg Glu Ile Thr Gly Tyr Leu Asn Ile Gln Ser
Trp Pro370 375 380Pro His Met His Asn Phe
Ser Val Phe Ser Asn Leu Thr Thr Ile Gly385 390
395 400Gly Arg Ser Leu Tyr Asn Arg Gly Phe Ser Leu
Leu Ile Met Lys Asn405 410 415Leu Asn Val
Thr Ser Leu Gly Phe Arg Ser Leu Lys Glu Ile Ser Ala420
425 430Gly Arg Ile Tyr Ile Ser Ala Asn Arg Gln Leu Cys
Tyr His His Ser435 440 445Leu Asn Trp Thr
Lys Val Leu Arg Gly Pro Thr Glu Glu Arg Leu Asp450 455
460Ile Lys His Asn Arg Pro Arg Arg Asp Cys Val Ala Glu Gly
Lys Val465 470 475 480Cys
Asp Pro Leu Cys Ser Ser Gly Gly Cys Trp Gly Pro Gly Pro Gly485
490 495Gln Cys Leu Ser Cys Arg Asn Tyr Ser Arg Gly
Gly Val Cys Val Thr500 505 510His Cys Asn
Phe Leu Asn Gly Glu Pro Arg Glu Phe Ala His Glu Ala515
520 525Glu Cys Phe Ser Cys His Pro Glu Cys Gln Pro Met
Glu Gly Thr Ala530 535 540Thr Cys Asn Gly
Ser Gly Ser Asp Thr Cys Ala Gln Cys Ala His Phe545 550
555 560Arg Asp Gly Pro His Cys Val Ser Ser
Cys Pro His Gly Val Leu Gly565 570 575Ala
Lys Gly Pro Ile Tyr Lys Tyr Pro Asp Val Gln Asn Glu Cys Arg580
585 590Pro Cys His Glu Asn Cys Thr Gln Gly Cys Lys
Gly Pro Glu Leu Gln595 600 605Asp Cys Leu
Gly Gln Thr Leu Val Leu Ile Gly Lys Thr His Leu Thr610
615 620Gly Xaa6251952585DNAArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 195aagcttgaat
tcccaccatg aagccggcga caggactttg ggtctgggtg agccttctcg 60tggcggcggg
gaccgtccag cccagcgatt ctcagtcagt gtgtgcagga acggagaata 120aactgagctc
tctctctgac ctggaacagc agtaccgagc cttgcgcaag tactatgaaa 180actgtgaggt
tgtcatgggc aacctggaga taaccagcat tgagcacaac cgggacctct 240ccttcctgcg
gtctgttcga gaagtcacag gctacgtgtt agtggctctt aatcagtttc 300gttacctgcc
tctggagaat ttacgcatta ttcgtgggac aaaactttat gaggatcgat 360atgccttggc
aatattttta aactacagaa aagatggaaa ctttggactt caagaacttg 420gattaaagaa
cttgacagaa atcctaaatg gtggagtcta tgtagaccag aacaaattcc 480tttgttatgc
agacaccatt cattggcaag atattgttcg gaacccatgg ccttccaact 540tgactcttgt
gtcaacaaat ggtagttcag gatgtggacg ttgccataag tcctgtactg 600gccgttgctg
gggacccaca gaaaatcatt gccagacttt gacaaggacg gtgtgtgcag 660aacaatgtga
cggcagatgc tacggacctt acgtcagtga ctgctgccat cgagaatgtg 720ctggaggctg
ctcaggacct aaggacacag actgctttgc ctgcatgaat ttcaatgaca 780gtggagcatg
tgttactcag tgtccccaaa cctttgtcta caatccaacc acctttcaac 840tggagcacaa
tttcaatgca aagtacacat atggagcatt ctgtgtcaag aaatgtccac 900ataactttgt
ggtagattcc agttcttgtg tgcgtgcctg ccctagttcc aagatggaag 960tagaagaaaa
tgggattaaa atgtgtaaac cttgcactga catttgccca aaggcttgtg 1020atggcattgg
cacaggatca ttgatgtcag ctcagactgt ggattccagt aacattgaca 1080aattcataaa
ctgtaccaag atcaatggga atttgatctt tctagtcact ggtattcatg 1140gggaccctta
caatgcaatt gaagccatag acccagagaa actgaacgtc tttcggacag 1200tcagagagat
aacaggtttc ctgaacatac agtcatggcc accaaacatg actgacttca 1260gtgttttttc
taacctggtg accattggtg gaagagtact ctatagtggc ctgtccttgc 1320ttatcctcaa
gcaacagggc atcacctctc tacagttcca gtccctgaag gaaatcagcg 1380caggaaacat
ctatattact gacaacagca acctgtgtta ttatcatacc attaactgga 1440caacactctt
cagcacaatc aaccagagaa tagtaatccg ggacaacaga aaagctgaaa 1500attgtactgc
tgaaggaatg gtgtgcaacc atctgtgttc cagtgatggc tgttggggac 1560ctgggccaga
ccaatgtctg tcgtgtcgcc gcttcagtag aggaaggatc tgcatagagt 1620cttgtaacct
ctatgatggt gaatttcggg agtttgagaa tggctccatc tgtgtggagt 1680gtgaccccca
gtgtgagaag atggaagatg gcctcctcac atgccatgga ccgggtcctg 1740acaactgtac
aaagtgctct cattttaaag atggcccaaa ctgtgtggaa aaatgtccag 1800atggcttaca
gggggcaaac agtttcattt tcaagtatgc tgatccagat cgggagtgcc 1860acccatgcca
tccaaactgc acccaagggt gtaacggtcc cactagtcat gactgcattt 1920actacccatg
gacgggccat tccactttac cacaacatgc tagaactccc gggtgcgtat 1980ccggtgacac
cattgtaatg actagtggcg ggccccgcac tgtggctgaa ctggagggca 2040aaccgttcac
cgcactgatt cgcggctctg gctacccatg cccctcaggt ttcttccgca 2100cctgtgaacg
tgacgtatat gatctgcgta cacgtgaggg tcattgctta cgtttgaccc 2160atgatcaccg
tgttctggtg atggatggtg gcctggaatg gcgtgccgcg ggtgaactgg 2220aacgcggcga
ccgcctggtg atggatgatg cagctggcga gtttccggca ctggcaacct 2280tccgtggcct
gcgtggcgct ggccgccagg atgtttatga cgctactgtt tacggtgcta 2340gcgcattcac
tgctaatggc ttcattgtac acgcatgtgg cgagcagccc gggaccggtc 2400tgaactcagg
cctcacgaca aatcctggtg tatccgcttg gcaggtcaac acagcttata 2460ctgcgggaca
attggtcaca tataacggca agacgtataa atgtttgcag ccccacacct 2520ccttggcagg
atgggaacca tccaacgttc ctgccttgtg gcagcttcaa tgactcgagc 2580ggccg
2585196851PRTArtificial Sequenceartificial construct relating to Homo
Sapiens immunoglobulin 196Met Lys Pro Ala Thr Gly Leu Trp Val Trp
Val Ser Leu Leu Val Ala1 5 10
15Ala Gly Thr Val Gln Pro Ser Asp Ser Gln Ser Val Cys Ala Gly Thr20
25 30Glu Asn Lys Leu Ser Ser Leu Ser Asp
Leu Glu Gln Gln Tyr Arg Ala35 40 45Leu
Arg Lys Tyr Tyr Glu Asn Cys Glu Val Val Met Gly Asn Leu Glu50
55 60Ile Thr Ser Ile Glu His Asn Arg Asp Leu Ser
Phe Leu Arg Ser Val65 70 75
80Arg Glu Val Thr Gly Tyr Val Leu Val Ala Leu Asn Gln Phe Arg Tyr85
90 95Leu Pro Leu Glu Asn Leu Arg Ile Ile
Arg Gly Thr Lys Leu Tyr Glu100 105 110Asp
Arg Tyr Ala Leu Ala Ile Phe Leu Asn Tyr Arg Lys Asp Gly Asn115
120 125Phe Gly Leu Gln Glu Leu Gly Leu Lys Asn Leu
Thr Glu Ile Leu Asn130 135 140Gly Gly Val
Tyr Val Asp Gln Asn Lys Phe Leu Cys Tyr Ala Asp Thr145
150 155 160Ile His Trp Gln Asp Ile Val
Arg Asn Pro Trp Pro Ser Asn Leu Thr165 170
175Leu Val Ser Thr Asn Gly Ser Ser Gly Cys Gly Arg Cys His Lys Ser180
185 190Cys Thr Gly Arg Cys Trp Gly Pro Thr
Glu Asn His Cys Gln Thr Leu195 200 205Thr
Arg Thr Val Cys Ala Glu Gln Cys Asp Gly Arg Cys Tyr Gly Pro210
215 220Tyr Val Ser Asp Cys Cys His Arg Glu Cys Ala
Gly Gly Cys Ser Gly225 230 235
240Pro Lys Asp Thr Asp Cys Phe Ala Cys Met Asn Phe Asn Asp Ser
Gly245 250 255Ala Cys Val Thr Gln Cys Pro
Gln Thr Phe Val Tyr Asn Pro Thr Thr260 265
270Phe Gln Leu Glu His Asn Phe Asn Ala Lys Tyr Thr Tyr Gly Ala Phe275
280 285Cys Val Lys Lys Cys Pro His Asn Phe
Val Val Asp Ser Ser Ser Cys290 295 300Val
Arg Ala Cys Pro Ser Ser Lys Met Glu Val Glu Glu Asn Gly Ile305
310 315 320Lys Met Cys Lys Pro Cys
Thr Asp Ile Cys Pro Lys Ala Cys Asp Gly325 330
335Ile Gly Thr Gly Ser Leu Met Ser Ala Gln Thr Val Asp Ser Ser
Asn340 345 350Ile Asp Lys Phe Ile Asn Cys
Thr Lys Ile Asn Gly Asn Leu Ile Phe355 360
365Leu Val Thr Gly Ile His Gly Asp Pro Tyr Asn Ala Ile Glu Ala Ile370
375 380Asp Pro Glu Lys Leu Asn Val Phe Arg
Thr Val Arg Glu Ile Thr Gly385 390 395
400Phe Leu Asn Ile Gln Ser Trp Pro Pro Asn Met Thr Asp Phe
Ser Val405 410 415Phe Ser Asn Leu Val Thr
Ile Gly Gly Arg Val Leu Tyr Ser Gly Leu420 425
430Ser Leu Leu Ile Leu Lys Gln Gln Gly Ile Thr Ser Leu Gln Phe
Gln435 440 445Ser Leu Lys Glu Ile Ser Ala
Gly Asn Ile Tyr Ile Thr Asp Asn Ser450 455
460Asn Leu Cys Tyr Tyr His Thr Ile Asn Trp Thr Thr Leu Phe Ser Thr465
470 475 480Ile Asn Gln Arg
Ile Val Ile Arg Asp Asn Arg Lys Ala Glu Asn Cys485 490
495Thr Ala Glu Gly Met Val Cys Asn His Leu Cys Ser Ser Asp
Gly Cys500 505 510Trp Gly Pro Gly Pro Asp
Gln Cys Leu Ser Cys Arg Arg Phe Ser Arg515 520
525Gly Arg Ile Cys Ile Glu Ser Cys Asn Leu Tyr Asp Gly Glu Phe
Arg530 535 540Glu Phe Glu Asn Gly Ser Ile
Cys Val Glu Cys Asp Pro Gln Cys Glu545 550
555 560Lys Met Glu Asp Gly Leu Leu Thr Cys His Gly Pro
Gly Pro Asp Asn565 570 575Cys Thr Lys Cys
Ser His Phe Lys Asp Gly Pro Asn Cys Val Glu Lys580 585
590Cys Pro Asp Gly Leu Gln Gly Ala Asn Ser Phe Ile Phe Lys
Tyr Ala595 600 605Asp Pro Asp Arg Glu Cys
His Pro Cys His Pro Asn Cys Thr Gln Gly610 615
620Cys Asn Gly Pro Thr Ser His Asp Cys Ile Tyr Tyr Pro Trp Thr
Gly625 630 635 640His Ser
Thr Leu Pro Gln His Ala Arg Thr Pro Gly Cys Val Ser Gly645
650 655Asp Thr Ile Val Met Thr Ser Gly Gly Pro Arg Thr
Val Ala Glu Leu660 665 670Glu Gly Lys Pro
Phe Thr Ala Leu Ile Arg Gly Ser Gly Tyr Pro Cys675 680
685Pro Ser Gly Phe Phe Arg Thr Cys Glu Arg Asp Val Tyr Asp
Leu Arg690 695 700Thr Arg Glu Gly His Cys
Leu Arg Leu Thr His Asp His Arg Val Leu705 710
715 720Val Met Asp Gly Gly Leu Glu Trp Arg Ala Ala
Gly Glu Leu Glu Arg725 730 735Gly Asp Arg
Leu Val Met Asp Asp Ala Ala Gly Glu Phe Pro Ala Leu740
745 750Ala Thr Phe Arg Gly Leu Arg Gly Ala Gly Arg Gln
Asp Val Tyr Asp755 760 765Ala Thr Val Tyr
Gly Ala Ser Ala Phe Thr Ala Asn Gly Phe Ile Val770 775
780His Ala Cys Gly Glu Gln Pro Gly Thr Gly Leu Asn Ser Gly
Leu Thr785 790 795 800Thr
Asn Pro Gly Val Ser Ala Trp Gln Val Asn Thr Ala Tyr Thr Ala805
810 815Gly Gln Leu Val Thr Tyr Asn Gly Lys Thr Tyr
Lys Cys Leu Gln Pro820 825 830His Thr Ser
Leu Ala Gly Trp Glu Pro Ser Asn Val Pro Ala Leu Trp835
840 845Gln Leu Gln850197826PRTArtificial
Sequenceartificial construct relating to Homo Sapiens
immunoglobulin 197Gln Ser Val Cys Ala Gly Thr Glu Asn Lys Leu Ser Ser Leu
Ser Asp1 5 10 15Leu Glu
Gln Gln Tyr Arg Ala Leu Arg Lys Tyr Tyr Glu Asn Cys Glu20
25 30Val Val Met Gly Asn Leu Glu Ile Thr Ser Ile Glu
His Asn Arg Asp35 40 45Leu Ser Phe Leu
Arg Ser Val Arg Glu Val Thr Gly Tyr Val Leu Val50 55
60Ala Leu Asn Gln Phe Arg Tyr Leu Pro Leu Glu Asn Leu Arg
Ile Ile65 70 75 80Arg
Gly Thr Lys Leu Tyr Glu Asp Arg Tyr Ala Leu Ala Ile Phe Leu85
90 95Asn Tyr Arg Lys Asp Gly Asn Phe Gly Leu Gln
Glu Leu Gly Leu Lys100 105 110Asn Leu Thr
Glu Ile Leu Asn Gly Gly Val Tyr Val Asp Gln Asn Lys115
120 125Phe Leu Cys Tyr Ala Asp Thr Ile His Trp Gln Asp
Ile Val Arg Asn130 135 140Pro Trp Pro Ser
Asn Leu Thr Leu Val Ser Thr Asn Gly Ser Ser Gly145 150
155 160Cys Gly Arg Cys His Lys Ser Cys Thr
Gly Arg Cys Trp Gly Pro Thr165 170 175Glu
Asn His Cys Gln Thr Leu Thr Arg Thr Val Cys Ala Glu Gln Cys180
185 190Asp Gly Arg Cys Tyr Gly Pro Tyr Val Ser Asp
Cys Cys His Arg Glu195 200 205Cys Ala Gly
Gly Cys Ser Gly Pro Lys Asp Thr Asp Cys Phe Ala Cys210
215 220Met Asn Phe Asn Asp Ser Gly Ala Cys Val Thr Gln
Cys Pro Gln Thr225 230 235
240Phe Val Tyr Asn Pro Thr Thr Phe Gln Leu Glu His Asn Phe Asn Ala245
250 255Lys Tyr Thr Tyr Gly Ala Phe Cys Val
Lys Lys Cys Pro His Asn Phe260 265 270Val
Val Asp Ser Ser Ser Cys Val Arg Ala Cys Pro Ser Ser Lys Met275
280 285Glu Val Glu Glu Asn Gly Ile Lys Met Cys Lys
Pro Cys Thr Asp Ile290 295 300Cys Pro Lys
Ala Cys Asp Gly Ile Gly Thr Gly Ser Leu Met Ser Ala305
310 315 320Gln Thr Val Asp Ser Ser Asn
Ile Asp Lys Phe Ile Asn Cys Thr Lys325 330
335Ile Asn Gly Asn Leu Ile Phe Leu Val Thr Gly Ile His Gly Asp Pro340
345 350Tyr Asn Ala Ile Glu Ala Ile Asp Pro
Glu Lys Leu Asn Val Phe Arg355 360 365Thr
Val Arg Glu Ile Thr Gly Phe Leu Asn Ile Gln Ser Trp Pro Pro370
375 380Asn Met Thr Asp Phe Ser Val Phe Ser Asn Leu
Val Thr Ile Gly Gly385 390 395
400Arg Val Leu Tyr Ser Gly Leu Ser Leu Leu Ile Leu Lys Gln Gln
Gly405 410 415Ile Thr Ser Leu Gln Phe Gln
Ser Leu Lys Glu Ile Ser Ala Gly Asn420 425
430Ile Tyr Ile Thr Asp Asn Ser Asn Leu Cys Tyr Tyr His Thr Ile Asn435
440 445Trp Thr Thr Leu Phe Ser Thr Ile Asn
Gln Arg Ile Val Ile Arg Asp450 455 460Asn
Arg Lys Ala Glu Asn Cys Thr Ala Glu Gly Met Val Cys Asn His465
470 475 480Leu Cys Ser Ser Asp Gly
Cys Trp Gly Pro Gly Pro Asp Gln Cys Leu485 490
495Ser Cys Arg Arg Phe Ser Arg Gly Arg Ile Cys Ile Glu Ser Cys
Asn500 505 510Leu Tyr Asp Gly Glu Phe Arg
Glu Phe Glu Asn Gly Ser Ile Cys Val515 520
525Glu Cys Asp Pro Gln Cys Glu Lys Met Glu Asp Gly Leu Leu Thr Cys530
535 540His Gly Pro Gly Pro Asp Asn Cys Thr
Lys Cys Ser His Phe Lys Asp545 550 555
560Gly Pro Asn Cys Val Glu Lys Cys Pro Asp Gly Leu Gln Gly
Ala Asn565 570 575Ser Phe Ile Phe Lys Tyr
Ala Asp Pro Asp Arg Glu Cys His Pro Cys580 585
590His Pro Asn Cys Thr Gln Gly Cys Asn Gly Pro Thr Ser His Asp
Cys595 600 605Ile Tyr Tyr Pro Trp Thr Gly
His Ser Thr Leu Pro Gln His Ala Arg610 615
620Thr Pro Gly Cys Val Ser Gly Asp Thr Ile Val Met Thr Ser Gly Gly625
630 635 640Pro Arg Thr Val
Ala Glu Leu Glu Gly Lys Pro Phe Thr Ala Leu Ile645 650
655Arg Gly Ser Gly Tyr Pro Cys Pro Ser Gly Phe Phe Arg Thr
Cys Glu660 665 670Arg Asp Val Tyr Asp Leu
Arg Thr Arg Glu Gly His Cys Leu Arg Leu675 680
685Thr His Asp His Arg Val Leu Val Met Asp Gly Gly Leu Glu Trp
Arg690 695 700Ala Ala Gly Glu Leu Glu Arg
Gly Asp Arg Leu Val Met Asp Asp Ala705 710
715 720Ala Gly Glu Phe Pro Ala Leu Ala Thr Phe Arg Gly
Leu Arg Gly Ala725 730 735Gly Arg Gln Asp
Val Tyr Asp Ala Thr Val Tyr Gly Ala Ser Ala Phe740 745
750Thr Ala Asn Gly Phe Ile Val His Ala Cys Gly Glu Gln Pro
Gly Thr755 760 765Gly Leu Asn Ser Gly Leu
Thr Thr Asn Pro Gly Val Ser Ala Trp Gln770 775
780Val Asn Thr Ala Tyr Thr Ala Gly Gln Leu Val Thr Tyr Asn Gly
Lys785 790 795 800Thr Tyr
Lys Cys Leu Gln Pro His Thr Ser Leu Ala Gly Trp Glu Pro805
810 815Ser Asn Val Pro Ala Leu Trp Gln Leu Gln820
825198627PRTArtificial Sequenceartificial construct relating to
Homo Sapiens immunoglobulin 198Gln Ser Val Cys Ala Gly Thr Glu Asn
Lys Leu Ser Ser Leu Ser Asp1 5 10
15Leu Glu Gln Gln Tyr Arg Ala Leu Arg Lys Tyr Tyr Glu Asn Cys
Glu20 25 30Val Val Met Gly Asn Leu Glu
Ile Thr Ser Ile Glu His Asn Arg Asp35 40
45Leu Ser Phe Leu Arg Ser Val Arg Glu Val Thr Gly Tyr Val Leu Val50
55 60Ala Leu Asn Gln Phe Arg Tyr Leu Pro Leu
Glu Asn Leu Arg Ile Ile65 70 75
80Arg Gly Thr Lys Leu Tyr Glu Asp Arg Tyr Ala Leu Ala Ile Phe
Leu85 90 95Asn Tyr Arg Lys Asp Gly Asn
Phe Gly Leu Gln Glu Leu Gly Leu Lys100 105
110Asn Leu Thr Glu Ile Leu Asn Gly Gly Val Tyr Val Asp Gln Asn Lys115
120 125Phe Leu Cys Tyr Ala Asp Thr Ile His
Trp Gln Asp Ile Val Arg Asn130 135 140Pro
Trp Pro Ser Asn Leu Thr Leu Val Ser Thr Asn Gly Ser Ser Gly145
150 155 160Cys Gly Arg Cys His Lys
Ser Cys Thr Gly Arg Cys Trp Gly Pro Thr165 170
175Glu Asn His Cys Gln Thr Leu Thr Arg Thr Val Cys Ala Glu Gln
Cys180 185 190Asp Gly Arg Cys Tyr Gly Pro
Tyr Val Ser Asp Cys Cys His Arg Glu195 200
205Cys Ala Gly Gly Cys Ser Gly Pro Lys Asp Thr Asp Cys Phe Ala Cys210
215 220Met Asn Phe Asn Asp Ser Gly Ala Cys
Val Thr Gln Cys Pro Gln Thr225 230 235
240Phe Val Tyr Asn Pro Thr Thr Phe Gln Leu Glu His Asn Phe
Asn Ala245 250 255Lys Tyr Thr Tyr Gly Ala
Phe Cys Val Lys Lys Cys Pro His Asn Phe260 265
270Val Val Asp Ser Ser Ser Cys Val Arg Ala Cys Pro Ser Ser Lys
Met275 280 285Glu Val Glu Glu Asn Gly Ile
Lys Met Cys Lys Pro Cys Thr Asp Ile290 295
300Cys Pro Lys Ala Cys Asp Gly Ile Gly Thr Gly Ser Leu Met Ser Ala305
310 315 320Gln Thr Val Asp
Ser Ser Asn Ile Asp Lys Phe Ile Asn Cys Thr Lys325 330
335Ile Asn Gly Asn Leu Ile Phe Leu Val Thr Gly Ile His Gly
Asp Pro340 345 350Tyr Asn Ala Ile Glu Ala
Ile Asp Pro Glu Lys Leu Asn Val Phe Arg355 360
365Thr Val Arg Glu Ile Thr Gly Phe Leu Asn Ile Gln Ser Trp Pro
Pro370 375 380Asn Met Thr Asp Phe Ser Val
Phe Ser Asn Leu Val Thr Ile Gly Gly385 390
395 400Arg Val Leu Tyr Ser Gly Leu Ser Leu Leu Ile Leu
Lys Gln Gln Gly405 410 415Ile Thr Ser Leu
Gln Phe Gln Ser Leu Lys Glu Ile Ser Ala Gly Asn420 425
430Ile Tyr Ile Thr Asp Asn Ser Asn Leu Cys Tyr Tyr His Thr
Ile Asn435 440 445Trp Thr Thr Leu Phe Ser
Thr Ile Asn Gln Arg Ile Val Ile Arg Asp450 455
460Asn Arg Lys Ala Glu Asn Cys Thr Ala Glu Gly Met Val Cys Asn
His465 470 475 480Leu Cys
Ser Ser Asp Gly Cys Trp Gly Pro Gly Pro Asp Gln Cys Leu485
490 495Ser Cys Arg Arg Phe Ser Arg Gly Arg Ile Cys Ile
Glu Ser Cys Asn500 505 510Leu Tyr Asp Gly
Glu Phe Arg Glu Phe Glu Asn Gly Ser Ile Cys Val515 520
525Glu Cys Asp Pro Gln Cys Glu Lys Met Glu Asp Gly Leu Leu
Thr Cys530 535 540His Gly Pro Gly Pro Asp
Asn Cys Thr Lys Cys Ser His Phe Lys Asp545 550
555 560Gly Pro Asn Cys Val Glu Lys Cys Pro Asp Gly
Leu Gln Gly Ala Asn565 570 575Ser Phe Ile
Phe Lys Tyr Ala Asp Pro Asp Arg Glu Cys His Pro Cys580
585 590His Pro Asn Cys Thr Gln Gly Cys Asn Gly Pro Thr
Ser His Asp Cys595 600 605Ile Tyr Tyr Pro
Trp Thr Gly His Ser Thr Leu Pro Gln His Ala Arg610 615
620Thr Pro Xaa625199628PRTArtificial Sequenceartificial
construct relating to Homo Sapiens immunoglobulin 199Gln Ser Val
Cys Ala Gly Thr Glu Asn Lys Leu Ser Ser Leu Ser Asp1 5
10 15Leu Glu Gln Gln Tyr Arg Ala Leu Arg
Lys Tyr Tyr Glu Asn Cys Glu20 25 30Val
Val Met Gly Asn Leu Glu Ile Thr Ser Ile Glu His Asn Arg Asp35
40 45Leu Ser Phe Leu Arg Ser Val Arg Glu Val Thr
Gly Tyr Val Leu Val50 55 60Ala Leu Asn
Gln Phe Arg Tyr Leu Pro Leu Glu Asn Leu Arg Ile Ile65 70
75 80Arg Gly Thr Lys Leu Tyr Glu Asp
Arg Tyr Ala Leu Ala Ile Phe Leu85 90
95Asn Tyr Arg Lys Asp Gly Asn Phe Gly Leu Gln Glu Leu Gly Leu Lys100
105 110Asn Leu Thr Glu Ile Leu Asn Gly Gly Val
Tyr Val Asp Gln Asn Lys115 120 125Phe Leu
Cys Tyr Ala Asp Thr Ile His Trp Gln Asp Ile Val Arg Asn130
135 140Pro Trp Pro Ser Asn Leu Thr Leu Val Ser Thr Asn
Gly Ser Ser Gly145 150 155
160Cys Gly Arg Cys His Lys Ser Cys Thr Gly Arg Cys Trp Gly Pro Thr165
170 175Glu Asn His Cys Gln Thr Leu Thr Arg
Thr Val Cys Ala Glu Gln Cys180 185 190Asp
Gly Arg Cys Tyr Gly Pro Tyr Val Ser Asp Cys Cys His Arg Glu195
200 205Cys Ala Gly Gly Cys Ser Gly Pro Lys Asp Thr
Asp Cys Phe Ala Cys210 215 220Met Asn Phe
Asn Asp Ser Gly Ala Cys Val Thr Gln Cys Pro Gln Thr225
230 235 240Phe Val Tyr Asn Pro Thr Thr
Phe Gln Leu Glu His Asn Phe Asn Ala245 250
255Lys Tyr Thr Tyr Gly Ala Phe Cys Val Lys Lys Cys Pro His Asn Phe260
265 270Val Val Asp Ser Ser Ser Cys Val Arg
Ala Cys Pro Ser Ser Lys Met275 280 285Glu
Val Glu Glu Asn Gly Ile Lys Met Cys Lys Pro Cys Thr Asp Ile290
295 300Cys Pro Lys Ala Cys Asp Gly Ile Gly Thr Gly
Ser Leu Met Ser Ala305 310 315
320Gln Thr Val Asp Ser Ser Asn Ile Asp Lys Phe Ile Asn Cys Thr
Lys325 330 335Ile Asn Gly Asn Leu Ile Phe
Leu Val Thr Gly Ile His Gly Asp Pro340 345
350Tyr Asn Ala Ile Glu Ala Ile Asp Pro Glu Lys Leu Asn Val Phe Arg355
360 365Thr Val Arg Glu Ile Thr Gly Phe Leu
Asn Ile Gln Ser Trp Pro Pro370 375 380Asn
Met Thr Asp Phe Ser Val Phe Ser Asn Leu Val Thr Ile Gly Gly385
390 395 400Arg Val Leu Tyr Ser Gly
Leu Ser Leu Leu Ile Leu Lys Gln Gln Gly405 410
415Ile Thr Ser Leu Gln Phe Gln Ser Leu Lys Glu Ile Ser Ala Gly
Asn420 425 430Ile Tyr Ile Thr Asp Asn Ser
Asn Leu Cys Tyr Tyr His Thr Ile Asn435 440
445Trp Thr Thr Leu Phe Ser Thr Ile Asn Gln Arg Ile Val Ile Arg Asp450
455 460Asn Arg Lys Ala Glu Asn Cys Thr Ala
Glu Gly Met Val Cys Asn His465 470 475
480Leu Cys Ser Ser Asp Gly Cys Trp Gly Pro Gly Pro Asp Gln
Cys Leu485 490 495Ser Cys Arg Arg Phe Ser
Arg Gly Arg Ile Cys Ile Glu Ser Cys Asn500 505
510Leu Tyr Asp Gly Glu Phe Arg Glu Phe Glu Asn Gly Ser Ile Cys
Val515 520 525Glu Cys Asp Pro Gln Cys Glu
Lys Met Glu Asp Gly Leu Leu Thr Cys530 535
540His Gly Pro Gly Pro Asp Asn Cys Thr Lys Cys Ser His Phe Lys Asp545
550 555 560Gly Pro Asn Cys
Val Glu Lys Cys Pro Asp Gly Leu Gln Gly Ala Asn565 570
575Ser Phe Ile Phe Lys Tyr Ala Asp Pro Asp Arg Glu Cys His
Pro Cys580 585 590His Pro Asn Cys Thr Gln
Gly Cys Asn Gly Pro Thr Ser His Asp Cys595 600
605Ile Tyr Tyr Pro Trp Thr Gly His Ser Thr Leu Pro Gln His Ala
Arg610 615 620Thr Pro Gly Xaa625
User Contributions:
comments("1"); ?> comment_form("1"); ?>Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
User Contributions:
Comment about this patent or add new information about this topic: