Patent application title: METHODS, COMPOSITIONS, AND KITS FOR DETECTION OF ASPERGILLOSIS
Inventors:
IPC8 Class: AG01N33573FI
USPC Class:
1 1
Class name:
Publication date: 2017-02-09
Patent application number: 20170038383
Abstract:
Provided herein are methods for detecting an Aspergillus protease in a
sample, diagnosing a subject with aspergillosis caused by an Aspergillus
infection based on the presence of an Aspergillus protease in a sample,
and methods of aspergillosis treatment that incorporate these diagnostic
methods. In certain embodiments, the Aspergillus protease is Asp f2, and
the Aspergillus infection is caused A. fumigatus, A. flavus, A.
versicolor, A. niger, or A. terreus. Also provided herein are antibodies
and kits for use in these methods, including novel antibodies specific
for Asp f2.Claims:
1. A method for detecting Aspergillus Asp f2 in a biological sample
comprising contacting the sample with an Asp f2 protease substrate that
can be cleaved by Asp f2, wherein cleavage of the protease substrate
indicates the presence of Asp f2 in the sample; and wherein the protease
substrate is selected from the group consisting of a protein or peptide
comprising the amino acid sequence of SEQ ID NO:5, an elastin protein or
peptide, and a collagen protein or peptide.
2. The method of claim 1, wherein the sample is enriched for Asp f2 prior to contact with the Asp f2 protease substrate by contacting the sample with an Asp f2-specific antibody or antigen binding fragment.
3. The method of claim 2, wherein the Asp f2-specific antibody is immobilized on an enrichment matrix.
4. The method of claim 1, wherein the protease substrate is conjugated to one or more fluorophores.
5. A method for diagnosing aspergillosis in a subject comprising: (a) obtaining a biological sample from the subject; (b) detecting the presence of Aspergillus Asp f2 in the sample by contacting the sample with an Asp f2 protease substrate that can be cleaved by Asp f2, wherein cleavage of the protease substrate indicates the presence of Asp f2 in the sample; and (c) diagnosing the subject with aspergillosis if Asp f2 is present in the sample, wherein the protease substrate is selected from the group consisting of a protein or peptide comprising the amino acid sequence of SEQ ID NO:5, an elastin protein or peptide, and a collagen protein or peptide.
6. The method of claim 5, further comprising (d) administering one or more therapeutic agents for the treatment of aspergillosis.
7. The method of claim 6, wherein the sample is enriched for Asp f2 prior to contact with the Asp f2 protease substrate by contacting the sample with an Asp f2-specific antibody or antigen binding fragment.
8. The method of claim 7, wherein the Asp f2-specific antibody is immobilized on an enrichment matrix.
9. The method of claim 5, wherein the protease substrate is conjugated to one or more fluorophores.
10. The method of claim 5, wherein the aspergillosis is the result of infection with an Aspergillus species selected from the group consisting of A. fumigatus, A. flavus, A. versicolor, A. niger, and A. terreus.
11. A method for the treatment of aspergillosis in a subject in need thereof comprising: (a) obtaining a biological sample from the subject; (b) detecting the presence of Aspergillus Asp f2 in the sample by contacting the sample with an Asp f2 protease substrate that can be cleaved by Asp f2, wherein cleavage of the protease substrate indicates the presence of Asp f2 in the sample; and (c) administering one or more therapeutic agents for the treatment of aspergillosis if Asp f2 is detected in the sample, and wherein the protease substrate is selected from the group consisting of a protein or peptide comprising the amino acid sequence of SEQ ID NO:5, an elastin protein or peptide, and a collagen protein or peptide.
12. The method of claim 11, wherein the sample is enriched for Asp f2 prior to contact with the Asp f2 protease substrate by contacting the sample with an Asp f2-specific antibody or antigen binding fragment.
13. The method of claim 12, wherein the Asp f2-specific antibody is immobilized on an enrichment matrix.
14. The method of claim 11, wherein the protease substrate is conjugated to one or more fluorophores.
15. The method of claim 11, wherein the aspergillosis is the result of infection with an Aspergillus species selected from the group consisting of A. fumigatus, A. flavus, A. versicolor, A. niger, and A. terreus.
16. A kit for performing the method of any one of claims 1, 5, and 11.
17. An Asp f2 protease substrate that can be cleaved by Asp f2, the protease substrate comprising the amino acid sequence of SEQ ID NO:5.
18. The protease substrate of claim 17, wherein the protease substrate is conjugated to one or more fluorophores.
Description:
RELATED APPLICATIONS
[0001] The present application is a continuation of U.S. application Ser. No. 14/217,004, filed Mar. 17, 2014, issuing as U.S. Pat. No. 9,416,395, which claims priority to U.S. Provisional Application No. 61/798,640, filed Mar. 15, 2013, the disclosure of which is incorporated by reference herein in its entirety, including drawings and sequence listing.
BACKGROUND
[0003] Aspergillus fumigatus is one of the most common Aspergillus species known to cause disease in humans. A. fumigatus is the most frequent cause of invasive fungal infection in immunosuppressed individuals, which include patients receiving immunosuppressive therapy for autoimmune or neoplastic disease, organ transplant recipients, and AIDS patients. A. fumigatus primarily causes invasive infection in the lung and represents a major cause of morbidity and mortality in these individuals. Additionally, A. fumigatus can cause chronic pulmonary infections or allergic disease in immunocompetent hosts. An A. fumigatus infection most commonly manifests as invasive pulmonary aspergillosis.
[0004] Only a few molecular assays exist for the detection of invasive fungal infections, including galactomannan and beta-glucan as well as DNA-based assays (e.g., PCR). The galactomannan and beta-glucan assays have limitations in terms of specificity and sensitivity. For example, antifungal medication is known to interfere with the galactomannan assay. A. fumigatus does not release much DNA into the circulation, which limits the utility of DNA-based assays. Therefore, there is a need for a sensitive assay to detect the presence of A. fumigatus in a sample from an infected patient.
SUMMARY
[0005] Provided herein are novel methods for detecting the presence of an Aspergillus protease such as Asp f2 in a sample and for diagnosing and treating a subject with aspergillosis caused by Aspergillus using this detection method. Also provided herein are kits for performing these methods, as well as antibodies specific to Aspergillus proteases such as Asp f2 and compositions, formulations, and kits comprising these antibodies.
[0006] Provided herein in certain embodiments are methods for detecting the presence of an Aspergillus protease in a biological sample comprising contacting the sample with a protease substrate comprising one or more Aspergillus protease cleavage sites, wherein cleavage of the protease substrate indicates the presence of the Aspergillus protease in the sample. In certain embodiments, the Aspergillus protease is Asp f2 or a homologue thereof. In certain embodiments, the biological sample is from a subject suspected of suffering from aspergillosis or previously diagnosed with aspergillosis. In certain embodiments, the sample is enriched for the Aspergillus protease prior to contact with the protease substrate, and in certain of these embodiments enrichment is carried out using an Aspergillus protease-specific antibody that is free in solution or immobilized to an enrichment matrix. In certain embodiments, the protease substrate is a protein or peptide comprising the amino acid sequence of SEQ ID NO: 5, an elastin protein or peptide, or a collagen protein or peptide. In certain embodiments, the protease substrate comprises one or more fluorophores, and in certain embodiments the protease substrate further comprises one or more acceptors. In certain of these embodiments, the fluorophore and/or acceptor is conjugated to the protease substrate via a peptide bond. In certain embodiments, the protease substrate is FluHSA2.
[0007] Provided herein in certain embodiments are methods for diagnosing aspergillosis in a subject comprising obtaining a biological sample from the subject, contacting the sample with a protease substrate comprising one or more Aspergillus protease cleavage sites, wherein cleavage of the protease substrate indicates the presence of the Aspergillus protease in the sample, and diagnosing the subject with aspergillosis if the Aspergillus protease is present in the sample. In certain embodiments, the Aspergillus protease is Asp f2 or a homologue thereof. In certain embodiments, the subject is suspected of suffering from aspergillosis or has been previously diagnosed with aspergillosis. In certain embodiments, the methods further comprise administering one or more therapeutic agents for treating aspergillosis. In certain embodiments, the sample is enriched for the Aspergillus protease prior to contact with the protease substrate, and in certain of these embodiments enrichment is carried out using an Aspergillus protease-specific antibody that is free in solution or immobilized to an enrichment matrix. In certain embodiments, the protease substrate is a protein or peptide comprising the amino acid sequence of SEQ ID NO:5, an elastin protein or peptide, or a collagen protein or peptide. In certain embodiments, the protease substrate comprises one or more fluorophores, and in certain embodiments the protease substrate further comprises one or more acceptors. In certain of these embodiments, the fluorophore and/or acceptor is conjugated to the protease substrate via a peptide bond. In certain embodiments, the protease substrate is FluHSA2.
[0008] Provided herein in certain embodiments are methods for treating aspergillosis in a subject comprising obtaining a biological sample from the subject, contacting the sample with a protease substrate comprising one or more Aspergillus protease cleavage sites, wherein cleavage of the protease substrate indicates the presence of the Aspergillus protease in the sample, and administering one or more therapeutic agents to the subject if the Aspergillus protease is determined to be present in the sample. In certain embodiments, the Aspergillus protease is Asp f2 or a homologue thereof. In certain embodiments, the therapeutic agent is a therapeutic agent that has not previously been administered to the subject. In other embodiments, the therapeutic agent is an agent that was previously administered to the subject, but administered at a higher dosage. In certain embodiments, the sample is enriched for the Aspergillus protease prior to contact with the protease substrate, and in certain of these embodiments enrichment is carried out using an Aspergillus protease-specific antibody or antigen binding fragment that is free in solution or immobilized to an enrichment matrix. In certain embodiments, the protease substrate is a protein or peptide comprising the amino acid sequence of SEQ ID NO:5, an elastin protein or peptide, or a collagen protein or peptide. In certain embodiments, the protease substrate comprises one or more fluorophores, and in certain embodiments the protease substrate further comprises one or more acceptors. In certain of these embodiments, the fluorophore and/or acceptor is conjugated to the protease substrate via a peptide bond. In certain embodiments, the protease substrate is FluHSA2.
[0009] Provided herein in certain embodiments are kits for use in detecting an Aspergillus protease in a sample, diagnosing a subject with aspergillosis based on the presence of an Aspergillus protease in a sample, or treating aspergillosis in a subject. In certain embodiments, the kits comprise instructions for use. In certain embodiments, the Aspergillus protease is Asp f2 or a homologue thereof. In certain embodiments, the kits provided herein comprise one or more antibodies or antigen binding fragments thereof that specifically bind an Aspergillus protease. In certain of these embodiments, the antibodies or antigen binding fragments thereof come pre-loaded on an enrichment matrix. In other embodiments, the antibodies or antigen binding fragments are not pre-loaded on an enrichment matrix. In certain of these embodiments, the kits may further comprise an enrichment matrix or components thereof on which a user may load the antibodies or antigen fragments thereof, such as beads to which the antibodies or antigen binding fragments may be immobilized. In certain embodiments, the kits comprise a protease substrate, and in certain of these embodiments the protease substrate is a protein or peptide comprising the amino acid sequence of SEQ ID NO: 5, an elastin protein or peptide, or a collagen protein or peptide. In certain embodiments, the protease substrate comprises one or more fluorophores, and in certain embodiments the protease substrate further comprises one or more acceptors. In certain of these embodiments, the fluorophore and/or acceptor is conjugated to the protease substrate via a peptide bond. In certain embodiments, the protease substrate is FluHSA2.
[0010] Provided herein in certain embodiments are antibodies or antigen binding fragments thereof that specifically bind an Aspergillus protease. In certain of these embodiments, the Aspergillus protease is Asp f2 or a homologue thereof. In certain embodiments, these antibodies or antigen binding fragments thereof comprise one or more CDR sequences comprising the amino acid sequences set forth in SEQ ID NOs: 8, 10, 12, 14, 16, or 18 or a combination thereof, and in certain of these embodiments the antibodies or antigen binding fragments thereof comprise a light chain variable region comprising the amino acid sequence of SEQ ID NO: 2 and/or a heavy chain variable region comprising the amino acid sequence of SEQ ID NO: 1. In other embodiments, the antibodies or antigen binding fragments thereof comprise the amino acid sequence of SEQ ID NO: 20 and/or 22. In certain embodiments, the antibodies are monoclonal, chimeric, or humanized antibodies, and in certain embodiments wherein the antibodies are monoclonal the antibodies are 5D7A1. In certain embodiments, the antigen binding fragments thereof is an scFv, F(ab')2, Fab, Fab' or Fv.
[0011] Provided herein in certain embodiments are methods for detecting the presence of an Aspergillus protease in a sample and for determining whether a subject is suffering from aspergillosis. The methods include the steps of exposing the sample containing the Aspergillus protease to an enrichment matrix comprising an Aspergillus protease-specific antibody that binds the Aspergillus protease and to a substrate composition comprising a protease substrate capable of eliciting a detectable fluorescence signal when modified by the Aspergillus protease, and measuring the level of change in the detectable fluorescence signal and detecting the presence of the Aspergillus protease when the level of change in the detectable fluorescence signal in the sample is elevated. When the level of change in the detectable fluorescence signal in the sample is elevated, the subject is treated for aspergillosis.
[0012] Provided herein in certain embodiments are methods for detecting the presence of an Aspergillus protease in a sample comprising (a) exposing the sample to an enrichment matrix comprising an Aspergillus protease-specific antibody or antigen-binding fragment thereof that specifically binds the Aspergillus protease, (b) exposing the sample to a protease substrate capable of eliciting a detectable fluorescence signal when modified by the Aspergillus protease, wherein the protease substrate comprises one or more Aspergillus protease cleavage sites, (c) measuring the level of change in the detectable fluorescence signal; and (d) detecting the presence of the Aspergillus protease when the level of change in the detectable fluorescence signal in the sample is elevated. In certain embodiments, the sample is exposed to the enrichment matrix prior to exposure to the protease substrate. In other embodiments, exposure to the enrichment matrix and the protease substrate occurs simultaneously. In certain of these embodiments, the Aspergillus protease is Asp f2 or a homologue thereof, and in certain of these embodiments the antibody or antigen binding fragment thereof specifically binds Asp f2 or a homologue thereof. In certain embodiments, the protease substrate is a circular substrate. In certain embodiments, the protease substrate is a protein or peptide comprising the amino acid sequence of SEQ ID NO: 5, an elastin protein or peptide, or a collagen protein or peptide, and in certain embodiments the protease substrate is FluHSA2. In certain embodiments, the protease substrate comprises at least one fluorophore conjugated via a peptide bond at or near the N-terminus and at least one acceptor conjugated at or near the C-terminus. In certain of these embodiments, the acceptor is a dark quencher. In certain embodiments, the step of exposure to the enrichment matrix and/or protease substrate occurs under conditions permitting binding of the Aspergillus protease to the antibody or antigen binding fragment thereof and modification of the protease substrate by the Aspergillus protease. In certain embodiments, the antibody or antigen binding fragment thereof comprises one or more CDRs selected from the group consisting of SEQ ID NOs: 8, 10, 12, 14, 16, and 18, and in certain embodiments the antibody or antigen binding fragments thereof comprises a variable heavy chain comprising the amino acid sequence of SEQ ID NO: 1 and/or a variable light chain comprising the amino acid sequence of SEQ ID NO: 2. In certain embodiments, the antibody is the monoclonal antibody 5D7A1. In certain embodiments, the level of change in the detectable fluorescence signal is elevated if it is significantly greater than the level of change in the detectable fluorescence signal from a negative control sample. In certain embodiments, the enrichment matrix further comprises a protease substrate-specific antibody or antigen binding fragment thereof that binds the protease substrate. In certain of these embodiments, the protease substrate-specific antibody or antigen binding fragment binds a protein or peptide component of the protease substrate. In other embodiments, the antibody or fragment thereof binds a fluorophore component of the protease substrate (e.g., an anti-fluorescein antibody) or an acceptor component of the protease substrate (e.g., an anti-DABCYL antibody). In certain embodiments, the enrichment matrix is an immunosorbent support comprised of loose beads or a fixed column.
[0013] Provided herein in certain embodiments are methods for determining whether a subject is suffering from aspergillosis comprising (a) exposing a biological sample from the subject to an enrichment matrix comprising an Aspergillus protease-specific antibody that binds an Aspergillus protease, (b) exposing the sample to a protease substrate capable of eliciting a detectable fluorescence signal when modified by the Aspergillus protease, wherein the protease substrate comprises one or more Aspergillus protease cleavage sites; and (c) measuring the level of change in the detectable fluorescence signal, wherein the subject is determined to be suffering from aspergillosis when the change in the detectable fluorescence signal in the sample is elevated. In certain embodiments, the methods further comprise (d) administering one or more therapeutic agents for treating aspergillosis. In certain embodiments, the sample is exposed to the enrichment matrix prior to exposure to the protease substrate. In other embodiments, exposure to the enrichment matrix and the protease substrate occurs simultaneously. In certain of these embodiments, the Aspergillus protease is Asp f2 or a homologue thereof, and in certain of these embodiments the antibody or antigen binding fragment thereof specifically binds Asp f2 or a homologue thereof. In certain embodiments, the protease substrate is a circular substrate. In certain embodiments, the protease substrate is a protein or peptide comprising the amino acid sequence of SEQ ID NO: 5, an elastin protein or peptide, or a collagen protein or peptide, and in certain embodiments the protease substrate is FluHSA2. In certain embodiments, the protease substrate comprises at least one fluorophore conjugated via a peptide bond at or near the N-terminus and at least one acceptor conjugated at or near the C-terminus. In certain of these embodiments, the acceptor is a dark quencher. In certain embodiments, the step of exposure to the enrichment matrix and/or protease substrate occurs under conditions permitting binding of the Aspergillus protease to the antibody or antigen binding fragment thereof and modification of the protease substrate by the Aspergillus protease. In certain embodiments, the antibody or antigen binding fragment thereof comprises one or more CDRs selected from the group consisting of SEQ ID NOs: 8, 10, 12, 14, 16, and 18, and in certain embodiments the antibody or antigen binding fragments thereof comprises a variable heavy chain comprising the amino acid sequence of SEQ ID NO: 1 and/or a variable light chain comprising the amino acid sequence of SEQ ID NO: 2. In certain embodiments, the antibody is the monoclonal antibody 5D7A1. In certain embodiments, the level of change in the detectable fluorescence signal is elevated if it is significantly greater than the level of change in the detectable fluorescence signal from a negative control sample. In certain embodiments, the enrichment matrix further comprises a protease substrate-specific antibody or antigen binding fragment thereof that binds the protease substrate. In certain of these embodiments, the protease substrate-specific antibody or antigen binding fragment binds a protein or peptide component of the protease substrate. In other embodiments, the antibody or fragment thereof binds a fluorophyl component of the protease substrate (e.g., an anti-fluorescein antibody) or an acceptor component of the protease substrate (e.g., an anti-DABCYL antibody). In certain embodiments, the enrichment matrix is an immunosorbent support comprised of loose beads or a fixed column.
[0014] Provided herein in certain embodiments is a protease substrate comprising a donor fluorophore, an acceptor having an absorbance spectrum overlapping the emission spectrum of the donor fluorophore, and one or more Aspergillus protease cleavage sites. In certain of these embodiments, the protease substrate comprises one or more cleavage sites for Asp f2 or a homologue thereof. In certain embodiments, the protease substrate is a circular substrate. In certain embodiments, the protease substrate is a protein or peptide comprising the amino acid sequence of SEQ ID NO: 5, an elastin protein or peptide, or a collagen protein or peptide, and in certain embodiments the protease substrate is FluHSA2. In certain embodiments, the fluorophore is conjugated to a protein or peptide component of the protease substrate via a peptide bond at or near the N-terminus, and in certain embodiments the acceptor is conjugated to a protein or peptide component of the protease substrate via a peptide bond at or near the C-terminus. In certain of these embodiments, the acceptor is a dark quencher.
[0015] Provided herein in certain embodiments are kits for detecting an Aspergillus protease. In some embodiments, the detection kit may be an enrichment matrix comprised of one or more immunoaffinity beads to which at least one Aspergillus protease specific antibody is bound and at least one substrate, where a detectable fluorescent signal is produced upon interaction of the substrate with the Aspergillus protease.
[0016] Provided herein in certain embodiments are kits for detecting an Aspergillus protease in a sample comprising (a) an enrichment matrix comprising one or more immunoaffinity beads to which at least one Aspergillus protease-specific antibody or antigen binding fragment is bound and (b) at least one protease substrate, wherein a detectable fluorescent signal is produced upon interaction of the protease substrate with the Aspergillus protease. In certain embodiments, the Aspergillus protease is Asp f2 or a homologue thereof, and in certain of these embodiments the antibody or antigen binding fragment thereof specifically binds Asp f2 or a homologue thereof. In certain embodiments, the antibody or antigen binding fragment thereof comprises one or more CDRs selected from the group consisting of SEQ ID NOs: 8, 10, 12, 14, 16, and 18, and in certain embodiments the antibody or antigen binding fragments thereof comprises a variable heavy chain comprising the amino acid sequence of SEQ ID NO: 1 and/or a variable light chain comprising the amino acid sequence of SEQ ID NO: 2. In certain embodiments, the antibody is the monoclonal antibody 5D7A1. In certain embodiments, the protease substrate is a circular substrate. In certain embodiments, the protease substrate is a protein or peptide comprising the amino acid sequence of SEQ ID NO: 5, an elastin protein or peptide, or a collagen protein or peptide, and in certain embodiments the protease substrate is FluHSA2. In certain embodiments, the protease substrate comprises at least one fluorophore conjugated via a peptide bond at or near the N-terminus and at least one acceptor conjugated at or near the C-terminus. In certain of these embodiments, the acceptor is a dark quencher. Also provided herein is the use of the kits provided herein for detecting an Aspergillus protease in a sample.
[0017] Provided herein in certain embodiments are antibodies and antigen binding fragments thereof that specifically bind an Aspergillus protease. In certain of these embodiments, the Aspergillus protease is Asp f2 or a homologue thereof, and in certain embodiments the Asp f2 or homologue thereof is glycosylated. In certain embodiments, the antibody or antigen binding fragment thereof comprises one or more CDRs selected from the group consisting of SEQ ID NOs: 8, 10, 12, 14, 16, and 18, and in certain embodiments the antibody or antigen binding fragments thereof comprises a variable heavy chain comprising the amino acid sequence of SEQ ID NO: 1 and/or a variable light chain comprising the amino acid sequence of SEQ ID NO: 2. In certain embodiments, the antibody is the monoclonal antibody 5D7A1. In certain embodiments, the antibody is a monoclonal antibody, a chimeric antibody, a humanized antibody, an affinity matured antibody, a human antibody, or a bispecific antibody. Also provided herein are compositions and formulations that comprise the antibodies and antigen binding fragments disclosed herein, as well as the use of these antibodies and fragments thereof, compositions, and formulations in methods and kits for detecting an Aspergillus protease in a sample.
BRIEF DESCRIPTION OF THE DRAWINGS
[0018] FIG. 1 shows the amino acid sequence and 5'-3' and 3'-5' DNA sequences of the variable heavy (VH) and variable light (VL) chains of the Asp f2-specific monoclonal antibody 5D7A1 that binds the native, glycosylated form of A. fumigatus Asp f2. The amino acid sequence of the variable heavy chain of 5D7A1 is marked with triangles (SEQ ID NO: 1) and the amino acid sequence of the variable light chain of 5D7A1 is marked with stars (SEQ ID NO: 2). The 5'-3' DNA sequence of 5D7A1 and surrounding vector region is marked with circles (SEQ ID NO: 24) and the 3'-5' complementary DNA strand is marked with squares (SEQ ID NO: 25). Note that although the figure shows the 3'-5' complementary DNA strand (see squares), SEQ ID NO: 25 in the sequence listing represents the reverse (i.e., 5'-3') complement of SEQ ID NO: 24. The DNA sequence was confirmed by mass spectrometric analysis of trypsin and chymotrypsin-derived peptides of the monoclonal antibody.
[0019] FIG. 2 shows the amino acid sequences of the VH chain (SEQ ID NO: 1) and VL chain (SEQ ID NO: 2) of the monoclonal antibody 5D7A1.
[0020] FIG. 3 shows DNA sequences encoding the VH chain (SEQ ID NO: 3) and VL chain (SEQ ID NO: 4) of the monoclonal antibody 5D7A1.
[0021] FIG. 4 shows the amino acid sequence of the FluHSA2 peptide (SEQ ID NO: 5). The sequence contains a fluorophore and a quencher. The 5-carboxyfluorescein (5-Fam) fluorophore is conjugated to the epsilon-amino group of threonine at the N-terminus and the 4-((4-(dimethylamino)phenyl)azo)benzoic acid (DABCYL) quencher is conjugated to the epsilon amino group of the lysine side chain at the C-terminus of the peptide.
[0022] FIG. 5 illustrates the mass spectrometric analysis (MALDI spectra) and primary structure of the FluHSA2 substrate (SEQ ID NO: 5) and its cleavage products produced by reaction with native Asp f2. The FluHSA2 was cleaved at four sites near the C-terminus of the peptide. In bold are the cysteine residues (represented by C) that form a disulfide bond which results in circularization of the substrate.
[0023] FIG. 6 illustrates the bead-based Asp f2 activity measured in bronchoalveolar lavage fluid (BALF) samples from City of Hope patients with either probable or proven aspergillosis, other mold infections, or no infection. The level of significance between patients with probable or proven aspergillosis and those that had other mold infections or were negative was p<0.0001. Statistical evaluation was performed by ANOVA. ****=significant, ns=non-significant.
[0024] FIG. 7 shows the amino acid sequence alignment of Asp f2 from A. fumigatus (gi|8300352) (SEQ ID NO: 27, marked with squares) and the deuterolysin from Aspergillus oryzae RIB40 (gi|94730401) (SEQ ID NO: 26, marked with circles). Identity between the sequences is 24% and homology is 35%.
[0025] FIG. 8 shows the amino acid sequence alignment of the Asp f2 from A. fumigatus (gi|83300352) (SEQ ID NO:29, marked with squares) and putative metalloprotease MEP20 from A. fumigatus (gi|780794) (SEQ ID NO:28, marked with circles). Identity between the sequences is 25% and homology is 39%.
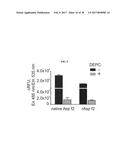
[0026] FIG. 9 shows the effect of diethylpyrocarbonate (DEPC) on proteolytic activity of native and recombinant forms of Asp f2. After one hour incubation with DEPC at 22.degree. C., Asp f2 protease activity is inhibited. Treatment without DEPC is indicated by the black bar and treatment with DEPC is indicated by the grey bar.
[0027] FIG. 10 shows the effects of various protease inhibitors and Zn.sup.2+ ions on proteolytic activity of native Asp f2. Samples were treated as indicated with iodoacetamide (IAA), aprotinin, pepstatin A, phenylmethanesulfonylfluoride (PMSF), ethylenediaminetetraacetic acid, disodium salt (EDTA), 1-10-phenantroline, soybean trypsin inhibitor (SBTI), tetrakis-(2-pyridylmethyl) ethylenediamine (TPEN), or Zn.sup.2+.
[0028] FIG. 11 shows the effects of various inhibitors and Zn.sup.2+ ions on proteolytic activity of recombinant Asp f2. Samples were treated as indicated with phenylmethanesulfonylfluoride (PMSF), ethylenediaminetetraacetic acid, disodium salt (EDTA), 1-10-phenantroline, aprotinin, pepstatin A, soybean trypsin inhibitor (SBTI), iodoacetamide (IAA), tetrakis-(2-pyridylmethyl) ethylenediamine (TPEN), or Zn.sup.2+.
[0029] FIG. 12 shows the full length amino acid sequence (1-310) of recombinant Asp f2 (SEQ ID NO: 30) and the analysis of Asp f2 histidines. Histidines H138, H186, H190, and H201 were mutated to alanine. H186 and H190 are located in the HRLYH motif.
[0030] FIG. 13 shows that the H186A Asp f2 mutant (white bar) displays significantly lower proteolytic activity with FluHSA2 compared to the original wild type Asp f2 construct (black bar).
[0031] FIG. 14 shows the fusion protein construct for Trx-SMT3-Asp f2 [aa 32-310] (where aa is amino acids). The Trx-SMT3 N-terminal linker contained the following from N- to C-terminus: Thioredoxin (Trx)-(glycine-serine-linker-His.sub.6)-(thrombin site)-(S-tag)-(enterokinase site)-(SMT3) and Asp f2 [aa 32-310]. Thioredoxin (Trx) was added to improve solubility of the construct and a yeast SUMO (Smt3) was added to help enhance expression and promote solubility of Asp f2.
[0032] FIG. 15 shows the amino acid and DNA sequence (5'-3' and 3'-5 of the fusion protein construct, Trx-SMT3-Asp f2 [aa 32-310]. The amino acid sequence of Trx-SMT3-Asp f2 is marked with triangles (SEQ ID NO: 6). The 5'-3' DNA sequence of Trx-SMT3-Asp f2 is marked with circles (SEQ ID NO: 7) and the 3'-5' complementary sequence strand is marked with squares (SEQ ID NO: 31). Note that although the figure shows the 3'-5' complementary DNA strand (see squares), SEQ ID NO: 31 in the sequence listing represents the reverse (i.e., 5'-3') complement of SEQ ID NO: 7.
[0033] FIG. 16 shows the amino acid sequence and the DNA sequence of the fusion protein construct, Trx-SMT3-Asp f2 [aa 32-310] (SEQ ID NOS: 6 and 7, respectively).
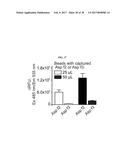
[0034] FIG. 17 illustrates the proteolytic activity of native Asp f2 and native Asp f3. Native Asp f2 and native Asp f3 were both tested with 25 .mu.l (white bars) and 50 .mu.l (black bars).
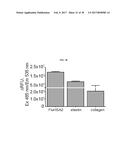
[0035] FIG. 18 illustrates the proteolytic activity of Asp f2 with different substrates including fluorogenic versions of elastin and collagen.
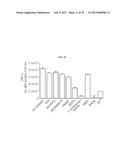
[0036] FIG. 19 shows a bead-based assay using different antibodies as indicated to assess Asp f2 proteolytic cleavage of the FITC-collagen substrate. Asp f2 showed the most robust proteolytic activity with the anti-Asp f2 antibody, 5D7A1, compared with other antibodies that are non-Asp f2 specific antibodies (4E17.1, anti-Asp f1, anti-Asp f3, and rabbit IgG).
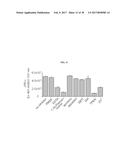
[0037] FIG. 20 shows a bead-based assay using different antibodies as indicated to assess Asp f2 proteolytic cleavage of the 5-Fam-elastin substrate. Asp f2 showed the most robust proteolytic activity with the anti-Asp f2 antibody, 5D7A1 compared with other antibodies that are non-Asp f2 specific antibodies (4E17.1, anti-Asp f1, anti-Asp f3, and rabbit IgG).
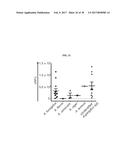
[0038] FIG. 21 shows Asp f2 proteolytic activity in aspergillosis patients for which Aspergillus species were identified as indicated by the clinical Microbiology lab at City of Hope. =A. fumigatus, .box-solid.=A. flavus, .tangle-solidup.=A. versicolor, .quadrature.=A. niger, =A. terreus, .largecircle.=unidentified Aspergillus species.
[0039] FIG. 22 shows the amino acid and DNA sequences of the variable heavy (VH) and variable light (VL) complementarity determining region (CDR) 1 regions of the monoclonal antibody, 5D7A1: VH CDR1 amino acid sequence (SEQ ID NO: 8); VH CDR1 DNA sequence (SEQ ID NO: 9); VL CDR1 amino acid sequence (SEQ ID NO: 10); and VL CDR1 DNA sequence (SEQ ID NO: 11).
[0040] FIG. 23 shows the amino acid and DNA sequences of the VH and VL CDR2 regions of the monoclonal antibody, 5D7A1: VH CDR2 amino acid sequence (SEQ ID NO: 12); VH CDR2 DNA sequence (SEQ ID NO: 13); VL CDR2 amino acid sequence (SEQ ID NO: 14); and VL CDR2 DNA sequence (SEQ ID NO: 15).
[0041] FIG. 24 shows the amino acid and DNA sequences of the VH and VL CDR3 regions of the monoclonal antibody, 5D7A1: VH CDR3 amino acid sequence (SEQ ID NO: 16); VH CDR3 DNA sequence (SEQ ID NO: 17); VL CDR3 amino acid sequence (SEQ ID NO: 18); and VL CDR3 DNA sequence (SEQ ID NO: 19).
[0042] FIG. 25 shows a ROC curve illustrating the sensitivity, specificity, and appropriate cut-off values for the presently disclosed Asp f2 activity assay. Solid line=aspergillosis, dotted line=all other mold infections.
[0043] FIG. 26A shows clinical galactomannan (GM) test results for diagnosis of aspergillosis infection.
[0044] FIG. 26B shows the presently disclosed Asp f2 activity assay for diagnosis of aspergillosis infection.
[0045] FIG. 27 shows the amino acid sequence of VL-G515-VH-Fc scFv and the DNA sequence of the VL-G515-VH-Fc scFv in the pEE12.4 vector. The amino acid sequence of VL-G515-VH-Fc scFv is marked with triangles (SEQ ID NO: 20). The 5'-3' DNA sequence of the VL-G515-VH-Fc scFv and surrounding pEE12.4 vector region is marked with circles (SEQ ID NO: 32) and the 3'-5' complementary strand is marked with squares (SEQ ID NO: 33). Note that although the figure shows the 3'-5' complementary DNA strand (see squares), SEQ ID NO: 33 in the sequence listing represents the reverse (i.e., 5'-3') complement of SEQ ID NO: 32.
[0046] FIG. 28 shows the amino acid of the V.sub.L-GS15-V.sub.H-Fc scFv and the DNA sequence of the scFv in the pEE12.4 vector.
[0047] FIG. 29 shows the DNA sequence of the VH-GS15-VL-Fc scFv in the pEE12.4 vector. The 5'-3' DNA sequence of the VH-GS15-VL-Fc scFv and surrounding pEE12.4 vector region is marked with circles (SEQ ID NO: 34) and the 3'-5' complementary strand is marked with squares (SEQ ID NO: 35). Note that although the figure shows the 3'-5' complementary DNA strand (see squares), SEQ ID NO: 35 in the sequence listing represents the reverse (i.e., 5'-3') complement of SEQ ID NO: 34.
[0048] FIG. 30 shows the amino acid of the V.sub.H-GS15-V.sub.L-Fc scFv and the DNA sequence of the scFv in the pEE12.4 vector.
[0049] FIG. 31A shows a Western blots for V.sub.H-GS15-V.sub.L-FC scFv binding to Asp f2. Asp f2-specific bands are marked with arrows. The following protein samples were loaded. M: molecular weight marker; 1: A. fumigatus culture filtrate of a 9-day culture; 2: A. fumigatus culture filtrate of another, shorter culture; 3: recombinant (r)SMT3-Asp f2. The strong band at .about.40 kDa represents unglycosylated Asp f2 that yields a stronger signal with 5D7A1 than with V.sub.H-GS15-V.sub.L-FC scFv.
[0050] FIG. 31B shows a Western blot for 5D7A1 binding to Asp f2. Asp f2-specific bands are marked with arrows. The following protein samples were loaded. M: molecular weight marker; 1: A. fumigatus culture filtrate of a 9-day culture; 2: A. fumigatus culture filtrate of another, shorter culture; 3: recombinant (r)SMT3-Asp f2. The strong band at .about.40 kDa represents unglycosylated Asp f2 that yields a stronger signal with 5D7A1 than with V.sub.H-GS15-V.sub.L-FC scFv.
DETAILED DESCRIPTION
[0051] As disclosed herein, the Aspergillus allergen Asp f2 has unexpectedly been found to possess zinc metalloprotease activity. Based on this finding, a novel activity-based method is provided herein for detecting Aspergillus proteases in a sample by contacting the sample with a protease substrate comprising one or more protease cleavage sites. Cleavage of the substrate indicates the presence of Aspergillus protease in the sample. In certain embodiments, this method can be used to diagnose a subject with aspergillosis caused by an Aspergillus infection by detecting the presence of a Aspergillus proteases in a biological sample from the subject. In other embodiments, this detection method can be incorporated into a method of treating aspergillosis caused by an Aspergillus infection. For example, detection of the presence of an Aspergillus protease in a biological sample from a subject can be used to determine whether to administer or re-administer a therapeutic agent, or to help determine an effective dosage for administering such an agent. Also provided herein are kits for carrying out the detection, diagnosis, and treatment methods disclosed herein. The methods and kits provided herein are superior to previously developed methods, in part because they provide equal or greater sensitivity with reduced time and cost. The methods disclosed herein may be performed in either a manual or automated format.
[0052] In certain embodiments, the methods and kits provided herein may be used to detect the presence of Aspergillus protease Asp f2. In other embodiments, the methods and kits provided herein may be used to detect the presence of other Aspergillus proteases, including but not limited to Aspergillus proteases comprising the amino acid motifs HRLYH (SEQ ID NO: 36) or HEXXH (SEQ ID NO: 37), or an amino acid sequence with significant sequence identity to these motifs. In certain of these embodiments, an amino acid sequence with significant sequence identity to these motifs comprises one or more conservative substitutions versus the motif.
[0053] In certain embodiments, the methods and kits provided herein may be used to detect an Aspergillus protease from Aspergillus fumigatus, including but not limited to Asp f2. In other embodiments, the methods and kits may be used to detect Aspergillus proteases such as Asp f2 or homologues thereof from other Aspergillus species, including for example A. nidulans, A. versicolor, A. niger, and A. terreus. Accordingly, the methods and kits disclosed herein may be used to diagnose a subject with aspergillosis caused by infection with A. fumigatus, A. nidulans, A. versicolor, A. niger, or A. terreus, or to treat aspergillosis caused by infection with A. fumigatus, A. nidulans, A. versicolor, A. niger, or A. terreus. In certain embodiments, the Aspergillus protease being detected is glycosylated at one or more locations within the protein. For example, in certain embodiments the methods and kits provided herein detect glycosylated Asp f2.
[0054] In certain embodiments of the methods and kits provided herein, a sample is enriched for an Aspergillus protease prior to or at the same time as contact with the protease substrate. In certain of these embodiments, enrichment is achieved by contacting the sample with one or more protease-specific antibodies or antigen binding fragments thereof. For example, where the Aspergillus protease is Asp f2, the sample may be contacted with one or more Asp f2-specific antibodies or antigen binding fragments thereof. In these embodiments, the antibodies or antigen binding fragments thereof may be monoclonal antibodies, chimeric antibodies, humanized antibodies, affinity matured antibodies, human antibodies, bispecific antibodies, or any antigen binding fragment thereof such as an scFv, F(ab')2, Fab, Fab' or Fv. In certain embodiments, the Aspergillus protease-specific antibody or antigen binding fragment thereof binds the Aspergillus protease in such a way such that the protease retains its catalytic activity after binding.
[0055] Provided herein in certain embodiments are Aspergillus protease-specific antibodies or antigen binding fragments thereof, as well as compositions, formulations, and kits comprising these antibodies or antigen binding fragments thereof and the use of these antibodies or antigen binding fragments thereof in the detection, diagnosis, and treatment methods disclosed herein. As discussed in the Examples below, the novel monoclonal antibody 5D7A1 was used to bind Asp f2 in a bead-based assay. Accordingly, in certain embodiments compositions, formulations, and kits are provided that comprise 5D7A1 alone or in combination with other components. In certain embodiments, the antibodies or antigen binding fragments thereof provided herein comprise the complete variable light or variable heavy chain of 5D7A1 or homologues thereof. In these embodiments, the antibodies or antigen binding fragments thereof comprise a light chain comprising the amino acid sequence of SEQ ID NO: 2, a heavy chain comprising the amino acid sequence of SEQ ID NO: 1, or a combination thereof. In other embodiments, the antibodies or antigen binding fragments thereof provided herein comprise a portion of the variable light or variable heavy chain of 5D7A1. In certain of these embodiments, the antibodies or antigen binding fragments thereof comprise one or more CDRs selected from the group consisting of SEQ ID NOs: 8, 10, 12, 14, 16, and 18 (FIGS. 22-24), or any combination thereof. In certain embodiments, an antigen binding fragment thereof as provided herein is an scFv, and in certain of these embodiments the scFv comprises the amino acid sequence of SEQ ID NO: 20 or 22 or one or more CDRs set forth therein (FIGS. 28, 30).
[0056] In certain embodiments where a sample is enriched for an Aspergillus protease using an Aspergillus protease-specific antibody or antigen binding fragment thereof, the antibody or antigen binding fragment thereof may be free in solution. In these embodiments, the Aspergillus protease-antibody conjugate resulting from binding may be removed from solution for testing with a protease substrate using methods well known in the art. In other embodiments, the antibody or antigen binding fragment thereof may be attached to or immobilized on an enrichment matrix such as an immunosorbent support of loose beads or a fixed column. In these embodiments, the enrichment matrix may comprise immunoaffinity beads including cyanogen-bromide (CNBr) activated Sepharose beads, protein-A, protein-G, or protein A/G conjugated Sepharose, agarose, or magnetic beads coupled and cross-linked to Aspergillus protease-specific antibodies or antigen binding fragments thereof.
[0057] In those embodiments of the methods and kits disclosed herein that utilize an enrichment matrix, formation of the Aspergillus protease-antibody conjugate results in the Aspergillus protease being attached to or immobilized on the enrichment matrix. For example, where a sample is being tested for the presence of Asp f2, the sample may be contacted with an enrichment matrix comprising one or more Asp f2-specific antibodies that bind Asp f2 to form an Asp f2-antibody conjugate. During or after formation of the Aspergillus protease-antibody conjugate, the enrichment matrix is contacted with the protease substrate. In certain embodiments, the enrichment matrix may comprise the protease substrate prior to addition of the sample. In other embodiments, the protease substrate may be added to the enrichment matrix after sample addition. In certain of these embodiments, the enrichment matrix may be washed prior to contact with the protease substrate in order to remove unbound Aspergillus protease and/or other proteins and molecules. If the Aspergillus protease is present in the enrichment matrix, the protease substrate will be cleaved upon contact with the enrichment matrix.
[0058] In certain embodiments of the methods disclosed herein, exposure of a sample to an enrichment matrix and to a protease substrate occurs under conditions permitting binding of Aspergillus protease to an antibody or antigen binding fragment thereof on the enrichment matrix and modification of the protease substrate by the Aspergillus protease. In certain embodiments, one or more of these steps may be carried out in the dark.
[0059] A protease substrate for use in the methods and kits disclosed herein can be any chemical, biochemical, or biological species or compound that reacts with or is capable of being modified by an Aspergillus protease. In certain embodiments, the protease substrate comprises one or more Aspergillus protease cleavage sites, and in certain of these embodiments the protease substrate may comprise one, two, three, four, or five or more cleavage sites. For example, where the Aspergillus protease is Asp f2, the protease substrate comprises one or more Asp f2 cleavage sites. In certain of these embodiments, the Asp f2 cleavage site comprises the amino acid sequence FSALK (SEQ ID NO: 38). As disclosed in the Examples below, Asp f2 is a protease that cleaves and/or degrades certain protein or peptide substrates, including the peptide of SEQ ID NO:5, collagen, and elastin. In certain embodiments of the methods and kits provided herein wherein the Aspergillus protease is Asp f2, the Asp f2 protease substrate is a peptide comprising the amino acid sequence FSALK (SEQ ID NO: 38), the amino acid sequence of SEQ ID NO: 5, a human serum albumin protein or portion thereof (e.g., a peptide), an elastin protein or portion thereof (e.g., a peptide), or a collagen protein or portion thereof (e.g., a peptide). In certain embodiments, the protease substrate is a circular substrate, which can increase the stability of the peptide by enhancing protection against digestion by other proteases. In certain of these embodiments, the circular substrate is formed through a disulfide bond between two cysteines located within the peptide. In other embodiments, the substrate may be circularized through amino acids other than cysteines. For example, a glutamic acid residue may be positioned at a location 5' of the protease cleavage site, and the peptide may be circularized via an N-terminus to glutamate side chain peptide bond. In other embodiments, the protease substrate is non-circular.
[0060] Cleavage or other modification of a protease substrate may be detected in a variety of ways well known in the art. For example, the protease substrate may be removed from solution or from an enrichment matrix and run on a gel to detect changes in size. In other embodiments, the protease substrate may be capable of eliciting a detectable fluorescent signal when modified by an Aspergillus protease. In these embodiments, cleavage may be detected based on this fluorescent signal. For example, in certain embodiments the protease substrate may comprise a donor fluorophore such as 5-carboxyfluorescein (5-Fam) or 4-methylumbelliferone (4-Mu) and an acceptor having an absorbance spectrum overlapping the emission spectrum of the donor fluorophore, including for example a dark quencher such as 4-(dimethylaminoazo)benzene-4-carboxy (DABCYL). In certain embodiments, the one or more fluorophores may be conjugated to the peptide or protein component of the protease substrate at or near the N-terminus of the peptide or protein. As used herein, "near the N-terminus" refers to any position on the peptide or protein component of the protease substrate that is within five amino acids of the N-terminus of the peptide or protein, while "near the C-terminus" refers to any position on the peptide or protein component of the protease substrate that is within five amino acids of the C-terminus of the peptide or protein. In certain of these embodiments, the fluorophore (e.g., 5-FAM) is conjugated to an alpha-amino group of an N-terminal amino acid, and in certain of these embodiments the N-terminal amino acid is a threonine residue. In other embodiments, the fluorophore may be conjugated to an epsilon-amino group of an N-terminal amino acid. Similarly, in certain embodiments the acceptor may be conjugated to the peptide or protein component at or near the C-terminus of the peptide or protein. In certain of these embodiments, the acceptor (e.g., DABCYL) is conjugated to an epsilon-amino group of a C-terminal amino acid, and in certain of these embodiments the C-terminal amino acid is a lysine residue.
[0061] In those embodiments where the fluorophore and/or acceptor are conjugated to the peptide or protein component of the protease substrate, conjugation may be via a peptide bond, which enhances the stability of the substrate. In certain embodiments wherein a protease substrate comprises a fluorophore and an acceptor, the acceptor (e.g., DABCYL) suppresses the fluorescence emission of the fluorophore when the protease substrate is intact and the acceptor and fluorophore remain close together. When the substrate is cleaved by the Aspergillus protease, the fluorophore and acceptor are separated and the fluorophore emits light energy upon excitation. Thus, protease substrate cleavage (and hence the presence of Aspergillus protease) is detected by contacting the protease substrate with an excitatory wavelength of light. If the protease substrate is cleaved (i.e., if the Aspergillus protease is present), the fluorophore will emit fluorescence upon excitation. If the protease substrate remains intact, the acceptor will suppress fluorescent emission. The protease substrate may be contacted with the excitatory wavelength in solution, on an enrichment matrix, or after removal or elution from an enrichment matrix. In certain embodiments, the protease substrate is part of a substrate composition comprising the protease substrate plus one or more additional components.
[0062] Detection of a fluorescent signal in conjunction with the methods disclosed herein may be performed using a handheld ultraviolet (UV) light, a fluorescence excitation and/or detecting tool, or any suitable commercially available fluorometer. In some embodiments, a Victor X2 multilabel plate reader (Perkin Elmer, Shelton, Conn.) may be used to detect fluorescence. In some embodiments, the level of the change in the detectable fluorescence signal is calculated as a change in relative fluorescence unit (RFU). In certain embodiments, the level of the change in the fluorescence signal may be classified as elevated (i.e., Aspergillus protease is present) when the level of the change in the signal is greater than or equal to a predetermined level of background fluorescence. In some embodiments, the predetermined level of background fluorescence may be the fluorescent level of an Aspergillus protease-free control sample. In some embodiments, the level of the change in the detectable fluorescence signal is elevated when the level is significantly greater than the level of change in detectable fluorescence signal from a negative control sample. As described herein, a "negative control" sample comprises a sample that does not comprise Aspergillus protease, for example a biological sample from a subject that is known to not be infected with Aspergillus. In other embodiments, the negative control sample may be treated with a metalloprotease inhibitor.
[0063] In certain embodiments, an enrichment matrix may comprise beads coupled and cross-linked to antibodies or antigen binding fragments thereof that bind a fluorophore and/or acceptor conjugated to the Aspergillus protease substrate. For example, cyanogen-bromide (CNBr)-activated Sepharose beads may be coupled and cross linked to anti-FITC antibodies that bind a fluorescent 5-Fam label conjugated to the Aspergillus protease substrate. In another example, the enrichment matrix comprises CNBr-activated Sepharose beads coupled and cross-linked to anti-DABCYL antibodies or antigen binding fragments thereof that bind DABCYL conjugated to the Aspergillus protease substrate. In certain embodiments, the enrichment matrix may further comprise anti-protease substrate specific antibodies or antigen binding fragments thereof. In certain embodiments, the anti-protease substrate specific antibodies or antigen binding fragments thereof may bind a portion of the protease substrate including a fluorophore or acceptor or amino acids of the peptide or protein component of the substrate sequence.
[0064] In certain embodiments, the enrichment matrix may comprise a double affinity matrix comprising beads coupled and cross-linked to antibodies or antigen binding fragments thereof that specifically bind the Aspergillus protease as described herein and beads coupled and cross-linked to antibodies or antigen binding fragments thereof that specifically bind the Aspergillus protease substrate or fluorophore and/or acceptors conjugated to the Aspergillus protease substrate as described herein.
[0065] In certain embodiments wherein the Aspergillus protease being detected is Asp f2, the protease substrate may consist of or comprise FluHSA2 (see Example 1 below). In these embodiments, the FluHSA2 protease substrate may be circular or non-circular. In certain of these embodiments, the methods and kits and utilize an enrichment matrix comprising CNBr-activated Sepharose beads coupled and cross linked to antibodies or antigen binding fragments thereof that bind a portion of FluHSA2.
[0066] In certain embodiments of the diagnosis and treatment methods provided herein, the methods are used to determine if a subject is currently suffering from aspergillosis caused by an Aspergillus infection. In other embodiments, the methods may be used to determine whether a subject has suffered from aspergillosis previously, or to predict whether a subject is likely to develop aspergillosis. In certain embodiments, a subject is suspected of suffering from aspergillosis due to the presence of one or more symptoms associated with aspergillosis. In certain embodiments, a subject has been previously diagnosed with aspergillosis.
[0067] In certain embodiments of the treatment methods provided herein, the steps of detecting Aspergillus protease are repeated at various timepoints following administration of a therapeutic agent or an increased dosage of a therapeutic agent. In this manner, the diagnostic methods disclosed herein can be used to monitor the efficacy of the therapeutic agent and/or to make adjustments to dosage. In certain embodiments, the steps of detecting Aspergillus protease are repeated at specific intervals until a specific therapeutic endpoint is reached, and in certain embodiments this therapeutic endpoint is the absence of detectable Aspergillus protease in a biological sample from the subject.
[0068] In certain embodiments of the methods and kits disclosed herein, a sample to be evaluated for the presence of Asp f2 is a biological sample, including but not limited to BALF, blood, serum, urine, saliva, or any other suitable biological sample that may contain Asp f2 in the presence of aspergillosis.
[0069] In certain embodiments of the kits provided herein, the kits comprise a protease substrate and instructions for carrying out the methods disclosed herein. In certain embodiments, the kits further comprise one or more Aspergillus protease-specific antibodies or antigen binding fragments thereof, and in certain of these embodiments the Aspergillus protease-specific antibodies or antigen binding fragments thereof are Asp f2-specific antibodies or antigen binding fragments thereof as disclosed herein. In certain embodiments, the kits comprise one or more Aspergillus protease substrate-specific antibodies or antigen binding fragments thereof, and in certain of these embodiments the Aspergillus protease substrate-specific antibodies or antigen binding fragments thereof are Asp f2-substrate specific antibodies or antigen binding fragments thereof as disclosed herein. In certain embodiments, the kits comprise a double affinity matrix as described herein. In certain embodiments, the kits comprise an enrichment matrix and instructions for use thereof. In certain of these embodiments, the enrichment matrix may be pre-loaded onto a solid column.
[0070] An example of an embodiment using an enrichment matrix is set forth in Example 1 below. 125 bronchoalveolar lavage fluid (BALF) samples from patients suspected of having an aspergillosis infection or another form of pulmonary mycosis were tested for the presence of Asp f2 using a bead-based assay. The Asp f2 antibody 5D7A1 was immobilized on beads, then contacted with the BALF samples. The presence of immobilized Asp f2-antibody complex on the beads was detected by contacting the beads with the novel fluorogenic peptide FluHSA2. The results of this experiment demonstrated that Asp f2 detection using the bead-based assay correlated with the clinical diagnosis of invasive aspergillosis, and confirmed that the presently disclosed methods are robust, inexpensive, and can be executed in a short timeframe (e.g., a few hours or less).
[0071] Provided herein in certain embodiments are detection and diagnosis methods that utilize the bead-based assay set forth in Example 1 or variations thereof (for example, using different fluorophores or protease substrates), as well as kits for carrying out such methods. In certain embodiments, these methods comprise obtaining a sample suspected of containing Asp f2 (e.g., a biological sample from a subject suspected of having aspergillosis due to Aspergillus infection), contacting the sample with an enrichment matrix comprising 5D7A1 or an Asp f2-binding fragment thereof immobilized on beads, contacting the enrichment matrix with the protease substrate such as FluHSA2, and measuring fluorescence at Excitation (Ex) 485 nm/Emission (Em) 535 nm. Also provided herein are treatment methods further comprising administering one or more aspergillosis therapeutic agents, or an increased dosage of a previously administered therapeutic agent, to a subject determined to have aspergillosis due to Aspergillus infection. In certain of these embodiments, the therapeutic agent may be selected from one or more antifungal medicines including, but not limited to, voriconazole, amphotericin B, echinocandins, and/or itraconazole, or immunosuppressive drugs. In other embodiments, the subject may be treated by undergoing surgery.
[0072] In certain embodiments of the kits provided herein, the kits comprise one or more components of the bead-based assay described in Example 1. In certain of these embodiments, the kits comprise an enrichment matrix comprised of one or more immunoaffinity beads to which at least one Aspergillus protease-specific antibody or antigen binding fragment thereof is bound and at least one protease substrate, wherein after interaction of the protease substrate with an Aspergillus protease a detectable fluorescent signal is produced. In certain embodiments, the immunoaffinity beads comprising Aspergillus protease-specific antibody may be lyophilized (freeze-dried) to allow for storage at 4.degree. C. for several months without loss of binding affinity. These ready-made beads reduce the time necessary to execute the assay provided by the kit. In some embodiments, the immunoaffinity beads may be lyophilized in the presence of 0.1 M ammonium bicarbonate buffer. In certain embodiments, the kit may include a control substrate that cannot be cleaved by the Aspergillus protease. In some embodiments, the kit may include an inhibitor of metalloprotease activity (e.g., the Zn metalloprotease inhibitor TPEN) as a control. In certain embodiments, the kit may include a positive control. For example, the kit may include a recombinant Asp f2 protein that may be used to test cleavage of the Aspergillus protease substrate provided in the kit. In certain embodiments, the Asp f2 protein may be glycosylated. In certain embodiments, the recombinant Asp f2 protein may be in an inactive form. In certain embodiments, the recombinant Asp f2 protein may comprise a fusion protein that may be activated after cleavage of an inactivating linker. For example, in certain embodiments, the Asp f2 protein may comprise SEQ ID NO: 6 as discussed in Example 2 or fragments or homologous proteins thereof. In certain of these embodiments, the inactive recombinant Asp f2 protein may be activated by cleaving the thioredoxin (Trx)-SMT3 N-terminal linker using ubiquitin-like specific protease (ULP1). In other embodiments, other enzymes may be used to cleave the inactivating linker such as thrombin or enterokinase. In certain embodiments, the kit may include recombinant Asp f2 protein in an active form. In certain embodiments, the recombinant Asp f2 protein used as a positive control in the kit comprises residues 32-310 of Asp f2 (see FIG. 12) or fragments or homologous proteins thereof.
[0073] According to some embodiments, the methods or kits provided herein may be used in a manual or automated format. In certain embodiments, the methods or kits may be used as a high-throughput detection system. In some embodiments, the methods or kits may be used as a high-throughput detection system for inhibitors of Aspergillus protease enzymatic activity. Such high-throughput detection systems are preferably automated for large-scale detection and testing, such as may be used in a diagnostic medical laboratory or in a manufacturing facility.
[0074] As used herein, a "subject" refers to a human or animal, including all mammals such as primates (particularly higher primates), sheep, dogs, rodents (e.g., mouse or rat), birds, guinea pigs, goats, pigs, cats, rabbits, and cows. In one embodiment, the subject is a penguin. In some embodiments, the subject is a human. In some embodiments, the subject is a patient.
[0075] As used herein the term "homologous" is used to refer to any amino acid or nucleotide sequence that displays at least 90% similarity with an amino acid or nucleotide sequence wherein the resulting protein still retains its desired functional properties.
[0076] As used herein, the term "significantly" or "significant" refers to a result that is statistically significant. In certain embodiments, statistical significance may be determined using any known test used to determine statistical significance. For example, a paired Student's t-test may be used to determine statistical significance. As described herein, a calculated p-value with a threshold of p<0.05 is considered statistically significant. In some embodiments, a calculated p-value with a threshold of p<0.0001 is considered statistically significant. In other embodiments, the term "significantly" or "significant" may be used to refer to a relative comparison between two or more experimental groups that are of interest. For example, if the results (i.e., change in detectable fluorescence signal or other measurable result) obtained from two experimental groups are found to be different by a factor of more than one, then this difference may be referred to as significant. In some embodiments, two or more groups may be significantly different if their experimental results are different by a factor of 2, 3, 4, 5, 6, 7, 8, 9, 10, or greater than 10.
[0077] The following examples are intended to illustrate various embodiments of the disclosure. As such, the specific embodiments discussed are not to be construed as limitations on the scope of the invention. It will be apparent to one skilled in the art that various equivalents, changes, and modifications may be made without departing from the scope of the disclosure, and it is understood that such equivalent embodiments are to be included herein. Further, all references cited in the disclosure are hereby incorporated by reference in their entirety, as if fully set forth herein.
[0078] The following examples are provided to better illustrate the claimed invention and are not to be interpreted as limiting the scope of the invention. To the extent that specific materials are mentioned, it is merely for purposes of illustration and is not intended to limit the invention. One skilled in the art may develop equivalent means or reactants without the exercise of inventive capacity and without departing from the scope of the invention.
EXAMPLES
Example 1
Detection of Proteolytic Activity of Asp f2 in Bronchoalveolar Lavage Fluid
[0079] A novel bead-based assay was developed for detecting Asp f2 proteolytic activity. An immobilization bead-based immuno-affinity matrix was prepared using the novel anti-Asp f2 monoclonal antibody 5D7A1 (FIGS. 1-3, SEQ ID NOS: 1-4). As demonstrated below, 5D7A1 binds the native, glycosylated form of Asp f2. Cyanogen bromide (CNBr)-activated sepharose beads were swelled in ice-cold HCl (1 mM). Beads (1.0 g) were incubated with 3.0 mg of 5D7A1 in Coupling Buffer (NaHCO.sub.3, 0.1 M and NaCl, 0.5 M, pH 8.3-8.5) and rotated for two hours at 22.degree. C. or for 16 hours at 4.degree. C. Beads were blocked with 0.1 M ethanolamine, pH 8.0, then washed with buffers of alternating pH (first with Coupling Buffer, pH 8.5, then with a low pH buffer (sodium acetate, 0.1 M, NaCl, 0.5 M, pH 4.0)). These dual washes were repeated four times. Lastly, beads were resuspended in ammonium bicarbonate (100 mM), lyophilized and stored at 4.degree. C.
[0080] 125 bronchoalveolar lavage fluid (BALF) samples from patients at City of Hope suspected of having an aspergillosis infection or another form of pulmonary mycosis were lyophilized to dryness and then re-dissolved in water in one tenth of the original BALF volume. Re-dissolved BALF (0.5 mL) was combined with 0.5 mL of Immunoprecipitation (IP) Binding Buffer (Tris base, 0.025 M, NaCl, 0.15M, NP-40, 1%, glycerol, 5%; pH 7.4). To immunoenrich active Asp f 2, antibody-coupled beads (200 .mu.L) were transferred into each tube of BALF/IP Binding Buffer or BALF-negative controls (0.9% NaCl: IP Binding Buffer, 1:1). Samples were rotated for 1 hour at 22.degree. C. followed by 16 hours rotation at 4.degree. C. The beads were washed twice with IP Binding buffer, three times with TBS (Tris base, 25 mM, NaCl, 150 mM, pH 7.2), once with NaCl (0.2 M), and finally three times with protease-free H.sub.2O. Beads were re-dissolved in H.sub.2O (2.times.100 .mu.L for duplicate or 3.times.100 .mu.L for triplicate measurements).
[0081] The enzymatic activity of immobilized Asp f2 was determined by cleavage of a novel fluorogenic peptide, FluHSA2 (FIGS. 4-5; [5-Fam]-TKCATESAVNRRPCFSALK-[DABCYL] SEQ ID NO: 5). FluHSA2 comprises the peptide of SEQ ID NO:5 conjugated to 5-Fam at the alpha-amino group of the threonine at the N-terminus of the peptide and to DABCYL at the epsilon amino group of the lysine side chain at the C-terminus of the peptide. Upon excitation, the DABCYL suppresses the fluorescence emission of the 5-Fam labeled peptides when the peptides are not cleaved and the fluorescent label and DABCYL remain close together. However, when the peptide is cleaved by Asp f2, the fluorescent label and DABCYL are separated and the fluorescent label emits light energy upon excitation. For each replicate (including negative controls), 100 .mu.L of the antibody coupled bead suspension was transferred into 0.5 mL reaction buffer (FluHSA2 (FIG. 4-5; SEQ ID NO: 5), 10 .mu.M, in sodium phosphate, 25 mM, NaCl, 100 mM, pH 7.2. Tubes were incubated in the dark for two hours at 37.degree. C. or for 16 hours at 22.degree. C. on a rotary shaker at 250 rpm. Fluorescence was measured at Excitation (Ex) 485 nm/Emission (Em) 535 nm in a black 96-well micro titer plate.
[0082] Results indicated that the FluHSA2 peptide was cleaved in samples from patients suspected of having aspergillosis, but remained uncleaved in samples from patients that had other mold infections or no infection (FIG. 6; p<0.0001). Thus, the novel antibody 5D7A1 is highly specific for the detection of aspergillosis. As such, this assay provides a useful tool for the clinical diagnosis of aspergillosis.
Example 2
Characterization of Zinc Metalloprotease Activity of Asp f2
[0083] Previous studies have not detected Asp f2 proteolytic activity, and Asp f2 has not previously been classified as a metalloprotease because it has very low (.about.15%) sequence identity to the metalloproteases of the Zincin-like family and lacks the HEXXH (SEQ ID NO: 37) signature that characterizes all metalloproteases (Amich et al., 2010). However, a sequence alignment of Asp f2 with deuterolysin from Aspergillus oryzae and the putative metalloprotease MEP20 from Aspergillus fumigatus shows that Asp f2 contains a motif similar to the HEXXH (SEQ ID NO: 37) motif, .sup.186HRLYH (SEQ ID NO: 36) (FIGS. 7 and 8, respectively). To investigate whether Asp f2 is a zinc metalloprotease, the bead-based assay described in Example 1 was used to evaluate Asp f2 protease activity in the presence of histidine modifiers and various chelators, as well as the activity of an Asp f2 with a mutation in the histidine residue of the HRLYH (SEQ ID NO: 36) motif. These experiments led to the unexpected finding that Asp f2 possesses zinc metalloproteolytic activity.
[0084] It was not possible to express proteolytically active Asp f2 directly in Escherichia coli bacteria because "active" Asp f2 lacks the signal peptide MAALLRLAVLLPLAAPLVATLPTSPVPIAAR (SEQ ID NO:39, residues 1-31) and kills the bacteria. Accordingly, active recombinant Asp f2 was generated using a fusion protein that expresses inactive recombinant Asp f2, which can then be activated after purification (FIGS. 14-16; SEQ ID NOS: 6 and 7). The fusion protein comprised an N-terminal linker (Trx-SMT3 N-terminal linker) region fused to Asp f2 residues 32-310 (codon optimized for expression in E. coli; FIG. 14) via a Ubiquitin-like-specific protease 1 (ULP1) cleavage site located between SMT3 and Asp f2. The Trx-SMT3 N-terminal linker contained the following from N- to C-terminus: thioredoxin (Trx)-glycine-serine-linker-His.sub.6-tag-linkers-another His.sub.6-tag-SMT3. Trx was added to improve solubility of the construct and a yeast SUMO (Smt3) was added to help enhance expression and promote solubility of Asp f2. Following purification using a His.sub.6-tag specific nickel-NTA column, Asp f2 was activated by cleaving the Trx-SMT3 N-terminal linker off using UPL1. This process yielded active recombinant Asp f2. The different cleavage sites in the linker region of the fusion protein construct can be used for cleavage by other enzymes (FIG. 14).
[0085] Diethylpyrocarbonate (DEPC) derivatization of histidines results in the chemical modification of histidines in proteins and is used to study the importance of histidyl residues in enzymes. Using the materials and methods described in Example 1, anti-Asp f2 (5D7A1)-bound beads were incubated with native or recombinant Asp f2 in the presence or absence of DEPC (30 .mu.M) and incubated for one hour at 22.degree. C. prior to reaction with FluHSA2. Samples were then incubated with 10 .mu.M of FluHSA2 for 16 hours at 22.degree. C. and fluorescence was measured as described above. Cleavage of FluHSA2 was detected in the presence or absence of DEPC to test the proteolytic activity of native and recombinant Asp f2. Results show that both native and recombinant Asp f2 cleave the FluHSA2 substrate, but are inactive in the presence of DEPC (FIG. 9). This result indicates that at least one histidine residue may be necessary for Asp f2 proteolytic activity, indicating that Asp f2 may be a metalloprotease.
[0086] To further investigate whether Asp f2 possesses zinc metalloprotease activity, a variety of protease inhibitors and zinc ions were tested to investigate their effects on native and recombinant Asp f2 activity (FIGS. 10 and 11, respectively). Anti-Asp f2 (5D7A1)-bound beads were incubated with native or recombinant Asp f2 in the presence or absence of 0.5 mM zinc chloride or 50 .mu.M of various inhibitors. Samples were incubated with FluHSA2 and fluorescence was measured as described above. Treatment with the zinc chelator, N,N,N',N'-tetrakis (2-pyridylmethyl) ethylenediamine (TPEN), the inhibitor of metallopeptidases, 1-10-phenanthroline, and the metal ion chelator, ethylenediaminetetraacetic acid (EDTA), suppressed both native and recombinant Asp f2 metalloprotease activity (FIGS. 10 and 11, respectively). Other protease inhibitors that are known to inhibit different families of proteases (e.g. cysteine peptidases (IAA), trypsin proteases (aprotinin and SBTI), aspartyl proteases (pepstatin A), and serine proteases (PMSF)), did not inhibit native or recombinant Asp f2 (FIGS. 10 and 11, respectively). These results further demonstrate that Asp f2 possesses zinc metalloprotease activity. Various Asp f2 histidine residues, including those located in the .sup.186HRLYH motif, were mutated to alanine (FIG. 12; H186A and H190A are located in the HRLYH motif), and mutant or wild-type Asp f2 was incubated with anti-Asp f2 (5D7A1)-bound beads as described above, followed by incubation with FluHSA2. Mutation of histidine 186 to alanine significantly decreased the proteolytic activity of Asp f2 compared with wild-type Asp f2 (FIG. 13).
Example 3
Characterization of Asp 2 Substrates
[0087] Cleavage specificity of the novel FluHSA peptide was tested using Asp f2 and another Aspergillus protein, Asp .English Pound.3. CNBr-activated sepharose beads were prepared as described in Example 1 and incubated with either the anti-Asp f2 monoclonal antibody 5D7A1 or an anti-Asp f3 rabbit polyclonal antibody. Anti-Asp f2-immobilized beads were incubated with native Asp f2 and anti-Asp f3-immobilized beads were incubated with native Asp f3. Samples were incubated with FluHSA2 (10 .mu.M) for 16 hours at 22.degree. C. Fluorescence was measured at Ex 485 nm/Em 535 nm in a black 96-well micro titer plate as described above. FluHSA2 was found to exhibit high specificity for cleavage by Asp f2 compared with Asp f3 (FIG. 17). This demonstrates the remarkable specificity that the FluHSA2 peptide has for the Aspergillus glycoprotein Asp f2.
[0088] The cleavage products of FluHSA2 after reaction with native Asp f2 were investigated by mass spectrometric analysis (MALDI spectra) (FIG. 5). Cleavage by Asp f2 occurred in the C-terminal linear tail section of the peptide resulting in at least 5 cleavage products (FIG. 5).
[0089] Peptides other than FluHSA2 were tested for cleavage by Asp f2. Anti-Asp f2 (5D7A1)-bound beads were incubated with native Asp f2, then incubated with either FluHSA2 (10 fluorescent labeled elastin protein (5-Fam-QXL520 labeled-elastin; SensoLyteGreen Elastase assay kit, #72178, Anaspec, Freemont, Calif., USA) or FITC conjugated collagen (#85111, Anaspec, Freemont, Calif., USA) (15 .mu.g/mL). Fluorescence was measured at Ex 485 nm/Em 535 nm in a black 96-well micro titer plate as described above. Results showed that native Asp f2 is also capable of degrading the human proteins collagen and elastin (FIG. 18). This demonstrates that active Asp f2 enriched on beads with 5D7A1 can cleave both fluorogenic elastin and collagen.
Example 4
Characterization of Asp f2 and Anti-Asp f2 Antibody Binding
[0090] The specificity of binding between Asp f2 and the anti-Asp f2 antibody 5D7A1 was further investigated using 5D7A1 and a variety of other antibodies known to bind Aspergillus Asp proteins other than Asp f2 (anti-Asp f1 (specific for Asp f1) and anti-Asp f3 (specific for Asp f3)) and proteases (4E17.1 (specific for botulinum neurotoxin serotype A heavy chain)) in the bead-based protease activity assay as described above. Polyclonal Rabbit IgG antibody was used as a non-specific IgG control. Antibodies were incubated with culture filtrate from A. fumigatus, and protease activity was detected using the 5-FAM-labeled elastin protein or FITC-conjugated collagen protein described above in Example 3. Fluorescence was measured at Ex 485 nm/Em 535 nm in a black 96-well micro titer plate as described above.
[0091] Results demonstrated that robust Asp f2 cleavage of the collagen and elastin substrates was detected in only those samples containing the anti-Asp f2 antibody 5D7A1 (FIG. 19, FITC-collagen substrate; FIG. 20, 5-Fam elastin substrate). Little to no activity was observed in samples containing antibodies that are non-Asp f2 specific antibodies (FIGS. 19 and 20). This assay demonstrates that Asp f2 and 5D7A1 bind each other specifically.
Example 5
Asp f2 Proteolytic Activity in Aspergillosis Patients
[0092] Most aspergillosis is caused by A. fumigatus, but there are other aspergilli that can cause the disease. Proteolytic activity of Asp f2 was investigated in BALF samples from aspergillosis patients for which Aspergillus species were identified as indicated by the clinical Microbiology lab at City of Hope. The different species identified were A. fumigatus, A. flavus, A. versicolor, A. niger, A. terreus, and other unidentified Aspergillus species. Results demonstrate that Asp f2 proteolytic activity may be useful for detecting other Aspergillus species as well (FIG. 21).
Example 6
Sensitivity, Specificity, and Cut-Off Values of Asp f2 Assay
[0093] A receiver operating characteristic (ROC) curve was generated to determine the sensitivity, specificity, and appropriate cut off values of the bead-based Asp f2 assay described herein (FIG. 25). The ROC analysis included clinical diagnostic data from a total of 144 patients, of which 27 had aspergillosis, 40 had other mold infections, and 77 were diagnosed negative for fungal infections. The ROC curve was used to determine the cutoff value for positive Asp f2 activity assay values. This cutoff value is .DELTA.RFU=8.8.times.10.sup.5, which corresponds to approximately 280 pmol/mL substrate (HSA2 peptide) cleaved by Asp f2 in 16 hours. The assay was found to have a specificity of 96.1% at a sensitivity of 77.8% for cases of aspergillosis versus a specificity of 72.7% at 57.5% sensitivity for all other mold infections, indicating that the assay is highly specific and sensitive for aspergillosis, and less specific and sensitive for other mold infections.
Example 7
Comparison of Asp f2 Activity Assay and Clinical Galactomannan (GM) Test Results
[0094] The Asp f2 activity assay disclosed herein was compared to clinical galactomannan (GM) test results. Clinical diagnosis was based on the EORTC/MSG criteria for probable and proven aspergillosis (DePauw 2008), including cytological, pathological, and microbiological examination of the patients by infectious disease physicians. The GM assay was performed by a clinical laboratory. A GM index cut off of >0.5 for patients positive with fungal infections was used in accordance with clinical practice. The GM assay had only 40% sensitivity in the sample group of City of Hope aspergillosis patients (FIG. 26A), while the Asp f2 activity assay achieved 77.8% sensitivity in the same group of patients (FIG. 26B). A cut-off value of >.DELTA.RFU=8.8.times.10.sup.5 (equivalent to 280 pmol/mL converted substrate in 16 hours) was used to determine aspergillosis positive samples.
Example 8
Generation of Recombinant Antibodies Based on 5D7A1 Variable Chain Sequences
[0095] A molecular cloning approach was used to generate single-chain variable fragments (scFvs) from the sequenced V.sub.L and V.sub.H chains of 5D7A1. Two types of scFvs were generated. In the first, the C-terminus of the V.sub.L chain is linked to the N-terminus of the V.sub.H chain via a glycine-serine linker (GS15). In the second, the sequential order of the V chains is reversed such that the C-terminus of the V.sub.H chain is linked to the N-terminus of the V.sub.L chain using the same GS15 linker.
[0096] Initial attempts at scFv expression resulted in insoluble products. To overcome this, scFvs were fused to a human Fc domain of IgG, and a human antibody signal peptide was added. The resultant DNA construct was cloned into a pEE12.4 vector (GS Gene Expression System; see WO06/111387) and expressed in Expi293F cells (derived from 293 cells, primary embryonic human kidney; Gibco, Life technologies) in Expi293 expression medium (Gibco, Life technologies). The resulting single chain monovalent antibody homologues were soluble and exhibited Asp f2 specific binding affinities. The DNA and amino acid sequences of the V.sub.L-GS15-V.sub.H-Fc scFv are set forth in FIGS. 27-28 and SEQ ID NOs: 20-21, and the sequences of the V.sub.H-GS15-V.sub.L-Fc scFv are set forth in FIGS. 29-30 and SEQ ID NOs: 22-23.
Example 9
Immunoaffinity Analysis of Recombinant scFvs
[0097] Western blots were performed to compare the binding specificity of 5D7A1 and the recombinant scFvs generated in Example 8. 5D7A1 and V.sub.H-GS15-V.sub.L-Fc scFv both bound native Asp f2 independent of the different glycosylation states responsible for the different gel migration patterns in lanes 1 and 2, and weakly bound recombinant SMT3-Asp f2 (FIG. 31). Unglycosylated Asp f2 showed a stronger signal for 5D7A1 than for the scFv.
[0098] As stated above, the foregoing are merely intended to illustrate the various embodiments of the present invention. As such, the specific modifications discussed above are not to be construed as limitations on the scope of the invention. It will be apparent to one skilled in the art that various equivalents, changes, and modifications may be made without departing from the scope of the invention, and it is understood that such equivalent embodiments are to be included herein. All references cited herein are incorporated by reference as if fully set forth herein.
REFERENCES
[0099] 1. Amich Eukaryot Cell 9:424-437 (2010)
[0100] 2. DePauw Clin Infect Dis 46:1813-1821(2008)
Sequence CWU
1
1
391139PRTArtificial SequencemAb 5D7A1 VH domain 1Leu Pro Glu Phe Glu Val
Gln Leu Glu Glu Ser Gly Ala Glu Leu Ala 1 5
10 15 Arg Pro Gly Ala Ser Val Lys Met Ser Cys Lys
Ala Ser Gly Tyr Thr 20 25
30 Phe Thr Thr Tyr Thr Met His Trp Val Lys Gln Arg Pro Gly Gln
Gly 35 40 45 Leu
Glu Trp Ile Gly Tyr Ile Asn Pro Ser Ser Gly Tyr Thr Asn Tyr 50
55 60 Asn Gln Lys Phe Lys Asp
Lys Ala Thr Leu Thr Ala Asn Lys Ser Ser 65 70
75 80 Ser Ile Gly Tyr Met Gln Leu Ser Ser Leu Thr
Ser Glu Asp Ser Ala 85 90
95 Leu Tyr Tyr Cys Leu Arg Arg Pro Tyr Arg Ser His Gly Gly Trp Phe
100 105 110 Phe Asp
Val Trp Gly Ala Gly Thr Thr Val Thr Val Ser Ser Ala Lys 115
120 125 Thr Thr Pro Pro Ser Val Tyr
Arg Ser Ser Lys 130 135
2145PRTArtificial SequencemAb 5D7A1 VL domain 2Met Glu Ser Gln Thr Gln
Val Phe Val Tyr Met Leu Leu Trp Leu Ser 1 5
10 15 Gly Val Asp Gly Asp Ile Val Met Thr Gln Ser
Gln Lys Phe Met Ser 20 25
30 Thr Ser Val Gly Asp Arg Val Ser Val Thr Cys Lys Ala Ser Gln
Asn 35 40 45 Val
Glu Thr Asn Val Ala Trp Tyr Gln Gln Lys Leu Gly Gln Ser Pro 50
55 60 Lys Ala Leu Ile Tyr Ser
Ala Ser Phe Arg Thr Ser Gly Val Pro Asp 65 70
75 80 Arg Phe Thr Gly Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr Ile Asn 85 90
95 Asn Val Gln Ser Glu Asp Leu Ala Glu Tyr Phe Cys Gln Gln Tyr Asn
100 105 110 Thr Tyr
Pro Leu Thr Phe Gly Ala Gly Thr Lys Leu Glu Leu Lys Arg 115
120 125 Ala Asp Ala Ala Pro Thr Val
Ser Ile Ser His His Pro Val Leu Glu 130 135
140 Ser 145 3417DNAArtificial SequencemAb 5D7A1 VH
domain 3cttccggaat tcgaggtcca gctggaggag tcaggggctg aattggcaag acctggggcc
60tcagtgaaga tgtcctgcaa ggcttctggc tacaccttta caacctacac gatgcactgg
120gtaaaacaga ggcctggaca gggtctggaa tggattggat acattaatcc tagtagtggt
180tatactaatt acaatcaaaa gttcaaagac aaggccacat tgactgcaaa caaatcctcc
240agtatcggct acatgcagct gagcagccta acatctgagg attctgcact ttattattgt
300ttaagaaggc cttataggtc ccacgggggc tggttcttcg atgtctgggg cgcagggacc
360acggtcaccg tctcctcagc caaaacgaca cccccatctg tctatagatc ttccaag
4174435DNAArtificial SequencemAb 5D7A1 VL domain 4atggagtcac agactcaggt
ctttgtatac atgttgctgt ggttgtctgg tgttgatgga 60gacattgtga tgacccagtc
tcaaaaattc atgtccacat cagtaggaga cagggtcagc 120gtcacctgca aggccagtca
gaatgtagag actaatgtag cctggtatca acagaaacta 180gggcaatctc ctaaagcact
gatttactcg gcatccttcc ggaccagtgg agtccctgat 240cgcttcacag gcagtggatc
tgggacagat ttcactctca ccatcaacaa tgtgcagtct 300gaagacttgg cagagtattt
ctgtcagcaa tataacacct atccgctcac gttcggtgct 360gggaccaagc tggagctgaa
acgggctgat gctgcaccaa ctgtatccat ctcccaccat 420ccagttctag aaagc
435519PRTArtificial
SequenceFluHSA2 peptide 5Thr Lys Cys Ala Thr Glu Ser Ala Val Asn Arg Arg
Pro Cys Phe Ser 1 5 10
15 Ala Leu Lys 6549PRTArtificial SequenceFusion protein construct
Trx-SMT3-Asp f2 6Met Ser Asp Lys Ile Ile His Leu Thr Asp Asp Ser Phe Asp
Thr Asp 1 5 10 15
Val Leu Lys Ala Asp Gly Ala Ile Leu Val Asp Phe Trp Ala Glu Trp
20 25 30 Cys Gly Pro Cys Lys
Met Ile Ala Pro Ile Leu Asp Glu Ile Ala Asp 35
40 45 Glu Tyr Gln Gly Lys Leu Thr Val Ala
Lys Leu Asn Ile Asp Gln Asn 50 55
60 Pro Gly Thr Ala Pro Lys Tyr Gly Ile Arg Gly Ile Pro
Thr Leu Leu 65 70 75
80 Leu Phe Lys Asn Gly Glu Val Ala Ala Thr Lys Val Gly Ala Leu Ser
85 90 95 Lys Gly Gln Leu
Lys Glu Phe Leu Asp Ala Asn Leu Ala Gly Ser Gly 100
105 110 Ser Gly His Met His His His His His
His Ser Ser Gly Leu Val Pro 115 120
125 Arg Gly Ser Gly Met Lys Glu Thr Ala Ala Ala Lys Phe Glu
Arg Gln 130 135 140
His Met Asp Ser Pro Asp Leu Gly Thr Asp Asp Asp Asp Lys Ala Met 145
150 155 160 Gly Gly His His His
His His His Gly Gly Gly Gly Met Ser Asp Ser 165
170 175 Glu Val Asn Gln Glu Ala Lys Pro Glu Val
Lys Pro Glu Val Lys Pro 180 185
190 Glu Thr His Ile Asn Leu Lys Val Ser Asp Gly Ser Ser Glu Ile
Phe 195 200 205 Phe
Lys Ile Lys Lys Thr Thr Pro Leu Arg Arg Leu Met Glu Ala Phe 210
215 220 Ala Lys Arg Gln Gly Lys
Glu Met Asp Ser Leu Arg Phe Leu Tyr Asp 225 230
235 240 Gly Ile Arg Ile Gln Ala Asp Gln Thr Pro Glu
Asp Leu Asp Met Glu 245 250
255 Asp Asn Asp Ile Ile Glu Ala His Arg Glu Gln Ile Gly Gly Ala Thr
260 265 270 Pro His
Glu Pro Val Phe Phe Ser Trp Asp Ala Gly Ala Val Thr Ser 275
280 285 Phe Pro Ile His Ser Ser Cys
Asn Ala Thr Gln Arg Arg Gln Ile Glu 290 295
300 Ala Gly Leu Asn Glu Ala Val Glu Leu Ala Arg His
Ala Lys Ala His 305 310 315
320 Ile Leu Arg Trp Gly Asn Glu Ser Glu Ile Tyr Arg Lys Tyr Phe Gly
325 330 335 Asn Arg Pro
Thr Met Glu Ala Val Gly Ala Tyr Asp Val Ile Val Asn 340
345 350 Gly Asp Lys Ala Asn Val Leu Phe
Arg Cys Asp Asn Pro Asp Gly Asn 355 360
365 Cys Ala Leu Glu Gly Trp Gly Gly His Trp Arg Gly Ala
Asn Ala Thr 370 375 380
Ser Glu Thr Val Ile Cys Asp Arg Ser Tyr Thr Thr Arg Arg Trp Leu 385
390 395 400 Val Ser Met Cys
Ser Gln Gly Tyr Thr Val Ala Gly Ser Glu Thr Asn 405
410 415 Thr Phe Trp Ala Ser Asp Leu Met His
Arg Leu Tyr His Val Pro Ala 420 425
430 Val Gly Gln Gly Trp Val Asp His Phe Ala Asp Gly Tyr Asp
Glu Val 435 440 445
Ile Ala Leu Ala Lys Ser Asn Gly Thr Glu Ser Thr His Asp Ser Glu 450
455 460 Ala Leu Gln Tyr Phe
Ala Leu Glu Ala Tyr Ala Phe Asp Ile Ala Ala 465 470
475 480 Pro Gly Val Gly Cys Ala Gly Glu Ser His
Gly Pro Asp Gln Gly His 485 490
495 Asp Thr Gly Ser Ala Ser Ala Pro Ala Ser Thr Ser Thr Ser Ser
Ser 500 505 510 Ser
Ser Gly Ser Gly Ser Gly Ala Thr Thr Thr Pro Thr Asp Ser Pro 515
520 525 Ser Ala Thr Ile Asp Val
Pro Ser Asn Cys His Thr His Glu Gly Gly 530 535
540 Gln Leu His Cys Thr 545
71650DNAArtificial SequenceFusion protein construct Trx-SMT3-Asp f2
7atgagcgata aaattattca cctgactgac gacagttttg acacggatgt actcaaagcg
60gacggggcga tcctcgtcga tttctgggca gagtggtgcg gtccgtgcaa aatgatcgcc
120ccgattctgg atgaaatcgc tgacgaatat cagggcaaac tgaccgttgc aaaactgaac
180atcgatcaaa accctggcac tgcgccgaaa tatggcatcc gtggtatccc gactctgctg
240ctgttcaaaa acggtgaagt ggcggcaacc aaagtgggtg cactgtctaa aggtcagttg
300aaagagttcc tcgacgctaa cctggccggt tctggttctg gccatatgca ccatcatcat
360catcattctt ctggtctggt gccacgcggt tctggtatga aagaaaccgc tgctgctaaa
420ttcgaacgcc agcacatgga cagcccagat ctgggtaccg acgacgacga caaggccatg
480ggaggtcatc atcatcacca tcatggtggt ggcggtatga gcgatagcga agttaatcaa
540gaagcaaaac cggaagttaa acctgaagtg aaaccggaaa cccatattaa cctgaaagtt
600agtgatggca gcagcgagat cttctttaaa atcaaaaaaa ccacaccgct gcgtcgtctg
660atggaagcat ttgcaaaacg tcagggtaaa gaaatggata gcctgcgttt tctgtatgat
720ggtattcgta ttcaggcaga tcagacaccg gaagatctgg atatggaaga taacgatatt
780atcgaagcac atcgtgagca gattggtggt gcaacaccgc atgaaccggt gttttttagc
840tgggatgccg gtgcagttac cagctttccg attcatagca gctgtaatgc aacccagcgt
900cgccagattg aagcaggtct gaatgaagca gttgaactgg cacgtcatgc aaaagcacat
960attctgcgtt ggggtaatga aagcgaaatc tatcgtaaat actttggcaa tcgtccgaca
1020atggaagccg ttggtgcata tgatgttatt gtgaatggtg ataaagccaa cgttctgttt
1080cgttgtgata atccggatgg taattgtgca ctggaaggtt ggggtggtca ttggcgtggt
1140gcaaatgcga ccagcgaaac cgttatttgt gatcgtagct ataccacccg tcgttggctg
1200gttagcatgt gtagccaggg ttataccgtt gcaggtagcg aaaccaatac cttttgggca
1260agcgatctga tgcatcgtct gtatcatgtt ccggcagttg gtcagggttg ggttgatcat
1320tttgcagatg gctatgatga agttattgca ctggcaaaaa gcaatggcac cgaaagcacc
1380catgatagtg aagcactgca gtattttgcc ctggaagcat atgcctttga tattgcagca
1440ccgggtgttg gttgtgccgg tgaaagtcat ggtccggatc agggtcatga taccggtagc
1500gcaagcgcac cggcaagcac cagcaccagc tcaagcagca gcggtagcgg ttcaggtgca
1560accaccaccc cgaccgatag cccgagcgca accattgatg ttccgagcaa ttgtcatacc
1620catgaaggtg gtcagctgca ttgtacctaa
165088PRTArtificial SequencemAb 5D7A1 VH CDR1 8Gly Tyr Thr Phe Thr Thr
Tyr Thr 1 5 924DNAArtificial SequencemAb
5D7A1 VH CDR1 9ggctacacct ttacaaccta cacg
24106PRTArtificial SequencemAb 5D7A1 VL CDR1 10Gln Asn Val Glu
Thr Asn 1 5 1118DNAArtificial SequencemAb 5D7A1 VL
CDR1 11cagaatgtag agactaat
18128PRTArtificial SequencemAb 5D7A1 VH CDR2 12Ile Asn Pro Ser Ser Gly
Tyr Thr 1 5 1324DNAArtificial SequencemAb
5D7A1 VH CDR2 13attaatccta gtagtggtta tact
24143PRTArtificial SequencemAb 5D7A1 VL CDR2 14Ser Ala Ser 1
159DNAArtificial SequencemAb 5D7A1 VL CDR2 15tcggcatcc
91615PRTArtificial
SequencemAb 5D7A1 VH CDR3 16Leu Arg Arg Pro Tyr Arg Ser His Gly Gly Trp
Phe Phe Asp Val 1 5 10
15 1745DNAArtificial SequencemAb 5D7A1 VH CDR3 17ttaagaaggc cttataggtc
ccacgggggc tggttcttcg atgtc 45189PRTArtificial
SequencemAb 5D7A1 VL CDR3 18Gln Gln Tyr Asn Thr Tyr Pro Leu Thr 1
5 1930DNAArtificial SequencemAb 5D7A1 VL CDR3
19tcagcaatat aacacctatc cgctcacgtt
3020506PRTArtificial SequenceVL-GS15-VH-Fc scFv 20Met Glu Thr Asp Thr Leu
Leu Leu Trp Val Leu Leu Leu Trp Val Pro 1 5
10 15 Gly Ser Thr Gly Asp Ile Val Met Thr Gln Ser
Gln Lys Phe Met Ser 20 25
30 Thr Ser Val Gly Asp Arg Val Ser Val Thr Cys Lys Ala Ser Gln
Asn 35 40 45 Val
Glu Thr Asn Val Ala Trp Tyr Gln Gln Lys Leu Gly Gln Ser Pro 50
55 60 Lys Ala Leu Ile Tyr Ser
Ala Ser Phe Arg Thr Ser Gly Val Pro Asp 65 70
75 80 Arg Phe Thr Gly Ser Gly Ser Gly Thr Asp Phe
Thr Leu Thr Ile Asn 85 90
95 Asn Val Gln Ser Glu Asp Leu Ala Glu Tyr Phe Cys Gln Gln Tyr Asn
100 105 110 Thr Tyr
Pro Leu Thr Phe Gly Ala Gly Thr Lys Leu Glu Leu Lys Ala 115
120 125 Met Ala Leu Gln Ala Ser Gly
Gly Gly Gly Ser Gly Gly Gly Gly Ser 130 135
140 Gly Gly Gly Gly Ser Ala Ser Glu Val Gln Leu Glu
Glu Ser Gly Ala 145 150 155
160 Glu Leu Ala Arg Pro Gly Ala Ser Val Lys Met Ser Cys Lys Ala Ser
165 170 175 Gly Tyr Thr
Phe Thr Thr Tyr Thr Met His Trp Val Lys Gln Arg Pro 180
185 190 Gly Gln Gly Leu Glu Trp Ile Gly
Tyr Ile Asn Pro Ser Ser Gly Tyr 195 200
205 Thr Asn Tyr Asn Gln Lys Phe Lys Asp Lys Ala Thr Leu
Thr Ala Asn 210 215 220
Lys Ser Ser Ser Ile Gly Tyr Met Gln Leu Ser Ser Leu Thr Ser Glu 225
230 235 240 Asp Ser Ala Leu
Tyr Tyr Cys Leu Arg Arg Pro Tyr Arg Ser His Gly 245
250 255 Gly Trp Phe Phe Asp Val Trp Gly Ala
Gly Thr Thr Val Thr Val Ser 260 265
270 Ser Val Glu Pro Lys Ser Cys Asp Lys Thr His Thr Cys Pro
Pro Cys 275 280 285
Pro Ala Pro Glu Leu Leu Gly Gly Pro Ser Val Phe Leu Phe Pro Pro 290
295 300 Lys Pro Lys Asp Thr
Leu Met Ile Ser Arg Thr Pro Glu Val Thr Cys 305 310
315 320 Val Val Val Asp Val Ser His Glu Asp Pro
Glu Val Lys Phe Asn Trp 325 330
335 Tyr Val Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg
Glu 340 345 350 Glu
Gln Tyr Asn Ser Thr Tyr Arg Val Val Ser Val Leu Thr Val Leu 355
360 365 His Gln Asp Trp Leu Asn
Gly Lys Glu Tyr Lys Cys Lys Val Ser Asn 370 375
380 Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile
Ser Lys Ala Lys Gly 385 390 395
400 Gln Pro Arg Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu
405 410 415 Leu Thr
Lys Asn Gln Val Ser Leu Thr Cys Leu Val Lys Gly Phe Tyr 420
425 430 Pro Ser Asp Ile Ala Val Glu
Trp Glu Ser Asn Gly Gln Pro Glu Asn 435 440
445 Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser Asp
Gly Ser Phe Phe 450 455 460
Leu Tyr Ser Lys Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn 465
470 475 480 Val Phe Ser
Cys Ser Val Met His Glu Ala Leu His Asn His Tyr Thr 485
490 495 Gln Lys Ser Leu Ser Leu Ser Pro
Gly Lys 500 505 211518DNAArtificial
SequenceVL-GS15-VH-Fc scFv within pEE12.4 vector 21atggaaaccg acaccctgct
gctgtgggtg ctgctgctgt gggtgccagg atctacaggc 60gacattgtga tgacccagtc
tcaaaaattc atgtccacat cagtaggaga cagggtcagc 120gtcacctgca aggccagtca
gaatgtagag actaatgtag cctggtatca acagaaacta 180gggcaatctc ctaaagcact
gatttactcg gcatccttcc ggaccagtgg agtccctgat 240cgcttcacag gcagtggatc
tgggacagat ttcactctca ccatcaacaa tgtgcagtct 300gaagacttgg cagagtattt
ctgtcagcaa tataacacct atccgctcac gttcggtgct 360gggaccaagc tggagctgaa
agccatggct ctgcaggcta gtggtggtgg tggttctggt 420ggtggtggtt ctggtggtgg
tggttctgct agcgaggtcc agctggagga gtcaggggct 480gaattggcaa gacctggggc
ctcagtgaag atgtcctgca aggcttctgg ctacaccttt 540acaacctaca cgatgcactg
ggtaaaacag aggcctggac agggtctgga atggattgga 600tacattaatc ctagtagtgg
ttatactaat tacaatcaaa agttcaaaga caaggccaca 660ttgactgcaa acaaatcctc
cagtatcggc tacatgcagc tgagcagcct aacatctgag 720gattctgcac tttattattg
tttaagaagg ccttataggt cccacggggg ctggttcttc 780gatgtctggg gcgcagggac
cacggtcacc gtctcctctg tagaacccaa atcttgcgac 840aaaactcaca catgcccacc
gtgcccagca cctgaactcc tggggggacc gtcagtcttc 900ctcttccccc caaaacccaa
ggacaccctc atgatctccc ggacccctga ggtcacatgc 960gtggtggtgg acgtgagcca
cgaagaccct gaggtcaagt tcaactggta cgtggacggc 1020gtggaggtgc ataatgccaa
gacaaagccg cgggaggagc agtacaacag cacgtaccgt 1080gtggtcagcg tcctcaccgt
cctgcaccag gactggctga atggcaagga gtacaagtgc 1140aaggtctcca acaaagccct
cccagccccc atcgagaaaa ccatctccaa agccaaaggg 1200cagccccgag aaccacaggt
gtacaccctg ccaccatcac gagatgagct gaccaagaac 1260caggtcagcc tgacctgcct
ggtcaaaggc ttctatccca gcgacatcgc cgtggagtgg 1320gagagcaatg ggcagccgga
gaacaactac aagaccacgc ctcccgtgct ggactccgac 1380ggctccttct tcctctacag
caagctcacc gtggacaaga gcaggtggca gcaggggaac 1440gtcttctcat gctccgtgat
gcatgaggct ctgcacaacc actacacgca gaagagcctc 1500tctctgtctc ccgggaaa
151822506PRTArtificial
SequenceVH-GS15-VL-Fc scFv 22Met Glu Thr Asp Thr Leu Leu Leu Trp Val Leu
Leu Leu Trp Val Pro 1 5 10
15 Gly Ser Thr Gly Glu Val Gln Leu Glu Glu Ser Gly Ala Glu Leu Ala
20 25 30 Arg Pro
Gly Ala Ser Val Lys Met Ser Cys Lys Ala Ser Gly Tyr Thr 35
40 45 Phe Thr Thr Tyr Thr Met His
Trp Val Lys Gln Arg Pro Gly Gln Gly 50 55
60 Leu Glu Trp Ile Gly Tyr Ile Asn Pro Ser Ser Gly
Tyr Thr Asn Tyr 65 70 75
80 Asn Gln Lys Phe Lys Asp Lys Ala Thr Leu Thr Ala Asn Lys Ser Ser
85 90 95 Ser Ile Gly
Tyr Met Gln Leu Ser Ser Leu Thr Ser Glu Asp Ser Ala 100
105 110 Leu Tyr Tyr Cys Leu Arg Arg Pro
Tyr Arg Ser His Gly Gly Trp Phe 115 120
125 Phe Asp Val Trp Gly Ala Gly Thr Thr Val Thr Val Ser
Ser Ala Met 130 135 140
Ala Leu Gln Ala Ser Gly Gly Gly Gly Ser Gly Gly Gly Gly Ser Gly 145
150 155 160 Gly Gly Gly Ser
Ala Ser Asp Ile Val Met Thr Gln Ser Gln Lys Phe 165
170 175 Met Ser Thr Ser Val Gly Asp Arg Val
Ser Val Thr Cys Lys Ala Ser 180 185
190 Gln Asn Val Glu Thr Asn Val Ala Trp Tyr Gln Gln Lys Leu
Gly Gln 195 200 205
Ser Pro Lys Ala Leu Ile Tyr Ser Ala Ser Phe Arg Thr Ser Gly Val 210
215 220 Pro Asp Arg Phe Thr
Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr 225 230
235 240 Ile Asn Asn Val Gln Ser Glu Asp Leu Ala
Glu Tyr Phe Cys Gln Gln 245 250
255 Tyr Asn Thr Tyr Pro Leu Thr Phe Gly Ala Gly Thr Lys Leu Glu
Leu 260 265 270 Lys
Val Glu Pro Lys Ser Cys Asp Lys Thr His Thr Cys Pro Pro Cys 275
280 285 Pro Ala Pro Glu Leu Leu
Gly Gly Pro Ser Val Phe Leu Phe Pro Pro 290 295
300 Lys Pro Lys Asp Thr Leu Met Ile Ser Arg Thr
Pro Glu Val Thr Cys 305 310 315
320 Val Val Val Asp Val Ser His Glu Asp Pro Glu Val Lys Phe Asn Trp
325 330 335 Tyr Val
Asp Gly Val Glu Val His Asn Ala Lys Thr Lys Pro Arg Glu 340
345 350 Glu Gln Tyr Asn Ser Thr Tyr
Arg Val Val Ser Val Leu Thr Val Leu 355 360
365 His Gln Asp Trp Leu Asn Gly Lys Glu Tyr Lys Cys
Lys Val Ser Asn 370 375 380
Lys Ala Leu Pro Ala Pro Ile Glu Lys Thr Ile Ser Lys Ala Lys Gly 385
390 395 400 Gln Pro Arg
Glu Pro Gln Val Tyr Thr Leu Pro Pro Ser Arg Asp Glu 405
410 415 Leu Thr Lys Asn Gln Val Ser Leu
Thr Cys Leu Val Lys Gly Phe Tyr 420 425
430 Pro Ser Asp Ile Ala Val Glu Trp Glu Ser Asn Gly Gln
Pro Glu Asn 435 440 445
Asn Tyr Lys Thr Thr Pro Pro Val Leu Asp Ser Asp Gly Ser Phe Phe 450
455 460 Leu Tyr Ser Lys
Leu Thr Val Asp Lys Ser Arg Trp Gln Gln Gly Asn 465 470
475 480 Val Phe Ser Cys Ser Val Met His Glu
Ala Leu His Asn His Tyr Thr 485 490
495 Gln Lys Ser Leu Ser Leu Ser Pro Gly Lys 500
505 231518DNAArtificial SequenceVL-GS15-VH-Fc scFv
within pEE12.4 vector 23atggaaaccg acaccctgct gctgtgggtg ctgctgctgt
gggtgccagg atctacaggc 60gaggtccagc tggaggagtc aggggctgaa ttggcaagac
ctggggcctc agtgaagatg 120tcctgcaagg cttctggcta cacctttaca acctacacga
tgcactgggt aaaacagagg 180cctggacagg gtctggaatg gattggatac attaatccta
gtagtggtta tactaattac 240aatcaaaagt tcaaagacaa ggccacattg actgcaaaca
aatcctccag tatcggctac 300atgcagctga gcagcctaac atctgaggat tctgcacttt
attattgttt aagaaggcct 360tataggtccc acgggggctg gttcttcgat gtctggggcg
cagggaccac ggtcaccgtc 420tcctcagcca tggctctgca ggctagtggt ggtggtggtt
ctggtggtgg tggttctggt 480ggtggtggtt ctgctagcga cattgtgatg acccagtctc
aaaaattcat gtccacatca 540gtaggagaca gggtcagcgt cacctgcaag gccagtcaga
atgtagagac taatgtagcc 600tggtatcaac agaaactagg gcaatctcct aaagcactga
tttactcggc atccttccgg 660accagtggag tccctgatcg cttcacaggc agtggatctg
ggacagattt cactctcacc 720atcaacaatg tgcagtctga agacttggca gagtatttct
gtcagcaata taacacctat 780ccgctcacgt tcggtgctgg gaccaagctg gagctgaaag
tagaacccaa atcttgcgac 840aaaactcaca catgcccacc gtgcccagca cctgaactcc
tggggggacc gtcagtcttc 900ctcttccccc caaaacccaa ggacaccctc atgatctccc
ggacccctga ggtcacatgc 960gtggtggtgg acgtgagcca cgaagaccct gaggtcaagt
tcaactggta cgtggacggc 1020gtggaggtgc ataatgccaa gacaaagccg cgggaggagc
agtacaacag cacgtaccgt 1080gtggtcagcg tcctcaccgt cctgcaccag gactggctga
atggcaagga gtacaagtgc 1140aaggtctcca acaaagccct cccagccccc atcgagaaaa
ccatctccaa agccaaaggg 1200cagccccgag aaccacaggt gtacaccctg cccccatccc
gggatgagct gaccaagaac 1260caggtcagcc tgacctgcct ggtcaaaggc ttctatccca
gcgacatcgc cgtggagtgg 1320gagagcaatg ggcagccgga gaacaactac aagaccacgc
ctcccgtgct ggactccgac 1380ggctccttct tcctctacag caagctcacc gtggacaaga
gcaggtggca gcaggggaac 1440gtcttctcat gctccgtgat gcatgaggct ctgcacaacc
actacacgca gaagagcctc 1500tctctgtctc ccgggaaa
1518241100DNAArtificial Sequence5D7A1 Antibody and
surrounding vector sequence 24tgattacgcc aagcttggta ccgagctcgg atccactagt
aacggccgcc agtgtgctgg 60aattcggctt cttccggaat tcgaggtcca gctggaggag
tcaggggctg aattggcaag 120acctggggcc tcagtgaaga tgtcctgcaa ggcttctggc
tacaccttta caacctacac 180gatgcactgg gtaaaacaga ggcctggaca gggtctggaa
tggattggat acattaatcc 240tagtagtggt tatactaatt acaatcaaaa gttcaaagac
aaggccacat tgactgcaaa 300caaatcctcc agtatcggct acatgcagct gagcagccta
acatctgagg attctgcact 360ttattattgt ttaagaaggc cttataggtc ccacgggggc
tggttcttcg atgtctgggg 420cgcagggacc acggtcaccg tctcctcagc caaaacgaca
cccccatctg tctatagatc 480ttccaagccg aattctgcag ctctagatgc atgctcgagc
ggccgccagt gtgatggata 540tctgcagaat tcggcttgat atccaccatg gagtcacaga
ctcaggtctt tgtatacatg 600ttgctgtggt tgtctggtgt tgatggagac attgtgatga
cccagtctca aaaattcatg 660tccacatcag taggagacag ggtcagcgtc acctgcaagg
ccagtcagaa tgtagagact 720aatgtagcct ggtatcaaca gaaactaggg caatctccta
aagcactgat ttactcggca 780tccttccgga ccagtggagt ccctgatcgc ttcacaggca
gtggatctgg gacagatttc 840actctcacca tcaacaatgt gcagtctgaa gacttggcag
agtatttctg tcagcaatat 900aacacctatc cgctcacgtt cggtgctggg accaagctgg
agctgaaacg ggctgatgct 960gcaccaactg tatccatctc ccaccatcca gttctagaaa
gccgaattcc agcacactgg 1020cggccgttac tagtggatcc gagctcggta ccaagcttgg
cgtaatcatg gtcatagctg 1080tttcctgtgt gaaattgtta
1100251100DNAArtificial Sequence5D7A1 Antibody and
surrounding vector sequence (Reverse Complement DNA) 25taacaatttc
acacaggaaa cagctatgac catgattacg ccaagcttgg taccgagctc 60ggatccacta
gtaacggccg ccagtgtgct ggaattcggc tttctagaac tggatggtgg 120gagatggata
cagttggtgc agcatcagcc cgtttcagct ccagcttggt cccagcaccg 180aacgtgagcg
gataggtgtt atattgctga cagaaatact ctgccaagtc ttcagactgc 240acattgttga
tggtgagagt gaaatctgtc ccagatccac tgcctgtgaa gcgatcaggg 300actccactgg
tccggaagga tgccgagtaa atcagtgctt taggagattg ccctagtttc 360tgttgatacc
aggctacatt agtctctaca ttctgactgg ccttgcaggt gacgctgacc 420ctgtctccta
ctgatgtgga catgaatttt tgagactggg tcatcacaat gtctccatca 480acaccagaca
accacagcaa catgtataca aagacctgag tctgtgactc catggtggat 540atcaagccga
attctgcaga tatccatcac actggcggcc gctcgagcat gcatctagag 600ctgcagaatt
cggcttggaa gatctataga cagatggggg tgtcgttttg gctgaggaga 660cggtgaccgt
ggtccctgcg ccccagacat cgaagaacca gcccccgtgg gacctataag 720gccttcttaa
acaataataa agtgcagaat cctcagatgt taggctgctc agctgcatgt 780agccgatact
ggaggatttg tttgcagtca atgtggcctt gtctttgaac ttttgattgt 840aattagtata
accactacta ggattaatgt atccaatcca ttccagaccc tgtccaggcc 900tctgttttac
ccagtgcatc gtgtaggttg taaaggtgta gccagaagcc ttgcaggaca 960tcttcactga
ggccccaggt cttgccaatt cagcccctga ctcctccagc tggacctcga 1020attccggaag
aagccgaatt ccagcacact ggcggccgtt actagtggat ccgagctcgg 1080taccaagctt
ggcgtaatca
110026171PRTAspergillus oryzaeMISC_FEATUREPortion of Deuterolysin Protein
(amino acids 179-349) 26Thr Asp Cys Lys Gly Asp Ala Glu Ser Ser Leu
Thr Thr Ala Leu Ser 1 5 10
15 Asn Ala Ala Lys Leu Ala Asn Gln Ala Ala Glu Ala Ala Glu Ser Gly
20 25 30 Asp Glu
Ser Lys Phe Glu Glu Tyr Phe Lys Thr Thr Asp Gln Gln Thr 35
40 45 Arg Thr Thr Val Ala Glu Arg
Leu Arg Ala Val Ala Lys Glu Ala Gly 50 55
60 Ser Thr Ser Gly Gly Ser Thr Thr Tyr His Cys Asn
Asp Pro Tyr Gly 65 70 75
80 Tyr Cys Glu Pro Asn Val Leu Ala Tyr Thr Leu Pro Ser Lys Asn Glu
85 90 95 Ile Ala Asn
Cys Asp Ile Tyr Tyr Ser Glu Leu Pro Pro Leu Ala Gln 100
105 110 Lys Cys His Ala Gln Asp Gln Ala
Thr Thr Thr Leu His Glu Phe Thr 115 120
125 His Ala Pro Gly Val Tyr Gln Pro Gly Thr Glu Asp Leu
Gly Tyr Gly 130 135 140
Tyr Asp Ala Ala Thr Gln Leu Ser Ala Gln Asp Ala Leu Asn Asn Ala 145
150 155 160 Asp Ser Tyr Ala
Leu Tyr Ala Asn Ala Ile Glu 165 170
27180PRTAspergillus fumigatusMISC_FEATUREPortion of Asp f2 protein (amino
acids 54-233) 27Ser Ser Cys Asn Ala Thr Gln Arg Arg Gln Ile Glu Ala Gly
Leu Asn 1 5 10 15
Glu Ala Val Glu Leu Ala Arg His Ala Lys Ala His Ile Leu Arg Trp
20 25 30 Gly Asn Glu Ser Glu
Ile Tyr Arg Lys Tyr Phe Gly Asn Arg Pro Thr 35
40 45 Met Glu Ala Val Gly Ala Tyr Asp Val
Ile Val Asn Gly Asp Lys Ala 50 55
60 Asn Val Leu Phe Arg Cys Asp Asn Pro Asp Gly Asn Cys
Ala Leu Glu 65 70 75
80 Gly Trp Gly Gly His Trp Arg Gly Ala Asn Ala Thr Ser Glu Thr Val
85 90 95 Ile Cys Asp Arg
Ser Tyr Thr Thr Arg Arg Trp Leu Val Ser Met Cys 100
105 110 Ser Gln Gly Tyr Thr Val Ala Gly Ser
Glu Thr Asn Thr Phe Trp Ala 115 120
125 Ser Asp Leu Met His Arg Leu Tyr His Val Pro Ala Val Gly
Gln Gly 130 135 140
Trp Val Asp His Phe Ala Asp Gly Tyr Asp Glu Val Ile Ala Leu Ala 145
150 155 160 Lys Ser Asn Gly Thr
Glu Ser Thr His Asp Ser Glu Ala Leu Gln Tyr 165
170 175 Phe Ala Leu Glu 180
28153PRTAspergillus fumigatusMISC_FEATUREPortion of MEP20 protein (amino
acids 176-328) 28Ala Ser Cys Ser Gly Ser Arg Ala Ser Ala Leu Ser Thr Ala
Leu Arg 1 5 10 15
Asn Ala Gly Ser Leu Ala Asn Ala Ala Ala Ser Ala Ala Ser Ser Gly
20 25 30 Ser Ser Thr Arg Phe
Gln Glu Tyr Phe Lys Thr Thr Ser Arg Arg Pro 35
40 45 Glu Asn Val Gly Gly Arg Phe Arg Ala
Val Gly Arg Glu Ala Ser Ser 50 55
60 Gln Ser Ser Gly Lys Thr Thr Tyr Tyr Cys Asn Asp Pro
Tyr Gly Tyr 65 70 75
80 Cys Asp Ser Asn Thr Leu Ala Tyr Thr Leu Pro Ser Ser Asn Leu Ile
85 90 95 Ala Asn Cys Asp
Ile Tyr Tyr Ser Tyr Leu Pro Ala Leu Thr Ser Ser 100
105 110 Cys His Ala Gln Asp Gln Ala Thr Thr
Thr Leu His Glu Phe Thr His 115 120
125 Ala Pro Ala Val Tyr Ser Pro Gly Thr Asp Asp Tyr Ala Tyr
Gly Tyr 130 135 140
Arg Ala Ser Thr Ala Leu Ser Ala Ser 145 150
29162PRTAspergillus fumigatusMISC_FEATUREPortion of Asp f2 protein (amino
acids 54-215) 29Ser Ser Cys Asn Ala Thr Gln Arg Arg Gln Ile Glu Ala Gly
Leu Asn 1 5 10 15
Glu Ala Val Glu Leu Ala Arg His Ala Lys Ala His Ile Leu Arg Trp
20 25 30 Gly Asn Glu Ser Glu
Ile Tyr Arg Lys Tyr Phe Gly Asn Arg Pro Thr 35
40 45 Met Glu Ala Val Gly Ala Tyr Asp Val
Ile Val Asn Gly Asp Lys Ala 50 55
60 Asn Val Leu Phe Arg Cys Asp Asn Pro Asp Gly Asn Cys
Ala Leu Glu 65 70 75
80 Gly Trp Gly Gly His Trp Arg Gly Ala Asn Ala Thr Ser Glu Thr Val
85 90 95 Ile Cys Asp Arg
Ser Tyr Thr Thr Arg Arg Trp Leu Val Ser Met Cys 100
105 110 Ser Gln Gly Tyr Thr Val Ala Gly Ser
Glu Thr Asn Thr Phe Trp Ala 115 120
125 Ser Asp Leu Met His Arg Leu Tyr His Val Pro Ala Val Gly
Gln Gly 130 135 140
Trp Val Asp His Phe Ala Asp Gly Tyr Asp Glu Val Ile Ala Leu Ala 145
150 155 160 Lys Ser
30316PRTAspergillus fumigatusMISC_FEATUREFull length recombinant Asp f2
30Met Ala Ala Leu Leu Arg Leu Ala Val Leu Leu Pro Leu Ala Ala Pro 1
5 10 15 Leu Val Ala Thr
Leu Pro Thr Ser Pro Val Pro Ile Ala Ala Arg Ala 20
25 30 Thr Pro His Glu Pro Val Phe Phe Ser
Trp Asp Ala Gly Ala Val Thr 35 40
45 Ser Phe Pro Ile His Ser Ser Cys Asn Ala Thr Gln Arg Arg
Gln Ile 50 55 60
Glu Ala Gly Leu Asn Glu Ala Val Glu Leu Ala Arg His Ala Lys Ala 65
70 75 80 His Ile Leu Arg Trp
Gly Asn Glu Ser Glu Ile Tyr Arg Lys Tyr Phe 85
90 95 Gly Asn Arg Pro Thr Met Glu Ala Val Gly
Ala Tyr Asp Val Ile Val 100 105
110 Asn Gly Asp Lys Ala Asn Val Leu Phe Arg Cys Asp Asn Pro Asp
Gly 115 120 125 Asn
Cys Ala Leu Glu Gly Trp Gly Gly His Trp Arg Gly Ala Asn Ala 130
135 140 Thr Ser Glu Thr Val Ile
Cys Asp Arg Ser Tyr Thr Thr Arg Arg Trp 145 150
155 160 Leu Val Ser Met Cys Ser Gln Gly Tyr Thr Val
Ala Gly Ser Glu Thr 165 170
175 Asn Thr Phe Trp Ala Ser Asp Leu Met His Arg Leu Tyr His Val Pro
180 185 190 Ala Val
Gly Gln Gly Trp Val Asp His Phe Ala Asp Gly Tyr Asp Glu 195
200 205 Val Ile Ala Leu Ala Lys Ser
Asn Gly Thr Glu Ser Thr His Asp Ser 210 215
220 Glu Ala Leu Gln Tyr Phe Ala Leu Glu Ala Tyr Ala
Phe Asp Ile Ala 225 230 235
240 Ala Pro Gly Val Gly Cys Ala Gly Glu Ser His Gly Pro Asp Gln Gly
245 250 255 His Asp Thr
Gly Ser Ala Ser Ala Pro Ala Ser Thr Ser Thr Ser Ser 260
265 270 Ser Ser Ser Gly Ser Gly Ser Gly
Ala Thr Thr Thr Pro Thr Asp Ser 275 280
285 Pro Ser Ala Thr Ile Asp Val Pro Ser Asn Cys His Thr
His Glu Gly 290 295 300
Gly Gln Leu His Cys Thr His His His His His His 305 310
315 311650DNAArtificial SequenceTrx-SMT3-Asp f2
fusion protein construct (Reverse complement DNA) 31ttaggtacaa
tgcagctgac caccttcatg ggtatgacaa ttgctcggaa catcaatggt 60tgcgctcggg
ctatcggtcg gggtggtggt tgcacctgaa ccgctaccgc tgctgcttga 120gctggtgctg
gtgcttgccg gtgcgcttgc gctaccggta tcatgaccct gatccggacc 180atgactttca
ccggcacaac caacacccgg tgctgcaata tcaaaggcat atgcttccag 240ggcaaaatac
tgcagtgctt cactatcatg ggtgctttcg gtgccattgc tttttgccag 300tgcaataact
tcatcatagc catctgcaaa atgatcaacc caaccctgac caactgccgg 360aacatgatac
agacgatgca tcagatcgct tgcccaaaag gtattggttt cgctacctgc 420aacggtataa
ccctggctac acatgctaac cagccaacga cgggtggtat agctacgatc 480acaaataacg
gtttcgctgg tcgcatttgc accacgccaa tgaccacccc aaccttccag 540tgcacaatta
ccatccggat tatcacaacg aaacagaacg ttggctttat caccattcac 600aataacatca
tatgcaccaa cggcttccat tgtcggacga ttgccaaagt atttacgata 660gatttcgctt
tcattacccc aacgcagaat atgtgctttt gcatgacgtg ccagttcaac 720tgcttcattc
agacctgctt caatctggcg acgctgggtt gcattacagc tgctatgaat 780cggaaagctg
gtaactgcac cggcatccca gctaaaaaac accggttcat gcggtgttgc 840accaccaatc
tgctcacgat gtgcttcgat aatatcgtta tcttccatat ccagatcttc 900cggtgtctga
tctgcctgaa tacgaatacc atcatacaga aaacgcaggc tatccatttc 960tttaccctga
cgttttgcaa atgcttccat cagacgacgc agcggtgtgg tttttttgat 1020tttaaagaag
atctcgctgc tgccatcact aactttcagg ttaatatggg tttccggttt 1080cacttcaggt
ttaacttccg gttttgcttc ttgattaact tcgctatcgc tcataccgcc 1140accaccatga
tggtgatgat gatgacctcc catggccttg tcgtcgtcgt cggtacccag 1200atctgggctg
tccatgtgct ggcgttcgaa tttagcagca gcggtttctt tcataccaga 1260accgcgtggc
accagaccag aagaatgatg atgatgatgg tgcatatggc cagaaccaga 1320accggccagg
ttagcgtcga ggaactcttt caactgacct ttagacagtg cacccacttt 1380ggttgccgcc
acttcaccgt ttttgaacag cagcagagtc gggataccac ggatgccata 1440tttcggcgca
gtgccagggt tttgatcgat gttcagtttt gcaacggtca gtttgccctg 1500atattcgtca
gcgatttcat ccagaatcgg ggcgatcatt ttgcacggac cgcaccactc 1560tgcccagaaa
tcgacgagga tcgccccgtc cgctttgagt acatccgtgt caaaactgtc 1620gtcagtcagg
tgaataattt tatcgctcat
1650321700DNAArtificial SequenceVL-GS15-VH-Fc scFv and surrounding
pEE12.4 vector sequence 32tagctgacag actaacagac tgttcctttc
catgggtctt ttctgcagtc accgtccttg 60acacgaagct tgccgccacc atggaaaccg
acaccctgct gctgtgggtg ctgctgctgt 120gggtgccagg atctacaggc gacattgtga
tgacccagtc tcaaaaattc atgtccacat 180cagtaggaga cagggtcagc gtcacctgca
aggccagtca gaatgtagag actaatgtag 240cctggtatca acagaaacta gggcaatctc
ctaaagcact gatttactcg gcatccttcc 300ggaccagtgg agtccctgat cgcttcacag
gcagtggatc tgggacagat ttcactctca 360ccatcaacaa tgtgcagtct gaagacttgg
cagagtattt ctgtcagcaa tataacacct 420atccgctcac gttcggtgct gggaccaagc
tggagctgaa agccatggct ctgcaggcta 480gtggtggtgg tggttctggt ggtggtggtt
ctggtggtgg tggttctgct agcgaggtcc 540agctggagga gtcaggggct gaattggcaa
gacctggggc ctcagtgaag atgtcctgca 600aggcttctgg ctacaccttt acaacctaca
cgatgcactg ggtaaaacag aggcctggac 660agggtctgga atggattgga tacattaatc
ctagtagtgg ttatactaat tacaatcaaa 720agttcaaaga caaggccaca ttgactgcaa
acaaatcctc cagtatcggc tacatgcagc 780tgagcagcct aacatctgag gattctgcac
tttattattg tttaagaagg ccttataggt 840cccacggggg ctggttcttc gatgtctggg
gcgcagggac cacggtcacc gtctcctctg 900tagaacccaa atcttgcgac aaaactcaca
catgcccacc gtgcccagca cctgaactcc 960tggggggacc gtcagtcttc ctcttccccc
caaaacccaa ggacaccctc atgatctccc 1020ggacccctga ggtcacatgc gtggtggtgg
acgtgagcca cgaagaccct gaggtcaagt 1080tcaactggta cgtggacggc gtggaggtgc
ataatgccaa gacaaagccg cgggaggagc 1140agtacaacag cacgtaccgt gtggtcagcg
tcctcaccgt cctgcaccag gactggctga 1200atggcaagga gtacaagtgc aaggtctcca
acaaagccct cccagccccc atcgagaaaa 1260ccatctccaa agccaaaggg cagccccgag
aaccacaggt gtacaccctg ccaccatcac 1320gagatgagct gaccaagaac caggtcagcc
tgacctgcct ggtcaaaggc ttctatccca 1380gcgacatcgc cgtggagtgg gagagcaatg
ggcagccgga gaacaactac aagaccacgc 1440ctcccgtgct ggactccgac ggctccttct
tcctctacag caagctcacc gtggacaaga 1500gcaggtggca gcaggggaac gtcttctcat
gctccgtgat gcatgaggct ctgcacaacc 1560actacacgca gaagagcctc tctctgtctc
ccgggaaata ggaattcatt gatcataatc 1620agccatacca catttgtaga ggttttactt
gctttaaaaa acctcccaca cctccccctg 1680aacctgaaac ataaaatgaa
1700331700DNAArtificial
SequenceVL-GS15-VH-Fc scFv and surrounding pEE12.4 vector sequence
(Reverse complement DNA) 33ttcattttat gtttcaggtt cagggggagg tgtgggaggt
tttttaaagc aagtaaaacc 60tctacaaatg tggtatggct gattatgatc aatgaattcc
tatttcccgg gagacagaga 120gaggctcttc tgcgtgtagt ggttgtgcag agcctcatgc
atcacggagc atgagaagac 180gttcccctgc tgccacctgc tcttgtccac ggtgagcttg
ctgtagagga agaaggagcc 240gtcggagtcc agcacgggag gcgtggtctt gtagttgttc
tccggctgcc cattgctctc 300ccactccacg gcgatgtcgc tgggatagaa gcctttgacc
aggcaggtca ggctgacctg 360gttcttggtc agctcatctc gtgatggtgg cagggtgtac
acctgtggtt ctcggggctg 420ccctttggct ttggagatgg ttttctcgat gggggctggg
agggctttgt tggagacctt 480gcacttgtac tccttgccat tcagccagtc ctggtgcagg
acggtgagga cgctgaccac 540acggtacgtg ctgttgtact gctcctcccg cggctttgtc
ttggcattat gcacctccac 600gccgtccacg taccagttga acttgacctc agggtcttcg
tggctcacgt ccaccaccac 660gcatgtgacc tcaggggtcc gggagatcat gagggtgtcc
ttgggttttg gggggaagag 720gaagactgac ggtcccccca ggagttcagg tgctgggcac
ggtgggcatg tgtgagtttt 780gtcgcaagat ttgggttcta cagaggagac ggtgaccgtg
gtccctgcgc cccagacatc 840gaagaaccag cccccgtggg acctataagg ccttcttaaa
caataataaa gtgcagaatc 900ctcagatgtt aggctgctca gctgcatgta gccgatactg
gaggatttgt ttgcagtcaa 960tgtggccttg tctttgaact tttgattgta attagtataa
ccactactag gattaatgta 1020tccaatccat tccagaccct gtccaggcct ctgttttacc
cagtgcatcg tgtaggttgt 1080aaaggtgtag ccagaagcct tgcaggacat cttcactgag
gccccaggtc ttgccaattc 1140agcccctgac tcctccagct ggacctcgct agcagaacca
ccaccaccag aaccaccacc 1200accagaacca ccaccaccac tagcctgcag agccatggct
ttcagctcca gcttggtccc 1260agcaccgaac gtgagcggat aggtgttata ttgctgacag
aaatactctg ccaagtcttc 1320agactgcaca ttgttgatgg tgagagtgaa atctgtccca
gatccactgc ctgtgaagcg 1380atcagggact ccactggtcc ggaaggatgc cgagtaaatc
agtgctttag gagattgccc 1440tagtttctgt tgataccagg ctacattagt ctctacattc
tgactggcct tgcaggtgac 1500gctgaccctg tctcctactg atgtggacat gaatttttga
gactgggtca tcacaatgtc 1560gcctgtagat cctggcaccc acagcagcag cacccacagc
agcagggtgt cggtttccat 1620ggtggcggca agcttcgtgt caaggacggt gactgcagaa
aagacccatg gaaaggaaca 1680gtctgttagt ctgtcagcta
1700341700DNAArtificial SequenceVH-GS15-VL-Fc scFv
and surrounding pEE12.4 vector sequence 34tagctgacag actaacagac
tgttcctttc catgggtctt ttctgcagtc accgtccttg 60acacgaagct tgccgccacc
atggaaaccg acaccctgct gctgtgggtg ctgctgctgt 120gggtgccagg atctacaggc
gaggtccagc tggaggagtc aggggctgaa ttggcaagac 180ctggggcctc agtgaagatg
tcctgcaagg cttctggcta cacctttaca acctacacga 240tgcactgggt aaaacagagg
cctggacagg gtctggaatg gattggatac attaatccta 300gtagtggtta tactaattac
aatcaaaagt tcaaagacaa ggccacattg actgcaaaca 360aatcctccag tatcggctac
atgcagctga gcagcctaac atctgaggat tctgcacttt 420attattgttt aagaaggcct
tataggtccc acgggggctg gttcttcgat gtctggggcg 480cagggaccac ggtcaccgtc
tcctcagcca tggctctgca ggctagtggt ggtggtggtt 540ctggtggtgg tggttctggt
ggtggtggtt ctgctagcga cattgtgatg acccagtctc 600aaaaattcat gtccacatca
gtaggagaca gggtcagcgt cacctgcaag gccagtcaga 660atgtagagac taatgtagcc
tggtatcaac agaaactagg gcaatctcct aaagcactga 720tttactcggc atccttccgg
accagtggag tccctgatcg cttcacaggc agtggatctg 780ggacagattt cactctcacc
atcaacaatg tgcagtctga agacttggca gagtatttct 840gtcagcaata taacacctat
ccgctcacgt tcggtgctgg gaccaagctg gagctgaaag 900tagaacccaa atcttgcgac
aaaactcaca catgcccacc gtgcccagca cctgaactcc 960tggggggacc gtcagtcttc
ctcttccccc caaaacccaa ggacaccctc atgatctccc 1020ggacccctga ggtcacatgc
gtggtggtgg acgtgagcca cgaagaccct gaggtcaagt 1080tcaactggta cgtggacggc
gtggaggtgc ataatgccaa gacaaagccg cgggaggagc 1140agtacaacag cacgtaccgt
gtggtcagcg tcctcaccgt cctgcaccag gactggctga 1200atggcaagga gtacaagtgc
aaggtctcca acaaagccct cccagccccc atcgagaaaa 1260ccatctccaa agccaaaggg
cagccccgag aaccacaggt gtacaccctg cccccatccc 1320gggatgagct gaccaagaac
caggtcagcc tgacctgcct ggtcaaaggc ttctatccca 1380gcgacatcgc cgtggagtgg
gagagcaatg ggcagccgga gaacaactac aagaccacgc 1440ctcccgtgct ggactccgac
ggctccttct tcctctacag caagctcacc gtggacaaga 1500gcaggtggca gcaggggaac
gtcttctcat gctccgtgat gcatgaggct ctgcacaacc 1560actacacgca gaagagcctc
tctctgtctc ccgggaaata ggaattcatt gatcataatc 1620agccatacca catttgtaga
ggttttactt gctttaaaaa acctcccaca cctccccctg 1680aacctgaaac ataaaatgaa
1700351700DNAArtificial
SequenceVH-GS15-VL-Fc scFv and surrounding pEE12.4 vector sequence
(Reverse complement DNA) 35ttcattttat gtttcaggtt cagggggagg tgtgggaggt
tttttaaagc aagtaaaacc 60tctacaaatg tggtatggct gattatgatc aatgaattcc
tatttcccgg gagacagaga 120gaggctcttc tgcgtgtagt ggttgtgcag agcctcatgc
atcacggagc atgagaagac 180gttcccctgc tgccacctgc tcttgtccac ggtgagcttg
ctgtagagga agaaggagcc 240gtcggagtcc agcacgggag gcgtggtctt gtagttgttc
tccggctgcc cattgctctc 300ccactccacg gcgatgtcgc tgggatagaa gcctttgacc
aggcaggtca ggctgacctg 360gttcttggtc agctcatccc gggatggggg cagggtgtac
acctgtggtt ctcggggctg 420ccctttggct ttggagatgg ttttctcgat gggggctggg
agggctttgt tggagacctt 480gcacttgtac tccttgccat tcagccagtc ctggtgcagg
acggtgagga cgctgaccac 540acggtacgtg ctgttgtact gctcctcccg cggctttgtc
ttggcattat gcacctccac 600gccgtccacg taccagttga acttgacctc agggtcttcg
tggctcacgt ccaccaccac 660gcatgtgacc tcaggggtcc gggagatcat gagggtgtcc
ttgggttttg gggggaagag 720gaagactgac ggtcccccca ggagttcagg tgctgggcac
ggtgggcatg tgtgagtttt 780gtcgcaagat ttgggttcta ctttcagctc cagcttggtc
ccagcaccga acgtgagcgg 840ataggtgtta tattgctgac agaaatactc tgccaagtct
tcagactgca cattgttgat 900ggtgagagtg aaatctgtcc cagatccact gcctgtgaag
cgatcaggga ctccactggt 960ccggaaggat gccgagtaaa tcagtgcttt aggagattgc
cctagtttct gttgatacca 1020ggctacatta gtctctacat tctgactggc cttgcaggtg
acgctgaccc tgtctcctac 1080tgatgtggac atgaattttt gagactgggt catcacaatg
tcgctagcag aaccaccacc 1140accagaacca ccaccaccag aaccaccacc accactagcc
tgcagagcca tggctgagga 1200gacggtgacc gtggtccctg cgccccagac atcgaagaac
cagcccccgt gggacctata 1260aggccttctt aaacaataat aaagtgcaga atcctcagat
gttaggctgc tcagctgcat 1320gtagccgata ctggaggatt tgtttgcagt caatgtggcc
ttgtctttga acttttgatt 1380gtaattagta taaccactac taggattaat gtatccaatc
cattccagac cctgtccagg 1440cctctgtttt acccagtgca tcgtgtaggt tgtaaaggtg
tagccagaag ccttgcagga 1500catcttcact gaggccccag gtcttgccaa ttcagcccct
gactcctcca gctggacctc 1560gcctgtagat cctggcaccc acagcagcag cacccacagc
agcagggtgt cggtttccat 1620ggtggcggca agcttcgtgt caaggacggt gactgcagaa
aagacccatg gaaaggaaca 1680gtctgttagt ctgtcagcta
1700365PRTArtificial Sequencemotif 36His Arg Leu
Tyr His 1 5 375PRTArtificial Sequencemotif 37His Glu Xaa
Xaa His 1 5 385PRTArtificial Sequencemotif 38Phe Ser Ala
Leu Lys 1 5 3931PRTAspergillus fumigatusMISC_FEATUREAsp
f2 signal peptide 39Met Ala Ala Leu Leu Arg Leu Ala Val Leu Leu Pro Leu
Ala Ala Pro 1 5 10 15
Leu Val Ala Thr Leu Pro Thr Ser Pro Val Pro Ile Ala Ala Arg
20 25 30
User Contributions:
Comment about this patent or add new information about this topic: