Patent application title: Polycistronic Vector for Human Induced Pluripotent Stem Cell Production
Inventors:
Tim Townes (Birmingham, AL, US)
Kevin M. Pawlik (Birmingham, AL, US)
IPC8 Class: AC12N5074FI
USPC Class:
424 9321
Class name: Whole live micro-organism, cell, or virus containing genetically modified micro-organism, cell, or virus (e.g., transformed, fused, hybrid, etc.) eukaryotic cell
Publication date: 2016-03-17
Patent application number: 20160076000
Abstract:
Methods of producing induced pluripotent stem (iPS) cells are provided.
For example, a method of producing an iPS cell from a differentiated
cell, which includes transforming the differentiated cell with a first
vector comprising a nucleic acid sequence comprising a nucleic acid
sequence encoding an Oct4, a nucleic acid sequence encoding a Sox2, and a
nucleic acid sequence encoding a Klf4. Each of the nucleic acid sequences
are separated from each other by a first and second viral 2A sequence.
The method described can further comprise culturing the transformed cell
under conditions that allow for the production of an iPS cell and
isolating the cultured iPS cell.Claims:
1. A method of producing an induced pluripotent stem (iPS) cell from a
differentiated cell comprising transforming the differentiated cell with
a first vector, wherein the first vector comprises a nucleic acid
sequence comprising (i) a nucleic acid sequence encoding an Oct4, (ii) a
nucleic acid sequence encoding a Sox2, and (iii) a nucleic acid sequence
encoding a Klf4, wherein each of the nucleic acid sequences, (i)-(iii),
are separated by a first and second nucleic acid encoding a viral 2A
sequence.
2. The method of claim 1, wherein the vector comprises SEQ ID NO:7.
3. The method of claim 1, wherein the vector comprises a nucleic acid sequence encoding SEQ ID NO:9.
4. The method of claim 1, further comprising culturing the transformed cell under conditions that allow for the production of a population of iPS cells.
5. The method of claim 1, further comprising isolating the population of iPS cells.
6. (canceled)
7. (canceled)
8. (canceled)
9. (canceled)
10. (canceled)
11. (canceled)
12. The method of claim 1, wherein the differentiated cell is a mammalian cell.
13. The method of claim 12, wherein the mammalian cell is a human cell.
14. The method of claim 13, wherein the mammalian cell is selected from the group consisting of a(n) epithelial cell, keratinocyte, fibroblast, hepatocyte, neuron, osteoblast, myocyte, kidney cell, lung cell, thyroid cell, and pancreatic cell.
15. The method of claim 14, wherein the mammalian cell is a keratinocyte.
16. (canceled)
17. (canceled)
18. (canceled)
19. The method of claim 1, wherein the first and second nucleic acid sequences encoding a viral 2A sequence comprises a nucleic acid sequence encoding the amino acid sequence ATNFSLLKQAGDVEENPGP (SEQ ID NO:2) or EGRGSLLTCGDVEENPGP (SEQ ID NO:3).
20. The method of claim 1, wherein the first nucleic acid sequence encoding a viral 2A sequence comprises a nucleic acid sequence encoding the amino acid sequence ATNFSLLKQAGDVEENPGP (SEQ ID NO:2) and the second nucleic acid sequence encoding a viral 2A sequence comprises a nucleic acid sequence encoding the amino acid sequence EGRGSLLTCGDVEENPGP (SEQ ID NO:3).
21. The method of claim 1, wherein the first vector is a plasmid, an adenoviral vector or a retroviral vector.
22. The method of claim 21, wherein the retroviral vector is a lentiviral vector.
23. The method of claim 22, wherein the lentiviral vector is a lentiviral SIN vector.
24. The method of claim 21, wherein the retroviral vector comprises a 3' long terminal repeat.
25. The method of claim 24, wherein the retroviral vector further comprises a loxP sequence.
26. The method of claim 25, wherein the loxP sequence is in a 3' long terminal repeat of the lentiviral vector.
27. The method of claim 25, further comprising transforming the iPS cell with a second vector, wherein the second vector comprises a nucleic acid encoding a Cre recombinase, wherein expression of the Cre recombinase results in the deletion of the first vector from the genome of the iPS cells.
28. The method of claim 27, further comprising isolating a population of iPS cells lacking the first vector.
29. An isolated iPS cell produced by the method described in claim 28.
30. The method of claim 1, further comprising correcting a genetic mutation in the differentiated cell, wherein the first vector further comprises a nucleic acid sequence comprising an unmutated nucleic acid sequence of interest and homologous nucleic acid sequences flanking the genetic mutation to be corrected.
31. The method of claim 30, wherein the genetic mutation is a mutation in the nucleic acid sequence encoding β-globin, the nucleic acid sequence encoding cystic fibrosis transmembrane conductance regulator, the nucleic acid sequence encoding phenylalanine hydroxylase, and the nucleic acid sequence encoding dystrophin.
32. The method of claim 31, wherein the genetic mutation is a mutation in the nucleic acid sequence encoding β-globin.
33. The method of claim 32, wherein the mutation in the nucleic acid sequence encoding β-globin results in a glutamic acid to valine substitution at the sixth amino acid of the β-globin protein.
34. The method of claim 33, wherein the glutamic acid to valine substitution is caused by an A to T transversion at base pair +20 relative to the A(+1) of the ATG start codon of the nucleic acid sequence encoding β-globin.
35. The method of claim 30, wherein the first vector further comprises a first and second loxP sequence.
36. The method of claim 35, wherein the first vector further comprises a nucleic acid sequence encoding a Cre recombinase operably linked to an inducible promoter.
37. The method of claim 36, wherein the inducible promoter comprises a Nanog-responsive thymidine kinase promoter.
38. The method of claim 30, wherein the first vector comprises SEQ ID NO:44.
39-70. (canceled)
71. A method of treating or preventing a disease associated with a genetic mutation in a subject, the method comprising: (a) selecting a subject with a disease associated with a genetic mutation; (b) isolating differentiated cells from the subject; (c) transforming the differentiated cells with a vector comprising an unmutated nucleic acid sequence of interest; (d) culturing the transformed cells under conditions that allow for the production of a population of iPS cells; (e) screening the iPS cells for correction of the genetic mutation; and (f) administering the iPS cells to the subject, wherein administration of the iPS cells treats or prevents the disease associated with the genetic mutation in the subject.
72. The method of claim 71, wherein the vector comprises a nucleic acid sequence comprising (i) an unmutated nucleic acid sequence of interest and homologous nucleic acid sequences flanking the genetic mutation, (ii) a nucleic acid sequence encoding a Cre recombinase operably linked to an inducible promoter, (iii) a first and second loxP sequence, (iv) a nucleic acid sequence encoding an Oct4, (v) a nucleic acid sequence encoding a Sox2, and (vi) a nucleic acid sequence encoding a Klf4, wherein each of the nucleic acid sequences, (iv)-(vi), are separated by a first and second nucleic acid sequence encoding a viral 2A sequence.
73. The method of claim 72, wherein the inducible promoter comprises a Nanog-responsive thymidine kinase promoter.
74. The method of claim 71, wherein the disease caused by the mutation in the genome is selected from the group consisting of sickle cell disease, thalassemia, cystic fibrosis, phenylketonuria, and Duchenne muscular dystrophy.
75. The method of claim 74, wherein the disease is sickle cell disease.
76. The method of claim 75, wherein the vector comprises SEQ ID NO:44.
77. The method of claim 71, wherein the differentiated cell is selected from the group consisting of a(n) epithelial cell, keratinocyte, fibroblast, hepatocyte, neuron, osteoblast, myocyte, kidney cell, lung cell, thyroid cell, and pancreatic cell.
78. The method of claim 77, wherein the differentiated cell is a keratinocyte.
Description:
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application is a divisional of U.S. application Ser. No. 13/480,753, filed May 25, 2012, which is a divisional of U.S. application Ser. No. 12/640,767, filed Dec. 17, 2009, which claims the benefit of U.S. Provisional Application No. 61/138,260, filed on Dec. 17, 2008, all of which are incorporated herein in their entireties by this reference.
BACKGROUND
[0003] Embryonic stem (ES) cells have the ability to grow indefinitely while maintaining pluripotency and the ability to differentiate into a multitude of different cell types. Because of these two qualities, human ES cell therapies have been proposed for regenerative medicine and tissue replacement after injury or disease. However, there are ethical difficulties regarding the use of human embryos for the isolation of human ES cells as well as problems with tissue rejection following transplantation of foreign ES cells in patients.
SUMMARY
[0004] Methods of producing induced pluripotent stem (iPS) cells are provided. For example, methods of producing an iPS cell from a differentiated cell are provided. The methods include the step of transforming the differentiated cell with a first vector comprising a nucleic acid sequence comprising a nucleic acid sequence encoding an Oct4, a nucleic acid sequence encoding a Sox2, and a nucleic acid sequence encoding a Klf4. Each of the nucleic acid sequences are separated by a first and second nucleic acid sequence encoding a viral 2A sequence.
[0005] Also provided are methods of producing an iPS cell, wherein the vector used to produce the cell is deleted from the genome of the iPS cell. For example, the methods include the step of transforming the differentiated cell with a first vector comprising a nucleic acid sequence comprising a nucleic acid sequence encoding an Oct4, a nucleic acid sequence encoding a Sox2, and a nucleic acid sequence encoding a Klf4. Each of the nucleic acid sequences are separated by a first and second nucleic acid sequence encoding a viral 2A sequence. The vector further comprises a loxP sequence. The methods further include the step of transforming the iPS cell with a second vector. The second vector comprises a nucleic acid sequence encoding a Cre recombinase. Expression of the Cre recombinase results in the deletion of the first retroviral vector from the genome of the cells.
[0006] Also provided are vectors comprising a nucleic acid sequence encoding an Oct4, a nucleic acid sequence encoding a Sox2, and a nucleic acid sequence encoding a Klf4, and cells comprising the vector. Each of the nucleic acid sequences are separated from each other by a first and second nucleic acid sequence encoding a viral 2A sequence.
[0007] Also provided are kits comprising a first vector and a second vector. The first vector comprises a nucleic acid sequence encoding an Oct4, a nucleic acid sequence encoding a Sox2, and a nucleic acid sequence encoding a Klf4. Each of the nucleic acid sequences are separated from each other by a first and second viral 2A sequence. The second vector comprises a nucleic acid sequence encoding a Cre recombinase.
[0008] Further provided are methods of treating or preventing a disease associated with a genetic mutation in a subject. The methods comprise selecting a subject with a disease associated with a genetic mutation; isolating differentiated cells from the subject; transforming the differentiated cells with a vector comprising an unmutated nucleic acid sequence of interest; culturing the transformed cells under conditions that allow for the production of a population of iPS cells; screening the iPS cells for correction of the genetic mutation; and administering the iPS cells to the subject, wherein administration of the iPS cells treats or prevents the disease associated with the genetic mutation in the subject. The vector comprises a nucleic acid sequence comprising (i) an unmutated nucleic acid sequence of interest and homologous nucleic acid sequences flanking the genetic mutation, (ii) a nucleic acid sequence encoding a Cre recombinase operably linked to an inducible promoter, (iii) a first and second loxP sequence, (iv) a nucleic acid sequence encoding an Oct4, (v) a nucleic acid sequence encoding a Sox2, and (vi) a nucleic acid sequence encoding a Klf4. Each of the nucleic acid sequences, (iv)-(vi), are separated by a first and second nucleic acid sequence encoding a viral 2A sequence.
DESCRIPTION OF DRAWINGS
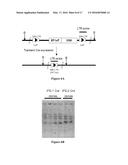
[0009] FIGS. 1A and 1B show the Oct4, Sox2, Klf4 (OSK) lentiviral vector for reprogramming adult skin fibroblasts to iPS cells. FIG. 1A shows a diagram of the vector. FIG. 1B shows the amino acid sequence of the 2A polypeptide with a 3-amino acid GSG linker (SEQ ID NO:1)
[0010] FIGS. 2A and 2B show images of iPS cell colonies. FIG. 2A shows immunofluorescent images of iPS cell colonies stained for Nanog and SSEA1 expression. FIG. 2B shows images of iPS cell colonies stained for alkaline phosphatase expression with iPS-1 Cre1 representing a typical colony after Cre recombinase mediated deletion of the OSK vector.
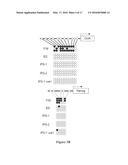
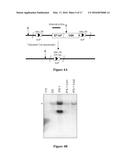
[0011] FIGS. 3A and 3B show RT-PCR analysis and Bisulfite sequence analysis of isolated iPS cells. FIG. 3A shows a gel of RT-PCR assays of polycistronic OSK RNA and endogenous Oct4, Sox2, Klf4, Nanog and Cripto RNA in iPS cells from 3 independent colonies (iPS-1, iPS-2, and iPS-3) and from iPS-1 cells post Cre recombinase mediated deletion of the OSK lentiviral vector (iPS-1 Cre1). FIG. 3B shows bisulfite sequencing of the endogenous and Oct4 and Nanog promoters in iPS-1, iPS-2, and iPS-1 Cre1 cells. Filled circles represent methylated CpGs and open circles represent unmethylated CpGs.
[0012] FIGS. 4A and 4B show a vector map and Southern blot hybridization of iPS-1 cellular DNA. FIG. 4A shows a map of the OSK vector pre- and post-Cre expression. K represents KpnI cleavage sites. The probe binding site is shown. FIG. 4B shows a Southern Blot demonstrating that iPS-1 cells contain 4 copies of the OSK lentiviral vector, and iPS-1 Cre1 cells contain no copies of the vector after transient Cre expression.
[0013] FIGS. 5A-5C show teratomas and chimeras derived from iPS cells. FIG. 5A shows teratomas containing tissue derived from all three germ layers in NOD/SCID IL-2 γR-/-mice injected with isolated iPS cells. a, intestine-like epithelium, with pancreatic acini in iPS-3 teratoma; b, respiratory epithelium; c, skeletal muscle; d, bone, with hyaline cartilage in iPS-2 teratoma; e, nervous tissue; f, skin-like stratified squamous epithelium. FIG. 5B shows chimeric embryos that were obtained following injection of iPS-1 Cre1 and iPS-1 Cre2 cells into wild type blastocysts. The top panel is a gel showing PCR products demonstrating chimeric embryos as iPS cells contain the human β-globin gene as a marker. FIG. 5C shows an adult chimeric animal (right) compared to an adult non-chimeric littermate (left).
[0014] FIGS. 6A and 6B show a vector map and Southern blot hybridization of iPS-1 and iPS-2 cellular DNA after OSK vector deletion. FIG. 6A shows a map of the OSK vector pre- and post-Cre expression. The probe binding site is shown. FIG. 6B shows a Southern blot demonstrating that iPS-1 Cre cells contain 4 insertion sites and iPS-2 Cre cells contain 3 insertion sites.
[0015] FIGS. 7A-G show the nucleotide (SEQ ID NO:7 for top strand and SEQ ID NO:8 for bottom strand) and amino acid (SEQ ID NO:9) sequences of the polycistron encoded by the vector. Underlined and labeled are primers used to create the polycistron. The Oct4, Sox2, Klf4 and PTV1 2A sequences are denoted.
[0016] FIG. 8 shows a brightfield image of an iPS cell colony derived from human keratinocytes using a polycistronic lentiviral vector.
[0017] FIG. 9 shows a schematic of a method to correct a β-globin mutation found in sickle cell disease with concomitant formation of iPS cells. The βs-globin locus is depicted at the top of the figure. The βs-globin locus has a single nucleotide, A to T transversion in the first exon. The targeting vector is depicted in the middle of the figure. The vector contains the normal GAG codon in the first exon flanked by sequences to effect homologous recombination. A herpes simplex virus thymidine kinase (HSV tk) gene is located outside of the sequences used to effect homologous recombination. Integrated between the homology arms is a floxed cassette (loxP site on either side of cassette) consisting of a Nanog-responsive (NBS) thymidine kinase (TK) promoter driving expression of Cre recombinase and the EF1α promoter driving expression of the Oct4-Sox2-Klf4 polycistronic sequence. The dashed lines show where the homologous recombination occurs. After homologous recombination occurs, the endogenous Nanog gene is expressed. Nanog binds to the NBS sites and forces Cre recombinase expression. Cre recombinase excises the floxed cassette and leaves behind a correct β-globin locus with a single loxP site in between exons 2 and 3 of β-globin.
DETAILED DESCRIPTION
[0018] A number of studies have been published detailing the production of induced pluripotent stem (iPS) cells from differentiated, embryonic and adult, mammalian cells (Takahashi and Yamanaka, Cell 1126:663-76 (2006); Meissner et al., Nat. Biotech. 25(10):1177-81 (2007); Takahashi et al., Cell 131:861-72 (2007); and Park et al., Nature 451:141-7 (2008)). In each of these publications, four transcription factors, Oct-3/4, Sox2, Klf4, and c-Myc, were introduced to the differentiated cells through retroviral transduction to produce iPS cells from differentiated somatic cells. Alternatively, it was found that another combination of factors, which include Oct-3/4, Sox2, Nanog, and Lin28, were capable of reprogramming somatic cells to iPS cells that exhibit the essential characteristics of embryonic stem (ES) cells (Yu et al., Science 18:1917-20 (2007)).
[0019] Oct4 and Sox2 are core transcription factors that function in the maintenance of pluripotency in early embryos and embryonic stem (ES) cells (Nichols et al., Cell 95:379-391 (1998); Niwa et al., Nat. Genet. 24:372-6 (2000); and Avilion et al., Gene Dev. 17:126-40 (2003)). Klf4 has been shown to contribute to the long-term maintenance of the ES cell phenotype and the rapid proliferation of ES cells in culture (Li et al., Blood 105:635-7 (2005)). Nanog is a transcription factor that is important in early development and stem cell pluripotency as it activates ES cell critical factors and represses differentiation-promoting genes (Wang et al., Proc. Natl. Acad. Sci. USA 105:6326-31 (2008)). Lin28 is a marker of undifferentiated human embryonic stem cells and has been shown to bind mRNAs in the cytoplasm as well as block the production of mature let-7 microRNA in mouse embryonic stem cells (Balzer and Moss, RNA Biology 4:16-25 (2007); Viswanathan et al., Science 320:97-100 (2008)). The c-Myc protein is also a transcription factor, as well as a tumor-related factor, and has many targets that enhance proliferation and transformation (Adhikary and Eilers, Nat. Rev. Mol. Cell. Bio. 6:635-45 (2005)) with many of these downstream targets potentially having roles in the generation of iPS cells. Additionally, c-Myc may globally induce histone acetylation (Fernandez et al., Genes Dev. 17:1115-29 (2003)), to allow other transcription factors to bind to their specific target loci. In the case of iPS cell production, expression of c-Myc would result in histone acetylation, thus allowing Oct3/4 and Sox2 to target the genes necessary to create a stem cell-like cell.
[0020] The use of retroviruses to incorporate Oct3/4, Sox2, Klf4, and c-Myc into the cells is both advantageous and deleterious. The advantages of using a retrovirus is that the virus integrates into the genome of the cell and thus is genetically transferred to the progeny when the cell undergoes cell division. This allows for the continued expression of these factors as differentiated cells undergo the transition to an iPS cell. In spite of these advantages, Takahashi et al. found that each iPS clone contained three to six retroviral integrations for each factor, creating the possibility of more than 20 retroviral integration sites per iPS clone, which increases the risk of tumorigenesis (Takahashi et al., Cell 131:861-72 (2007)). In fact, approximately 20% of mice derived from iPS cells developed tumors. This was attributable, at least in part, to the reactivation of the c-Myc retrovirus (Okita et al., Nature 448:313-7 (2007)).
[0021] The methods and compositions provided herein are designed to produce iPS cells that reduce the risk of insertional mutagenesis by allowing for the removal or deletion of vectors once the iPS cells have been generated or by using vectors that do not integrate into the cellular genome.
[0022] As used herein, the term induced pluripotent stem (iPS) cell encompasses any cell that has been reprogrammed to phenotypically resemble a pluripotent stem cell. An iPS cell is derived from a non-pluripotent cell but is capable of reproducing itself. An iPS cell is also capable of terminal differentiation into a cell-type normally found in the relevant system, tissue, or organ. An iPS cell is similar to an ES cell in morphology, proliferation, and pluripotency. For example, an iPS cell and an ES cell express the same markers. Examples of these markers include Oct3/4, Nanog, E-Ras, Cripto, Dax1, Fgf4, stage-specific embryonic antigen 1 (SSEA1), SSEA3, SSEA4, alkaline phosphatase, tumor-related antigen (TRA)-1-60, TRA-1-81, and Zfp296.
[0023] Provided herein are vectors for producing iPS cells. Thus, provided herein is a first vector comprising a nucleic acid sequence encoding an Oct4, a nucleic acid sequence encoding a Sox2, and a nucleic acid sequence encoding a Klf4. Each of the nucleic acid sequences are separated by a first and second nucleic acid sequence encoding a viral 2A sequence. The first nucleic acid sequence encoding a viral 2A sequence is the same as or different from the second nucleic acid sequence encoding a viral 2A sequence. Optionally, the first vector comprises SEQ ID NO:7. Optionally, the first vector comprises a nucleic acid sequence encoding SEQ ID NO:9. Optionally, the first vector comprises SEQ ID NO:43. The vector comprising SEQ ID NO:43 was deposited with the American Type Culture Collection, 10801 University Boulevard, Manassas, Va. 20110-2209 in accordance with the Budapest Treaty on Oct. 6, 2009, and has accession number PTA-10385.
[0024] Optionally, Oct4, Sox2, and Klf4 are human. Optionally, Oct4, Sox2, and Klf4 are non-human (e.g., rodent, canine, or feline). There are a variety of sequences that are disclosed on Genbank, at www.pubmed.gov and these sequences and others are herein incorporated by reference in their entireties as are individual subsequences or fragments contained therein. As used herein, Oct4 refers to the Oct4 transcription factor and homologs, variants, and isoforms thereof. For example, the nucleotide and amino acid sequences of human Oct4 can be found at GenBank Accession Nos. BC 117435 and AAI17436.1, respectively. Optionally, the nucleotide and amino acid sequences of human Oct4 isoform 1 can be found at GenBank Accession Nos. NM--002701.4 and NP--002692.2, respectively. The nucleotide and amino acid sequences for human Oct4 isoform 2 can be found at GenBank Accession Nos. NM--203289.3 and NP--976034.3, respectively. As used herein, Sox2 refers to the Sox2 transcription factor and homologs, variants, and isoforms thereof. The nucleotide and amino acid sequences of human Sox2 can be found at GenBank Accession Nos. BC013923 and AAH13923.1, respectively. Optionally, the nucleotide and amino acid sequences of human Sox2 can be found at GenBank Accession Nos. NM--003106.2 and NP--003097.1, respectively. As used herein, Klf4 refers to the Klf4 transcription factor and homologs, variants, and isoforms thereof. The nucleotide and amino acid sequences of human Klf4 can be found at GenBank Accession Nos. BC029923 and AAH29923.1, respectively. Optionally, the nucleotide and amino acid sequences of human Klf4 can be found at GenBank Accession Nos. NM--004235.4 and NP--004226.3, respectively. Thus provided are the nucleotide sequences of Oct4, Sox2, and Klf4 comprising a nucleotide sequence at least about 70%, 75%, 80%, 85%, 90%, 95%, 98%, 99% or more identical to the nucleotide sequence of the aforementioned GenBank Accession Numbers. Also provided are amino acid sequences of Oct4, Sox2, and Klf4 comprising an amino acid sequence at least about 70%, 75%, 80%, 85%, 90%, 95%, 98%, 99% or more identical to the sequences of the aforementioned GenBank Accession Numbers.
[0025] Nucleic acids that encode the polypeptide sequences, variants, and fragments thereof are disclosed. These sequences include all degenerate sequences related to a specific protein sequence, i.e., all nucleic acids having a sequence that encodes one particular protein sequence as well as all nucleic acids, including degenerate nucleic acids, encoding the disclosed variants and derivatives of the protein sequences. Thus, while each particular nucleic acid sequence may not be written out herein, it is understood that each and every sequence is in fact disclosed and described herein through the disclosed protein sequences.
[0026] As used herein, the term peptide, polypeptide or protein is used to mean a molecule comprised of two or more amino acids linked by a peptide bond. Protein, peptide, and polypeptide are also used herein interchangeably to refer to amino acid sequences. It should be recognized that the term polypeptide or protein is not used herein to suggest a particular size or number of amino acids comprising the molecule and that a polypeptide of the disclosure can contain up to several amino acid residues or more.
[0027] As with all peptides, polypeptides, and proteins, including fragments thereof, it is understood that additional modifications in the amino acid sequence of the variant Oct4, Sox2, and Klf4 polypeptides can occur that do not alter the nature or function of the peptides, polypeptides, or proteins. Such modifications include conservative amino acids substitutions and are discussed in greater detail below.
[0028] The polypeptides provided herein have a desired function. Oct4 and Sox2 are core transcription factors that regulate the expression of a defined set of target genes to maintain the pluripotency associated with ES cells. Klf4 is a transcription factor that regulates the expression of a defined set of target genes to maintain the long-term ES cell phenotype as well as to drive the proliferation of ES cells. The polypeptides are tested for their desired activity using the in vitro assays described herein.
[0029] The polypeptides described herein can be further modified and varied so long as the desired function is maintained. It is understood that one way to define any known modifications and derivatives or those that might arise, of the disclosed genes and proteins herein is through defining the modifications and derivatives in terms of identity to specific known sequences. Specifically disclosed are polypeptides which have at least 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83 , 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99 percent identity to Oct4, Sox2, and Klf4 and variants provided herein. Those of skill in the art readily understand how to determine the identity of two polypeptides. For example, the identity can be calculated after aligning the two sequences so that the identity is at its highest level.
[0030] Another way of calculating identity can be performed by published algorithms. Optimal alignment of sequences for comparison may be conducted by the local identity algorithm of Smith and Waterman, Adv. Appl. Math 2:482 (1981), by the identity alignment algorithm of Needleman and Wunsch, J. Mol. Biol. 48:443 (1970), by the search for similarity method of Pearson and Lipman, Proc. Natl. Acad. Sci. USA 85:2444 (1988), by computerized implementations of these algorithms (GAP, BESTFIT, FASTA, and TFASTA in the Wisconsin Genetics Software Package, Genetics Computer Group, 575 Science Dr., Madison, Wis.), or by inspection.
[0031] The same types of identity can be obtained for nucleic acids by, for example, the algorithms disclosed in Zuker, Science 244:48-52 (1989); Jaeger et al., Proc. Natl. Acad. Sci. USA 86:7706-10 (1989); Jaeger et al., Methods Enzymol. 183:281-306 (1989), which are herein incorporated by reference for at least material related to nucleic acid alignment. It is understood that any of the methods typically can be used and that in certain instances the results of these various methods may differ, but the skilled artisan understands if identity is found with at least one of these methods, the sequences would be said to have the stated identity and to be disclosed herein.
[0032] Protein modifications include amino acid sequence modifications. Modifications in amino acid sequence may arise naturally as allelic variations (e.g., due to genetic polymorphism), may arise due to environmental influence (e.g., by exposure to ultraviolet light), or may be produced by human intervention (e.g., by mutagenesis of cloned DNA sequences), such as induced point, deletion, insertion, and substitution mutants. These modifications can result in changes in the amino acid sequence, provide silent mutations, modify a restriction site, or provide other specific mutations. Amino acid sequence modifications typically fall into one or more of three classes: substitutional, insertional, or deletional modifications. Insertions include amino and/or terminal fusions as well as intrasequence insertions of single or multiple amino acid residues. Insertions ordinarily will be smaller insertions than those of amino or carboxyl terminal fusions, for example, on the order of one to four residues. Deletions are characterized by the removal of one or more amino acid residues from the protein sequence. Typically, no more than about from 2 to 6 residues are deleted at any one site within the protein molecule. Amino acid substitutions are typically of single residues, but can occur at a number of different locations at once; insertions usually will be on the order of about from 1 to 10 amino acid residues; and deletions will range about from 1 to 30 residues. Deletions or insertions preferably are made in adjacent pairs, i.e., a deletion of 2 residues or insertion of 2 residues. Substitutions, deletions, insertions or any combination thereof may be combined to arrive at a final construct. The mutations must not place the sequence out of reading frame and preferably will not create complementary regions that could produce secondary mRNA structure. Substitutional modifications are those in which at lease one residue has been removed and a different residues inserted in its place. Such substitutions generally are made in accordance with the following Table 1 and are referred to as conservative substitutions.
TABLE-US-00001 TABLE 1 Amino Acid Substitutions Substitutions Amino Acid (others are known in the art) Ala Ser, Gly, Cys Arg Lys, Gln, Met, Ile Asn Gln, His, Glu, Asp Asp Glu, Asn, Gln Cys Ser, Met, Thr Gln Asn, Lys, Glu, Asp Glu Asp, Asn, Gln Gly Pro, Ala His Asn, Gln Ile Leu, Val, Met Leu Ile, Val, Met Lys Arg, Gln, Met, Ile Met Leu, Ile, Val Phe Met, Leu, Tyr, Trp, His Ser Thr, Met, Cys Thr Ser, Met, Val Trp Tyr, Phe Tyr Trp, Phe, His Val Ile, Leu, Met
[0033] Modifications, including the specific amino acid substitutions, are made by known methods. By way of example, modifications are made by site specific mutagenesis of nucleotides in the DNA encoding the protein, thereby producing DNA encoding the modification, and thereafter expressing the DNA in recombinant cell culture. Techniques for making substitution mutations at predetermined sites in DNA having a known sequence are well known, for example M13 primer mutagenesis and PCR mutagenesis.
[0034] Optionally, the vector comprises its various components in any order. Examples include from the 5' end, a nucleic acid sequence encoding a first polypeptide, the first nucleic acid encoding a viral 2A sequence, a nucleic acid encoding a second polypeptide, the second nucleic acid sequence encoding a viral 2A sequence, and a nucleic acid sequence encoding a third polypeptide. The first nucleic acid sequence encoding a viral 2A sequence is the same as or different from the second nucleic acid sequence encoding a viral 2A sequence. The first, second, and third polypeptides are selected from the group consisting of Oct4, Sox2, and Klf4, and the first, second, and third polypeptides are different from each other. Thus, for example, the first polypeptide is Oct4, the second polypeptide is Sox2, and the third polypeptide is Klf4. By way of another example, the first polypeptide is Sox2, the second polypeptide is Oct4, and the third polypeptide is Klf4.
[0035] The vector comprises in order from the 5' end, a nucleic acid sequence encoding an Oct4, a first nucleic acid sequence encoding a viral 2A sequence, a nucleic acid sequence encoding a Sox2, a second nucleic acid sequence encoding a viral 2A sequence, and a nucleic acid sequence encoding a Klf4. Optionally, the vector comprises in order from the 5' end, a nucleic acid sequence encoding an Oct4, a first nucleic acid sequence encoding a viral 2A sequence, a nucleic acid sequence encoding a Klf4, a second nucleic acid sequence encoding a viral 2A sequence, and a nucleic acid sequence encoding a Sox2. Optionally, the vector comprises in order from the 5' end, a nucleic acid sequence encoding a Sox2, a first nucleic acid sequence encoding a viral 2A sequence, a nucleic acid sequence encoding an Oct4, a second nucleic acid sequence encoding a viral 2A sequence, and a nucleic acid sequence encoding a Klf4. Optionally, the vector comprises in order from the 5' end, a nucleic acid sequence encoding a Sox2, a first nucleic acid sequence encoding a viral 2A sequence, a nucleic acid sequence encoding a Klf4, a second nucleic acid sequence encoding a viral 2A sequence, and a nucleic acid sequence encoding an Oct4. Optionally, the vector comprises in order from the 5' end, a nucleic acid sequence encoding a Klf4, a first nucleic acid sequence encoding a viral 2A sequence, a nucleic acid sequence encoding an Oct4, a second nucleic acid sequence encoding a viral 2A sequence, and a nucleic acid sequence encoding a Sox2. Optionally, the vector comprises in order from the 5' end, a nucleic acid sequence encoding a Klf4, a first nucleic acid sequence encoding a viral 2A sequence, a nucleic acid sequence encoding a Sox2, a second nucleic acid sequence encoding a viral 2A sequence, and a nucleic acid sequence encoding an Oct4.
[0036] A common strategy of positive-strand RNA viruses is to encode some, or all, of their proteins in the form of a polyprotein translated from one RNA molecule. Viruses have adapted multiple methods to allow for the production of individual protein molecules from a polyprotein. In the case of picornaviruses, all of the proteins are encoded in a single open reading frame. The picornaviral polyproteins undergo a cleavage event between the major domains of the viral genome, which are separated by viral 2A sequences. Viral 2A sequences allow for the translation of multiple polypeptides in a multicistronic RNA molecule by stimulating peptide cleavage between the polypeptides without disengaging the ribosome. The use of viral 2A sequences to produce multiple proteins from a multicistronic message is known, see, e.g., Donnelly et al., J. Gen. Virol. 82:1013-25 (2001); Donnelly et al., J. Gen. Virol. 82:1027-41 (2001); Chinnasamy et al., Virol. J. 3:14 (2006); Holstet al., Nat. Protoc. 1(1):406-17 (2006); and Szymczak et al., Nat. Biotechnol. 22(5):589-94 (2004).
[0037] Optionally, the first and second nucleic acid sequences encoding a viral 2A sequence is a picornaviral, a tetraviral 2A sequence, or a combination thereof. Optionally, the picornaviral 2A sequences are selected from the group consisting of the Enteroviral 2A sequences, Rhinoviral 2A sequences, Cardioviral 2A sequences, Aphthoviral 2A sequences, Hepatoviral 2A sequences, Erboviral 2A sequences, Kobuviral 2A sequences, Teschoviral 2A sequences, and the Parechoviral 2A sequences. Optionally, the tetraviral 2A sequences are selected from Betatetraviral 2A sequences or Omegatetraviral 2A sequences. Optionally, the first and second nucleic acid sequences encoding a viral 2A sequence are picornaviral 2A sequences. Optionally, the first and second nucleic acid sequence encoding a viral 2A sequence is a Teschoviral 2A sequence. Optionally, the first nucleic acid sequence encoding a viral 2A sequence is a Cardioviral 2A sequence, and the second nucleic acid sequence encoding a viral 2A sequence is a Hepatoviral 2A sequence. Optionally, the first and second nucleic acid sequences encoding a viral 2A sequence are tetraviridae 2A sequences. Optionally, the first and second nucleic acid sequences encoding a viral 2A sequence is a Betatetraviral 2A sequence. Optionally, the first nucleic acid sequence encoding a viral 2A sequence is a Betatetraviral 2A sequence, and the second nucleic acid sequence encoding a viral 2A sequence is an Omegatetraviral 2A sequence. Optionally, the first nucleic acid sequence encoding a viral 2A sequence is a picornaviral 2A sequence, and the second nucleic acid sequence encoding a viral 2A sequence is a tetraviridae 2A sequence. Optionally, the first nucleic acid sequence encoding a viral 2A sequence is a Teschoviral 2A sequence, and the second nucleic acid sequence encoding a viral 2A sequence is a Betatetraviral 2A sequence. Optionally, the first nucleic acid sequence encoding a viral 2A sequence is a tetraviridae 2A sequence, and the second nucleic acid sequence encoding a viral 2A sequence is a picornaviral 2A sequence. Optionally, the first nucleic acid sequence encoding a viral 2A sequence is a Betatetraviral 2A sequence, and the second nucleic acid sequence encoding a viral 2A sequence is a Teschoviral 2A sequence. Optionally, the first and second nucleic acid sequences encoding a viral 2A sequence comprise a nucleic acid sequence encoding the amino acid sequence ATNFSLLKQAGDVEENPGP (SEQ ID NO:2). Optionally, the first and second nucleic acid sequences encoding a viral 2A sequence comprise a nucleic acid sequence encoding the amino acid sequence EGRGSLLTCGDVEENPGP (SEQ ID NO:3). Optionally the first nucleic acid sequence encoding a viral 2A sequence comprises a nucleic acid sequence encoding the amino acid sequence ATNFSLLKQAGDVEENPGP (SEQ ID NO:2), and the second nucleic acid sequence encoding a viral 2A sequence comprises a nucleic acid sequence encoding the amino acid sequence EGRGSLLTCGDVEENPGP (SEQ ID NO:3).
[0038] Optionally the first and second nucleic acid sequences encoding a viral 2A sequence comprises a nucleic acid sequence encoding an amino acid linker. The amino acid linker can be 1 to 10 amino acids in length. The amino acid linker can be 1 to 5 amino acids in length. The amino acid linker can be 1 to 3 amino acids in length. The amino acid linker is preferably 3 amino acids in length. The amino acid linker is, for example, GSG (SEQ ID NO:4). Optionally the first and second nucleic acid sequences encoding a viral 2A sequence with an amino acid linker comprise a nucleic acid sequence encoding the amino acid sequence GSGATNFSLLKQAGDVEENPGP (SEQ ID NO:1). Optionally the first and second nucleic acid sequences encoding a viral 2A sequence with an amino acid linker comprise a nucleic acid sequence encoding the amino acid sequence GSGEGRGSLLTCGDVEENPGP (SEQ ID NO:5).
[0039] The provided vector, for example, can be a retroviral vector. Retroviral vectors are able to integrate efficiently into the genomic DNA of cells. Integration into the genomic DNA allows for the continuous expression of the transgene and additionally allows for the transmission of the transgene to progeny cells when the cells divide. Another advantage of retroviral vectors is that they have the ability of being able to transduce a wide range of cell types from different animal species. Examples of retroviral vectors are known. See, e.g., Coffin et al., Retorviruses, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y. (1997).
[0040] Optionally, the retroviral vector is a lentiviral vector. Lentiviral vectors are capable of infecting non-dividing cells. Optionally, the lentiviral vector is a lentiviral self-inactivating (SIN) vector. Lentiviral SIN vectors overcome the risk of activating cellular oncogenes when they are randomly integrated into the host genome. The lentiviral SIN vector is generated by deleting viral enhancer and promoter sequences within the vector, so that integration into the genome does not result in the activation of cellular oncogenes driven by the viral promoter and enhancer sequences. Methods of making and using the lentiviral SIN vectors are known. See, e.g., Miyoshi et al., J. Virol. 72(10):8150-7 (1998) and Zufferey et al., J. Virol. 72(12):9873-80 (1998).
[0041] Optionally, the retroviral vector contains a loxP sequence (e.g., ATAACTTCGTATAATGTATGCTATACGAAGTTAT (SEQ ID NO:6)). The loxP nucleic acid sequence is generally a 34 base pair nucleic acid sequence derived from Bacteriophage P1 that is used in combination with Cre recombinase to allow for site specific recombination. When a nucleic acid sequence contains a loxP sequence, the location of the loxP sequence is referred to as a loxP site. Usually, a nucleic acid sequence contains two loxP sites. The loxP sites are located on either side of a nucleic acid sequence to be removed from, for example, the genome of a cell. Expression of Cre recombinase in the cell promotes a recombination event that results in the deletion of the genomic DNA that is present in between the loxP sites. Specifically, the Cre recombinase binds and catalyzes the cleavage and strand exchange of DNA at two loxP sites, excising the nucleic acid between the loxP sites, and leaving a single loxP site in the genome. Examples of the Cre/lox system are known. See, e.g., Sauer, Methods 14(4):381-92 (1998); Florin et al., Genesis 38(3):139-44; and Schnutgen et al., Nat. Biotechnol. 21(5):562-5 (2003).
[0042] Optionally, the loxP sequence is located in the 3' long terminal repeat of the vector. Retroviral integration into the genome of a cell occurs in a three part process. First the retroviral RNA is reverse transcribed by a virally encoded RNA reverse transcriptase to form a RNA-DNA hybrid helix. The reverse transcriptase uses the newly synthesized DNA as a template to synthesize the complementary DNA, while degrading the RNA template. The resulting DNA duplex is integrated into the genome of the cell with the loxP sequence in the 3' long terminal repeat of the retroviral vector copied into the 5' long terminal repeat during reverse transcription and then integrated into the genome. This provides a loxP sequence at either end of the integrated lentiviral vector; therefore, making it possible to remove the integrated retroviral vector by expression of Cre recombinase. Optionally, provided is a second vector comprising a nucleic acid encoding a Cre recombinase. Expression of the Cre recombinase results in the deletion of the first vector from the genome of the iPS cells.
[0043] Optionally, the vector is designed to correct a genetic mutation associated with a disease and to produce induced pluripotent stem (iPS) cells. The vector comprises a nucleic acid sequence comprising (i) a nucleic acid sequence encoding an Oct4, (ii) a nucleic acid sequence encoding a Sox2, and (iii) a nucleic acid sequence encoding a Klf4. Each of the nucleic acid sequences, (i)-(iii), are separated by a first and second nucleic acid sequence encoding a viral 2A sequence. The first nucleic acid sequence encoding a viral 2A sequence is the same as or different from the second nucleic acid sequence encoding a viral 2A sequence. The vector further comprises an unmutated nucleic acid sequence of interest and homologous nucleic acid sequences flanking the genetic mutation. An unmutated nucleic acid sequence of interest is a nucleic acid sequence lacking the genetic mutation associated with the disease. Optionally, the unmutated nucleic acid sequence of interest comprises the nucleic acid sequence encoding β-globin. Optionally, the vector further comprises a first and second loxP sequence. Optionally, the vector further comprises a nucleic acid sequence encoding a Cre recombinase operably linked to an inducible promoter. The inducible promoter, for example, can comprise a Nanog-responsive thymidine kinase promoter. Optionally, the vector can comprise a selectable marker. Optionally, the vector comprises SEQ ID NO:44.
[0044] Optionally, the nucleic acid comprising a nucleic acid sequence encoding an Oct4, a nucleic acid sequence encoding a Sox2, and a nucleic acid sequence encoding a Klf4, wherein the nucleic acid sequences are separated by a first and second nucleic acid sequence encoding a viral 2A sequence is administered by another type of vector comprising the nucleic acid. The vector based delivery is largely broken down into two classes: viral based delivery systems and non-viral based delivery systems. Such methods are known in the art and are readily adaptable for use with the methods described herein.
[0045] Provided herein are viral based expression vectors comprising the disclosed nucleic acid. Viral based delivery systems can, for example, include Adenoviral vectors, Adeno-associated viral vectors, Herpes viral vectors, Vaccinia viral vectors, Polio viral vectors, Sindbis viral vectors, and any other RNA viral vectors. Also useful are any viral families that share the properties of these listed viruses and vectors that make them suitable for use as vectors. The construction of replication-defective adenoviruses has been described (Berkner et al., J. Virology 61:1213-20 (1987); Massie et al., Mol. Cell. Biol. 6:2872-83 (1986); Haj-Ahmad et al., J. Virology 57:267-74 (1986); Davidson et al., J. Virology 61:1226-39 (1987); Zhang et al., BioTechniques 15:868-72 (1993)). The viral vectors are limited in the extent to which they can spread to other cell types, since they can replicate within an initial infected cell but are unable to form new infectious viral particles. Recombinant adenoviruses have been shown to achieve high efficiency after direct, in vivo delivery to airway epithelium, hepatocytes, vascular endothelium, CNS parenchyma and a number of other tissue sites. Other useful systems include, for example, replicating and host-restricted non-replicating vaccinia virus vectors.
[0046] Provided herein are also non-viral based expression vectors comprising the disclosed nucleic acids. Suitable vector backbones include, for example, plasmids, artificial chromosomes, BACs, YACs, or PACs. Numerous vectors and expression systems are commercially available from such corporations as Novagen (Madison, Wis.), Clonetech (Palo Alto, Calif.), Stratagene (La Jolla, Calif.), and Invitrogen/Life Technologies (Carlsbad, Calif.). Vectors typically contain one or more regulatory regions. Regulatory regions include, without limitation, promoter sequences, enhancer sequences, response elements, protein recognition sites, inducible elements, protein binding sequences, 5' and 3' untranslated regions (UTRs), transcriptional start sites, termination sequences, polyadenylation sequences, and introns.
[0047] Any of the vectors provided herein can have a promoter sequence that drives the expression of the nucleic acid sequence comprising a nucleic acid sequence encoding a an Oct4, a nucleic acid sequence encoding a Sox2, and a nucleic acid sequence encoding a Klf4. Each of the nucleic acid sequences are separated from each other by a first and second viral 2A sequence. The first viral 2A sequence is the same as or different from the second viral 2A sequence. Preferred promoters controlling transcription from vectors in mammalian host cells may be obtained from various sources, for example, the genomes of viruses such as polyoma, Simian Virus 40 (SV40), adenovirus, retroviruses, hepatitis B virus and most preferably cytomegalovirus, or from heterologous mammalian promoters, e.g. beta actin promoter or EF1 promoter, or from hybrid or chimeric promoters (e.g., cytomegalovirus promoter fused to the beta actin promoter). The early and late promoters of the SV40 virus are conveniently obtained as an SV40 restriction fragment which also contains the SV40 viral origin of replication. The immediate early promoter of the human cytomegalovirus is conveniently obtained as a HindIII E restriction fragment. Of course, promoters from the host cell or related species also are useful herein.
[0048] The promoter can be an inducible promoter (e.g. chemically or physically regulated promoter). A chemically regulated promoter can, for example, be regulated by the presence of alcohol, tetracycline, a steroid, or a metal. A physically regulated promoter can, for example, be regulated by environmental factors, such as temperature and light. The promoter can be a cell type specific promoter (e.g. neuronal-specific, renal-specific, cardio-specific, liver-specific, or muscle-specific). A cell-type specific promoter is only expressed in the cell-type in which it is intended to be expressed. The promoter can be a promoter that is expressed independent of cell type. Examples of promoters that can be expressed independent of cell type include the cytomegalovirus (CMV) promoter, the Raus sarcoma virus (RSV) promoter, the adenoviral E1A promoter, and the EF-la promoter. The promoter is preferably the EF-la promoter.
[0049] Enhancer generally refers to a sequence of DNA that functions at no fixed distance from the transcription start site and can be either 5' or 3' to the transcription unit. Furthermore, enhancers can be within an intron as well as within the coding sequence itself. They are usually between 10 and 300 base pairs in length, and they function in cis. Enhancers usually function to increase transcription from nearby promoters. Enhancers can also contain response elements that mediate the regulation of transcription. While many enhancer sequences are now known from mammalian genes (globin, elastase, albumin, fetoprotein and insulin), typically one will use an enhancer from a eukaryotic cell virus for general expression. Preferred examples are the SV40 enhancer on the late side of the replication origin, the cytomegalovirus early promoter enhancer, the polyoma enhancer on the late side of the replication origin, and adenovirus enhancers.
[0050] The vectors also can include, for example, origins of replication, scaffold attachment regions (SARs), and/or markers. A marker gene can confer a selectable phenotype, e.g., antibiotic resistance, on a cell. This marker product is used to determine if the gene has been delivered to the cell and once delivered is being expressed. Examples of marker genes include the E. coli lacZ gene, which encodes B galactosidase, green fluorescent protein (GFP), and luciferase. Examples of suitable selectable markers for mammalian cells are dihydrofolate reductase (DHFR), thymidine kinase, neomycin, neomycin analog G418, hygromycin, blasticidin, and puromycin. When such selectable markers are successfully transferred into a mammalian host cell, the transformed mammalian host cell can survive if placed under selective pressure. In addition, an expression vector can include a tag sequence designed to facilitate manipulation or detection (e.g., purification or localization) of the expressed polypeptide. Tag sequences, such as green fluorescent protein (GFP), glutathione S-transferase (GST), polyhistidine, c-myc, hemagglutinin, or FLAG® tag (Kodak, New Haven, Conn.) sequences typically are expressed as a fusion with the encoded polypeptide. Such tags can be inserted anywhere within the polypeptide, including at either the carboxyl or amino terminus.
[0051] Provided herein are methods for the production of iPS cells from differentiated cells. The methods include transforming the differentiated cell with a first vector comprising a nucleic acid sequence comprising a nucleic acid sequence encoding an Oct4, a nucleic acid sequence encoding a Sox2, and a nucleic acid sequence encoding a Klf4. Each of the nucleic acid sequences are separated by a first and second nucleic acid sequence encoding a viral 2A sequence. The first nucleic acid sequence encoding a viral 2A sequence can be the same as or different from the second nucleic acid sequence encoding a viral 2A sequence. Optionally, the method further includes transforming the differentiated cell with a second vector comprising a nucleic acid sequence encoding a c-Myc. Optionally, the first vector comprises a nucleic acid sequence comprising a nucleic acid sequence encoding an Oct4, a nucleic acid sequence encoding a Sox2, a nucleic acid sequence encoding a Klf4, and a nucleic acid sequence encoding a c-Myc. Each of the nucleic acid sequences are separated by a first, second, and third nucleic acid sequence encoding a viral 2A sequence. The first nucleic acid sequence encoding a viral 2A sequence can be the same as or different from the second nucleic acid sequence encoding a viral 2A sequence. The second nucleic acid sequence encoding a viral 2A sequence can be the same as or different from the third nucleic acid sequence encoding a viral 2A sequence. Optionally, the first vector comprises a nucleic acid sequence comprising a nucleic acid sequence encoding an Oct4, a nucleic acid sequence encoding a Sox2, and a nucleic acid sequence encoding a Nanog, wherein the nucleic acid sequences are each separated by a first and second nucleic acid sequence encoding a viral 2A sequence. The first nucleic acid sequence encoding a viral 2A sequence can be the same as or different from the second nucleic acid encoding a viral 2A sequence. The method further includes transforming the differentiated cell with a second vector comprising a nucleic acid sequence encoding a Lin28. Optionally, the first vector comprises a nucleic acid sequence comprising a nucleic acid sequence encoding an Oct4, a nucleic acid sequence encoding a Sox2, a nucleic acid sequence encoding a Nanog, and a nucleic acid sequence encoding a Lin28. Each of the nucleic acid sequences are separated by a first, second, and third nucleic acid sequence encoding a viral 2A sequence. The first nucleic acid sequence encoding a viral 2A sequence can be the same as or different from the second nucleic acid sequence encoding a viral 2A sequence. The second nucleic acid sequence encoding a viral 2A sequence can be the same as or different from the third nucleic acid sequence encoding a viral 2A sequence.
[0052] As used herein, the term transforming is used broadly to define a method of inserting a vector into a target cell. This can be accomplished, for example, by transfecting the vector into a target cell. Transfecting a vector into a target cell can be accomplished through the use of carriers, which can be divided into three primary classes: (cationic) polymers, liposomes, and nanoparticles. Examples of cationic polymers are DEAE-dextran and polyethylenimine, which bind the negatively charged vector and allows for the vector to be taken up by the cell through endocytosis. Liposomes are small, membrane-bounded bodies that fuse with the cell membrane and allow for the release of the vector into the cell. Nanoparticles are coupled to the vector and are shot directly into the nucleus of a cell using a gene gun. Transfections can further be divided into two categories: stable and transient transfections. Stable transfections result in the vector being permanently introduced into the cell and can be accomplished through the use of selectable marker, e.g., antibiotic resistance, as discussed herein. Transient transfections result in the vector being introduced temporarily to the cell. Alternatively, if the vector is a viral vector, it can be transfected into a host cell to produce virus, and the virus can be harvested and used to transduce the vector into the target cell. Transfection and transduction protocols are known. See, e.g., Sambrook et al., Molecular Cloning: A Laboratory Manual, 3rd Ed., Cold Spring Harbor Press, Cold Spring Harbor, N.Y. (2001); Ausubel et al. (eds.), Current Protocols in Molecular Biology, John Wiley & Sons, Hoboken, N.J. (2004).
[0053] The differentiated cell can, for example, be obtained from a subject. The differentiated cell can be obtained and cultured from the subject by a variety of methods known and described, e.g., in Schantz and Ng, A Manual for Primary Human Cell Culture, World Scientific, Hackensack, N.J. (2004); and Human Cell Culture Protocols 2nd Edition, (Ed. Picot, J), Humana Press, Totowa, N.J. (2004).
[0054] Optionally, the differentiated cell is a mammalian cell. The mammalian cell is optionally a human cell. Mammalian cells suitable for use in the claimed methods, include, but are not limited to epithelial cells, keratinocytes, fibroblasts, hepatocytes, neurons, osteoblasts, myocytes, kidney cells, lung cells, thyroid cells, and pancreatic cells.
[0055] Optionally, the methods further comprise culturing the transformed cell under conditions that allow for the isolation of an iPS cell or a population of iPS cells. For example, transformed cells (e.g., transformed keratinocytes) can be cultured under conditions with relatively high calcium levels. Specifically, prior to transfection, the differentiated cells are cultured under conditions with low calcium levels in the range of 0.01 mM to 0.1 mM. After transformation, the transformed cells are cultured under conditions with high calcium levels in the range of 1.0 mM to 2.0 mM. The high calcium levels promote the death of any untransformed differentiated cells but allow the survival of transformed cells that have undergone the transition to generate iPS cells. Alternatively, the transformed cells can be cultured under conditions that allow for the production of iPS cells through selection based on drug resistance. For example, the transformed vector contains a gene that will provide the transformed cells drug resistance (e.g., blasticidin, zeomycin, hygromycin, or neomycin resistance). Culturing untransformed cells in media supplemented with the selected drug promotes cell death. Culturing the transformed cells in media supplemented with the selected drug allows for the production of iPS cells.
[0056] Also provided are methods of producing iPS cells from differentiated cells comprising transforming the differentiated cells with a first retroviral vector comprising a loxP site in the 3' long terminal repeat of the vector and a nucleic acid sequence comprising a nucleic acid sequence encoding an Oct4, a nucleic acid sequence encoding a Sox2, and a nucleic acid sequence encoding a Klf4 (or any of the nucleic acid sequences described above). The nucleic acid sequences are separated from each other by a first and second nucleic acid sequence encoding a viral 2A sequence. The first nucleic acid sequence encoding a viral 2A sequence can be the same as or different from the second nucleic acid sequence encoding a viral 2A sequence. The method further comprises culturing the transformed cells under conditions that allow for the production of an iPS cell. The method can further comprise transforming the iPS cell with a second vector comprising a nucleic acid sequence encoding a Cre recombinase. Expression of the Cre recombinase results in the deletion of the first vector from the genome of the iPS cell, with the exception of a SIN LTR containing a loxP sequence. Deletion of the first vector from the genome of the iPS cell avoids or reduces the risk of insertional mutagenesis caused by the insertion of the vector into the genome. The method can further comprise isolating a population of the iPS cells lacking the first vector. The iPS cells isolated by this method are physically different from iPS cells produced by other methods, as these iPS cells lack the genomically integrated retroviral vector used to create the iPS cell.
[0057] Also provided are methods of correcting a genetic mutation of a differentiated cell prior to producing an iPS cell from the differentiated cell. The methods comprise transforming a differentiated cell with a vector comprising a nucleic acid sequence comprising (i) a nucleic acid sequence encoding an Oct4, (ii) a nucleic acid sequence encoding a Sox2, and (iii) a nucleic acid sequence encoding a Klf4, wherein each of the nucleic acid sequences, (i)-(iii), are separated by a first and second nucleic acid sequence encoding a viral 2A sequence. The vector further comprises a nucleic acid sequence comprising an unmutated nucleic acid sequence of interest and homologous nucleic acid sequences flanking the genetic mutation. Optionally, the vector further comprises a first and second loxP sequence. Optionally, the vector further comprises a nucleic acid sequence encoding a Cre recombinase operably linked to an inducible promoter. The inducible promoter can, for example, comprise a Nanog-responsive thymidine kinase promoter. Optionally, the vector comprises SEQ ID NO:44.
[0058] Optionally, the genetic mutation is a mutation in the nucleic acid sequence encoding β-globin, the nucleic acid sequence encoding cystic fibrosis transmembrane conductance regulator, the nucleic acid sequence encoding phenylalanine hydroxylase, and/or the nucleic acid sequence encoding dystrophin.
[0059] Optionally, the genetic mutation is a mutation in the nucleic acid sequence encoding β-globin. The mutation in the nucleic acid sequence encoding β-globin can, for example, result in a glutamic acid to valine substitution at the sixth amino acid of the β-globin protein. The glutamic acid to valine substitution can, for example, be caused by an A to T transversion at base pair +20 relative to the A(+1) of the ATG start codon of the nucleic acid sequence encoding β-globin. β-globin is used throughout as an example.
[0060] Further provided are iPS cells produced by these methods. iPS cells produced by these methods can, for example, be identified based on morphological characteristics of the cell (e.g., cell shape, cell composition, cellular organelle shape, and cell size). An iPS cell produced by these methods can be identified based on the expression of ES cell markers. ES cell markers can, for example, include Oct3/4, Nanog, E-Ras, Cripto, Dax1, Sox2, Fgf4, stage-specific embryonic antigen 1 (SSEA1), SSEA3, SSEA4, alkaline phosphatase, tumor-related antigen (TRA)-1-60, TRA-1-81, and Zfp296. Optionally, an iPS cell produced by these methods can be identified by comparing CpG methylation patterns in gene promoters of nontransformed, transformed, and ES cells. Optionally, an iPS cell produced by these methods can be identified based on the ability to form a teratoma comprised of cells derived from the endoderm, mesoderm, and ectoderm in an immunocompromised mouse. An iPS cell can be identified by a combination of cell morphological characteristics, expression of ES cell markers, CpG methylation patterns, and the ability to form a teratoma in an immunocompromised mouse.
[0061] Examples of analytical techniques useful in determining the expression of ES cell markers include reverse transcription-polymerase chain reaction (RT-PCR), quantitative real-time-PCR (qRT-PCR), one step PCR, RNase protection assay, primer extension assay, microarray analysis, gene chip, in situ hybridization, immunohistochemistry, Northern blot, Western blot, enzyme-linked immunosorbent assay (ELISA), enzyme immunoassay (EIA), radioimmunoassay (RIA), or protein array. These techniques are known. See, e.g., Sambrook et al., Molecular Cloning: A Laboratory Manual, 3rd Ed., Cold Spring Harbor Press, Cold Spring Harbor, N.Y. (2001).
[0062] Further provided are kits consisting of any of the first vectors described and a second vector comprising a nucleic acid sequence encoding a Cre recombinase. Optionally, the first vector comprises a nucleic acid sequence comprising a nucleic acid sequence encoding an Oct4, a nucleic acid encoding a Sox2, and a nucleic acid sequence encoding a Klf4. Each of the nucleic acid sequences are separated by a first and second viral 2A sequence. The first viral 2A sequence is the same as or different from the second viral 2A sequence. Optionally, directions to produce an iPS cell from a differentiated cell, a culture plate for producing the iPS cells, and/or containers for the vector or vectors are included in the kit.
[0063] Also provided herein, are methods of treating or preventing a disease or disorder in a subject at risk of developing a disease or disorder. The methods comprise isolating differentiated cells from the subject and transforming the differentiated cells with a first vector comprising a nucleic acid comprising a nucleic acid sequence encoding an Oct4, a nucleic acid sequence encoding a Sox2, and a nucleic acid sequence encoding a Klf4. Each of the nucleic acid sequences are separated by a first and second nucleic acid sequence encoding a viral 2A sequence. The first nucleic acid sequence encoding a viral 2A sequence can be the same as or different from the second nucleic acid sequence encoding a viral 2A sequence. The vector may further comprise a nucleic acid sequence comprising a therapeutic agent. Alternatively, the transformed cells may be transformed with a second vector comprising a nucleic acid sequence comprising a therapeutic agent. The method further comprises isolating a population of the iPS cells. The method further comprises administering to the subject the isolated population of iPS cells that are expressing the therapeutic agent.
[0064] The therapeutic agent can be an RNA molecule, a protein, or a DNA molecule. An RNA molecule can, for example, comprise an antisense RNA molecule, a ribozyme, a small interfering RNA (siRNA) that mediates RNA interference (RNAi), or a microRNA (miRNA) that mediates miRNA-induced translational repression. In the event the therapeutic agent is a protein, the protein can be a receptor, a signaling molecule, a transcription factor, a factor that promotes or inhibits apoptosis, a DNA replication factor, an enzyme, a structural protein, a neural protein, a heat shock protein, or a histone. In the event that the therapeutic agent is a DNA molecule, the DNA molecule can correct a defective or mutated DNA sequence within the genome of the subject. Ordinary skill in the art determines which therapeutic agents are expressed to treat a subject with or at risk of developing a disease or disorder.
[0065] Also provided are methods of treating or preventing a disease associated with a genetic mutation in a subject. The methods comprise selecting a subject with a disease associated with the genetic mutation; isolating differentiated cells from the subject; transforming the differentiated cells with a vector comprising an unmutated nucleic acid sequence of interest; culturing the transformed cells under conditions that allow for the production of a population of iPS cells; screening the iPS cells for correction of the genetic mutation; and administering an effective amount of the iPS cells to the subject. Administration of the iPS cells treats or prevents the disease associated with the genetic mutation in the subject. The vector comprising the unmutated nucleic acid sequence of interest is capable of correcting the genetic mutation associated with the disease and is capable of inducing pluripotent stem (iPS) cells. Optionally, the vector comprises a nucleic acid sequence comprising (i) an unmutated nucleic acid sequence of interest and homologous nucleic acid sequences flanking the genetic mutation, (ii) a nucleic acid sequence encoding a Cre recombinase operably linked to an inducible promoter, (iii) a first and second loxP sequence, (iv) a nucleic acid sequence encoding an Oct4, (v) a nucleic acid sequence encoding a Sox2, and (vi) a nucleic acid sequence encoding a Klf4. Each of the nucleic acid sequences, (iv)-(vi), are separated by a first and second nucleic acid sequence encoding a viral 2A sequence. The first nucleic acid sequence encoding a viral 2A sequence can be the same as or different from the second nucleic acid sequence encoding a viral 2A sequence. Optionally, the inducible promoter comprises a Nanog-responsive thymidine kinase promoter. Optionally, the vector comprises SEQ ID NO:44.
[0066] Examples of analytical techniques useful in screening an iPS cell for correction of the genetic mutation include any DNA-based sequencing assay, reverse transcription-polymerase chain reaction (RT-PCR), quantitative real-time-PCR (qRT-PCR), RNase protection assay, Southern blot, Northern blot, and restriction length polymorphism (RFLP) analysis. These techniques are known. See, e.g., Sambrook et al., Molecular Cloning: A Laboratory Manual, 3rd Ed., Cold Spring Harbor Press, Cold Spring Harbor, N.Y. (2001).
[0067] Optionally, administration of the isolated iPS cells to the subject can be done after the isolated iPS cells have been differentiated to specific types of stem cells (e.g., hematopoietic stem cells). Administration of the differentiated iPS cells to the subject can be done systemically (e.g., injection of iPS cells into the circulatory system) or it can be localized to an organ or tissue (e.g., injection of iPS cells or delivery of stem cells, optionally, on or in a scaffold/matrix to specified organ or tissue). Thus, the administered iPS cells are designed so they interact with the tissue or organ or with target cells. The method of administration is determined by one of skill in the art to be consistent with the treatment of the disease or disorder that the subject has or is at risk of developing.
[0068] Optionally, the differentiated cell is selected from the group consisting of a(n) epithelial cell, keratinocyte, fibroblast, hepatocyte, neuron, osteoblast, myocyte, kidney cell, lung cell, thyroid cell, and pancreatic cell. Optionally, the differentiated cell is a keratinocyte.
[0069] The disease associated with a genetic mutation can, for example, be selected from the group consisting of sickle cell disease, thalassemia, cystic fibrosis, phenylketonuria, and Duchenne muscular dystrophy. The genetic mutation can be corrected via targeted gene replacement and the disease is amenable to a gene/cell therapy approach.
[0070] As used herein, a subject can be a vertebrate, more specifically a mammal (e.g., a human, horse, pig, rabbit, dog, sheep, goat, non-human primate, cow, cat, guinea pig or rodent), a fish, a bird or a reptile or an amphibian. The term does not denote a particular age or sex. Thus, adult and newborn subjects, as well as fetuses, whether male or female, are intended to be covered. As used herein, patient or subject may be used interchangeably and can refer to a subject with or at risk of developing a disease or disorder. The term patient or subject includes human and veterinary subjects.
[0071] A subject at risk of developing a disease or disorder can be genetically predisposed to the disease or condition, e.g., have a mutation in a gene that causes the disease or disorder or have a family history of the disease or disorder. Additionally, a subject at risk of developing a disease or disorder may have symptoms or signs of early onset for the disease or condition. A subject with a disease or disorder has one or more symptoms of the disease or disorder or has been diagnosed with the disease or disorder.
[0072] According to the methods taught herein, the subject is administered an effective amount of the therapeutic agent and/or iPS cells. The terms effective amount and effective dosage are used interchangeably. The term effective amount is defined as any amount necessary to produce a desired physiologic response. Effective amounts and schedules for administering the therapeutic agent and/or iPS cells may be determined empirically, and making such determination is within the skill in the art. The dosage ranges for administration are those large enough to produce the desired effect in which one or more symptoms of the disease or disorder are affected (e.g., reduced or delayed). The dosage should not be so large as to cause substantial adverse side effects, such as unwanted cross-reactions, anaphylactic reactions, and the like. Generally, the dosage will vary with the age, condition, sex, type of disease, the extent of the disease or disorder, route of administration, or whether other drugs are included in the regimen, and can be determined by one or skill in the art. The dosage can be adjusted by the individual physician in the event of any contraindications. Dosages can vary, and can be administered in one or more dose administrations daily, for one or several days. Guidance can be found in the literature for appropriate dosages for given classes of pharmaceutical products.
[0073] As used herein the terms treatment, treat, or treating refer to a method of reducing the effects of a disease or condition or one or more symptoms of the disease or condition. Thus in the disclosed method, treatment can refer to a 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or 100% reduction in the severity of an established disease or condition or one or more symptoms of the disease or condition. For example, a method for treating a disease is considered to be a treatment if there is a 10% reduction in one or more symptoms of the disease in a treated subject as compared to a control. A control can refer to an untreated subject. Alternatively, a control can comprise samples from the subject prior to treatment (i.e., the levels of one or more symptoms of the disease in the subject are determined prior to treatment and compared to the levels of one or more symtpoms of the disease in the subject after treatment). Thus the reduction can be a 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100%, or any percent reduction in between 10% and 100% as compared to native or control levels. It is understood that treatment does not necessarily refer to a cure or complete ablation of the disease, condition, or symptoms of the disease or condition.
[0074] As used herein, the terms prevent, preventing, and prevention of a disease or disorder refers to an action, for example, administration of a therapeutic agent, that occurs before or at about the same time a subject begins to show one or more symptoms of the disease or disorder, wherein the administration inhibits or delays onset or exacerbation of one or more symptoms of the disease or disorder. As used herein, references to decreasing, reducing, or inhibiting include a change of 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90% or greater as compared to a control level. Such terms can include but do not necessarily include complete elimination.
[0075] Disclosed are materials, compositions, and components that can be used for, can be used in conjunction with, can be used in preparation for, or are products of the disclosed methods and compositions. These and other materials are disclosed herein, and it is understood that when combinations, subsets, interactions, groups, etc. of these materials are disclosed that while specific reference of each various individual and collective combinations and permutation of these compounds may not be explicitly disclosed, each is specifically contemplated and described herein. For example, if a method is disclosed and discussed and a number of modifications that can be made to a number of molecules including the method are discussed, each and every combination and permutation of the method, and the modifications that are possible are specifically contemplated unless specifically indicated to the contrary. Likewise, any subset or combination of these is also specifically contemplated and disclosed. This concept applies to all aspects of this disclosure including, but not limited to, steps in methods of using the disclosed compositions. Thus, if there are a variety of additional steps that can be performed it is understood that each of these additional steps can be performed with any specific method steps or combination of method steps of the disclosed methods, and that each such combination or subset of combinations is specifically contemplated and should be considered disclosed.
[0076] Publications cited herein and the material for which they are cited are hereby specifically incorporated by reference in their entireties.
[0077] The examples below are intended to further illustrate certain aspects of the methods and compositions described herein, and are not intended to limit the scope of the claims.
EXAMPLES
General Methods
Production of OSK Polycistronic Lentiviral Vectors
[0078] The complete nucleotide sequence of pKP332 (the OSK polycistronic lentiviral vector) is given by SEQ ID NO:43. The pKP332 vector was deposited with the American Type Culture Collection, 10801 University Boulevard, Manassas, Va. 20110-2209 in accordance with the Budapest Treaty on Oct. 6, 2009, and has accession number PTA-10385. The complete nucleotide and amino acid map of the polycistron encoded by the vector used is given by SEQ ID NO:7 (top strand) and SEQ ID NO:9, respectively (FIG. 7). Construction of the polycistron using PTV1 2A sequences and fusion PCR was performed essentially as described (Holst et al., Nature Protocols 1:406-17 (2006)). Briefly, human Oct4 cDNA (Open Biosystems Clone 40125986) (Open Biosystems; Huntsville, Ala.) was PCR amplified and modified with primers OCT4-F: cacacagcggccgcatttaaatccaccatggcgggacacctggcttc (SEQ ID NO:10) and OCT4-R: agaggacgaacgaaattgtctctcttcaagcaccgaggcaaacttacgtaccctctcgg (SEQ ID NO:11) to contain Not I and Swa I restriction sites at the 5' end and a Kozak consensus sequence. At the 3' end, the Oct4 stop codon was eliminated and replaced with nucleotides (nt) from PTV 1 2A that will form a 22-nt overlap with the 5' end of the Sox2 amplicon. Human Sox2 cDNA (Open Biosystems Clone 2823424) (Open Biosystems; Huntsville, Ala.) was PCR amplified and modified with primers SOX2-F: ctctgttaaagcaagcaggagatgttgaagaaaaccccgggcctatgtacaacatgatggagacgg (SEQ ID NO:12) and SOX2-R: agaggacgaacgaaattgtctctcttcaagcaccgaggcctagggtacacactctccccgtcac (SEQ ID NO:13) to overlap with the 3' end of the Oct4 amplicon and to append 2A nt sequences upstream of the Sox2 ATG. At the 3' end, the Sox2 stop codon was eliminated and replaced with nt from PTV 1 2A that will form a 22-nt overlap with the 5' end of the Klf4 amplicon. Human Klf4 cDNA (Open Biosystems Clone 5111134) (Open Biosystems; Huntsville, Ala.) was PCR amplified and modified with primers KLF4-F: ctctgttaaagcaagcaggagatgttgaagaaaaccccgggcctatggctgtcagcgacgcgc (SEQ ID NO:14) and KLF4-R: gtgtgtcagctgtaaatttaaatttttacggagaagtacacatt (SEQ ID NO:15) to overlap with the 3' end of the Sox2 amplicon and to append 2A nt sequences upstream of the Klf4 ATG. At the 3' end, the Klf4 stop codon was retained and Swa I and Sal I restriction sites were added. After PCR, the individual amplicons were gel purified and used in a three-element fusion PCR at a 1:100:1 (Oct4:Sox2:Klf4) molar ratio along with primers OCT4-F (SEQ ID NO:10) and KLF4-R (SEQ ID NO:15) to produce a 3623 base pair (bp) amplicon containing the polycistron. The polycistron was gel purified and cloned into the general cloning vector pKP114 using the NotI and SalI restriction sites to produce pKP330 and sequenced for authenticity. Subsequently, the polycistron was removed from pKP330 as a Swa I (Roche; Indianapolis, Ind.) fragment and subcloned into a Swa I site downstream of the EF1α promoter in the lentiviral vector pDL 171 (Levasseur et al., Blood 102:4312-9 (2003)) to produce the OSK polycistronic lentiviral vector pKP332, which was sequenced for authenticity.
[0079] By the same strategy, a second polycistronic lentival vector, pKP333, was produced that substitutes the PTV1 2A peptide between Sox2 and Klf4 with the Thosea asigna virus 18 amino acid 2A-like sequence and a GSG linker (underlined): GSGEGRGSLLT CGDVEENPGP (SEQ ID NO:5).
[0080] The complete nucleotide sequence of pKP360 (the OSK polycistronic lentiviral vector designed to correct β-globin mutation) is given by SEQ ID NO:44. To create this vector, a 6938 base pair (bp) loxP-SalI-NBS-TK-Cre/GFP-EF1a-OCT4-2A-SOX2-2A-KLF4-AscI-loxP DNA fragment is inserted into the second intron of the human β-globin gene contained within a bacterial artificial chromosome (BAC) by recombineering in DY380 E. coli cells. In a second recombineering step, a capture vector containing an MC1-driven herpes simplex virus thymidine kinase (HSV tk) gene is used to extract a 16,890 bp sequence from the BAC. The captured sequence consists of 5602 bp of human β-globin 5' homology, the 6938 bp insert sequence, and 4350 bp of human β-globin 3' homology. The first and second β-globin exons are contained within the 5' homology and the third exon is contained within the 3' homology. pKP360 contains a unique NotI restriction site at nucleotide #21049 for vector linearization prior to transfection. The HSV tk gene is used as a negative selection marker for random integration of the vector. Briefly, following transfection with pKP360 of differentiated cells isolated from a sickle cell disease (SCD) patient, 3 classes of cells results: (1) cells that do not receive the vector; these cells remain differentiated and eventually die in culture due to a limited replicative life span; (2) cells that integrate the vector in a non-targeted location; these cells could become iPS cells but will be selected against by gancyclovir because they contain the HSV tk gene; and (3) cells that integrate the vector by homologous recombination into the β-globin locus; these cells have lost the HSV tk marker and will therefore survive gancyclovir selection to become iPS cells with a corrected β-globin gene.
[0081] PCR reactions were performed using PrimeStar polymerase (Takara Bio Inc.; Otsu, Shiga, Japan). All of the oligos used in this study were synthesized by Integrated DNA Technologies (IDT; Coralville, Iowa) and all DNA gel extractions were performed using QIAquick Gel Extraction Kits (Qiagen; Valencia, Calif.).
Cell Culture and Viral Infections
[0082] Embryonic stem (ES) and induced pluripotent stem (iPS) cells were cultured on irradiated murine embryonic fibroblasts (MEFs) in ES cell media consisting of DMEM supplemented with 1× non-essential amino acids, 1× penicillin-streptomycin, 1× L-glutamine (Mediatech; Manassas, Va.), 1× nucleosides (Chemicon; Temecula, Calif.), 15% Fetal Bovine Serum (FBS) (Hyclone; Logan, Utah), 2-ME (Sigma; St. Louis, Mo.) and Leukemia Inhibitory Factor (LIF) (laboratory preparation).
[0083] For preparation of lentivirus, 140 μg of the polycistronic vector (pKP332), 70 μg of the envelope plasmid (pMDG), and 105 μg of the packaging plasmid (pCMBVdR8.9.1) were co-transfected into 1.7×107 293T cells by the CaCl2 method as previously described (Levasseur et al., Blood 102:4312-9 (2003)). Virus-containing supernatant was collected 2 days after transfection, passed through a 0.45 μm filter and concentrated by centrifugation at 26,000 rpm for 90 minutes at 8° C. in an SW-28 rotor using a Beckman XL-100 ultracentrifuge (Beckman; Fullerton, Calif.).
[0084] For iPS cell induction, 3×105 mouse tail-tip fibroblasts (TTFs) were seeded onto one well of a 6-well plate. The next day, 2.5 μL of the concentrated virus was mixed with 2 mL of ES cell medium containing 8 μg/mL polybrene and added to the TTFs. Forty-eight hours later, the TTFs were trypsinized and transferred to a 100 mm dish without MEFs and continuously cultured on the same dish for 3 weeks with daily media changes. Potential iPS cell colonies started to appear after 2-3 weeks. These colonies were individually picked and expanded on MEFs for analysis.
[0085] To remove the integrated lentiviral and polycistronic sequences, iPS cells were either electroporated with a Cre-expressing plasmid (pCAGGS-Cre) or infected with a Cre-expressing adenovirus (rAd-Cre-IE). Individual colonies were picked and Cre-mediated removal of floxed sequences was verified by PCR and southern blot analysis.
[0086] For the construction of rAd-Cre-IE (rAd-Cre-IRES-EGFP), Cre cDNA was PCR amplified from pCAGGS-Cre and inserted between the NheI and EcoRI sites of the expression vector pEC-IE, which contains an IRES-EGFP downstream of the MCS. The Cre-IE expression cassette is flanked by attL1 and attL2 sites, thus allowing transfer of the Cre-IE sequence from pEC-IE to pAd/p1-DEST (Invitrogen; Carlsbad, Calif.) by the LR reaction. The recombinant adenovirus was packaged in 293A cells according to the manufacturer's instructions.
[0087] Primary human keratinocytes were isolated from a patient skin biopsy. Briefly, the biopsied tissue was placed into Keratinocyte-SFM (9K-SFM; Invitrogen; Carlsbad, Calif.) supplemented with 10 mg/ml Dispase and 2× Antibiotics/Antimycotics (CELLnTEC CnT-ABM) and incubated overnight at 4° C. The next day, the keratinocyte-containing epidermal layer was isolated from the fibroblast-containing dermal layer with forceps and then trypsinized for 20 minutes at room temperature. Cell clumps were triturated with a pipet and then centrifuged at 200×g for 5 minutes. Cells were resuspended in K-SFM and 1× Antibiotics/Antimycotics, transferred to one well of a six-well plate, and incubated at 37° C. with daily media changes. For transduction, 3×105 keratinocytes were seeded into one well of a six-well plate in K-SFM. The next day the media was removed and replaced with 2 ml of K-SFM containing 5 mg/ml of polybrene and the polycistronic lentivirus. After 24 hours, the transduced cells were trypsinized, centrifuged, resuspended in K-SFM and transferred into a 10 cm tissue culture dish containing γ-irradiated CF-1 murine embryonic fibroblasts (MEFs). The next day, the medium was changed to human ES cell medium (DMEM/F-12, 20% Knockout SR, 2 mM L-glutamine, 1× Pen/Strep, 1× nonessential amino acids (all from Invitrogen; Carlsbad, Calif.), 0.5 mM β-mercaptoethanol (Sigma; St. Louis, Mo.), and 4 ng/ml bFGF (Calbiochem; San Diego, Calif.)). Cells were incubated at 37° C. with daily media changes and after 10 days, CF-1 conditioned medium was added. iPS colonies appeared after about 30 days.
[0088] With the exception of the pKP332 construction, all of the PCRs performed used ExTaq polymerase (Takara Bio Inc.; Otsu, Shiga, Japan). All of the sequencing was performed by the Genomics Core Facility of the Howell and Elizabeth Heflin Center for Human Genetics of the University of Alabama at Birmingham using the BigDye Terminator v3.1 Cycle Sequencing Ready Reaction kit as per the manufacture's instructions (Applied Biosystems; Foster City, Calif.). The sequencing products were run following standard protocols on an Applied Biosystems 3730 Genetic Analyzer with POP-7 polymer.
Immunostaining and AP Staining
[0089] iPS cells were cultured on cover slips pretreated with FBS, fixed with 4% paraformaldehyde and permeabilized with 0.5% Triton X-100. Cells were stained with DAPI and primary antibodies against Nanog and SSEA1 (R&D Systems; Minneapolis, Minn.) and incubated with fluorophore-labeled secondary antibodies (Jackson Immunoresearch; West Grove, Pa.).
[0090] For AP staining, 100-200 iPS cells were seeded onto one well of a six-well plate and cultured for one week. iPS cells were then stained using the Vector Blue Alkaline Phosphatase Substrate Kit III (Vector Laboratories; Burlingame, Calif.) according to the manufacturer's instructions.
RT-PCR Analysis
[0091] Total RNA was isolated from cells with Trizol reagent (Invitrogen; Carlsbad, Calif.). RNA was pretreated with RQ1 RNase-free DNase (Promega; Madison, Wis.) and reverse transcribed with SuperScript First-Strand Synthesis System (Invitrogen; Carlsbad, Calif.) using oligo d(T)n. Primers for PCR amplification of the cDNA were: polycistronic transgene F, gatgaactgaccaggcacta (SEQ ID NO:16) and polycistronic transgene R, gattatcggaattccctcgag (SEQ ID NO:17); Nanog F, accaaaggatgaagtgcaag (SEQ ID NO:18) and Nanog R, agttttgctgcaactgtacg (SEQ ID NO:19); Oct4 F, agcttgggctagagaaggat (SEQ ID NO:20) and Oct4 R, tcagtttgaatgcatgggag (SEQ ID NO:21); Sox2 F, tgcacatggcccagcacta (SEQ ID NO:22) and Sox2 R, ttctccagttcgcagtccag (SEQ ID NO:23); Cripto F, aacttgctgtctgaatggag (SEQ ID NO:24) and Cripto R, tttgaggtcctggtccatca (SEQ ID NO:25); Klf4 F, cagcagggactgtcaccctg (SEQ ID NO:26) and Klf4 R, ggtcacatccactacgtgggat (SEQ ID NO:27); and Natl F, ggagagtgcgattgcagaag (SEQ ID NO:28) and Natl R, ggtcacatccactacgtggga (SEQ ID NO:29).
Bisulfite Modification and Sequencing
[0092] Bisulfite treatment of DNA was performed with the CpGenome Fast DNA Modification Kit (Chemicon; Temecula, Calif.) according to the manufacturer's instructions. The Oct4 and Nanog gene promoter regions were amplified by nested PCR using the Oct4 primers F1, gttgttttgttttggttttggatat (SEQ ID NO:30), Oct4 F2, atgggttgaaatattgggtttattta (SEQ ID NO:31) and Oct4 R, ccaccctctaaccttaacctctaac (SEQ ID NO:32) or the Nanog primers F1, gaggatgttttttaagtttttttt (SEQ ID NO:33), Nanog F2, aatgtttatggtggattttgtaggt (SEQ ID NO:34) and Nanog R, cccacactcatatcaatataataac (SEQ ID NO:35). Amplified PCR products were purified using a QIAgen Gel Extraction Kit (Qiagen; Valencia, Calif.), cloned into a Topo TA vector (Invitrogen; Carlsbad, Calif.), and sequenced with T7 and Ml3R primers.
Southern Blot Analysis
[0093] Ten μg of genomic DNA were digested with BamHI or KpnI (Roche; Indianapolis, Ind.), separated on a 0.8% agarose gel and blotted onto Hybond-N.sup.+ membrane (Amersham Biosciences; Piscataway, N.J.). The polycistronic vector served as template to PCR amplify a 0.3 kb SIN LTR probe using the primers SIN LTR F, gctcggtacctttaagaccaatgac (SEQ ID NO:36) and SIN LTR R, atgctgctagagattttccacactg (SEQ ID NO:37). To produce the internal probe, the polycistronic vector was digested with SalI and XhoI (Roche; Indianapolis, Ind.) and the 1 kb fragment containing the EF1α promoter was gel purified. Probes were labeled using the Random Primed DNA Labeling Kit (Roche; Indianapolis, Ind.) with 32P-α-dCTP and blots were hybridized in MiracleHyb solution (Stratagene; La Jolla, Calif.).
Inverse PCR
[0094] One to two μg of total genomic DNA were digested with the tetranucleotide-recognizing restriction enzymes MseI or AluI (New England Biolabs (NEB); Ipswich, Mass.). The digested fragments were diluted and incubated with T4 DNA Ligase (Roche; Indianapolis, Ind.) to obtain self-ligated monomers, which were then linearized with the hexanucleotide-recognizing restriction enzymes NcoI or XmnI (NEB; Ipswich, Mass.). These fragments were isolated by ethanol precipitation and used as templates in PCR reactions using the primers 5LentiR1, tgaattgatcccatcttgtcttcg (SEQ ID NO:38) and SLentiF1, tgctgctttttgcttgtactgg (SEQ ID NO:39). PCR products were run on a 2% agarose gel in the presence of ethidium bromide (0.5 μg/mL). All bands visible under UV light were gel purified and sequenced.
Teratoma Formation
[0095] One million iPS cells in a 100 μL volume of PBS were injected via a 21 G needle into the dorsal flanks of SCID mice. Teratomas were recovered 4-5 weeks postinjection and processed for histological analysis.
Production and Analysis of Chimeric Mice
[0096] C57BL/6 blastocysts were injected with iPS cells and then transferred to pseudopregnant CD-1 females. After two weeks, embryos were collected for photographs and analyzed for chimerism using PCR. Embryos were individually minced and lysed overnight at 55° C. in a solution of Proteinase K and SDS. DNA was then purified from the lysate by phenol/chloroform extraction and ethanol precipitation. PCR was performed using the primers mbeta KI F, ttgagcaatgtggacagagaagg (SEQ ID NO:40), mbeta KI R, gtcagaagcaaatgtgaggagca (SEQ ID NO:41) and 1400gamma R, aattctggcttatcggaggcaag (SEQ ID NO:42).
Example 1
iPS Cells Produced by Transduction of Polycistronic Oct4, Sox2, Klf4 (OSK) Vector
[0097] FIG. 1A illustrates the lentiviral vector constructed for transduction of adult skin fibroblasts. Human Oct4, Sox2 and Klf4 cDNAs (OSK) were linked with porcine teschovirus-1 (PTV1) 2A sequences that function as cis-acting hydrolase elements (CHYSELs) to trigger ribosome skipping (Donnelly et al., J. Gen. Virol. 82:1013-25 (2001); Chinnasamy et al., Virol. J. 3:14 (2006)). The 2A peptide sequences (FIG. 1B) are cleaved during translation and produce Oct4 and Sox2 proteins containing an additional 21 amino acids at the carboxy-termini. A single proline is also appended to the amino-termini of Sox2 and Klf4. The OSK polycistron was subcloned downstream of an EF1α promoter in a self-inactivating (SIN) lentiviral vector containing a loxP site in the truncated 3' LTR (Zuffferey et al., J. Virol. 72:9873-80 (1998); Levasseur et al., Blood 102:4312-9 (2003)). After lentivirus production, one million adult skin fibroblasts derived from tail tips of humanized sickle mice were transduced with the polycistronic vector, and four colonies with highly defined borders and tightly packed cells were picked at 19 to 30 days post-transduction. These colonies were expanded and stained for alkaline phosphatase, Nanog and SSEA1, which are characteristic markers of pluripotent stem cells. FIGS. 2A and 2B illustrate the staining pattern of typical colonies (iPS-1 and iPS-2). The colonies stained intensely for alkaline phosphatase and strongly with antibodies to Nanog and SSEA1.
[0098] Reverse transcription-polymerase chain reaction (RT-PCR) assays for expression of additional iPS cell markers are shown in FIG. 3A. iPS-1, -2, and -3 cells expressed polycistronic OSK RNA and endogenous Oct4, Sox2, Klf4, Nanog and Cripto RNA (FIG. 3A). Consistent with these results, bisulfite sequencing of the endogenous Oct4 and Nanog promoters in iPS-1 and iPS-2 cells demonstrated effective demethylation of these sequences (FIG. 3B). CpGs in the endogenous Oct4 and Nanog promoters of tail tip fibroblasts (TTFs) were highly methylated (FIG. 3B) and endogenous Oct4, Sox2, Nanog and Cripto RNAs were not detected (FIG. 3A).
[0099] When these iPS cells were injected into the dorsal flanks of nonobese diabetic (NOD)/SCID IL-2 γR-/-mice, teratomas containing tissue derived from all three germ layers were obtained (FIG. 5A). These results demonstrate that the polycistronic OSK lentiviral vector effectively reprograms adult skin fibroblasts to induced pluripotent stem cells.
Example 2
Removal of Polycistronic OSK Vector from iPS Cell Genome by Exogenous Cre Recombinase Expression
[0100] The polycistronic vector was deleted by electroporation of iPS cells with a Cre recombinase-expressing plasmid or by infection of iPS cells with adenovirus that expresses Cre recombinase (Adeno/Cre). Subsequently, individual colonies were picked, expanded and iPS cell DNA was analyzed by Southern blot hybridization (FIG. 4B). DNA isolated before (iPS-1) and after (iPS-1 Cre) Cre expression was digested with Kpn I, which cuts once within the OSK polycistron, and probed with a DNA fragment containing EF1α sequences. Four bands are observed for iPS-1 DNA indicating that four copies of the polycistronic OSK vector are integrated into the genome (also see FIG. 6B, iPS-2 cells contain 3 copies of the vector). None of these four bands are observed in iPS-1 Cre DNA; only a band representing endogenous EF1α sequences is detected. These results demonstrate that transient Cre expression effectively deletes all copies of the polycistronic OSK lentiviral vector.
[0101] Junctions of the four iPS-1 insertion sites were cloned by inverse PCR and sequenced (Pawlik et al., Gene 165:173-81 (1995); Silver and Keerikatte, J. Virol. 63:1924-8 (1989)). Table 2 lists the locations of these sites. Three of the insertion sites are within introns, and one is located in an intergenic region that is 2 megabases (Mb) downstream of the transcription start site (TSS) of the NMBr gene and 1 Mb upstream of the TSS of the Cited2 gene. These results demonstrate that iPS cells can be readily obtained by this procedure without interruption of coding sequences, promoters or known regulatory elements. Cloning and sequencing of the insertion sites from iPS-1 Cre cells demonstrated that only the 291 base pair (bp) 3' LTR of the polycistronic vector remains in the genome. This small SIN LTR does not contain a promoter or enhancer; therefore, the probability of insertional activation or inactivation of endogenous genes is low.
TABLE-US-00002 TABLE 2 OSK lentiviral integration sites. iPS Clones No: Chrom. Gene Name Gene ID Location Base from TSS iPS-1 1 CH2 RAB14 MGI:1915615 Intron +8,129 2 CH8 Cadherin 13 MGI:99551 Intron +24,738 3 CH10 Cbp/p300-interacting MGI:1306784 Intergenic -966,513 transactivator 4 CH14 F-box protein 34 MGI:1926188 Intron +52,366 iPS-2 1 CH5 Ribokinase MGI:1918586 Intron +38,503 2 CH15 Estrogen receptor-binding MGI:1859920 Intron +20,439 fragment associated gene 9 3 CH15 Angiopoietin 1 MGI:108448 Intron +21,069
[0102] FIGS. 2A and 2B demonstrate that iPS-1 Cre cells continue to stain positive for alkaline phosphatase, Nanog and SSEA1 after OSK deletion, and FIG. 3A demonstrates that expression of endogenous Oct4, Sox2, Klf4, Nanog and Cripto was maintained in the absence of OSK expression. As expected, the endogenous Oct4 and Nanog promoters remained demethylated after OSK deletion (FIG. 3B).
[0103] Finally, two iPS-1 Cre cell lines were injected into wild-type blastocysts, and these blastocysts were transferred into the uteri of pseudo-pregnant female mice. After two weeks, embryos were analyzed for chimerism by PCR with primers specific for human and mouse β-globin genes. FIG. 5B demonstrates that several high-level chimeras were obtained; most tissues of these embryos were derived from iPS-1 Cre cells which contain only human β-globin genes. One pregnancy was allowed to proceed to term, and FIG. 5C shows an adult high-level chimera (right) derived from iPS-1 Cre 2 cells. These results demonstrate that adult skin fibroblasts can be effectively reprogrammed to iPS cells with the polycistronic lentiviral vector and that tissues from all three germ layers can be derived from these cells.
Example 3
iPS Cells Derived from Human Keratinocytes
[0104] To determine whether iPS cells were produced from primary human keratinocytes, primary human keratinocytes were cultured from a patient skin biopsy. The cultured cells were transduced with the vector described above. After 24 hours, the transduced cells were trypsinized, centrifuged, resuspended in media and transferred into a tissue culture dish containing murine embryonic fibroblasts (MEFs). After about 30 days in culture, iPS colonies were produced. The iPS cells from the human keratinocytes were sustainable in culture and were capable of multiple passages. FIG. 8 shows a brightfield image of one of the iPS cell colonies produced. The iPS cell colony was stained with SSEA-4, which is an antibody that recognizes human embryonic stem cells, but not differentiated cells, to confirm the presence of embryonic stem cells comprising the iPS cell colony. The same iPS colony was stained with DAPI, which is a general nuclear stain, to confirm the presence of nuclei in the cells of the iPS cell colony.
Example 4
Correction of Sickle Cell Disease (SCD) with Concomitant Formation of iPS Cells
[0105] FIG. 9 shows a schematic of a method to correct a βs-globin mutation in a cell from a subject with sickle cell disease (SCD) while dedifferentiating the cell to a pluripotent state. The method is applicable to a range of genetic mutations.
[0106] To determine whether the β-globin locus of a subject with SCD is corrected, cells from a human subject with SCD are collected and expanded in culture. The mutated βs-globin locus is depicted at the top of FIG. 9. The βs-globin mutation is a single nucleotide, A to T transversion, that changes the normal GAG codon to a GTG codon in exon 1 of β-globin. As a result, the sixth amino acid of the βs-globin is a valine instead of the normal glutamic acid.
[0107] Once the cells are expanded in culture, the targeting vector (middle of FIG. 9) is introduced into the cells from the subject with SCD. The vector contains the normal GAG nucleotide sequence in the first exon and flanking sequences to effect homologous recombination within the target locus. A herpes simplex virus thymidine kinase (HSV tk) gene is located outside of the sequences used to effect homologous recombination. Integrated between the flanking homology arms is a floxed cassette consisting of a Nanog-responsive thymidine kinase promoter driving expression of a Cre recombinase and the EF1α promoter driving expression of the Oct4-Sox2-Klf4 polycistronic sequence. Alternatively, the floxed cassette can contain a marker gene that can either be an addition to the polycistron or have its own promoter. The marker can be used as a positive selection to select cells that have incorporated the vector.
[0108] The targeting vector homologously recombines with the mutated βs-globin locus incorporating the corrected GAG codon. The Oct4-Sox2-Klf4 polycistron is expressed, resulting in the dedifferentiation of the cells. While Oct4, Sox2, and Klf4 are expressed from the EF1α promoter, the TK promoter remains silent. Once the cell begins to dedifferentiate, the endogenous Nanog gene is expressed. Expression of Nanog results in the activation of the TK promoter, which is Nanog responsive. Activation of the TK promoter results in the expression of Cre recombinase. Cre recombinase binds to the loxP sites to effect the deletion of the floxed cassette, resulting in a corrected β-globin locus containing a single loxP site in between the second and third exons of the corrected β-globin locus (bottom of FIG. 9). Excision of the floxed cassette is important for two reasons: (1) it prevents the disregulation of the corrected β-globin gene, and (2) it halts the expression of the vector-introduced reprogramming factors, as their continued expression inhibits the reprogramming process.
Sequence CWU
1
1
44122PRTArtificial sequenceSynthetic construct 1Gly Ser Gly Ala Thr Asn
Phe Ser Leu Leu Lys Gln Ala Gly Asp Val 1 5
10 15 Glu Glu Asn Pro Gly Pro 20
219PRTArtificial sequenceSynthetic construct 2Ala Thr Asn Phe Ser Leu
Leu Lys Gln Ala Gly Asp Val Glu Glu Asn 1 5
10 15 Pro Gly Pro 318PRTArtificial
sequenceSynthetic construct 3Glu Gly Arg Gly Ser Leu Leu Thr Cys Gly Asp
Val Glu Glu Asn Pro 1 5 10
15 Gly Pro 43PRTArtificial sequenceSynthetic construct 4Gly Ser
Gly 1 521PRTArtificial sequenceSynthetic construct 5Gly Ser Gly
Glu Gly Arg Gly Ser Leu Leu Thr Cys Gly Asp Val Glu 1 5
10 15 Glu Asn Pro Gly Pro
20 634DNAArtificial sequenceSynthetic construct 6ataacttcgt
ataatgtatg ctatacgaag ttat
3473623DNAArtificial sequenceSynthetic construct 7cacacagcgg ccgcatttaa
atccaccatg gcgggacacc tggcttcgga tttcgccttc 60tcgccccctc caggtggtgg
aggtgatggg ccaggggggc cggagccggg ctgggttgat 120cctcggacct ggctaagctt
ccaaggccct cctggagggc caggaatcgg gccgggggtt 180gggccaggct ctgaggtgtg
ggggattccc ccatgccccc cgccgtatga gttctgtggg 240gggatggcgt actgtgggcc
ccaggttgga gtggggctag tgccccaagg cggcttggag 300acctctcagc ctgagggcga
agcaggagtc ggggtggaga gcaactccga tggggcctcc 360ccggagccct gcaccgtcac
ccctggtgcc gtgaagctgg agaaggagaa gctggagcaa 420aacccggagg agtcccagga
catcaaagct ctgcagaaag aactcgagca atttgccaag 480ctcctgaagc agaagaggat
caccctggga tatacacagg ccgatgtggg gctcaccctg 540ggggttctat ttgggaaggt
attcagccaa acgaccatct gccgctttga ggctctgcag 600cttagcttca agaacatgtg
taagctgcgg cccttgctgc agaagtgggt ggaggaagct 660gacaacaatg aaaatcttca
ggagatatgc aaagcagaaa ccctcgtgca ggcccgaaag 720agaaagcgaa ccagtatcga
gaaccgagtg agaggcaacc tggagaattt gttcctgcag 780tgcccgaaac ccacactgca
gcagatcagc cacatcgccc agcagcttgg gctcgagaag 840gatgtggtcc gagtgtggtt
ctgtaaccgg cgccagaagg gcaagcgatc aagcagcgac 900tatgcacaac gagaggattt
tgaggctgct gggtctcctt tctcaggggg accagtgtcc 960tttcctctgg ccccagggcc
ccattttggt accccaggct atgggagccc tcacttcact 1020gcactgtact cctcggtccc
tttccctgag ggggaagcct ttccccctgt ctccgtcacc 1080actctgggct ctcccatgca
ttcaaacgga tccggagcca cgaacttctc tctgttaaag 1140caagcaggag atgttgaaga
aaaccccggg cctatgtaca acatgatgga gacggagctg 1200aagccgccgg gcccgcagca
aacttcgggg ggcggcggcg gcaactccac cgcggcggcg 1260gccggcggca accagaaaaa
cagcccggac cgcgtcaagc ggcccatgaa tgccttcatg 1320gtgtggtccc gcgggcagcg
gcgcaagatg gcccaggaga accccaagat gcacaactcg 1380gagatcagca agcgcctggg
cgccgagtgg aaacttttgt cggagacgga gaagcggccg 1440ttcatcgacg aggctaagcg
gctgcgagcg ctgcacatga aggagcaccc ggattataaa 1500taccggcccc ggcggaaaac
caagacgctc atgaagaagg ataagtacac gctgcccggc 1560gggctgctgg cccccggcgg
caatagcatg gcgagcgggg tcggggtggg cgccggcctg 1620ggcgcgggcg tgaaccagcg
catggacagt tacgcgcaca tgaacggctg gagcaacggc 1680agctacagca tgatgcagga
ccagctgggc tacccgcagc acccgggcct caatgcgcac 1740ggcgcagcgc agatgcagcc
catgcaccgc tacgacgtga gcgccctgca gtacaactcc 1800atgaccagct cgcagaccta
catgaacggc tcgcccacct acagcatgtc ctactcgcag 1860cagggcaccc ctggcatggc
tcttggctcc atgggttcgg tggtcaagtc cgaggccagc 1920tccagccccc ctgtggttac
ctcttcctcc cactccaggg cgccctgcca ggccggggac 1980ctccgggaca tgatcagcat
gtatctcccc ggcgccgagg tgccggaacc cgccgccccc 2040agcagacttc acatgtccca
gcactaccag agcggcccgg tgcccggcac ggccattaac 2100ggcacactgc ccctctcaca
catgggatcc ggagccacga acttctctct gttaaagcaa 2160gcaggagatg ttgaagaaaa
ccccgggcct atggctgtca gcgacgcgct gctcccatct 2220ttctccacgt tcgcgtctgg
cccggcggga agggagaaga cactgcgtca agcaggtgcc 2280ccgaataacc gctggcggga
ggagctctcc cacatgaagc gacttccccc agtgcttccc 2340ggccgcccct atgacctggc
ggcggcgacc gtggccacag acctggagag cggcggagcc 2400ggtgcggctt gcggcggtag
caacctggcg cccctacctc ggagagagac cgaggagttc 2460aacgatctcc tggacctgga
ctttattctc tccaattcgc tgacccatcc tccggagtca 2520gtggccgcca ccgtgtcctc
gtcagcgtca gcctcctctt cgtcgtcgcc gtcgagcagc 2580ggccctgcca gcgcgccctc
cacctgcagc ttcacctatc cgatccgggc cgggaacgac 2640ccgggcgtgg cgccgggcgg
cacgggcgga ggcctcctct atggcaggga gtccgctccc 2700cctccgacgg ctcccttcaa
cctggcggac atcaacgacg tgagcccctc gggcggcttc 2760gtggccgagc tcctgcggcc
agaattggac ccggtgtaca ttccgccgca gcagccgcag 2820ccgccaggtg gcgggctgat
gggcaagttc gtgctgaagg cgtcgctgag cgcccctggc 2880agcgagtacg gcagcccgtc
ggtcatcagc gtcagcaaag gcagccctga cggcagccac 2940ccggtggtgg tggcgcccta
caacggcggg ccgccgcgca cgtgccccaa gatcaagcag 3000gaggcggtct cttcgtgcac
ccacttgggc gctggacccc ctctcagcaa tggccaccgg 3060ccggctgcac acgacttccc
cctggggcgg cagctcccca gcaggactac cccgaccctg 3120ggtcttgagg aagtgctgag
cagcagggac tgtcaccctg ccctgccgct tcctcccggc 3180ttccatcccc acccggggcc
caattaccca tccttcctgc ccgatcagat gcagccgcaa 3240gtcccgccgc tccattacca
agagctcatg ccacccggtt cctgcatgcc agaggagccc 3300aagccaaaga ggggaagacg
atcgtggccc cggaaaagga ccgccaccca cacttgtgat 3360tacgcgggct gcggcaaaac
ctacacaaag agttcccatc tcaaggcaca cctgcgaacc 3420cacacaggtg agaaacctta
ccactgtgac tgggacggct gtggatggaa attcgcccgc 3480tcagatgaac tgaccaggca
ctaccgtaaa cacacggggc accgcccgtt ccagtgccaa 3540aaatgcgacc gagcattttc
caggtcggac cacctcgcct tacacatgaa gaggcatttt 3600taaatttaaa tgtcgactgt
gtg 362383623DNAArtificial
sequencesynthetic construct 8gtgtgtcgcc ggcgtaaatt taggacctac cgccctgtgg
accgaagcct aaagcggaag 60agcgggggag gtccaccacc tccactaccc ggtccccccg
gcctcggccc gacccaacta 120ggagcctgga ccgattcgaa ggttccggga ggacctcccg
gtccttagcc cggcccccaa 180cccggtccga gactccacac cccctaaggg ggtacggggg
gcggcatact caagacaccc 240ccctaccgca tgacacccgg ggtccaacct caccccgatc
acggggttcc gccgaacctc 300tggagagtcg gactcccgct tcgtcctcag ccccacctct
cgttgaggct accccggagg 360ggcctcggga cgtggcagtg gggaccacgg cacttcgacc
tcttcctctt cgacctcgtt 420ttgggcctcc tcagggtcct gtagtttcga gacgtctttc
ttgagctcgt taaacggttc 480gaggacttcg tcttctccta gtgggaccct atatgtgtcc
ggctacaccc cgagtgggac 540ccccaagata aacccttcca taagtcggtt tgctggtaga
cggcgaaact ccgagacgtc 600gaatcgaagt tcttgtacac attcgacgcc gggaacgacg
tcttcaccca cctccttcga 660ctgttgttac ttttagaagt cctctatacg tttcgtcttt
gggagcacgt ccgggctttc 720tctttcgctt ggtcatagct cttggctcac tctccgttgg
acctcttaaa caaggacgtc 780acgggctttg ggtgtgacgt cgtctagtcg gtgtagcggg
tcgtcgaacc cgagctcttc 840ctacaccagg ctcacaccaa gacattggcc gcggtcttcc
cgttcgctag ttcgtcgctg 900atacgtgttg ctctcctaaa actccgacga cccagaggaa
agagtccccc tggtcacagg 960aaaggagacc ggggtcccgg ggtaaaacca tggggtccga
taccctcggg agtgaagtga 1020cgtgacatga ggagccaggg aaagggactc ccccttcgga
aagggggaca gaggcagtgg 1080tgagacccga gagggtacgt aagtttgcct aggcctcggt
gcttgaagag agacaatttc 1140gttcgtcctc tacaacttct tttggggccc ggatacatgt
tgtactacct ctgcctcgac 1200ttcggcggcc cgggcgtcgt ttgaagcccc ccgccgccgc
cgttgaggtg gcgccgccgc 1260cggccgccgt tggtcttttt gtcgggcctg gcgcagttcg
ccgggtactt acggaagtac 1320cacaccaggg cgcccgtcgc cgcgttctac cgggtcctct
tggggttcta cgtgttgagc 1380ctctagtcgt tcgcggaccc gcggctcacc tttgaaaaca
gcctctgcct cttcgccggc 1440aagtagctgc tccgattcgc cgacgctcgc gacgtgtact
tcctcgtggg cctaatattt 1500atggccgggg ccgccttttg gttctgcgag tacttcttcc
tattcatgtg cgacgggccg 1560cccgacgacc gggggccgcc gttatcgtac cgctcgcccc
agccccaccc gcggccggac 1620ccgcgcccgc acttggtcgc gtacctgtca atgcgcgtgt
acttgccgac ctcgttgccg 1680tcgatgtcgt actacgtcct ggtcgacccg atgggcgtcg
tgggcccgga gttacgcgtg 1740ccgcgtcgcg tctacgtcgg gtacgtggcg atgctgcact
cgcgggacgt catgttgagg 1800tactggtcga gcgtctggat gtacttgccg agcgggtgga
tgtcgtacag gatgagcgtc 1860gtcccgtggg gaccgtaccg agaaccgagg tacccaagcc
accagttcag gctccggtcg 1920aggtcggggg gacaccaatg gagaaggagg gtgaggtccc
gcgggacggt ccggcccctg 1980gaggccctgt actagtcgta catagagggg ccgcggctcc
acggccttgg gcggcggggg 2040tcgtctgaag tgtacagggt cgtgatggtc tcgccgggcc
acgggccgtg ccggtaattg 2100ccgtgtgacg gggagagtgt gtaccctagg cctcggtgct
tgaagagaga caatttcgtt 2160cgtcctctac aacttctttt ggggcccgga taccgacagt
cgctgcgcga cgagggtaga 2220aagaggtgca agcgcagacc gggccgccct tccctcttct
gtgacgcagt tcgtccacgg 2280ggcttattgg cgaccgccct cctcgagagg gtgtacttcg
ctgaaggggg tcacgaaggg 2340ccggcgggga tactggaccg ccgccgctgg caccggtgtc
tggacctctc gccgcctcgg 2400ccacgccgaa cgccgccatc gttggaccgc ggggatggag
cctctctctg gctcctcaag 2460ttgctagagg acctggacct gaaataagag aggttaagcg
actgggtagg aggcctcagt 2520caccggcggt ggcacaggag cagtcgcagt cggaggagaa
gcagcagcgg cagctcgtcg 2580ccgggacggt cgcgcgggag gtggacgtcg aagtggatag
gctaggcccg gcccttgctg 2640ggcccgcacc gcggcccgcc gtgcccgcct ccggaggaga
taccgtccct caggcgaggg 2700ggaggctgcc gagggaagtt ggaccgcctg tagttgctgc
actcggggag cccgccgaag 2760caccggctcg aggacgccgg tcttaacctg ggccacatgt
aaggcggcgt cgtcggcgtc 2820ggcggtccac cgcccgacta cccgttcaag cacgacttcc
gcagcgactc gcggggaccg 2880tcgctcatgc cgtcgggcag ccagtagtcg cagtcgtttc
cgtcgggact gccgtcggtg 2940ggccaccacc accgcgggat gttgccgccc ggcggcgcgt
gcacggggtt ctagttcgtc 3000ctccgccaga gaagcacgtg ggtgaacccg cgacctgggg
gagagtcgtt accggtggcc 3060ggccgacgtg tgctgaaggg ggaccccgcc gtcgaggggt
cgtcctgatg gggctgggac 3120ccagaactcc ttcacgactc gtcgtccctg acagtgggac
gggacggcga aggagggccg 3180aaggtagggg tgggccccgg gttaatgggt aggaaggacg
ggctagtcta cgtcggcgtt 3240cagggcggcg aggtaatggt tctcgagtac ggtgggccaa
ggacgtacgg tctcctcggg 3300ttcggtttct ccccttctgc tagcaccggg gccttttcct
ggcggtgggt gtgaacacta 3360atgcgcccga cgccgttttg gatgtgtttc tcaagggtag
agttccgtgt ggacgcttgg 3420gtgtgtccac tctttggaat ggtgacactg accctgccga
cacctacctt taagcgggcg 3480agtctacttg actggtccgt gatggcattt gtgtgccccg
tggcgggcaa ggtcacggtt 3540tttacgctgg ctcgtaaaag gtccagcctg gtggagcgga
atgtgtactt ctccgtaaaa 3600atttaaattt acagctgaca cac
362391191PRTArtificial sequenceSynthetic construct
9Met Ala Gly His Leu Ala Ser Asp Phe Ala Phe Ser Pro Pro Pro Gly 1
5 10 15 Gly Gly Gly Asp
Gly Pro Gly Gly Pro Glu Pro Gly Trp Val Asp Pro 20
25 30 Arg Thr Trp Leu Ser Phe Gln Gly Pro
Pro Gly Gly Pro Gly Ile Gly 35 40
45 Pro Gly Val Gly Pro Gly Ser Glu Val Trp Gly Ile Pro Pro
Cys Pro 50 55 60
Pro Pro Tyr Glu Phe Cys Gly Gly Met Ala Tyr Cys Gly Pro Gln Val 65
70 75 80 Gly Val Gly Leu Val
Pro Gln Gly Gly Leu Glu Thr Ser Gln Pro Glu 85
90 95 Gly Glu Ala Gly Val Gly Val Glu Ser Asn
Ser Asp Gly Ala Ser Pro 100 105
110 Glu Pro Cys Thr Val Thr Pro Gly Ala Val Lys Leu Glu Lys Glu
Lys 115 120 125 Leu
Glu Gln Asn Pro Glu Glu Ser Gln Asp Ile Lys Ala Leu Gln Lys 130
135 140 Glu Leu Glu Gln Phe Ala
Lys Leu Leu Lys Gln Lys Arg Ile Thr Leu 145 150
155 160 Gly Tyr Thr Gln Ala Asp Val Gly Leu Thr Leu
Gly Val Leu Phe Gly 165 170
175 Lys Val Phe Ser Gln Thr Thr Ile Cys Arg Phe Glu Ala Leu Gln Leu
180 185 190 Ser Phe
Lys Asn Met Cys Lys Leu Arg Pro Leu Leu Gln Lys Trp Val 195
200 205 Glu Glu Ala Asp Asn Asn Glu
Asn Leu Gln Glu Ile Cys Lys Ala Glu 210 215
220 Thr Leu Val Gln Ala Arg Lys Arg Lys Arg Thr Ser
Ile Glu Asn Arg 225 230 235
240 Val Arg Gly Asn Leu Glu Asn Leu Phe Leu Gln Cys Pro Lys Pro Thr
245 250 255 Leu Gln Gln
Ile Ser His Ile Ala Gln Gln Leu Gly Leu Glu Lys Asp 260
265 270 Val Val Arg Val Trp Phe Cys Asn
Arg Arg Gln Lys Gly Lys Arg Ser 275 280
285 Ser Ser Asp Tyr Ala Gln Arg Glu Asp Phe Glu Ala Ala
Gly Ser Pro 290 295 300
Phe Ser Gly Gly Pro Val Ser Phe Pro Leu Ala Pro Gly Pro His Phe 305
310 315 320 Gly Thr Pro Gly
Tyr Gly Ser Pro His Phe Thr Ala Leu Tyr Ser Ser 325
330 335 Val Pro Phe Pro Glu Gly Glu Ala Phe
Pro Pro Val Ser Val Thr Thr 340 345
350 Leu Gly Ser Pro Met His Ser Asn Gly Ser Gly Ala Thr Asn
Phe Ser 355 360 365
Leu Leu Lys Gln Ala Gly Asp Val Glu Glu Asn Pro Gly Pro Met Tyr 370
375 380 Asn Met Met Glu Thr
Glu Leu Lys Pro Pro Gly Pro Gln Gln Thr Ser 385 390
395 400 Gly Gly Gly Gly Gly Asn Ser Thr Ala Ala
Ala Ala Gly Gly Asn Gln 405 410
415 Lys Asn Ser Pro Asp Arg Val Lys Arg Pro Met Asn Ala Phe Met
Val 420 425 430 Trp
Ser Arg Gly Gln Arg Arg Lys Met Ala Gln Glu Asn Pro Lys Met 435
440 445 His Asn Ser Glu Ile Ser
Lys Arg Leu Gly Ala Glu Trp Lys Leu Leu 450 455
460 Ser Glu Thr Glu Lys Arg Pro Phe Ile Asp Glu
Ala Lys Arg Leu Arg 465 470 475
480 Ala Leu His Met Lys Glu His Pro Asp Tyr Lys Tyr Arg Pro Arg Arg
485 490 495 Lys Thr
Lys Thr Leu Met Lys Lys Asp Lys Tyr Thr Leu Pro Gly Gly 500
505 510 Leu Leu Ala Pro Gly Gly Asn
Ser Met Ala Ser Gly Val Gly Val Gly 515 520
525 Ala Gly Leu Gly Ala Gly Val Asn Gln Arg Met Asp
Ser Tyr Ala His 530 535 540
Met Asn Gly Trp Ser Asn Gly Ser Tyr Ser Met Met Gln Asp Gln Leu 545
550 555 560 Gly Tyr Pro
Gln His Pro Gly Leu Asn Ala His Gly Ala Ala Gln Met 565
570 575 Gln Pro Met His Arg Tyr Asp Val
Ser Ala Leu Gln Tyr Asn Ser Met 580 585
590 Thr Ser Ser Gln Thr Tyr Met Asn Gly Ser Pro Thr Tyr
Ser Met Ser 595 600 605
Tyr Ser Gln Gln Gly Thr Pro Gly Met Ala Leu Gly Ser Met Gly Ser 610
615 620 Val Val Lys Ser
Glu Ala Ser Ser Ser Pro Pro Val Val Thr Ser Ser 625 630
635 640 Ser His Ser Arg Ala Pro Cys Gln Ala
Gly Asp Leu Arg Asp Met Ile 645 650
655 Ser Met Tyr Leu Pro Gly Ala Glu Val Pro Glu Pro Ala Ala
Pro Ser 660 665 670
Arg Leu His Met Ser Gln His Tyr Gln Ser Gly Pro Val Pro Gly Thr
675 680 685 Ala Ile Asn Gly
Thr Leu Pro Leu Ser His Met Gly Ser Gly Ala Thr 690
695 700 Asn Phe Ser Leu Leu Lys Gln Ala
Gly Asp Val Glu Glu Asn Pro Gly 705 710
715 720 Pro Met Ala Val Ser Asp Ala Leu Leu Pro Ser Phe
Ser Thr Phe Ala 725 730
735 Ser Gly Pro Ala Gly Arg Glu Lys Thr Leu Arg Gln Ala Gly Ala Pro
740 745 750 Asn Asn Arg
Trp Arg Glu Glu Leu Ser His Met Lys Arg Leu Pro Pro 755
760 765 Val Leu Pro Gly Arg Pro Tyr Asp
Leu Ala Ala Ala Thr Val Ala Thr 770 775
780 Asp Leu Glu Ser Gly Gly Ala Gly Ala Ala Cys Gly Gly
Ser Asn Leu 785 790 795
800 Ala Pro Leu Pro Arg Arg Glu Thr Glu Glu Phe Asn Asp Leu Leu Asp
805 810 815 Leu Asp Phe Ile
Leu Ser Asn Ser Leu Thr His Pro Pro Glu Ser Val 820
825 830 Ala Ala Thr Val Ser Ser Ser Ala Ser
Ala Ser Ser Ser Ser Ser Pro 835 840
845 Ser Ser Ser Gly Pro Ala Ser Ala Pro Ser Thr Cys Ser Phe
Thr Tyr 850 855 860
Pro Ile Arg Ala Gly Asn Asp Pro Gly Val Ala Pro Gly Gly Thr Gly 865
870 875 880 Gly Gly Leu Leu Tyr
Gly Arg Glu Ser Ala Pro Pro Pro Thr Ala Pro 885
890 895 Phe Asn Leu Ala Asp Ile Asn Asp Val Ser
Pro Ser Gly Gly Phe Val 900 905
910 Ala Glu Leu Leu Arg Pro Glu Leu Asp Pro Val Tyr Ile Pro Pro
Gln 915 920 925 Gln
Pro Gln Pro Pro Gly Gly Gly Leu Met Gly Lys Phe Val Leu Lys 930
935 940 Ala Ser Leu Ser Ala Pro
Gly Ser Glu Tyr Gly Ser Pro Ser Val Ile 945 950
955 960 Ser Val Ser Lys Gly Ser Pro Asp Gly Ser His
Pro Val Val Val Ala 965 970
975 Pro Tyr Asn Gly Gly Pro Pro Arg Thr Cys Pro Lys Ile Lys Gln Glu
980 985 990 Ala Val
Ser Ser Cys Thr His Leu Gly Ala Gly Pro Pro Leu Ser Asn 995
1000 1005 Gly His Arg Pro Ala
Ala His Asp Phe Pro Leu Gly Arg Gln Leu 1010 1015
1020 Pro Ser Arg Thr Thr Pro Thr Leu Gly Leu
Glu Glu Val Leu Ser 1025 1030 1035
Ser Arg Asp Cys His Pro Ala Leu Pro Leu Pro Pro Gly Phe His
1040 1045 1050 Pro His
Pro Gly Pro Asn Tyr Pro Ser Phe Leu Pro Asp Gln Met 1055
1060 1065 Gln Pro Gln Val Pro Pro Leu
His Tyr Gln Glu Leu Met Pro Pro 1070 1075
1080 Gly Ser Cys Met Pro Glu Glu Pro Lys Pro Lys Arg
Gly Arg Arg 1085 1090 1095
Ser Trp Pro Arg Lys Arg Thr Ala Thr His Thr Cys Asp Tyr Ala 1100
1105 1110 Gly Cys Gly Lys Thr
Tyr Thr Lys Ser Ser His Leu Lys Ala His 1115 1120
1125 Leu Arg Thr His Thr Gly Glu Lys Pro Tyr
His Cys Asp Trp Asp 1130 1135 1140
Gly Cys Gly Trp Lys Phe Ala Arg Ser Asp Glu Leu Thr Arg His
1145 1150 1155 Tyr Arg
Lys His Thr Gly His Arg Pro Phe Gln Cys Gln Lys Cys 1160
1165 1170 Asp Arg Ala Phe Ser Arg Ser
Asp His Leu Ala Leu His Met Lys 1175 1180
1185 Arg His Phe 1190 1047DNAArtificial
sequenceSynthetic construct 10cacacagcgg ccgcatttaa atccaccatg gcgggacacc
tggcttc 471159DNAArtificial sequenceSynthetic construct
11agaggacgaa cgaaattgtc tctcttcaag caccgaggca aacttacgta ccctctcgg
591266DNAArtificial sequenceSynthetic construct 12ctctgttaaa gcaagcagga
gatgttgaag aaaaccccgg gcctatgtac aacatgatgg 60agacgg
661364DNAArtificial
sequenceSynthetic construct 13agaggacgaa cgaaattgtc tctcttcaag caccgaggcc
tagggtacac actctccccg 60tcac
641463DNAArtificial sequenceSynthetic construct
14ctctgttaaa gcaagcagga gatgttgaag aaaaccccgg gcctatggct gtcagcgacg
60cgc
631544DNAArtificial sequenceSynthetic construct 15gtgtgtcagc tgtaaattta
aatttttacg gagaagtaca catt 441644DNAArtificial
sequenceSynthetic construct 16gtgtgtcagc tgtaaattta aatttttacg gagaagtaca
catt 441721DNAArtificial sequenceSynthetic construct
17gattatcgga attccctcga g
211819DNAArtificial sequenceSynthetic construct 18ccaaaggatg aagtgcaag
191920DNAArtificial
sequenceSynthetic construct 19agttttgctg caactgtacg
202020DNAArtificial sequenceSynthetic construct
20agcttgggct agagaaggat
202120DNAArtificial sequenceSynthetic construct 21tcagtttgaa tgcatgggag
202219DNAArtificial
sequenceSynthetic construct 22tgcacatggc ccagcacta
192320DNAArtificial sequenceSynthetic construct
23ttctccagtt cgcagtccag
202420DNAArtificial sequenceSynthetic construct 24aacttgctgt ctgaatggag
202520DNAArtificial
sequenceSynthetic construct 25tttgaggtcc tggtccatca
202620DNAArtificial sequenceSynthetic construct
26cagcagggac tgtcaccctg
202722DNAArtificial sequenceSynthetic construct 27ggtcacatcc actacgtggg
at 222820DNAArtificial
sequenceSynthetic construct 28ggagagtgcg attgcagaag
202921DNAArtificial sequenceSynthetic construct
29ggtcacatcc actacgtggg a
213025DNAArtificial sequenceSynthetic construct 30gttgttttgt tttggttttg
gatat 253126DNAArtificial
sequenceSynthetic construct 31atgggttgaa atattgggtt tattta
263225DNAArtificial sequenceSynthetic construct
32ccaccctcta accttaacct ctaac
253324DNAArtificial sequenceSynthetic construct 33gaggatgttt tttaagtttt
tttt 243425DNAArtificial
sequenceSynthetic construct 34aatgtttatg gtggattttg taggt
253525DNAArtificial sequenceSynthetic construct
35cccacactca tatcaatata ataac
253625DNAArtificial sequenceSynthetic construct 36gctcggtacc tttaagacca
atgac 253725DNAArtificial
sequenceSynthetic construct 37atgctgctag agattttcca cactg
253824DNAArtificial sequenceSynthetic construct
38tgaattgatc ccatcttgtc ttcg
243922DNAArtificial sequenceSynthetic construct 39tgctgctttt tgcttgtact
gg 224023DNAArtificial
sequenceSynthetic construct 40ttgagcaatg tggacagaga agg
234123DNAArtificial sequenceSynthetic construct
41gtcagaagca aatgtgagga gca
234223DNAArtificial sequenceSynthetic construct 42aattctggct tatcggaggc
aag 234313281DNAArtificial
sequenceSynthetic construct 43gttggaaggg ctaattcact cccaaagaag acaagatatc
cttgatctgt ggatctacca 60cacacaaggc tacttccctg attagcagaa ctacacacca
gggccagggg tcagatatcc 120actgaccttt ggatggtgct acaagctagt accagttgag
ccagataagg tagaagaggc 180caataaagga gagaacacca gcttgttaca ccctgtgagc
ctgcatggga tggatgaccc 240ggagagagaa gtgttagagt ggaggtttga cagccgccta
gcatttcatc acgtggcccg 300agagctgcat ccggagtact tcaagaactg ctgatatcga
gcttgctaca agggactttc 360cgctggggac tttccaggga ggcgtggcct gggcgggact
ggggagtggc gagccctcag 420atcctgcata taagcagctg ctttttgcct gtactgggtc
tctctggtta gaccagatct 480gagcctggga gctctctggc taactaggga acccactgct
taagcctcaa taaagcttgc 540cttgagtgct tcaagtagtg tgtgcccgtc tgttgtgtga
ctctggtaac tagagatccc 600tcagaccctt ttagtcagtg tggaaaatct ctagcagtgg
cgcccgaaca gggacttgaa 660agcgaaaggg aaaccagagg agctctctcg acgcaggact
cggcttgctg aagcgcgcac 720ggcaagaggc gaggggcggc gactggtgag tacgccaaaa
attttgacta gcggaggcta 780gaaggagaga gatgggtgcg agagcgtcag tattaagcgg
gggagaatta gatcgcgatg 840ggaaaaaatt cggttaaggc cagggggaaa gaaaaaatat
aaattaaaac atatagtatg 900ggcaagcagg gagctagaac gattcgcagt taatcctggc
ctgttagaaa catcagaagg 960ctgtagacaa atactgggac agctacaacc atcccttcag
acaggatcag aagaacttag 1020atcattatat aatacagtag caaccctcta ttgtgtgcat
caaaggatag agataaaaga 1080caccaaggaa gctttagaca agatagagga agagcaaaac
aaaagtaaga ccaccgcaca 1140gcaagcggcc gctgatcttc agacctggag gaggagatat
gagggacaat tggagaagtg 1200aattatataa atataaagta gtaaaaattg aaccattagg
agtagcaccc accaaggcaa 1260agagaagagt ggtgcagaga gaaaaaagag cagtgggaat
aggagctttg ttccttgggt 1320tcttgggagc agcaggaagc actatgggcg cagcgtcaat
gacgctgacg gtacaggcca 1380gacaattatt gtctggtata gtgcagcagc agaacaattt
gctgagggct attgaggcgc 1440aacagcatct gttgcaactc acagtctggg gcatcaagca
gctccaggca agaatcctgg 1500ctgtggaaag atacctaaag gatcaacagc tcctggggat
ttggggttgc tctggaaaac 1560tcatttgcac cactgctgtg ccttggaatg ctagttggag
taataaatct ctggaacaga 1620tttggaatca cacgacctgg atggagtggg acagagaaat
taacaattac acaagcttaa 1680tacactcctt aattgaagaa tcgcaaaacc agcaagaaaa
gaatgaacaa gaattattgg 1740aattagataa atgggcaagt ttgtggaatt ggtttaacat
aacaaattgg ctgtggtata 1800taaaattatt cataatgata gtaggaggct tggtaggttt
aagaatagtt tttgctgtac 1860tttctatagt gaatagagtt aggcagggat attcaccatt
atcgtttcag acccacctcc 1920caaccccgag gggacccgac aggcccgaag gaatagaaga
agaaggtgga gagagagaca 1980gagacagatc cattcgatta gtgaacggat ctcgacggta
tcgatgtcga cgataagctt 2040tgcaaagatg gataaagttt taaacagaga ggaatctttg
cagctaatgg accttctagg 2100tcttgaaagg agtgggaatt ggctccggtg cccgtcagtg
ggcagagcgc acatcgccca 2160cagtccccga gaagttgggg ggaggggtcg gcaattgaac
cggtgcctag agaaggtggc 2220gcggggtaaa ctgggaaagt gatgtcgtgt actggctccg
cctttttccc gagggtgggg 2280gagaaccgta tataagtgca gtagtcgccg tgaacgttct
ttttcgcaac gggtttgccg 2340ccagaacaca ggtaagtgcc gtgtgtggtt cccgcgggcc
tggcctcttt acgggttatg 2400gcccttgcgt gccttgaatt acttccactg gctgcagtac
gtgattcttg atcccgagct 2460tcgggttgga agtgggtggg agagttcgag gccttgcgct
taaggagccc cttcgcctcg 2520tgcttgagtt gaggcctggc ctgggcgctg gggccgccgc
gtgcgaatct ggtggcacct 2580tcgcgcctgt ctcgctgctt tcgataagtc tctagccatt
taaaattttt gatgacctgc 2640tgcgacgctt tttttctggc aagatagtct tgtaaatgcg
ggccaagatc tgcacactgg 2700tatttcggtt tttggggccg cgggcggcga cggggcccgt
gcgtcccagc gcacatgttc 2760ggcgaggcgg ggcctgcgag cgcggccacc gagaatcgga
cgggggtagt ctcaagctgg 2820ccggcctgct ctggtgcctg gcctcgcgcc gccgtgtatc
gccccgccct gggcggcaag 2880gctggcccgg tcggcaccag ttgcgtgagc ggaaagatgg
ccgcttcccg gccctgctgc 2940agggagctca aaatggagga cgcggcgctc gggagagcgg
gcgggtgagt cacccacaca 3000aaggaaaagg gcctttccgt cctcagccgt cgcttcatgt
gactccacgg agtaccgggc 3060gccgtccagg cacctcgatt agttctcgag cttttggagt
acgtcgtctt taggttgggg 3120ggaggggttt tatgcgatgg agtttcccca cactgagtgg
gtggagactg aagttaggcc 3180agcttggcac ttgatgtaat tctccttgga atttgccctt
tttgagtttg gatcttggtt 3240cattctcaag cctcagacag tggttcaaag tttttttctt
ccatttcagg tgtcgtgagg 3300aatttcgaca tttaaatcca ccatggcggg acacctggct
tcggatttcg ccttctcgcc 3360ccctccaggt ggtggaggtg atgggccagg ggggccggag
ccgggctggg ttgatcctcg 3420gacctggcta agcttccaag gccctcctgg agggccagga
atcgggccgg gggttgggcc 3480aggctctgag gtgtggggga ttcccccatg ccccccgccg
tatgagttct gtggggggat 3540ggcgtactgt gggccccagg ttggagtggg gctagtgccc
caaggcggct tggagacctc 3600tcagcctgag ggcgaagcag gagtcggggt ggagagcaac
tccgatgggg cctccccgga 3660gccctgcacc gtcacccctg gtgccgtgaa gctggagaag
gagaagctgg agcaaaaccc 3720ggaggagtcc caggacatca aagctctgca gaaagaactc
gagcaatttg ccaagctcct 3780gaagcagaag aggatcaccc tgggatatac acaggccgat
gtggggctca ccctgggggt 3840tctatttggg aaggtattca gccaaacgac catctgccgc
tttgaggctc tgcagcttag 3900cttcaagaac atgtgtaagc tgcggccctt gctgcagaag
tgggtggagg aagctgacaa 3960caatgaaaat cttcaggaga tatgcaaagc agaaaccctc
gtgcaggccc gaaagagaaa 4020gcgaaccagt atcgagaacc gagtgagagg caacctggag
aatttgttcc tgcagtgccc 4080gaaacccaca ctgcagcaga tcagccacat cgcccagcag
cttgggctcg agaaggatgt 4140ggtccgagtg tggttctgta accggcgcca gaagggcaag
cgatcaagca gcgactatgc 4200acaacgagag gattttgagg ctgctgggtc tcctttctca
gggggaccag tgtcctttcc 4260tctggcccca gggccccatt ttggtacccc aggctatggg
agccctcact tcactgcact 4320gtactcctcg gtccctttcc ctgaggggga agcctttccc
cctgtctccg tcaccactct 4380gggctctccc atgcattcaa acggatccgg agccacgaac
ttctctctgt taaagcaagc 4440aggagatgtt gaagaaaacc ccgggcctat gtacaacatg
atggagacgg agctgaagcc 4500gccgggcccg cagcaaactt cggggggcgg cggcggcaac
tccaccgcgg cggcggccgg 4560cggcaaccag aaaaacagcc cggaccgcgt caagcggccc
atgaatgcct tcatggtgtg 4620gtcccgcggg cagcggcgca agatggccca ggagaacccc
aagatgcaca actcggagat 4680cagcaagcgc ctgggcgccg agtggaaact tttgtcggag
acggagaagc ggccgttcat 4740cgacgaggct aagcggctgc gagcgctgca catgaaggag
cacccggatt ataaataccg 4800gccccggcgg aaaaccaaga cgctcatgaa gaaggataag
tacacgctgc ccggcgggct 4860gctggccccc ggcggcaata gcatggcgag cggggtcggg
gtgggcgccg gcctgggcgc 4920gggcgtgaac cagcgcatgg acagttacgc gcacatgaac
ggctggagca acggcagcta 4980cagcatgatg caggaccagc tgggctaccc gcagcacccg
ggcctcaatg cgcacggcgc 5040agcgcagatg cagcccatgc accgctacga cgtgagcgcc
ctgcagtaca actccatgac 5100cagctcgcag acctacatga acggctcgcc cacctacagc
atgtcctact cgcagcaggg 5160cacccctggc atggctcttg gctccatggg ttcggtggtc
aagtccgagg ccagctccag 5220cccccctgtg gttacctctt cctcccactc cagggcgccc
tgccaggccg gggacctccg 5280ggacatgatc agcatgtatc tccccggcgc cgaggtgccg
gaacccgccg cccccagcag 5340acttcacatg tcccagcact accagagcgg cccggtgccc
ggcacggcca ttaacggcac 5400actgcccctc tcacacatgg gatccggagc cacgaacttc
tctctgttaa agcaagcagg 5460agatgttgaa gaaaaccccg ggcctatggc tgtcagcgac
gcgctgctcc catctttctc 5520cacgttcgcg tctggcccgg cgggaaggga gaagacactg
cgtcaagcag gtgccccgaa 5580taaccgctgg cgggaggagc tctcccacat gaagcgactt
cccccagtgc ttcccggccg 5640cccctatgac ctggcggcgg cgaccgtggc cacagacctg
gagagcggcg gagccggtgc 5700ggcttgcggc ggtagcaacc tggcgcccct acctcggaga
gagaccgagg agttcaacga 5760tctcctggac ctggacttta ttctctccaa ttcgctgacc
catcctccgg agtcagtggc 5820cgccaccgtg tcctcgtcag cgtcagcctc ctcttcgtcg
tcgccgtcga gcagcggccc 5880tgccagcgcg ccctccacct gcagcttcac ctatccgatc
cgggccggga acgacccggg 5940cgtggcgccg ggcggcacgg gcggaggcct cctctatggc
agggagtccg ctccccctcc 6000gacggctccc ttcaacctgg cggacatcaa cgacgtgagc
ccctcgggcg gcttcgtggc 6060cgagctcctg cggccagaat tggacccggt gtacattccg
ccgcagcagc cgcagccgcc 6120aggtggcggg ctgatgggca agttcgtgct gaaggcgtcg
ctgagcgccc ctggcagcga 6180gtacggcagc ccgtcggtca tcagcgtcag caaaggcagc
cctgacggca gccacccggt 6240ggtggtggcg ccctacaacg gcgggccgcc gcgcacgtgc
cccaagatca agcaggaggc 6300ggtctcttcg tgcacccact tgggcgctgg accccctctc
agcaatggcc accggccggc 6360tgcacacgac ttccccctgg ggcggcagct ccccagcagg
actaccccga ccctgggtct 6420tgaggaagtg ctgagcagca gggactgtca ccctgccctg
ccgcttcctc ccggcttcca 6480tccccacccg gggcccaatt acccatcctt cctgcccgat
cagatgcagc cgcaagtccc 6540gccgctccat taccaagagc tcatgccacc cggttcctgc
atgccagagg agcccaagcc 6600aaagagggga agacgatcgt ggccccggaa aaggaccgcc
acccacactt gtgattacgc 6660gggctgcggc aaaacctaca caaagagttc ccatctcaag
gcacacctgc gaacccacac 6720aggtgagaaa ccttaccact gtgactggga cggctgtgga
tggaaattcg cccgctcaga 6780tgaactgacc aggcactacc gtaaacacac ggggcaccgc
ccgttccagt gccaaaaatg 6840cgaccgagca ttttccaggt cggaccacct cgccttacac
atgaagaggc atttttaaat 6900ttaaatttaa ttaatctcga cggtatcggt taacttttaa
aagaaaaggg gggattgggg 6960ggtacagtgc aggggaaaga atagtagaca taatagcaac
agacatacaa actaaagaat 7020tacaaaaaca aattacaaaa attcaaaatt ttccgatcac
gagactagcc tcgagggaat 7080tccgataatc aacctctgga ttacaaaatt tgtgaaagat
tgactggtat tcttaactat 7140gttgctcctt ttacgctatg tggatacgct gctttaatgc
ctttgtatca tgctattgct 7200tcccgtatgg ctttcatttt ctcctccttg tataaatcct
ggttgctgtc tctttatgag 7260gagttgtggc ccgttgtcag gcaacgtggc gtggtgtgca
ctgtgtttgc tgacgcaacc 7320cccactggtt ggggcattgc caccacctgt cagctccttt
ccgggacttt cgctttcccc 7380ctccctattg ccacggcgga actcatcgcc gcctgccttg
cccgctgctg gacaggggct 7440cggctgttgg gcactgacaa ttccgtggtg ttgtcgggga
agctgacgtc ctttccatgg 7500ctgctcgcct gtgttgccac ctggattctg cgcgggacgt
ccttctgcta cgtcccttcg 7560gccctcaatc cagcggacct tccttcccgc ggcctgctgc
cggctctgcg gcctcttccg 7620cgtcttcgcc ttcgccctca gacgagtcgg atctcccttt
gggccgcctc cccgcatcgg 7680gaattcgctc aagcttcgaa ttaattctgc agagctcggt
acctttaaga ccaatgactt 7740acaaggcagc tgtagatctt agccactttt taaaagaaaa
ggggggactg gaagggctaa 7800ttcactccca acgaagacaa gatgggatca attcaccatg
ggaataactt cgtatagcat 7860acattatacg aagttatgct gctttttgct tgtactgggt
ctctctggtt agaccagatc 7920tgagcctggg agctctctgg ctaactaggg aacccactgc
ttaagcctca ataaagcttg 7980ccttgagtgc ttcaagtagt gtgtgcccgt ctgttgtgtg
actctggtaa ctagagatcc 8040ctcagaccct tttagtcagt gtggaaaatc tctagcagca
tctagaatta attccgtgta 8100ttctatagtg tcacctaaat cgtatgtgta tgatacataa
ggttatgtat taattgtagc 8160cgcgttctaa cgacaatatg tacaagccta attgtgtagc
atctggctta ctgaagcaga 8220ccctatcatc tctctcgtaa actgccgtca gagtcggttt
ggttggacga accttctgag 8280tttctggtaa cgccgtcccg cacccggaaa tggtcagcga
accaatcagc agggtcatcg 8340ctagccagat cctctacgcc ggacgcatcg tggccggcat
caccggcgcc acaggtgcgg 8400ttgctggcgc ctatatcgcc gacatcaccg atggggaaga
tcgggctcgc cacttcgggc 8460tcatgagcgc ttgtttcggc gtgggtatgg tggcaggccc
cgtggccggg ggactgttgg 8520gcgccatctc cttgcatgca ccattccttg cggcggcggt
gctcaacggc ctcaacctac 8580tactgggctg cttcctaatg caggagtcgc ataagggaga
gcgtcgaatg gtgcactctc 8640agtacaatct gctctgatgc cgcatagtta agccagcccc
gacacccgcc aacacccgct 8700gacgcgccct gacgggcttg tctgctcccg gcatccgctt
acagacaagc tgtgaccgtc 8760tccgggagct gcatgtgtca gaggttttca ccgtcatcac
cgaaacgcgc gagacgaaag 8820ggcctcgtga tacgcctatt tttataggtt aatgtcatga
taataatggt ttcttagacg 8880tcaggtggca cttttcgggg aaatgtgcgc ggaaccccta
tttgtttatt tttctaaata 8940cattcaaata tgtatccgct catgagacaa taaccctgat
aaatgcttca ataatattga 9000aaaaggaaga gtatgagtat tcaacatttc cgtgtcgccc
ttattccctt ttttgcggca 9060ttttgccttc ctgtttttgc tcacccagaa acgctggtga
aagtaaaaga tgctgaagat 9120cagttgggtg cacgagtggg ttacatcgaa ctggatctca
acagcggtaa gatccttgag 9180agttttcgcc ccgaagaacg ttttccaatg atgagcactt
ttaaagttct gctatgtggc 9240gcggtattat cccgtattga cgccgggcaa gagcaactcg
gtcgccgcat acactattct 9300cagaatgact tggttgagta ctcaccagtc acagaaaagc
atcttacgga tggcatgaca 9360gtaagagaat tatgcagtgc tgccataacc atgagtgata
acactgcggc caacttactt 9420ctgacaacga tcggaggacc gaaggagcta accgcttttt
tgcacaacat gggggatcat 9480gtaactcgcc ttgatcgttg ggaaccggag ctgaatgaag
ccataccaaa cgacgagcgt 9540gacaccacga tgcctgtagc aatggcaaca acgttgcgca
aactattaac tggcgaacta 9600cttactctag cttcccggca acaattaata gactggatgg
aggcggataa agttgcagga 9660ccacttctgc gctcggccct tccggctggc tggtttattg
ctgataaatc tggagccggt 9720gagcgtgggt ctcgcggtat cattgcagca ctggggccag
atggtaagcc ctcccgtatc 9780gtagttatct acacgacggg gagtcaggca actatggatg
aacgaaatag acagatcgct 9840gagataggtg cctcactgat taagcattgg taactgtcag
accaagttta ctcatatata 9900ctttagattg atttaaaact tcatttttaa tttaaaagga
tctaggtgaa gatccttttt 9960gataatctca tgaccaaaat cccttaacgt gagttttcgt
tccactgagc gtcagacccc 10020gtagaaaaga tcaaaggatc ttcttgagat cctttttttc
tgcgcgtaat ctgctgcttg 10080caaacaaaaa aaccaccgct accagcggtg gtttgtttgc
cggatcaaga gctaccaact 10140ctttttccga aggtaactgg cttcagcaga gcgcagatac
caaatactgt ccttctagtg 10200tagccgtagt taggccacca cttcaagaac tctgtagcac
cgcctacata cctcgctctg 10260ctaatcctgt taccagtggc tgctgccagt ggcgataagt
cgtgtcttac cgggttggac 10320tcaagacgat agttaccgga taaggcgcag cggtcgggct
gaacgggggg ttcgtgcaca 10380cagcccagct tggagcgaac gacctacacc gaactgagat
acctacagcg tgagcattga 10440gaaagcgcca cgcttcccga agggagaaag gcggacaggt
atccggtaag cggcagggtc 10500ggaacaggag agcgcacgag ggagcttcca gggggaaacg
cctggtatct ttatagtcct 10560gtcgggtttc gccacctctg acttgagcgt cgatttttgt
gatgctcgtc aggggggcgg 10620agcctatgga aaaacgccag caacgcggcc tttttacggt
tcctggcctt ttgctggcct 10680tttgctcaca tgttctttcc tgcgttatcc cctgattctg
tggataaccg tattaccgcc 10740tttgagtgag ctgataccgc tcgccgcagc cgaacgaccg
agcgcagcga gtcagtgagc 10800gaggaagcgg aagagcgccc aatacgcaaa ccgcctctcc
ccgcgcgttg gccgattcat 10860taatgcagct gtggaatgtg tgtcagttag ggtgtggaaa
gtccccaggc tccccagcag 10920gcagaagtat gcaaagcatg catctcaatt agtcagcaac
caggtgtgga aagtccccag 10980gctccccagc aggcagaagt atgcaaagca tgcatctcaa
ttagtcagca accatagtcc 11040cgcccctaac tccgcccatc ccgcccctaa ctccgcccag
ttccgcccat tctccgcccc 11100atggctgact aatttttttt atttatgcag aggccgaggc
cgcctcggcc tctgagctat 11160tccagaagta gtgaggaggc ttttttggag gcctaggctt
ttgcaaaaag cttggacaca 11220agacaggctt gcgagatatg tttgagaata ccactttatc
ccgcgtcagg gagaggcagt 11280gcgtaaaaag acgcggactc atgtgaaata ctggttttta
gtgcgccaga tctctataat 11340ctcgcgcaac ctattttccc ctcgaacact ttttaagccg
tagataaaca ggctgggaca 11400cttcacatga gcgaaaaata catcgtcacc tgggacatgt
tgcagatcca tgcacgtaaa 11460ctcgcaagcc gactgatgcc ttctgaacaa tggaaaggca
ttattgccgt aagccgtggc 11520ggtctgtacc gggtgcgtta ctggcgcgtg aactgggtat
tcgtcatgtc gataccgttt 11580gtatttccag ctacgatcac gacaaccagc gcgagcttaa
agtgctgaaa cgcgcagaag 11640gcgatggcga aggcttcatc gttattgatg acctggtgga
taccggtggt actgcggttg 11700cgattcgtga aatgtatcca aaagcgcact ttgtcaccat
cttcgcaaaa ccggctggtc 11760gtccgctggt tgatgactat gttgttgata tcccgcaaga
tacctggatt gaacagccgt 11820gggatatggg cgtcgtattc gtcccgccaa tctccggtcg
ctaatctttt caacgcctgg 11880cactgccggg cgttgttctt tttaacttca ggcgggttac
aatagtttcc agtaagtatt 11940ctggaggctg catccatgac acaggcaaac ctgagcgaaa
ccctgttcaa accccgcttt 12000aaacatcctg aaacctcgac gctagtccgc cgctttaatc
acggcgcaca accgcctgtg 12060cagtcggccc ttgatggtaa aaccatccct cactggtatc
gcatgattaa ccgtctgatg 12120tggatctggc gcggcattga cccacgcgaa atcctcgacg
tccaggcacg tattgtgatg 12180agcgatgccg aacgtaccga cgatgattta tacgatacgg
tgattggcta ccgtggcggc 12240aactggattt atgagtgggc cccggatctt tgtgaaggaa
ccttacttct gtggtgtgac 12300ataattggac aaactaccta cagagattta aagctctaag
gtaaatataa aatttttaag 12360tgtataatgt gttaaactac tgattctaat tgtttgtgta
ttttagattc caacctatgg 12420aactgatgaa tgggagcagt ggtggaatgc ctttaatgag
gaaaacctgt tttgctcaga 12480agaaatgcca tctagtgatg atgaggctac tgctgactct
caacattcta ctcctccaaa 12540aaagaagaga aaggtagaag accccaagga ctttccttca
gaattgctaa gttttttgag 12600tcatgctgtg tttagtaata gaactcttgc ttgctttgct
atttacacca caaaggaaaa 12660agctgcactg ctatacaaga aaattatgga aaaatattct
gtaaccttta taagtaggca 12720taacagttat aatcataaca tactgttttt tcttactcca
cacaggcata gagtgtctgc 12780tattaataac tatgctcaaa aattgtgtac ctttagcttt
ttaatttgta aaggggttaa 12840taaggaatat ttgatgtata gtgccttgac tagagatcat
aatcagccat accacatttg 12900tagaggtttt acttgcttta aaaaacctcc cacacctccc
cctgaacctg aaacataaaa 12960tgaatgcaat tgttgttgtt aacttgttta ttgcagctta
taatggttac aaataaagca 13020atagcatcac aaatttcaca aataaagcat ttttttcact
gcattctagt tgtggtttgt 13080ccaaactcat caatgtatct tatcatgtct ggatcaactg
gataactcaa gctaaccaaa 13140atcatcccaa acttcccacc ccatacccta ttaccactgc
caattaccta gtggtttcat 13200ttactctaaa cctgtgattc ctctgaatta ttttcatttt
aaagaaattg tatttgttaa 13260atatgtacta caaacttagt a
132814421697DNAArtificial sequenceSynthetic
construct 44cgggaagcgt ggcgctttct catagctcac gctgtaggta tctcagttcg
gtgtaggtcg 60ttcgctccaa gctgggctgt gtgcacgaac cccccgttca gcccgaccgc
tgcgccttat 120ccggtaacta tcgtcttgag tccaacccgg taagacacga cttatcgcca
ctggcagcag 180ccactggtaa caggattagc agagcgaggt atgtaggcgg tgctacagag
ttcttgaagt 240ggtggcctaa ctacggctac actagaagga cagtatttgg tatctgcgct
ctgctgaagc 300cagttacctt cggaaaaaga gttggtagct cttgatccgg caaacaaacc
accgctggta 360gcggtggttt ttttgtttgc aagcagcaga ttacgcgcag aaaaaaagga
tctcaagaag 420atcctttgat cttttctacg gggtctgacg ctcagtggaa cgaaaactca
cgttaaggga 480ttttggtcat gagattatca aaaaggatct tcacctagat ccttttaaat
taaaaatgaa 540gttttaaatc aatctaaagt atatatgagt aaacttggtc tgacagttac
caatgcttaa 600tcagtgaggc acctatctca gcgatctgtc tatttcgttc atccatagtt
gcctgactcc 660ccgtcgtgta gataactacg atacgggagg gcttaccatc tggccccagt
gctgcaatga 720taccgcgaga cccacgctca ccggctccag atttatcagc aataaaccag
ccagccggaa 780gggccgagcg cagaagtggt cctgcaactt tatccgcctc catccagtct
attaattgtt 840gccgggaagc tagagtaagt agttcgccag ttaatagttt gcgcaacgtt
gttgccattg 900ctacaggcat cgtggtgtca cgctcgtcgt ttggtatggc ttcattcagc
tccggttccc 960aacgatcaag gcgagttaca tgatccccca tgttgtgcaa aaaagcggtt
agctccttcg 1020gtcctccgat cgttgtcaga agtaagttgg ccgcagtgtt atcactcatg
gttatggcag 1080cactgcataa ttctcttact gtcatgccat ccgtaagatg cttttctgtg
actggtgagt 1140actcaaccaa gtcattctga gaatagtgta tgcggcgacc gagttgctct
tgcccggcgt 1200caatacggga taataccgcg ccacatagca gaactttaaa agtgctcatc
attggaaaac 1260gttcttcggg gcgaaaactc tcaaggatct taccgctgtt gagatccagt
tcgatgtaac 1320ccactcgtgc acccaactga tcttcagcat cttttacttt caccagcgtt
tctgggtgag 1380caaaaacagg aaggcaaaat gccgcaaaaa agggaataag ggcgacacgg
aaatgttgaa 1440tactcatact cttccttttt caatattatt gaagcattta tcagggttat
tgtctcatga 1500gcggatacat atttgaatgt atttagaaaa ataaacaaat aggggttccg
cgcacatttc 1560cccgaaaagt gccacctaaa ttgtaagcgt taatattttg ttaaaattcg
cgttaaattt 1620ttgttaaatc agctcatttt ttaaccaata ggccgaaatc ggcaaaatcc
cttataaatc 1680aaaagaatag accgagatag ggttgagtgt tgttccagtt tggaacaaga
gtccactatt 1740aaagaacgtg gactccaacg tcaaagggcg aaaaaccgtc tatcagggcg
atggcccact 1800acgtgaacca tcaccctaat caagtttttt ggggtcgagg tgccgtaaag
cactaaatcg 1860gaaccctaaa gggagccccc gatttagagc ttgacgggga aagccggcga
acgtggcgag 1920aaaggaaggg aagaaagcga aaggagcggg cgctagggcg ctggcaagtg
tagcggtcac 1980gctgcgcgta accaccacac ccgccgcgct taatgcgccg ctacagggcg
cgtcccattc 2040gccattcagg ctgcgcaact gttgggaagg gcgatcggtg cgggcctctt
cgctattacg 2100ccagctggcg aaagggggat gtgctgcaag gcgattaagt tgggtaacgc
cagggttttc 2160ccagtcacga cgttgtaaaa cgacggccag tgaattgtaa tacgactcac
tatagggcga 2220attgggtacc gggccccccc tcgagcagtg tggttttcaa gaggaagcaa
aaagcctctc 2280cacccaggcc tggaatgttt ccacccaatg tcgagcagtg tggttttgca
agaggaagca 2340aaaagcctct ccacccaggc ctggaatgtt tccacccaat gtcgagcaaa
ccccgcccag 2400cgtcttgtca ttggcgaatt cgaacacgca gatgcagtcg gggcggcgcg
gtccgaggtc 2460cacttcgcat attaaggtga cgcgtgtggc ctcgaacacc gagcgaccct
gcagcgaccc 2520gcttaacagc gtcaacagcg tgccgcagat cttggtggcg tgaaactccc
gcacctcttc 2580ggccagcgcc ttgtagaagc gcgtatggct tcgtaccccg gccatcaaca
cgcgtctgcg 2640ttcgaccagg ctgcgcgttc tcgcggccat agcaaccgac gtacggcgtt
gcgccctcgc 2700cggcagcaag aagccacgga agtccgcccg gagcagaaaa tgcccacgct
actgcgggtt 2760tatatagacg gtccccacgg gatggggaaa accaccacca cgcaactgct
ggtggccctg 2820ggttcgcgcg acgatatcgt ctacgtaccc gagccgatga cttactggcg
ggtgctgggg 2880gcttccgaga caatcgcgaa catctacacc acacaacacc gcctcgacca
gggtgagata 2940tcggccgggg acgcggcggt ggtaatgaca agcgcccaga taacaatggg
catgccttat 3000gccgtgaccg acgccgttct ggctcctcat atcggggggg aggctgggag
ctcacatgcc 3060ccgcccccgg ccctcaccct catcttcgac cgccatccca tcgccgccct
cctgtgctac 3120ccggccgcgc ggtaccttat gggcagcatg accccccagg ccgtgctggc
gttcgtggcc 3180ctcatcccgc cgaccttgcc cggcaccaac atcgtgcttg gggcccttcc
ggaggacaga 3240cacatcgacc gcctggccaa acgccagcgc cccggcgagc ggctggacct
ggctatgctg 3300gctgcgattc gccgcgttta cgggctactt gccaatacgg tgcggtatct
gcagtgcggc 3360gggtcgtggc gggaggactg gggacagctt tcggggacgg ccgtgccgcc
ccagggtgcc 3420gagccccaga gcaacgcggg cccacgaccc catatcgggg acacgttatt
taccctgttt 3480cgggcccccg agttgctggc ccccaacggc gacctgtata acgtgtttgc
ctgggccttg 3540gacgtcttgg ccaaacgcct ccgttccatg cacgtcttta tcctggatta
cgaccaatcg 3600cccgccggct gccgggacgc cctgctgcaa cttacctccg ggatggtcca
gacccacgtc 3660accacccccg gctccatacc gacgatatgc gacctggcgc gcacgtttgc
ccgggagatg 3720ggggaggcta actgaaacac ggaaggagac aataccggaa ggaacccgcg
ctatgacggc 3780aataaaaaga cagaataaaa cgcacgggtg ttgggtcgtt tgttcataaa
cgcggggttc 3840ggtcccaggg ctggcactct gtcgataccc caccgagacc ccattggggc
caatacgccc 3900gcgtttcttc cttttcccca ccccaccccc caagttcggg tgaaggccca
gggctcgcag 3960ccaacgtcgg ggcggcaggc cctgccatag ccactggccc cgtgggttag
ggacggggtc 4020ccccatgggg aatggtttat ggttcgtggg ggttattatt ttgggcgttg
cgtggggtca 4080ggtccacgac cctaagcttg atatcgaatt cctgcagccc gggggatcct
cctccttcct 4140ttgcctgcac attgtagccc ataatactat accccatcaa gtgttcctgc
tccaagaaat 4200agcttcctcc tcttacttgc cccagaacat ctctgtaaag aatttcctct
tatcttccca 4260tatttcagtc aagattcatt gctcacgtat tacttgtgac ctctcttgac
cccagccaca 4320ataaacttct ctatactacc caaaaaatct ttccaaaccc tccccgacac
catattttta 4380tatttttctt atttatttca tgcacacaca cacactccgt gctttataag
caattctgcc 4440tattctctac cttcttacaa tgcctactgt gcctcatatt aaattcatca
atgggcagaa 4500agaaaatatt tattcaagaa aacagtgaat gaatgaacga atgagtaaat
gagtaaatga 4560aggaatgatt attccttgct ttagaacttc tggaattaga ggacaatatt
aataatacca 4620tcgcacagtg tttctttgtt gttaatgcta caacatacaa agaggaagca
tgcagtaaac 4680aaccgaacag ttatttcctt tctgatcata ggagtaatat ttttttcctt
gagcacattt 4740ttgccatagg taaaattaga aggattttta gaactttctc agttgtatac
atttttaaaa 4800atctgtatta tatgcatgtt gattaatttt aaacttactt gaatacctaa
acagaatctg 4860ttgtttcctt gtgtttgaaa gtgctttcac agtaactctg tctgtactgc
cagaatatac 4920tgacaatgtg ttatagttaa ctgttttgat cacaacattt tgaattgact
ggcagcagaa 4980gctcttttta tatccatgtg ttttccttaa gtcattatac atagtaggca
tgagactctt 5040tatactgaat aagatattta ggaaccactg gtttacatat cagaagcaga
gctactcagg 5100gcattttggg gaagatcact ttcacattcc tgagcatagg gaagttctca
taagagtaag 5160atattaaaag gagatacttg tgtggtattc gaaagacagt aagagagatt
gtagacctta 5220tgatcttgat agggaaaaca aactacattc ctttctccaa aagtcaaaaa
aaaagagcaa 5280atatagctta ctataccttc tattcctaca ccattagaag tagtcagtga
gtctaggcaa 5340gatgttggcc ctaaaaatcc aaataccaga gaattcatga gaacatcacc
tggatgggac 5400atgtgccgag caacacaatt actatatgct aggcattgct atcttcatat
tgaagatgag 5460gaggtcaaga gatgaaaaaa gacttggcac cttgttgtta tattaaaatt
atttgttaga 5520gtagagcttt tgtaagagtc taggagtgtg ggagctaaat gatgatacac
atggacacaa 5580agaatagatc aacagacacc caggcctact tgagggttga gggtgggaag
agggagacga 5640tgaaaaagaa cctattgggt attaagttca tcactgagtg atgaaataat
ctgtacatca 5700agacccagtg atatgcaatt tacctatata acttgtacat gtacccccaa
atttaaaata 5760aagttaaaac aaagtatagg aatggaatta attcctcaag atttggcttt
aattttattt 5820gataatttat caaatggttg tttttctttt ctcactatgg cgttgcttta
taaactatgt 5880tcagtatgtc tgaatgaaag ggtgtgtgtg tgtgtgaaag agagggagag
aggaagggaa 5940gagaggacgt aataatgtga atttgagttc atgaaaattt ttcaataaaa
taatttaatg 6000tcaggagaat taagcctaat agtctcctaa atcatccatc tcttgagctt
cagagcagtc 6060ctctgaatta atgcctacat gtttgtaaag ggtgttcaga ctgaagccaa
gattctacct 6120ctaaagagat gcaatctcaa atttatctga agactgtacc tctgctctcc
ataaattgac 6180accatggccc acttaatgag gttaaaaaaa agctaattct gaatgaaaat
ctgagcccag 6240tggaggaaat attaatgaac aaggtgcaga ctgaaatata aattttctgt
aataattatg 6300catatacttt agcaaagttc tgtctatgtt gactttattg cttttggtaa
gaaatacaac 6360tttttaaagt gaactaaact atcctatttc caaactattt tgtgtgtgtg
cggtttgttt 6420ctatgggttc tggttttctt ggagcatttt tatttcattt taattaatta
attctgagag 6480ctgctgagtt gtgtttactg agagattgtg tatctgcgag agaagtctgt
agcaagtagc 6540tagactgtgc ttgacctagg aacatataca gtagattgct aaaatgtctc
acttggggaa 6600ttttagacta aacagtagag catgtataaa aatactctag tcaagtgctg
cttttgaaac 6660aaatgataaa accacactcc catagatgag tgtcatgatt ttcatggagg
aagttaatat 6720tcatcctcta agtataccca gactagggcc attctgatat aaaacattag
gacttaagaa 6780agattaatag actggagtaa aggaaatgga cctctgtctc tctcgctgtc
tcttttttga 6840ggacttgtgt gtgtgtgtgt gtgtgtgtgt gtgtgtgttg tggtcagtgg
ggctggaata 6900aaagtagaat agacctgcac ctgctgtggc atccattcac agagtagaag
caagctcaca 6960atagtgaaga tgtcagtaag cttgaatagt ttttcaggaa ctttgaatgc
tgatttagat 7020ttgaaactga ggctctgacc ataaccaaat ttgcactatt tattgcttct
tgaaacttat 7080ttgcctggta tgcctgggct tttgatggtc ttagtatagc ttgcagcctt
gtccctgcag 7140ggtattatgg gtaatagaaa gaaaagtctg cgttacactc tagtcacact
aagtaactac 7200cattggaaaa gcaacccctg ccttgaagcc aggatgatgg tatctgcagc
agttgccaac 7260acaagagaag gatccatagt tcatcattta aaaaagaaaa caaaatagaa
aaaggaaaac 7320tatttctgag cataagaagt tgtagggtaa gtctttaaga aggtgacaat
ttctgccaat 7380caggatttca aagctcttgc tttgacaatt ttggtctttc agaatactat
aaatataacc 7440tatattataa tttcataaag tctgtgcatt ttctttgacc caggatattt
gcaaaagaca 7500tattcaaact tccgcagaac actttatttc acatatacat gcctcttata
tcagggatgt 7560gaaacagggt cttgaaaact gtctaaatct aaaacaatgc taatgcaggt
ttaaatttaa 7620taaaataaaa tccaaaatct aacagccaag tcaaatctgt atgttttaac
atttaaaata 7680ttttaaagac gtcttttccc aggattcaac atgtgaaatc ttttctcagg
gatacacgtg 7740tgcctagatc ctcattgctt tagtttttta cagaggaatg aatataaaaa
gaaaatactt 7800aaattttatc cctcttacct ctataatcat acataggcat aattttttaa
cctaggctcc 7860agatagccat agaagaacca aacactttct gcgtgtgtga gaataatcag
agtgagattt 7920tttcacaagt acctgatgag ggttgagaca ggtagaaaaa gtgagagatc
tctatttatt 7980tagcaataat agagaaagca tttaagagaa taaagcaatg gaaataagaa
atttgtaaat 8040ttccttctga taactagaaa tagaggatcc agtttctttt ggttaaccta
aattttattt 8100cattttattg ttttatttta ttttatttta ttttattttg tgtaatcgta
gtttcagagt 8160gttagagctg aaaggaagaa gtaggagaaa catgcaaagt aaaagtataa
cactttcctt 8220actaaaccga ctgggtttcc aggtaggggc aggattcagg atgactgaca
gggcccttag 8280ggaacactga gaccctacgc tgacctcata aatgcttgct acctttgctg
ttttaattac 8340atcttttaat agcaggaagc agaactctgc acttcaaaag tttttcctca
cctgaggagt 8400taatttagta caaggggaaa aagtacaggg ggatgggaga aaggcgatca
cgttgggaag 8460ctatagagaa agaagagtaa attttagtaa aggaggttta aacaaacaaa
atataaagag 8520aaataggaac ttgaatcaag gaaatgattt taaaacgcag tattcttagt
ggactagagg 8580aaaaaaataa tctgagccaa gtagaagacc ttttcccctc ctacccctac
tttctaagtc 8640acagaggctt tttgttcccc cagacactct tgcagattag tccaggcaga
aacagttaga 8700tgtccccagt taacctccta tttgacacca ctgattaccc cattgatagt
cacactttgg 8760gttgtaagtg actttttatt tatttgtatt tttgactgca ttaagaggtc
tctagttttt 8820tatctcttgt ttcccaaaac ctaataagta actaatgcac agagcacatt
gatttgtatt 8880tattctattt ttagacataa tttattagca tgcatgagca aattaagaaa
aacaacaaca 8940aatgaatgca tatatatgta tatgtatgtg tgtatatata cacatatata
tatatatttt 9000ttttcttttc ttaccagaag gttttaatcc aaataaggag aagatatgct
tagaactgag 9060gtagagtttt catccattct gtcctgtaag tattttgcat attctggaga
cgcaggaaga 9120gatccatcta catatcccaa agctgaatta tggtagacaa agctcttcca
cttttagtgc 9180atcaatttct tatttgtgta ataagaaaat tgggaaaacg atcttcaata
tgcttaccaa 9240gctgtgattc caaatattac gtaaatacac ttgcaaagga ggatgttttt
agtagcaatt 9300tgtactgatg gtatggggcc aagagatata tcttagaggg agggctgagg
gtttgaagtc 9360caactcctaa gccagtgcca gaagagccaa ggacaggtac ggctgtcatc
acttagacct 9420caccctgtgg agccacaccc tagggttggc caatctactc ccaggagcag
ggagggcagg 9480agccagggct gggcataaaa gtcagggcag agccatctat tgcttacatt
tgcttctgac 9540acaactgtgt tcactagcaa cctcaaacag acaccatggt gcacctgact
cctgaggaga 9600agtctgccgt tactgccctg tggggcaagg tgaacgtgga tgaagttggt
ggtgaggccc 9660tgggcaggtt ggtatcaagg ttacaagaca ggtttaagga gaccaataga
aactgggcat 9720gtggagacag agatagtgga tccataactt cgtatagcat acattatacg
aagttatgtc 9780gacactagtg tcgagtcgcc gattaagtac tgtcgagtcg ccgattaagt
actgtcgagt 9840cgccgattaa gtactgtcga gtcgccgatt aagtactgtc gagtcgccga
ttaagtactg 9900tcgagccgag gtccacttcg catattaagg tgacgcgtgt ggcctcgaac
accgagcgac 9960cctgcagcga cccgcttaac ctgcagggcc gccaccatgg ccaatttact
gaccgtacac 10020caaaatttgc ctgcattacc ggtcgatgca acgagtgatg aggttcgcaa
gaacctgatg 10080gacatgttca gggatcgcca ggcgttttct gagcatacct ggaaaatgct
tctgtccgtt 10140tgccggtcgt gggcggcatg gtgcaagttg aataaccgga aatggtttcc
cgcagaacct 10200gaagatgttc gcgattatct tctatatctt caggcgcgcg gtctggcagt
aaaaactatc 10260cagcaacatt tgggccagct aaacatgctt catcgtcggt ccgggctgcc
acgaccaagt 10320gacagcaatg ctgtttcact ggttatgcgg cggatccgaa aagaaaacgt
tgatgccggt 10380gaacgtgcaa aacaggctct agcgttcgaa cgcactgatt tcgaccaggt
tcgttcactc 10440atggaaaata gcgatcgctg ccaggatata cgtaatctgg catttctggg
gattgcttat 10500aacaccctgt tacgtatagc cgaaattgcc aggatcaggg ttaaagatat
ctcacgtact 10560gacggtggga gaatgttaat ccatattggc agaacgaaaa cgctggttag
caccgcaggt 10620gtagagaagg cacttagcct gggggtaact aaactggtcg agcgatggat
ttccgtctct 10680ggtgtagctg atgatccgaa taactacctg ttttgccggg tcagaaaaaa
tggtgttgcc 10740gcgccatctg ccaccagcca gctatcaact cgcgccctgg aagggatttt
tgaagcaact 10800catcgattga tttacggcgc taaggatgac tctggtcaga gatacctggc
ctggtctgga 10860cacagtgccc gtgtcggagc cgcgcgagat atggcccgcg ctggagtttc
aataccggag 10920atcatgcaag ctggtggctg gaccaatgta aatattgtca tgaactatat
ccgtaacctg 10980gatagtgaaa caggggcaat ggtgcgcctg ctggaagatg gcgatggacc
ggtcgccacc 11040atggtgagca agggcgagga gctgttcacc ggggtggtgc ccatcctggt
cgagctggac 11100ggcgacgtaa acggccacaa gttcagcgtg tccggcgagg gcgagggcga
tgccacctac 11160ggcaagctga ccctgaagtt catctgcacc accggcaagc tgcccgtgcc
ctggcccacc 11220ctcgtgacca ccctgaccta cggcgtgcag tgcttcagcc gctaccccga
ccacatgaag 11280cagcacgact tcttcaagtc cgccatgccc gaaggctacg tccaggagcg
caccatcttc 11340ttcaaggacg acggcaacta caagacccgc gccgaggtga agttcgaggg
cgacaccctg 11400gtgaaccgca tcgagctgaa gggcatcgac ttcaaggagg acggcaacat
cctggggcac 11460aagctggagt acaactacaa cagccacaac gtctatatca tggccgacaa
gcagaagaac 11520ggcatcaagg tgaacttcaa gatccgccac aacatcgagg acggcagcgt
gcagctcgcc 11580gaccactacc agcagaacac ccccatcggc gacggccccg tgctgctgcc
cgacaaccac 11640tacctgagca cccagtccgc cctgagcaaa gaccccaacg agaagcgcga
tcacatggtc 11700ctgctggagt tcgtgaccgc cgccgggatc actctcggca tggacgagct
gtacaagtaa 11760catatgctcg acgataagct ttgcaaagat ggataaagtt ttaaacagag
aggaatcttt 11820gcagctaatg gaccttctag gtcttgaaag gagtgggaat tggctccggt
gcccgtcagt 11880gggcagagcg cacatcgccc acagtccccg agaagttggg gggaggggtc
ggcaattgaa 11940ccggtgccta gagaaggtgg cgcggggtaa actgggaaag tgatgtcgtg
tactggctcc 12000gcctttttcc cgagggtggg ggagaaccgt atataagtgc agtagtcgcc
gtgaacgttc 12060tttttcgcaa cgggtttgcc gccagaacac aggtaagtgc cgtgtgtggt
tcccgcgggc 12120ctggcctctt tacgggttat ggcccttgcg tgccttgaat tacttccact
ggctgcagta 12180cgtgattctt gatcccgagc ttcgggttgg aagtgggtgg gagagttcga
ggccttgcgc 12240ttaaggagcc ccttcgcctc gtgcttgagt tgaggcctgg cctgggcgct
ggggccgccg 12300cgtgcgaatc tggtggcacc ttcgcgcctg tctcgctgct ttcgataagt
ctctagccat 12360ttaaaatttt tgatgacctg ctgcgacgct ttttttctgg caagatagtc
ttgtaaatgc 12420gggccaagat ctgcacactg gtatttcggt ttttggggcc gcgggcggcg
acggggcccg 12480tgcgtcccag cgcacatgtt cggcgaggcg gggcctgcga gcgcggccac
cgagaatcgg 12540acgggggtag tctcaagctg gccggcctgc tctggtgcct ggcctcgcgc
cgccgtgtat 12600cgccccgccc tgggcggcaa ggctggcccg gtcggcacca gttgcgtgag
cggaaagatg 12660gccgcttccc ggccctgctg cagggagctc aaaatggagg acgcggcgct
cgggagagcg 12720ggcgggtgag tcacccacac aaaggaaaag ggcctttccg tcctcagccg
tcgcttcatg 12780tgactccacg gagtaccggg cgccgtccag gcacctcgat tagttctcga
gcttttggag 12840tacgtcgtct ttaggttggg gggaggggtt ttatgcgatg gagtttcccc
acactgagtg 12900ggtggagact gaagttaggc cagcttggca cttgatgtaa ttctccttgg
aatttgccct 12960ttttgagttt ggatcttggt tcattctcaa gcctcagaca gtggttcaaa
gtttttttct 13020tccatttcag gtgtcgtgag gaatttcgac atttaaatcc accatggcgg
gacacctggc 13080ttcggatttc gccttctcgc cccctccagg tggtggaggt gatgggccag
gggggccgga 13140gccgggctgg gttgatcctc ggacctggct aagcttccaa ggccctcctg
gagggccagg 13200aatcgggccg ggggttgggc caggctctga ggtgtggggg attcccccat
gccccccgcc 13260gtatgagttc tgtgggggga tggcgtactg tgggccccag gttggagtgg
ggctagtgcc 13320ccaaggcggc ttggagacct ctcagcctga gggcgaagca ggagtcgggg
tggagagcaa 13380ctccgatggg gcctccccgg agccctgcac cgtcacccct ggtgccgtga
agctggagaa 13440ggagaagctg gagcaaaacc cggaggagtc ccaggacatc aaagctctgc
agaaagaact 13500cgagcaattt gccaagctcc tgaagcagaa gaggatcacc ctgggatata
cacaggccga 13560tgtggggctc accctggggg ttctatttgg gaaggtattc agccaaacga
ccatctgccg 13620ctttgaggct ctgcagctta gcttcaagaa catgtgtaag ctgcggccct
tgctgcagaa 13680gtgggtggag gaagctgaca acaatgaaaa tcttcaggag atatgcaaag
cagaaaccct 13740cgtgcaggcc cgaaagagaa agcgaaccag tatcgagaac cgagtgagag
gcaacctgga 13800gaatttgttc ctgcagtgcc cgaaacccac actgcagcag atcagccaca
tcgcccagca 13860gcttgggctc gagaaggatg tggtccgagt gtggttctgt aaccggcgcc
agaagggcaa 13920gcgatcaagc agcgactatg cacaacgaga ggattttgag gctgctgggt
ctcctttctc 13980agggggacca gtgtcctttc ctctggcccc agggccccat tttggtaccc
caggctatgg 14040gagccctcac ttcactgcac tgtactcctc ggtccctttc cctgaggggg
aagcctttcc 14100ccctgtctcc gtcaccactc tgggctctcc catgcattca aacggatccg
gagccacgaa 14160cttctctctg ttaaagcaag caggagatgt tgaagaaaac cccgggccta
tgtacaacat 14220gatggagacg gagctgaagc cgccgggccc gcagcaaact tcggggggcg
gcggcggcaa 14280ctccaccgcg gcggcggccg gcggcaacca gaaaaacagc ccggaccgcg
tcaagcggcc 14340catgaatgcc ttcatggtgt ggtcccgcgg gcagcggcgc aagatggccc
aggagaaccc 14400caagatgcac aactcggaga tcagcaagcg cctgggcgcc gagtggaaac
ttttgtcgga 14460gacggagaag cggccgttca tcgacgaggc taagcggctg cgagcgctgc
acatgaagga 14520gcacccggat tataaatacc ggccccggcg gaaaaccaag acgctcatga
agaaggataa 14580gtacacgctg cccggcgggc tgctggcccc cggcggcaat agcatggcga
gcggggtcgg 14640ggtgggcgcc ggcctgggcg cgggcgtgaa ccagcgcatg gacagttacg
cgcacatgaa 14700cggctggagc aacggcagct acagcatgat gcaggaccag ctgggctacc
cgcagcaccc 14760gggcctcaat gcgcacggcg cagcgcagat gcagcccatg caccgctacg
acgtgagcgc 14820cctgcagtac aactccatga ccagctcgca gacctacatg aacggctcgc
ccacctacag 14880catgtcctac tcgcagcagg gcacccctgg catggctctt ggctccatgg
gttcggtggt 14940caagtccgag gccagctcca gcccccctgt ggttacctct tcctcccact
ccagggcgcc 15000ctgccaggcc ggggacctcc gggacatgat cagcatgtat ctccccggcg
ccgaggtgcc 15060ggaacccgcc gcccccagca gacttcacat gtcccagcac taccagagcg
gcccggtgcc 15120cggcacggcc attaacggca cactgcccct ctcacacatg ggatccggag
ccacgaactt 15180ctctctgtta aagcaagcag gagatgttga agaaaacccc gggcctatgg
ctgtcagcga 15240cgcgctgctc ccatctttct ccacgttcgc gtctggcccg gcgggaaggg
agaagacact 15300gcgtcaagca ggtgccccga ataaccgctg gcgggaggag ctctcccaca
tgaagcgact 15360tcccccagtg cttcccggcc gcccctatga cctggcggcg gcgaccgtgg
ccacagacct 15420ggagagcggc ggagccggtg cggcttgcgg cggtagcaac ctggcgcccc
tacctcggag 15480agagaccgag gagttcaacg atctcctgga cctggacttt attctctcca
attcgctgac 15540ccatcctccg gagtcagtgg ccgccaccgt gtcctcgtca gcgtcagcct
cctcttcgtc 15600gtcgccgtcg agcagcggcc ctgccagcgc gccctccacc tgcagcttca
cctatccgat 15660ccgggccggg aacgacccgg gcgtggcgcc gggcggcacg ggcggaggcc
tcctctatgg 15720cagggagtcc gctccccctc cgacggctcc cttcaacctg gcggacatca
acgacgtgag 15780cccctcgggc ggcttcgtgg ccgagctcct gcggccagaa ttggacccgg
tgtacattcc 15840gccgcagcag ccgcagccgc caggtggcgg gctgatgggc aagttcgtgc
tgaaggcgtc 15900gctgagcgcc cctggcagcg agtacggcag cccgtcggtc atcagcgtca
gcaaaggcag 15960ccctgacggc agccacccgg tggtggtggc gccctacaac ggcgggccgc
cgcgcacgtg 16020ccccaagatc aagcaggagg cggtctcttc gtgcacccac ttgggcgctg
gaccccctct 16080cagcaatggc caccggccgg ctgcacacga cttccccctg gggcggcagc
tccccagcag 16140gactaccccg accctgggtc ttgaggaagt gctgagcagc agggactgtc
accctgccct 16200gccgcttcct cccggcttcc atccccaccc ggggcccaat tacccatcct
tcctgcccga 16260tcagatgcag ccgcaagtcc cgccgctcca ttaccaagag ctcatgccac
ccggttcctg 16320catgccagag gagcccaagc caaagagggg aagacgatcg tggccccgga
aaaggaccgc 16380cacccacact tgtgattacg cgggctgcgg caaaacctac acaaagagtt
cccatctcaa 16440ggcacacctg cgaacccaca caggtgagaa accttaccac tgtgactggg
acggctgtgg 16500atggaaattc gcccgctcag atgaactgac caggcactac cgtaaacaca
cggggcaccg 16560cccgttccag tgccaaaaat gcgaccgagc attttccagg tcggaccacc
tcgccttaca 16620catgaagagg catttttaag gcgcgccata acttcgtata gcatacatta
tacgaagtta 16680tctgcaggaa gactcttggg tttctgatag gcactgactc tctctgccta
ttggtctatt 16740ttcccaccct taggctgctg gtggtctacc cttggaccca gaggttcttt
gagtcctttg 16800gggatctgtc cactcctgat gctgttatgg gcaaccctaa ggtgaaggct
catggcaaga 16860aagtgctcgg tgcctttagt gatggcctgg ctcacctgga caacctcaag
ggcacctttg 16920ccacactgag tgagctgcac tgtgacaagc tgcacgtgga tcctgagaac
ttcagggtga 16980gtctatggga cccttgatgt tttctttccc cttcttttct atggttaagt
tcatgtcata 17040ggaaggggag aagtaacagg gtacagttta gaatgggaaa cagacgaatg
attgcatcag 17100tgtggaagtc tcaggatcgt tttagtttct tttatttgct gttcataaca
attgttttct 17160tttgtttaat tcttgctttc tttttttttc ttctccgcaa tttttactat
tatacttaat 17220gccttaacat tgtgtataac aaaaggaaat atctctgaga tacattaagt
aacttaaaaa 17280aaaactttac acagtctgcc tagtacatta ctatttggaa tatatgtgtg
cttatttgca 17340tattcataat ctccctactt tattttcttt tatttttaat tgatacataa
tcattataca 17400tatttatggg ttaaagtgta atgttttaat atgtgtacac atattgacca
aatcagggta 17460attttgcatt tgtaatttta aaaaatgctt tcttctttta atatactttt
ttgtttatct 17520tatttctaat actttcccta atctctttct ttcagggcaa taatgataca
atgtatcatg 17580cctctttgca ccattctaaa gaataacagt gataatttct gggttaaggc
aatagcaata 17640tttctgcata taaatatttc tgcatataaa ttgtaactga tgtaagaggt
ttcatattgc 17700taatagcagc tacaatccag ctaccattct gcttttattt tatggttggg
ataaggctgg 17760attattctga gtccaagcta ggcccttttg ctaatcatgt tcatacctct
tatcttcctc 17820ccacagctcc tgggcaacgt gctggtctgt gtgctggccc atcactttgg
caaagaattc 17880accccaccag tgcaggctgc ctatcagaaa gtggtggctg gtgtggctaa
tgccctggcc 17940cacaagtatc actaagctcg ctttcttgct gtccaatttc tattaaaggt
tcctttgttc 18000cctaagtcca actactaaac tgggggatat tatgaagggc cttgagcatc
tggattctgc 18060ctaataaaaa acatttattt tcattgcaat gatgtattta aattatttct
gaatatttta 18120ctaaaaaggg aatgtgggag gtcagtgcat ttaaaacata aagaaatgaa
gagctagttc 18180aaaccttggg aaaatacact atatcttaaa ctccatgaaa gaaggtgagg
ctgcaaacag 18240ctaatgcaca ttggcaacag ccctgatgcc tatgccttat tcatccctca
gaaaaggatt 18300caagtagagg cttgatttgg aggttaaagt tttgctatgc tgtattttac
attacttatt 18360gttttagctg tcctcatgaa tgtcttttca ctacccattt gcttatcctg
catctctcag 18420ccttgactcc actcagttct cttgcttaga gataccacct ttcccctgaa
gtgttccttc 18480catgttttac ggcgagatgg tttctcctcg cctggccact cagccttagt
tgtctctgtt 18540gtcttataga ggtctacttg aagaaggaaa aacagggggc atggtttgac
tgtcctgtga 18600gcccttcttc cctgcctccc ccactcacag tgacccggaa tctgcagtgc
tagtctcccg 18660gaactatcac tctttcacag tctgctttgg aaggactggg cttagtatga
aaagttagga 18720ctgagaagaa tttgaaaggg ggctttttgt agcttgatat tcactactgt
cttattaccc 18780tatcataggc ccaccccaaa tggaagtccc attcttcctc aggatgttta
agattagcat 18840tcaggaagag atcagaggtc tgctggctcc cttatcatgt cccttatggt
gcttctggct 18900ctgcagttat tagcatagtg ttaccatcaa ccaccttaac ttcatttttc
ttattcaata 18960cctaggtagg tagatgctag attctggaaa taaaatatga gtctcaagtg
gtccttgtcc 19020tctctcccag tcaaattctg aatctagttg gcaagattct gaaatcaagg
catataatca 19080gtaataagtg atgatagaag ggtatataga agaattttat tatatgagag
ggtgaaacct 19140aaaatgaaat gaaatcagac ccttgtctta caccataaac aaaaataaat
ttgaatgggt 19200taaagaatta aactaagacc taaaaccata aaaattttta aagaaatcaa
aagaagaaaa 19260ttctaatatt catgttgcag ccgttttttg aatttgatat gagaagcaaa
ggcaacaaaa 19320ggaaaaataa agaagtgagg ctacatcaaa ctaaaaaatt tccacacaaa
aaagaaaaca 19380atgaacaaat gaaaggtgaa ccatgaaatg gcatatttgc aaaccaaata
tttcttaaat 19440attttggtta atatccaaaa tatataagaa acacagatga ttcaataaca
aacaaaaaat 19500taaaaatagg aaaataaaaa aattaaaaag aagaaaatcc tgccatttat
gcgagaattg 19560atgaacctgg aggatgtaaa actaagaaaa ataagcctga cacaaaaaga
caaatactac 19620acaaccttgc tcatatgtga aacataaaaa agtcactctc atggaaacag
acagtagagg 19680tatggtttcc aggggttggg ggtgggagaa tcaggaaact attactcaaa
gggtataaaa 19740tttcagttat gtgggatgaa taaattctag atatctaatg tacagcatcg
tgactgtagt 19800taattgtact gtaagtatat ttaaaatttg caaagagagt agattttttt
gtttttttag 19860atggagtttt gctcttgttg tccaggctgg agtgcaatgg caagatcttg
gctcactgca 19920acctccgcct cctgggttca agcaaatctc ctgcctcagc ctcccgagta
gctgggatta 19980caggcatgcg acaccatgcc cagctaattt tgtattttta gtagagacgg
ggtttctcca 20040tgttggtcag gctgatccgc ctcctcggcc accaaagggc tgggattaca
ggcgtgacca 20100ccgggcctgg ccgagagtag atcttaaaag catttaccac aagaaaaagg
taactatgtg 20160agataatggg tatgttaatt agcttgattg tggtaatcat ttcacaaggt
atacatatat 20220taaaacatca tgttgtacac cttaaatata tacaattttt atttgtgaat
gatacctcaa 20280taaagttgaa gaataataaa aaagaataga catcacatga attaaaaaac
taaaaaataa 20340aaaaatgcat cttgatgatt agaattgcat tcttgatttt tcagatacaa
atatccattt 20400gactgtttac tcttttccaa aacaatacaa taaattttag cactttatct
tcattttccc 20460cttcccaatc tataatttta tatatatata ttttagatat tttgtatagt
tttactccct 20520agattttcta gtgttattat taaatagtga agaaatgttt acacttatgt
acaaaatgtt 20580ttgcatgctt ttcttcattt ctaacattct ctctaagttt attctatttt
ttcctgatta 20640tccttaatat tatctctttc tgctggaaat atattgttac ttttggttta
tctaaaaatg 20700gcttcatttt cttcattcta aaatcatgtt aaattaatac cactcatgtg
taagtaagat 20760agtggaataa atagaaatcc aaaaactaaa tctcacaaaa tataataatg
tgatatataa 20820aaatatagct tttaaattta gcttggaaat aaaaaacaaa cagtaattga
acaactatac 20880tttttgaaaa gagtaaagtg aaatgcttaa ctgcatatac cacaatcgat
tacacaatta 20940ggtgtgaagg taaaattcag tcacgaaaaa actagaataa aaatatggga
agacatgtat 21000ataatcttag agataacagt gttatttaat tatcaactag ttctagagcg
gccgccaccg 21060cggtggagct ccagcttttg ttccctttag tgagggttaa tttcgagctt
ggcgtaatca 21120tggtcatagc tgtttcctgt gtgaaattgt tatccgctca caattccaca
caacatacga 21180gccggaagca taaagtgtaa agcctggggt gcctaatgag tgagctaact
cacattaatt 21240gcgttgcgct cactgcccgc tttccagtcg ggaaacctgt cgtgccagct
gcattaatga 21300atcggccaac gcgcggggag aggcggtttg cgtattgggc gctcttccgc
ttcctcgctc 21360actgactcgc tgcgctcggt cgttcggctg cggcgagcgg tatcagctca
ctcaaaggcg 21420gtaatacggt tatccacaga atcaggggat aacgcaggaa agaacatgtg
agcaaaaggc 21480cagcaaaagg ccaggaaccg taaaaaggcc gcgttgctgg cgtttttcca
taggctccgc 21540ccccctgacg agcatcacaa aaatcgacgc tcaagtcaga ggtggcgaaa
cccgacagga 21600ctataaagat accaggcgtt tccccctgga agctccctcg tgcgctctcc
tgttccgacc 21660ctgccgctta ccggatacct gtccgccttt ctccctt
21697
User Contributions:
Comment about this patent or add new information about this topic: