Patent application title: Methods For Identifying Analgesic Agents
Inventors:
IPC8 Class: AA61K31407FI
USPC Class:
514409
Class name: Five-membered hetero ring containing at least one nitrogen ring atom (e.g., 1,2,3-triazoles, etc.) the five-membered hetero ring consists of one nitrogen and four carbons spiro ring system
Publication date: 2016-06-16
Patent application number: 20160166541
Abstract:
The present invention relates to the discovery that mutations in SCN9A
are causative of Congenital Indifference to Pain (CIP) in humans. The
invention also relates to methods of using compounds small organic
compounds to treat pain as well as to predict the effect of analgesic
agents in patients, such as those possessing gain of function mutations.Claims:
1. A method of treating pain in a patient having a gain of function
nucleotide polymorphism in an SCN9A gene, comprising administering to
said patient a sodium channel blocking agent that selectively inhibits
SCN9A polypeptide.
2. The method of claim 1, wherein said gain of function polymorphism results in an SCN9A polypeptide having a tryptophan at amino acid 1150.
3. The method of claim 1, wherein said sodium channel blocking agent is compound A.
4. The method of claim 1, wherein said SCN9A gene without said polymorphism has the nucleotide sequence of SEQ ID NO: 3.
5. The method of claim 1, wherein said SCN9A polypeptide has the amino acid sequence of SEQ ID NO: 2.
6. A method of predicting the analgesic effect of a sodium channel blocking agent that selectively inhibits SCN9A polypeptide in a mammal, comprising: (a) determining in said mammal the presence of a gain of function polymorphism in an SCN9A gene of said mammal, (b) administering to said mammal an effective amount of Compound A, and (c) determining a positive analgesic effect in said mammal due to said administering of step (b), wherein said positive analgesic effect in step (c) indicates a positive analgesic effect by said sodium channel blocking agent.
7. The method of claim 6, wherein said gain of function polymorphism results in an SCN9A polypeptide having a tryptophan at amino acid 1150.
8. The method of claim 6, wherein said sodium channel blocking agent is an analog of compound A.
9. The method of claim 6, wherein said SCN9A gene without said polymorphism has the nucleotide sequence of SEQ ID NO: 3.
10. The method of claim 6, wherein said SCN9A polypeptide has the amino acid sequence of SEQ ID NO: 2.
11. The method of claim 6, wherein said mammal is a human being.
Description:
PRIORITY DATA
[0001] This application is a continuation-in-part of U.S. application Ser. No. 12/799,506, filed 26 Apr. 2010, which is a continuation of U.S. application Ser. No. 12/590,935, filed 17 Nov. 2009, now U.S. Pat. No. 8,410,054, which is a divisional of U.S. application Ser. No. 10/369,909, filed 19 Feb. 2003, now U.S. Pat. No. 7,659,082, which claims priority of U.S. Provisional Application No. 60/357,964, filed 19 Feb. 2002, and 60/429,836, filed 26 Nov. 2002, the disclosures of all of which are hereby incorporated by reference in their entirety.
FIELD OF THE INVENTION
[0002] The present invention relates to the discovery that mutations in SCN9A are causative of Congenital Indifference to Pain in humans. The invention also relates to methods of utilizing the SCN9A gene and expression products thereof for the screening and identification of therapeutic agents, including small organic compounds, which are selective only for SCN9A, and are useful in the treatment of pain. The invention also relates to methods of using these compounds to treat or otherwise ameliorate pain.
BACKGROUND OF THE INVENTION
[0003] This invention is concerned with a hereditary pain disorder known as "Congenital Indifference To Pain" (also referred to herein as "C-I-P"). It is an extremely rare but interesting medical condition. It is an autosomal recessive disorder that interferes with the normal perception of pain (see, Landrieu, P. S. G., and Allaire, C. Ann. Neurology, 27 (5):574-58 (1990) and Comings D E and Amromin G D, 1974. Neurology. September; 24(9):838-48.)
[0004] The description of the condition is fascinating. Patients are essentially completely indifferent to sensations that would cause pain in most individuals; yet at the same time they are quite able to distinguish between other sensations, such as thermal (hot/cold) and tactile (sharp/dull) sensations. Tendon reflex and vibration recognition is normal. Patients do not perceive inflammatory pain or dental pain even though there is a normal flare response to intradermal histidine injection, indicating that the peripheral arc reflex is intact. (Landrieu et al, 1990) A general description of C-I-P can be found at Online Mendelian Inheritance in Man (OMIM) reference*243000.
[0005] It is highly intriguing that the genetic mutation that underlies this disorder has such a specific and selective medical consequence. The inventors recognize that identification of the mutant gene which is associated with the disorder will provide a therapeutic target against which novel therapeutic agents can act. The inventors recognize that therapeutic agents which are highly selective for this therapeutic target, if they can be designed to mimic the effect of the mutation, have the potential to induce the same kind of analgesia, in the general population, as experienced by C-I-P patients.
[0006] It thus serves to examine in detail the physiological consequences of C-I-P. One C-I-P patient (3 years old, female) had no pain sensation but had normal thermal and tactile sensation and deep sensation. She appeared normal physically, although having tissue damage presumably resulting from the disorder, such as a shorter tongue, lack of some fingertips and multiple scars on fingers due to old burns. She learned to walk at 18 months (somewhat delayed). (Guillermo A. and A. Grinspan, "Congenital indifference to pain, a propos of a case with antecedents of consanguinity". Rev Neurol (Paris), 1970. 123(6): 434-5). In a follow up examination at age 34, patient appeared normal. Intelligence was normal. A broken bone not identified by the patient had resulted in an orthopedic deformity. Patient had a dental prosthesis. Sweating was normal. Tendon reflex and vibration recognition was normal. Position sense was normal. Patient was able to discriminate between "hot and cold" and between "sharp and dull" but discrimination between "touch" and "pinprick" was attenuated. Her maternal grandfather and her paternal grandmother were first cousins who had a common cousin who was apparently indifferent to pain. This consanguinity is in support of autosomal recessive inheritance.
[0007] Sweating, blood pressure and other autonomic responses are quite normal. No histological abnormalities are present in the peripheral nerves. (see Comings et al (1974) and Hirsch E, Moye D, Dimon J H 3rd. South Med J, 88(8):851-857 (1995)).
[0008] Still, to date, no genes or genetic loci responsible for C-I-P have been reported. Karyotype analysis of one C-I-P patient appeared normal. (Amguerra-Escobio, 2001, unpublished observations). One of the confusing aspects that has prevented identification of genes in the past is that there are other patients who have other hereditary pain disorders which have not been properly distinguished from autosomal recessive C-I-P. This makes it significantly harder to identify the common underlying genetic cause of the disorder.
[0009] For example, there are autosomal dominant and sporadic cases of congenital indifference to pain. The inventors believe that many of these cases have a different genetic basis than autosomal recessive C-I-P. Reports of these cases may be found at Becak et al, Acta Genet. Statist. Med. 14:133-142 (1964); Comings et al (1974); and in Landrieu et al, (1990).
[0010] Further, there is a very different from of the disease known as Hereditary Sensory and Autonomic Neuropathy (HSAN). HSAN is the classic "insensitivity" to pain. It has been classified into types I-V (Reilly, M. M. 1998. J. Neurol. 245(1):6-13). Each of these conditions is linked to an inherited neuropathy of sensory nerves. Some of these conditions have been linked to, genes or genetic loci, however, none of these are believed to correspond to the genetic basis underlying C-I-P.
[0011] It should be recognized that "insensitivity" and "indifference" are often used interchangeably in the art, but they should be properly distinguished for better understanding of the pathology of the disorder. The term "insensitivity" refers to situations in which sensory pathways are altered leading to lack of painful sensation while "indifference" refers to situations wherein the sensory pathway appears normal but there is a lack of reaction to a painful stimulus. The key distinction between these groupings is that the former are inherited neuropathies in which neuropathy is the primary part of the disease and the sensory pathway is the sole or primary part of the neuropathy, whereas in the latter, the sensory pathway appears, at a microscopic and macroscopic level, to be intact and normal. (Comings et al 1974)
[0012] Because this invention is directed towards genes, proteins and other tools which are targets for analgesic agents, an additional area of relevant background relates to existing analgesic agents and their therapeutic targets, where known.
[0013] Therapeutic agents for treatment of pain fall into two main classes--the NSAIDs (non-steroidal anti-inflammatory drugs) and the opioids. NSAIDs treat pain in a way similar to the mechanism of aspirin, the most well-known and oldest member of the class. Common NSAIDs include acetaminophen, ibuprofen and naproxen. These drugs mainly inhibit the body's ability to synthesize prostaglandins. The common mechanism of action for all NSAIDs is the inhibition of the enzyme cyclooxgenase (COX). A major commercial success has been achieved with specific inhibitors of COX-2, such as Celebrex.TM. from Pharmacia/Pfizer, and Vioxx.TM. from Merck & Co. The recently launched Bextra.TM. from Pharmacia/Pfizer, which is highly selective for inhibition of COX-2 over COX-1, is also expected to become commercial success.
[0014] Opioids act through the opioid receptor family. These drugs include the weak opioids such as codeine and Tylenol 3, and strong opioids such as morphine and methadone. Some are long acting, others are of short duration. Opioid analgesics have a tendency to addiction and dependency, and so are not ideal for long-term or chronic pain management.
[0015] Outside of the NSAIDs and opiods, there are a number of other suggested analgesic agents in clinical trials (i.e. not yet approved for marketing) which are believed to have alternative targets. Some clinical trials are attempting to establish that central neuropathic pain may respond to ion channel blockers such as blockers of calcium, sodium and/or NMDA (N-methyl-D-aspartate) channels. For example, in development are low affinity NMDA channel blocking agents for the treatment of neuropathic pain. The literature provides substantial pre-clinical electrophysiological evidence in support of the use of NMDA antagonists in the treatment of neuropathic pain. Such agents also may find use in the control of pain after tolerance to opioid analgesia occurs, particularly in cancer patients.
[0016] In accordance with the present invention, the identification of the hereditary basis for Congenital Indifference To Pain will be a key step for developing novel therapeutic agents because it has the potential to be a novel therapeutic target. This therapeutic target can be used to identify and discover more effective analgesics. Discovery of the target will also provide new methods and compositions for diagnosis of C-I-P and for distinguishing between types of inherited pain disorders.
BRIEF SUMMARY OF THE INVENTION
[0017] The present invention relates to the discovery that the human gene SCN9A (also known as Nav 1.7) when mutated, results in congenital indifference to pain.
[0018] In one aspect, the invention relates to a method of treating pain in a patient having a gain of function nucleotide polymorphism in an SCN9A gene, or amino acid replacement in a polypeptide, such as an SCN9A polypeptide, comprising administering to said patient a sodium channel blocking agent that selectively inhibits SCN9A polypeptide.
[0019] In another aspect, the present invention relates to a method of predicting the analgesic effect of a sodium channel blocking agent that selectively inhibits SCN9A polypeptide in a mammal, comprising:
[0020] (a) determining in said mammal the presence of a gain of function polymorphism in an SCN9A gene of said mammal, or an amino acid replacement in an SCN9A polypeptide,
[0021] (b) administering to said mammal an effective amount of a sodium channel blocking agent, preferably an agent that is selective for SCN9A, and
[0022] (c) determining a positive analgesic effect in said mammal due to the administering of step (b),
[0023] wherein said positive analgesic effect in step (c) indicates a positive analgesic effect by said sodium channel blocking agent.
[0024] In separate embodiments of these methods, the gain of function polymorphism results in an SCN9A polypeptide having a tryptophan at amino acid 1150 (as, for example, in SEQ ID NO: 2) and the sodium channel blocking agent is Compound A. In another embodiment, the SCN9A gene without said polymorphism has the nucleotide sequence of SEQ ID NO: 2 or 3.
BRIEF DESCRIPTION OF THE DRAWINGS
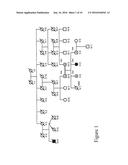
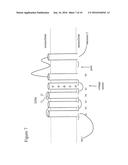
[0025] FIG. 1 shows the pedigree for a family designated CIP-10 showing family relationships and haplotypes. Squares and circles represent males and females, respectively. Filled symbols indicate individuals with CIP. Symbols with an "N" indicate individuals diagnosed as normal, and clear symbols with a question mark indicate individuals who have not been diagnosed
[0026] FIG. 2 shows the pedigree for a family designated CIP-14. Symbols as in FIG. 1.
[0027] FIG. 3 shows the pedigree for a family designated CIP-08. Symbols as in FIG. 1.
[0028] FIG. 4a. Nucleotide sequence of the 5067G>A SCN9A mutation in CIP patient CIP-10-503 (SEQ ID NO: 23). The patient is homozygous for a G>A substitution at nt 5067 (numbered relative to the initiator methionine in Genbank record NM_002977). This corresponds to a change to a stop codon at tryptophan 1689. The wild-type amino acid sequence from an unaffected patient is shown on top (SEQ ID NO: 22) and the wild-type nucleotide sequence as SEQ ID NO: 21. The amino acid sequence for the mutant is given as SEQ ID NO: 24. FIG. 4b. Nucleotide sequence of the 984C>A SCN9A mutation in CIP patient CIP-14-A005 (SEQ ID NO: 27). The patient is homozygous for a C>A substitution at nt 984. This corresponds to a change to a stop codon at tyrosine 328. The wild-type amino acid sequence from an unaffected patient is shown on top (SEQ ID NO: 26) with the wild-type nucleotide sequence above it (SEQ ID NO: 25). The mutant amino acid sequence is SEQ ID NO: 28. FIG. 4c. Nucleotide sequence of the 2488C>T SCN9A mutation in CIP patient CIP-08-11:01 (SEQ ID NO: 31). The patient is homozygous for a C>T substitution at nt 2488. This corresponds to a change to a stop codon at arginine 830. The wild-type sequence from an unaffected patient is shown on top (SEQ ID NO: 30) with the wild-type nucleotide sequence above it (SEQ ID NO: 29). The mutant amino acid sequence is SEQ ID NO: 32.
[0029] FIG. 5a. Genomic arrangement of alternatively spliced exon 5N and 5A of the human SCN9A gene. The neonatal (N) and adult (A) isoforms are generated by a mutually exclusive alternative splicing mechanism. The length in base pairs of the introns and exons are shown. FIG. 5b. Formation of SCN9A 5N (neonatal) isoform generated by splicing. FIG. 5c. Formation of SCN9A 5A (adult) isoform generated by splicing. Bent lines indicated portions of the pre-mRNA which are removed to generate the mature mRNA for translation.
[0030] FIG. 6. Comparison of the alternatively spliced exons of other sodium channels with the two variants of human SCN9A. The cDNA sequence and the predicted amino acid sequence of the alternatively spliced exon 5 of the A and N isoforms of the human (SEQ ID NO: 33 and 35, respectively) and rabbit (SEQ ID NO: 36 and 37, respectively) SCN9A, human (SEQ ID NO: 42 and 43, respectively) and rat (SEQ ID NO: 44 and 45, respectively) SCN3A, human SCN2A2 (SEQ ID NO: 38 and 39, respectively), and rat (SEQ ID NO: 40 and 41, respectively) SCN2A1 are shown. Isoform-specific differences in the amino acid sequence are indicated below the nucleotide sequence (SEQ ID NO: 34 is the human SCN9A isoform A).
[0031] FIG. 7. The proposed transmembrane topography of the first domain of a sodium channel protein. The 30 amino acids specified by these exon 5 variants encode a short extracellular portion of the S3 transmembrane helix, the short extracellular loop between S3 and S4, and most of the S4 transmembrane helix, which is thought to be involved in sensing changes in membrane potential. The + symbols refer to the repeating basic (positively charged) residues within the S4 transmembrane segment. Amino acid 206 of SCN9A is shown as an aspartate residue (A isoforms); it would be an uncharged residue in the N isoforms (i.e. asparagine in SCN9A and SCN2A; and serine in SCN3A).
[0032] FIG. 8. Illustration of nucleotide and amino acid sequence of alternative exon 11B. Alternative splice donor sites for exon 11 found in SCN9A isoform are indicated as upstream site and downstream site. Arrow indicates the approximate position of the additional amino acids in the secondary structure of the sodium channel protein.
[0033] FIG. 9. Evolutionary conservation of the additional amino acids of Exon 11B in other members of the sodium channel gene family (SCN9A, SCN1A, SCN8A). Here, the alignment shows human SCN9A (SEQ ID NO: 114), rat SCN9A (SEQ ID NO: 115), mouse SCN9A (SEQ ID NO: 116), rabbit SCN9A (SEQ ID NO: 117), human SCN1A (SEQ ID NO: 118), rat SCN1A (SEQ ID NO: 119), human SCN8A (SEQ ID NO: 120), rat SCN8A (SEQ ID NO: 121) and mouse SCN8A (SEQ ID NO: 122).
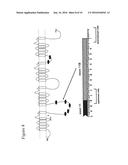
[0034] FIG. 10. Expression of SCN9A mRNA in various tissues of the central nervous system (CNS) and other organs. A representative gel is shown. To control for difference in the amounts of RNA among samples, pixel density relative to an internal .beta.-actin standard was calculated. The graph shows the results of 3 relative quantitative RT-PCR experiments relative to a dorsal root ganglia (DRG) positive control.
SEQUENCE LISTING
[0035] This application incorporates a sequence listing that has been filed electronically with the application as a text file only.
DETAILED DESCRIPTION OF THE INVENTION
[0036] The present invention provides a novel therapeutic target related to pain response and methods of using this target to identify useful analgesic agents, as well as methods for treating pain in human patients.
Description of SCN9A
[0037] In one aspect, the present invention identifies a gene and its corresponding protein related to pain indifference, one such embodiment being the SCN9A gene. The SCN9A gene and protein may be known to those skilled in the art under a variety of names, including voltage-gated sodium channel type IX alpha polypeptide, neuroendocrine sodium channel, peripheral sodium channel 1, Schwann cell sodium channel, NENA, NE-Na, hNE-Na, hNE, PN1, hPN1, NaS, Nas, Na.sub.v1.7, hNav1.7, and Na(v)1.7. Preliminary versions of the cDNA and amino acid sequences are found at GenBank Accession No. NM_002977.1 or XM_011955; The gene is described further at OMIM (On-line Mendelian Inheritance in Man) reference*603415.
[0038] The correct wild type nucleic acid sequence of the two isoforms of SCN9A (herein called "neo-natal" and "adult") is herein provided for the first time. The neo-natal SCN9A nucleic acid sequence is shown in SEQ ID NO: 1 with the correct neo-natal wild type amino acid sequence set forth at SEQ ID NO: 2. Table 1 identifies where the correct wild type sequence of neo-natal SCN9A set forth in SEQ ID NO: 1 diverges from the previously available public sequences of neo-natal SCN9A. The distinction between neo-natal and adult forms is a tentative description based on alternate Exon 5 (i.e. Exon 5N (for neo-natal) and Exon 5A (for adult)) further described, below. A nucleic acid sequence encoding a predicted adult splice variant of SCN9A employing exon 5A is found in SEQ ID NO: 3. The corresponding amino acid sequence of the predicted splice variant of SCN9A employing exon 5A is found in SEQ ID NO: 4. A nucleic acid sequence employing an alternate splice donor site at the end of Exon 11, herein called Exon 11B, is set forth at SEQ ID No. 5. The corresponding amino acid sequence of the predicted splice variant of SCN9A employing Exon 11B is found at SEQ ID No. 6. The sequences listed in Table 1, shown below, represent SEQ ID NO: 46 to 111, respectively.
[0039] The present invention provides a method of treating pain in a patient having a gain of function nucleotide polymorphism in an SCN9A gene, comprising administering to said patient a sodium channel blocking agent that selectively inhibits SCN9A polypeptide.
[0040] The present invention also provides a method of predicting the analgesic effect of a sodium channel blocking agent that selectively inhibits SCN9A polypeptide in a mammal, comprising:
[0041] (a) determining in said mammal the presence of a gain of function polymorphismin an SCN9A gene of said mammal,
[0042] (b) administering to said mammal an effective amount of Compound A, and
[0043] (c) determining a positive analgesic effect in said mammal due to said administering of step (b),
[0044] wherein said positive analgesic effect in step (c) indicates a positive analgesic effect by said sodium channel blocking agent.
[0045] The present invention discloses that an agent, especially Compound A, that selectively inhibits SCN9A containing a gain of function mutation, such as the R1150W mutation of SEQ ID NO: 2, is a predictor of analgesic activity of other sodium channel blocking agents that selectively inhibit SCN9A polypeptide in mammals, such as humans, containing gain of function mutations. Analgesic activity by compound A for a gain of function mutant polypeptide thus indicates that other SCN9A analgesic agents should also show analgesic activity with such mutants and serves to identify other susceptible gain of function mutants.
TABLE-US-00001 TABLE 1 Variation in relation Sequence variations to a (numbered relative Exon/ Variation in relation protein to initiator methionine) intron to a cNDA sequences sequence Differences between all 27 6182_6185delGATT not Public AAAAGTGATTGATTCAGTTTT samples or genomic applicable sequence: TTG sequence and Genbank Correct AAAAGTGATTCAGTTTTTTG entry NM_002977.1 sequence: Mutations 8 984C > A Y328X More CAGAGGGGTACACCTGTGTGA common: Less CAGAGGGGTAAACCTGTGTGA common: 15 2488C > T R830X More CTTTTAGCTCCGAGTCTTCAA common: Less CTTTTAGCTCTGAGTCTTCAA common: 26 5067G > A W1689X More CTGCTGGCTGGGATGGATTGC common: Less CTGCTGGCTGAGATGGATTGC common: Polymorphisms 1 174A > G not More CTGGCAAACAACTGCGCTTCA applicable common: Less CTGGCAAACAGCTGCCCTTCA common: 3 444G > A not More TGAATAACCCGCCGGACTGGA applicable common: Less TGAATAACCCACCGGACTGGA common: 9 1119C > T not More CGCTGCGTGCCGCTGGCAAAA applicable common: Less CGCTGCGTGCTGCTGGCAAAA common: 9 1266G > A not More AAGAATTAGAGTTTCAACAGA applicable common: Less AAGAATTAGAATTTCAAGAGA common: 9 1287A > T not More TGTTAGACCGACTTAAAAAAG applicable common: Less TGTTAGACCGTCTTAAAAAAG common: 18 3448C > T R1150W More GTTGTGTACGGAGGTTCTC common: Less GTTGTGTATGGAGGTTCTC common: 26 4779G > T not More CGTATTTTGTGTGCCCTACCC applicable common: Less CGTATTTTGTTTCCCCTACCC common: Intron 2 377 + 93_94delTGlnsGA not More TTTGCACCTTTGAAGACTCTG applicable common: G Less TTTGCACCTTGAAAGACTCTG common: G Intron 2 378 - 90A > T not More ATTTTTTTCTAAGGAAAAGTT applicable common: Less ATTTTTTTCTTAGGAAAAGTT common: Intron 3 478 - 219T > C not More ATATTCTTAGTTATTTCAAGT applicable common: Less ATATTCTTAGCTATTTCAAGT common: Intron 4 598 + 25T > C not More CTTTTTGAAATGGCAAATTTA applicable common: Less CTTTTTGAAACGGCAAATTTA common: Intron 4 598 + 591T > G not More TCAGAAAAATTGATTTTTACA applicable common: Less TCAGAAAAATGGATTTTTACA common: Intron 6 902 - 118T > C not More GGATGCATATTGCCTGGGACC applicable common: Less GGATGCATATCGCCTGGGACC common: Intron 8 1107 + 41A > G not More TTTGAATGGCATATGTACCTG applicable common: Less TTTGAATGGCGTATGTACCTG common: Intron 8 1107 + 48T > C not More GGCATATGTATCTGGTGTATG applicable common: Less GGCATATGTACCTGGTGTATG common: Intron 9 1314 + 76C > T not More GCGGTATATGCTTGGCCTTCT applicable common: Less GCGGTATATGTTTGGCCTTCT common: Intron 9 1314 + 199C > T not More CCCATAATCACCTCACTGCAT applicable common: Less CCCATAATCATCTCACTGCAT common: Intron 10 1603 - 368T > C not More TTTGTGAAGCTTGGGGATTGA applicable common: Less TTTGTGAAGCCTGGGGATTGA common: Intron 11 c.1942-2_3insT not More ATTTTTTTTTTAGGGCACGAC applicable common: C Less ATTTTTTTTTTTAGGGCACGA common: CC Intron 12 2072 - 14T > C not More ATGTTCTCTGTTTTTTTCTCC applicable common: Less ATGTTCTCTGCTTTTTTCTCC common: Intron 19 3769 - 70T > C not More TAGTGAGTTTTAGAATTGACT applicable common: Less TAGTGAGTTTCAGAATTGACT common: Intron 19 3769 - 4A > G not More TGTTATTTTTATAGGTTTCTT applicable common: Less TGTTATTTTTGTAGGTTTCTT common: Intron 21 4173 + 58_59delAA not More CGAAGGATATAAGTTATTCTT applicable common: T Less CGAAGGATATGTTATTCTTT common: Intron 21 4173 + 77C > A not More TTTAAATAGTCTATTAATTAT applicable common: Less TTTAAATAGTATATTAATTAT common: Intron 21 4174 - 146insA not More TTTAAAAAAATCTTTACATT applicable common: Less TTTAAAAAAAATCTTTACATT common: Intron 21 4174 - 196A > G not More ATAATTAACTAGGACTAAGAT applicable common: Less ATAATTAACTGGGACTAAGAT common: Intron 23 4365 + 99A > G not More TTCATGATTAATTTTATTAGA applicable common: Less TTCATGATTAGTTTTATTAGA common: Intron 23 4366 - 13_14insGTTT not More TTTTTTGTTTCTTTACCTTG applicable common: Less TTTTTTGTTTGTTTCTTTACC common: TTG Intron 25 474 + 16A > T not More AATATTTATTATTCAGATTTT applicable common: Less AATATTTATTTTTCAGATTTT common:
[0046] In preferred embodiments of these methods, the gain of function polymorphism results in an SCN9A polypeptide having a tryptophan at amino acid 1150, in place of the arginine that is otherwise present, and the sodium channel blocking agent is compound A. In other embodiments, the SCN9A gene without said polymorphism has the nucleotide sequence of SEQ ID NO: 3 and in one example of such a polymorphism it has the nucleotide sequence of SEQ ID NO: 1. In one embodiment, the SCN9A polypeptide has the amino acid sequence of SEQ ID NO: 2. In a preferred embodiment, the patient or mammal is a human being.
[0047] As used herein, a gain of function mutation refers to a mutation wherein the SCN9A polypeptide exhibits increased sodium transport that results in heightened sensitivity to pain. For example, human patients with a gain of function mutation in their SCN9A polypeptide, and therefore a gain of function polymorphism in their SCN9A gene encoding said polypeptide, report an increased pain score as described in examples 3 and 4.
[0048] As used herein, "pain score" is based on "pain intensity" as measured by trial subjects using an 11-point numerical rating scale (0-10, where 0=no pain and 10=worst pain imaginable, i.e., one that is incapacitating and may require the use of strong analgesic agents, or even hospitalization). This scale is also known as a Likert scale, a type of psychometric scale using a questionnaire to determine patient responses, and is widely used in pain clinical trials.
[0049] As measured herein, patients actively responded twice each day (once around the time they woke up and again around 8:00 PM.) In the morning they were asked to rate the previous night's pain intensity. In the evening, they were asked to provide their pain intensity for three different time periods throughout the day (between morning call and noon, between noon and 4:00 PM, and between 4:00 PM and the evening call). Patients were asked to enter the worst pain they experienced for the requested interval using the scale: 0 (no pain)-10 (worst pain you've ever imagined). For example, a value of greater than 3 might require some type of analgesic. A value of between 7 and 9 was considered severe.
[0050] In applying the methods of the present invention, Compound A is a compound having the chemical structure
##STR00001##
[0051] The structures of compounds that are analogs of this compound as well as methods of making and using said compound(s) are described in detail in U.S. Pub. No. US 2006/0252812 A1 (published 9 Nov. 2006) and explicitly in U.S. Application Publication No. US 2010/0331386 A1 (published 30 Dec. 2010), the disclosures of which are incorporated herein by reference in their entirety.
[0052] The present invention further provides at SEQ ID No. 7, a first mutation in the gene encoding this protein found in a first family (CIP-10) at G5067A (herein shown in the 5N/11 isoform, although 5A and 11B isoforms are also expected at different developmental stages). The mutation results in a truncation described as W1689X (SEQ ID NO: 8). The present invention also provides a second mutation in the gene encoding SCN9A which is responsible for congenital indifference to pain in a different family (CIP-14). The nucleic acid sequence of this second mutation, C984A, is set out in SEQ ID NO: 9 (herein shown in the 5N/11 isoform) and the corresponding mutant protein, Y328X, is described in SEQ ID NO: 10. The present invention also provides a third mutation in the gene encoding SCN9A which is responsible for congenital indifference to pain in a third family (CIP-08). The nucleic acid sequence of this second mutation, C2488T, is set out in SEQ ID NO 11 (herein shown in the 5N/11 isoform) and the corresponding mutant protein, R830X, is described in SEQ ID NO: 12.
[0053] This invention also provides human BAC clone RP11-437H3 (AC108146) which carries the complete human genomic fragment bearing the SCN9A gene. This BAC may be obtained commercially from well known sources. A short part of this BAC bearing the nucleotide sequence from the 3' end of exon 4 to the 5' end of exon 6, including exon 5A and exon 5N, is set forth at SEQ ID No. 13. The promoter region of SCN9A, including both genomic sequence and the 5'UTR of SCN9A is set forth at SEQ ID No. 14.
[0054] This invention also provides SCN9A from mouse (M. musculus) and various isoforms in Table 2 below.
[0055] The invention discloses several novel isoforms and mutants of human and mouse SCN9A.
TABLE-US-00002 TABLE 2 Isoforms of Mouse SCN9A disclosed SEQ ID No. Mouse SCN9A isoform 15 Mouse 11B nt 16 Mouse 11B aa 17 Mouse 5A nt 18 Mouse 5A aa 19 Mouse 5N nt 20 Mouse 5N aa nt = nucleotide, aa = amino acid
[0056] In aspects of the invention relating to the use of SCN9A, such as in screening assays, or for the development of antibodies, etc., etc., the inventors recognize that those skilled in the art may prefer to use forms of SCN9A corresponding to the sequences disclosed herein, although not necessarily the same. For example, screening assays may utilize SCN9A from a different organism, preferably a vertebrate, and most preferably from a mammalian species. The shared technical features of these forms of SCN9A, are that, when expressed, they have sodium channel activity, and that they share functional similarity with SCN9A, such as may be determined by those skilled in the art. Thus the invention encompasses the use of, for example, sheep, dog or horse SCN9A, for the same purposes as set out more specifically herein for human or mouse SCN9A.
[0057] As used herein, the term "correspond" means that the gene has the indicated nucleotide sequence or that it encodes the same RNA as would be encoded by the indicated sequence, including splice variants thereof.
[0058] Because of the processing that may take place in transforming the initial RNA transcript into the final mRNA, the sequences disclosed herein may represent less than the full genomic sequence. They may also represent sequences derived from alternate splicing of exons, ribosomal and/or transfer RNAs. Consequently, the genes present in the cell (and representing the genomic sequences) and the sequences disclosed herein, which are mostly cDNA sequences, may be identical or may be such that the cDNAs contain less than the full genomic sequence. Such genes and cDNA sequences are still considered corresponding sequences because they both encode similar RNA sequences. Thus, by way of non-limiting example only, a gene that encodes an RNA transcript, which is then processed into a shorter mRNA, is deemed to encode both such RNAs and therefore encodes an RNA complementary to (using the usual Watson-Crick complementarity rules), or that would otherwise be encoded by, a cDNA (for example, a sequence as disclosed herein). (Those skilled in the art understand that the word "encode" and its derivatives mean, in this field "can be transcribed into".) Thus, the sequences disclosed herein correspond to genes contained in the cells and are used to determine relative levels of expression because they represent the same sequences or are complementary to RNAs encoded by these genes. Such genes also include different alleles and splice variants that may occur in the cells used in the processes of the invention.
[0059] Thus, the polynucleotides, such as the genes disclosed herein, for use in the screening assays of the invention "correspond to" the polynucleotide encoding SCN9A mRNA (processed or unprocessed, including naturally occurring splice variants and alleles) at least 60%, preferably at least 70%, even more preferably at least 80%, or even at least 85%, most preferably at least 90%, or even at least 95%, or most especially at least 98%, with the especially preferred embodiment of identical to, and especially having the sequence of, an RNA that would be encoded by, or be complementary to, such as by hybridization under reasonably stringent conditions, with a SCN9A polynucleotide (SEQ ID NO: 1). In addition, sequences encoding the same polypeptides and proteins as any of these sequences, regardless of the percent identity of such sequences, are also specifically contemplated by any of the methods of the present invention that rely on any or all of said sequences, regardless of how they are otherwise described or limited. Thus, any such sequences are available for use in carrying out any of the methods disclosed according to the invention. Such sequences also include any open reading frames, as defined herein, present within an SCN9A polynucleotide.
[0060] As used herein, the term "percent identity" or "percent identical," when referring to a sequence, means that a sequence is compared to a claimed or described sequence after alignment of the sequence to be compared (the "Compared Sequence") with the described or claimed sequence (the "Reference Sequence"). The Percent Identity is then determined according to the following formula:
Percent Identity=100 [1-(C/R)]
wherein C is the number of differences between the Reference Sequence and the Compared Sequence over the length of alignment between the Reference Sequence and the Compared Sequence wherein (i) each base or amino acid in the Reference Sequence that does not have a corresponding aligned base or amino acid in the Compared Sequence and (ii) each gap in the Reference Sequence and (iii) each aligned base or amino acid in the Reference Sequence that is different from an aligned base or amino acid in the Compared Sequence, constitutes a difference; and R is the number of bases or amino acids in the Reference Sequence over the length of the alignment with the Compared Sequence with any gap created in the Reference Sequence also being counted as a base or amino acid.
[0061] If an alignment exists between the Compared Sequence and the Reference Sequence for which the percent identity as calculated above is about equal to or greater than a specified minimum Percent Identity then the Compared Sequence has the specified minimum percent identity to the Reference Sequence even though alignments may exist in which the hereinabove calculated Percent Identity is less than the specified Percent Identity.
[0062] As used herein, the terms "portion," "segment," and "fragment," when used in relation to polypeptides, refer to a continuous sequence of nucleotide residues, sequence forms a subset of a larger sequence. Such terms include the products produced by treatment of said polynucleotides with any of the common endonucleases, or any stretch of polynucleotides that could be synthetically synthesized. These may include exonic and intronic sequences of the corresponding genes.
[0063] Identification of SCN9A Modulating Agents
[0064] The present invention readily affords different means for identification of SCN9A modulating agents which are useful as therapeutic agents. One such protocol involves the screening of chemical agents for ability to modulate the activity of SCN9A thereby identifying an SCN9A modulating agent. SCN9A modulating agents so identified are then tested in a variety of in vivo models so as to determine if they alleviate pain, especially chronic pain or other disorders. In a preferred embodiment the agent modulates the activity of SCN9A downwards.
[0065] Thus, in one aspect the present invention relates to a method for identifying an agent that modulates the activity of a polynucleotide whose expression contributes to pain sensation or whose non-expression contributes to lack of pain sensation, comprising:
[0066] a) contacting under physiological conditions a chemical agent with a polynucleotide corresponding to a promoter of the SCN9A gene, preferably having the sequence of SEQ. ID NO. 14, under conditions promoting such contacting; and
[0067] b) detecting a change in the expression of said polynucleotide as a result of said contacting;
[0068] thereby identifying an agent that modulates said polynucleotide activity.
[0069] Such modulation is preferably a decrease in expression. In preferred embodiments, such expression is measured by measuring the amount of an expression product encoded by said polynucleotide, most preferably an RNA or a polypeptide. In a preferred embodiment the promoter sequence is operably linked to a reporter gene, and the assay measures relative expression of the reporter gene or its gene product.
[0070] In preferred embodiments, the polynucleotide whose expression is to be measured or monitored is present in an intact cell, preferably a mammalian cell, most preferably a neuronal cell, and may include a recombinant cell. In preferred embodiments, such an intact cell is a cell that has been engineered to comprise said polynucleotide, such as by genetic engineering, most preferably wherein the cell does not express the subject gene or polynucleotide absent having been engineered to do so.
[0071] Agents that alter the activity of SCN9A or active fragments or portions of said gene, or that may modulate the activity of polypeptides encoded by SCN9A, or polypeptides that act as transcription factors to modulate the activity of such genes, or other related gene segments, such as enhancers or other regulatory genetic elements that modulate the activity of SCN9A, acting either in cis or trans fashion, are thereby identified and may prove useful in treating chronic and other types of pain. Related genes might include those with high sequence homology, perhaps at least 98%, to the sequences disclosed herein and would, preferably, have sequences identical to the sequences disclosed herein.
[0072] In another aspect, the present invention relates to a method for identifying an agent that modulates the activity of a polypeptide encoded by a polynucleotide as disclosed herein, comprising:
[0073] a) contacting under physiological conditions a chemical agent with a polypeptide encoded by a polynucleotide corresponding to a SCN9A gene, preferably having the sequence of SEQ. ID NO. 1, 3, 5, 15, 17, 19; and
[0074] b) detecting a change in the activity of said polypeptide as a result of said contacting;
[0075] thereby identifying an agent that modulates said polypeptide activity.
[0076] In a preferred embodiment, the observed change in activity in step (b) is a decrease in activity, most preferably wherein said change in activity is the result of binding to or interacting with said polypeptide by said chemical agent of step (b), especially where the polypeptide is an ion channel, and the ion channel is blocked by said binding. In a further preferred embodiment, the polypeptide is a sodium channel, and said blocking is voltage dependent. A preferred therapeutic agent may be an irreversible binding agent or it may be a reversible binding agent.
[0077] In additional preferred embodiments, the polypeptide is part of a lipid bilayer, such as an intact cell, preferably a mammalian cell, such as a neuronal cell, most preferably a recombinant cell. In one such embodiment, a cell that has been engineered to comprise said polypeptide, including by genetic engineering, especially where the cell does not possess the polypeptide absent said engineering.
[0078] In another embodiment, the polypeptide is part of a cell, tissue, cell-line, immortalized cell or the like, which carries the gene and that expresses it, either naturally or upon induction. In this embodiment, the expressed gene is non-recombinant. As a non-limiting example, the PC-12 cell line derived from a transplantable rat pheochromocytoma responds reversibly to nerve growth factor (NGF) by induction of the neuronal phenotype and SCN9A expression. See Toledo-Aral J J, et al. Proc Natl Acad Sci USA. 1997 Feb. 18; 94(4):1527-32.
[0079] In one preferred embodiment of such method, the SCN9A polypeptide is a polypeptide corresponding to a polypeptide having the amino acid sequence of SEQ ID NO: 2, 4, 6, 16, 18, or 20.
[0080] Thus, the present invention specifically contemplates embodiments in which the cell is engineered by other than genetic engineering, such as where the activity of a polypeptide is to be enhanced and the cell has been engineered to contain, or have on its surface, said polypeptide but wherein the polypeptide is present due to physical insertion of the polypeptide into the membrane or cytoplasm of the cell and not through expression of a gene contained in the cell. Such engineering includes direct insertion of the polypeptide into the lipid bilayer, such as where a lipid bilayer comprising the polypeptide is fused to a membrane, such as that of an intact cell, resulting in the cell membrane comprising the polypeptide. Methods well known in the art, such as use of polyethylene glycol, viruses, and the like, are available to effect such insertions and the details of such procedures need not be further described herein.
[0081] In one preferred embodiment of such method, the polypeptide is a polypeptide that reacts with an antibody that reacts with, or is specific for, a polypeptide having an amino acid sequence at least 95% identical to, more preferably at least 98% identical to, the sequence of SEQ ID NO: 2 and where any difference in amino acid sequence is due only to conservative amino acid substitutions. In an especially preferred embodiment, the polypeptide has the amino acid sequence of SEQ ID NO: 2.
[0082] In a further aspect, the present invention relates to a method for identifying an analgesic agent, comprising:
[0083] a) administering to an animal an agent found to have activity using an assay or screening method as disclosed herein, and
[0084] b) detecting in said animal a decrease in response to a pain stimulus following said administering and due thereto,
[0085] thereby identifying an analgesic agent.
[0086] Preferably, the animal is a mammal, such as a human being. In specific embodiments, the pain stimulus is a heat stimulus and reaction or sensitivity to hot and/or cold may be measured. In another embodiment, an electrical stimulus may be used. In all cases, the stimulus may be represented as a sharp or dull sensation. In some cases, the animal may otherwise react normally to such stimulus so that a decrease in normal response due to the test agent is being measured whereas in other cases the animal may initially possess a heightened sensitivity to the stimulus prior to administering the test agent. In all cases, observation of an analgesic effect need not necessarily involve a reduced sensitivity or response to pain but may involve simply a reduced sensation of a particular stimulus. The analgesics identified by the methods of the invention may induce general analgesia in an animal or may have more localized analgesic or anesthetic effects. As used herein, the term "analgesia" refers to a reduction in, or absence of, pain in response to a stimulus that would otherwise be painful.
[0087] Further embodiments of the assays useful for identifying modulators of SCN9A are set out further below in this specification.
[0088] Selectivity for SCN9A Over Other Sodium Channels
[0089] The present invention specifically contemplates the identification of chemical agents, especially small organic molecules, that inhibit the expression of an SCN9A gene or the activity of an SCN9A polypeptide, with high specificity and/or selectivity and that have limited effect on other sodium channel genes and/or expression products. The inventors recognize on the basis of their discovery that selective inhibition of SCN9A as opposed to any of the other sodium channels leads to the therapeutically desirable outcomes observed in C--I-P. Because the other sodium channels are implicated in other essential physiological processes, such as heart activity, muscle contraction, and various neurological processes, it is highly desirable to avoid modulation of these other sodium channels.
[0090] This invention thus sets forth, for the first time, a scientifically justified rationale for the pursuit and discovery of sodium channel blockers which are highly selective for SCN9A for use as analgesics and to induce other symptomatic responses observed in C-I-P. The invention herein provides a solution to the problems posed in Anger et al. (2001. J. Med. Chem. 44(2):115-137) about the lack of information concerning which sodium channel sub-type to pursue and about the significant uncertainty in the art regarding the extent of selectivity required.
[0091] Also, by identifying SCN9A as the therapeutic target of choice in humans, the invention overcomes prior art teachings which suggested, but did not prove, that SCN9A may be involved in essential bodily functions such as bladder control and temperature control. SCN9A was believed to be the primary sodium channel of unmyelinated sensory and autonomic nerve cells. This prior art work led the field to believe that SCN9A was to be avoided as a target for therapeutic modulation because such modulation would negatively effect these essential bodily functions. However, the instant invention teaches, for the first time, that in humans, modulation of SCN9A can provide a highly desirable result, namely the treatment of pain, evidently without other physiological or autonomic impacts.
[0092] Thus, in a preferred embodiment of the invention, the compound identified which modulates SCN9A gene expression or SCN9A polypeptide activity is selective for SCN9A as opposed to other sodium channel alpha subunits (i.e. SCN1A to SCN8A, SCN10A to SCN12A). Such selectivity of modulation is preferably at least 10%, 50%, 100%, 10 times, 20 times, 100 times, 1000 times, 10,000 times or higher for SCN9A over any other sodium channel alpha subunit. Alternatively, a compound which is selective for SCN9A may not demonstrate an absolute preference for SCN9A, but the compound may show a preference for modulating SCN9A compared to any other sodium channel which preference is greater than any other prior art sodium channel blocker.
[0093] In another preferred embodiment, the methods disclosed herein for identifying an agent that modulates, preferably inhibits, expression or activity of a gene or polypeptide corresponding to SCN9A comprise first identifying such agent and then testing such agent for effects on expression or activity of at least one other sodium channel gene or polypeptide, as the case may be, preferably at least two other such genes, or polypeptides, with little or no effect. A preferred compound identified herein normally demonstrates no more than 25% of the effect that any compound heretofore measured has on the other sodium channel gene and/or polypeptide, preferably no more than 10% of such activity, most preferably no more than 5% of such activity and especially exhibiting no more than 1% of such activity. In such cases, the activity may be measured as the rate at which sodium ions are transported under the actions of the polypeptide or the amount of RNA expressed by said gene, especially on a weight-to-weight basis, or the nanomole ratio of RNA products produced by transcription of the gene.
[0094] Thus, the agents contemplated by the present invention are highly selective for the SCN9A gene or protein and administration of such an agent to a human or other animal in need thereof generates a temporary condition which mimics the Congenital Indifference to Pain phenotype to the highest degree possible, or aspects thereof. For example, it mimics the effects of one or more of the mutated forms of the gene as disclosed herein. In a preferred embodiment, the agent generates indifference to pain, but does not diminish sensations which are available to patients with CIP, such as but not limited to heat/cold sensing, pressure sensing, sharp/dull sensations, and does not diminish physiological responses such as sweating, heart rate, muscle activity and the like. For example, indifference to pain without any concomitant numbness or loss of sensation would represent a useful manifestation of the results to be achieved.
[0095] It is a preferred embodiment of this invention to use the screening assays of the invention to identify potential therapeutic agents, or analogs thereof, that are selective for the SCN9A sodium channel ahead of other sodium channels, in humans and other animals. It will therefore be apparent to those skilled in the art that a series of assays for measuring differential interaction with other sodium channels would provide the tools necessary to identify selective agents for SCN9A. In this embodiment, a first screening assay employing SCN9A is used to identify compounds which inhibit SCN9A activity. A suite of secondary assays are then employed, each containing one or more different sodium channels selected from among Na.sub.v1.1 (SCN1A); Na.sub.v1.2 (SCN2A), Na.sub.v1.3 (SCN3A), Na.sub.v1.4 (SCN4A), Na.sub.v1.5 (SCN5A), Na.sub.v1.6 (SCN8A), Na.sub.v1.8 (SCN10A), Na.sub.v1.9 (SCN11A), and Na.sub.x (SCN6A, SCN7A) (for consensus nomenclature see Goldin, A L. 2001. Ann. Rev. Physiol. 63:871-894). Compounds which were found to inhibit SCN9A expression or activity, are systematically tested against the assays for the other sodium channels. Those compounds which are more selective for SCN9A over other channels than current sodium channel blocking compounds used in the art are preferred compounds.
[0096] The other sodium channels can be incorporated into screening assays according to methods disclosed herein, or as disclosed elsewhere in the art or as later discovered or invented. As such, the Na.sub.v1.1, Na.sub.v1.2 and Na.sub.v1.3 screening assays set out in WO 01/38564 (Rouleau et al, published 31 May 2001) may be used to effectively distinguish compounds which are broadly selective for sodium channels from those that are specific and selective exclusively for SCN9A.
[0097] Some of the standard screening assays for a protein like SCN9A are set out in U.S. Pat. No. 6,110,672 (a US counterpart of WO 96/14077), incorporated herein by reference in its entirety. These assays provide for basic analysis of agents which interact with a protein like SCN9A (although the reference provides no guidance on achieving selectivity among sodium channels). The genetic and phenotypic information provided in the instant patent application now provides therapeutic agents which effectively mimic the Congenital Indifference to Pain phenotype, by selecting those compounds which are selective for SCN9A over other sodium channel family members. The invention provides a compound which, while not necessarily being selective for the peripheral nervous system over the central nervous system (as prescribed in WO 96/14077), the compound is specific for SCN9A ahead of all other sodium channels. Such a compound provides analgesic relief without side-effects such as nausea, tremor and irritability.
[0098] The benefits of the present invention and the value of selectivity for SCN9A over other sodium channels for the treatment of pain is further illustrated in Table 4 below. Table 4 sets out the known phenotypic or disease consequences of mutations in various known sodium channels in rodents and humans. As can be seen, modulation of the activity of almost all the other sodium channels results in severe and undesirable physiological consequences. For example, inhibitory mutations in human SCN1A result in a type of epilepsy. Gain of function mutations in rodent SCN2A result in behavioral disorders. Loss of function mutations in rodent SCN2A result in severe neurological disorders. SCN2A mutations in humans also result in a type of epilepsy. The list leads to the conclusion that a compound which selectively inhibits SCN9A will be a useful treatment for pain, and will not cause the undesirable' consequences of modulation of the other sodium channels.
[0099] The emphasis noted herein on the preferred selectivity for SCN9A over other sodium channels, should also be understood to include selectivity over other ion channels and related proteins and genes. Preferred selective SCN9A modulating agents have limited or no effect on the activity of ion channels such as potassium, calcium, ion co-transporters and the like, nor does it affect the hERG channel or other physiologically relevant channels. The emphasis on sodium channel selectivity is based on the high degree of similarity among sodium channels among which selectivity is technically more challenging to achieve than with less closely related proteins/genes.
TABLE-US-00003 TABLE 4 Human Disease or Gene Protein Rodent Phenotype Phenotype Reference SCN1A Na(v)1.1 Unknown Generalized epilepsy with 1, 2, 3, febrile seizures plus type II (OMIM 604233); severe myoclonic epilepsy of infancy (OMIM 607208) SCN2A2, Na(v)1.2 Transgenic Mouse - Generalized epilepsy with 4, 25 SCN2A seizures, focal motor febrile seizures plus (OMIM abnormalities, behavioral 604233) arrest and stereotyped repetitive behaviors; Knock-Out Mouse - perinatal lethal with severe hypoxia and massive neuronal apoptosis in the brainstem SCN3A Na(v)1.3 Unknown Unknown SCN4A Na(v)1.4 Unknown Hyper-(and hypo-) kalemic 10, 11 periodic paralysis (OMIM 170500, 170400 paramyotonia congenita (OMIM 168300, 168350) SCN5A Na(v)1.5 Knock-In Mice -arrhythmias; Long QT syndrome-3 12, 13, 14, Knock-Out mice (-/-) - (OMIM 603830), Brugada 15, 16, 17, intrauterine lethality with syndrome (601144); heart 26 defective ventricular block (OMIM 113900; morphogenesis; paroxysmal familial Knock-Out mice (+/-) - ventricular fibrillation (OMIM arrhythmias 603829) SCN8A Na(v)1.6 Various spontaneous Unknown 5, 6, 7, 8, 9 mutations result in: motor endplate disease - progressive paralysis and juvenile lethality in C57BL/6J mice; Viable adults with dystonia in C3H mice; Jolting mice - cerebellar ataxia; DMU mice - skeletal and cardiac muscle degeneration; heterozygotes unaffected SCN10A Na(v)1.8 Knock-out mice mechanical Unknown 18, 19, 20, analgesia, reduced thermal 21, 22 hyperalgesia, and delayed inflammatory hyperalgesia, but normal neuropathic pain perception (heterozygotes like wild-type); Knock-down mice - reduced inflammatory hyperalgesia and neuropathic pain (note discrepancy with above) SCN12A, Na(v)1.9 Knock-down in rat: No effect Unknown 23 SCN11A on nerve-injury induced behavior responses SCN6A, Na(x) Knock-out: Altered salt Unknown 24 SCN7A intake SCN9A Na(v)1.7 Unknown Congenital indifference to This work pain (OMIM 243000)
[0100] This invention also establishes that improved therapeutic agents for the treatment of pain must take advantage of the dynamic state of a sodium channel. Therapeutic agents can now be identified with entirely novel modes of action against the channel. A sodium channel cycles from the open (activated) state to the closed (inactivated) state, then to the resting (closed but capable of being activated) state, where it waits for an electrical signal to convert to the open state and allow passage of sodium ions. The transition from open to closed may be by fast inactivation or slow inactivation. Previous knowledge of the SCN9A channel did not allow for the identification of which state or transition state of the channel was preferred for therapeutic intervention. The instant invention now establishes that the Y328X or the W1689X mutation causes the desired channel behavior to generate the desirable indifference to pain in humans. An irreversible binding agent may be as suitable for use as a reversible binding agent. This invention therefore provides mutant proteins which may be used in the screening assays to assist in the identification of therapeutic agents or their analogs which mimic the truncation mutations of the invention. It also teaches a functional description about what the binding agent/blocking agent must do to achieve analgesia.
[0101] Further Assays
[0102] Stated broadly, the screening assays of the invention simplify the identification, evaluation and development of classes of compounds which are suitable for use as analgesic agents and for the reduction of adverse pain responses to disease, such as chronic disease, for example cancer. In general, these screening methods provide a ready means for selecting either natural product extracts or synthetic compounds of interest from a large population (i.e. a chemical library, for example, one produced by combinatorial means). As stated previously, an assay is first designed which measures SCN9A expression or activity. Exemplary methods useful for the identification of such compounds are detailed herein, although those skilled in the art will be aware of alternative means. In a first step, compounds are sequentially tested against the assay to determine whether they influence a measurable biological activity of the assay.
[0103] Functional assays may be based one or more of the diverse measurable biological activities of a gene or polypeptide corresponding to SCN9A. "SCN9A activity" as used herein, especially relating to screening assays, is to be interpreted broadly and contemplates all directly or indirectly measurable and identifiable biological activities of the SCN9A gene and protein. Relating to the purified SCN9A protein, SCN9A activity includes, but is not limited to, all those biological processes, interactions, binding behavior, binding-activity relationships, pKa, pD, enzyme kinetics, stability, and functional assessments of the protein. Relating to SCN9A activity in cell fractions, reconstituted cell fractions or whole cells, these activities include, but are not limited to the rate at which the SCN9A channel transports sodium, guanidine, lithium or other ions across a membrane, or the dynamics of this transport (such as voltage dependence, rate of transition between states) and all measurable consequences of these effects, including cell growth, development or behavior and other direct or indirect effects of SCN9A activity. Relating to SCN9A genes and transcription, SCN9A activity includes the rate, scale or scope of transcription of genomic DNA to generate RNA; the effect of regulatory proteins on such transcription, the effect of modulators of such regulatory proteins on such transcription; plus the stability and behavior of mRNA transcripts, post-transcription processing, mRNA amounts and turnover, and all measurements of translation of the mRNA into polypeptide sequences. Relating to SCN9A activity in organisms, this includes but is not limited biological activities which are identified by their absence or deficiency in disease processes or disorders caused by aberrant SCN9A biological activity in those organisms. Broadly speaking, SCN9A biological activity can be determined by all these and other means for analyzing biological properties of proteins and genes that are known in the art.
[0104] The invention therefore provides numerous assays which measure an activity of SCN9A and are useful for the testing of chemical compounds to identify which ones effect such activity. The invention also invites those skilled in the art to develop further SCN9A activity assays which go beyond those disclosed herein, for use in the screening compound libraries.
[0105] A typical assay uses patch-clamp techniques to study the behavior of channels. Such techniques are known to those skilled in the art, and may be developed, using current technologies, into low or medium throughput assays for evaluating compounds for their ability to modulate sodium channel behavior.
[0106] A competitive binding assay with known sodium channel toxins such as tetrodotoxin, alpha-scorpion toxins, and the like, may be suitable for identifying potential therapeutic agents with high selectivity for SCN9A.
[0107] In still another assay, uptake of radioactive isotopes into or out of a vesicle can be measured. The vesicles are separated from the extra-vesicular medium and the radioactivity in the vesicles and in the medium is quantitated and compared.
[0108] Other typical assays employ drug screening technology such as (but not limited to) radioactive, colorimetric or fluorescent based measurements. A classic colorimetric assay measures the ability of a dye to change color in response to changes in assay conditions resulting from the activity of the polypeptides. A useful instrument for the study of ion channels, including sodium channels is the ICR 8000 available from Aurora Biomed Inc. (Vancouver, BC).
[0109] Functional drug screening assays can also be based upon the ability of SCN9A polypeptides to interact with other proteins. Such interacting proteins can be identified by a variety of methods known in the art, including, for example, radioimmunoprecipitation, co-immunoprecipitation, co-purification, and yeast two-hybrid screening. Such interactions can be further assayed by means including but not limited to fluorescence polarization or scintillation proximity methods. Drug screens can also be based upon functions of the polypeptides deduced upon X-ray crystallography of the protein and comparison of their 3-D structure to that of proteins with known functions. Drug screens can be based upon a function or feature apparent upon creation of a transgenic or knockout mouse, or upon overexpression of the protein or protein fragment in mammalian cells in vitro. Moreover, expression of mammalian (e.g., human) polypeptides in yeast or C. elegans allows for screening of candidate compounds in wild-type and mutant backgrounds, as well as screens for mutations that enhance or suppress a low pain sensitivity phenotype. Modifier screens can also be performed in transgenic or knock-out mice.
[0110] Additionally, drug screening assays can also be based upon polypeptide functions deduced upon antisense interference with the gene function. Intracellular localization of SCN9A polypeptides, or effects which occur upon a change in intracellular localization of such proteins, can also be used as an assay for drug screening. Immunocytochemical methods may be used to determine the exact location of the pain-related polypeptides.
[0111] Polypeptides encoded by the polynucleotides disclosed herein can be used as an antigen to raise antibodies, including monoclonal antibodies. Such antibodies will be useful for a wide variety of purposes, including but not limited to functional studies and the development of drug screening assays and diagnostics. Monitoring the influence of agents (e.g., small organic compounds) on the expression or biological activity of the pain-related polypeptides identified according to the invention can be applied not only in basic drug screening, but also in clinical trials. For example, the effectiveness of an agent determined by a screening assay as described herein to increase or decrease gene expression, protein levels, or biological activity can be monitored in clinical trails of subjects exhibiting symptoms of chronic or persistent pain due to inadequate gene expression, protein levels, or biological activity (for example, the individuals studied herein. Alternatively, the effectiveness of an agent determined by a screening assay to modulate expression of SCN9A, as well as structurally and functionally related genes, including genes with high homology thereto, and including protein levels, or biological activity can be monitored in clinical trials of subjects exhibiting decreased altered gene expression, protein levels, or biological activity. In such clinical trials, the expression or activity of the genes or polypeptides disclosed herein and, preferably, other genes that have been implicated in, for example, congenital indifference to pain stimuli, can be used to ascertain the effectiveness of a particular analgesic drug.
[0112] As a non-limiting example, genes that are modulated in cells by treatment with an agent (e.g., compound, drug or small molecule) that modulates the activity of the SCN9A gene, or any expression products thereof, or polypeptides that modulate the activity of SCN9A (e.g., identified in a screening assay as described herein) can be identified. Preferably, such cells are recombinant cells engineered to express a polynucleotide or polypeptide as disclosed herein. Thus, such recombinant cells are prepared and RNA isolated and analyzed for the levels of expression of SCN9A after contacting said cells with agents that may have analgesic properties. The levels of gene expression can be quantified by Northern blot analysis or RT-PCR, or, alternatively, by measuring the amount of protein produced, by one of a number of methods known in the art, or by measuring the levels of biological activity of polypeptides encoded thereby or other genes. In this way, the gene expression can serve as a marker, indicative of the physiological response of the cells to the agent. Accordingly, this response state may be determined before, and at various points during, treatment of the individual with the agent.
[0113] In another aspect, the invention provides a method for computationally identifying a compound having analgesic properties. The method involves (a) determining crystal structure and preferably the active site of a SCN9A protein (i.e. through x-ray crystallography or other techniques); and (b) through computational modeling, identifying a compound which interacts with the active site, thereby identifying a compound, or its analog, as a compound which is useful for modulating the activity of such a polypeptide. Useful screening assays may also be performed in silico using available computerized databases for the identification of such compounds.
[0114] In a preferred embodiment, the present invention provides a method for monitoring the effectiveness of treatment of a subject with an agent (e.g., an agonist, antagonist, peptidomimetic, protein, peptide, nucleic acid, small molecule, or other drug candidate identified by the screening assays described herein) including the steps of (i) determining that a patient exhibits discomfort due to a disease or disorder that causes some type of painful stimulus; (ii) administering an effective amount of an agent identified using one of the screening assays disclosed herein; (iii) ascertaining a reduction to pain or other stimuli following said administration and (iv) altering the administration of the agent to the subject accordingly. For example, increased administration of the agent may be desirable to decrease the expression or activity of gene or encoded polypeptide, i.e., to increase the effectiveness of the agent.
[0115] Where the patient is non-human, biopsy samples can be taken to show a decrease in gene expression, such as by measuring levels of protein, mRNA, or genomic DNA post-administration samples and comparing the level of expression or activity of said protein, mRNA, or genomic DNA in the pre-administration sample with that of the corresponding post administration sample or samples, thereby showing the effects of drug administration on one or more of the genes disclosed herein and concomitant reduction in pain response and/or sensitivity.
[0116] The gene disclosed herein as being involved in congenital indifference to pain in an animal can be used, or a fragment thereof can be used, as a tool to express a protein, where such genes encode a protein, in an appropriate cell in vitro, or can be cloned into expression vectors which can be used to produce large enough amounts of protein to use in in vitro assays for drug screening. Expression systems which may be employed include baculovirus, herpes virus, adenovirus, adeno-associated virus, bacterial systems, and eucaryotic systems such as CHO cells, HEK cells and other cells commonly available. Naked DNA and DNA-liposome complexes can also be used.
[0117] Candidate modulators may be purified (or substantially purified) molecules or may be one component of a mixture of compounds (e.g., an extract or supernatant obtained from cells). In a mixed compound assay, SCN9A activity is tested against progressively smaller subsets of the candidate compound pool (e.g., produced by standard purification techniques, e.g., HPLC or FPLC; Ausubel et al.) until a single compound or minimal compound mixture is demonstrated to modulate gene or protein activity or expression in a manner having analgesic effects.
[0118] Specific compounds which will modulate the gene expression or gene transcript levels in a cell of SCN9A include antisense nucleic acids, ribozymes and other nucleic acid compositions which specifically hybridize with SCN9A (including exons or introns of such genes, promoters, 3'tails, etc.). These specific compounds are compounds of the invention, and are useful for treating the diseases discussed previously. Design and manufacturing of such compounds are well known to those skilled in the art.
[0119] Specific compounds which modulate the activity of a SCN9A polypeptide include antibodies (polyclonal or monoclonal) which specifically bind to an epitope of said polypeptide. These specific compounds are compounds of the invention, and are useful for inducing resistance of tolerance to pain stimuli. Design and manufacturing of such compounds are well known to those skilled in the art.
[0120] Specific compounds which modulate the activity of SCN9A in the body include gene therapy vectors comprising all or a part of the SCN9A gene sequence or mutant SCN9A sequence. As is well known to those skilled in the art, gene therapy allows the delivery of the SCN9A gene in an organism to cells where it is taken up and expressed, thus changing the level or amount of SCN9A protein in such cell. These vectors thereby modulate the activity of SCN9A in the body and are useful for the therapeutic indications disclosed herein.
[0121] In accordance with the foregoing, the present invention provides the amino acid sequence of a protein, designated SCN9A, that is a known ion channel structure found in neuronal cells (SEQ ID NO: 2), and which is associated with hereditary transmission of indifference to pain. In addition, a mutation, here a truncation, has been found in this sequence derived from individuals found to have such indifference to pain. Thus, agents that mimic the phenotypic effects of this truncation, such as aberrant protein structure and decreased, or absent, function represent candidate compounds for evaluation as analgesic agents. In addition, agents that have the effect of reducing the half-life of such polypeptide in cells would also act to induce decreased sensitivity to pain and thereby achieve analgesia.
[0122] By way of non-limiting example, cells expressing a wild-type SCN9A polypeptide are transiently metabolically labeled during translation, contacted with a candidate compound, and the half-life of the polypeptide is determined using standard techniques. Compounds that decrease the half-life of the polypeptide are useful compounds in the present invention.
[0123] In other embodiments, treatment with an antagonist of the invention may be combined with other analgesics to achieve a combined, possibly even synergistic, effect.
[0124] The ability of analgesic compounds to modulate polypeptides as disclosed herein, such as SCN9A, can be determined by any number of different binding assays, including use of a solid support, either as part of a column or as a batch procedure. Such support may be composed of plastic or glass, and includes standard resins and resin beads. Such assays are also available to test the ability of polypeptides, such as SCN9A, including mutated and/or truncated forms thereof, to bind to such test compounds or to other proteins present in cells, thereby identifying modulators of SCN9A activity according to the invention.
[0125] In one such assay for which the polypeptides encoded by genes disclosed herein are useful, the polypeptide (or a polypeptide fragment thereof or an epitope-tagged form or fragment thereof) is bound to a suitable support (e.g., nitrocellulose or an antibody or a metal agarose column in the case of, for example, a his-tagged form of said polypeptide). Binding to the support is preferably done under conditions that allow proteins associated with the polypeptide to remain associated with it. Such conditions may include use of buffers that minimize interference with protein-protein interactions. If desired, other proteins (e.g., a cell lysate) are added, and allowed time to associate with the polypeptide. The immobilized polypeptide is then washed to remove proteins or other cell constituents that may be non-specifically associated with it the polypeptide or the support. The immobilized polypeptide can then be used for multiple purposes. In a compound screening embodiment, compounds can be tested for their ability to interfere with interactions between SCN9A and other bound molecules (which are presumably SCN9A interacting proteins). Compounds which can successfully displace interacting proteins are thereby identified as SCN9A modulating agents of the invention.
[0126] In an alternative embodiment designed to identify the SCN9A interacting proteins, the immobilized polypeptide is dissociated from its support, and proteins bound to it are released (for example, by heating), or, alternatively, associated proteins are released from the polypeptide without releasing the latter polypeptide from the support. The released proteins and other cell constituents can be analyzed, for example, by SDS-PAGE gel electrophoresis, Western blotting and detection with specific antibodies, phospho-amino acid analysis, protease digestion, protein sequencing, or isoelectric focusing. Normal and mutant forms of such polypeptide can be employed in these assays to gain additional information about which part of the polypeptide a given factor is binding to. In addition, when incompletely purified polypeptide is employed, comparison of the normal and mutant forms of the protein can be used to help distinguish true binding proteins. Such an assay can be performed using a purified or semipurified protein or other molecule that is known to interact with a polypeptide encoded by a SCN9A polynucleotide.
[0127] This assay may include the following steps.
[0128] 1. Harvest the encoded polypeptide and couple a suitable fluorescent label to it;
[0129] 2. Label an interacting protein (or other molecule) with a second, different fluorescent label. Use dyes that will produce different quenching patterns when they are in close proximity to each other vs. when they are physically separate (i.e., dyes that quench each other when they are close together but fluoresce when they are not in close proximity);
[0130] 3. Expose the interacting molecule to the immobilized polypeptide in the presence or absence of a compound being tested for its ability to interfere with an interaction between the two; and
[0131] 4. Collect fluorescent readout data.
[0132] An alternative assay for such protein interaction is the Fluorescent Resonance Energy Transfer (FRET) assay. This assay can be performed as follows.
[0133] 1. Provide the encoded protein or a suitable polypeptide fragment thereof and couple a suitable FRET donor (e.g., nitro-benzoxadiazole (NBD)) to it;
[0134] 2. Label an interacting protein (or other molecule) with a FRET acceptor (e.g., rhodamine);
[0135] 3. Expose the acceptor-labeled interacting molecule to the donor-labeled polypeptide in the presence or absence of a compound being tested for its ability to interfere with an interaction between the two; and
[0136] 4. Measure fluorescence resonance energy transfer.
[0137] Quenching and FRET assays are related. Either one can be applied in a given case, depending on which pair of fluorophores is used in the assay.
[0138] One or more of the genes disclosed herein may act by altering membrane permeability, such as the permeability of membranes to ions, an example being the SCN9A gene disclosed herein, Such activity may be assayed for using, vesicles, such as liposomes or intact cells, wherein such structures comprise one or more of the polypeptides of the invention, which polypeptides are expressed in such vesicle, preferably an intact cell, such as a mammalian recombinant cell, and the permeability of the membrane of the cell is determined in the presence or absence of such expression. In the same way, such permeability can then be assayed in the presence and absence of chemical agents known to modulate the activity of one or more of the genes disclosed herein. Thus, the utility of these agents in enhancing the activity of proteins known to affect such membrane transport can be readily determined. In the same way, the ability of these agents to affect the transport of other molecules, such as lipids, amino acids, and the like, across such membranes is readily determined.
[0139] In performing such assays, the test cell, such as a mammalian recombinant cell expressing SCN9A, or a polynucleotide corresponding to such gene, is loaded with a reporter molecule (such as a fluorescent ion indicator whose fluorescent properties change when it binds a particular ion) that can detect ions (to observe outward movement), or alternatively, the external medium is loaded with such a molecule (to observe inward movement). A molecule which exhibits differential properties when it is inside the vesicle compared to when it is outside the vesicle is preferred. For example, a molecule that has quenching properties when it is at high concentration but not when it is at another low concentration would be suitable. The movement of the charged molecule (either its ability to move or the kinetics of its movement) in the presence or absence of a compound being tested for its ability to affect this process can be determined.
[0140] Throughput of test compounds is an important consideration in the choice of screening assay to be used. In some strategies, where hundreds of thousands of compounds are to be tested, it is not desirable to use low throughput means. In other cases, however, low throughput is satisfactory to identify important differences between a limited number of compounds. Often it will be necessary to combine assay types to identify preferred SCN9A modulating compounds
[0141] In brief, electrophysiology using patch clamp techniques is accepted as a gold standard for detailed characterization of sodium channel compound binding. There is a manual low-throughput screening (LTS) method which can compare 2-10 compounds per day; a recently developed system for automated medium-throughput screening (MTS) at 20-50 patches (i.e. compounds) per day; and a technology reportedly in development from Molecular Devices Corporation (Sunnyvale, Calif.) which permits automated high-throughput screening (HTS) at 1000-3000 patches (i.e. compounds) per day.
[0142] Binding Assays are also available, however these are of only limited functional value and information, content. Designs include traditional radioactive filter based binding assays or the confocal based fluorescent system available from Evotec OAI group of companies (Hamburg, Germany), both of which are HTS.
[0143] Radioactive flux assays can also be used. In this assay, channels are stabilized in an open state with veratridine, and channel blockers are identified by their ability to prevent ion influx. The assay can use radioactive 22[Na] and 14[C] guanidinium ions as tracers. FlashPlate & Cytostar-T plates in living cells avoids separation steps and are suitable for HTS. Scintillation plate technology has also advanced this method to HTS suitability. Because of the functional aspects of the assay, the information content is reasonably good.
[0144] Yet another format measures the redistribution of membrane potential using the FLIPR system membrane potential kit (HTS) available from Molecular Dynamics (a division of Amersham Biosciences, Piscataway, N.J.). Again, in this assay channels are stabilized in an open state with veratridine, and channel blockers are identified. The assay can use radioactive 22[Na] and 14[C] guanidinium ions as tracers. This method is limited to slow membrane potential changes. Some problems may result from the fluorescent background of compounds. Test compounds may also directly influence the fluidity of the cell membrane and lead to an increase in intracellular dye concentrations. Still, because of the functional aspects of the assay, the information content is reasonably good.
[0145] Sodium dyes can be used to measure the rate or amount of sodium ion influx through a channel. This type of assay provides a very high information content regarding potential channel blockers. The assay is functional and would measure Na influx directly. CoroNa Red, SBFI and/or sodium green (Molecular Probes, Inc. Eugene Oreg.) can be used to measure Na influx; all are Na responsive dyes. They can be used in combination with the FLIPR instrument. The use of these dyes in a screen has not been previously described in the literature. Calcium dyes may also have potential in this format.
[0146] In a further embodiment, FRET based voltage sensors are used to measure the ability of a test compound to directly block Na influx. Commercially available HIS systems include the VIPR.TM. II FRET system (Aurora Biosciences Corporation, San Diego, Calif., a division of Vertex Pharmaceuticals, Inc.) which may be used in conjunction with FRET dyes, also available from Aurora Biosciences. This assay measures sub-second responses to voltage changes. There is no requirement for a modifier of channel function. The assay measures depolarization and hyperpolarizations, and provides ratiometric outputs for quantification. A somewhat less expensive MTS version of this assay employs the FLEXstation.TM. (Molecular Devices Corporation) in conjunction with FRET dyes from Aurora Biosciences.
[0147] The present invention also relates to assays that may employ transcription factors for one or more of the genes disclosed herein. The effect of a test compound on the relative ability of such transcription factor to modulate transcription of the SCN9A gene is assessed by means of such an assay. In accordance with the disclosure herein, untranslated regions and promoter regions of SCN9A are provided or readily obtained. Such genomic or untranslated regions may be included in plasmids comprising the identified gene, such as in assays to identify compounds which modulate expression thereof. In one such assay, 1-5 kilobases of upstream genomic region of SCN9A is operably linked (i.e. ligated) to a reporter gene, and incorporated into an expression plasmid. The plasmid is transfected into a cell, and the recombinant cell is exposed to test compound(s). Those compounds which increase or decrease the expression of the reporter gene are then modulators of the SCN9A gene/protein, and are considered therapeutic agents of the invention.
[0148] Those skilled in the art are familiar with typical and easily measured reporter genes, such as luciferase, chloramphenicol acetyl-transferase (CAT), and other luminescent or fluorescent assays.
[0149] Thus, in one aspect the present invention relates to a method for identifying an agent that modulates the activity of a polynucleotide whose expression contributes to pain sensation or whose non-expression contributes to lack of pain sensation, comprising:
[0150] a) contacting under physiological conditions a chemical agent with a polynucleotide corresponding to a promoter of the SCN9A gene, preferably having the sequence of SEQ. ID NO. 14, under conditions promoting such contacting; and
[0151] b) detecting a change in the expression of said polynucleotide as a result of said contacting;
[0152] thereby identifying an agent that modulates said polynucleotide activity.
[0153] Medicinal Chemistry and Lead Optimization
[0154] While this invention discloses a wide variety of assays for measuring the effect of a compound on SCN9A expression or activity, it is important to note that this compound detection is merely the first step in the industrial process of identification of an approvable therapeutic agent. The library screening accomplished with the first screening assay may be low, medium or high-throughput screening. It identifies "hits" or individual compounds from the library which cause the desired modulation of SCN9A expression or activity. The hits are further evaluated at a chemical structure level, and, if possible, are organized according to shared core structures, which presumably define chemical features required to achieve the desired modulation of SCN9A. This process, sometimes called lead identification, may involve a structure-activity relationship (or SAR) analysis. Various cycles of medicinal chemistry or focussed library generation may then be employed to generate multiple analogs of such core structures in a process called lead optimization. Finally, those skilled in the art know how to identify and test those preferred analogs (optimized leads) which have improved characteristics as therapeutic agents, generally through a series of in vitro and in vivo analyses.
[0155] In general, novel agents having SCN9A modulating properties are identified from large libraries of both natural product or synthetic (or semi-synthetic) extracts or chemical libraries according to methods known in the art. Those skilled in the field or drug discovery and development will understand that the precise source of test extracts or compounds is not critical to the screening procedure(s) of the invention. Accordingly, virtually any number of chemical extracts or compounds can be screened using the exemplary methods described herein. Examples of such extracts or compounds include, but are not limited to, plant-, fungal-, prokaryotic- or animal-based extracts, fermentation broths, and synthetic compounds, as well as modification of existing compounds. Numerous methods are also available for generating random or directed synthesis (e.g., semi-synthesis or total synthesis) of any number of chemical compounds, including, but not limited to, saccharide-, lipid-, peptide-, and nucleic acid-based compounds. Synthetic compound libraries are commercially available from Brandon Associates (Merrimack, N.H.) and Aldrich Chemical (Milwaukee, Wis.). Alternatively, libraries of natural compounds in the form of bacterial, fungal, plant, and animal extracts are commercially available from a number of sources, including Biotics (Sussex, UK), Xenova (Slough, UK), Harbor Branch Oceangraphics Institute (Ft. Pierce, Fla.), and PharmaMar, U.S.A. (Cambridge, Mass.). In addition, natural and synthetically produced libraries are produced, if desired, according to methods known in the art, e.g., by standard extraction and fractionation methods. Furthermore, if desired, any library or compound is readily modified using standard chemical, physical, or biochemical methods.
[0156] In addition, those skilled in the art of drug discovery and development readily understand that methods for dereplication (e.g., taxonomic dereplication, biological dereplication, and chemical dereplication, or any combination thereof) or the elimination of replicates or repeats of materials already known for their analgesic and/or anesthetic activities should be employed whenever possible.
[0157] When a crude extract is found to have SCN9A modulating activity, further fractionation of the positive lead extract is necessary to isolate chemical constituent responsible for the observed effect. Thus, the goal of the extraction, fractionation, and purification process is the careful characterization and identification of a chemical entity within the crude extract having potential therapeutic activities. The same assays described herein for the detection of activities in mixtures of compounds can be used to purify the active component and to test derivatives thereof. Methods of fractionation and purification of such heterogeneous extracts are known in the art.
[0158] In an alternative strategy, if a compound is identified which is known to modulate expression or activity of 8 polynucleotide or polypeptide corresponding to SCN9A, analogs of that compound can be developed and tested for improved selectivity, potency, binding affinity or the like against the target gene/protein. According to the invention, such analogs will also be tested against other sodium channels (such as NaV 1.1 to 1.6 and NaV 1.8 to 1.12) to identify analogs which are preferentially selective towards blocking NaV 1.7 and which do not interact with the others. Such improved analogs, which are compounds of the invention, are expected to demonstrate reduced side effects and improved analgesic effects in human patients as compared to their parent compounds.
[0159] In an alternative embodiment, an investigator may desire to initiate research with known sodium channel blockers and to develop analogs of the blocker which are more selective for SCN9A. Known sodium channel modulators that may be selectively improved to increase specificity for SCN9A include:
[0160] Improved analogs may also include compounds with improved stability, biodistribution, pharmacokinetics or other desirable features for therapeutic agents which are not directly related to modulation of the therapeutic target.
[0161] In a preferred embodiment, the improved analog of the invention is effectively delivered, either by physiological means or assisted means, to cells of the body expressing the SCN9A protein.
[0162] Compounds identified as having potential therapeutic value are subsequently analyzed using any standard in vitro assay or in vivo animal model for the disease indication known in the art.
TABLE-US-00004 TABLE 3 Drug/clinical candidate Company AWD-140-190 ASTA Medica AG AWD-33-173 ASTA Medica AG BIA-2-024 Boehringer Ingelheim BIA-2-093 Boehringer Ingelheim Co-102862 University of Saskatchewan Conopeptides Cognetix Inc DCUKA Lohocla Research Corp felbamate Carter-Wallace Inc fosphenytoin InterX Research Corp GW-273293 Glaxo Wellcome plc lamotrigine Glaxo Wellcome pic OROS (phenytoin) ALZA Corp oxcarbazepine Novartis AG rufinamide Novartis AG safinamide Pharmacia & Upjohn AB topiramate Johnson & Johnson valproate semisodlum Abbott Laboratories vinpocetine Richter Gedeon VG ZM-227189 Zeneca Group plc zonisamide Dainippon Pharmaceutical Co Ltd.
[0163] Method of Treatment
[0164] In a further aspect, the present invention relates to a method for treating a condition in an animal afflicted with a source of chronic pain comprising administering to said animal an effective amount of an analgesic agent first identified by an assay method of the invention. Preferably, said animal is a human patient, such as a patient afflicted with a chronic ailment, such as a cancerous condition.
[0165] Therapeutic Use of Selective SCN9A Modulators
[0166] In a preferred embodiment, the agent generates indifference to pain, but does not diminish sensations which are available to patients with CIP, such as but not limited to hot/cold sensing, pressure sensing and sharp/dull sensations. For example, indifference to pain without any concomitant numbness or loss of sensation would represent an important use of the compounds of the invention. In an alternative embodiment, a preferred therapeutic agent is useful in humans or other animals for treating pain, inducing analgesia or anesthesia, or another disorder which is connected to the C-I-P phenotype.
[0167] Compounds first identified as useful in reducing sensitivity to pain stimuli using one or more of the assays of the invention may be administered with a pharmaceutically-acceptable diluent, carrier, or excipient, in unit dosage form. Conventional pharmaceutical practice may be employed to provide suitable formulations or compositions to administer such compositions to patients. Although oral administration is preferred, any appropriate route of administration may be employed, for example, intravenous, parenteral, subcutaneous, intramuscular, intracranial, intraorbital, ophthalmic, intraventricular, intracapsular, intraspinal, intrathecal, epidural, intracisternal, intraperitoneal, intranasal, or aerosol administration. Therapeutic formulations may be in the form of liquid solutions or suspension; for oral administration, formulations may be in the form of tablets or capsules; and for intranasal formulations, in the form of powders, nasal drops, or aerosols.
[0168] Methods well known in the art for making formulations are found in, for example, Remington: The Science and Practice of Pharmacy, (19th ed.) ed. A. R. Gennaro A R., 1995, Mack Publishing Company, Easton, Pa. Formulations for parenteral administration may, for example, contain excipients, sterile water, or saline, polyalkylene glycols such as polyethylene glycol, oils of vegetable origin, or hydrogenated napthalenes. Biocompatible, biodegradable lactide polymer, lactide/glycolide copolymer, or polyoxyethylene-polyoxypropylene copolymers may be used to control the release of the compounds. Other potentially useful parenteral delivery systems for agonists of the invention include ethylenevinyl acetate copolymer particles, osmotic pumps, implantable infusion systems, and liposomes. Formulations for inhalation may contain excipients, or example, lactose, or may be aqueous solutions containing, for example, polyoxyethylene-9-lauryl ether, glycocholate and deoxycholate, or may be oily solutions for administration in the form of nasal drops, or as a gel.
[0169] The methods of the invention simplify the evaluation, identification and development of active agents for the treatment and prevention of conditions of chronic or other types of pain while not necessarily treating the causative condition. Of course, both may be treated simultaneously as contemplated by the invention. For example, an analgesic agent identified by one of the screening methods disclosed herein may be administered along with an agent intended to treat a coincident conditions, such as where analgesic and antitumor agents are given together or contemporaneously.
[0170] Process of Data Transfer
[0171] The present invention also relates to a process that comprises a method for producing a product comprising identifying an agent according to one of the disclosed processes for identifying such an agent (i.e., the therapeutic agents identified according to the assay procedures disclosed herein) wherein said product is the data collected with respect to said agent as a result of said identification process, or assay, and wherein said data is sufficient to convey the chemical character and/or structure and/or properties of said agent. For example, the present invention specifically contemplates a situation whereby a user of an assay of the invention may use the assay to screen for compounds having the desired enzyme modulating activity and, having identified the compound, then conveys that information (i.e., information as to structure, dosage, etc) to another user who then utilizes the information to reproduce the agent and administer it for therapeutic or research purposes according to the invention. For example, the user of the assay (user 1) may screen a number of test compounds without knowing the structure or identity of the compounds (such as where a number of code numbers are used the first user is simply given samples labeled with said code numbers) and, after performing the screening process, using one or more assay processes of the present invention, then imparts to a second user (user 2), verbally or in writing or some equivalent fashion, sufficient information to identify the compounds having a particular modulating activity (for example, the code number with the corresponding results). This transmission of information from user 1 to user 2 is specifically contemplated by the present invention.
[0172] In one embodiment of the foregoing, the present invention relates to a method for producing test data with respect to the SCN9A polynucleotide modulating activity of a compound, comprising
[0173] a) contacting a chemical agent with a polynucleotide corresponding to a SCN9A polynucleotide, or corresponding to a SCN9A promoter sequence polynucleotide and under conditions promoting expression of such polynucleotide;
[0174] b) detecting a change in the expression of said polynucleotide as a result of said contacting; and
[0175] (c) producing test data with respect to the SCN9A polynucleotide modulating activity of said compound based on a change in the expression of the determined SCN9A polynucleotide modulating activity indicating such modulating activity.
Diagnostics and Pharmacogenomics
[0176] In a further embodiment, the invention relates to diagnostic and pharmacogenomic compounds, kits and methods. This aspect relates to analysis SCN9A for the diagnosis of insensitivity or indifference to pain, other pain disorder, or in the selection of a therapeutic agent for a patient (i.e. pharmacogenomics).
[0177] For example, nucleic acid analysis can be used to identify the G5067A or the C984A mutation thus confirming the diagnosis of congenital indifference to pain. Many nucleic acid diagnostic techniques are well known to those skilled in the art. Such techniques include DNA sequencing, hybridization probing, single stranded conformational analysis, PCR based techniques such as mismatch amplification, and myriad other well known methods. All such analysis can be performed on a small sample of blood, saliva, urine or other tissue provided by the patient.
[0178] Alternatively, protein based analyses such as antibody based assays (Elisa, Radioimmunoassay and the like) can be employed to identify the expression, amount or presence or absence of a mutant SCN9A protein, such as the W1689X or Y328X mutant.
[0179] Gene expression, both comparable and absolute, as well as biological activity, and mutational analysis can each serve as a diagnostic tool for pain disorders; thus determination of the amount of SCN9A mRNA can be used to diagnose the presence or absence of a mutation correlated with such pain disorder.
[0180] Pharmacogenomics deals with clinically significant hereditary variations in the response to drugs due to altered drug disposition and abnormal action in affected persons (Eichelbaum, M., Clin. Exp. Pharmacol. Physiol., 23:983-985, 1996; Linder, M. W., Clin. Chem., 43:254-266, 1997). In general, two types of pharmacogenetic conditions can be differentiated. Genetic conditions transmitted as a single factor altering the way drugs act on the body (altered drug action) or genetic conditions transmitted as single factors altering the way the body acts on drugs (altered drug metabolism). Altered drug action may occur in a patient having a polymorphism (e.g., a single nucleotide polymorphism or SNP) in promoter, intronic, or exonic sequences of SCN9A. Thus, determining the presence and prevalence of polymorphisms allow for prediction of a patient's response to a particular therapeutic agent.
[0181] This pharmacogenomic analysis can lead to the tailoring of drug treatments according to patient genotype, including prediction of side effects upon administration of therapeutic agents, particularly therapeutic agents for treating disorders disclosed in this specification. Pharmacogenomics allows for the selection of agents (e.g., drugs) for therapeutic or prophylactic treatment of an individual based on the genotype of the individual (e.g., the genotype of the individual is examined to determine the ability of the individual to respond to a particular agent).
[0182] Diagnostics employing a gene or protein corresponding to SCN9A can also be useful in selecting patients for clinical trials of a potential therapeutic agent. Patients can be stratified according to the DNA or protein sequence of SCN9A and their response to drug treatment can be evaluated. Such stratification can greatly reduce the number of patients required to establish efficacy for a potential therapeutic agent.
[0183] Sequences 1 to 20 have the following identifications:
SEQ ID NO: 1--Homo sapiens sodium channel, voltage-gated, type IX, alpha polypeptide (SCN9A), mRNA sequence SEQ ID NO: 2--Homo sapiens sodium channel, voltage-gated, type IX, alpha polypeptide (SCN9A), protein sequence SEQ ID NO: 3--Homo sapiens sodium channel, voltage-gated, type IX, alpha polypeptide (SCN9A), mRNA sequence, predicted splice variant SEQ 4--SCN9A predicted splice variant SEQ 5--Homo sapiens sodium channel, Voltage-gated, type IX, alpha polypeptide (SCN9A), mRNA, alternately spliced transcript using exon 5N and alternate splice donor site for exon 11 (11B) SEQ 6--Homo sapiens sodium channel, voltage-gated, type IX, alpha polypeptide (SCN9A), protein sequence, alternately spliced isoform using exon 5N and alternate splice donor site for exon 11 (11B) SEQ 7--Homo sapiens sodium channel, voltage-gated, type IX, alpha polypeptide (SCN9A), mRNA sequence, CIP-10 mutant SEQ 8--SCN9A W1689X protein sequence SEQ 9--Homo sapiens sodium channel, voltage-gated, type IX, alpha polypeptide (SCN9A), mRNA sequence, CIP-14 mutant SEQ ID NO: 10--SCN9A Y328X protein sequence SEQ ID NO: 11--Homo sapiens sodium channel, voltage-gated, type IX, alpha polypeptide (SCN9A), mRNA sequence, c.2488C>T mutation in CIP-08 SEQ ID NO: 12--Homo sapiens sodium channel, voltage-gated, type IX, alpha polypeptide (SCN9A), protein sequence, R830X CIP-08 mutation SEQ ID No. 13--Partial Sequence from human BAC clone RP11-437H3 (AC108146) showing genomic arrangement of Exon 5A and Exon 5N The first exon is the 3' end of Exon 4 (residues 1-16). Next Exon is Exon 5N (residues 1479-1570). The one following is Exon 5A (residues 1686-1777). The final exon is the 5' end of Exon 6 (residues 2506-2521). SEQ ID NO: 14--human genomic sequence containing the SCN9A promoter region ending with the ATG of the initiator methionine codon; a partial cDNA sequence appears as residues 9913-9963. SEQ ID NO 15--predicted mouse SCN9A 11B isoform SEQ ID NO: 16--predicted mouse SCN9A 11B isoform SEQ ID NO: 17--predicted mouse SCN9A 5A isoform SEQ ID NO: 18--predicted_1 mouse SCN9A 5A isoform SEQ ID NO: 19--predicted mouse SCN9A 5N isoform SEQ ID NO: 20--predicted mouse SCN9A 5N isoform
[0184] In carrying out the procedures of the present invention it is of course to be understood that reference to particular buffers, media, reagents, cells, culture conditions and the like are not intended to be limiting, but are to be read so as to include all related materials that one of ordinary skill in the art would recognize as being of interest or value in the particular context in which that discussion is presented. For example, it is often possible to substitute one buffer system or culture medium for another and still achieve similar, if not identical, results. Those of skill in the art will have sufficient knowledge of such systems and methodologies so as to be able, without undue experimentation, to make such substitutions as will optimally serve their purposes in using the methods and procedures disclosed herein.
[0185] The invention is described in more detail in the following non-limiting examples. It is to be understood that these methods and examples in no way limit the invention to the embodiments described herein and that other embodiments and uses will no doubt suggest themselves to those skilled in the art.
Example 1
Identification of the Genetic Mutation Responsible for Congenital Indifference to Pain (CIP) in Humans
[0186] We collected a multigenerational family with 3 affecteds (CIP-14) FIG. 1 and a smaller family with 1 affected (CIP-10) FIG. 2 reported previously (Guillermo A. and A. Grinspan. 1970. Rev Neurol (Paris) 123(6): p. 434-5.)
[0187] 31 family members from CIP-14 were genotyped at 763 autosomal markers and at 48 X markers. A genome-wide two-point analysis identified 8 regions with LOD scores >1.0. A region on chromosome 2 had two consecutive markers--D2S2330-D2S335--spanning 6 cM, with positive scores. A LOD score of 1.51 at zero recombination was obtained at D2S2330. Multi-point analysis of this region was consistent with linkage to CIP. The CIP-14 pedigree contains 3 affecteds, one issued from a consanguineous marriage; and two expected to share one allele identical by descent with the first case and one allele identical by descent between themselves. Because the pedigree comes from an isolated population, it can be expected that there would be linkage disequilibrium at the disease gene, and that all 3 affecteds would share alleles. However, as there was only one allele that could be assumed to be inherited identically by descent to the 3 affecteds, a scoring method was designed to assign scores to allele sharing taking that into consideration. The allele sharing algorithm identified 4 regions as having two-point LOD scores >1.0, and identified 3 additional regions that had not been excluded in two-point linkage analysis.
[0188] Haplotype construction using 12 polymorphic markers in candidate regions identified by 2-point linkage analysis and allele-sharing methods suggested that only the D2S2330-D2S335 region was consistent with linkage to CIP. All 3 affecteds shared one chromosome in common inherited from 3006/3009. Individuals 0000 and 0002 share another chromosome in common inherited from a 3004/3005. A005 has a chromosome introduced through 1020 that appears to be identical by state (and potentially identical by descent) to the one inherited from 3006/3009, which may be explained by the consanguinity of the population. The homozygous haplotypes in the 3 affecteds could represent multiple copies of a single ancestral chromosome. Recombination with the proximal marker D2S306, and the distal marker, D2S2188, is seen in 2002 and 0002/A006, respectively, provided centromeric and telomeric boundaries for the disease gene at the 2q24-31 locus.
[0189] Definition of the CIP Minimal Genomic Region
[0190] The interval between D2S306, and D2S2188 corresponds to a genetic size of .about.16 cM. To increase the resolution of the interval, we performed fine mapping with 32 additional markers, including several novel dinucleotide-repeat markers identified from genomic sequence. Haplotype construction using 41 polymorphic markers revealed one proximal and 5 distal recombinations. Of the 6 recombinant individuals, one was affected and 5 were carriers. The additional data narrowed the boundaries defined in 2002 and A006 to approximately 11.2 MB, between CA1AC010876 and D2S1267.
[0191] To assess whether the disease in CIP-10 maps to the 2q24-31 locus, five family members were genotyped using the 41 polymorphic markers described above. The proband was homozygous for markers between D2S2299 and D2S2177, and the genotypic data was consistent with linkage to CIP. Haplotype construction revealed a candidate interval of .about.19 cM defined by inferred recombination events that must have occurred in the untyped generations.
Use of genetic data from CIP-14 and CIP-10 together provided a centromeric boundary at CA1AC010876 (based a meiotic recombination in CIP14-2002) and a telomeric boundary at D2S2177 (based on the haplotype sharing data of CIP-10).
Mutation Analysis
[0192] We next embarked on a positional cloning effort. According to the UCSC August 2001 genomic sequence assembly, the CIP genomic region contains 26 known genes, five of which code for alpha polypeptides of voltage-gated sodium channels. We initiated sequencing of the coding regions of SCN9A and SCN3A in CIP-14-A005 and CIP-10-503, theorizing that selective expression in dorsal root ganglia and upregulation in models of neuropathic pain, respectively, made them interesting candidate genes. In CIP-10-503, we detected a homozygous nucleotide change of G>A at nt 5067 in SCN9A that resulted in a change to a stop codon at tryptophan 1689 (FIG. 4a). The stop codon results in deletion of 289 amino acids at the C-terminal of SCN9A, including the last transmembrane domain of the fourth channel domain. In CIP-14-A005 we detected a homozygous nucleotide change of 984C>A in SCN9A that resulted in a change to a stop codon at tyrosine 328 (FIG. 4b). We developed PCR-RFLP assays and showed that both mutations cosegregated perfectly with the disease and the affected haplotypes. Both mutations were absent in 142 control chromosomes from individuals of similar ancestral background. A third mutation was identified in a CIP proband descended from first cousins in another previously identified family. FIG. 3 (See Lievre J A, et al. 1968. Bull Mem Soc Med Hop Paris. 1968 Mar. 15; 119(5):447-56.) Patient CIP-08-II:1 is homozygous for a C>T substitution at nt 2488 that corresponds to a change to a stop codon at arginine 830 (FIG. 4c).
[0193] Alternative Exons 5N and 5A
[0194] The rabbit SCN9A gene (NaS), the rat and human SCN2A and SCN3A genes, and the human SCN8A gene contain two alternatively spliced exons encoding segments S3 and S4 of channel domain I. (See Belcher S M, et al. Proc Natl Acad Sci USA. 1995 Nov. 21; 92(24):11034-8; Sarao R, et al. Nucleic Acids Res. 1991 Oct. 25; 19(20):5673-9; Gustafson T A, et al. J Biol Chem. 1993 Sep. 5; 268(25):18648-53; Lu C M, and Brown G B. J Mol Neurosci 1998 February; 10(1):67-70; and Plummer N W, et al. Genomics. 1998 Dec. 1; 54(2):287-96.) The two exons are separated by introns of .about.90-150 bp. The isoforms are believed to be developmentally regulated, and without being bound to theory, it is suggested that the upstream exon (N) is predominantly expressed during the neonatal period, while the downstream exon (A) is expressed in adult brain. To determine whether this organization is conserved in human SCN9A, we aligned the genomic sequence from BAC clone RP11-437H3 (Genbank accession # AC108146) SEQ ID No. 13, with the human and rabbit coding sequences, and found that two potential copies of exon 5 are separated by 115 bp (FIG. 5). The exons contain two predicted amino acid differences. Alternative splicing for these exons is illustrated in FIG. 5. Residue 201 in the second exon is valine rather than leucine and residue 206 (corresponding to 209 in SCN2A and 208 in SCN3A) is aspartic acid rather than asparagine. A species and channel comparison in FIG. 6 shows the conserved relationship between isoforms. Because their physical arrangement and amino acid sequence correspond to those in other sodium channel genes, they are designated exons 5N and 5A. It has been suggested that the proximity of the residue 206 in these exons and the positively charged residues of the voltage sensor in transmembrane segment IS4 (see FIG. 7) may alter the voltage dependence or permeability of the neonatal and adult channels. Plummer NW, supra.
[0195] We used PCR to amplify the region of human SCN9A described above from total RNA of adult dorsal root ganglia (DRG) to examine exon usage within intact cDNAs. Primers designed to specifically bind cDNAs containing either exon 5A or exon 5N produced fragments of the expected size and sequencing confirmed mutually exclusive use of both exons.
[0196] Alternative Splice Donor for Exon 11
[0197] Two clones containing exons 1-13 of SCN9A amplified from human DRG were sequenced. One clone contained an additional 33 nucleotides located between the sequences of exon 11 and exon 12. To identify the origin of the extra 33 bp in the novel transcript, we examined the genomic sequence of the human SCN9A gene. Two alternative splice donor sites separated by 33 bp were identified. Splicing at the downstream site generates the novel transcript that encodes a protein with 11 additional amino acids in cytoplasmic loop 1 (FIG. 8). The sequence of the additional amino acids is conserved in rat and rabbit SCN9A and additional members of the channel gene family (FIG. 9).
Example 2
RT-PCR Analysis of Relative SCN9A Expression
[0198] cDNAs from a variety of human tissues were used to perform relative quantitative RT-PCR with primers and conditions that specifically amplified the SCN9A transcript (FIG. 10). Experiments to determine the tissue distribution of SCN9A were performed with three separate primer sets. Dorsal root ganglia was included as a positive control and .beta.-actin was included as an internal standard. We found highest levels of SCN9A relative to the .beta.-actin standard in dorsal root ganglia and medulla oblongata. Lower levels of SCN9A product were detected in the samples from temporal lobe, thymus, hippocampus, cerebral cortex, amygdala, frontal lobe, spinal cord, occipital lobe, cerebellum, thalamus, and ovary. SCN9A expression was not detected in cDNA from parietal lobe, corpus callosum, cerebral peduncles, pons, thyroid, or placenta. The sequences of the RT-PCR products amplified from dorsal root ganglia and medulla oblongata were identical to human SCN9A cDNA. Previous studies have reported low levels of SCN9A in the CNS relative to the PNS, but we have detected significant levels in the CNS, and particularly high levels in the medulla oblongata.
[0199] This expression profile teaches those skilled in the art where compounds which modulate SCN9A gene expression or SCN9A protein activity are likely to have effect. For example, expression of SCN9A in the CNS may support a role for this gene in central perception of pain, although alternative theories are suggested. It also is strongly suggestive of the other therapeutic indications for SCN9A modulating agents.
Example 3
Sequencing of Exon 19 of Scn9a from Participants in Study
[0200] This was a phase 2a, randomized, double-blind, placebo-controlled, two-period crossover study to evaluate the safety, tolerability, preliminary efficacy and systemic exposure of topical application of compound A in subjects with post herpetic neuralgia. The purpose was to investigate the prevalence of the polymorphism (R1150W) mutation in exon 19 of the SCN9A gene in patients who participated in the Compound A-201 clinical study and determine if the presence or absence of this mutation correlated with responsiveness or non responsiveness in any of the efficacy measures investigated when treating these subjects PHN pain with Compound A ointment.
[0201] The polymorphism R1150W (rs6746030) is a mutation with a global population minor allele frequency (MAF) of 0.099 (dbSNP) and has been linked to increased pain sensitivity (F Reimann, et al., PNAS, Vol. 107, pp. 5148-5153 (2010)). It therefore constitutes a gain-of-function (GOF) variant. It has also been linked to multiple regional pain and is involved in genetic susceptibility to pain but not osteoarthritis. (see Valdes et al., Arthritis Care and research, Vol. 63, pp. 440-444 (2011)
[0202] Compound A-201 ointment is a novel, potent, voltage-dependent sodium channel (NaV) blocker being developed for the treatment of various pain indications. Investigation of the presence of the R1150W polymorphism in patients with PHN treated with Compound A-201 may lead us to a better understanding of the populations of patients experiencing refractory PHN and the response to Compound A in patients with this polymorphism.
[0203] Blood samples were taken from 58 participants following obtaining consent for pharmacogenetic testing. Blood was collected in a 10 ml EthyleneDiamineTetraAcetate (EDTA) tube, mixed by inverting 8-10 times and then frozen immediately at -20.degree. C. Six subjects were identified as having left the study prior to receiving a dose of Compound A-201 and samples from these subjects were not processed any further. DNA was extracted from 52 tubes using the Gentra Puregene Blood Kit from Qiagen (CAT# D-5000) according to the manufacturer's protocol.
[0204] PCR was carried out with primers designed from P7 for Exon 19 of SCN9A (Table 1) and using the PCR mix and cycling as outlined in appendix A. Samples were sent to Seqwright for sequencing. All samples were amplified and sequenced in duplicate to ensure result accuracy.
TABLE-US-00005 TABLE 5 Primer Sequences Primer name Sequence P157_SCN9A_1150WF GGAGTATCACAGAAAGCAGTTTGC P157_SCN9A_1150WR TGTCATTGGCACTAATCATAGGG
[0205] All sequencing analysis was carried out using Sequencher version 4.10.1 and aligned to a reference sequence taken from the SCN9A entry in Ensembl build 37 (Ensembl).
TABLE-US-00006 TABLE 6 Results for each individual screened for R1150W Subject Genotype Result 001-001 C/C Wildtype 001-002 C/T R1150W 001-003 C/C Wildtype 001-004 C/C Wildtype 001-005 C/C Wildtype 001-011 C/C Wildtype 003-003 C/C Wildtype 004-001 C/C Wildtype 005-001 C/C Wildtype 005-002 C/C Wildtype 005-005 C/C Wildtype 005-007 C/C Wildtype 005-008 C/T R1150W 005-009 C/C Wildtype 005-011 C/C Wildtype 005-012 C/C Wildtype 005-013 C/C Wildtype 005-014 C/T R1150W 008-001 C/C Wildtype 008-002 C/C Wildtype 008-005 C/C Wildtype 008-006 C/C Wildtype 008-007 C/C Wildtype 009-001 C/C Wildtype 010-002 C/C Wildtype 011-001 C/C Wildtype 011-002 C/C Wildtype 011-004 C/C Wildtype 011-005 C/C Wildtype 011-006 C/C Wildtype 011-009 C/C Wildtype 012-001 C/C Wildtype 012-003 C/C Wildtype 012-004 C/C Wildtype 014-002 C/T R1150W 014-004 C/C Wildtype 014-006 C/T R1150W 015-001 C/C Wildtype 015-003 C/C Wildtype 015-004 C/C Wildtype 016-005 C/C Wildtype 016-008 C/T R1150W 016-010 C/C Wildtype 016-012 C/C Wildtype 016-014 C/T R1150W 016-015 C/C Wildtype 016-019 C/T R1150W 017-002 C/T R1150W 017-004 C/C Wildtype 026-001 C/C Wildtype 026-002 C/C Wildtype 027-002 C/C Wildtype
[0206] Nine individuals were found to be heterozygous for the polymorphism R1150W. Forty three individuals were found to not have the polymorphism. Table 6 lists all the individuals tested and their results for the polymorphism.
[0207] Nine individuals out of the 52 subjects tested have the polymorphism R1150W. This is a MAF of 0.17 which is higher than the dbSNP global MAF for this polymorphism but within range of tested populations (0.022-0.284).
TABLE-US-00007 TABLE 7 PCR Mix Volume Reagent 9.2 ul H.sub.2O 4.0 ul 5x buffer 1.6 ul 25 mM MgCl.sub.2 0.05 ul (each) dNTPs 0.15 ul Taq (2500 u/ml)
PCR Cycling
1.times.--95.degree. C. for 5 min
14.times.--95.degree. C. for 1 min
[0208] 70.degree. C. for 1 min (-1.degree. C. per cycle)
[0209] 72.degree. C. for 1 min
39.times.--95.degree. C. for 1 min
[0209]
[0210] 58.degree. C. for 1 min
[0211] 72.degree. C. for 1 min
1.times.--72.degree. C. for 10 min
Example 4
Clinical Study Report
[0212] This was a randomized, placebo-controlled, 2-period crossover study consisting of an up to 3-week Screening/washout Period; two 1-week, single-blind, Placebo Run-in Periods; a 1-week inter-treatment washout period; and two 3-week, double-blind Treatment Periods. It was a Phase 2a, Randomized, Double-blind, Placebo-controlled, Two-Period Crossover Study to Evaluate the Safety, Tolerability, Preliminary Efficacy, and Systemic Exposure of Topical Compound A in Subjects with Postherpetic Neuralgia.
[0213] The objectives of the study were to compare the safety and efficacy of Compound A (in the form of an ointment) to that of placebo (also an ointment) for the relief of pain in subjects with post-herpetic neuralgia (PHN) and to evaluate the extent of systemic exposure of Compound A following topical application of ointment in subjects with PHN.
[0214] During a first Placebo Run-in Periods, subjects applied placebo ointment twice daily (bid, i.e., "bis in die" or twice a day)) for 7 days. The area of application was to cover the most painful skin segments up to 400 cm.sup.2. Subjects recorded their pain score at 4 specific times each day (upon waking, 12 noon.+-.1 hour, 4 PM.+-.1 hour, and 8 PM.+-.1 hour) and reported these scores using the Medpace ClinTrak Interactive Voice Response System (CTIVRS). The use of any rescue medication (1000 mg acetaminophen every 4 to 6 hours, up to 4000 mg/day) was recorded in the rescue medication diary.
[0215] Subjects who reported average daily pain scores on the 11-point Likert Numerical Rating Scale (NRS) for at least 4 days during the first Placebo Run-in Period were eligible for Randomization to treatment and entered into Treatment Period 1. Subjects were randomly assigned (in a 1:1 ratio) to one of the following treatment sequences: ointment/placebo or placebo/ointment. Subjects received the first specified treatment during Treatment Period 1 and the second specified treatment during Treatment Period 2.
[0216] During each 3-week, double-blind Treatment Period, subjects applied study medication bid to cover the most painful skin segments up to 400 cm.sup.2. As in the Placebo Run-in Period, subjects continued to record their pain scores at 4 specific times each day and report their scores twice a day using the CTIVRS. The use of any rescue medication was recorded in a rescue medication diary. Subjects were assessed for safety, signs of local irritation, the presence of numbness (decreased sensitization), and the presence of allodynia (pain induced by a gentle brush touch or other pressure or light touch) at the beginning and end of each Treatment Period. In addition, subjects provided a weekly blood sample for pharmacokinetic (PK) analyses during both treatment and washout periods.
[0217] Upon completing Treatment Period 1, subjects began a 1-week washout prior to starting the second period of the crossover study. During the washout, subjects did not apply any study medication (placebo or Compound A) but continued to record pain scores using the CTIVRS. The use of any rescue medication was recorded.
[0218] After the washout, subjects entered the second single-blind, Placebo Run-in Period, which was followed by Treatment Period 2. Subjects returned for a follow-up visit 1 week after completing Treatment Period 2.
[0219] Duration of treatment was 42 days of randomized treatment (21 days of placebo and 21 days of Compound A) plus 14 days of placebo treatment (a 1-week, Placebo Run-in Period prior to each Treatment Period). A total of 54 subjects were used in the study after initial screening procedures were completed.
[0220] The population for this study included male and female subjects between 18 and 80 years of age with persistent pain for more than 6 months from the appearance of herpes zoster rash that was not located on the face, above the hairline of the scalp, and/or in proximity to mucous membranes. In this study, after initial evaluation and examination, of the 52 subjects whose DNA was studied (see Table 6 above), 45 subjects were found efficacy evaluable based on results with DNA and consent to participate further. Of these, 37 had the wild type allele (see SEQ ID NO: 4) and 8 had the R1150W variant (see SEQ ID NO: 2). These 45 subjects were referred to as the "efficacy evaluable population."
[0221] Compound A was administered as an 8% ointment formulated for topical administration. The placebo ointment contained identical excipients and was identical in appearance to Compound A ointment.
[0222] The primary efficacy parameter was change in mean daily pain score from baseline, measured on an 11-point Likert NRS, between the last week of Compound A treatment and the last week of placebo treatment for each subject.
[0223] Secondary efficacy parameters included a change in mean daily pain score, measured on an 11-point Likert NRS, from baseline to the first, second, and third week of the Treatment Period and the proportion of subjects achieving 1-point improvement on the 11-point Likert NRS during treatment with Compound A compared to placebo. Also measured were the proportion of subjects achieving at least 50% improvement in mean daily pain score during treatment compared to placebo as well as the proportion of subjects achieving at least 30% improvement in mean daily pain score during Compound A treatment compared to placebo.
[0224] Safety assessments included adverse events, clinical laboratory measurements, electrocardiograms (ECGs), vital signs (including body weight), and physical examinations.
[0225] The primary efficacy variable was the change in mean daily pain score from baseline to the last week (Week 3) of the Treatment Period. Baseline for Treatment Period 1 was the average of the 7 mean daily pain measurements prior to Randomization and baseline for Treatment Period 2 was the average of 7 mean daily pain measurements.
[0226] The primary efficacy hypothesis was that the mean change in mean daily pain scores from baseline to the end of the Treatment Period would differ between Compound A and placebo treatment. This hypothesis was tested using a mixed effects analysis of covariance (ANCOVA) model with treatment, period, and treatment sequence as fixed effects, subject within treatment sequence as a random effect, and the baseline pain score as a covariate. The least-squares means, standard errors, and the 2-tailed 95% confidence intervals for each treatment group were presented.
[0227] The secondary efficacy parameter of change in mean daily pain scores from baseline to Week 1, Week 2, Week 2 with LOCF, Week 3, and over the entire Treatment Period (Weeks 1-3 combined) was analyzed in a similar manner as the primary efficacy variable.
[0228] All other efficacy variables were summarized using similar methods as for the primary efficacy variable. Efficacy variables were summarized by area of application (5100 cm.sup.2, 101 cm.sup.2 to 200 cm.sup.2, 201 cm.sup.2 to 300 cm.sup.2, and 301 cm.sup.2 to 400 cm.sup.2). Other subgroup analyses (such as demographic variables, disease duration, concomitant medication, etc.) for efficacy variables were performed, as needed. An additional analysis was performed in which all pain scores collected during weeks when subject study medication utilization was <80% or >125% were excluded.
[0229] Blood samples taken at Visits 3 through 12 were assayed for Compound A levels using a validated bioanalytical method. These values were summarized for each visit during the Treatment Period in which the subject was treated with Compound A.
[0230] The planned efficacy analysis of the primary endpoint of change in mean daily pain score, measured on an 11-point Likert NRS, did not result in statistically significant or clinically meaningful differences between Compound A and placebo treatments at any time point. However, the planned secondary analyses of the proportions of subjects achieving at least 50% improvement in mean daily pain score demonstrated a statistically significant difference between Compound A and placebo. In the Efficacy Evaluable Population, more subjects achieved at least 50% improvement with Compound A than with placebo at Week 3 with LOCF (p=0.0039). A similar trend was seen in the proportion of subjects achieving at least 30% improvement at Week 3 with LOCF (p=0.0784). In addition, the difference between treatments in the proportion of subjects achieving at least 30% improvement overall was statistically significant (p=0.0213). Results were similar for the Per-Protocol Population, with the proportions of subjects achieving at least 30% and at least 50% improvement at Week 3 with LOCF achieving statistical significance (p=0.0490 and p=0.0078, respectively). The differences between treatments in the proportion of subjects achieving 1-point improvement in mean daily pain score were not statistically significant for either population.
[0231] In addition, post hoc responder analyses were produced to assess the effect of the Treatment Period on improvement in mean daily pain score. Although there were no clear differences between treatments during Treatment Period 1, subjects receiving Compound A showed greater improvement in mean daily pain score compared to subjects receiving placebo during Treatment Period 2. Similarly, during Treatment Period 2, more subjects achieved at least 30% (p=0.0154) and at least 50% (p=0.0122) improvement in pain score after 3 weeks of Compound A treatment compared to placebo (such subjects were dubbed "responders").
[0232] Overall, there was less rescue medication usage with Compound A treatment than with placebo treatment. The mean total number of pills of rescue medication taken was lower with Compound A treatment compared with placebo treatment (24.8 pills versus 29.2 pills).
[0233] NPSI score analysis showed greater decreases in the overall score with Compound A treatment than with placebo treatment, but the difference between treatments was not statistically significant. There were no apparent differences between treatments in change from baseline to Week 3 with LOCF in mean DSIS score and Patient Global Impression of Change. However, results of exploratory subgroup analyses indicated a trend towards more favorable outcomes on Compound A treatment.
[0234] As expected, PK analyses demonstrated a low systemic exposure with the current formulation and dosing regimen of Compound A. Throughout Compound A treatment, approximately 70% of subjects had Compound A levels above the detectable limit, but these levels were significantly lower than those previously shown to mediate a systemic analgesic effect. Low residual plasma concentrations were measured during placebo treatment for some subjects whose first treatment was Compound A. However, the clinical efficacy data do not indicate that these very low levels interfered with the pain intensity results of the study. The placebo response was minimal for those subjects who were dosed with placebo in Treatment Period 2 (the Compound A/placebo sequence) with only 1 subject demonstrating a 50% or greater improvement and 3 subjects demonstrating a 30% or greater improvement.
[0235] Compound A (in ointment) was safe and well tolerated in subjects with PHN. The overall incidence of TEAEs (Treatment-Emergent Adverse Events) was 50.8% with placebo treatment and 53.2% with Compound A treatment. The overall incidence of study medication-related TEAEs was 30.2% with placebo treatment and 17.7% with Compound A treatment. The majority of subjects experienced TEAEs that were considered by the Investigator to be mild in severity. Only 3 (4.4%) subjects experienced at least 1 severe TEAE (2 [3.2%] subjects during placebo treatment and 1 [1.6%] subject during Compound A treatment).
[0236] For the efficacy evaluable population, there was a greater mean improvement in R1150W carriers than in wildtype subjects as shown in Table 7:
TABLE-US-00008 TABLE 7 Mean Improvement in R1150W Carriers Compound A Pacebo Subject Base Change % Change Base Change % Change 001-002 0.93 -0.89 -96.15 4.43 -2.69 -60.75 005-008 4.25 -0.82 -19.33 3.68 -0.43 -11.65 005-014 3.28 -1.57 -47.76 4.07 -0.92 -22.48 014-002 8.50 0.36 4.20 5.86 -1.36 -23.17 014-006 7.00 0.00 0.00 6.50 0.54 8.24 016-014 5.46 -3.57 -65.36 8.22 -2.03 -24.68 016-019 9.25 -4.00 -43.24 1.25 0.06 4.76 017-002 4.18 -3.14 -75.21 2.49 -0.55 -22.13
[0237] In general, application site reactions were more common with placebo treatment than with Compound A treatment, suggesting that the presence of Compound A in the ointment may have some local protective or relieving effect.
[0238] Compound A treatment resulted in statistically significant proportions of subjects achieving at least 50% improvement in mean pain score compared to placebo.
[0239] The mean change in the above pain scores was as in Table 8:
TABLE-US-00009 TABLE 8 Mean Change in Pain Score: SCN9A Compound A Placebo Difference in Mean R1150W -1.70 -0.92 -0.78 Wildtype -0.73 -1.04 -0.31
[0240] Of the >30% responders to Compound A, 62.5% exhibited the R1150W variant versus 35.1% for the wildtype. Comparison of % response in subjects with the R1150W variant versus wildtype gave the following results:
TABLE-US-00010 TABLE 9 R1150W versus WT Responders Non-Responders Pla- Pla- Gene Cpd A % cebo % Cpd A % cebo % R1150W 5/18 27.8 1/9 11.1 3/27 11.1 7/36 19.4 Wildtype 13/18 72.2 8/9 88.9 24/27 88.9 29/36 80.4
[0241] Thus, for those subjects showing >30% improvement in pain score after 3 weeks (responders), the percentage of subjects with the R1150W variant was much greater than that for the placebo treated group whereas for the wildtype, the percentages were similar, although placebo was higher. This clearly shows the increase in successful treatment of subjects with the R1150W variant using Compound A (relative to placebo) versus wildtype.
REFERENCE LIST
[0242] 1. Claes, L. et al. De novo mutations in the sodium-channel gene SCN1A cause severe myoclonic epilepsy of infancy. Am. J. Hum. Genet. 68, 1327-1332 (2001).
[0243] Rouleau, G., et al. Loci for idiopathic generalized epilepsy, mapping to chromosome 2, mutations thereof and method using the same to assess, diagnose, prognose or treat epilepsy. PCT Patent Publication WO 01/38564 published May 31, 2001, priority date Nov. 25, 1999. Also see: Escayg, A. et al. Mutations of SCN1A, encoding a neuronal sodium channel, in two families with GEFS+2. Nat. Genet. 24, 343-345 (2000).
[0244] 3. Lossin, C., Wang, D. W., Rhodes, T. H., Vanoye, C. G., & George, A. L., Jr. Molecular basis of an inherited epilepsy. Neuron 34, 877-884 (2002).
[0245] 4. Planells-Cases, R. et al. Neuronal death and perinatal lethality in voltage-gated sodium channel alpha(II)-deficient mice. Biophys. J. 78, 2878-2891 (2000).
[0246] 5. Burgess, D. L. et al. Mutation of a new sodium channel gene, Scn8a, in the mouse mutant `motor endplate disease`. Nat. Genet. 10, 461-465 (1995).
[0247] 6. Kohrman, D. C., Harris, J. B., & Meisler, M. H. Mutation detection in the med and medJ alleles of the sodium channel Scn8a. Unusual splicing due to a minor class AT-AC intron. J. Biol. Chem. 271, 17576-17581 (1996).
[0248] 7. Kohrman, D. C., Smith, M. R., Goldin, A. L., Harris, J., & Meisler, M. H. A missense mutation in the sodium channel Scn8a is responsible for cerebellar ataxia in the mouse mutant jolting. J. Neurosci. 16, 5993-5999 (1996).
[0249] 8. De Repentigny, Y. et al. Pathological and genetic analysis of the degenerating muscle (dmu) mouse: a new allele of Scn8a. Hum. Mol. Genet. 10, 1819-1827 (2001).
[0250] 9. Sprunger, L. K., Escayg, A., Tallaksen-Greene, S., Albin, R. L., & Meisler, M. H. Dystonia associated with mutation of the neuronal sodium channel Scn8a and identification of the modifier locus Scnm1 on mouse chromosome 3. Hum. Mol. Genet. 8, 471-479 (1999).
[0251] 10. Rojas, C. V. et al. A Met-to-Val mutation in the skeletal muscle Na+ channel alpha-subunit in hyperkalaemic periodic paralysis. Nature 354, 387-389 (1991).
[0252] 11. Bendahhou, S., Cummins, T. R., Kula, R. W., Fu, Y. H., & Ptacek, L. J. Impairment of slow inactivation as a common mechanism for periodic paralysis in DIIS4-S5. Neurology 58, 1266-1272 (2002).
[0253] 12. Nuyens, D. et al. Abrupt rate accelerations or premature beats cause life-threatening arrhythmias in mice with long-QT3 syndrome. Nat. Med. 7, 1021-1027 (2001).
[0254] 13. Papadatos, G. A. et al. Slowed conduction and ventricular tachycardia after targeted disruption of the cardiac sodium channel gene Scn5a. Proc. Natl. Acad. Sci. U.S.A 99, 6210-6215 (2002).
[0255] 14. Wang, Q. et al. SCN5A mutations associated with an inherited cardiac arrhythmia, long QT syndrome. Cell 80, 805-811 (1995).
[0256] 15. Chen, Q. et al. Genetic basis and molecular mechanism for idiopathic ventricular fibrillation. Nature 392, 293-296 (1998).
[0257] 16. Schott, J. J. et al. Cardiac conduction defects associate with mutations in SCN5A. Nat. Genet. 23, 20-21 (1999).
[0258] 17. Bennett, P. B., Yazawa, K., Makita, N., & George, A. L., Jr. Molecular mechanism for an inherited cardiac arrhythmia. Nature 376, 683-685 (1995).
[0259] 18. Akopian, A. N. et al. The tetrodotoxin-resistant sodium channel SNS has a specialized function in pain pathways. Nat. Neurosci. 2, 541-548 (1999).
[0260] 19. Lai, J. et al. Inhibition of neuropathic pain by decreased expression of the tetrodotoxin-resistant sodium channel, NaV1.8. Pain 95, 143-152 (2002).
[0261] 20. Khasar, S. G., Gold, M. S., & Levine, J. D. A tetrodotoxin-resistant sodium current mediates inflammatory pain in the rat. Neurosci. Lett. 256, 17-20 (1998).
[0262] 21. Laird, J. M Souslova, V., Wood, J. N., & Cervero, F. Deficits in visceral pain and referred hyperalgesia in Nav1.8 (SNS/PN3)-null mice. J. Neurosci. 22, 8352-8356 (2002).
[0263] 22. Kerr, B. J., Souslova, V., McMahon, S. B., & Wood, J. N. A role for the TTX-resistant sodium channel Nav 1.8 in NGF-induced hyperalgesia, but not neuropathic pain. Neuroreport 12, 3077-3080 (2001).
[0264] 23. Porreca, F. et al. A comparison of the potential role of the tetrodotoxin-insensitive sodium channels, PN3/SNS and NaN/SNS2, in rat models of chronic pain. Proc. Natl. Acad. Sci. U.S.A 96, 7640-7644 (1999).
[0265] 24. Watanabe, E. et al. Nav2/NaG channel is involved in control of salt-intake behavior in the CNS. J. Neurosci. 20, 7743-7751 (2000).
[0266] 25. Sugawara, T. et al. A missense mutation of the Na+ channel alpha II subunit gene Na(v)1.2 in a patient with febrile and afebrile seizures causes channel dysfunction. Proc. Natl. Acad. Sci. U.S.A 98, 6384-6389 (2001).
[0267] 26. Akai, J. et al. A novel SCN5A mutation associated with idiopathic ventricular fibrillation without typical ECG findings of Brugada syndrome. FEBS. Lett. 479, 29-34 (2000).
Sequence CWU
1
1
12216367DNAHomo sapiens 1ctcttatgtg aggagctgaa gaggaattaa aatatacagg
atgaaaagat ggcaatgttg 60cctcccccag gacctcagag ctttgtccat ttcacaaaac
agtctcttgc cctcattgaa 120caacgcattg ctgaaagaaa atcaaaggaa cccaaagaag
aaaagaaaga tgatgatgaa 180gaagccccaa agccaagcag tgacttggaa gctggcaaac
agctgccctt catctatggg 240gacattcctc ccggcatggt gtcagagccc ctggaggact
tggaccccta ctatgcagac 300aaaaagactt tcatagtatt gaacaaaggg aaaacaatct
tccgtttcaa tgccacacct 360gctttatata tgctttctcc tttcagtcct ctaagaagaa
tatctattaa gattttagta 420cactccttat tcagcatgct catcatgtgc actattctga
caaactgcat atttatgacc 480atgaataacc caccggactg gaccaaaaat gtcgagtaca
cttttactgg aatatatact 540tttgaatcac ttgtaaaaat ccttgcaaga ggcttctgtg
taggagaatt cacttttctt 600cgtgacccgt ggaactggct ggattttgtc gtcattgttt
ttgcgtattt aacagaattt 660gtaaacctag gcaatgtttc agctcttcga actttcagag
tattgagagc tttgaaaact 720atttctgtaa tcccaggcct gaagacaatt gtaggggctt
tgatccagtc agtgaagaag 780ctttctgatg tcatgatcct gactgtgttc tgtctgagtg
tgtttgcact aattggacta 840cagctgttca tgggaaacct gaagcataaa tgttttcgaa
attcacttga aaataatgaa 900acattagaaa gcataatgaa taccctagag agtgaagaag
actttagaaa atatttttat 960tacttggaag gatccaaaga tgctctcctt tgtggtttca
gcacagattc aggtcagtgt 1020ccagaggggt acacctgtgt gaaaattggc agaaaccctg
attatggcta cacgagcttt 1080gacactttca gctgggcctt cttagccttg tttaggctaa
tgacccaaga ttactgggaa 1140aacctttacc aacagacgct gcgtgctgct ggcaaaacct
acatgatctt ctttgtcgta 1200gtgattttcc tgggctcctt ttatctaata aacttgatcc
tggctgtggt tgccatggca 1260tatgaagaac agaaccaggc aaacattgaa gaagctaaac
agaaagaatt agaatttcaa 1320cagatgttag accgtcttaa aaaagagcaa gaagaagctg
aggcaattgc agcggcagcg 1380gctgaatata caagtattag gagaagcaga attatgggcc
tctcagagag ttcttctgaa 1440acatccaaac tgagctctaa aagtgctaaa gaaagaagaa
acagaagaaa gaaaaagaat 1500caaaagaagc tctccagtgg agaggaaaag ggagatgctg
agaaattgtc gaaatcagaa 1560tcagaggaca gcatcagaag aaaaagtttc caccttggtg
tcgaagggca taggcgagca 1620catgaaaaga ggttgtctac ccccaatcag tcaccactca
gcattcgtgg ctccttgttt 1680tctgcaaggc gaagcagcag aacaagtctt tttagtttca
aaggcagagg aagagatata 1740ggatctgaga ctgaatttgc cgatgatgag cacagcattt
ttggagacaa tgagagcaga 1800aggggctcac tgtttgtgcc ccacagaccc caggagcgac
gcagcagtaa catcagccaa 1860gccagtaggt ccccaccaat gctgccggtg aacgggaaaa
tgcacagtgc tgtggactgc 1920aacggtgtgg tctccctggt tgatggacgc tcagccctca
tgctccccaa tggacagctt 1980ctgccagagg gcacgaccaa tcaaatacac aagaaaaggc
gttgtagttc ctatctcctt 2040tcagaggata tgctgaatga tcccaacctc agacagagag
caatgagtag agcaagcata 2100ttaacaaaca ctgtggaaga acttgaagag tccagacaaa
aatgtccacc ttggtggtac 2160agatttgcac acaaattctt gatctggaat tgctctccat
attggataaa attcaaaaag 2220tgtatctatt ttattgtaat ggatcctttt gtagatcttg
caattaccat ttgcatagtt 2280ttaaacacat tatttatggc tatggaacac cacccaatga
ctgaggaatt caaaaatgta 2340cttgctatag gaaatttggt ctttactgga atctttgcag
ctgaaatggt attaaaactg 2400attgccatgg atccatatga gtatttccaa gtaggctgga
atatttttga cagccttatt 2460gtgactttaa gtttagtgga gctctttcta gcagatgtgg
aaggattgtc agttctgcga 2520tcattcagac tgctccgagt cttcaagttg gcaaaatcct
ggccaacatt gaacatgctg 2580attaagatca ttggtaactc agtaggggct ctaggtaacc
tcaccttagt gttggccatc 2640atcgtcttca tttttgctgt ggtcggcatg cagctctttg
gtaagagcta caaagaatgt 2700gtctgcaaga tcaatgatga ctgtacgctc ccacggtggc
acatgaacga cttcttccac 2760tccttcctga ttgtgttccg cgtgctgtgt ggagagtgga
tagagaccat gtgggactgt 2820atggaggtcg ctggtcaagc tatgtgcctt attgtttaca
tgatggtcat ggtcattgga 2880aacctggtgg tcctaaacct atttctggcc ttattattga
gctcatttag ttcagacaat 2940cttacagcaa ttgaagaaga ccctgatgca aacaacctcc
agattgcagt gactagaatt 3000aaaaagggaa taaattatgt gaaacaaacc ttacgtgaat
ttattctaaa agcattttcc 3060aaaaagccaa agatttccag ggagataaga caagcagaag
atctgaatac taagaaggaa 3120aactatattt ctaaccatac acttgctgaa atgagcaaag
gtcacaattt cctcaaggaa 3180aaagataaaa tcagtggttt tggaagcagc gtggacaaac
acttgatgga agacagtgat 3240ggtcaatcat ttattcacaa tcccagcctc acagtgacag
tgccaattgc acctggggaa 3300tccgatttgg aaaatatgaa tgctgaggaa cttagcagtg
attcggatag tgaatacagc 3360aaagtgagat taaaccggtc aagctcctca gagtgcagca
cagttgataa ccctttgcct 3420ggagaaggag aagaagcaga ggctgaacct atgaattccg
atgagccaga ggcctgtttc 3480acagatggtt gtgtatggag gttctcatgc tgccaagtta
acatagagtc agggaaagga 3540aaaatctggt ggaacatcag gaaaacctgc tacaagattg
ttgaacacag ttggtttgaa 3600agcttcattg tcctcatgat cctgctcagc agtggtgccc
tggcttttga agatatttat 3660attgaaagga aaaagaccat taagattatc ctggagtatg
cagacaagat cttcacttac 3720atcttcattc tggaaatgct tctaaaatgg atagcatatg
gttataaaac atatttcacc 3780aatgcctggt gttggctgga tttcctaatt gttgatgttt
ctttggttac tttagtggca 3840aacactcttg gctactcaga tcttggcccc attaaatccc
ttcggacact gagagcttta 3900agacctctaa gagccttatc tagatttgaa ggaatgaggg
tcgttgtgaa tgcactcata 3960ggagcaattc cttccatcat gaatgtgcta cttgtgtgtc
ttatattctg gctgatattc 4020agcatcatgg gagtaaattt gtttgctggc aagttctatg
agtgtattaa caccacagat 4080gggtcacggt ttcctgcaag tcaagttcca aatcgttccg
aatgttttgc ccttatgaat 4140gttagtcaaa atgtgcgatg gaaaaacctg aaagtgaact
ttgataatgt cggacttggt 4200tacctatctc tgcttcaagt tgcaactttt aagggatgga
cgattattat gtatgcagca 4260gtggattctg ttaatgtaga caagcagccc aaatatgaat
atagcctcta catgtatatt 4320tattttgtcg tctttatcat ctttgggtca ttcttcactt
tgaacttgtt cattggtgtc 4380atcatagata atttcaacca acagaaaaag aagcttggag
gtcaagacat ctttatgaca 4440gaagaacaga agaaatacta taatgcaatg aaaaagctgg
ggtccaagaa gccacaaaag 4500ccaattcctc gaccagggaa caaaatccaa ggatgtatat
ttgacctagt gacaaatcaa 4560gcctttgata ttagtatcat ggttcttatc tgtctcaaca
tggtaaccat gatggtagaa 4620aaggagggtc aaagtcaaca tatgactgaa gttttatatt
ggataaatgt ggtttttata 4680atccttttca ctggagaatg tgtgctaaaa ctgatctccc
tcagacacta ctacttcact 4740gtaggatgga atatttttga ttttgtggtt gtgattatct
ccattgtagg tatgtttcta 4800gctgatttga ttgaaacgta ttttgtgtcc cctaccctgt
tccgagtgat ccgtcttgcc 4860aggattggcc gaatcctacg tctagtcaaa ggagcaaagg
ggatccgcac gctgctcttt 4920gctttgatga tgtcccttcc tgcgttgttt aacatcggcc
tcctgctctt cctggtcatg 4980ttcatctacg ccatctttgg aatgtccaac tttgcctatg
ttaaaaagga agatggaatt 5040aatgacatgt tcaattttga gacctttggc aacagtatga
tttgcctgtt ccaaattaca 5100acctctgctg gctgggatgg attgctagca cctattctta
acagtaagcc acccgactgt 5160gacccaaaaa aagttcatcc tggaagttca gttgaaggag
actgtggtaa cccatctgtt 5220ggaatattct actttgttag ttatatcatc atatccttcc
tggttgtggt gaacatgtac 5280attgcagtca tactggagaa ttttagtgtt gccactgaag
aaagtactga acctctgagt 5340gaggatgact ttgagatgtt ctatgaggtt tgggagaagt
ttgatcccga tgcgacccag 5400tttatagagt tctctaaact ctctgatttt gcagctgccc
tggatcctcc tcttctcata 5460gcaaaaccca acaaagtcca gctcattgcc atggatctgc
ccatggttag tggtgaccgg 5520atccattgtc ttgacatctt atttgctttt acaaagcgtg
ttttgggtga gagtggggag 5580atggattctc ttcgttcaca gatggaagaa aggttcatgt
ctgcaaatcc ttccaaagtg 5640tcctatgaac ccatcacaac cacactaaaa cggaaacaag
aggatgtgtc tgctactgtc 5700attcagcgtg cttatagacg ttaccgctta aggcaaaatg
tcaaaaatat atcaagtata 5760tacataaaag atggagacag agatgatgat ttactcaata
aaaaagatat ggcttttgat 5820aatgttaatg agaactcaag tccagaaaaa acagatgcca
cttcatccac cacctctcca 5880ccttcatatg atagtgtaac aaagccagac aaagagaaat
atgaacaaga cagaacagaa 5940aaggaagaca aagggaaaga cagcaaggaa agcaaaaaat
agagcttcat ttttgatata 6000ttgtttacag cctgtgaaag tgatttattt gtgttaataa
aactcttttg aggaagtcta 6060tgccaaaatc ctttttatca aaatattctc gaaggcagtg
cagtcactaa ctctgatttc 6120ctaagaaagg tgggcagcat tagcagatgg ttatttttgc
actgatgatt ctttaagaat 6180cgtaagagaa ctctgtagga attattgatt atagcataca
aaagtgattc agttttttgg 6240tttttaataa atcagaagac catgtagaaa acttttacat
ctgccttgtc atcttttcac 6300aggattgtaa ttagtcttgt ttcccatgta aataaacaac
acacgcatac agaaaaaaaa 6360aaaaaaa
636721977PRTHomo sapiens 2Met Ala Met Leu Pro Pro
Pro Gly Pro Gln Ser Phe Val His Phe Thr 1 5
10 15 Lys Gln Ser Leu Ala Leu Ile Glu Gln Arg Ile
Ala Glu Arg Lys Ser 20 25 30
Lys Glu Pro Lys Glu Glu Lys Lys Asp Asp Asp Glu Glu Ala Pro Lys
35 40 45 Pro Ser
Ser Asp Leu Glu Ala Gly Lys Gln Leu Pro Phe Ile Tyr Gly 50
55 60 Asp Ile Pro Pro Gly Met Val
Ser Glu Pro Leu Glu Asp Leu Asp Pro 65 70
75 80 Tyr Tyr Ala Asp Lys Lys Thr Phe Ile Val Leu Asn
Lys Gly Lys Thr 85 90
95 Ile Phe Arg Phe Asn Ala Thr Pro Ala Leu Tyr Met Leu Ser Pro Phe
100 105 110 Ser Pro Leu
Arg Arg Ile Ser Ile Lys Ile Leu Val His Ser Leu Phe 115
120 125 Ser Met Leu Ile Met Cys Thr Ile
Leu Thr Asn Cys Ile Phe Met Thr 130 135
140 Met Asn Asn Pro Pro Asp Trp Thr Lys Asn Val Glu Tyr
Thr Phe Thr 145 150 155
160 Gly Ile Tyr Thr Phe Glu Ser Leu Val Lys Ile Leu Ala Arg Gly Phe
165 170 175 Cys Val Gly Glu
Phe Thr Phe Leu Arg Asp Pro Trp Asn Trp Leu Asp 180
185 190 Phe Val Val Ile Val Phe Ala Tyr Leu
Thr Glu Phe Val Asn Leu Gly 195 200
205 Asn Val Ser Ala Leu Arg Thr Phe Arg Val Leu Arg Ala Leu
Lys Thr 210 215 220
Ile Ser Val Ile Pro Gly Leu Lys Thr Ile Val Gly Ala Leu Ile Gln 225
230 235 240 Ser Val Lys Lys Leu
Ser Asp Val Met Ile Leu Thr Val Phe Cys Leu 245
250 255 Ser Val Phe Ala Leu Ile Gly Leu Gln Leu
Phe Met Gly Asn Leu Lys 260 265
270 His Lys Cys Phe Arg Asn Ser Leu Glu Asn Asn Glu Thr Leu Glu
Ser 275 280 285 Ile
Met Asn Thr Leu Glu Ser Glu Glu Asp Phe Arg Lys Tyr Phe Tyr 290
295 300 Tyr Leu Glu Gly Ser Lys
Asp Ala Leu Leu Cys Gly Phe Ser Thr Asp 305 310
315 320 Ser Gly Gln Cys Pro Glu Gly Tyr Thr Cys Val
Lys Ile Gly Arg Asn 325 330
335 Pro Asp Tyr Gly Tyr Thr Ser Phe Asp Thr Phe Ser Trp Ala Phe Leu
340 345 350 Ala Leu
Phe Arg Leu Met Thr Gln Asp Tyr Trp Glu Asn Leu Tyr Gln 355
360 365 Gln Thr Leu Arg Ala Ala Gly
Lys Thr Tyr Met Ile Phe Phe Val Val 370 375
380 Val Ile Phe Leu Gly Ser Phe Tyr Leu Ile Asn Leu
Ile Leu Ala Val 385 390 395
400 Val Ala Met Ala Tyr Glu Glu Gln Asn Gln Ala Asn Ile Glu Glu Ala
405 410 415 Lys Gln Lys
Glu Leu Glu Phe Gln Gln Met Leu Asp Arg Leu Lys Lys 420
425 430 Glu Gln Glu Glu Ala Glu Ala Ile
Ala Ala Ala Ala Ala Glu Tyr Thr 435 440
445 Ser Ile Arg Arg Ser Arg Ile Met Gly Leu Ser Glu Ser
Ser Ser Glu 450 455 460
Thr Ser Lys Leu Ser Ser Lys Ser Ala Lys Glu Arg Arg Asn Arg Arg 465
470 475 480 Lys Lys Lys Asn
Gln Lys Lys Leu Ser Ser Gly Glu Glu Lys Gly Asp 485
490 495 Ala Glu Lys Leu Ser Lys Ser Glu Ser
Glu Asp Ser Ile Arg Arg Lys 500 505
510 Ser Phe His Leu Gly Val Glu Gly His Arg Arg Ala His Glu
Lys Arg 515 520 525
Leu Ser Thr Pro Asn Gln Ser Pro Leu Ser Ile Arg Gly Ser Leu Phe 530
535 540 Ser Ala Arg Arg Ser
Ser Arg Thr Ser Leu Phe Ser Phe Lys Gly Arg 545 550
555 560 Gly Arg Asp Ile Gly Ser Glu Thr Glu Phe
Ala Asp Asp Glu His Ser 565 570
575 Ile Phe Gly Asp Asn Glu Ser Arg Arg Gly Ser Leu Phe Val Pro
His 580 585 590 Arg
Pro Gln Glu Arg Arg Ser Ser Asn Ile Ser Gln Ala Ser Arg Ser 595
600 605 Pro Pro Met Leu Pro Val
Asn Gly Lys Met His Ser Ala Val Asp Cys 610 615
620 Asn Gly Val Val Ser Leu Val Asp Gly Arg Ser
Ala Leu Met Leu Pro 625 630 635
640 Asn Gly Gln Leu Leu Pro Glu Gly Thr Thr Asn Gln Ile His Lys Lys
645 650 655 Arg Arg
Cys Ser Ser Tyr Leu Leu Ser Glu Asp Met Leu Asn Asp Pro 660
665 670 Asn Leu Arg Gln Arg Ala Met
Ser Arg Ala Ser Ile Leu Thr Asn Thr 675 680
685 Val Glu Glu Leu Glu Glu Ser Arg Gln Lys Cys Pro
Pro Trp Trp Tyr 690 695 700
Arg Phe Ala His Lys Phe Leu Ile Trp Asn Cys Ser Pro Tyr Trp Ile 705
710 715 720 Lys Phe Lys
Lys Cys Ile Tyr Phe Ile Val Met Asp Pro Phe Val Asp 725
730 735 Leu Ala Ile Thr Ile Cys Ile Val
Leu Asn Thr Leu Phe Met Ala Met 740 745
750 Glu His His Pro Met Thr Glu Glu Phe Lys Asn Val Leu
Ala Ile Gly 755 760 765
Asn Leu Val Phe Thr Gly Ile Phe Ala Ala Glu Met Val Leu Lys Leu 770
775 780 Ile Ala Met Asp
Pro Tyr Glu Tyr Phe Gln Val Gly Trp Asn Ile Phe 785 790
795 800 Asp Ser Leu Ile Val Thr Leu Ser Leu
Val Glu Leu Phe Leu Ala Asp 805 810
815 Val Glu Gly Leu Ser Val Leu Arg Ser Phe Arg Leu Leu Arg
Val Phe 820 825 830
Lys Leu Ala Lys Ser Trp Pro Thr Leu Asn Met Leu Ile Lys Ile Ile
835 840 845 Gly Asn Ser Val
Gly Ala Leu Gly Asn Leu Thr Leu Val Leu Ala Ile 850
855 860 Ile Val Phe Ile Phe Ala Val Val
Gly Met Gln Leu Phe Gly Lys Ser 865 870
875 880 Tyr Lys Glu Cys Val Cys Lys Ile Asn Asp Asp Cys
Thr Leu Pro Arg 885 890
895 Trp His Met Asn Asp Phe Phe His Ser Phe Leu Ile Val Phe Arg Val
900 905 910 Leu Cys Gly
Glu Trp Ile Glu Thr Met Trp Asp Cys Met Glu Val Ala 915
920 925 Gly Gln Ala Met Cys Leu Ile Val
Tyr Met Met Val Met Val Ile Gly 930 935
940 Asn Leu Val Val Leu Asn Leu Phe Leu Ala Leu Leu Leu
Ser Ser Phe 945 950 955
960 Ser Ser Asp Asn Leu Thr Ala Ile Glu Glu Asp Pro Asp Ala Asn Asn
965 970 975 Leu Gln Ile Ala
Val Thr Arg Ile Lys Lys Gly Ile Asn Tyr Val Lys 980
985 990 Gln Thr Leu Arg Glu Phe Ile Leu
Lys Ala Phe Ser Lys Lys Pro Lys 995 1000
1005 Ile Ser Arg Glu Ile Arg Gln Ala Glu Asp Leu
Asn Thr Lys Lys 1010 1015 1020
Glu Asn Tyr Ile Ser Asn His Thr Leu Ala Glu Met Ser Lys Gly
1025 1030 1035 His Asn Phe
Leu Lys Glu Lys Asp Lys Ile Ser Gly Phe Gly Ser 1040
1045 1050 Ser Val Asp Lys His Leu Met Glu
Asp Ser Asp Gly Gln Ser Phe 1055 1060
1065 Ile His Asn Pro Ser Leu Thr Val Thr Val Pro Ile Ala
Pro Gly 1070 1075 1080
Glu Ser Asp Leu Glu Asn Met Asn Ala Glu Glu Leu Ser Ser Asp 1085
1090 1095 Ser Asp Ser Glu Tyr
Ser Lys Val Arg Leu Asn Arg Ser Ser Ser 1100 1105
1110 Ser Glu Cys Ser Thr Val Asp Asn Pro Leu
Pro Gly Glu Gly Glu 1115 1120 1125
Glu Ala Glu Ala Glu Pro Met Asn Ser Asp Glu Pro Glu Ala Cys
1130 1135 1140 Phe Thr
Asp Gly Cys Val Trp Arg Phe Ser Cys Cys Gln Val Asn 1145
1150 1155 Ile Glu Ser Gly Lys Gly Lys
Ile Trp Trp Asn Ile Arg Lys Thr 1160 1165
1170 Cys Tyr Lys Ile Val Glu His Ser Trp Phe Glu Ser
Phe Ile Val 1175 1180 1185
Leu Met Ile Leu Leu Ser Ser Gly Ala Leu Ala Phe Glu Asp Ile 1190
1195 1200 Tyr Ile Glu Arg Lys
Lys Thr Ile Lys Ile Ile Leu Glu Tyr Ala 1205 1210
1215 Asp Lys Ile Phe Thr Tyr Ile Phe Ile Leu
Glu Met Leu Leu Lys 1220 1225 1230
Trp Ile Ala Tyr Gly Tyr Lys Thr Tyr Phe Thr Asn Ala Trp Cys
1235 1240 1245 Trp Leu
Asp Phe Leu Ile Val Asp Val Ser Leu Val Thr Leu Val 1250
1255 1260 Ala Asn Thr Leu Gly Tyr Ser
Asp Leu Gly Pro Ile Lys Ser Leu 1265 1270
1275 Arg Thr Leu Arg Ala Leu Arg Pro Leu Arg Ala Leu
Ser Arg Phe 1280 1285 1290
Glu Gly Met Arg Val Val Val Asn Ala Leu Ile Gly Ala Ile Pro 1295
1300 1305 Ser Ile Met Asn Val
Leu Leu Val Cys Leu Ile Phe Trp Leu Ile 1310 1315
1320 Phe Ser Ile Met Gly Val Asn Leu Phe Ala
Gly Lys Phe Tyr Glu 1325 1330 1335
Cys Ile Asn Thr Thr Asp Gly Ser Arg Phe Pro Ala Ser Gln Val
1340 1345 1350 Pro Asn
Arg Ser Glu Cys Phe Ala Leu Met Asn Val Ser Gln Asn 1355
1360 1365 Val Arg Trp Lys Asn Leu Lys
Val Asn Phe Asp Asn Val Gly Leu 1370 1375
1380 Gly Tyr Leu Ser Leu Leu Gln Val Ala Thr Phe Lys
Gly Trp Thr 1385 1390 1395
Ile Ile Met Tyr Ala Ala Val Asp Ser Val Asn Val Asp Lys Gln 1400
1405 1410 Pro Lys Tyr Glu Tyr
Ser Leu Tyr Met Tyr Ile Tyr Phe Val Val 1415 1420
1425 Phe Ile Ile Phe Gly Ser Phe Phe Thr Leu
Asn Leu Phe Ile Gly 1430 1435 1440
Val Ile Ile Asp Asn Phe Asn Gln Gln Lys Lys Lys Leu Gly Gly
1445 1450 1455 Gln Asp
Ile Phe Met Thr Glu Glu Gln Lys Lys Tyr Tyr Asn Ala 1460
1465 1470 Met Lys Lys Leu Gly Ser Lys
Lys Pro Gln Lys Pro Ile Pro Arg 1475 1480
1485 Pro Gly Asn Lys Ile Gln Gly Cys Ile Phe Asp Leu
Val Thr Asn 1490 1495 1500
Gln Ala Phe Asp Ile Ser Ile Met Val Leu Ile Cys Leu Asn Met 1505
1510 1515 Val Thr Met Met Val
Glu Lys Glu Gly Gln Ser Gln His Met Thr 1520 1525
1530 Glu Val Leu Tyr Trp Ile Asn Val Val Phe
Ile Ile Leu Phe Thr 1535 1540 1545
Gly Glu Cys Val Leu Lys Leu Ile Ser Leu Arg His Tyr Tyr Phe
1550 1555 1560 Thr Val
Gly Trp Asn Ile Phe Asp Phe Val Val Val Ile Ile Ser 1565
1570 1575 Ile Val Gly Met Phe Leu Ala
Asp Leu Ile Glu Thr Tyr Phe Val 1580 1585
1590 Ser Pro Thr Leu Phe Arg Val Ile Arg Leu Ala Arg
Ile Gly Arg 1595 1600 1605
Ile Leu Arg Leu Val Lys Gly Ala Lys Gly Ile Arg Thr Leu Leu 1610
1615 1620 Phe Ala Leu Met Met
Ser Leu Pro Ala Leu Phe Asn Ile Gly Leu 1625 1630
1635 Leu Leu Phe Leu Val Met Phe Ile Tyr Ala
Ile Phe Gly Met Ser 1640 1645 1650
Asn Phe Ala Tyr Val Lys Lys Glu Asp Gly Ile Asn Asp Met Phe
1655 1660 1665 Asn Phe
Glu Thr Phe Gly Asn Ser Met Ile Cys Leu Phe Gln Ile 1670
1675 1680 Thr Thr Ser Ala Gly Trp Asp
Gly Leu Leu Ala Pro Ile Leu Asn 1685 1690
1695 Ser Lys Pro Pro Asp Cys Asp Pro Lys Lys Val His
Pro Gly Ser 1700 1705 1710
Ser Val Glu Gly Asp Cys Gly Asn Pro Ser Val Gly Ile Phe Tyr 1715
1720 1725 Phe Val Ser Tyr Ile
Ile Ile Ser Phe Leu Val Val Val Asn Met 1730 1735
1740 Tyr Ile Ala Val Ile Leu Glu Asn Phe Ser
Val Ala Thr Glu Glu 1745 1750 1755
Ser Thr Glu Pro Leu Ser Glu Asp Asp Phe Glu Met Phe Tyr Glu
1760 1765 1770 Val Trp
Glu Lys Phe Asp Pro Asp Ala Thr Gln Phe Ile Glu Phe 1775
1780 1785 Ser Lys Leu Ser Asp Phe Ala
Ala Ala Leu Asp Pro Pro Leu Leu 1790 1795
1800 Ile Ala Lys Pro Asn Lys Val Gln Leu Ile Ala Met
Asp Leu Pro 1805 1810 1815
Met Val Ser Gly Asp Arg Ile His Cys Leu Asp Ile Leu Phe Ala 1820
1825 1830 Phe Thr Lys Arg Val
Leu Gly Glu Ser Gly Glu Met Asp Ser Leu 1835 1840
1845 Arg Ser Gln Met Glu Glu Arg Phe Met Ser
Ala Asn Pro Ser Lys 1850 1855 1860
Val Ser Tyr Glu Pro Ile Thr Thr Thr Leu Lys Arg Lys Gln Glu
1865 1870 1875 Asp Val
Ser Ala Thr Val Ile Gln Arg Ala Tyr Arg Arg Tyr Arg 1880
1885 1890 Leu Arg Gln Asn Val Lys Asn
Ile Ser Ser Ile Tyr Ile Lys Asp 1895 1900
1905 Gly Asp Arg Asp Asp Asp Leu Leu Asn Lys Lys Asp
Met Ala Phe 1910 1915 1920
Asp Asn Val Asn Glu Asn Ser Ser Pro Glu Lys Thr Asp Ala Thr 1925
1930 1935 Ser Ser Thr Thr Ser
Pro Pro Ser Tyr Asp Ser Val Thr Lys Pro 1940 1945
1950 Asp Lys Glu Lys Tyr Glu Gln Asp Arg Thr
Glu Lys Glu Asp Lys 1955 1960 1965
Gly Lys Asp Ser Lys Glu Ser Lys Lys 1970
1975 36367DNAHomo sapiens 3ctcttatgtg aggagctgaa gaggaattaa
aatatacagg atgaaaagat ggcaatgttg 60cctcccccag gacctcagag ctttgtccat
ttcacaaaac agtctcttgc cctcattgaa 120caacgcattg ctgaaagaaa atcaaaggaa
cccaaagaag aaaagaaaga tgatgatgaa 180gaagccccaa agccaagcag tgacttggaa
gctggcaaac aactgccctt catctatggg 240gacattcctc ccggcatggt gtcagagccc
ctggaggact tggaccccta ctatgcagac 300aaaaagactt tcatagtatt gaacaaaggg
aaaacaatct tccgtttcaa tgccacacct 360gctttatata tgctttctcc tttcagtcct
ctaagaagaa tatctattaa gattttagta 420cactccttat tcagcatgct catcatgtgc
actattctga caaactgcat atttatgacc 480atgaataacc cgccggactg gaccaaaaat
gtcgagtaca cttttactgg aatatatact 540tttgaatcac ttgtaaaaat ccttgcaaga
ggcttctgtg taggagaatt cacttttctt 600cgtgacccgt ggaactggct ggattttgtc
gtcattgttt ttgcatatgt gacagagttt 660gtggacctgg gcaatgtctc agcattgaga
acattcagag ttctccgagc attgaaaaca 720atttcagtca ttccaggcct gaagacaatt
gtaggggctt tgatccagtc agtgaagaag 780ctttctgatg tcatgatcct gactgtgttc
tgtctgagtg tgtttgcact aattggacta 840cagctgttca tgggaaacct gaagcataaa
tgttttcgaa attcacttga aaataatgaa 900acattagaaa gcataatgaa taccctagag
agtgaagaag actttagaaa atatttttat 960tacttggaag gatccaaaga tgctctcctt
tgtggtttca gcacagattc aggtcagtgt 1020ccagaggggt acacctgtgt gaaaattggc
agaaaccctg attatggcta cacgagcttt 1080gacactttca gctgggcctt cttagccttg
tttaggctaa tgacccaaga ttactgggaa 1140aacctttacc aacagacgct gcgtgccgct
ggcaaaacct acatgatctt ctttgtcgta 1200gtgattttcc tgggctcctt ttatctaata
aacttgatcc tggctgtggt tgccatggca 1260tatgaagaac agaaccaggc aaacattgaa
gaagctaaac agaaagaatt agagtttcaa 1320cagatgttag accgacttaa aaaagagcaa
gaagaagctg aggcaattgc agcggcagcg 1380gctgaatata caagtattag gagaagcaga
attatgggcc tctcagagag ttcttctgaa 1440acatccaaac tgagctctaa aagtgctaaa
gaaagaagaa acagaagaaa gaaaaagaat 1500caaaagaagc tctccagtgg agaggaaaag
ggagatgctg agaaattgtc gaaatcagaa 1560tcagaggaca gcatcagaag aaaaagtttc
caccttggtg tcgaagggca taggcgagca 1620catgaaaaga ggttgtctac ccccaatcag
tcaccactca gcattcgtgg ctccttgttt 1680tctgcaaggc gaagcagcag aacaagtctt
tttagtttca aaggcagagg aagagatata 1740ggatctgaga ctgaatttgc cgatgatgag
cacagcattt ttggagacaa tgagagcaga 1800aggggctcac tgtttgtgcc ccacagaccc
caggagcgac gcagcagtaa catcagccaa 1860gccagtaggt ccccaccaat gctgccggtg
aacgggaaaa tgcacagtgc tgtggactgc 1920aacggtgtgg tctccctggt tgatggacgc
tcagccctca tgctccccaa tggacagctt 1980ctgccagagg gcacgaccaa tcaaatacac
aagaaaaggc gttgtagttc ctatctcctt 2040tcagaggata tgctgaatga tcccaacctc
agacagagag caatgagtag agcaagcata 2100ttaacaaaca ctgtggaaga acttgaagag
tccagacaaa aatgtccacc ttggtggtac 2160agatttgcac acaaattctt gatctggaat
tgctctccat attggataaa attcaaaaag 2220tgtatctatt ttattgtaat ggatcctttt
gtagatcttg caattaccat ttgcatagtt 2280ttaaacacat tatttatggc tatggaacac
cacccaatga ctgaggaatt caaaaatgta 2340cttgctatag gaaatttggt ctttactgga
atctttgcag ctgaaatggt attaaaactg 2400attgccatgg atccatatga gtatttccaa
gtaggctgga atatttttga cagccttatt 2460gtgactttaa gtttagtgga gctctttcta
gcagatgtgg aaggattgtc agttctgcga 2520tcattcagac tgctccgagt cttcaagttg
gcaaaatcct ggccaacatt gaacatgctg 2580attaagatca ttggtaactc agtaggggct
ctaggtaacc tcaccttagt gttggccatc 2640atcgtcttca tttttgctgt ggtcggcatg
cagctctttg gtaagagcta caaagaatgt 2700gtctgcaaga tcaatgatga ctgtacgctc
ccacggtggc acatgaacga cttcttccac 2760tccttcctga ttgtgttccg cgtgctgtgt
ggagagtgga tagagaccat gtgggactgt 2820atggaggtcg ctggtcaagc tatgtgcctt
attgtttaca tgatggtcat ggtcattgga 2880aacctggtgg tcctaaacct atttctggcc
ttattattga gctcatttag ttcagacaat 2940cttacagcaa ttgaagaaga ccctgatgca
aacaacctcc agattgcagt gactagaatt 3000aaaaagggaa taaattatgt gaaacaaacc
ttacgtgaat ttattctaaa agcattttcc 3060aaaaagccaa agatttccag ggagataaga
caagcagaag atctgaatac taagaaggaa 3120aactatattt ctaaccatac acttgctgaa
atgagcaaag gtcacaattt cctcaaggaa 3180aaagataaaa tcagtggttt tggaagcagc
gtggacaaac acttgatgga agacagtgat 3240ggtcaatcat ttattcacaa tcccagcctc
acagtgacag tgccaattgc acctggggaa 3300tccgatttgg aaaatatgaa tgctgaggaa
cttagcagtg attcggatag tgaatacagc 3360aaagtgagat taaaccggtc aagctcctca
gagtgcagca cagttgataa ccctttgcct 3420ggagaaggag aagaagcaga ggctgaacct
atgaattccg atgagccaga ggcctgtttc 3480acagatggtt gtgtacggag gttctcatgc
tgccaagtta acatagagtc agggaaagga 3540aaaatctggt ggaacatcag gaaaacctgc
tacaagattg ttgaacacag ttggtttgaa 3600agcttcattg tcctcatgat cctgctcagc
agtggtgccc tggcttttga agatatttat 3660attgaaagga aaaagaccat taagattatc
ctggagtatg cagacaagat cttcacttac 3720atcttcattc tggaaatgct tctaaaatgg
atagcatatg gttataaaac atatttcacc 3780aatgcctggt gttggctgga tttcctaatt
gttgatgttt ctttggttac tttagtggca 3840aacactcttg gctactcaga tcttggcccc
attaaatccc ttcggacact gagagcttta 3900agacctctaa gagccttatc tagatttgaa
ggaatgaggg tcgttgtgaa tgcactcata 3960ggagcaattc cttccatcat gaatgtgcta
cttgtgtgtc ttatattctg gctgatattc 4020agcatcatgg gagtaaattt gtttgctggc
aagttctatg agtgtattaa caccacagat 4080gggtcacggt ttcctgcaag tcaagttcca
aatcgttccg aatgttttgc ccttatgaat 4140gttagtcaaa atgtgcgatg gaaaaacctg
aaagtgaact ttgataatgt cggacttggt 4200tacctatctc tgcttcaagt tgcaactttt
aagggatgga cgattattat gtatgcagca 4260gtggattctg ttaatgtaga caagcagccc
aaatatgaat atagcctcta catgtatatt 4320tattttgtcg tctttatcat ctttgggtca
ttcttcactt tgaacttgtt cattggtgtc 4380atcatagata atttcaacca acagaaaaag
aagcttggag gtcaagacat ctttatgaca 4440gaagaacaga agaaatacta taatgcaatg
aaaaagctgg ggtccaagaa gccacaaaag 4500ccaattcctc gaccagggaa caaaatccaa
ggatgtatat ttgacctagt gacaaatcaa 4560gcctttgata ttagtatcat ggttcttatc
tgtctcaaca tggtaaccat gatggtagaa 4620aaggagggtc aaagtcaaca tatgactgaa
gttttatatt ggataaatgt ggtttttata 4680atccttttca ctggagaatg tgtgctaaaa
ctgatctccc tcagacacta ctacttcact 4740gtaggatgga atatttttga ttttgtggtt
gtgattatct ccattgtagg tatgtttcta 4800gctgatttga ttgaaacgta ttttgtgtcc
cctaccctgt tccgagtgat ccgtcttgcc 4860aggattggcc gaatcctacg tctagtcaaa
ggagcaaagg ggatccgcac gctgctcttt 4920gctttgatga tgtcccttcc tgcgttgttt
aacatcggcc tcctgctctt cctggtcatg 4980ttcatctacg ccatctttgg aatgtccaac
tttgcctatg ttaaaaagga agatggaatt 5040aatgacatgt tcaattttga gacctttggc
aacagtatga tttgcctgtt ccaaattaca 5100acctctgctg gctgggatgg attgctagca
cctattctta acagtaagcc acccgactgt 5160gacccaaaaa aagttcatcc tggaagttca
gttgaaggag actgtggtaa cccatctgtt 5220ggaatattct actttgttag ttatatcatc
atatccttcc tggttgtggt gaacatgtac 5280attgcagtca tactggagaa ttttagtgtt
gccactgaag aaagtactga acctctgagt 5340gaggatgact ttgagatgtt ctatgaggtt
tgggagaagt ttgatcccga tgcgacccag 5400tttatagagt tctctaaact ctctgatttt
gcagctgccc tggatcctcc tcttctcata 5460gcaaaaccca acaaagtcca gctcattgcc
atggatctgc ccatggttag tggtgaccgg 5520atccattgtc ttgacatctt atttgctttt
acaaagcgtg ttttgggtga gagtggggag 5580atggattctc ttcgttcaca gatggaagaa
aggttcatgt ctgcaaatcc ttccaaagtg 5640tcctatgaac ccatcacaac cacactaaaa
cggaaacaag aggatgtgtc tgctactgtc 5700attcagcgtg cttatagacg ttaccgctta
aggcaaaatg tcaaaaatat atcaagtata 5760tacataaaag atggagacag agatgatgat
ttactcaata aaaaagatat ggcttttgat 5820aatgttaatg agaactcaag tccagaaaaa
acagatgcca cttcatccac cacctctcca 5880ccttcatatg atagtgtaac aaagccagac
aaagagaaat atgaacaaga cagaacagaa 5940aaggaagaca aagggaaaga cagcaaggaa
agcaaaaaat agagcttcat ttttgatata 6000ttgtttacag cctgtgaaag tgatttattt
gtgttaataa aactcttttg aggaagtcta 6060tgccaaaatc ctttttatca aaatattctc
gaaggcagtg cagtcactaa ctctgatttc 6120ctaagaaagg tgggcagcat tagcagatgg
ttatttttgc actgatgatt ctttaagaat 6180cgtaagagaa ctctgtagga attattgatt
atagcataca aaagtgattc agttttttgg 6240tttttaataa atcagaagac catgtagaaa
acttttacat ctgccttgtc atcttttcac 6300aggattgtaa ttagtcttgt ttcccatgta
aataaacaac acacgcatac agaaaaaaaa 6360aaaaaaa
636741977PRTHomo sapiens 4Met Ala Met
Leu Pro Pro Pro Gly Pro Gln Ser Phe Val His Phe Thr 1 5
10 15 Lys Gln Ser Leu Ala Leu Ile Glu
Gln Arg Ile Ala Glu Arg Lys Ser 20 25
30 Lys Glu Pro Lys Glu Glu Lys Lys Asp Asp Asp Glu Glu
Ala Pro Lys 35 40 45
Pro Ser Ser Asp Leu Glu Ala Gly Lys Gln Leu Pro Phe Ile Tyr Gly 50
55 60 Asp Ile Pro Pro
Gly Met Val Ser Glu Pro Leu Glu Asp Leu Asp Pro 65 70
75 80 Tyr Tyr Ala Asp Lys Lys Thr Phe Ile
Val Leu Asn Lys Gly Lys Thr 85 90
95 Ile Phe Arg Phe Asn Ala Thr Pro Ala Leu Tyr Met Leu Ser
Pro Phe 100 105 110
Ser Pro Leu Arg Arg Ile Ser Ile Lys Ile Leu Val His Ser Leu Phe
115 120 125 Ser Met Leu Ile
Met Cys Thr Ile Leu Thr Asn Cys Ile Phe Met Thr 130
135 140 Met Asn Asn Pro Pro Asp Trp Thr
Lys Asn Val Glu Tyr Thr Phe Thr 145 150
155 160 Gly Ile Tyr Thr Phe Glu Ser Leu Val Lys Ile Leu
Ala Arg Gly Phe 165 170
175 Cys Val Gly Glu Phe Thr Phe Leu Arg Asp Pro Trp Asn Trp Leu Asp
180 185 190 Phe Val Val
Ile Val Phe Ala Tyr Val Thr Glu Phe Val Asp Leu Gly 195
200 205 Asn Val Ser Ala Leu Arg Thr Phe
Arg Val Leu Arg Ala Leu Lys Thr 210 215
220 Ile Ser Val Ile Pro Gly Leu Lys Thr Ile Val Gly Ala
Leu Ile Gln 225 230 235
240 Ser Val Lys Lys Leu Ser Asp Val Met Ile Leu Thr Val Phe Cys Leu
245 250 255 Ser Val Phe Ala
Leu Ile Gly Leu Gln Leu Phe Met Gly Asn Leu Lys 260
265 270 His Lys Cys Phe Arg Asn Ser Leu Glu
Asn Asn Glu Thr Leu Glu Ser 275 280
285 Ile Met Asn Thr Leu Glu Ser Glu Glu Asp Phe Arg Lys Tyr
Phe Tyr 290 295 300
Tyr Leu Glu Gly Ser Lys Asp Ala Leu Leu Cys Gly Phe Ser Thr Asp 305
310 315 320 Ser Gly Gln Cys Pro
Glu Gly Tyr Thr Cys Val Lys Ile Gly Arg Asn 325
330 335 Pro Asp Tyr Gly Tyr Thr Ser Phe Asp Thr
Phe Ser Trp Ala Phe Leu 340 345
350 Ala Leu Phe Arg Leu Met Thr Gln Asp Tyr Trp Glu Asn Leu Tyr
Gln 355 360 365 Gln
Thr Leu Arg Ala Ala Gly Lys Thr Tyr Met Ile Phe Phe Val Val 370
375 380 Val Ile Phe Leu Gly Ser
Phe Tyr Leu Ile Asn Leu Ile Leu Ala Val 385 390
395 400 Val Ala Met Ala Tyr Glu Glu Gln Asn Gln Ala
Asn Ile Glu Glu Ala 405 410
415 Lys Gln Lys Glu Leu Glu Phe Gln Gln Met Leu Asp Arg Leu Lys Lys
420 425 430 Glu Gln
Glu Glu Ala Glu Ala Ile Ala Ala Ala Ala Ala Glu Tyr Thr 435
440 445 Ser Ile Arg Arg Ser Arg Ile
Met Gly Leu Ser Glu Ser Ser Ser Glu 450 455
460 Thr Ser Lys Leu Ser Ser Lys Ser Ala Lys Glu Arg
Arg Asn Arg Arg 465 470 475
480 Lys Lys Lys Asn Gln Lys Lys Leu Ser Ser Gly Glu Glu Lys Gly Asp
485 490 495 Ala Glu Lys
Leu Ser Lys Ser Glu Ser Glu Asp Ser Ile Arg Arg Lys 500
505 510 Ser Phe His Leu Gly Val Glu Gly
His Arg Arg Ala His Glu Lys Arg 515 520
525 Leu Ser Thr Pro Asn Gln Ser Pro Leu Ser Ile Arg Gly
Ser Leu Phe 530 535 540
Ser Ala Arg Arg Ser Ser Arg Thr Ser Leu Phe Ser Phe Lys Gly Arg 545
550 555 560 Gly Arg Asp Ile
Gly Ser Glu Thr Glu Phe Ala Asp Asp Glu His Ser 565
570 575 Ile Phe Gly Asp Asn Glu Ser Arg Arg
Gly Ser Leu Phe Val Pro His 580 585
590 Arg Pro Gln Glu Arg Arg Ser Ser Asn Ile Ser Gln Ala Ser
Arg Ser 595 600 605
Pro Pro Met Leu Pro Val Asn Gly Lys Met His Ser Ala Val Asp Cys 610
615 620 Asn Gly Val Val Ser
Leu Val Asp Gly Arg Ser Ala Leu Met Leu Pro 625 630
635 640 Asn Gly Gln Leu Leu Pro Glu Gly Thr Thr
Asn Gln Ile His Lys Lys 645 650
655 Arg Arg Cys Ser Ser Tyr Leu Leu Ser Glu Asp Met Leu Asn Asp
Pro 660 665 670 Asn
Leu Arg Gln Arg Ala Met Ser Arg Ala Ser Ile Leu Thr Asn Thr 675
680 685 Val Glu Glu Leu Glu Glu
Ser Arg Gln Lys Cys Pro Pro Trp Trp Tyr 690 695
700 Arg Phe Ala His Lys Phe Leu Ile Trp Asn Cys
Ser Pro Tyr Trp Ile 705 710 715
720 Lys Phe Lys Lys Cys Ile Tyr Phe Ile Val Met Asp Pro Phe Val Asp
725 730 735 Leu Ala
Ile Thr Ile Cys Ile Val Leu Asn Thr Leu Phe Met Ala Met 740
745 750 Glu His His Pro Met Thr Glu
Glu Phe Lys Asn Val Leu Ala Ile Gly 755 760
765 Asn Leu Val Phe Thr Gly Ile Phe Ala Ala Glu Met
Val Leu Lys Leu 770 775 780
Ile Ala Met Asp Pro Tyr Glu Tyr Phe Gln Val Gly Trp Asn Ile Phe 785
790 795 800 Asp Ser Leu
Ile Val Thr Leu Ser Leu Val Glu Leu Phe Leu Ala Asp 805
810 815 Val Glu Gly Leu Ser Val Leu Arg
Ser Phe Arg Leu Leu Arg Val Phe 820 825
830 Lys Leu Ala Lys Ser Trp Pro Thr Leu Asn Met Leu Ile
Lys Ile Ile 835 840 845
Gly Asn Ser Val Gly Ala Leu Gly Asn Leu Thr Leu Val Leu Ala Ile 850
855 860 Ile Val Phe Ile
Phe Ala Val Val Gly Met Gln Leu Phe Gly Lys Ser 865 870
875 880 Tyr Lys Glu Cys Val Cys Lys Ile Asn
Asp Asp Cys Thr Leu Pro Arg 885 890
895 Trp His Met Asn Asp Phe Phe His Ser Phe Leu Ile Val Phe
Arg Val 900 905 910
Leu Cys Gly Glu Trp Ile Glu Thr Met Trp Asp Cys Met Glu Val Ala
915 920 925 Gly Gln Ala Met
Cys Leu Ile Val Tyr Met Met Val Met Val Ile Gly 930
935 940 Asn Leu Val Val Leu Asn Leu Phe
Leu Ala Leu Leu Leu Ser Ser Phe 945 950
955 960 Ser Ser Asp Asn Leu Thr Ala Ile Glu Glu Asp Pro
Asp Ala Asn Asn 965 970
975 Leu Gln Ile Ala Val Thr Arg Ile Lys Lys Gly Ile Asn Tyr Val Lys
980 985 990 Gln Thr Leu
Arg Glu Phe Ile Leu Lys Ala Phe Ser Lys Lys Pro Lys 995
1000 1005 Ile Ser Arg Glu Ile Arg
Gln Ala Glu Asp Leu Asn Thr Lys Lys 1010 1015
1020 Glu Asn Tyr Ile Ser Asn His Thr Leu Ala Glu
Met Ser Lys Gly 1025 1030 1035
His Asn Phe Leu Lys Glu Lys Asp Lys Ile Ser Gly Phe Gly Ser
1040 1045 1050 Ser Val Asp
Lys His Leu Met Glu Asp Ser Asp Gly Gln Ser Phe 1055
1060 1065 Ile His Asn Pro Ser Leu Thr Val
Thr Val Pro Ile Ala Pro Gly 1070 1075
1080 Glu Ser Asp Leu Glu Asn Met Asn Ala Glu Glu Leu Ser
Ser Asp 1085 1090 1095
Ser Asp Ser Glu Tyr Ser Lys Val Arg Leu Asn Arg Ser Ser Ser 1100
1105 1110 Ser Glu Cys Ser Thr
Val Asp Asn Pro Leu Pro Gly Glu Gly Glu 1115 1120
1125 Glu Ala Glu Ala Glu Pro Met Asn Ser Asp
Glu Pro Glu Ala Cys 1130 1135 1140
Phe Thr Asp Gly Cys Val Arg Arg Phe Ser Cys Cys Gln Val Asn
1145 1150 1155 Ile Glu
Ser Gly Lys Gly Lys Ile Trp Trp Asn Ile Arg Lys Thr 1160
1165 1170 Cys Tyr Lys Ile Val Glu His
Ser Trp Phe Glu Ser Phe Ile Val 1175 1180
1185 Leu Met Ile Leu Leu Ser Ser Gly Ala Leu Ala Phe
Glu Asp Ile 1190 1195 1200
Tyr Ile Glu Arg Lys Lys Thr Ile Lys Ile Ile Leu Glu Tyr Ala 1205
1210 1215 Asp Lys Ile Phe Thr
Tyr Ile Phe Ile Leu Glu Met Leu Leu Lys 1220 1225
1230 Trp Ile Ala Tyr Gly Tyr Lys Thr Tyr Phe
Thr Asn Ala Trp Cys 1235 1240 1245
Trp Leu Asp Phe Leu Ile Val Asp Val Ser Leu Val Thr Leu Val
1250 1255 1260 Ala Asn
Thr Leu Gly Tyr Ser Asp Leu Gly Pro Ile Lys Ser Leu 1265
1270 1275 Arg Thr Leu Arg Ala Leu Arg
Pro Leu Arg Ala Leu Ser Arg Phe 1280 1285
1290 Glu Gly Met Arg Val Val Val Asn Ala Leu Ile Gly
Ala Ile Pro 1295 1300 1305
Ser Ile Met Asn Val Leu Leu Val Cys Leu Ile Phe Trp Leu Ile 1310
1315 1320 Phe Ser Ile Met Gly
Val Asn Leu Phe Ala Gly Lys Phe Tyr Glu 1325 1330
1335 Cys Ile Asn Thr Thr Asp Gly Ser Arg Phe
Pro Ala Ser Gln Val 1340 1345 1350
Pro Asn Arg Ser Glu Cys Phe Ala Leu Met Asn Val Ser Gln Asn
1355 1360 1365 Val Arg
Trp Lys Asn Leu Lys Val Asn Phe Asp Asn Val Gly Leu 1370
1375 1380 Gly Tyr Leu Ser Leu Leu Gln
Val Ala Thr Phe Lys Gly Trp Thr 1385 1390
1395 Ile Ile Met Tyr Ala Ala Val Asp Ser Val Asn Val
Asp Lys Gln 1400 1405 1410
Pro Lys Tyr Glu Tyr Ser Leu Tyr Met Tyr Ile Tyr Phe Val Val 1415
1420 1425 Phe Ile Ile Phe Gly
Ser Phe Phe Thr Leu Asn Leu Phe Ile Gly 1430 1435
1440 Val Ile Ile Asp Asn Phe Asn Gln Gln Lys
Lys Lys Leu Gly Gly 1445 1450 1455
Gln Asp Ile Phe Met Thr Glu Glu Gln Lys Lys Tyr Tyr Asn Ala
1460 1465 1470 Met Lys
Lys Leu Gly Ser Lys Lys Pro Gln Lys Pro Ile Pro Arg 1475
1480 1485 Pro Gly Asn Lys Ile Gln Gly
Cys Ile Phe Asp Leu Val Thr Asn 1490 1495
1500 Gln Ala Phe Asp Ile Ser Ile Met Val Leu Ile Cys
Leu Asn Met 1505 1510 1515
Val Thr Met Met Val Glu Lys Glu Gly Gln Ser Gln His Met Thr 1520
1525 1530 Glu Val Leu Tyr Trp
Ile Asn Val Val Phe Ile Ile Leu Phe Thr 1535 1540
1545 Gly Glu Cys Val Leu Lys Leu Ile Ser Leu
Arg His Tyr Tyr Phe 1550 1555 1560
Thr Val Gly Trp Asn Ile Phe Asp Phe Val Val Val Ile Ile Ser
1565 1570 1575 Ile Val
Gly Met Phe Leu Ala Asp Leu Ile Glu Thr Tyr Phe Val 1580
1585 1590 Ser Pro Thr Leu Phe Arg Val
Ile Arg Leu Ala Arg Ile Gly Arg 1595 1600
1605 Ile Leu Arg Leu Val Lys Gly Ala Lys Gly Ile Arg
Thr Leu Leu 1610 1615 1620
Phe Ala Leu Met Met Ser Leu Pro Ala Leu Phe Asn Ile Gly Leu 1625
1630 1635 Leu Leu Phe Leu Val
Met Phe Ile Tyr Ala Ile Phe Gly Met Ser 1640 1645
1650 Asn Phe Ala Tyr Val Lys Lys Glu Asp Gly
Ile Asn Asp Met Phe 1655 1660 1665
Asn Phe Glu Thr Phe Gly Asn Ser Met Ile Cys Leu Phe Gln Ile
1670 1675 1680 Thr Thr
Ser Ala Gly Trp Asp Gly Leu Leu Ala Pro Ile Leu Asn 1685
1690 1695 Ser Lys Pro Pro Asp Cys Asp
Pro Lys Lys Val His Pro Gly Ser 1700 1705
1710 Ser Val Glu Gly Asp Cys Gly Asn Pro Ser Val Gly
Ile Phe Tyr 1715 1720 1725
Phe Val Ser Tyr Ile Ile Ile Ser Phe Leu Val Val Val Asn Met 1730
1735 1740 Tyr Ile Ala Val Ile
Leu Glu Asn Phe Ser Val Ala Thr Glu Glu 1745 1750
1755 Ser Thr Glu Pro Leu Ser Glu Asp Asp Phe
Glu Met Phe Tyr Glu 1760 1765 1770
Val Trp Glu Lys Phe Asp Pro Asp Ala Thr Gln Phe Ile Glu Phe
1775 1780 1785 Ser Lys
Leu Ser Asp Phe Ala Ala Ala Leu Asp Pro Pro Leu Leu 1790
1795 1800 Ile Ala Lys Pro Asn Lys Val
Gln Leu Ile Ala Met Asp Leu Pro 1805 1810
1815 Met Val Ser Gly Asp Arg Ile His Cys Leu Asp Ile
Leu Phe Ala 1820 1825 1830
Phe Thr Lys Arg Val Leu Gly Glu Ser Gly Glu Met Asp Ser Leu 1835
1840 1845 Arg Ser Gln Met Glu
Glu Arg Phe Met Ser Ala Asn Pro Ser Lys 1850 1855
1860 Val Ser Tyr Glu Pro Ile Thr Thr Thr Leu
Lys Arg Lys Gln Glu 1865 1870 1875
Asp Val Ser Ala Thr Val Ile Gln Arg Ala Tyr Arg Arg Tyr Arg
1880 1885 1890 Leu Arg
Gln Asn Val Lys Asn Ile Ser Ser Ile Tyr Ile Lys Asp 1895
1900 1905 Gly Asp Arg Asp Asp Asp Leu
Leu Asn Lys Lys Asp Met Ala Phe 1910 1915
1920 Asp Asn Val Asn Glu Asn Ser Ser Pro Glu Lys Thr
Asp Ala Thr 1925 1930 1935
Ser Ser Thr Thr Ser Pro Pro Ser Tyr Asp Ser Val Thr Lys Pro 1940
1945 1950 Asp Lys Glu Lys Tyr
Glu Gln Asp Arg Thr Glu Lys Glu Asp Lys 1955 1960
1965 Gly Lys Asp Ser Lys Glu Ser Lys Lys
1970 1975 56397DNAHomo sapiens 5ctcttatgtg
aggagctgaa gaggaattaa aatatacagg atgaaaagat ggcaatgttg 60cctcccccag
gacctcagag ctttgtccat ttcacaaaac agtctcttgc cctcattgaa 120caacgcattg
ctgaaagaaa atcaaaggaa cccaaagaag aaaagaaaga tgatgatgaa 180gaagccccaa
agccaagcag tgacttggaa gctggcaaac aactgccctt catctatggg 240gacattcctc
ccggcatggt gtcagagccc ctggaggact tggaccccta ctatgcagac 300aaaaagactt
tcatagtatt gaacaaaggg aaaacaatct tccgtttcaa tgccacacct 360gctttatata
tgctttctcc tttcagtcct ctaagaagaa tatctattaa gattttagta 420cactccttat
tcagcatgct catcatgtgc actattctga caaactgcat atttatgacc 480atgaataacc
cgccggactg gaccaaaaat gtcgagtaca cttttactgg aatatatact 540tttgaatcac
ttgtaaaaat ccttgcaaga ggcttctgtg taggagaatt cacttttctt 600cgtgacccgt
ggaactggct ggattttgtc gtcattgttt ttgcgtattt aacagaattt 660gtaaacctag
gcaatgtttc agctcttcga actttcagag tattgagagc tttgaaaact 720atttctgtaa
tcccaggcct gaagacaatt gtaggggctt tgatccagtc agtgaagaag 780ctttctgatg
tcatgatcct gactgtgttc tgtctgagtg tgtttgcact aattggacta 840cagctgttca
tgggaaacct gaagcataaa tgttttcgaa attcacttga aaataatgaa 900acattagaaa
gcataatgaa taccctagag agtgaagaag actttagaaa atatttttat 960tacttggaag
gatccaaaga tgctctcctt tgtggtttca gcacagattc aggtcagtgt 1020ccagaggggt
acacctgtgt gaaaattggc agaaaccctg attatggcta cacgagcttt 1080gacactttca
gctgggcctt cttagccttg tttaggctaa tgacccaaga ttactgggaa 1140aacctttacc
aacagacgct gcgtgccgct ggcaaaacct acatgatctt ctttgtcgta 1200gtgattttcc
tgggctcctt ttatctaata aacttgatcc tggctgtggt tgccatggca 1260tatgaagaac
agaaccaggc aaacattgaa gaagctaaac agaaagaatt agagtttcaa 1320cagatgttag
accgacttaa aaaagagcaa gaagaagctg aggcaattgc agcggcagcg 1380gctgaatata
caagtattag gagaagcaga attatgggcc tctcagagag ttcttctgaa 1440acatccaaac
tgagctctaa aagtgctaaa gaaagaagaa acagaagaaa gaaaaagaat 1500caaaagaagc
tctccagtgg agaggaaaag ggagatgctg agaaattgtc gaaatcagaa 1560tcagaggaca
gcatcagaag aaaaagtttc caccttggtg tcgaagggca taggcgagca 1620catgaaaaga
ggttgtctac ccccaatcag tcaccactca gcattcgtgg ctccttgttt 1680tctgcaaggc
gaagcagcag aacaagtctt tttagtttca aaggcagagg aagagatata 1740ggatctgaga
ctgaatttgc cgatgatgag cacagcattt ttggagacaa tgagagcaga 1800aggggctcac
tgtttgtgcc ccacagaccc caggagcgac gcagcagtaa catcagccaa 1860gccagtaggt
ccccaccaat gctgccggtg aacgggaaaa tgcacagtgc tgtggactgc 1920aacggtgtgg
tctccctggt tgatggacgc tcagccctca tgctccccaa tggacagctt 1980ctgccagagg
tgataataga taaggcaact tctgatgaca gcggcacgac caatcaaata 2040cacaagaaaa
ggcgttgtag ttcctatctc ctttcagagg atatgctgaa tgatcccaac 2100ctcagacaga
gagcaatgag tagagcaagc atattaacaa acactgtgga agaacttgaa 2160gagtccagac
aaaaatgtcc accttggtgg tacagatttg cacacaaatt cttgatctgg 2220aattgctctc
catattggat aaaattcaaa aagtgtatct attttattgt aatggatcct 2280tttgtagatc
ttgcaattac catttgcata gttttaaaca cattatttat ggctatggaa 2340caccacccaa
tgactgagga attcaaaaat gtacttgcta taggaaattt ggtctttact 2400ggaatctttg
cagctgaaat ggtattaaaa ctgattgcca tggatccata tgagtatttc 2460caagtaggct
ggaatatttt tgacagcctt attgtgactt taagtttagt ggagctcttt 2520ctagcagatg
tggaaggatt gtcagttctg cgatcattca gactgctccg agtcttcaag 2580ttggcaaaat
cctggccaac attgaacatg ctgattaaga tcattggtaa ctcagtaggg 2640gctctaggta
acctcacctt agtgttggcc atcatcgtct tcatttttgc tgtggtcggc 2700atgcagctct
ttggtaagag ctacaaagaa tgtgtctgca agatcaatga tgactgtacg 2760ctcccacggt
ggcacatgaa cgacttcttc cactccttcc tgattgtgtt ccgcgtgctg 2820tgtggagagt
ggatagagac catgtgggac tgtatggagg tcgctggtca agctatgtgc 2880cttattgttt
acatgatggt catggtcatt ggaaacctgg tggtcctaaa cctatttctg 2940gccttattat
tgagctcatt tagttcagac aatcttacag caattgaaga agaccctgat 3000gcaaacaacc
tccagattgc agtgactaga attaaaaagg gaataaatta tgtgaaacaa 3060accttacgtg
aatttattct aaaagcattt tccaaaaagc caaagatttc cagggagata 3120agacaagcag
aagatctgaa tactaagaag gaaaactata tttctaacca tacacttgct 3180gaaatgagca
aaggtcacaa tttcctcaag gaaaaagata aaatcagtgg ttttggaagc 3240agcgtggaca
aacacttgat ggaagacagt gatggtcaat catttattca caatcccagc 3300ctcacagtga
cagtgccaat tgcacctggg gaatccgatt tggaaaatat gaatgctgag 3360gaacttagca
gtgattcgga tagtgaatac agcaaagtga gattaaaccg gtcaagctcc 3420tcagagtgca
gcacagttga taaccctttg cctggagaag gagaagaagc agaggctgaa 3480cctatgaatt
ccgatgagcc agaggcctgt ttcacagatg gttgtgtacg gaggttctca 3540tgctgccaag
ttaacataga gtcagggaaa ggaaaaatct ggtggaacat caggaaaacc 3600tgctacaaga
ttgttgaaca cagttggttt gaaagcttca ttgtcctcat gatcctgctc 3660agcagtggtg
ccctggcttt tgaagatatt tatattgaaa ggaaaaagac cattaagatt 3720atcctggagt
atgcagacaa gatcttcact tacatcttca ttctggaaat gcttctaaaa 3780tggatagcat
atggttataa aacatatttc accaatgcct ggtgttggct ggatttccta 3840attgttgatg
tttctttggt tactttagtg gcaaacactc ttggctactc agatcttggc 3900cccattaaat
cccttcggac actgagagct ttaagacctc taagagcctt atctagattt 3960gaaggaatga
gggtcgttgt gaatgcactc ataggagcaa ttccttccat catgaatgtg 4020ctacttgtgt
gtcttatatt ctggctgata ttcagcatca tgggagtaaa tttgtttgct 4080ggcaagttct
atgagtgtat taacaccaca gatgggtcac ggtttcctgc aagtcaagtt 4140ccaaatcgtt
ccgaatgttt tgcccttatg aatgttagtc aaaatgtgcg atggaaaaac 4200ctgaaagtga
actttgataa tgtcggactt ggttacctat ctctgcttca agttgcaact 4260tttaagggat
ggacgattat tatgtatgca gcagtggatt ctgttaatgt agacaagcag 4320cccaaatatg
aatatagcct ctacatgtat atttattttg tcgtctttat catctttggg 4380tcattcttca
ctttgaactt gttcattggt gtcatcatag ataatttcaa ccaacagaaa 4440aagaagcttg
gaggtcaaga catctttatg acagaagaac agaagaaata ctataatgca 4500atgaaaaagc
tggggtccaa gaagccacaa aagccaattc ctcgaccagg gaacaaaatc 4560caaggatgta
tatttgacct agtgacaaat caagcctttg atattagtat catggttctt 4620atctgtctca
acatggtaac catgatggta gaaaaggagg gtcaaagtca acatatgact 4680gaagttttat
attggataaa tgtggttttt ataatccttt tcactggaga atgtgtgcta 4740aaactgatct
ccctcagaca ctactacttc actgtaggat ggaatatttt tgattttgtg 4800gttgtgatta
tctccattgt aggtatgttt ctagctgatt tgattgaaac gtattttgtg 4860tcccctaccc
tgttccgagt gatccgtctt gccaggattg gccgaatcct acgtctagtc 4920aaaggagcaa
aggggatccg cacgctgctc tttgctttga tgatgtccct tcctgcgttg 4980tttaacatcg
gcctcctgct cttcctggtc atgttcatct acgccatctt tggaatgtcc 5040aactttgcct
atgttaaaaa ggaagatgga attaatgaca tgttcaattt tgagaccttt 5100ggcaacagta
tgatttgcct gttccaaatt acaacctctg ctggctggga tggattgcta 5160gcacctattc
ttaacagtaa gccacccgac tgtgacccaa aaaaagttca tcctggaagt 5220tcagttgaag
gagactgtgg taacccatct gttggaatat tctactttgt tagttatatc 5280atcatatcct
tcctggttgt ggtgaacatg tacattgcag tcatactgga gaattttagt 5340gttgccactg
aagaaagtac tgaacctctg agtgaggatg actttgagat gttctatgag 5400gtttgggaga
agtttgatcc cgatgcgacc cagtttatag agttctctaa actctctgat 5460tttgcagctg
ccctggatcc tcctcttctc atagcaaaac ccaacaaagt ccagctcatt 5520gccatggatc
tgcccatggt tagtggtgac cggatccatt gtcttgacat cttatttgct 5580tttacaaagc
gtgttttggg tgagagtggg gagatggatt ctcttcgttc acagatggaa 5640gaaaggttca
tgtctgcaaa tccttccaaa gtgtcctatg aacccatcac aaccacacta 5700aaacggaaac
aagaggatgt gtctgctact gtcattcagc gtgcttatag acgttaccgc 5760ttaaggcaaa
atgtcaaaaa tatatcaagt atatacataa aagatggaga cagagatgat 5820gatttactca
ataaaaaaga tatggctttt gataatgtta atgagaactc aagtccagaa 5880aaaacagatg
ccacttcatc caccacctct ccaccttcat atgatagtgt aacaaagcca 5940gacaaagaga
aatatgaaca agacagaaca gaaaaggaag acaaagggaa agacagcaag 6000gaaagcaaaa
aatagagctt catttttgat atattgttta cagcctgtga aagtgattta 6060tttgtgttaa
taaaactctt ttgaggaagt ctatgccaaa atccttttta tcaaaatatt 6120ctcgaaggca
gtgcagtcac taactctgat ttcctaagaa aggtgggcag cattagcaga 6180tggttatttt
tgcactgatg attctttaag aatcgtaaga gaactctgta ggaattattg 6240attatagcat
acaaaagtga ttcagttttt tggtttttaa taaatcagaa gaccatgtag 6300aaaactttta
catctgcctt gtcatctttt cacaggattg taattagtct tgtttcccat 6360gtaaataaac
aacacacgca tacagaaaaa aaaaaaa 639761988PRTHomo
sapiens 6Met Ala Met Leu Pro Pro Pro Gly Pro Gln Ser Phe Val His Phe Thr
1 5 10 15 Lys Gln
Ser Leu Ala Leu Ile Glu Gln Arg Ile Ala Glu Arg Lys Ser 20
25 30 Lys Glu Pro Lys Glu Glu Lys
Lys Asp Asp Asp Glu Glu Ala Pro Lys 35 40
45 Pro Ser Ser Asp Leu Glu Ala Gly Lys Gln Leu Pro
Phe Ile Tyr Gly 50 55 60
Asp Ile Pro Pro Gly Met Val Ser Glu Pro Leu Glu Asp Leu Asp Pro 65
70 75 80 Tyr Tyr Ala
Asp Lys Lys Thr Phe Ile Val Leu Asn Lys Gly Lys Thr 85
90 95 Ile Phe Arg Phe Asn Ala Thr Pro
Ala Leu Tyr Met Leu Ser Pro Phe 100 105
110 Ser Pro Leu Arg Arg Ile Ser Ile Lys Ile Leu Val His
Ser Leu Phe 115 120 125
Ser Met Leu Ile Met Cys Thr Ile Leu Thr Asn Cys Ile Phe Met Thr 130
135 140 Met Asn Asn Pro
Pro Asp Trp Thr Lys Asn Val Glu Tyr Thr Phe Thr 145 150
155 160 Gly Ile Tyr Thr Phe Glu Ser Leu Val
Lys Ile Leu Ala Arg Gly Phe 165 170
175 Cys Val Gly Glu Phe Thr Phe Leu Arg Asp Pro Trp Asn Trp
Leu Asp 180 185 190
Phe Val Val Ile Val Phe Ala Tyr Leu Thr Glu Phe Val Asn Leu Gly
195 200 205 Asn Val Ser Ala
Leu Arg Thr Phe Arg Val Leu Arg Ala Leu Lys Thr 210
215 220 Ile Ser Val Ile Pro Gly Leu Lys
Thr Ile Val Gly Ala Leu Ile Gln 225 230
235 240 Ser Val Lys Lys Leu Ser Asp Val Met Ile Leu Thr
Val Phe Cys Leu 245 250
255 Ser Val Phe Ala Leu Ile Gly Leu Gln Leu Phe Met Gly Asn Leu Lys
260 265 270 His Lys Cys
Phe Arg Asn Ser Leu Glu Asn Asn Glu Thr Leu Glu Ser 275
280 285 Ile Met Asn Thr Leu Glu Ser Glu
Glu Asp Phe Arg Lys Tyr Phe Tyr 290 295
300 Tyr Leu Glu Gly Ser Lys Asp Ala Leu Leu Cys Gly Phe
Ser Thr Asp 305 310 315
320 Ser Gly Gln Cys Pro Glu Gly Tyr Thr Cys Val Lys Ile Gly Arg Asn
325 330 335 Pro Asp Tyr Gly
Tyr Thr Ser Phe Asp Thr Phe Ser Trp Ala Phe Leu 340
345 350 Ala Leu Phe Arg Leu Met Thr Gln Asp
Tyr Trp Glu Asn Leu Tyr Gln 355 360
365 Gln Thr Leu Arg Ala Ala Gly Lys Thr Tyr Met Ile Phe Phe
Val Val 370 375 380
Val Ile Phe Leu Gly Ser Phe Tyr Leu Ile Asn Leu Ile Leu Ala Val 385
390 395 400 Val Ala Met Ala Tyr
Glu Glu Gln Asn Gln Ala Asn Ile Glu Glu Ala 405
410 415 Lys Gln Lys Glu Leu Glu Phe Gln Gln Met
Leu Asp Arg Leu Lys Lys 420 425
430 Glu Gln Glu Glu Ala Glu Ala Ile Ala Ala Ala Ala Ala Glu Tyr
Thr 435 440 445 Ser
Ile Arg Arg Ser Arg Ile Met Gly Leu Ser Glu Ser Ser Ser Glu 450
455 460 Thr Ser Lys Leu Ser Ser
Lys Ser Ala Lys Glu Arg Arg Asn Arg Arg 465 470
475 480 Lys Lys Lys Asn Gln Lys Lys Leu Ser Ser Gly
Glu Glu Lys Gly Asp 485 490
495 Ala Glu Lys Leu Ser Lys Ser Glu Ser Glu Asp Ser Ile Arg Arg Lys
500 505 510 Ser Phe
His Leu Gly Val Glu Gly His Arg Arg Ala His Glu Lys Arg 515
520 525 Leu Ser Thr Pro Asn Gln Ser
Pro Leu Ser Ile Arg Gly Ser Leu Phe 530 535
540 Ser Ala Arg Arg Ser Ser Arg Thr Ser Leu Phe Ser
Phe Lys Gly Arg 545 550 555
560 Gly Arg Asp Ile Gly Ser Glu Thr Glu Phe Ala Asp Asp Glu His Ser
565 570 575 Ile Phe Gly
Asp Asn Glu Ser Arg Arg Gly Ser Leu Phe Val Pro His 580
585 590 Arg Pro Gln Glu Arg Arg Ser Ser
Asn Ile Ser Gln Ala Ser Arg Ser 595 600
605 Pro Pro Met Leu Pro Val Asn Gly Lys Met His Ser Ala
Val Asp Cys 610 615 620
Asn Gly Val Val Ser Leu Val Asp Gly Arg Ser Ala Leu Met Leu Pro 625
630 635 640 Asn Gly Gln Leu
Leu Pro Glu Val Ile Ile Asp Lys Ala Thr Ser Asp 645
650 655 Asp Ser Gly Thr Thr Asn Gln Ile His
Lys Lys Arg Arg Cys Ser Ser 660 665
670 Tyr Leu Leu Ser Glu Asp Met Leu Asn Asp Pro Asn Leu Arg
Gln Arg 675 680 685
Ala Met Ser Arg Ala Ser Ile Leu Thr Asn Thr Val Glu Glu Leu Glu 690
695 700 Glu Ser Arg Gln Lys
Cys Pro Pro Trp Trp Tyr Arg Phe Ala His Lys 705 710
715 720 Phe Leu Ile Trp Asn Cys Ser Pro Tyr Trp
Ile Lys Phe Lys Lys Cys 725 730
735 Ile Tyr Phe Ile Val Met Asp Pro Phe Val Asp Leu Ala Ile Thr
Ile 740 745 750 Cys
Ile Val Leu Asn Thr Leu Phe Met Ala Met Glu His His Pro Met 755
760 765 Thr Glu Glu Phe Lys Asn
Val Leu Ala Ile Gly Asn Leu Val Phe Thr 770 775
780 Gly Ile Phe Ala Ala Glu Met Val Leu Lys Leu
Ile Ala Met Asp Pro 785 790 795
800 Tyr Glu Tyr Phe Gln Val Gly Trp Asn Ile Phe Asp Ser Leu Ile Val
805 810 815 Thr Leu
Ser Leu Val Glu Leu Phe Leu Ala Asp Val Glu Gly Leu Ser 820
825 830 Val Leu Arg Ser Phe Arg Leu
Leu Arg Val Phe Lys Leu Ala Lys Ser 835 840
845 Trp Pro Thr Leu Asn Met Leu Ile Lys Ile Ile Gly
Asn Ser Val Gly 850 855 860
Ala Leu Gly Asn Leu Thr Leu Val Leu Ala Ile Ile Val Phe Ile Phe 865
870 875 880 Ala Val Val
Gly Met Gln Leu Phe Gly Lys Ser Tyr Lys Glu Cys Val 885
890 895 Cys Lys Ile Asn Asp Asp Cys Thr
Leu Pro Arg Trp His Met Asn Asp 900 905
910 Phe Phe His Ser Phe Leu Ile Val Phe Arg Val Leu Cys
Gly Glu Trp 915 920 925
Ile Glu Thr Met Trp Asp Cys Met Glu Val Ala Gly Gln Ala Met Cys 930
935 940 Leu Ile Val Tyr
Met Met Val Met Val Ile Gly Asn Leu Val Val Leu 945 950
955 960 Asn Leu Phe Leu Ala Leu Leu Leu Ser
Ser Phe Ser Ser Asp Asn Leu 965 970
975 Thr Ala Ile Glu Glu Asp Pro Asp Ala Asn Asn Leu Gln Ile
Ala Val 980 985 990
Thr Arg Ile Lys Lys Gly Ile Asn Tyr Val Lys Gln Thr Leu Arg Glu
995 1000 1005 Phe Ile Leu
Lys Ala Phe Ser Lys Lys Pro Lys Ile Ser Arg Glu 1010
1015 1020 Ile Arg Gln Ala Glu Asp Leu Asn
Thr Lys Lys Glu Asn Tyr Ile 1025 1030
1035 Ser Asn His Thr Leu Ala Glu Met Ser Lys Gly His Asn
Phe Leu 1040 1045 1050
Lys Glu Lys Asp Lys Ile Ser Gly Phe Gly Ser Ser Val Asp Lys 1055
1060 1065 His Leu Met Glu Asp
Ser Asp Gly Gln Ser Phe Ile His Asn Pro 1070 1075
1080 Ser Leu Thr Val Thr Val Pro Ile Ala Pro
Gly Glu Ser Asp Leu 1085 1090 1095
Glu Asn Met Asn Ala Glu Glu Leu Ser Ser Asp Ser Asp Ser Glu
1100 1105 1110 Tyr Ser
Lys Val Arg Leu Asn Arg Ser Ser Ser Ser Glu Cys Ser 1115
1120 1125 Thr Val Asp Asn Pro Leu Pro
Gly Glu Gly Glu Glu Ala Glu Ala 1130 1135
1140 Glu Pro Met Asn Ser Asp Glu Pro Glu Ala Cys Phe
Thr Asp Gly 1145 1150 1155
Cys Val Arg Arg Phe Ser Cys Cys Gln Val Asn Ile Glu Ser Gly 1160
1165 1170 Lys Gly Lys Ile Trp
Trp Asn Ile Arg Lys Thr Cys Tyr Lys Ile 1175 1180
1185 Val Glu His Ser Trp Phe Glu Ser Phe Ile
Val Leu Met Ile Leu 1190 1195 1200
Leu Ser Ser Gly Ala Leu Ala Phe Glu Asp Ile Tyr Ile Glu Arg
1205 1210 1215 Lys Lys
Thr Ile Lys Ile Ile Leu Glu Tyr Ala Asp Lys Ile Phe 1220
1225 1230 Thr Tyr Ile Phe Ile Leu Glu
Met Leu Leu Lys Trp Ile Ala Tyr 1235 1240
1245 Gly Tyr Lys Thr Tyr Phe Thr Asn Ala Trp Cys Trp
Leu Asp Phe 1250 1255 1260
Leu Ile Val Asp Val Ser Leu Val Thr Leu Val Ala Asn Thr Leu 1265
1270 1275 Gly Tyr Ser Asp Leu
Gly Pro Ile Lys Ser Leu Arg Thr Leu Arg 1280 1285
1290 Ala Leu Arg Pro Leu Arg Ala Leu Ser Arg
Phe Glu Gly Met Arg 1295 1300 1305
Val Val Val Asn Ala Leu Ile Gly Ala Ile Pro Ser Ile Met Asn
1310 1315 1320 Val Leu
Leu Val Cys Leu Ile Phe Trp Leu Ile Phe Ser Ile Met 1325
1330 1335 Gly Val Asn Leu Phe Ala Gly
Lys Phe Tyr Glu Cys Ile Asn Thr 1340 1345
1350 Thr Asp Gly Ser Arg Phe Pro Ala Ser Gln Val Pro
Asn Arg Ser 1355 1360 1365
Glu Cys Phe Ala Leu Met Asn Val Ser Gln Asn Val Arg Trp Lys 1370
1375 1380 Asn Leu Lys Val Asn
Phe Asp Asn Val Gly Leu Gly Tyr Leu Ser 1385 1390
1395 Leu Leu Gln Val Ala Thr Phe Lys Gly Trp
Thr Ile Ile Met Tyr 1400 1405 1410
Ala Ala Val Asp Ser Val Asn Val Asp Lys Gln Pro Lys Tyr Glu
1415 1420 1425 Tyr Ser
Leu Tyr Met Tyr Ile Tyr Phe Val Val Phe Ile Ile Phe 1430
1435 1440 Gly Ser Phe Phe Thr Leu Asn
Leu Phe Ile Gly Val Ile Ile Asp 1445 1450
1455 Asn Phe Asn Gln Gln Lys Lys Lys Leu Gly Gly Gln
Asp Ile Phe 1460 1465 1470
Met Thr Glu Glu Gln Lys Lys Tyr Tyr Asn Ala Met Lys Lys Leu 1475
1480 1485 Gly Ser Lys Lys Pro
Gln Lys Pro Ile Pro Arg Pro Gly Asn Lys 1490 1495
1500 Ile Gln Gly Cys Ile Phe Asp Leu Val Thr
Asn Gln Ala Phe Asp 1505 1510 1515
Ile Ser Ile Met Val Leu Ile Cys Leu Asn Met Val Thr Met Met
1520 1525 1530 Val Glu
Lys Glu Gly Gln Ser Gln His Met Thr Glu Val Leu Tyr 1535
1540 1545 Trp Ile Asn Val Val Phe Ile
Ile Leu Phe Thr Gly Glu Cys Val 1550 1555
1560 Leu Lys Leu Ile Ser Leu Arg His Tyr Tyr Phe Thr
Val Gly Trp 1565 1570 1575
Asn Ile Phe Asp Phe Val Val Val Ile Ile Ser Ile Val Gly Met 1580
1585 1590 Phe Leu Ala Asp Leu
Ile Glu Thr Tyr Phe Val Ser Pro Thr Leu 1595 1600
1605 Phe Arg Val Ile Arg Leu Ala Arg Ile Gly
Arg Ile Leu Arg Leu 1610 1615 1620
Val Lys Gly Ala Lys Gly Ile Arg Thr Leu Leu Phe Ala Leu Met
1625 1630 1635 Met Ser
Leu Pro Ala Leu Phe Asn Ile Gly Leu Leu Leu Phe Leu 1640
1645 1650 Val Met Phe Ile Tyr Ala Ile
Phe Gly Met Ser Asn Phe Ala Tyr 1655 1660
1665 Val Lys Lys Glu Asp Gly Ile Asn Asp Met Phe Asn
Phe Glu Thr 1670 1675 1680
Phe Gly Asn Ser Met Ile Cys Leu Phe Gln Ile Thr Thr Ser Ala 1685
1690 1695 Gly Trp Asp Gly Leu
Leu Ala Pro Ile Leu Asn Ser Lys Pro Pro 1700 1705
1710 Asp Cys Asp Pro Lys Lys Val His Pro Gly
Ser Ser Val Glu Gly 1715 1720 1725
Asp Cys Gly Asn Pro Ser Val Gly Ile Phe Tyr Phe Val Ser Tyr
1730 1735 1740 Ile Ile
Ile Ser Phe Leu Val Val Val Asn Met Tyr Ile Ala Val 1745
1750 1755 Ile Leu Glu Asn Phe Ser Val
Ala Thr Glu Glu Ser Thr Glu Pro 1760 1765
1770 Leu Ser Glu Asp Asp Phe Glu Met Phe Tyr Glu Val
Trp Glu Lys 1775 1780 1785
Phe Asp Pro Asp Ala Thr Gln Phe Ile Glu Phe Ser Lys Leu Ser 1790
1795 1800 Asp Phe Ala Ala Ala
Leu Asp Pro Pro Leu Leu Ile Ala Lys Pro 1805 1810
1815 Asn Lys Val Gln Leu Ile Ala Met Asp Leu
Pro Met Val Ser Gly 1820 1825 1830
Asp Arg Ile His Cys Leu Asp Ile Leu Phe Ala Phe Thr Lys Arg
1835 1840 1845 Val Leu
Gly Glu Ser Gly Glu Met Asp Ser Leu Arg Ser Gln Met 1850
1855 1860 Glu Glu Arg Phe Met Ser Ala
Asn Pro Ser Lys Val Ser Tyr Glu 1865 1870
1875 Pro Ile Thr Thr Thr Leu Lys Arg Lys Gln Glu Asp
Val Ser Ala 1880 1885 1890
Thr Val Ile Gln Arg Ala Tyr Arg Arg Tyr Arg Leu Arg Gln Asn 1895
1900 1905 Val Lys Asn Ile Ser
Ser Ile Tyr Ile Lys Asp Gly Asp Arg Asp 1910 1915
1920 Asp Asp Leu Leu Asn Lys Lys Asp Met Ala
Phe Asp Asn Val Asn 1925 1930 1935
Glu Asn Ser Ser Pro Glu Lys Thr Asp Ala Thr Ser Ser Thr Thr
1940 1945 1950 Ser Pro
Pro Ser Tyr Asp Ser Val Thr Lys Pro Asp Lys Glu Lys 1955
1960 1965 Tyr Glu Gln Asp Arg Thr Glu
Lys Glu Asp Lys Gly Lys Asp Ser 1970 1975
1980 Lys Glu Ser Lys Lys 1985 7
6367DNAHomo sapiens 7ctcttatgtg aggagctgaa gaggaattaa aatatacagg
atgaaaagat ggcaatgttg 60cctcccccag gacctcagag ctttgtccat ttcacaaaac
agtctcttgc cctcattgaa 120caacgcattg ctgaaagaaa atcaaaggaa cccaaagaag
aaaagaaaga tgatgatgaa 180gaagccccaa agccaagcag tgacttggaa gctggcaaac
agctgccctt catctatggg 240gacattcctc ccggcatggt gtcagagccc ctggaggact
tggaccccta ctatgcagac 300aaaaagactt tcatagtatt gaacaaaggg aaaacaatct
tccgtttcaa tgccacacct 360gctttatata tgctttctcc tttcagtcct ctaagaagaa
tatctattaa gattttagta 420cactccttat tcagcatgct catcatgtgc actattctga
caaactgcat atttatgacc 480atgaataacc caccggactg gaccaaaaat gtcgagtaca
cttttactgg aatatatact 540tttgaatcac ttgtaaaaat ccttgcaaga ggcttctgtg
taggagaatt cacttttctt 600cgtgacccgt ggaactggct ggattttgtc gtcattgttt
ttgcgtattt aacagaattt 660gtaaacctag gcaatgtttc agctcttcga actttcagag
tattgagagc tttgaaaact 720atttctgtaa tcccaggcct gaagacaatt gtaggggctt
tgatccagtc agtgaagaag 780ctttctgatg tcatgatcct gactgtgttc tgtctgagtg
tgtttgcact aattggacta 840cagctgttca tgggaaacct gaagcataaa tgttttcgaa
attcacttga aaataatgaa 900acattagaaa gcataatgaa taccctagag agtgaagaag
actttagaaa atatttttat 960tacttggaag gatccaaaga tgctctcctt tgtggtttca
gcacagattc aggtcagtgt 1020ccagaggggt acacctgtgt gaaaattggc agaaaccctg
attatggcta cacgagcttt 1080gacactttca gctgggcctt cttagccttg tttaggctaa
tgacccaaga ttactgggaa 1140aacctttacc aacagacgct gcgtgctgct ggcaaaacct
acatgatctt ctttgtcgta 1200gtgattttcc tgggctcctt ttatctaata aacttgatcc
tggctgtggt tgccatggca 1260tatgaagaac agaaccaggc aaacattgaa gaagctaaac
agaaagaatt agaatttcaa 1320cagatgttag accgtcttaa aaaagagcaa gaagaagctg
aggcaattgc agcggcagcg 1380gctgaatata caagtattag gagaagcaga attatgggcc
tctcagagag ttcttctgaa 1440acatccaaac tgagctctaa aagtgctaaa gaaagaagaa
acagaagaaa gaaaaagaat 1500caaaagaagc tctccagtgg agaggaaaag ggagatgctg
agaaattgtc gaaatcagaa 1560tcagaggaca gcatcagaag aaaaagtttc caccttggtg
tcgaagggca taggcgagca 1620catgaaaaga ggttgtctac ccccaatcag tcaccactca
gcattcgtgg ctccttgttt 1680tctgcaaggc gaagcagcag aacaagtctt tttagtttca
aaggcagagg aagagatata 1740ggatctgaga ctgaatttgc cgatgatgag cacagcattt
ttggagacaa tgagagcaga 1800aggggctcac tgtttgtgcc ccacagaccc caggagcgac
gcagcagtaa catcagccaa 1860gccagtaggt ccccaccaat gctgccggtg aacgggaaaa
tgcacagtgc tgtggactgc 1920aacggtgtgg tctccctggt tgatggacgc tcagccctca
tgctccccaa tggacagctt 1980ctgccagagg gcacgaccaa tcaaatacac aagaaaaggc
gttgtagttc ctatctcctt 2040tcagaggata tgctgaatga tcccaacctc agacagagag
caatgagtag agcaagcata 2100ttaacaaaca ctgtggaaga acttgaagag tccagacaaa
aatgtccacc ttggtggtac 2160agatttgcac acaaattctt gatctggaat tgctctccat
attggataaa attcaaaaag 2220tgtatctatt ttattgtaat ggatcctttt gtagatcttg
caattaccat ttgcatagtt 2280ttaaacacat tatttatggc tatggaacac cacccaatga
ctgaggaatt caaaaatgta 2340cttgctatag gaaatttggt ctttactgga atctttgcag
ctgaaatggt attaaaactg 2400attgccatgg atccatatga gtatttccaa gtaggctgga
atatttttga cagccttatt 2460gtgactttaa gtttagtgga gctctttcta gcagatgtgg
aaggattgtc agttctgcga 2520tcattcagac tgctccgagt cttcaagttg gcaaaatcct
ggccaacatt gaacatgctg 2580attaagatca ttggtaactc agtaggggct ctaggtaacc
tcaccttagt gttggccatc 2640atcgtcttca tttttgctgt ggtcggcatg cagctctttg
gtaagagcta caaagaatgt 2700gtctgcaaga tcaatgatga ctgtacgctc ccacggtggc
acatgaacga cttcttccac 2760tccttcctga ttgtgttccg cgtgctgtgt ggagagtgga
tagagaccat gtgggactgt 2820atggaggtcg ctggtcaagc tatgtgcctt attgtttaca
tgatggtcat ggtcattgga 2880aacctggtgg tcctaaacct atttctggcc ttattattga
gctcatttag ttcagacaat 2940cttacagcaa ttgaagaaga ccctgatgca aacaacctcc
agattgcagt gactagaatt 3000aaaaagggaa taaattatgt gaaacaaacc ttacgtgaat
ttattctaaa agcattttcc 3060aaaaagccaa agatttccag ggagataaga caagcagaag
atctgaatac taagaaggaa 3120aactatattt ctaaccatac acttgctgaa atgagcaaag
gtcacaattt cctcaaggaa 3180aaagataaaa tcagtggttt tggaagcagc gtggacaaac
acttgatgga agacagtgat 3240ggtcaatcat ttattcacaa tcccagcctc acagtgacag
tgccaattgc acctggggaa 3300tccgatttgg aaaatatgaa tgctgaggaa cttagcagtg
attcggatag tgaatacagc 3360aaagtgagat taaaccggtc aagctcctca gagtgcagca
cagttgataa ccctttgcct 3420ggagaaggag aagaagcaga ggctgaacct atgaattccg
atgagccaga ggcctgtttc 3480acagatggtt gtgtatggag gttctcatgc tgccaagtta
acatagagtc agggaaagga 3540aaaatctggt ggaacatcag gaaaacctgc tacaagattg
ttgaacacag ttggtttgaa 3600agcttcattg tcctcatgat cctgctcagc agtggtgccc
tggcttttga agatatttat 3660attgaaagga aaaagaccat taagattatc ctggagtatg
cagacaagat cttcacttac 3720atcttcattc tggaaatgct tctaaaatgg atagcatatg
gttataaaac atatttcacc 3780aatgcctggt gttggctgga tttcctaatt gttgatgttt
ctttggttac tttagtggca 3840aacactcttg gctactcaga tcttggcccc attaaatccc
ttcggacact gagagcttta 3900agacctctaa gagccttatc tagatttgaa ggaatgaggg
tcgttgtgaa tgcactcata 3960ggagcaattc cttccatcat gaatgtgcta cttgtgtgtc
ttatattctg gctgatattc 4020agcatcatgg gagtaaattt gtttgctggc aagttctatg
agtgtattaa caccacagat 4080gggtcacggt ttcctgcaag tcaagttcca aatcgttccg
aatgttttgc ccttatgaat 4140gttagtcaaa atgtgcgatg gaaaaacctg aaagtgaact
ttgataatgt cggacttggt 4200tacctatctc tgcttcaagt tgcaactttt aagggatgga
cgattattat gtatgcagca 4260gtggattctg ttaatgtaga caagcagccc aaatatgaat
atagcctcta catgtatatt 4320tattttgtcg tctttatcat ctttgggtca ttcttcactt
tgaacttgtt cattggtgtc 4380atcatagata atttcaacca acagaaaaag aagcttggag
gtcaagacat ctttatgaca 4440gaagaacaga agaaatacta taatgcaatg aaaaagctgg
ggtccaagaa gccacaaaag 4500ccaattcctc gaccagggaa caaaatccaa ggatgtatat
ttgacctagt gacaaatcaa 4560gcctttgata ttagtatcat ggttcttatc tgtctcaaca
tggtaaccat gatggtagaa 4620aaggagggtc aaagtcaaca tatgactgaa gttttatatt
ggataaatgt ggtttttata 4680atccttttca ctggagaatg tgtgctaaaa ctgatctccc
tcagacacta ctacttcact 4740gtaggatgga atatttttga ttttgtggtt gtgattatct
ccattgtagg tatgtttcta 4800gctgatttga ttgaaacgta ttttgtgtcc cctaccctgt
tccgagtgat ccgtcttgcc 4860aggattggcc gaatcctacg tctagtcaaa ggagcaaagg
ggatccgcac gctgctcttt 4920gctttgatga tgtcccttcc tgcgttgttt aacatcggcc
tcctgctctt cctggtcatg 4980ttcatctacg ccatctttgg aatgtccaac tttgcctatg
ttaaaaagga agatggaatt 5040aatgacatgt tcaattttga gacctttggc aacagtatga
tttgcctgtt ccaaattaca 5100acctctgctg gctgagatgg attgctagca cctattctta
acagtaagcc acccgactgt 5160gacccaaaaa aagttcatcc tggaagttca gttgaaggag
actgtggtaa cccatctgtt 5220ggaatattct actttgttag ttatatcatc atatccttcc
tggttgtggt gaacatgtac 5280attgcagtca tactggagaa ttttagtgtt gccactgaag
aaagtactga acctctgagt 5340gaggatgact ttgagatgtt ctatgaggtt tgggagaagt
ttgatcccga tgcgacccag 5400tttatagagt tctctaaact ctctgatttt gcagctgccc
tggatcctcc tcttctcata 5460gcaaaaccca acaaagtcca gctcattgcc atggatctgc
ccatggttag tggtgaccgg 5520atccattgtc ttgacatctt atttgctttt acaaagcgtg
ttttgggtga gagtggggag 5580atggattctc ttcgttcaca gatggaagaa aggttcatgt
ctgcaaatcc ttccaaagtg 5640tcctatgaac ccatcacaac cacactaaaa cggaaacaag
aggatgtgtc tgctactgtc 5700attcagcgtg cttatagacg ttaccgctta aggcaaaatg
tcaaaaatat atcaagtata 5760tacataaaag atggagacag agatgatgat ttactcaata
aaaaagatat ggcttttgat 5820aatgttaatg agaactcaag tccagaaaaa acagatgcca
cttcatccac cacctctcca 5880ccttcatatg atagtgtaac aaagccagac aaagagaaat
atgaacaaga cagaacagaa 5940aaggaagaca aagggaaaga cagcaaggaa agcaaaaaat
agagcttcat ttttgatata 6000ttgtttacag cctgtgaaag tgatttattt gtgttaataa
aactcttttg aggaagtcta 6060tgccaaaatc ctttttatca aaatattctc gaaggcagtg
cagtcactaa ctctgatttc 6120ctaagaaagg tgggcagcat tagcagatgg ttatttttgc
actgatgatt ctttaagaat 6180cgtaagagaa ctctgtagga attattgatt atagcataca
aaagtgattc agttttttgg 6240tttttaataa atcagaagac catgtagaaa acttttacat
ctgccttgtc atcttttcac 6300aggattgtaa ttagtcttgt ttcccatgta aataaacaac
acacgcatac agaaaaaaaa 6360aaaaaaa
636781688PRTHomo sapiens 8Met Ala Met Leu Pro Pro
Pro Gly Pro Gln Ser Phe Val His Phe Thr 1 5
10 15 Lys Gln Ser Leu Ala Leu Ile Glu Gln Arg Ile
Ala Glu Arg Lys Ser 20 25
30 Lys Glu Pro Lys Glu Glu Lys Lys Asp Asp Asp Glu Glu Ala Pro
Lys 35 40 45 Pro
Ser Ser Asp Leu Glu Ala Gly Lys Gln Leu Pro Phe Ile Tyr Gly 50
55 60 Asp Ile Pro Pro Gly Met
Val Ser Glu Pro Leu Glu Asp Leu Asp Pro 65 70
75 80 Tyr Tyr Ala Asp Lys Lys Thr Phe Ile Val Leu
Asn Lys Gly Lys Thr 85 90
95 Ile Phe Arg Phe Asn Ala Thr Pro Ala Leu Tyr Met Leu Ser Pro Phe
100 105 110 Ser Pro
Leu Arg Arg Ile Ser Ile Lys Ile Leu Val His Ser Leu Phe 115
120 125 Ser Met Leu Ile Met Cys Thr
Ile Leu Thr Asn Cys Ile Phe Met Thr 130 135
140 Met Asn Asn Pro Pro Asp Trp Thr Lys Asn Val Glu
Tyr Thr Phe Thr 145 150 155
160 Gly Ile Tyr Thr Phe Glu Ser Leu Val Lys Ile Leu Ala Arg Gly Phe
165 170 175 Cys Val Gly
Glu Phe Thr Phe Leu Arg Asp Pro Trp Asn Trp Leu Asp 180
185 190 Phe Val Val Ile Val Phe Ala Tyr
Leu Thr Glu Phe Val Asn Leu Gly 195 200
205 Asn Val Ser Ala Leu Arg Thr Phe Arg Val Leu Arg Ala
Leu Lys Thr 210 215 220
Ile Ser Val Ile Pro Gly Leu Lys Thr Ile Val Gly Ala Leu Ile Gln 225
230 235 240 Ser Val Lys Lys
Leu Ser Asp Val Met Ile Leu Thr Val Phe Cys Leu 245
250 255 Ser Val Phe Ala Leu Ile Gly Leu Gln
Leu Phe Met Gly Asn Leu Lys 260 265
270 His Lys Cys Phe Arg Asn Ser Leu Glu Asn Asn Glu Thr Leu
Glu Ser 275 280 285
Ile Met Asn Thr Leu Glu Ser Glu Glu Asp Phe Arg Lys Tyr Phe Tyr 290
295 300 Tyr Leu Glu Gly Ser
Lys Asp Ala Leu Leu Cys Gly Phe Ser Thr Asp 305 310
315 320 Ser Gly Gln Cys Pro Glu Gly Tyr Thr Cys
Val Lys Ile Gly Arg Asn 325 330
335 Pro Asp Tyr Gly Tyr Thr Ser Phe Asp Thr Phe Ser Trp Ala Phe
Leu 340 345 350 Ala
Leu Phe Arg Leu Met Thr Gln Asp Tyr Trp Glu Asn Leu Tyr Gln 355
360 365 Gln Thr Leu Arg Ala Ala
Gly Lys Thr Tyr Met Ile Phe Phe Val Val 370 375
380 Val Ile Phe Leu Gly Ser Phe Tyr Leu Ile Asn
Leu Ile Leu Ala Val 385 390 395
400 Val Ala Met Ala Tyr Glu Glu Gln Asn Gln Ala Asn Ile Glu Glu Ala
405 410 415 Lys Gln
Lys Glu Leu Glu Phe Gln Gln Met Leu Asp Arg Leu Lys Lys 420
425 430 Glu Gln Glu Glu Ala Glu Ala
Ile Ala Ala Ala Ala Ala Glu Tyr Thr 435 440
445 Ser Ile Arg Arg Ser Arg Ile Met Gly Leu Ser Glu
Ser Ser Ser Glu 450 455 460
Thr Ser Lys Leu Ser Ser Lys Ser Ala Lys Glu Arg Arg Asn Arg Arg 465
470 475 480 Lys Lys Lys
Asn Gln Lys Lys Leu Ser Ser Gly Glu Glu Lys Gly Asp 485
490 495 Ala Glu Lys Leu Ser Lys Ser Glu
Ser Glu Asp Ser Ile Arg Arg Lys 500 505
510 Ser Phe His Leu Gly Val Glu Gly His Arg Arg Ala His
Glu Lys Arg 515 520 525
Leu Ser Thr Pro Asn Gln Ser Pro Leu Ser Ile Arg Gly Ser Leu Phe 530
535 540 Ser Ala Arg Arg
Ser Ser Arg Thr Ser Leu Phe Ser Phe Lys Gly Arg 545 550
555 560 Gly Arg Asp Ile Gly Ser Glu Thr Glu
Phe Ala Asp Asp Glu His Ser 565 570
575 Ile Phe Gly Asp Asn Glu Ser Arg Arg Gly Ser Leu Phe Val
Pro His 580 585 590
Arg Pro Gln Glu Arg Arg Ser Ser Asn Ile Ser Gln Ala Ser Arg Ser
595 600 605 Pro Pro Met Leu
Pro Val Asn Gly Lys Met His Ser Ala Val Asp Cys 610
615 620 Asn Gly Val Val Ser Leu Val Asp
Gly Arg Ser Ala Leu Met Leu Pro 625 630
635 640 Asn Gly Gln Leu Leu Pro Glu Gly Thr Thr Asn Gln
Ile His Lys Lys 645 650
655 Arg Arg Cys Ser Ser Tyr Leu Leu Ser Glu Asp Met Leu Asn Asp Pro
660 665 670 Asn Leu Arg
Gln Arg Ala Met Ser Arg Ala Ser Ile Leu Thr Asn Thr 675
680 685 Val Glu Glu Leu Glu Glu Ser Arg
Gln Lys Cys Pro Pro Trp Trp Tyr 690 695
700 Arg Phe Ala His Lys Phe Leu Ile Trp Asn Cys Ser Pro
Tyr Trp Ile 705 710 715
720 Lys Phe Lys Lys Cys Ile Tyr Phe Ile Val Met Asp Pro Phe Val Asp
725 730 735 Leu Ala Ile Thr
Ile Cys Ile Val Leu Asn Thr Leu Phe Met Ala Met 740
745 750 Glu His His Pro Met Thr Glu Glu Phe
Lys Asn Val Leu Ala Ile Gly 755 760
765 Asn Leu Val Phe Thr Gly Ile Phe Ala Ala Glu Met Val Leu
Lys Leu 770 775 780
Ile Ala Met Asp Pro Tyr Glu Tyr Phe Gln Val Gly Trp Asn Ile Phe 785
790 795 800 Asp Ser Leu Ile Val
Thr Leu Ser Leu Val Glu Leu Phe Leu Ala Asp 805
810 815 Val Glu Gly Leu Ser Val Leu Arg Ser Phe
Arg Leu Leu Arg Val Phe 820 825
830 Lys Leu Ala Lys Ser Trp Pro Thr Leu Asn Met Leu Ile Lys Ile
Ile 835 840 845 Gly
Asn Ser Val Gly Ala Leu Gly Asn Leu Thr Leu Val Leu Ala Ile 850
855 860 Ile Val Phe Ile Phe Ala
Val Val Gly Met Gln Leu Phe Gly Lys Ser 865 870
875 880 Tyr Lys Glu Cys Val Cys Lys Ile Asn Asp Asp
Cys Thr Leu Pro Arg 885 890
895 Trp His Met Asn Asp Phe Phe His Ser Phe Leu Ile Val Phe Arg Val
900 905 910 Leu Cys
Gly Glu Trp Ile Glu Thr Met Trp Asp Cys Met Glu Val Ala 915
920 925 Gly Gln Ala Met Cys Leu Ile
Val Tyr Met Met Val Met Val Ile Gly 930 935
940 Asn Leu Val Val Leu Asn Leu Phe Leu Ala Leu Leu
Leu Ser Ser Phe 945 950 955
960 Ser Ser Asp Asn Leu Thr Ala Ile Glu Glu Asp Pro Asp Ala Asn Asn
965 970 975 Leu Gln Ile
Ala Val Thr Arg Ile Lys Lys Gly Ile Asn Tyr Val Lys 980
985 990 Gln Thr Leu Arg Glu Phe Ile Leu
Lys Ala Phe Ser Lys Lys Pro Lys 995 1000
1005 Ile Ser Arg Glu Ile Arg Gln Ala Glu Asp Leu
Asn Thr Lys Lys 1010 1015 1020
Glu Asn Tyr Ile Ser Asn His Thr Leu Ala Glu Met Ser Lys Gly
1025 1030 1035 His Asn Phe
Leu Lys Glu Lys Asp Lys Ile Ser Gly Phe Gly Ser 1040
1045 1050 Ser Val Asp Lys His Leu Met Glu
Asp Ser Asp Gly Gln Ser Phe 1055 1060
1065 Ile His Asn Pro Ser Leu Thr Val Thr Val Pro Ile Ala
Pro Gly 1070 1075 1080
Glu Ser Asp Leu Glu Asn Met Asn Ala Glu Glu Leu Ser Ser Asp 1085
1090 1095 Ser Asp Ser Glu Tyr
Ser Lys Val Arg Leu Asn Arg Ser Ser Ser 1100 1105
1110 Ser Glu Cys Ser Thr Val Asp Asn Pro Leu
Pro Gly Glu Gly Glu 1115 1120 1125
Glu Ala Glu Ala Glu Pro Met Asn Ser Asp Glu Pro Glu Ala Cys
1130 1135 1140 Phe Thr
Asp Gly Cys Val Trp Arg Phe Ser Cys Cys Gln Val Asn 1145
1150 1155 Ile Glu Ser Gly Lys Gly Lys
Ile Trp Trp Asn Ile Arg Lys Thr 1160 1165
1170 Cys Tyr Lys Ile Val Glu His Ser Trp Phe Glu Ser
Phe Ile Val 1175 1180 1185
Leu Met Ile Leu Leu Ser Ser Gly Ala Leu Ala Phe Glu Asp Ile 1190
1195 1200 Tyr Ile Glu Arg Lys
Lys Thr Ile Lys Ile Ile Leu Glu Tyr Ala 1205 1210
1215 Asp Lys Ile Phe Thr Tyr Ile Phe Ile Leu
Glu Met Leu Leu Lys 1220 1225 1230
Trp Ile Ala Tyr Gly Tyr Lys Thr Tyr Phe Thr Asn Ala Trp Cys
1235 1240 1245 Trp Leu
Asp Phe Leu Ile Val Asp Val Ser Leu Val Thr Leu Val 1250
1255 1260 Ala Asn Thr Leu Gly Tyr Ser
Asp Leu Gly Pro Ile Lys Ser Leu 1265 1270
1275 Arg Thr Leu Arg Ala Leu Arg Pro Leu Arg Ala Leu
Ser Arg Phe 1280 1285 1290
Glu Gly Met Arg Val Val Val Asn Ala Leu Ile Gly Ala Ile Pro 1295
1300 1305 Ser Ile Met Asn Val
Leu Leu Val Cys Leu Ile Phe Trp Leu Ile 1310 1315
1320 Phe Ser Ile Met Gly Val Asn Leu Phe Ala
Gly Lys Phe Tyr Glu 1325 1330 1335
Cys Ile Asn Thr Thr Asp Gly Ser Arg Phe Pro Ala Ser Gln Val
1340 1345 1350 Pro Asn
Arg Ser Glu Cys Phe Ala Leu Met Asn Val Ser Gln Asn 1355
1360 1365 Val Arg Trp Lys Asn Leu Lys
Val Asn Phe Asp Asn Val Gly Leu 1370 1375
1380 Gly Tyr Leu Ser Leu Leu Gln Val Ala Thr Phe Lys
Gly Trp Thr 1385 1390 1395
Ile Ile Met Tyr Ala Ala Val Asp Ser Val Asn Val Asp Lys Gln 1400
1405 1410 Pro Lys Tyr Glu Tyr
Ser Leu Tyr Met Tyr Ile Tyr Phe Val Val 1415 1420
1425 Phe Ile Ile Phe Gly Ser Phe Phe Thr Leu
Asn Leu Phe Ile Gly 1430 1435 1440
Val Ile Ile Asp Asn Phe Asn Gln Gln Lys Lys Lys Leu Gly Gly
1445 1450 1455 Gln Asp
Ile Phe Met Thr Glu Glu Gln Lys Lys Tyr Tyr Asn Ala 1460
1465 1470 Met Lys Lys Leu Gly Ser Lys
Lys Pro Gln Lys Pro Ile Pro Arg 1475 1480
1485 Pro Gly Asn Lys Ile Gln Gly Cys Ile Phe Asp Leu
Val Thr Asn 1490 1495 1500
Gln Ala Phe Asp Ile Ser Ile Met Val Leu Ile Cys Leu Asn Met 1505
1510 1515 Val Thr Met Met Val
Glu Lys Glu Gly Gln Ser Gln His Met Thr 1520 1525
1530 Glu Val Leu Tyr Trp Ile Asn Val Val Phe
Ile Ile Leu Phe Thr 1535 1540 1545
Gly Glu Cys Val Leu Lys Leu Ile Ser Leu Arg His Tyr Tyr Phe
1550 1555 1560 Thr Val
Gly Trp Asn Ile Phe Asp Phe Val Val Val Ile Ile Ser 1565
1570 1575 Ile Val Gly Met Phe Leu Ala
Asp Leu Ile Glu Thr Tyr Phe Val 1580 1585
1590 Ser Pro Thr Leu Phe Arg Val Ile Arg Leu Ala Arg
Ile Gly Arg 1595 1600 1605
Ile Leu Arg Leu Val Lys Gly Ala Lys Gly Ile Arg Thr Leu Leu 1610
1615 1620 Phe Ala Leu Met Met
Ser Leu Pro Ala Leu Phe Asn Ile Gly Leu 1625 1630
1635 Leu Leu Phe Leu Val Met Phe Ile Tyr Ala
Ile Phe Gly Met Ser 1640 1645 1650
Asn Phe Ala Tyr Val Lys Lys Glu Asp Gly Ile Asn Asp Met Phe
1655 1660 1665 Asn Phe
Glu Thr Phe Gly Asn Ser Met Ile Cys Leu Phe Gln Ile 1670
1675 1680 Thr Thr Ser Ala Gly 1685
96367DNAHomo sapiens 9ctcttatgtg aggagctgaa gaggaattaa
aatatacagg atgaaaagat ggcaatgttg 60cctcccccag gacctcagag ctttgtccat
ttcacaaaac agtctcttgc cctcattgaa 120caacgcattg ctgaaagaaa atcaaaggaa
cccaaagaag aaaagaaaga tgatgatgaa 180gaagccccaa agccaagcag tgacttggaa
gctggcaaac agctgccctt catctatggg 240gacattcctc ccggcatggt gtcagagccc
ctggaggact tggaccccta ctatgcagac 300aaaaagactt tcatagtatt gaacaaaggg
aaaacaatct tccgtttcaa tgccacacct 360gctttatata tgctttctcc tttcagtcct
ctaagaagaa tatctattaa gattttagta 420cactccttat tcagcatgct catcatgtgc
actattctga caaactgcat atttatgacc 480atgaataacc caccggactg gaccaaaaat
gtcgagtaca cttttactgg aatatatact 540tttgaatcac ttgtaaaaat ccttgcaaga
ggcttctgtg taggagaatt cacttttctt 600cgtgacccgt ggaactggct ggattttgtc
gtcattgttt ttgcgtattt aacagaattt 660gtaaacctag gcaatgtttc agctcttcga
actttcagag tattgagagc tttgaaaact 720atttctgtaa tcccaggcct gaagacaatt
gtaggggctt tgatccagtc agtgaagaag 780ctttctgatg tcatgatcct gactgtgttc
tgtctgagtg tgtttgcact aattggacta 840cagctgttca tgggaaacct gaagcataaa
tgttttcgaa attcacttga aaataatgaa 900acattagaaa gcataatgaa taccctagag
agtgaagaag actttagaaa atatttttat 960tacttggaag gatccaaaga tgctctcctt
tgtggtttca gcacagattc aggtcagtgt 1020ccagaggggt aaacctgtgt gaaaattggc
agaaaccctg attatggcta cacgagcttt 1080gacactttca gctgggcctt cttagccttg
tttaggctaa tgacccaaga ttactgggaa 1140aacctttacc aacagacgct gcgtgctgct
ggcaaaacct acatgatctt ctttgtcgta 1200gtgattttcc tgggctcctt ttatctaata
aacttgatcc tggctgtggt tgccatggca 1260tatgaagaac agaaccaggc aaacattgaa
gaagctaaac agaaagaatt agaatttcaa 1320cagatgttag accgtcttaa aaaagagcaa
gaagaagctg aggcaattgc agcggcagcg 1380gctgaatata caagtattag gagaagcaga
attatgggcc tctcagagag ttcttctgaa 1440acatccaaac tgagctctaa aagtgctaaa
gaaagaagaa acagaagaaa gaaaaagaat 1500caaaagaagc tctccagtgg agaggaaaag
ggagatgctg agaaattgtc gaaatcagaa 1560tcagaggaca gcatcagaag aaaaagtttc
caccttggtg tcgaagggca taggcgagca 1620catgaaaaga ggttgtctac ccccaatcag
tcaccactca gcattcgtgg ctccttgttt 1680tctgcaaggc gaagcagcag aacaagtctt
tttagtttca aaggcagagg aagagatata 1740ggatctgaga ctgaatttgc cgatgatgag
cacagcattt ttggagacaa tgagagcaga 1800aggggctcac tgtttgtgcc ccacagaccc
caggagcgac gcagcagtaa catcagccaa 1860gccagtaggt ccccaccaat gctgccggtg
aacgggaaaa tgcacagtgc tgtggactgc 1920aacggtgtgg tctccctggt tgatggacgc
tcagccctca tgctccccaa tggacagctt 1980ctgccagagg gcacgaccaa tcaaatacac
aagaaaaggc gttgtagttc ctatctcctt 2040tcagaggata tgctgaatga tcccaacctc
agacagagag caatgagtag agcaagcata 2100ttaacaaaca ctgtggaaga acttgaagag
tccagacaaa aatgtccacc ttggtggtac 2160agatttgcac acaaattctt gatctggaat
tgctctccat attggataaa attcaaaaag 2220tgtatctatt ttattgtaat ggatcctttt
gtagatcttg caattaccat ttgcatagtt 2280ttaaacacat tatttatggc tatggaacac
cacccaatga ctgaggaatt caaaaatgta 2340cttgctatag gaaatttggt ctttactgga
atctttgcag ctgaaatggt attaaaactg 2400attgccatgg atccatatga gtatttccaa
gtaggctgga atatttttga cagccttatt 2460gtgactttaa gtttagtgga gctctttcta
gcagatgtgg aaggattgtc agttctgcga 2520tcattcagac tgctccgagt cttcaagttg
gcaaaatcct ggccaacatt gaacatgctg 2580attaagatca ttggtaactc agtaggggct
ctaggtaacc tcaccttagt gttggccatc 2640atcgtcttca tttttgctgt ggtcggcatg
cagctctttg gtaagagcta caaagaatgt 2700gtctgcaaga tcaatgatga ctgtacgctc
ccacggtggc acatgaacga cttcttccac 2760tccttcctga ttgtgttccg cgtgctgtgt
ggagagtgga tagagaccat gtgggactgt 2820atggaggtcg ctggtcaagc tatgtgcctt
attgtttaca tgatggtcat ggtcattgga 2880aacctggtgg tcctaaacct atttctggcc
ttattattga gctcatttag ttcagacaat 2940cttacagcaa ttgaagaaga ccctgatgca
aacaacctcc agattgcagt gactagaatt 3000aaaaagggaa taaattatgt gaaacaaacc
ttacgtgaat ttattctaaa agcattttcc 3060aaaaagccaa agatttccag ggagataaga
caagcagaag atctgaatac taagaaggaa 3120aactatattt ctaaccatac acttgctgaa
atgagcaaag gtcacaattt cctcaaggaa 3180aaagataaaa tcagtggttt tggaagcagc
gtggacaaac acttgatgga agacagtgat 3240ggtcaatcat ttattcacaa tcccagcctc
acagtgacag tgccaattgc acctggggaa 3300tccgatttgg aaaatatgaa tgctgaggaa
cttagcagtg attcggatag tgaatacagc 3360aaagtgagat taaaccggtc aagctcctca
gagtgcagca cagttgataa ccctttgcct 3420ggagaaggag aagaagcaga ggctgaacct
atgaattccg atgagccaga ggcctgtttc 3480acagatggtt gtgtatggag gttctcatgc
tgccaagtta acatagagtc agggaaagga 3540aaaatctggt ggaacatcag gaaaacctgc
tacaagattg ttgaacacag ttggtttgaa 3600agcttcattg tcctcatgat cctgctcagc
agtggtgccc tggcttttga agatatttat 3660attgaaagga aaaagaccat taagattatc
ctggagtatg cagacaagat cttcacttac 3720atcttcattc tggaaatgct tctaaaatgg
atagcatatg gttataaaac atatttcacc 3780aatgcctggt gttggctgga tttcctaatt
gttgatgttt ctttggttac tttagtggca 3840aacactcttg gctactcaga tcttggcccc
attaaatccc ttcggacact gagagcttta 3900agacctctaa gagccttatc tagatttgaa
ggaatgaggg tcgttgtgaa tgcactcata 3960ggagcaattc cttccatcat gaatgtgcta
cttgtgtgtc ttatattctg gctgatattc 4020agcatcatgg gagtaaattt gtttgctggc
aagttctatg agtgtattaa caccacagat 4080gggtcacggt ttcctgcaag tcaagttcca
aatcgttccg aatgttttgc ccttatgaat 4140gttagtcaaa atgtgcgatg gaaaaacctg
aaagtgaact ttgataatgt cggacttggt 4200tacctatctc tgcttcaagt tgcaactttt
aagggatgga cgattattat gtatgcagca 4260gtggattctg ttaatgtaga caagcagccc
aaatatgaat atagcctcta catgtatatt 4320tattttgtcg tctttatcat ctttgggtca
ttcttcactt tgaacttgtt cattggtgtc 4380atcatagata atttcaacca acagaaaaag
aagcttggag gtcaagacat ctttatgaca 4440gaagaacaga agaaatacta taatgcaatg
aaaaagctgg ggtccaagaa gccacaaaag 4500ccaattcctc gaccagggaa caaaatccaa
ggatgtatat ttgacctagt gacaaatcaa 4560gcctttgata ttagtatcat ggttcttatc
tgtctcaaca tggtaaccat gatggtagaa 4620aaggagggtc aaagtcaaca tatgactgaa
gttttatatt ggataaatgt ggtttttata 4680atccttttca ctggagaatg tgtgctaaaa
ctgatctccc tcagacacta ctacttcact 4740gtaggatgga atatttttga ttttgtggtt
gtgattatct ccattgtagg tatgtttcta 4800gctgatttga ttgaaacgta ttttgtgtcc
cctaccctgt tccgagtgat ccgtcttgcc 4860aggattggcc gaatcctacg tctagtcaaa
ggagcaaagg ggatccgcac gctgctcttt 4920gctttgatga tgtcccttcc tgcgttgttt
aacatcggcc tcctgctctt cctggtcatg 4980ttcatctacg ccatctttgg aatgtccaac
tttgcctatg ttaaaaagga agatggaatt 5040aatgacatgt tcaattttga gacctttggc
aacagtatga tttgcctgtt ccaaattaca 5100acctctgctg gctgggatgg attgctagca
cctattctta acagtaagcc acccgactgt 5160gacccaaaaa aagttcatcc tggaagttca
gttgaaggag actgtggtaa cccatctgtt 5220ggaatattct actttgttag ttatatcatc
atatccttcc tggttgtggt gaacatgtac 5280attgcagtca tactggagaa ttttagtgtt
gccactgaag aaagtactga acctctgagt 5340gaggatgact ttgagatgtt ctatgaggtt
tgggagaagt ttgatcccga tgcgacccag 5400tttatagagt tctctaaact ctctgatttt
gcagctgccc tggatcctcc tcttctcata 5460gcaaaaccca acaaagtcca gctcattgcc
atggatctgc ccatggttag tggtgaccgg 5520atccattgtc ttgacatctt atttgctttt
acaaagcgtg ttttgggtga gagtggggag 5580atggattctc ttcgttcaca gatggaagaa
aggttcatgt ctgcaaatcc ttccaaagtg 5640tcctatgaac ccatcacaac cacactaaaa
cggaaacaag aggatgtgtc tgctactgtc 5700attcagcgtg cttatagacg ttaccgctta
aggcaaaatg tcaaaaatat atcaagtata 5760tacataaaag atggagacag agatgatgat
ttactcaata aaaaagatat ggcttttgat 5820aatgttaatg agaactcaag tccagaaaaa
acagatgcca cttcatccac cacctctcca 5880ccttcatatg atagtgtaac aaagccagac
aaagagaaat atgaacaaga cagaacagaa 5940aaggaagaca aagggaaaga cagcaaggaa
agcaaaaaat agagcttcat ttttgatata 6000ttgtttacag cctgtgaaag tgatttattt
gtgttaataa aactcttttg aggaagtcta 6060tgccaaaatc ctttttatca aaatattctc
gaaggcagtg cagtcactaa ctctgatttc 6120ctaagaaagg tgggcagcat tagcagatgg
ttatttttgc actgatgatt ctttaagaat 6180cgtaagagaa ctctgtagga attattgatt
atagcataca aaagtgattc agttttttgg 6240tttttaataa atcagaagac catgtagaaa
acttttacat ctgccttgtc atcttttcac 6300aggattgtaa ttagtcttgt ttcccatgta
aataaacaac acacgcatac agaaaaaaaa 6360aaaaaaa
636710327PRTHomo sapiens 10Met Ala Met
Leu Pro Pro Pro Gly Pro Gln Ser Phe Val His Phe Thr 1 5
10 15 Lys Gln Ser Leu Ala Leu Ile Glu
Gln Arg Ile Ala Glu Arg Lys Ser 20 25
30 Lys Glu Pro Lys Glu Glu Lys Lys Asp Asp Asp Glu Glu
Ala Pro Lys 35 40 45
Pro Ser Ser Asp Leu Glu Ala Gly Lys Gln Leu Pro Phe Ile Tyr Gly 50
55 60 Asp Ile Pro Pro
Gly Met Val Ser Glu Pro Leu Glu Asp Leu Asp Pro 65 70
75 80 Tyr Tyr Ala Asp Lys Lys Thr Phe Ile
Val Leu Asn Lys Gly Lys Thr 85 90
95 Ile Phe Arg Phe Asn Ala Thr Pro Ala Leu Tyr Met Leu Ser
Pro Phe 100 105 110
Ser Pro Leu Arg Arg Ile Ser Ile Lys Ile Leu Val His Ser Leu Phe
115 120 125 Ser Met Leu Ile
Met Cys Thr Ile Leu Thr Asn Cys Ile Phe Met Thr 130
135 140 Met Asn Asn Pro Pro Asp Trp Thr
Lys Asn Val Glu Tyr Thr Phe Thr 145 150
155 160 Gly Ile Tyr Thr Phe Glu Ser Leu Val Lys Ile Leu
Ala Arg Gly Phe 165 170
175 Cys Val Gly Glu Phe Thr Phe Leu Arg Asp Pro Trp Asn Trp Leu Asp
180 185 190 Phe Val Val
Ile Val Phe Ala Tyr Leu Thr Glu Phe Val Asn Leu Gly 195
200 205 Asn Val Ser Ala Leu Arg Thr Phe
Arg Val Leu Arg Ala Leu Lys Thr 210 215
220 Ile Ser Val Ile Pro Gly Leu Lys Thr Ile Val Gly Ala
Leu Ile Gln 225 230 235
240 Ser Val Lys Lys Leu Ser Asp Val Met Ile Leu Thr Val Phe Cys Leu
245 250 255 Ser Val Phe Ala
Leu Ile Gly Leu Gln Leu Phe Met Gly Asn Leu Lys 260
265 270 His Lys Cys Phe Arg Asn Ser Leu Glu
Asn Asn Glu Thr Leu Glu Ser 275 280
285 Ile Met Asn Thr Leu Glu Ser Glu Glu Asp Phe Arg Lys Tyr
Phe Tyr 290 295 300
Tyr Leu Glu Gly Ser Lys Asp Ala Leu Leu Cys Gly Phe Ser Thr Asp 305
310 315 320 Ser Gly Gln Cys Pro
Glu Gly 325 116367DNAHomo sapiens 11ctcttatgtg
aggagctgaa gaggaattaa aatatacagg atgaaaagat ggcaatgttg 60cctcccccag
gacctcagag ctttgtccat ttcacaaaac agtctcttgc cctcattgaa 120caacgcattg
ctgaaagaaa atcaaaggaa cccaaagaag aaaagaaaga tgatgatgaa 180gaagccccaa
agccaagcag tgacttggaa gctggcaaac aactgccctt catctatggg 240gacattcctc
ccggcatggt gtcagagccc ctggaggact tggaccccta ctatgcagac 300aaaaagactt
tcatagtatt gaacaaaggg aaaacaatct tccgtttcaa tgccacacct 360gctttatata
tgctttctcc tttcagtcct ctaagaagaa tatctattaa gattttagta 420cactccttat
tcagcatgct catcatgtgc actattctga caaactgcat atttatgacc 480atgaataacc
cgccggactg gaccaaaaat gtcgagtaca cttttactgg aatatatact 540tttgaatcac
ttgtaaaaat ccttgcaaga ggcttctgtg taggagaatt cacttttctt 600cgtgacccgt
ggaactggct ggattttgtc gtcattgttt ttgcgtattt aacagaattt 660gtaaacctag
gcaatgtttc agctcttcga actttcagag tattgagagc tttgaaaact 720atttctgtaa
tcccaggcct gaagacaatt gtaggggctt tgatccagtc agtgaagaag 780ctttctgatg
tcatgatcct gactgtgttc tgtctgagtg tgtttgcact aattggacta 840cagctgttca
tgggaaacct gaagcataaa tgttttcgaa attcacttga aaataatgaa 900acattagaaa
gcataatgaa taccctagag agtgaagaag actttagaaa atatttttat 960tacttggaag
gatccaaaga tgctctcctt tgtggtttca gcacagattc aggtcagtgt 1020ccagaggggt
acacctgtgt gaaaattggc agaaaccctg attatggcta cacgagcttt 1080gacactttca
gctgggcctt cttagccttg tttaggctaa tgacccaaga ttactgggaa 1140aacctttacc
aacagacgct gcgtgccgct ggcaaaacct acatgatctt ctttgtcgta 1200gtgattttcc
tgggctcctt ttatctaata aacttgatcc tggctgtggt tgccatggca 1260tatgaagaac
agaaccaggc aaacattgaa gaagctaaac agaaagaatt agagtttcaa 1320cagatgttag
accgacttaa aaaagagcaa gaagaagctg aggcaattgc agcggcagcg 1380gctgaatata
caagtattag gagaagcaga attatgggcc tctcagagag ttcttctgaa 1440acatccaaac
tgagctctaa aagtgctaaa gaaagaagaa acagaagaaa gaaaaagaat 1500caaaagaagc
tctccagtgg agaggaaaag ggagatgctg agaaattgtc gaaatcagaa 1560tcagaggaca
gcatcagaag aaaaagtttc caccttggtg tcgaagggca taggcgagca 1620catgaaaaga
ggttgtctac ccccaatcag tcaccactca gcattcgtgg ctccttgttt 1680tctgcaaggc
gaagcagcag aacaagtctt tttagtttca aaggcagagg aagagatata 1740ggatctgaga
ctgaatttgc cgatgatgag cacagcattt ttggagacaa tgagagcaga 1800aggggctcac
tgtttgtgcc ccacagaccc caggagcgac gcagcagtaa catcagccaa 1860gccagtaggt
ccccaccaat gctgccggtg aacgggaaaa tgcacagtgc tgtggactgc 1920aacggtgtgg
tctccctggt tgatggacgc tcagccctca tgctccccaa tggacagctt 1980ctgccagagg
gcacgaccaa tcaaatacac aagaaaaggc gttgtagttc ctatctcctt 2040tcagaggata
tgctgaatga tcccaacctc agacagagag caatgagtag agcaagcata 2100ttaacaaaca
ctgtggaaga acttgaagag tccagacaaa aatgtccacc ttggtggtac 2160agatttgcac
acaaattctt gatctggaat tgctctccat attggataaa attcaaaaag 2220tgtatctatt
ttattgtaat ggatcctttt gtagatcttg caattaccat ttgcatagtt 2280ttaaacacat
tatttatggc tatggaacac cacccaatga ctgaggaatt caaaaatgta 2340cttgctatag
gaaatttggt ctttactgga atctttgcag ctgaaatggt attaaaactg 2400attgccatgg
atccatatga gtatttccaa gtaggctgga atatttttga cagccttatt 2460gtgactttaa
gtttagtgga gctctttcta gcagatgtgg aaggattgtc agttctgcga 2520tcattcagac
tgctctgagt cttcaagttg gcaaaatcct ggccaacatt gaacatgctg 2580attaagatca
ttggtaactc agtaggggct ctaggtaacc tcaccttagt gttggccatc 2640atcgtcttca
tttttgctgt ggtcggcatg cagctctttg gtaagagcta caaagaatgt 2700gtctgcaaga
tcaatgatga ctgtacgctc ccacggtggc acatgaacga cttcttccac 2760tccttcctga
ttgtgttccg cgtgctgtgt ggagagtgga tagagaccat gtgggactgt 2820atggaggtcg
ctggtcaagc tatgtgcctt attgtttaca tgatggtcat ggtcattgga 2880aacctggtgg
tcctaaacct atttctggcc ttattattga gctcatttag ttcagacaat 2940cttacagcaa
ttgaagaaga ccctgatgca aacaacctcc agattgcagt gactagaatt 3000aaaaagggaa
taaattatgt gaaacaaacc ttacgtgaat ttattctaaa agcattttcc 3060aaaaagccaa
agatttccag ggagataaga caagcagaag atctgaatac taagaaggaa 3120aactatattt
ctaaccatac acttgctgaa atgagcaaag gtcacaattt cctcaaggaa 3180aaagataaaa
tcagtggttt tggaagcagc gtggacaaac acttgatgga agacagtgat 3240ggtcaatcat
ttattcacaa tcccagcctc acagtgacag tgccaattgc acctggggaa 3300tccgatttgg
aaaatatgaa tgctgaggaa cttagcagtg attcggatag tgaatacagc 3360aaagtgagat
taaaccggtc aagctcctca gagtgcagca cagttgataa ccctttgcct 3420ggagaaggag
aagaagcaga ggctgaacct atgaattccg atgagccaga ggcctgtttc 3480acagatggtt
gtgtacggag gttctcatgc tgccaagtta acatagagtc agggaaagga 3540aaaatctggt
ggaacatcag gaaaacctgc tacaagattg ttgaacacag ttggtttgaa 3600agcttcattg
tcctcatgat cctgctcagc agtggtgccc tggcttttga agatatttat 3660attgaaagga
aaaagaccat taagattatc ctggagtatg cagacaagat cttcacttac 3720atcttcattc
tggaaatgct tctaaaatgg atagcatatg gttataaaac atatttcacc 3780aatgcctggt
gttggctgga tttcctaatt gttgatgttt ctttggttac tttagtggca 3840aacactcttg
gctactcaga tcttggcccc attaaatccc ttcggacact gagagcttta 3900agacctctaa
gagccttatc tagatttgaa ggaatgaggg tcgttgtgaa tgcactcata 3960ggagcaattc
cttccatcat gaatgtgcta cttgtgtgtc ttatattctg gctgatattc 4020agcatcatgg
gagtaaattt gtttgctggc aagttctatg agtgtattaa caccacagat 4080gggtcacggt
ttcctgcaag tcaagttcca aatcgttccg aatgttttgc ccttatgaat 4140gttagtcaaa
atgtgcgatg gaaaaacctg aaagtgaact ttgataatgt cggacttggt 4200tacctatctc
tgcttcaagt tgcaactttt aagggatgga cgattattat gtatgcagca 4260gtggattctg
ttaatgtaga caagcagccc aaatatgaat atagcctcta catgtatatt 4320tattttgtcg
tctttatcat ctttgggtca ttcttcactt tgaacttgtt cattggtgtc 4380atcatagata
atttcaacca acagaaaaag aagcttggag gtcaagacat ctttatgaca 4440gaagaacaga
agaaatacta taatgcaatg aaaaagctgg ggtccaagaa gccacaaaag 4500ccaattcctc
gaccagggaa caaaatccaa ggatgtatat ttgacctagt gacaaatcaa 4560gcctttgata
ttagtatcat ggttcttatc tgtctcaaca tggtaaccat gatggtagaa 4620aaggagggtc
aaagtcaaca tatgactgaa gttttatatt ggataaatgt ggtttttata 4680atccttttca
ctggagaatg tgtgctaaaa ctgatctccc tcagacacta ctacttcact 4740gtaggatgga
atatttttga ttttgtggtt gtgattatct ccattgtagg tatgtttcta 4800gctgatttga
ttgaaacgta ttttgtgtcc cctaccctgt tccgagtgat ccgtcttgcc 4860aggattggcc
gaatcctacg tctagtcaaa ggagcaaagg ggatccgcac gctgctcttt 4920gctttgatga
tgtcccttcc tgcgttgttt aacatcggcc tcctgctctt cctggtcatg 4980ttcatctacg
ccatctttgg aatgtccaac tttgcctatg ttaaaaagga agatggaatt 5040aatgacatgt
tcaattttga gacctttggc aacagtatga tttgcctgtt ccaaattaca 5100acctctgctg
gctgggatgg attgctagca cctattctta acagtaagcc acccgactgt 5160gacccaaaaa
aagttcatcc tggaagttca gttgaaggag actgtggtaa cccatctgtt 5220ggaatattct
actttgttag ttatatcatc atatccttcc tggttgtggt gaacatgtac 5280attgcagtca
tactggagaa ttttagtgtt gccactgaag aaagtactga acctctgagt 5340gaggatgact
ttgagatgtt ctatgaggtt tgggagaagt ttgatcccga tgcgacccag 5400tttatagagt
tctctaaact ctctgatttt gcagctgccc tggatcctcc tcttctcata 5460gcaaaaccca
acaaagtcca gctcattgcc atggatctgc ccatggttag tggtgaccgg 5520atccattgtc
ttgacatctt atttgctttt acaaagcgtg ttttgggtga gagtggggag 5580atggattctc
ttcgttcaca gatggaagaa aggttcatgt ctgcaaatcc ttccaaagtg 5640tcctatgaac
ccatcacaac cacactaaaa cggaaacaag aggatgtgtc tgctactgtc 5700attcagcgtg
cttatagacg ttaccgctta aggcaaaatg tcaaaaatat atcaagtata 5760tacataaaag
atggagacag agatgatgat ttactcaata aaaaagatat ggcttttgat 5820aatgttaatg
agaactcaag tccagaaaaa acagatgcca cttcatccac cacctctcca 5880ccttcatatg
atagtgtaac aaagccagac aaagagaaat atgaacaaga cagaacagaa 5940aaggaagaca
aagggaaaga cagcaaggaa agcaaaaaat agagcttcat ttttgatata 6000ttgtttacag
cctgtgaaag tgatttattt gtgttaataa aactcttttg aggaagtcta 6060tgccaaaatc
ctttttatca aaatattctc gaaggcagtg cagtcactaa ctctgatttc 6120ctaagaaagg
tgggcagcat tagcagatgg ttatttttgc actgatgatt ctttaagaat 6180cgtaagagaa
ctctgtagga attattgatt atagcataca aaagtgattc agttttttgg 6240tttttaataa
atcagaagac catgtagaaa acttttacat ctgccttgtc atcttttcac 6300aggattgtaa
ttagtcttgt ttcccatgta aataaacaac acacgcatac agaaaaaaaa 6360aaaaaaa
636712829PRTHomo
sapiens 12Met Ala Met Leu Pro Pro Pro Gly Pro Gln Ser Phe Val His Phe Thr
1 5 10 15 Lys Gln
Ser Leu Ala Leu Ile Glu Gln Arg Ile Ala Glu Arg Lys Ser 20
25 30 Lys Glu Pro Lys Glu Glu Lys
Lys Asp Asp Asp Glu Glu Ala Pro Lys 35 40
45 Pro Ser Ser Asp Leu Glu Ala Gly Lys Gln Leu Pro
Phe Ile Tyr Gly 50 55 60
Asp Ile Pro Pro Gly Met Val Ser Glu Pro Leu Glu Asp Leu Asp Pro 65
70 75 80 Tyr Tyr Ala
Asp Lys Lys Thr Phe Ile Val Leu Asn Lys Gly Lys Thr 85
90 95 Ile Phe Arg Phe Asn Ala Thr Pro
Ala Leu Tyr Met Leu Ser Pro Phe 100 105
110 Ser Pro Leu Arg Arg Ile Ser Ile Lys Ile Leu Val His
Ser Leu Phe 115 120 125
Ser Met Leu Ile Met Cys Thr Ile Leu Thr Asn Cys Ile Phe Met Thr 130
135 140 Met Asn Asn Pro
Pro Asp Trp Thr Lys Asn Val Glu Tyr Thr Phe Thr 145 150
155 160 Gly Ile Tyr Thr Phe Glu Ser Leu Val
Lys Ile Leu Ala Arg Gly Phe 165 170
175 Cys Val Gly Glu Phe Thr Phe Leu Arg Asp Pro Trp Asn Trp
Leu Asp 180 185 190
Phe Val Val Ile Val Phe Ala Tyr Leu Thr Glu Phe Val Asn Leu Gly
195 200 205 Asn Val Ser Ala
Leu Arg Thr Phe Arg Val Leu Arg Ala Leu Lys Thr 210
215 220 Ile Ser Val Ile Pro Gly Leu Lys
Thr Ile Val Gly Ala Leu Ile Gln 225 230
235 240 Ser Val Lys Lys Leu Ser Asp Val Met Ile Leu Thr
Val Phe Cys Leu 245 250
255 Ser Val Phe Ala Leu Ile Gly Leu Gln Leu Phe Met Gly Asn Leu Lys
260 265 270 His Lys Cys
Phe Arg Asn Ser Leu Glu Asn Asn Glu Thr Leu Glu Ser 275
280 285 Ile Met Asn Thr Leu Glu Ser Glu
Glu Asp Phe Arg Lys Tyr Phe Tyr 290 295
300 Tyr Leu Glu Gly Ser Lys Asp Ala Leu Leu Cys Gly Phe
Ser Thr Asp 305 310 315
320 Ser Gly Gln Cys Pro Glu Gly Tyr Thr Cys Val Lys Ile Gly Arg Asn
325 330 335 Pro Asp Tyr Gly
Tyr Thr Ser Phe Asp Thr Phe Ser Trp Ala Phe Leu 340
345 350 Ala Leu Phe Arg Leu Met Thr Gln Asp
Tyr Trp Glu Asn Leu Tyr Gln 355 360
365 Gln Thr Leu Arg Ala Ala Gly Lys Thr Tyr Met Ile Phe Phe
Val Val 370 375 380
Val Ile Phe Leu Gly Ser Phe Tyr Leu Ile Asn Leu Ile Leu Ala Val 385
390 395 400 Val Ala Met Ala Tyr
Glu Glu Gln Asn Gln Ala Asn Ile Glu Glu Ala 405
410 415 Lys Gln Lys Glu Leu Glu Phe Gln Gln Met
Leu Asp Arg Leu Lys Lys 420 425
430 Glu Gln Glu Glu Ala Glu Ala Ile Ala Ala Ala Ala Ala Glu Tyr
Thr 435 440 445 Ser
Ile Arg Arg Ser Arg Ile Met Gly Leu Ser Glu Ser Ser Ser Glu 450
455 460 Thr Ser Lys Leu Ser Ser
Lys Ser Ala Lys Glu Arg Arg Asn Arg Arg 465 470
475 480 Lys Lys Lys Asn Gln Lys Lys Leu Ser Ser Gly
Glu Glu Lys Gly Asp 485 490
495 Ala Glu Lys Leu Ser Lys Ser Glu Ser Glu Asp Ser Ile Arg Arg Lys
500 505 510 Ser Phe
His Leu Gly Val Glu Gly His Arg Arg Ala His Glu Lys Arg 515
520 525 Leu Ser Thr Pro Asn Gln Ser
Pro Leu Ser Ile Arg Gly Ser Leu Phe 530 535
540 Ser Ala Arg Arg Ser Ser Arg Thr Ser Leu Phe Ser
Phe Lys Gly Arg 545 550 555
560 Gly Arg Asp Ile Gly Ser Glu Thr Glu Phe Ala Asp Asp Glu His Ser
565 570 575 Ile Phe Gly
Asp Asn Glu Ser Arg Arg Gly Ser Leu Phe Val Pro His 580
585 590 Arg Pro Gln Glu Arg Arg Ser Ser
Asn Ile Ser Gln Ala Ser Arg Ser 595 600
605 Pro Pro Met Leu Pro Val Asn Gly Lys Met His Ser Ala
Val Asp Cys 610 615 620
Asn Gly Val Val Ser Leu Val Asp Gly Arg Ser Ala Leu Met Leu Pro 625
630 635 640 Asn Gly Gln Leu
Leu Pro Glu Gly Thr Thr Asn Gln Ile His Lys Lys 645
650 655 Arg Arg Cys Ser Ser Tyr Leu Leu Ser
Glu Asp Met Leu Asn Asp Pro 660 665
670 Asn Leu Arg Gln Arg Ala Met Ser Arg Ala Ser Ile Leu Thr
Asn Thr 675 680 685
Val Glu Glu Leu Glu Glu Ser Arg Gln Lys Cys Pro Pro Trp Trp Tyr 690
695 700 Arg Phe Ala His Lys
Phe Leu Ile Trp Asn Cys Ser Pro Tyr Trp Ile 705 710
715 720 Lys Phe Lys Lys Cys Ile Tyr Phe Ile Val
Met Asp Pro Phe Val Asp 725 730
735 Leu Ala Ile Thr Ile Cys Ile Val Leu Asn Thr Leu Phe Met Ala
Met 740 745 750 Glu
His His Pro Met Thr Glu Glu Phe Lys Asn Val Leu Ala Ile Gly 755
760 765 Asn Leu Val Phe Thr Gly
Ile Phe Ala Ala Glu Met Val Leu Lys Leu 770 775
780 Ile Ala Met Asp Pro Tyr Glu Tyr Phe Gln Val
Gly Trp Asn Ile Phe 785 790 795
800 Asp Ser Leu Ile Val Thr Leu Ser Leu Val Glu Leu Phe Leu Ala Asp
805 810 815 Val Glu
Gly Leu Ser Val Leu Arg Ser Phe Arg Leu Leu 820
825 13 2521DNAHomo sapiens 13tcgtcattgt
ttttgcgtaa gtactttcag ctttttgaaa cggcaaattt atgaaaatct 60caggcagcac
agtgttccaa taacagcaaa cctctggggt ctgatttttc cagcatggaa 120tgtctgtatt
taaaggaaaa tggaagctat atgaaagcta gtcattgata ggcaaatact 180aaatttttca
aaaagactat ctcccactca gaggtatgtg acatcattga gattgatcat 240cttcctccat
gctcagcact tggatggata gtctatgcag tgccaattag gtttacactt 300tttccaagtt
cttgtttaaa aatgaataaa acctataaga agcacacaaa aaactcaggc 360ctttaaatac
aagtacagtg ctctttatat taccctgtat tattcaatta ttttatattt 420ttgtattaaa
atataaaagt atatatttgt aaaagagctg tcactgctta ctaagcctaa 480gattaatgca
ttgtatacac attatatata cactcagttt attttcaagg aaaatattac 540attacagtat
aattttaatt tattcataat tgttactctt ttcccaaatg tatttatcag 600aaaaatggat
ttttacaatc tcatatctag atattctatt ttccgttaat gactttgaat 660tatcaaagta
atccaggttc actggagcca cctagacaat acagaagggt cttaagattc 720ctaatatccg
attccttcat ccctgcctaa cctcttttcc atcaggctcc ctccttttga 780ggtcacatat
gatggggcag ttccatctgc tacctgctgc taagaaaaca ttgtgcaaag 840aagactgcct
cttctcatag gtctcctttt tttcctattg gtacaattca aatactacaa 900aattcacaat
tttaaagtat acaattcagt gtttttcagt atattgacaa acttgcgcaa 960catcaccact
acctaattag agaagtttaa caaccccaaa tagaaactct gtactcatta 1020gcagtcactt
caatttcctt ctctccctgg ccccttggaa accactcatc tatgttttat 1080ctctatggat
ttgtctattt ggattttttt atgaatggaa atatcaaaca gtatgcagcc 1140ttttgtgtct
gacacttagc atactatttt caatgtttta taaaaaatta aaagtctaaa 1200ttaccataat
gttaagggtc caacaattga aacatttagt tgtttacgca tttaagtgta 1260tgaatccaat
ttaataatat ataatatctg tattttgttt gcatcaaatg agtcaaatca 1320atgctaatta
aattaagtta gttatgatag caaaaaagac ttgcagaact atacatttaa 1380aatagaagcc
ccaaacgtag aaaatacctt gactaaaggc tcaggtaagt atcatagact 1440ctatctaaat
tctgaataat tctgatttaa ttctacaggt atttaacaga atttgtaaac 1500ctaggcaatg
tttcagctct tcgaactttc agagtattga gagctttgaa aactatttct 1560gtaatcccag
gtaagaagta attggtgtga agcattaggc cactcataac tccaactatt 1620tgggagttgt
tctttgttcg tttgtgtgtg tgtcgtcact gtgtgtataa actcccctat 1680tacagatatg
tgacagagtt tgtggacctg ggcaatgtct cagcattgag aacattcaga 1740gttctccgag
cattgaaaac aatttcagtc attccaggtg agagtggggt taaacaccag 1800ggctgacttt
gatcttttga aagaaagaca taaaaaaaac ctttctttat gcaattttaa 1860ttaaatttga
tttcttctgt tttccacaat cattatcaaa agtcaacctt tgagtttaat 1920caattgcatg
ggtctttagg atgaggatga ctacttggct cacatctcaa cttgaaatct 1980tccctcctgt
ctgagggttt gtttttccat ttgcttatac aggaaggaaa atttcaatcc 2040attttaattg
attttaaaat acttaggaaa gagatcataa atatcagcat ggggatattg 2100catgacattg
ttttttattt tttctcagca aaccatttct ttattgaaaa ttattacaag 2160gtgtcactat
atgctgtaat ttgttttggt gccatttata gagaagtggt ttgtttgaca 2220aacattagaa
cttacaaaaa ttatttttaa agtggtttga atagccaaat tgttcatttt 2280gacaacctag
gtattctttg agaatatgtt cctattttta accatttgct gtaatatata 2340aaaacaaagt
atatataaaa tagcaaaccc agaaggcttt ttttgaatta ggttacttaa 2400ggtcattgat
ttgatataat gcatgacttt ctaggaaagc ttgtgttttt tgtttcgatt 2460cagaggcttt
atgtcattac aaacactttt ttctcccatt ttcaggcctg aagacaattg 2520t
2521149963DNAHomo
sapiens 14aaatattaaa acattgaaaa actttttttt accctttttg gtttatcatc
ttccttctct 60ccttcaactt ccattttttt cattcagttt ttcccctcat tccaactctt
ctttctttaa 120ttgaacataa tagagactcg gagttcaaaa tatgtgttgt ttctattgaa
ttgatagatt 180tttttttttt ttttgagaca gggtctctgt ctgtcatcca agctggaccg
cagtggtgtg 240atcatggctc actgcagact caacctcccc aggctcaggt gatcctccca
ccttagctac 300tctcaagcag ctgagactac aggtgcacag ctccatgcct ggctaatttt
tgtacttttc 360ctagagacag ggtgtcactg tgttgcccat gctgttcttg aactcctgcc
ctcaagtgat 420tcacccgcct tggcctccga aaatgttgga attacaggca cgagccactg
tgcctggtct 480attgaatgga atttatgttt atcctttcca attttaaaat agttaacttt
aaagttgtgt 540attttacatt cattgtcttc aaaccaaaca aaatccaaaa acttttctac
tttttgaatt 600gaattttttc tctccttttg aataataaaa cctggtcatt gatgtatgtg
tactcatctg 660atatttttga tttaacaaaa tcaaatggag ggttcacgtt agtattttat
gctttttaaa 720agtagacatg cttatgctct tccctaaaac ttttatttca gcattactta
gataagccct 780ggcatctgta tttttttagg ttacacagat gatttttttg tataccctgg
tcaatagcca 840cagtggagta tacctgcatt ctgatatata cactcttgac aaccatagag
acaaatgata 900cttagctgca tttttaacat tacctaacat aagtgaaagg gattttaaaa
cagtagtatt 960ttggaaaaca gcagataata aaacattaat tttgtgttgt gggttattgg
tattaagtat 1020catgtaatgc ccacatctca aagctcacta tggaagaatt ggtgtcagat
aataggaaga 1080ttgatattgt tgaaaaccat gcattcagca agccagccat tggtagcacc
caatgaagtg 1140ttcagtattc atctaagcag agaaagtaga agacacaatc tttactgtaa
ctttgagtgt 1200ttgcagtgtg tcagatggtc ccttgaagaa gaaatattaa taaaaatgaa
ttgcaaattg 1260tctccaattt acctgaaaga gatcaaaata atgtatctaa atatatggtg
tttgaaaggt 1320gtagagtcag tccatatttt tcaagcccag tgaaggtaac agaacagttt
ctacttaagc 1380agctgtttgt tggtttcttt ggctcaagag agaggtataa tatcttcttt
tattttacag 1440ctttttaatc ataataataa ctattataaa attaacaaat gcacatggta
actaattcaa 1500acaatttttg ctttatccct ttaaagccat tatctaggga ataatcactt
cccgcttttt 1560taaaaaaaag tttttctgat attttccttt ttatctctaa ataatatgct
tatgtctcat 1620ttttatttca tcaacttatg taatgttttc taactttcta atttgaaaat
gaggaattta 1680ttttatttat gtaaactttt ctctttctgc cccattctca aattccaagc
agttatgttt 1740tcatttctaa gtaattctat gggttggatt tataccttta aatggaatat
acctgaacct 1800ctatttcttg ttacttaaac tttagccagg atgcactaac tgccttcaat
gtaagataag 1860gggtcagctc tccaactttt ccctcaccac taatccccct ctacctctca
atttttgtaa 1920gctattccat tacttttaca ctgataatat ttaacacata tatgtgtata
tcttgtcttt 1980gtgtaaataa agttctataa ttgtacttac attttatata tgctttggta
attggcacat 2040ttgtgagatg attaactgat acaatttaac aatattataa ttatgtatgt
atttttcatt 2100ttaaaacaag taacatggtt tgatctagag gaggaattta aacctctcat
tcaaactctg 2160ctacttgaag aacacatcaa actcaagtgg atcctctctg cttcaattct
accagttgtt 2220caagttcatg cctcattttg aattgattcc tatttggaca atatggctat
ttacacgttc 2280taaagaaaca aattcccttt gcttttattc acactatttg tgaaacatct
tgtattctat 2340gtgttttaga agtacttttt tttttttgct ctaccttaat ccaaagtaac
ttaaaaaaaa 2400aagataaaaa ggagttaaat tttgaatttt cttataccta gaaatatatt
ttgtttgcca 2460tcacacttga taataggttg tctgataata gaatttgatt caaaatattt
tctctcatca 2520cactgaaagc attgattgtg ttctagtgtc aattatggct aaggaaattt
taatggcaat 2580ctaattttca ttctttttag ttagttatct tatttttgga agattataaa
atcttgtttc 2640tattgttaat gctctaaact ttcacaagaa tggaactttt aaattgatct
tcttttgtac 2700ttttagcttg atgactcttg gtgaattttc tgccattact ttgtttatta
tttcctatct 2760actatttgct gctctcttga gactaatgtc gaacaacatg gattattctg
tatcttctaa 2820gttttatttt gtactttttt aatttttaaa attctcctaa aatatataga
aaaatctaag 2880atatagaact ccatgcattt ttacatatac atattttccc atgtaaccat
cacatagatc 2940aaggtataga acgtttctag cacctgagaa tgctcactgg tgccccattc
tggtaaagtg 3000ccccttttgg gtaaagtaaa aggtaactac aattatgact tctccttttc
agcaagcatt 3060ctactgaaat tttttgtttc acaaagcatg acctttaccc agaattgatg
ttgttgtttt 3120tggtagccta tttttttttt ttaataaaag atactctgtc tcctcaaatt
tctctaggaa 3180tactaacttt aacttctcaa tatcccacaa tgtttgcttt tttttttcta
aagttaaaag 3240tttggtctgt tcatcttagt ctttcaattt tatgtttggt agatcttcat
taccaattta 3300ttcttgataa tctattaata tttatggaag aaaggccagg ttggtgttgg
ccaggtttgc 3360atctgtcttt atacaggtct gtttcccaga caaatcctct cttgaaagag
agggatgact 3420gaaaggtctg tgtatataca tgtatggtgt actgacaggc aggctacatt
ttaggatgta 3480tggatgagga gtagataagg gagtcacctt tccctcgact ccaccctggt
tctagctaaa 3540atggacatgg ataccctagg agtcctacct taattaatgc agatattatc
cttttggtga 3600tagcttggag ataaaccact ctgctacata tttaacatag agagggaagt
gggacaatgt 3660tttctgcaac attattgaat aaataatcca tggattaatc aacttgtttt
ctgatctaca 3720tcacatattt gccatagaaa tctgtgactt ctctgctaga aagaaattct
aacattctca 3780gatgagtgtt ctagactgta gcttctgctt tgatctcctc actagggtaa
tcttatgatt 3840tatgataact ttttttgtct tcctctggtt tttagtgata ttatctaaca
tttcgccact 3900gtacatcttt gtttttactg caatggaaat ttgaaggcca gagagtgtga
taaacacatg 3960ggataagacc attttcatga actgggagct gacattgctt tgactggaaa
gatgtgtgaa 4020gaaaaataga tgctagatag acaaatacat gatatgatat ttggacagac
atagatccat 4080agatggataa gcatagataa aacatatcta tatatatgtg tgtgtctgca
tacatagaaa 4140ataagaaaga gttcagaata aatatttcca tattgatttt aactatagga
aatcttagat 4200acaagaataa cacattcatt ttttaatttt agtagttgcc taagtattct
ctaatatgag 4260attaatggtg agtgggtaag atagtaatga gaaaacaaaa gtactgcctt
tatatgcaac 4320attgggattt cattctaaag aagatacaat catagtaatg ttaataaaag
tggaaatgta 4380gcaatatatt tttcaaaatt gtaaattaaa catgacaaga ataaatattg
attgatttca 4440ttgactatat aaaattaggt caaataattt taaatgattt ttttcaaagt
atattttctt 4500tcttctggat agaaaaacat gacattttcg ttgtacaaat ttgaaaagta
aagtataaag 4560acataaaaat aatccataat ctcttaacta gggctaaact ttgttaaatg
cattgtgtat 4620ttaaattaaa tatcatttta aaaattatta ttaatttaca tctatctgtg
tgtataacct 4680ggattccact tattatgcat tatattttta taattcattt aattcattaa
atagtctttg 4740aacgctctct ttcaaagctg ctcattattt cattatacat tcattacttc
atttagtgta 4800acccttactg ttaggatttt ggattgatta taatttttca atattataaa
aatattgtag 4860taaatatatt tatacatgtt agaggacacc tgagattact gtctttgaat
aaattcctag 4920aaggaaaatt atttggcaat ataaacgaat tcacgtgttt aaggcttttg
atagatatta 4980caaaattacc ctcaaaaagt ttgtacactt tatttattta tttgagacgg
agtctcgctc 5040tattgcccag gctggagtgc agtggcacga tcttggctca ctgcaagctc
cgcctcctgg 5100gttcacgcca ttctctgcct cagcctcccg agcagctggg actataggcg
cctgccaaca 5160tgcctggcta atttttttgt acttttagta gagacagggt ttcacagtgt
tcgccaggtt 5220ggtctcgatc tcctgacctc atgatctgcc tgcctgggcc tcccaaagtt
ctgggattat 5280aggcgtgagc caccgcaccc agacagtaca cattatttat tactttcatc
cagacaggca 5340acactgattt cccctataac ctctctaata tcactattac ttttgttaaa
atctttgcta 5400atgattaata ggccaaattt ggttccccat gtttatttta atatttgttt
ctttgattac 5460tagtgaggtt tgacatggtt acaagtgttt attaaccatt tgtattttta
gataattatg 5520ttttttacta aattttatga gatatattaa tatctttatt cattaaagga
aaagcacttc 5580atctgtcatt tgtattgtac tttttttatt ttgtggatgc cttccacttt
taaaaacctt 5640cattgttatg agattaaatc tagcattctt tatctttttc gattttgcct
ttggtgtaat 5700gctttacaag tcctctctac tctcacatta aataagtatt cattttcact
tttattagaa 5760tgtttgagct ttatttttcc atttaaatat ttactccacc agaaatctca
aatcttaaac 5820taaatgcctg actaatgaaa actcctgtgg cctgtttatc cagaagacta
agggaagagg 5880aaagaagttg tagtcacaca taggtcctgc cccatcattg catcttcttt
gtgtgtgttg 5940gtgaagtgtg aggctccatt tttacttatg ttgccctctg taaccctgtc
agcctatcat 6000cttgccaccc cactggcctg gtacaataac ttccaagcaa actcagatgc
caactaaagc 6060tgctcttctt caaagctata aaattagtaa aatgaaagga agtttcacct
ctccaaacta 6120gaggatgatg atgaaatctg atcaaggtct cttcaacttc catttcacag
atcatgatgt 6180taagcaggtg cttagtgtct tcctttaatt tagcttgatc agggaaatat
tgcccatgca 6240tggtaccact agcattgacc accctttgcc tctgcttttt tcatatactt
tagtcattct 6300ggagctgtgt tgagcatgtg catgggggtt gggggagaca gagacagagg
agaataggaa 6360tattggaaga atcaaactgg taatggacat ctgagttcac aacatctttg
tacctagatt 6420tctgtcaaat aattcttgaa agagatggta ccctccagtt gaatgtagaa
tctacaaaaa 6480caccagttat tcaagccaac ttttcttcaa gagccacagt atgtctacac
tgaaagtcta 6540gtgaggcaag ttatcccgtc ttctcacctt tatgatttcc aaaagtgttg
aggaatctcc 6600aaagcacaag tggcaactgc tttgatttga ttcattgatt tgtttccttt
ttactaaatc 6660tcttcctatt ctgaggacaa ttttcacata ttttaggatt caaaattctt
aatgcatatt 6720catagtagtt ttattaagat ataatttaca caccataaaa aatcaccatt
ttaaagtgta 6780tcactcagtg gtttttagta tgttcacaaa ggtatgcaac aatcatcatt
atgtagtttt 6840agaatatttt tatcacccca aaagaagccc catactcttt aaccattaac
cacataacca 6900tcaataatca cttcctatta cacccagcct ctgacaaaaa atactctaca
ttttgtttct 6960ggggatttgc ctattctgga catttcataa gatggaatca tatcgtatgt
ggccttttgg 7020ctctgtgttc tttcacgggg ttttcatcac atccatgttg taacatgtac
ctatacttca 7080tttgttttta ttgctaaata ctaattcagt gtttgaatat cccacattcc
tttatccatt 7140tattcattgg tgagtaatga ggttgtgttt actttttggc tagtatgaat
aatgctgctg 7200aacattcatg taaacgttct tgtgtggaca tatgttttaa attcccttga
gtctgtacca 7260agtagtggaa ttgccgagtc ataacttact ctttaccttt cagaatcatc
ctacttttga 7320atgtgtatat gaaatatccc ttatttttca ctacattctt taaaaacctt
tatcctaact 7380ttggtttcca tcaaaataca ggcatacttc attttatggg gctttacttt
attgcacttc 7440aagatactgc attctttaca agttgaaggt ttgtggcaac cctgcattaa
gcaagtctgt 7500cagcgtaatt ttttcaacag catgtgctca cttcatgcat ctgtgtcaca
ctttggcaat 7560tctcacaata tttcaacttt gtaattgcca ttatttaatt attactgtat
gtgttatggt 7620gatctatgat cagtgatctt tgttgttact gtggtaattg ttttggggtg
ctatgaacca 7680catccacata agacagcaaa caatcattaa atgtgacgtg ttctgtttca
gcatccctat 7740ctctctccct ttcctcaggc ttccctgttc cctgacacac aacgatgtta
aaattaggcc 7800aattagtaac ccattcccac ccctaatgtt caagtaaagg aagaattgcg
catttctcac 7860tttaaatcag aagctagaaa tgattaagct tagtgaggaa ggcatgttga
aagccaagat 7920aggccaaaat ctaggcctct tgcactaagt acttagccaa gttgtgaatg
caaaggaaac 7980attcttgaag gaaatcaaat gtgctactcc agtgaacaca tgaatgataa
gaaaacaaaa 8040cagccttatt gcaaatatag agaaagtttt agtgatctag atagaagatc
aaccagccac 8100aacatttcct taagccaaag cctaatccac agcaaggcca taactccaat
tgtatgaagg 8160ctgagaggtg aagaagctgg aaaaaaaagg tttgaagctg gcagacggta
ccccatgaga 8220ttgaaggaaa gaagtcactg ccataacata aaaatgcaag gtgaaacagc
aagtgccaat 8280ggagaagatg cagcaagtta ttcagaagat ctagctaaga tcattgataa
gagtggttac 8340attaaacaac agatttgcaa tgaaggtcta acagccttct actagaagaa
tatattatag 8400ctagaaagga gaagtcaatg tctggcttca aagcttcaac gaacaggctg
actctcttgt 8460taggggataa tgcagctggt gactttaaat tgaagcaaat gctaatttat
cgttcagaaa 8520atcttagaat tatattattt aattattcag aatattaata attatttact
attcagaaaa 8580ttacataata attatattat ttactattca gaaaattaca taataattat
attatttact 8640attcagaaaa ttaaataatt atattcttta ctattcagaa aattaaataa
taactatatt 8700atttaattat tcagaaaatt aaataataat tatattattt aatcattcag
aaaatctctt 8760aagaattata ctaaatctac tctgcctgtg ctctataaat agaacaacaa
agcctggatg 8820acagcacgtc tgtttacagc atggttcact aaatatttta ggcccagtgt
tgatacttat 8880agctcaggaa aaaaaaaaaa gatttctttc aaaatatcac tgcctattga
caatgtgcct 8940ggtcactcaa gagctctaac ggagaggtac aagggcatta atgttatttt
catgcctgct 9000aacgctatcc attctgtagc ctatgggcca aagagtaatt tcaactttta
agtcttattt 9060tttaataaat acattttata agactatagc tgccatagat agtgattcct
ctgatgggtc 9120tgggaaaagt aaattgaaaa tcttctggaa aggattcacc attctagatg
ttattagaac 9180attaatgatt tgtgggagga gggcaatata caaacattaa taggaaattg
gaatgagttg 9240attccaactc tcatggatga ctatgagagg tttaagactt cagtgatgga
aagaactgca 9300gatgtggtgg aaatagcaag agaattagaa ttaagagatg gagcctaaaa
ttgtgactga 9360attgctgcca tctcgcggta acttgaatgg atgaggagtt gcttctgatt
gataagcaaa 9420gaaagtggtt tcgtgagatg gaatcttctc ctggtcaaga tgttatgaac
attgttgaag 9480tgacagcaga ggatttagaa tattacataa gtttagttgg taaaacagca
tcaggctgga 9540gagtggtgct accaaaggca ttatcataaa gacttttaaa agaggaaagt
tgagttgaat 9600tgcaccaaag gtaaatatag acgaaaagaa ggggaattta ggaaaattta
ctcagaaaat 9660tgtcattttt tactttaaag ctatgaagcg taaacttcaa atttctgagt
ataaagatgg 9720ttagaaggca cgaaagataa gtcccgccca ttgcctgaca catagtaatc
ccttaacaag 9780ctgataattc tagaacatgt taccttttgt agttgaaact aaaaatctgt
ttatgttgtt 9840attattagtt tttaatgggc ctttcttggc aggcaaatag ttaagtcttt
atttctttgt 9900ttccatccag gcctcttatg tgaggagctg aagaggaatt aaaatataca
ggatgaaaag 9960atg
9963159298DNAMus musculus 15tacaggatga gaagatggcg atgttgcctc
cgccaggacc tcagagcttc gttcacttca 60caaaacagtc ccttgccctc attgaacaac
gcatttctga agaaaaagcc aagggacaca 120aagatgaaaa gaaagatgat gaggaagaag
gtcccaagcc cagtagtgac ttggaagcag 180gtaaacagct acccttcatc tacggagaca
ttcccccggg aatggtgtca gagcccctgg 240aggacctgga cccatactat gcagacaaaa
aaacttttat agtattgaac aaagggaaag 300caatcttccg tttcaacgcc acccctgctt
tgtacatgct gtctcccttc agtcctctca 360gaagaatatc tattaagatt ttagtgcact
ccttattcag catgctaatc atgtgcacaa 420ttctgacaaa ctgcatattc atgaccatga
gcaaccctcc agattggacc aaaaacgtag 480agtacacttt tactgggata tatacttttg
aatcactcat aaaaatcctt gcaagaggct 540tttgcgtggg cgaattcacc ttcctccgtg
acccttggaa ctggctggac tttgttgtca 600ttgtttttgc gtatttaaca gaatttgtaa
acctaggcaa tgtttcagct cttcgaactt 660tcagagtctt gagagctttg aaaactattt
ctgtaattcc aggactaaaa accatcgtgg 720gggccctgat ccaatcagtg aagaagctct
ctgacgtcat gatcctcact gtgttctgtc 780tcagtgtgtt cgcactaatt ggactacaac
tgtttatggg caacttgaag cataaatgtt 840tccggaagga ccttgagcag aatgaaacat
tagaaagcat catgagtact gctgagagtg 900aagaagaatt gaaaagatat ttttattact
tggagggatc caaagatgct cttctttgcg 960gtttcagcac agattcaggg cagtgtcctg
aagggtacga gtgtgtgaca gctggcagaa 1020acccagatta tggctacaca agctttgaca
cgttcggctg ggccttcttg gccttgtttc 1080ggctaatgac tcaggactac tgggagaacc
tttatcaaca gacactgcgt gctgctggca 1140aaacctacat gattttcttt gtcgtggtga
tatttctggg atccttttac ctgataaact 1200tgatcctggc tgtggtagcc atggcgtacg
aggaacagaa ccaggccaac atcgaagaag 1260ctaaacagaa agagttagaa tttcagcaga
tgttagaccg actcaaaaaa gagcaggaag 1320aagccgaggc gatcgctgca gccgctgctg
agtacacgag tttagggcgg agcaggatta 1380tgggactctc tgagagctct tcagaaacct
ccaggctgag ctcaaagagt gccaaggaga 1440gaagaaaccg aagaaagaaa aaaaaacaga
agctgtccag tggcgaggaa aagggtgatg 1500atgagaagct gtccaagtca gggtcagagg
aaagcatccg aaagaaaagc ttccatctcg 1560gcgtggaagg gcaccaccgg gccagggaaa
agaggctgtc cacccccaac cagtcaccac 1620tcagcattcg tgggtccttg ttttctgcca
ggcgcagcag cagaacaagt ctcttcagtt 1680ttaaggggcg aggaagagat ctgggatctg
aaacggaatt tgctgatgat gagcatagca 1740tttttggaga caacgagagc agaaggggtt
cactatttgt accccataga ccccgggagc 1800ggcgcagcag taacatcagc caggccagta
ggtccccacc agtgctgccg gtgaacggga 1860agatgcacag tgcagtggac tgcaatggcg
tggtgtcgct tgttgatgga ccctcagccc 1920tcatgctccc caatggacag cttcttccag
aggtgataat agataaggca acttccgacg 1980acagcggcac aactaatcag atgcgtaaaa
aaaggctctc tagttcttac tttttgtctg 2040aggacatgct gaatgaccca catctcaggc
aaagggccat gagcagagca agcattctaa 2100ccaacacagt agaagaactt gaagaatcta
gacaaaaatg tccaccatgg tggtacagat 2160ttgctcacac atttttaatc tggaattgtt
ctccatattg gataaaattc aaaaagttca 2220tctattttat tgtaatggat ccttttgtag
atcttgcaat taccatttgc atagttttaa 2280acaccttgtt tatggctatg gagcaccatc
caatgacgga tgaattcaaa aatgtacttg 2340cagtcgggaa cctggtcttc acagggatct
tcgcagctga aatggtactg aagttaatag 2400ccatggatcc ctatgaatat ttccaagtag
ggtggaatat ttttgacagc ctgattgtga 2460cgttgagttt ggtggagctt ttcctagcag
atgtggaagg attatcagtc ctgcggtcct 2520ttagattgct gcgagtcttc aagttggcaa
aatcctggcc cacactgaat atgctcatta 2580agatcatcgg caactcggtg ggcgcactgg
gcaacctgac cctggtgctg gccatcatcg 2640tcttcatttt tgccgtggtc ggcatgcagc
tgtttggaaa gagctacaag gagtgtgttt 2700gcaagatcaa cgagaactgc aagctcccac
gctggcacat gaacgacttc ttccactcct 2760tcctgatcgt gttccgtgtg ctgtgtgggg
agtggataga gaccatgtgg gactgcatgg 2820aggttgcggg ccagaccatg tgccttattg
tttacatgat ggtcatggtg attgggaacc 2880ttgtggtcct gaacctgttt ctggctttat
tactgagttc ctttagttct gacaatctta 2940cagcaattga agaagacacc gacgcaaaca
acctccagat tgcagtggcc agaattaaaa 3000gagggatcaa ttatgtgaaa cagaccctgc
gtgaattcat tctaaagtca ttttccaaaa 3060agccaaaggg ctccaaggac acaaaacgaa
cagcggatcc caacaacaaa agagaaaact 3120atatctcaaa ccgtaccctt gcggagataa
gcaaagatca caatttcctc aaagaaaagg 3180ataagatcag tggttttagc agcagtctag
acaaaagctt tatggatgaa aacgattacc 3240agtcctttat tcataatccc agcctcacag
tgacagtgcc cattgcacct ggggagtctg 3300atttggagaa tatgaacaca gaagagctta
gcagtgactc agatagtgac tacagcaaag 3360agagacggaa ccgatcaagt tcttcagagt
gcagcacagt tgataaccct ctgccaggag 3420aagaggaggc agaagctgag cctatcaatg
cagatgagcc cgaagcctgt tttacagatg 3480gctgtgtgag gagatttcca tgctgccaag
ttaacataga ctccgggaaa gggaaagttt 3540ggtggaccat ccggaagacc tgctacagga
tagtggagca cagctggttt gaaagcttca 3600ttgttctcat gatcctgctc agcagtggag
ctctggcttt tgaggatata tatattgaaa 3660agaaaaagac cattaagatt atcctggagt
atgctgacaa gatattcacc tacatcttca 3720ttctggaaat gcttctaaaa tgggtggcat
acgggtataa aacatatttc actaatgcct 3780ggtgttggct ggacttctta attgttgatg
tgtctctagt tactttagta gccaacactc 3840ttggctactc agaccttggc cccattaaat
ctctacggac actgagggcc ctaagacccc 3900taagagcttt gtctagattt gaaggaatga
gggtagtggt caacgcactc ataggagcaa 3960tcccttccat catgaatgtg cttcttgtgt
gccttatatt ctggctaata tttagcatca 4020tgggagtcaa tctgtttgct ggcaagttct
atgagtgtgt taacaccaca gatggctcac 4080gattttctgt atctcaagtt gcaaaccgtt
ctgagtgttt tgccctgatg aatgttagtg 4140gaaatgtgcg atggaaaaac ctgaaagtaa
acttcgataa cgttggactt ggttacctgt 4200cgctgcttca agttgcaacg ttcaagggct
ggatggatat tatgtatgca gcagttgact 4260ctgttaatgt aaatgcacaa ccaatatatg
aatacaacct ctacatgtac atttattttg 4320tcatcttcat catctttggc tcattcttca
ctttgaactt gttcattggt gtcatcatcg 4380ataatttcaa ccaacagaag aagaagcttg
gaggtcaaga tatctttatg acagaagaac 4440agaagaaata ctataatgca atgaagaagc
tggggtccaa gaaaccacaa aaaccaattc 4500cgaggccagg gaacaaattc caaggatgca
tatttgactt agtgacaaac caagcttttg 4560atatcaccat catggttctt atctgcctca
atatggtaac catgatggta gaaaaagagg 4620ggcaaactga ctacatgagt tttgtgctat
actggatcaa cgtggtcttc atcatcctgt 4680tcactgggga gtgtgtgctg aagctgatct
ctctcaggca ttactacttc actgtgggat 4740ggaacatttt tgattttgtg gtagtgatcc
tctccattgt aggaatgttt ctcgctgaga 4800tgatagagaa gtatttcgtg tctcctaccc
tgttccgagt cattcgcctg gccaggattg 4860gacgaatcct acgcctgatc aaaggcgcca
aggggatccg cacgctgctc tttgctctga 4920tgatgtccct tcctgcgctg ttcaacatcg
gcctcctgct tttcctcgtc atgttcatct 4980acgccatctt tgggatgtcc aactttgcct
acgttaaaaa ggaagctgga attaatgaca 5040tgttcaactt tgagaccttc ggcaacagca
tgatctgcct gttccaaatc accacctctg 5100cgggctggga tggactgctg gcccccatcc
tcaacagtgc acctcctgac tgtgacccaa 5160aaaaggttca cccaggaagt tcagtggaag
gggactgtgg aaatccatct gtgggaattt 5220tctactttgt cagctacatc atcatatcct
tcctggttgt ggtgaacatg tacattgctg 5280tcatcctgga gaacttcagc gttgccacag
aagaaagtac tgagcccctg agtgaggacg 5340actttgagat gttctacgaa gtctgggaga
agttcgaccc tgacgccacc cagttcatag 5400agttctgcaa gctctctgac tttgcagcag
ccctggatcc tcccctcctc atcgcaaagc 5460caaacaaagt ccagctcatt gccatggacc
tgcccatggt gagtggagac cgcatccact 5520gcctggacat cttatttgct tttacaaagc
gggtcctggg cgagagcgga gagatggatt 5580cccttcgttc acagatggaa gaaaggttta
tgtcagccaa tccttctaaa gtgtcctatg 5640agcccatcac aaccacactg aagcgaaaac
aagaggatgt atctgcgact atcattcagc 5700gtgcttacag acggtaccgc cttaggcaaa
acgtcaagaa tatatcaagt atatatataa 5760aagatggaga cagagatgac gatttgccca
ataaagaaga tatagttttt gataatgtta 5820acgagaactc aagtccagaa aagacagatg
caacagcctc taccatctct ccaccttcct 5880atgacagtgt cacaaagcca gatcaagaga
aatatgaaac agacaaaacg gagaaggaag 5940acaaagagaa agacgaaagc aggaaataga
gcttcggttt tgatacactg tttacagcct 6000gcgaaggtga ctcactcgtg ttaataagac
tcttttacgg aggtctatgc caaactcttt 6060ttatcaaata ttctcaaagg cagcacagcc
actagctctg atccagtgaa acaagagaga 6120agcatttaca catggctact ttttgcgttg
gtcaatgatt ctttaagaat tgtgcatgta 6180actctacagg gaataatcat tattgcaatc
aagggtgact taatgatttt aaatatcaga 6240aaaccacata gaacattttc tcttttgcct
ccatttcttt ccctagattc taagtagatg 6300tgtacccatg tgaatataga aattcaggcg
cacatgctca cagtcacaaa cacaaacagg 6360attagctgtg atttggaatt cgatgtaaat
atttcacctg tgatttgcaa tgaaattcct 6420tgtaaaagaa atgcgaatta gtgatgaagg
ttttgtgaaa acatcttatc attagggagt 6480cagaatttct gtccataaag aattcagttt
atattttgag gtgctgaaac ttatcctaca 6540ttgcatcaaa atcaatttat aggtatctgt
aaaatgtcat gggactgaaa aacatatata 6600ggctacttgt ttaagaaatg gctttcattc
atatagatag gcattcacct tgatttatgg 6660acatctttgg cattttgtga tcacatgatt
cttccacaaa attgcttagc tggaacttca 6720ggcacacatc acagagaaca gctacccagt
cttatgcccc tctctgtttg tacaataatc 6780acagagcttg aaacattatt tgaactataa
atatcaggtt tctccacata gacatatgaa 6840tattgttaac agaaaaaaat tttatttaca
gcgttttatt tactaatatt tatccaatct 6900agtttgccca atgagacagc tcatgactca
catctgaaag ccagttgcca catttatctt 6960cttatgtaac tttggtttgt catactttat
gtctaagcaa attgaatgtc tcctttctaa 7020tgagatgtac cctgaaatgc agttaggtac
ttgatacttt agcgcttgtt tgagcagatg 7080actggagcac agtgtggact gcatctctta
aatacaatcc ttaatgtgtt tggcagcttc 7140tcaggttaca aggaacaccc ggcttttagt
gtctatctgt tcaccaggtg tttagtatga 7200atgaaacggc attcaaagag tgagtctcac
tggcttgctt tattactgat gccttcccta 7260tggagaatta atcctctgaa gccccattat
gtccccctgt aaataatgta gatgtcactt 7320ccttcttaat attctaatcc atactgtgaa
atcgattttg catttatcgg tcaaatagag 7380cattttgaga tagttggagt taccctgcca
aggattagaa atctacttca tgtttttaaa 7440gtacttgtta aaaatgaacg accctggcac
attctctcat aattttattc cagccatgtg 7500aaatctttct tctaaacact ttatccttgc
ggaggaaaaa aaaaatgagc tgatgagcca 7560tttaagcaca aaggggcttt atttagaaga
ttccaagggg gaactttgaa gtaaatatat 7620aaaacatact tcatcaattt gcctataaaa
ctaaaagagg aacacaggaa tattgataaa 7680ataagtcatt aaaacacatt ctttatttct
tgcccagttt aaaagaaaga actaaacatc 7740cctagagaga ggaagagaca tagagagaga
gtgactaata gaaagaggga gagaagaaac 7800aaggcaccaa ggacaaaaag agataattag
acagaacttg tccaggtttt cacactatgt 7860gctctgtcca gtaccgtaca caagaacctc
tttccaaata tttgtcctaa ggctctaaga 7920agttaagtac gaggctgaag gttgaataca
actgtcttta atcattaaca gtttggggag 7980ctacttttaa acgtctatgg aagatgccaa
gcagtggtaa gccagacaat acagagcact 8040gcatatctgt caagcagctg aaatatgttt
gggcaactta atggtgagcc acacaaaaca 8100tccatttgta acaattttaa tacattcaat
taagaaacca ggatttttat tattttgcac 8160ccataaaaat ataactatat tgttcatttt
tattgataga gtatgtgtga atcttattga 8220ttatctgtaa tttactatta atgtttttac
agtgactgtt ttttttgtgt gtgtaaactt 8280aatatatgtc agcaactggt tcctcaacac
aatttttttt agcattacaa aaaaatgaac 8340aggtataaag gttctctttt ttctacatca
tgttgaacat attttgttct gaattacata 8400gttttaaatg taatattaag ttttatattc
atatatgttt aacatcaaaa tcactactta 8460tgacattgtt atcaatttaa aaaatagtat
ttgacactag gatagcattt aattaaagct 8520aaaaagctta caccccattt catgttgatt
agtgtttgga ctaactctaa aatgtcatca 8580atggaagcta gtcactgaaa ttattttatc
tattgtcata gaatggtgac tacccaaaaa 8640atataagtta gcattaaata gaagaaagcg
tacgtgacca caaatccatg cacagggttg 8700tgtgaagaca ggagaacctc atttttctgt
tttgtctctt tccactgtgt aaaaagtcta 8760catctgtggg ctatttctaa attcaaattg
tcacaatttg caatcataaa tgtttagcat 8820actttgtaga attttgatag ttttgtaaaa
gagtgaaaaa caaatgcata tgtaaataaa 8880gcagcccata ctagcagatt cctcaaatgt
taatatgtaa ataaagcagc ccttactagc 8940agattcatca aatgttaata tgtaaataaa
gcagtcctta ttagcagatt tgtcatatgt 9000taaggggagt aatgataagg aggcaactaa
atcaggatgg tcagtaactg atctgggttt 9060agaactgtgt ttggagccat caatttttaa
atatatgttc tcactatgtt attagttgtc 9120tgaagaagca atcaagaatt gctcccagaa
aatgagtaag tagccatgaa tatatgaatg 9180ctgtttacag aacccataga cctatgaatg
ctcaaaatgt ttgggtttgt caaaaaatta 9240cattgtagtt atacttgata cttaaaaact
gttaatagag tctaaaataa aagtcgct 9298161984PRTMus musculus 16Met Ala
Met Leu Pro Pro Pro Gly Pro Gln Ser Phe Val His Phe Thr 1 5
10 15 Lys Gln Ser Leu Ala Leu Ile
Glu Gln Arg Ile Ser Glu Glu Lys Ala 20 25
30 Lys Gly His Lys Asp Glu Lys Lys Asp Asp Glu Glu
Glu Gly Pro Lys 35 40 45
Pro Ser Ser Asp Leu Glu Ala Gly Lys Gln Leu Pro Phe Ile Tyr Gly
50 55 60 Asp Ile Pro
Pro Gly Met Val Ser Glu Pro Leu Glu Asp Leu Asp Pro 65
70 75 80 Tyr Tyr Ala Asp Lys Lys Thr
Phe Ile Val Leu Asn Lys Gly Lys Ala 85
90 95 Ile Phe Arg Phe Asn Ala Thr Pro Ala Leu Tyr
Met Leu Ser Pro Phe 100 105
110 Ser Pro Leu Arg Arg Ile Ser Ile Lys Ile Leu Val His Ser Leu
Phe 115 120 125 Ser
Met Leu Ile Met Cys Thr Ile Leu Thr Asn Cys Ile Phe Met Thr 130
135 140 Met Ser Asn Pro Pro Asp
Trp Thr Lys Asn Val Glu Tyr Thr Phe Thr 145 150
155 160 Gly Ile Tyr Thr Phe Glu Ser Leu Ile Lys Ile
Leu Ala Arg Gly Phe 165 170
175 Cys Val Gly Glu Phe Thr Phe Leu Arg Asp Pro Trp Asn Trp Leu Asp
180 185 190 Phe Val
Val Ile Val Phe Ala Tyr Leu Thr Glu Phe Val Asn Leu Gly 195
200 205 Asn Val Ser Ala Leu Arg Thr
Phe Arg Val Leu Arg Ala Leu Lys Thr 210 215
220 Ile Ser Val Ile Pro Gly Leu Lys Thr Ile Val Gly
Ala Leu Ile Gln 225 230 235
240 Ser Val Lys Lys Leu Ser Asp Val Met Ile Leu Thr Val Phe Cys Leu
245 250 255 Ser Val Phe
Ala Leu Ile Gly Leu Gln Leu Phe Met Gly Asn Leu Lys 260
265 270 His Lys Cys Phe Arg Lys Asp Leu
Glu Gln Asn Glu Thr Leu Glu Ser 275 280
285 Ile Met Ser Thr Ala Glu Ser Glu Glu Glu Leu Lys Arg
Tyr Phe Tyr 290 295 300
Tyr Leu Glu Gly Ser Lys Asp Ala Leu Leu Cys Gly Phe Ser Thr Asp 305
310 315 320 Ser Gly Gln Cys
Pro Glu Gly Tyr Glu Cys Val Thr Ala Gly Arg Asn 325
330 335 Pro Asp Tyr Gly Tyr Thr Ser Phe Asp
Thr Phe Gly Trp Ala Phe Leu 340 345
350 Ala Leu Phe Arg Leu Met Thr Gln Asp Tyr Trp Glu Asn Leu
Tyr Gln 355 360 365
Gln Thr Leu Arg Ala Ala Gly Lys Thr Tyr Met Ile Phe Phe Val Val 370
375 380 Val Ile Phe Leu Gly
Ser Phe Tyr Leu Ile Asn Leu Ile Leu Ala Val 385 390
395 400 Val Ala Met Ala Tyr Glu Glu Gln Asn Gln
Ala Asn Ile Glu Glu Ala 405 410
415 Lys Gln Lys Glu Leu Glu Phe Gln Gln Met Leu Asp Arg Leu Lys
Lys 420 425 430 Glu
Gln Glu Glu Ala Glu Ala Ile Ala Ala Ala Ala Ala Glu Tyr Thr 435
440 445 Ser Leu Gly Arg Ser Arg
Ile Met Gly Leu Ser Glu Ser Ser Ser Glu 450 455
460 Thr Ser Arg Leu Ser Ser Lys Ser Ala Lys Glu
Arg Arg Asn Arg Arg 465 470 475
480 Lys Lys Lys Lys Gln Lys Leu Ser Ser Gly Glu Glu Lys Gly Asp Asp
485 490 495 Glu Lys
Leu Ser Lys Ser Gly Ser Glu Glu Ser Ile Arg Lys Lys Ser 500
505 510 Phe His Leu Gly Val Glu Gly
His His Arg Ala Arg Glu Lys Arg Leu 515 520
525 Ser Thr Pro Asn Gln Ser Pro Leu Ser Ile Arg Gly
Ser Leu Phe Ser 530 535 540
Ala Arg Arg Ser Ser Arg Thr Ser Leu Phe Ser Phe Lys Gly Arg Gly 545
550 555 560 Arg Asp Leu
Gly Ser Glu Thr Glu Phe Ala Asp Asp Glu His Ser Ile 565
570 575 Phe Gly Asp Asn Glu Ser Arg Arg
Gly Ser Leu Phe Val Pro His Arg 580 585
590 Pro Arg Glu Arg Arg Ser Ser Asn Ile Ser Gln Ala Ser
Arg Ser Pro 595 600 605
Pro Val Leu Pro Val Asn Gly Lys Met His Ser Ala Val Asp Cys Asn 610
615 620 Gly Val Val Ser
Leu Val Asp Gly Pro Ser Ala Leu Met Leu Pro Asn 625 630
635 640 Gly Gln Leu Leu Pro Glu Val Ile Ile
Asp Lys Ala Thr Ser Asp Asp 645 650
655 Ser Gly Thr Thr Asn Gln Met Arg Lys Lys Arg Leu Ser Ser
Ser Tyr 660 665 670
Phe Leu Ser Glu Asp Met Leu Asn Asp Pro His Leu Arg Gln Arg Ala
675 680 685 Met Ser Arg Ala
Ser Ile Leu Thr Asn Thr Val Glu Glu Leu Glu Glu 690
695 700 Ser Arg Gln Lys Cys Pro Pro Trp
Trp Tyr Arg Phe Ala His Thr Phe 705 710
715 720 Leu Ile Trp Asn Cys Ser Pro Tyr Trp Ile Lys Phe
Lys Lys Phe Ile 725 730
735 Tyr Phe Ile Val Met Asp Pro Phe Val Asp Leu Ala Ile Thr Ile Cys
740 745 750 Ile Val Leu
Asn Thr Leu Phe Met Ala Met Glu His His Pro Met Thr 755
760 765 Asp Glu Phe Lys Asn Val Leu Ala
Val Gly Asn Leu Val Phe Thr Gly 770 775
780 Ile Phe Ala Ala Glu Met Val Leu Lys Leu Ile Ala Met
Asp Pro Tyr 785 790 795
800 Glu Tyr Phe Gln Val Gly Trp Asn Ile Phe Asp Ser Leu Ile Val Thr
805 810 815 Leu Ser Leu Val
Glu Leu Phe Leu Ala Asp Val Glu Gly Leu Ser Val 820
825 830 Leu Arg Ser Phe Arg Leu Leu Arg Val
Phe Lys Leu Ala Lys Ser Trp 835 840
845 Pro Thr Leu Asn Met Leu Ile Lys Ile Ile Gly Asn Ser Val
Gly Ala 850 855 860
Leu Gly Asn Leu Thr Leu Val Leu Ala Ile Ile Val Phe Ile Phe Ala 865
870 875 880 Val Val Gly Met Gln
Leu Phe Gly Lys Ser Tyr Lys Glu Cys Val Cys 885
890 895 Lys Ile Asn Glu Asn Cys Lys Leu Pro Arg
Trp His Met Asn Asp Phe 900 905
910 Phe His Ser Phe Leu Ile Val Phe Arg Val Leu Cys Gly Glu Trp
Ile 915 920 925 Glu
Thr Met Trp Asp Cys Met Glu Val Ala Gly Gln Thr Met Cys Leu 930
935 940 Ile Val Tyr Met Met Val
Met Val Ile Gly Asn Leu Val Val Leu Asn 945 950
955 960 Leu Phe Leu Ala Leu Leu Leu Ser Ser Phe Ser
Ser Asp Asn Leu Thr 965 970
975 Ala Ile Glu Glu Asp Thr Asp Ala Asn Asn Leu Gln Ile Ala Val Ala
980 985 990 Arg Ile
Lys Arg Gly Ile Asn Tyr Val Lys Gln Thr Leu Arg Glu Phe 995
1000 1005 Ile Leu Lys Ser Phe
Ser Lys Lys Pro Lys Gly Ser Lys Asp Thr 1010 1015
1020 Lys Arg Thr Ala Asp Pro Asn Asn Lys Arg
Glu Asn Tyr Ile Ser 1025 1030 1035
Asn Arg Thr Leu Ala Glu Ile Ser Lys Asp His Asn Phe Leu Lys
1040 1045 1050 Glu Lys
Asp Lys Ile Ser Gly Phe Ser Ser Ser Leu Asp Lys Ser 1055
1060 1065 Phe Met Asp Glu Asn Asp Tyr
Gln Ser Phe Ile His Asn Pro Ser 1070 1075
1080 Leu Thr Val Thr Val Pro Ile Ala Pro Gly Glu Ser
Asp Leu Glu 1085 1090 1095
Asn Met Asn Thr Glu Glu Leu Ser Ser Asp Ser Asp Ser Asp Tyr 1100
1105 1110 Ser Lys Glu Arg Arg
Asn Arg Ser Ser Ser Ser Glu Cys Ser Thr 1115 1120
1125 Val Asp Asn Pro Leu Pro Gly Glu Glu Glu
Ala Glu Ala Glu Pro 1130 1135 1140
Ile Asn Ala Asp Glu Pro Glu Ala Cys Phe Thr Asp Gly Cys Val
1145 1150 1155 Arg Arg
Phe Pro Cys Cys Gln Val Asn Ile Asp Ser Gly Lys Gly 1160
1165 1170 Lys Val Trp Trp Thr Ile Arg
Lys Thr Cys Tyr Arg Ile Val Glu 1175 1180
1185 His Ser Trp Phe Glu Ser Phe Ile Val Leu Met Ile
Leu Leu Ser 1190 1195 1200
Ser Gly Ala Leu Ala Phe Glu Asp Ile Tyr Ile Glu Lys Lys Lys 1205
1210 1215 Thr Ile Lys Ile Ile
Leu Glu Tyr Ala Asp Lys Ile Phe Thr Tyr 1220 1225
1230 Ile Phe Ile Leu Glu Met Leu Leu Lys Trp
Val Ala Tyr Gly Tyr 1235 1240 1245
Lys Thr Tyr Phe Thr Asn Ala Trp Cys Trp Leu Asp Phe Leu Ile
1250 1255 1260 Val Asp
Val Ser Leu Val Thr Leu Val Ala Asn Thr Leu Gly Tyr 1265
1270 1275 Ser Asp Leu Gly Pro Ile Lys
Ser Leu Arg Thr Leu Arg Ala Leu 1280 1285
1290 Arg Pro Leu Arg Ala Leu Ser Arg Phe Glu Gly Met
Arg Val Val 1295 1300 1305
Val Asn Ala Leu Ile Gly Ala Ile Pro Ser Ile Met Asn Val Leu 1310
1315 1320 Leu Val Cys Leu Ile
Phe Trp Leu Ile Phe Ser Ile Met Gly Val 1325 1330
1335 Asn Leu Phe Ala Gly Lys Phe Tyr Glu Cys
Val Asn Thr Thr Asp 1340 1345 1350
Gly Ser Arg Phe Ser Val Ser Gln Val Ala Asn Arg Ser Glu Cys
1355 1360 1365 Phe Ala
Leu Met Asn Val Ser Gly Asn Val Arg Trp Lys Asn Leu 1370
1375 1380 Lys Val Asn Phe Asp Asn Val
Gly Leu Gly Tyr Leu Ser Leu Leu 1385 1390
1395 Gln Val Ala Thr Phe Lys Gly Trp Met Asp Ile Met
Tyr Ala Ala 1400 1405 1410
Val Asp Ser Val Asn Val Asn Ala Gln Pro Ile Tyr Glu Tyr Asn 1415
1420 1425 Leu Tyr Met Tyr Ile
Tyr Phe Val Ile Phe Ile Ile Phe Gly Ser 1430 1435
1440 Phe Phe Thr Leu Asn Leu Phe Ile Gly Val
Ile Ile Asp Asn Phe 1445 1450 1455
Asn Gln Gln Lys Lys Lys Leu Gly Gly Gln Asp Ile Phe Met Thr
1460 1465 1470 Glu Glu
Gln Lys Lys Tyr Tyr Asn Ala Met Lys Lys Leu Gly Ser 1475
1480 1485 Lys Lys Pro Gln Lys Pro Ile
Pro Arg Pro Gly Asn Lys Phe Gln 1490 1495
1500 Gly Cys Ile Phe Asp Leu Val Thr Asn Gln Ala Phe
Asp Ile Thr 1505 1510 1515
Ile Met Val Leu Ile Cys Leu Asn Met Val Thr Met Met Val Glu 1520
1525 1530 Lys Glu Gly Gln Thr
Asp Tyr Met Ser Phe Val Leu Tyr Trp Ile 1535 1540
1545 Asn Val Val Phe Ile Ile Leu Phe Thr Gly
Glu Cys Val Leu Lys 1550 1555 1560
Leu Ile Ser Leu Arg His Tyr Tyr Phe Thr Val Gly Trp Asn Ile
1565 1570 1575 Phe Asp
Phe Val Val Val Ile Leu Ser Ile Val Gly Met Phe Leu 1580
1585 1590 Ala Glu Met Ile Glu Lys Tyr
Phe Val Ser Pro Thr Leu Phe Arg 1595 1600
1605 Val Ile Arg Leu Ala Arg Ile Gly Arg Ile Leu Arg
Leu Ile Lys 1610 1615 1620
Gly Ala Lys Gly Ile Arg Thr Leu Leu Phe Ala Leu Met Met Ser 1625
1630 1635 Leu Pro Ala Leu Phe
Asn Ile Gly Leu Leu Leu Phe Leu Val Met 1640 1645
1650 Phe Ile Tyr Ala Ile Phe Gly Met Ser Asn
Phe Ala Tyr Val Lys 1655 1660 1665
Lys Glu Ala Gly Ile Asn Asp Met Phe Asn Phe Glu Thr Phe Gly
1670 1675 1680 Asn Ser
Met Ile Cys Leu Phe Gln Ile Thr Thr Ser Ala Gly Trp 1685
1690 1695 Asp Gly Leu Leu Ala Pro Ile
Leu Asn Ser Ala Pro Pro Asp Cys 1700 1705
1710 Asp Pro Lys Lys Val His Pro Gly Ser Ser Val Glu
Gly Asp Cys 1715 1720 1725
Gly Asn Pro Ser Val Gly Ile Phe Tyr Phe Val Ser Tyr Ile Ile 1730
1735 1740 Ile Ser Phe Leu Val
Val Val Asn Met Tyr Ile Ala Val Ile Leu 1745 1750
1755 Glu Asn Phe Ser Val Ala Thr Glu Glu Ser
Thr Glu Pro Leu Ser 1760 1765 1770
Glu Asp Asp Phe Glu Met Phe Tyr Glu Val Trp Glu Lys Phe Asp
1775 1780 1785 Pro Asp
Ala Thr Gln Phe Ile Glu Phe Cys Lys Leu Ser Asp Phe 1790
1795 1800 Ala Ala Ala Leu Asp Pro Pro
Leu Leu Ile Ala Lys Pro Asn Lys 1805 1810
1815 Val Gln Leu Ile Ala Met Asp Leu Pro Met Val Ser
Gly Asp Arg 1820 1825 1830
Ile His Cys Leu Asp Ile Leu Phe Ala Phe Thr Lys Arg Val Leu 1835
1840 1845 Gly Glu Ser Gly Glu
Met Asp Ser Leu Arg Ser Gln Met Glu Glu 1850 1855
1860 Arg Phe Met Ser Ala Asn Pro Ser Lys Val
Ser Tyr Glu Pro Ile 1865 1870 1875
Thr Thr Thr Leu Lys Arg Lys Gln Glu Asp Val Ser Ala Thr Ile
1880 1885 1890 Ile Gln
Arg Ala Tyr Arg Arg Tyr Arg Leu Arg Gln Asn Val Lys 1895
1900 1905 Asn Ile Ser Ser Ile Tyr Ile
Lys Asp Gly Asp Arg Asp Asp Asp 1910 1915
1920 Leu Pro Asn Lys Glu Asp Ile Val Phe Asp Asn Val
Asn Glu Asn 1925 1930 1935
Ser Ser Pro Glu Lys Thr Asp Ala Thr Ala Ser Thr Ile Ser Pro 1940
1945 1950 Pro Ser Tyr Asp Ser
Val Thr Lys Pro Asp Gln Glu Lys Tyr Glu 1955 1960
1965 Thr Asp Lys Thr Glu Lys Glu Asp Lys Glu
Lys Asp Glu Ser Arg 1970 1975 1980
Lys 179265DNAMus musculus 17tacaggatga gaagatggcg atgttgcctc
cgccaggacc tcagagcttc gttcacttca 60caaaacagtc ccttgccctc attgaacaac
gcatttctga agaaaaagcc aagggacaca 120aagatgaaaa gaaagatgat gaggaagaag
gtcccaagcc cagtagtgac ttggaagcag 180gtaaacagct acccttcatc tacggagaca
ttcccccggg aatggtgtca gagcccctgg 240aggacctgga cccatactat gcagacaaaa
aaacttttat agtattgaac aaagggaaag 300caatcttccg tttcaacgcc acccctgctt
tgtacatgct gtctcccttc agtcctctca 360gaagaatatc tattaagatt ttagtgcact
ccttattcag catgctaatc atgtgcacaa 420ttctgacaaa ctgcatattc atgaccatga
gcaaccctcc agattggacc aaaaacgtag 480agtacacttt tactgggata tatacttttg
aatcactcat aaaaatcctt gcaagaggct 540tttgcgtggg cgaattcacc ttcctccgtg
acccttggaa ctggctggac tttgttgtca 600ttgtttttgc atatgtgaca gagtttgtgg
acctgggcaa tgtctcagcg ttgagaacat 660tcagagttct ccgagcattg aaaacaatat
cagtcattcc aggactaaaa accatcgtgg 720gggccctgat ccaatcagtg aagaagctct
ctgacgtcat gatcctcact gtgttctgtc 780tcagtgtgtt cgcactaatt ggactacaac
tgtttatggg caacttgaag cataaatgtt 840tccggaagga ccttgagcag aatgaaacat
tagaaagcat catgagtact gctgagagtg 900aagaagaatt gaaaagatat ttttattact
tggagggatc caaagatgct cttctttgcg 960gtttcagcac agattcaggg cagtgtcctg
aagggtacga gtgtgtgaca gctggcagaa 1020acccagatta tggctacaca agctttgaca
cgttcggctg ggccttcttg gccttgtttc 1080ggctaatgac tcaggactac tgggagaacc
tttatcaaca gacactgcgt gctgctggca 1140aaacctacat gattttcttt gtcgtggtga
tatttctggg atccttttac ctgataaact 1200tgatcctggc tgtggtagcc atggcgtacg
aggaacagaa ccaggccaac atcgaagaag 1260ctaaacagaa agagttagaa tttcagcaga
tgttagaccg actcaaaaaa gagcaggaag 1320aagccgaggc gatcgctgca gccgctgctg
agtacacgag tttagggcgg agcaggatta 1380tgggactctc tgagagctct tcagaaacct
ccaggctgag ctcaaagagt gccaaggaga 1440gaagaaaccg aagaaagaaa aaaaaacaga
agctgtccag tggcgaggaa aagggtgatg 1500atgagaagct gtccaagtca gggtcagagg
aaagcatccg aaagaaaagc ttccatctcg 1560gcgtggaagg gcaccaccgg gccagggaaa
agaggctgtc cacccccaac cagtcaccac 1620tcagcattcg tgggtccttg ttttctgcca
ggcgcagcag cagaacaagt ctcttcagtt 1680ttaaggggcg aggaagagat ctgggatctg
aaacggaatt tgctgatgat gagcatagca 1740tttttggaga caacgagagc agaaggggtt
cactatttgt accccataga ccccgggagc 1800ggcgcagcag taacatcagc caggccagta
ggtccccacc agtgctgccg gtgaacggga 1860agatgcacag tgcagtggac tgcaatggcg
tggtgtcgct tgttgatgga ccctcagccc 1920tcatgctccc caatggacag cttcttccag
agggcacaac taatcagatg cgtaaaaaaa 1980ggctctctag ttcttacttt ttgtctgagg
acatgctgaa tgacccacat ctcaggcaaa 2040gggccatgag cagagcaagc attctaacca
acacagtaga agaacttgaa gaatctagac 2100aaaaatgtcc accatggtgg tacagatttg
ctcacacatt tttaatctgg aattgttctc 2160catattggat aaaattcaaa aagttcatct
attttattgt aatggatcct tttgtagatc 2220ttgcaattac catttgcata gttttaaaca
ccttgtttat ggctatggag caccatccaa 2280tgacggatga attcaaaaat gtacttgcag
tcgggaacct ggtcttcaca gggatcttcg 2340cagctgaaat ggtactgaag ttaatagcca
tggatcccta tgaatatttc caagtagggt 2400ggaatatttt tgacagcctg attgtgacgt
tgagtttggt ggagcttttc ctagcagatg 2460tggaaggatt atcagtcctg cggtccttta
gattgctgcg agtcttcaag ttggcaaaat 2520cctggcccac actgaatatg ctcattaaga
tcatcggcaa ctcggtgggc gcactgggca 2580acctgaccct ggtgctggcc atcatcgtct
tcatttttgc cgtggtcggc atgcagctgt 2640ttggaaagag ctacaaggag tgtgtttgca
agatcaacga gaactgcaag ctcccacgct 2700ggcacatgaa cgacttcttc cactccttcc
tgatcgtgtt ccgtgtgctg tgtggggagt 2760ggatagagac catgtgggac tgcatggagg
ttgcgggcca gaccatgtgc cttattgttt 2820acatgatggt catggtgatt gggaaccttg
tggtcctgaa cctgtttctg gctttattac 2880tgagttcctt tagttctgac aatcttacag
caattgaaga agacaccgac gcaaacaacc 2940tccagattgc agtggccaga attaaaagag
ggatcaatta tgtgaaacag accctgcgtg 3000aattcattct aaagtcattt tccaaaaagc
caaagggctc caaggacaca aaacgaacag 3060cggatcccaa caacaaaaga gaaaactata
tctcaaaccg tacccttgcg gagataagca 3120aagatcacaa tttcctcaaa gaaaaggata
agatcagtgg ttttagcagc agtctagaca 3180aaagctttat ggatgaaaac gattaccagt
cctttattca taatcccagc ctcacagtga 3240cagtgcccat tgcacctggg gagtctgatt
tggagaatat gaacacagaa gagcttagca 3300gtgactcaga tagtgactac agcaaagaga
gacggaaccg atcaagttct tcagagtgca 3360gcacagttga taaccctctg ccaggagaag
aggaggcaga agctgagcct atcaatgcag 3420atgagcccga agcctgtttt acagatggct
gtgtgaggag atttccatgc tgccaagtta 3480acatagactc cgggaaaggg aaagtttggt
ggaccatccg gaagacctgc tacaggatag 3540tggagcacag ctggtttgaa agcttcattg
ttctcatgat cctgctcagc agtggagctc 3600tggcttttga ggatatatat attgaaaaga
aaaagaccat taagattatc ctggagtatg 3660ctgacaagat attcacctac atcttcattc
tggaaatgct tctaaaatgg gtggcatacg 3720ggtataaaac atatttcact aatgcctggt
gttggctgga cttcttaatt gttgatgtgt 3780ctctagttac tttagtagcc aacactcttg
gctactcaga ccttggcccc attaaatctc 3840tacggacact gagggcccta agacccctaa
gagctttgtc tagatttgaa ggaatgaggg 3900tagtggtcaa cgcactcata ggagcaatcc
cttccatcat gaatgtgctt cttgtgtgcc 3960ttatattctg gctaatattt agcatcatgg
gagtcaatct gtttgctggc aagttctatg 4020agtgtgttaa caccacagat ggctcacgat
tttctgtatc tcaagttgca aaccgttctg 4080agtgttttgc cctgatgaat gttagtggaa
atgtgcgatg gaaaaacctg aaagtaaact 4140tcgataacgt tggacttggt tacctgtcgc
tgcttcaagt tgcaacgttc aagggctgga 4200tggatattat gtatgcagca gttgactctg
ttaatgtaaa tgcacaacca atatatgaat 4260acaacctcta catgtacatt tattttgtca
tcttcatcat ctttggctca ttcttcactt 4320tgaacttgtt cattggtgtc atcatcgata
atttcaacca acagaagaag aagcttggag 4380gtcaagatat ctttatgaca gaagaacaga
agaaatacta taatgcaatg aagaagctgg 4440ggtccaagaa accacaaaaa ccaattccga
ggccagggaa caaattccaa ggatgcatat 4500ttgacttagt gacaaaccaa gcttttgata
tcaccatcat ggttcttatc tgcctcaata 4560tggtaaccat gatggtagaa aaagaggggc
aaactgacta catgagtttt gtgctatact 4620ggatcaacgt ggtcttcatc atcctgttca
ctggggagtg tgtgctgaag ctgatctctc 4680tcaggcatta ctacttcact gtgggatgga
acatttttga ttttgtggta gtgatcctct 4740ccattgtagg aatgtttctc gctgagatga
tagagaagta tttcgtgtct cctaccctgt 4800tccgagtcat tcgcctggcc aggattggac
gaatcctacg cctgatcaaa ggcgccaagg 4860ggatccgcac gctgctcttt gctctgatga
tgtcccttcc tgcgctgttc aacatcggcc 4920tcctgctttt cctcgtcatg ttcatctacg
ccatctttgg gatgtccaac tttgcctacg 4980ttaaaaagga agctggaatt aatgacatgt
tcaactttga gaccttcggc aacagcatga 5040tctgcctgtt ccaaatcacc acctctgcgg
gctgggatgg actgctggcc cccatcctca 5100acagtgcacc tcctgactgt gacccaaaaa
aggttcaccc aggaagttca gtggaagggg 5160actgtggaaa tccatctgtg ggaattttct
actttgtcag ctacatcatc atatccttcc 5220tggttgtggt gaacatgtac attgctgtca
tcctggagaa cttcagcgtt gccacagaag 5280aaagtactga gcccctgagt gaggacgact
ttgagatgtt ctacgaagtc tgggagaagt 5340tcgaccctga cgccacccag ttcatagagt
tctgcaagct ctctgacttt gcagcagccc 5400tggatcctcc cctcctcatc gcaaagccaa
acaaagtcca gctcattgcc atggacctgc 5460ccatggtgag tggagaccgc atccactgcc
tggacatctt atttgctttt acaaagcggg 5520tcctgggcga gagcggagag atggattccc
ttcgttcaca gatggaagaa aggtttatgt 5580cagccaatcc ttctaaagtg tcctatgagc
ccatcacaac cacactgaag cgaaaacaag 5640aggatgtatc tgcgactatc attcagcgtg
cttacagacg gtaccgcctt aggcaaaacg 5700tcaagaatat atcaagtata tatataaaag
atggagacag agatgacgat ttgcccaata 5760aagaagatat agtttttgat aatgttaacg
agaactcaag tccagaaaag acagatgcaa 5820cagcctctac catctctcca ccttcctatg
acagtgtcac aaagccagat caagagaaat 5880atgaaacaga caaaacggag aaggaagaca
aagagaaaga cgaaagcagg aaatagagct 5940tcggttttga tacactgttt acagcctgcg
aaggtgactc actcgtgtta ataagactct 6000tttacggagg tctatgccaa actcttttta
tcaaatattc tcaaaggcag cacagccact 6060agctctgatc cagtgaaaca agagagaagc
atttacacat ggctactttt tgcgttggtc 6120aatgattctt taagaattgt gcatgtaact
ctacagggaa taatcattat tgcaatcaag 6180ggtgacttaa tgattttaaa tatcagaaaa
ccacatagaa cattttctct tttgcctcca 6240tttctttccc tagattctaa gtagatgtgt
acccatgtga atatagaaat tcaggcgcac 6300atgctcacag tcacaaacac aaacaggatt
agctgtgatt tggaattcga tgtaaatatt 6360tcacctgtga tttgcaatga aattccttgt
aaaagaaatg cgaattagtg atgaaggttt 6420tgtgaaaaca tcttatcatt agggagtcag
aatttctgtc cataaagaat tcagtttata 6480ttttgaggtg ctgaaactta tcctacattg
catcaaaatc aatttatagg tatctgtaaa 6540atgtcatggg actgaaaaac atatataggc
tacttgttta agaaatggct ttcattcata 6600tagataggca ttcaccttga tttatggaca
tctttggcat tttgtgatca catgattctt 6660ccacaaaatt gcttagctgg aacttcaggc
acacatcaca gagaacagct acccagtctt 6720atgcccctct ctgtttgtac aataatcaca
gagcttgaaa cattatttga actataaata 6780tcaggtttct ccacatagac atatgaatat
tgttaacaga aaaaaatttt atttacagcg 6840ttttatttac taatatttat ccaatctagt
ttgcccaatg agacagctca tgactcacat 6900ctgaaagcca gttgccacat ttatcttctt
atgtaacttt ggtttgtcat actttatgtc 6960taagcaaatt gaatgtctcc tttctaatga
gatgtaccct gaaatgcagt taggtacttg 7020atactttagc gcttgtttga gcagatgact
ggagcacagt gtggactgca tctcttaaat 7080acaatcctta atgtgtttgg cagcttctca
ggttacaagg aacacccggc ttttagtgtc 7140tatctgttca ccaggtgttt agtatgaatg
aaacggcatt caaagagtga gtctcactgg 7200cttgctttat tactgatgcc ttccctatgg
agaattaatc ctctgaagcc ccattatgtc 7260cccctgtaaa taatgtagat gtcacttcct
tcttaatatt ctaatccata ctgtgaaatc 7320gattttgcat ttatcggtca aatagagcat
tttgagatag ttggagttac cctgccaagg 7380attagaaatc tacttcatgt ttttaaagta
cttgttaaaa atgaacgacc ctggcacatt 7440ctctcataat tttattccag ccatgtgaaa
tctttcttct aaacacttta tccttgcgga 7500ggaaaaaaaa aatgagctga tgagccattt
aagcacaaag gggctttatt tagaagattc 7560caagggggaa ctttgaagta aatatataaa
acatacttca tcaatttgcc tataaaacta 7620aaagaggaac acaggaatat tgataaaata
agtcattaaa acacattctt tatttcttgc 7680ccagtttaaa agaaagaact aaacatccct
agagagagga agagacatag agagagagtg 7740actaatagaa agagggagag aagaaacaag
gcaccaagga caaaaagaga taattagaca 7800gaacttgtcc aggttttcac actatgtgct
ctgtccagta ccgtacacaa gaacctcttt 7860ccaaatattt gtcctaaggc tctaagaagt
taagtacgag gctgaaggtt gaatacaact 7920gtctttaatc attaacagtt tggggagcta
cttttaaacg tctatggaag atgccaagca 7980gtggtaagcc agacaataca gagcactgca
tatctgtcaa gcagctgaaa tatgtttggg 8040caacttaatg gtgagccaca caaaacatcc
atttgtaaca attttaatac attcaattaa 8100gaaaccagga tttttattat tttgcaccca
taaaaatata actatattgt tcatttttat 8160tgatagagta tgtgtgaatc ttattgatta
tctgtaattt actattaatg tttttacagt 8220gactgttttt tttgtgtgtg taaacttaat
atatgtcagc aactggttcc tcaacacaat 8280tttttttagc attacaaaaa aatgaacagg
tataaaggtt ctcttttttc tacatcatgt 8340tgaacatatt ttgttctgaa ttacatagtt
ttaaatgtaa tattaagttt tatattcata 8400tatgtttaac atcaaaatca ctacttatga
cattgttatc aatttaaaaa atagtatttg 8460acactaggat agcatttaat taaagctaaa
aagcttacac cccatttcat gttgattagt 8520gtttggacta actctaaaat gtcatcaatg
gaagctagtc actgaaatta ttttatctat 8580tgtcatagaa tggtgactac ccaaaaaata
taagttagca ttaaatagaa gaaagcgtac 8640gtgaccacaa atccatgcac agggttgtgt
gaagacagga gaacctcatt tttctgtttt 8700gtctctttcc actgtgtaaa aagtctacat
ctgtgggcta tttctaaatt caaattgtca 8760caatttgcaa tcataaatgt ttagcatact
ttgtagaatt ttgatagttt tgtaaaagag 8820tgaaaaacaa atgcatatgt aaataaagca
gcccatacta gcagattcct caaatgttaa 8880tatgtaaata aagcagccct tactagcaga
ttcatcaaat gttaatatgt aaataaagca 8940gtccttatta gcagatttgt catatgttaa
ggggagtaat gataaggagg caactaaatc 9000aggatggtca gtaactgatc tgggtttaga
actgtgtttg gagccatcaa tttttaaata 9060tatgttctca ctatgttatt agttgtctga
agaagcaatc aagaattgct cccagaaaat 9120gagtaagtag ccatgaatat atgaatgctg
tttacagaac ccatagacct atgaatgctc 9180aaaatgtttg ggtttgtcaa aaaattacat
tgtagttata cttgatactt aaaaactgtt 9240aatagagtct aaaataaaag tcgct
9265181973PRTMus musculus 18Met Ala Met
Leu Pro Pro Pro Gly Pro Gln Ser Phe Val His Phe Thr 1 5
10 15 Lys Gln Ser Leu Ala Leu Ile Glu
Gln Arg Ile Ser Glu Glu Lys Ala 20 25
30 Lys Gly His Lys Asp Glu Lys Lys Asp Asp Glu Glu Glu
Gly Pro Lys 35 40 45
Pro Ser Ser Asp Leu Glu Ala Gly Lys Gln Leu Pro Phe Ile Tyr Gly 50
55 60 Asp Ile Pro Pro
Gly Met Val Ser Glu Pro Leu Glu Asp Leu Asp Pro 65 70
75 80 Tyr Tyr Ala Asp Lys Lys Thr Phe Ile
Val Leu Asn Lys Gly Lys Ala 85 90
95 Ile Phe Arg Phe Asn Ala Thr Pro Ala Leu Tyr Met Leu Ser
Pro Phe 100 105 110
Ser Pro Leu Arg Arg Ile Ser Ile Lys Ile Leu Val His Ser Leu Phe
115 120 125 Ser Met Leu Ile
Met Cys Thr Ile Leu Thr Asn Cys Ile Phe Met Thr 130
135 140 Met Ser Asn Pro Pro Asp Trp Thr
Lys Asn Val Glu Tyr Thr Phe Thr 145 150
155 160 Gly Ile Tyr Thr Phe Glu Ser Leu Ile Lys Ile Leu
Ala Arg Gly Phe 165 170
175 Cys Val Gly Glu Phe Thr Phe Leu Arg Asp Pro Trp Asn Trp Leu Asp
180 185 190 Phe Val Val
Ile Val Phe Ala Tyr Val Thr Glu Phe Val Asp Leu Gly 195
200 205 Asn Val Ser Ala Leu Arg Thr Phe
Arg Val Leu Arg Ala Leu Lys Thr 210 215
220 Ile Ser Val Ile Pro Gly Leu Lys Thr Ile Val Gly Ala
Leu Ile Gln 225 230 235
240 Ser Val Lys Lys Leu Ser Asp Val Met Ile Leu Thr Val Phe Cys Leu
245 250 255 Ser Val Phe Ala
Leu Ile Gly Leu Gln Leu Phe Met Gly Asn Leu Lys 260
265 270 His Lys Cys Phe Arg Lys Asp Leu Glu
Gln Asn Glu Thr Leu Glu Ser 275 280
285 Ile Met Ser Thr Ala Glu Ser Glu Glu Glu Leu Lys Arg Tyr
Phe Tyr 290 295 300
Tyr Leu Glu Gly Ser Lys Asp Ala Leu Leu Cys Gly Phe Ser Thr Asp 305
310 315 320 Ser Gly Gln Cys Pro
Glu Gly Tyr Glu Cys Val Thr Ala Gly Arg Asn 325
330 335 Pro Asp Tyr Gly Tyr Thr Ser Phe Asp Thr
Phe Gly Trp Ala Phe Leu 340 345
350 Ala Leu Phe Arg Leu Met Thr Gln Asp Tyr Trp Glu Asn Leu Tyr
Gln 355 360 365 Gln
Thr Leu Arg Ala Ala Gly Lys Thr Tyr Met Ile Phe Phe Val Val 370
375 380 Val Ile Phe Leu Gly Ser
Phe Tyr Leu Ile Asn Leu Ile Leu Ala Val 385 390
395 400 Val Ala Met Ala Tyr Glu Glu Gln Asn Gln Ala
Asn Ile Glu Glu Ala 405 410
415 Lys Gln Lys Glu Leu Glu Phe Gln Gln Met Leu Asp Arg Leu Lys Lys
420 425 430 Glu Gln
Glu Glu Ala Glu Ala Ile Ala Ala Ala Ala Ala Glu Tyr Thr 435
440 445 Ser Leu Gly Arg Ser Arg Ile
Met Gly Leu Ser Glu Ser Ser Ser Glu 450 455
460 Thr Ser Arg Leu Ser Ser Lys Ser Ala Lys Glu Arg
Arg Asn Arg Arg 465 470 475
480 Lys Lys Lys Lys Gln Lys Leu Ser Ser Gly Glu Glu Lys Gly Asp Asp
485 490 495 Glu Lys Leu
Ser Lys Ser Gly Ser Glu Glu Ser Ile Arg Lys Lys Ser 500
505 510 Phe His Leu Gly Val Glu Gly His
His Arg Ala Arg Glu Lys Arg Leu 515 520
525 Ser Thr Pro Asn Gln Ser Pro Leu Ser Ile Arg Gly Ser
Leu Phe Ser 530 535 540
Ala Arg Arg Ser Ser Arg Thr Ser Leu Phe Ser Phe Lys Gly Arg Gly 545
550 555 560 Arg Asp Leu Gly
Ser Glu Thr Glu Phe Ala Asp Asp Glu His Ser Ile 565
570 575 Phe Gly Asp Asn Glu Ser Arg Arg Gly
Ser Leu Phe Val Pro His Arg 580 585
590 Pro Arg Glu Arg Arg Ser Ser Asn Ile Ser Gln Ala Ser Arg
Ser Pro 595 600 605
Pro Val Leu Pro Val Asn Gly Lys Met His Ser Ala Val Asp Cys Asn 610
615 620 Gly Val Val Ser Leu
Val Asp Gly Pro Ser Ala Leu Met Leu Pro Asn 625 630
635 640 Gly Gln Leu Leu Pro Glu Gly Thr Thr Asn
Gln Met Arg Lys Lys Arg 645 650
655 Leu Ser Ser Ser Tyr Phe Leu Ser Glu Asp Met Leu Asn Asp Pro
His 660 665 670 Leu
Arg Gln Arg Ala Met Ser Arg Ala Ser Ile Leu Thr Asn Thr Val 675
680 685 Glu Glu Leu Glu Glu Ser
Arg Gln Lys Cys Pro Pro Trp Trp Tyr Arg 690 695
700 Phe Ala His Thr Phe Leu Ile Trp Asn Cys Ser
Pro Tyr Trp Ile Lys 705 710 715
720 Phe Lys Lys Phe Ile Tyr Phe Ile Val Met Asp Pro Phe Val Asp Leu
725 730 735 Ala Ile
Thr Ile Cys Ile Val Leu Asn Thr Leu Phe Met Ala Met Glu 740
745 750 His His Pro Met Thr Asp Glu
Phe Lys Asn Val Leu Ala Val Gly Asn 755 760
765 Leu Val Phe Thr Gly Ile Phe Ala Ala Glu Met Val
Leu Lys Leu Ile 770 775 780
Ala Met Asp Pro Tyr Glu Tyr Phe Gln Val Gly Trp Asn Ile Phe Asp 785
790 795 800 Ser Leu Ile
Val Thr Leu Ser Leu Val Glu Leu Phe Leu Ala Asp Val 805
810 815 Glu Gly Leu Ser Val Leu Arg Ser
Phe Arg Leu Leu Arg Val Phe Lys 820 825
830 Leu Ala Lys Ser Trp Pro Thr Leu Asn Met Leu Ile Lys
Ile Ile Gly 835 840 845
Asn Ser Val Gly Ala Leu Gly Asn Leu Thr Leu Val Leu Ala Ile Ile 850
855 860 Val Phe Ile Phe
Ala Val Val Gly Met Gln Leu Phe Gly Lys Ser Tyr 865 870
875 880 Lys Glu Cys Val Cys Lys Ile Asn Glu
Asn Cys Lys Leu Pro Arg Trp 885 890
895 His Met Asn Asp Phe Phe His Ser Phe Leu Ile Val Phe Arg
Val Leu 900 905 910
Cys Gly Glu Trp Ile Glu Thr Met Trp Asp Cys Met Glu Val Ala Gly
915 920 925 Gln Thr Met Cys
Leu Ile Val Tyr Met Met Val Met Val Ile Gly Asn 930
935 940 Leu Val Val Leu Asn Leu Phe Leu
Ala Leu Leu Leu Ser Ser Phe Ser 945 950
955 960 Ser Asp Asn Leu Thr Ala Ile Glu Glu Asp Thr Asp
Ala Asn Asn Leu 965 970
975 Gln Ile Ala Val Ala Arg Ile Lys Arg Gly Ile Asn Tyr Val Lys Gln
980 985 990 Thr Leu Arg
Glu Phe Ile Leu Lys Ser Phe Ser Lys Lys Pro Lys Gly 995
1000 1005 Ser Lys Asp Thr Lys Arg
Thr Ala Asp Pro Asn Asn Lys Arg Glu 1010 1015
1020 Asn Tyr Ile Ser Asn Arg Thr Leu Ala Glu Ile
Ser Lys Asp His 1025 1030 1035
Asn Phe Leu Lys Glu Lys Asp Lys Ile Ser Gly Phe Ser Ser Ser
1040 1045 1050 Leu Asp Lys
Ser Phe Met Asp Glu Asn Asp Tyr Gln Ser Phe Ile 1055
1060 1065 His Asn Pro Ser Leu Thr Val Thr
Val Pro Ile Ala Pro Gly Glu 1070 1075
1080 Ser Asp Leu Glu Asn Met Asn Thr Glu Glu Leu Ser Ser
Asp Ser 1085 1090 1095
Asp Ser Asp Tyr Ser Lys Glu Arg Arg Asn Arg Ser Ser Ser Ser 1100
1105 1110 Glu Cys Ser Thr Val
Asp Asn Pro Leu Pro Gly Glu Glu Glu Ala 1115 1120
1125 Glu Ala Glu Pro Ile Asn Ala Asp Glu Pro
Glu Ala Cys Phe Thr 1130 1135 1140
Asp Gly Cys Val Arg Arg Phe Pro Cys Cys Gln Val Asn Ile Asp
1145 1150 1155 Ser Gly
Lys Gly Lys Val Trp Trp Thr Ile Arg Lys Thr Cys Tyr 1160
1165 1170 Arg Ile Val Glu His Ser Trp
Phe Glu Ser Phe Ile Val Leu Met 1175 1180
1185 Ile Leu Leu Ser Ser Gly Ala Leu Ala Phe Glu Asp
Ile Tyr Ile 1190 1195 1200
Glu Lys Lys Lys Thr Ile Lys Ile Ile Leu Glu Tyr Ala Asp Lys 1205
1210 1215 Ile Phe Thr Tyr Ile
Phe Ile Leu Glu Met Leu Leu Lys Trp Val 1220 1225
1230 Ala Tyr Gly Tyr Lys Thr Tyr Phe Thr Asn
Ala Trp Cys Trp Leu 1235 1240 1245
Asp Phe Leu Ile Val Asp Val Ser Leu Val Thr Leu Val Ala Asn
1250 1255 1260 Thr Leu
Gly Tyr Ser Asp Leu Gly Pro Ile Lys Ser Leu Arg Thr 1265
1270 1275 Leu Arg Ala Leu Arg Pro Leu
Arg Ala Leu Ser Arg Phe Glu Gly 1280 1285
1290 Met Arg Val Val Val Asn Ala Leu Ile Gly Ala Ile
Pro Ser Ile 1295 1300 1305
Met Asn Val Leu Leu Val Cys Leu Ile Phe Trp Leu Ile Phe Ser 1310
1315 1320 Ile Met Gly Val Asn
Leu Phe Ala Gly Lys Phe Tyr Glu Cys Val 1325 1330
1335 Asn Thr Thr Asp Gly Ser Arg Phe Ser Val
Ser Gln Val Ala Asn 1340 1345 1350
Arg Ser Glu Cys Phe Ala Leu Met Asn Val Ser Gly Asn Val Arg
1355 1360 1365 Trp Lys
Asn Leu Lys Val Asn Phe Asp Asn Val Gly Leu Gly Tyr 1370
1375 1380 Leu Ser Leu Leu Gln Val Ala
Thr Phe Lys Gly Trp Met Asp Ile 1385 1390
1395 Met Tyr Ala Ala Val Asp Ser Val Asn Val Asn Ala
Gln Pro Ile 1400 1405 1410
Tyr Glu Tyr Asn Leu Tyr Met Tyr Ile Tyr Phe Val Ile Phe Ile 1415
1420 1425 Ile Phe Gly Ser Phe
Phe Thr Leu Asn Leu Phe Ile Gly Val Ile 1430 1435
1440 Ile Asp Asn Phe Asn Gln Gln Lys Lys Lys
Leu Gly Gly Gln Asp 1445 1450 1455
Ile Phe Met Thr Glu Glu Gln Lys Lys Tyr Tyr Asn Ala Met Lys
1460 1465 1470 Lys Leu
Gly Ser Lys Lys Pro Gln Lys Pro Ile Pro Arg Pro Gly 1475
1480 1485 Asn Lys Phe Gln Gly Cys Ile
Phe Asp Leu Val Thr Asn Gln Ala 1490 1495
1500 Phe Asp Ile Thr Ile Met Val Leu Ile Cys Leu Asn
Met Val Thr 1505 1510 1515
Met Met Val Glu Lys Glu Gly Gln Thr Asp Tyr Met Ser Phe Val 1520
1525 1530 Leu Tyr Trp Ile Asn
Val Val Phe Ile Ile Leu Phe Thr Gly Glu 1535 1540
1545 Cys Val Leu Lys Leu Ile Ser Leu Arg His
Tyr Tyr Phe Thr Val 1550 1555 1560
Gly Trp Asn Ile Phe Asp Phe Val Val Val Ile Leu Ser Ile Val
1565 1570 1575 Gly Met
Phe Leu Ala Glu Met Ile Glu Lys Tyr Phe Val Ser Pro 1580
1585 1590 Thr Leu Phe Arg Val Ile Arg
Leu Ala Arg Ile Gly Arg Ile Leu 1595 1600
1605 Arg Leu Ile Lys Gly Ala Lys Gly Ile Arg Thr Leu
Leu Phe Ala 1610 1615 1620
Leu Met Met Ser Leu Pro Ala Leu Phe Asn Ile Gly Leu Leu Leu 1625
1630 1635 Phe Leu Val Met Phe
Ile Tyr Ala Ile Phe Gly Met Ser Asn Phe 1640 1645
1650 Ala Tyr Val Lys Lys Glu Ala Gly Ile Asn
Asp Met Phe Asn Phe 1655 1660 1665
Glu Thr Phe Gly Asn Ser Met Ile Cys Leu Phe Gln Ile Thr Thr
1670 1675 1680 Ser Ala
Gly Trp Asp Gly Leu Leu Ala Pro Ile Leu Asn Ser Ala 1685
1690 1695 Pro Pro Asp Cys Asp Pro Lys
Lys Val His Pro Gly Ser Ser Val 1700 1705
1710 Glu Gly Asp Cys Gly Asn Pro Ser Val Gly Ile Phe
Tyr Phe Val 1715 1720 1725
Ser Tyr Ile Ile Ile Ser Phe Leu Val Val Val Asn Met Tyr Ile 1730
1735 1740 Ala Val Ile Leu Glu
Asn Phe Ser Val Ala Thr Glu Glu Ser Thr 1745 1750
1755 Glu Pro Leu Ser Glu Asp Asp Phe Glu Met
Phe Tyr Glu Val Trp 1760 1765 1770
Glu Lys Phe Asp Pro Asp Ala Thr Gln Phe Ile Glu Phe Cys Lys
1775 1780 1785 Leu Ser
Asp Phe Ala Ala Ala Leu Asp Pro Pro Leu Leu Ile Ala 1790
1795 1800 Lys Pro Asn Lys Val Gln Leu
Ile Ala Met Asp Leu Pro Met Val 1805 1810
1815 Ser Gly Asp Arg Ile His Cys Leu Asp Ile Leu Phe
Ala Phe Thr 1820 1825 1830
Lys Arg Val Leu Gly Glu Ser Gly Glu Met Asp Ser Leu Arg Ser 1835
1840 1845 Gln Met Glu Glu Arg
Phe Met Ser Ala Asn Pro Ser Lys Val Ser 1850 1855
1860 Tyr Glu Pro Ile Thr Thr Thr Leu Lys Arg
Lys Gln Glu Asp Val 1865 1870 1875
Ser Ala Thr Ile Ile Gln Arg Ala Tyr Arg Arg Tyr Arg Leu Arg
1880 1885 1890 Gln Asn
Val Lys Asn Ile Ser Ser Ile Tyr Ile Lys Asp Gly Asp 1895
1900 1905 Arg Asp Asp Asp Leu Pro Asn
Lys Glu Asp Ile Val Phe Asp Asn 1910 1915
1920 Val Asn Glu Asn Ser Ser Pro Glu Lys Thr Asp Ala
Thr Ala Ser 1925 1930 1935
Thr Ile Ser Pro Pro Ser Tyr Asp Ser Val Thr Lys Pro Asp Gln 1940
1945 1950 Glu Lys Tyr Glu Thr
Asp Lys Thr Glu Lys Glu Asp Lys Glu Lys 1955 1960
1965 Asp Glu Ser Arg Lys 1970
199265DNAMus musculus 19tacaggatga gaagatggcg atgttgcctc cgccaggacc
tcagagcttc gttcacttca 60caaaacagtc ccttgccctc attgaacaac gcatttctga
agaaaaagcc aagggacaca 120aagatgaaaa gaaagatgat gaggaagaag gtcccaagcc
cagtagtgac ttggaagcag 180gtaaacagct acccttcatc tacggagaca ttcccccggg
aatggtgtca gagcccctgg 240aggacctgga cccatactat gcagacaaaa aaacttttat
agtattgaac aaagggaaag 300caatcttccg tttcaacgcc acccctgctt tgtacatgct
gtctcccttc agtcctctca 360gaagaatatc tattaagatt ttagtgcact ccttattcag
catgctaatc atgtgcacaa 420ttctgacaaa ctgcatattc atgaccatga gcaaccctcc
agattggacc aaaaacgtag 480agtacacttt tactgggata tatacttttg aatcactcat
aaaaatcctt gcaagaggct 540tttgcgtggg cgaattcacc ttcctccgtg acccttggaa
ctggctggac tttgttgtca 600ttgtttttgc gtatttaaca gaatttgtaa acctaggcaa
tgtttcagct cttcgaactt 660tcagagtctt gagagctttg aaaactattt ctgtaattcc
aggactaaaa accatcgtgg 720gggccctgat ccaatcagtg aagaagctct ctgacgtcat
gatcctcact gtgttctgtc 780tcagtgtgtt cgcactaatt ggactacaac tgtttatggg
caacttgaag cataaatgtt 840tccggaagga ccttgagcag aatgaaacat tagaaagcat
catgagtact gctgagagtg 900aagaagaatt gaaaagatat ttttattact tggagggatc
caaagatgct cttctttgcg 960gtttcagcac agattcaggg cagtgtcctg aagggtacga
gtgtgtgaca gctggcagaa 1020acccagatta tggctacaca agctttgaca cgttcggctg
ggccttcttg gccttgtttc 1080ggctaatgac tcaggactac tgggagaacc tttatcaaca
gacactgcgt gctgctggca 1140aaacctacat gattttcttt gtcgtggtga tatttctggg
atccttttac ctgataaact 1200tgatcctggc tgtggtagcc atggcgtacg aggaacagaa
ccaggccaac atcgaagaag 1260ctaaacagaa agagttagaa tttcagcaga tgttagaccg
actcaaaaaa gagcaggaag 1320aagccgaggc gatcgctgca gccgctgctg agtacacgag
tttagggcgg agcaggatta 1380tgggactctc tgagagctct tcagaaacct ccaggctgag
ctcaaagagt gccaaggaga 1440gaagaaaccg aagaaagaaa aaaaaacaga agctgtccag
tggcgaggaa aagggtgatg 1500atgagaagct gtccaagtca gggtcagagg aaagcatccg
aaagaaaagc ttccatctcg 1560gcgtggaagg gcaccaccgg gccagggaaa agaggctgtc
cacccccaac cagtcaccac 1620tcagcattcg tgggtccttg ttttctgcca ggcgcagcag
cagaacaagt ctcttcagtt 1680ttaaggggcg aggaagagat ctgggatctg aaacggaatt
tgctgatgat gagcatagca 1740tttttggaga caacgagagc agaaggggtt cactatttgt
accccataga ccccgggagc 1800ggcgcagcag taacatcagc caggccagta ggtccccacc
agtgctgccg gtgaacggga 1860agatgcacag tgcagtggac tgcaatggcg tggtgtcgct
tgttgatgga ccctcagccc 1920tcatgctccc caatggacag cttcttccag agggcacaac
taatcagatg cgtaaaaaaa 1980ggctctctag ttcttacttt ttgtctgagg acatgctgaa
tgacccacat ctcaggcaaa 2040gggccatgag cagagcaagc attctaacca acacagtaga
agaacttgaa gaatctagac 2100aaaaatgtcc accatggtgg tacagatttg ctcacacatt
tttaatctgg aattgttctc 2160catattggat aaaattcaaa aagttcatct attttattgt
aatggatcct tttgtagatc 2220ttgcaattac catttgcata gttttaaaca ccttgtttat
ggctatggag caccatccaa 2280tgacggatga attcaaaaat gtacttgcag tcgggaacct
ggtcttcaca gggatcttcg 2340cagctgaaat ggtactgaag ttaatagcca tggatcccta
tgaatatttc caagtagggt 2400ggaatatttt tgacagcctg attgtgacgt tgagtttggt
ggagcttttc ctagcagatg 2460tggaaggatt atcagtcctg cggtccttta gattgctgcg
agtcttcaag ttggcaaaat 2520cctggcccac actgaatatg ctcattaaga tcatcggcaa
ctcggtgggc gcactgggca 2580acctgaccct ggtgctggcc atcatcgtct tcatttttgc
cgtggtcggc atgcagctgt 2640ttggaaagag ctacaaggag tgtgtttgca agatcaacga
gaactgcaag ctcccacgct 2700ggcacatgaa cgacttcttc cactccttcc tgatcgtgtt
ccgtgtgctg tgtggggagt 2760ggatagagac catgtgggac tgcatggagg ttgcgggcca
gaccatgtgc cttattgttt 2820acatgatggt catggtgatt gggaaccttg tggtcctgaa
cctgtttctg gctttattac 2880tgagttcctt tagttctgac aatcttacag caattgaaga
agacaccgac gcaaacaacc 2940tccagattgc agtggccaga attaaaagag ggatcaatta
tgtgaaacag accctgcgtg 3000aattcattct aaagtcattt tccaaaaagc caaagggctc
caaggacaca aaacgaacag 3060cggatcccaa caacaaaaga gaaaactata tctcaaaccg
tacccttgcg gagataagca 3120aagatcacaa tttcctcaaa gaaaaggata agatcagtgg
ttttagcagc agtctagaca 3180aaagctttat ggatgaaaac gattaccagt cctttattca
taatcccagc ctcacagtga 3240cagtgcccat tgcacctggg gagtctgatt tggagaatat
gaacacagaa gagcttagca 3300gtgactcaga tagtgactac agcaaagaga gacggaaccg
atcaagttct tcagagtgca 3360gcacagttga taaccctctg ccaggagaag aggaggcaga
agctgagcct atcaatgcag 3420atgagcccga agcctgtttt acagatggct gtgtgaggag
atttccatgc tgccaagtta 3480acatagactc cgggaaaggg aaagtttggt ggaccatccg
gaagacctgc tacaggatag 3540tggagcacag ctggtttgaa agcttcattg ttctcatgat
cctgctcagc agtggagctc 3600tggcttttga ggatatatat attgaaaaga aaaagaccat
taagattatc ctggagtatg 3660ctgacaagat attcacctac atcttcattc tggaaatgct
tctaaaatgg gtggcatacg 3720ggtataaaac atatttcact aatgcctggt gttggctgga
cttcttaatt gttgatgtgt 3780ctctagttac tttagtagcc aacactcttg gctactcaga
ccttggcccc attaaatctc 3840tacggacact gagggcccta agacccctaa gagctttgtc
tagatttgaa ggaatgaggg 3900tagtggtcaa cgcactcata ggagcaatcc cttccatcat
gaatgtgctt cttgtgtgcc 3960ttatattctg gctaatattt agcatcatgg gagtcaatct
gtttgctggc aagttctatg 4020agtgtgttaa caccacagat ggctcacgat tttctgtatc
tcaagttgca aaccgttctg 4080agtgttttgc cctgatgaat gttagtggaa atgtgcgatg
gaaaaacctg aaagtaaact 4140tcgataacgt tggacttggt tacctgtcgc tgcttcaagt
tgcaacgttc aagggctgga 4200tggatattat gtatgcagca gttgactctg ttaatgtaaa
tgcacaacca atatatgaat 4260acaacctcta catgtacatt tattttgtca tcttcatcat
ctttggctca ttcttcactt 4320tgaacttgtt cattggtgtc atcatcgata atttcaacca
acagaagaag aagcttggag 4380gtcaagatat ctttatgaca gaagaacaga agaaatacta
taatgcaatg aagaagctgg 4440ggtccaagaa accacaaaaa ccaattccga ggccagggaa
caaattccaa ggatgcatat 4500ttgacttagt gacaaaccaa gcttttgata tcaccatcat
ggttcttatc tgcctcaata 4560tggtaaccat gatggtagaa aaagaggggc aaactgacta
catgagtttt gtgctatact 4620ggatcaacgt ggtcttcatc atcctgttca ctggggagtg
tgtgctgaag ctgatctctc 4680tcaggcatta ctacttcact gtgggatgga acatttttga
ttttgtggta gtgatcctct 4740ccattgtagg aatgtttctc gctgagatga tagagaagta
tttcgtgtct cctaccctgt 4800tccgagtcat tcgcctggcc aggattggac gaatcctacg
cctgatcaaa ggcgccaagg 4860ggatccgcac gctgctcttt gctctgatga tgtcccttcc
tgcgctgttc aacatcggcc 4920tcctgctttt cctcgtcatg ttcatctacg ccatctttgg
gatgtccaac tttgcctacg 4980ttaaaaagga agctggaatt aatgacatgt tcaactttga
gaccttcggc aacagcatga 5040tctgcctgtt ccaaatcacc acctctgcgg gctgggatgg
actgctggcc cccatcctca 5100acagtgcacc tcctgactgt gacccaaaaa aggttcaccc
aggaagttca gtggaagggg 5160actgtggaaa tccatctgtg ggaattttct actttgtcag
ctacatcatc atatccttcc 5220tggttgtggt gaacatgtac attgctgtca tcctggagaa
cttcagcgtt gccacagaag 5280aaagtactga gcccctgagt gaggacgact ttgagatgtt
ctacgaagtc tgggagaagt 5340tcgaccctga cgccacccag ttcatagagt tctgcaagct
ctctgacttt gcagcagccc 5400tggatcctcc cctcctcatc gcaaagccaa acaaagtcca
gctcattgcc atggacctgc 5460ccatggtgag tggagaccgc atccactgcc tggacatctt
atttgctttt acaaagcggg 5520tcctgggcga gagcggagag atggattccc ttcgttcaca
gatggaagaa aggtttatgt 5580cagccaatcc ttctaaagtg tcctatgagc ccatcacaac
cacactgaag cgaaaacaag 5640aggatgtatc tgcgactatc attcagcgtg cttacagacg
gtaccgcctt aggcaaaacg 5700tcaagaatat atcaagtata tatataaaag atggagacag
agatgacgat ttgcccaata 5760aagaagatat agtttttgat aatgttaacg agaactcaag
tccagaaaag acagatgcaa 5820cagcctctac catctctcca ccttcctatg acagtgtcac
aaagccagat caagagaaat 5880atgaaacaga caaaacggag aaggaagaca aagagaaaga
cgaaagcagg aaatagagct 5940tcggttttga tacactgttt acagcctgcg aaggtgactc
actcgtgtta ataagactct 6000tttacggagg tctatgccaa actcttttta tcaaatattc
tcaaaggcag cacagccact 6060agctctgatc cagtgaaaca agagagaagc atttacacat
ggctactttt tgcgttggtc 6120aatgattctt taagaattgt gcatgtaact ctacagggaa
taatcattat tgcaatcaag 6180ggtgacttaa tgattttaaa tatcagaaaa ccacatagaa
cattttctct tttgcctcca 6240tttctttccc tagattctaa gtagatgtgt acccatgtga
atatagaaat tcaggcgcac 6300atgctcacag tcacaaacac aaacaggatt agctgtgatt
tggaattcga tgtaaatatt 6360tcacctgtga tttgcaatga aattccttgt aaaagaaatg
cgaattagtg atgaaggttt 6420tgtgaaaaca tcttatcatt agggagtcag aatttctgtc
cataaagaat tcagtttata 6480ttttgaggtg ctgaaactta tcctacattg catcaaaatc
aatttatagg tatctgtaaa 6540atgtcatggg actgaaaaac atatataggc tacttgttta
agaaatggct ttcattcata 6600tagataggca ttcaccttga tttatggaca tctttggcat
tttgtgatca catgattctt 6660ccacaaaatt gcttagctgg aacttcaggc acacatcaca
gagaacagct acccagtctt 6720atgcccctct ctgtttgtac aataatcaca gagcttgaaa
cattatttga actataaata 6780tcaggtttct ccacatagac atatgaatat tgttaacaga
aaaaaatttt atttacagcg 6840ttttatttac taatatttat ccaatctagt ttgcccaatg
agacagctca tgactcacat 6900ctgaaagcca gttgccacat ttatcttctt atgtaacttt
ggtttgtcat actttatgtc 6960taagcaaatt gaatgtctcc tttctaatga gatgtaccct
gaaatgcagt taggtacttg 7020atactttagc gcttgtttga gcagatgact ggagcacagt
gtggactgca tctcttaaat 7080acaatcctta atgtgtttgg cagcttctca ggttacaagg
aacacccggc ttttagtgtc 7140tatctgttca ccaggtgttt agtatgaatg aaacggcatt
caaagagtga gtctcactgg 7200cttgctttat tactgatgcc ttccctatgg agaattaatc
ctctgaagcc ccattatgtc 7260cccctgtaaa taatgtagat gtcacttcct tcttaatatt
ctaatccata ctgtgaaatc 7320gattttgcat ttatcggtca aatagagcat tttgagatag
ttggagttac cctgccaagg 7380attagaaatc tacttcatgt ttttaaagta cttgttaaaa
atgaacgacc ctggcacatt 7440ctctcataat tttattccag ccatgtgaaa tctttcttct
aaacacttta tccttgcgga 7500ggaaaaaaaa aatgagctga tgagccattt aagcacaaag
gggctttatt tagaagattc 7560caagggggaa ctttgaagta aatatataaa acatacttca
tcaatttgcc tataaaacta 7620aaagaggaac acaggaatat tgataaaata agtcattaaa
acacattctt tatttcttgc 7680ccagtttaaa agaaagaact aaacatccct agagagagga
agagacatag agagagagtg 7740actaatagaa agagggagag aagaaacaag gcaccaagga
caaaaagaga taattagaca 7800gaacttgtcc aggttttcac actatgtgct ctgtccagta
ccgtacacaa gaacctcttt 7860ccaaatattt gtcctaaggc tctaagaagt taagtacgag
gctgaaggtt gaatacaact 7920gtctttaatc attaacagtt tggggagcta cttttaaacg
tctatggaag atgccaagca 7980gtggtaagcc agacaataca gagcactgca tatctgtcaa
gcagctgaaa tatgtttggg 8040caacttaatg gtgagccaca caaaacatcc atttgtaaca
attttaatac attcaattaa 8100gaaaccagga tttttattat tttgcaccca taaaaatata
actatattgt tcatttttat 8160tgatagagta tgtgtgaatc ttattgatta tctgtaattt
actattaatg tttttacagt 8220gactgttttt tttgtgtgtg taaacttaat atatgtcagc
aactggttcc tcaacacaat 8280tttttttagc attacaaaaa aatgaacagg tataaaggtt
ctcttttttc tacatcatgt 8340tgaacatatt ttgttctgaa ttacatagtt ttaaatgtaa
tattaagttt tatattcata 8400tatgtttaac atcaaaatca ctacttatga cattgttatc
aatttaaaaa atagtatttg 8460acactaggat agcatttaat taaagctaaa aagcttacac
cccatttcat gttgattagt 8520gtttggacta actctaaaat gtcatcaatg gaagctagtc
actgaaatta ttttatctat 8580tgtcatagaa tggtgactac ccaaaaaata taagttagca
ttaaatagaa gaaagcgtac 8640gtgaccacaa atccatgcac agggttgtgt gaagacagga
gaacctcatt tttctgtttt 8700gtctctttcc actgtgtaaa aagtctacat ctgtgggcta
tttctaaatt caaattgtca 8760caatttgcaa tcataaatgt ttagcatact ttgtagaatt
ttgatagttt tgtaaaagag 8820tgaaaaacaa atgcatatgt aaataaagca gcccatacta
gcagattcct caaatgttaa 8880tatgtaaata aagcagccct tactagcaga ttcatcaaat
gttaatatgt aaataaagca 8940gtccttatta gcagatttgt catatgttaa ggggagtaat
gataaggagg caactaaatc 9000aggatggtca gtaactgatc tgggtttaga actgtgtttg
gagccatcaa tttttaaata 9060tatgttctca ctatgttatt agttgtctga agaagcaatc
aagaattgct cccagaaaat 9120gagtaagtag ccatgaatat atgaatgctg tttacagaac
ccatagacct atgaatgctc 9180aaaatgtttg ggtttgtcaa aaaattacat tgtagttata
cttgatactt aaaaactgtt 9240aatagagtct aaaataaaag tcgct
9265201973PRTMus musculus 20Met Ala Met Leu Pro Pro
Pro Gly Pro Gln Ser Phe Val His Phe Thr 1 5
10 15 Lys Gln Ser Leu Ala Leu Ile Glu Gln Arg Ile
Ser Glu Glu Lys Ala 20 25
30 Lys Gly His Lys Asp Glu Lys Lys Asp Asp Glu Glu Glu Gly Pro
Lys 35 40 45 Pro
Ser Ser Asp Leu Glu Ala Gly Lys Gln Leu Pro Phe Ile Tyr Gly 50
55 60 Asp Ile Pro Pro Gly Met
Val Ser Glu Pro Leu Glu Asp Leu Asp Pro 65 70
75 80 Tyr Tyr Ala Asp Lys Lys Thr Phe Ile Val Leu
Asn Lys Gly Lys Ala 85 90
95 Ile Phe Arg Phe Asn Ala Thr Pro Ala Leu Tyr Met Leu Ser Pro Phe
100 105 110 Ser Pro
Leu Arg Arg Ile Ser Ile Lys Ile Leu Val His Ser Leu Phe 115
120 125 Ser Met Leu Ile Met Cys Thr
Ile Leu Thr Asn Cys Ile Phe Met Thr 130 135
140 Met Ser Asn Pro Pro Asp Trp Thr Lys Asn Val Glu
Tyr Thr Phe Thr 145 150 155
160 Gly Ile Tyr Thr Phe Glu Ser Leu Ile Lys Ile Leu Ala Arg Gly Phe
165 170 175 Cys Val Gly
Glu Phe Thr Phe Leu Arg Asp Pro Trp Asn Trp Leu Asp 180
185 190 Phe Val Val Ile Val Phe Ala Tyr
Leu Thr Glu Phe Val Asn Leu Gly 195 200
205 Asn Val Ser Ala Leu Arg Thr Phe Arg Val Leu Arg Ala
Leu Lys Thr 210 215 220
Ile Ser Val Ile Pro Gly Leu Lys Thr Ile Val Gly Ala Leu Ile Gln 225
230 235 240 Ser Val Lys Lys
Leu Ser Asp Val Met Ile Leu Thr Val Phe Cys Leu 245
250 255 Ser Val Phe Ala Leu Ile Gly Leu Gln
Leu Phe Met Gly Asn Leu Lys 260 265
270 His Lys Cys Phe Arg Lys Asp Leu Glu Gln Asn Glu Thr Leu
Glu Ser 275 280 285
Ile Met Ser Thr Ala Glu Ser Glu Glu Glu Leu Lys Arg Tyr Phe Tyr 290
295 300 Tyr Leu Glu Gly Ser
Lys Asp Ala Leu Leu Cys Gly Phe Ser Thr Asp 305 310
315 320 Ser Gly Gln Cys Pro Glu Gly Tyr Glu Cys
Val Thr Ala Gly Arg Asn 325 330
335 Pro Asp Tyr Gly Tyr Thr Ser Phe Asp Thr Phe Gly Trp Ala Phe
Leu 340 345 350 Ala
Leu Phe Arg Leu Met Thr Gln Asp Tyr Trp Glu Asn Leu Tyr Gln 355
360 365 Gln Thr Leu Arg Ala Ala
Gly Lys Thr Tyr Met Ile Phe Phe Val Val 370 375
380 Val Ile Phe Leu Gly Ser Phe Tyr Leu Ile Asn
Leu Ile Leu Ala Val 385 390 395
400 Val Ala Met Ala Tyr Glu Glu Gln Asn Gln Ala Asn Ile Glu Glu Ala
405 410 415 Lys Gln
Lys Glu Leu Glu Phe Gln Gln Met Leu Asp Arg Leu Lys Lys 420
425 430 Glu Gln Glu Glu Ala Glu Ala
Ile Ala Ala Ala Ala Ala Glu Tyr Thr 435 440
445 Ser Leu Gly Arg Ser Arg Ile Met Gly Leu Ser Glu
Ser Ser Ser Glu 450 455 460
Thr Ser Arg Leu Ser Ser Lys Ser Ala Lys Glu Arg Arg Asn Arg Arg 465
470 475 480 Lys Lys Lys
Lys Gln Lys Leu Ser Ser Gly Glu Glu Lys Gly Asp Asp 485
490 495 Glu Lys Leu Ser Lys Ser Gly Ser
Glu Glu Ser Ile Arg Lys Lys Ser 500 505
510 Phe His Leu Gly Val Glu Gly His His Arg Ala Arg Glu
Lys Arg Leu 515 520 525
Ser Thr Pro Asn Gln Ser Pro Leu Ser Ile Arg Gly Ser Leu Phe Ser 530
535 540 Ala Arg Arg Ser
Ser Arg Thr Ser Leu Phe Ser Phe Lys Gly Arg Gly 545 550
555 560 Arg Asp Leu Gly Ser Glu Thr Glu Phe
Ala Asp Asp Glu His Ser Ile 565 570
575 Phe Gly Asp Asn Glu Ser Arg Arg Gly Ser Leu Phe Val Pro
His Arg 580 585 590
Pro Arg Glu Arg Arg Ser Ser Asn Ile Ser Gln Ala Ser Arg Ser Pro
595 600 605 Pro Val Leu Pro
Val Asn Gly Lys Met His Ser Ala Val Asp Cys Asn 610
615 620 Gly Val Val Ser Leu Val Asp Gly
Pro Ser Ala Leu Met Leu Pro Asn 625 630
635 640 Gly Gln Leu Leu Pro Glu Gly Thr Thr Asn Gln Met
Arg Lys Lys Arg 645 650
655 Leu Ser Ser Ser Tyr Phe Leu Ser Glu Asp Met Leu Asn Asp Pro His
660 665 670 Leu Arg Gln
Arg Ala Met Ser Arg Ala Ser Ile Leu Thr Asn Thr Val 675
680 685 Glu Glu Leu Glu Glu Ser Arg Gln
Lys Cys Pro Pro Trp Trp Tyr Arg 690 695
700 Phe Ala His Thr Phe Leu Ile Trp Asn Cys Ser Pro Tyr
Trp Ile Lys 705 710 715
720 Phe Lys Lys Phe Ile Tyr Phe Ile Val Met Asp Pro Phe Val Asp Leu
725 730 735 Ala Ile Thr Ile
Cys Ile Val Leu Asn Thr Leu Phe Met Ala Met Glu 740
745 750 His His Pro Met Thr Asp Glu Phe Lys
Asn Val Leu Ala Val Gly Asn 755 760
765 Leu Val Phe Thr Gly Ile Phe Ala Ala Glu Met Val Leu Lys
Leu Ile 770 775 780
Ala Met Asp Pro Tyr Glu Tyr Phe Gln Val Gly Trp Asn Ile Phe Asp 785
790 795 800 Ser Leu Ile Val Thr
Leu Ser Leu Val Glu Leu Phe Leu Ala Asp Val 805
810 815 Glu Gly Leu Ser Val Leu Arg Ser Phe Arg
Leu Leu Arg Val Phe Lys 820 825
830 Leu Ala Lys Ser Trp Pro Thr Leu Asn Met Leu Ile Lys Ile Ile
Gly 835 840 845 Asn
Ser Val Gly Ala Leu Gly Asn Leu Thr Leu Val Leu Ala Ile Ile 850
855 860 Val Phe Ile Phe Ala Val
Val Gly Met Gln Leu Phe Gly Lys Ser Tyr 865 870
875 880 Lys Glu Cys Val Cys Lys Ile Asn Glu Asn Cys
Lys Leu Pro Arg Trp 885 890
895 His Met Asn Asp Phe Phe His Ser Phe Leu Ile Val Phe Arg Val Leu
900 905 910 Cys Gly
Glu Trp Ile Glu Thr Met Trp Asp Cys Met Glu Val Ala Gly 915
920 925 Gln Thr Met Cys Leu Ile Val
Tyr Met Met Val Met Val Ile Gly Asn 930 935
940 Leu Val Val Leu Asn Leu Phe Leu Ala Leu Leu Leu
Ser Ser Phe Ser 945 950 955
960 Ser Asp Asn Leu Thr Ala Ile Glu Glu Asp Thr Asp Ala Asn Asn Leu
965 970 975 Gln Ile Ala
Val Ala Arg Ile Lys Arg Gly Ile Asn Tyr Val Lys Gln 980
985 990 Thr Leu Arg Glu Phe Ile Leu Lys
Ser Phe Ser Lys Lys Pro Lys Gly 995 1000
1005 Ser Lys Asp Thr Lys Arg Thr Ala Asp Pro Asn
Asn Lys Arg Glu 1010 1015 1020
Asn Tyr Ile Ser Asn Arg Thr Leu Ala Glu Ile Ser Lys Asp His
1025 1030 1035 Asn Phe Leu
Lys Glu Lys Asp Lys Ile Ser Gly Phe Ser Ser Ser 1040
1045 1050 Leu Asp Lys Ser Phe Met Asp Glu
Asn Asp Tyr Gln Ser Phe Ile 1055 1060
1065 His Asn Pro Ser Leu Thr Val Thr Val Pro Ile Ala Pro
Gly Glu 1070 1075 1080
Ser Asp Leu Glu Asn Met Asn Thr Glu Glu Leu Ser Ser Asp Ser 1085
1090 1095 Asp Ser Asp Tyr Ser
Lys Glu Arg Arg Asn Arg Ser Ser Ser Ser 1100 1105
1110 Glu Cys Ser Thr Val Asp Asn Pro Leu Pro
Gly Glu Glu Glu Ala 1115 1120 1125
Glu Ala Glu Pro Ile Asn Ala Asp Glu Pro Glu Ala Cys Phe Thr
1130 1135 1140 Asp Gly
Cys Val Arg Arg Phe Pro Cys Cys Gln Val Asn Ile Asp 1145
1150 1155 Ser Gly Lys Gly Lys Val Trp
Trp Thr Ile Arg Lys Thr Cys Tyr 1160 1165
1170 Arg Ile Val Glu His Ser Trp Phe Glu Ser Phe Ile
Val Leu Met 1175 1180 1185
Ile Leu Leu Ser Ser Gly Ala Leu Ala Phe Glu Asp Ile Tyr Ile 1190
1195 1200 Glu Lys Lys Lys Thr
Ile Lys Ile Ile Leu Glu Tyr Ala Asp Lys 1205 1210
1215 Ile Phe Thr Tyr Ile Phe Ile Leu Glu Met
Leu Leu Lys Trp Val 1220 1225 1230
Ala Tyr Gly Tyr Lys Thr Tyr Phe Thr Asn Ala Trp Cys Trp Leu
1235 1240 1245 Asp Phe
Leu Ile Val Asp Val Ser Leu Val Thr Leu Val Ala Asn 1250
1255 1260 Thr Leu Gly Tyr Ser Asp Leu
Gly Pro Ile Lys Ser Leu Arg Thr 1265 1270
1275 Leu Arg Ala Leu Arg Pro Leu Arg Ala Leu Ser Arg
Phe Glu Gly 1280 1285 1290
Met Arg Val Val Val Asn Ala Leu Ile Gly Ala Ile Pro Ser Ile 1295
1300 1305 Met Asn Val Leu Leu
Val Cys Leu Ile Phe Trp Leu Ile Phe Ser 1310 1315
1320 Ile Met Gly Val Asn Leu Phe Ala Gly Lys
Phe Tyr Glu Cys Val 1325 1330 1335
Asn Thr Thr Asp Gly Ser Arg Phe Ser Val Ser Gln Val Ala Asn
1340 1345 1350 Arg Ser
Glu Cys Phe Ala Leu Met Asn Val Ser Gly Asn Val Arg 1355
1360 1365 Trp Lys Asn Leu Lys Val Asn
Phe Asp Asn Val Gly Leu Gly Tyr 1370 1375
1380 Leu Ser Leu Leu Gln Val Ala Thr Phe Lys Gly Trp
Met Asp Ile 1385 1390 1395
Met Tyr Ala Ala Val Asp Ser Val Asn Val Asn Ala Gln Pro Ile 1400
1405 1410 Tyr Glu Tyr Asn Leu
Tyr Met Tyr Ile Tyr Phe Val Ile Phe Ile 1415 1420
1425 Ile Phe Gly Ser Phe Phe Thr Leu Asn Leu
Phe Ile Gly Val Ile 1430 1435 1440
Ile Asp Asn Phe Asn Gln Gln Lys Lys Lys Leu Gly Gly Gln Asp
1445 1450 1455 Ile Phe
Met Thr Glu Glu Gln Lys Lys Tyr Tyr Asn Ala Met Lys 1460
1465 1470 Lys Leu Gly Ser Lys Lys Pro
Gln Lys Pro Ile Pro Arg Pro Gly 1475 1480
1485 Asn Lys Phe Gln Gly Cys Ile Phe Asp Leu Val Thr
Asn Gln Ala 1490 1495 1500
Phe Asp Ile Thr Ile Met Val Leu Ile Cys Leu Asn Met Val Thr 1505
1510 1515 Met Met Val Glu Lys
Glu Gly Gln Thr Asp Tyr Met Ser Phe Val 1520 1525
1530 Leu Tyr Trp Ile Asn Val Val Phe Ile Ile
Leu Phe Thr Gly Glu 1535 1540 1545
Cys Val Leu Lys Leu Ile Ser Leu Arg His Tyr Tyr Phe Thr Val
1550 1555 1560 Gly Trp
Asn Ile Phe Asp Phe Val Val Val Ile Leu Ser Ile Val 1565
1570 1575 Gly Met Phe Leu Ala Glu Met
Ile Glu Lys Tyr Phe Val Ser Pro 1580 1585
1590 Thr Leu Phe Arg Val Ile Arg Leu Ala Arg Ile Gly
Arg Ile Leu 1595 1600 1605
Arg Leu Ile Lys Gly Ala Lys Gly Ile Arg Thr Leu Leu Phe Ala 1610
1615 1620 Leu Met Met Ser Leu
Pro Ala Leu Phe Asn Ile Gly Leu Leu Leu 1625 1630
1635 Phe Leu Val Met Phe Ile Tyr Ala Ile Phe
Gly Met Ser Asn Phe 1640 1645 1650
Ala Tyr Val Lys Lys Glu Ala Gly Ile Asn Asp Met Phe Asn Phe
1655 1660 1665 Glu Thr
Phe Gly Asn Ser Met Ile Cys Leu Phe Gln Ile Thr Thr 1670
1675 1680 Ser Ala Gly Trp Asp Gly Leu
Leu Ala Pro Ile Leu Asn Ser Ala 1685 1690
1695 Pro Pro Asp Cys Asp Pro Lys Lys Val His Pro Gly
Ser Ser Val 1700 1705 1710
Glu Gly Asp Cys Gly Asn Pro Ser Val Gly Ile Phe Tyr Phe Val 1715
1720 1725 Ser Tyr Ile Ile Ile
Ser Phe Leu Val Val Val Asn Met Tyr Ile 1730 1735
1740 Ala Val Ile Leu Glu Asn Phe Ser Val Ala
Thr Glu Glu Ser Thr 1745 1750 1755
Glu Pro Leu Ser Glu Asp Asp Phe Glu Met Phe Tyr Glu Val Trp
1760 1765 1770 Glu Lys
Phe Asp Pro Asp Ala Thr Gln Phe Ile Glu Phe Cys Lys 1775
1780 1785 Leu Ser Asp Phe Ala Ala Ala
Leu Asp Pro Pro Leu Leu Ile Ala 1790 1795
1800 Lys Pro Asn Lys Val Gln Leu Ile Ala Met Asp Leu
Pro Met Val 1805 1810 1815
Ser Gly Asp Arg Ile His Cys Leu Asp Ile Leu Phe Ala Phe Thr 1820
1825 1830 Lys Arg Val Leu Gly
Glu Ser Gly Glu Met Asp Ser Leu Arg Ser 1835 1840
1845 Gln Met Glu Glu Arg Phe Met Ser Ala Asn
Pro Ser Lys Val Ser 1850 1855 1860
Tyr Glu Pro Ile Thr Thr Thr Leu Lys Arg Lys Gln Glu Asp Val
1865 1870 1875 Ser Ala
Thr Ile Ile Gln Arg Ala Tyr Arg Arg Tyr Arg Leu Arg 1880
1885 1890 Gln Asn Val Lys Asn Ile Ser
Ser Ile Tyr Ile Lys Asp Gly Asp 1895 1900
1905 Arg Asp Asp Asp Leu Pro Asn Lys Glu Asp Ile Val
Phe Asp Asn 1910 1915 1920
Val Asn Glu Asn Ser Ser Pro Glu Lys Thr Asp Ala Thr Ala Ser 1925
1930 1935 Thr Ile Ser Pro Pro
Ser Tyr Asp Ser Val Thr Lys Pro Asp Gln 1940 1945
1950 Glu Lys Tyr Glu Thr Asp Lys Thr Glu Lys
Glu Asp Lys Glu Lys 1955 1960 1965
Asp Glu Ser Arg Lys 1970 2160DNAHomo sapiens
21ttccaaatta caacctctgc tggctgggat ggattgctag cacctattct taacagtaag
602220PRTHomo sapiens 22Phe Gln Ile Thr Thr Ser Ala Gly Trp Asp Gly Leu
Leu Ala Pro Ile 1 5 10
15 Leu Asn Ser Lys 20 2360DNAHomo sapiens 23ttccaaatta
caacctctgc tggctgagat ggattgctag cacctattct taacagtaag 60248PRTHomo
sapiens 24Phe Gln Ile Thr Thr Ser Ala Gly 1 5
2560DNAHomo sapiens 25tcaggtcagt gtccagaggg gtacacctgt gtgaaaattg
gcagaaaccc tgattatggc 602620PRTHomo sapiens 26Ser Gly Gln Cys Pro Glu
Gly Tyr Thr Cys Val Lys Ile Gly Arg Asn 1 5
10 15 Pro Asp Tyr Gly 20 2760DNAHomo
sapiens 27tcaggtcagt gtccagaggg gtaaacctgt gtgaaaattg gcagaaaccc
tgattatggc 60287PRTHomo sapiens 28Ser Gly Gln Cys Pro Glu Gly 1
5 2960DNAHomo sapiens 29tcagttctgc gatcattcag
actgctccga gtcttcaagt tggcaaaatc ctggccaaca 603020PRTHomo sapiens
30Ser Val Leu Arg Ser Phe Arg Leu Leu Arg Val Phe Lys Leu Ala Lys 1
5 10 15 Ser Trp Pro Thr
20 3160DNAHomo sapiens 31tcagttctgc gatcattcag actgctctga
gtcttcaagt tggcaaaatc ctggccaaca 60329PRTHomo sapiens 32Ser Val Leu
Arg Ser Phe Arg Leu Leu 1 5 3392DNAHomo
sapiens 33atatgtgaca gagtttgtgg acctgggcaa tgtctcagca ttgagaacat
tcagagttct 60ccgagcattg aaaacaattt cagtcattcc ag
923431PRTHomo sapiens 34Tyr Val Thr Glu Phe Val Asp Leu Gly
Asn Val Ser Ala Leu Arg Thr 1 5 10
15 Phe Arg Val Leu Arg Ala Leu Lys Thr Ile Ser Val Ile Pro
Gly 20 25 30
3592DNAHomo sapiens 35gtatttaaca gaatttgtaa acctaggcaa tgtttcagct
cttcgaactt tcagagtatt 60gagagctttg aaaactattt ctgtaatccc ag
923692DNALepus americanus 36atatgtgacg gagtttgtgg
acctgggcaa tgtctcagcg ttgagaacat tcagagttct 60ccgagctttg aaaacaattt
cagtcattcc ag 923792DNALepus americanus
37gtatttaaca gaatttgtaa acctaggcaa tgtttcagct cttcgaactt tcagagtctt
60gagagctttg aaaactattt ctgtaatccc ag
923892DNAHomo sapiens 38atatgtgaca gagtttgtgg acctgggcaa tgtctcagcg
ttgagaacat tcagagttct 60ccgagcattg aaaacaattt cagtcattcc ag
923992DNAHomo sapiens 39gtatgtaaca gaatttgtaa
acctaggcaa tgtttcagct cttcgaactt tcagagtctt 60gagagctttg aaaactattt
ctgtaattcc ag 924092DNARattus
norvegicus 40atatgtgaca gagtttgtgg acctgggcaa tgtctcagcg ctgagaacgt
tcagagttct 60ccgagcattg aaaacaatat cagtcattcc ag
924192DNARattus norvegicus 41gtatgtaaca gaatttgtaa
acctaggcaa tgtttcagct cttcgaactt tcagagtctt 60gagagctttg aaaactattt
ctgtaattcc ag 924292DNAHomo sapiens
42atatgtgaca gagtttgtgg acctgggcaa tgtctcagcg ttgagaacat tcagagttct
60ccgagcactg aaaacaattt cagtcattcc ag
924392DNAHomo sapiens 43gtatgtaaca gaatttgtaa gcctaggcaa tgtttcagcc
cttcgaactt tcagagtctt 60gagagctctg aaaactattt ctgtaattcc ag
924492DNARattus norvegicus 44atatgtgaca
gagtttgtgg acctgggcaa tgtctcagcg ctgagaacgt tcagagttct 60ccgagcattg
aaaacaatat cagtcattcc ag
924592DNARattus norvegicus 45gtatgtaaca gaatttgtaa gcctaggcaa tgtttcagcc
cttcgaactt tcagagtctt 60gcgagctctg aaaactattt ctgtaattcc ag
924624DNAHomo sapiens 46aaaagtgatt gattcagttt
tttg 244720DNAHomo sapiens
47aaaagtgatt cagttttttg
204821DNAHomo sapiens 48cagaggggta cacctgtgtg a
214921DNAHomo sapiens 49cagaggggta aacctgtgtg a
215021DNAHomo sapiens
50cttttagctc cgagtcttca a
215121DNAHomo sapiens 51cttttagctc tgagtcttca a
215222DNAHomo sapiens 52ctgctggctg ggatggattt gc
225322DNAHomo sapiens
53ctgctggctg agatggattt gc
225421DNAHomo sapiens 54ctggcaaaca actgcccttc a
215521DNAHomo sapiens 55ctggcaaaca gctgcccttc a
215621DNAHomo sapiens
56tgaataaccc gccggactgg a
215721DNAHomo sapiens 57tgaataaccc accggactgg a
215821DNAHomo sapiens 58cgctgcgtgc cgctggcaaa a
215921DNAHomo sapiens
59cgctgcgtgc tgctggcaaa a
216021DNAHomo sapiens 60aagaattaga gtttcaacag a
216121DNAHomo sapiens 61aagaattaga atttcaacag a
216221DNAHomo sapiens
62tgttagaccg acttaaaaaa g
216321DNAHomo sapiens 63tgttagaccg tcttaaaaaa g
216419DNAHomo sapiens 64gttgtgtacg gaggttctc
196519DNAHomo sapiens
65gttgtgtatg gaggttctc
196621DNAHomo sapiens 66cgtattttgt gtcccctacc c
216721DNAHomo sapiens 67cgtattttgt ttcccctacc c
216822DNAHomo sapiens
68tttgcacctt tgaagactct gg
226922DNAHomo sapiens 69tttgcacctt gaaagactct gg
227021DNAHomo sapiens 70atttttttct aaggaaaagt t
217121DNAHomo sapiens
71atttttttct taggaaaagt t
217221DNAHomo sapiens 72atattcttag ttatttcaag t
217321DNAHomo sapiens 73atattcttag ctatttcaag t
217421DNAHomo sapiens
74ctttttgaaa tggcaaattt a
217521DNAHomo sapiens 75ctttttgaaa cggcaaattt a
217621DNAHomo sapiens 76tcagaaaaat tgatttttac a
217721DNAHomo sapiens
77tcagaaaaat ggatttttac a
217821DNAHomo sapiens 78ggatgcatat tgcctgggac c
217921DNAHomo sapiens 79ggatgcatat cgcctgggac c
218021DNAHomo sapiens
80tttgaatggc atatgtacct g
218121DNAHomo sapiens 81tttgaatggc gtatgtacct g
218221DNAHomo sapiens 82ggcatatgta tctggtgtat g
218321DNAHomo sapiens
83ggcatatgta cctggtgtat g
218421DNAHomo sapiens 84gcggtatatg cttggccttc t
218521DNAHomo sapiens 85gcggtatatg tttggccttc t
218621DNAHomo sapiens
86cccataatca cctcactgca t
218721DNAHomo sapiens 87cccataatca tctcactgca t
218821DNAHomo sapiens 88tttgtgaagc ttggggattg a
218921DNAHomo sapiens
89tttgtgaagc ctggggattg a
219022DNAHomo sapiens 90attttttttt tagggcacga cc
229123DNAHomo sapiens 91attttttttt ttagggcacg acc
239221DNAHomo sapiens
92atgttctctg tttttttctc c
219321DNAHomo sapiens 93atgttctctg cttttttctc c
219421DNAHomo sapiens 94tagtgagttt tagaattgac t
219521DNAHomo sapiens
95tagtgagttt cagaattgac t
219621DNAHomo sapiens 96tgttattttt ataggtttct t
219721DNAHomo sapiens 97tgttattttt gtaggtttct t
219822DNAHomo sapiens
98cgaaggatat aagttattct tt
229920DNAHomo sapiens 99cgaaggatat gttattcttt
2010021DNAHomo sapiens 100tttaaatagt ctattaatta t
2110121DNAHomo sapiens
101tttaaatagt atattaatta t
2110220DNAHomo sapiens 102tttaaaaaaa tctttacatt
2010321DNAHomo sapiens 103tttaaaaaaa atctttacat t
2110421DNAHomo sapiens
104ataattaact aggactaaga t
2110521DNAHomo sapiens 105ataattaact gggactaaga t
2110621DNAHomo sapiens 106ttcatgatta attttattag a
2110721DNAHomo sapiens
107ttcatgatta gttttattag a
2110820DNAHomo sapiens 108ttttttgttt ctttaccttg
2010924DNAHomo sapiens 109ttttttgttt gtttctttac
cttg 2411021DNAHomo sapiens
110aatatttatt attcagattt t
2111121DNAHomo sapiens 111aatatttatt tttcagattt t
2111251DNAHomo sapiens 112ctgccagagg tgataataga
taaggcaact tctgatgaca gcgtaaggac g 5111314PRTHomo sapiens
113Leu Pro Glu Val Ile Ile Asp Lys Ala Thr Ser Asp Asp Ser 1
5 10 11411PRTHomo sapiens
114Val Ile Ile Asp Lys Ala Thr Ser Asp Asp Ser 1 5
10 11511PRTRattus rattus 115Val Ile Ile Asp Lys Ala Thr
Ser Asp Asp Ser 1 5 10
11611PRTMus musculus 116Val Ile Ile Asp Lys Ala Thr Ser Asp Asp Ser 1
5 10 11711PRTRabbit 117Val Ile Ile Asp
Lys Ala Thr Ser Asp Asp Ser 1 5 10
11811PRTHomo sapiens 118Val Ile Ile Asp Lys Pro Ala Thr Asp Asp Asn 1
5 1011911PRTRattus rattus 119Val Ile Ile Asp
Lys Pro Ala Thr Asp Asp Asn 1 5
1012011PRTHomo sapiens 120Val Lys Ile Asp Lys Ala Ala Thr Asp Asp Ser 1
5 10 12110PRTRattus rattus 121Val
Lys Ile Asp Lys Ala Ala Thr Asp Ser 1 5
10 12210PRTMus musculus 122Val Lys Ile Asp Lys Ala Ala Thr Asp Ser
1 5 10
User Contributions:
Comment about this patent or add new information about this topic: