Patent application title: NOVEL INFLUENZA HEMAGGLUTININ PROTEIN-BASED VACCINES
Inventors:
Gary J. Nabel (Washington, DC, US)
Masaru Kanekiyo (Silver Spring, MD, US)
Chih-Jen Wei (Potomac, MD, US)
Patrick M. Mctamney (Bethesda, MD, US)
Hadi M. Yassine (Rockville, MD, US)
Jeffrey C. Boyington (Clarksburg, MD, US)
IPC8 Class: AG01N33569FI
USPC Class:
435 5
Class name: Chemistry: molecular biology and microbiology measuring or testing process involving enzymes or micro-organisms; composition or test strip therefore; processes of forming such composition or test strip involving virus or bacteriophage
Publication date: 2014-03-13
Patent application number: 20140072958
Abstract:
Novel vaccines are provided that elicit broadly neutralizing
anti-influenza antibodies. Some vaccines comprise nanoparticles that
display hemagglutinin trimers from influenza virus on their surface. The
nanoparticles comprise fusion proteins comprising a monomeric subunit of
ferritin joined to at least a portion of an influenza hemagglutinin
protein. Some portions comprise the ectodomain while some portions are
limited to the stem region. The fusion proteins self-assemble to form the
hemagglutinin-displaying nanoparticles. Some vaccines comprise only the
stem region of an influenza hemagglutinin protein joined to a
trimerization domain. Such vaccines can be used to vaccinate an
individual against infection by heterologous influenza viruses and
influenza virus that are antigenically divergent from the virus from
which the nanoparticle hemagglutinin protein was obtained. Also provided
are fusion proteins and nucleic acid molecules encoding such proteins.
Finally, also provided are assays using nanoparticles of the invention to
detect anti-influenza antibodies.Claims:
1. A method for detecting anti-influenza virus antibodies in a sample,
the method comprising: a. contacting at least a portion of the sample
with a nanoparticle comprising a fusion protein, wherein the fusion
protein comprises at least 25 contiguous amino acids from a monomeric
ferritin subunit protein joined to at least one epitope from an influenza
hemagglutinin (HA) protein such that the nanoparticle comprises trimers
of the influenza virus HA protein epitope on its surface, and wherein the
at least a portion of the sample and the nanoparticle are contacted under
conditions suitable for forming a complex between the nanoparticle and
the anti-influenza virus antibodies, if present; and, b. analyzing the
contacted sample for the presence of a nanoparticle/antibody complex,
wherein the presence of such a complex indicates the sample contains
anti-influenza antibodies.
2. The method of claim 1, wherein the monomeric ferritin subunit protein is selected from the group consisting of a bacterial ferritin, a plant ferritin, an algal ferritin and a mammalian ferritin.
3. The nanoparticle of claim 1, wherein the monomeric ferritin subunit protein comprises at least 25 contiguous amino acids an amino acid sequence selected from the group consisting of SEQ ID NO:2 and SEQ ID NO:5, where in the fusion protein is capable of self-assembling into nanoparticles.
4. The nanoparticle of claim 1, wherein the hemagglutinin protein is from an influenza virus selected from the group consisting of A/New Calcdonia/20/1999 (1999 NC, H1), A/California/04/2009 (2009 CA, H1), A/Singapore/1/1957 (1957 Sing, H2), A/Hong Kong/1/1968 (1968 HK, H3), A/Brisbane/10/2007 (2007 Bris, H3), A/Indonesia/05/2005 (2005 Indo, H5), B/Florida/4/2006 (2006 Flo, B), A/Perth/16/2009 (2009 Per, H3), A/Brisbane/59/2007 (2007 Bris, H1), B/Brisbane/60/2008 (2008 Bris, B).
5. The nanoparticle of claim 1, wherein the fusion protein comprises an amino acid sequence at least 80% identical to a sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, SEQ ID NO:68, wherein the nanoparticle elicits an immune response against an influenza virus.
6. The nanoparticle of claim 1, wherein the nanoparticle comprises a second fusion protein comprising a second influenza hemagglutinin protein, wherein the first and second influenza hemagglutinin proteins are from different types, sub-types or strains of influenza viruses.
7. The method of claim 1, wherein the step of analyzing comprises an assay selected from the group consisting of a hemagglutinin inhibition assay, an immunodiffusion assay, an enzyme-linked immunoassay, a radioimmunoassay, a fluorescence immunoassay, a chemiluminescent assay, a lateral flow assay, a flow-through assay, a precipitation assay, an immunoprecipitation assay, a BioCoreJ assay (e.g., using colloidal gold), an immunodot assay (e.g., CMG=s Immunodot System, Fribourg, Switzerland), an immunoblot assay (e.g., a western blot), an phosphorescence assay, a chromatography assay, a PAGe-based assay, a surface plasmon resonance assay, a spectrophotometric assay, and an electronic sensory assay.
8. A method for identifying an individual having anti-influenza antibodies, comprising: a. contacting at least a portion of a sample from an individual with a nanoparticle comprising a fusion protein, wherein the fusion protein comprises at least 25 contiguous amino acids from a monomeric ferritin subunit protein joined to at least one epitope from an influenza hemagglutinin (HA) protein such that the nanoparticle comprises trimers of the influenza virus HA protein epitope on its surface, and wherein the at least a portion of the sample and the nanoparticle are contacted under conditions suitable for forming a complex between the nanoparticle and the anti-influenza virus antibodies, if present; b. analyzing the contacted sample for the presence of a nanoparticle/antibody complex, wherein the presence of such a complex indicates the individual has anti-influenza antibodies.
9. The method of claim 8, wherein the monomeric ferritin subunit protein is selected from the group consisting of a bacterial ferritin, a plant ferritin, an algal ferritin and a mammalian ferritin.
10. The nanoparticle of claim 8, wherein the monomeric ferritin subunit protein comprises at least 25 contiguous amino acids an amino acid sequence selected from the group consisting of SEQ ID NO:2 and SEQ ID NO:5, where in the fusion protein is capable of self-assembling into nanoparticles.
11. The nanoparticle of claim 8, wherein the hemagglutinin protein is from an influenza virus selected from the group consisting of A/New Calcdonia/20/1999 (1999 NC, H1), A/California/04/2009 (2009 CA, H1), A/Singapore/1/1957 (1957 Sing, H2), A/Hong Kong/1/1968 (1968 HK, H3), A/Brisbane/10/2007 (2007 Bris, H3), A/Indonesia/05/2005 (2005 Indo, H5), B/Florida/4/2006 (2006 Flo, B), A/Perth/16/2009 (2009 Per, H3), A/Brisbane/59/2007 (2007 Bris, H1), B/Brisbane/60/2008 (2008 Bris, B).
12. The nanoparticle of claim 8, wherein the fusion protein comprises an amino acid sequence at least 80% identical to a sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, SEQ ID NO:68, wherein the nanoparticle elicits an immune response against an influenza virus.
13. The nanoparticle of claim 8, wherein the nanoparticle comprises a second fusion protein comprising a second influenza hemagglutinin protein, wherein the first and second influenza hemagglutinin proteins are from different types, sub-types or strains of influenza viruses.
14. The method of claim 8, wherein the step of analyzing comprises an assay selected from the group consisting of a hemagglutinin inhibition assay, an immunodiffusion assay, an enzyme-linked immunoassay, a radioimmunoassay, a fluorescence immunoassay, a chemiluminescent assay, a lateral flow assay, a flow-through assay, a precipitation assay, an immunoprecipitation assay, a BioCoreJ assay (e.g., using colloidal gold), an immunodot assay (e.g., CMG=s Immunodot System, Fribourg, Switzerland), an immunoblot assay (e.g., a western blot), an phosphorescence assay, a chromatography assay, a PAGe-based assay, a surface plasmon resonance assay, a spectrophotometric assay, and an electronic sensory assay.
15. A method for measuring the response of an individual to a vaccine, comprising: a. administering to the individual a vaccine for influenza virus; b. contacting at least a portion of a sample from the individual with a nanoparticle comprising a fusion protein, wherein the fusion protein comprises at least 25 contiguous amino acids from a monomeric ferritin subunit protein joined to at least one epitope from an influenza hemagglutinin (HA) protein such that the nanoparticle comprises trimers of the influenza virus HA protein epitope on its surface, and wherein the at least a portion of the sample and the nanoparticle are contacted under conditions suitable for forming a complex between the nanoparticle and the anti-influenza virus antibodies, if present; and, c. determining the level of antibody present in the sample by determining the level of nanoparticle/antibody complex; wherein an increase in the level of antibody in the sample over the pre-vaccination level of antibody indicates the vaccine induced an immune response in the individual.
16. The method of claim 15, wherein the monomeric ferritin subunit protein is selected from the group consisting of a bacterial ferritin, a plant ferritin, an algal ferritin and a mammalian ferritin.
17. The nanoparticle of claim 15, wherein the monomeric ferritin subunit protein comprises at least 25 contiguous amino acids an amino acid sequence selected from the group consisting of SEQ ID NO:2 and SEQ ID NO:5, where in the fusion protein is capable of self-assembling into nanoparticles.
18. The nanoparticle of claim 15, wherein the hemagglutinin protein is from an influenza virus selected from the group consisting of A/New Calcdonia/20/1999 (1999 NC, H1), A/California/04/2009 (2009 CA, H1), A/Singapore/1/1957 (1957 Sing, H2), A/Hong Kong/1/1968 (1968 HK, H3), A/Brisbane/10/2007 (2007 Bris, H3), A/Indonesia/05/2005 (2005 Indo, H5), B/Florida/4/2006 (2006 Flo, B), A/Perth/16/2009 (2009 Per, H3), A/Brisbane/59/2007 (2007 Bris, H1), B/Brisbane/60/2008 (2008 Bris, B).
19. The nanoparticle of claim 15, wherein the fusion protein comprises an amino acid sequence at least 80% identical to a sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, SEQ ID NO:68, wherein the nanoparticle elicits an immune response against an influenza virus.
20. The method of claim 15, wherein the step of analyzing comprises an assay selected from the group consisting of a hemagglutinin inhibition assay, an immunodiffusion assay, an enzyme-linked immunoassay, a radioimmunoassay, a fluorescence immunoassay, a chemiluminescent assay, a lateral flow assay, a flow-through assay, a precipitation assay, an immunoprecipitation assay, a BioCoreJ assay (e.g., using colloidal gold), an immunodot assay (e.g., CMG=s Immunodot System, Fribourg, Switzerland), an immunoblot assay (e.g., a western blot), an phosphorescence assay, a chromatography assay, a PAGe-based assay, a surface plasmon resonance assay, a spectrophotometric assay, and an electronic sensory assay.
Description:
CROSS REFERENCE TO RELATED APPLICATIONS
[0001] This application claims priority under 35 U.S.C. §119(e) to P.C.T. Patent Application Serial No. PCT/US12/56822 entitled "NOVEL INFLUENZA HEMAGGLUTININ PROTEIN-BASED VACCINES" filed Sep. 24, 2012; which claims priority to U.S. Provisional Patent Application Ser. No. 61/661,209 entitled "SELF-ASSEMBLED FERRITIN NANOPARTICLES EXPRESSING HEMAGGLUTININ AS AN INFLUENZA VACCINE" filed Jun. 18, 2012; and U.S. Provisional Patent Application Ser. No. 61/538,663 entitled "SELF-ASSEMBLED FERRITIN NANOPARTICLES EXPRESSING HEMAGGLUTININ AS AN INFLUENZA VACCINE" filed Sep. 23, 2011, all of which are incorporated herein in their entirety by this reference.
SUMMARY OF THE INVENTION
[0002] The present invention provides novel hemagglutinin protein-based influenza vaccines that are easily manufactured, potent, and which elicit broadly neutralizing influenza antibodies. In particular, the present invention provides influenza hemagglutinin proteins, and portions thereof, that are useful in inducing the production of neutralizing antibodies. It also provides novel HA-ferritin nanoparticle (np) vaccines. Such nanoparticles comprise fusion proteins, each of which comprises a monomeric subunit of ferritin joined to an immunogenic portion of an influenza hemagglutinin protein. Because such nanoparticles display influenza hemagglutinin protein on their surface, they can be used to vaccinate an individual against influenza virus.
[0003] In one embodiment, the invention is a nanoparticle that comprises a fusion protein, and in this embodiment the fusion protein comprises at least 25 contiguous amino acids from a monomeric ferritin subunit protein joined to a first influenza hemagglutinin (HA) protein, such that the nanoparticle comprises influenza virus HA protein trimers on its surface. The nanoparticle can form an octahedron, which can consist of 24 subunits having 432 symmetry. Further, the monomeric ferritin subunit protein can be selected from a bacterial ferritin, a plant ferritin, an algal ferritin, an insect ferritin, a fungal ferritin and a mammalian ferritin, and in a preferred embodiment, is a Helicobacter pylori ferritin protein.
[0004] In this embodiment, the monomeric ferritin subunit protein can comprise at least 25 contiguous amino acids of an amino acid sequence selected from SEQ ID NO:2 and SEQ ID NO:5 or can comprise an amino acid at least about 80% identical, at least about 85% identical, at least about 90% identical, at least about 95% identical, at least about 97% identical, at least about 99% identical to those sequences or can comprise those sequences. In another embodiment, the monomeric subunit comprises a region corresponding to amino acids 5-167 of SEQ ID NO:2.
[0005] In this embodiment, the hemagglutinin protein can comprise at least 25 contiguous amino acids from the hemagglutinin protein of an influenza virus selected from A/New Calcdonia/20/1999 (1999 NC, H1), A/California/04/2009 (2009 CA, H1), A/Singapore/1/1957 (1957 Sing, H2), A/Hong Kong/1/1968 (1968 HK, H3), A/Brisbane/10/2007 (2007 Bris, H3), A/Indonesia/05/2005 (2005 Indo, H5), B/Florida/4/2006 (2006 Flo, B), A/Perth/16/2009 (2009 Per, H3), A/Brisbane/59/2007 (2007 Bris, H1), B/Brisbane/60/2008 (2008 Bris, B). Also, the hemagglutinin protein can comprise an amino acid sequence that is selected from the amino acid sequences of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38 or one that is at least 80% identical, at least about 85% identical, at least about 90% identical, at least about 95% identical, at least about 97% identical, at least about 99% identical thereto. Alternatively, the hemagglutinin protein can comprise an amino acid sequence that is selected from the amino acid sequences of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95 and SEQ ID NO:98 or one that is at least 80% identical, at least about 85% identical, at least about 90% identical, at least about 95% identical, at least about 97% identical, at least about 99% identical thereto.
[0006] In this embodiment, the hemagglutinin protein can be capable of eliciting an immune response to a protein comprising an amino acid sequence selected from SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38 or it can comprise a region selected from a region capable of allowing formation of a hemagglutinin trimer, a stem region, an ectodomain, and a region comprising the amino acid sequence from the amino acid residue immediately distal to the last amino acid of the second helical coiled coil to the amino acid residue proximal to the first amino acid of the transmembrane domain.
[0007] The hemagglutinin protein can also comprise a hemagglutinin spike domain, a region corresponding to amino acids 1-519 of SEQ ID NO:8 or an amino acid sequence selected from the group consisting of amino acids 1-519 of SEQ ID NO:8 and SEQ ID NO:11.
[0008] In this embodiment, the fusion protein can comprise a linker sequence.
[0009] In this embodiment, the nanoparticle can elicit an immune response against a stem region of influenza hemagglutinin, a spike of influenza hemagglutinin, an influenza virus strain that is heterologous to the strain influenza virus from which the hemagglutinin protein was obtained or an influenza virus that is antigenically divergent from the influenza virus from which the hemagglutinin protein was obtained.
[0010] In this embodiment, the fusion protein can comprise an amino acid sequence at least about 80% identical, at least about 85% identical, at least about 90% identical, at least about 95% identical, at least about 97% identical, at least about 99% identical to a sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, and SEQ ID NO:68, wherein the nanoparticle elicits an immune response against an influenza virus or can comprise an amino acid sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, and SEQ ID NO:68. The fusion protein can also comprise an amino acid sequence at least 80% identical, at least about 85% identical, at least about 90% identical, at least about 95% identical, at least about 97% identical, at least about 99% identical to a sequence selected from the group consisting of SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128, wherein the nanoparticle elicits an immune response against an influenza virus.
[0011] In this embodiment, the nanoparticle can comprise a second fusion protein comprising a second influenza hemagglutinin protein, wherein the first and second influenza hemagglutinin proteins are from different Types, from different sub-types or different strains of influenza viruses.
[0012] Another embodiment of the present invention is a vaccine composition comprising any of the foregoing nanoparticle. The vaccine composition can further comprise at least one additional nanoparticle that comprises at least one hemagglutinin protein from a different strain of influenza than the first hemagglutinin protein and the second hemagglutinin protein.
[0013] A further embodiment of the invention is a method to produce a vaccine against influenza virus. The method includes expressing a fusion protein comprising a monomeric ferritin protein joined to an influenza hemagglutinin protein under conditions such that the fusion proteins form a nanoparticle displaying hemagglutinin trimers on its surface and recovering the nanoparticle.
[0014] The invention also includes a method to vaccinate an individual against influenza that includes administering a nanoparticle to an individual such that the nanoparticle elicits an immune response against influenza virus. In this embodiment, the nanoparticle comprises a monomeric subunit of ferritin joined to an influenza hemagglutinin protein and the nanoparticle displays influenza hemagglutinin trimers on its surface. In this embodiment, the nanoparticle can elicit an immune response to an influenza virus strain that is heterologous to the sub-type or strain of or that is antigenically divergent from the influenza virus from which the hemagglutinin protein was obtained.
[0015] This method can further include administering to the individual a first vaccine composition and then at a later time, administering a second vaccine composition comprising a nanoparticle that comprises an HA-SS-ferritin fusion protein. The HA SS-ferritin fusion protein can comprise an amino acid sequence selected from SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95 and SEQ ID NO:98 or one that is at least 80% identical, at least 85% identical, at least 90% identical, at least 95% identical, at least 97% identical or at least 99% identical thereto, wherein the HA 55-ferritin fusion protein elicits an immune response to an influenza virus. The HA 55-ferritin fusion protein can comprise an amino acid sequence selected from the group consisting of SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128, or one at least 80% identical, at least 85% identical, at least 90% identical, at least 95% identical, at least 97% identical or at least 99% identical thereto, wherein the HA 55-ferritin fusion protein elicits an immune response to an influenza virus.
[0016] In this method, the first vaccine composition can comprise a nanoparticle comprising an ectodomain from the hemagglutinin protein of an influenza virus selected from the group consisting of A/New Calcdonia/20/1999 (1999 NC, H1), A/California/04/2009 (2009 CA, H1), A/Singapore/1/1957 (1957 Sing, H2), A/Hong Kong/1/1968 (1968 HK, H3), A/Brisbane/10/2007 (2007 Bris, H3), A/Indonesia/05/2005 (2005 Indo, H5), B/Florida/4/2006 (2006 Flo, B), A/Perth/16/2009 (2009 Per, H3), A/Brisbane/59/2007 (2007 Bris, H1), B/Brisbane/60/2008 (2008 Bris, B). Alternatively, the hemagglutinin of the first vaccine composition protein can comprise an amino acid sequence selected from SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38 or one that is at least about 80% identical, at least about 85% identical, at least about 90% identical, at least about 95% identical, at least about 97% identical, at least about 99% identical thereto. Further, the first vaccine composition can comprise an HA-ferritin fusion protein comprising an amino acid sequence selected from SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, and SEQ ID NO:68 or an amino acid sequence that is at least 80% identical, at least about 85% identical, at least about 90% identical, at least about 95% identical, at least about 97% identical, at least about 99% identical thereto, wherein the nanoparticle elicits an immune response against an influenza virus.
[0017] Administration of the boosting composition is generally weeks or months after administration of the priming composition.
[0018] A further embodiment of the present invention is a fusion protein comprising a monomeric ferritin subunit protein joined to an influenza hemagglutinin protein. The monomeric ferritin subunit protein can be selected from a bacterial ferritin, a plant ferritin, an algal ferritin, an insect ferritin, a fungal ferritin and a mammalian ferritin or can be a monomeric subunit of a Helicobacter pylori ferritin protein. The monomeric ferritin subunit protein can comprise a domain that allows the fusion protein to self-assemble into nanoparticles. In this embodiment, the monomeric ferritin subunit protein can comprise SEQ ID NO:2 or SEQ ID NO:5 or comprise at least 25 contiguous amino acids from or be at least about 80% identical, at least about 85% identical, at least about 90% identical, at least about 95% identical, at least about 97% identical, at least about 99% to a sequence selected from SEQ ID NO:2 and SEQ ID NO:5 and the fusion protein can be capable of self-assembling into nanoparticles. Additionally, the monomeric subunit can comprise a region corresponding to amino acids 5-167 of SEQ ID NO:2.
[0019] In this embodiment, the hemagglutinin protein can comprise at least 25 amino acids from an influenza virus selected from A/New Calcdonia/20/1999 (1999 NC, H1), A/California/04/2009 (2009 CA, H1), A/Singapore/1/1957 (1957 Sing, H2), A/Hong Kong/1/1968 (1968 HK, H3), A/Brisbane/10/2007 (2007 Bris, H3), A/Indonesia/05/2005 (2005 Indo, H5), B/Florida/4/2006 (2006 Flo, B), A/Perth/16/2009 (2009 Per, H3), A/Brisbane/59/2007 (2007 Bris, H1), and B/Brisbane/60/2008 (2008 Bris, B). Alternatively, the hemagglutinin protein can be capable of eliciting an immune response to a protein comprising an amino acid sequence selected from SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38 or one that is at least about 80% identical, at least about 85% identical, at least about 90% identical, at least about 95% identical, at least about 97% identical, at least about 99% thereto.
[0020] In this embodiment, the fusion protein can comprise an amino acid sequence selected from SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, and SEQ ID NO:68 or one that is at least about 80% identical, at least about 85% identical, at least about 90% identical, at least about 95% identical, at least about 97% identical, at least about 99% thereto.
[0021] Further in this embodiment, the hemagglutinin protein can comprise a region selected from a region capable of allowing trimerization of the hemagglutinin protein, a stem region, an ectodomain, and a region comprising the amino acid sequence from the amino acid residue immediately distal to the last amino acid of the second helical coiled coil to the amino acid residue proximal to the first amino acid of the transmembrane domain. The hemagglutinin protein alternatively can comprise a region corresponding to amino acids 1-519 of SEQ ID NO:8, an amino acid sequence selected from the group consisting of amino acids 1-519 of SEQ ID NO:8 and SEQ ID NO:11, or a hemagglutinin spike domain. Further, the hemagglutinin protein can comprise the stem region from an influenza virus selected from A/New Calcdonia/20/1999 (1999 NC, H1), A/California/04/2009 (2009 CA, H1), A/Singapore/1/1957 (1957 Sing, H2), A/Hong Kong/1/1968 (1968 HK, H3), A/Brisbane/10/2007 (2007 Bris, H3), A/Indonesia/05/2005 (2005 Indo, H5), B/Florida/4/2006 (2006 Flo, B), A/Perth/16/2009 (2009 Per, H3), A/Brisbane/59/2007 (2007 Bris, H1), or B/Brisbane/60/2008 (2008 Bris, B). The hemagglutinin protein can also comprise an amino acid sequence at least about 80% identical, at least about 85% identical, at least about 90% identical, at least about 95% identical, at least about 97% identical, at least about 99% to SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98.
[0022] In this embodiment, the fusion protein can comprise one or more linker sequences or an amino acid sequence of selected from the group consisting of SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128 or a sequence that is at least about 80% identical, at least about 85% identical, at least about 90% identical, at least about 95% identical, at least about 97% identical, at least about 99% thereto.
[0023] A further embodiment of the present invention is a nucleic acid molecule encoding any of the fusion proteins described above. In this embodiment, the nucleic acid molecule can be functionally linked to a promoter. Other embodiments of the invention include recombinant cells and viruses that comprise such nucleic acid molecules.
[0024] Another embodiment of the invention is a protein comprising an amino acid sequence at least 80% identical, at least about 85% identical, at least about 90% identical, at least about 95% identical, at least about 97% identical, at least about 99% to an amino acid selected from SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98, wherein the protein is joined to one or more trimerization domains. In this embodiment, the protein can be joined to at least a portion of the head region of an influenza hemagglutinin protein, comprise one or more linker regions or elicit an immune response against an influenza virus. A further embodiment is a nucleic acid molecule encoding such a protein.
BACKGROUND
[0025] Protective immune responses induced by vaccination against influenza virus are primarily directed to the viral hemagglutinin (HA) protein, which is a glycoprotein on the surface of the virus responsible for interaction of the virus with host cell receptors. HA proteins on the virus surface are trimers of hemagglutinin protein monomers that are enzymatically cleaved to yield amino-terminal HA1 and carboxy-terminal HA2 polypeptides. The globular head consists exclusively of the major portion of the HA1 polypeptide, whereas the stem that anchors the hemagglutinin protein into the viral lipid envelope is comprised of HA2 and part of HA1. The globular head of a hemagglutinin protein includes two domains: the receptor binding domain (RBD), an ˜148-amino acid residue domain that includes the sialic acid-binding site, and the vestigial esterase domain, a smaller ˜75-amino acid residue region just below the RBD. The top part of the RBD adjacent to the 2,6-sialic acid recognition sites includes a large region (amino acids 131-143, 170-182, 205-215 and 257-262, 1918 numbering) (referred to herein as the RBD-A region) of over 6000 A2 per trimer that is 95% conserved between A/South Carolina/1/1918 (1918 SC) and A/California/04/2009 (2009 CA) pandemic strains. The globular head includes several antigenic sites that include immunodominant epitopes. Examples include the Sa, Sb, Ca1, Ca2 and Cb antigenic sites (see, for example, Caton A J et al, 1982, Cell 31, 417-427). The RBD-A region includes the Sa antigenic site and part of the Sb antigenic site.
[0026] Antibodies against influenza often target variable antigenic sites in the globular head of HA, which surround a conserved sialic acid binding site, and thus, neutralize only antigenically closely related viruses. The variability of the HA head is due to the constant antigenic drift of influenza viruses and is responsible for seasonal endemics of influenza. In contrast, gene segments of the viral genome can undergo reassortment (antigenic shift) in host species, creating new viruses with altered antigenicity that are capable of becoming pandemics [Salomon, R. et al. Cell 136, 402-410 (2009)]. Until now, each year, influenza vaccine is updated to reflect the predicted HA and neuraminidase (NA) for upcoming circulating viruses.
[0027] Current vaccine strategies for influenza use either a chemically inactivated or a live attenuated influenza virus. Both vaccines are generally produced in embryonated eggs which present major manufacturing limitations due to the time consuming process and limited production capacity. Another more critical limitation of current vaccines is its highly strain-specific efficacy. These challenges became glaring obvious during emergence of the 2009 H1N1 pandemic, thus validating the necessity for new vaccine platforms capable of overcoming these limitations. Virus-like particles represent one of such alternative approaches and are currently being evaluated in clinical trials [Roldao, A. et al. Expert Rev Vaccines 9, 1149-1176 (2010); Sheridan, C. Nat Biotechnol 27, 489-491 (2009)]. Instead of embryonated eggs, VLPs that often comprise HA, NA and matrix protein 1 (M1) can be mass-produced in mammalian or insect cell expression systems [Haynes, J. R. Expert Rev Vaccines 8, 435-445 (2009)]. The advantages of this approach are its particulate, multivalent nature and the authentic display of properly folded, trimeric HA spikes that faithfully mimic the infectious virion. In contrast, by the nature of its assembly, the enveloped VLPs contain a small but finite host cell component that may present potential safety, immunogenicity challenges following repeated use of this platform [Wu, C. Y. et al. PLoS One 5, e9784 (2010)]. Moreover, the immunity induced by the VLPs is essentially the same as current vaccines do, and thus, does not likely improve both potency and breadth of vaccine-induced protective immunity. In addition to VLPs, a recombinant HA protein has also been evaluated in humans [Treanor, J. J. et al. Vaccine 19, 1732-1737 (2001); Treanor, J. J. JAMA 297, 1577-1582 (2007)], though the ability to induce protective neutralizing antibody titers are limited. The recombinant HA proteins used in those trials were produced in insect cells and might not form native trimer preferentially [Stevens, J. Science 303, 1866-1870 (2004)].
[0028] Recently, entirely new classes of broadly neutralizing antibodies against influenza viruses were isolated. One class of antibodies recognizes the highly conserved HA stem [Corti, D. et al. J Clin Invest 120, 1663-1673 (2010); Ekiert, D. C. et al. Science 324, 246-251 (2009); Kashyap, A. K. et al. Proc Natl Acad Sci USA 105, 5986-5991 (2008); Okuno, Y. et al. J Virol 67, 2552-2558 (1993); Sui, J. et al. Nat Struct Mol Biol 16, 265-273 (2009); Ekiert, D. C. et al. Science 333, 843-850 (2011); Corti, D. et al. Science 333, 850-856 (2011)], and another class of antibodies precisely recognizes the sialic acid binding site of the RBD on the variable HA head [Whittle, J. R. et al. Proc Natl Acad Sci USA 108, 14216-14221 (2011); Krause, J. C. et al. J Virol 85, 10905-10908 (2011)]. Unlike strain-specific antibodies, those antibodies are capable of neutralizing multiple antigenically distinct viruses, and hence inducing such antibodies has been a focus of next generation universal vaccine [Nabel, G. J. et al. Nat Med 16, 1389-1391 (2010)]. However, robustly eliciting these antibodies with such heterologous neutralizing profile by vaccination has been difficult [Steel, J. et al. MBio 1, e0018 (2010); Wang, T. T. et al. PLoS Pathog 6, e1000796 (2010); Wei, C. J. et al. Science 329, 1060-1064 (2010)].
[0029] Despite several alternatives to conventional influenza vaccines, advances in biotechnology in past decades have allowed engineering of biological materials to be exploited for the generation of novel vaccine platforms. Ferritin, an iron storage protein found in almost all living organisms, is an example which has been extensively studied and engineered for a number of potential biochemical/biomedical purposes [Iwahori, K. U.S. Patent 2009/0233377 (2009); Meldrum, F. C. et al. Science 257, 522-523 (1992); Naitou, M. et al. U.S. Patent 2011/0038025 (2011); Yamashita, I. Biochim Biophys Acta 1800, 846-857 (2010)], including a potential vaccine platform for displaying exogenous epitope peptides [Carter, D. C. et al. U.S. Patent 2006/0251679 (2006); Li, C. Q. et al. Industrial Biotechnol 2, 143-147 (2006)]. Its use as a vaccine platform is particularly interesting because of its self-assembly and multivalent presentation of antigen which induces stronger B cell responses than monovalent form as well as induce T-cell independent antibody responses [Bachmann, M. F. et al. Annu Rev Immunol 15, 235-270 (1997); Dintzis, H. M. et al. Proc Natl Acad Sci USA 73, 3671-3675 (1976)]. Further, the molecular architecture of ferritin, which consists of 24 subunits assembling into an octahedral cage with 432 symmetry has the potential to display multimeric antigens on its surface.
[0030] There remains a need for an efficacious influenza vaccine that provides robust protection against influenza virus. There particularly remains a need for an influenza vaccine that protects individuals from heterologous strains of influenza virus, including evolving seasonal and pandemic influenza virus strains of the future. The present invention meets this need by providing a novel HA-ferritin nanoparticle (HA-ferritin np) influenza vaccine that is easily manufactured, potent, and elicits broadly neutralizing influenza antibodies.
BRIEF DESCRIPTION OF THE FIGURES
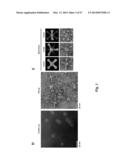
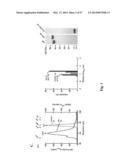
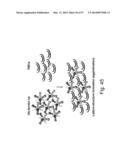
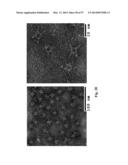
[0031] FIG. 1. Molecular design and construction of ferritin particles displaying influenza virus hemagglutinin. (a) Ribbon diagram of a subunit of H. pylori nonheme ferritin (PDB: 3bve) (left). Amino- and carboxyl-termini are labeled as N and C, respectively. Three ferritin subunits surrounding an octahedral three-fold axis are shown as a ribbon diagram (middle). Residue Asp5 is indicated. The octahedral assembly of the ferritin particle (viewed at 8 Å resolution along an octahedral three-fold axis) and A/Solomon Islands/3/2006 (H1N1) HA trimer (PDB: 3sm5) (viewed down from membrane proximal side) (right). The measured average distance between the Asp5 residues in each ferritin subunit surrounding an octahedral three-fold axis is shown as a triangle. The same equilateral triangle (a=b=c=28 Å) is also drawn on the HA trimer (right). (b) Schematic representation of the HA-ferritin expression vector used for protein production. (c) Chromatogram of the size exclusion chromatography of ferritin nanoparticles (np) and HA-np (left). Molecular weights (kDa) of calibration standards are indicated above the curves with vertical lines. Calculated molecular weights for ferritin nanoparticles and HA-np were 419 and 2,165 kDa, respectively, and were within 10% of the predicted molecular weights (408 and 2,040 kDa, respectively). Particle size distribution (radius) of purified ferritin nanoparticles and HA-np was determined by dynamic light scattering (middle). Measured mean diameters (d) are indicated. The polydispersity indices of purified ferritin np and HA-np were 0.035 and 0.011, respectively. Purified HA trimer (thrombin uncleaved), HA-np and ferritin nanoparticles were analyzed by SDS-PAGE (right). (d) Negatively stained transmission electron microscopy images of ferritin nanoparticles (left) and HA-np (right). Images were originally recorded at 67,000× magnification. (e) Models representing octahedral four-, three- and two-fold symmetries of HA-ferritin np (top panels) and actual TEM image (bottom panels) are shown. Visible HA spikes are numbered in the images.
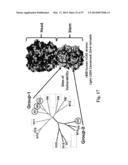
[0032] FIG. 2. Genetic and structural comparison of ferritins. (a) Phylogenetic tree analysis of ferritins found in RSCB PDB. Twenty-two sequences contain 16 ferritins including Vc (Vibrio cholerae), Ec (E. coli), Hp (H. pylori), Af (Archaeoglobus fulgidus), Pf (Pyrococcus furiosus), Tm (Thermatoga maritime), Pm (Pseudo-nitzschia multiseries), Tn (L) (Trichoplusia ni light chain), Soybean (chloroplastic), Horse (L) (light chain), Human (L), (H) and (M) (light, heavy chains and mitochondrial, respectively), Mouse (L) (light chain), and Frog (M) and (L) (middle and lower subunits, respectively), and 6 bacterioferritins (B) including Mt (B) (Mycobacterium tuberculosis), Pa (B) (Pseudomonas aeruginosa), Rs (B) (Rhodabacter sphaeroides), Bm (B) (Brucella melitensis), Av (B) (Azobacter vinelandii), and Ec (B). Protein sequences were aligned using Clustal W2 (www.ebi.ac.uk/Tools/msa/clustalw2) with Gonnet matrix and a phylogenetic tree was generated with the Phylodendron program (http://iubio.bio.indiana.edu/treeapp/treeprint-form.html) using the neighbor joining method. (b) Comparison of surface exposed residues between H. pylori and mouse (light chain) (left) or human (light chain) (middle), and mouse and human (light chains) (right). Conservation of surface exposed residues was rendered by UCSF Chimera using a protein sequence alignment generated by Clustal W2. Conserved and varied residues between the two ferritins are shown as light and dark residues, respectively. PDB files 3bve (H. pylori) (left and middle) and 1h96 (mouse light chain) (right) were used for surface rendering.
[0033] FIG. 3. Antigenic characterization of HA-ferritin np. (a) Binding of mAbs directed to globular head and stem of HA was measured by ELISA. Equal amount (200 ng of HA per well) of HA trimer (), TIV (.box-solid.), HA-ferritin ( ) or Ferr (equimolar amount as HA-Ferr) (◯) were coated on the plates and wells were probed with anti-head mAb (3u-u) and anti-stem mAb CR6261. The half maximal effective concentrations (EC50) of binding were calculated for each antibody and showed as ng ml-1 (b) Inhibition of antibody-mediated neutralization of 1999 NC pseudotyped virus by using HA trimer, HA-Ferr or Ferr as a competitor. Inhibition of neutralization was plotted as percent inhibition respect to no competitor control. The anti-stem neutralizing mAbs, F10 (left) and CR6261 (right) were used at 3.125 and 25 μg ml-1, respectively. Competitor proteins were added to the reactions at a final concentration of 20 μg ml-1.
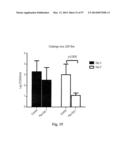
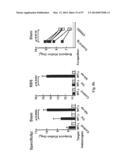
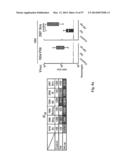
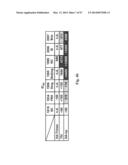
[0034] FIG. 4. Immune responses in HA-np-immunized mice. (a) HAI (left), IC90 neutralization (middle), and anti-HA ab endpoint titers (right) against 1999 NC HA after two immunizations with 0.17 μg (amount of H1 HA) of TIV or HA-np with or without Ribi adjuvant and a 3-week interval. The immune sera were collected 2 weeks after the second immunization. The data are presented as box-and-whiskers plots (boxed from lower to upper quartile with whiskers from minimum to maximum) with lines at the mean (n=5). (b) Neutralization breadth of the immune sera elicited by HA-trimer, TIV, or HA-np. An additional group of mice (n=4) was immunized twice with 20 μg of trimeric HA protein using Ribi adjuvant and a 4-week interval. The immune sera were collected 2 weeks after the second immunization. IC50 neutralization titers against a panel of H1N1 pseudotyped viruses were determined. (c) Cellular and humoral immune responses against H. pylori (top) and mouse (bottom) ferritins. Mice were immunized twice with 1.67 μg (amount of H1 HA) of TIV or HA-np, or 0.57 μg of ferritin nanoparticles (equimolar to HA-np) using Ribi adjuvant and a 3-week interval. The splenocytes and immune sera were harvested 11 days after the second immunization. Cytokine-producing CD4.sup.+ and CD8.sup.+ T cells were measured by ICS (left), and ab response was detected by ELISA (right). All cells expressing IFN-γ, TNFα, or IL-2 were identified as cytokine.sup.+ cells. The percentage of cytokine.sup.+ cells in CD4.sup.+ and CD8.sup.+ T cells that were activated in response to stimulation with specific peptides covering the entire H. pylori or mouse ferritins (heavy and light chains combined) were plotted. Recombinant H. pylori and purified mouse (liver) ferritins were used for detecting anti-ferritin ab responses. The data are presented as box-and-whiskers plots with lines at the mean (n=5).
[0035] FIG. 5. Successive immunization of HA-nanoparticles in mice. Mice were pre-immunized with 1.67 μg (amount of HA) of 2009 Perth (H3) HA-nanoparticles or 0.57 μg (equimolar to HA-nanoparticle) of empty ferritin nanoparticles at week 0 and then immunized with 1.67 μg (amount of HA) of 1999 NC (H1) HA-nanoparticles at week 3. Ribi was used as an adjuvant. Another group of mice was immunized with 1999 NC (H1) HA-nanoparticles without pre-immunization of empty ferritin nanoparticles or H3 HA-nanoparticles. (a) Ab responses to H. pylori ferritin (left) and 2009 Perth H3 HA (right). Immune sera collected 2 weeks after the immunization with H3 HA-nanoparticles or empty ferritin nanoparticles were analyzed by ELISA. (b) Immune responses to 1999 NC (H1) after 1999 NC (H1) HA-nanoparticle immunization. Naive mice or mice with pre-immunity to ferritin or H3 HA were immunized with H1 HA-nanoparticles at week 3 and HAI (left), IC90 neutralization (middle) and ELISA (right) Ab titers were measured 2 weeks after the immunization. The data are presented as box-and-whiskers plots with lines at the mean (n=5).
[0036] FIG. 6. Development of trivalent HA-np. (a) HA-np consisting of HAs from 2009 CA (H1), 2009 Perth (H3) or 2006 FL (type B) were purified and visualized by TEM. (b) HAI titers against 2009 CA (H1N1) and 2009 Perth (H3N2) viruses in immunized mice. Mice were immunized twice with 1.67 μg (amount of HA) of monovalent H1, monovalent H3, monovalent type B, or 5.0 μg (total amount of HA) of trivalent HA-np or TIV (2011-2012 season) using Ribi adjuvant with a 3-week interval. Immune sera were collected 2 weeks after the second immunization. The data are presented as box-and-whiskers plots with lines at the mean (n=5).
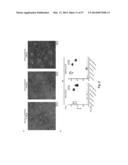
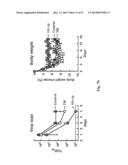
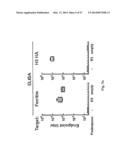
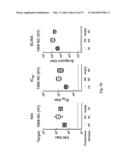
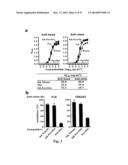
[0037] FIG. 7. Protective immunity induced in ferrets immunized with the HA-np. Ferrets were immunized twice with PBS (control), 7.5 ug (2.5 ug of H1 HA) of TIV or 2.5 ug (amount of HA) 1999 NC HA-np using Ribi adjuvant and a 4-week interval. Control animals received PBS. (a) HA1 (left), IC90 neutralization (middle), and anti-HA ab endpoint titers (right) in immunized ferrets against homologous 1999 NC HA were determined. Immune sera were collected 3 and 2 weeks after the first (R. Salomon, R. G. Webster, The influenza virus enigma. Cell 136, 402-410 (2009) and second (L. C. Lambert, A. S. Fauci, Influenza vaccines for the future. N Engl J Med 363, 2036-2044 (2010)) immunizations, respectively. The data are presented as box and whisker plots with lines at the mean (n=6). (b) Protection of immunized ferrets from an unmatched 2007 Bris virus challenge. Ferrets were challenged with 106.5 50% egg infectious dose (EID50) of 2007 Bris virus 5 weeks after the second immunization. Virus titers in the nasal washes from 1, 3 and 5 days post challenge were determined by a 50% tissue culture infectious dose (TCID50) assay (left). One of six ferrets in the TIV-immunized group showed measurable virus on day 5. Virus titers in 4 out of 6 ferrets on day 3 and 6 out of 6 ferrets on day 5 in the HA-np-immunized group were under the detection limit (<102). The mean viral loads with standard deviation (s.d.) at each time point were plotted (n=6). Change in the body weight after the virus challenge was also monitored (right). Each data point represents the mean percent change in body weight from the pre-challenge (day 0). The mean body weight changes with standard error (s.e.) at each time point were plotted (n=6).
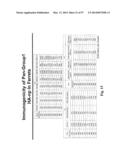
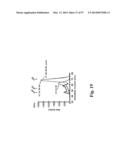
[0038] FIG. 8. Improved neutralization breadth and detection of stem- and RBS-directed abs. (a) Neutralization breadth of immune sera in ferrets. IC50 neutralization titers against a panel of H1N1 pseudotyped viruses (left) and HAI titers against 1934 PR8 and 2007 Bris H1N1 viruses (right) were determined. The HAI titers are presented as box-and-whiskers plots with lines at the mean (n=6). (b) Stem- and RBS-directed abs elicited by HA-np immunization. Ferret immune sera (diluted 1:100) were pre-absorbed with ΔStem HA-expressing cells and their binding to WT or ΔStem HA were analyzed by ELISA (left). The immune sera (diluted 1:1,000) were pre-absorbed with ΔRBS HA-expressing cells and their binding to WT or ΔRBS HA were analyzed by ELISA (middle). The mean endpoint dilution titers were plotted with s.d. (n=6). Competition ELISA with stem-directed mAb CR6261 (right). The immune sera pre-absorbed with ΔStem were tested for binding to HA in the presence of an isotype control IgG or CR6261. Each symbol represents the titer of an individual ferret (n=6). (c) Neutralization competition with WT, ΔStem or ΔRBS HA protein (left). The neutralization of HA-np immune sera against 1986 Sing, 1995 Beijing, 1999 NC and 2007 Bris was measured in the presence of irrelevant protein (control), WT, ΔStem or ΔRBS HA as a competitor. Percent neutralizations at serum dilution 1:200 (1986 Sing, 2007 Bris), 1:800 (1995 Beijing) or 1:3,200 (1999 NC) were plotted. Each symbol represents the individual ferret serum and mean is indicated as a red line with s.d. (n=6 for 1986 Sing, 1995 Beijing and 1999 NC; n=3 for 2007 Bris). The relative contribution of the stem- and RBS-directed neutralization was determined by the inhibition of neutralization for each competitor protein (right). Mean percent contributions in neutralizing each virus were plotted as pile-up bars (n=6).
[0039] FIG. 9. Characterization of ΔRBS HA probe. (a) Crystal structure of HA (A/Solomon Islands/3/2006) complex with an anti-RBS mAb CH65 Fab (PDB: 3sm5) (J. R. Whittle et al., Broadly neutralizing human antibody that recognizes the receptor-binding pocket of influenza virus hemagglutinin. Proc Natl Acad Sci USA 108, 14216-14221 (2011)) (left). Close up view of CH65 contact area (right). The residue HA1 190 which has been mutated to be glycosylated in ΔRBS mutant is highlighted. The CH65 Fab-bound HA1 protomer is darkened. (b) Characterization of the soluble trimer of WT and ΔRBS HAs from 1999 NC and 2007 Bris. The WT and ΔRBS HA proteins were immunoprecipitated with anti-RBS (CH65), stem (CR6261) and control (anti-HIV, VRC01) mAbs. Immune complexes were then dissolved in Lamini buffer and analyzed by SDS-PAGE. Antibody heavy and light chains are labeled as HC and LC, respectively.
[0040] FIG. 10. Purification of HA-np. HA-np were purified by routine iodixanol gradient ultracentrifugation routinely. Fractions containing HA-np were confirmed by SDS-PAGE and Western blotting using a mAb against 1999 NC HA. The HA-np were enriched in the fraction with density of ˜1.15 g/ml.
[0041] FIG. 11. Protocol for immunization of mice and ferrets using pan-group 1 HA-ferritin np. Mice were injected intramuscularly twice (Week 0 and week 4) with PBS (control) or 6.8 ug (1.7 ug of each HA-ferritin np) pan-group 1 vaccine in Ribi. Ferrets were injected intramuscularly twice (Week 0 and week 4) with PBS (control) or 10 ug (2.5 ug of each HA-ferritin np) pan-group 1 vaccine in Ribi.
[0042] FIG. 12. Neutralization activity of mouse antisera against Group1 HA pseudotyped viruses. Neutralization activity of murine antisera from control or pan Group1 HA-np immunized mice against the indicated HA pseudotyped viruses. IC50 titers are shown for all panels.
[0043] FIG. 13. Neutralization activity of ferret antisera against Group1 HA pseudotyped viruses. Neutralization activity of ferret antisera from control or pan Group1 HA np immunized ferrets against the indicated HA pseudotyped viruses. IC50 titers are shown for all panels.
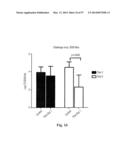
[0044] FIG. 14. H1 hemaggulutination inhbitions (HAI) assays were performed using the sera obtained from the ferritin immunization studies. These studies were performed using actual H1 virus, and H2 and H5 HAI were performed using HA-ferritin np.
[0045] FIG. 15. Protection of ferrets from viral challenge with Influenza A/Brisbane/59/2007 Brisbane (H1N1) (2007 Bris). Two groups of ferrets (n=6 for control and n=5 for pan-group1 immune) were immunized with pan Group1 HA np vaccine or PBS (control) and challenged with heterologous 2007 Bris virus (106.5 EID50). Virus titers were measured in nasal swabs collected on day 3 and day 5 post challenge. Titers were determined using end-point titration in MDCK cells.
[0046] FIG. 16. Protection of ferrets from viral challenge with Influenza A/Mexico/2009 (H1N1) (2009 Mex). Two groups of ferrets (n=6) were immunized with pan Group1 HA np vaccine or PBS (control) and challenged with heterologous 2009 Mex virus (106.5 EID50). Virus titers were measured in nasal swabs collected on day 3 and day 5 post challenge. Titers were determined using end-point titration in MDCK cells.
[0047] FIG. 17. Conservation of the influenza HA stem region. (left, right) Neutralizing antibodies that react with both Group 1 and Group 2 viruses act at the sites of vulnerability shown in the Figure. (Right) Space filling model of influenza HA protein illustrating amino acid sequence conservation in over 800 human H1N1 strains. Light residues indicate residues that are 100% conserved. Dark residues as indicate sites of variation.
[0048] FIG. 18. Design of HA Stabilized Stem protein. (A) Schematic of the HA SS (bottom) in comparison to HA (top). HA SS was constructed by inserting a GWG linker between residues 42 and 314 of HA1 RBD head, a gp41 post-fusion trimerization motif inserted in place of residues 59 through 93 of HA2, a GG linker between HA2 and the gp41 HR2 helix and an NGTGGGSG linker between the two gp41 helices. The gene sequence of H1 NC 99 SS is provided in the supplemental materials. (B) Trimeric and monomeric representation of HA (PDB entry 1RU7) in comparison to the HA SS model. Coloring is respective to above panel, with the monomeric representation also illustrating the CR6261 epitope as yellow and HA residues which are omitted in the stabilized HA stem as grey. (C) CR6261, FI6v3, and the germline of the VH1-69 Ab 70-5B03 have similar affinity to HA and SS by ELISA. HA SS competes with CR6261 (D) binding to HA and (E) neutralization of H1N1 pseudovirus similar to soluble HA trimer.
[0049] FIG. 19. Size exclusion chromatogram of HA and HA SS probes. Calibration standards are shown above the curves as vertical lines.
[0050] FIG. 20. Electron microscopic analysis of nanoparticles. Purified SS-np were subjected to transmission electron microscopic analysis. The samples were negatively stained with ammonium molybdate and images were recorded on a Tecnai T12 microscope (FEI) at 80 kV with a CCD camera (AMT Corp.). Images of lower (left) and higher (right) magnifications are shown. The SS spikes were protruding perpendicularly from the particle core and clearly visible.
[0051] FIG. 21. Antigenic characterization of HA SS-ferritin np. The ability of purified HA SS and HA SS-np to bind to monoclonal Abs CR6261 (left) and FI6v3 (right) was characterized by ELISA. HA and HIV gp120 proteins served as controls.
[0052] FIG. 22. Immune sera of mice immunized heterologously with HA-np prime and HA SS-np boost are reactive to the conserved HA stem epitope. Antibodies elicited by vaccination target the conserved HA stem epitope as individual mice possess differential binding (a minimum of 2-fold difference in endpoint dilution) between wt and Δstem HA variants. The percentage of mice displaying differential binding is given above matched wt and Δstem constructs. Error bars represent standard error.
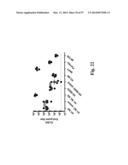
[0053] FIG. 23. Immune sera of mice immunized with HA SS neutralizes diverse pseudovirus stains. IC50 values are shown for individual mice against H1 homosubtypic strains and H2, H5 and H9 group-1 heterosubtypic strains. Dashed lines represents the lowest dilution assayed (50). Error bars represent standard error.
[0054] FIG. 24. Boosting with HA SS-np increases neutralizing titers in ferrets against H1N1 New Calendonia. Pseudovirus neutralizing titers were calculated for preimmune, HA FL-np primed, and HA SS-np boosted sera from individual mice. Error bars represent standard deviation of values.
[0055] FIG. 25. Map and sequence of CMV8x/R-H1NC HA(517)_SGG-egm and the related HA-ferritin fusion protein.
[0056] FIG. 26. Map and sequence of CMV8x/R-H1CA HA(518)_SGG-egm and the related HA-ferritin fusion protein.
[0057] FIG. 27. Map and sequence of CMV8x/R-H2Sing HA(514)_SGG-egm and the related HA-ferritin fusion protein.
[0058] FIG. 28. Map and sequence of CMV8x/R-H3HK HA(519)_SGG-egm and the related HA-ferritin fusion protein.
[0059] FIG. 29. Map and sequence of CMV8x/R-H3Bris HA(519)_SGG-egm and the related HA-ferritin fusion protein.
[0060] FIG. 30. Map and sequence of CMV8x/R-H5Indo HA(520)_SGG-egm and the related HA-ferritin fusion protein.
[0061] FIG. 31. Map and sequence of CMV8x/R-B.Florida HA(534)_SGG-egm and the related HA-ferritin fusion protein.
[0062] FIG. 32. Map and sequence of CMV8x/R-H3Perth HA(519)_SGG-egm and the related HA-ferritin fusion protein.
[0063] FIG. 33. Map and sequence of CMV8x/R-H1Bris HA(517)_SGG-egm and the related HA-ferritin fusion protein.
[0064] FIG. 34. Map and sequence of CMV8x/R-B.Bris HA(535)_SGG-egm and the related HA-ferritin fusion protein.
[0065] FIG. 35. Map and sequence of CMV8x/R--H1NC SS Gen4.55_SGG-egm and the related HA SS-ferritin fusion protein.
[0066] FIG. 36. Map and sequence of CMV/R H1 CA SS/Gen4.55/ferritin.
[0067] FIG. 37. Map and sequence of CMV/R H1 Bris SS/Gen4.55/ferritin.
[0068] FIG. 38. Map and sequence of CMV/R H2 Sing SS/Gen4.55/ferritin.
[0069] FIG. 39. Map and sequence of CMV/R H3 Bris SS/Gen4.55/ferritin.
[0070] FIG. 40. Map and sequence of CMV/R H3 Perth SS/Gen4.55/ferritin.
[0071] FIG. 41. Map and sequence of CMV/R H3 HK68 SS/Gen4.55/ferritin.
[0072] FIG. 42. Map and sequence of CMV/R H5 Indo SS/Gen4.55/ferritin.
[0073] FIG. 43. Map and sequence of CMV/R B Bris SS/Gen4.55/ferritin.
[0074] FIG. 44. Map and sequence of CMV/R B FL SS/Gen4.55/ferritin.
[0075] FIG. 45. Illustration showing hemagglutination reaction.
[0076] FIG. 46. Comparison of the hemagglutination activities of HA-ferritin nanoparticle and H5N1 vaccine. Inactivated vaccine (A) and HA-ferritin nanoparticles (B) were serially diluted in PBS and incubated with 0.5% chicken red blood cells (RBCs) for 30 minutes at room temperature. A series of dilutions were also performed using PBS as a control (C). Hemagglutination mediated through sialic acid moiety on RBC surface and the sialic acid-binding site on HA was determined by formation of lattice-structure of multiple RBCs (lack of red dot in the well).
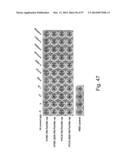
[0077] FIG. 47. Mutation in sialic acid-binding site (ΔSA; Y98F) diminishes hemagglutination of HA-ferritin nanoparticles. Hemagglutination activity of HA-ferritin nanoparticles was diminished by a mutation in the sialic acid-binding site on HA.
[0078] FIG. 48. Titration of antibodies inhibiting hemagglutination using two assay systems (traditional HAI assay and nanoparticle-based HAI assay). Sera from twenty-four ferrets were pretreated with receptor-destroying enzyme (RDE) and heat inactivated were titrated by using A/New Calcdonia/20/99 (H1N1) virus or HA-np having the same HA. The samples include 6 negative (pre-immune) and 24 test (post-immune) sera. The titers are shown as scattered dots (left) and comparative (middle) plots. Correlation between the two assays was assessed (right).
DETAILED DESCRIPTION OF THE INVENTION
[0079] The present invention relates to a novel vaccine for influenza virus. More specifically, the present invention relates to novel, influenza hemagglutinin protein-based vaccines that elicit an immune response against a broad range of influenza viruses. It also relates to self-assembling ferritin-based, nanoparticles that display immunogenic portions of influenza hemagglutinin protein on their surface. Such nanoparticles are useful for vaccinating individuals against influenza virus. Accordingly, the present invention also relates to fusion proteins for producing such nanoparticles and nucleic acid molecules encoding such proteins. Additionally, the present invention relates to, methods of producing nanoparticles of the present invention, and methods of using such nanoparticles to vaccinate individuals.
[0080] Before the present invention is further described, it is to be understood that this invention is not limited to particular embodiments described, as such may, of course, vary. It is also to be understood that the terminology used herein is for the purpose of describing particular embodiments only, and is not intended to be limiting, since the scope of the present invention will be limited only by the claims.
[0081] It must be noted that as used herein and in the appended claims, the singular forms "a," "an," and "the" include plural referents unless the context clearly dictates otherwise. For example, a nucleic acid molecule refers to one or more nucleic acid molecules. As such, the terms "a", "an", "one or more" and "at least one" can be used interchangeably. Similarly the terms "comprising", "including" and "having" can be used interchangeably. It is further noted that the claims may be drafted to exclude any optional element. As such, this statement is intended to serve as antecedent basis for use of such exclusive terminology as "solely," "only" and the like in connection with the recitation of claim elements, or use of a "negative" limitation.
[0082] In addition to the above, unless specifically defined otherwise, the following terms and phrases, which are common to the various embodiments disclosed herein, are defined as follows:
[0083] As used herein, the term immunogenic refers to the ability of a specific protein, or a specific region thereof, to elicit an immune response to the specific protein, or to proteins comprising an amino acid sequence having a high degree of identity with the specific protein. According to the present invention, two proteins having a high degree of identity have amino acid sequences at least 80% identical, at least 85% identical, at least 87% identical, at least 90% identical, at least 92% identical, at least 94% identical, at least 96% identical, at least 98% identical or at least 99% identical.
[0084] As used herein, an immune response to a vaccine, or nanoparticle, of the present invention is the development in a subject of a humoral and/or a cellular immune response to a hemagglutinin protein present in the vaccine. For purposes of the present invention, a "humoral immune response" refers to an immune response mediated by antibody molecules, including secretory (IgA) or IgG molecules, while a "cellular immune response" is one mediated by T-lymphocytes and/or other white blood cells. One important aspect of cellular immunity involves an antigen-specific response by cytolytic T-cells ("CTL"s). CTLs have specificity for peptide antigens that are presented in association with proteins encoded by the major histocompatibility complex (MHC) and expressed on the surfaces of cells. CTLs help induce and promote the destruction of intracellular microbes, or the lysis of cells infected with such microbes. Another aspect of cellular immunity involves an antigen-specific response by helper T-cells. Helper T-cells act to help stimulate the function, and focus the activity of, nonspecific effector cells against cells displaying peptide antigens in association with MHC molecules on their surface. A cellular immune response also refers to the production of cytokines, chemokines and other such molecules produced by activated T-cells and/or other white blood cells, including those derived from CD4+ and CD8+ T-cells.
[0085] Thus, an immunological response may be one that stimulates CTLs, and/or the production or activation of helper T-cells. The production of chemokines and/or cytokines may also be stimulated. The vaccine may also elicit an antibody-mediated immune response. Hence, an immunological response may include one or more of the following effects: the production of antibodies (e.g., IgA or IgG) by B-cells; and/or the activation of suppressor, cytotoxic, or helper T-cells and/or T-cells directed specifically to a hemagglutinin protein present in the vaccine. These responses may serve to neutralize infectivity, and/or mediate antibody-complement, or antibody dependent cell cytotoxicity (ADCC) to provide protection to an immunized individual. Such responses can be determined using standard immunoassays and neutralization assays, well known in the art.
[0086] According to the present invention all nomenclature used to classify influenza virus is that commonly used by those skilled in the art. Thus, a Type, or Group, of influenza virus refers to influenza Type A, influenza Type B or influenza type C. It is understood by those skilled in the art that the designation of a virus as a specific Type relates to sequence difference in the respective M1 (matrix) protein or NP (nucleoprotein). Type A influenza viruses are further divided into Group1 and Group 2. These Groups are further divided into subtypes, which refers to classification of a virus based on the sequence of its HA protein. Examples of current commonly recognized subtypes are H1, H2, H3, H4, H5, H6, H7, H8, H8, H10, H11, H12, H13, H14, H15 or H16. Group 1 influenza subtypes are H1, H2, H5, H7 and H9. Group 2 influenza subtypes are H4, H4, H6, H8, H10, H11, H12, H13, H14, H15 and H16. Finally, the term strain refers to viruses within a subtype that differ from one another in that they have small, genetic variations in their genome.
[0087] As used herein, neutralizing antibodies are antibodies that prevent influenza virus from completing one round of replication. As defined herein, one round of replication refers the life cycle of the virus, starting with attachment of the virus to a host cell and ending with budding of newly formed virus from the host cell. This life cycle includes, but is not limited to, the steps of attaching to a cell, entering a cell, cleavage and rearrangement of the HA protein, fusion of the viral membrane with the endosomal membrane, release of viral ribonucleoproteins into the cytoplasm, formation of new viral particles and budding of viral particles from the host cell membrane.
[0088] As used herein, broadly neutralizing antibodies are antibodies that neutralize more than one type, subtype and/or strain of influenza virus. For example, broadly neutralizing antibodies elicited against an HA protein from a Type A influenza virus may neutralize a Type B or Type C virus. As a further example, broadly neutralizing antibodies elicited against an HA protein from Group I influenza virus may neutralize a Group 2 μm. AS an additional example, broadly neutralizing antibodies elicited against an HA protein from one sub-type or strain of virus, may neutralize another sub-type or strain of virus. For example, broadly neutralizing antibodies elicited against an HA protein from an H1 influenza virus may neutralize viruses from one or more sub-types selected from the group consisting of H2, H3, H4, H5, H6, H7, H8, H8, H10, H11, H12, H13, H14, H15 or H16.
[0089] As used herein, an influenza hemagglutinin protein, or HA protein, refers to a full-length influenza hemagglutinin protein or any portion thereof, that is capable of eliciting an immune response. Preferred HA proteins are those that are capable of forming a trimer. An epitope of a full-length influenza hemagglutinin protein refers to a portion of such protein that can elicit a neutralizing antibody response against the homologous influenza strain, i.e., a strain from which the HA is derived. In some embodiments, such an epitope can also elicit a neutralizing antibody response against a heterologous influenza strain, i.e., a strain having an HA that is not identical to that of the HA of the immunogen.
[0090] With regard to hemagglutinin proteins, it is understood by those skilled in the art that hemagglutinin proteins from different influenza viruses may have different lengths due to mutations (insertions, deletions) in the protein. Thus, reference to a corresponding region refers to a region of another proteins that is identical, or nearly so (e.g., at least 95%, identical, at least 98% identical or at least 99% identical), in sequence, structure and/or function to the region being compared. For example, with regard to the stem region of a hemagglutinin protein, the corresponding region in another hemagglutinin protein may not have the same residue numbers, but will have a nearly identical sequence and will perform the same function. To better clarify sequences comparisons between viruses, numbering systems are used by those in the field, which relate amino acid positions to a reference sequence. Thus, corresponding amino acid residues in hemagglutinin proteins from different strains of influenza may not have the same residue number with respect to their distance from the n-terminal amino acid of the protein. For example, using the H3 numbering system, reference to residue 100 in A/New Calcdonia/20/1999 (1999 NC, H1) does not mean it is the 100th residue from the N-terminal amino acid. Instead, residue 100 of A/New Calcdonia/20/1999 (1999 NC, H1) aligns with residue 100 of influenza H3N2 strain. The use of such numbering systems is understood by those skilled in the art. Unless otherwise noted, reference to amino acids in hemagglutinin proteins herein is made using the H3 numbering system.
[0091] According to the present invention, a trimerization domain is a series of amino acids that when joined (also referred to as fused) to a protein or peptide, allow the fusion protein to interact with other fusion proteins containing the trimerization domain, such that a trimeric structure is formed. Any known trimerization domain can be used in the present invention. Examples of trimerization domains include, but are not limited to, the HIV-1 gp41 trimerization domain, the SIV gp41 trimerization domain, the Ebola virus gp-2 trimerization domain, the HTLV-1 gp-21 trimerization domain, the T4 fibritin trimerization domain (i.e., foldon), the yeast heat shock transcription factor trimerization domain, and the human collagen trimerization domain.
[0092] As used herein, a variant refers to a protein, or nucleic acid molecule, the sequence of which is similar, but not identical to, a reference sequence, wherein the activity of the variant protein (or the protein encoded by the variant nucleic acid molecule) is not significantly altered. These variations in sequence can be naturally occurring variations or they can be engineered through the use of genetic engineering technique know to those skilled in the art. Examples of such techniques are found in Sambrook J, Fritsch E F, Maniatis T et al., in Molecular Cloning--A Laboratory Manual, 2nd Edition, Cold Spring Harbor Laboratory Press, 1989, pp. 9.31-9.57), or in Current Protocols in Molecular Biology, John Wiley & Sons, N.Y. (1989), 6.3.1-6.3.6, both of which are incorporated herein by reference in their entirety.
[0093] With regard to variants, any type of alteration in the amino acid, or nucleic acid, sequence is permissible so long as the resulting variant protein retains the ability to elicit neutralizing antibodies against an influenza virus. Examples of such variations include, but are not limited to, deletions, insertions, substitutions and combinations thereof. For example, with regard to proteins, it is well understood by those skilled in the art that one or more (e.g., 2, 3, 4, 5, 6, 7, 8, 9 or 10), amino acids can often be removed from the amino and/or carboxy terminal ends of a protein without significantly affecting the activity of that protein. Similarly, one or more (e.g., 2, 3, 4, 5, 6, 7, 8, 9 or 10) amino acids can often be inserted into a protein without significantly affecting the activity of the protein.
[0094] As noted, variant proteins of the present invention can contain amino acid substitutions relative to the influenza HA proteins disclosed herein. Any amino acid substitution is permissible so long as the activity of the protein is not significantly affected. In this regard, it is appreciated in the art that amino acids can be classified into groups based on their physical properties. Examples of such groups include, but are not limited to, charged amino acids, uncharged amino acids, polar uncharged amino acids, and hydrophobic amino acids. Preferred variants that contain substitutions are those in which an amino acid is substituted with an amino acid from the same group. Such substitutions are referred to as conservative substitutions.
[0095] Naturally occurring residues may be divided into classes based on common side chain properties:
[0096] 1) hydrophobic: Met, Ala, Val, Leu, Ile;
[0097] 2) neutral hydrophilic: Cys, Ser, Thr;
[0098] 3) acidic: Asp, Glu;
[0099] 4) basic: Asn, Gln, His, Lys, Arg;
[0100] 5) residues that influence chain orientation: Gly, Pro; and
[0101] 6) aromatic: Trp, Tyr, Phe.
[0102] For example, non-conservative substitutions may involve the exchange of a member of one of these classes for a member from another class.
[0103] In making amino acid changes, the hydropathic index of amino acids may be considered. Each amino acid has been assigned a hydropathic index on the basis of its hydrophobicity and charge characteristics. The hydropathic indices are: isoleucine (+4.5); valine (+4.2); leucine (+3.8); phenylalanine (+2.8); cysteine/cystine (+2.5); methionine (+1.9); alanine (+1.8); glycine (-0.4); threonine (-0.7); serine (-0.8); tryptophan (-0.9); tyrosine (-1.3); proline (-1.6); histidine (-3.2); glutamate (-3.5); glutamine (-3.5); aspartate (-3.5); asparagine (-3.5); lysine (-3.9); and arginine (-4.5). The importance of the hydropathic amino acid index in conferring interactive biological function on a protein is generally understood in the art (Kyte et al., 1982, J. Mol. Biol. 157:105-31). It is known that certain amino acids may be substituted for other amino acids having a similar hydropathic index or score and still retain a similar biological activity. In making changes based upon the hydropathic index, the substitution of amino acids whose hydropathic indices are within ±2 is preferred, those within ±1 are particularly preferred, and those within ±0.5 are even more particularly preferred.
[0104] It is also understood in the art that the substitution of like amino acids can be made effectively on the basis of hydrophilicity, particularly where the biologically functionally equivalent protein or peptide thereby created is intended for use in immunological invention, as in the present case. The greatest local average hydrophilicity of a protein, as governed by the hydrophilicity of its adjacent amino acids, correlates with its immunogenicity and antigenicity, i.e., with a biological property of the protein. The following hydrophilicity values have been assigned to these amino acid residues: arginine (+3.0); lysine (+3.0); aspartate (+3.0±1); glutamate (+3.0±1); serine (+0.3); asparagine (+0.2); glutamine (+0.2); glycine (0); threonine (-0.4); proline (-0.5±1); alanine (-0.5); histidine (-0.5); cysteine (-1.0); methionine (-1.3); valine (-1.5); leucine (-1.8); isoleucine (-1.8); tyrosine (-2.3); phenylalanine (-2.5); and tryptophan (-3.4). In making changes based upon similar hydrophilicity values, the substitution of amino acids whose hydrophilicity values are within ±2 is preferred, those within ±1 are particularly preferred, and those within ±0.5 are even more particularly preferred. One may also identify epitopes from primary amino acid sequences on the basis of hydrophilicity.
[0105] Desired amino acid substitutions (whether conservative or non-conservative) can be determined by those skilled in the art at the time such substitutions are desired. For example, amino acid substitutions can be used to identify important residues of the HA protein, or to increase or decrease the immunogenicity, solubility or stability of the HA proteins described herein. Exemplary amino acid substitutions are shown below in Table 1.
TABLE-US-00001 TABLE 1 Amino Acid Substitutions Original Amino Acid Exemplary Substitutions Ala Val, Leu, Ile Arg Lys, Gln, Asn Asn Gln Asp Glu Cys Ser, Ala Gln Asn Glu Asp Gly Pro, Ala His Asn, Gln, Lys, Arg Ile Leu, Val, Met, Ala Leu Ile, Val, Met, Ala Lys Arg, Gln, Asn Met Leu, Phe, Ile Phe Leu, Val, Ile, Ala, Tyr Pro Ala Ser Thr, Ala, Cys Thr Ser Trp Tyr, Phe Tyr Trp, Phe, Thr, Ser Val Ile, Met, Leu, Phe, Ala
[0106] As used herein, the phrase significantly affect a proteins activity refers to a decrease in the activity of a protein by at least 10%, at least 20%, at least 30%, at least 40% or at least 50%. With regard to the present invention, such an activity may be measured, for example, as the ability of a protein to elicit neutralizing antibodies against an influenza virus. Such activity may be measured by measuring the titer of such antibodies against influenza virus, or by measuring the number of types, subtypes or strains neutralized by the elicited antibodies. Methods of determining antibody titers and methods of performing virus neutralization assays are known to those skilled in the art. In addition to the activities described above, other activities that may be measured include the ability to agglutinate red blood cells and the binding affinity of the protein for a cell. Methods of measuring such activities are known to those skilled in the art.
[0107] As used herein, a fusion protein is a recombinant protein containing amino acid sequence from at least two unrelated proteins that have been joined together, via a peptide bond, to make a single protein. The unrelated amino acid sequences can be joined directly to each other or they can be joined using a linker sequence. As used herein, proteins are unrelated, if their amino acid sequences are not normally found joined together via a peptide bond in their natural environment(s) (e.g., inside a cell). For example, the amino acid sequences of monomeric subunits that make up ferritin, and the amino acid sequences of influenza hemagglutinin proteins are not normally found joined together via a peptide bond.
[0108] The terms individual, subject, and patient are well-recognized in the art, and are herein used interchangeably to refer to any human or other animal susceptible to influenza infection. Examples include, but are not limited to, humans and other primates, including non-human primates such as chimpanzees and other apes and monkey species; farm animals such as cattle, sheep, pigs, seals, goats and horses; domestic mammals such as dogs and cats; laboratory animals including rodents such as mice, rats and guinea pigs; birds, including domestic, wild and game birds such as chickens, turkeys and other gallinaceous birds, ducks, geese, and the like. The terms individual, subject, and patient by themselves, do not denote a particular age, sex, race, and the like. Thus, individuals of any age, whether male or female, are intended to be covered by the present disclosure and include, but are not limited to the elderly, adults, children, babies, infants, and toddlers. Likewise, the methods of the present invention can be applied to any race, including, for example, Caucasian (white), African-American (black), Native American, Native Hawaiian, Hispanic, Latino, Asian, and European. An infected subject is a subject that is known to have influenza virus in their body.
[0109] As used herein, a vaccinated subject is a subject that has been administered a vaccine that is intended to provide a protective effect against an influenza virus.
[0110] As used herein, the terms exposed, exposure, and the like, indicate the subject has come in contact with a person of animal that is known to be infected with an influenza virus.
[0111] The publications discussed herein are provided solely for their disclosure prior to the filing date of the present application. Nothing herein is to be construed as an admission that the present invention is not entitled to antedate such publication by virtue of prior invention. Further, the dates of publication provided may be different from the actual publication dates, which may need to be independently confirmed.
[0112] Unless defined otherwise, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention belongs. Although any methods and materials similar or equivalent to those described herein can also be used in the practice or testing of the present invention, the preferred methods and materials are now described. All publications mentioned herein are incorporated herein by reference to disclose and describe the methods and/or materials in connection with which the publications are cited.
[0113] It is appreciated that certain features of the invention, which are, for clarity, described in the context of separate embodiments, may also be provided in combination in a single embodiment. Conversely, various features of the invention, which are, for brevity, described in the context of a single embodiment, may also be provided separately or in any suitable sub-combination. All combinations of the embodiments are specifically embraced by the present invention and are disclosed herein just as if each and every combination was individually and explicitly disclosed. In addition, all sub-combinations are also specifically embraced by the present invention and are disclosed herein just as if each and every such sub-combination was individually and explicitly disclosed herein.
[0114] According to the present invention, vaccines are provided that elicit a broad immune response against influenza viruses. Some vaccines disclosed herein may elicit an immune response against the entire HA protein, while others may elicit an immune response against a specific region or portion of an influenza HA protein. Moreover, the inventors have discovered that specific fusion proteins comprising portions of hemagglutinin protein are useful for eliciting a broad immune response against influenza viruses. Each of these embodiments will now be disclosed in detail below.
Vaccines Against the Stem Region of Influenza HA Protein
[0115] As stated previously, the amino acid sequence of the stem region of the hemagglutinin protein is highly conserved across types, sub-types and strains of influenza viruses and contains a site of vulnerability for group 1 viruses. Thus, an immune response directed this region of the HA protein may protect individuals against influenza viruses from several types, sub-types and/or strains.
[0116] Consequently, one embodiment of the present invention is a protein that elicits an immune response against the stem region of an influenza HA protein. In one embodiment, the immune response can be directed against the stem region of an HA protein from a virus selected from the group consisting of influenza A viruses, influenza B viruses and influenza C viruses. In one embodiment, the immune response can be directed against the stem region of an HA protein from a virus selected from the group consisting of an H1 influenza virus, an H2 influenza virus, an influenza H3 virus, an influenza H4 virus, an influenza H5 virus, an influenza H6 virus, an H7 influenza virus, an H8 influenza virus, an H9 influenza virus, an H10 influenza virus, an H11 influenza virus, an H12 influenza virus, an H13 influenza virus, an H14 influenza virus, an H15 influenza virus and an H16 influenza virus. In one embodiment, the immune response can be directed against the stem region of an HA protein from a strain of virus selected from the group of viruses listed in Table 2.
TABLE-US-00002 TABLE 2 SEQ ID NO Comments FERRITIN 1 Coding sequence for ferritin monomeric subunit protein from H. pylori 2 Amino acid sequence encoded by SEQ ID NO: 1 3 Complement of SEQ ID NO1 4 Nucleic acid sequence encoding amino acids 5-167 from SEQ ID NO: 2; Asn19 has been replaced with Gln 5 Amino acid sequence encoded by SEQ ID NO: 3 6 Complement of SEQ ID NO3 FULL LENGTH HA 7 Nucleic acid sequence encoding full length hemagglutinin protein from A/New Caledonia/20/1999 (1999 NC, H1)(GenBank: AY289929) 8 Amino acid sequence encoded by SEQ ID NO: 7 (full length hemagglutinin protein from A/New Caledonia/20/1999 (1999 NC, H1)(GenBank: AY289929)) 9 Complement of SEQ ID NO: 7 ECTODOMAINS 10 Nucleic acid sequence encoding ectodomain from hemagglutinin protein from A/New Caledonia/20/1999 (1999 NC, H1). 11 Amino acid sequence encoded by SEQ ID NO: 10 (ectodomain from hemagglutinin protein from A/New Caledonia/20/1999 (1999 NC, H1). Amino acids 1-517 from SEQ ID NO: 8. 12 Complement of SEQ ID NO: 10 13 Nucleic acid sequence encoding ectodomain from hemagglutinin protein from A/California/04/2009 (2009 CA, H1). 14 Amino acid sequence encoded by SEQ ID NO: 13 (ectodomain from hemagglutinin protein from A/California/04/2009 (2009 CA, H1)) 15 Complement of SEQ ID NO: 13 16 Nucleic acid sequence encoding ectodomain from hemagglutinin protein from A/Singapore/1/1957 (1957 Sing, H2). 17 Amino acid sequence encoded by SEQ ID NO: 16 (ectodomain from hemagglutinin protein from A/Singapore/1/1957 (1957 Sing, H2)) 18 Complement of SEQ ID NO: 16 19 Nucleic acid sequence encoding ectodomain from hemagglutinin protein from A/Hong Kong/1/1968 (1968 HK, H3). 20 Amino acid sequence encoded by SEQ ID NO: 19) ectodomain from hemagglutinin protein from A/Hong Kong/1/1968 (1968 HK, H3)) 21 Complement of SEQ ID NO: 19 22 Nucleic acid sequence encoding ectodomain from hemagglutinin protein from A/Brisbane/10/2007 (2007 Bris, H3). 23 Amino acid sequence encoded by SEQ ID NO: 22 (ectodomain from hemagglutinin protein from A/Brisbane/10/2007 (2007 Bris, H3)) 24 Complement of SEQ ID NO: 22. 25 Nucleic acid sequence encoding ectodomain from hemagglutinin protein from A/Indonesia/05/2005 (2005 Indo, H5) 26 Amino acid sequence encoded by SEQ ID NO: 25 (ectodomain from hemagglutinin protein from A/Indonesia/05/2005 (2005 Indo, H5)) 27 Complement of SEQ ID NO: 25 28 Nucleic acid sequence encoding ectodomain from hemagglutinin protein from B/Florida/4/2006 (2006 Flo, B) 29 Amino acid sequence encoded by SEQ ID NO: 28 (ectodomain from hemagglutinin protein from B/Florida/4/2006 (2006 Flo, B)) 30 Complement of SEQ ID NO: 28 31 Nucleic acid sequence encoding ectodomain from hemagglutinin protein from A/Perth/16/2009 (2009 Per, H3) 32 Amino acid sequence encoded by SEQ ID NO: 31 (ectodomain from hemagglutinin protein from A/Perth/16/2009 (2009 Per, H3)) 33 Complement of SEQ ID NO: 31 34 Nucleic acid sequence encoding ectodomain from hemagglutinin protein from A/Brisbane/59/2007 (2007 Bris, H1) 35 Amino acid sequence encoded by SEQ ID NO: 34 (ectodomain from hemagglutinin protein from A/Brisbane/59/2007 (2007 Bris, H1)) 36 Complement of SEQ ID NO: 34 37 Nucleic acid sequence encoding ectodomain from hemagglutinin protein from B/Brisbane/60/2008 (2008 Bris, B) 38 Amino acid sequence encoded by SEQ ID NO: 37 (ectodomain from hemagglutinin protein from B/Brisbane/60/2008 (2008 Bris, B)) 39 Complement of SEQ ID NO: 37 FERRITIN-HA ECTODOMAIN FUSION 40 Nucleic acid sequence encoding SEQ ID NO: 41 41 Amino acid sequence of ferritin-HA fusion (ectodomain from hemagglutinin protein from A/New Caledonia/20/1999 (1999 NC, H1)) 42 Complement of SEQ ID NO: 40 43 Nucleic acid sequence encoding SEQ ID NO: 44 44 Amino acid sequence of ferritin-HA fusion (ectodomain from hemagglutinin protein from A/California/04/2009 (2009 CA, H1)) 45 Complement of SEQ ID NO: 43 46 Nucleic acid sequence encoding SEQ ID NO: 47 47 Amino acid sequence of ferritin-HA fusion (ectodomain from hemagglutinin protein from A/Singapore/1/1957 (1957 Sing, H2)) 48 Complement of SEQ ID NO: 46 49 Nucleic acid sequence encoding SEQ ID NO: 50 50 Amino acid sequence of ferritin-HA fusion (ectodomain from hemagglutinin protein from A/Hong Kong/1/1968 (1968 HK, H3)) 51 Complement of SEQ ID NO: 49 52 Nucleic acid sequence encoding SEQ ID NO: 53 53 Amino acid sequence of ferritin-HA fusion (ectodomain from hemagglutinin protein from A/Brisbane/10/2007 (2007 Bris, H3)) 54 Complement of SEQ ID NO: 52 55 Nucleic acid sequence encoding SEQ ID NO: 56 56 Amino acid sequence of ferritin-HA fusion (ectodomain from hemagglutinin protein from A/Indonesia/05/2005 (2005 Indo, H5)) 57 Compliment of SEQ ID NO: 55 58 Nucleic acid sequence encoding SEQ ID NO: 59 59 Amino acid sequence of ferritin-HA fusion protein (ectodomain from hemagglutinin protein from B/Florida/4/2006 (2006 Flo, B)) 60 Complement of SEQ ID NO: 58 61 Nucleic acid sequence encoding SEQ ID NO: 62 62 Amino acid sequence of ferritin-HA fusion protein (ectodomain from hemagglutinin protein from A/Perth/16/2009 (2009 Per, H3)) 63 Complement of SEQ ID NO: 61 64 Nucleic acid sequence encoding SEQ ID NO: 65 65 Amino acid sequence of ferritin-HA fusion protein (ectodomain from hemagglutinin protein from A/Brisbane/59/2007 (2007 Bris, H1)) 66 Complement of SEQ ID NO: 64 67 Nucleic acid sequence encoding SEQ ID NO: 68 68 Amino acid sequence of ferritin-HA fusion protein (ectodomain from hemagglutinin protein from B/Brisbane/60/2008 (2008 Bris, B)) 69 Complement of SEQ ID NO: 67 STEM REGION 70 Nucleic acid molecule encoding SEQ ID NO: 71 71 Amino acid sequence of stem region from A/New Caledonia/20/1999 (1999 NC, H1)(GenBank: AY289929) 72 Complement of SEQ ID NO: 70 73 Nucleic acid sequence encoding SEQ ID NO: 74 74 Amino acid sequence of stem region from A/California/04/2009 (2009 CA, H1) 75 Complement of SEQ ID NO: 73 76 Nucleic acid sequence encoding SEQ ID NO: 77 77 Amino acid sequence of stem region from A/Singapore/1/1957 (1957 Sing, H2) 78 Complement of SEQ ID NO: 76 79 Nucleic acid sequence encoding SEQ ID NO: 80 80 Amino acid sequence of stem region from A/Hong Kong/1/1968 (1968 HK, H3) 81 Complement of SEQ ID NO: 79 82 Nucleic acid sequence encoding SEQ ID NO: 83 83 Amino acid sequence of stem region from A/Brisbane/10/2007 (2007 Bris, H3) 84 Complement of SEQ ID NO: 82 85 Nucleic acid sequence encoding SEQ ID NO: 86 86 Amino acid sequence of stem region from A/Indonesia/05/2005 (2005 Indo, H5) 87 Complement of SEQ ID NO: 85 88 Nucleic acid sequence encoding SEQ ID NO: 89 89 Amino acid sequence of stem region from B/Florida/4/2006 (2006 Flo, B) 90 Complement of SEQ ID NO: 88 91 Nucleic acid sequence encoding SEQ ID NO: 92 92 Amino acid sequence of stem region from A/Perth/16/2009 (2009 Per, H3) 93 Complement of SEQ ID NO: 91 94 Nucleic acid sequence encoding SEQ ID NO: 95 95 Amino acid sequence of stem region from A/Brisbane/59/2007 (2007 Bris, H1) 96 Complement of SEQ ID NO: 94 97 Nucleic acid sequence encoding SEQ ID NO: 98 98 Amino acid sequence of stem region from B/Brisbane/60/2008 (2008 Bris, B) 99 Complement of SEQ ID NO: 97 FERRITIN-HA STEM REGION FUSION 100 Nucleic acid sequence encoding SEQ ID NO: 101 101 Amino acid sequence of ferritin-HA stem region fusion protein A/New Caledonia/20/1999 (1999 NC, H1) 102 Complement of SEQ ID NO: 100 103 Nucleic acid sequence encoding SEQ ID NO: 104 104 Amino acid sequence of ferritin-HA stem region fusion protein (H1 CA) 105 Complement of SEQ ID NO: 103 106 Nucleic acid sequence encoding SEQ ID NO: 107 107 Amino acid sequence of ferritin-HA stem region fusion protein (H2 Sing) 108 Complement of SEQ ID NO: 106 109 Nucleic acid sequence encoding SEQ ID NO: 110 110 Amino acid sequence of ferritin-HA stem region fusion protein (H3 Hong Kong) 111 Complement of SEQ ID NO: 109 112 Nucleic acid sequence encoding SEQ ID NO: 113 113 Amino acid sequence of ferritin-HA stem region fusion protein (H5 Indonesia) 114 Complement of SEQ ID NO: 112 115 Nucleic acid sequence encoding SEQ ID NO: 116 116 Amino acid sequence of ferritin-HA stem region fusion protein (A/Brisbane/59/2007 (2007 Bris, H1)) 117 Complement of SEQ ID NO: 115 118 Nucleic acid sequence encoding SEQ ID NO: 119 119 Amino acid sequence of ferritin-HA stem region fusion protein (A/Brisbane/10/2007 (2007 Bris, H3)) 120 Complement of SEQ ID NO: 118 121 Nucleic acid sequence encoding SEQ ID NO: 122 122 Amino acid sequence of ferritin-HA stem region fusion protein (A/Perth/16/2009 (2009 Per, H3)) 123 Complement of SEQ ID NO: 121 124 Nucleic acid sequence encoding SEQ ID NO: 125 125 Amino acid sequence of ferritin-HA stem region fusion protein (B/Brisbane/60/2008 (2008 Bris, B) 126 Complement of SEQ ID NO: 124 127 Nucleic acid sequence encoding SEQ ID NO: 128 128 Amino acid sequence of ferritin-HA stem region fusion protein (B/Florida/4/2006 (2006 Flo, B)) 129 Complement of SEQ ID NO: 127
[0117] One type of immune response is a B-cell response, which results in the production of antibodies against the antigen that elicited the immune response. Thus, one embodiment of the present invention is a protein that elicits antibodies that bind to the stem region of influenza HA protein from a virus selected from the group consisting of influenza A viruses, influenza B viruses and influenza C viruses. One embodiment of the present invention is a protein that elicits antibodies that bind to the stem region of influenza HA protein selected from the group consisting of an H1 influenza virus HA protein, an H2 influenza virus HA protein, an influenza H3 virus HA protein, an influenza H4 virus HA protein, an influenza H5 virus HA protein, an influenza H6 virus HA protein, an H7 influenza virus HA protein, an H8 influenza virus HA protein, an H9 influenza virus HA protein, an H10 influenza virus HA protein HA protein, an H11 influenza virus HA protein, an H12 influenza virus HA protein, an H13 influenza virus HA protein, an H14 influenza virus HA protein, an H15 influenza virus HA protein and an H16 influenza virus HA protein. One embodiment of the present invention is a protein that elicits antibodies that bind to the stem region of influenza HA protein from a strain of virus selected from the viruses listed in Table 2.
[0118] While all antibodies are capable of binding to the antigen which elicited the immune response that resulted in antibody production, preferred antibodies are those that neutralize an influenza virus. Thus, one embodiment of the present invention is a protein that elicits neutralizing antibodies that bind to the stem region of influenza HA protein from a virus selected from the group consisting of influenza A viruses, influenza B viruses and influenza C viruses. One embodiment of the present invention is a protein that elicits neutralizing antibodies that bind to the stem region of influenza HA protein selected from the group consisting of an H1 influenza virus HA protein, an H2 influenza virus HA protein, an influenza H3 virus HA protein, an influenza H4 virus HA protein, an influenza H5 virus HA protein, an influenza H6 virus HA protein, an H7 influenza virus HA protein, an H8 influenza virus HA protein, an H9 influenza virus HA protein, an H10 influenza virus HA protein HA protein, an H11 influenza virus HA protein, an H12 influenza virus HA protein, an H13 influenza virus HA protein, an H14 influenza virus HA protein, an H15 influenza virus HA protein and an H16 influenza virus HA protein. One embodiment of the present invention is a protein that elicits neutralizing antibodies that bind to the stem region of influenza HA protein from a strain of virus selected from the viruses listed in Table 2. One embodiment of the present invention is a protein that elicits neutralizing antibodies that bind to a protein comprising an amino acid sequence at least 80% identical to a sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95 and SEQ ID NO:98. One embodiment of the present invention is a protein that elicits neutralizing antibodies that bind to a protein comprising an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95 and SEQ ID NO:98.
[0119] Neutralizing antibodies elicited by proteins of the present invention can neutralize viral infections by affecting any step in the life cycle of the virus. For example, neutralizing antibodies may prevent an influenza virus from attaching to a cell, entering a cell, releasing viral ribonucleoproteins into the cytoplasm, forming new viral particles in the infected cell and budding new viral particles from the infected host cell membrane. In one embodiment, neutralizing antibodies elicited by proteins of the present invention prevent influenza virus from attaching to the host cell. In one embodiment, neutralizing antibodies elicited by proteins of the present invention prevent influenza virus from entering the host cell. In one embodiment, neutralizing antibodies elicited by proteins of the present invention prevent fusion of viral membranes with endosomal membranes. In one embodiment, neutralizing antibodies elicited by proteins of the present invention prevent release of ribonucleoproteins into the cytoplasm of the host cell. In one embodiment, neutralizing antibodies elicited by proteins of the present invention prevent assembly of new virus in the infected host cell. In one embodiment, neutralizing antibodies elicited by proteins of the present invention prevent release of newly formed virus from the infected host cell.
[0120] Because the amino acid sequence of the stem region of influenza virus is highly conserved, neutralizing antibodies elicited by proteins of the present invention may be broadly neutralizing. That is, neutralizing antibodies elicited by proteins of the present invention may neutralize influenza viruses of more than one type, subtype and/or strain. Thus, one embodiment of the present invention is a protein that elicits broadly neutralizing antibodies that bind the stem region of influenza HA protein. One embodiment is a protein that elicits antibodies that bind the stem region of an HA protein from more than one type of influenza virus selected from the group consisting of influenza type A viruses, influenza type B viruses and influenza type C viruses. One embodiment is a protein that elicits antibodies that bind the stem region of an HA protein from more than one sub-type of influenza virus selected from the group consisting of an H1 influenza virus, an H2 influenza virus, an influenza H3 virus, an influenza H4 virus, an influenza H5 virus, an influenza H6 virus, an H7 influenza virus, an H8 influenza virus, an H9 influenza virus, an H10 influenza virus, an H11 influenza virus, an H12 influenza virus, an H13 influenza virus, an H14 influenza virus, an H15 influenza virus and an H16 influenza virus. One embodiment is a protein that elicits antibodies that bind the stem region of an HA protein from more than strain of influenza virus. One embodiment is a protein that elicits antibodies that bind the stem region of an HA protein from more than one strain of influenza virus selected from the viruses listed in Table 2. One embodiment of the present invention is a protein that elicits antibodies that bind more than one protein comprising an amino acid sequence at least 80% identical to a sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95 and SEQ ID NO:98. One embodiment of the present invention is a protein that elicits neutralizing antibodies that bind to more than one protein comprising an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95 and SEQ ID NO:98.
[0121] Particularly useful proteins of the present invention are those comprising an immunogenic portion of an influenza HA protein. Thus, one embodiment of the present invention is a protein comprising at least one immunogenic portion from the stem region of influenza HA protein, wherein the protein elicits neutralizing antibodies against an influenza virus. Such a protein is referred to as a stem-region protein (or a stem-region immunogen). One embodiment of the present invention is a protein comprising at least one immunogenic portion from the stem region of an HA protein from a virus selected from the group consisting of influenza type A viruses, influenza type B viruses and influenza type C viruses, wherein the protein elicits neutralizing antibodies against an influenza virus. One embodiment of the present invention is a protein comprising at least one immunogenic portion from the stem region of an HA protein selected from the group consisting of an H1 influenza virus HA protein, an H2 influenza virus HA protein, an influenza H3 virus HA protein, an influenza H4 virus HA protein, an influenza H5 virus HA protein, an influenza H6 virus HA protein, an H7 influenza virus HA protein, an H8 influenza virus HA protein, an H9 influenza virus HA protein, an H10 influenza virus HA protein HA protein, an H11 influenza virus HA protein, an H12 influenza virus HA protein, an H13 influenza virus HA protein, an H14 influenza virus HA protein, an H15 influenza virus HA protein and an H16 influenza virus HA protein. One embodiment of the present invention is a protein comprising at least one immunogenic portion from the stem region of an HA protein from the viruses listed in Table 2. One embodiment of the present invention is a protein comprising at least one immunogenic portion from a protein comprising an amino acid sequence at least 80% identical to a sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95 and SEQ ID NO:98. One embodiment of the present invention is a protein comprising at least one immunogenic portion from a protein comprising an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95 and SEQ ID NO:98. In one embodiment, such proteins comprising immunogenic portions of the HA protein elicit the production of broadly neutralizing antibodies against influenza virus.
[0122] Immunogenic portions of proteins comprise epitopes, which are clusters of amino acid residues that are recognized by the immune system, thereby eliciting an immune response. Such epitopes may consist of contiguous amino acids residues (i.e., amino acid residues that are adjacent to one another in the protein), or they may consist of non-contiguous amino acid residues (i.e., amino acid residues that are not adjacent one another in the protein) but which are in close special proximity in the finally folded protein. It is well understood by those skilled in the art that epitopes require a minimum of six amino acid residues in order to be recognized by the immune system. Thus, in one embodiment the immunogenic portion from the influenza HA protein comprises at least one epitope. One embodiment of the present invention is a protein comprising at least 6 amino acids, at least 10 amino acids, at least 25 amino acids, at least 50 amino acids, at least 75 amino acids or at least 100 amino acids from the stem region of influenza HA protein. One embodiment of the present invention is a protein comprising at least 6 amino acids, at least 10 amino acids, at least 25 amino acids, at least 50 amino acids, at least 75 amino acids or at least 100 amino acids from the stem region of an HA protein from a virus selected from the group consisting of influenza type A viruses, influenza type B viruses and influenza type C viruses. One embodiment of the present invention is a protein comprising at least 6 amino acids, at least 10 amino acids, at least 25 amino acids, at least 50 amino acids, at least 75 amino acids or at least 100 amino acids from the stem region of an HA protein selected from the group consisting an H1 influenza virus HA protein, an H2 influenza virus HA protein, an influenza H3 virus HA protein, an influenza H4 virus HA protein, an influenza H5 virus HA protein, an influenza H6 virus HA protein, an H7 influenza virus HA protein, an H8 influenza virus HA protein, an H9 influenza virus HA protein, an H10 influenza virus HA protein HA protein, an H11 influenza virus HA protein, an H12 influenza virus HA protein, an H13 influenza virus HA protein, an H14 influenza virus HA protein, an H15 influenza virus HA protein and an H16 influenza virus HA protein. One embodiment of the present invention is a protein comprising at least 6 amino acids, at least 10 amino acids, at least 25 amino acids, at least 50 amino acids, at least 75 amino acids or at least 100 amino acids from the stem region of an HA protein from a strain of virus selected from the viruses listed in Table 2. In one embodiment, the amino acids are contiguous amino acids from the stem region of the HA protein. In one embodiment, such proteins comprising at least 6 amino acids, at least 10 amino acids, at least 25 amino acids, at least 50 amino acids, at least 75 amino acids or at least 100 amino acids from the stem region of an HA protein elicit the production of broadly neutralizing antibodies against influenza virus. One embodiment of the present invention is a protein comprising at least 6 amino acids, at least 10 amino acids, at least 25 amino acids, at least 50 amino acids, at least 75 amino acids or at least 100 amino acids from the stem region of an HA protein comprising an amino acid sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. In one embodiment, the amino acids are contiguous amino acids from the stem region of the HA protein. In one embodiment, the amino acids are non-contiguous, but are in close spatial proximity in the final protein.
[0123] While the present application discloses the use of stem regions from several exemplary HA proteins having specific sequences, the invention may also be practiced using stem regions from proteins comprising variations of the disclosed HA sequences. Thus, one embodiment of the present invention is a stem-region protein comprising an amino acid sequence at least 80%, at least 85%, at least 90%, at least 92%, at least 94%, at least 96%, at least 98% or at least 99% identical the stem region of an HA protein comprising an amino acid sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. One embodiment of the present invention is a stem-region protein comprising an amino acid sequence at least 80%, at least 85%, at least 90%, at least 92%, at least 94%, at least 96%, at least 98% or at least 99% identical to a sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98. One embodiment of the present invention is a stem-region protein comprising the stem region of an HA protein comprising an amino acid sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. One embodiment of the present invention is a stem-region protein comprising an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98.
[0124] While the proteins disclosed thus far may elicit broadly neutralizing antibodies against an influenza virus, the inventors have discovered that such proteins are more stable and easier to purify when they exist in a trimeric form. Thus, one embodiment is a protein comprising the stem-region protein of the present invention joined to a trimerization domain. In one embodiment, the stem region is from an HA protein comprising an amino acid sequence at least 80% identical, at least 85% identical, at least 90% identical, at least 92% identical, at least 94% identical, at least 96% identical, at least 98% identical or at least 99% identical to a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. In one embodiment, the stem region is from an HA protein comprising an amino acid sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. In one embodiment, the stem region protein comprises an amino acid sequence at least 80% identical, at least 85% identical, at least 90% identical, at least 92% identical, at least 94% identical, at least 96% identical, at least 98% identical or at least 99% identical to an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98. In one embodiment, the stem region protein comprises an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98. In one embodiment, the trimerization domain is selected from the group consisting of the HIV-1 gp41 trimerization domain, the SIV gp41 trimerization domain, the Ebola virus gp-2 trimerization domain, the HTLV-1 gp-21 trimerization domain, the T4 fibritin trimerization domain (i.e., foldon), the yeast heat shock transcription factor trimerization domain, and the human collagen trimerization domain. In one embodiment, the trimerization domain is an HIV gp41 trimerization domain.
[0125] The inventors have also found that, in some instances, stem region proteins of the present invention may be more stable when joined to at least part of the head region of the HA protein. Thus, one embodiment of the present invention is a protein comprising a stem region protein joined to the head region of an HA protein and a trimerization domain. In one embodiment, the stem region protein is from an HA protein comprising an amino acid sequence at least 80% identical, at least 85% identical, at least 90% identical, at least 92% identical, at least 94% identical, at least 96% identical, at least 98% identical or at least 99% identical to a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. In one embodiment, the stem region protein is from an HA protein comprising an amino acid sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. In one embodiment, the stem region protein comprises an amino acid sequence at least 80% identical, at least 85% identical, at least 90% identical, at least 92% identical, at least 94% identical, at least 96% identical, at least 98% identical or at least 99% identical to an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98. In one embodiment, the stem region protein comprises an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98.
[0126] In some embodiments of the present invention, the various protein domains (e.g., stem region protein, trimerization domain, head region, etc.) may be joined directly to one another. In other embodiments, it may be necessary to employ linkers (also referred to as a spacer sequences) so that the various domains are in the proper special orientation. The linker sequence is designed to position the hemagglutinin protein in such a way to that it maintains the ability to elicit an immune response to the influenza virus. Linker sequences of the present invention comprise amino acids. Preferable amino acids to use are those having small side chains and/or those which are not charged. Such amino acids are less likely to interfere with proper folding and activity of the fusion protein. Accordingly, preferred amino acids to use in linker sequences, either alone or in combination are serine, glycine and alanine Examples of such linker sequences include, but are not limited to, SGG, GSG, GG and NGTGGSG. Amino acids can be added or subtracted as needed. Those skilled in the art are capable of determining appropriate linker sequences for proteins of the present invention.
[0127] One embodiment of the present invention is a fusion protein comprising a stem region protein joined to at least a portion of the head region of an HA protein and a trimerization domain, wherein the fusion protein comprises one or more linker sequences. In one embodiment, the stem region protein is from an HA protein comprising an amino acid sequence at least 80% identical, at least 85% identical, at least 90% identical, at least 92% identical, at least 94% identical, at least 96% identical, at least 98% identical or at least 99% identical to a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. In one embodiment, the stem region protein is from an HA protein comprising an amino acid sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. In one embodiment, the stem region protein comprises an amino acid sequence at least 80% identical, at least 85% identical, at least 90% identical, at least 92% identical, at least 94% identical, at least 96% identical, at least 98% identical or at least 99% identical to an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98. In one embodiment, the stem region protein comprises an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98. In one embodiment, the linker is selected from the group consisting of GG, GSG and NGTGGSG. In one embodiment, the protein elicits antibodies that neutralize at least one virus that is a different Type, sub-type or strain than the Type, sub-type or strain of the virus from which the HA protein was obtained.
Vaccines Comprising HA-Ferritin Fusion Proteins
[0128] The inventors have also discovered that fusion of influenza HA protein with ferritin protein (HA-ferritin fusion protein) results in a vaccine that elicits a robust immune response to influenza virus. Such HA-ferritin fusion proteins self-assemble into nanoparticles that display immunogenic portions of influenza hemagglutinin protein on their surface. These nanoparticles are useful for vaccinating individuals against a broad range of influenza viruses. Thus, one embodiment of the present invention is an HA-ferritin fusion protein comprising a monomeric ferritin subunit disclosed herein joined to an influenza hemagglutinin protein disclosed herein, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles.
[0129] Ferritin is a globular protein found in all animals, bacteria, and plants, that acts primarily to control the rate and location of polynuclear Fe(III)2O3 formation through the transportation of hydrated iron ions and protons to and from a mineralized core. The globular form of ferritin is made up of monomeric subunit proteins (also referred to as monomeric ferritin subunits), which are polypeptides having a molecule weight of approximately 17-20 kDa. An example of the sequence of one such monomeric ferritin subunit is represented by SEQ ID NO:2. Each monomeric ferritin subunit has the topology of a helix bundle which includes a four antiparallel helix motif, with a fifth shorter helix (the c-terminal helix) lying roughly perpendicular to the long axis of the 4 helix bundle. According to convention, the helices are labeled `A, B, C, and D & E` from the N-terminus respectively. The N-terminal sequence lies adjacent to the capsid three-fold axis and extends to the surface, while the E helices pack together at the four-fold axis with the C-terminus extending into the particle core. The consequence of this packing creates two pores on the capsid surface. It is expected that one or both of these pores represent the point by which the hydrated iron diffuses into and out of the capsid. Following production, these monomeric ferritin subunit proteins self-assemble into the globular ferritin protein. Thus, the globular form of ferritin comprises 24 monomeric, ferritin subunit proteins, and has a capsid-like structure having 432 symmetry.
[0130] According to the present invention, a monomeric ferritin subunit of the present invention is a full length, single polypeptide of a ferritin protein, or any portion thereof, which is capable of directing self-assembly of monomeric ferritin subunits into the globular form of the protein. Amino acid sequences from monomeric ferritin subunits of any known ferritin protein can be used to produce fusion proteins of the present invention, so long as the monomeric ferritin subunit is capable of self-assembling into a nanoparticle displaying hemagglutinin on its surface. In one embodiment, the monomeric subunit is from a ferritin protein selected from the group consisting of a bacterial ferritin protein, a plant ferritin protein, an algal ferritin protein, an insect ferritin protein, a fungal ferritin protein and a mammalian ferritin protein. In one embodiment, the ferritin protein is from Helicobacter pylori.
[0131] HA-ferritin fusion proteins of the present invention need not comprise the full-length sequence of a monomeric subunit polypeptide of a ferritin protein. Portions, or regions, of the monomeric ferritin subunit protein can be utilized so long as the portion comprises an amino acid sequence that directs self-assembly of monomeric ferritin subunits into the globular form of the protein. One example of such a region is located between amino acids 5 and 167 of the Helicobacter pylori ferritin protein. More specific regions are described in Zhang, Y. Self-Assembly in the Ferritin Nano-Cage Protein Super Family. 2011, Int. J. Mol. Sci., 12, 5406-5421, which is incorporated herein by reference in its entirety.
[0132] One embodiment of the present invention is an HA-ferritin fusion protein comprising an HA protein of the present invention joined to at least 25 contiguous amino acids, at least 50 contiguous amino acids, at least 75 contiguous amino acids, at least 100 contiguous amino acids, or at least 150 contiguous amino acids from a monomeric ferritin subunit, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles. One embodiment of the present invention is an HA-ferritin fusion protein comprising an HA protein of the present invention joined to at least 25 contiguous amino acids, at least 50 contiguous amino acids, at least 75 contiguous amino acids, at least 100 contiguous amino acids, or at least 150 contiguous amino acids from the region of a ferritin protein corresponding to the amino acid sequences of the Helicobacter pylori ferritin monomeric subunit that direct self-assembly of the monomeric subunits into the globular form of the ferritin protein, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles. One embodiment of the present invention is an HA-ferritin fusion protein comprising an HA protein of the present invention joined to at least 25 contiguous amino acids, at least 50 contiguous amino acids, at least 75 contiguous amino acids, at least 100 contiguous amino acids, or at least 150 contiguous amino acids from SEQ ID NO:2 that are capable of directing self-assembly of the monomeric subunits into the globular ferritin protein, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles. One embodiment of the present invention is an HA-ferritin fusion protein comprising an HA-protein of the present invention joined to at least 25 contiguous amino acids, at least 50 contiguous amino acids, at least 75 contiguous amino acids, at least 100 contiguous amino acids, or at least 150 contiguous amino acids from amino acid residues 5-167 of SEQ ID NO:2, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles. One embodiment of the present invention is an HA-ferritin fusion protein comprising an HA protein of the present invention joined to at least 25 contiguous amino acids, at least 50 contiguous amino acids, at least 75 contiguous amino acids, at least 100 contiguous amino acids, or at least 150 contiguous amino acids from SEQ ID NO:5, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles. One embodiment of the present invention is an HA-ferritin fusion protein comprising an HA protein of the present invention joined to amino acid residues 5-167 from SEQ ID NO:2, or SEQ ID NO:5, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles. As has been previously discussed, it is well-known in the art that some variations can be made in the amino acid sequence of a protein without affecting the activity of the protein. Such variations include insertion of amino acid residues, deletions of amino acid residues, and substitutions of amino acid residues. Thus, in one embodiment, the sequence of the monomeric ferritin subunit is divergent enough from the sequence of a ferritin subunit naturally found in a mammal, such that when the variant monomeric ferritin subunit is introduced into the mammal, it does not result in the production of antibodies that react with the mammal's natural ferritin protein. According to the present invention, such a monomeric subunit is referred to as immunogenically neutral. One embodiment of the present invention is an HA-ferritin fusion protein comprising an HA protein of the present invention joined to an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, and at least 97% identical to the amino acid sequence of a monomeric ferritin subunit that is responsible for directing self-assembly of the monomeric ferritin subunits into the globular form of the protein, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles. In one embodiment, the HA-ferritin fusion protein comprises a polypeptide sequence identical in sequence to a monomeric ferritin subunit. One embodiment of the present invention is an HA-ferritin fusion protein comprising an HA protein of the present invention joined to an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, and at least 97% identical to the amino acid sequence of a monomeric ferritin subunit from Helicobacter pylori, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles. One embodiment of the present invention is an HA-ferritin fusion protein comprising an HA protein of the present invention joined to an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, and at least 97% identical to a sequence selected from amino acid residues 5-167 from SEQ ID NO:2 and SEQ ID NO:5, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles. One embodiment of the present invention is an HA-ferritin fusion protein comprising an HA protein of the present invention joined to a sequence selected from amino acid residues 5-167 from SEQ ID NO:2 and SEQ ID NO:5.
[0133] In some embodiments, it may be useful to engineer mutations into the amino acid sequences of proteins of the present invention. For example, it may be useful to alter sites such as enzyme recognition sites or glycosylation sites in the monomeric ferritin subunit, the trimerization domain, or linker sequences, in order to give the fusion protein beneficial properties (e.g., solubility, half-life, mask portions of the protein from immune surveillance). In this regard, it is known that the monomeric subunit of ferritin is not glycosylated naturally. However, it can be glycosylated if it is expressed as a secreted protein in mammalian or yeast cells. Thus, in one embodiment, potential N-linked glycosylation sites in the amino acid sequences from the monomeric ferritin subunit are mutated so that the mutated ferritin subunit sequences are no longer glycosylated at the mutated site. One such sequence of a mutated monomeric ferritin subunit is represented by SEQ ID NO:5.
[0134] According to the present invention, the hemagglutinin protein portion of HA-ferritin fusion proteins of the present invention can be from any influenza virus, so long as the HA-ferritin fusion protein elicits an immune response against an influenza virus. Thus, one embodiment of the preset invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence from an HA protein from an influenza A virus, an influenza B virus or an influenza C virus. One embodiment of the preset invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence from an influenza A Group 1 virus HA protein. One embodiment of the preset invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence from an influenza A Group 2 virus HA protein. One embodiment of the preset invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence from an HA protein selected from the group consisting of an H1 influenza virus HA protein, an H2 influenza virus HA protein, an H5 influenza virus HA protein, an H7 virus influenza HA protein and an H9 influenza virus HA protein. One embodiment of the preset invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence from an HA protein selected from the group consisting of an H3 influenza virus HA protein, an H4 influenza virus HA protein, an H6 influenza virus HA protein, an H8 influenza virus HA protein, an H10 influenza virus HA protein, an H11 influenza virus HA protein, an H12 influenza virus HA protein, an H13 influenza virus HA protein, an H14 influenza virus HA protein, an H15 influenza virus HA protein, and an H15 influenza virus HA protein. One embodiment of the preset invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence of an HA protein from a virus listed in Table 2.
[0135] Preferred hemagglutinin proteins to use in constructing HA-ferritin fusion proteins of the present invention are those that elicit an immune response against an influenza virus. Even more preferred hemagglutinin proteins are those that are capable of eliciting antibodies to an influenza virus. One embodiment of the present invention is an HA-ferritin fusion protein that elicits antibodies to a virus selected from the group consisting of influenza A viruses, influenza B viruses and influenza C viruses. One embodiment of the present invention is a HA-ferritin fusion protein that elicits antibodies to a virus selected from the group consisting of an H1 influenza virus, an H2 influenza virus, an influenza H3 virus, an influenza H4 virus, an influenza H5 virus, an influenza H6 virus, an H7 influenza virus, an H8 influenza virus, an H9 influenza virus, an H10 influenza virus, an H11 influenza virus, an H12 influenza virus, an H13 influenza virus, an H14 influenza virus, an H15 influenza virus and an H16 influenza virus. One embodiment of the present invention is an HA-ferritin fusion protein that elicits antibodies to a virus listed in Table 2. Preferred antibodies elicited by HA-ferritin fusion proteins of the present invention are those that neutralize an influenza virus. Thus, one embodiment of the present invention is an HA-ferritin fusion protein that elicits neutralizing antibodies to a virus selected from the group consisting of influenza A viruses, influenza B viruses and influenza C viruses. One embodiment of the present invention is an HA-ferritin fusion protein that elicits neutralizing antibodies to a virus having a subtype selected from the group consisting of an H1 influenza virus, an H2 influenza virus, an influenza H3 virus, an influenza H4 virus, an influenza H5 virus, an influenza H6 virus, an H7 influenza virus, an H8 influenza virus, an H9 influenza virus, an H10 influenza virus, an H11 influenza virus, an H12 influenza virus, an H13 influenza virus, an H14 influenza virus, an H15 influenza virus and an H16 influenza virus. One embodiment of the present invention is an HA-ferritin fusion protein that elicits neutralizing antibodies to a virus listed in Table 2.
[0136] As has been discussed, neutralizing antibodies elicited by a HA-ferritin fusion protein of the present invention can neutralize viral infections by affecting any step in the life cycle of the virus. Thus, in one embodiment of the present invention, an HA-ferritin fusion protein elicits neutralizing antibodies that prevent influenza virus from attaching to the host cell. In one embodiment of the present invention, an HA-ferritin fusion protein may elicit neutralizing antibodies that prevent influenza virus from entering the host cell. In one embodiment of the present invention, an HA-ferritin fusion protein may elicit neutralizing antibodies that prevent fusion of viral membranes with endosomal membranes. In one embodiment of the present invention, an HA-ferritin fusion protein may elicit neutralizing antibodies that prevent influenza virus from releasing ribonucleoproteins into the cytoplasm of the host cell. In one embodiment of the present invention, an HA-ferritin fusion protein may elicit neutralizing antibodies that prevent assembly of new virus in the infected host cell. In one embodiment of the present invention, an HA-ferritin fusion protein may elicit neutralizing antibodies that prevent release of newly formed virus from the infected host cell.
[0137] Preferred HA-ferritin fusion proteins of the present invention are those that elicit broadly neutralizing antibodies. Thus, one embodiment is an HA-ferritin fusion protein that elicits antibodies that neutralizes more than one type of influenza virus selected from the group consisting of influenza type A viruses, influenza type B viruses and influenza type C viruses. One embodiment is an HA-ferritin fusion protein that elicits antibodies that neutralize more than one sub-type of influenza virus selected from the group consisting of an H1 influenza virus, an H2 influenza virus, an influenza H3 virus, an influenza H4 virus, an influenza H5 virus, an influenza H6 virus, an H7 influenza virus, an H8 influenza virus, an H9 influenza virus, an H10 influenza virus, an H11 influenza virus, an H12 influenza virus, an H13 influenza virus, an H14 influenza virus, an H15 influenza virus and an H16 influenza virus. One embodiment is an HA-ferritin protein that elicits antibodies that neutralize from more than one strain of influenza virus selected from the viruses listed in Table 2.
[0138] It will be understood by those skilled in the art that particularly useful HA-ferritin useful proteins of the present invention are those comprising an immunogenic portion of influenza HA protein. Thus, one embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to at least one immunogenic portion of an influenza HA protein. One embodiment of the present invention is an HA-ferritin protein comprising a ferritin protein of the present invention joined to at least one immunogenic portion of an HA protein from a virus selected from the group consisting of influenza type A viruses, influenza type B viruses and influenza type C viruses. One embodiment of the present invention is an HA-ferritin protein comprising a ferritin protein of the present invention joined to at least one immunogenic portion of an HA protein selected from the group consisting of an H1 influenza virus HA protein, an H2 influenza virus HA protein, an H5 influenza virus HA protein, an H7 virus influenza HA protein and an H9 influenza virus HA protein. One embodiment of the preset invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to at least one immunogenic portion of an HA protein selected from the group consisting of an H3 influenza virus HA protein, an H4 influenza virus HA protein, an H6 influenza virus HA protein, an H8 influenza virus HA protein, an H10 influenza virus HA protein, an H11 influenza virus HA protein, an H12 influenza virus HA protein, an H13 influenza virus HA protein, an H14 influenza virus HA protein, an H15 influenza virus HA protein, and an H16 influenza virus HA protein, joined to a ferritin protein of the present invention. One embodiment of the present invention is an HA-ferritin protein comprising a ferritin protein of the present invention joined to at least one immunogenic portion of an HA protein from virus listed in Table 2. In one embodiment, an HA-ferritin fusion protein comprising an immunogenic portion of an HA protein elicits the production of broadly neutralizing antibodies against influenza virus.
[0139] Immunogenic portions of proteins comprise epitopes, which are clusters of amino acid residues that are recognized by the immune system, thus eliciting an immune response. Such epitopes may consist of contiguous amino acids residues (i.e., amino acid residues that are adjacent to one another in the protein), or they may consist of non-contiguous amino acid residues (i.e., amino acid residues that are not adjacent one another in the protein) but which are in close special proximity in the finally folded protein. It is well understood by those skilled in the art that such epitopes require a minimum of six amino acid residues in order to be recognized by the immune system. Thus, one embodiment of the present invention is an HA-ferritin fusion comprising an immunogenic portion from the influenza HA protein, wherein the immunogenic portion comprises at least one epitope.
[0140] It is known in the art that some variation in a protein sequence can be tolerated without significantly affecting the activity of the protein. Thus, one embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence that is a variant of an HA protein from a virus selected from the group consisting of influenza Type A viruses influenza Type B viruses and influenza type C viruses. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the sequence of a HA protein from a virus selected from the group consisting of influenza Type A viruses influenza Type B viruses and influenza type C viruses, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the sequence of a HA protein selected from the group consisting an H1 influenza virus HA protein, an H2 influenza virus HA protein, an H5 influenza virus HA protein, an H7 virus influenza HA protein and an H9 influenza virus HA protein, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the sequence of a HA protein selected from the group consisting of an H3 influenza virus HA protein, an H4 influenza virus HA protein, an H6 influenza virus HA protein, an H8 influenza virus HA protein, an H10 influenza virus HA protein, an H11 influenza virus HA protein, an H12 influenza virus HA protein, an H13 influenza virus HA protein, an H14 influenza virus HA protein, an H15 influenza virus HA protein, and an H16 influenza virus HA protein, joined to a ferritin protein of the present invention, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the sequence of a HA protein from a virus listed in Table 2, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to amino acid sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38.
[0141] One embodiment of the present invention is an HA-ferritin fusion protein comprising an amino acid sequence at least 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to a sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, and SEQ ID NO:68. One embodiment of the present invention is an HA-ferritin fusion protein comprising an amino acid sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, and SEQ ID NO:68.
[0142] It is known in the art that influenza hemagglutinin proteins have various regions, or domains, each possessing specific activities. For example, the globular head extends out from the lipid membrane and comprises two domains: the receptor binding domain (RBD) and the vestigial esterase domain. The RB domain is involved in binding of the HA protein to receptors. The globular head also includes several antigenic sites that include immunodominant epitopes. The stem region is responsible for anchoring the HA protein into the viral lipid envelope. Thus, it will be understood by those skilled in the art that HA-ferritin fusion proteins of the present invention need not comprise the entire sequence of the HA protein. Instead, an HA-ferritin fusion protein can comprise only those portions, regions, domains, and the like, that contain the necessary activities for practicing the present invention. For example, an HA-ferritin fusion protein may contain only those amino acid sequences from the HA protein that contain antigenic sites, epitopes, immunodominant epitopes, and the like.
[0143] One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, at least 200 amino acids, at least 300 amino acids, at least 400 amino acids, or at least 500 amino acids from an HA protein from a virus selected from the group consisting of influenza Type A viruses influenza Type B viruses and influenza type C viruses, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, at least 200 amino acids, at least 300 amino acids, at least 400 amino acids, or at least 500 amino acids from an HA protein selected from the group consisting an H1 influenza virus HA protein, an H2 influenza virus HA protein, an H5 influenza virus HA protein, an H7 virus influenza HA protein and an H9 influenza virus HA protein, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, at least 200 amino acids, at least 300 amino acids, at least 400 amino acids, or at least 500 amino acids from an HA protein selected from the group consisting of an H3 influenza virus HA protein, an H4 influenza virus HA protein, an H6 influenza virus HA protein, an H8 influenza virus HA protein, an H10 influenza virus HA protein, an H11 influenza virus HA protein, an H12 influenza virus HA protein, an H13 influenza virus HA protein, an H14 influenza virus HA protein, an H15 influenza virus HA protein, and an H16 influenza virus HA protein, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, at least 200 amino acids, at least 300 amino acids, at least 400 amino acids, or at least 500 amino acids from and HA protein from a virus listed in Table 2, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against in influenza virus. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, at least 200 amino acids, at least 300 amino acids, at least 400 amino acids, or at least 500 amino acids from a protein consisting of a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38.
[0144] One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to at least one domain from a HA protein from a virus listed in Table 2, wherein the domain is selected from the group consisting of an ectodomain, an RDB domain, a stem domain, and a domain comprising the region stretching from the amino acid residue immediately distal to the last amino acid of second helical coil to the amino acid residue proximal to the first amino acid of the transmembrane domain. According to the present invention, an ectodomain of an influenza hemagglutinin protein refers to the portion of the hemagglutinin protein that lies outside its transmembrane domain. In one embodiment, the HA-ferritin fusion protein comprises a ferritin protein of the present invention joined to a region of a HA protein from a virus listed in Table 2, wherein the region consists of the amino acid immediately distal to the last amino acid of the second helical coiled coil and proximal to the first amino acid of the transmembrane domain. In one embodiment, the HA-ferritin fusion protein comprises a ferritin protein of the present invention joined to a region of a HA protein from a virus listed in Table 2, wherein the region comprises an amino acid sequence distal to the second helical coiled coil and proximal to the transmembrane domain. In one embodiment, the HA-ferritin fusion protein comprises a ferritin protein of the present invention joined to the ectodomain of a HA protein from a virus listed in Table 2. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38.
[0145] The stem region of an influenza HA protein is a particularly useful domain for constructing fusion proteins of the present invention. Thus, one embodiment of the present invention is a ferritin protein of the present invention joined to at least one immunogenic portion from the stem region of influenza HA protein. According to the preset invention, such a protein is referred to an HA SS-ferritin fusion protein. As used herein, the HA stem region of the hemagglutinin protein consists of the amino acids from the membrane up to the head region of the protein. More specifically, the stem region consists of the amino terminal amino acid up to the cysteine at position 52, and all residues after the cysteine residue at position 277 (using standard H3 numbering). Sequences of exemplary stem regions are represented by SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95 and SEQ ID NO:98.
[0146] One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, or at least 200 amino acids from the stem region of an HA protein from a virus selected from the group consisting of influenza Type A viruses influenza Type B viruses and influenza type C viruses, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, or at least 200 amino acids from the stem region of an HA protein selected from the group consisting an H1 influenza virus HA protein, an H2 influenza virus HA protein, an H5 influenza virus HA protein, an H7 virus influenza HA protein and an H9 influenza virus HA protein, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, or at least 200 amino acids from the stem region of an HA protein selected from the group consisting of an H3 influenza virus HA protein, an H4 influenza virus HA protein, an H6 influenza virus HA protein, an H8 influenza virus HA protein, an H10 influenza virus HA protein, an H11 influenza virus HA protein, an H12 influenza virus HA protein, an H13 influenza virus HA protein, an H14 influenza virus HA protein, an H15 influenza virus HA protein, and an H16 influenza virus HA protein, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, or at least 200 amino acids from the stem region of an HA protein from a virus listed in Table 2, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against in influenza virus. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, or at least 200 amino acids from the stem region of an HA protein comprising a sequence selected from the group consisting of SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, or at least 200 amino acids from the stem region comprising a sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95 and SEQ ID NO:98.
[0147] One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the sequence of the stem region of an HA protein from a virus selected from the group consisting of influenza Type A viruses influenza Type B viruses and influenza type C viruses, wherein the Ha-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the sequence of the stem region of an HA protein selected from the group consisting an H1 influenza virus HA protein, an H2 influenza virus HA protein, an H5 influenza virus HA protein, an H7 virus influenza HA protein and an H9 influenza virus HA protein, wherein the Ha-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the sequence of the stem region of an HA protein selected from the group consisting of an H3 influenza virus HA protein, an H4 influenza virus HA protein, an H6 influenza virus HA protein, an H8 influenza virus HA protein, an H10 influenza virus HA protein, an H11 influenza virus HA protein, an H12 influenza virus HA protein, an H13 influenza virus HA protein, an H14 influenza virus HA protein, an H15 influenza virus HA protein, and an H16 influenza virus HA protein, joined to a ferritin protein of the present invention, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the sequence of the stem region of an HA protein from a virus listed in Table 2, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the stem region of an HA protein comprising an amino acid sequence selected from the group consisting of SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38., wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95 and SEQ ID NO:98, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95 and SEQ ID NO:98.
[0148] As has been described for stem region proteins of the present invention, the inventors have found that HA-ferritin fusion proteins are more stable and easier to purify when they exist in a trimeric form. Thus, in one embodiment of the present invention the HA portion of the HA-ferritin fusion protein is joined to one or more trimerization domains. In one embodiment, the HA protein comprises an amino acid sequence at least 80% identical, at least 85% identical, at least 90% identical, at least 92% identical, at least 94% identical, at least 96% identical, at least 98% identical or at least 99% identical to a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38, joined to one or more trimerization domains. In one embodiment, the HA protein comprises an amino acid sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38 joined to one or more trimerization domains. In one embodiment, the HA protein comprises an amino acid sequence at least 80% identical, at least 85% identical, at least 90% identical, at least 92% identical, at least 94% identical, at least 96% identical, at least 98% identical or at least 99% identical to an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98 joined to one or more trimerization domains. In one embodiment, the HA protein comprises an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98 joined to one or more trimerization domains. In one embodiment, the trimerization domain is selected from the group consisting of the HIV-1 gp41 trimerization domain, the SIV gp41 trimerization domain, the Ebola virus gp-2 trimerization domain, the HTLV-1 gp-21 trimerization domain, the T4 fibritin trimerization domain (i.e., foldon), the yeast heat shock transcription factor trimerization domain, and the human collagen trimerization domain. In one embodiment, the trimerization domain is an HIV gp41 trimerization domain.
[0149] Additionally, the inventors have found that, in some instances, HA-ferritin fusion proteins in which the HA portion is limited to HA stem region sequences may be more stable when joined to at least part of the head region of the HA protein. Thus, one embodiment of the present invention is an HA 55-ferritin fusion protein, wherein, the HA portion of the fusion protein is joined to an amino acid sequence from at least a portion of an HA protein head region.
[0150] HA-ferritin proteins of the present invention are constructed by joining ferritin proteins of the present invention with HA proteins of the present invention. In addition, HA-ferritin fusion proteins may contain other domains (e.g., stem region protein, trimerization domain, head region, etc.) that improve the functionality of the final HA-ferritin fusion protein. In some embodiments, joining of the various proteins and/or domains can be done such that the sequences are directly linked. In other embodiments, it may be necessary to employ linkers (also referred to as a spacer sequences) between the various proteins and/or domains so that the so that they are in the proper special orientation. More specifically, linker sequence can be inserted so that the hemagglutinin protein is positioned in such a way to maintain the ability to elicit an immune response to the influenza virus. Linker sequences of the present invention comprise amino acids. Preferable amino acids to use are those having small side chains and/or those which are not charged. Such amino acids are less likely to interfere with proper folding and activity of the fusion protein. Accordingly, preferred amino acids to use in linker sequences, either alone or in combination are serine, glycine and alanine Examples of such linker sequences include, but are not limited to, SGG, GSG, GG and NGTGGSG. Amino acids can be added or subtracted as needed. Those skilled in the art are capable of determining appropriate linker sequences for proteins of the present invention.
[0151] In accordance with the invention, suitable portions of the hemagglutinin protein can be joined to the ferritin protein either as an exocapsid product by fusion with the N-terminal sequence lying adjacent to the capsid three-fold axis, as an endocapsid product by fusion with the C-terminus extending inside the capsid core, or a combination thereof. In one embodiment, the hemagglutinin portion of the fusion protein is joined to the N-terminal sequence of the ferritin portion of the fusion protein.
[0152] One embodiment of the present invention is an HA-ferritin fusion protein comprising an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128, wherein the HA-ferritin fusion protein elicits the production of neutralizing antibodies against an influenza protein. One embodiment of the present invention is an HA-ferritin fusion protein comprising SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128.
[0153] Proteins of the present invention are encoded by nucleic acid molecules of the present invention. In addition, they are expressed by nucleic acid constructs of the present invention. As used herein a nucleic acid construct is a recombinant expression vector, i.e., a vector linked to a nucleic acid molecule encoding a protein such that the nucleic acid molecule can effect expression of the protein when the nucleic acid construct is administered to, for example, a subject or an organ, tissue or cell. The vector also enables transport of the nucleic acid molecule to a cell within an environment, such as, but not limited to, an organism, tissue, or cell culture. A nucleic acid construct of the present disclosure is produced by human intervention. The nucleic acid construct can be DNA, RNA or variants thereof. The vector can be a DNA plasmid, a viral vector, or other vector. In one embodiment, a vector can be a cytomegalovirus (CMV), retrovirus, adenovirus, adeno-associated virus, herpes virus, vaccinia virus, poliovirus, sindbis virus, or any other DNA or RNA virus vector. In one embodiment, a vector can be a pseudotyped lentiviral or retroviral vector. In one embodiment, a vector can be a DNA plasmid. In one embodiment, a vector can be a DNA plasmid comprising viral components and plasmid components to enable nucleic acid molecule delivery and expression. Methods for the construction of nucleic acid constructs of the present disclosure are well known. See, for example, Molecular Cloning: a Laboratory Manual, 3rd edition, Sambrook et al. 2001 Cold Spring Harbor Laboratory Press, and Current Protocols in Molecular Biology, Ausubel et al. eds., John Wiley & Sons, 1994. In one embodiment, the vector is a DNA plasmid, such as a CMV/R plasmid such as CMV/R or CMV/R 8 KB (also referred to herein as CMV/R 8 kb). Examples of CMV/R and CMV/R 8 kb are provided herein. CMV/R is also described in U.S. Pat. No. 7,094,598 B2, issued Aug. 22, 2006.
[0154] As used herein, a nucleic acid molecule comprises a nucleic acid sequence that encodes a stem region immunogen, a ferritin monomeric subunit, a hemagglutinin protein, and/or an HA-ferritin fusion protein of the present invention. A nucleic acid molecule can be produced recombinantly, synthetically, or by a combination of recombinant and synthetic procedures. A nucleic acid molecule of the disclosure can have a wild-type nucleic acid sequence or a codon-modified nucleic acid sequence to, for example, incorporate codons better recognized by the human translation system. In one embodiment, a nucleic acid molecule can be genetically-engineered to introduce, or eliminate, codons encoding different amino acids, such as to introduce codons that encode an N-linked glycosylation site. Methods to produce nucleic acid molecules of the disclosure are known in the art, particularly once the nucleic acid sequence is know. It is to be appreciated that a nucleic acid construct can comprise one nucleic acid molecule or more than one nucleic acid molecule. It is also to be appreciated that a nucleic acid molecule can encode one protein or more than one protein.
[0155] Preferred nucleic acid molecules are those that encode a stem-region protein, a ferritin monomeric subunit, a hemagglutinin protein, and/or an HA-ferritin fusion protein comprising a monomeric subunit of a ferritin protein joined to an influenza hemagglutinin protein. Thus, one embodiment of the present invention is a nucleic acid molecule comprising a nucleic acid sequence encoding a protein that comprises a monomeric subunit of a ferritin protein joined to an influenza hemagglutinin protein. In one embodiment, the monomeric subunit of ferritin is from the ferritin protein of Helicobacter pylori. In one embodiment, the monomeric subunit comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 92%, at least 94%, at least 96%, at least 98% or at least 99% identical to a sequence selected from the group consisting of SEQ ID NO:2 and SEQ ID NO:5. In one embodiment, the monomeric subunit comprises an amino acid sequence selected from the group consisting of SEQ ID NO:2 and SEQ ID NO:5. In one embodiment the influenza hemagglutinin protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 92%, at least 94%, at least 96%, at least 98% or at least 99% identical to a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. In one embodiment the influenza hemagglutinin protein comprises a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. In one embodiment the influenza hemagglutinin protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 92%, at least 94%, at least 96%, at least 98% or at least 99% identical to a sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98. In one embodiment the influenza hemagglutinin protein comprises a sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98. In one embodiment the influenza hemagglutinin protein comprises at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, or at least 200 amino acids from a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. In one embodiment the influenza hemagglutinin protein comprises at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, or at least 200 amino acids from a sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98.
[0156] One embodiment of the present invention is a nucleic acid molecule comprising a nucleic sequence encoding a protein comprising an amino acid sequence at least 80%, at least 85%, at least 90%, at least 92%, at least 94%, at least 96%, at least 98% or at least 99% identical to a sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, and SEQ ID NO:68. One embodiment of the present invention is a nucleic acid molecule comprising a nucleic sequence encoding an amino acid sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, and SEQ ID NO:68.
[0157] One embodiment of the present invention is a nucleic acid molecule comprising a nucleic sequence encoding a protein comprising an amino acid sequence at least 80%, at least 85%, at least 90%, at least 92%, at least 94%, at least 96%, at least 98% or at least 99% identical to SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128. One embodiment of the present invention is a nucleic acid molecule comprising a nucleic sequence encoding an amino acid sequence selected from the group consisting of SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128.
[0158] Also embodied in the present invention are nucleic acid sequences that are variants of nucleic acid sequence encoding protein of the present invention. Such variants include nucleotide insertions, deletions, and substitutions, so long as they do not affect the ability of fusion proteins of the present invention to self-assemble into nanoparticles, or significantly affect the ability of the hemagglutinin portion of fusion proteins to elicit an immune response to an influenza virus. Thus, one embodiment of the present invention is a nucleic acid molecule encoding a fusion protein of the present invention, wherein the monomeric subunit is encoded by a nucleotide sequence at least 85%, at least 90%, at least 95%, or at least 97% identical to a nucleotide sequence selected from the group consisting of SEQ ID NO:1 and SEQ ID NO:4. One embodiment of the present invention is a nucleic acid molecule encoding an HA-ferritin fusion protein of the present invention, wherein the HA protein is encoded by a nucleotide sequence at least 85%, at least 90%, at least 95%, at least 97% identical or at least 99% identical to a nucleic acid sequence encoding an amino acid sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. One embodiment of the present invention is a nucleic acid molecule encoding an HA-ferritin fusion protein of the present invention, wherein the HA protein is encoded by a nucleotide sequence at least 85%, at least 90%, at least 95%, at least 97% identical or at least 99% identical to a nucleic acid sequence encoding an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98.
[0159] One embodiment of the present invention is a nucleic acid molecule comprising a nucleic acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to a nucleic acid sequence selected from the group consisting of SEQ ID NO:7, SEQ ID NO:10, SEQ ID NO:13, SEQ ID NO:16, SEQ ID NO:19, SEQ ID NO:22, SEQ ID NO:25, SEQ ID NO:28, SEQ ID NO:31, SEQ ID NO:34, and SEQ ID NO:37. One embodiment of the present invention is a nucleic acid molecule comprising a nucleic acid sequence selected from the group consisting of SEQ ID NO:7, SEQ ID NO:10, SEQ ID NO:13, SEQ ID NO:16, SEQ ID NO:19, SEQ ID NO:22, SEQ ID NO:25, SEQ ID NO:28, SEQ ID NO:31, SEQ ID NO:34, and SEQ ID NO:37.
[0160] One embodiment of the present invention is a nucleic acid molecule comprising a nucleic acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to a nucleic acid sequence selected from the group consisting of SEQ ID NO:40, SEQ ID NO:43, SEQ ID NO:46, SEQ ID NO:49, SEQ ID NO:52, SEQ ID NO:55, SEQ ID NO:58, SEQ ID NO:61, SEQ ID NO:64, and SEQ ID NO:67. One embodiment of the present invention is a nucleic acid molecule comprising a nucleic acid sequence selected from the group consisting of SEQ ID NO:40, SEQ ID NO:43, SEQ ID NO:46, SEQ ID NO:49, SEQ ID NO:52, SEQ ID NO:55, SEQ ID NO:58, SEQ ID NO:61, SEQ ID NO:64, and SEQ ID NO:67.
[0161] One embodiment of the present invention is a nucleic acid molecule comprising a nucleic acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to a nucleic acid sequence selected from the group consisting of SEQ ID NO:70, SEQ ID NO:73, SEQ ID NO:76, SEQ ID NO:79, SEQ ID NO:82, SEQ ID NO:85, SEQ ID NO:88, SEQ ID NO:91, SEQ ID NO:94, and SEQ ID NO:97. One embodiment of the present invention is a nucleic acid molecule comprising a nucleic acid sequence selected from the group consisting of SEQ ID NO:70, SEQ ID NO:73, SEQ ID NO:76, SEQ ID NO:79, SEQ ID NO:82, SEQ ID NO:85, SEQ ID NO:88, SEQ ID NO:91, SEQ ID NO:94, and SEQ ID NO:97.
[0162] One embodiment of the present invention is a nucleic acid molecule comprising a nucleic acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to a nucleic acid sequence selected from the group consisting of SEQ ID NO:100, SEQ ID NO:103, SEQ ID NO:106, SEQ ID NO:109, SEQ ID NO:112, SEQ ID NO:115, SEQ ID NO:121, SEQ ID NO:124, and SEQ ID NO:127. One embodiment of the present invention is a nucleic acid molecule comprising a nucleic acid sequence selected from the group consisting of SEQ ID NO:100, SEQ ID NO:103, SEQ ID NO:106, SEQ ID NO:109, SEQ ID NO:112, SEQ ID NO:115, SEQ ID NO:121, SEQ ID NO:124, and SEQ ID NO:127.
[0163] Also encompassed by the present invention are expression systems for producing fusion proteins of the present invention. In one embodiment, nucleic acid molecules of the present invention are operationally linked to a promoter. As used herein, operationally linked means that proteins encoded by the linked nucleic acid molecules can be expressed when the linked promoter is activated. Promoters useful for practicing the present invention are known to those skilled in the art. One embodiment of the present invention is a recombinant cell comprising a nucleic acid molecule of the present invention. One embodiment of the present invention is a recombinant virus comprising a nucleic acid molecule of the present invention.
[0164] As indicated above, the recombinant production of the ferritin fusion proteins of the present invention can take place using any suitable conventional recombinant technology currently known in the field. For example, molecular cloning a fusion protein, such as ferritin with a suitable protein such as the recombinant influenza hemagglutinin protein, can be carried out via expression in E. coli with the suitable monomeric subunit protein, such as the helicobacter pylori ferritin monomeric subunit. The construct may then be transformed into protein expression cells, grown to suitable size, and induced to produce the fusion protein.
[0165] As has been described, because HA-ferritin fusion proteins of the present invention comprise a monomeric subunit of ferritin, they can self-assemble. According to the present invention, the supramolecule resulting from such self-assembly is referred to as a hemagglutinin expressing ferritin based nanoparticle. For ease of discussion, the hemagglutinin expressing ferritin based nanoparticle will simply be referred to as a, or the, nanoparticle (np). Nanoparticles of the present invention have the same structural characteristics as the ferritin proteins described earlier. That is, they contain 24 subunits and have 432 symmetry. In the case of nanoparticles of the present invention, the subunits are the fusion proteins comprising a ferritin monomeric subunit joined to an influenza hemagglutinin protein. Such nanoparticles display at least a portion of the hemagglutinin protein on their surface as hemagglutinin trimers. In such a construction, the hemagglutinin trimer is accessible to the immune system and thus can elicit an immune response. Thus, one embodiment of the present invention is a nanoparticle comprising an HA-ferritin fusion protein, wherein the fusion protein comprises a monomeric ferritin subunit joined to an influenza hemagglutinin protein. In one embodiment, the nanoparticle is an octahedron. In one embodiment, the nanoparticle displays the hemagglutinin protein on its surface as a hemagglutinin trimer. In one embodiment, the influenza hemagglutinin protein is capable of eliciting neutralizing antibodies to an influenza virus. In one embodiment, the monomeric ferritin subunit comprises at least 50 amino acids, at least 100 amino acids, or at least 150 amino acids from an amino acid sequence selected from the group consisting of SEQ ID NO:2 and SEQ ID NO:5, and/or comprises an amino acid sequence at least 85%, at least 90%, at least 95%, at least 97% at least 99% identical to an amino acid sequence selected from the group consisting of SEQ ID NO:2 and SEQ ID NO:5. In one embodiment, the monomeric ferritin subunit comprises SEQ ID NO:2 or SEQ ID NO:5.
[0166] In one embodiment, the influenza hemagglutinin protein comprises at least one epitope from an influenza hemagglutinin protein listed in Table 2. In one embodiment, the influenza hemagglutinin protein comprises at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, at least 200 amino acids, at least 300 amino acids, at least 400 amino acids, or at least 500 amino acids from a hemagglutinin protein of a virus listed in Table 2. In one embodiment, the hemagglutinin protein comprises at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, at least 200 amino acids, at least 300 amino acids, at least 400 amino acids, or at least 500 amino acids from a protein consisting of a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. In one embodiment, the hemagglutinin protein comprises a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38.
[0167] In one embodiment, the influenza hemagglutinin protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, at least 97%, at least 99% identical to the sequence of an hemagglutinin protein from a virus listed in Table 2. In one embodiment, the influenza hemagglutinin protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, at least 97%, at least 99% identical to a protein sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38.
[0168] In one embodiment, the hemagglutinin protein comprises at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, at least 200 amino acids, at least 300 amino acids, at least 400 amino acids, or at least 500 amino acids from a protein consisting of a sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98. In one embodiment, the influenza hemagglutinin protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, at least 97%, at least 99% identical to a protein sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98. In one embodiment, the hemagglutinin protein comprises a sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98.
[0169] In one embodiment, the HA-ferritin fusion protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, at least 97%, at least 99% identical to a protein sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, and SEQ ID NO:68. In one embodiment, the HA-ferritin fusion protein comprises an amino acid sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, and SEQ ID NO:68. In one embodiment, the HA-ferritin fusion protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, at least 97%, at least 99% identical to SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128. In one embodiment, the HA-ferritin fusion protein comprises SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128.
[0170] Because stem region immunogens, HA-ferritin fusion proteins and nanoparticles of the present invention can elicit an immune response to an influenza virus, they can be used as vaccines to protect individuals against infection by influenza virus. According to the present invention a vaccine can be a stem region immunogen, an HA-ferritin fusion protein, or a nanoparticle of the present invention. Thus, one embodiment of the present invention is a vaccine comprising a stem region immunogen, an HA-ferritin fusion protein, or a nanoparticle of the present invention. Vaccines of the present invention can also contain other components such as adjuvants, buffers and the like. Although any adjuvant can be used, preferred embodiments can contain: chemical adjuvants such as aluminum phosphate, benzyalkonium chloride, ubenimex, and QS21; genetic adjuvants such as the IL-2 gene or fragments thereof, the granulocyte macrophage colony-stimulating factor (GM-CSF) gene or fragments thereof, the IL-18 gene or fragments thereof, the chemokine (C--C motif) ligand 21 (CCL21) gene or fragments thereof, the IL-6 gene or fragments thereof, CpG, LPS, TLR agonists, and other immune stimulatory genes; protein adjuvants such IL-2 or fragments thereof, the granulocyte macrophage colony-stimulating factor (GM-CSF) or fragments thereof, IL-18 or fragments thereof, the chemokine (C--C motif) ligand 21 (CCL21) or fragments thereof, IL-6 or fragments thereof, CpG, LPS, TLR agonists and other immune stimulatory cytokines or fragments thereof; lipid adjuvants such as cationic liposomes, N3 (cationic lipid), monophosphoryl lipid A (MPL1); other adjuvants including cholera toxin, enterotoxin, Fms-like tyrosine kinase-3 ligand (Flt-3L), bupivacaine, marcaine, and levamisole.
[0171] One embodiment of the disclosure is a ferritin-based nanoparticle vaccine that includes more than one influenza hemagglutinin protein. Such a vaccine can include a combination of different influenza hemagglutinin proteins, either on a single nanoparticle or as a mixture of nanoparticles, at least two of which have a unique influenza hemagglutinin proteins. A multivalent vaccine can comprise as many influenza hemagglutinin proteins as necessary in order to result in production of the immune response necessary to protect against a desired breadth of virus strains. In one embodiment, the vaccine comprises a hemagglutinin protein from at least two different influenza strains (bi-valent). In one embodiment, the vaccine comprises a hemagglutinin protein from at least three different influenza strains (tri-valent). In one embodiment, the vaccine comprises a hemagglutinin protein from at least four different influenza strains (tetra-valent). In one embodiment, the vaccine comprises a hemagglutinin protein from at least five different influenza strains (penta-valent). In one embodiment, the vaccine comprises a hemagglutinin protein from at least six different influenza strains (hexa-valent). In various embodiments, a vaccine comprises a hemagglutinin protein from each of 7, 8, 9, or 10 different strains of influenza virus. An example of such a combination is a ferritin-based nanoparticle vaccine that comprises influenza A group 1 hemagglutinin protein, an influenza A group 2 hemagglutinin protein, and an influenza B hemagglutinin protein. In one embodiment, the influenza hemagglutinin proteins are H1 HA, H3 HA, and B HA. In one embodiment, the influenza hemagglutinin proteins are those included in the 2011-2012 influenza vaccine. Another example of a multivalent vaccine is a ferritin based nanoparticle vaccine that comprises hemagglutinin proteins from four different influenza viruses. In one embodiment, the multivalent vaccine comprises hemagglutinin proteins from H1 A/NC/20/1999, H1 A/CA/04/2009, H2 A/Singapore/1/1957 and H5 A/Indonesia/05/2005. Such a vaccine is described in Example 2.
[0172] One embodiment of the present invention is a method to vaccinate an individual against influenza virus, the method comprising administering a nanoparticle to an individual such that an immune response against influenza virus is produced in the individual, wherein the nanoparticle comprises a monomeric subunit from ferritin joined to an influenza hemagglutinin protein, and wherein the nanoparticle displays the influenza hemagglutinin on its surface. In one embodiment, the nanoparticle is a monovalent nanoparticle. In one embodiment, the nanoparticle is multivalent nanoparticle. Another embodiment of the present invention is a method to vaccinate an individual against infection with influenza virus, the method comprising:
[0173] a) obtaining a nanoparticle comprising monomeric subunits, wherein the monomeric subunits comprise a ferritin protein joined to an influenza hemagglutinin protein, and wherein the nanoparticle displays the influenza hemagglutinin on its surface; and,
[0174] b) administering the nanoparticle to an individual such that an immune response against an influenza virus is produced.
[0175] One embodiment of the present invention is a method to vaccinate an individual against influenza virus, the method comprising administering a vaccine of the embodiments to an individual such that an immune response against influenza virus is produced in the individual, wherein the vaccine comprises at least one nanoparticle comprising a monomeric subunit from ferritin joined to an influenza hemagglutinin protein, and wherein the nanoparticle displays the influenza hemagglutinin on its surface. In one embodiment, the vaccine is a stem region immunogen. In one embodiment, the vaccine is a nanoparticle. In one embodiment, the vaccine is a monovalent vaccine. In one embodiment, the vaccine is multivalent vaccine. Another embodiment of the present invention is a method to vaccinate an individual against infection with influenza virus, the method comprising:
[0176] a) obtaining a vaccine comprising at least one nanoparticle comprising an HA-ferritin fusion protein, wherein the fusion protein comprises a ferritin protein joined to an influenza HA protein, and wherein the nanoparticle displays the influenza HA on its surface; and,
[0177] b) administering the vaccine to an individual such that an immune response against an influenza virus is produced.
[0178] In one embodiment, the nanoparticle is a monovalent nanoparticle. In one embodiment, the nanoparticle is multivalent nanoparticle.
[0179] In one embodiment, the nanoparticle is an octahedron. In one embodiment, the influenza hemagglutinin protein is capable of eliciting neutralizing antibodies to an influenza virus. In one embodiment, the influenza HA protein is capable of eliciting broadly neutralizing antibodies to an influenza virus. In one embodiment, the ferritin portion of the fusion protein comprise at least 50 amino acids, at least 100 amino acids, or at least 150 amino acids from an amino acid sequence selected from the group consisting of SEQ ID NO:2 and SEQ ID NO:5, and/or comprises an amino acid sequence at least 85%, at least 90%, at least 95%, at least 97% at least 99% identical to an amino acid sequence selected from the group consisting of SEQ ID NO:2 and SEQ ID NO:5. In one embodiment, the HA portion of the fusion protein comprises at least one epitope from an influenza hemagglutinin protein listed in Table 2. In one embodiment, the HA portion of the fusion protein comprises at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, at least 200 amino acids, at least 300 amino acids, at least 400 amino acids, or at least 500 amino acids from a hemagglutinin protein of a virus listed in Table 2. In one embodiment, the HA portion of the fusion protein comprises at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, at least 200 amino acids, at least 300 amino acids, at least 400 amino acids, or at least 500 amino acids from a protein consisting of a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. In one embodiment, the HA portion of the fusion protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, at least 97%, at least 99% identical to the sequence of an HA protein from a virus listed in Table 2. In one embodiment, the HA portion of the fusion protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, at least 97%, at least 99% identical to a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. In one embodiment, the HA portion of the fusion protein comprises at least 25 amino acids, at least 50 amino acids, at least 75 amino acids, at least 100 amino acids, at least 150 amino acids, at least 200 amino acids, at least 300 amino acids, at least 400 amino acids, or at least 500 amino acids from a sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98. In one embodiment, the HA portion of the fusion protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, at least 97%, at least 99% identical to a sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95, and SEQ ID NO:98. In one embodiment, the HA-ferritin fusion protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, at least 97%, at least 99% identical to a protein sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, and SEQ ID NO:68. In one embodiment, the HA-ferritin fusion protein comprises an amino acid sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, and SEQ ID NO:68. In one embodiment, the HA-ferritin fusion protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, at least 97%, at least 99% identical to SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128. In one embodiment, the HA-ferritin fusion protein comprises SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128.
[0180] Vaccines of the present invention can be used to vaccinate individuals using a prime/boost protocol. Such a protocol is described in U.S. Patent Publication No. 20110177122, which is incorporated herein by reference in its entirety. In such a protocol, a first vaccine composition may be administered to the individual (prime) and then after a period of time, a second vaccine composition may be administered to the individual (boost). Administration of the boosting composition is generally weeks or months after administration of the priming composition, preferably about 2-3 weeks or 4 weeks, or 8 weeks, or 16 weeks, or 20 weeks, or 24 weeks, or 28 weeks, or 32 weeks. In one embodiment, the boosting composition is formulated for administration about 1 week, or 2 weeks, or 3 weeks, or 4 weeks, or 5 weeks, or 6 weeks, or 7 weeks, or 8 weeks, or 9 weeks, or 16 weeks, or 20 weeks, or 24 weeks, or 28 weeks, or 32 weeks after administration of the priming composition.
[0181] The first and second vaccine compositions can be, but need not be, the same composition. Thus, in one embodiment of the present invention, the step of administering the vaccine comprises administering a first vaccine composition, and then at a later time, administering a second vaccine composition. In one embodiment, the first vaccine composition comprises a nanoparticle comprising an HA-ferritin fusion protein of the present invention. In one embodiment, the first vaccine composition comprises a nanoparticle comprising an ectodomain from the hemagglutinin protein of an influenza virus selected from the group consisting of A/New Calcdonia/20/1999 (1999 NC, H1), A/California/04/2009 (2009 CA, H1), A/Singapore/1/1957 (1957 Sing, H2), A/Hong Kong/1/1968 (1968 HK, H3), A/Brisbane/10/2007 (2007 Bris, H3), A/Indonesia/05/2005 (2005 Indo, H5), B/Florida/4/2006 (2006 Flo, B), A/Perth/16/2009 (2009 Per, H3), A/Brisbane/59/2007 (2007 Bris, H1), B/Brisbane/60/2008 (2008 Bris, B). In one embodiment, the hemagglutinin of the first vaccine composition comprises an amino acid sequence at least about 80% identical to an amino acid sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38. In one embodiment, the first vaccine composition comprises an HA-ferritin fusion protein comprising an amino acid sequence at least 80% identical, at least 85% identical, at least 90% identical, at least 95% identical, at least 97% identical or at least 99% identical to a sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, and SEQ ID NO:68, wherein the nanoparticle elicits an immune response against an influenza virus. In one embodiment, the first vaccine composition comprises an HA-ferritin fusion protein comprising an amino acid sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, and SEQ ID NO:68. In one embodiment, the second vaccine composition comprises a nanoparticle comprising an HA SS-ferritin fusion protein of the present invention. In one embodiment, the HA SS-ferritin fusion protein comprises an amino acid sequence at least 80% identical, at least 85% identical, at least 90% identical, at least 95% identical, at least 97% identical or at least 99% identical to a sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95 and SEQ ID NO:98. In one embodiment, the HA SS-ferritin fusion protein comprises an amino acid sequence selected from the group consisting of SEQ ID NO:71, SEQ ID NO:74, SEQ ID NO:77, SEQ ID NO:80, SEQ ID NO:83, SEQ ID NO:86, SEQ ID NO:89, SEQ ID NO:92, SEQ ID NO:95 and SEQ ID NO:98. In one embodiment, the HA 55-ferritin fusion protein comprises an amino acid sequence at least 80% identical, at least 85% identical, at least 90% identical, at least 95% identical, at least 97% identical or at least 99% identical to SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128, wherein the HA 55-ferritin fusion protein elicits an immune response to an influenza virus. In one embodiment, the HA SS-ferritin fusion protein comprises SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128. In one embodiment, the individual is at risk for infection with influenza virus. In one embodiment, the individual has been exposed to influenza virus. As used herein, the terms exposed, exposure, and the like, indicate the subject has come in contact with a person of animal that is known to be infected with an influenza virus. Vaccines of the present invention may be administered using techniques well known to those in the art. Techniques for formulation and administration may be found, for example, in "Remington's Pharmaceutical Sciences", 18th ed., 1990, Mack Publishing Co., Easton, Pa. Vaccines may be administered by means including, but not limited to, traditional syringes, needleless injection devices, or microprojectile bombardment gene guns. Suitable routes of administration include, but are not limited to, parenteral delivery, such as intramuscular, intradermal, subcutaneous, intramedullary injections, as well as, intrathecal, direct intraventricular, intravenous, intraperitoneal, intranasal, or intraocular injections, just to name a few. For injection, the compounds of one embodiment of the invention may be formulated in aqueous solutions, preferably in physiologically compatible buffers such as Hanks' solution, Ringer's solution, or physiological saline buffer.
[0182] In one embodiment, vaccines, or nanoparticles, of the present invention can be used to protect an individual against infection by heterologous influenza virus. That is, a vaccine made using hemagglutinin protein from one strain of influenza virus is capable of protecting an individual against infection by different strains of influenza. For example, a vaccine made using hemagglutinin protein from influenza A/New Calcdonia/20/1999 (1999 NC, H1), can be used to protect an individual against infection by an influenza virus including, but not limited to A/New Calcdonia/20/1999 (1999 NC, H1), A/California/04/2009 (2009 CA, H1), A/Singapore/1/1957 (1957 Sing, H2), A/Hong Kong/1/1968 (1968 HK, H3), A/Brisbane/10/2007 (2007 Bris, H3), A/Indonesia/05/2005 (2005 indo, H5), A/Perth/16/2009 (2009 Per, H3), and/or A/Brisbane/59/2007 (2007 Bris, H1).
[0183] In one embodiment, vaccines, or nanoparticles, of the present invention can be used to protect an individual against infection by an antigenically divergent influenza virus. Antigenically divergent refers to the tendency of a strain of influenza virus to mutate over time, thereby changing the amino acids that are displayed to the immune system. Such mutation over time is also referred to as antigenic drift. Thus, for example, a vaccine made using hemagglutinin protein from a A/New Calcdonia/20/1999 (1999 NC, H1) strain of influenza virus is capable of protecting an individual against infection by earlier, antigenically divergent New Calcdonia strains of influenza, and by evolving (or diverging) influenza strains of the future.
[0184] Because nanoparticles of the present invention display hemagglutinin proteins that are antigenically similar to an intact hemagglutinin, they can be used in assays for detecting antibodies against influenza virus (anti-influenza antibodies).
[0185] Thus, one embodiment of the present invention is a method for detecting anti-influenza virus antibodies using nanoparticles of the present invention. A detection method of the present invention can generally be accomplished by:
[0186] a. contacting at least a portion of a sample being tested for the presence of anti-influenza antibodies with a nanoparticle of the present invention; and,
[0187] b. detecting the presence of a nanoparticle/antibody complex;
[0188] wherein the presence of a nanoparticle/antibody complex indicates that the sample contains anti-influenza antibodies.
[0189] In one embodiment of the present invention, a sample is obtained, or collected, from an individual to be tested for the presence of anti-influenza virus antibodies. The individual may or may not be suspected of having anti-influenza antibodies or of having been exposed to influenza virus. A sample is any specimen obtained from the individual that can be used to test for the presence of anti-influenza virus antibodies. A preferred sample is a body fluid that can be used to detect the presence of anti-influenza virus antibodies. Examples of body fluids that may be used to practice the present method include, but are not limited to, blood, plasma, serum, lacrimal fluid and saliva. Those skilled in the art can readily identify samples appropriate for practicing the disclosed methods.
[0190] Blood, or blood-derived fluids such as plasma, serum, and the like, are particularly suitable as the sample. Such samples can be collected and prepared from individuals using methods known in the art. The sample may be refrigerated or frozen before assay.
[0191] Any nanoparticle of the present invention can be used to practice the disclosed method as long as the nanoparticle binds to anti-influenza virus antibodies. Useful nanoparticles, and methods of their production, have been described in detail herein. In a preferred embodiment, the nanoparticle comprises a fusion protein, wherein the fusion protein comprises at least 25, at least 50, at least 75, at least 100, or at least 150 contiguous amino acids from a monomeric ferritin subunit protein joined to (fused to) at least one epitope from an influenza hemagglutinin protein (i.e., an HA-ferritin fusion protein) such that the nanoparticle comprises trimers of the influenza virus HA protein epitope on its surface, and wherein the fusion protein is capable of self-assembling into nanoparticles. In one embodiment the at least 25, at least 50, at least 75, at least 100, or at least 150 contiguous amino acids are from the region of a monomeric ferritin protein corresponding to the amino acid sequences of the Helicobacter pylori ferritin monomeric subunit that direct self-assembly of the monomeric subunits into the globular form of the ferritin protein. In one embodiment the at least 25, at least 50, at least 75, at least 100, or at least 150 contiguous amino acids are from SEQ ID NO:2, and are capable of directing self-assembly of the monomeric subunits into the globular ferritin protein. In one embodiment the at least 25, at least 50, at least 75, at least 100, or at least 150 contiguous amino acids are from amino acid residues 5-167 of SEQ ID NO:2, or from SEQ ID NO:5, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles.
[0192] In one embodiment the nanoparticle comprises a fusion protein comprising an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, or at least 97% identical to the amino acid sequence of a monomeric ferritin subunit that is responsible for directing self-assembly of the monomeric ferritin subunits into the globular form of the protein, wherein the fusion protein is capable of self-assembling into nanoparticles. In one embodiment, the fusion protein comprises a polypeptide sequence identical in sequence to a monomeric ferritin subunit. In one embodiment the fusion protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, or at least 97% identical to the amino acid sequence of a monomeric ferritin subunit from Helicobacter pylori, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles. In one embodiment the fusion protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, or at least 97% identical to the amino acid sequence of to a sequence selected from the group consisting of (1) amino acid residues 5-167 from SEQ ID NO:2 and (2) SEQ ID NO:5, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles. In one embodiment the fusion protein comprises a sequence selected from the group consisting of (1) amino acid residues 5-167 from SEQ ID NO:2 and (2) SEQ ID NO:5.
[0193] In one embodiment, the nanoparticle comprises a fusion protein comprising a ferritin protein of the present invention joined to at least one immunogenic portion of an HA protein from a virus selected from the group consisting of influenza type A viruses, influenza type B viruses and influenza type C viruses. In one embodiment the fusion protein comprises a ferritin protein of the present invention joined to at least one immunogenic portion of an HA protein selected from the group consisting of an H1 influenza virus HA protein, an H2 influenza virus HA protein, H3 influenza virus HA protein, an H4 influenza virus HA protein, an H5 influenza virus HA protein, an H6 influenza virus HA protein, an H7 virus influenza HA protein, an H8 influenza virus HA protein, an H9 influenza virus HA protein, an H10 influenza virus HA protein, an H11 influenza virus HA protein, an H12 influenza virus HA protein, an H13 influenza virus HA protein, an H14 influenza virus HA protein, an H15 influenza virus HA protein, and an H16 influenza virus HA protein. In one embodiment, the fusion protein comprises a ferritin protein of the present invention joined to at least one immunogenic portion of an HA protein from virus listed in Table 2. In, one embodiment the immunogenic portion comprises at least one epitope.
[0194] In one embodiment, the nanoparticle comprises a fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence that is a variant of an HA protein from a virus selected from the group consisting of influenza Type A viruses influenza type B viruses and influenza type C viruses, wherein the fusion protein is capable of selectively binding anti-influenza antibodies. In one embodiment, the fusion protein comprises a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the sequence of a HA protein from a virus selected from the group consisting of influenza Type A viruses influenza Type B viruses and influenza type C viruses, wherein the fusion protein is capable of selectively binding anti-influenza antibodies. In one embodiment, the fusion protein comprises a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the sequence of a HA protein selected from the group consisting an H1 influenza virus HA protein, an H2 influenza virus HA protein, H3 influenza virus HA protein, an H4 influenza virus HA protein, an H5 influenza virus HA protein, an H6 influenza virus HA protein, an H7 virus influenza HA protein, an H8 influenza virus HA protein, an H9 influenza virus HA protein, an H10 influenza virus HA protein, an H11 influenza virus HA protein, an H12 influenza virus HA protein, an H13 influenza virus HA protein, an H14 influenza virus HA protein, an H15 influenza virus HA protein, and an H16 influenza virus HA protein, wherein the fusion protein is capable of selectively binding anti-influenza antibodies. In one embodiment, the fusion protein comprises a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the sequence of a HA protein from a virus listed in Table 2, wherein the fusion protein is capable of selectively binding anti-influenza antibodies. In one embodiment, the fusion protein comprises a ferritin protein of the present invention joined to amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38, wherein the fusion protein is capable of selectively binding anti-influenza antibodies. In one embodiment, the fusion protein comprises a ferritin protein of the present invention joined to amino acid sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38, wherein the fusion protein is capable of selectively binding anti-influenza antibodies.
[0195] In one embodiment the nanoparticle comprises a fusion protein comprising an amino acid sequence at least 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to a sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, SEQ ID NO:68, SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128, wherein the fusion protein is capable of selectively binding anti-influenza antibodies. In one embodiment the fusion protein comprises an amino acid sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, SEQ ID NO:68, SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128.
[0196] As used herein, the term contacting refers to the introduction of a sample being tested for the presence of anti-influenza antibodies to a nanoparticle of the present invention, for example, by combining or mixing the sample and the nanoparticle of the present invention, such that the nanoparticle is able to come into physical contact with antibodies in the sample, if present. When anti-influenza virus antibodies are present in the sample, an antibody/nanoparticle complex is then formed. Such complex formation refers to the ability of an anti-influenza virus antibodies to selectively bind to the HA portion of the fusion protein in the nanoparticle in order to form a stable complex that can be detected. Binding of anti-influenza virus antibodies in the sample to the nanoparticle is accomplished under conditions suitable to form a complex. Such conditions (e.g., appropriate concentrations, buffers, temperatures, reaction times) as well as methods to optimize such conditions are known to those skilled in the art. Binding can be measured using a variety of methods standard in the art including, but not limited to, agglutination assays, precipitation assays, enzyme immunoassays (e.g., ELISA), immunoprecipitation assays, immunoblot assays and other immunoassays as described, for example, in Sambrook et al., Molecular Cloning: A Laboratory Manual, (Cold Spring Harbor Labs Press, 1989), and Harlow et al., Antibodies, a Laboratory Manual (Cold Spring Harbor Labs Press, 1988), both of which are incorporated by reference herein in their entirety. These references also provide examples of complex formation conditions.
[0197] As used herein, the phrases selectively binds hemagglutinin, selective binding to hemagglutinin, and the like, refer to the ability of an antibody to preferentially bind a HA protein as opposed to binding proteins unrelated to HA, or non-protein components in the sample or assay. An antibody that selectively binds HA is one that binds albumin but does not significantly bind other molecules or components that may be present in the sample or assay. Significant binding, is considered, for example, binding of an anti-HA antibody to a non-HA molecule with an affinity or avidity great enough to interfere with the ability of the assay to detect and/or determine the level of, anti-influenza antibodies in the sample. Examples of other molecules and compounds that may be present in the sample, or the assay, include, but are not limited to, non-HA proteins, such as albumin, lipids and carbohydrates.
[0198] In one embodiment, an anti-influenza virus antibody/nanoparticle complex, also referred to herein as an antibody/nanoparticle complex, can be formed in solution. In one embodiment an antibody/nanoparticle complex can be formed in which the nanoparticle is immobilized on (e.g., coated onto) a substrate. Immobilization techniques are known to those skilled in the art. Suitable substrate materials include, but are not limited to, plastic, glass, gel, celluloid, fabric, paper, and particulate materials. Examples of substrate materials include, but are not limited to, latex, polystyrene, nylon, nitrocellulose, agarose, cotton, PVDF (poly-vinylidene-fluoride), and magnetic resin. Suitable shapes for substrate material include, but are not limited to, a well (e.g., microtiter dish well), a microtiter plate, a dipstick, a strip, a bead, a lateral flow apparatus, a membrane, a filter, a tube, a dish, a celluloid-type matrix, a magnetic particle, and other particulates. Particularly preferred substrates include, for example, an ELISA plate, a dipstick, an immunodot strip, a radioimmunoassay plate, an agarose bead, a plastic bead, a latex bead, a cotton thread, a plastic chip, an immunoblot membrane, an immunoblot paper and a flow-through membrane. In one embodiment, a substrate, such as a particulate, can include a detectable marker. For descriptions of examples of substrate materials, see, for example, Kemeny, D. M. (1991) A Practical Guide to ELISA, Pergamon Press, Elmsford, N.Y. pp 33-44, and Price, C. and Newman, D. eds. Principles and Practice of Immunoassay, 2nd edition (1997) Stockton Press, NY, N.Y., both of which are incorporated herein by reference in their entirety.
[0199] In accordance with the present invention, once formed, an anti-influenza virus antibody/nanoparticle complex is detected. Detection can be qualitative, quantitative, or semi-quantitative. As used herein, the phrases detecting complex formation, detecting the complex, and the like, refer to identifying the presence of anti-influenza virus antibody complexed with the nanoparticle. If complexes are formed, the amount of complexes formed can, but need not be, quantified. Complex formation, or selective binding, between a putative anti-influenza virus antibody and a nanoparticle can be measured (i.e., detected, determined) using a variety of methods standard in the art (see, for example, Sambrook et al. supra.), examples of which are disclosed herein. A complex can be detected in a variety of ways including, but not limited to use of one or more of the following assays: a hemagglutination inhibition assay, a radial diffusion assay, an enzyme-linked immunoassay, a competitive enzyme-linked immunoassay, a radioimmunoassay, a fluorescence immunoassay, a chemiluminescent assay, a lateral flow assay, a flow-through assay, a particulate-based assay (e.g., using particulates such as, but not limited to, magnetic particles or plastic polymers, such as latex or polystyrene beads), an immunoprecipitation assay, a BioCoreJ assay (e.g., using colloidal gold), an immunodot assay (e.g., CMG=s Immunodot System, Fribourg, Switzerland), and an immunoblot assay (e.g., a western blot), an phosphorescence assay, a flow-through assay, a chromatography assay, a PAGe-based assay, a surface plasmon resonance assay, a spectrophotometric assay, and an electronic sensory assay. Such assays are well known to those skilled in the art.
[0200] Assays can be used to give qualitative or quantitative results depending on how they are used. Some assays, such as agglutination, particulate separation, and precipitation assays, can be observed visually (e.g., either by eye or by a machines, such as a densitometer or spectrophotometer) without the need for a detectable marker.
[0201] In other assays, conjugation (i.e., attachment) of a detectable marker to the nanoparticle, or to a reagent that selectively binds to the nanoparticle, aids in detecting complex formation. A detectable marker can be conjugated to the nanoparticle, or nanoparticle-binding reagent, at a site that does not interfere with ability of the nanoparticle to bind to an anti-influenza virus antibody. Methods of conjugation are known to those of skill in the art. Examples of detectable markers include, but are not limited to, a radioactive label, a fluorescent label, a chemiluminescent label, a chromophoric label, an enzyme label, a phosphorescent label, an electronic label; a metal sol label, a colored bead, a physical label, or a ligand. A ligand refers to a molecule that binds selectively to another molecule. Preferred detectable markers include, but are not limited to, fluorescein, a radioisotope, a phosphatase (e.g., alkaline phosphatase), biotin, avidin, a peroxidase (e.g., horseradish peroxidase), beta-galactosidase, and biotin-related compounds or avidin-related compounds (e.g., streptavidin or ImmunoPure7 NeutrAvidin).
[0202] In one embodiment, an antibody/nanoparticle complex can be detected by contacting a sample with a specific compound, such as an antibody, that binds to an anti-influenza antibody, ferritin, or to the antibody/nanoparticle complex, conjugated to a detectable marker. A detectable marker can be conjugated to the specific compound in such a manner as not to block the ability of the compound to bind to the complex being detected. Preferred detectable markers include, but are not limited to, fluorescein, a radioisotope, a phosphatase (e.g., alkaline phosphatase), biotin, avidin, a peroxidase (e.g., horseradish peroxidase), beta-galactosidase, and biotin-related compounds or avidin-related compounds (e.g., streptavidin or ImmunoPure7 NeutrAvidin).
[0203] In another embodiment, a complex is detected by contacting the complex with an indicator molecule. Suitable indicator molecules include molecules that can bind to the anti-influenza virus antibody/nanoparticle complex, the anti-influenza virus antibody, or the nanoparticle. As such, an indicator molecule can comprise, for example, a reagent that binds the anti-influenza virus antibody, such as an antibody that recognizes immunoglobulins. Preferred indicator molecules that are antibodies include, for example, antibodies reactive with the antibodies from species of individual in which the anti-influenza virus antibodies are produced. An indicator molecule itself can be attached to a detectable marker of the present invention. For example, an antibody can be conjugated to biotin, horseradish peroxidase, alkaline phosphatase or fluorescein.
[0204] The present invention can further comprise one or more layers and/or types of secondary molecules or other binding molecules capable of detecting the presence of an indicator molecule. For example, an untagged (i.e., not conjugated to a detectable marker) secondary antibody that selectively binds to an indicator molecule can be bound to a tagged (i.e., conjugated to a detectable marker) tertiary antibody that selectively binds to the secondary antibody. Suitable secondary antibodies, tertiary antibodies and other secondary or tertiary molecules can be readily selected by those skilled in the art. Preferred tertiary molecules can also be selected by those skilled in the art based upon the characteristics of the secondary molecule. The same strategy can be applied for subsequent layers.
[0205] Preferably, the indicator molecule is conjugated to a detectable marker. A developing agent is added, if required, and the substrate is submitted to a detection device for analysis. In some protocols, washing steps are added after one or both complex formation steps in order to remove excess reagents. If such steps are used, they involve conditions known to those skilled in the art such that excess reagents are removed but the complex is retained.
[0206] While the assays described thus far directly detect the antibody/nanoparticle complex, it is also possible to detect the complex using indirect methods. One example of an indirect assay is the hemagglutination inhibition (HAI) assay, which is a standard assay used to identify the levels of influenza virus in the serum of people thought to be infected with the virus. General methods for performing hemagglutinin inhibition assays are taught herein and are also disclosed in, for example, (see, for example, the WHO Reference on Animal Influenza Diagnosis and Surveillance, 2002, Department of Communicable Disease Surveillance and Response, World Health Organization).
[0207] The HAI assay is based on the fact that hemagglutinin protein on influenza virus is able to bind sialic acid molecules on red blood cells. Thus, when influenza virus is mixed with red blood cells, the virus and the red blood cells bind together forming a lattice (see FIG. 45), with the result that the lattice settles to the bottom of the reaction vessel (e.g., the well of an 96-well plate) forming a diffuse, red color over the entire surface. In contrast, if the reaction contains anything that interferes with binding of the HA protein to the sialic acid molecules on the red blood cells (such as anti-HA antibodies), formation of the lattice is prevented causing the red blood cells to fall out of solution and pool in the bottom of the container. In the latter case, the red blood cells form a characteristic red dot or "button" in the bottom of the well (see, for example, FIG. 46). In the HAI assay, a sample to be tested for the presence of anti-influenza antibodies is added to the influenza virus/red blood cell mixture (the assay mix) and the effect on button formation is observed. If the test sample contains anti-influenza antibodies, they will bind to the virus in the assay mixture, preventing, or disrupting, formation of the lattice causing the red blood cells to fall out of solution, pool in the well and form a button (or dot). The titer of the antibody present in the test sample is determined by counting how far the sample can be diluted before its hemagglutination inhibiting activity is lost. For example, if a sample diluted 1:1000 stil produces a dot nd the next dilution (1:2000) results in lattice formation (no dot) then the titer of antibody in the sample is 1:1000. Such methods of determining antibody titers are known to those skilled in the art.
[0208] A problem with the HAI assay is that it requires the use of live influenza virus, which due to the inherent dangers of growing such virus, requires the use of BSL2 and BSL3 facilities. However, such problems can be alleviated by using nanoparticles of the present invention in the HAI assay since they are fully recombinant and are made without the need to produce potentially dangerous live virus in eggs or in cell cultures. Moreover, because the fusion proteins making up the nanoparticles are recombinant, they offer the opportunity to test the effect of mutations in the HA protein that would otherwise inactivate viral replication. Thus, a HAI assay using nanoparticles of the present invention offers improvements and benefits not found in the currently available HAI assay.
[0209] Accordingly, in one embodiment, detection of an anti-influenza virus antibody/nanoparticle complex is conducted using a hemagglutination inhibition assay, wherein the hemagglutination inhibition assay is performed using nanoparticles of the present invention instead of influenza virus. More specifically, one embodiment of the present invention is a method for detecting anti-influenza virus antibodies in a sample, the method comprising:
[0210] a. contacting at least a portion of the sample with a nanoparticle of the present invention and with red blood cells comprising sialic acid molecules, to form a test mixture;
[0211] b. analyzing the test mixture for the presence of pooled red blood cells (i.e., a button), wherein the presence of pooled red blood cells (i.e., a button) in indicative of the presence anti-influenza antibodies in the sample.
[0212] As has been described, the presence of anti-influenza antibodies in the test sample (at least at sufficient levels) causes the red blood cells to fall out of solution and pool, thereby forming a "button" (or dot) at the bottom of the vessel (e.g., ELISA well) housing the test mixture. Thus, formation of a button in the vessel containing the test mixture is indicative of the presence of anti-influenza antibodies in the test sample.
[0213] The level of red blood cell pooling (i.e., button formation) can be determined using any of the techniques known in the art for conducting hemagglutinin inhibition assays. In some embodiments, the level of button formation may be determined by simple visual inspection using the naked eye. In some embodiments, the level of button formation may be determined using a magnifying device such as, for example, a dissecting scope) or a microscope. In some embodiments, the level of button formation may be determined using a device such as, for example, a spectrophotometer or a refractometer. Methods of detecting or measuring button formation are known to those skilled in the art.
[0214] Any nanoparticle of the present invention can be used to practice hemagglutination inhibition assays of the present invention as long as it is capable of binding to sialic acid residues and to anti-influenza virus antibodies. Useful nanoparticles and methods of their production have been disclosed in detail herein. In a preferred embodiment, the nanoparticle comprises a fusion protein, wherein the fusion protein comprises at least 25, at least 50, at least 75, at least 100, or at least 150 contiguous amino acids from a monomeric ferritin subunit protein joined to at least one epitope from an influenza hemagglutinin protein (i.e., an HA-ferritin fusion protein) such that the nanoparticle comprises trimers of the influenza virus HA protein epitope on its surface, and wherein the fusion protein is capable of self-assembling into nanoparticles. In one embodiment the at least 25, at least 50, at least 75, at least 100, or at least 150 contiguous amino acids are from the region of a monomeric ferritin protein corresponding to the amino acid sequences of the Helicobacter pylori ferritin monomeric subunit that direct self-assembly of the monomeric subunits into the globular form of the ferritin protein. In one embodiment the at least 25, at least 50, at least 75, at least 100, or at least 150 contiguous amino acids are from SEQ ID NO:2, and are capable of directing self-assembly of the monomeric subunits into the globular ferritin protein. In one embodiment the at least 25, at least 50, at least 75, at least 100, or at least 150 contiguous amino acids are from amino acid residues 5-167 of SEQ ID NO:2, or from SEQ ID NO:5, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles.
[0215] In one embodiment the nanoparticle comprises a fusion protein comprising an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, or at least 97% identical to the amino acid sequence of a monomeric ferritin subunit that is responsible for directing self-assembly of the monomeric ferritin subunits into the globular form of the protein, wherein the fusion protein is capable of self-assembling into nanoparticles. In one embodiment, the fusion protein comprises a polypeptide sequence identical in sequence to a monomeric ferritin subunit. In one embodiment the fusion protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, or at least 97% identical to the amino acid sequence of a monomeric ferritin subunit from Helicobacter pylori, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles. In one embodiment the fusion protein comprises an amino acid sequence at least 80%, at least 85%, at least 90%, at least 95%, or at least 97% identical to the amino acid sequence of to a sequence selected from the group consisting of (1) amino acid residues 5-167 from SEQ ID NO:2 and (2) SEQ ID NO:5, wherein the HA-ferritin fusion protein is capable of self-assembling into nanoparticles. In one embodiment the fusion protein comprises a sequence selected from the group consisting of (1) amino acid residues 5-167 from SEQ ID NO:2 and (2) SEQ ID NO:5.
[0216] In one embodiment, the nanoparticle comprises a fusion protein comprising a ferritin protein of the present invention joined to at least one immunogenic portion of an HA protein from a virus selected from the group consisting of influenza type A viruses, influenza type B viruses and influenza type C viruses. In one embodiment the fusion protein comprises a ferritin protein of the present invention joined to at least one immunogenic portion of an HA protein selected from the group consisting of an H1 influenza virus HA protein, an H2 influenza virus HA protein, H3 influenza virus HA protein, an H4 influenza virus HA protein, an H5 influenza virus HA protein, an H6 influenza virus HA protein, an H7 virus influenza HA protein, an H8 influenza virus HA protein, an H9 influenza virus HA protein, an H10 influenza virus HA protein, an H11 influenza virus HA protein, an H12 influenza virus HA protein, an H13 influenza virus HA protein, an H14 influenza virus HA protein, an H15 influenza virus HA protein, and an H16 influenza virus HA protein. In one embodiment, the fusion protein comprises a ferritin protein of the present invention joined to at least one immunogenic portion of an HA protein from an influenza virus of the Victoria of Yamagata lineage. In one embodiment, the fusion protein comprises a ferritin protein of the present invention joined to at least one immunogenic portion of an HA protein from virus listed in Table 2. In, one embodiment the immunogenic portion comprises at least one epitope.
[0217] In one embodiment, the nanoparticle comprises a fusion protein comprising a ferritin protein of the present invention joined to an amino acid sequence that is a variant of an HA protein from a virus selected from the group consisting of influenza Type A viruses influenza Type B viruses and influenza type C viruses, wherein the fusion protein is capable of selectively binding anti-influenza antibodies. In one embodiment, the fusion protein comprises a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the sequence of a HA protein from a virus selected from the group consisting of influenza Type A viruses influenza Type B viruses and influenza type C viruses, wherein the fusion protein is capable of selectively binding anti-influenza antibodies. In one embodiment, the fusion protein comprises a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the sequence of a HA protein selected from the group consisting an H1 influenza virus HA protein, an H2 influenza virus HA protein, H3 influenza virus HA protein, an H4 influenza virus HA protein, an H5 influenza virus HA protein, an H6 influenza virus HA protein, an H7 virus influenza HA protein, an H8 influenza virus HA protein, an H9 influenza virus HA protein, an H10 influenza virus HA protein, an H11 influenza virus HA protein, an H12 influenza virus HA protein, an H13 influenza virus HA protein, an H14 influenza virus HA protein, an H15 influenza virus HA protein, and an H16 influenza virus HA protein, wherein the fusion protein is capable of selectively binding anti-influenza antibodies. In one embodiment, the fusion protein comprises a ferritin protein of the present invention joined to an amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to the sequence of a HA protein from a virus listed in Table 2, wherein the fusion protein is capable of selectively binding anti-influenza antibodies. In one embodiment, the fusion protein comprises a ferritin protein of the present invention joined to amino acid sequence at least about 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to a sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38, wherein the fusion protein is capable of selectively binding anti-influenza antibodies. In one embodiment, the fusion protein comprises a ferritin protein of the present invention joined to amino acid sequence selected from the group consisting of SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:23, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, and SEQ ID NO:38, wherein the fusion protein is capable of selectively binding anti-influenza antibodies.
[0218] In one embodiment the nanoparticle comprises a fusion protein comprising an amino acid sequence at least 80%, at least about 85%, at least about 90%, at least about 95%, at least about 97% or at least about 99% identical to a sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, SEQ ID NO:68, SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128, wherein the fusion protein is capable of selectively binding anti-influenza antibodies. In one embodiment the fusion protein comprises an amino acid sequence selected from the group consisting of SEQ ID NO:41, SEQ ID NO:44, SEQ ID NO:47, SEQ ID NO:50, SEQ ID NO:53, SEQ ID NO:56, SEQ ID NO:59, SEQ ID NO:62, SEQ ID NO:65, SEQ ID NO:68, SEQ ID NO:101, SEQ ID NO:104 SEQ ID NO:107 SEQ ID NO:110 SEQ ID NO:113 SEQ ID NO:116 SEQ ID NO:119 SEQ ID NO:122 SEQ ID NO:125 and SEQ ID NO:128.
[0219] Any red blood cell can be used to perform a HAI assay of the present invention as long as the red blood cells comprise sialic acid residues that are able to bind to influenza hemagglutinin protein. Examples of suitable red bloods cells include chicken red bloods cells, and turkey red bloods cells? Those skilled in the art are capable of identifying red blood cells useful for practicing the disclosed invention. Likewise, conditions suitable for formation of a complex between a nanoparticle and an anti-influenza antibody, if present, are known to those skilled in the art.
[0220] Because assays of the present invention can detect anti-influenza virus antibodies in a sample, including a blood sample, such assays can be used to identify individuals having anti-influenza antibodies. Thus, one embodiment of the present invention is a method to identify an individual having anti-influenza virus antibodies, the method comprising:
[0221] a. contacting a sample from an individual being tested for anti-influenza antibodies with a nanoparticle of the present invention; and,
[0222] b. analyzing the contacted sample for the presence of a nanoparticle/antibody complex
[0223] wherein the presence of a nanoparticle/antibody complex indicates the individual has anti-influenza antibodies.
[0224] Any of the disclosed assay formats can be used to conduct the disclosed method. Examples of useful assay formats include, but are not limited to, a hemagglutination inhibition assay, a radial diffusion assay, an enzyme-linked immunoassay, a competitive enzyme-linked immunoassay, a radioimmunoassay, a fluorescence immunoassay, a chemiluminescent assay, a lateral flow assay, a flow-through assay, a particulate-based assay (e.g., using particulates such as, but not limited to, magnetic particles or plastic polymers, such as latex or polystyrene beads), an immunoprecipitation assay, a BioCoreJ assay (e.g., using colloidal gold), an immunodot assay (e.g., CMG=s Immunodot System, Fribourg, Switzerland), and an immunoblot assay (e.g., a western blot), an phosphorescence assay, a flow-through assay, a chromatography assay, a PAGe-based assay, a surface plasmon resonance assay, a spectrophotometric assay, and an electronic sensory assay.
[0225] If no anti-influenza antibodies are detected in the sample, such a result indicates the individual does not have anti-influenza virus antibodies. The individual being tested may or may not be suspected of having antibodies to influenza virus. The disclosed methods may also be used to determine if an individual has been exposed to one or more specific type, group, sub-group or strain of influenza virus. To make such a determination, a sample is obtained from an individual that has tested negative for antibodies (i.e., lacked antibodies) to one or more specific type, group, sub-group or strain of influenza virus sometime in their past (e.g., greater than about 1 year, greater than about 2 years, greater than about 3 years, greater than about 4 years, greater than about 5 years, etc.). The sample is then tested for the presence of anti-influenza virus antibodies to one or more type, group, sub-group or strain, of influenza virus using a nanoparticle-based assay of the present invention. If the assay indicates the presence of such antibodies, the individual is then identified as having been exposed to one or more type, group sub-group or strain, of influenza virus sometime after the test identifying them as negative for anti-influenza antibodies. Thus, one embodiment of the present invention is method to identify an individual that has been exposed to influenza virus, the method comprising:
[0226] a. contacting at least a portion of a sample from an individual being tested for anti-influenza antibodies with a nanoparticle of the present invention; and,
[0227] b. analyzing the contacted sample for the presence or level of a antibody/nanoparticle complex, wherein the presence or level of antibody/nanoparticle complex indicates the presence or level of recent anti-influenza antibodies;
[0228] c. comparing the recent anti-influenza antibody level with a past anti-influenza antibody level;
[0229] wherein an increase in the recent anti-influenza antibody level over the past anti-influenza antibody level indicates the individual has been exposed to influenza virus subsequent to determination of the past anti-influenza antibody level.
[0230] Methods of the present invention are also useful for determining the response of an individual to a vaccine. Thus, one embodiment is a method for measuring the response of an individual to an influenza vaccine, the method comprising:
[0231] a. administering to the individual a vaccine for influenza virus;
[0232] b. contacting at least a portion of a sample from the individual with a nanoparticle of the present invention;
[0233] c. analyzing the contacted sample for the presence or level of a antibody/nanoparticle complex, wherein the presence or level of antibody/nanoparticle complex indicates the presence or level of recent anti-influenza antibodies
[0234] wherein an increase in the level of antibody in the sample over the pre-vaccination level of antibody in the individual indicates the vaccine induced an immune response in the individual.
[0235] The influenza vaccine administered to the individual may, but need not, comprise a vaccine of the present invention, as long as the nanoparticle comprises an HA protein that can bind an anti-influenza antibody induced by the administered vaccine. Methods of administering influenza vaccines are known to those of skill in the art.
[0236] Analysis of the sample obtained from the individual may be performed using any of the disclosed assay formats. In one embodiment, analysis of the sample is performed using an assay format selected from the group consisting of, a hemagglutination inhibition assay, a radial diffusion assay, an enzyme-linked immunoassay, a competitive enzyme-linked immunoassay, a radioimmunoassay, a fluorescence immunoassay, a chemiluminescent assay, a lateral flow assay, a flow-through assay, a particulate-based assay (e.g., using particulates such as, but not limited to, magnetic particles or plastic polymers, such as latex or polystyrene beads), an immunoprecipitation assay, a BioCoreJ assay (e.g., using colloidal gold), an immunodot assay (e.g., CMG=s Immunodot System, Fribourg, Switzerland), and an immunoblot assay (e.g., a western blot), an phosphorescence assay, a flow-through assay, a chromatography assay, a PAGe-based assay, a surface plasmon resonance assay, a spectrophotometric assay, and an electronic sensory assay.
[0237] In one embodiment, the method includes a step of determining the level of anti-influenza antibody present in the individual prior to administering the vaccine. However, it is also possible to determine the level of anti-influenza antibody present in the individual from prior medical records, if such information is available.
[0238] While not necessary to perform the disclosed method, it may be preferable to wait some period of time between the step of administering the vaccine and the step of determining the level of anti-influenza antibody in the individual. In one embodiment, determination of the level of anti-influenza antibodies present in the individual is performed at least 1 day, at least 2 days, at least 3 days, at least 4 days, at least 5 days, at least 6 days, at least one week, at least two weeks, at least three weeks, at least four weeks, at least two months, at least three months or at least six months, following administration of the vaccine.
[0239] The present invention also includes kits suitable for detecting anti-influenza antibodies. Suitable means of detection include the techniques disclosed herein, utilizing nanoparticles of the present invention. Kits may also comprise a detectable marker, such as an antibody that selectively binds to the nanoparticle, or other indicator molecules. The kit can also contain associated components, such as, but not limited to, buffers, labels, containers, inserts, tubings, vials, syringes and the like.
EXAMPLES
[0240] The following examples are put forth so as to provide those of ordinary skill in the art with a complete disclosure and description of how to make and use the embodiments, and are not intended to limit the scope of what the inventors regard as their invention nor are they intended to represent that the experiments below are all or the only experiments performed. Efforts have been made to ensure accuracy with respect to numbers used (e.g. amounts, temperature, etc.) but some experimental errors and deviations should be accounted for. Unless indicated otherwise, parts are parts by weight, molecular weight is weight average molecular weight, and temperature is in degrees Celsius. Standard abbreviations are used.
Example 1
Design and Production of Ferritin-Based Nanoparticles Expressing Influenza Virus HA
[0241] This Example demonstrates the ability of HA-ferritin fusion proteins to form nanoparticles. Analysis of ferritin structure suggested that it was possible to insert a heterologous protein, specifically influenza virus HA, so that it mimics a physiologically relevant trimeric viral spike (FIG. 1A). Ferritin forms a nearly spherical particle consisting of 24 subunits arranged with octahedral symmetry around a hollow interior. The symmetry of the ferritin nanoparticles includes eight three-fold axes on the surface. The aspartic acid (Asp) at residue 5 near the NH2 terminus is readily accessible, and the distance (28 Å) between each Asp5 on the three-fold axis is almost identical to the distance between the central axes of each HA2 subunit of trimeric HA (FIG. 1A, right).
Vector Construction.
[0242] The HA-ferritin fusion proteins were constructed by joining the ectodomain of A/New Calcdonia/20/1999 (1999 NC) HA to ferritin (FIG. 1B). Specifically, the gene encoding H. pylori nonheme iron-containing ferritin (GenBank NP--223316) with a point mutation (N19Q) to abolish a potential N-linked glycosylation site was synthesized by PCR-based accurate synthesis (M. F. Bachmann, R. M. Zinkernagel, Neutralizing antiviral B cell responses. Annu Rev Immunol 15, 235-270 (1997)) using human-preferred codons. The human CD5 leader sequence and a serine-glycine-glycine (SGG) spacer were joined to the gene fragment encoding ferritin (residues 5-167) to generate a secreted protein. The plasmids encoding various influenza virus HAs, including A/South Carolina/1/1918 (1918 SC), GenBank AF117241; A/Puerto Rico/8/1934 (1934 PR8), UniProt P03452; A/Singapore/6/1986 (1986 Sing), GenBank AB038395; A/Beijing/262/1995 (1995 Beijing), GenBank AAP34323; A/New Calcdonia/20/1999 (1999 NC), GenBank AY289929; A/Solomon Islands/3/2006 (2006 SI), GenBank ABU99109; A/Brisbane/59/2007 (2007 Bris), GenBank ACA28844; A/California/04/2009 (2009 CA), GenBank ACP41105; A/Perth/16/2009 (H3 2009 Perth), GenBank ACS71642; B/Florida/04/2006 (B 2006 Florida), GenBank ACA33493 and their corresponding NAs with human preferred codons were synthesized as previously reported (C. J. Wei et al., Induction of broadly neutralizing H1N1 influenza antibodies by vaccination. Science 329, 1060-1064 (2010)). The gene fragments encoding HAs (residues HA1 1-HA2 174, H3 numbering) from 1999 NC HA, 2009 CA HA, 2009 Perth H3 and 2006 Florida B were amplified and joined to the ferritin gene fragment (residues 5-167) with an SGG linker to give rise to the HA-ferritin fusion gene. To produce soluble trimeric HA, the 1999 NC HA gene fragment (residues HA1 1-HA2 174, H3 numbering) was joined to a thrombin cleavage site followed by a foldon trimerization motif and a poly-histidine tag as described previously (A. S. Xiong et al., PCR-based accurate synthesis of long DNA sequences. Nat Protoc 1, 791-797 (2006)). Both full length and soluble forms of 1999 NC ΔStem (C. J. Wei et al., Induction of broadly neutralizing H1N1 influenza antibodies by vaccination. Science 329, 1060-1064 (2010)) and ΔRBS HA mutants were generated by introducing an N-linked glycosylation site at residues HA2 45 (145N/G47T) and HA1 190 (Q192T), respectively. The soluble form of 2007 Bris ΔRBS HA mutant was generated by introducing an N-linked glycosylation site at the same site. All genes were then cloned into mammalian expression vectors for efficient expression (C. J. Wei et al., Comparative efficacy of neutralizing antibodies elicited by recombinant hemagglutinin proteins from avian H5N1 influenza virus. J Virol 82, 6200-6208 (2008)). Plasmids encoding the mAbs, CR6261 (D. C. Ekiert et al., Antibody recognition of a highly conserved influenza virus epitope. Science 324, 246-251 (2009)), CH65 (J. R. Whittle et al., Broadly neutralizing human antibody that recognizes the receptor-binding pocket of influenza virus hemagglutinin. Proc Natl Acad Sci USA 108, 14216-14221 (2011)) and a single-chain variable fragment F10 (J. Sui et al., Structural and functional bases for broad-spectrum neutralization of avian and human influenza A viruses. Nat Struct Mol Biol 16, 265-273 (2009)) were also synthesized as described by C. J. Wei et al., (Science 329, 1060-1064 (2010).
Protein Biosyntheses and Purifications.
[0243] To produce ferritin nanoparticles, HA-np and trimeric HA, the expression vectors were transfected into 293F cells (Invitrogen), a human embryonic kidney cell line using 293fectin (Invitrogen) according to the manufacturer's instructions. Matched NAs were co-transfected at 20:1 HA:NA (wt:wt). The cells were grown in Freestyle 293 expression medium (Invitrogen) and the culture supernatants were collected 4 days post-transfection by centrifugation and filtered through a 0.22 μm pore filter unit (Nalgene) to remove cell debris. The supernatants were concentrated with a 30 kDa molecular weight cut-off filter unit (Pall Corp.) and then buffer exchanged to a Tris buffer (20 mM Tris, 50 mM NaCl, pH 7.5 for ferritin nanoparticles; 20 mM Tris, 500 mM NaCl, pH 7.5 for HA-np). The ferritin nanoparticles were purified by ion-exchange chromatography using a HiLoad 16/10 Q Sepharose HP column (GE Healthcare). The HA-np were purified by affinity column chromatography using Erythrina cristagalli agglutinin (ECA, coral tree lectin; EY Laboratories, Inc.) specific for galactose β(1,4) N-acetylglucosamine. The ferritin nanoparticles and HA-np were further purified by size exclusion chromatography with a Superose 6 PG XK 16/70 column (GE Healthcare) in PBS. The peak fraction was collected and used for further studies. The molecular weights of the ferritin nanoparticle and HA-np were calculated based on two equations generated by least squares linear regression on a semi-log plot using gel filtration low and high molecular weight standards (Bio-Rad), respectively. The yield of 1999 NC HA-np is ˜4 mg liter-1 and appears stable at 4° C. or frozen at -80° C. The trimeric HA proteins were purified as described by A. S. Xiong et at (Nat Protoc 1, 791-797 (2006)) with slight modifications. Briefly, HA proteins were first purified by affinity chromatography using Ni Sepharose HP resin (GE Healthcare), and then were separated by size exclusion chromatography with a HiLoad 16/60 Superdex 200 PG column (GE Healthcare). To remove the foldon trimerization motif and poly-histidine tag, HA proteins were digested with thrombin (EMD Chemicals, Inc.) (3 U mg ml-1) overnight at 4° C. Undigested proteins were removed by passing over Ni Sepharose HP resin and the digested HAs were purified on a HiLoad 16/60 Superdex 200 PG column. All purified proteins were verified by SDS-PAGE. Protein purity and size distribution were examined by dynamic light scattering using a DynaPro system (Wyatt Technology). All human mAbs and a single-chain variable fragment were also produced in 293F cells and purified as described previously (C. J. Wei et al., Induction of broadly neutralizing H1N1 influenza antibodies by vaccination. Science 329, 1060-1064 (2010); W. P. Kong et al., Protective immunity to lethal challenge of the 1918 pandemic influenza virus by vaccination. Proc Natl Acad Sci USA 103, 15987-15991 (2006)). MAbs against 1999 NC HA were purified from hybridoma supernatants as previously described (C. J. Wei et al., Induction of broadly neutralizing H1N1 influenza antibodies by vaccination. Science 329, 1060-1064 (2010)).
Iodixanol-Based Gradient Centrifugation.
[0244] Alternatively, HA-np were purified by iodixanol gradient ultracentrifugation (FIG. 10) routinely used for virus and VLP purifications (C. J. Wei et al., Cross-neutralization of 1918 and 2009 influenza viruses: role of glycans in viral evolution and vaccine design. Sci Transl Med 2, 24ra21 (2010)). Fractions containing HA np were confirmed by SDS-PAGE and Western blotting using a mAb against 1999 NC HA.
Electron Microscopic Analysis.
[0245] Purified ferritin nanoparticles and HA-np were subjected to transmission electron microscopic analysis. The samples were negatively stained with phosphotungstic acid (ferritin nanoparticles) or ammonium molybdate (HA-np) and images were recorded on a Tecnai T12 microscope (FEI) at 80 kV with a CCD camera (AMT Corp.).
Analysis of HA-Ferritin np.
[0246] Among the various ferritins, Helicobacter (H.) pylori nonheme ferritin (K. J. Cho et al., The crystal structure of ferritin from Helicobacter pylori reveals unusual conformational changes for iron uptake. J Mol Biol 390, 83-98 (2009)) was selected as a prototype because of its highly divergent sequence compared to mammalian ferritins (FIG. 2), thus minimizing the likelihood of inducing autoimmunity after vaccination. The final purification step for recombinant HA-ferritin was size exclusion chromatography (FIG. 1C, left) and dynamic light scattering was used to confirm that both ferritin and HA-ferritin self-assembled into supramolecules with diameters of 14.61 and 37.23 nm, respectively (FIG. 1C, middle). HA-ferritin and ferritin subunits from these nanoparticles migrated at the expected respective molecular weights of 85 and 17 kDa by SDS-PAGE compared to 68 kDa for purified HA (FIG. 1C, right). While the morphology of the ferritin nanoparticles was smooth, as visualized by transmission electron microscopy (TEM), HA-ferritin formed np that exhibited clearly visible spikes around the spherical core (FIG. 1D, Ferritin np vs. HA-np). Remarkably, the placement of these spikes clearly illustrated the octahedral symmetry of the HA-np design. Octahedral two-, three- and four-fold axes were distinctly observed in the TEM image (FIG. 1E, right). These data demonstrated the formation of HA spikes on self-assembling HA-ferritin nanoparticles. More importantly, this design enabled HAs from different subtypes or influenza B viruses to be readily joined to a ferritin core without substantial modification.
Example 2
Antigenicity and Immunogenicity of HA-np in Mice
[0247] To verify the antigenicity of the HA spikes on the np, HA-ferritin np were analyzed for their ability to react with anti-HA head ab and a conformation-dependent monoclonal ab (mAb), CR6261, that recognizes a highly conserved structure in the trimeric HA stem and neutralizes diverse influenza A group 1 viruses D. C. Ekiert et al., Antibody recognition of a highly conserved influenza virus epitope. Science 324, 246-251 (2009)), using ELISA and a virus neutralization assay.
Analysis by ELISA.
[0248] Purified trimeric HA, HA-np, and TIV (2 μg of H1 HA ml-1), ferritin nanoparticles (0.68 μg ml-1 for FIG. 3 or 2 μg ml-1 for the rest), mouse liver ferritin (2 μg ml-1, Alpha Diagnostic International, Inc.), ΔStem and ΔRBS HA trimer (2 μg ml-1) were coated (100 μl/well) onto MaxiSorp® plates (Nunc) and the wells were probed with the anti-HA mAbs, anti-mouse liver ferritin IgG (Alpha Diagnostic International, Inc.) or immune sera followed by peroxidase-conjugated secondary antibodies (anti-mouse IgG and anti-human IgG, SouthernBiotech; anti-ferret IgG, Rockland Immunochemicals, Inc.). The wells were developed using a SureBlue chromogen (KPL) and the reaction was stopped by adding 0.5 M sulfuric acid. For the ELISA-based competition assay, HA trimer (2 μg ml-1) was coated onto the plates. Plates were incubated with an anti-stem mAb, CR6261 (8 μg ml-1) or an isotype control Ab, VRC01 (8 μg ml-1) (Z. Y. Yang et al., Immunization by avian H5 influenza hemagglutinin mutants with altered receptor binding specificity. Science 317, 825-828 (2007); X. Wu et al., Rational design of envelope identifies broadly neutralizing human monoclonal antibodies to HIV-1. Science 329, 856-861 (2010)) before adding serially diluted pre-absorbed ferret immune sera. The wells were probed with anti-ferret IgG and developed as described above. Absorbance at 450 nm was measured by SpectraMax M2e (Molecular Devices). The endpoint titers were determined by calculating the intersection of the observed binding curve and the absorbance threshold (four times background).
Neutralization Assays.
[0249] HA/NA-pseudotyped lentiviral vectors encoding luciferase were used. Immune sera used for the assay were pretreated with RDE as described above. Pre-titrated pseudotyped viruses (Gag p24≈6.25 ng ml-1) were incubated with serially diluted sera for 20 minutes at room temperature and added to 293A cells (10,000 cells/well in a 96-well plate; 50 μl/well; in triplicate). Plates were then washed and replaced with fresh media 2 hours later, and luciferase activity was measured after 24 hours. For the protein competition assay, neutralizing activity of the mAbs F10, CR6261 or immune sera was measured in the presence of competitor proteins, trimeric HA (WT, ΔStem or ΔRBS), HA-np, ferritin nanoparticles or irrelevant protein (HIV-1 gp120) at final concentration of 20 and 25 μg ml-1 for mAbs and immune sera, respectively. The HA-np was able to bind to anti-head or anti-stem mAbs with affinities similar to trimeric HA or trivalent inactivated vaccine (TIV) containing the same 1999 NC HA at equimolar concentrations of HA, in contrast to a ferritin nanoparticle control (FIG. 3A). Analogous to trimeric HA, the HA-np also blocked neutralization by CR6261 and another stem-directed mAb, F10 (4 J. Sui et al., Structural and functional bases for broad-spectrum neutralization of avian and human influenza A viruses. Nat Struct Mol Biol 16, 265-273 (2009)) (FIG. 3B). These results indicated that HA molecules on the HA-np antigenically resembled the physiological trimeric viral spike.
Example 3
Immunogenicity of HA-Ferritin np In Vivo
[0250] This Example demonstrates the ability of HA-ferritin np of the present invention to elicit neutralizing antibodies.
[0251] To assess the immunogenicity of the HA-ferritin np in vivo, mice were immunized twice with HA-np or TIV's from the 2006-2007 season, with HAs from A/New Calcdonia/20/1999 (H1N1), A/Wisconsin/67/2005 (H3N2) and B/Malaysia/2504/04 (type B), or from the 2011-2012 season, with HAs from A/California/07/09-like (H1N1), A/Perth/16/09 (H3N2) and B/Brisbane/60/08 (type B). Briefly, female BALB/c mice (6-8 weeks old; Charles River Laboratories) were immunized (5 mice/group) intramuscularly with 5 or 0.5 μg (1.67 or 0.17 μg of H1 HA) of TIV, 2.24 or 0.22 μg (1.67 or 0.17 μg of HA) of HA-np or 0.57 μg of ferritin nanoparticles (equimolar to 2.24 μg of HA-np) in 100 μl of PBS or in 100 μl of 50% (v/v) mixture of Ribi adjuvant (Sigma) in PBS at weeks 0 and 3. A group of BALB/c mice (n=4) was immunized with 20 μg of trimeric HA (thrombin cleaved) in 100 μl of 50% (v/v) mixture of Ribi adjuvant in PBS at weeks 0 and 4. For the experiment using trivalent HA-np, mice were immunized (n=5) with 6.72 μg (1.67 μg of each HA component) of trivalent HA-np in 100 μl of 50% (v/v) mixture of Ribi adjuvant in PBS at weeks 0 and 3. Blood samples were collected prior to the first dose, and at 2 weeks after each immunization.
[0252] The resulting antibody titers were determined as described in Example 2. The HA-np induced significantly higher HAI titers than TIV (FIG. 4A, left; p<0.0001), and a similar effect was observed in the neutralization assay and ELISA (FIG. 4A, middle and right; p<0.0001). For example, neutralization titers elicited by HA-np as assessed by the concentration of ab needed to inhibit viral entry by 90% (IC90) were ˜34 times higher than TIV (FIG. 4A, middle). Because higher titers were observed in groups with the adjuvant Ribi, further comparisons were performed with this adjuvant. Neutralization against a panel of H1N1 strains revealed not only increased potency but also enhanced breadth stimulated by HA-np compared with TIV or trimeric HA (FIG. 4B). Neutralization against two highly divergent H1N1 viruses, A/Puerto Rico/8/1934 (1934 PR8) and A/Singapore/6/1986 (1986 Sing) were only observed in mice immunized with the HA-np, and the titer against the contemporary virus A/Brisbane/59/2007 (2007 Bris) was more than one log higher in mice immunized with HA-np than with TIV (FIG. 4B).
[0253] To assess whether the preexisting immune responses to ferritin nanoparticles or to other HA subtypes would attenuate the immunogenicity of the subsequent immunization of HA-np, mice were pre-immunized with either H3 (A/Perth/16/09, 2009 Perth) HA-np or empty ferritin nanoparticles to elicit anti-H3 HA and/or anti-H. pylori ferritin immune responses (FIG. 5A). These animals were then immunized with H1 (1999 NC) HA-np. Comparable HAI, IC90 neutralization and ELISA titers against 1999 NC HA were observed in naive animals as well as in groups pre-immunized with H3 HA-np or empty ferritin nanoparticles (FIG. 5b). These results indicated that preexisting anti-H. pylori ferritin immunity did not diminish the HA-specific ab response.
Example 4
Lack of Autoreactivity of H. pylori Ferritin Nanoparticles
[0254] This Example demonstrates analyzes the ability of HA-ferritin np of the present invention to elicit an auto-immune response against autologous ferritin in mice.
[0255] Although the overall structural architecture and physiological functions of ferritin are conserved across organisms, murine ferritin has only 27% amino acid sequence identity to H. pylori ferritin. This homology nonetheless raised the possibility that immunization with H. pylori ferritin in mice might abrogate immune tolerance and induce autoimmunity. To address this concern, CD4, CD8 T-cell and ab responses against both murine and H. pylori ferritins were analyzed by intracellular cytokine staining (ICS) and ELISA in mice immunized with HA-np. ELISAs were performed according to the procedure in Example 2. For intracellular cytokine analysis, CD4.sup.+ and CD8.sup.+ T-cell responses were evaluated for interferon-γ (IFN-γ), tumor necrosis factor α (TNFα), and interleukin-2 (IL-2) as described by T. Zhou et al. (Science 329, 811-817 (2010)). Individual peptide pools (15-mer overlapping by 11 residues, 2.5 μg ml-1 for each peptide) covering H. pylori ferritin or mouse ferritin light and heavy chains were used to stimulate cells. After stimulation, cells were fixed, permeabilized and stained using anti-mouse CD3, CD4, CD8, IFN-γ, TNFα and IL-2 mAbs (BD Pharmingen) together with aqua blue dye for live/dead stain (Invitrogen). The data were collected by LSR II Flow Cytometer (BD Biosciences) and IFN-γ-, TNFα- and IL-2-positive cells in the CD4.sup.+ and CD8.sup.+ cell populations were analyzed with FlowJo software (Tree Star).
[0256] Although an increase in the ICS staining of CD4.sup.+ T cells stimulated with H. pylori ferritin peptides (FIG. 4C, top left) was observed, no increases in the CD4.sup.+ and CD8.sup.+ ICS responses were seen with murine ferritin peptide stimulation (FIG. 4C, bottom left and middle). In addition, while high titers (>106) of anti-H. pylori ferritin abs were detected in ferritin nanoparticle- and HA-np-immune sera, abs to mouse ferritin were undetectable (FIG. 4C, right). These results demonstrate that HA-ferritin np of the present invention do not elicit autoreactivity to autologous ferritin in mice.
Example 5
Generation of Trivalent HA-np and Immunogenicity in Mice
[0257] The Example analyzes whether multivalent HA-np were similar in immunogenicity to monovalent np.
[0258] HA-np expressing HAs from H1 (A/California/04/09, 2009 CA), H3 (2009 Perth) or influenza B (B/Florida/04/06, 2006 FL) were generated. The 2009 CA (H1)-, 2009 Perth (H3)- and 2006 FL (type B)-HA-np self-assembled and displayed the same morphology observed for 1999 NC HA-np (FIG. 6A). Trivalent HA-np were generated by combining three monovalent HA-np, and their immunogenicity was compared to a seasonal TIV containing the same H1 and H3 strains and a mismatched type B (B/Brisbane/60/08). HAI titers against homologous H1N1 and H3N2 viruses were significantly increased in animals immunized with trivalent HA-np relative to TIV-immunized animals (FIG. 6B; p=0.0125 and 0.0036, respectively). When compared to animals immunized with the corresponding monovalent HA-np, HAI titers against 2009 CA (H1) and 2009 Perth (H3) induced by trivalent HA-np were comparable (FIG. 6B). These results demonstrate that no substantial antigenic competition between H1 and H3 HA-np was observed with a trivalent HA-np vaccine.
Example 6
Cross-Protective Immunity Elicited by HA-np in Ferrets
[0259] This Example demonstrates that vaccination of ferrets with 1999 NC HA-np elicits a protective immunity similar to that observed in human disease.
[0260] Male Fitch ferrets (6 months old; Triple F Farms), seronegative for exposure to H1N1, H3N2 and type B influenza viruses, were housed and cared for at BIOQUAL, Inc. (Rockville, Md.). Prior to study start, a temperature transponder (Biomedic Data Systems, Inc.) was implanted into the neck of each ferret. Ferrets were immunized (6 ferrets/group) intramuscularly with 500 μl of PBS, 7.5 μg (2.5 μg of H1 HA) of TIV or 3.35 μg (2.5 μg of HA) of HA-np in 500 μl of 50% (v/v) mixture of Ribi adjuvant in PBS at weeks 0 and 4. Blood was collected 3 and 2 weeks after the first and the second immunization, respectively.
[0261] Three weeks after the first immunization, all ferrets receiving HA-np generated protective HAI titers against homologous H1 1999 NC virus (>1:40), while only 50% (3/6) of TIV-immunized ferrets induced HAI titers greater than 1:40 (FIG. 7A, left; p=0.0056). The same trend was also observed for both neutralization and anti-HA ab titers (FIG. 7A, middle and right; p=0.0047 and p=0.0045, respectively), documenting the superior potency of HA-np in a second species. After boosting, the HAI and IC90 neutralization titers of the HA-np-immune sera were ˜10-fold higher than those of TIV-immunized ferrets (FIG. 7A, left and middle; 457±185 vs. 5760±1541, p=0.0066, and 598±229 vs. 5515±1074, p=0.0012, respectively). A similar enhancement in HA-np vs. TIV immunization was also observed by ELISA titers (FIG. 7A, right; p=0.0038). Remarkably, a single immunization with HA-np induced immune responses comparable to two immunizations with TIV (FIG. 7A).
[0262] To determine whether HA-np could confer protection against an unmatched H1N1 virus, five weeks after the last immunization ferrets immunized with 1999 NC HA-np or TIV containing the same H1 HA were challenged with 106.5 EID50 of 2007 Bris virus. (1999 NC and 2007 Bris viruses are 8 years apart and their antigenic characteristics are sufficiently different to require the production of two different vaccines to confer protection in humans.) The virus was expanded in embryonated chicken eggs from a seed stock obtained from CDC (Atlanta, Ga.) and has a titer of 106.5 EID50 ml-1. The virus stock was inoculated intranasally into ferrets, which had been anesthetized with ketamine/xylazine, in a volume of 500 μl per nostril. The ferrets were observed for clinical signs twice daily and weight and temperature measurements recorded daily by technicians blind to the treatment groups. Nasal washes were obtained on days 1, 3 and 5 and infectious viral titers were determined by TCID50 assay using MDCK cells as described previously (C. J. Wei et al., Induction of broadly neutralizing H1N1 influenza antibodies by vaccination. Science 329, 1060-1064 (2010)).
[0263] Ferrets immunized with HA-np showed a significant reduction in viral shedding beginning 1 day after challenge compared to the sham control group (FIG. 7B, left; p=0.0259). At the same time point, no reduction in viral shedding was seen in the TIV-immunized group. Four of six animals immunized with HA-np had no detectable viral load after 3 days and by day 5, all animals in this group cleared the virus, while all animals in the sham control group still had detectable virus (FIG. 7B). In addition, HA-np-immunized ferrets suffered less body weight loss compared to the TIV-immunized and sham control groups (FIG. 7B, right). These results demonstrate faster virus clearance in ferrets immunized with HA-np than with TIV and further demonstrate that HA-np effectively induced cross-protective immunity in vaccinated ferrets.
Example 7
Induction of Two Types of Neutralizing abs (nAbs) in Ferrets
[0264] This Example demonstrates the breadth and specificity of nAbs in ferret immune sera.
[0265] IC50 neutralization titers against 1986 Sing, A/Beijing/262/1995 (1995 Beijing), A/Solomon Islands/3/2006 (2006 SI) and 2007 Bris were significantly higher in animals immunized with HA-np compared to immunization with TIV (FIG. 8A, left). This enhanced breadth was due not only to a quantitative increase in overall ab titer (˜9-fold against matched virus) but also reflected a qualitative difference in the types of abs elicited (>40-fold enhancement against an unmatched strain). To determine whether the cross-reactivity induced by HA-np was due to nAbs to the conserved HA stem epitope, ferret immune sera were pre-absorbed with cells expressing a stem mutant (ΔStem) HA to remove non-stem directed antibodies. Briefly, ferret immune sera taken 2 weeks after the second immunization were subjected to the assay. The plasmids encoding for ΔStem and ΔRBS HAs were transfected into 293F cells. Three days after transfection, the cells were analyzed by flow cytometry to confirm expression of HA on the cell surface and used for serum absorption. One ml of the immune sera diluted at 1:100 and 1:1,000 was incubated with 100 μl of pre-washed ΔStem and ΔRBS HA-expressing 293F cell pellets, respectively. After incubating for 1 hour at 4° C., supernatants were harvested by centrifugation and binding to WT and mutant HAs was examined by ELISA previously described (C. J. Wei et al., Induction of broadly neutralizing H1N1 influenza antibodies by vaccination. Science 329, 1060-1064 (2010)). The ΔStem HA-pre-absorbed sera were also used for competition ELISA.
[0266] Stem-specific abs were detected in HA-np-immunized ferrets (6/6) in greater frequency and magnitude than TIV-immune ferrets (2/6) (FIG. 8B, left; p=0.0056). Moreover, binding of these pre-absorbed sera to HA was inhibited by CR6261 mAb (FIG. 8B, right; p=0.0019), further documenting the specificity of HA-np immune sera to the stem epitope. The HAI titers against heterologous 2007 Bris virus were also significantly higher in ferrets immunized with HA-np (6/6, 1:80-1:640) than with TIV (3/6, 1:40-1:80) (FIG. 8A, right; p=0.0054). Interestingly, in contrast to a previous study in which DNA prime/TIV boost was used to elicit anti-stem broadly neutralizing abs (bnAbs) (C. J. Wei et al., Induction of broadly neutralizing H1N1 influenza antibodies by vaccination. Science 329, 1060-1064 (2010)), sera from animals immunized with HA-np showed HAI ab titers against a highly divergent 1934 PR8 strain, with titers ≧1:40 in all ferrets. However, no HAI titers against 1934 PR8 were detected in TIV-immunized ferrets (FIG. 8A, right). These data suggested that the HA-np vaccine might elicit another class of nAb directed towards the conserved RBS in the HA head.
[0267] To determine whether HA-np elicited abs against RBS, an RBS mutant HA (ΔRBS) was generated by introducing a glycosylation site in the sialic acid binding pocket at residue 190 (FIG. 9) (D. Lingwood et al., Structural and genetic basis for development of broadly neutralizing influenza antibodies. Nature, in press). Ferret immune sera were absorbed with ΔRBS HA-expressing cells to remove abs to HA outside of this region and tested for binding against WT or ΔRBS HA. RBS-directed abs were detected with titers of >1:2,000 in all HA-np-immunized ferrets, but only 1 out of 6 ferrets that received TIV (FIG. 8B, middle).
[0268] To define the relative contributions of these stem and RBS abs to the breadth of neutralization, neutralization assays were performed in the presence of competitor proteins: WT, ΔStem or ΔRBS HA. In the presence of excess ΔStem HA, only stem-directed abs can neutralize viruses; similarly, ΔRBS HA interferes with all antibodies in the serum except those proximal to the RBS. The relative contribution of stem- and RBS-directed neutralization was measured as activity remaining in the presence of the respective competitor HA. For example, with 2007 Bris, ΔRBS HA only partially inhibited neutralization, while either WT or ΔStem HA almost completely abolished the neutralization activity of the sera; hence, the neutralization against 2007 Bris was due almost entirely to RBS-directed abs (FIG. 8C). Four H1N1 strains were tested in this assay. The pattern of neutralization inhibition varied by strain. Neutralization of 1999 NC or 2007 Bris was mediated predominantly by RBS-directed abs. However, neutralization of 1986 Sing was due mainly to stem-directed abs. Interestingly, the neutralization of 1995 Beijing was more complex. Both stem- and RBS-directed abs contributed to neutralization of this virus (FIG. 8C).
[0269] These results demonstrate that HA-np induce both known types of bnAbs-stem-directed and RBS-directed. Together, these abs contribute to the breadth and potency of the immune sera elicited by HA-np. The synergy between them explains mechanistically the observed superior efficacy of the HA-np vaccine and decreases the likelihood of viral escape mutations from either antibody alone.
[0270] Taken together the above-disclosed Examples demonstrate that a ferritin-based nanoparticle is able to present trimeric HA in its native fold, rigidly and symmetrically, with sufficient spacing to ensure optimal access to potential bnAbs directed to the stem. They also demonstrate that the nanoparticles have enhanced immunogenicity and an expanded neutralization breadth to both stem and RBD antibodies.
Example 8
Immunization of Mice and Ferrets Using a Tetravalent Vaccine
[0271] This Example demonstrates the ability of a multivalent vaccine to elicit an immune response against several strains and sub-types of influenza virus.
[0272] The ability of a pan-group 1 vaccine to stimulate neutralizing antibodies against a variety of influenza viruses was tested in mice and ferrets using a protocol similar to that described in Example 1, and outlined in FIG. 11. Briefly, a pan-group 1 HA-ferritin np vaccine was produced by combining four different monovalent HA-ferritin np vaccines. Specifically, HA-ferritin np, each expressing either H1 A/NC/20/1999, H1 A/CA/04/2009, H2 A/Singapore/1/1957 or H5 A/Indonesia/05/2005, were combined to produce a single vaccine containing all four HA proteins. Mice were immunized twice in a four week interval using 6.8 ug total of the pan-group 1 vaccine (1.7 ug of each HA-ferritin np) in Ribi. Ferrets were immunized twice in a four week interval using 10 ug total of the pan-group 1 vaccine (2.5 ug of each HA-ferritin np) in Ribi. Blood was obtained from the immunized animals and the titer of neutralizing antibodies against various influenza viruses measured. The results of this analysis are shown in FIGS. 12-14. Immunized ferrets were also challenged with either influenza A/Brisbane/59/2007 Brisbane (H1N1) (2207 Bris) (FIG. 15) or influenza A/Mexico/2009 (H1N1) (2009 Mex) (FIG. 16) and the resulting virus titers measured on day 3 and 5 post-challenge.
Example 9
Design and Construction HA-Ferritin Stem-Region Fusion Proteins
[0273] This Example demonstrates the construction of HA-ferritin proteins and nanoparticles that present the stem region of the influenza HA protein.
[0274] As illustrated in FIG. 17, the stem region of the influenza HA protein is highly conserved among different influenza strains, and possesses a site of vulnerability for Group 1 viruses. Thus, a vaccine that elicits neutralizing antibodies against the stem region of the influenza HA protein should be broadly neutralizing. A nanoparticle displaying the stem region of the influenza stem region was constructed as a vaccine.
Design of an HA-Stabilized Stem Fusion Protein
[0275] An HA-stabilized stem fusion protein (HA SS) was constructed as follows: residues 43-313 of the head domain of HA1 were replace with a Gly-Trp-Gly linker. The membrane distal end of HA2 (residues 59 to 93) was replaced by an HIV-1 Bal gp41 HR2 helix followed by a six residue glycine-rich linker (Asn-Gly-Thr-Gly-Gly-Gly-Ser-Gly) and the gp41 HR1 helix. The HR1 helix of gp41 was added in frame with helix C of HA2 so as to generate a long central chimeric helix. The resulting six helix bundle sitting atop the modified hemagglutinin stem provides stability to the SS trimer in lieu of the missing head residues. A schematic of the resulting protein is shown in FIG. 18A, while a ribbon diagram is shown in FIG. 18B. A second trimerization domain consisting of a 28 residue T4 foldon domain was joined to the membrane proximal C-terminus of HA2. The HA 55-ferritin nanoparticle (HA SS-np) protein was generated by joining residue 174 (H3 numbering) of HA SS to H. pylori ferritin (residues 5-167) with a Ser-Gly-Gly linker.
[0276] In constructing HA-SS fusion proteins, genes encoding wild-type HA proteins (A/Puerto Rico/8/1934 (H1 1934 PR8), A/Singapore/6/1986 (H1 1986 Sing), A/New Calcdonia/20/1999 (H1 1999 NC), A/Brisbane/59/2007 (H1 2007 Bris), A/Vietnam/1203/2004 (H5 2004 VN), A/Canada/720/05 (H2 2005 CAN), A/Hong Kong/1/1968 (H3 1968 HK), A/Hong Kong/1073/1999 (H9 1999 HK) and their corresponding NAs, H1 NC 99 SS, RSC3 HIV gp120 control protein, and all Abs (CR6261, F16v3, and VRC01) were synthesized with human preferred codons as previously described (Wei et al. Science 2010, 329(5995):1060-4). Helicobacter pylori nonheme iron-containing ferritin (GenBank NP--223316) with a point mutation (N19Q) to abolish a potential N-linked glycosylation site was synthesized by PCR-based accurate synthesis (Xiong et al. Nat Protoc 2006, 1(2):791-797) using human-preferred codons. Coding sequences for the human CD5 leader sequence and a serine-glycine-glycine (SGG) spacer were joined to the gene fragment encoding ferritin (residues 5-167) to generate a secreted protein. HA and HA SS-- np fusion proteins were generated by overlap PCR by joining the HA ectodomains at residue HA2 174 (H3 numbering) to H. pylori ferritin (residues 5-167) with a Ser-Gly-Gly linker. Stem mutant probes Δstem (glycosylation insertion into the CR6261 binding epitope at position 45 in HA2; H3 numbering) which prevent binding at the conserved H1 stem epitope were generated using site directed mutagenesis. Genes encoding these proteins were cloned into a CMVR plasmid backbone for efficient mammalian cell expression.
Protein Expression and Purification
[0277] Plasmids encoding soluble proteins were transfected (HA ectodomain genes were cotransfected with the corresponding NA encoding plasmids) into the human embryonic kidney cell line 293F and isolated from expression supernatants 72-96 hrs post-transfection. All HA and HA SS trimeric proteins were purified first by metal chelation affinity chromatography and then by size exclusion chromatography as previously described (Wei et al. J Virol. 2008, 82(13):6200-8). IgG Abs were purified using a Protein G affinity column (GE Healthcare). The HA- and HA SS-np were purified by affinity column chromatography using Erythrina cristagalli agglutinin (ECA, coral tree lectin; EY Laboratories, Inc.) specific for galactose β(1,4) N-acetylglucosamine and Galanthus nivalis agglutinin (GNA, snowdrop lectin; EY Laboratories, Inc.) specific for α(1,3) and α(1,6) linked high mannose structures, respectively. HA- and HA SS-np were further purified by size exclusion chromatography with a Superose 6 PG XK 16/70 column (GE Healthcare) in PBS (FIG. 19).
HA SS-Ferritin Characterization.
[0278] HA SS-ferritin np were visualized by electron microscopy. Briefly, purified HA SS-np were negatively stained with phosphotungstic acid and ammonium molybdate, respectively, and images were recorded on a Tecnai T12 microscope (FEI) at 80 kV with a CCD camera (AMT Corp.). The results of this analysis are shown in FIG. 20. IN addition, the ability of purified HA SS and HA SS-np to bind to monoclonal Abs CR6261 and FI6v3 (1.7×10-4 to 10 μg/mL) was characterized by ELISA. HA and HIV gp120 proteins served as controls. Ab binding was detected by peroxidase-conjugated goat anti-human IgG. The results of this analysis, which are shown in FIG. 21, demonstrate that HASS-ferritin is antigenically similar to HA protein.
Example 10
Immune Response to HA SS-Ferritin Nanoparticles
[0279] This Example demonstrates the immune response generated in animals following immunization with HA SS-ferritin np.
[0280] BALB/c mice were immunized twice intramuscularly with protein (2 or 10 μg each) formulated with Ribi adjuvant system (Sigma) at a 3 week interval. Mice received either homologous (HA SS-np prime and boost) or heterologous (HA-np prime and HA SS-np boost) immunizations. Ferrets were immunized three times intramuscularly with HA SS-np (10 μg each) formulated with Ribi adjuvant system (Sigma) at weeks 0, 4 and 14. Serum was collected from animals 2 weeks after each immunization and 1 week prior to the first immunization and heat inactivated (30 min at 56° C.).
[0281] Pre- and post-immune sera from immunized mice and ferrets were assayed for binding to HA and HA SS by ELISA. Briefly, sera were serially diluted (diluted 50 to 2.3×106) and assayed for reactivity to soluble trimeric HA and HA SS proteins, as well as control proteins (200 ng/well with molar equivalents plated according to HA SS). Binding was detected by peroxidase conjugated anti-mouse or anti-ferret IgG, respectively. Endpoint dilutions were determined from nonlinear fit dose-response curves using a detection limit of 2× background absorbance. The result from this analysis are shown in FIG. 22 and demonstrate that stem specific cross-reactive antibodies which recognize the conserved stem-epitope are elicited by HA SS-np vaccination.
[0282] Sera were also analyzed for neutralization of pseudotyped recombinant lentiviruses expressing wild-type HA with the corresponding NA with a luciferase reporter gene as previously described (Wei et al. Science 2010, 329(5995):1060-4) following pretreatment with receptor-destroying enzyme (RDE II; Denka Seiken Co., Ltd.). Psuedotype neutralization competition of ferret serum was performed by incubating serially diluted serum in the presence of either H1 1999 NC SS, H1 1999 NC SS Δstem probe or gp120 control (10 μg/mL) for 1 hr (RT) before addition to pseudotyped recombinant lentiviruses and assaying for neutralization. The results from this analysis are shown in FIG. 23 and demonstrate that vaccination with HA SS-np elicits neutralizing antibodies against various group-1 strains.
Example 11
Immune Response to HA SS-Ferritin Heterologous Immunization Boost
[0283] This example demonstrates that HA SS-np can be utilized to boost antibodies directed to the conserved stem epitope.
[0284] BALB/c mice were immunized twice intramuscularly with heterologous ferritin proteins (HA-np prime and HA SS-np boost; 2 μg each) formulated with Ribi adjuvant system (Sigma) at a 3 week interval. Serum was collected from animals 2 weeks after each immunization and 1 week prior to the first immunization and heat inactivated (30 min at 56° C.).
[0285] Pre- and post-immune sera from immunized mice were assayed for binding to HA and HA SS by ELISA. Briefly, sera were serially diluted (diluted 50 to 2.3×106) and assayed for reactivity to soluble trimeric HA and HA SS proteins, as well as control proteins (200 ng/well with molar equivalents plated according to HA SS). Binding was detected by peroxidase conjugated anti-mouse or anti-ferret IgG, respectively. Endpoint dilutions were determined from nonlinear fit dose-response curves using a detection limit of 2× background absorbance. The results from this analysis are shown in FIG. 22 and demonstrate that cross-reactive stem-epitope specific antibodies are being elicited.
[0286] Sera were also analyzed for neutralization of pseudotyped recombinant lentiviruses expressing wild-type HA with the corresponding NA with a luciferase reporter gene as previously described (Wei et al. Science 2010, 329(5995):1060-4) following pretreatment with receptor-destroying enzyme (RDE II; Denka Seiken Co., Ltd.). The results from this analysis are shown in FIG. 24 and demonstrate that mice which have preexisting stem antibodies titers can be boosted with HA SS-np.
[0287] While the present invention has been described with reference to the specific embodiments thereof, it should be understood by those skilled in the art that various changes may be made and equivalents may be substituted without departing from the true spirit and scope of the invention. In addition, many modifications may be made to adapt a particular situation, material, composition of matter, process, process step or steps, to the objective, spirit and scope of the present invention. All such modifications are intended to be within the scope of the claims.
Example 12
Use of HA-Ferritin Nanoparticles in a Hemagglutination Inhibition Assay
[0288] This Example demonstrates the ability of HA-ferritin nanoparticles of the invention to agglutinate red blood cells as well as the utility of such nanoparticles in performing hemagglutination inhibition assays.
[0289] To compare the ability of HA-ferritin nanoparticles to agglutinate red blood cells, agglutination assays were performed using either inactivated influenza virus of HA-ferritin nanoparticles. Briefly, using 96-well plates, inactivated vaccine and HA-ferritin nanoparticles were each serially diluted in PBS and incubated with 0.5% chicken RBCs (vol/vol using PBS) for 30 minutes at room temperature. Hemagglutination mediated through sialic acid moiety on RBC surface and the sialic acid-binding site on HA was determined by lack of a red dot in the well, which indicates formation of a lattice-structure comprising RBCs and either virus or HA-ferritin nanoparticles. The results of this analysis, which are shown in FIG. 46, demonstrate that the ability of HA-ferritin nanoparticles of the present invention to agglutinate red blood cells is equivalent to that of inactivated influenza virus.
[0290] The importance of the HA protein sialic acid binding domain on the ability of the nanoparticles to agglutinate RBCs was examined by constructing nanoparticles using HA-ferritin fusion proteins in which the sialic acid binding domain was disrupted by a mutation. These nanoparticles were then used in an agglutination assay, as described above. The results of this analysis, which are shown in FIG. 46, demonstrate that the sialic acid binding domain is essential to the ability of the nanoparticles to agglutinate RBCs.
[0291] The ability of HA-ferritin nanoparticles to be used in a hemagglutinin inhibition assay was compared to that of live virus as follows:
HAI Assay Using Live Virus
[0292] Seed stocks of influenza viruses were obtained from the CDC (Atlanta, Ga.) and the viruses expanded in embryonated chicken eggs or in Madin-Darby canine kidney (MDCK) cells. Serum samples were pretreated with receptor-destroying enzyme (RDE II; Denka Seiken Co., Ltd.) overnight at 37° C. followed by heat inactivation at 56° C. for 30 minutes. HAI assays were conducted in V-bottom 96-well plates (Corning, Inc.) using four hemagglutinating units of virus per well and 0.5% turkey or chicken red blood cells (RBCs). Sera pretreated with RDE were serially diluted in PBS (25 μl) and mixed with virus (25 μl) for 30 min at room temperature. RBCs (50 μl) were then added to the reaction and incubated for another 30 min at room temperature. Hemagglutination was determined by visual inspection of the wells for the presence (inhibition; anti-influenza antibodies present) or absence (lack of inhibition; no anti-influenza antibodies present) of a red dot in the bottom of the well.
HAI Assay Using HA-Ferritin Nanoparticles
[0293] HA-ferritin nanoparticles were produced using the methods disclosed herein. Briefly, vectors expressing influenza HA were transfected into 293F cells (Invitrogen) using 293fectin (Invitrogen) according to the manufacturer's instructions. Matched neuraminidase (NA) vectors were co-transfected at a ratio of 20:1 HA:NA (wt:wt) (NA cleaves the sialic acid from the HA to prevent the HA proteins from binding to each other and forming aggregates). The cells were grown in Freestyle 293 expression medium (Invitrogen) and the culture supernatants were collected 4 days post-transfection. The supernatants were concentrated and then buffer exchanged to PBS. The HA ferritin-nanoparticles were purified by affinity column chromatography using Erythrina cristagalli agglutinin (ECA, coral tree lectin; EY Laboratories, Inc.) specific for galactose β(1,4) N-acetylglucosamine. The HA ferritin-nanoparticles were further purified by size exclusion chromatography with a Superose 6 PG XK 16/70 column (GE Healthcare) in PBS. HAI assays were performed as described above, using V-bottom 96-well plates (Corning, Inc.) and four hemagglutinating units of HA ferritin-nanoparticles per well.
[0294] The HA-ferritin nanoparticles were then tested for their ability to work in a HAI assay. Briefly, serum samples were pretreated with receptor-destroying enzyme (RDE II; Denka Seiken Co., Ltd.) overnight at 37° C. followed by heat inactivation at 56° C. for 30 minutes. HAI assays were conducted in V-bottom 96-well plates (Corning, Inc.) using four hemagglutinating units of HA ferritin nanoparticles per well and 0.5% turkey or chicken red blood cells (RBCs). Sera pretreated with RDE were serially diluted in PBS (25 μl) and mixed with HA ferritin nanoparticles (25 μl) for 30 min at room temperature. RBCs (50 μl) were then added to the reaction and incubated for another 30 min at room temperature. Hemagglutination was determined by visual inspection of the wells for the presence (inhibition; anti-influenza antibodies present) or absence (lack of inhibition; no anti-influenza antibodies present) of a red dot in the bottom of the well.
[0295] The results of these studies, which are shown in FIG. 48, demonstrate that the level of anti-influenza antibodies obtained using the HA-ferritin nanoparticle HAI assay is highly correlated with the level obtained using a conventional virus HAI assay.
Sequence CWU
1
1
1291504DNAHelicobacter pylori 1atgttatcaa aagacatcat taagttgcta aacgaacaag
tgaataagga aatgaactct 60tccaacttgt atatgagcat gagttcatgg tgctataccc
atagcttaga tggcgcgggg 120cttttcttgt ttgaccatgc ggctgaagaa tacgagcatg
ctaaaaagct tattatcttc 180ttgaatgaaa acaatgtgcc tgtgcaattg accagcatca
gcgcgcctga gcataagttt 240gaaggtttga ctcaaatttt ccaaaaagcc tatgaacatg
agcaacacat cagcgagtct 300attaacaata tcgtagatca cgccataaaa agcaaagatc
atgcgacttt caatttcttg 360caatggtatg tggctgaaca gcatgaagaa gaagtgcttt
tcaaggatat tttggataaa 420attgagttga ttggtaatga aaaccatggc ttgtatttag
ccgatcagta tgtcaaaggg 480atcgctaaaa gcaggaaatc ttaa
5042167PRTHelicobacter pylori 2Met Leu Ser Lys Asp
Ile Ile Lys Leu Leu Asn Glu Gln Val Asn Lys 1 5
10 15 Glu Met Asn Ser Ser Asn Leu Tyr Met Ser
Met Ser Ser Trp Cys Tyr 20 25
30 Thr His Ser Leu Asp Gly Ala Gly Leu Phe Leu Phe Asp His Ala
Ala 35 40 45 Glu
Glu Tyr Glu His Ala Lys Lys Leu Ile Ile Phe Leu Asn Glu Asn 50
55 60 Asn Val Pro Val Gln Leu
Thr Ser Ile Ser Ala Pro Glu His Lys Phe 65 70
75 80 Glu Gly Leu Thr Gln Ile Phe Gln Lys Ala Tyr
Glu His Glu Gln His 85 90
95 Ile Ser Glu Ser Ile Asn Asn Ile Val Asp His Ala Ile Lys Ser Lys
100 105 110 Asp His
Ala Thr Phe Asn Phe Leu Gln Trp Tyr Val Ala Glu Gln His 115
120 125 Glu Glu Glu Val Leu Phe Lys
Asp Ile Leu Asp Lys Ile Glu Leu Ile 130 135
140 Gly Asn Glu Asn His Gly Leu Tyr Leu Ala Asp Gln
Tyr Val Lys Gly 145 150 155
160 Ile Ala Lys Ser Arg Lys Ser 165
3504DNAHelicobacter pylori 3ttaagatttc ctgcttttag cgatcccttt gacatactga
tcggctaaat acaagccatg 60gttttcatta ccaatcaact caattttatc caaaatatcc
ttgaaaagca cttcttcttc 120atgctgttca gccacatacc attgcaagaa attgaaagtc
gcatgatctt tgctttttat 180ggcgtgatct acgatattgt taatagactc gctgatgtgt
tgctcatgtt cataggcttt 240ttggaaaatt tgagtcaaac cttcaaactt atgctcaggc
gcgctgatgc tggtcaattg 300cacaggcaca ttgttttcat tcaagaagat aataagcttt
ttagcatgct cgtattcttc 360agccgcatgg tcaaacaaga aaagccccgc gccatctaag
ctatgggtat agcaccatga 420actcatgctc atatacaagt tggaagagtt catttcctta
ttcacttgtt cgtttagcaa 480cttaatgatg tcttttgata acat
5044489DNAHelicobacter pylori 4gacatcatca
agctgctgaa cgagcaggtg aacaaggaga tgcagagcag caacctgtac 60atgagcatga
gcagctggtg ctacacccac agcctggacg gcgccggcct gttcctgttc 120gaccacgccg
ccgaggagta cgagcacgcc aagaagctga tcatcttcct gaacgagaac 180aacgtgcccg
tgcagctgac cagcatcagc gcccccgagc acaagttcga gggcctgacc 240cagatcttcc
agaaggccta cgagcacgag cagcacatca gcgagagcat caacaacatc 300gtggaccacg
ccatcaagag caaggaccac gccaccttca acttcctgca gtggtacgtg 360gccgagcagc
acgaggagga ggtgctgttc aaggacatcc tggacaagat cgagctgatc 420ggcaacgaga
accacggcct gtacctggcc gaccagtacg tgaagggcat cgccaagagc 480aggaagagc
4895163PRTHelicobacter pylori 5Asp Ile Ile Lys Leu Leu Asn Glu Gln Val
Asn Lys Glu Met Gln Ser 1 5 10
15 Ser Asn Leu Tyr Met Ser Met Ser Ser Trp Cys Tyr Thr His Ser
Leu 20 25 30 Asp
Gly Ala Gly Leu Phe Leu Phe Asp His Ala Ala Glu Glu Tyr Glu 35
40 45 His Ala Lys Lys Leu Ile
Ile Phe Leu Asn Glu Asn Asn Val Pro Val 50 55
60 Gln Leu Thr Ser Ile Ser Ala Pro Glu His Lys
Phe Glu Gly Leu Thr 65 70 75
80 Gln Ile Phe Gln Lys Ala Tyr Glu His Glu Gln His Ile Ser Glu Ser
85 90 95 Ile Asn
Asn Ile Val Asp His Ala Ile Lys Ser Lys Asp His Ala Thr 100
105 110 Phe Asn Phe Leu Gln Trp Tyr
Val Ala Glu Gln His Glu Glu Glu Val 115 120
125 Leu Phe Lys Asp Ile Leu Asp Lys Ile Glu Leu Ile
Gly Asn Glu Asn 130 135 140
His Gly Leu Tyr Leu Ala Asp Gln Tyr Val Lys Gly Ile Ala Lys Ser 145
150 155 160 Arg Lys Ser
6489DNAHelicobacter pylori 6gctcttcctg ctcttggcga tgcccttcac gtactggtcg
gccaggtaca ggccgtggtt 60ctcgttgccg atcagctcga tcttgtccag gatgtccttg
aacagcacct cctcctcgtg 120ctgctcggcc acgtaccact gcaggaagtt gaaggtggcg
tggtccttgc tcttgatggc 180gtggtccacg atgttgttga tgctctcgct gatgtgctgc
tcgtgctcgt aggccttctg 240gaagatctgg gtcaggccct cgaacttgtg ctcgggggcg
ctgatgctgg tcagctgcac 300gggcacgttg ttctcgttca ggaagatgat cagcttcttg
gcgtgctcgt actcctcggc 360ggcgtggtcg aacaggaaca ggccggcgcc gtccaggctg
tgggtgtagc accagctgct 420catgctcatg tacaggttgc tgctctgcat ctccttgttc
acctgctcgt tcagcagctt 480gatgatgtc
48971695DNAInfluenza virus 7atgaaggcca aactgctggt
gctgctgtgt acctttaccg ccacctacgc cgacacaatc 60tgtatcggct accacgccaa
caatagcacc gacaccgtgg atacagtgct ggagaagaac 120gtgaccgtga cccactctgt
gaacctgctg gaggacagcc acaatggcaa gctgtgtctg 180ctgaaaggca ttgcccctct
gcagctgggc aattgttctg tggccggatg gattctgggc 240aaccccgagt gtgagctgct
gatttctaag gagagctgga gctacatcgt ggagaccccc 300aatcctgaga atggcacctg
ctaccctggc tacttcgccg attacgagga gctgcgcgag 360cagctgtcta gcgtgtccag
cttcgagaga ttcgagatct tccccaagga gtccagctgg 420cctaatcaca cagtgacagg
cgtgtctgcc agctgtagcc acaacggcaa aagcagcttc 480taccggaacc tgctgtggct
gacaggcaag aatggcctgt accccaacct gagcaagagc 540tacgtgaaca acaaggaaaa
ggaagtgctg gtgctgtggg gagtgcacca ccctcccaac 600atcggaaatc agcgggccct
gtaccacaca gagaacgcct atgtgagcgt ggtgtccagc 660cactacagca gaagattcac
ccccgagatc gccaagagac ccaaagtgag agaccaggag 720ggccggatca attactactg
gaccctgctg gagcctggcg ataccatcat cttcgaggcc 780aacggcaatc tgatcgcccc
ttggtatgcc tttgccctga gcagaggctt tggcagcggc 840atcatcacaa gcaacgcccc
catggatgag tgtgatgcca agtgccagac acctcagggc 900gccatcaata gcagcctgcc
cttccagaat gtgcaccctg tgaccatcgg cgagtgcccc 960aagtatgtga gaagcgccaa
gctgagaatg gtgaccggcc tgagaaacat ccctagcatc 1020cagagcagag gactgtttgg
agccatcgcc ggattcatcg agggaggatg gacaggcatg 1080gtggatggct ggtacggcta
ccaccaccag aatgagcagg gctctggata tgccgccgat 1140cagaagtcta cccagaacgc
catcaacggc atcaccaaca aggtgaacag cgtgatcgag 1200aagatgaaca cccagtttac
cgctgtgggc aaggagttca acaagctgga gcggaggatg 1260gagaacctga acaagaaggt
ggacgacggc tttctggaca tctggaccta caatgccgaa 1320ctcctggtcc tcctcgagaa
tgagaggacc ctggacttcc acgacagcaa cgtgaagaac 1380ctgtatgaga aggtgaagag
ccagctgaag aacaacgcca aggagatcgg caacggctgc 1440ttcgagttct accacaagtg
taacaacgag tgtatggaga gcgtgaagaa cggcacctac 1500gactacccta agtacagcga
ggagagcaag ctgaaccggg agaagatcga tggcgtgaag 1560ctggagagca tgggcgtgta
tcagatcctg gccatctaca gcacagtggc ctcttctctg 1620gtgctgctgg tgtctctggg
cgccatctcc ttttggatgt gctccaacgg cagcctgcag 1680tgcaggatct gtatc
16958565PRTInfluenza virus
8Met Lys Ala Lys Leu Leu Val Leu Leu Cys Thr Phe Thr Ala Thr Tyr 1
5 10 15 Ala Asp Thr Ile
Cys Ile Gly Tyr His Ala Asn Asn Ser Thr Asp Thr 20
25 30 Val Asp Thr Val Leu Glu Lys Asn Val
Thr Val Thr His Ser Val Asn 35 40
45 Leu Leu Glu Asp Ser His Asn Gly Lys Leu Cys Leu Leu Lys
Gly Ile 50 55 60
Ala Pro Leu Gln Leu Gly Asn Cys Ser Val Ala Gly Trp Ile Leu Gly 65
70 75 80 Asn Pro Glu Cys Glu
Leu Leu Ile Ser Lys Glu Ser Trp Ser Tyr Ile 85
90 95 Val Glu Thr Pro Asn Pro Glu Asn Gly Thr
Cys Tyr Pro Gly Tyr Phe 100 105
110 Ala Asp Tyr Glu Glu Leu Arg Glu Gln Leu Ser Ser Val Ser Ser
Phe 115 120 125 Glu
Arg Phe Glu Ile Phe Pro Lys Glu Ser Ser Trp Pro Asn His Thr 130
135 140 Val Thr Gly Val Ser Ala
Ser Cys Ser His Asn Gly Lys Ser Ser Phe 145 150
155 160 Tyr Arg Asn Leu Leu Trp Leu Thr Gly Lys Asn
Gly Leu Tyr Pro Asn 165 170
175 Leu Ser Lys Ser Tyr Val Asn Asn Lys Glu Lys Glu Val Leu Val Leu
180 185 190 Trp Gly
Val His His Pro Pro Asn Ile Gly Asn Gln Arg Ala Leu Tyr 195
200 205 His Thr Glu Asn Ala Tyr Val
Ser Val Val Ser Ser His Tyr Ser Arg 210 215
220 Arg Phe Thr Pro Glu Ile Ala Lys Arg Pro Lys Val
Arg Asp Gln Glu 225 230 235
240 Gly Arg Ile Asn Tyr Tyr Trp Thr Leu Leu Glu Pro Gly Asp Thr Ile
245 250 255 Ile Phe Glu
Ala Asn Gly Asn Leu Ile Ala Pro Trp Tyr Ala Phe Ala 260
265 270 Leu Ser Arg Gly Phe Gly Ser Gly
Ile Ile Thr Ser Asn Ala Pro Met 275 280
285 Asp Glu Cys Asp Ala Lys Cys Gln Thr Pro Gln Gly Ala
Ile Asn Ser 290 295 300
Ser Leu Pro Phe Gln Asn Val His Pro Val Thr Ile Gly Glu Cys Pro 305
310 315 320 Lys Tyr Val Arg
Ser Ala Lys Leu Arg Met Val Thr Gly Leu Arg Asn 325
330 335 Ile Pro Ser Ile Gln Ser Arg Gly Leu
Phe Gly Ala Ile Ala Gly Phe 340 345
350 Ile Glu Gly Gly Trp Thr Gly Met Val Asp Gly Trp Tyr Gly
Tyr His 355 360 365
His Gln Asn Glu Gln Gly Ser Gly Tyr Ala Ala Asp Gln Lys Ser Thr 370
375 380 Gln Asn Ala Ile Asn
Gly Ile Thr Asn Lys Val Asn Ser Val Ile Glu 385 390
395 400 Lys Met Asn Thr Gln Phe Thr Ala Val Gly
Lys Glu Phe Asn Lys Leu 405 410
415 Glu Arg Arg Met Glu Asn Leu Asn Lys Lys Val Asp Asp Gly Phe
Leu 420 425 430 Asp
Ile Trp Thr Tyr Asn Ala Glu Leu Leu Val Leu Leu Glu Asn Glu 435
440 445 Arg Thr Leu Asp Phe His
Asp Ser Asn Val Lys Asn Leu Tyr Glu Lys 450 455
460 Val Lys Ser Gln Leu Lys Asn Asn Ala Lys Glu
Ile Gly Asn Gly Cys 465 470 475
480 Phe Glu Phe Tyr His Lys Cys Asn Asn Glu Cys Met Glu Ser Val Lys
485 490 495 Asn Gly
Thr Tyr Asp Tyr Pro Lys Tyr Ser Glu Glu Ser Lys Leu Asn 500
505 510 Arg Glu Lys Ile Asp Gly Val
Lys Leu Glu Ser Met Gly Val Tyr Gln 515 520
525 Ile Leu Ala Ile Tyr Ser Thr Val Ala Ser Ser Leu
Val Leu Leu Val 530 535 540
Ser Leu Gly Ala Ile Ser Phe Trp Met Cys Ser Asn Gly Ser Leu Gln 545
550 555 560 Cys Arg Ile
Cys Ile 565 91695DNAInfluenza virus 9gatacagatc
ctgcactgca ggctgccgtt ggagcacatc caaaaggaga tggcgcccag 60agacaccagc
agcaccagag aagaggccac tgtgctgtag atggccagga tctgatacac 120gcccatgctc
tccagcttca cgccatcgat cttctcccgg ttcagcttgc tctcctcgct 180gtacttaggg
tagtcgtagg tgccgttctt cacgctctcc atacactcgt tgttacactt 240gtggtagaac
tcgaagcagc cgttgccgat ctccttggcg ttgttcttca gctggctctt 300caccttctca
tacaggttct tcacgttgct gtcgtggaag tccagggtcc tctcattctc 360gaggaggacc
aggagttcgg cattgtaggt ccagatgtcc agaaagccgt cgtccacctt 420cttgttcagg
ttctccatcc tccgctccag cttgttgaac tccttgccca cagcggtaaa 480ctgggtgttc
atcttctcga tcacgctgtt caccttgttg gtgatgccgt tgatggcgtt 540ctgggtagac
ttctgatcgg cggcatatcc agagccctgc tcattctggt ggtggtagcc 600gtaccagcca
tccaccatgc ctgtccatcc tccctcgatg aatccggcga tggctccaaa 660cagtcctctg
ctctggatgc tagggatgtt tctcaggccg gtcaccattc tcagcttggc 720gcttctcaca
tacttggggc actcgccgat ggtcacaggg tgcacattct ggaagggcag 780gctgctattg
atggcgccct gaggtgtctg gcacttggca tcacactcat ccatgggggc 840gttgcttgtg
atgatgccgc tgccaaagcc tctgctcagg gcaaaggcat accaaggggc 900gatcagattg
ccgttggcct cgaagatgat ggtatcgcca ggctccagca gggtccagta 960gtaattgatc
cggccctcct ggtctctcac tttgggtctc ttggcgatct cgggggtgaa 1020tcttctgctg
tagtggctgg acaccacgct cacataggcg ttctctgtgt ggtacagggc 1080ccgctgattt
ccgatgttgg gagggtggtg cactccccac agcaccagca cttccttttc 1140cttgttgttc
acgtagctct tgctcaggtt ggggtacagg ccattcttgc ctgtcagcca 1200cagcaggttc
cggtagaagc tgcttttgcc gttgtggcta cagctggcag acacgcctgt 1260cactgtgtga
ttaggccagc tggactcctt ggggaagatc tcgaatctct cgaagctgga 1320cacgctagac
agctgctcgc gcagctcctc gtaatcggcg aagtagccag ggtagcaggt 1380gccattctca
ggattggggg tctccacgat gtagctccag ctctccttag aaatcagcag 1440ctcacactcg
gggttgccca gaatccatcc ggccacagaa caattgccca gctgcagagg 1500ggcaatgcct
ttcagcagac acagcttgcc attgtggctg tcctccagca ggttcacaga 1560gtgggtcacg
gtcacgttct tctccagcac tgtatccacg gtgtcggtgc tattgttggc 1620gtggtagccg
atacagattg tgtcggcgta ggtggcggta aaggtacaca gcagcaccag 1680cagtttggcc
ttcat
1695101551DNAInfluenza virus 10atgaaggcca aactgctggt gctgctgtgt
acctttaccg ccacctacgc cgacacaatc 60tgtatcggct accacgccaa caatagcacc
gacaccgtgg atacagtgct ggagaagaac 120gtgaccgtga cccactctgt gaacctgctg
gaggacagcc acaatggcaa gctgtgtctg 180ctgaaaggca ttgcccctct gcagctgggc
aattgttctg tggccggatg gattctgggc 240aaccccgagt gtgagctgct gatttctaag
gagagctgga gctacatcgt ggagaccccc 300aatcctgaga atggcacctg ctaccctggc
tacttcgccg attacgagga gctgcgcgag 360cagctgtcta gcgtgtccag cttcgagaga
ttcgagatct tccccaagga gtccagctgg 420cctaatcaca cagtgacagg cgtgtctgcc
agctgtagcc acaacggcaa aagcagcttc 480taccggaacc tgctgtggct gacaggcaag
aatggcctgt accccaacct gagcaagagc 540tacgtgaaca acaaggaaaa ggaagtgctg
gtgctgtggg gagtgcacca ccctcccaac 600atcggaaatc agcgggccct gtaccacaca
gagaacgcct atgtgagcgt ggtgtccagc 660cactacagca gaagattcac ccccgagatc
gccaagagac ccaaagtgag agaccaggag 720ggccggatca attactactg gaccctgctg
gagcctggcg ataccatcat cttcgaggcc 780aacggcaatc tgatcgcccc ttggtatgcc
tttgccctga gcagaggctt tggcagcggc 840atcatcacaa gcaacgcccc catggatgag
tgtgatgcca agtgccagac acctcagggc 900gccatcaata gcagcctgcc cttccagaat
gtgcaccctg tgaccatcgg cgagtgcccc 960aagtatgtga gaagcgccaa gctgagaatg
gtgaccggcc tgagaaacat ccctagcatc 1020cagagcagag gactgtttgg agccatcgcc
ggattcatcg agggaggatg gacaggcatg 1080gtggatggct ggtacggcta ccaccaccag
aatgagcagg gctctggata tgccgccgat 1140cagaagtcta cccagaacgc catcaacggc
atcaccaaca aggtgaacag cgtgatcgag 1200aagatgaaca cccagtttac cgctgtgggc
aaggagttca acaagctgga gcggaggatg 1260gagaacctga acaagaaggt ggacgacggc
tttctggaca tctggaccta caatgccgaa 1320ctcctggtcc tcctcgagaa tgagaggacc
ctggacttcc acgacagcaa cgtgaagaac 1380ctgtatgaga aggtgaagag ccagctgaag
aacaacgcca aggagatcgg caacggctgc 1440ttcgagttct accacaagtg taacaacgag
tgtatggaga gcgtgaagaa cggcacctac 1500gactacccta agtacagcga ggagagcaag
ctgaaccggg agaagatcga t 155111517PRTInfluenza virus 11Met Lys
Ala Lys Leu Leu Val Leu Leu Cys Thr Phe Thr Ala Thr Tyr 1 5
10 15 Ala Asp Thr Ile Cys Ile Gly
Tyr His Ala Asn Asn Ser Thr Asp Thr 20 25
30 Val Asp Thr Val Leu Glu Lys Asn Val Thr Val Thr
His Ser Val Asn 35 40 45
Leu Leu Glu Asp Ser His Asn Gly Lys Leu Cys Leu Leu Lys Gly Ile
50 55 60 Ala Pro Leu
Gln Leu Gly Asn Cys Ser Val Ala Gly Trp Ile Leu Gly 65
70 75 80 Asn Pro Glu Cys Glu Leu Leu
Ile Ser Lys Glu Ser Trp Ser Tyr Ile 85
90 95 Val Glu Thr Pro Asn Pro Glu Asn Gly Thr Cys
Tyr Pro Gly Tyr Phe 100 105
110 Ala Asp Tyr Glu Glu Leu Arg Glu Gln Leu Ser Ser Val Ser Ser
Phe 115 120 125 Glu
Arg Phe Glu Ile Phe Pro Lys Glu Ser Ser Trp Pro Asn His Thr 130
135 140 Val Thr Gly Val Ser Ala
Ser Cys Ser His Asn Gly Lys Ser Ser Phe 145 150
155 160 Tyr Arg Asn Leu Leu Trp Leu Thr Gly Lys Asn
Gly Leu Tyr Pro Asn 165 170
175 Leu Ser Lys Ser Tyr Val Asn Asn Lys Glu Lys Glu Val Leu Val Leu
180 185 190 Trp Gly
Val His His Pro Pro Asn Ile Gly Asn Gln Arg Ala Leu Tyr 195
200 205 His Thr Glu Asn Ala Tyr Val
Ser Val Val Ser Ser His Tyr Ser Arg 210 215
220 Arg Phe Thr Pro Glu Ile Ala Lys Arg Pro Lys Val
Arg Asp Gln Glu 225 230 235
240 Gly Arg Ile Asn Tyr Tyr Trp Thr Leu Leu Glu Pro Gly Asp Thr Ile
245 250 255 Ile Phe Glu
Ala Asn Gly Asn Leu Ile Ala Pro Trp Tyr Ala Phe Ala 260
265 270 Leu Ser Arg Gly Phe Gly Ser Gly
Ile Ile Thr Ser Asn Ala Pro Met 275 280
285 Asp Glu Cys Asp Ala Lys Cys Gln Thr Pro Gln Gly Ala
Ile Asn Ser 290 295 300
Ser Leu Pro Phe Gln Asn Val His Pro Val Thr Ile Gly Glu Cys Pro 305
310 315 320 Lys Tyr Val Arg
Ser Ala Lys Leu Arg Met Val Thr Gly Leu Arg Asn 325
330 335 Ile Pro Ser Ile Gln Ser Arg Gly Leu
Phe Gly Ala Ile Ala Gly Phe 340 345
350 Ile Glu Gly Gly Trp Thr Gly Met Val Asp Gly Trp Tyr Gly
Tyr His 355 360 365
His Gln Asn Glu Gln Gly Ser Gly Tyr Ala Ala Asp Gln Lys Ser Thr 370
375 380 Gln Asn Ala Ile Asn
Gly Ile Thr Asn Lys Val Asn Ser Val Ile Glu 385 390
395 400 Lys Met Asn Thr Gln Phe Thr Ala Val Gly
Lys Glu Phe Asn Lys Leu 405 410
415 Glu Arg Arg Met Glu Asn Leu Asn Lys Lys Val Asp Asp Gly Phe
Leu 420 425 430 Asp
Ile Trp Thr Tyr Asn Ala Glu Leu Leu Val Leu Leu Glu Asn Glu 435
440 445 Arg Thr Leu Asp Phe His
Asp Ser Asn Val Lys Asn Leu Tyr Glu Lys 450 455
460 Val Lys Ser Gln Leu Lys Asn Asn Ala Lys Glu
Ile Gly Asn Gly Cys 465 470 475
480 Phe Glu Phe Tyr His Lys Cys Asn Asn Glu Cys Met Glu Ser Val Lys
485 490 495 Asn Gly
Thr Tyr Asp Tyr Pro Lys Tyr Ser Glu Glu Ser Lys Leu Asn 500
505 510 Arg Glu Lys Ile Asp
515 121551DNAInfluenza virus 12atcgatcttc tcccggttca gcttgctctc
ctcgctgtac ttagggtagt cgtaggtgcc 60gttcttcacg ctctccatac actcgttgtt
acacttgtgg tagaactcga agcagccgtt 120gccgatctcc ttggcgttgt tcttcagctg
gctcttcacc ttctcataca ggttcttcac 180gttgctgtcg tggaagtcca gggtcctctc
attctcgagg aggaccagga gttcggcatt 240gtaggtccag atgtccagaa agccgtcgtc
caccttcttg ttcaggttct ccatcctccg 300ctccagcttg ttgaactcct tgcccacagc
ggtaaactgg gtgttcatct tctcgatcac 360gctgttcacc ttgttggtga tgccgttgat
ggcgttctgg gtagacttct gatcggcggc 420atatccagag ccctgctcat tctggtggtg
gtagccgtac cagccatcca ccatgcctgt 480ccatcctccc tcgatgaatc cggcgatggc
tccaaacagt cctctgctct ggatgctagg 540gatgtttctc aggccggtca ccattctcag
cttggcgctt ctcacatact tggggcactc 600gccgatggtc acagggtgca cattctggaa
gggcaggctg ctattgatgg cgccctgagg 660tgtctggcac ttggcatcac actcatccat
gggggcgttg cttgtgatga tgccgctgcc 720aaagcctctg ctcagggcaa aggcatacca
aggggcgatc agattgccgt tggcctcgaa 780gatgatggta tcgccaggct ccagcagggt
ccagtagtaa ttgatccggc cctcctggtc 840tctcactttg ggtctcttgg cgatctcggg
ggtgaatctt ctgctgtagt ggctggacac 900cacgctcaca taggcgttct ctgtgtggta
cagggcccgc tgatttccga tgttgggagg 960gtggtgcact ccccacagca ccagcacttc
cttttccttg ttgttcacgt agctcttgct 1020caggttgggg tacaggccat tcttgcctgt
cagccacagc aggttccggt agaagctgct 1080tttgccgttg tggctacagc tggcagacac
gcctgtcact gtgtgattag gccagctgga 1140ctccttgggg aagatctcga atctctcgaa
gctggacacg ctagacagct gctcgcgcag 1200ctcctcgtaa tcggcgaagt agccagggta
gcaggtgcca ttctcaggat tgggggtctc 1260cacgatgtag ctccagctct ccttagaaat
cagcagctca cactcggggt tgcccagaat 1320ccatccggcc acagaacaat tgcccagctg
cagaggggca atgcctttca gcagacacag 1380cttgccattg tggctgtcct ccagcaggtt
cacagagtgg gtcacggtca cgttcttctc 1440cagcactgta tccacggtgt cggtgctatt
gttggcgtgg tagccgatac agattgtgtc 1500ggcgtaggtg gcggtaaagg tacacagcag
caccagcagt ttggccttca t 1551131554DNAInfluenza virus
13atgaaggcca tcctggtggt gctgctgtac accttcgcca ccgccaacgc cgacaccctg
60tgcatcggct accacgccaa caacagcacc gacaccgtgg acaccgtgct ggagaagaac
120gtgaccgtga cccacagcgt gaacctgctg gaggacaagc acaacggcaa gctgtgcaag
180ctgcggggcg tggcccccct gcacctgggc aagtgcaaca tcgccggctg gattctgggc
240aaccccgagt gcgagagcct gagcaccgcc agcagctgga gctacatcgt ggagaccccc
300agcagcgaca acggcacctg ctaccccggc gacttcatcg actacgagga gctgcgggag
360cagctgagca gcgtgagcag cttcgagcgg ttcgagatct tccccaagac cagcagctgg
420cccaaccacg acagcaacaa gggcgtgacc gccgcctgcc cccacgccgg cgccaagagc
480ttctacaaga acctgatctg gctggtgaag aagggcaaca gctaccccaa gctgagcaag
540agctacatca acgacaaggg caaggaggtg ctggtgctgt ggggcatcca ccaccccagc
600accagcgccg accagcagag cctgtaccag aacgccgaca cctacgtgtt cgtgggcagc
660agccggtaca gcaagaagtt caagcccgag atcgccatcc ggcccaaggt gcgggaccag
720gagggccgga tgaactacta ctggaccctg gtggagcccg gcgacaagat caccttcgag
780gccaccggca acctggtggt gccccggtac gccttcgcca tggagcggaa cgccggcagc
840ggcatcatca tcagcgacac ccccgtgcac gactgcaaca ccacctgcca gacccccaag
900ggcgccatca acaccagcct gcccttccag aacatccacc ccatcaccat cggcaagtgc
960cccaagtacg tgaagagcac caagctgcgg ctggccaccg gcctgcggaa catccccagc
1020atccagagcc ggggcctgtt cggcgccatc gccggcttca tcgagggcgg ctggaccggc
1080atggtggacg gctggtacgg ctaccaccac cagaacgagc agggcagcgg ctacgccgcc
1140gacctgaaga gcacccagaa cgccatcgac gagatcacca acaaggtgaa cagcgtgatc
1200gagaagatga acacccagtt caccgccgtg ggcaaggagt tcaaccacct ggagaagcgg
1260atcgagaacc tgaacaagaa ggtggacgac ggcttcctgg acatctggac ctacaacgcc
1320gagctgctgg tgctgctgga gaacgagcgg accctggact accacgacag caacgtgaag
1380aacctgtacg agaaggtgcg gagccagctg aagaacaacg ccaaggagat cggcaacggc
1440tgcttcgagt tctaccacaa gtgcgacaac acctgcatgg agagcgtgaa gaacggcacc
1500tacgactacc ccaagtacag cgaggaggcc aagctgaacc gggaggagat cgac
155414518PRTInfluenza virus 14Met Lys Ala Ile Leu Val Val Leu Leu Tyr Thr
Phe Ala Thr Ala Asn 1 5 10
15 Ala Asp Thr Leu Cys Ile Gly Tyr His Ala Asn Asn Ser Thr Asp Thr
20 25 30 Val Asp
Thr Val Leu Glu Lys Asn Val Thr Val Thr His Ser Val Asn 35
40 45 Leu Leu Glu Asp Lys His Asn
Gly Lys Leu Cys Lys Leu Arg Gly Val 50 55
60 Ala Pro Leu His Leu Gly Lys Cys Asn Ile Ala Gly
Trp Ile Leu Gly 65 70 75
80 Asn Pro Glu Cys Glu Ser Leu Ser Thr Ala Ser Ser Trp Ser Tyr Ile
85 90 95 Val Glu Thr
Pro Ser Ser Asp Asn Gly Thr Cys Tyr Pro Gly Asp Phe 100
105 110 Ile Asp Tyr Glu Glu Leu Arg Glu
Gln Leu Ser Ser Val Ser Ser Phe 115 120
125 Glu Arg Phe Glu Ile Phe Pro Lys Thr Ser Ser Trp Pro
Asn His Asp 130 135 140
Ser Asn Lys Gly Val Thr Ala Ala Cys Pro His Ala Gly Ala Lys Ser 145
150 155 160 Phe Tyr Lys Asn
Leu Ile Trp Leu Val Lys Lys Gly Asn Ser Tyr Pro 165
170 175 Lys Leu Ser Lys Ser Tyr Ile Asn Asp
Lys Gly Lys Glu Val Leu Val 180 185
190 Leu Trp Gly Ile His His Pro Ser Thr Ser Ala Asp Gln Gln
Ser Leu 195 200 205
Tyr Gln Asn Ala Asp Thr Tyr Val Phe Val Gly Ser Ser Arg Tyr Ser 210
215 220 Lys Lys Phe Lys Pro
Glu Ile Ala Ile Arg Pro Lys Val Arg Asp Gln 225 230
235 240 Glu Gly Arg Met Asn Tyr Tyr Trp Thr Leu
Val Glu Pro Gly Asp Lys 245 250
255 Ile Thr Phe Glu Ala Thr Gly Asn Leu Val Val Pro Arg Tyr Ala
Phe 260 265 270 Ala
Met Glu Arg Asn Ala Gly Ser Gly Ile Ile Ile Ser Asp Thr Pro 275
280 285 Val His Asp Cys Asn Thr
Thr Cys Gln Thr Pro Lys Gly Ala Ile Asn 290 295
300 Thr Ser Leu Pro Phe Gln Asn Ile His Pro Ile
Thr Ile Gly Lys Cys 305 310 315
320 Pro Lys Tyr Val Lys Ser Thr Lys Leu Arg Leu Ala Thr Gly Leu Arg
325 330 335 Asn Ile
Pro Ser Ile Gln Ser Arg Gly Leu Phe Gly Ala Ile Ala Gly 340
345 350 Phe Ile Glu Gly Gly Trp Thr
Gly Met Val Asp Gly Trp Tyr Gly Tyr 355 360
365 His His Gln Asn Glu Gln Gly Ser Gly Tyr Ala Ala
Asp Leu Lys Ser 370 375 380
Thr Gln Asn Ala Ile Asp Glu Ile Thr Asn Lys Val Asn Ser Val Ile 385
390 395 400 Glu Lys Met
Asn Thr Gln Phe Thr Ala Val Gly Lys Glu Phe Asn His 405
410 415 Leu Glu Lys Arg Ile Glu Asn Leu
Asn Lys Lys Val Asp Asp Gly Phe 420 425
430 Leu Asp Ile Trp Thr Tyr Asn Ala Glu Leu Leu Val Leu
Leu Glu Asn 435 440 445
Glu Arg Thr Leu Asp Tyr His Asp Ser Asn Val Lys Asn Leu Tyr Glu 450
455 460 Lys Val Arg Ser
Gln Leu Lys Asn Asn Ala Lys Glu Ile Gly Asn Gly 465 470
475 480 Cys Phe Glu Phe Tyr His Lys Cys Asp
Asn Thr Cys Met Glu Ser Val 485 490
495 Lys Asn Gly Thr Tyr Asp Tyr Pro Lys Tyr Ser Glu Glu Ala
Lys Leu 500 505 510
Asn Arg Glu Glu Ile Asp 515 151554DNAInfluenza virus
15gtcgatctcc tcccggttca gcttggcctc ctcgctgtac ttggggtagt cgtaggtgcc
60gttcttcacg ctctccatgc aggtgttgtc gcacttgtgg tagaactcga agcagccgtt
120gccgatctcc ttggcgttgt tcttcagctg gctccgcacc ttctcgtaca ggttcttcac
180gttgctgtcg tggtagtcca gggtccgctc gttctccagc agcaccagca gctcggcgtt
240gtaggtccag atgtccagga agccgtcgtc caccttcttg ttcaggttct cgatccgctt
300ctccaggtgg ttgaactcct tgcccacggc ggtgaactgg gtgttcatct tctcgatcac
360gctgttcacc ttgttggtga tctcgtcgat ggcgttctgg gtgctcttca ggtcggcggc
420gtagccgctg ccctgctcgt tctggtggtg gtagccgtac cagccgtcca ccatgccggt
480ccagccgccc tcgatgaagc cggcgatggc gccgaacagg ccccggctct ggatgctggg
540gatgttccgc aggccggtgg ccagccgcag cttggtgctc ttcacgtact tggggcactt
600gccgatggtg atggggtgga tgttctggaa gggcaggctg gtgttgatgg cgcccttggg
660ggtctggcag gtggtgttgc agtcgtgcac gggggtgtcg ctgatgatga tgccgctgcc
720ggcgttccgc tccatggcga aggcgtaccg gggcaccacc aggttgccgg tggcctcgaa
780ggtgatcttg tcgccgggct ccaccagggt ccagtagtag ttcatccggc cctcctggtc
840ccgcaccttg ggccggatgg cgatctcggg cttgaacttc ttgctgtacc ggctgctgcc
900cacgaacacg taggtgtcgg cgttctggta caggctctgc tggtcggcgc tggtgctggg
960gtggtggatg ccccacagca ccagcacctc cttgcccttg tcgttgatgt agctcttgct
1020cagcttgggg tagctgttgc ccttcttcac cagccagatc aggttcttgt agaagctctt
1080ggcgccggcg tgggggcagg cggcggtcac gcccttgttg ctgtcgtggt tgggccagct
1140gctggtcttg gggaagatct cgaaccgctc gaagctgctc acgctgctca gctgctcccg
1200cagctcctcg tagtcgatga agtcgccggg gtagcaggtg ccgttgtcgc tgctgggggt
1260ctccacgatg tagctccagc tgctggcggt gctcaggctc tcgcactcgg ggttgcccag
1320aatccagccg gcgatgttgc acttgcccag gtgcaggggg gccacgcccc gcagcttgca
1380cagcttgccg ttgtgcttgt cctccagcag gttcacgctg tgggtcacgg tcacgttctt
1440ctccagcacg gtgtccacgg tgtcggtgct gttgttggcg tggtagccga tgcacagggt
1500gtcggcgttg gcggtggcga aggtgtacag cagcaccacc aggatggcct tcat
1554161542DNAInfluenza virus 16atggccatca tctacctgat cctgctgttt
acagctgtgc ggggcgatca gatctgtatc 60ggctaccacg ccaacaatag caccgagaag
gtggacacca tcctggaaag aaatgtgacc 120gtgacccacg ccaaggatat tctggaaaag
acccacaacg gcaagctgtg caagctgaat 180ggcattcctc ctctggaact gggcgattgt
tctattgctg gctggctgct gggaaatcct 240gagtgcgata gactgctgtc tgtgcctgag
tggagctaca tcatggaaaa agagaaccct 300agggacggac tgtgttaccc cggcagcttc
aacgattacg aggaactgaa gcacctgctg 360tccagcgtga agcacttcga gaaagtgaag
atcctgccca aggatagatg gacccagcat 420acaacaacag gcggaagcag agcttgtgct
gtgtccggca accccagctt cttcagaaat 480atggtctggc tgaccaagaa gggctctaat
tatcctgtgg ccaagggcag ctacaataat 540acaagcggcg agcagatgct gattatttgg
ggcgtgcacc accctaatga tgagacagag 600cagagaaccc tgtaccagaa tgtgggcaca
tacgtgtctg tgggcaccag cacactgaat 660aagagaagca cccccgatat tgccaccaga
cccaaagtga atggacaggg cggcagaatg 720gaattttcct ggaccctgct ggatatgtgg
gacaccatca actttgagag caccgggaat 780ctgattgccc ctgagtacgg cttcaagatc
agcaagagag gcagcagcgg catcatgaaa 840acagagggca ccctggaaaa ctgtgaaacc
aagtgtcaga cacctctggg cgccattaat 900accaccctgc ccttccataa tgtgcaccct
ctgacaatcg gcgagtgccc taagtacgtg 960aagtctgaga aactggtgct ggccacagga
ctgagaaatg tgccccagat cgagtcaaga 1020ggcctgtttg gagccattgc cggctttatt
gaaggcggat ggcagggaat ggtggatggg 1080tggtacggct atcaccacag caatgatcag
ggatctggct atgccgccga taaagagagc 1140acccagaagg cctttgacgg catcaccaac
aaagtgaaca gcgtgatcga gaagatgaac 1200acccagtttg aggccgtggg caaagagttc
agcaatctgg aaagacggct ggaaaacctg 1260aacaagaaaa tggaagatgg cttcctggac
gtgtggacat ataatgccga gctgctggtg 1320ctgatggaaa acgagaggac cctggacttt
cacgacagca acgtgaagaa cctgtacgac 1380aaagtgcgga tgcagctgag agacaatgtg
aaagagctgg gcaacggctg ctttgagttc 1440taccacaagt gcgacgacga gtgcatgaat
agcgtgaaga acggcaccta cgactaccct 1500aagtatgagg aagagagcaa gctgaacaga
aacgagatca ag 154217514PRTInfluenza virus 17Met Ala
Ile Ile Tyr Leu Ile Leu Leu Phe Thr Ala Val Arg Gly Asp 1 5
10 15 Gln Ile Cys Ile Gly Tyr His
Ala Asn Asn Ser Thr Glu Lys Val Asp 20 25
30 Thr Ile Leu Glu Arg Asn Val Thr Val Thr His Ala
Lys Asp Ile Leu 35 40 45
Glu Lys Thr His Asn Gly Lys Leu Cys Lys Leu Asn Gly Ile Pro Pro
50 55 60 Leu Glu Leu
Gly Asp Cys Ser Ile Ala Gly Trp Leu Leu Gly Asn Pro 65
70 75 80 Glu Cys Asp Arg Leu Leu Ser
Val Pro Glu Trp Ser Tyr Ile Met Glu 85
90 95 Lys Glu Asn Pro Arg Asp Gly Leu Cys Tyr Pro
Gly Ser Phe Asn Asp 100 105
110 Tyr Glu Glu Leu Lys His Leu Leu Ser Ser Val Lys His Phe Glu
Lys 115 120 125 Val
Lys Ile Leu Pro Lys Asp Arg Trp Thr Gln His Thr Thr Thr Gly 130
135 140 Gly Ser Arg Ala Cys Ala
Val Ser Gly Asn Pro Ser Phe Phe Arg Asn 145 150
155 160 Met Val Trp Leu Thr Lys Lys Gly Ser Asn Tyr
Pro Val Ala Lys Gly 165 170
175 Ser Tyr Asn Asn Thr Ser Gly Glu Gln Met Leu Ile Ile Trp Gly Val
180 185 190 His His
Pro Asn Asp Glu Thr Glu Gln Arg Thr Leu Tyr Gln Asn Val 195
200 205 Gly Thr Tyr Val Ser Val Gly
Thr Ser Thr Leu Asn Lys Arg Ser Thr 210 215
220 Pro Asp Ile Ala Thr Arg Pro Lys Val Asn Gly Gln
Gly Gly Arg Met 225 230 235
240 Glu Phe Ser Trp Thr Leu Leu Asp Met Trp Asp Thr Ile Asn Phe Glu
245 250 255 Ser Thr Gly
Asn Leu Ile Ala Pro Glu Tyr Gly Phe Lys Ile Ser Lys 260
265 270 Arg Gly Ser Ser Gly Ile Met Lys
Thr Glu Gly Thr Leu Glu Asn Cys 275 280
285 Glu Thr Lys Cys Gln Thr Pro Leu Gly Ala Ile Asn Thr
Thr Leu Pro 290 295 300
Phe His Asn Val His Pro Leu Thr Ile Gly Glu Cys Pro Lys Tyr Val 305
310 315 320 Lys Ser Glu Lys
Leu Val Leu Ala Thr Gly Leu Arg Asn Val Pro Gln 325
330 335 Ile Glu Ser Arg Gly Leu Phe Gly Ala
Ile Ala Gly Phe Ile Glu Gly 340 345
350 Gly Trp Gln Gly Met Val Asp Gly Trp Tyr Gly Tyr His His
Ser Asn 355 360 365
Asp Gln Gly Ser Gly Tyr Ala Ala Asp Lys Glu Ser Thr Gln Lys Ala 370
375 380 Phe Asp Gly Ile Thr
Asn Lys Val Asn Ser Val Ile Glu Lys Met Asn 385 390
395 400 Thr Gln Phe Glu Ala Val Gly Lys Glu Phe
Ser Asn Leu Glu Arg Arg 405 410
415 Leu Glu Asn Leu Asn Lys Lys Met Glu Asp Gly Phe Leu Asp Val
Trp 420 425 430 Thr
Tyr Asn Ala Glu Leu Leu Val Leu Met Glu Asn Glu Arg Thr Leu 435
440 445 Asp Phe His Asp Ser Asn
Val Lys Asn Leu Tyr Asp Lys Val Arg Met 450 455
460 Gln Leu Arg Asp Asn Val Lys Glu Leu Gly Asn
Gly Cys Phe Glu Phe 465 470 475
480 Tyr His Lys Cys Asp Asp Glu Cys Met Asn Ser Val Lys Asn Gly Thr
485 490 495 Tyr Asp
Tyr Pro Lys Tyr Glu Glu Glu Ser Lys Leu Asn Arg Asn Glu 500
505 510 Ile Lys 181542DNAInfluenza
virus 18cttgatctcg tttctgttca gcttgctctc ttcctcatac ttagggtagt cgtaggtgcc
60gttcttcacg ctattcatgc actcgtcgtc gcacttgtgg tagaactcaa agcagccgtt
120gcccagctct ttcacattgt ctctcagctg catccgcact ttgtcgtaca ggttcttcac
180gttgctgtcg tgaaagtcca gggtcctctc gttttccatc agcaccagca gctcggcatt
240atatgtccac acgtccagga agccatcttc cattttcttg ttcaggtttt ccagccgtct
300ttccagattg ctgaactctt tgcccacggc ctcaaactgg gtgttcatct tctcgatcac
360gctgttcact ttgttggtga tgccgtcaaa ggccttctgg gtgctctctt tatcggcggc
420atagccagat ccctgatcat tgctgtggtg atagccgtac cacccatcca ccattccctg
480ccatccgcct tcaataaagc cggcaatggc tccaaacagg cctcttgact cgatctgggg
540cacatttctc agtcctgtgg ccagcaccag tttctcagac ttcacgtact tagggcactc
600gccgattgtc agagggtgca cattatggaa gggcagggtg gtattaatgg cgcccagagg
660tgtctgacac ttggtttcac agttttccag ggtgccctct gttttcatga tgccgctgct
720gcctctcttg ctgatcttga agccgtactc aggggcaatc agattcccgg tgctctcaaa
780gttgatggtg tcccacatat ccagcagggt ccaggaaaat tccattctgc cgccctgtcc
840attcactttg ggtctggtgg caatatcggg ggtgcttctc ttattcagtg tgctggtgcc
900cacagacacg tatgtgccca cattctggta cagggttctc tgctctgtct catcattagg
960gtggtgcacg ccccaaataa tcagcatctg ctcgccgctt gtattattgt agctgccctt
1020ggccacagga taattagagc ccttcttggt cagccagacc atatttctga agaagctggg
1080gttgccggac acagcacaag ctctgcttcc gcctgttgtt gtatgctggg tccatctatc
1140cttgggcagg atcttcactt tctcgaagtg cttcacgctg gacagcaggt gcttcagttc
1200ctcgtaatcg ttgaagctgc cggggtaaca cagtccgtcc ctagggttct ctttttccat
1260gatgtagctc cactcaggca cagacagcag tctatcgcac tcaggatttc ccagcagcca
1320gccagcaata gaacaatcgc ccagttccag aggaggaatg ccattcagct tgcacagctt
1380gccgttgtgg gtcttttcca gaatatcctt ggcgtgggtc acggtcacat ttctttccag
1440gatggtgtcc accttctcgg tgctattgtt ggcgtggtag ccgatacaga tctgatcgcc
1500ccgcacagct gtaaacagca ggatcaggta gatgatggcc at
1542191557DNAInfluenza virus 19atgaaaacca tcattgccct gagctacatc
ttttgtctgg ctctgggcca ggatctgccc 60ggcaatgata atagcaccgc caccctgtgt
ctgggacacc acgccgtgcc taatggcacc 120ctggtgaaaa ccattaccga cgaccagatc
gaagtgacca atgccaccga gctggtgcag 180agcagcagca ccggcaagat ctgcaacaac
ccccacagaa tcctggatgg catcgactgt 240accctgatcg atgccctgct gggcgatcct
cactgcgacg tgttccagaa cgagacatgg 300gacctgttcg tggagagaag caaggccttc
agcaactgct acccctacga tgtgcccgat 360tacgcctctc tgagaagcct ggtggccagc
agcggcacac tggaattcat caccgagggc 420tttacctgga caggcgtgac ccagaatggc
ggcagcaatg cctgtaaaag aggccctggc 480agcggcttct tcagcagact gaactggctg
accaagtccg gcagcaccta ccctgtgctg 540aacgtgacca tgcccaacaa cgacaacttc
gacaagctgt acatctgggg cgtgcaccac 600cctagcacca atcaggaaca gaccagcctg
tacgtgcagg ccagcggcag agtgaccgtg 660tctaccagac ggtcccagca gaccatcatc
cccaacatcg agtcaagacc ttgggtgcgc 720ggcctgagca gcagaatcag catctactgg
accatcgtga aacctggcga cgtgctggtg 780atcaacagca atggcaacct gatcgccccc
agaggctact tcaagatgcg gaccggcaag 840agcagcatca tgagaagcga cgcccccatc
gatacctgta tcagcgagtg catcaccccc 900aacggcagca tccccaacga caagcccttc
cagaacgtga acaagatcac ctacggcgcc 960tgccctaagt acgtgaagca gaacaccctg
aagctggcca ccggcatgag aaatgtgccc 1020gagaagcaga caagaggcct gtttggcgcc
attgccggct ttatcgagaa cggctgggag 1080ggcatgatcg atgggtggta cggcttcaga
caccagaatt ctgagggcac aggacaggcc 1140gccgatctga agtctacaca ggccgccatc
gaccagatca acggcaagct gaacagagtg 1200atcgagaaaa ccaacgagaa gttccaccag
atcgagaaag aattcagcga ggtggagggc 1260agaatccagg acctggaaaa atacgtggag
gacaccaaga tcgacctgtg gagctacaat 1320gccgaactgc tggtcgccct ggaaaaccag
cacaccatcg acctgaccga cagcgagatg 1380aataagctgt tcgaaaagac cagacggcag
ctgagagaaa acgccgagga catgggcaac 1440ggctgcttca agatctacca caagtgcgac
aacgcctgca tcgagagcat cagaaacggc 1500acctacgacc acgatgtgta cagggacgag
gccctgaaca acagattcca gatcaag 155720519PRTInfluenza virus 20Met Lys
Thr Ile Ile Ala Leu Ser Tyr Ile Phe Cys Leu Ala Leu Gly 1 5
10 15 Gln Asp Leu Pro Gly Asn Asp
Asn Ser Thr Ala Thr Leu Cys Leu Gly 20 25
30 His His Ala Val Pro Asn Gly Thr Leu Val Lys Thr
Ile Thr Asp Asp 35 40 45
Gln Ile Glu Val Thr Asn Ala Thr Glu Leu Val Gln Ser Ser Ser Thr
50 55 60 Gly Lys Ile
Cys Asn Asn Pro His Arg Ile Leu Asp Gly Ile Asp Cys 65
70 75 80 Thr Leu Ile Asp Ala Leu Leu
Gly Asp Pro His Cys Asp Val Phe Gln 85
90 95 Asn Glu Thr Trp Asp Leu Phe Val Glu Arg Ser
Lys Ala Phe Ser Asn 100 105
110 Cys Tyr Pro Tyr Asp Val Pro Asp Tyr Ala Ser Leu Arg Ser Leu
Val 115 120 125 Ala
Ser Ser Gly Thr Leu Glu Phe Ile Thr Glu Gly Phe Thr Trp Thr 130
135 140 Gly Val Thr Gln Asn Gly
Gly Ser Asn Ala Cys Lys Arg Gly Pro Gly 145 150
155 160 Ser Gly Phe Phe Ser Arg Leu Asn Trp Leu Thr
Lys Ser Gly Ser Thr 165 170
175 Tyr Pro Val Leu Asn Val Thr Met Pro Asn Asn Asp Asn Phe Asp Lys
180 185 190 Leu Tyr
Ile Trp Gly Val His His Pro Ser Thr Asn Gln Glu Gln Thr 195
200 205 Ser Leu Tyr Val Gln Ala Ser
Gly Arg Val Thr Val Ser Thr Arg Arg 210 215
220 Ser Gln Gln Thr Ile Ile Pro Asn Ile Glu Ser Arg
Pro Trp Val Arg 225 230 235
240 Gly Leu Ser Ser Arg Ile Ser Ile Tyr Trp Thr Ile Val Lys Pro Gly
245 250 255 Asp Val Leu
Val Ile Asn Ser Asn Gly Asn Leu Ile Ala Pro Arg Gly 260
265 270 Tyr Phe Lys Met Arg Thr Gly Lys
Ser Ser Ile Met Arg Ser Asp Ala 275 280
285 Pro Ile Asp Thr Cys Ile Ser Glu Cys Ile Thr Pro Asn
Gly Ser Ile 290 295 300
Pro Asn Asp Lys Pro Phe Gln Asn Val Asn Lys Ile Thr Tyr Gly Ala 305
310 315 320 Cys Pro Lys Tyr
Val Lys Gln Asn Thr Leu Lys Leu Ala Thr Gly Met 325
330 335 Arg Asn Val Pro Glu Lys Gln Thr Arg
Gly Leu Phe Gly Ala Ile Ala 340 345
350 Gly Phe Ile Glu Asn Gly Trp Glu Gly Met Ile Asp Gly Trp
Tyr Gly 355 360 365
Phe Arg His Gln Asn Ser Glu Gly Thr Gly Gln Ala Ala Asp Leu Lys 370
375 380 Ser Thr Gln Ala Ala
Ile Asp Gln Ile Asn Gly Lys Leu Asn Arg Val 385 390
395 400 Ile Glu Lys Thr Asn Glu Lys Phe His Gln
Ile Glu Lys Glu Phe Ser 405 410
415 Glu Val Glu Gly Arg Ile Gln Asp Leu Glu Lys Tyr Val Glu Asp
Thr 420 425 430 Lys
Ile Asp Leu Trp Ser Tyr Asn Ala Glu Leu Leu Val Ala Leu Glu 435
440 445 Asn Gln His Thr Ile Asp
Leu Thr Asp Ser Glu Met Asn Lys Leu Phe 450 455
460 Glu Lys Thr Arg Arg Gln Leu Arg Glu Asn Ala
Glu Asp Met Gly Asn 465 470 475
480 Gly Cys Phe Lys Ile Tyr His Lys Cys Asp Asn Ala Cys Ile Glu Ser
485 490 495 Ile Arg
Asn Gly Thr Tyr Asp His Asp Val Tyr Arg Asp Glu Ala Leu 500
505 510 Asn Asn Arg Phe Gln Ile Lys
515 211557DNAInfluenza virus 21cttgatctgg
aatctgttgt tcagggcctc gtccctgtac acatcgtggt cgtaggtgcc 60gtttctgatg
ctctcgatgc aggcgttgtc gcacttgtgg tagatcttga agcagccgtt 120gcccatgtcc
tcggcgtttt ctctcagctg ccgtctggtc ttttcgaaca gcttattcat 180ctcgctgtcg
gtcaggtcga tggtgtgctg gttttccagg gcgaccagca gttcggcatt 240gtagctccac
aggtcgatct tggtgtcctc cacgtatttt tccaggtcct ggattctgcc 300ctccacctcg
ctgaattctt tctcgatctg gtggaacttc tcgttggttt tctcgatcac 360tctgttcagc
ttgccgttga tctggtcgat ggcggcctgt gtagacttca gatcggcggc 420ctgtcctgtg
ccctcagaat tctggtgtct gaagccgtac cacccatcga tcatgccctc 480ccagccgttc
tcgataaagc cggcaatggc gccaaacagg cctcttgtct gcttctcggg 540cacatttctc
atgccggtgg ccagcttcag ggtgttctgc ttcacgtact tagggcaggc 600gccgtaggtg
atcttgttca cgttctggaa gggcttgtcg ttggggatgc tgccgttggg 660ggtgatgcac
tcgctgatac aggtatcgat gggggcgtcg cttctcatga tgctgctctt 720gccggtccgc
atcttgaagt agcctctggg ggcgatcagg ttgccattgc tgttgatcac 780cagcacgtcg
ccaggtttca cgatggtcca gtagatgctg attctgctgc tcaggccgcg 840cacccaaggt
cttgactcga tgttggggat gatggtctgc tgggaccgtc tggtagacac 900ggtcactctg
ccgctggcct gcacgtacag gctggtctgt tcctgattgg tgctagggtg 960gtgcacgccc
cagatgtaca gcttgtcgaa gttgtcgttg ttgggcatgg tcacgttcag 1020cacagggtag
gtgctgccgg acttggtcag ccagttcagt ctgctgaaga agccgctgcc 1080agggcctctt
ttacaggcat tgctgccgcc attctgggtc acgcctgtcc aggtaaagcc 1140ctcggtgatg
aattccagtg tgccgctgct ggccaccagg cttctcagag aggcgtaatc 1200gggcacatcg
taggggtagc agttgctgaa ggccttgctt ctctccacga acaggtccca 1260tgtctcgttc
tggaacacgt cgcagtgagg atcgcccagc agggcatcga tcagggtaca 1320gtcgatgcca
tccaggattc tgtgggggtt gttgcagatc ttgccggtgc tgctgctctg 1380caccagctcg
gtggcattgg tcacttcgat ctggtcgtcg gtaatggttt tcaccagggt 1440gccattaggc
acggcgtggt gtcccagaca cagggtggcg gtgctattat cattgccggg 1500cagatcctgg
cccagagcca gacaaaagat gtagctcagg gcaatgatgg ttttcat
1557221557DNAInfluenza virus 22atgaaaacca tcattgccct gagctacatc
ctgtgcctgg tgttcacaca gaagctgccc 60ggcaacgata atagcaccgc cacactgtgt
ctgggacacc acgccgtgcc taatggcacc 120atcgtgaaaa caatcaccaa cgaccagatc
gaagtgacca atgccacaga gctggtgcag 180agcagcagca caggcgagat ctgtgacagc
ccccaccaga tcctggatgg cgagaactgt 240accctgatcg atgccctgct gggcgatcct
cagtgcgacg gcttccagaa caagaaatgg 300gacctgttcg tggagagaag caaggcctac
agcaactgct acccctacga cgtgcctgat 360tacgccagcc tgagaagcct ggtggcctct
agcggcaccc tggaattcaa caacgagagc 420ttcaactgga ccggcgtgac acagaatggc
accagcagcg cctgcatcag acggtccaac 480aacagcttct tcagtagact gaattggctg
acccacctga agttcaagta ccccgccctg 540aacgtgacca tgcccaacaa tgagaagttc
gacaagctgt acatctgggg agtgcaccac 600cctggcaccg acaacgatca gatcttccct
tacgcccagg ccagcggcag aatcaccgtg 660tccaccaaga gaagccagca gaccgtgatc
cccaatatcg gcagcagacc cagagtgcgg 720aacatcccca gcaggatcag catctactgg
acaatcgtga agcctggcga catcctgctg 780atcaacagca ccggcaacct gatcgcccct
cggggctact ttaagatcag aagcggcaag 840agcagcatca tgagatccga cgcccccatc
ggcaagtgca acagcgagtg catcacccca 900aacggcagca tccccaacga caagcccttc
cagaacgtga acaggatcac ctacggcgcc 960tgccctagat acgtgaagca gaacaccctg
aagctggcca ccggcatgag aaatgtgccc 1020gagaagcaga ccagaggcat ctttggcgcc
attgccggct ttatcgagaa tggctgggag 1080ggaatggtgg atgggtggta cggcttcaga
caccagaata gcgagggaat tggacaggcc 1140gccgatctga aatctaccca ggccgccatc
gaccagatca acggcaagct gaacaggctg 1200atcggcaaga ccaacgagaa gttccaccag
atcgagaaag aattcagcga ggtggagggc 1260agaatccagg acctggaaaa atacgtggag
gacaccaaga tcgacctgtg gagctacaat 1320gccgaactgc tggtcgccct ggaaaaccag
cacacaattg atctgacaga cagtgagatg 1380aataagctgt tcgagaaaac caagaagcag
ctgagagaaa acgccgagga catgggcaac 1440ggctgcttca agatctacca caagtgcgac
aacgcctgca tcggcagcat cagaaacggc 1500acctacgacc acgacgtgta cagagatgag
gccctgaaca accggtttca gatcaag 155723519PRTInfluenza virus 23Met Lys
Thr Ile Ile Ala Leu Ser Tyr Ile Leu Cys Leu Val Phe Thr 1 5
10 15 Gln Lys Leu Pro Gly Asn Asp
Asn Ser Thr Ala Thr Leu Cys Leu Gly 20 25
30 His His Ala Val Pro Asn Gly Thr Ile Val Lys Thr
Ile Thr Asn Asp 35 40 45
Gln Ile Glu Val Thr Asn Ala Thr Glu Leu Val Gln Ser Ser Ser Thr
50 55 60 Gly Glu Ile
Cys Asp Ser Pro His Gln Ile Leu Asp Gly Glu Asn Cys 65
70 75 80 Thr Leu Ile Asp Ala Leu Leu
Gly Asp Pro Gln Cys Asp Gly Phe Gln 85
90 95 Asn Lys Lys Trp Asp Leu Phe Val Glu Arg Ser
Lys Ala Tyr Ser Asn 100 105
110 Cys Tyr Pro Tyr Asp Val Pro Asp Tyr Ala Ser Leu Arg Ser Leu
Val 115 120 125 Ala
Ser Ser Gly Thr Leu Glu Phe Asn Asn Glu Ser Phe Asn Trp Thr 130
135 140 Gly Val Thr Gln Asn Gly
Thr Ser Ser Ala Cys Ile Arg Arg Ser Asn 145 150
155 160 Asn Ser Phe Phe Ser Arg Leu Asn Trp Leu Thr
His Leu Lys Phe Lys 165 170
175 Tyr Pro Ala Leu Asn Val Thr Met Pro Asn Asn Glu Lys Phe Asp Lys
180 185 190 Leu Tyr
Ile Trp Gly Val His His Pro Gly Thr Asp Asn Asp Gln Ile 195
200 205 Phe Pro Tyr Ala Gln Ala Ser
Gly Arg Ile Thr Val Ser Thr Lys Arg 210 215
220 Ser Gln Gln Thr Val Ile Pro Asn Ile Gly Ser Arg
Pro Arg Val Arg 225 230 235
240 Asn Ile Pro Ser Arg Ile Ser Ile Tyr Trp Thr Ile Val Lys Pro Gly
245 250 255 Asp Ile Leu
Leu Ile Asn Ser Thr Gly Asn Leu Ile Ala Pro Arg Gly 260
265 270 Tyr Phe Lys Ile Arg Ser Gly Lys
Ser Ser Ile Met Arg Ser Asp Ala 275 280
285 Pro Ile Gly Lys Cys Asn Ser Glu Cys Ile Thr Pro Asn
Gly Ser Ile 290 295 300
Pro Asn Asp Lys Pro Phe Gln Asn Val Asn Arg Ile Thr Tyr Gly Ala 305
310 315 320 Cys Pro Arg Tyr
Val Lys Gln Asn Thr Leu Lys Leu Ala Thr Gly Met 325
330 335 Arg Asn Val Pro Glu Lys Gln Thr Arg
Gly Ile Phe Gly Ala Ile Ala 340 345
350 Gly Phe Ile Glu Asn Gly Trp Glu Gly Met Val Asp Gly Trp
Tyr Gly 355 360 365
Phe Arg His Gln Asn Ser Glu Gly Ile Gly Gln Ala Ala Asp Leu Lys 370
375 380 Ser Thr Gln Ala Ala
Ile Asp Gln Ile Asn Gly Lys Leu Asn Arg Leu 385 390
395 400 Ile Gly Lys Thr Asn Glu Lys Phe His Gln
Ile Glu Lys Glu Phe Ser 405 410
415 Glu Val Glu Gly Arg Ile Gln Asp Leu Glu Lys Tyr Val Glu Asp
Thr 420 425 430 Lys
Ile Asp Leu Trp Ser Tyr Asn Ala Glu Leu Leu Val Ala Leu Glu 435
440 445 Asn Gln His Thr Ile Asp
Leu Thr Asp Ser Glu Met Asn Lys Leu Phe 450 455
460 Glu Lys Thr Lys Lys Gln Leu Arg Glu Asn Ala
Glu Asp Met Gly Asn 465 470 475
480 Gly Cys Phe Lys Ile Tyr His Lys Cys Asp Asn Ala Cys Ile Gly Ser
485 490 495 Ile Arg
Asn Gly Thr Tyr Asp His Asp Val Tyr Arg Asp Glu Ala Leu 500
505 510 Asn Asn Arg Phe Gln Ile Lys
515 241557DNAInfluenza virus 24cttgatctga
aaccggttgt tcagggcctc atctctgtac acgtcgtggt cgtaggtgcc 60gtttctgatg
ctgccgatgc aggcgttgtc gcacttgtgg tagatcttga agcagccgtt 120gcccatgtcc
tcggcgtttt ctctcagctg cttcttggtt ttctcgaaca gcttattcat 180ctcactgtct
gtcagatcaa ttgtgtgctg gttttccagg gcgaccagca gttcggcatt 240gtagctccac
aggtcgatct tggtgtcctc cacgtatttt tccaggtcct ggattctgcc 300ctccacctcg
ctgaattctt tctcgatctg gtggaacttc tcgttggtct tgccgatcag 360cctgttcagc
ttgccgttga tctggtcgat ggcggcctgg gtagatttca gatcggcggc 420ctgtccaatt
ccctcgctat tctggtgtct gaagccgtac cacccatcca ccattccctc 480ccagccattc
tcgataaagc cggcaatggc gccaaagatg cctctggtct gcttctcggg 540cacatttctc
atgccggtgg ccagcttcag ggtgttctgc ttcacgtatc tagggcaggc 600gccgtaggtg
atcctgttca cgttctggaa gggcttgtcg ttggggatgc tgccgtttgg 660ggtgatgcac
tcgctgttgc acttgccgat gggggcgtcg gatctcatga tgctgctctt 720gccgcttctg
atcttaaagt agccccgagg ggcgatcagg ttgccggtgc tgttgatcag 780caggatgtcg
ccaggcttca cgattgtcca gtagatgctg atcctgctgg ggatgttccg 840cactctgggt
ctgctgccga tattggggat cacggtctgc tggcttctct tggtggacac 900ggtgattctg
ccgctggcct gggcgtaagg gaagatctga tcgttgtcgg tgccagggtg 960gtgcactccc
cagatgtaca gcttgtcgaa cttctcattg ttgggcatgg tcacgttcag 1020ggcggggtac
ttgaacttca ggtgggtcag ccaattcagt ctactgaaga agctgttgtt 1080ggaccgtctg
atgcaggcgc tgctggtgcc attctgtgtc acgccggtcc agttgaagct 1140ctcgttgttg
aattccaggg tgccgctaga ggccaccagg cttctcaggc tggcgtaatc 1200aggcacgtcg
taggggtagc agttgctgta ggccttgctt ctctccacga acaggtccca 1260tttcttgttc
tggaagccgt cgcactgagg atcgcccagc agggcatcga tcagggtaca 1320gttctcgcca
tccaggatct ggtgggggct gtcacagatc tcgcctgtgc tgctgctctg 1380caccagctct
gtggcattgg tcacttcgat ctggtcgttg gtgattgttt tcacgatggt 1440gccattaggc
acggcgtggt gtcccagaca cagtgtggcg gtgctattat cgttgccggg 1500cagcttctgt
gtgaacacca ggcacaggat gtagctcagg gcaatgatgg ttttcat
1557251560DNAInfluenza virus 25atggaaaaga tcgtgctgct gctggccatt
gtgagcctgg tgaagagcga ccagatctgc 60attggctacc acgccaacaa tagcacagag
caggtggaca ccatcatgga aaaaaacgtg 120accgtgaccc acgctcagga catcctggaa
aagacccaca acggcaagct gtgtgatctg 180gacggcgtga agcctctgat cctgagagat
tgtagcgtgg ctggatggct gctgggcaac 240cctatgtgcg acgagttcat caacgtgccc
gagtggagct atatcgtgga gaaggccaac 300cccaccaacg atctgtgtta ccccggcagc
ttcaacgatt acgaggaact gaagcacctg 360ctgtcccgga tcaaccactt cgagaagatc
cagatcatcc ccaagtcctc ttggagcgat 420cacgaagcct ctagcggagt gtctagcgcc
tgtccttacc tgggcagccc cagcttcttc 480agaaacgtgg tgtggctgat caagaagaac
agcacctacc ccaccatcaa gaagagctac 540aacaacacca accaggaaga tctgctggtc
ctgtggggaa tccaccaccc taatgatgcc 600gccgagcaga ccagactgta ccagaacccc
accacctata tcagcatcgg caccagcacc 660ctgaatcaga gactggtgcc caagatcgcc
accagatcca aggtgaacgg ccagagcggc 720aggatggaat tcttctggac catcctgaag
cccaacgacg ccatcaactt cgagagcaac 780ggcaacttta tcgcccctga gtacgcctac
aagatcgtga agaagggcga cagcgccatc 840atgaagagcg agctggaata cggcaactgc
aacaccaagt gccagacacc tatgggcgcc 900atcaacagca gcatgccctt ccacaacatc
caccctctga ccatcggcga gtgccctaag 960tacgtgaaga gcaacagact ggtgctggcc
acaggcctga gaaatagccc ccagcgggag 1020agcagaagaa agaagagggg cctgtttgga
gccatcgccg gctttattga aggcggctgg 1080cagggaatgg tggatggctg gtacggctac
caccacagca atgagcaggg ctctggatat 1140gccgccgaca aagagtctac ccagaaggcc
atcgacggcg tcaccaacaa ggtgaacagc 1200atcatcgaca agatgaacac ccagttcgag
gctgtgggca gagagttcaa caacctggaa 1260cggcggatcg agaacctgaa caagaaaatg
gaagatggct tcctggatgt gtggacctac 1320aatgccgaac tgctggtgct gatggaaaac
gagcggaccc tggacttcca cgacagcaac 1380gtgaagaacc tgtacgacaa agtgcggctg
cagctgagag acaacgccaa agagctgggc 1440aacggctgct tcgagttcta ccacaagtgc
gacaacgagt gcatggaaag catcaggaac 1500ggcacctaca actaccctca gtacagcgag
gaagccaggc tgaagaggga agagatcagc 156026520PRTInfluenza virus 26Met Glu
Lys Ile Val Leu Leu Leu Ala Ile Val Ser Leu Val Lys Ser 1 5
10 15 Asp Gln Ile Cys Ile Gly Tyr
His Ala Asn Asn Ser Thr Glu Gln Val 20 25
30 Asp Thr Ile Met Glu Lys Asn Val Thr Val Thr His
Ala Gln Asp Ile 35 40 45
Leu Glu Lys Thr His Asn Gly Lys Leu Cys Asp Leu Asp Gly Val Lys
50 55 60 Pro Leu Ile
Leu Arg Asp Cys Ser Val Ala Gly Trp Leu Leu Gly Asn 65
70 75 80 Pro Met Cys Asp Glu Phe Ile
Asn Val Pro Glu Trp Ser Tyr Ile Val 85
90 95 Glu Lys Ala Asn Pro Thr Asn Asp Leu Cys Tyr
Pro Gly Ser Phe Asn 100 105
110 Asp Tyr Glu Glu Leu Lys His Leu Leu Ser Arg Ile Asn His Phe
Glu 115 120 125 Lys
Ile Gln Ile Ile Pro Lys Ser Ser Trp Ser Asp His Glu Ala Ser 130
135 140 Ser Gly Val Ser Ser Ala
Cys Pro Tyr Leu Gly Ser Pro Ser Phe Phe 145 150
155 160 Arg Asn Val Val Trp Leu Ile Lys Lys Asn Ser
Thr Tyr Pro Thr Ile 165 170
175 Lys Lys Ser Tyr Asn Asn Thr Asn Gln Glu Asp Leu Leu Val Leu Trp
180 185 190 Gly Ile
His His Pro Asn Asp Ala Ala Glu Gln Thr Arg Leu Tyr Gln 195
200 205 Asn Pro Thr Thr Tyr Ile Ser
Ile Gly Thr Ser Thr Leu Asn Gln Arg 210 215
220 Leu Val Pro Lys Ile Ala Thr Arg Ser Lys Val Asn
Gly Gln Ser Gly 225 230 235
240 Arg Met Glu Phe Phe Trp Thr Ile Leu Lys Pro Asn Asp Ala Ile Asn
245 250 255 Phe Glu Ser
Asn Gly Asn Phe Ile Ala Pro Glu Tyr Ala Tyr Lys Ile 260
265 270 Val Lys Lys Gly Asp Ser Ala Ile
Met Lys Ser Glu Leu Glu Tyr Gly 275 280
285 Asn Cys Asn Thr Lys Cys Gln Thr Pro Met Gly Ala Ile
Asn Ser Ser 290 295 300
Met Pro Phe His Asn Ile His Pro Leu Thr Ile Gly Glu Cys Pro Lys 305
310 315 320 Tyr Val Lys Ser
Asn Arg Leu Val Leu Ala Thr Gly Leu Arg Asn Ser 325
330 335 Pro Gln Arg Glu Ser Arg Arg Lys Lys
Arg Gly Leu Phe Gly Ala Ile 340 345
350 Ala Gly Phe Ile Glu Gly Gly Trp Gln Gly Met Val Asp Gly
Trp Tyr 355 360 365
Gly Tyr His His Ser Asn Glu Gln Gly Ser Gly Tyr Ala Ala Asp Lys 370
375 380 Glu Ser Thr Gln Lys
Ala Ile Asp Gly Val Thr Asn Lys Val Asn Ser 385 390
395 400 Ile Ile Asp Lys Met Asn Thr Gln Phe Glu
Ala Val Gly Arg Glu Phe 405 410
415 Asn Asn Leu Glu Arg Arg Ile Glu Asn Leu Asn Lys Lys Met Glu
Asp 420 425 430 Gly
Phe Leu Asp Val Trp Thr Tyr Asn Ala Glu Leu Leu Val Leu Met 435
440 445 Glu Asn Glu Arg Thr Leu
Asp Phe His Asp Ser Asn Val Lys Asn Leu 450 455
460 Tyr Asp Lys Val Arg Leu Gln Leu Arg Asp Asn
Ala Lys Glu Leu Gly 465 470 475
480 Asn Gly Cys Phe Glu Phe Tyr His Lys Cys Asp Asn Glu Cys Met Glu
485 490 495 Ser Ile
Arg Asn Gly Thr Tyr Asn Tyr Pro Gln Tyr Ser Glu Glu Ala 500
505 510 Arg Leu Lys Arg Glu Glu Ile
Ser 515 520 271560DNAInfluenza virus 27gctgatctct
tccctcttca gcctggcttc ctcgctgtac tgagggtagt tgtaggtgcc 60gttcctgatg
ctttccatgc actcgttgtc gcacttgtgg tagaactcga agcagccgtt 120gcccagctct
ttggcgttgt ctctcagctg cagccgcact ttgtcgtaca ggttcttcac 180gttgctgtcg
tggaagtcca gggtccgctc gttttccatc agcaccagca gttcggcatt 240gtaggtccac
acatccagga agccatcttc cattttcttg ttcaggttct cgatccgccg 300ttccaggttg
ttgaactctc tgcccacagc ctcgaactgg gtgttcatct tgtcgatgat 360gctgttcacc
ttgttggtga cgccgtcgat ggccttctgg gtagactctt tgtcggcggc 420atatccagag
ccctgctcat tgctgtggtg gtagccgtac cagccatcca ccattccctg 480ccagccgcct
tcaataaagc cggcgatggc tccaaacagg cccctcttct ttcttctgct 540ctcccgctgg
gggctatttc tcaggcctgt ggccagcacc agtctgttgc tcttcacgta 600cttagggcac
tcgccgatgg tcagagggtg gatgttgtgg aagggcatgc tgctgttgat 660ggcgcccata
ggtgtctggc acttggtgtt gcagttgccg tattccagct cgctcttcat 720gatggcgctg
tcgcccttct tcacgatctt gtaggcgtac tcaggggcga taaagttgcc 780gttgctctcg
aagttgatgg cgtcgttggg cttcaggatg gtccagaaga attccatcct 840gccgctctgg
ccgttcacct tggatctggt ggcgatcttg ggcaccagtc tctgattcag 900ggtgctggtg
ccgatgctga tataggtggt ggggttctgg tacagtctgg tctgctcggc 960ggcatcatta
gggtggtgga ttccccacag gaccagcaga tcttcctggt tggtgttgtt 1020gtagctcttc
ttgatggtgg ggtaggtgct gttcttcttg atcagccaca ccacgtttct 1080gaagaagctg
gggctgccca ggtaaggaca ggcgctagac actccgctag aggcttcgtg 1140atcgctccaa
gaggacttgg ggatgatctg gatcttctcg aagtggttga tccgggacag 1200caggtgcttc
agttcctcgt aatcgttgaa gctgccgggg taacacagat cgttggtggg 1260gttggccttc
tccacgatat agctccactc gggcacgttg atgaactcgt cgcacatagg 1320gttgcccagc
agccatccag ccacgctaca atctctcagg atcagaggct tcacgccgtc 1380cagatcacac
agcttgccgt tgtgggtctt ttccaggatg tcctgagcgt gggtcacggt 1440cacgtttttt
tccatgatgg tgtccacctg ctctgtgcta ttgttggcgt ggtagccaat 1500gcagatctgg
tcgctcttca ccaggctcac aatggccagc agcagcacga tcttttccat
1560281602DNAInfluenza virus 28atgaaggcca tcatcgtgct gctgatggtg
gtgaccagca acgccgatag aatctgcacc 60ggcatcacca gcagcaatag cccccatgtg
gtgaaaacag ccacccaggg cgaagtgaat 120gtgacaggcg tgatccctct gaccaccacc
cccaccaaga gctacttcgc caacctgaag 180ggcaccagaa ccagaggcaa gctgtgcccc
gattgcctga actgcaccga tctggatgtg 240gctctgggca gacctatgtg tgtgggcacc
acaccatctg ccaaggccag catcctgcac 300gaagtgaagc ctgtgaccag cggctgcttc
cccatcatgc acgaccggac caagatcaga 360cagctgccca acctgctgag aggctacgag
aacatccggc tgtccaccca gaatgtgatc 420gatgccgaga aagcccctgg cggaccttat
agactgggca ccagcggctc ttgtcccaat 480gccacctcca agagcggctt ttttgccaca
atggcctggg ccgtgcctaa ggacaacaac 540aagaacgcca ccaaccctct gaccgtggag
gtgccctaca tctgtacaga gggcgaggat 600cagatcacag tgtggggctt ccacagcgac
gacaagaccc agatgaagaa cctgtacggc 660gacagcaacc cccagaagtt taccagcagc
gccaatggcg tgaccaccca ctacgtgtcc 720cagatcggca gctttcccga tcagacagag
gatggcggac tgcctcagtc tggcaggatc 780gtggtggact acatgatgca gaagcctggc
aagaccggca ccatcgtgta tcagagaggc 840gtgctgctgc ctcagaaagt gtggtgtgcc
agcggcaggt ctaaagtgat caagggcagc 900ctgcctctga ttggcgaggc cgactgtctg
cacgaaaagt acggcggcct gaacaagagc 960aagccctact acacaggcga gcacgccaag
gccatcggca attgccccat ctgggtgaaa 1020acccccctga agctggccaa tggcaccaag
tacagacctc ccgccaagct gctgaaagag 1080agaggcttct ttggcgccat tgccggattt
ctggaaggcg gctgggaggg aatgattgcc 1140ggctggcacg gctatacatc tcatggggcc
catggcgtgg ctgtggccgc cgatctgaag 1200tctacccagg aagccatcaa caagatcacc
aagaacctga acagcctgag cgagctggaa 1260gtgaagaatc tgcagagact gagcggcgcc
atggatgagc tgcacaacga gatcctggaa 1320ctggacgaga aagtggatga tctccgcgcc
gatacaattt cctcccagat tgaactggcc 1380gtgctgctgt ccaacgaggg catcatcaac
agcgaggatg aacacctgct ggccctggaa 1440cggaagctga agaagatgct gggcccttct
gccgtggaga tcggcaacgg ctgcttcgag 1500acaaagcaca agtgcaacca gacctgcctg
gatagaatcg ccgctggcac cttcaatgcc 1560ggcgagttca gcctgcctac cttcgacagc
ctgaatatca cc 160229534PRTInfluenza virus 29Met Lys
Ala Ile Ile Val Leu Leu Met Val Val Thr Ser Asn Ala Asp 1 5
10 15 Arg Ile Cys Thr Gly Ile Thr
Ser Ser Asn Ser Pro His Val Val Lys 20 25
30 Thr Ala Thr Gln Gly Glu Val Asn Val Thr Gly Val
Ile Pro Leu Thr 35 40 45
Thr Thr Pro Thr Lys Ser Tyr Phe Ala Asn Leu Lys Gly Thr Arg Thr
50 55 60 Arg Gly Lys
Leu Cys Pro Asp Cys Leu Asn Cys Thr Asp Leu Asp Val 65
70 75 80 Ala Leu Gly Arg Pro Met Cys
Val Gly Thr Thr Pro Ser Ala Lys Ala 85
90 95 Ser Ile Leu His Glu Val Lys Pro Val Thr Ser
Gly Cys Phe Pro Ile 100 105
110 Met His Asp Arg Thr Lys Ile Arg Gln Leu Pro Asn Leu Leu Arg
Gly 115 120 125 Tyr
Glu Asn Ile Arg Leu Ser Thr Gln Asn Val Ile Asp Ala Glu Lys 130
135 140 Ala Pro Gly Gly Pro Tyr
Arg Leu Gly Thr Ser Gly Ser Cys Pro Asn 145 150
155 160 Ala Thr Ser Lys Ser Gly Phe Phe Ala Thr Met
Ala Trp Ala Val Pro 165 170
175 Lys Asp Asn Asn Lys Asn Ala Thr Asn Pro Leu Thr Val Glu Val Pro
180 185 190 Tyr Ile
Cys Thr Glu Gly Glu Asp Gln Ile Thr Val Trp Gly Phe His 195
200 205 Ser Asp Asp Lys Thr Gln Met
Lys Asn Leu Tyr Gly Asp Ser Asn Pro 210 215
220 Gln Lys Phe Thr Ser Ser Ala Asn Gly Val Thr Thr
His Tyr Val Ser 225 230 235
240 Gln Ile Gly Ser Phe Pro Asp Gln Thr Glu Asp Gly Gly Leu Pro Gln
245 250 255 Ser Gly Arg
Ile Val Val Asp Tyr Met Met Gln Lys Pro Gly Lys Thr 260
265 270 Gly Thr Ile Val Tyr Gln Arg Gly
Val Leu Leu Pro Gln Lys Val Trp 275 280
285 Cys Ala Ser Gly Arg Ser Lys Val Ile Lys Gly Ser Leu
Pro Leu Ile 290 295 300
Gly Glu Ala Asp Cys Leu His Glu Lys Tyr Gly Gly Leu Asn Lys Ser 305
310 315 320 Lys Pro Tyr Tyr
Thr Gly Glu His Ala Lys Ala Ile Gly Asn Cys Pro 325
330 335 Ile Trp Val Lys Thr Pro Leu Lys Leu
Ala Asn Gly Thr Lys Tyr Arg 340 345
350 Pro Pro Ala Lys Leu Leu Lys Glu Arg Gly Phe Phe Gly Ala
Ile Ala 355 360 365
Gly Phe Leu Glu Gly Gly Trp Glu Gly Met Ile Ala Gly Trp His Gly 370
375 380 Tyr Thr Ser His Gly
Ala His Gly Val Ala Val Ala Ala Asp Leu Lys 385 390
395 400 Ser Thr Gln Glu Ala Ile Asn Lys Ile Thr
Lys Asn Leu Asn Ser Leu 405 410
415 Ser Glu Leu Glu Val Lys Asn Leu Gln Arg Leu Ser Gly Ala Met
Asp 420 425 430 Glu
Leu His Asn Glu Ile Leu Glu Leu Asp Glu Lys Val Asp Asp Leu 435
440 445 Arg Ala Asp Thr Ile Ser
Ser Gln Ile Glu Leu Ala Val Leu Leu Ser 450 455
460 Asn Glu Gly Ile Ile Asn Ser Glu Asp Glu His
Leu Leu Ala Leu Glu 465 470 475
480 Arg Lys Leu Lys Lys Met Leu Gly Pro Ser Ala Val Glu Ile Gly Asn
485 490 495 Gly Cys
Phe Glu Thr Lys His Lys Cys Asn Gln Thr Cys Leu Asp Arg 500
505 510 Ile Ala Ala Gly Thr Phe Asn
Ala Gly Glu Phe Ser Leu Pro Thr Phe 515 520
525 Asp Ser Leu Asn Ile Thr 530
301602DNAInfluenza virus 30ggtgatattc aggctgtcga aggtaggcag gctgaactcg
ccggcattga aggtgccagc 60ggcgattcta tccaggcagg tctggttgca cttgtgcttt
gtctcgaagc agccgttgcc 120gatctccacg gcagaagggc ccagcatctt cttcagcttc
cgttccaggg ccagcaggtg 180ttcatcctcg ctgttgatga tgccctcgtt ggacagcagc
acggccagtt caatctggga 240ggaaattgta tcggcgcgga gatcatccac tttctcgtcc
agttccagga tctcgttgtg 300cagctcatcc atggcgccgc tcagtctctg cagattcttc
acttccagct cgctcaggct 360gttcaggttc ttggtgatct tgttgatggc ttcctgggta
gacttcagat cggcggccac 420agccacgcca tgggccccat gagatgtata gccgtgccag
ccggcaatca ttccctccca 480gccgccttcc agaaatccgg caatggcgcc aaagaagcct
ctctctttca gcagcttggc 540gggaggtctg tacttggtgc cattggccag cttcaggggg
gttttcaccc agatggggca 600attgccgatg gccttggcgt gctcgcctgt gtagtagggc
ttgctcttgt tcaggccgcc 660gtacttttcg tgcagacagt cggcctcgcc aatcagaggc
aggctgccct tgatcacttt 720agacctgccg ctggcacacc acactttctg aggcagcagc
acgcctctct gatacacgat 780ggtgccggtc ttgccaggct tctgcatcat gtagtccacc
acgatcctgc cagactgagg 840cagtccgcca tcctctgtct gatcgggaaa gctgccgatc
tgggacacgt agtgggtggt 900cacgccattg gcgctgctgg taaacttctg ggggttgctg
tcgccgtaca ggttcttcat 960ctgggtcttg tcgtcgctgt ggaagcccca cactgtgatc
tgatcctcgc cctctgtaca 1020gatgtagggc acctccacgg tcagagggtt ggtggcgttc
ttgttgttgt ccttaggcac 1080ggcccaggcc attgtggcaa aaaagccgct cttggaggtg
gcattgggac aagagccgct 1140ggtgcccagt ctataaggtc cgccaggggc tttctcggca
tcgatcacat tctgggtgga 1200cagccggatg ttctcgtagc ctctcagcag gttgggcagc
tgtctgatct tggtccggtc 1260gtgcatgatg gggaagcagc cgctggtcac aggcttcact
tcgtgcagga tgctggcctt 1320ggcagatggt gtggtgccca cacacatagg tctgcccaga
gccacatcca gatcggtgca 1380gttcaggcaa tcggggcaca gcttgcctct ggttctggtg
cccttcaggt tggcgaagta 1440gctcttggtg ggggtggtgg tcagagggat cacgcctgtc
acattcactt cgccctgggt 1500ggctgttttc accacatggg ggctattgct gctggtgatg
ccggtgcaga ttctatcggc 1560gttgctggtc accaccatca gcagcacgat gatggccttc
at 1602311493DNAInfluenza virus 31atgacaactc
aacagccacg ctctgcttgg ggcaccatgc cgtccctaac gggaccattg 60tgaaaaccat
tactaacgat cagatagagg tgactaatgc caccgagctg gtgcaaagta 120gctccacagg
agagatctgc gatagtcccc accagattct ggacggaaag aattgtacgc 180tgatcgacgc
gctgttgggc gaccctcagt gtgacggatt tcagaataag aagtgggatc 240tgtttgtgga
aaggtcaaag gcttattcaa attgctaccc ttacgatgtg cctgattatg 300ccagcctgcg
gtccctcgtc gcgtctagtg ggactctgga gttcaacaac gagtcattta 360actggactgg
cgttacacag aacgggacta gttccgcttg cataaggaga agcaaaaata 420gtttcttcag
cagactgaat tggctgacac atctgaactt caagtaccct gcactgaatg 480taaccatgcc
caacaacgag cagttcgata agctttacat ttggggagtt catcatcctg 540gcactgacaa
ggatcagatc tttctgtatg cccaggcttc cggcaggatt accgtgtcta 600caaagagaag
ccagcaaact gtgtctccca atatcggcag tagacccaga gtacggaaca 660tccctagtcg
catcagtatt tactggacca tcgtgaaacc aggcgatatt ctcctgatta 720acagtactgg
caacctgatc gccccccggg gatactttaa aatccgctct ggaaagtcct 780ccattatgag
atcagatgca ccgatcggaa aatgcaactc tgagtgtatc acacccaatg 840ggagcattcc
caatgacaaa cctttccaga acgttaatcg aataacttat ggggcctgtc 900cacggtacgt
gaagcaaaat accttgaaac tggcgaccgg tatgcgcaat gtccccgaaa 960aacagacccg
cgggatattt ggggctatcg caggctttat cgagaatggc tgggaaggga 1020tggtggatgg
ttggtatggt tttagacatc aaaactccga aggcagaggc caggctgccg 1080atctcaagag
cacgcaggcc gctatagatc agatcaatgg aaagctcaac agactgatcg 1140ggaaaaccaa
cgaaaaattc catcagatcg agaaagagtt ctccgaagtc gaggggcgca 1200tacaggacct
ggagaagtat gttgaggata caaagattga tctgtggtcc tacaatgccg 1260agctgctggt
ggctctggag aatcagcaca ctattgacct gaccgattca gagatgaaca 1320aactttttga
gaagacgaag aagcagctta gagaaaatgc agaggacatg gggaacggat 1380gctttaaaat
atatcataag tgtgataatg cctgcatcgg atcaattaga aatggtacct 1440atgatcacga
tgtttacagg gacgaagcgc tgaataacag gttccagata aaa
149332519PRTInfluenza virus 32Met Lys Thr Ile Ile Ala Leu Ser Tyr Ile Leu
Cys Leu Val Phe Ala 1 5 10
15 Gln Lys Leu Pro Gly Asn Asp Asn Ser Thr Ala Thr Leu Cys Leu Gly
20 25 30 His His
Ala Val Pro Asn Gly Thr Ile Val Lys Thr Ile Thr Asn Asp 35
40 45 Gln Ile Glu Val Thr Asn Ala
Thr Glu Leu Val Gln Ser Ser Ser Thr 50 55
60 Gly Glu Ile Cys Asp Ser Pro His Gln Ile Leu Asp
Gly Lys Asn Cys 65 70 75
80 Thr Leu Ile Asp Ala Leu Leu Gly Asp Pro Gln Cys Asp Gly Phe Gln
85 90 95 Asn Lys Lys
Trp Asp Leu Phe Val Glu Arg Ser Lys Ala Tyr Ser Asn 100
105 110 Cys Tyr Pro Tyr Asp Val Pro Asp
Tyr Ala Ser Leu Arg Ser Leu Val 115 120
125 Ala Ser Ser Gly Thr Leu Glu Phe Asn Asn Glu Ser Phe
Asn Trp Thr 130 135 140
Gly Val Thr Gln Asn Gly Thr Ser Ser Ala Cys Ile Arg Arg Ser Lys 145
150 155 160 Asn Ser Phe Phe
Ser Arg Leu Asn Trp Leu Thr His Leu Asn Phe Lys 165
170 175 Tyr Pro Ala Leu Asn Val Thr Met Pro
Asn Asn Glu Gln Phe Asp Lys 180 185
190 Leu Tyr Ile Trp Gly Val His His Pro Gly Thr Asp Lys Asp
Gln Ile 195 200 205
Phe Leu Tyr Ala Gln Ala Ser Gly Arg Ile Thr Val Ser Thr Lys Arg 210
215 220 Ser Gln Gln Thr Val
Ser Pro Asn Ile Gly Ser Arg Pro Arg Val Arg 225 230
235 240 Asn Ile Pro Ser Arg Ile Ser Ile Tyr Trp
Thr Ile Val Lys Pro Gly 245 250
255 Asp Ile Leu Leu Ile Asn Ser Thr Gly Asn Leu Ile Ala Pro Arg
Gly 260 265 270 Tyr
Phe Lys Ile Arg Ser Gly Lys Ser Ser Ile Met Arg Ser Asp Ala 275
280 285 Pro Ile Gly Lys Cys Asn
Ser Glu Cys Ile Thr Pro Asn Gly Ser Ile 290 295
300 Pro Asn Asp Lys Pro Phe Gln Asn Val Asn Arg
Ile Thr Tyr Gly Ala 305 310 315
320 Cys Pro Arg Tyr Val Lys Gln Asn Thr Leu Lys Leu Ala Thr Gly Met
325 330 335 Arg Asn
Val Pro Glu Lys Gln Thr Arg Gly Ile Phe Gly Ala Ile Ala 340
345 350 Gly Phe Ile Glu Asn Gly Trp
Glu Gly Met Val Asp Gly Trp Tyr Gly 355 360
365 Phe Arg His Gln Asn Ser Glu Gly Arg Gly Gln Ala
Ala Asp Leu Lys 370 375 380
Ser Thr Gln Ala Ala Ile Asp Gln Ile Asn Gly Lys Leu Asn Arg Leu 385
390 395 400 Ile Gly Lys
Thr Asn Glu Lys Phe His Gln Ile Glu Lys Glu Phe Ser 405
410 415 Glu Val Glu Gly Arg Ile Gln Asp
Leu Glu Lys Tyr Val Glu Asp Thr 420 425
430 Lys Ile Asp Leu Trp Ser Tyr Asn Ala Glu Leu Leu Val
Ala Leu Glu 435 440 445
Asn Gln His Thr Ile Asp Leu Thr Asp Ser Glu Met Asn Lys Leu Phe 450
455 460 Glu Lys Thr Lys
Lys Gln Leu Arg Glu Asn Ala Glu Asp Met Gly Asn 465 470
475 480 Gly Cys Phe Lys Ile Tyr His Lys Cys
Asp Asn Ala Cys Ile Gly Ser 485 490
495 Ile Arg Asn Gly Thr Tyr Asp His Asp Val Tyr Arg Asp Glu
Ala Leu 500 505 510
Asn Asn Arg Phe Gln Ile Lys 515
331493DNAInfluenza virus 33ttttatctgg aacctgttat tcagcgcttc gtccctgtaa
acatcgtgat cataggtacc 60atttctaatt gatccgatgc aggcattatc acacttatga
tatattttaa agcatccgtt 120ccccatgtcc tctgcatttt ctctaagctg cttcttcgtc
ttctcaaaaa gtttgttcat 180ctctgaatcg gtcaggtcaa tagtgtgctg attctccaga
gccaccagca gctcggcatt 240gtaggaccac agatcaatct ttgtatcctc aacatacttc
tccaggtcct gtatgcgccc 300ctcgacttcg gagaactctt tctcgatctg atggaatttt
tcgttggttt tcccgatcag 360tctgttgagc tttccattga tctgatctat agcggcctgc
gtgctcttga gatcggcagc 420ctggcctctg ccttcggagt tttgatgtct aaaaccatac
caaccatcca ccatcccttc 480ccagccattc tcgataaagc ctgcgatagc cccaaatatc
ccgcgggtct gtttttcggg 540gacattgcgc ataccggtcg ccagtttcaa ggtattttgc
ttcacgtacc gtggacaggc 600cccataagtt attcgattaa cgttctggaa aggtttgtca
ttgggaatgc tcccattggg 660tgtgatacac tcagagttgc attttccgat cggtgcatct
gatctcataa tggaggactt 720tccagagcgg attttaaagt atccccgggg ggcgatcagg
ttgccagtac tgttaatcag 780gagaatatcg cctggtttca cgatggtcca gtaaatactg
atgcgactag ggatgttccg 840tactctgggt ctactgccga tattgggaga cacagtttgc
tggcttctct ttgtagacac 900ggtaatcctg ccggaagcct gggcatacag aaagatctga
tccttgtcag tgccaggatg 960atgaactccc caaatgtaaa gcttatcgaa ctgctcgttg
ttgggcatgg ttacattcag 1020tgcagggtac ttgaagttca gatgtgtcag ccaattcagt
ctgctgaaga aactattttt 1080gcttctcctt atgcaagcgg aactagtccc gttctgtgta
acgccagtcc agttaaatga 1140ctcgttgttg aactccagag tcccactaga cgcgacgagg
gaccgcaggc tggcataatc 1200aggcacatcg taagggtagc aatttgaata agcctttgac
ctttccacaa acagatccca 1260cttcttattc tgaaatccgt cacactgagg gtcgcccaac
agcgcgtcga tcagcgtaca 1320attctttccg tccagaatct ggtggggact atcgcagatc
tctcctgtgg agctactttg 1380caccagctcg gtggcattag tcacctctat ctgatcgtta
gtaatggttt tcacaatggt 1440cccgttaggg acggcatggt gccccaagca gagcgtggct
gttgagttgt cat 1493341551DNAInfluenza virus 34atgaaagtga
agctgctggt gctgctgtgt acctttaccg ccacctacgc cgataccatc 60tgtatcggct
accacgccaa caatagcacc gacaccgtgg ataccgtgct ggaaaagaac 120gtgaccgtga
cccacagcgt gaacctgctg gaaaacagcc acaacggcaa gctgtgtctg 180ctgaaaggca
ttgcccctct gcagctggga aattgtagcg tggccggctg gattctgggc 240aatcctgagt
gcgagctgct gatttccaaa gagtcctggt cctacatcgt ggagaagccc 300aaccctgaga
atggcacctg ctaccctggc cacttcgccg attacgagga actgagagaa 360cagctgtcca
gcgtgtccag cttcgagaga ttcgagatct tccccaaaga gagcagctgg 420cccaatcata
cagtgaccgg cgtgagcgcc tcttgtagcc acaatggcga gagcagcttc 480tacagaaacc
tgctgtggct gaccggcaag aacggcctgt accccaacct gagcaagagc 540tacgccaaca
acaaagaaaa agaagtgctg gtcctctggg gagtgcacca ccctcctaac 600atcggcatcc
agaaggccct gtaccacacc gagaatgcct acgtgtccgt ggtgtccagc 660cactacagca
gaaagttcac ccccgagatc gccaaaagac ccaaagtgcg ggaccaggaa 720ggcaggatca
actactactg gaccctgctg gaacctggcg acaccatcat cttcgaggcc 780aacggcaatc
tgatcgcccc tagatacgcc tttgccctga gcagaggctt tggcagcggc 840atcatcaaca
gcaacgcccc catggacaag tgtgacgcca agtgtcagac accacaggga 900gctatcaata
gcagcctgcc cttccagaat gtgcaccctg tgaccatcgg cgagtgtcct 960aaatacgtgc
ggagcgccaa gctgagaatg gtgaccggcc tgaggaatat ccccagcatc 1020cagagcagag
gcctgtttgg cgccattgcc ggctttatcg agggcggatg gacaggcatg 1080gtggatgggt
ggtacggcta ccaccaccag aatgagcagg gatctggcta tgccgccgat 1140cagaagagca
cccagaacgc catcaacggc atcaccaaca aagtgaacag cgtgatcgag 1200aagatgaaca
cccagttcac cgccgtgggc aaagagttca acaagctgga acggcggatg 1260gaaaacctga
acaagaaggt ggacgacggc ttcatcgaca tctggaccta caacgccgaa 1320ctcctggtcc
tcctggaaaa tgagaggacc ctggacttcc acgacagcaa cgtgaagaac 1380ctgtacgaga
aagtgaagag ccagctgaag aacaacgcca aagagatcgg caacggctgc 1440ttcgagttct
accacaagtg caacgacgag tgcatggaaa gcgtgaagaa cggcacctac 1500gactacccca
agtacagcga ggaaagcaag ctgaaccggg agaagatcga t
155135517PRTInfluenza virus 35Met Lys Val Lys Leu Leu Val Leu Leu Cys Thr
Phe Thr Ala Thr Tyr 1 5 10
15 Ala Asp Thr Ile Cys Ile Gly Tyr His Ala Asn Asn Ser Thr Asp Thr
20 25 30 Val Asp
Thr Val Leu Glu Lys Asn Val Thr Val Thr His Ser Val Asn 35
40 45 Leu Leu Glu Asn Ser His Asn
Gly Lys Leu Cys Leu Leu Lys Gly Ile 50 55
60 Ala Pro Leu Gln Leu Gly Asn Cys Ser Val Ala Gly
Trp Ile Leu Gly 65 70 75
80 Asn Pro Glu Cys Glu Leu Leu Ile Ser Lys Glu Ser Trp Ser Tyr Ile
85 90 95 Val Glu Lys
Pro Asn Pro Glu Asn Gly Thr Cys Tyr Pro Gly His Phe 100
105 110 Ala Asp Tyr Glu Glu Leu Arg Glu
Gln Leu Ser Ser Val Ser Ser Phe 115 120
125 Glu Arg Phe Glu Ile Phe Pro Lys Glu Ser Ser Trp Pro
Asn His Thr 130 135 140
Val Thr Gly Val Ser Ala Ser Cys Ser His Asn Gly Glu Ser Ser Phe 145
150 155 160 Tyr Arg Asn Leu
Leu Trp Leu Thr Gly Lys Asn Gly Leu Tyr Pro Asn 165
170 175 Leu Ser Lys Ser Tyr Ala Asn Asn Lys
Glu Lys Glu Val Leu Val Leu 180 185
190 Trp Gly Val His His Pro Pro Asn Ile Gly Ile Gln Lys Ala
Leu Tyr 195 200 205
His Thr Glu Asn Ala Tyr Val Ser Val Val Ser Ser His Tyr Ser Arg 210
215 220 Lys Phe Thr Pro Glu
Ile Ala Lys Arg Pro Lys Val Arg Asp Gln Glu 225 230
235 240 Gly Arg Ile Asn Tyr Tyr Trp Thr Leu Leu
Glu Pro Gly Asp Thr Ile 245 250
255 Ile Phe Glu Ala Asn Gly Asn Leu Ile Ala Pro Arg Tyr Ala Phe
Ala 260 265 270 Leu
Ser Arg Gly Phe Gly Ser Gly Ile Ile Asn Ser Asn Ala Pro Met 275
280 285 Asp Lys Cys Asp Ala Lys
Cys Gln Thr Pro Gln Gly Ala Ile Asn Ser 290 295
300 Ser Leu Pro Phe Gln Asn Val His Pro Val Thr
Ile Gly Glu Cys Pro 305 310 315
320 Lys Tyr Val Arg Ser Ala Lys Leu Arg Met Val Thr Gly Leu Arg Asn
325 330 335 Ile Pro
Ser Ile Gln Ser Arg Gly Leu Phe Gly Ala Ile Ala Gly Phe 340
345 350 Ile Glu Gly Gly Trp Thr Gly
Met Val Asp Gly Trp Tyr Gly Tyr His 355 360
365 His Gln Asn Glu Gln Gly Ser Gly Tyr Ala Ala Asp
Gln Lys Ser Thr 370 375 380
Gln Asn Ala Ile Asn Gly Ile Thr Asn Lys Val Asn Ser Val Ile Glu 385
390 395 400 Lys Met Asn
Thr Gln Phe Thr Ala Val Gly Lys Glu Phe Asn Lys Leu 405
410 415 Glu Arg Arg Met Glu Asn Leu Asn
Lys Lys Val Asp Asp Gly Phe Ile 420 425
430 Asp Ile Trp Thr Tyr Asn Ala Glu Leu Leu Val Leu Leu
Glu Asn Glu 435 440 445
Arg Thr Leu Asp Phe His Asp Ser Asn Val Lys Asn Leu Tyr Glu Lys 450
455 460 Val Lys Ser Gln
Leu Lys Asn Asn Ala Lys Glu Ile Gly Asn Gly Cys 465 470
475 480 Phe Glu Phe Tyr His Lys Cys Asn Asp
Glu Cys Met Glu Ser Val Lys 485 490
495 Asn Gly Thr Tyr Asp Tyr Pro Lys Tyr Ser Glu Glu Ser Lys
Leu Asn 500 505 510
Arg Glu Lys Ile Asp 515 361551DNAInfluenza virus
36atcgatcttc tcccggttca gcttgctttc ctcgctgtac ttggggtagt cgtaggtgcc
60gttcttcacg ctttccatgc actcgtcgtt gcacttgtgg tagaactcga agcagccgtt
120gccgatctct ttggcgttgt tcttcagctg gctcttcact ttctcgtaca ggttcttcac
180gttgctgtcg tggaagtcca gggtcctctc attttccagg aggaccagga gttcggcgtt
240gtaggtccag atgtcgatga agccgtcgtc caccttcttg ttcaggtttt ccatccgccg
300ttccagcttg ttgaactctt tgcccacggc ggtgaactgg gtgttcatct tctcgatcac
360gctgttcact ttgttggtga tgccgttgat ggcgttctgg gtgctcttct gatcggcggc
420atagccagat ccctgctcat tctggtggtg gtagccgtac cacccatcca ccatgcctgt
480ccatccgccc tcgataaagc cggcaatggc gccaaacagg cctctgctct ggatgctggg
540gatattcctc aggccggtca ccattctcag cttggcgctc cgcacgtatt taggacactc
600gccgatggtc acagggtgca cattctggaa gggcaggctg ctattgatag ctccctgtgg
660tgtctgacac ttggcgtcac acttgtccat gggggcgttg ctgttgatga tgccgctgcc
720aaagcctctg ctcagggcaa aggcgtatct aggggcgatc agattgccgt tggcctcgaa
780gatgatggtg tcgccaggtt ccagcagggt ccagtagtag ttgatcctgc cttcctggtc
840ccgcactttg ggtcttttgg cgatctcggg ggtgaacttt ctgctgtagt ggctggacac
900cacggacacg taggcattct cggtgtggta cagggccttc tggatgccga tgttaggagg
960gtggtgcact ccccagagga ccagcacttc tttttctttg ttgttggcgt agctcttgct
1020caggttgggg tacaggccgt tcttgccggt cagccacagc aggtttctgt agaagctgct
1080ctcgccattg tggctacaag aggcgctcac gccggtcact gtatgattgg gccagctgct
1140ctctttgggg aagatctcga atctctcgaa gctggacacg ctggacagct gttctctcag
1200ttcctcgtaa tcggcgaagt ggccagggta gcaggtgcca ttctcagggt tgggcttctc
1260cacgatgtag gaccaggact ctttggaaat cagcagctcg cactcaggat tgcccagaat
1320ccagccggcc acgctacaat ttcccagctg cagaggggca atgcctttca gcagacacag
1380cttgccgttg tggctgtttt ccagcaggtt cacgctgtgg gtcacggtca cgttcttttc
1440cagcacggta tccacggtgt cggtgctatt gttggcgtgg tagccgatac agatggtatc
1500ggcgtaggtg gcggtaaagg tacacagcag caccagcagc ttcactttca t
1551371605DNAInfluenza virus 37atgaaggcca tcatcgtgct gctgatggtg
gtcacaagca acgccgatag aatctgtacc 60ggcatcacca gcagcaatag ccctcacgtc
gtgaaaacag ctacacaggg cgaagtgaat 120gtgaccggcg tgatccctct gaccacaaca
cctacaaaga gccacttcgc caatctgaag 180ggcacagaga caagaggcaa gctgtgtccc
aagtgcctga attgcacaga tctggatgtg 240gctctgggca gacctaagtg tacaggcaaa
atccctagcg ccagagtgtc cattctgcat 300gaagtgcgac ctgtgaccag cggctgtttt
cctattatgc acgaccggac caagatcaga 360cagctgccta atctgctgag aggctacgag
cacatcagac tgagcaccca caatgtgatc 420aacgccgaaa atgctcctgg cggcccttat
aagatcggca catctggcag ctgccccaac 480attacaaatg gcaatggctt ctttgccacc
atggcttggg ccgtgcctaa gaacgataag 540aacaagaccg ccaccaaccc cctgacaatc
gaggtgccat atatctgtac agagggcgag 600gatcagatca ccgtgtgggg atttcacagc
gacaacgaaa cacagatggc caagctgtac 660ggcgatagca agcctcagaa gtttaccagc
tctgccaatg gcgtgaccac acactatgtg 720tctcagatcg gcggcttccc taatcagaca
gaagatggcg gactgcctca gtctggaaga 780atcgtggtgg attacatggt gcagaagtct
ggcaagaccg gcaccatcac atatcagaga 840ggaatcctgc tgccccagaa agtgtggtgc
gcttctggaa gatccaaagt gatcaagggc 900agcctgcctc tgattggaga agccgattgt
ctgcacgaga aatacggcgg cctgaacaag 960agcaagcctt actatacagg cgagcacgcc
aaggccatcg gcaattgtcc tatttgggtc 1020aagacccctc tgaagctggc caatggcaca
aagtatagac ctccagccaa gctgctgaaa 1080gagagaggct tttttggagc tatcgccggc
tttctggaag gcggatggga gggaatgatt 1140gctggatggc atggctacac atctcatggc
gcacatggcg tggcagtggc tgctgatctg 1200aaatctacac aggaagccat caacaagatc
accaagaacc tgaacagcct gagcgagctg 1260gaagtgaaga atctgcagag actgtctggc
gccatggacg aactgcacaa tgagatcctg 1320gaactggacg agaaggtgga cgatctgaga
gccgatacaa tcagcagcca gattgaactg 1380gctgtgctgc tgtctaacga gggcatcatc
aatagcgagg acgaacatct gctggccctg 1440gaaagaaagc tgaagaagat gctgggacct
agcgccgtgg aaatcggcaa tggatgcttt 1500gagacaaagc acaagtgcaa ccagacctgc
ctggatagaa ttgccgccgg aacatttgat 1560gccggcgagt tttctctgcc caccttcgat
agcctgaata tcaca 160538535PRTInfluenza virus 38Met Lys
Ala Ile Ile Val Leu Leu Met Val Val Thr Ser Asn Ala Asp 1 5
10 15 Arg Ile Cys Thr Gly Ile Thr
Ser Ser Asn Ser Pro His Val Val Lys 20 25
30 Thr Ala Thr Gln Gly Glu Val Asn Val Thr Gly Val
Ile Pro Leu Thr 35 40 45
Thr Thr Pro Thr Lys Ser His Phe Ala Asn Leu Lys Gly Thr Glu Thr
50 55 60 Arg Gly Lys
Leu Cys Pro Lys Cys Leu Asn Cys Thr Asp Leu Asp Val 65
70 75 80 Ala Leu Gly Arg Pro Lys Cys
Thr Gly Lys Ile Pro Ser Ala Arg Val 85
90 95 Ser Ile Leu His Glu Val Arg Pro Val Thr Ser
Gly Cys Phe Pro Ile 100 105
110 Met His Asp Arg Thr Lys Ile Arg Gln Leu Pro Asn Leu Leu Arg
Gly 115 120 125 Tyr
Glu His Ile Arg Leu Ser Thr His Asn Val Ile Asn Ala Glu Asn 130
135 140 Ala Pro Gly Gly Pro Tyr
Lys Ile Gly Thr Ser Gly Ser Cys Pro Asn 145 150
155 160 Ile Thr Asn Gly Asn Gly Phe Phe Ala Thr Met
Ala Trp Ala Val Pro 165 170
175 Lys Asn Asp Lys Asn Lys Thr Ala Thr Asn Pro Leu Thr Ile Glu Val
180 185 190 Pro Tyr
Ile Cys Thr Glu Gly Glu Asp Gln Ile Thr Val Trp Gly Phe 195
200 205 His Ser Asp Asn Glu Thr Gln
Met Ala Lys Leu Tyr Gly Asp Ser Lys 210 215
220 Pro Gln Lys Phe Thr Ser Ser Ala Asn Gly Val Thr
Thr His Tyr Val 225 230 235
240 Ser Gln Ile Gly Gly Phe Pro Asn Gln Thr Glu Asp Gly Gly Leu Pro
245 250 255 Gln Ser Gly
Arg Ile Val Val Asp Tyr Met Val Gln Lys Ser Gly Lys 260
265 270 Thr Gly Thr Ile Thr Tyr Gln Arg
Gly Ile Leu Leu Pro Gln Lys Val 275 280
285 Trp Cys Ala Ser Gly Arg Ser Lys Val Ile Lys Gly Ser
Leu Pro Leu 290 295 300
Ile Gly Glu Ala Asp Cys Leu His Glu Lys Tyr Gly Gly Leu Asn Lys 305
310 315 320 Ser Lys Pro Tyr
Tyr Thr Gly Glu His Ala Lys Ala Ile Gly Asn Cys 325
330 335 Pro Ile Trp Val Lys Thr Pro Leu Lys
Leu Ala Asn Gly Thr Lys Tyr 340 345
350 Arg Pro Pro Ala Lys Leu Leu Lys Glu Arg Gly Phe Phe Gly
Ala Ile 355 360 365
Ala Gly Phe Leu Glu Gly Gly Trp Glu Gly Met Ile Ala Gly Trp His 370
375 380 Gly Tyr Thr Ser His
Gly Ala His Gly Val Ala Val Ala Ala Asp Leu 385 390
395 400 Lys Ser Thr Gln Glu Ala Ile Asn Lys Ile
Thr Lys Asn Leu Asn Ser 405 410
415 Leu Ser Glu Leu Glu Val Lys Asn Leu Gln Arg Leu Ser Gly Ala
Met 420 425 430 Asp
Glu Leu His Asn Glu Ile Leu Glu Leu Asp Glu Lys Val Asp Asp 435
440 445 Leu Arg Ala Asp Thr Ile
Ser Ser Gln Ile Glu Leu Ala Val Leu Leu 450 455
460 Ser Asn Glu Gly Ile Ile Asn Ser Glu Asp Glu
His Leu Leu Ala Leu 465 470 475
480 Glu Arg Lys Leu Lys Lys Met Leu Gly Pro Ser Ala Val Glu Ile Gly
485 490 495 Asn Gly
Cys Phe Glu Thr Lys His Lys Cys Asn Gln Thr Cys Leu Asp 500
505 510 Arg Ile Ala Ala Gly Thr Phe
Asp Ala Gly Glu Phe Ser Leu Pro Thr 515 520
525 Phe Asp Ser Leu Asn Ile Thr 530
535 391605DNAInfluenza virus 39tgtgatattc aggctatcga aggtgggcag
agaaaactcg ccggcatcaa atgttccggc 60ggcaattcta tccaggcagg tctggttgca
cttgtgcttt gtctcaaagc atccattgcc 120gatttccacg gcgctaggtc ccagcatctt
cttcagcttt ctttccaggg ccagcagatg 180ttcgtcctcg ctattgatga tgccctcgtt
agacagcagc acagccagtt caatctggct 240gctgattgta tcggctctca gatcgtccac
cttctcgtcc agttccagga tctcattgtg 300cagttcgtcc atggcgccag acagtctctg
cagattcttc acttccagct cgctcaggct 360gttcaggttc ttggtgatct tgttgatggc
ttcctgtgta gatttcagat cagcagccac 420tgccacgcca tgtgcgccat gagatgtgta
gccatgccat ccagcaatca ttccctccca 480tccgccttcc agaaagccgg cgatagctcc
aaaaaagcct ctctctttca gcagcttggc 540tggaggtcta tactttgtgc cattggccag
cttcagaggg gtcttgaccc aaataggaca 600attgccgatg gccttggcgt gctcgcctgt
atagtaaggc ttgctcttgt tcaggccgcc 660gtatttctcg tgcagacaat cggcttctcc
aatcagaggc aggctgccct tgatcacttt 720ggatcttcca gaagcgcacc acactttctg
gggcagcagg attcctctct gatatgtgat 780ggtgccggtc ttgccagact tctgcaccat
gtaatccacc acgattcttc cagactgagg 840cagtccgcca tcttctgtct gattagggaa
gccgccgatc tgagacacat agtgtgtggt 900cacgccattg gcagagctgg taaacttctg
aggcttgcta tcgccgtaca gcttggccat 960ctgtgtttcg ttgtcgctgt gaaatcccca
cacggtgatc tgatcctcgc cctctgtaca 1020gatatatggc acctcgattg tcagggggtt
ggtggcggtc ttgttcttat cgttcttagg 1080cacggcccaa gccatggtgg caaagaagcc
attgccattt gtaatgttgg ggcagctgcc 1140agatgtgccg atcttataag ggccgccagg
agcattttcg gcgttgatca cattgtgggt 1200gctcagtctg atgtgctcgt agcctctcag
cagattaggc agctgtctga tcttggtccg 1260gtcgtgcata ataggaaaac agccgctggt
cacaggtcgc acttcatgca gaatggacac 1320tctggcgcta gggattttgc ctgtacactt
aggtctgccc agagccacat ccagatctgt 1380gcaattcagg cacttgggac acagcttgcc
tcttgtctct gtgcccttca gattggcgaa 1440gtggctcttt gtaggtgttg tggtcagagg
gatcacgccg gtcacattca cttcgccctg 1500tgtagctgtt ttcacgacgt gagggctatt
gctgctggtg atgccggtac agattctatc 1560ggcgttgctt gtgaccacca tcagcagcac
gatgatggcc ttcat 1605402058DNAArtificial
SequenceSynthetic 40atgaaggcca aactgctggt gctgctgtgt acctttaccg
ccacctacgc cgacacaatc 60tgtatcggct accacgccaa caatagcacc gacaccgtgg
atacagtgct ggagaagaac 120gtgaccgtga cccactctgt gaacctgctg gaggacagcc
acaatggcaa gctgtgtctg 180ctgaaaggca ttgcccctct gcagctgggc aattgttctg
tggccggatg gattctgggc 240aaccccgagt gtgagctgct gatttctaag gagagctgga
gctacatcgt ggagaccccc 300aatcctgaga atggcacctg ctaccctggc tacttcgccg
attacgagga gctgcgcgag 360cagctgtcta gcgtgtccag cttcgagaga ttcgagatct
tccccaagga gtccagctgg 420cctaatcaca cagtgacagg cgtgtctgcc agctgtagcc
acaacggcaa aagcagcttc 480taccggaacc tgctgtggct gacaggcaag aatggcctgt
accccaacct gagcaagagc 540tacgtgaaca acaaggaaaa ggaagtgctg gtgctgtggg
gagtgcacca ccctcccaac 600atcggaaatc agcgggccct gtaccacaca gagaacgcct
atgtgagcgt ggtgtccagc 660cactacagca gaagattcac ccccgagatc gccaagagac
ccaaagtgag agaccaggag 720ggccggatca attactactg gaccctgctg gagcctggcg
ataccatcat cttcgaggcc 780aacggcaatc tgatcgcccc ttggtatgcc tttgccctga
gcagaggctt tggcagcggc 840atcatcacaa gcaacgcccc catggatgag tgtgatgcca
agtgccagac acctcagggc 900gccatcaata gcagcctgcc cttccagaat gtgcaccctg
tgaccatcgg cgagtgcccc 960aagtatgtga gaagcgccaa gctgagaatg gtgaccggcc
tgagaaacat ccctcagagg 1020gagaccagag gactgtttgg agccatcgcc ggattcatcg
agggaggatg gacaggcatg 1080gtggatggct ggtacggcta ccaccaccag aatgagcagg
gctctggata tgccgccgat 1140cagaagtcta cccagaacgc catcaacggc atcaccaaca
aggtgaacag cgtgatcgag 1200aagatgaaca cccagtttac cgctgtgggc aaggagttca
acaagctgga gcggaggatg 1260gagaacctga acaagaaggt ggacgacggc tttctggaca
tctggaccta caatgccgaa 1320ctcctggtcc tcctcgagaa tgagaggacc ctggacttcc
acgacagcaa cgtgaagaac 1380ctgtatgaga aggtgaagag ccagctgaag aacaacgcca
aggagatcgg caacggctgc 1440ttcgagttct accacaagtg taacaacgag tgtatggaga
gcgtgaagaa cggcacctac 1500gactacccta agtacagcga ggagagcaag ctgaaccggg
agaagatcga ttccggaggc 1560gacatcatca agctgctgaa cgagcaggtg aacaaggaga
tgcagagcag caacctgtac 1620atgagcatga gcagctggtg ctacacccac agcctggacg
gcgccggcct gttcctgttc 1680gaccacgccg ccgaggagta cgagcacgcc aagaagctga
tcatcttcct gaacgagaac 1740aacgtgcccg tgcagctgac cagcatcagc gcccccgagc
acaagttcga gggcctgacc 1800cagatcttcc agaaggccta cgagcacgag cagcacatca
gcgagagcat caacaacatc 1860gtggaccacg ccatcaagag caaggaccac gccaccttca
acttcctgca gtggtacgtg 1920gccgagcagc acgaggagga ggtgctgttc aaggacatcc
tggacaagat cgagctgatc 1980ggcaacgaga accacggcct gtacctggcc gaccagtacg
tgaagggcat cgccaagagc 2040aggaagagcg gatcctag
205841685PRTArtificial SequenceSynthetic 41Met Lys
Ala Lys Leu Leu Val Leu Leu Cys Thr Phe Thr Ala Thr Tyr 1 5
10 15 Ala Asp Thr Ile Cys Ile Gly
Tyr His Ala Asn Asn Ser Thr Asp Thr 20 25
30 Val Asp Thr Val Leu Glu Lys Asn Val Thr Val Thr
His Ser Val Asn 35 40 45
Leu Leu Glu Asp Ser His Asn Gly Lys Leu Cys Leu Leu Lys Gly Ile
50 55 60 Ala Pro Leu
Gln Leu Gly Asn Cys Ser Val Ala Gly Trp Ile Leu Gly 65
70 75 80 Asn Pro Glu Cys Glu Leu Leu
Ile Ser Lys Glu Ser Trp Ser Tyr Ile 85
90 95 Val Glu Thr Pro Asn Pro Glu Asn Gly Thr Cys
Tyr Pro Gly Tyr Phe 100 105
110 Ala Asp Tyr Glu Glu Leu Arg Glu Gln Leu Ser Ser Val Ser Ser
Phe 115 120 125 Glu
Arg Phe Glu Ile Phe Pro Lys Glu Ser Ser Trp Pro Asn His Thr 130
135 140 Val Thr Gly Val Ser Ala
Ser Cys Ser His Asn Gly Lys Ser Ser Phe 145 150
155 160 Tyr Arg Asn Leu Leu Trp Leu Thr Gly Lys Asn
Gly Leu Tyr Pro Asn 165 170
175 Leu Ser Lys Ser Tyr Val Asn Asn Lys Glu Lys Glu Val Leu Val Leu
180 185 190 Trp Gly
Val His His Pro Pro Asn Ile Gly Asn Gln Arg Ala Leu Tyr 195
200 205 His Thr Glu Asn Ala Tyr Val
Ser Val Val Ser Ser His Tyr Ser Arg 210 215
220 Arg Phe Thr Pro Glu Ile Ala Lys Arg Pro Lys Val
Arg Asp Gln Glu 225 230 235
240 Gly Arg Ile Asn Tyr Tyr Trp Thr Leu Leu Glu Pro Gly Asp Thr Ile
245 250 255 Ile Phe Glu
Ala Asn Gly Asn Leu Ile Ala Pro Trp Tyr Ala Phe Ala 260
265 270 Leu Ser Arg Gly Phe Gly Ser Gly
Ile Ile Thr Ser Asn Ala Pro Met 275 280
285 Asp Glu Cys Asp Ala Lys Cys Gln Thr Pro Gln Gly Ala
Ile Asn Ser 290 295 300
Ser Leu Pro Phe Gln Asn Val His Pro Val Thr Ile Gly Glu Cys Pro 305
310 315 320 Lys Tyr Val Arg
Ser Ala Lys Leu Arg Met Val Thr Gly Leu Arg Asn 325
330 335 Ile Pro Gln Arg Glu Thr Arg Gly Leu
Phe Gly Ala Ile Ala Gly Phe 340 345
350 Ile Glu Gly Gly Trp Thr Gly Met Val Asp Gly Trp Tyr Gly
Tyr His 355 360 365
His Gln Asn Glu Gln Gly Ser Gly Tyr Ala Ala Asp Gln Lys Ser Thr 370
375 380 Gln Asn Ala Ile Asn
Gly Ile Thr Asn Lys Val Asn Ser Val Ile Glu 385 390
395 400 Lys Met Asn Thr Gln Phe Thr Ala Val Gly
Lys Glu Phe Asn Lys Leu 405 410
415 Glu Arg Arg Met Glu Asn Leu Asn Lys Lys Val Asp Asp Gly Phe
Leu 420 425 430 Asp
Ile Trp Thr Tyr Asn Ala Glu Leu Leu Val Leu Leu Glu Asn Glu 435
440 445 Arg Thr Leu Asp Phe His
Asp Ser Asn Val Lys Asn Leu Tyr Glu Lys 450 455
460 Val Lys Ser Gln Leu Lys Asn Asn Ala Lys Glu
Ile Gly Asn Gly Cys 465 470 475
480 Phe Glu Phe Tyr His Lys Cys Asn Asn Glu Cys Met Glu Ser Val Lys
485 490 495 Asn Gly
Thr Tyr Asp Tyr Pro Lys Tyr Ser Glu Glu Ser Lys Leu Asn 500
505 510 Arg Glu Lys Ile Asp Ser Gly
Gly Asp Ile Ile Lys Leu Leu Asn Glu 515 520
525 Gln Val Asn Lys Glu Met Gln Ser Ser Asn Leu Tyr
Met Ser Met Ser 530 535 540
Ser Trp Cys Tyr Thr His Ser Leu Asp Gly Ala Gly Leu Phe Leu Phe 545
550 555 560 Asp His Ala
Ala Glu Glu Tyr Glu His Ala Lys Lys Leu Ile Ile Phe 565
570 575 Leu Asn Glu Asn Asn Val Pro Val
Gln Leu Thr Ser Ile Ser Ala Pro 580 585
590 Glu His Lys Phe Glu Gly Leu Thr Gln Ile Phe Gln Lys
Ala Tyr Glu 595 600 605
His Glu Gln His Ile Ser Glu Ser Ile Asn Asn Ile Val Asp His Ala 610
615 620 Ile Lys Ser Lys
Asp His Ala Thr Phe Asn Phe Leu Gln Trp Tyr Val 625 630
635 640 Ala Glu Gln His Glu Glu Glu Val Leu
Phe Lys Asp Ile Leu Asp Lys 645 650
655 Ile Glu Leu Ile Gly Asn Glu Asn His Gly Leu Tyr Leu Ala
Asp Gln 660 665 670
Tyr Val Lys Gly Ile Ala Lys Ser Arg Lys Ser Gly Ser 675
680 685 422058DNAArtificial SequenceSynthetic
42ctaggatccg ctcttcctgc tcttggcgat gcccttcacg tactggtcgg ccaggtacag
60gccgtggttc tcgttgccga tcagctcgat cttgtccagg atgtccttga acagcacctc
120ctcctcgtgc tgctcggcca cgtaccactg caggaagttg aaggtggcgt ggtccttgct
180cttgatggcg tggtccacga tgttgttgat gctctcgctg atgtgctgct cgtgctcgta
240ggccttctgg aagatctggg tcaggccctc gaacttgtgc tcgggggcgc tgatgctggt
300cagctgcacg ggcacgttgt tctcgttcag gaagatgatc agcttcttgg cgtgctcgta
360ctcctcggcg gcgtggtcga acaggaacag gccggcgccg tccaggctgt gggtgtagca
420ccagctgctc atgctcatgt acaggttgct gctctgcatc tccttgttca cctgctcgtt
480cagcagcttg atgatgtcgc ctccggaatc gatcttctcc cggttcagct tgctctcctc
540gctgtactta gggtagtcgt aggtgccgtt cttcacgctc tccatacact cgttgttaca
600cttgtggtag aactcgaagc agccgttgcc gatctccttg gcgttgttct tcagctggct
660cttcaccttc tcatacaggt tcttcacgtt gctgtcgtgg aagtccaggg tcctctcatt
720ctcgaggagg accaggagtt cggcattgta ggtccagatg tccagaaagc cgtcgtccac
780cttcttgttc aggttctcca tcctccgctc cagcttgttg aactccttgc ccacagcggt
840aaactgggtg ttcatcttct cgatcacgct gttcaccttg ttggtgatgc cgttgatggc
900gttctgggta gacttctgat cggcggcata tccagagccc tgctcattct ggtggtggta
960gccgtaccag ccatccacca tgcctgtcca tcctccctcg atgaatccgg cgatggctcc
1020aaacagtcct ctggtctccc tctgagggat gtttctcagg ccggtcacca ttctcagctt
1080ggcgcttctc acatacttgg ggcactcgcc gatggtcaca gggtgcacat tctggaaggg
1140caggctgcta ttgatggcgc cctgaggtgt ctggcacttg gcatcacact catccatggg
1200ggcgttgctt gtgatgatgc cgctgccaaa gcctctgctc agggcaaagg cataccaagg
1260ggcgatcaga ttgccgttgg cctcgaagat gatggtatcg ccaggctcca gcagggtcca
1320gtagtaattg atccggccct cctggtctct cactttgggt ctcttggcga tctcgggggt
1380gaatcttctg ctgtagtggc tggacaccac gctcacatag gcgttctctg tgtggtacag
1440ggcccgctga tttccgatgt tgggagggtg gtgcactccc cacagcacca gcacttcctt
1500ttccttgttg ttcacgtagc tcttgctcag gttggggtac aggccattct tgcctgtcag
1560ccacagcagg ttccggtaga agctgctttt gccgttgtgg ctacagctgg cagacacgcc
1620tgtcactgtg tgattaggcc agctggactc cttggggaag atctcgaatc tctcgaagct
1680ggacacgcta gacagctgct cgcgcagctc ctcgtaatcg gcgaagtagc cagggtagca
1740ggtgccattc tcaggattgg gggtctccac gatgtagctc cagctctcct tagaaatcag
1800cagctcacac tcggggttgc ccagaatcca tccggccaca gaacaattgc ccagctgcag
1860aggggcaatg cctttcagca gacacagctt gccattgtgg ctgtcctcca gcaggttcac
1920agagtgggtc acggtcacgt tcttctccag cactgtatcc acggtgtcgg tgctattgtt
1980ggcgtggtag ccgatacaga ttgtgtcggc gtaggtggcg gtaaaggtac acagcagcac
2040cagcagtttg gccttcat
2058432061DNAArtificial SequenceSynthetic 43atgaaggcca tcctggtggt
gctgctgtac accttcgcca ccgccaacgc cgacaccctg 60tgcatcggct accacgccaa
caacagcacc gacaccgtgg acaccgtgct ggagaagaac 120gtgaccgtga cccacagcgt
gaacctgctg gaggacaagc acaacggcaa gctgtgcaag 180ctgcggggcg tggcccccct
gcacctgggc aagtgcaaca tcgccggctg gattctgggc 240aaccccgagt gcgagagcct
gagcaccgcc agcagctgga gctacatcgt ggagaccccc 300agcagcgaca acggcacctg
ctaccccggc gacttcatcg actacgagga gctgcgggag 360cagctgagca gcgtgagcag
cttcgagcgg ttcgagatct tccccaagac cagcagctgg 420cccaaccacg acagcaacaa
gggcgtgacc gccgcctgcc cccacgccgg cgccaagagc 480ttctacaaga acctgatctg
gctggtgaag aagggcaaca gctaccccaa gctgagcaag 540agctacatca acgacaaggg
caaggaggtg ctggtgctgt ggggcatcca ccaccccagc 600accagcgccg accagcagag
cctgtaccag aacgccgaca cctacgtgtt cgtgggcagc 660agccggtaca gcaagaagtt
caagcccgag atcgccatcc ggcccaaggt gcgggaccag 720gagggccgga tgaactacta
ctggaccctg gtggagcccg gcgacaagat caccttcgag 780gccaccggca acctggtggt
gccccggtac gccttcgcca tggagcggaa cgccggcagc 840ggcatcatca tcagcgacac
ccccgtgcac gactgcaaca ccacctgcca gacccccaag 900ggcgccatca acaccagcct
gcccttccag aacatccacc ccatcaccat cggcaagtgc 960cccaagtacg tgaagagcac
caagctgcgg ctggccaccg gcctgcggaa catccccagc 1020atccagagcc ggggcctgtt
cggcgccatc gccggcttca tcgagggcgg ctggaccggc 1080atggtggacg gctggtacgg
ctaccaccac cagaacgagc agggcagcgg ctacgccgcc 1140gacctgaaga gcacccagaa
cgccatcgac gagatcacca acaaggtgaa cagcgtgatc 1200gagaagatga acacccagtt
caccgccgtg ggcaaggagt tcaaccacct ggagaagcgg 1260atcgagaacc tgaacaagaa
ggtggacgac ggcttcctgg acatctggac ctacaacgcc 1320gagctgctgg tgctgctgga
gaacgagcgg accctggact accacgacag caacgtgaag 1380aacctgtacg agaaggtgcg
gagccagctg aagaacaacg ccaaggagat cggcaacggc 1440tgcttcgagt tctaccacaa
gtgcgacaac acctgcatgg agagcgtgaa gaacggcacc 1500tacgactacc ccaagtacag
cgaggaggcc aagctgaacc gggaggagat cgactccgga 1560ggcgacatca tcaagctgct
gaacgagcag gtgaacaagg agatgcagag cagcaacctg 1620tacatgagca tgagcagctg
gtgctacacc cacagcctgg acggcgccgg cctgttcctg 1680ttcgaccacg ccgccgagga
gtacgagcac gccaagaagc tgatcatctt cctgaacgag 1740aacaacgtgc ccgtgcagct
gaccagcatc agcgcccccg agcacaagtt cgagggcctg 1800acccagatct tccagaaggc
ctacgagcac gagcagcaca tcagcgagag catcaacaac 1860atcgtggacc acgccatcaa
gagcaaggac cacgccacct tcaacttcct gcagtggtac 1920gtggccgagc agcacgagga
ggaggtgctg ttcaaggaca tcctggacaa gatcgagctg 1980atcggcaacg agaaccacgg
cctgtacctg gccgaccagt acgtgaaggg catcgccaag 2040agcaggaaga gcggatccta g
206144686PRTArtificial
SequenceSynthetic 44Met Lys Ala Ile Leu Val Val Leu Leu Tyr Thr Phe Ala
Thr Ala Asn 1 5 10 15
Ala Asp Thr Leu Cys Ile Gly Tyr His Ala Asn Asn Ser Thr Asp Thr
20 25 30 Val Asp Thr Val
Leu Glu Lys Asn Val Thr Val Thr His Ser Val Asn 35
40 45 Leu Leu Glu Asp Lys His Asn Gly Lys
Leu Cys Lys Leu Arg Gly Val 50 55
60 Ala Pro Leu His Leu Gly Lys Cys Asn Ile Ala Gly Trp
Ile Leu Gly 65 70 75
80 Asn Pro Glu Cys Glu Ser Leu Ser Thr Ala Ser Ser Trp Ser Tyr Ile
85 90 95 Val Glu Thr Pro
Ser Ser Asp Asn Gly Thr Cys Tyr Pro Gly Asp Phe 100
105 110 Ile Asp Tyr Glu Glu Leu Arg Glu Gln
Leu Ser Ser Val Ser Ser Phe 115 120
125 Glu Arg Phe Glu Ile Phe Pro Lys Thr Ser Ser Trp Pro Asn
His Asp 130 135 140
Ser Asn Lys Gly Val Thr Ala Ala Cys Pro His Ala Gly Ala Lys Ser 145
150 155 160 Phe Tyr Lys Asn Leu
Ile Trp Leu Val Lys Lys Gly Asn Ser Tyr Pro 165
170 175 Lys Leu Ser Lys Ser Tyr Ile Asn Asp Lys
Gly Lys Glu Val Leu Val 180 185
190 Leu Trp Gly Ile His His Pro Ser Thr Ser Ala Asp Gln Gln Ser
Leu 195 200 205 Tyr
Gln Asn Ala Asp Thr Tyr Val Phe Val Gly Ser Ser Arg Tyr Ser 210
215 220 Lys Lys Phe Lys Pro Glu
Ile Ala Ile Arg Pro Lys Val Arg Asp Gln 225 230
235 240 Glu Gly Arg Met Asn Tyr Tyr Trp Thr Leu Val
Glu Pro Gly Asp Lys 245 250
255 Ile Thr Phe Glu Ala Thr Gly Asn Leu Val Val Pro Arg Tyr Ala Phe
260 265 270 Ala Met
Glu Arg Asn Ala Gly Ser Gly Ile Ile Ile Ser Asp Thr Pro 275
280 285 Val His Asp Cys Asn Thr Thr
Cys Gln Thr Pro Lys Gly Ala Ile Asn 290 295
300 Thr Ser Leu Pro Phe Gln Asn Ile His Pro Ile Thr
Ile Gly Lys Cys 305 310 315
320 Pro Lys Tyr Val Lys Ser Thr Lys Leu Arg Leu Ala Thr Gly Leu Arg
325 330 335 Asn Ile Pro
Ser Ile Gln Ser Arg Gly Leu Phe Gly Ala Ile Ala Gly 340
345 350 Phe Ile Glu Gly Gly Trp Thr Gly
Met Val Asp Gly Trp Tyr Gly Tyr 355 360
365 His His Gln Asn Glu Gln Gly Ser Gly Tyr Ala Ala Asp
Leu Lys Ser 370 375 380
Thr Gln Asn Ala Ile Asp Glu Ile Thr Asn Lys Val Asn Ser Val Ile 385
390 395 400 Glu Lys Met Asn
Thr Gln Phe Thr Ala Val Gly Lys Glu Phe Asn His 405
410 415 Leu Glu Lys Arg Ile Glu Asn Leu Asn
Lys Lys Val Asp Asp Gly Phe 420 425
430 Leu Asp Ile Trp Thr Tyr Asn Ala Glu Leu Leu Val Leu Leu
Glu Asn 435 440 445
Glu Arg Thr Leu Asp Tyr His Asp Ser Asn Val Lys Asn Leu Tyr Glu 450
455 460 Lys Val Arg Ser Gln
Leu Lys Asn Asn Ala Lys Glu Ile Gly Asn Gly 465 470
475 480 Cys Phe Glu Phe Tyr His Lys Cys Asp Asn
Thr Cys Met Glu Ser Val 485 490
495 Lys Asn Gly Thr Tyr Asp Tyr Pro Lys Tyr Ser Glu Glu Ala Lys
Leu 500 505 510 Asn
Arg Glu Glu Ile Asp Ser Gly Gly Asp Ile Ile Lys Leu Leu Asn 515
520 525 Glu Gln Val Asn Lys Glu
Met Gln Ser Ser Asn Leu Tyr Met Ser Met 530 535
540 Ser Ser Trp Cys Tyr Thr His Ser Leu Asp Gly
Ala Gly Leu Phe Leu 545 550 555
560 Phe Asp His Ala Ala Glu Glu Tyr Glu His Ala Lys Lys Leu Ile Ile
565 570 575 Phe Leu
Asn Glu Asn Asn Val Pro Val Gln Leu Thr Ser Ile Ser Ala 580
585 590 Pro Glu His Lys Phe Glu Gly
Leu Thr Gln Ile Phe Gln Lys Ala Tyr 595 600
605 Glu His Glu Gln His Ile Ser Glu Ser Ile Asn Asn
Ile Val Asp His 610 615 620
Ala Ile Lys Ser Lys Asp His Ala Thr Phe Asn Phe Leu Gln Trp Tyr 625
630 635 640 Val Ala Glu
Gln His Glu Glu Glu Val Leu Phe Lys Asp Ile Leu Asp 645
650 655 Lys Ile Glu Leu Ile Gly Asn Glu
Asn His Gly Leu Tyr Leu Ala Asp 660 665
670 Gln Tyr Val Lys Gly Ile Ala Lys Ser Arg Lys Ser Gly
Ser 675 680 685
452061DNAArtificial SequenceSynthetic 45ctaggatccg ctcttcctgc tcttggcgat
gcccttcacg tactggtcgg ccaggtacag 60gccgtggttc tcgttgccga tcagctcgat
cttgtccagg atgtccttga acagcacctc 120ctcctcgtgc tgctcggcca cgtaccactg
caggaagttg aaggtggcgt ggtccttgct 180cttgatggcg tggtccacga tgttgttgat
gctctcgctg atgtgctgct cgtgctcgta 240ggccttctgg aagatctggg tcaggccctc
gaacttgtgc tcgggggcgc tgatgctggt 300cagctgcacg ggcacgttgt tctcgttcag
gaagatgatc agcttcttgg cgtgctcgta 360ctcctcggcg gcgtggtcga acaggaacag
gccggcgccg tccaggctgt gggtgtagca 420ccagctgctc atgctcatgt acaggttgct
gctctgcatc tccttgttca cctgctcgtt 480cagcagcttg atgatgtcgc ctccggagtc
gatctcctcc cggttcagct tggcctcctc 540gctgtacttg gggtagtcgt aggtgccgtt
cttcacgctc tccatgcagg tgttgtcgca 600cttgtggtag aactcgaagc agccgttgcc
gatctccttg gcgttgttct tcagctggct 660ccgcaccttc tcgtacaggt tcttcacgtt
gctgtcgtgg tagtccaggg tccgctcgtt 720ctccagcagc accagcagct cggcgttgta
ggtccagatg tccaggaagc cgtcgtccac 780cttcttgttc aggttctcga tccgcttctc
caggtggttg aactccttgc ccacggcggt 840gaactgggtg ttcatcttct cgatcacgct
gttcaccttg ttggtgatct cgtcgatggc 900gttctgggtg ctcttcaggt cggcggcgta
gccgctgccc tgctcgttct ggtggtggta 960gccgtaccag ccgtccacca tgccggtcca
gccgccctcg atgaagccgg cgatggcgcc 1020gaacaggccc cggctctgga tgctggggat
gttccgcagg ccggtggcca gccgcagctt 1080ggtgctcttc acgtacttgg ggcacttgcc
gatggtgatg gggtggatgt tctggaaggg 1140caggctggtg ttgatggcgc ccttgggggt
ctggcaggtg gtgttgcagt cgtgcacggg 1200ggtgtcgctg atgatgatgc cgctgccggc
gttccgctcc atggcgaagg cgtaccgggg 1260caccaccagg ttgccggtgg cctcgaaggt
gatcttgtcg ccgggctcca ccagggtcca 1320gtagtagttc atccggccct cctggtcccg
caccttgggc cggatggcga tctcgggctt 1380gaacttcttg ctgtaccggc tgctgcccac
gaacacgtag gtgtcggcgt tctggtacag 1440gctctgctgg tcggcgctgg tgctggggtg
gtggatgccc cacagcacca gcacctcctt 1500gcccttgtcg ttgatgtagc tcttgctcag
cttggggtag ctgttgccct tcttcaccag 1560ccagatcagg ttcttgtaga agctcttggc
gccggcgtgg gggcaggcgg cggtcacgcc 1620cttgttgctg tcgtggttgg gccagctgct
ggtcttgggg aagatctcga accgctcgaa 1680gctgctcacg ctgctcagct gctcccgcag
ctcctcgtag tcgatgaagt cgccggggta 1740gcaggtgccg ttgtcgctgc tgggggtctc
cacgatgtag ctccagctgc tggcggtgct 1800caggctctcg cactcggggt tgcccagaat
ccagccggcg atgttgcact tgcccaggtg 1860caggggggcc acgccccgca gcttgcacag
cttgccgttg tgcttgtcct ccagcaggtt 1920cacgctgtgg gtcacggtca cgttcttctc
cagcacggtg tccacggtgt cggtgctgtt 1980gttggcgtgg tagccgatgc acagggtgtc
ggcgttggcg gtggcgaagg tgtacagcag 2040caccaccagg atggccttca t
2061462049DNAArtificial
SequenceSynthetic 46atggccatca tctacctgat cctgctgttt acagctgtgc
ggggcgatca gatctgtatc 60ggctaccacg ccaacaatag caccgagaag gtggacacca
tcctggaaag aaatgtgacc 120gtgacccacg ccaaggatat tctggaaaag acccacaacg
gcaagctgtg caagctgaat 180ggcattcctc ctctggaact gggcgattgt tctattgctg
gctggctgct gggaaatcct 240gagtgcgata gactgctgtc tgtgcctgag tggagctaca
tcatggaaaa agagaaccct 300agggacggac tgtgttaccc cggcagcttc aacgattacg
aggaactgaa gcacctgctg 360tccagcgtga agcacttcga gaaagtgaag atcctgccca
aggatagatg gacccagcat 420acaacaacag gcggaagcag agcttgtgct gtgtccggca
accccagctt cttcagaaat 480atggtctggc tgaccaagaa gggctctaat tatcctgtgg
ccaagggcag ctacaataat 540acaagcggcg agcagatgct gattatttgg ggcgtgcacc
accctaatga tgagacagag 600cagagaaccc tgtaccagaa tgtgggcaca tacgtgtctg
tgggcaccag cacactgaat 660aagagaagca cccccgatat tgccaccaga cccaaagtga
atggacaggg cggcagaatg 720gaattttcct ggaccctgct ggatatgtgg gacaccatca
actttgagag caccgggaat 780ctgattgccc ctgagtacgg cttcaagatc agcaagagag
gcagcagcgg catcatgaaa 840acagagggca ccctggaaaa ctgtgaaacc aagtgtcaga
cacctctggg cgccattaat 900accaccctgc ccttccataa tgtgcaccct ctgacaatcg
gcgagtgccc taagtacgtg 960aagtctgaga aactggtgct ggccacagga ctgagaaatg
tgccccagat cgagtcaaga 1020ggcctgtttg gagccattgc cggctttatt gaaggcggat
ggcagggaat ggtggatggg 1080tggtacggct atcaccacag caatgatcag ggatctggct
atgccgccga taaagagagc 1140acccagaagg cctttgacgg catcaccaac aaagtgaaca
gcgtgatcga gaagatgaac 1200acccagtttg aggccgtggg caaagagttc agcaatctgg
aaagacggct ggaaaacctg 1260aacaagaaaa tggaagatgg cttcctggac gtgtggacat
ataatgccga gctgctggtg 1320ctgatggaaa acgagaggac cctggacttt cacgacagca
acgtgaagaa cctgtacgac 1380aaagtgcgga tgcagctgag agacaatgtg aaagagctgg
gcaacggctg ctttgagttc 1440taccacaagt gcgacgacga gtgcatgaat agcgtgaaga
acggcaccta cgactaccct 1500aagtatgagg aagagagcaa gctgaacaga aacgagatca
agtccggagg cgacatcatc 1560aagctgctga acgagcaggt gaacaaggag atgcagagca
gcaacctgta catgagcatg 1620agcagctggt gctacaccca cagcctggac ggcgccggcc
tgttcctgtt cgaccacgcc 1680gccgaggagt acgagcacgc caagaagctg atcatcttcc
tgaacgagaa caacgtgccc 1740gtgcagctga ccagcatcag cgcccccgag cacaagttcg
agggcctgac ccagatcttc 1800cagaaggcct acgagcacga gcagcacatc agcgagagca
tcaacaacat cgtggaccac 1860gccatcaaga gcaaggacca cgccaccttc aacttcctgc
agtggtacgt ggccgagcag 1920cacgaggagg aggtgctgtt caaggacatc ctggacaaga
tcgagctgat cggcaacgag 1980aaccacggcc tgtacctggc cgaccagtac gtgaagggca
tcgccaagag caggaagagc 2040ggatcctag
204947682PRTArtificial SequenceSynthetic 47Met Ala
Ile Ile Tyr Leu Ile Leu Leu Phe Thr Ala Val Arg Gly Asp 1 5
10 15 Gln Ile Cys Ile Gly Tyr His
Ala Asn Asn Ser Thr Glu Lys Val Asp 20 25
30 Thr Ile Leu Glu Arg Asn Val Thr Val Thr His Ala
Lys Asp Ile Leu 35 40 45
Glu Lys Thr His Asn Gly Lys Leu Cys Lys Leu Asn Gly Ile Pro Pro
50 55 60 Leu Glu Leu
Gly Asp Cys Ser Ile Ala Gly Trp Leu Leu Gly Asn Pro 65
70 75 80 Glu Cys Asp Arg Leu Leu Ser
Val Pro Glu Trp Ser Tyr Ile Met Glu 85
90 95 Lys Glu Asn Pro Arg Asp Gly Leu Cys Tyr Pro
Gly Ser Phe Asn Asp 100 105
110 Tyr Glu Glu Leu Lys His Leu Leu Ser Ser Val Lys His Phe Glu
Lys 115 120 125 Val
Lys Ile Leu Pro Lys Asp Arg Trp Thr Gln His Thr Thr Thr Gly 130
135 140 Gly Ser Arg Ala Cys Ala
Val Ser Gly Asn Pro Ser Phe Phe Arg Asn 145 150
155 160 Met Val Trp Leu Thr Lys Lys Gly Ser Asn Tyr
Pro Val Ala Lys Gly 165 170
175 Ser Tyr Asn Asn Thr Ser Gly Glu Gln Met Leu Ile Ile Trp Gly Val
180 185 190 His His
Pro Asn Asp Glu Thr Glu Gln Arg Thr Leu Tyr Gln Asn Val 195
200 205 Gly Thr Tyr Val Ser Val Gly
Thr Ser Thr Leu Asn Lys Arg Ser Thr 210 215
220 Pro Asp Ile Ala Thr Arg Pro Lys Val Asn Gly Gln
Gly Gly Arg Met 225 230 235
240 Glu Phe Ser Trp Thr Leu Leu Asp Met Trp Asp Thr Ile Asn Phe Glu
245 250 255 Ser Thr Gly
Asn Leu Ile Ala Pro Glu Tyr Gly Phe Lys Ile Ser Lys 260
265 270 Arg Gly Ser Ser Gly Ile Met Lys
Thr Glu Gly Thr Leu Glu Asn Cys 275 280
285 Glu Thr Lys Cys Gln Thr Pro Leu Gly Ala Ile Asn Thr
Thr Leu Pro 290 295 300
Phe His Asn Val His Pro Leu Thr Ile Gly Glu Cys Pro Lys Tyr Val 305
310 315 320 Lys Ser Glu Lys
Leu Val Leu Ala Thr Gly Leu Arg Asn Val Pro Gln 325
330 335 Ile Glu Ser Arg Gly Leu Phe Gly Ala
Ile Ala Gly Phe Ile Glu Gly 340 345
350 Gly Trp Gln Gly Met Val Asp Gly Trp Tyr Gly Tyr His His
Ser Asn 355 360 365
Asp Gln Gly Ser Gly Tyr Ala Ala Asp Lys Glu Ser Thr Gln Lys Ala 370
375 380 Phe Asp Gly Ile Thr
Asn Lys Val Asn Ser Val Ile Glu Lys Met Asn 385 390
395 400 Thr Gln Phe Glu Ala Val Gly Lys Glu Phe
Ser Asn Leu Glu Arg Arg 405 410
415 Leu Glu Asn Leu Asn Lys Lys Met Glu Asp Gly Phe Leu Asp Val
Trp 420 425 430 Thr
Tyr Asn Ala Glu Leu Leu Val Leu Met Glu Asn Glu Arg Thr Leu 435
440 445 Asp Phe His Asp Ser Asn
Val Lys Asn Leu Tyr Asp Lys Val Arg Met 450 455
460 Gln Leu Arg Asp Asn Val Lys Glu Leu Gly Asn
Gly Cys Phe Glu Phe 465 470 475
480 Tyr His Lys Cys Asp Asp Glu Cys Met Asn Ser Val Lys Asn Gly Thr
485 490 495 Tyr Asp
Tyr Pro Lys Tyr Glu Glu Glu Ser Lys Leu Asn Arg Asn Glu 500
505 510 Ile Lys Ser Gly Gly Asp Ile
Ile Lys Leu Leu Asn Glu Gln Val Asn 515 520
525 Lys Glu Met Gln Ser Ser Asn Leu Tyr Met Ser Met
Ser Ser Trp Cys 530 535 540
Tyr Thr His Ser Leu Asp Gly Ala Gly Leu Phe Leu Phe Asp His Ala 545
550 555 560 Ala Glu Glu
Tyr Glu His Ala Lys Lys Leu Ile Ile Phe Leu Asn Glu 565
570 575 Asn Asn Val Pro Val Gln Leu Thr
Ser Ile Ser Ala Pro Glu His Lys 580 585
590 Phe Glu Gly Leu Thr Gln Ile Phe Gln Lys Ala Tyr Glu
His Glu Gln 595 600 605
His Ile Ser Glu Ser Ile Asn Asn Ile Val Asp His Ala Ile Lys Ser 610
615 620 Lys Asp His Ala
Thr Phe Asn Phe Leu Gln Trp Tyr Val Ala Glu Gln 625 630
635 640 His Glu Glu Glu Val Leu Phe Lys Asp
Ile Leu Asp Lys Ile Glu Leu 645 650
655 Ile Gly Asn Glu Asn His Gly Leu Tyr Leu Ala Asp Gln Tyr
Val Lys 660 665 670
Gly Ile Ala Lys Ser Arg Lys Ser Gly Ser 675 680
482049DNAArtificial SequenceSynthetic 48ctaggatccg ctcttcctgc
tcttggcgat gcccttcacg tactggtcgg ccaggtacag 60gccgtggttc tcgttgccga
tcagctcgat cttgtccagg atgtccttga acagcacctc 120ctcctcgtgc tgctcggcca
cgtaccactg caggaagttg aaggtggcgt ggtccttgct 180cttgatggcg tggtccacga
tgttgttgat gctctcgctg atgtgctgct cgtgctcgta 240ggccttctgg aagatctggg
tcaggccctc gaacttgtgc tcgggggcgc tgatgctggt 300cagctgcacg ggcacgttgt
tctcgttcag gaagatgatc agcttcttgg cgtgctcgta 360ctcctcggcg gcgtggtcga
acaggaacag gccggcgccg tccaggctgt gggtgtagca 420ccagctgctc atgctcatgt
acaggttgct gctctgcatc tccttgttca cctgctcgtt 480cagcagcttg atgatgtcgc
ctccggactt gatctcgttt ctgttcagct tgctctcttc 540ctcatactta gggtagtcgt
aggtgccgtt cttcacgcta ttcatgcact cgtcgtcgca 600cttgtggtag aactcaaagc
agccgttgcc cagctctttc acattgtctc tcagctgcat 660ccgcactttg tcgtacaggt
tcttcacgtt gctgtcgtga aagtccaggg tcctctcgtt 720ttccatcagc accagcagct
cggcattata tgtccacacg tccaggaagc catcttccat 780tttcttgttc aggttttcca
gccgtctttc cagattgctg aactctttgc ccacggcctc 840aaactgggtg ttcatcttct
cgatcacgct gttcactttg ttggtgatgc cgtcaaaggc 900cttctgggtg ctctctttat
cggcggcata gccagatccc tgatcattgc tgtggtgata 960gccgtaccac ccatccacca
ttccctgcca tccgccttca ataaagccgg caatggctcc 1020aaacaggcct cttgactcga
tctggggcac atttctcagt cctgtggcca gcaccagttt 1080ctcagacttc acgtacttag
ggcactcgcc gattgtcaga gggtgcacat tatggaaggg 1140cagggtggta ttaatggcgc
ccagaggtgt ctgacacttg gtttcacagt tttccagggt 1200gccctctgtt ttcatgatgc
cgctgctgcc tctcttgctg atcttgaagc cgtactcagg 1260ggcaatcaga ttcccggtgc
tctcaaagtt gatggtgtcc cacatatcca gcagggtcca 1320ggaaaattcc attctgccgc
cctgtccatt cactttgggt ctggtggcaa tatcgggggt 1380gcttctctta ttcagtgtgc
tggtgcccac agacacgtat gtgcccacat tctggtacag 1440ggttctctgc tctgtctcat
cattagggtg gtgcacgccc caaataatca gcatctgctc 1500gccgcttgta ttattgtagc
tgcccttggc cacaggataa ttagagccct tcttggtcag 1560ccagaccata tttctgaaga
agctggggtt gccggacaca gcacaagctc tgcttccgcc 1620tgttgttgta tgctgggtcc
atctatcctt gggcaggatc ttcactttct cgaagtgctt 1680cacgctggac agcaggtgct
tcagttcctc gtaatcgttg aagctgccgg ggtaacacag 1740tccgtcccta gggttctctt
tttccatgat gtagctccac tcaggcacag acagcagtct 1800atcgcactca ggatttccca
gcagccagcc agcaatagaa caatcgccca gttccagagg 1860aggaatgcca ttcagcttgc
acagcttgcc gttgtgggtc ttttccagaa tatccttggc 1920gtgggtcacg gtcacatttc
tttccaggat ggtgtccacc ttctcggtgc tattgttggc 1980gtggtagccg atacagatct
gatcgccccg cacagctgta aacagcagga tcaggtagat 2040gatggccat
2049492064DNAArtificial
SequenceSynthetic 49atgaaaacca tcattgccct gagctacatc ttttgtctgg
ctctgggcca ggatctgccc 60ggcaatgata atagcaccgc caccctgtgt ctgggacacc
acgccgtgcc taatggcacc 120ctggtgaaaa ccattaccga cgaccagatc gaagtgacca
atgccaccga gctggtgcag 180agcagcagca ccggcaagat ctgcaacaac ccccacagaa
tcctggatgg catcgactgt 240accctgatcg atgccctgct gggcgatcct cactgcgacg
tgttccagaa cgagacatgg 300gacctgttcg tggagagaag caaggccttc agcaactgct
acccctacga tgtgcccgat 360tacgcctctc tgagaagcct ggtggccagc agcggcacac
tggaattcat caccgagggc 420tttacctgga caggcgtgac ccagaatggc ggcagcaatg
cctgtaaaag aggccctggc 480agcggcttct tcagcagact gaactggctg accaagtccg
gcagcaccta ccctgtgctg 540aacgtgacca tgcccaacaa cgacaacttc gacaagctgt
acatctgggg cgtgcaccac 600cctagcacca atcaggaaca gaccagcctg tacgtgcagg
ccagcggcag agtgaccgtg 660tctaccagac ggtcccagca gaccatcatc cccaacatcg
agtcaagacc ttgggtgcgc 720ggcctgagca gcagaatcag catctactgg accatcgtga
aacctggcga cgtgctggtg 780atcaacagca atggcaacct gatcgccccc agaggctact
tcaagatgcg gaccggcaag 840agcagcatca tgagaagcga cgcccccatc gatacctgta
tcagcgagtg catcaccccc 900aacggcagca tccccaacga caagcccttc cagaacgtga
acaagatcac ctacggcgcc 960tgccctaagt acgtgaagca gaacaccctg aagctggcca
ccggcatgag aaatgtgccc 1020gagaagcaga caagaggcct gtttggcgcc attgccggct
ttatcgagaa cggctgggag 1080ggcatgatcg atgggtggta cggcttcaga caccagaatt
ctgagggcac aggacaggcc 1140gccgatctga agtctacaca ggccgccatc gaccagatca
acggcaagct gaacagagtg 1200atcgagaaaa ccaacgagaa gttccaccag atcgagaaag
aattcagcga ggtggagggc 1260agaatccagg acctggaaaa atacgtggag gacaccaaga
tcgacctgtg gagctacaat 1320gccgaactgc tggtcgccct ggaaaaccag cacaccatcg
acctgaccga cagcgagatg 1380aataagctgt tcgaaaagac cagacggcag ctgagagaaa
acgccgagga catgggcaac 1440ggctgcttca agatctacca caagtgcgac aacgcctgca
tcgagagcat cagaaacggc 1500acctacgacc acgatgtgta cagggacgag gccctgaaca
acagattcca gatcaagtcc 1560ggaggcgaca tcatcaagct gctgaacgag caggtgaaca
aggagatgca gagcagcaac 1620ctgtacatga gcatgagcag ctggtgctac acccacagcc
tggacggcgc cggcctgttc 1680ctgttcgacc acgccgccga ggagtacgag cacgccaaga
agctgatcat cttcctgaac 1740gagaacaacg tgcccgtgca gctgaccagc atcagcgccc
ccgagcacaa gttcgagggc 1800ctgacccaga tcttccagaa ggcctacgag cacgagcagc
acatcagcga gagcatcaac 1860aacatcgtgg accacgccat caagagcaag gaccacgcca
ccttcaactt cctgcagtgg 1920tacgtggccg agcagcacga ggaggaggtg ctgttcaagg
acatcctgga caagatcgag 1980ctgatcggca acgagaacca cggcctgtac ctggccgacc
agtacgtgaa gggcatcgcc 2040aagagcagga agagcggatc ctag
206450687PRTArtificial SequenceSynthetic 50Met Lys
Thr Ile Ile Ala Leu Ser Tyr Ile Phe Cys Leu Ala Leu Gly 1 5
10 15 Gln Asp Leu Pro Gly Asn Asp
Asn Ser Thr Ala Thr Leu Cys Leu Gly 20 25
30 His His Ala Val Pro Asn Gly Thr Leu Val Lys Thr
Ile Thr Asp Asp 35 40 45
Gln Ile Glu Val Thr Asn Ala Thr Glu Leu Val Gln Ser Ser Ser Thr
50 55 60 Gly Lys Ile
Cys Asn Asn Pro His Arg Ile Leu Asp Gly Ile Asp Cys 65
70 75 80 Thr Leu Ile Asp Ala Leu Leu
Gly Asp Pro His Cys Asp Val Phe Gln 85
90 95 Asn Glu Thr Trp Asp Leu Phe Val Glu Arg Ser
Lys Ala Phe Ser Asn 100 105
110 Cys Tyr Pro Tyr Asp Val Pro Asp Tyr Ala Ser Leu Arg Ser Leu
Val 115 120 125 Ala
Ser Ser Gly Thr Leu Glu Phe Ile Thr Glu Gly Phe Thr Trp Thr 130
135 140 Gly Val Thr Gln Asn Gly
Gly Ser Asn Ala Cys Lys Arg Gly Pro Gly 145 150
155 160 Ser Gly Phe Phe Ser Arg Leu Asn Trp Leu Thr
Lys Ser Gly Ser Thr 165 170
175 Tyr Pro Val Leu Asn Val Thr Met Pro Asn Asn Asp Asn Phe Asp Lys
180 185 190 Leu Tyr
Ile Trp Gly Val His His Pro Ser Thr Asn Gln Glu Gln Thr 195
200 205 Ser Leu Tyr Val Gln Ala Ser
Gly Arg Val Thr Val Ser Thr Arg Arg 210 215
220 Ser Gln Gln Thr Ile Ile Pro Asn Ile Glu Ser Arg
Pro Trp Val Arg 225 230 235
240 Gly Leu Ser Ser Arg Ile Ser Ile Tyr Trp Thr Ile Val Lys Pro Gly
245 250 255 Asp Val Leu
Val Ile Asn Ser Asn Gly Asn Leu Ile Ala Pro Arg Gly 260
265 270 Tyr Phe Lys Met Arg Thr Gly Lys
Ser Ser Ile Met Arg Ser Asp Ala 275 280
285 Pro Ile Asp Thr Cys Ile Ser Glu Cys Ile Thr Pro Asn
Gly Ser Ile 290 295 300
Pro Asn Asp Lys Pro Phe Gln Asn Val Asn Lys Ile Thr Tyr Gly Ala 305
310 315 320 Cys Pro Lys Tyr
Val Lys Gln Asn Thr Leu Lys Leu Ala Thr Gly Met 325
330 335 Arg Asn Val Pro Glu Lys Gln Thr Arg
Gly Leu Phe Gly Ala Ile Ala 340 345
350 Gly Phe Ile Glu Asn Gly Trp Glu Gly Met Ile Asp Gly Trp
Tyr Gly 355 360 365
Phe Arg His Gln Asn Ser Glu Gly Thr Gly Gln Ala Ala Asp Leu Lys 370
375 380 Ser Thr Gln Ala Ala
Ile Asp Gln Ile Asn Gly Lys Leu Asn Arg Val 385 390
395 400 Ile Glu Lys Thr Asn Glu Lys Phe His Gln
Ile Glu Lys Glu Phe Ser 405 410
415 Glu Val Glu Gly Arg Ile Gln Asp Leu Glu Lys Tyr Val Glu Asp
Thr 420 425 430 Lys
Ile Asp Leu Trp Ser Tyr Asn Ala Glu Leu Leu Val Ala Leu Glu 435
440 445 Asn Gln His Thr Ile Asp
Leu Thr Asp Ser Glu Met Asn Lys Leu Phe 450 455
460 Glu Lys Thr Arg Arg Gln Leu Arg Glu Asn Ala
Glu Asp Met Gly Asn 465 470 475
480 Gly Cys Phe Lys Ile Tyr His Lys Cys Asp Asn Ala Cys Ile Glu Ser
485 490 495 Ile Arg
Asn Gly Thr Tyr Asp His Asp Val Tyr Arg Asp Glu Ala Leu 500
505 510 Asn Asn Arg Phe Gln Ile Lys
Ser Gly Gly Asp Ile Ile Lys Leu Leu 515 520
525 Asn Glu Gln Val Asn Lys Glu Met Gln Ser Ser Asn
Leu Tyr Met Ser 530 535 540
Met Ser Ser Trp Cys Tyr Thr His Ser Leu Asp Gly Ala Gly Leu Phe 545
550 555 560 Leu Phe Asp
His Ala Ala Glu Glu Tyr Glu His Ala Lys Lys Leu Ile 565
570 575 Ile Phe Leu Asn Glu Asn Asn Val
Pro Val Gln Leu Thr Ser Ile Ser 580 585
590 Ala Pro Glu His Lys Phe Glu Gly Leu Thr Gln Ile Phe
Gln Lys Ala 595 600 605
Tyr Glu His Glu Gln His Ile Ser Glu Ser Ile Asn Asn Ile Val Asp 610
615 620 His Ala Ile Lys
Ser Lys Asp His Ala Thr Phe Asn Phe Leu Gln Trp 625 630
635 640 Tyr Val Ala Glu Gln His Glu Glu Glu
Val Leu Phe Lys Asp Ile Leu 645 650
655 Asp Lys Ile Glu Leu Ile Gly Asn Glu Asn His Gly Leu Tyr
Leu Ala 660 665 670
Asp Gln Tyr Val Lys Gly Ile Ala Lys Ser Arg Lys Ser Gly Ser 675
680 685 512064DNAArtificial
SequenceSynthetic 51ctaggatccg ctcttcctgc tcttggcgat gcccttcacg
tactggtcgg ccaggtacag 60gccgtggttc tcgttgccga tcagctcgat cttgtccagg
atgtccttga acagcacctc 120ctcctcgtgc tgctcggcca cgtaccactg caggaagttg
aaggtggcgt ggtccttgct 180cttgatggcg tggtccacga tgttgttgat gctctcgctg
atgtgctgct cgtgctcgta 240ggccttctgg aagatctggg tcaggccctc gaacttgtgc
tcgggggcgc tgatgctggt 300cagctgcacg ggcacgttgt tctcgttcag gaagatgatc
agcttcttgg cgtgctcgta 360ctcctcggcg gcgtggtcga acaggaacag gccggcgccg
tccaggctgt gggtgtagca 420ccagctgctc atgctcatgt acaggttgct gctctgcatc
tccttgttca cctgctcgtt 480cagcagcttg atgatgtcgc ctccggactt gatctggaat
ctgttgttca gggcctcgtc 540cctgtacaca tcgtggtcgt aggtgccgtt tctgatgctc
tcgatgcagg cgttgtcgca 600cttgtggtag atcttgaagc agccgttgcc catgtcctcg
gcgttttctc tcagctgccg 660tctggtcttt tcgaacagct tattcatctc gctgtcggtc
aggtcgatgg tgtgctggtt 720ttccagggcg accagcagtt cggcattgta gctccacagg
tcgatcttgg tgtcctccac 780gtatttttcc aggtcctgga ttctgccctc cacctcgctg
aattctttct cgatctggtg 840gaacttctcg ttggttttct cgatcactct gttcagcttg
ccgttgatct ggtcgatggc 900ggcctgtgta gacttcagat cggcggcctg tcctgtgccc
tcagaattct ggtgtctgaa 960gccgtaccac ccatcgatca tgccctccca gccgttctcg
ataaagccgg caatggcgcc 1020aaacaggcct cttgtctgct tctcgggcac atttctcatg
ccggtggcca gcttcagggt 1080gttctgcttc acgtacttag ggcaggcgcc gtaggtgatc
ttgttcacgt tctggaaggg 1140cttgtcgttg gggatgctgc cgttgggggt gatgcactcg
ctgatacagg tatcgatggg 1200ggcgtcgctt ctcatgatgc tgctcttgcc ggtccgcatc
ttgaagtagc ctctgggggc 1260gatcaggttg ccattgctgt tgatcaccag cacgtcgcca
ggtttcacga tggtccagta 1320gatgctgatt ctgctgctca ggccgcgcac ccaaggtctt
gactcgatgt tggggatgat 1380ggtctgctgg gaccgtctgg tagacacggt cactctgccg
ctggcctgca cgtacaggct 1440ggtctgttcc tgattggtgc tagggtggtg cacgccccag
atgtacagct tgtcgaagtt 1500gtcgttgttg ggcatggtca cgttcagcac agggtaggtg
ctgccggact tggtcagcca 1560gttcagtctg ctgaagaagc cgctgccagg gcctctttta
caggcattgc tgccgccatt 1620ctgggtcacg cctgtccagg taaagccctc ggtgatgaat
tccagtgtgc cgctgctggc 1680caccaggctt ctcagagagg cgtaatcggg cacatcgtag
gggtagcagt tgctgaaggc 1740cttgcttctc tccacgaaca ggtcccatgt ctcgttctgg
aacacgtcgc agtgaggatc 1800gcccagcagg gcatcgatca gggtacagtc gatgccatcc
aggattctgt gggggttgtt 1860gcagatcttg ccggtgctgc tgctctgcac cagctcggtg
gcattggtca cttcgatctg 1920gtcgtcggta atggttttca ccagggtgcc attaggcacg
gcgtggtgtc ccagacacag 1980ggtggcggtg ctattatcat tgccgggcag atcctggccc
agagccagac aaaagatgta 2040gctcagggca atgatggttt tcat
2064522064DNAArtificial SequenceSynthetic
52atgaaaacca tcattgccct gagctacatc ctgtgcctgg tgttcacaca gaagctgccc
60ggcaacgata atagcaccgc cacactgtgt ctgggacacc acgccgtgcc taatggcacc
120atcgtgaaaa caatcaccaa cgaccagatc gaagtgacca atgccacaga gctggtgcag
180agcagcagca caggcgagat ctgtgacagc ccccaccaga tcctggatgg cgagaactgt
240accctgatcg atgccctgct gggcgatcct cagtgcgacg gcttccagaa caagaaatgg
300gacctgttcg tggagagaag caaggcctac agcaactgct acccctacga cgtgcctgat
360tacgccagcc tgagaagcct ggtggcctct agcggcaccc tggaattcaa caacgagagc
420ttcaactgga ccggcgtgac acagaatggc accagcagcg cctgcatcag acggtccaac
480aacagcttct tcagtagact gaattggctg acccacctga agttcaagta ccccgccctg
540aacgtgacca tgcccaacaa tgagaagttc gacaagctgt acatctgggg agtgcaccac
600cctggcaccg acaacgatca gatcttccct tacgcccagg ccagcggcag aatcaccgtg
660tccaccaaga gaagccagca gaccgtgatc cccaatatcg gcagcagacc cagagtgcgg
720aacatcccca gcaggatcag catctactgg acaatcgtga agcctggcga catcctgctg
780atcaacagca ccggcaacct gatcgcccct cggggctact ttaagatcag aagcggcaag
840agcagcatca tgagatccga cgcccccatc ggcaagtgca acagcgagtg catcacccca
900aacggcagca tccccaacga caagcccttc cagaacgtga acaggatcac ctacggcgcc
960tgccctagat acgtgaagca gaacaccctg aagctggcca ccggcatgag aaatgtgccc
1020gagaagcaga ccagaggcat ctttggcgcc attgccggct ttatcgagaa tggctgggag
1080ggaatggtgg atgggtggta cggcttcaga caccagaata gcgagggaat tggacaggcc
1140gccgatctga aatctaccca ggccgccatc gaccagatca acggcaagct gaacaggctg
1200atcggcaaga ccaacgagaa gttccaccag atcgagaaag aattcagcga ggtggagggc
1260agaatccagg acctggaaaa atacgtggag gacaccaaga tcgacctgtg gagctacaat
1320gccgaactgc tggtcgccct ggaaaaccag cacacaattg atctgacaga cagtgagatg
1380aataagctgt tcgagaaaac caagaagcag ctgagagaaa acgccgagga catgggcaac
1440ggctgcttca agatctacca caagtgcgac aacgcctgca tcggcagcat cagaaacggc
1500acctacgacc acgacgtgta cagagatgag gccctgaaca accggtttca gatcaagtcc
1560ggaggcgaca tcatcaagct gctgaacgag caggtgaaca aggagatgca gagcagcaac
1620ctgtacatga gcatgagcag ctggtgctac acccacagcc tggacggcgc cggcctgttc
1680ctgttcgacc acgccgccga ggagtacgag cacgccaaga agctgatcat cttcctgaac
1740gagaacaacg tgcccgtgca gctgaccagc atcagcgccc ccgagcacaa gttcgagggc
1800ctgacccaga tcttccagaa ggcctacgag cacgagcagc acatcagcga gagcatcaac
1860aacatcgtgg accacgccat caagagcaag gaccacgcca ccttcaactt cctgcagtgg
1920tacgtggccg agcagcacga ggaggaggtg ctgttcaagg acatcctgga caagatcgag
1980ctgatcggca acgagaacca cggcctgtac ctggccgacc agtacgtgaa gggcatcgcc
2040aagagcagga agagcggatc ctag
206453687PRTArtificial SequenceSynthetic 53Met Lys Thr Ile Ile Ala Leu
Ser Tyr Ile Leu Cys Leu Val Phe Thr 1 5
10 15 Gln Lys Leu Pro Gly Asn Asp Asn Ser Thr Ala
Thr Leu Cys Leu Gly 20 25
30 His His Ala Val Pro Asn Gly Thr Ile Val Lys Thr Ile Thr Asn
Asp 35 40 45 Gln
Ile Glu Val Thr Asn Ala Thr Glu Leu Val Gln Ser Ser Ser Thr 50
55 60 Gly Glu Ile Cys Asp Ser
Pro His Gln Ile Leu Asp Gly Glu Asn Cys 65 70
75 80 Thr Leu Ile Asp Ala Leu Leu Gly Asp Pro Gln
Cys Asp Gly Phe Gln 85 90
95 Asn Lys Lys Trp Asp Leu Phe Val Glu Arg Ser Lys Ala Tyr Ser Asn
100 105 110 Cys Tyr
Pro Tyr Asp Val Pro Asp Tyr Ala Ser Leu Arg Ser Leu Val 115
120 125 Ala Ser Ser Gly Thr Leu Glu
Phe Asn Asn Glu Ser Phe Asn Trp Thr 130 135
140 Gly Val Thr Gln Asn Gly Thr Ser Ser Ala Cys Ile
Arg Arg Ser Asn 145 150 155
160 Asn Ser Phe Phe Ser Arg Leu Asn Trp Leu Thr His Leu Lys Phe Lys
165 170 175 Tyr Pro Ala
Leu Asn Val Thr Met Pro Asn Asn Glu Lys Phe Asp Lys 180
185 190 Leu Tyr Ile Trp Gly Val His His
Pro Gly Thr Asp Asn Asp Gln Ile 195 200
205 Phe Pro Tyr Ala Gln Ala Ser Gly Arg Ile Thr Val Ser
Thr Lys Arg 210 215 220
Ser Gln Gln Thr Val Ile Pro Asn Ile Gly Ser Arg Pro Arg Val Arg 225
230 235 240 Asn Ile Pro Ser
Arg Ile Ser Ile Tyr Trp Thr Ile Val Lys Pro Gly 245
250 255 Asp Ile Leu Leu Ile Asn Ser Thr Gly
Asn Leu Ile Ala Pro Arg Gly 260 265
270 Tyr Phe Lys Ile Arg Ser Gly Lys Ser Ser Ile Met Arg Ser
Asp Ala 275 280 285
Pro Ile Gly Lys Cys Asn Ser Glu Cys Ile Thr Pro Asn Gly Ser Ile 290
295 300 Pro Asn Asp Lys Pro
Phe Gln Asn Val Asn Arg Ile Thr Tyr Gly Ala 305 310
315 320 Cys Pro Arg Tyr Val Lys Gln Asn Thr Leu
Lys Leu Ala Thr Gly Met 325 330
335 Arg Asn Val Pro Glu Lys Gln Thr Arg Gly Ile Phe Gly Ala Ile
Ala 340 345 350 Gly
Phe Ile Glu Asn Gly Trp Glu Gly Met Val Asp Gly Trp Tyr Gly 355
360 365 Phe Arg His Gln Asn Ser
Glu Gly Ile Gly Gln Ala Ala Asp Leu Lys 370 375
380 Ser Thr Gln Ala Ala Ile Asp Gln Ile Asn Gly
Lys Leu Asn Arg Leu 385 390 395
400 Ile Gly Lys Thr Asn Glu Lys Phe His Gln Ile Glu Lys Glu Phe Ser
405 410 415 Glu Val
Glu Gly Arg Ile Gln Asp Leu Glu Lys Tyr Val Glu Asp Thr 420
425 430 Lys Ile Asp Leu Trp Ser Tyr
Asn Ala Glu Leu Leu Val Ala Leu Glu 435 440
445 Asn Gln His Thr Ile Asp Leu Thr Asp Ser Glu Met
Asn Lys Leu Phe 450 455 460
Glu Lys Thr Lys Lys Gln Leu Arg Glu Asn Ala Glu Asp Met Gly Asn 465
470 475 480 Gly Cys Phe
Lys Ile Tyr His Lys Cys Asp Asn Ala Cys Ile Gly Ser 485
490 495 Ile Arg Asn Gly Thr Tyr Asp His
Asp Val Tyr Arg Asp Glu Ala Leu 500 505
510 Asn Asn Arg Phe Gln Ile Lys Ser Gly Gly Asp Ile Ile
Lys Leu Leu 515 520 525
Asn Glu Gln Val Asn Lys Glu Met Gln Ser Ser Asn Leu Tyr Met Ser 530
535 540 Met Ser Ser Trp
Cys Tyr Thr His Ser Leu Asp Gly Ala Gly Leu Phe 545 550
555 560 Leu Phe Asp His Ala Ala Glu Glu Tyr
Glu His Ala Lys Lys Leu Ile 565 570
575 Ile Phe Leu Asn Glu Asn Asn Val Pro Val Gln Leu Thr Ser
Ile Ser 580 585 590
Ala Pro Glu His Lys Phe Glu Gly Leu Thr Gln Ile Phe Gln Lys Ala
595 600 605 Tyr Glu His Glu
Gln His Ile Ser Glu Ser Ile Asn Asn Ile Val Asp 610
615 620 His Ala Ile Lys Ser Lys Asp His
Ala Thr Phe Asn Phe Leu Gln Trp 625 630
635 640 Tyr Val Ala Glu Gln His Glu Glu Glu Val Leu Phe
Lys Asp Ile Leu 645 650
655 Asp Lys Ile Glu Leu Ile Gly Asn Glu Asn His Gly Leu Tyr Leu Ala
660 665 670 Asp Gln Tyr
Val Lys Gly Ile Ala Lys Ser Arg Lys Ser Gly Ser 675
680 685 542064DNAArtificial SequenceSynthetic
54ctaggatccg ctcttcctgc tcttggcgat gcccttcacg tactggtcgg ccaggtacag
60gccgtggttc tcgttgccga tcagctcgat cttgtccagg atgtccttga acagcacctc
120ctcctcgtgc tgctcggcca cgtaccactg caggaagttg aaggtggcgt ggtccttgct
180cttgatggcg tggtccacga tgttgttgat gctctcgctg atgtgctgct cgtgctcgta
240ggccttctgg aagatctggg tcaggccctc gaacttgtgc tcgggggcgc tgatgctggt
300cagctgcacg ggcacgttgt tctcgttcag gaagatgatc agcttcttgg cgtgctcgta
360ctcctcggcg gcgtggtcga acaggaacag gccggcgccg tccaggctgt gggtgtagca
420ccagctgctc atgctcatgt acaggttgct gctctgcatc tccttgttca cctgctcgtt
480cagcagcttg atgatgtcgc ctccggactt gatctgaaac cggttgttca gggcctcatc
540tctgtacacg tcgtggtcgt aggtgccgtt tctgatgctg ccgatgcagg cgttgtcgca
600cttgtggtag atcttgaagc agccgttgcc catgtcctcg gcgttttctc tcagctgctt
660cttggttttc tcgaacagct tattcatctc actgtctgtc agatcaattg tgtgctggtt
720ttccagggcg accagcagtt cggcattgta gctccacagg tcgatcttgg tgtcctccac
780gtatttttcc aggtcctgga ttctgccctc cacctcgctg aattctttct cgatctggtg
840gaacttctcg ttggtcttgc cgatcagcct gttcagcttg ccgttgatct ggtcgatggc
900ggcctgggta gatttcagat cggcggcctg tccaattccc tcgctattct ggtgtctgaa
960gccgtaccac ccatccacca ttccctccca gccattctcg ataaagccgg caatggcgcc
1020aaagatgcct ctggtctgct tctcgggcac atttctcatg ccggtggcca gcttcagggt
1080gttctgcttc acgtatctag ggcaggcgcc gtaggtgatc ctgttcacgt tctggaaggg
1140cttgtcgttg gggatgctgc cgtttggggt gatgcactcg ctgttgcact tgccgatggg
1200ggcgtcggat ctcatgatgc tgctcttgcc gcttctgatc ttaaagtagc cccgaggggc
1260gatcaggttg ccggtgctgt tgatcagcag gatgtcgcca ggcttcacga ttgtccagta
1320gatgctgatc ctgctgggga tgttccgcac tctgggtctg ctgccgatat tggggatcac
1380ggtctgctgg cttctcttgg tggacacggt gattctgccg ctggcctggg cgtaagggaa
1440gatctgatcg ttgtcggtgc cagggtggtg cactccccag atgtacagct tgtcgaactt
1500ctcattgttg ggcatggtca cgttcagggc ggggtacttg aacttcaggt gggtcagcca
1560attcagtcta ctgaagaagc tgttgttgga ccgtctgatg caggcgctgc tggtgccatt
1620ctgtgtcacg ccggtccagt tgaagctctc gttgttgaat tccagggtgc cgctagaggc
1680caccaggctt ctcaggctgg cgtaatcagg cacgtcgtag gggtagcagt tgctgtaggc
1740cttgcttctc tccacgaaca ggtcccattt cttgttctgg aagccgtcgc actgaggatc
1800gcccagcagg gcatcgatca gggtacagtt ctcgccatcc aggatctggt gggggctgtc
1860acagatctcg cctgtgctgc tgctctgcac cagctctgtg gcattggtca cttcgatctg
1920gtcgttggtg attgttttca cgatggtgcc attaggcacg gcgtggtgtc ccagacacag
1980tgtggcggtg ctattatcgt tgccgggcag cttctgtgtg aacaccaggc acaggatgta
2040gctcagggca atgatggttt tcat
2064552067DNAArtificial SequenceSynthetic 55atggaaaaga tcgtgctgct
gctggccatt gtgagcctgg tgaagagcga ccagatctgc 60attggctacc acgccaacaa
tagcacagag caggtggaca ccatcatgga aaaaaacgtg 120accgtgaccc acgctcagga
catcctggaa aagacccaca acggcaagct gtgtgatctg 180gacggcgtga agcctctgat
cctgagagat tgtagcgtgg ctggatggct gctgggcaac 240cctatgtgcg acgagttcat
caacgtgccc gagtggagct atatcgtgga gaaggccaac 300cccaccaacg atctgtgtta
ccccggcagc ttcaacgatt acgaggaact gaagcacctg 360ctgtcccgga tcaaccactt
cgagaagatc cagatcatcc ccaagtcctc ttggagcgat 420cacgaagcct ctagcggagt
gtctagcgcc tgtccttacc tgggcagccc cagcttcttc 480agaaacgtgg tgtggctgat
caagaagaac agcacctacc ccaccatcaa gaagagctac 540aacaacacca accaggaaga
tctgctggtc ctgtggggaa tccaccaccc taatgatgcc 600gccgagcaga ccagactgta
ccagaacccc accacctata tcagcatcgg caccagcacc 660ctgaatcaga gactggtgcc
caagatcgcc accagatcca aggtgaacgg ccagagcggc 720aggatggaat tcttctggac
catcctgaag cccaacgacg ccatcaactt cgagagcaac 780ggcaacttta tcgcccctga
gtacgcctac aagatcgtga agaagggcga cagcgccatc 840atgaagagcg agctggaata
cggcaactgc aacaccaagt gccagacacc tatgggcgcc 900atcaacagca gcatgccctt
ccacaacatc caccctctga ccatcggcga gtgccctaag 960tacgtgaaga gcaacagact
ggtgctggcc acaggcctga gaaatagccc ccagcgggag 1020agcagaagaa agaagagggg
cctgtttgga gccatcgccg gctttattga aggcggctgg 1080cagggaatgg tggatggctg
gtacggctac caccacagca atgagcaggg ctctggatat 1140gccgccgaca aagagtctac
ccagaaggcc atcgacggcg tcaccaacaa ggtgaacagc 1200atcatcgaca agatgaacac
ccagttcgag gctgtgggca gagagttcaa caacctggaa 1260cggcggatcg agaacctgaa
caagaaaatg gaagatggct tcctggatgt gtggacctac 1320aatgccgaac tgctggtgct
gatggaaaac gagcggaccc tggacttcca cgacagcaac 1380gtgaagaacc tgtacgacaa
agtgcggctg cagctgagag acaacgccaa agagctgggc 1440aacggctgct tcgagttcta
ccacaagtgc gacaacgagt gcatggaaag catcaggaac 1500ggcacctaca actaccctca
gtacagcgag gaagccaggc tgaagaggga agagatcagc 1560tccggaggcg acatcatcaa
gctgctgaac gagcaggtga acaaggagat gcagagcagc 1620aacctgtaca tgagcatgag
cagctggtgc tacacccaca gcctggacgg cgccggcctg 1680ttcctgttcg accacgccgc
cgaggagtac gagcacgcca agaagctgat catcttcctg 1740aacgagaaca acgtgcccgt
gcagctgacc agcatcagcg cccccgagca caagttcgag 1800ggcctgaccc agatcttcca
gaaggcctac gagcacgagc agcacatcag cgagagcatc 1860aacaacatcg tggaccacgc
catcaagagc aaggaccacg ccaccttcaa cttcctgcag 1920tggtacgtgg ccgagcagca
cgaggaggag gtgctgttca aggacatcct ggacaagatc 1980gagctgatcg gcaacgagaa
ccacggcctg tacctggccg accagtacgt gaagggcatc 2040gccaagagca ggaagagcgg
atcctag 206756688PRTArtificial
SequenceSynthetic 56Met Glu Lys Ile Val Leu Leu Leu Ala Ile Val Ser Leu
Val Lys Ser 1 5 10 15
Asp Gln Ile Cys Ile Gly Tyr His Ala Asn Asn Ser Thr Glu Gln Val
20 25 30 Asp Thr Ile Met
Glu Lys Asn Val Thr Val Thr His Ala Gln Asp Ile 35
40 45 Leu Glu Lys Thr His Asn Gly Lys Leu
Cys Asp Leu Asp Gly Val Lys 50 55
60 Pro Leu Ile Leu Arg Asp Cys Ser Val Ala Gly Trp Leu
Leu Gly Asn 65 70 75
80 Pro Met Cys Asp Glu Phe Ile Asn Val Pro Glu Trp Ser Tyr Ile Val
85 90 95 Glu Lys Ala Asn
Pro Thr Asn Asp Leu Cys Tyr Pro Gly Ser Phe Asn 100
105 110 Asp Tyr Glu Glu Leu Lys His Leu Leu
Ser Arg Ile Asn His Phe Glu 115 120
125 Lys Ile Gln Ile Ile Pro Lys Ser Ser Trp Ser Asp His Glu
Ala Ser 130 135 140
Ser Gly Val Ser Ser Ala Cys Pro Tyr Leu Gly Ser Pro Ser Phe Phe 145
150 155 160 Arg Asn Val Val Trp
Leu Ile Lys Lys Asn Ser Thr Tyr Pro Thr Ile 165
170 175 Lys Lys Ser Tyr Asn Asn Thr Asn Gln Glu
Asp Leu Leu Val Leu Trp 180 185
190 Gly Ile His His Pro Asn Asp Ala Ala Glu Gln Thr Arg Leu Tyr
Gln 195 200 205 Asn
Pro Thr Thr Tyr Ile Ser Ile Gly Thr Ser Thr Leu Asn Gln Arg 210
215 220 Leu Val Pro Lys Ile Ala
Thr Arg Ser Lys Val Asn Gly Gln Ser Gly 225 230
235 240 Arg Met Glu Phe Phe Trp Thr Ile Leu Lys Pro
Asn Asp Ala Ile Asn 245 250
255 Phe Glu Ser Asn Gly Asn Phe Ile Ala Pro Glu Tyr Ala Tyr Lys Ile
260 265 270 Val Lys
Lys Gly Asp Ser Ala Ile Met Lys Ser Glu Leu Glu Tyr Gly 275
280 285 Asn Cys Asn Thr Lys Cys Gln
Thr Pro Met Gly Ala Ile Asn Ser Ser 290 295
300 Met Pro Phe His Asn Ile His Pro Leu Thr Ile Gly
Glu Cys Pro Lys 305 310 315
320 Tyr Val Lys Ser Asn Arg Leu Val Leu Ala Thr Gly Leu Arg Asn Ser
325 330 335 Pro Gln Arg
Glu Ser Arg Arg Lys Lys Arg Gly Leu Phe Gly Ala Ile 340
345 350 Ala Gly Phe Ile Glu Gly Gly Trp
Gln Gly Met Val Asp Gly Trp Tyr 355 360
365 Gly Tyr His His Ser Asn Glu Gln Gly Ser Gly Tyr Ala
Ala Asp Lys 370 375 380
Glu Ser Thr Gln Lys Ala Ile Asp Gly Val Thr Asn Lys Val Asn Ser 385
390 395 400 Ile Ile Asp Lys
Met Asn Thr Gln Phe Glu Ala Val Gly Arg Glu Phe 405
410 415 Asn Asn Leu Glu Arg Arg Ile Glu Asn
Leu Asn Lys Lys Met Glu Asp 420 425
430 Gly Phe Leu Asp Val Trp Thr Tyr Asn Ala Glu Leu Leu Val
Leu Met 435 440 445
Glu Asn Glu Arg Thr Leu Asp Phe His Asp Ser Asn Val Lys Asn Leu 450
455 460 Tyr Asp Lys Val Arg
Leu Gln Leu Arg Asp Asn Ala Lys Glu Leu Gly 465 470
475 480 Asn Gly Cys Phe Glu Phe Tyr His Lys Cys
Asp Asn Glu Cys Met Glu 485 490
495 Ser Ile Arg Asn Gly Thr Tyr Asn Tyr Pro Gln Tyr Ser Glu Glu
Ala 500 505 510 Arg
Leu Lys Arg Glu Glu Ile Ser Ser Gly Gly Asp Ile Ile Lys Leu 515
520 525 Leu Asn Glu Gln Val Asn
Lys Glu Met Gln Ser Ser Asn Leu Tyr Met 530 535
540 Ser Met Ser Ser Trp Cys Tyr Thr His Ser Leu
Asp Gly Ala Gly Leu 545 550 555
560 Phe Leu Phe Asp His Ala Ala Glu Glu Tyr Glu His Ala Lys Lys Leu
565 570 575 Ile Ile
Phe Leu Asn Glu Asn Asn Val Pro Val Gln Leu Thr Ser Ile 580
585 590 Ser Ala Pro Glu His Lys Phe
Glu Gly Leu Thr Gln Ile Phe Gln Lys 595 600
605 Ala Tyr Glu His Glu Gln His Ile Ser Glu Ser Ile
Asn Asn Ile Val 610 615 620
Asp His Ala Ile Lys Ser Lys Asp His Ala Thr Phe Asn Phe Leu Gln 625
630 635 640 Trp Tyr Val
Ala Glu Gln His Glu Glu Glu Val Leu Phe Lys Asp Ile 645
650 655 Leu Asp Lys Ile Glu Leu Ile Gly
Asn Glu Asn His Gly Leu Tyr Leu 660 665
670 Ala Asp Gln Tyr Val Lys Gly Ile Ala Lys Ser Arg Lys
Ser Gly Ser 675 680 685
572067DNAArtificial SequenceSynthetic 57ctaggatccg ctcttcctgc
tcttggcgat gcccttcacg tactggtcgg ccaggtacag 60gccgtggttc tcgttgccga
tcagctcgat cttgtccagg atgtccttga acagcacctc 120ctcctcgtgc tgctcggcca
cgtaccactg caggaagttg aaggtggcgt ggtccttgct 180cttgatggcg tggtccacga
tgttgttgat gctctcgctg atgtgctgct cgtgctcgta 240ggccttctgg aagatctggg
tcaggccctc gaacttgtgc tcgggggcgc tgatgctggt 300cagctgcacg ggcacgttgt
tctcgttcag gaagatgatc agcttcttgg cgtgctcgta 360ctcctcggcg gcgtggtcga
acaggaacag gccggcgccg tccaggctgt gggtgtagca 420ccagctgctc atgctcatgt
acaggttgct gctctgcatc tccttgttca cctgctcgtt 480cagcagcttg atgatgtcgc
ctccggagct gatctcttcc ctcttcagcc tggcttcctc 540gctgtactga gggtagttgt
aggtgccgtt cctgatgctt tccatgcact cgttgtcgca 600cttgtggtag aactcgaagc
agccgttgcc cagctctttg gcgttgtctc tcagctgcag 660ccgcactttg tcgtacaggt
tcttcacgtt gctgtcgtgg aagtccaggg tccgctcgtt 720ttccatcagc accagcagtt
cggcattgta ggtccacaca tccaggaagc catcttccat 780tttcttgttc aggttctcga
tccgccgttc caggttgttg aactctctgc ccacagcctc 840gaactgggtg ttcatcttgt
cgatgatgct gttcaccttg ttggtgacgc cgtcgatggc 900cttctgggta gactctttgt
cggcggcata tccagagccc tgctcattgc tgtggtggta 960gccgtaccag ccatccacca
ttccctgcca gccgccttca ataaagccgg cgatggctcc 1020aaacaggccc ctcttctttc
ttctgctctc ccgctggggg ctatttctca ggcctgtggc 1080cagcaccagt ctgttgctct
tcacgtactt agggcactcg ccgatggtca gagggtggat 1140gttgtggaag ggcatgctgc
tgttgatggc gcccataggt gtctggcact tggtgttgca 1200gttgccgtat tccagctcgc
tcttcatgat ggcgctgtcg cccttcttca cgatcttgta 1260ggcgtactca ggggcgataa
agttgccgtt gctctcgaag ttgatggcgt cgttgggctt 1320caggatggtc cagaagaatt
ccatcctgcc gctctggccg ttcaccttgg atctggtggc 1380gatcttgggc accagtctct
gattcagggt gctggtgccg atgctgatat aggtggtggg 1440gttctggtac agtctggtct
gctcggcggc atcattaggg tggtggattc cccacaggac 1500cagcagatct tcctggttgg
tgttgttgta gctcttcttg atggtggggt aggtgctgtt 1560cttcttgatc agccacacca
cgtttctgaa gaagctgggg ctgcccaggt aaggacaggc 1620gctagacact ccgctagagg
cttcgtgatc gctccaagag gacttgggga tgatctggat 1680cttctcgaag tggttgatcc
gggacagcag gtgcttcagt tcctcgtaat cgttgaagct 1740gccggggtaa cacagatcgt
tggtggggtt ggccttctcc acgatatagc tccactcggg 1800cacgttgatg aactcgtcgc
acatagggtt gcccagcagc catccagcca cgctacaatc 1860tctcaggatc agaggcttca
cgccgtccag atcacacagc ttgccgttgt gggtcttttc 1920caggatgtcc tgagcgtggg
tcacggtcac gtttttttcc atgatggtgt ccacctgctc 1980tgtgctattg ttggcgtggt
agccaatgca gatctggtcg ctcttcacca ggctcacaat 2040ggccagcagc agcacgatct
tttccat 2067582109DNAArtificial
SequenceSynthetic 58atgaaggcca tcatcgtgct gctgatggtg gtgaccagca
acgccgatag aatctgcacc 60ggcatcacca gcagcaatag cccccatgtg gtgaaaacag
ccacccaggg cgaagtgaat 120gtgacaggcg tgatccctct gaccaccacc cccaccaaga
gctacttcgc caacctgaag 180ggcaccagaa ccagaggcaa gctgtgcccc gattgcctga
actgcaccga tctggatgtg 240gctctgggca gacctatgtg tgtgggcacc acaccatctg
ccaaggccag catcctgcac 300gaagtgaagc ctgtgaccag cggctgcttc cccatcatgc
acgaccggac caagatcaga 360cagctgccca acctgctgag aggctacgag aacatccggc
tgtccaccca gaatgtgatc 420gatgccgaga aagcccctgg cggaccttat agactgggca
ccagcggctc ttgtcccaat 480gccacctcca agagcggctt ttttgccaca atggcctggg
ccgtgcctaa ggacaacaac 540aagaacgcca ccaaccctct gaccgtggag gtgccctaca
tctgtacaga gggcgaggat 600cagatcacag tgtggggctt ccacagcgac gacaagaccc
agatgaagaa cctgtacggc 660gacagcaacc cccagaagtt taccagcagc gccaatggcg
tgaccaccca ctacgtgtcc 720cagatcggca gctttcccga tcagacagag gatggcggac
tgcctcagtc tggcaggatc 780gtggtggact acatgatgca gaagcctggc aagaccggca
ccatcgtgta tcagagaggc 840gtgctgctgc ctcagaaagt gtggtgtgcc agcggcaggt
ctaaagtgat caagggcagc 900ctgcctctga ttggcgaggc cgactgtctg cacgaaaagt
acggcggcct gaacaagagc 960aagccctact acacaggcga gcacgccaag gccatcggca
attgccccat ctgggtgaaa 1020acccccctga agctggccaa tggcaccaag tacagacctc
ccgccaagct gctgaaagag 1080agaggcttct ttggcgccat tgccggattt ctggaaggcg
gctgggaggg aatgattgcc 1140ggctggcacg gctatacatc tcatggggcc catggcgtgg
ctgtggccgc cgatctgaag 1200tctacccagg aagccatcaa caagatcacc aagaacctga
acagcctgag cgagctggaa 1260gtgaagaatc tgcagagact gagcggcgcc atggatgagc
tgcacaacga gatcctggaa 1320ctggacgaga aagtggatga tctccgcgcc gatacaattt
cctcccagat tgaactggcc 1380gtgctgctgt ccaacgaggg catcatcaac agcgaggatg
aacacctgct ggccctggaa 1440cggaagctga agaagatgct gggcccttct gccgtggaga
tcggcaacgg ctgcttcgag 1500acaaagcaca agtgcaacca gacctgcctg gatagaatcg
ccgctggcac cttcaatgcc 1560ggcgagttca gcctgcctac cttcgacagc ctgaatatca
cctccggagg cgacatcatc 1620aagctgctga acgagcaggt gaacaaggag atgcagagca
gcaacctgta catgagcatg 1680agcagctggt gctacaccca cagcctggac ggcgccggcc
tgttcctgtt cgaccacgcc 1740gccgaggagt acgagcacgc caagaagctg atcatcttcc
tgaacgagaa caacgtgccc 1800gtgcagctga ccagcatcag cgcccccgag cacaagttcg
agggcctgac ccagatcttc 1860cagaaggcct acgagcacga gcagcacatc agcgagagca
tcaacaacat cgtggaccac 1920gccatcaaga gcaaggacca cgccaccttc aacttcctgc
agtggtacgt ggccgagcag 1980cacgaggagg aggtgctgtt caaggacatc ctggacaaga
tcgagctgat cggcaacgag 2040aaccacggcc tgtacctggc cgaccagtac gtgaagggca
tcgccaagag caggaagagc 2100ggatcctag
210959702PRTArtificial SequenceSynthetic 59Met Lys
Ala Ile Ile Val Leu Leu Met Val Val Thr Ser Asn Ala Asp 1 5
10 15 Arg Ile Cys Thr Gly Ile Thr
Ser Ser Asn Ser Pro His Val Val Lys 20 25
30 Thr Ala Thr Gln Gly Glu Val Asn Val Thr Gly Val
Ile Pro Leu Thr 35 40 45
Thr Thr Pro Thr Lys Ser Tyr Phe Ala Asn Leu Lys Gly Thr Arg Thr
50 55 60 Arg Gly Lys
Leu Cys Pro Asp Cys Leu Asn Cys Thr Asp Leu Asp Val 65
70 75 80 Ala Leu Gly Arg Pro Met Cys
Val Gly Thr Thr Pro Ser Ala Lys Ala 85
90 95 Ser Ile Leu His Glu Val Lys Pro Val Thr Ser
Gly Cys Phe Pro Ile 100 105
110 Met His Asp Arg Thr Lys Ile Arg Gln Leu Pro Asn Leu Leu Arg
Gly 115 120 125 Tyr
Glu Asn Ile Arg Leu Ser Thr Gln Asn Val Ile Asp Ala Glu Lys 130
135 140 Ala Pro Gly Gly Pro Tyr
Arg Leu Gly Thr Ser Gly Ser Cys Pro Asn 145 150
155 160 Ala Thr Ser Lys Ser Gly Phe Phe Ala Thr Met
Ala Trp Ala Val Pro 165 170
175 Lys Asp Asn Asn Lys Asn Ala Thr Asn Pro Leu Thr Val Glu Val Pro
180 185 190 Tyr Ile
Cys Thr Glu Gly Glu Asp Gln Ile Thr Val Trp Gly Phe His 195
200 205 Ser Asp Asp Lys Thr Gln Met
Lys Asn Leu Tyr Gly Asp Ser Asn Pro 210 215
220 Gln Lys Phe Thr Ser Ser Ala Asn Gly Val Thr Thr
His Tyr Val Ser 225 230 235
240 Gln Ile Gly Ser Phe Pro Asp Gln Thr Glu Asp Gly Gly Leu Pro Gln
245 250 255 Ser Gly Arg
Ile Val Val Asp Tyr Met Met Gln Lys Pro Gly Lys Thr 260
265 270 Gly Thr Ile Val Tyr Gln Arg Gly
Val Leu Leu Pro Gln Lys Val Trp 275 280
285 Cys Ala Ser Gly Arg Ser Lys Val Ile Lys Gly Ser Leu
Pro Leu Ile 290 295 300
Gly Glu Ala Asp Cys Leu His Glu Lys Tyr Gly Gly Leu Asn Lys Ser 305
310 315 320 Lys Pro Tyr Tyr
Thr Gly Glu His Ala Lys Ala Ile Gly Asn Cys Pro 325
330 335 Ile Trp Val Lys Thr Pro Leu Lys Leu
Ala Asn Gly Thr Lys Tyr Arg 340 345
350 Pro Pro Ala Lys Leu Leu Lys Glu Arg Gly Phe Phe Gly Ala
Ile Ala 355 360 365
Gly Phe Leu Glu Gly Gly Trp Glu Gly Met Ile Ala Gly Trp His Gly 370
375 380 Tyr Thr Ser His Gly
Ala His Gly Val Ala Val Ala Ala Asp Leu Lys 385 390
395 400 Ser Thr Gln Glu Ala Ile Asn Lys Ile Thr
Lys Asn Leu Asn Ser Leu 405 410
415 Ser Glu Leu Glu Val Lys Asn Leu Gln Arg Leu Ser Gly Ala Met
Asp 420 425 430 Glu
Leu His Asn Glu Ile Leu Glu Leu Asp Glu Lys Val Asp Asp Leu 435
440 445 Arg Ala Asp Thr Ile Ser
Ser Gln Ile Glu Leu Ala Val Leu Leu Ser 450 455
460 Asn Glu Gly Ile Ile Asn Ser Glu Asp Glu His
Leu Leu Ala Leu Glu 465 470 475
480 Arg Lys Leu Lys Lys Met Leu Gly Pro Ser Ala Val Glu Ile Gly Asn
485 490 495 Gly Cys
Phe Glu Thr Lys His Lys Cys Asn Gln Thr Cys Leu Asp Arg 500
505 510 Ile Ala Ala Gly Thr Phe Asn
Ala Gly Glu Phe Ser Leu Pro Thr Phe 515 520
525 Asp Ser Leu Asn Ile Thr Ser Gly Gly Asp Ile Ile
Lys Leu Leu Asn 530 535 540
Glu Gln Val Asn Lys Glu Met Gln Ser Ser Asn Leu Tyr Met Ser Met 545
550 555 560 Ser Ser Trp
Cys Tyr Thr His Ser Leu Asp Gly Ala Gly Leu Phe Leu 565
570 575 Phe Asp His Ala Ala Glu Glu Tyr
Glu His Ala Lys Lys Leu Ile Ile 580 585
590 Phe Leu Asn Glu Asn Asn Val Pro Val Gln Leu Thr Ser
Ile Ser Ala 595 600 605
Pro Glu His Lys Phe Glu Gly Leu Thr Gln Ile Phe Gln Lys Ala Tyr 610
615 620 Glu His Glu Gln
His Ile Ser Glu Ser Ile Asn Asn Ile Val Asp His 625 630
635 640 Ala Ile Lys Ser Lys Asp His Ala Thr
Phe Asn Phe Leu Gln Trp Tyr 645 650
655 Val Ala Glu Gln His Glu Glu Glu Val Leu Phe Lys Asp Ile
Leu Asp 660 665 670
Lys Ile Glu Leu Ile Gly Asn Glu Asn His Gly Leu Tyr Leu Ala Asp
675 680 685 Gln Tyr Val Lys
Gly Ile Ala Lys Ser Arg Lys Ser Gly Ser 690 695
700 602109DNAArtificial SequenceSynthetic 60ctaggatccg
ctcttcctgc tcttggcgat gcccttcacg tactggtcgg ccaggtacag 60gccgtggttc
tcgttgccga tcagctcgat cttgtccagg atgtccttga acagcacctc 120ctcctcgtgc
tgctcggcca cgtaccactg caggaagttg aaggtggcgt ggtccttgct 180cttgatggcg
tggtccacga tgttgttgat gctctcgctg atgtgctgct cgtgctcgta 240ggccttctgg
aagatctggg tcaggccctc gaacttgtgc tcgggggcgc tgatgctggt 300cagctgcacg
ggcacgttgt tctcgttcag gaagatgatc agcttcttgg cgtgctcgta 360ctcctcggcg
gcgtggtcga acaggaacag gccggcgccg tccaggctgt gggtgtagca 420ccagctgctc
atgctcatgt acaggttgct gctctgcatc tccttgttca cctgctcgtt 480cagcagcttg
atgatgtcgc ctccggaggt gatattcagg ctgtcgaagg taggcaggct 540gaactcgccg
gcattgaagg tgccagcggc gattctatcc aggcaggtct ggttgcactt 600gtgctttgtc
tcgaagcagc cgttgccgat ctccacggca gaagggccca gcatcttctt 660cagcttccgt
tccagggcca gcaggtgttc atcctcgctg ttgatgatgc cctcgttgga 720cagcagcacg
gccagttcaa tctgggagga aattgtatcg gcgcggagat catccacttt 780ctcgtccagt
tccaggatct cgttgtgcag ctcatccatg gcgccgctca gtctctgcag 840attcttcact
tccagctcgc tcaggctgtt caggttcttg gtgatcttgt tgatggcttc 900ctgggtagac
ttcagatcgg cggccacagc cacgccatgg gccccatgag atgtatagcc 960gtgccagccg
gcaatcattc cctcccagcc gccttccaga aatccggcaa tggcgccaaa 1020gaagcctctc
tctttcagca gcttggcggg aggtctgtac ttggtgccat tggccagctt 1080caggggggtt
ttcacccaga tggggcaatt gccgatggcc ttggcgtgct cgcctgtgta 1140gtagggcttg
ctcttgttca ggccgccgta cttttcgtgc agacagtcgg cctcgccaat 1200cagaggcagg
ctgcccttga tcactttaga cctgccgctg gcacaccaca ctttctgagg 1260cagcagcacg
cctctctgat acacgatggt gccggtcttg ccaggcttct gcatcatgta 1320gtccaccacg
atcctgccag actgaggcag tccgccatcc tctgtctgat cgggaaagct 1380gccgatctgg
gacacgtagt gggtggtcac gccattggcg ctgctggtaa acttctgggg 1440gttgctgtcg
ccgtacaggt tcttcatctg ggtcttgtcg tcgctgtgga agccccacac 1500tgtgatctga
tcctcgccct ctgtacagat gtagggcacc tccacggtca gagggttggt 1560ggcgttcttg
ttgttgtcct taggcacggc ccaggccatt gtggcaaaaa agccgctctt 1620ggaggtggca
ttgggacaag agccgctggt gcccagtcta taaggtccgc caggggcttt 1680ctcggcatcg
atcacattct gggtggacag ccggatgttc tcgtagcctc tcagcaggtt 1740gggcagctgt
ctgatcttgg tccggtcgtg catgatgggg aagcagccgc tggtcacagg 1800cttcacttcg
tgcaggatgc tggccttggc agatggtgtg gtgcccacac acataggtct 1860gcccagagcc
acatccagat cggtgcagtt caggcaatcg gggcacagct tgcctctggt 1920tctggtgccc
ttcaggttgg cgaagtagct cttggtgggg gtggtggtca gagggatcac 1980gcctgtcaca
ttcacttcgc cctgggtggc tgttttcacc acatgggggc tattgctgct 2040ggtgatgccg
gtgcagattc tatcggcgtt gctggtcacc accatcagca gcacgatgat 2100ggccttcat
2109612064DNAArtificial SequenceSynthetic 61atgaaaacca taattgcgct
gtcctacata ctgtgtctgg tgtttgccca gaaactgccg 60ggcaatgaca actcaacagc
cacgctctgc ttggggcacc atgccgtccc taacgggacc 120attgtgaaaa ccattactaa
cgatcagata gaggtgacta atgccaccga gctggtgcaa 180agtagctcca caggagagat
ctgcgatagt ccccaccaga ttctggacgg aaagaattgt 240acgctgatcg acgcgctgtt
gggcgaccct cagtgtgacg gatttcagaa taagaagtgg 300gatctgtttg tggaaaggtc
aaaggcttat tcaaattgct acccttacga tgtgcctgat 360tatgccagcc tgcggtccct
cgtcgcgtct agtgggactc tggagttcaa caacgagtca 420tttaactgga ctggcgttac
acagaacggg actagttccg cttgcataag gagaagcaaa 480aatagtttct tcagcagact
gaattggctg acacatctga acttcaagta ccctgcactg 540aatgtaacca tgcccaacaa
cgagcagttc gataagcttt acatttgggg agttcatcat 600cctggcactg acaaggatca
gatctttctg tatgcccagg cttccggcag gattaccgtg 660tctacaaaga gaagccagca
aactgtgtct cccaatatcg gcagtagacc cagagtacgg 720aacatcccta gtcgcatcag
tatttactgg accatcgtga aaccaggcga tattctcctg 780attaacagta ctggcaacct
gatcgccccc cggggatact ttaaaatccg ctctggaaag 840tcctccatta tgagatcaga
tgcaccgatc ggaaaatgca actctgagtg tatcacaccc 900aatgggagca ttcccaatga
caaacctttc cagaacgtta atcgaataac ttatggggcc 960tgtccacggt acgtgaagca
aaataccttg aaactggcga ccggtatgcg caatgtcccc 1020gaaaaacaga cccgcgggat
atttggggct atcgcaggct ttatcgagaa tggctgggaa 1080gggatggtgg atggttggta
tggttttaga catcaaaact ccgaaggcag aggccaggct 1140gccgatctca agagcacgca
ggccgctata gatcagatca atggaaagct caacagactg 1200atcgggaaaa ccaacgaaaa
attccatcag atcgagaaag agttctccga agtcgagggg 1260cgcatacagg acctggagaa
gtatgttgag gatacaaaga ttgatctgtg gtcctacaat 1320gccgagctgc tggtggctct
ggagaatcag cacactattg acctgaccga ttcagagatg 1380aacaaacttt ttgagaagac
gaagaagcag cttagagaaa atgcagagga catggggaac 1440ggatgcttta aaatatatca
taagtgtgat aatgcctgca tcggatcaat tagaaatggt 1500acctatgatc acgatgttta
cagggacgaa gcgctgaata acaggttcca gataaaatcc 1560ggaggcgaca tcatcaagct
gctgaacgag caggtgaaca aggagatgca gagcagcaac 1620ctgtacatga gcatgagcag
ctggtgctac acccacagcc tggacggcgc cggcctgttc 1680ctgttcgacc acgccgccga
ggagtacgag cacgccaaga agctgatcat cttcctgaac 1740gagaacaacg tgcccgtgca
gctgaccagc atcagcgccc ccgagcacaa gttcgagggc 1800ctgacccaga tcttccagaa
ggcctacgag cacgagcagc acatcagcga gagcatcaac 1860aacatcgtgg accacgccat
caagagcaag gaccacgcca ccttcaactt cctgcagtgg 1920tacgtggccg agcagcacga
ggaggaggtg ctgttcaagg acatcctgga caagatcgag 1980ctgatcggca acgagaacca
cggcctgtac ctggccgacc agtacgtgaa gggcatcgcc 2040aagagcagga agagcggatc
ctag 206462687PRTArtificial
SequenceSynthetic 62Met Lys Thr Ile Ile Ala Leu Ser Tyr Ile Leu Cys Leu
Val Phe Ala 1 5 10 15
Gln Lys Leu Pro Gly Asn Asp Asn Ser Thr Ala Thr Leu Cys Leu Gly
20 25 30 His His Ala Val
Pro Asn Gly Thr Ile Val Lys Thr Ile Thr Asn Asp 35
40 45 Gln Ile Glu Val Thr Asn Ala Thr Glu
Leu Val Gln Ser Ser Ser Thr 50 55
60 Gly Glu Ile Cys Asp Ser Pro His Gln Ile Leu Asp Gly
Lys Asn Cys 65 70 75
80 Thr Leu Ile Asp Ala Leu Leu Gly Asp Pro Gln Cys Asp Gly Phe Gln
85 90 95 Asn Lys Lys Trp
Asp Leu Phe Val Glu Arg Ser Lys Ala Tyr Ser Asn 100
105 110 Cys Tyr Pro Tyr Asp Val Pro Asp Tyr
Ala Ser Leu Arg Ser Leu Val 115 120
125 Ala Ser Ser Gly Thr Leu Glu Phe Asn Asn Glu Ser Phe Asn
Trp Thr 130 135 140
Gly Val Thr Gln Asn Gly Thr Ser Ser Ala Cys Ile Arg Arg Ser Lys 145
150 155 160 Asn Ser Phe Phe Ser
Arg Leu Asn Trp Leu Thr His Leu Asn Phe Lys 165
170 175 Tyr Pro Ala Leu Asn Val Thr Met Pro Asn
Asn Glu Gln Phe Asp Lys 180 185
190 Leu Tyr Ile Trp Gly Val His His Pro Gly Thr Asp Lys Asp Gln
Ile 195 200 205 Phe
Leu Tyr Ala Gln Ala Ser Gly Arg Ile Thr Val Ser Thr Lys Arg 210
215 220 Ser Gln Gln Thr Val Ser
Pro Asn Ile Gly Ser Arg Pro Arg Val Arg 225 230
235 240 Asn Ile Pro Ser Arg Ile Ser Ile Tyr Trp Thr
Ile Val Lys Pro Gly 245 250
255 Asp Ile Leu Leu Ile Asn Ser Thr Gly Asn Leu Ile Ala Pro Arg Gly
260 265 270 Tyr Phe
Lys Ile Arg Ser Gly Lys Ser Ser Ile Met Arg Ser Asp Ala 275
280 285 Pro Ile Gly Lys Cys Asn Ser
Glu Cys Ile Thr Pro Asn Gly Ser Ile 290 295
300 Pro Asn Asp Lys Pro Phe Gln Asn Val Asn Arg Ile
Thr Tyr Gly Ala 305 310 315
320 Cys Pro Arg Tyr Val Lys Gln Asn Thr Leu Lys Leu Ala Thr Gly Met
325 330 335 Arg Asn Val
Pro Glu Lys Gln Thr Arg Gly Ile Phe Gly Ala Ile Ala 340
345 350 Gly Phe Ile Glu Asn Gly Trp Glu
Gly Met Val Asp Gly Trp Tyr Gly 355 360
365 Phe Arg His Gln Asn Ser Glu Gly Arg Gly Gln Ala Ala
Asp Leu Lys 370 375 380
Ser Thr Gln Ala Ala Ile Asp Gln Ile Asn Gly Lys Leu Asn Arg Leu 385
390 395 400 Ile Gly Lys Thr
Asn Glu Lys Phe His Gln Ile Glu Lys Glu Phe Ser 405
410 415 Glu Val Glu Gly Arg Ile Gln Asp Leu
Glu Lys Tyr Val Glu Asp Thr 420 425
430 Lys Ile Asp Leu Trp Ser Tyr Asn Ala Glu Leu Leu Val Ala
Leu Glu 435 440 445
Asn Gln His Thr Ile Asp Leu Thr Asp Ser Glu Met Asn Lys Leu Phe 450
455 460 Glu Lys Thr Lys Lys
Gln Leu Arg Glu Asn Ala Glu Asp Met Gly Asn 465 470
475 480 Gly Cys Phe Lys Ile Tyr His Lys Cys Asp
Asn Ala Cys Ile Gly Ser 485 490
495 Ile Arg Asn Gly Thr Tyr Asp His Asp Val Tyr Arg Asp Glu Ala
Leu 500 505 510 Asn
Asn Arg Phe Gln Ile Lys Ser Gly Gly Asp Ile Ile Lys Leu Leu 515
520 525 Asn Glu Gln Val Asn Lys
Glu Met Gln Ser Ser Asn Leu Tyr Met Ser 530 535
540 Met Ser Ser Trp Cys Tyr Thr His Ser Leu Asp
Gly Ala Gly Leu Phe 545 550 555
560 Leu Phe Asp His Ala Ala Glu Glu Tyr Glu His Ala Lys Lys Leu Ile
565 570 575 Ile Phe
Leu Asn Glu Asn Asn Val Pro Val Gln Leu Thr Ser Ile Ser 580
585 590 Ala Pro Glu His Lys Phe Glu
Gly Leu Thr Gln Ile Phe Gln Lys Ala 595 600
605 Tyr Glu His Glu Gln His Ile Ser Glu Ser Ile Asn
Asn Ile Val Asp 610 615 620
His Ala Ile Lys Ser Lys Asp His Ala Thr Phe Asn Phe Leu Gln Trp 625
630 635 640 Tyr Val Ala
Glu Gln His Glu Glu Glu Val Leu Phe Lys Asp Ile Leu 645
650 655 Asp Lys Ile Glu Leu Ile Gly Asn
Glu Asn His Gly Leu Tyr Leu Ala 660 665
670 Asp Gln Tyr Val Lys Gly Ile Ala Lys Ser Arg Lys Ser
Gly Ser 675 680 685
632064DNAArtificial SequenceSynthetic 63ctaggatccg ctcttcctgc tcttggcgat
gcccttcacg tactggtcgg ccaggtacag 60gccgtggttc tcgttgccga tcagctcgat
cttgtccagg atgtccttga acagcacctc 120ctcctcgtgc tgctcggcca cgtaccactg
caggaagttg aaggtggcgt ggtccttgct 180cttgatggcg tggtccacga tgttgttgat
gctctcgctg atgtgctgct cgtgctcgta 240ggccttctgg aagatctggg tcaggccctc
gaacttgtgc tcgggggcgc tgatgctggt 300cagctgcacg ggcacgttgt tctcgttcag
gaagatgatc agcttcttgg cgtgctcgta 360ctcctcggcg gcgtggtcga acaggaacag
gccggcgccg tccaggctgt gggtgtagca 420ccagctgctc atgctcatgt acaggttgct
gctctgcatc tccttgttca cctgctcgtt 480cagcagcttg atgatgtcgc ctccggattt
tatctggaac ctgttattca gcgcttcgtc 540cctgtaaaca tcgtgatcat aggtaccatt
tctaattgat ccgatgcagg cattatcaca 600cttatgatat attttaaagc atccgttccc
catgtcctct gcattttctc taagctgctt 660cttcgtcttc tcaaaaagtt tgttcatctc
tgaatcggtc aggtcaatag tgtgctgatt 720ctccagagcc accagcagct cggcattgta
ggaccacaga tcaatctttg tatcctcaac 780atacttctcc aggtcctgta tgcgcccctc
gacttcggag aactctttct cgatctgatg 840gaatttttcg ttggttttcc cgatcagtct
gttgagcttt ccattgatct gatctatagc 900ggcctgcgtg ctcttgagat cggcagcctg
gcctctgcct tcggagtttt gatgtctaaa 960accataccaa ccatccacca tcccttccca
gccattctcg ataaagcctg cgatagcccc 1020aaatatcccg cgggtctgtt tttcggggac
attgcgcata ccggtcgcca gtttcaaggt 1080attttgcttc acgtaccgtg gacaggcccc
ataagttatt cgattaacgt tctggaaagg 1140tttgtcattg ggaatgctcc cattgggtgt
gatacactca gagttgcatt ttccgatcgg 1200tgcatctgat ctcataatgg aggactttcc
agagcggatt ttaaagtatc cccggggggc 1260gatcaggttg ccagtactgt taatcaggag
aatatcgcct ggtttcacga tggtccagta 1320aatactgatg cgactaggga tgttccgtac
tctgggtcta ctgccgatat tgggagacac 1380agtttgctgg cttctctttg tagacacggt
aatcctgccg gaagcctggg catacagaaa 1440gatctgatcc ttgtcagtgc caggatgatg
aactccccaa atgtaaagct tatcgaactg 1500ctcgttgttg ggcatggtta cattcagtgc
agggtacttg aagttcagat gtgtcagcca 1560attcagtctg ctgaagaaac tatttttgct
tctccttatg caagcggaac tagtcccgtt 1620ctgtgtaacg ccagtccagt taaatgactc
gttgttgaac tccagagtcc cactagacgc 1680gacgagggac cgcaggctgg cataatcagg
cacatcgtaa gggtagcaat ttgaataagc 1740ctttgacctt tccacaaaca gatcccactt
cttattctga aatccgtcac actgagggtc 1800gcccaacagc gcgtcgatca gcgtacaatt
ctttccgtcc agaatctggt ggggactatc 1860gcagatctct cctgtggagc tactttgcac
cagctcggtg gcattagtca cctctatctg 1920atcgttagta atggttttca caatggtccc
gttagggacg gcatggtgcc ccaagcagag 1980cgtggctgtt gagttgtcat tgcccggcag
tttctgggca aacaccagac acagtatgta 2040ggacagcgca attatggttt tcat
2064642058DNAArtificial
SequenceSynthetic 64atgaaagtga agctgctggt gctgctgtgt acctttaccg
ccacctacgc cgataccatc 60tgtatcggct accacgccaa caatagcacc gacaccgtgg
ataccgtgct ggaaaagaac 120gtgaccgtga cccacagcgt gaacctgctg gaaaacagcc
acaacggcaa gctgtgtctg 180ctgaaaggca ttgcccctct gcagctggga aattgtagcg
tggccggctg gattctgggc 240aatcctgagt gcgagctgct gatttccaaa gagtcctggt
cctacatcgt ggagaagccc 300aaccctgaga atggcacctg ctaccctggc cacttcgccg
attacgagga actgagagaa 360cagctgtcca gcgtgtccag cttcgagaga ttcgagatct
tccccaaaga gagcagctgg 420cccaatcata cagtgaccgg cgtgagcgcc tcttgtagcc
acaatggcga gagcagcttc 480tacagaaacc tgctgtggct gaccggcaag aacggcctgt
accccaacct gagcaagagc 540tacgccaaca acaaagaaaa agaagtgctg gtcctctggg
gagtgcacca ccctcctaac 600atcggcatcc agaaggccct gtaccacacc gagaatgcct
acgtgtccgt ggtgtccagc 660cactacagca gaaagttcac ccccgagatc gccaaaagac
ccaaagtgcg ggaccaggaa 720ggcaggatca actactactg gaccctgctg gaacctggcg
acaccatcat cttcgaggcc 780aacggcaatc tgatcgcccc tagatacgcc tttgccctga
gcagaggctt tggcagcggc 840atcatcaaca gcaacgcccc catggacaag tgtgacgcca
agtgtcagac accacaggga 900gctatcaata gcagcctgcc cttccagaat gtgcaccctg
tgaccatcgg cgagtgtcct 960aaatacgtgc ggagcgccaa gctgagaatg gtgaccggcc
tgaggaatat ccccagcatc 1020cagagcagag gcctgtttgg cgccattgcc ggctttatcg
agggcggatg gacaggcatg 1080gtggatgggt ggtacggcta ccaccaccag aatgagcagg
gatctggcta tgccgccgat 1140cagaagagca cccagaacgc catcaacggc atcaccaaca
aagtgaacag cgtgatcgag 1200aagatgaaca cccagttcac cgccgtgggc aaagagttca
acaagctgga acggcggatg 1260gaaaacctga acaagaaggt ggacgacggc ttcatcgaca
tctggaccta caacgccgaa 1320ctcctggtcc tcctggaaaa tgagaggacc ctggacttcc
acgacagcaa cgtgaagaac 1380ctgtacgaga aagtgaagag ccagctgaag aacaacgcca
aagagatcgg caacggctgc 1440ttcgagttct accacaagtg caacgacgag tgcatggaaa
gcgtgaagaa cggcacctac 1500gactacccca agtacagcga ggaaagcaag ctgaaccggg
agaagatcga ttccggaggc 1560gacatcatca agctgctgaa cgagcaggtg aacaaggaga
tgcagagcag caacctgtac 1620atgagcatga gcagctggtg ctacacccac agcctggacg
gcgccggcct gttcctgttc 1680gaccacgccg ccgaggagta cgagcacgcc aagaagctga
tcatcttcct gaacgagaac 1740aacgtgcccg tgcagctgac cagcatcagc gcccccgagc
acaagttcga gggcctgacc 1800cagatcttcc agaaggccta cgagcacgag cagcacatca
gcgagagcat caacaacatc 1860gtggaccacg ccatcaagag caaggaccac gccaccttca
acttcctgca gtggtacgtg 1920gccgagcagc acgaggagga ggtgctgttc aaggacatcc
tggacaagat cgagctgatc 1980ggcaacgaga accacggcct gtacctggcc gaccagtacg
tgaagggcat cgccaagagc 2040aggaagagcg gatcctag
205865685PRTArtificial SequenceSynthetic 65Met Lys
Val Lys Leu Leu Val Leu Leu Cys Thr Phe Thr Ala Thr Tyr 1 5
10 15 Ala Asp Thr Ile Cys Ile Gly
Tyr His Ala Asn Asn Ser Thr Asp Thr 20 25
30 Val Asp Thr Val Leu Glu Lys Asn Val Thr Val Thr
His Ser Val Asn 35 40 45
Leu Leu Glu Asn Ser His Asn Gly Lys Leu Cys Leu Leu Lys Gly Ile
50 55 60 Ala Pro Leu
Gln Leu Gly Asn Cys Ser Val Ala Gly Trp Ile Leu Gly 65
70 75 80 Asn Pro Glu Cys Glu Leu Leu
Ile Ser Lys Glu Ser Trp Ser Tyr Ile 85
90 95 Val Glu Lys Pro Asn Pro Glu Asn Gly Thr Cys
Tyr Pro Gly His Phe 100 105
110 Ala Asp Tyr Glu Glu Leu Arg Glu Gln Leu Ser Ser Val Ser Ser
Phe 115 120 125 Glu
Arg Phe Glu Ile Phe Pro Lys Glu Ser Ser Trp Pro Asn His Thr 130
135 140 Val Thr Gly Val Ser Ala
Ser Cys Ser His Asn Gly Glu Ser Ser Phe 145 150
155 160 Tyr Arg Asn Leu Leu Trp Leu Thr Gly Lys Asn
Gly Leu Tyr Pro Asn 165 170
175 Leu Ser Lys Ser Tyr Ala Asn Asn Lys Glu Lys Glu Val Leu Val Leu
180 185 190 Trp Gly
Val His His Pro Pro Asn Ile Gly Ile Gln Lys Ala Leu Tyr 195
200 205 His Thr Glu Asn Ala Tyr Val
Ser Val Val Ser Ser His Tyr Ser Arg 210 215
220 Lys Phe Thr Pro Glu Ile Ala Lys Arg Pro Lys Val
Arg Asp Gln Glu 225 230 235
240 Gly Arg Ile Asn Tyr Tyr Trp Thr Leu Leu Glu Pro Gly Asp Thr Ile
245 250 255 Ile Phe Glu
Ala Asn Gly Asn Leu Ile Ala Pro Arg Tyr Ala Phe Ala 260
265 270 Leu Ser Arg Gly Phe Gly Ser Gly
Ile Ile Asn Ser Asn Ala Pro Met 275 280
285 Asp Lys Cys Asp Ala Lys Cys Gln Thr Pro Gln Gly Ala
Ile Asn Ser 290 295 300
Ser Leu Pro Phe Gln Asn Val His Pro Val Thr Ile Gly Glu Cys Pro 305
310 315 320 Lys Tyr Val Arg
Ser Ala Lys Leu Arg Met Val Thr Gly Leu Arg Asn 325
330 335 Ile Pro Ser Ile Gln Ser Arg Gly Leu
Phe Gly Ala Ile Ala Gly Phe 340 345
350 Ile Glu Gly Gly Trp Thr Gly Met Val Asp Gly Trp Tyr Gly
Tyr His 355 360 365
His Gln Asn Glu Gln Gly Ser Gly Tyr Ala Ala Asp Gln Lys Ser Thr 370
375 380 Gln Asn Ala Ile Asn
Gly Ile Thr Asn Lys Val Asn Ser Val Ile Glu 385 390
395 400 Lys Met Asn Thr Gln Phe Thr Ala Val Gly
Lys Glu Phe Asn Lys Leu 405 410
415 Glu Arg Arg Met Glu Asn Leu Asn Lys Lys Val Asp Asp Gly Phe
Ile 420 425 430 Asp
Ile Trp Thr Tyr Asn Ala Glu Leu Leu Val Leu Leu Glu Asn Glu 435
440 445 Arg Thr Leu Asp Phe His
Asp Ser Asn Val Lys Asn Leu Tyr Glu Lys 450 455
460 Val Lys Ser Gln Leu Lys Asn Asn Ala Lys Glu
Ile Gly Asn Gly Cys 465 470 475
480 Phe Glu Phe Tyr His Lys Cys Asn Asp Glu Cys Met Glu Ser Val Lys
485 490 495 Asn Gly
Thr Tyr Asp Tyr Pro Lys Tyr Ser Glu Glu Ser Lys Leu Asn 500
505 510 Arg Glu Lys Ile Asp Ser Gly
Gly Asp Ile Ile Lys Leu Leu Asn Glu 515 520
525 Gln Val Asn Lys Glu Met Gln Ser Ser Asn Leu Tyr
Met Ser Met Ser 530 535 540
Ser Trp Cys Tyr Thr His Ser Leu Asp Gly Ala Gly Leu Phe Leu Phe 545
550 555 560 Asp His Ala
Ala Glu Glu Tyr Glu His Ala Lys Lys Leu Ile Ile Phe 565
570 575 Leu Asn Glu Asn Asn Val Pro Val
Gln Leu Thr Ser Ile Ser Ala Pro 580 585
590 Glu His Lys Phe Glu Gly Leu Thr Gln Ile Phe Gln Lys
Ala Tyr Glu 595 600 605
His Glu Gln His Ile Ser Glu Ser Ile Asn Asn Ile Val Asp His Ala 610
615 620 Ile Lys Ser Lys
Asp His Ala Thr Phe Asn Phe Leu Gln Trp Tyr Val 625 630
635 640 Ala Glu Gln His Glu Glu Glu Val Leu
Phe Lys Asp Ile Leu Asp Lys 645 650
655 Ile Glu Leu Ile Gly Asn Glu Asn His Gly Leu Tyr Leu Ala
Asp Gln 660 665 670
Tyr Val Lys Gly Ile Ala Lys Ser Arg Lys Ser Gly Ser 675
680 685 662058DNAArtificial SequenceSynthetic
66ctaggatccg ctcttcctgc tcttggcgat gcccttcacg tactggtcgg ccaggtacag
60gccgtggttc tcgttgccga tcagctcgat cttgtccagg atgtccttga acagcacctc
120ctcctcgtgc tgctcggcca cgtaccactg caggaagttg aaggtggcgt ggtccttgct
180cttgatggcg tggtccacga tgttgttgat gctctcgctg atgtgctgct cgtgctcgta
240ggccttctgg aagatctggg tcaggccctc gaacttgtgc tcgggggcgc tgatgctggt
300cagctgcacg ggcacgttgt tctcgttcag gaagatgatc agcttcttgg cgtgctcgta
360ctcctcggcg gcgtggtcga acaggaacag gccggcgccg tccaggctgt gggtgtagca
420ccagctgctc atgctcatgt acaggttgct gctctgcatc tccttgttca cctgctcgtt
480cagcagcttg atgatgtcgc ctccggaatc gatcttctcc cggttcagct tgctttcctc
540gctgtacttg gggtagtcgt aggtgccgtt cttcacgctt tccatgcact cgtcgttgca
600cttgtggtag aactcgaagc agccgttgcc gatctctttg gcgttgttct tcagctggct
660cttcactttc tcgtacaggt tcttcacgtt gctgtcgtgg aagtccaggg tcctctcatt
720ttccaggagg accaggagtt cggcgttgta ggtccagatg tcgatgaagc cgtcgtccac
780cttcttgttc aggttttcca tccgccgttc cagcttgttg aactctttgc ccacggcggt
840gaactgggtg ttcatcttct cgatcacgct gttcactttg ttggtgatgc cgttgatggc
900gttctgggtg ctcttctgat cggcggcata gccagatccc tgctcattct ggtggtggta
960gccgtaccac ccatccacca tgcctgtcca tccgccctcg ataaagccgg caatggcgcc
1020aaacaggcct ctgctctgga tgctggggat attcctcagg ccggtcacca ttctcagctt
1080ggcgctccgc acgtatttag gacactcgcc gatggtcaca gggtgcacat tctggaaggg
1140caggctgcta ttgatagctc cctgtggtgt ctgacacttg gcgtcacact tgtccatggg
1200ggcgttgctg ttgatgatgc cgctgccaaa gcctctgctc agggcaaagg cgtatctagg
1260ggcgatcaga ttgccgttgg cctcgaagat gatggtgtcg ccaggttcca gcagggtcca
1320gtagtagttg atcctgcctt cctggtcccg cactttgggt cttttggcga tctcgggggt
1380gaactttctg ctgtagtggc tggacaccac ggacacgtag gcattctcgg tgtggtacag
1440ggccttctgg atgccgatgt taggagggtg gtgcactccc cagaggacca gcacttcttt
1500ttctttgttg ttggcgtagc tcttgctcag gttggggtac aggccgttct tgccggtcag
1560ccacagcagg tttctgtaga agctgctctc gccattgtgg ctacaagagg cgctcacgcc
1620ggtcactgta tgattgggcc agctgctctc tttggggaag atctcgaatc tctcgaagct
1680ggacacgctg gacagctgtt ctctcagttc ctcgtaatcg gcgaagtggc cagggtagca
1740ggtgccattc tcagggttgg gcttctccac gatgtaggac caggactctt tggaaatcag
1800cagctcgcac tcaggattgc ccagaatcca gccggccacg ctacaatttc ccagctgcag
1860aggggcaatg cctttcagca gacacagctt gccgttgtgg ctgttttcca gcaggttcac
1920gctgtgggtc acggtcacgt tcttttccag cacggtatcc acggtgtcgg tgctattgtt
1980ggcgtggtag ccgatacaga tggtatcggc gtaggtggcg gtaaaggtac acagcagcac
2040cagcagcttc actttcat
2058672112DNAArtificial SequenceSynthetic 67atgaaggcca tcatcgtgct
gctgatggtg gtcacaagca acgccgatag aatctgtacc 60ggcatcacca gcagcaatag
ccctcacgtc gtgaaaacag ctacacaggg cgaagtgaat 120gtgaccggcg tgatccctct
gaccacaaca cctacaaaga gccacttcgc caatctgaag 180ggcacagaga caagaggcaa
gctgtgtccc aagtgcctga attgcacaga tctggatgtg 240gctctgggca gacctaagtg
tacaggcaaa atccctagcg ccagagtgtc cattctgcat 300gaagtgcgac ctgtgaccag
cggctgtttt cctattatgc acgaccggac caagatcaga 360cagctgccta atctgctgag
aggctacgag cacatcagac tgagcaccca caatgtgatc 420aacgccgaaa atgctcctgg
cggcccttat aagatcggca catctggcag ctgccccaac 480attacaaatg gcaatggctt
ctttgccacc atggcttggg ccgtgcctaa gaacgataag 540aacaagaccg ccaccaaccc
cctgacaatc gaggtgccat atatctgtac agagggcgag 600gatcagatca ccgtgtgggg
atttcacagc gacaacgaaa cacagatggc caagctgtac 660ggcgatagca agcctcagaa
gtttaccagc tctgccaatg gcgtgaccac acactatgtg 720tctcagatcg gcggcttccc
taatcagaca gaagatggcg gactgcctca gtctggaaga 780atcgtggtgg attacatggt
gcagaagtct ggcaagaccg gcaccatcac atatcagaga 840ggaatcctgc tgccccagaa
agtgtggtgc gcttctggaa gatccaaagt gatcaagggc 900agcctgcctc tgattggaga
agccgattgt ctgcacgaga aatacggcgg cctgaacaag 960agcaagcctt actatacagg
cgagcacgcc aaggccatcg gcaattgtcc tatttgggtc 1020aagacccctc tgaagctggc
caatggcaca aagtatagac ctccagccaa gctgctgaaa 1080gagagaggct tttttggagc
tatcgccggc tttctggaag gcggatggga gggaatgatt 1140gctggatggc atggctacac
atctcatggc gcacatggcg tggcagtggc tgctgatctg 1200aaatctacac aggaagccat
caacaagatc accaagaacc tgaacagcct gagcgagctg 1260gaagtgaaga atctgcagag
actgtctggc gccatggacg aactgcacaa tgagatcctg 1320gaactggacg agaaggtgga
cgatctgaga gccgatacaa tcagcagcca gattgaactg 1380gctgtgctgc tgtctaacga
gggcatcatc aatagcgagg acgaacatct gctggccctg 1440gaaagaaagc tgaagaagat
gctgggacct agcgccgtgg aaatcggcaa tggatgcttt 1500gagacaaagc acaagtgcaa
ccagacctgc ctggatagaa ttgccgccgg aacatttgat 1560gccggcgagt tttctctgcc
caccttcgat agcctgaata tcacatccgg aggcgacatc 1620atcaagctgc tgaacgagca
ggtgaacaag gagatgcaga gcagcaacct gtacatgagc 1680atgagcagct ggtgctacac
ccacagcctg gacggcgccg gcctgttcct gttcgaccac 1740gccgccgagg agtacgagca
cgccaagaag ctgatcatct tcctgaacga gaacaacgtg 1800cccgtgcagc tgaccagcat
cagcgccccc gagcacaagt tcgagggcct gacccagatc 1860ttccagaagg cctacgagca
cgagcagcac atcagcgaga gcatcaacaa catcgtggac 1920cacgccatca agagcaagga
ccacgccacc ttcaacttcc tgcagtggta cgtggccgag 1980cagcacgagg aggaggtgct
gttcaaggac atcctggaca agatcgagct gatcggcaac 2040gagaaccacg gcctgtacct
ggccgaccag tacgtgaagg gcatcgccaa gagcaggaag 2100agcggatcct ag
211268703PRTArtificial
SequenceSynthetic 68Met Lys Ala Ile Ile Val Leu Leu Met Val Val Thr Ser
Asn Ala Asp 1 5 10 15
Arg Ile Cys Thr Gly Ile Thr Ser Ser Asn Ser Pro His Val Val Lys
20 25 30 Thr Ala Thr Gln
Gly Glu Val Asn Val Thr Gly Val Ile Pro Leu Thr 35
40 45 Thr Thr Pro Thr Lys Ser His Phe Ala
Asn Leu Lys Gly Thr Glu Thr 50 55
60 Arg Gly Lys Leu Cys Pro Lys Cys Leu Asn Cys Thr Asp
Leu Asp Val 65 70 75
80 Ala Leu Gly Arg Pro Lys Cys Thr Gly Lys Ile Pro Ser Ala Arg Val
85 90 95 Ser Ile Leu His
Glu Val Arg Pro Val Thr Ser Gly Cys Phe Pro Ile 100
105 110 Met His Asp Arg Thr Lys Ile Arg Gln
Leu Pro Asn Leu Leu Arg Gly 115 120
125 Tyr Glu His Ile Arg Leu Ser Thr His Asn Val Ile Asn Ala
Glu Asn 130 135 140
Ala Pro Gly Gly Pro Tyr Lys Ile Gly Thr Ser Gly Ser Cys Pro Asn 145
150 155 160 Ile Thr Asn Gly Asn
Gly Phe Phe Ala Thr Met Ala Trp Ala Val Pro 165
170 175 Lys Asn Asp Lys Asn Lys Thr Ala Thr Asn
Pro Leu Thr Ile Glu Val 180 185
190 Pro Tyr Ile Cys Thr Glu Gly Glu Asp Gln Ile Thr Val Trp Gly
Phe 195 200 205 His
Ser Asp Asn Glu Thr Gln Met Ala Lys Leu Tyr Gly Asp Ser Lys 210
215 220 Pro Gln Lys Phe Thr Ser
Ser Ala Asn Gly Val Thr Thr His Tyr Val 225 230
235 240 Ser Gln Ile Gly Gly Phe Pro Asn Gln Thr Glu
Asp Gly Gly Leu Pro 245 250
255 Gln Ser Gly Arg Ile Val Val Asp Tyr Met Val Gln Lys Ser Gly Lys
260 265 270 Thr Gly
Thr Ile Thr Tyr Gln Arg Gly Ile Leu Leu Pro Gln Lys Val 275
280 285 Trp Cys Ala Ser Gly Arg Ser
Lys Val Ile Lys Gly Ser Leu Pro Leu 290 295
300 Ile Gly Glu Ala Asp Cys Leu His Glu Lys Tyr Gly
Gly Leu Asn Lys 305 310 315
320 Ser Lys Pro Tyr Tyr Thr Gly Glu His Ala Lys Ala Ile Gly Asn Cys
325 330 335 Pro Ile Trp
Val Lys Thr Pro Leu Lys Leu Ala Asn Gly Thr Lys Tyr 340
345 350 Arg Pro Pro Ala Lys Leu Leu Lys
Glu Arg Gly Phe Phe Gly Ala Ile 355 360
365 Ala Gly Phe Leu Glu Gly Gly Trp Glu Gly Met Ile Ala
Gly Trp His 370 375 380
Gly Tyr Thr Ser His Gly Ala His Gly Val Ala Val Ala Ala Asp Leu 385
390 395 400 Lys Ser Thr Gln
Glu Ala Ile Asn Lys Ile Thr Lys Asn Leu Asn Ser 405
410 415 Leu Ser Glu Leu Glu Val Lys Asn Leu
Gln Arg Leu Ser Gly Ala Met 420 425
430 Asp Glu Leu His Asn Glu Ile Leu Glu Leu Asp Glu Lys Val
Asp Asp 435 440 445
Leu Arg Ala Asp Thr Ile Ser Ser Gln Ile Glu Leu Ala Val Leu Leu 450
455 460 Ser Asn Glu Gly Ile
Ile Asn Ser Glu Asp Glu His Leu Leu Ala Leu 465 470
475 480 Glu Arg Lys Leu Lys Lys Met Leu Gly Pro
Ser Ala Val Glu Ile Gly 485 490
495 Asn Gly Cys Phe Glu Thr Lys His Lys Cys Asn Gln Thr Cys Leu
Asp 500 505 510 Arg
Ile Ala Ala Gly Thr Phe Asp Ala Gly Glu Phe Ser Leu Pro Thr 515
520 525 Phe Asp Ser Leu Asn Ile
Thr Ser Gly Gly Asp Ile Ile Lys Leu Leu 530 535
540 Asn Glu Gln Val Asn Lys Glu Met Gln Ser Ser
Asn Leu Tyr Met Ser 545 550 555
560 Met Ser Ser Trp Cys Tyr Thr His Ser Leu Asp Gly Ala Gly Leu Phe
565 570 575 Leu Phe
Asp His Ala Ala Glu Glu Tyr Glu His Ala Lys Lys Leu Ile 580
585 590 Ile Phe Leu Asn Glu Asn Asn
Val Pro Val Gln Leu Thr Ser Ile Ser 595 600
605 Ala Pro Glu His Lys Phe Glu Gly Leu Thr Gln Ile
Phe Gln Lys Ala 610 615 620
Tyr Glu His Glu Gln His Ile Ser Glu Ser Ile Asn Asn Ile Val Asp 625
630 635 640 His Ala Ile
Lys Ser Lys Asp His Ala Thr Phe Asn Phe Leu Gln Trp 645
650 655 Tyr Val Ala Glu Gln His Glu Glu
Glu Val Leu Phe Lys Asp Ile Leu 660 665
670 Asp Lys Ile Glu Leu Ile Gly Asn Glu Asn His Gly Leu
Tyr Leu Ala 675 680 685
Asp Gln Tyr Val Lys Gly Ile Ala Lys Ser Arg Lys Ser Gly Ser 690
695 700 692112DNAArtificial
SequenceSynthetic 69ctaggatccg ctcttcctgc tcttggcgat gcccttcacg
tactggtcgg ccaggtacag 60gccgtggttc tcgttgccga tcagctcgat cttgtccagg
atgtccttga acagcacctc 120ctcctcgtgc tgctcggcca cgtaccactg caggaagttg
aaggtggcgt ggtccttgct 180cttgatggcg tggtccacga tgttgttgat gctctcgctg
atgtgctgct cgtgctcgta 240ggccttctgg aagatctggg tcaggccctc gaacttgtgc
tcgggggcgc tgatgctggt 300cagctgcacg ggcacgttgt tctcgttcag gaagatgatc
agcttcttgg cgtgctcgta 360ctcctcggcg gcgtggtcga acaggaacag gccggcgccg
tccaggctgt gggtgtagca 420ccagctgctc atgctcatgt acaggttgct gctctgcatc
tccttgttca cctgctcgtt 480cagcagcttg atgatgtcgc ctccggatgt gatattcagg
ctatcgaagg tgggcagaga 540aaactcgccg gcatcaaatg ttccggcggc aattctatcc
aggcaggtct ggttgcactt 600gtgctttgtc tcaaagcatc cattgccgat ttccacggcg
ctaggtccca gcatcttctt 660cagctttctt tccagggcca gcagatgttc gtcctcgcta
ttgatgatgc cctcgttaga 720cagcagcaca gccagttcaa tctggctgct gattgtatcg
gctctcagat cgtccacctt 780ctcgtccagt tccaggatct cattgtgcag ttcgtccatg
gcgccagaca gtctctgcag 840attcttcact tccagctcgc tcaggctgtt caggttcttg
gtgatcttgt tgatggcttc 900ctgtgtagat ttcagatcag cagccactgc cacgccatgt
gcgccatgag atgtgtagcc 960atgccatcca gcaatcattc cctcccatcc gccttccaga
aagccggcga tagctccaaa 1020aaagcctctc tctttcagca gcttggctgg aggtctatac
tttgtgccat tggccagctt 1080cagaggggtc ttgacccaaa taggacaatt gccgatggcc
ttggcgtgct cgcctgtata 1140gtaaggcttg ctcttgttca ggccgccgta tttctcgtgc
agacaatcgg cttctccaat 1200cagaggcagg ctgcccttga tcactttgga tcttccagaa
gcgcaccaca ctttctgggg 1260cagcaggatt cctctctgat atgtgatggt gccggtcttg
ccagacttct gcaccatgta 1320atccaccacg attcttccag actgaggcag tccgccatct
tctgtctgat tagggaagcc 1380gccgatctga gacacatagt gtgtggtcac gccattggca
gagctggtaa acttctgagg 1440cttgctatcg ccgtacagct tggccatctg tgtttcgttg
tcgctgtgaa atccccacac 1500ggtgatctga tcctcgccct ctgtacagat atatggcacc
tcgattgtca gggggttggt 1560ggcggtcttg ttcttatcgt tcttaggcac ggcccaagcc
atggtggcaa agaagccatt 1620gccatttgta atgttggggc agctgccaga tgtgccgatc
ttataagggc cgccaggagc 1680attttcggcg ttgatcacat tgtgggtgct cagtctgatg
tgctcgtagc ctctcagcag 1740attaggcagc tgtctgatct tggtccggtc gtgcataata
ggaaaacagc cgctggtcac 1800aggtcgcact tcatgcagaa tggacactct ggcgctaggg
attttgcctg tacacttagg 1860tctgcccaga gccacatcca gatctgtgca attcaggcac
ttgggacaca gcttgcctct 1920tgtctctgtg cccttcagat tggcgaagtg gctctttgta
ggtgttgtgg tcagagggat 1980cacgccggtc acattcactt cgccctgtgt agctgttttc
acgacgtgag ggctattgct 2040gctggtgatg ccggtacaga ttctatcggc gttgcttgtg
accaccatca gcagcacgat 2100gatggccttc at
211270837DNAInfluenza virus 70atgaaggcca agctgctggt
gctgctgtgc acctttaccg ccacctacgc cgacaccatc 60tgcattggct accacgccaa
caacagcacc gacaccgtgg ataccgtgct ggaaaagaac 120gtgaccgtga cccacagcgt
gaacctggga tccggactga gaatggtcac cggcctgaga 180aacatcccca gcatccagag
cagaggcctg tttggagcca ttgccggctt tattgagggc 240ggatggaccg gaatggtgga
tgggtggtac ggctaccacc accagaatga gcagggctct 300ggctatgccg ccgatcagaa
gtctacccag aacgccatca acggcatcac caacaaagtg 360aacagcgtga tcgagaagat
gggcggcgat cctgaatggg acagagagat caacaactac 420accagcatca tctacagcct
gatcgaggaa agccagaacc agcaggaaaa cggcacaggc 480ggcggatctg gaattgtgca
gcagcagaac aacctgctga gagccattga ggcccagcag 540catctgctgc agctgacagt
gtggggcatc aagcagctgc agacctacaa tgccgagctg 600ctggtcctcc tggaaaacga
gagaaccctg gacttccacg acagcaacgt gaagaacctg 660tacgagaaag tgaagtccca
gctgaagaac aacgccaaag agatcggcaa cggctgcttc 720gagttctacc acaagtgcaa
caacgagtgc atggaaagcg tgaagaacgg cacctacgac 780taccccaagt acagcgagga
aagcaagctg aacagagaga agatcgactc cggaggc 83771279PRTInfluenza virus
71Met Lys Ala Lys Leu Leu Val Leu Leu Cys Thr Phe Thr Ala Thr Tyr 1
5 10 15 Ala Asp Thr Ile
Cys Ile Gly Tyr His Ala Asn Asn Ser Thr Asp Thr 20
25 30 Val Asp Thr Val Leu Glu Lys Asn Val
Thr Val Thr His Ser Val Asn 35 40
45 Leu Gly Ser Gly Leu Arg Met Val Thr Gly Leu Arg Asn Ile
Pro Ser 50 55 60
Ile Gln Ser Arg Gly Leu Phe Gly Ala Ile Ala Gly Phe Ile Glu Gly 65
70 75 80 Gly Trp Thr Gly Met
Val Asp Gly Trp Tyr Gly Tyr His His Gln Asn 85
90 95 Glu Gln Gly Ser Gly Tyr Ala Ala Asp Gln
Lys Ser Thr Gln Asn Ala 100 105
110 Ile Asn Gly Ile Thr Asn Lys Val Asn Ser Val Ile Glu Lys Met
Gly 115 120 125 Gly
Asp Pro Glu Trp Asp Arg Glu Ile Asn Asn Tyr Thr Ser Ile Ile 130
135 140 Tyr Ser Leu Ile Glu Glu
Ser Gln Asn Gln Gln Glu Asn Gly Thr Gly 145 150
155 160 Gly Gly Ser Gly Ile Val Gln Gln Gln Asn Asn
Leu Leu Arg Ala Ile 165 170
175 Glu Ala Gln Gln His Leu Leu Gln Leu Thr Val Trp Gly Ile Lys Gln
180 185 190 Leu Gln
Thr Tyr Asn Ala Glu Leu Leu Val Leu Leu Glu Asn Glu Arg 195
200 205 Thr Leu Asp Phe His Asp Ser
Asn Val Lys Asn Leu Tyr Glu Lys Val 210 215
220 Lys Ser Gln Leu Lys Asn Asn Ala Lys Glu Ile Gly
Asn Gly Cys Phe 225 230 235
240 Glu Phe Tyr His Lys Cys Asn Asn Glu Cys Met Glu Ser Val Lys Asn
245 250 255 Gly Thr Tyr
Asp Tyr Pro Lys Tyr Ser Glu Glu Ser Lys Leu Asn Arg 260
265 270 Glu Lys Ile Asp Ser Gly Gly
275 72837DNAInfluenza virus 72gcctccggag tcgatcttct
ctctgttcag cttgctttcc tcgctgtact tggggtagtc 60gtaggtgccg ttcttcacgc
tttccatgca ctcgttgttg cacttgtggt agaactcgaa 120gcagccgttg ccgatctctt
tggcgttgtt cttcagctgg gacttcactt tctcgtacag 180gttcttcacg ttgctgtcgt
ggaagtccag ggttctctcg ttttccagga ggaccagcag 240ctcggcattg taggtctgca
gctgcttgat gccccacact gtcagctgca gcagatgctg 300ctgggcctca atggctctca
gcaggttgtt ctgctgctgc acaattccag atccgccgcc 360tgtgccgttt tcctgctggt
tctggctttc ctcgatcagg ctgtagatga tgctggtgta 420gttgttgatc tctctgtccc
attcaggatc gccgcccatc ttctcgatca cgctgttcac 480tttgttggtg atgccgttga
tggcgttctg ggtagacttc tgatcggcgg catagccaga 540gccctgctca ttctggtggt
ggtagccgta ccacccatcc accattccgg tccatccgcc 600ctcaataaag ccggcaatgg
ctccaaacag gcctctgctc tggatgctgg ggatgtttct 660caggccggtg accattctca
gtccggatcc caggttcacg ctgtgggtca cggtcacgtt 720cttttccagc acggtatcca
cggtgtcggt gctgttgttg gcgtggtagc caatgcagat 780ggtgtcggcg taggtggcgg
taaaggtgca cagcagcacc agcagcttgg ccttcat 83773828DNAInfluenza virus
73atgaaggcta tcctggtggt gctgctgtac acctttgcca ccgccaatgc cgacaccctg
60tgtattggct accacgccaa caacagcacc gacaccgtgg ataccgtgct ggaaaagaac
120gtgaccgtga cccacagcgt gaacctgggc tccggcctga gactggccac cggcctgaga
180aacatcccca gcattcagag cagaggcctg tttggagcca ttgccggctt tattgagggc
240ggatggaccg gaatggtgga tgggtggtac ggctaccacc accagaatga gcagggctct
300ggctatgccg ccgacctgaa gtctacccag aacgccatcg acgagatcac caacaaagtg
360aacagcgtga tcgagaagat gggcggctgg gacccatggg acagagagat caacaactac
420accagcatca tctacagcct gatcgaggaa agccagaacc agcaggaaaa cggcacaggc
480ggcggatctg gaattgtgca gcagcagaac aacctgctga gagccattga ggcccagcag
540catctgctgc agctgacagt gtggggcatc aagcagctgc agacctacaa cgccgagctg
600ctggtgctgc tcgagaatga gagaaccctg gactaccacg acagcaacgt gaagaacctg
660tacgagaaag tgcggagcca gctgaagaac aacgccaaag agatcggcaa cggctgcttc
720gagttctacc acaagtgcga caatacctgc atggaaagcg tgaagaacgg cacctacgac
780taccccaagt acagcgagga agccaagctg aaccgggaag agatcgat
82874276PRTInfluenza virus 74Met Lys Ala Ile Leu Val Val Leu Leu Tyr Thr
Phe Ala Thr Ala Asn 1 5 10
15 Ala Asp Thr Leu Cys Ile Gly Tyr His Ala Asn Asn Ser Thr Asp Thr
20 25 30 Val Asp
Thr Val Leu Glu Lys Asn Val Thr Val Thr His Ser Val Asn 35
40 45 Leu Gly Ser Gly Leu Arg Leu
Ala Thr Gly Leu Arg Asn Ile Pro Ser 50 55
60 Ile Gln Ser Arg Gly Leu Phe Gly Ala Ile Ala Gly
Phe Ile Glu Gly 65 70 75
80 Gly Trp Thr Gly Met Val Asp Gly Trp Tyr Gly Tyr His His Gln Asn
85 90 95 Glu Gln Gly
Ser Gly Tyr Ala Ala Asp Leu Lys Ser Thr Gln Asn Ala 100
105 110 Ile Asp Glu Ile Thr Asn Lys Val
Asn Ser Val Ile Glu Lys Met Gly 115 120
125 Gly Trp Asp Pro Trp Asp Arg Glu Ile Asn Asn Tyr Thr
Ser Ile Ile 130 135 140
Tyr Ser Leu Ile Glu Glu Ser Gln Asn Gln Gln Glu Asn Gly Thr Gly 145
150 155 160 Gly Gly Ser Gly
Ile Val Gln Gln Gln Asn Asn Leu Leu Arg Ala Ile 165
170 175 Glu Ala Gln Gln His Leu Leu Gln Leu
Thr Val Trp Gly Ile Lys Gln 180 185
190 Leu Gln Thr Tyr Asn Ala Glu Leu Leu Val Leu Leu Glu Asn
Glu Arg 195 200 205
Thr Leu Asp Tyr His Asp Ser Asn Val Lys Asn Leu Tyr Glu Lys Val 210
215 220 Arg Ser Gln Leu Lys
Asn Asn Ala Lys Glu Ile Gly Asn Gly Cys Phe 225 230
235 240 Glu Phe Tyr His Lys Cys Asp Asn Thr Cys
Met Glu Ser Val Lys Asn 245 250
255 Gly Thr Tyr Asp Tyr Pro Lys Tyr Ser Glu Glu Ala Lys Leu Asn
Arg 260 265 270 Glu
Glu Ile Asp 275 75828DNAInfluenza virus 75atcgatctct
tcccggttca gcttggcttc ctcgctgtac ttggggtagt cgtaggtgcc 60gttcttcacg
ctttccatgc aggtattgtc gcacttgtgg tagaactcga agcagccgtt 120gccgatctct
ttggcgttgt tcttcagctg gctccgcact ttctcgtaca ggttcttcac 180gttgctgtcg
tggtagtcca gggttctctc attctcgagc agcaccagca gctcggcgtt 240gtaggtctgc
agctgcttga tgccccacac tgtcagctgc agcagatgct gctgggcctc 300aatggctctc
agcaggttgt tctgctgctg cacaattcca gatccgccgc ctgtgccgtt 360ttcctgctgg
ttctggcttt cctcgatcag gctgtagatg atgctggtgt agttgttgat 420ctctctgtcc
catgggtccc agccgcccat cttctcgatc acgctgttca ctttgttggt 480gatctcgtcg
atggcgttct gggtagactt caggtcggcg gcatagccag agccctgctc 540attctggtgg
tggtagccgt accacccatc caccattccg gtccatccgc cctcaataaa 600gccggcaatg
gctccaaaca ggcctctgct ctgaatgctg gggatgtttc tcaggccggt 660ggccagtctc
aggccggagc ccaggttcac gctgtgggtc acggtcacgt tcttttccag 720cacggtatcc
acggtgtcgg tgctgttgtt ggcgtggtag ccaatacaca gggtgtcggc 780attggcggtg
gcaaaggtgt acagcagcac caccaggata gccttcat
82876822DNAInfluenza virus 76atggccatca tctacctgat tctgctgttt acagccgtca
gaggcgatca gatctgtatt 60ggctaccacg ccaacaatag caccgagaaa gtggatacca
tcctggaaag aaatgtgaca 120gtgacacacg ccaaggatat tggatcagga ctggtgctgg
ctacaggact gagaaatgtg 180cctcagattg agagcagagg cctgtttgga gccattgctg
gctttattga aggcggatgg 240cagggaatga ttgatgggtg gtacggctac caccactcta
atgatcaggg atctggatat 300gccgccgaca aagaatctac acagaaagcc ttcgacggca
tcaccaacaa agtgaatagc 360gtgatcgaga agatgggcgg agatcccgaa tgggacagag
agatcaacaa ctacaccagc 420atcatctaca gcctgatcga ggaaagccag aatcagcagg
aaaatggaac aggcggagga 480tctggaattg tgcagcagca gaacaatctg ctgagagcta
ttgaagctca gcagcatctg 540ctgaatctga cagtgtgggg aatcaaacag ctgcagacat
acaatgctga gctgctggtg 600ctgatggaaa atgagagaac cctggacttc cacgacagca
atgtgaagaa cctgtacgac 660aaagtgcgga tgcagctgag agacaatgtg aaagaactgg
gcaatggctg cttcgagttc 720taccacaagt gcgacgatga gtgtatgaac agcgtgaaga
acggcaccta cgactaccct 780aagtacgagg aagagagcaa gctgaacaga aatgagatca
ag 82277274PRTInfluenza virus 77Met Ala Ile Ile Tyr
Leu Ile Leu Leu Phe Thr Ala Val Arg Gly Asp 1 5
10 15 Gln Ile Cys Ile Gly Tyr His Ala Asn Asn
Ser Thr Glu Lys Val Asp 20 25
30 Thr Ile Leu Glu Arg Asn Val Thr Val Thr His Ala Lys Asp Ile
Gly 35 40 45 Ser
Gly Leu Val Leu Ala Thr Gly Leu Arg Asn Val Pro Gln Ile Glu 50
55 60 Ser Arg Gly Leu Phe Gly
Ala Ile Ala Gly Phe Ile Glu Gly Gly Trp 65 70
75 80 Gln Gly Met Ile Asp Gly Trp Tyr Gly Tyr His
His Ser Asn Asp Gln 85 90
95 Gly Ser Gly Tyr Ala Ala Asp Lys Glu Ser Thr Gln Lys Ala Phe Asp
100 105 110 Gly Ile
Thr Asn Lys Val Asn Ser Val Ile Glu Lys Met Gly Gly Asp 115
120 125 Pro Glu Trp Asp Arg Glu Ile
Asn Asn Tyr Thr Ser Ile Ile Tyr Ser 130 135
140 Leu Ile Glu Glu Ser Gln Asn Gln Gln Glu Asn Gly
Thr Gly Gly Gly 145 150 155
160 Ser Gly Ile Val Gln Gln Gln Asn Asn Leu Leu Arg Ala Ile Glu Ala
165 170 175 Gln Gln His
Leu Leu Asn Leu Thr Val Trp Gly Ile Lys Gln Leu Gln 180
185 190 Thr Tyr Asn Ala Glu Leu Leu Val
Leu Met Glu Asn Glu Arg Thr Leu 195 200
205 Asp Phe His Asp Ser Asn Val Lys Asn Leu Tyr Asp Lys
Val Arg Met 210 215 220
Gln Leu Arg Asp Asn Val Lys Glu Leu Gly Asn Gly Cys Phe Glu Phe 225
230 235 240 Tyr His Lys Cys
Asp Asp Glu Cys Met Asn Ser Val Lys Asn Gly Thr 245
250 255 Tyr Asp Tyr Pro Lys Tyr Glu Glu Glu
Ser Lys Leu Asn Arg Asn Glu 260 265
270 Ile Lys 78822DNAInfluenza virus 78cttgatctca tttctgttca
gcttgctctc ttcctcgtac ttagggtagt cgtaggtgcc 60gttcttcacg ctgttcatac
actcatcgtc gcacttgtgg tagaactcga agcagccatt 120gcccagttct ttcacattgt
ctctcagctg catccgcact ttgtcgtaca ggttcttcac 180attgctgtcg tggaagtcca
gggttctctc attttccatc agcaccagca gctcagcatt 240gtatgtctgc agctgtttga
ttccccacac tgtcagattc agcagatgct gctgagcttc 300aatagctctc agcagattgt
tctgctgctg cacaattcca gatcctccgc ctgttccatt 360ttcctgctga ttctggcttt
cctcgatcag gctgtagatg atgctggtgt agttgttgat 420ctctctgtcc cattcgggat
ctccgcccat cttctcgatc acgctattca ctttgttggt 480gatgccgtcg aaggctttct
gtgtagattc tttgtcggcg gcatatccag atccctgatc 540attagagtgg tggtagccgt
accacccatc aatcattccc tgccatccgc cttcaataaa 600gccagcaatg gctccaaaca
ggcctctgct ctcaatctga ggcacatttc tcagtcctgt 660agccagcacc agtcctgatc
caatatcctt ggcgtgtgtc actgtcacat ttctttccag 720gatggtatcc actttctcgg
tgctattgtt ggcgtggtag ccaatacaga tctgatcgcc 780tctgacggct gtaaacagca
gaatcaggta gatgatggcc at 82279858DNAInfluenza virus
79atgaagacca tcatcgccct gagctacatc ttctgcctgg ccctgggcca ggacctgccc
60ggcaacgaca acagcaccgc caccctgtgc ctgggccacc acgccgtgcc caacggcacc
120ctggtgaaga ccatcaccga cgaccagatc gaggtgacca acgccaccga gctgggctcc
180ggcctgaagc tggccaccgg catgcggaac gtgcccgaga agcagacccg gggcctgttc
240ggcgccatcg ccggcttcat cgagaacggc tgggagggca tgatcgacgg ctggtacggc
300ttccggcacc agaacagcga gggcaccggc caggccgccg acctgaagag cacccaggcc
360gccatcgacc agatcaacgg caagctgaac cgggtgatcg agaagaccgg cggcgatccc
420gagtgggacc gggagatcaa caactacacc agcatcatct acagcctgat cgaggagagc
480cagaaccagc aggagaacgg caccggcggc ggcagcggca tcgtgcagca gcagaacaac
540ctgctgcggg ccatcgaggc ccagcagcac ctgctgcagc tgaccgtgtg gggcatcaag
600cagctgcaga gctacaacgc cgagctgctg gtggccctgg agaaccagca caccatcgac
660ctgaccgaca gcgagatgaa caagctgttc gagaagaccc ggcggcagct gcgggagaac
720gccgaggaca tgggcaacgg ctgcttcaag atctaccaca agtgcgacaa cgcctgcatc
780gagagcatcc ggaacggcac ctacgaccac gacgtgtacc gggacgaggc cctgaacaac
840cggttccaga tcaagggc
85880286PRTInfluenza virus 80Met Lys Thr Ile Ile Ala Leu Ser Tyr Ile Phe
Cys Leu Ala Leu Gly 1 5 10
15 Gln Asp Leu Pro Gly Asn Asp Asn Ser Thr Ala Thr Leu Cys Leu Gly
20 25 30 His His
Ala Val Pro Asn Gly Thr Leu Val Lys Thr Ile Thr Asp Asp 35
40 45 Gln Ile Glu Val Thr Asn Ala
Thr Glu Leu Gly Ser Gly Leu Lys Leu 50 55
60 Ala Thr Gly Met Arg Asn Val Pro Glu Lys Gln Thr
Arg Gly Leu Phe 65 70 75
80 Gly Ala Ile Ala Gly Phe Ile Glu Asn Gly Trp Glu Gly Met Ile Asp
85 90 95 Gly Trp Tyr
Gly Phe Arg His Gln Asn Ser Glu Gly Thr Gly Gln Ala 100
105 110 Ala Asp Leu Lys Ser Thr Gln Ala
Ala Ile Asp Gln Ile Asn Gly Lys 115 120
125 Leu Asn Arg Val Ile Glu Lys Thr Gly Gly Asp Pro Glu
Trp Asp Arg 130 135 140
Glu Ile Asn Asn Tyr Thr Ser Ile Ile Tyr Ser Leu Ile Glu Glu Ser 145
150 155 160 Gln Asn Gln Gln
Glu Asn Gly Thr Gly Gly Gly Ser Gly Ile Val Gln 165
170 175 Gln Gln Asn Asn Leu Leu Arg Ala Ile
Glu Ala Gln Gln His Leu Leu 180 185
190 Gln Leu Thr Val Trp Gly Ile Lys Gln Leu Gln Ser Tyr Asn
Ala Glu 195 200 205
Leu Leu Val Ala Leu Glu Asn Gln His Thr Ile Asp Leu Thr Asp Ser 210
215 220 Glu Met Asn Lys Leu
Phe Glu Lys Thr Arg Arg Gln Leu Arg Glu Asn 225 230
235 240 Ala Glu Asp Met Gly Asn Gly Cys Phe Lys
Ile Tyr His Lys Cys Asp 245 250
255 Asn Ala Cys Ile Glu Ser Ile Arg Asn Gly Thr Tyr Asp His Asp
Val 260 265 270 Tyr
Arg Asp Glu Ala Leu Asn Asn Arg Phe Gln Ile Lys Gly 275
280 285 81858DNAInfluenza virus 81gcccttgatc
tggaaccggt tgttcagggc ctcgtcccgg tacacgtcgt ggtcgtaggt 60gccgttccgg
atgctctcga tgcaggcgtt gtcgcacttg tggtagatct tgaagcagcc 120gttgcccatg
tcctcggcgt tctcccgcag ctgccgccgg gtcttctcga acagcttgtt 180catctcgctg
tcggtcaggt cgatggtgtg ctggttctcc agggccacca gcagctcggc 240gttgtagctc
tgcagctgct tgatgcccca cacggtcagc tgcagcaggt gctgctgggc 300ctcgatggcc
cgcagcaggt tgttctgctg ctgcacgatg ccgctgccgc cgccggtgcc 360gttctcctgc
tggttctggc tctcctcgat caggctgtag atgatgctgg tgtagttgtt 420gatctcccgg
tcccactcgg gatcgccgcc ggtcttctcg atcacccggt tcagcttgcc 480gttgatctgg
tcgatggcgg cctgggtgct cttcaggtcg gcggcctggc cggtgccctc 540gctgttctgg
tgccggaagc cgtaccagcc gtcgatcatg ccctcccagc cgttctcgat 600gaagccggcg
atggcgccga acaggccccg ggtctgcttc tcgggcacgt tccgcatgcc 660ggtggccagc
ttcaggccgg agcccagctc ggtggcgttg gtcacctcga tctggtcgtc 720ggtgatggtc
ttcaccaggg tgccgttggg cacggcgtgg tggcccaggc acagggtggc 780ggtgctgttg
tcgttgccgg gcaggtcctg gcccagggcc aggcagaaga tgtagctcag 840ggcgatgatg
gtcttcat
85882867DNAInfluenza virus 82atgaaaacca tcattgccct gagctacatc ctgtgcctgg
tgttcacaca gaagctgccc 60ggcaacgata atagcaccgc cacactgtgt ctgggacacc
acgccgtgcc taatggcacc 120atcgtgaaaa caatcaccaa cgaccagatc gaagtgacca
atgccacaga gctgggctcc 180ggcctgaagc tggccaccgg catgagaaat gtgcccgaga
agcagaccag aggcatcttt 240ggcgccattg ccggctttat cgagaatggc tgggagggaa
tggtggatgg gtggtacggc 300ttcagacacc agaatagcga gggaattgga caggccgccg
atctgaaatc tacccaggcc 360gccatcgacc agatcaacgg caagctgaac aggctgatcg
gcaagaccgg cggcgatccc 420gagtgggacc gggagatcaa caactacacc agcatcatct
acagcctgat cgaggagagc 480cagaaccagc aggagaacgg caccggcggc ggcagcggca
tcgtgcagca gcagaacaac 540ctgctgcggg ccatcgaggc ccagcagcac ctgctgcagc
tgaccgtgtg gggcatcaag 600cagctgcaga gctacaatgc cgaactgctg gtcgccctgg
aaaaccagca cacaattgat 660ctgacagaca gtgagatgaa taagctgttc gagaaaacca
agaagcagct gagagaaaac 720gccgaggaca tgggcaacgg ctgcttcaag atctaccaca
agtgcgacaa cgcctgcatc 780ggcagcatca gaaacggcac ctacgaccac gacgtgtaca
gagatgaggc cctgaacaac 840cggtttcaga tcaagggctc cggaggc
86783289PRTInfluenza virus 83Met Lys Thr Ile Ile
Ala Leu Ser Tyr Ile Leu Cys Leu Val Phe Thr 1 5
10 15 Gln Lys Leu Pro Gly Asn Asp Asn Ser Thr
Ala Thr Leu Cys Leu Gly 20 25
30 His His Ala Val Pro Asn Gly Thr Ile Val Lys Thr Ile Thr Asn
Asp 35 40 45 Gln
Ile Glu Val Thr Asn Ala Thr Glu Leu Gly Ser Gly Leu Lys Leu 50
55 60 Ala Thr Gly Met Arg Asn
Val Pro Glu Lys Gln Thr Arg Gly Ile Phe 65 70
75 80 Gly Ala Ile Ala Gly Phe Ile Glu Asn Gly Trp
Glu Gly Met Val Asp 85 90
95 Gly Trp Tyr Gly Phe Arg His Gln Asn Ser Glu Gly Ile Gly Gln Ala
100 105 110 Ala Asp
Leu Lys Ser Thr Gln Ala Ala Ile Asp Gln Ile Asn Gly Lys 115
120 125 Leu Asn Arg Leu Ile Gly Lys
Thr Gly Gly Asp Pro Glu Trp Asp Arg 130 135
140 Glu Ile Asn Asn Tyr Thr Ser Ile Ile Tyr Ser Leu
Ile Glu Glu Ser 145 150 155
160 Gln Asn Gln Gln Glu Asn Gly Thr Gly Gly Gly Ser Gly Ile Val Gln
165 170 175 Gln Gln Asn
Asn Leu Leu Arg Ala Ile Glu Ala Gln Gln His Leu Leu 180
185 190 Gln Leu Thr Val Trp Gly Ile Lys
Gln Leu Gln Ser Tyr Asn Ala Glu 195 200
205 Leu Leu Val Ala Leu Glu Asn Gln His Thr Ile Asp Leu
Thr Asp Ser 210 215 220
Glu Met Asn Lys Leu Phe Glu Lys Thr Lys Lys Gln Leu Arg Glu Asn 225
230 235 240 Ala Glu Asp Met
Gly Asn Gly Cys Phe Lys Ile Tyr His Lys Cys Asp 245
250 255 Asn Ala Cys Ile Gly Ser Ile Arg Asn
Gly Thr Tyr Asp His Asp Val 260 265
270 Tyr Arg Asp Glu Ala Leu Asn Asn Arg Phe Gln Ile Lys Gly
Ser Gly 275 280 285
Gly 84867DNAInfluenza virus 84gcctccggag cccttgatct gaaaccggtt gttcagggcc
tcatctctgt acacgtcgtg 60gtcgtaggtg ccgtttctga tgctgccgat gcaggcgttg
tcgcacttgt ggtagatctt 120gaagcagccg ttgcccatgt cctcggcgtt ttctctcagc
tgcttcttgg ttttctcgaa 180cagcttattc atctcactgt ctgtcagatc aattgtgtgc
tggttttcca gggcgaccag 240cagttcggca ttgtagctct gcagctgctt gatgccccac
acggtcagct gcagcaggtg 300ctgctgggcc tcgatggccc gcagcaggtt gttctgctgc
tgcacgatgc cgctgccgcc 360gccggtgccg ttctcctgct ggttctggct ctcctcgatc
aggctgtaga tgatgctggt 420gtagttgttg atctcccggt cccactcggg atcgccgccg
gtcttgccga tcagcctgtt 480cagcttgccg ttgatctggt cgatggcggc ctgggtagat
ttcagatcgg cggcctgtcc 540aattccctcg ctattctggt gtctgaagcc gtaccaccca
tccaccattc cctcccagcc 600attctcgata aagccggcaa tggcgccaaa gatgcctctg
gtctgcttct cgggcacatt 660tctcatgccg gtggccagct tcaggccgga gcccagctct
gtggcattgg tcacttcgat 720ctggtcgttg gtgattgttt tcacgatggt gccattaggc
acggcgtggt gtcccagaca 780cagtgtggcg gtgctattat cgttgccggg cagcttctgt
gtgaacacca ggcacaggat 840gtagctcagg gcaatgatgg ttttcat
86785837DNAInfluenza virus 85atggagaaga tcgtgctgct
gctggccatc gtgagcctgg tgaagagcga ccagatctgc 60atcggctacc acgccaacaa
cagcaccgag caggtggaca ccatcatgga gaagaacgtg 120accgtgaccc acgcccagga
catcggctcc ggcctggtgc tggccaccgg cctgcggaac 180agcccccagc gggagagccg
gcggaagaag cggggcctgt tcggcgccat cgccggcttc 240atcgagggcg gctggcaggg
catggtggac ggctggtacg gctaccacca cagcaacgag 300cagggcagcg gctacgccgc
cgacaaggag agcacccaga aggccatcga cggcgtgacc 360aacaaggtga acagcatcat
cgacaagatg ggcggcgatc ccgagtggga ccgggagatc 420aacaactaca ccagcatcat
ctacagcctg atcgaggaga gccagaacca gcaggagaac 480ggcaccggcg gcggcagcgg
catcgtgcag cagcagaaca acctgctgcg ggccatcgag 540gcccagcagc acctgctgca
gctgaccgtg tggggcatca agcagctgca gacctacaac 600gccgagctgc tggtgctgat
ggagaacgag cggaccctgg acttccacga cagcaacgtg 660aagaacctgt acgacaaggt
gcggctgcag ctgcgggaca acgccaagga gctgggcaac 720ggctgcttcg agttctacca
caagtgcgac aacgagtgca tggagagcat ccggaacggc 780acctacaact acccccagta
cagcgaggag gcccggctga agcgggagga gatcagc 83786279PRTInfluenza virus
86Met Glu Lys Ile Val Leu Leu Leu Ala Ile Val Ser Leu Val Lys Ser 1
5 10 15 Asp Gln Ile Cys
Ile Gly Tyr His Ala Asn Asn Ser Thr Glu Gln Val 20
25 30 Asp Thr Ile Met Glu Lys Asn Val Thr
Val Thr His Ala Gln Asp Ile 35 40
45 Gly Ser Gly Leu Val Leu Ala Thr Gly Leu Arg Asn Ser Pro
Gln Arg 50 55 60
Glu Ser Arg Arg Lys Lys Arg Gly Leu Phe Gly Ala Ile Ala Gly Phe 65
70 75 80 Ile Glu Gly Gly Trp
Gln Gly Met Val Asp Gly Trp Tyr Gly Tyr His 85
90 95 His Ser Asn Glu Gln Gly Ser Gly Tyr Ala
Ala Asp Lys Glu Ser Thr 100 105
110 Gln Lys Ala Ile Asp Gly Val Thr Asn Lys Val Asn Ser Ile Ile
Asp 115 120 125 Lys
Met Gly Gly Asp Pro Glu Trp Asp Arg Glu Ile Asn Asn Tyr Thr 130
135 140 Ser Ile Ile Tyr Ser Leu
Ile Glu Glu Ser Gln Asn Gln Gln Glu Asn 145 150
155 160 Gly Thr Gly Gly Gly Ser Gly Ile Val Gln Gln
Gln Asn Asn Leu Leu 165 170
175 Arg Ala Ile Glu Ala Gln Gln His Leu Leu Gln Leu Thr Val Trp Gly
180 185 190 Ile Lys
Gln Leu Gln Thr Tyr Asn Ala Glu Leu Leu Val Leu Met Glu 195
200 205 Asn Glu Arg Thr Leu Asp Phe
His Asp Ser Asn Val Lys Asn Leu Tyr 210 215
220 Asp Lys Val Arg Leu Gln Leu Arg Asp Asn Ala Lys
Glu Leu Gly Asn 225 230 235
240 Gly Cys Phe Glu Phe Tyr His Lys Cys Asp Asn Glu Cys Met Glu Ser
245 250 255 Ile Arg Asn
Gly Thr Tyr Asn Tyr Pro Gln Tyr Ser Glu Glu Ala Arg 260
265 270 Leu Lys Arg Glu Glu Ile Ser
275 87837DNAInfluenza virus 87gctgatctcc tcccgcttca
gccgggcctc ctcgctgtac tgggggtagt tgtaggtgcc 60gttccggatg ctctccatgc
actcgttgtc gcacttgtgg tagaactcga agcagccgtt 120gcccagctcc ttggcgttgt
cccgcagctg cagccgcacc ttgtcgtaca ggttcttcac 180gttgctgtcg tggaagtcca
gggtccgctc gttctccatc agcaccagca gctcggcgtt 240gtaggtctgc agctgcttga
tgccccacac ggtcagctgc agcaggtgct gctgggcctc 300gatggcccgc agcaggttgt
tctgctgctg cacgatgccg ctgccgccgc cggtgccgtt 360ctcctgctgg ttctggctct
cctcgatcag gctgtagatg atgctggtgt agttgttgat 420ctcccggtcc cactcgggat
cgccgcccat cttgtcgatg atgctgttca ccttgttggt 480cacgccgtcg atggccttct
gggtgctctc cttgtcggcg gcgtagccgc tgccctgctc 540gttgctgtgg tggtagccgt
accagccgtc caccatgccc tgccagccgc cctcgatgaa 600gccggcgatg gcgccgaaca
ggccccgctt cttccgccgg ctctcccgct gggggctgtt 660ccgcaggccg gtggccagca
ccaggccgga gccgatgtcc tgggcgtggg tcacggtcac 720gttcttctcc atgatggtgt
ccacctgctc ggtgctgttg ttggcgtggt agccgatgca 780gatctggtcg ctcttcacca
ggctcacgat ggccagcagc agcacgatct tctccat 83788837DNAInfluenza virus
88atgaaggcca tcatcgtgct gctgatggtg gtgaccagca acgccgatag aatctgcacc
60ggcatcacca gcagcaatag cccccatgtg gtgaaaacag ccacccaggg cgaagtgaat
120gtgacaggcg tgatccctct gggatcagga ctgaagctgg ccaatggcac caagtacaga
180cctcccgcca agctgctgaa agagagaggc ttctttggcg ccattgccgg atttctggaa
240ggcggctggg agggaatgat tgccggctgg cacggctata catctcatgg ggcccatggc
300gtggctgtgg ccgccgatct gaagtctacc caggaagcca tcaacaagat caccaagaac
360ctgaacagcc tgagcgagct ggaaggaggc gaccccgagt gggatcgcga aatcaacaac
420tacacatcta tcatctacag tctgattgag gaaagccaga accagcagga gaatgggact
480gggggaggct ccggaatcgt gcagcagcag aacaatctgc tgcgagccat tgaagctcag
540cagcacctgc tgcagctgac agtgtggggc atcaagcagc tgcaggggtc ccagattgaa
600ctggccgtgc tgctgtccaa cgagggcatc atcaacagcg aggatgaaca cctgctggcc
660ctggaacgga agctgaagaa gatgctgggc ccttctgccg tggagatcgg caacggctgc
720ttcgagacaa agcacaagtg caaccagacc tgcctggata gaatcgccgc tggcaccttc
780aatgccggcg agttcagcct gcctaccttc gacagcctga atatcacctc cggaggc
83789279PRTInfluenza virus 89Met Lys Ala Ile Ile Val Leu Leu Met Val Val
Thr Ser Asn Ala Asp 1 5 10
15 Arg Ile Cys Thr Gly Ile Thr Ser Ser Asn Ser Pro His Val Val Lys
20 25 30 Thr Ala
Thr Gln Gly Glu Val Asn Val Thr Gly Val Ile Pro Leu Gly 35
40 45 Ser Gly Leu Lys Leu Ala Asn
Gly Thr Lys Tyr Arg Pro Pro Ala Lys 50 55
60 Leu Leu Lys Glu Arg Gly Phe Phe Gly Ala Ile Ala
Gly Phe Leu Glu 65 70 75
80 Gly Gly Trp Glu Gly Met Ile Ala Gly Trp His Gly Tyr Thr Ser His
85 90 95 Gly Ala His
Gly Val Ala Val Ala Ala Asp Leu Lys Ser Thr Gln Glu 100
105 110 Ala Ile Asn Lys Ile Thr Lys Asn
Leu Asn Ser Leu Ser Glu Leu Glu 115 120
125 Gly Gly Asp Pro Glu Trp Asp Arg Glu Ile Asn Asn Tyr
Thr Ser Ile 130 135 140
Ile Tyr Ser Leu Ile Glu Glu Ser Gln Asn Gln Gln Glu Asn Gly Thr 145
150 155 160 Gly Gly Gly Ser
Gly Ile Val Gln Gln Gln Asn Asn Leu Leu Arg Ala 165
170 175 Ile Glu Ala Gln Gln His Leu Leu Gln
Leu Thr Val Trp Gly Ile Lys 180 185
190 Gln Leu Gln Gly Ser Gln Ile Glu Leu Ala Val Leu Leu Ser
Asn Glu 195 200 205
Gly Ile Ile Asn Ser Glu Asp Glu His Leu Leu Ala Leu Glu Arg Lys 210
215 220 Leu Lys Lys Met Leu
Gly Pro Ser Ala Val Glu Ile Gly Asn Gly Cys 225 230
235 240 Phe Glu Thr Lys His Lys Cys Asn Gln Thr
Cys Leu Asp Arg Ile Ala 245 250
255 Ala Gly Thr Phe Asn Ala Gly Glu Phe Ser Leu Pro Thr Phe Asp
Ser 260 265 270 Leu
Asn Ile Thr Ser Gly Gly 275 90837DNAInfluenza
virus 90gcctccggag gtgatattca ggctgtcgaa ggtaggcagg ctgaactcgc cggcattgaa
60ggtgccagcg gcgattctat ccaggcaggt ctggttgcac ttgtgctttg tctcgaagca
120gccgttgccg atctccacgg cagaagggcc cagcatcttc ttcagcttcc gttccagggc
180cagcaggtgt tcatcctcgc tgttgatgat gccctcgttg gacagcagca cggccagttc
240aatctgggac ccctgcagct gcttgatgcc ccacactgtc agctgcagca ggtgctgctg
300agcttcaatg gctcgcagca gattgttctg ctgctgcacg attccggagc ctcccccagt
360cccattctcc tgctggttct ggctttcctc aatcagactg tagatgatag atgtgtagtt
420gttgatttcg cgatcccact cggggtcgcc tccttccagc tcgctcaggc tgttcaggtt
480cttggtgatc ttgttgatgg cttcctgggt agacttcaga tcggcggcca cagccacgcc
540atgggcccca tgagatgtat agccgtgcca gccggcaatc attccctccc agccgccttc
600cagaaatccg gcaatggcgc caaagaagcc tctctctttc agcagcttgg cgggaggtct
660gtacttggtg ccattggcca gcttcagtcc tgatcccaga gggatcacgc ctgtcacatt
720cacttcgccc tgggtggctg ttttcaccac atgggggcta ttgctgctgg tgatgccggt
780gcagattcta tcggcgttgc tggtcaccac catcagcagc acgatgatgg ccttcat
83791867DNAInfluenza virus 91atgaaaacca taattgcgct gtcctacata ctgtgtctgg
tgtttgccca gaaactgccg 60ggcaatgaca actcaacagc cacgctctgc ttggggcacc
atgccgtccc taacgggacc 120attgtgaaaa ccattactaa cgatcagata gaggtgacta
atgccaccga gctgggctcc 180ggcttgaaac tggcgaccgg tatgcgcaat gtccccgaaa
aacagacccg cgggatattt 240ggggctatcg caggctttat cgagaatggc tgggaaggga
tggtggatgg ttggtatggt 300tttagacatc aaaactccga aggcagaggc caggctgccg
atctcaagag cacgcaggcc 360gctatagatc agatcaatgg aaagctcaac agactgatcg
ggaaaaccgg cggcgatccc 420gagtgggacc gggagatcaa caactacacc agcatcatct
acagcctgat cgaggagagc 480cagaaccagc aggagaacgg caccggcggc ggcagcggca
tcgtgcagca gcagaacaac 540ctgctgcggg ccatcgaggc ccagcagcac ctgctgcagc
tgaccgtgtg gggcatcaag 600cagctgcagt cctacaatgc cgagctgctg gtggctctgg
agaatcagca cactattgac 660ctgaccgatt cagagatgaa caaacttttt gagaagacga
agaagcagct tagagaaaat 720gcagaggaca tggggaacgg atgctttaaa atatatcata
agtgtgataa tgcctgcatc 780ggatcaatta gaaatggtac ctatgatcac gatgtttaca
gggacgaagc gctgaataac 840aggttccaga taaaaggctc cggaggc
86792289PRTInfluenza virus 92Met Lys Thr Ile Ile
Ala Leu Ser Tyr Ile Leu Cys Leu Val Phe Ala 1 5
10 15 Gln Lys Leu Pro Gly Asn Asp Asn Ser Thr
Ala Thr Leu Cys Leu Gly 20 25
30 His His Ala Val Pro Asn Gly Thr Ile Val Lys Thr Ile Thr Asn
Asp 35 40 45 Gln
Ile Glu Val Thr Asn Ala Thr Glu Leu Gly Ser Gly Leu Lys Leu 50
55 60 Ala Thr Gly Met Arg Asn
Val Pro Glu Lys Gln Thr Arg Gly Ile Phe 65 70
75 80 Gly Ala Ile Ala Gly Phe Ile Glu Asn Gly Trp
Glu Gly Met Val Asp 85 90
95 Gly Trp Tyr Gly Phe Arg His Gln Asn Ser Glu Gly Arg Gly Gln Ala
100 105 110 Ala Asp
Leu Lys Ser Thr Gln Ala Ala Ile Asp Gln Ile Asn Gly Lys 115
120 125 Leu Asn Arg Leu Ile Gly Lys
Thr Gly Gly Asp Pro Glu Trp Asp Arg 130 135
140 Glu Ile Asn Asn Tyr Thr Ser Ile Ile Tyr Ser Leu
Ile Glu Glu Ser 145 150 155
160 Gln Asn Gln Gln Glu Asn Gly Thr Gly Gly Gly Ser Gly Ile Val Gln
165 170 175 Gln Gln Asn
Asn Leu Leu Arg Ala Ile Glu Ala Gln Gln His Leu Leu 180
185 190 Gln Leu Thr Val Trp Gly Ile Lys
Gln Leu Gln Ser Tyr Asn Ala Glu 195 200
205 Leu Leu Val Ala Leu Glu Asn Gln His Thr Ile Asp Leu
Thr Asp Ser 210 215 220
Glu Met Asn Lys Leu Phe Glu Lys Thr Lys Lys Gln Leu Arg Glu Asn 225
230 235 240 Ala Glu Asp Met
Gly Asn Gly Cys Phe Lys Ile Tyr His Lys Cys Asp 245
250 255 Asn Ala Cys Ile Gly Ser Ile Arg Asn
Gly Thr Tyr Asp His Asp Val 260 265
270 Tyr Arg Asp Glu Ala Leu Asn Asn Arg Phe Gln Ile Lys Gly
Ser Gly 275 280 285
Gly 93867DNAInfluenza virus 93gcctccggag ccttttatct ggaacctgtt attcagcgct
tcgtccctgt aaacatcgtg 60atcataggta ccatttctaa ttgatccgat gcaggcatta
tcacacttat gatatatttt 120aaagcatccg ttccccatgt cctctgcatt ttctctaagc
tgcttcttcg tcttctcaaa 180aagtttgttc atctctgaat cggtcaggtc aatagtgtgc
tgattctcca gagccaccag 240cagctcggca ttgtaggact gcagctgctt gatgccccac
acggtcagct gcagcaggtg 300ctgctgggcc tcgatggccc gcagcaggtt gttctgctgc
tgcacgatgc cgctgccgcc 360gccggtgccg ttctcctgct ggttctggct ctcctcgatc
aggctgtaga tgatgctggt 420gtagttgttg atctcccggt cccactcggg atcgccgccg
gttttcccga tcagtctgtt 480gagctttcca ttgatctgat ctatagcggc ctgcgtgctc
ttgagatcgg cagcctggcc 540tctgccttcg gagttttgat gtctaaaacc ataccaacca
tccaccatcc cttcccagcc 600attctcgata aagcctgcga tagccccaaa tatcccgcgg
gtctgttttt cggggacatt 660gcgcataccg gtcgccagtt tcaagccgga gcccagctcg
gtggcattag tcacctctat 720ctgatcgtta gtaatggttt tcacaatggt cccgttaggg
acggcatggt gccccaagca 780gagcgtggct gttgagttgt cattgcccgg cagtttctgg
gcaaacacca gacacagtat 840gtaggacagc gcaattatgg ttttcat
86794837DNAInfluenza virus 94atgaaagtga agctgctggt
gctgctgtgt acctttaccg ccacctacgc cgataccatc 60tgtatcggct accacgccaa
caatagcacc gacaccgtgg ataccgtgct ggaaaagaac 120gtgaccgtga cccacagcgt
gaacctggga tcaggactga gaatggtgac cggcctgagg 180aatatcccca gcatccagag
cagaggcctg tttggcgcca ttgccggctt tatcgagggc 240ggatggacag gcatggtgga
tgggtggtac ggctaccacc accagaatga gcagggatct 300ggctatgccg ccgatcagaa
gagcacccag aacgccatca acggcatcac caacaaagtg 360aacagcgtga tcgagaagat
gggcggcgat cctgaatggg acagagagat caacaactac 420accagcatca tctacagcct
gatcgaggaa agccagaacc agcaggaaaa cggcacaggc 480ggcggatctg gaattgtgca
gcagcagaac aacctgctga gagccattga ggcccagcag 540catctgctgc agctgacagt
gtggggcatc aagcagctgc agacctacaa cgccgaactc 600ctggtcctcc tggaaaatga
gaggaccctg gacttccacg acagcaacgt gaagaacctg 660tacgagaaag tgaagagcca
gctgaagaac aacgccaaag agatcggcaa cggctgcttc 720gagttctacc acaagtgcaa
cgacgagtgc atggaaagcg tgaagaacgg cacctacgac 780taccccaagt acagcgagga
aagcaagctg aaccgggaga agatcgattc cggaggc 83795276PRTInfluenza virus
95Met Lys Val Lys Leu Leu Val Leu Leu Cys Thr Phe Thr Ala Thr Tyr 1
5 10 15 Ala Asp Thr Ile
Cys Ile Gly Tyr His Ala Asn Asn Ser Thr Asp Thr 20
25 30 Val Asp Thr Val Leu Glu Lys Asn Val
Thr Val Thr His Ser Val Asn 35 40
45 Leu Gly Ser Gly Leu Arg Met Val Thr Gly Leu Arg Asn Ile
Pro Ser 50 55 60
Ile Gln Ser Arg Gly Leu Phe Gly Ala Ile Ala Gly Phe Ile Glu Gly 65
70 75 80 Gly Trp Thr Gly Met
Val Asp Gly Trp Tyr Gly Tyr His His Gln Asn 85
90 95 Glu Gln Gly Ser Gly Tyr Ala Ala Asp Gln
Lys Ser Thr Gln Asn Ala 100 105
110 Ile Asn Gly Ile Thr Asn Lys Val Asn Ser Val Ile Glu Lys Met
Gly 115 120 125 Gly
Asp Pro Glu Trp Asp Arg Glu Ile Asn Asn Tyr Thr Ser Ile Ile 130
135 140 Tyr Ser Leu Ile Glu Glu
Ser Gln Asn Gln Gln Glu Asn Gly Thr Gly 145 150
155 160 Gly Gly Ser Gly Ile Val Gln Gln Gln Asn Asn
Leu Leu Arg Ala Ile 165 170
175 Glu Ala Gln Gln His Leu Leu Gln Leu Thr Val Trp Gly Ile Lys Gln
180 185 190 Leu Gln
Thr Tyr Asn Ala Glu Leu Leu Val Leu Leu Glu Asn Glu Arg 195
200 205 Thr Leu Asp Phe His Asp Ser
Asn Val Lys Asn Leu Tyr Glu Lys Val 210 215
220 Lys Ser Gln Leu Lys Asn Asn Ala Lys Glu Ile Gly
Asn Gly Cys Phe 225 230 235
240 Glu Phe Tyr His Lys Cys Asn Asp Glu Cys Met Glu Ser Val Lys Asn
245 250 255 Gly Thr Tyr
Asp Tyr Pro Lys Tyr Ser Glu Glu Ser Lys Leu Asn Arg 260
265 270 Glu Lys Ile Asp 275
96837DNAInfluenza virus 96gcctccggaa tcgatcttct cccggttcag cttgctttcc
tcgctgtact tggggtagtc 60gtaggtgccg ttcttcacgc tttccatgca ctcgtcgttg
cacttgtggt agaactcgaa 120gcagccgttg ccgatctctt tggcgttgtt cttcagctgg
ctcttcactt tctcgtacag 180gttcttcacg ttgctgtcgt ggaagtccag ggtcctctca
ttttccagga ggaccaggag 240ttcggcgttg taggtctgca gctgcttgat gccccacact
gtcagctgca gcagatgctg 300ctgggcctca atggctctca gcaggttgtt ctgctgctgc
acaattccag atccgccgcc 360tgtgccgttt tcctgctggt tctggctttc ctcgatcagg
ctgtagatga tgctggtgta 420gttgttgatc tctctgtccc attcaggatc gccgcccatc
ttctcgatca cgctgttcac 480tttgttggtg atgccgttga tggcgttctg ggtgctcttc
tgatcggcgg catagccaga 540tccctgctca ttctggtggt ggtagccgta ccacccatcc
accatgcctg tccatccgcc 600ctcgataaag ccggcaatgg cgccaaacag gcctctgctc
tggatgctgg ggatattcct 660caggccggtc accattctca gtcctgatcc caggttcacg
ctgtgggtca cggtcacgtt 720cttttccagc acggtatcca cggtgtcggt gctattgttg
gcgtggtagc cgatacagat 780ggtatcggcg taggtggcgg taaaggtaca cagcagcacc
agcagcttca ctttcat 83797837DNAInfluenza virus 97atgaaggcca
tcatcgtgct gctgatggtg gtcacaagca acgccgatag aatctgtacc 60ggcatcacca
gcagcaatag ccctcacgtc gtgaaaacag ctacacaggg cgaagtgaat 120gtgaccggcg
tgatccctct gggatcagga ctgaagctgg ccaatggcac aaagtataga 180cctccagcca
agctgctgaa agagagaggc ttttttggag ctatcgccgg ctttctggaa 240ggcggatggg
agggaatgat tgctggatgg catggctaca catctcatgg cgcacatggc 300gtggcagtgg
ctgctgatct gaaatctaca caggaagcca tcaacaagat caccaagaac 360ctgaacagcc
tgagcgagct ggaaggaggc gaccccgagt gggatcgcga aatcaacaac 420tacacatcta
tcatctacag tctgattgag gaaagccaga accagcagga gaatgggact 480gggggaggct
ccggaatcgt gcagcagcag aacaatctgc tgcgagccat tgaagctcag 540cagcacctgc
tgcagctgac agtgtggggc atcaagcagc tgcaggggag ccagattgaa 600ctggctgtgc
tgctgtctaa cgagggcatc atcaatagcg aggacgaaca tctgctggcc 660ctggaaagaa
agctgaagaa gatgctggga cctagcgccg tggaaatcgg caatggatgc 720tttgagacaa
agcacaagtg caaccagacc tgcctggata gaattgccgc cggaacattt 780gatgccggcg
agttttctct gcccaccttc gatagcctga atatcacatc cggaggc
83798279PRTInfluenza virus 98Met Lys Ala Ile Ile Val Leu Leu Met Val Val
Thr Ser Asn Ala Asp 1 5 10
15 Arg Ile Cys Thr Gly Ile Thr Ser Ser Asn Ser Pro His Val Val Lys
20 25 30 Thr Ala
Thr Gln Gly Glu Val Asn Val Thr Gly Val Ile Pro Leu Gly 35
40 45 Ser Gly Leu Lys Leu Ala Asn
Gly Thr Lys Tyr Arg Pro Pro Ala Lys 50 55
60 Leu Leu Lys Glu Arg Gly Phe Phe Gly Ala Ile Ala
Gly Phe Leu Glu 65 70 75
80 Gly Gly Trp Glu Gly Met Ile Ala Gly Trp His Gly Tyr Thr Ser His
85 90 95 Gly Ala His
Gly Val Ala Val Ala Ala Asp Leu Lys Ser Thr Gln Glu 100
105 110 Ala Ile Asn Lys Ile Thr Lys Asn
Leu Asn Ser Leu Ser Glu Leu Glu 115 120
125 Gly Gly Asp Pro Glu Trp Asp Arg Glu Ile Asn Asn Tyr
Thr Ser Ile 130 135 140
Ile Tyr Ser Leu Ile Glu Glu Ser Gln Asn Gln Gln Glu Asn Gly Thr 145
150 155 160 Gly Gly Gly Ser
Gly Ile Val Gln Gln Gln Asn Asn Leu Leu Arg Ala 165
170 175 Ile Glu Ala Gln Gln His Leu Leu Gln
Leu Thr Val Trp Gly Ile Lys 180 185
190 Gln Leu Gln Gly Ser Gln Ile Glu Leu Ala Val Leu Leu Ser
Asn Glu 195 200 205
Gly Ile Ile Asn Ser Glu Asp Glu His Leu Leu Ala Leu Glu Arg Lys 210
215 220 Leu Lys Lys Met Leu
Gly Pro Ser Ala Val Glu Ile Gly Asn Gly Cys 225 230
235 240 Phe Glu Thr Lys His Lys Cys Asn Gln Thr
Cys Leu Asp Arg Ile Ala 245 250
255 Ala Gly Thr Phe Asp Ala Gly Glu Phe Ser Leu Pro Thr Phe Asp
Ser 260 265 270 Leu
Asn Ile Thr Ser Gly Gly 275 99837DNAInfluenza
virus 99gcctccggat gtgatattca ggctatcgaa ggtgggcaga gaaaactcgc cggcatcaaa
60tgttccggcg gcaattctat ccaggcaggt ctggttgcac ttgtgctttg tctcaaagca
120tccattgccg atttccacgg cgctaggtcc cagcatcttc ttcagctttc tttccagggc
180cagcagatgt tcgtcctcgc tattgatgat gccctcgtta gacagcagca cagccagttc
240aatctggctc ccctgcagct gcttgatgcc ccacactgtc agctgcagca ggtgctgctg
300agcttcaatg gctcgcagca gattgttctg ctgctgcacg attccggagc ctcccccagt
360cccattctcc tgctggttct ggctttcctc aatcagactg tagatgatag atgtgtagtt
420gttgatttcg cgatcccact cggggtcgcc tccttccagc tcgctcaggc tgttcaggtt
480cttggtgatc ttgttgatgg cttcctgtgt agatttcaga tcagcagcca ctgccacgcc
540atgtgcgcca tgagatgtgt agccatgcca tccagcaatc attccctccc atccgccttc
600cagaaagccg gcgatagctc caaaaaagcc tctctctttc agcagcttgg ctggaggtct
660atactttgtg ccattggcca gcttcagtcc tgatcccaga gggatcacgc cggtcacatt
720cacttcgccc tgtgtagctg ttttcacgac gtgagggcta ttgctgctgg tgatgccggt
780acagattcta tcggcgttgc ttgtgaccac catcagcagc acgatgatgg ccttcat
8371001332DNAArtificial SequenceSynthetic 100atgaaggcca agctgctggt
gctgctgtgc acctttaccg ccacctacgc cgacaccatc 60tgcattggct accacgccaa
caacagcacc gacaccgtgg ataccgtgct ggaaaagaac 120gtgaccgtga cccacagcgt
gaacctggga tccggactga gaatggtcac cggcctgaga 180aacatcccca gcatccagag
cagaggcctg tttggagcca ttgccggctt tattgagggc 240ggatggaccg gaatggtgga
tgggtggtac ggctaccacc accagaatga gcagggctct 300ggctatgccg ccgatcagaa
gtctacccag aacgccatca acggcatcac caacaaagtg 360aacagcgtga tcgagaagat
gggcggcgat cctgaatggg acagagagat caacaactac 420accagcatca tctacagcct
gatcgaggaa agccagaacc agcaggaaaa cggcacaggc 480ggcggatctg gaattgtgca
gcagcagaac aacctgctga gagccattga ggcccagcag 540catctgctgc agctgacagt
gtggggcatc aagcagctgc agacctacaa tgccgagctg 600ctggtcctcc tggaaaacga
gagaaccctg gacttccacg acagcaacgt gaagaacctg 660tacgagaaag tgaagtccca
gctgaagaac aacgccaaag agatcggcaa cggctgcttc 720gagttctacc acaagtgcaa
caacgagtgc atggaaagcg tgaagaacgg cacctacgac 780taccccaagt acagcgagga
aagcaagctg aacagagaga agatcgactc cggaggcgac 840atcatcaagc tgctgaacga
gcaggtgaac aaggagatgc agagcagcaa cctgtacatg 900agcatgagca gctggtgcta
cacccacagc ctggacggcg ccggcctgtt cctgttcgac 960cacgccgccg aggagtacga
gcacgccaag aagctgatca tcttcctgaa cgagaacaac 1020gtgcccgtgc agctgaccag
catcagcgcc cccgagcaca agttcgaggg cctgacccag 1080atcttccaga aggcctacga
gcacgagcag cacatcagcg agagcatcaa caacatcgtg 1140gaccacgcca tcaagagcaa
ggaccacgcc accttcaact tcctgcagtg gtacgtggcc 1200gagcagcacg aggaggaggt
gctgttcaag gacatcctgg acaagatcga gctgatcggc 1260aacgagaacc acggcctgta
cctggccgac cagtacgtga agggcatcgc caagagcagg 1320aagagcggat cc
1332101444PRTArtificial
SequenceSynthetic 101Met Lys Ala Lys Leu Leu Val Leu Leu Cys Thr Phe Thr
Ala Thr Tyr 1 5 10 15
Ala Asp Thr Ile Cys Ile Gly Tyr His Ala Asn Asn Ser Thr Asp Thr
20 25 30 Val Asp Thr Val
Leu Glu Lys Asn Val Thr Val Thr His Ser Val Asn 35
40 45 Leu Gly Ser Gly Leu Arg Met Val Thr
Gly Leu Arg Asn Ile Pro Ser 50 55
60 Ile Gln Ser Arg Gly Leu Phe Gly Ala Ile Ala Gly Phe
Ile Glu Gly 65 70 75
80 Gly Trp Thr Gly Met Val Asp Gly Trp Tyr Gly Tyr His His Gln Asn
85 90 95 Glu Gln Gly Ser
Gly Tyr Ala Ala Asp Gln Lys Ser Thr Gln Asn Ala 100
105 110 Ile Asn Gly Ile Thr Asn Lys Val Asn
Ser Val Ile Glu Lys Met Gly 115 120
125 Gly Asp Pro Glu Trp Asp Arg Glu Ile Asn Asn Tyr Thr Ser
Ile Ile 130 135 140
Tyr Ser Leu Ile Glu Glu Ser Gln Asn Gln Gln Glu Asn Gly Thr Gly 145
150 155 160 Gly Gly Ser Gly Ile
Val Gln Gln Gln Asn Asn Leu Leu Arg Ala Ile 165
170 175 Glu Ala Gln Gln His Leu Leu Gln Leu Thr
Val Trp Gly Ile Lys Gln 180 185
190 Leu Gln Thr Tyr Asn Ala Glu Leu Leu Val Leu Leu Glu Asn Glu
Arg 195 200 205 Thr
Leu Asp Phe His Asp Ser Asn Val Lys Asn Leu Tyr Glu Lys Val 210
215 220 Lys Ser Gln Leu Lys Asn
Asn Ala Lys Glu Ile Gly Asn Gly Cys Phe 225 230
235 240 Glu Phe Tyr His Lys Cys Asn Asn Glu Cys Met
Glu Ser Val Lys Asn 245 250
255 Gly Thr Tyr Asp Tyr Pro Lys Tyr Ser Glu Glu Ser Lys Leu Asn Arg
260 265 270 Glu Lys
Ile Asp Ser Gly Gly Asp Ile Ile Lys Leu Leu Asn Glu Gln 275
280 285 Val Asn Lys Glu Met Gln Ser
Ser Asn Leu Tyr Met Ser Met Ser Ser 290 295
300 Trp Cys Tyr Thr His Ser Leu Asp Gly Ala Gly Leu
Phe Leu Phe Asp 305 310 315
320 His Ala Ala Glu Glu Tyr Glu His Ala Lys Lys Leu Ile Ile Phe Leu
325 330 335 Asn Glu Asn
Asn Val Pro Val Gln Leu Thr Ser Ile Ser Ala Pro Glu 340
345 350 His Lys Phe Glu Gly Leu Thr Gln
Ile Phe Gln Lys Ala Tyr Glu His 355 360
365 Glu Gln His Ile Ser Glu Ser Ile Asn Asn Ile Val Asp
His Ala Ile 370 375 380
Lys Ser Lys Asp His Ala Thr Phe Asn Phe Leu Gln Trp Tyr Val Ala 385
390 395 400 Glu Gln His Glu
Glu Glu Val Leu Phe Lys Asp Ile Leu Asp Lys Ile 405
410 415 Glu Leu Ile Gly Asn Glu Asn His Gly
Leu Tyr Leu Ala Asp Gln Tyr 420 425
430 Val Lys Gly Ile Ala Lys Ser Arg Lys Ser Gly Ser
435 440 1021332DNAArtificial
SequenceSynthetic 102ggatccgctc ttcctgctct tggcgatgcc cttcacgtac
tggtcggcca ggtacaggcc 60gtggttctcg ttgccgatca gctcgatctt gtccaggatg
tccttgaaca gcacctcctc 120ctcgtgctgc tcggccacgt accactgcag gaagttgaag
gtggcgtggt ccttgctctt 180gatggcgtgg tccacgatgt tgttgatgct ctcgctgatg
tgctgctcgt gctcgtaggc 240cttctggaag atctgggtca ggccctcgaa cttgtgctcg
ggggcgctga tgctggtcag 300ctgcacgggc acgttgttct cgttcaggaa gatgatcagc
ttcttggcgt gctcgtactc 360ctcggcggcg tggtcgaaca ggaacaggcc ggcgccgtcc
aggctgtggg tgtagcacca 420gctgctcatg ctcatgtaca ggttgctgct ctgcatctcc
ttgttcacct gctcgttcag 480cagcttgatg atgtcgcctc cggagtcgat cttctctctg
ttcagcttgc tttcctcgct 540gtacttgggg tagtcgtagg tgccgttctt cacgctttcc
atgcactcgt tgttgcactt 600gtggtagaac tcgaagcagc cgttgccgat ctctttggcg
ttgttcttca gctgggactt 660cactttctcg tacaggttct tcacgttgct gtcgtggaag
tccagggttc tctcgttttc 720caggaggacc agcagctcgg cattgtaggt ctgcagctgc
ttgatgcccc acactgtcag 780ctgcagcaga tgctgctggg cctcaatggc tctcagcagg
ttgttctgct gctgcacaat 840tccagatccg ccgcctgtgc cgttttcctg ctggttctgg
ctttcctcga tcaggctgta 900gatgatgctg gtgtagttgt tgatctctct gtcccattca
ggatcgccgc ccatcttctc 960gatcacgctg ttcactttgt tggtgatgcc gttgatggcg
ttctgggtag acttctgatc 1020ggcggcatag ccagagccct gctcattctg gtggtggtag
ccgtaccacc catccaccat 1080tccggtccat ccgccctcaa taaagccggc aatggctcca
aacaggcctc tgctctggat 1140gctggggatg tttctcaggc cggtgaccat tctcagtccg
gatcccaggt tcacgctgtg 1200ggtcacggtc acgttctttt ccagcacggt atccacggtg
tcggtgctgt tgttggcgtg 1260gtagccaatg cagatggtgt cggcgtaggt ggcggtaaag
gtgcacagca gcaccagcag 1320cttggccttc at
13321031332DNAArtificial SequenceSynthetic
103atgaaggcta tcctggtggt gctgctgtac acctttgcca ccgccaatgc cgacaccctg
60tgtattggct accacgccaa caacagcacc gacaccgtgg ataccgtgct ggaaaagaac
120gtgaccgtga cccacagcgt gaacctgggc tccggcctga gactggccac cggcctgaga
180aacatcccca gcattcagag cagaggcctg tttggagcca ttgccggctt tattgagggc
240ggatggaccg gaatggtgga tgggtggtac ggctaccacc accagaatga gcagggctct
300ggctatgccg ccgacctgaa gtctacccag aacgccatcg acgagatcac caacaaagtg
360aacagcgtga tcgagaagat gggcggctgg gacccatggg acagagagat caacaactac
420accagcatca tctacagcct gatcgaggaa agccagaacc agcaggaaaa cggcacaggc
480ggcggatctg gaattgtgca gcagcagaac aacctgctga gagccattga ggcccagcag
540catctgctgc agctgacagt gtggggcatc aagcagctgc agacctacaa cgccgagctg
600ctggtgctgc tcgagaatga gagaaccctg gactaccacg acagcaacgt gaagaacctg
660tacgagaaag tgcggagcca gctgaagaac aacgccaaag agatcggcaa cggctgcttc
720gagttctacc acaagtgcga caatacctgc atggaaagcg tgaagaacgg cacctacgac
780taccccaagt acagcgagga agccaagctg aaccgggaag agatcgattc cggaggcgac
840atcatcaagc tgctgaacga gcaggtgaac aaggagatgc agagcagcaa cctgtacatg
900agcatgagca gctggtgcta cacccacagc ctggacggcg ccggcctgtt cctgttcgac
960cacgccgccg aggagtacga gcacgccaag aagctgatca tcttcctgaa cgagaacaac
1020gtgcccgtgc agctgaccag catcagcgcc cccgagcaca agttcgaggg cctgacccag
1080atcttccaga aggcctacga gcacgagcag cacatcagcg agagcatcaa caacatcgtg
1140gaccacgcca tcaagagcaa ggaccacgcc accttcaact tcctgcagtg gtacgtggcc
1200gagcagcacg aggaggaggt gctgttcaag gacatcctgg acaagatcga gctgatcggc
1260aacgagaacc acggcctgta cctggccgac cagtacgtga agggcatcgc caagagcagg
1320aagagcggat cc
1332104444PRTArtificial SequenceSynthetic 104Met Lys Ala Ile Leu Val Val
Leu Leu Tyr Thr Phe Ala Thr Ala Asn 1 5
10 15 Ala Asp Thr Leu Cys Ile Gly Tyr His Ala Asn
Asn Ser Thr Asp Thr 20 25
30 Val Asp Thr Val Leu Glu Lys Asn Val Thr Val Thr His Ser Val
Asn 35 40 45 Leu
Gly Ser Gly Leu Arg Leu Ala Thr Gly Leu Arg Asn Ile Pro Ser 50
55 60 Ile Gln Ser Arg Gly Leu
Phe Gly Ala Ile Ala Gly Phe Ile Glu Gly 65 70
75 80 Gly Trp Thr Gly Met Val Asp Gly Trp Tyr Gly
Tyr His His Gln Asn 85 90
95 Glu Gln Gly Ser Gly Tyr Ala Ala Asp Leu Lys Ser Thr Gln Asn Ala
100 105 110 Ile Asp
Glu Ile Thr Asn Lys Val Asn Ser Val Ile Glu Lys Met Gly 115
120 125 Gly Trp Asp Pro Trp Asp Arg
Glu Ile Asn Asn Tyr Thr Ser Ile Ile 130 135
140 Tyr Ser Leu Ile Glu Glu Ser Gln Asn Gln Gln Glu
Asn Gly Thr Gly 145 150 155
160 Gly Gly Ser Gly Ile Val Gln Gln Gln Asn Asn Leu Leu Arg Ala Ile
165 170 175 Glu Ala Gln
Gln His Leu Leu Gln Leu Thr Val Trp Gly Ile Lys Gln 180
185 190 Leu Gln Thr Tyr Asn Ala Glu Leu
Leu Val Leu Leu Glu Asn Glu Arg 195 200
205 Thr Leu Asp Tyr His Asp Ser Asn Val Lys Asn Leu Tyr
Glu Lys Val 210 215 220
Arg Ser Gln Leu Lys Asn Asn Ala Lys Glu Ile Gly Asn Gly Cys Phe 225
230 235 240 Glu Phe Tyr His
Lys Cys Asp Asn Thr Cys Met Glu Ser Val Lys Asn 245
250 255 Gly Thr Tyr Asp Tyr Pro Lys Tyr Ser
Glu Glu Ala Lys Leu Asn Arg 260 265
270 Glu Glu Ile Asp Ser Gly Gly Asp Ile Ile Lys Leu Leu Asn
Glu Gln 275 280 285
Val Asn Lys Glu Met Gln Ser Ser Asn Leu Tyr Met Ser Met Ser Ser 290
295 300 Trp Cys Tyr Thr His
Ser Leu Asp Gly Ala Gly Leu Phe Leu Phe Asp 305 310
315 320 His Ala Ala Glu Glu Tyr Glu His Ala Lys
Lys Leu Ile Ile Phe Leu 325 330
335 Asn Glu Asn Asn Val Pro Val Gln Leu Thr Ser Ile Ser Ala Pro
Glu 340 345 350 His
Lys Phe Glu Gly Leu Thr Gln Ile Phe Gln Lys Ala Tyr Glu His 355
360 365 Glu Gln His Ile Ser Glu
Ser Ile Asn Asn Ile Val Asp His Ala Ile 370 375
380 Lys Ser Lys Asp His Ala Thr Phe Asn Phe Leu
Gln Trp Tyr Val Ala 385 390 395
400 Glu Gln His Glu Glu Glu Val Leu Phe Lys Asp Ile Leu Asp Lys Ile
405 410 415 Glu Leu
Ile Gly Asn Glu Asn His Gly Leu Tyr Leu Ala Asp Gln Tyr 420
425 430 Val Lys Gly Ile Ala Lys Ser
Arg Lys Ser Gly Ser 435 440
1051332DNAArtificial SequenceSynthetic 105ggatccgctc ttcctgctct
tggcgatgcc cttcacgtac tggtcggcca ggtacaggcc 60gtggttctcg ttgccgatca
gctcgatctt gtccaggatg tccttgaaca gcacctcctc 120ctcgtgctgc tcggccacgt
accactgcag gaagttgaag gtggcgtggt ccttgctctt 180gatggcgtgg tccacgatgt
tgttgatgct ctcgctgatg tgctgctcgt gctcgtaggc 240cttctggaag atctgggtca
ggccctcgaa cttgtgctcg ggggcgctga tgctggtcag 300ctgcacgggc acgttgttct
cgttcaggaa gatgatcagc ttcttggcgt gctcgtactc 360ctcggcggcg tggtcgaaca
ggaacaggcc ggcgccgtcc aggctgtggg tgtagcacca 420gctgctcatg ctcatgtaca
ggttgctgct ctgcatctcc ttgttcacct gctcgttcag 480cagcttgatg atgtcgcctc
cggaatcgat ctcttcccgg ttcagcttgg cttcctcgct 540gtacttgggg tagtcgtagg
tgccgttctt cacgctttcc atgcaggtat tgtcgcactt 600gtggtagaac tcgaagcagc
cgttgccgat ctctttggcg ttgttcttca gctggctccg 660cactttctcg tacaggttct
tcacgttgct gtcgtggtag tccagggttc tctcattctc 720gagcagcacc agcagctcgg
cgttgtaggt ctgcagctgc ttgatgcccc acactgtcag 780ctgcagcaga tgctgctggg
cctcaatggc tctcagcagg ttgttctgct gctgcacaat 840tccagatccg ccgcctgtgc
cgttttcctg ctggttctgg ctttcctcga tcaggctgta 900gatgatgctg gtgtagttgt
tgatctctct gtcccatggg tcccagccgc ccatcttctc 960gatcacgctg ttcactttgt
tggtgatctc gtcgatggcg ttctgggtag acttcaggtc 1020ggcggcatag ccagagccct
gctcattctg gtggtggtag ccgtaccacc catccaccat 1080tccggtccat ccgccctcaa
taaagccggc aatggctcca aacaggcctc tgctctgaat 1140gctggggatg tttctcaggc
cggtggccag tctcaggccg gagcccaggt tcacgctgtg 1200ggtcacggtc acgttctttt
ccagcacggt atccacggtg tcggtgctgt tgttggcgtg 1260gtagccaata cacagggtgt
cggcattggc ggtggcaaag gtgtacagca gcaccaccag 1320gatagccttc at
13321061326DNAArtificial
SequenceSynthetic 106atggccatca tctacctgat tctgctgttt acagccgtca
gaggcgatca gatctgtatt 60ggctaccacg ccaacaatag caccgagaaa gtggatacca
tcctggaaag aaatgtgaca 120gtgacacacg ccaaggatat tggatcagga ctggtgctgg
ctacaggact gagaaatgtg 180cctcagattg agagcagagg cctgtttgga gccattgctg
gctttattga aggcggatgg 240cagggaatga ttgatgggtg gtacggctac caccactcta
atgatcaggg atctggatat 300gccgccgaca aagaatctac acagaaagcc ttcgacggca
tcaccaacaa agtgaatagc 360gtgatcgaga agatgggcgg agatcccgaa tgggacagag
agatcaacaa ctacaccagc 420atcatctaca gcctgatcga ggaaagccag aatcagcagg
aaaatggaac aggcggagga 480tctggaattg tgcagcagca gaacaatctg ctgagagcta
ttgaagctca gcagcatctg 540ctgaatctga cagtgtgggg aatcaaacag ctgcagacat
acaatgctga gctgctggtg 600ctgatggaaa atgagagaac cctggacttc cacgacagca
atgtgaagaa cctgtacgac 660aaagtgcgga tgcagctgag agacaatgtg aaagaactgg
gcaatggctg cttcgagttc 720taccacaagt gcgacgatga gtgtatgaac agcgtgaaga
acggcaccta cgactaccct 780aagtacgagg aagagagcaa gctgaacaga aatgagatca
agtccggagg cgacatcatc 840aagctgctga acgagcaggt gaacaaggag atgcagagca
gcaacctgta catgagcatg 900agcagctggt gctacaccca cagcctggac ggcgccggcc
tgttcctgtt cgaccacgcc 960gccgaggagt acgagcacgc caagaagctg atcatcttcc
tgaacgagaa caacgtgccc 1020gtgcagctga ccagcatcag cgcccccgag cacaagttcg
agggcctgac ccagatcttc 1080cagaaggcct acgagcacga gcagcacatc agcgagagca
tcaacaacat cgtggaccac 1140gccatcaaga gcaaggacca cgccaccttc aacttcctgc
agtggtacgt ggccgagcag 1200cacgaggagg aggtgctgtt caaggacatc ctggacaaga
tcgagctgat cggcaacgag 1260aaccacggcc tgtacctggc cgaccagtac gtgaagggca
tcgccaagag caggaagagc 1320ggatcc
1326107442PRTArtificial SequenceSynthetic 107Met
Ala Ile Ile Tyr Leu Ile Leu Leu Phe Thr Ala Val Arg Gly Asp 1
5 10 15 Gln Ile Cys Ile Gly Tyr
His Ala Asn Asn Ser Thr Glu Lys Val Asp 20
25 30 Thr Ile Leu Glu Arg Asn Val Thr Val Thr
His Ala Lys Asp Ile Gly 35 40
45 Ser Gly Leu Val Leu Ala Thr Gly Leu Arg Asn Val Pro Gln
Ile Glu 50 55 60
Ser Arg Gly Leu Phe Gly Ala Ile Ala Gly Phe Ile Glu Gly Gly Trp 65
70 75 80 Gln Gly Met Ile Asp
Gly Trp Tyr Gly Tyr His His Ser Asn Asp Gln 85
90 95 Gly Ser Gly Tyr Ala Ala Asp Lys Glu Ser
Thr Gln Lys Ala Phe Asp 100 105
110 Gly Ile Thr Asn Lys Val Asn Ser Val Ile Glu Lys Met Gly Gly
Asp 115 120 125 Pro
Glu Trp Asp Arg Glu Ile Asn Asn Tyr Thr Ser Ile Ile Tyr Ser 130
135 140 Leu Ile Glu Glu Ser Gln
Asn Gln Gln Glu Asn Gly Thr Gly Gly Gly 145 150
155 160 Ser Gly Ile Val Gln Gln Gln Asn Asn Leu Leu
Arg Ala Ile Glu Ala 165 170
175 Gln Gln His Leu Leu Asn Leu Thr Val Trp Gly Ile Lys Gln Leu Gln
180 185 190 Thr Tyr
Asn Ala Glu Leu Leu Val Leu Met Glu Asn Glu Arg Thr Leu 195
200 205 Asp Phe His Asp Ser Asn Val
Lys Asn Leu Tyr Asp Lys Val Arg Met 210 215
220 Gln Leu Arg Asp Asn Val Lys Glu Leu Gly Asn Gly
Cys Phe Glu Phe 225 230 235
240 Tyr His Lys Cys Asp Asp Glu Cys Met Asn Ser Val Lys Asn Gly Thr
245 250 255 Tyr Asp Tyr
Pro Lys Tyr Glu Glu Glu Ser Lys Leu Asn Arg Asn Glu 260
265 270 Ile Lys Ser Gly Gly Asp Ile Ile
Lys Leu Leu Asn Glu Gln Val Asn 275 280
285 Lys Glu Met Gln Ser Ser Asn Leu Tyr Met Ser Met Ser
Ser Trp Cys 290 295 300
Tyr Thr His Ser Leu Asp Gly Ala Gly Leu Phe Leu Phe Asp His Ala 305
310 315 320 Ala Glu Glu Tyr
Glu His Ala Lys Lys Leu Ile Ile Phe Leu Asn Glu 325
330 335 Asn Asn Val Pro Val Gln Leu Thr Ser
Ile Ser Ala Pro Glu His Lys 340 345
350 Phe Glu Gly Leu Thr Gln Ile Phe Gln Lys Ala Tyr Glu His
Glu Gln 355 360 365
His Ile Ser Glu Ser Ile Asn Asn Ile Val Asp His Ala Ile Lys Ser 370
375 380 Lys Asp His Ala Thr
Phe Asn Phe Leu Gln Trp Tyr Val Ala Glu Gln 385 390
395 400 His Glu Glu Glu Val Leu Phe Lys Asp Ile
Leu Asp Lys Ile Glu Leu 405 410
415 Ile Gly Asn Glu Asn His Gly Leu Tyr Leu Ala Asp Gln Tyr Val
Lys 420 425 430 Gly
Ile Ala Lys Ser Arg Lys Ser Gly Ser 435 440
1081326DNAArtificial SequenceSynthetic 108ggatccgctc ttcctgctct
tggcgatgcc cttcacgtac tggtcggcca ggtacaggcc 60gtggttctcg ttgccgatca
gctcgatctt gtccaggatg tccttgaaca gcacctcctc 120ctcgtgctgc tcggccacgt
accactgcag gaagttgaag gtggcgtggt ccttgctctt 180gatggcgtgg tccacgatgt
tgttgatgct ctcgctgatg tgctgctcgt gctcgtaggc 240cttctggaag atctgggtca
ggccctcgaa cttgtgctcg ggggcgctga tgctggtcag 300ctgcacgggc acgttgttct
cgttcaggaa gatgatcagc ttcttggcgt gctcgtactc 360ctcggcggcg tggtcgaaca
ggaacaggcc ggcgccgtcc aggctgtggg tgtagcacca 420gctgctcatg ctcatgtaca
ggttgctgct ctgcatctcc ttgttcacct gctcgttcag 480cagcttgatg atgtcgcctc
cggacttgat ctcatttctg ttcagcttgc tctcttcctc 540gtacttaggg tagtcgtagg
tgccgttctt cacgctgttc atacactcat cgtcgcactt 600gtggtagaac tcgaagcagc
cattgcccag ttctttcaca ttgtctctca gctgcatccg 660cactttgtcg tacaggttct
tcacattgct gtcgtggaag tccagggttc tctcattttc 720catcagcacc agcagctcag
cattgtatgt ctgcagctgt ttgattcccc acactgtcag 780attcagcaga tgctgctgag
cttcaatagc tctcagcaga ttgttctgct gctgcacaat 840tccagatcct ccgcctgttc
cattttcctg ctgattctgg ctttcctcga tcaggctgta 900gatgatgctg gtgtagttgt
tgatctctct gtcccattcg ggatctccgc ccatcttctc 960gatcacgcta ttcactttgt
tggtgatgcc gtcgaaggct ttctgtgtag attctttgtc 1020ggcggcatat ccagatccct
gatcattaga gtggtggtag ccgtaccacc catcaatcat 1080tccctgccat ccgccttcaa
taaagccagc aatggctcca aacaggcctc tgctctcaat 1140ctgaggcaca tttctcagtc
ctgtagccag caccagtcct gatccaatat ccttggcgtg 1200tgtcactgtc acatttcttt
ccaggatggt atccactttc tcggtgctat tgttggcgtg 1260gtagccaata cagatctgat
cgcctctgac ggctgtaaac agcagaatca ggtagatgat 1320ggccat
13261091362DNAArtificial
SequenceSynthetic 109atgaagacca tcatcgccct gagctacatc ttctgcctgg
ccctgggcca ggacctgccc 60ggcaacgaca acagcaccgc caccctgtgc ctgggccacc
acgccgtgcc caacggcacc 120ctggtgaaga ccatcaccga cgaccagatc gaggtgacca
acgccaccga gctgggctcc 180ggcctgaagc tggccaccgg catgcggaac gtgcccgaga
agcagacccg gggcctgttc 240ggcgccatcg ccggcttcat cgagaacggc tgggagggca
tgatcgacgg ctggtacggc 300ttccggcacc agaacagcga gggcaccggc caggccgccg
acctgaagag cacccaggcc 360gccatcgacc agatcaacgg caagctgaac cgggtgatcg
agaagaccgg cggcgatccc 420gagtgggacc gggagatcaa caactacacc agcatcatct
acagcctgat cgaggagagc 480cagaaccagc aggagaacgg caccggcggc ggcagcggca
tcgtgcagca gcagaacaac 540ctgctgcggg ccatcgaggc ccagcagcac ctgctgcagc
tgaccgtgtg gggcatcaag 600cagctgcaga gctacaacgc cgagctgctg gtggccctgg
agaaccagca caccatcgac 660ctgaccgaca gcgagatgaa caagctgttc gagaagaccc
ggcggcagct gcgggagaac 720gccgaggaca tgggcaacgg ctgcttcaag atctaccaca
agtgcgacaa cgcctgcatc 780gagagcatcc ggaacggcac ctacgaccac gacgtgtacc
gggacgaggc cctgaacaac 840cggttccaga tcaagggctc cggaggcgac atcatcaagc
tgctgaacga gcaggtgaac 900aaggagatgc agagcagcaa cctgtacatg agcatgagca
gctggtgcta cacccacagc 960ctggacggcg ccggcctgtt cctgttcgac cacgccgccg
aggagtacga gcacgccaag 1020aagctgatca tcttcctgaa cgagaacaac gtgcccgtgc
agctgaccag catcagcgcc 1080cccgagcaca agttcgaggg cctgacccag atcttccaga
aggcctacga gcacgagcag 1140cacatcagcg agagcatcaa caacatcgtg gaccacgcca
tcaagagcaa ggaccacgcc 1200accttcaact tcctgcagtg gtacgtggcc gagcagcacg
aggaggaggt gctgttcaag 1260gacatcctgg acaagatcga gctgatcggc aacgagaacc
acggcctgta cctggccgac 1320cagtacgtga agggcatcgc caagagcagg aagagcggat
cc 1362110454PRTArtificial SequenceSynthetic 110Met
Lys Thr Ile Ile Ala Leu Ser Tyr Ile Phe Cys Leu Ala Leu Gly 1
5 10 15 Gln Asp Leu Pro Gly Asn
Asp Asn Ser Thr Ala Thr Leu Cys Leu Gly 20
25 30 His His Ala Val Pro Asn Gly Thr Leu Val
Lys Thr Ile Thr Asp Asp 35 40
45 Gln Ile Glu Val Thr Asn Ala Thr Glu Leu Gly Ser Gly Leu
Lys Leu 50 55 60
Ala Thr Gly Met Arg Asn Val Pro Glu Lys Gln Thr Arg Gly Leu Phe 65
70 75 80 Gly Ala Ile Ala Gly
Phe Ile Glu Asn Gly Trp Glu Gly Met Ile Asp 85
90 95 Gly Trp Tyr Gly Phe Arg His Gln Asn Ser
Glu Gly Thr Gly Gln Ala 100 105
110 Ala Asp Leu Lys Ser Thr Gln Ala Ala Ile Asp Gln Ile Asn Gly
Lys 115 120 125 Leu
Asn Arg Val Ile Glu Lys Thr Gly Gly Asp Pro Glu Trp Asp Arg 130
135 140 Glu Ile Asn Asn Tyr Thr
Ser Ile Ile Tyr Ser Leu Ile Glu Glu Ser 145 150
155 160 Gln Asn Gln Gln Glu Asn Gly Thr Gly Gly Gly
Ser Gly Ile Val Gln 165 170
175 Gln Gln Asn Asn Leu Leu Arg Ala Ile Glu Ala Gln Gln His Leu Leu
180 185 190 Gln Leu
Thr Val Trp Gly Ile Lys Gln Leu Gln Ser Tyr Asn Ala Glu 195
200 205 Leu Leu Val Ala Leu Glu Asn
Gln His Thr Ile Asp Leu Thr Asp Ser 210 215
220 Glu Met Asn Lys Leu Phe Glu Lys Thr Arg Arg Gln
Leu Arg Glu Asn 225 230 235
240 Ala Glu Asp Met Gly Asn Gly Cys Phe Lys Ile Tyr His Lys Cys Asp
245 250 255 Asn Ala Cys
Ile Glu Ser Ile Arg Asn Gly Thr Tyr Asp His Asp Val 260
265 270 Tyr Arg Asp Glu Ala Leu Asn Asn
Arg Phe Gln Ile Lys Gly Ser Gly 275 280
285 Gly Asp Ile Ile Lys Leu Leu Asn Glu Gln Val Asn Lys
Glu Met Gln 290 295 300
Ser Ser Asn Leu Tyr Met Ser Met Ser Ser Trp Cys Tyr Thr His Ser 305
310 315 320 Leu Asp Gly Ala
Gly Leu Phe Leu Phe Asp His Ala Ala Glu Glu Tyr 325
330 335 Glu His Ala Lys Lys Leu Ile Ile Phe
Leu Asn Glu Asn Asn Val Pro 340 345
350 Val Gln Leu Thr Ser Ile Ser Ala Pro Glu His Lys Phe Glu
Gly Leu 355 360 365
Thr Gln Ile Phe Gln Lys Ala Tyr Glu His Glu Gln His Ile Ser Glu 370
375 380 Ser Ile Asn Asn Ile
Val Asp His Ala Ile Lys Ser Lys Asp His Ala 385 390
395 400 Thr Phe Asn Phe Leu Gln Trp Tyr Val Ala
Glu Gln His Glu Glu Glu 405 410
415 Val Leu Phe Lys Asp Ile Leu Asp Lys Ile Glu Leu Ile Gly Asn
Glu 420 425 430 Asn
His Gly Leu Tyr Leu Ala Asp Gln Tyr Val Lys Gly Ile Ala Lys 435
440 445 Ser Arg Lys Ser Gly Ser
450 1111362DNAArtificial SequenceSynthetic
111ggatccgctc ttcctgctct tggcgatgcc cttcacgtac tggtcggcca ggtacaggcc
60gtggttctcg ttgccgatca gctcgatctt gtccaggatg tccttgaaca gcacctcctc
120ctcgtgctgc tcggccacgt accactgcag gaagttgaag gtggcgtggt ccttgctctt
180gatggcgtgg tccacgatgt tgttgatgct ctcgctgatg tgctgctcgt gctcgtaggc
240cttctggaag atctgggtca ggccctcgaa cttgtgctcg ggggcgctga tgctggtcag
300ctgcacgggc acgttgttct cgttcaggaa gatgatcagc ttcttggcgt gctcgtactc
360ctcggcggcg tggtcgaaca ggaacaggcc ggcgccgtcc aggctgtggg tgtagcacca
420gctgctcatg ctcatgtaca ggttgctgct ctgcatctcc ttgttcacct gctcgttcag
480cagcttgatg atgtcgcctc cggagccctt gatctggaac cggttgttca gggcctcgtc
540ccggtacacg tcgtggtcgt aggtgccgtt ccggatgctc tcgatgcagg cgttgtcgca
600cttgtggtag atcttgaagc agccgttgcc catgtcctcg gcgttctccc gcagctgccg
660ccgggtcttc tcgaacagct tgttcatctc gctgtcggtc aggtcgatgg tgtgctggtt
720ctccagggcc accagcagct cggcgttgta gctctgcagc tgcttgatgc cccacacggt
780cagctgcagc aggtgctgct gggcctcgat ggcccgcagc aggttgttct gctgctgcac
840gatgccgctg ccgccgccgg tgccgttctc ctgctggttc tggctctcct cgatcaggct
900gtagatgatg ctggtgtagt tgttgatctc ccggtcccac tcgggatcgc cgccggtctt
960ctcgatcacc cggttcagct tgccgttgat ctggtcgatg gcggcctggg tgctcttcag
1020gtcggcggcc tggccggtgc cctcgctgtt ctggtgccgg aagccgtacc agccgtcgat
1080catgccctcc cagccgttct cgatgaagcc ggcgatggcg ccgaacaggc cccgggtctg
1140cttctcgggc acgttccgca tgccggtggc cagcttcagg ccggagccca gctcggtggc
1200gttggtcacc tcgatctggt cgtcggtgat ggtcttcacc agggtgccgt tgggcacggc
1260gtggtggccc aggcacaggg tggcggtgct gttgtcgttg ccgggcaggt cctggcccag
1320ggccaggcag aagatgtagc tcagggcgat gatggtcttc at
13621121341DNAArtificial SequenceSynthetic 112atggagaaga tcgtgctgct
gctggccatc gtgagcctgg tgaagagcga ccagatctgc 60atcggctacc acgccaacaa
cagcaccgag caggtggaca ccatcatgga gaagaacgtg 120accgtgaccc acgcccagga
catcggctcc ggcctggtgc tggccaccgg cctgcggaac 180agcccccagc gggagagccg
gcggaagaag cggggcctgt tcggcgccat cgccggcttc 240atcgagggcg gctggcaggg
catggtggac ggctggtacg gctaccacca cagcaacgag 300cagggcagcg gctacgccgc
cgacaaggag agcacccaga aggccatcga cggcgtgacc 360aacaaggtga acagcatcat
cgacaagatg ggcggcgatc ccgagtggga ccgggagatc 420aacaactaca ccagcatcat
ctacagcctg atcgaggaga gccagaacca gcaggagaac 480ggcaccggcg gcggcagcgg
catcgtgcag cagcagaaca acctgctgcg ggccatcgag 540gcccagcagc acctgctgca
gctgaccgtg tggggcatca agcagctgca gacctacaac 600gccgagctgc tggtgctgat
ggagaacgag cggaccctgg acttccacga cagcaacgtg 660aagaacctgt acgacaaggt
gcggctgcag ctgcgggaca acgccaagga gctgggcaac 720ggctgcttcg agttctacca
caagtgcgac aacgagtgca tggagagcat ccggaacggc 780acctacaact acccccagta
cagcgaggag gcccggctga agcgggagga gatcagctcc 840ggaggcgaca tcatcaagct
gctgaacgag caggtgaaca aggagatgca gagcagcaac 900ctgtacatga gcatgagcag
ctggtgctac acccacagcc tggacggcgc cggcctgttc 960ctgttcgacc acgccgccga
ggagtacgag cacgccaaga agctgatcat cttcctgaac 1020gagaacaacg tgcccgtgca
gctgaccagc atcagcgccc ccgagcacaa gttcgagggc 1080ctgacccaga tcttccagaa
ggcctacgag cacgagcagc acatcagcga gagcatcaac 1140aacatcgtgg accacgccat
caagagcaag gaccacgcca ccttcaactt cctgcagtgg 1200tacgtggccg agcagcacga
ggaggaggtg ctgttcaagg acatcctgga caagatcgag 1260ctgatcggca acgagaacca
cggcctgtac ctggccgacc agtacgtgaa gggcatcgcc 1320aagagcagga agagcggatc c
1341113447PRTArtificial
SequenceSynthetic 113Met Glu Lys Ile Val Leu Leu Leu Ala Ile Val Ser Leu
Val Lys Ser 1 5 10 15
Asp Gln Ile Cys Ile Gly Tyr His Ala Asn Asn Ser Thr Glu Gln Val
20 25 30 Asp Thr Ile Met
Glu Lys Asn Val Thr Val Thr His Ala Gln Asp Ile 35
40 45 Gly Ser Gly Leu Val Leu Ala Thr Gly
Leu Arg Asn Ser Pro Gln Arg 50 55
60 Glu Ser Arg Arg Lys Lys Arg Gly Leu Phe Gly Ala Ile
Ala Gly Phe 65 70 75
80 Ile Glu Gly Gly Trp Gln Gly Met Val Asp Gly Trp Tyr Gly Tyr His
85 90 95 His Ser Asn Glu
Gln Gly Ser Gly Tyr Ala Ala Asp Lys Glu Ser Thr 100
105 110 Gln Lys Ala Ile Asp Gly Val Thr Asn
Lys Val Asn Ser Ile Ile Asp 115 120
125 Lys Met Gly Gly Asp Pro Glu Trp Asp Arg Glu Ile Asn Asn
Tyr Thr 130 135 140
Ser Ile Ile Tyr Ser Leu Ile Glu Glu Ser Gln Asn Gln Gln Glu Asn 145
150 155 160 Gly Thr Gly Gly Gly
Ser Gly Ile Val Gln Gln Gln Asn Asn Leu Leu 165
170 175 Arg Ala Ile Glu Ala Gln Gln His Leu Leu
Gln Leu Thr Val Trp Gly 180 185
190 Ile Lys Gln Leu Gln Thr Tyr Asn Ala Glu Leu Leu Val Leu Met
Glu 195 200 205 Asn
Glu Arg Thr Leu Asp Phe His Asp Ser Asn Val Lys Asn Leu Tyr 210
215 220 Asp Lys Val Arg Leu Gln
Leu Arg Asp Asn Ala Lys Glu Leu Gly Asn 225 230
235 240 Gly Cys Phe Glu Phe Tyr His Lys Cys Asp Asn
Glu Cys Met Glu Ser 245 250
255 Ile Arg Asn Gly Thr Tyr Asn Tyr Pro Gln Tyr Ser Glu Glu Ala Arg
260 265 270 Leu Lys
Arg Glu Glu Ile Ser Ser Gly Gly Asp Ile Ile Lys Leu Leu 275
280 285 Asn Glu Gln Val Asn Lys Glu
Met Gln Ser Ser Asn Leu Tyr Met Ser 290 295
300 Met Ser Ser Trp Cys Tyr Thr His Ser Leu Asp Gly
Ala Gly Leu Phe 305 310 315
320 Leu Phe Asp His Ala Ala Glu Glu Tyr Glu His Ala Lys Lys Leu Ile
325 330 335 Ile Phe Leu
Asn Glu Asn Asn Val Pro Val Gln Leu Thr Ser Ile Ser 340
345 350 Ala Pro Glu His Lys Phe Glu Gly
Leu Thr Gln Ile Phe Gln Lys Ala 355 360
365 Tyr Glu His Glu Gln His Ile Ser Glu Ser Ile Asn Asn
Ile Val Asp 370 375 380
His Ala Ile Lys Ser Lys Asp His Ala Thr Phe Asn Phe Leu Gln Trp 385
390 395 400 Tyr Val Ala Glu
Gln His Glu Glu Glu Val Leu Phe Lys Asp Ile Leu 405
410 415 Asp Lys Ile Glu Leu Ile Gly Asn Glu
Asn His Gly Leu Tyr Leu Ala 420 425
430 Asp Gln Tyr Val Lys Gly Ile Ala Lys Ser Arg Lys Ser Gly
Ser 435 440 445
1141341DNAArtificial SequenceSynthetic 114ggatccgctc ttcctgctct
tggcgatgcc cttcacgtac tggtcggcca ggtacaggcc 60gtggttctcg ttgccgatca
gctcgatctt gtccaggatg tccttgaaca gcacctcctc 120ctcgtgctgc tcggccacgt
accactgcag gaagttgaag gtggcgtggt ccttgctctt 180gatggcgtgg tccacgatgt
tgttgatgct ctcgctgatg tgctgctcgt gctcgtaggc 240cttctggaag atctgggtca
ggccctcgaa cttgtgctcg ggggcgctga tgctggtcag 300ctgcacgggc acgttgttct
cgttcaggaa gatgatcagc ttcttggcgt gctcgtactc 360ctcggcggcg tggtcgaaca
ggaacaggcc ggcgccgtcc aggctgtggg tgtagcacca 420gctgctcatg ctcatgtaca
ggttgctgct ctgcatctcc ttgttcacct gctcgttcag 480cagcttgatg atgtcgcctc
cggagctgat ctcctcccgc ttcagccggg cctcctcgct 540gtactggggg tagttgtagg
tgccgttccg gatgctctcc atgcactcgt tgtcgcactt 600gtggtagaac tcgaagcagc
cgttgcccag ctccttggcg ttgtcccgca gctgcagccg 660caccttgtcg tacaggttct
tcacgttgct gtcgtggaag tccagggtcc gctcgttctc 720catcagcacc agcagctcgg
cgttgtaggt ctgcagctgc ttgatgcccc acacggtcag 780ctgcagcagg tgctgctggg
cctcgatggc ccgcagcagg ttgttctgct gctgcacgat 840gccgctgccg ccgccggtgc
cgttctcctg ctggttctgg ctctcctcga tcaggctgta 900gatgatgctg gtgtagttgt
tgatctcccg gtcccactcg ggatcgccgc ccatcttgtc 960gatgatgctg ttcaccttgt
tggtcacgcc gtcgatggcc ttctgggtgc tctccttgtc 1020ggcggcgtag ccgctgccct
gctcgttgct gtggtggtag ccgtaccagc cgtccaccat 1080gccctgccag ccgccctcga
tgaagccggc gatggcgccg aacaggcccc gcttcttccg 1140ccggctctcc cgctgggggc
tgttccgcag gccggtggcc agcaccaggc cggagccgat 1200gtcctgggcg tgggtcacgg
tcacgttctt ctccatgatg gtgtccacct gctcggtgct 1260gttgttggcg tggtagccga
tgcagatctg gtcgctcttc accaggctca cgatggccag 1320cagcagcacg atcttctcca t
13411151332DNAArtificial
SequenceSynthetic 115atgaaagtga agctgctggt gctgctgtgt acctttaccg
ccacctacgc cgataccatc 60tgtatcggct accacgccaa caatagcacc gacaccgtgg
ataccgtgct ggaaaagaac 120gtgaccgtga cccacagcgt gaacctggga tcaggactga
gaatggtgac cggcctgagg 180aatatcccca gcatccagag cagaggcctg tttggcgcca
ttgccggctt tatcgagggc 240ggatggacag gcatggtgga tgggtggtac ggctaccacc
accagaatga gcagggatct 300ggctatgccg ccgatcagaa gagcacccag aacgccatca
acggcatcac caacaaagtg 360aacagcgtga tcgagaagat gggcggcgat cctgaatggg
acagagagat caacaactac 420accagcatca tctacagcct gatcgaggaa agccagaacc
agcaggaaaa cggcacaggc 480ggcggatctg gaattgtgca gcagcagaac aacctgctga
gagccattga ggcccagcag 540catctgctgc agctgacagt gtggggcatc aagcagctgc
agacctacaa cgccgaactc 600ctggtcctcc tggaaaatga gaggaccctg gacttccacg
acagcaacgt gaagaacctg 660tacgagaaag tgaagagcca gctgaagaac aacgccaaag
agatcggcaa cggctgcttc 720gagttctacc acaagtgcaa cgacgagtgc atggaaagcg
tgaagaacgg cacctacgac 780taccccaagt acagcgagga aagcaagctg aaccgggaga
agatcgattc cggaggcgac 840atcatcaagc tgctgaacga gcaggtgaac aaggagatgc
agagcagcaa cctgtacatg 900agcatgagca gctggtgcta cacccacagc ctggacggcg
ccggcctgtt cctgttcgac 960cacgccgccg aggagtacga gcacgccaag aagctgatca
tcttcctgaa cgagaacaac 1020gtgcccgtgc agctgaccag catcagcgcc cccgagcaca
agttcgaggg cctgacccag 1080atcttccaga aggcctacga gcacgagcag cacatcagcg
agagcatcaa caacatcgtg 1140gaccacgcca tcaagagcaa ggaccacgcc accttcaact
tcctgcagtg gtacgtggcc 1200gagcagcacg aggaggaggt gctgttcaag gacatcctgg
acaagatcga gctgatcggc 1260aacgagaacc acggcctgta cctggccgac cagtacgtga
agggcatcgc caagagcagg 1320aagagcggat cc
1332116444PRTArtificial SequenceSynthetic 116Met
Lys Val Lys Leu Leu Val Leu Leu Cys Thr Phe Thr Ala Thr Tyr 1
5 10 15 Ala Asp Thr Ile Cys Ile
Gly Tyr His Ala Asn Asn Ser Thr Asp Thr 20
25 30 Val Asp Thr Val Leu Glu Lys Asn Val Thr
Val Thr His Ser Val Asn 35 40
45 Leu Gly Ser Gly Leu Arg Met Val Thr Gly Leu Arg Asn Ile
Pro Ser 50 55 60
Ile Gln Ser Arg Gly Leu Phe Gly Ala Ile Ala Gly Phe Ile Glu Gly 65
70 75 80 Gly Trp Thr Gly Met
Val Asp Gly Trp Tyr Gly Tyr His His Gln Asn 85
90 95 Glu Gln Gly Ser Gly Tyr Ala Ala Asp Gln
Lys Ser Thr Gln Asn Ala 100 105
110 Ile Asn Gly Ile Thr Asn Lys Val Asn Ser Val Ile Glu Lys Met
Gly 115 120 125 Gly
Asp Pro Glu Trp Asp Arg Glu Ile Asn Asn Tyr Thr Ser Ile Ile 130
135 140 Tyr Ser Leu Ile Glu Glu
Ser Gln Asn Gln Gln Glu Asn Gly Thr Gly 145 150
155 160 Gly Gly Ser Gly Ile Val Gln Gln Gln Asn Asn
Leu Leu Arg Ala Ile 165 170
175 Glu Ala Gln Gln His Leu Leu Gln Leu Thr Val Trp Gly Ile Lys Gln
180 185 190 Leu Gln
Thr Tyr Asn Ala Glu Leu Leu Val Leu Leu Glu Asn Glu Arg 195
200 205 Thr Leu Asp Phe His Asp Ser
Asn Val Lys Asn Leu Tyr Glu Lys Val 210 215
220 Lys Ser Gln Leu Lys Asn Asn Ala Lys Glu Ile Gly
Asn Gly Cys Phe 225 230 235
240 Glu Phe Tyr His Lys Cys Asn Asp Glu Cys Met Glu Ser Val Lys Asn
245 250 255 Gly Thr Tyr
Asp Tyr Pro Lys Tyr Ser Glu Glu Ser Lys Leu Asn Arg 260
265 270 Glu Lys Ile Asp Ser Gly Gly Asp
Ile Ile Lys Leu Leu Asn Glu Gln 275 280
285 Val Asn Lys Glu Met Gln Ser Ser Asn Leu Tyr Met Ser
Met Ser Ser 290 295 300
Trp Cys Tyr Thr His Ser Leu Asp Gly Ala Gly Leu Phe Leu Phe Asp 305
310 315 320 His Ala Ala Glu
Glu Tyr Glu His Ala Lys Lys Leu Ile Ile Phe Leu 325
330 335 Asn Glu Asn Asn Val Pro Val Gln Leu
Thr Ser Ile Ser Ala Pro Glu 340 345
350 His Lys Phe Glu Gly Leu Thr Gln Ile Phe Gln Lys Ala Tyr
Glu His 355 360 365
Glu Gln His Ile Ser Glu Ser Ile Asn Asn Ile Val Asp His Ala Ile 370
375 380 Lys Ser Lys Asp His
Ala Thr Phe Asn Phe Leu Gln Trp Tyr Val Ala 385 390
395 400 Glu Gln His Glu Glu Glu Val Leu Phe Lys
Asp Ile Leu Asp Lys Ile 405 410
415 Glu Leu Ile Gly Asn Glu Asn His Gly Leu Tyr Leu Ala Asp Gln
Tyr 420 425 430 Val
Lys Gly Ile Ala Lys Ser Arg Lys Ser Gly Ser 435
440 1171332DNAArtificial SequenceSynthetic 117ggatccgctc
ttcctgctct tggcgatgcc cttcacgtac tggtcggcca ggtacaggcc 60gtggttctcg
ttgccgatca gctcgatctt gtccaggatg tccttgaaca gcacctcctc 120ctcgtgctgc
tcggccacgt accactgcag gaagttgaag gtggcgtggt ccttgctctt 180gatggcgtgg
tccacgatgt tgttgatgct ctcgctgatg tgctgctcgt gctcgtaggc 240cttctggaag
atctgggtca ggccctcgaa cttgtgctcg ggggcgctga tgctggtcag 300ctgcacgggc
acgttgttct cgttcaggaa gatgatcagc ttcttggcgt gctcgtactc 360ctcggcggcg
tggtcgaaca ggaacaggcc ggcgccgtcc aggctgtggg tgtagcacca 420gctgctcatg
ctcatgtaca ggttgctgct ctgcatctcc ttgttcacct gctcgttcag 480cagcttgatg
atgtcgcctc cggaatcgat cttctcccgg ttcagcttgc tttcctcgct 540gtacttgggg
tagtcgtagg tgccgttctt cacgctttcc atgcactcgt cgttgcactt 600gtggtagaac
tcgaagcagc cgttgccgat ctctttggcg ttgttcttca gctggctctt 660cactttctcg
tacaggttct tcacgttgct gtcgtggaag tccagggtcc tctcattttc 720caggaggacc
aggagttcgg cgttgtaggt ctgcagctgc ttgatgcccc acactgtcag 780ctgcagcaga
tgctgctggg cctcaatggc tctcagcagg ttgttctgct gctgcacaat 840tccagatccg
ccgcctgtgc cgttttcctg ctggttctgg ctttcctcga tcaggctgta 900gatgatgctg
gtgtagttgt tgatctctct gtcccattca ggatcgccgc ccatcttctc 960gatcacgctg
ttcactttgt tggtgatgcc gttgatggcg ttctgggtgc tcttctgatc 1020ggcggcatag
ccagatccct gctcattctg gtggtggtag ccgtaccacc catccaccat 1080gcctgtccat
ccgccctcga taaagccggc aatggcgcca aacaggcctc tgctctggat 1140gctggggata
ttcctcaggc cggtcaccat tctcagtcct gatcccaggt tcacgctgtg 1200ggtcacggtc
acgttctttt ccagcacggt atccacggtg tcggtgctat tgttggcgtg 1260gtagccgata
cagatggtat cggcgtaggt ggcggtaaag gtacacagca gcaccagcag 1320cttcactttc
at
13321181362DNAArtificial SequenceSynthetic 118atgaaaacca tcattgccct
gagctacatc ctgtgcctgg tgttcacaca gaagctgccc 60ggcaacgata atagcaccgc
cacactgtgt ctgggacacc acgccgtgcc taatggcacc 120atcgtgaaaa caatcaccaa
cgaccagatc gaagtgacca atgccacaga gctgggctcc 180ggcctgaagc tggccaccgg
catgagaaat gtgcccgaga agcagaccag aggcatcttt 240ggcgccattg ccggctttat
cgagaatggc tgggagggaa tggtggatgg gtggtacggc 300ttcagacacc agaatagcga
gggaattgga caggccgccg atctgaaatc tacccaggcc 360gccatcgacc agatcaacgg
caagctgaac aggctgatcg gcaagaccgg cggcgatccc 420gagtgggacc gggagatcaa
caactacacc agcatcatct acagcctgat cgaggagagc 480cagaaccagc aggagaacgg
caccggcggc ggcagcggca tcgtgcagca gcagaacaac 540ctgctgcggg ccatcgaggc
ccagcagcac ctgctgcagc tgaccgtgtg gggcatcaag 600cagctgcaga gctacaatgc
cgaactgctg gtcgccctgg aaaaccagca cacaattgat 660ctgacagaca gtgagatgaa
taagctgttc gagaaaacca agaagcagct gagagaaaac 720gccgaggaca tgggcaacgg
ctgcttcaag atctaccaca agtgcgacaa cgcctgcatc 780ggcagcatca gaaacggcac
ctacgaccac gacgtgtaca gagatgaggc cctgaacaac 840cggtttcaga tcaagggctc
cggaggcgac atcatcaagc tgctgaacga gcaggtgaac 900aaggagatgc agagcagcaa
cctgtacatg agcatgagca gctggtgcta cacccacagc 960ctggacggcg ccggcctgtt
cctgttcgac cacgccgccg aggagtacga gcacgccaag 1020aagctgatca tcttcctgaa
cgagaacaac gtgcccgtgc agctgaccag catcagcgcc 1080cccgagcaca agttcgaggg
cctgacccag atcttccaga aggcctacga gcacgagcag 1140cacatcagcg agagcatcaa
caacatcgtg gaccacgcca tcaagagcaa ggaccacgcc 1200accttcaact tcctgcagtg
gtacgtggcc gagcagcacg aggaggaggt gctgttcaag 1260gacatcctgg acaagatcga
gctgatcggc aacgagaacc acggcctgta cctggccgac 1320cagtacgtga agggcatcgc
caagagcagg aagagcggat cc 1362119454PRTArtificial
SequenceSynthetic 119Met Lys Thr Ile Ile Ala Leu Ser Tyr Ile Leu Cys Leu
Val Phe Thr 1 5 10 15
Gln Lys Leu Pro Gly Asn Asp Asn Ser Thr Ala Thr Leu Cys Leu Gly
20 25 30 His His Ala Val
Pro Asn Gly Thr Ile Val Lys Thr Ile Thr Asn Asp 35
40 45 Gln Ile Glu Val Thr Asn Ala Thr Glu
Leu Gly Ser Gly Leu Lys Leu 50 55
60 Ala Thr Gly Met Arg Asn Val Pro Glu Lys Gln Thr Arg
Gly Ile Phe 65 70 75
80 Gly Ala Ile Ala Gly Phe Ile Glu Asn Gly Trp Glu Gly Met Val Asp
85 90 95 Gly Trp Tyr Gly
Phe Arg His Gln Asn Ser Glu Gly Ile Gly Gln Ala 100
105 110 Ala Asp Leu Lys Ser Thr Gln Ala Ala
Ile Asp Gln Ile Asn Gly Lys 115 120
125 Leu Asn Arg Leu Ile Gly Lys Thr Gly Gly Asp Pro Glu Trp
Asp Arg 130 135 140
Glu Ile Asn Asn Tyr Thr Ser Ile Ile Tyr Ser Leu Ile Glu Glu Ser 145
150 155 160 Gln Asn Gln Gln Glu
Asn Gly Thr Gly Gly Gly Ser Gly Ile Val Gln 165
170 175 Gln Gln Asn Asn Leu Leu Arg Ala Ile Glu
Ala Gln Gln His Leu Leu 180 185
190 Gln Leu Thr Val Trp Gly Ile Lys Gln Leu Gln Ser Tyr Asn Ala
Glu 195 200 205 Leu
Leu Val Ala Leu Glu Asn Gln His Thr Ile Asp Leu Thr Asp Ser 210
215 220 Glu Met Asn Lys Leu Phe
Glu Lys Thr Lys Lys Gln Leu Arg Glu Asn 225 230
235 240 Ala Glu Asp Met Gly Asn Gly Cys Phe Lys Ile
Tyr His Lys Cys Asp 245 250
255 Asn Ala Cys Ile Gly Ser Ile Arg Asn Gly Thr Tyr Asp His Asp Val
260 265 270 Tyr Arg
Asp Glu Ala Leu Asn Asn Arg Phe Gln Ile Lys Gly Ser Gly 275
280 285 Gly Asp Ile Ile Lys Leu Leu
Asn Glu Gln Val Asn Lys Glu Met Gln 290 295
300 Ser Ser Asn Leu Tyr Met Ser Met Ser Ser Trp Cys
Tyr Thr His Ser 305 310 315
320 Leu Asp Gly Ala Gly Leu Phe Leu Phe Asp His Ala Ala Glu Glu Tyr
325 330 335 Glu His Ala
Lys Lys Leu Ile Ile Phe Leu Asn Glu Asn Asn Val Pro 340
345 350 Val Gln Leu Thr Ser Ile Ser Ala
Pro Glu His Lys Phe Glu Gly Leu 355 360
365 Thr Gln Ile Phe Gln Lys Ala Tyr Glu His Glu Gln His
Ile Ser Glu 370 375 380
Ser Ile Asn Asn Ile Val Asp His Ala Ile Lys Ser Lys Asp His Ala 385
390 395 400 Thr Phe Asn Phe
Leu Gln Trp Tyr Val Ala Glu Gln His Glu Glu Glu 405
410 415 Val Leu Phe Lys Asp Ile Leu Asp Lys
Ile Glu Leu Ile Gly Asn Glu 420 425
430 Asn His Gly Leu Tyr Leu Ala Asp Gln Tyr Val Lys Gly Ile
Ala Lys 435 440 445
Ser Arg Lys Ser Gly Ser 450 1201362DNAArtificial
SequenceSynthetic 120ggatccgctc ttcctgctct tggcgatgcc cttcacgtac
tggtcggcca ggtacaggcc 60gtggttctcg ttgccgatca gctcgatctt gtccaggatg
tccttgaaca gcacctcctc 120ctcgtgctgc tcggccacgt accactgcag gaagttgaag
gtggcgtggt ccttgctctt 180gatggcgtgg tccacgatgt tgttgatgct ctcgctgatg
tgctgctcgt gctcgtaggc 240cttctggaag atctgggtca ggccctcgaa cttgtgctcg
ggggcgctga tgctggtcag 300ctgcacgggc acgttgttct cgttcaggaa gatgatcagc
ttcttggcgt gctcgtactc 360ctcggcggcg tggtcgaaca ggaacaggcc ggcgccgtcc
aggctgtggg tgtagcacca 420gctgctcatg ctcatgtaca ggttgctgct ctgcatctcc
ttgttcacct gctcgttcag 480cagcttgatg atgtcgcctc cggagccctt gatctgaaac
cggttgttca gggcctcatc 540tctgtacacg tcgtggtcgt aggtgccgtt tctgatgctg
ccgatgcagg cgttgtcgca 600cttgtggtag atcttgaagc agccgttgcc catgtcctcg
gcgttttctc tcagctgctt 660cttggttttc tcgaacagct tattcatctc actgtctgtc
agatcaattg tgtgctggtt 720ttccagggcg accagcagtt cggcattgta gctctgcagc
tgcttgatgc cccacacggt 780cagctgcagc aggtgctgct gggcctcgat ggcccgcagc
aggttgttct gctgctgcac 840gatgccgctg ccgccgccgg tgccgttctc ctgctggttc
tggctctcct cgatcaggct 900gtagatgatg ctggtgtagt tgttgatctc ccggtcccac
tcgggatcgc cgccggtctt 960gccgatcagc ctgttcagct tgccgttgat ctggtcgatg
gcggcctggg tagatttcag 1020atcggcggcc tgtccaattc cctcgctatt ctggtgtctg
aagccgtacc acccatccac 1080cattccctcc cagccattct cgataaagcc ggcaatggcg
ccaaagatgc ctctggtctg 1140cttctcgggc acatttctca tgccggtggc cagcttcagg
ccggagccca gctctgtggc 1200attggtcact tcgatctggt cgttggtgat tgttttcacg
atggtgccat taggcacggc 1260gtggtgtccc agacacagtg tggcggtgct attatcgttg
ccgggcagct tctgtgtgaa 1320caccaggcac aggatgtagc tcagggcaat gatggttttc
at 13621211362DNAArtificial SequenceSynthetic
121atgaaaacca taattgcgct gtcctacata ctgtgtctgg tgtttgccca gaaactgccg
60ggcaatgaca actcaacagc cacgctctgc ttggggcacc atgccgtccc taacgggacc
120attgtgaaaa ccattactaa cgatcagata gaggtgacta atgccaccga gctgggctcc
180ggcttgaaac tggcgaccgg tatgcgcaat gtccccgaaa aacagacccg cgggatattt
240ggggctatcg caggctttat cgagaatggc tgggaaggga tggtggatgg ttggtatggt
300tttagacatc aaaactccga aggcagaggc caggctgccg atctcaagag cacgcaggcc
360gctatagatc agatcaatgg aaagctcaac agactgatcg ggaaaaccgg cggcgatccc
420gagtgggacc gggagatcaa caactacacc agcatcatct acagcctgat cgaggagagc
480cagaaccagc aggagaacgg caccggcggc ggcagcggca tcgtgcagca gcagaacaac
540ctgctgcggg ccatcgaggc ccagcagcac ctgctgcagc tgaccgtgtg gggcatcaag
600cagctgcagt cctacaatgc cgagctgctg gtggctctgg agaatcagca cactattgac
660ctgaccgatt cagagatgaa caaacttttt gagaagacga agaagcagct tagagaaaat
720gcagaggaca tggggaacgg atgctttaaa atatatcata agtgtgataa tgcctgcatc
780ggatcaatta gaaatggtac ctatgatcac gatgtttaca gggacgaagc gctgaataac
840aggttccaga taaaaggctc cggaggcgac atcatcaagc tgctgaacga gcaggtgaac
900aaggagatgc agagcagcaa cctgtacatg agcatgagca gctggtgcta cacccacagc
960ctggacggcg ccggcctgtt cctgttcgac cacgccgccg aggagtacga gcacgccaag
1020aagctgatca tcttcctgaa cgagaacaac gtgcccgtgc agctgaccag catcagcgcc
1080cccgagcaca agttcgaggg cctgacccag atcttccaga aggcctacga gcacgagcag
1140cacatcagcg agagcatcaa caacatcgtg gaccacgcca tcaagagcaa ggaccacgcc
1200accttcaact tcctgcagtg gtacgtggcc gagcagcacg aggaggaggt gctgttcaag
1260gacatcctgg acaagatcga gctgatcggc aacgagaacc acggcctgta cctggccgac
1320cagtacgtga agggcatcgc caagagcagg aagagcggat cc
1362122454PRTArtificial SequenceSynthetic 122Met Lys Thr Ile Ile Ala Leu
Ser Tyr Ile Leu Cys Leu Val Phe Ala 1 5
10 15 Gln Lys Leu Pro Gly Asn Asp Asn Ser Thr Ala
Thr Leu Cys Leu Gly 20 25
30 His His Ala Val Pro Asn Gly Thr Ile Val Lys Thr Ile Thr Asn
Asp 35 40 45 Gln
Ile Glu Val Thr Asn Ala Thr Glu Leu Gly Ser Gly Leu Lys Leu 50
55 60 Ala Thr Gly Met Arg Asn
Val Pro Glu Lys Gln Thr Arg Gly Ile Phe 65 70
75 80 Gly Ala Ile Ala Gly Phe Ile Glu Asn Gly Trp
Glu Gly Met Val Asp 85 90
95 Gly Trp Tyr Gly Phe Arg His Gln Asn Ser Glu Gly Arg Gly Gln Ala
100 105 110 Ala Asp
Leu Lys Ser Thr Gln Ala Ala Ile Asp Gln Ile Asn Gly Lys 115
120 125 Leu Asn Arg Leu Ile Gly Lys
Thr Gly Gly Asp Pro Glu Trp Asp Arg 130 135
140 Glu Ile Asn Asn Tyr Thr Ser Ile Ile Tyr Ser Leu
Ile Glu Glu Ser 145 150 155
160 Gln Asn Gln Gln Glu Asn Gly Thr Gly Gly Gly Ser Gly Ile Val Gln
165 170 175 Gln Gln Asn
Asn Leu Leu Arg Ala Ile Glu Ala Gln Gln His Leu Leu 180
185 190 Gln Leu Thr Val Trp Gly Ile Lys
Gln Leu Gln Ser Tyr Asn Ala Glu 195 200
205 Leu Leu Val Ala Leu Glu Asn Gln His Thr Ile Asp Leu
Thr Asp Ser 210 215 220
Glu Met Asn Lys Leu Phe Glu Lys Thr Lys Lys Gln Leu Arg Glu Asn 225
230 235 240 Ala Glu Asp Met
Gly Asn Gly Cys Phe Lys Ile Tyr His Lys Cys Asp 245
250 255 Asn Ala Cys Ile Gly Ser Ile Arg Asn
Gly Thr Tyr Asp His Asp Val 260 265
270 Tyr Arg Asp Glu Ala Leu Asn Asn Arg Phe Gln Ile Lys Gly
Ser Gly 275 280 285
Gly Asp Ile Ile Lys Leu Leu Asn Glu Gln Val Asn Lys Glu Met Gln 290
295 300 Ser Ser Asn Leu Tyr
Met Ser Met Ser Ser Trp Cys Tyr Thr His Ser 305 310
315 320 Leu Asp Gly Ala Gly Leu Phe Leu Phe Asp
His Ala Ala Glu Glu Tyr 325 330
335 Glu His Ala Lys Lys Leu Ile Ile Phe Leu Asn Glu Asn Asn Val
Pro 340 345 350 Val
Gln Leu Thr Ser Ile Ser Ala Pro Glu His Lys Phe Glu Gly Leu 355
360 365 Thr Gln Ile Phe Gln Lys
Ala Tyr Glu His Glu Gln His Ile Ser Glu 370 375
380 Ser Ile Asn Asn Ile Val Asp His Ala Ile Lys
Ser Lys Asp His Ala 385 390 395
400 Thr Phe Asn Phe Leu Gln Trp Tyr Val Ala Glu Gln His Glu Glu Glu
405 410 415 Val Leu
Phe Lys Asp Ile Leu Asp Lys Ile Glu Leu Ile Gly Asn Glu 420
425 430 Asn His Gly Leu Tyr Leu Ala
Asp Gln Tyr Val Lys Gly Ile Ala Lys 435 440
445 Ser Arg Lys Ser Gly Ser 450
1231362DNAArtificial SequenceSynthetic 123ggatccgctc ttcctgctct
tggcgatgcc cttcacgtac tggtcggcca ggtacaggcc 60gtggttctcg ttgccgatca
gctcgatctt gtccaggatg tccttgaaca gcacctcctc 120ctcgtgctgc tcggccacgt
accactgcag gaagttgaag gtggcgtggt ccttgctctt 180gatggcgtgg tccacgatgt
tgttgatgct ctcgctgatg tgctgctcgt gctcgtaggc 240cttctggaag atctgggtca
ggccctcgaa cttgtgctcg ggggcgctga tgctggtcag 300ctgcacgggc acgttgttct
cgttcaggaa gatgatcagc ttcttggcgt gctcgtactc 360ctcggcggcg tggtcgaaca
ggaacaggcc ggcgccgtcc aggctgtggg tgtagcacca 420gctgctcatg ctcatgtaca
ggttgctgct ctgcatctcc ttgttcacct gctcgttcag 480cagcttgatg atgtcgcctc
cggagccttt tatctggaac ctgttattca gcgcttcgtc 540cctgtaaaca tcgtgatcat
aggtaccatt tctaattgat ccgatgcagg cattatcaca 600cttatgatat attttaaagc
atccgttccc catgtcctct gcattttctc taagctgctt 660cttcgtcttc tcaaaaagtt
tgttcatctc tgaatcggtc aggtcaatag tgtgctgatt 720ctccagagcc accagcagct
cggcattgta ggactgcagc tgcttgatgc cccacacggt 780cagctgcagc aggtgctgct
gggcctcgat ggcccgcagc aggttgttct gctgctgcac 840gatgccgctg ccgccgccgg
tgccgttctc ctgctggttc tggctctcct cgatcaggct 900gtagatgatg ctggtgtagt
tgttgatctc ccggtcccac tcgggatcgc cgccggtttt 960cccgatcagt ctgttgagct
ttccattgat ctgatctata gcggcctgcg tgctcttgag 1020atcggcagcc tggcctctgc
cttcggagtt ttgatgtcta aaaccatacc aaccatccac 1080catcccttcc cagccattct
cgataaagcc tgcgatagcc ccaaatatcc cgcgggtctg 1140tttttcgggg acattgcgca
taccggtcgc cagtttcaag ccggagccca gctcggtggc 1200attagtcacc tctatctgat
cgttagtaat ggttttcaca atggtcccgt tagggacggc 1260atggtgcccc aagcagagcg
tggctgttga gttgtcattg cccggcagtt tctgggcaaa 1320caccagacac agtatgtagg
acagcgcaat tatggttttc at 13621241332DNAArtificial
SequenceSynthetic 124atgaaggcca tcatcgtgct gctgatggtg gtcacaagca
acgccgatag aatctgtacc 60ggcatcacca gcagcaatag ccctcacgtc gtgaaaacag
ctacacaggg cgaagtgaat 120gtgaccggcg tgatccctct gggatcagga ctgaagctgg
ccaatggcac aaagtataga 180cctccagcca agctgctgaa agagagaggc ttttttggag
ctatcgccgg ctttctggaa 240ggcggatggg agggaatgat tgctggatgg catggctaca
catctcatgg cgcacatggc 300gtggcagtgg ctgctgatct gaaatctaca caggaagcca
tcaacaagat caccaagaac 360ctgaacagcc tgagcgagct ggaaggaggc gaccccgagt
gggatcgcga aatcaacaac 420tacacatcta tcatctacag tctgattgag gaaagccaga
accagcagga gaatgggact 480gggggaggct ccggaatcgt gcagcagcag aacaatctgc
tgcgagccat tgaagctcag 540cagcacctgc tgcagctgac agtgtggggc atcaagcagc
tgcaggggag ccagattgaa 600ctggctgtgc tgctgtctaa cgagggcatc atcaatagcg
aggacgaaca tctgctggcc 660ctggaaagaa agctgaagaa gatgctggga cctagcgccg
tggaaatcgg caatggatgc 720tttgagacaa agcacaagtg caaccagacc tgcctggata
gaattgccgc cggaacattt 780gatgccggcg agttttctct gcccaccttc gatagcctga
atatcacatc cggaggcgac 840atcatcaagc tgctgaacga gcaggtgaac aaggagatgc
agagcagcaa cctgtacatg 900agcatgagca gctggtgcta cacccacagc ctggacggcg
ccggcctgtt cctgttcgac 960cacgccgccg aggagtacga gcacgccaag aagctgatca
tcttcctgaa cgagaacaac 1020gtgcccgtgc agctgaccag catcagcgcc cccgagcaca
agttcgaggg cctgacccag 1080atcttccaga aggcctacga gcacgagcag cacatcagcg
agagcatcaa caacatcgtg 1140gaccacgcca tcaagagcaa ggaccacgcc accttcaact
tcctgcagtg gtacgtggcc 1200gagcagcacg aggaggaggt gctgttcaag gacatcctgg
acaagatcga gctgatcggc 1260aacgagaacc acggcctgta cctggccgac cagtacgtga
agggcatcgc caagagcagg 1320aagagcggat cc
1332125444PRTArtificial SequenceSynthetic 125Met
Lys Ala Ile Ile Val Leu Leu Met Val Val Thr Ser Asn Ala Asp 1
5 10 15 Arg Ile Cys Thr Gly Ile
Thr Ser Ser Asn Ser Pro His Val Val Lys 20
25 30 Thr Ala Thr Gln Gly Glu Val Asn Val Thr
Gly Val Ile Pro Leu Gly 35 40
45 Ser Gly Leu Lys Leu Ala Asn Gly Thr Lys Tyr Arg Pro Pro
Ala Lys 50 55 60
Leu Leu Lys Glu Arg Gly Phe Phe Gly Ala Ile Ala Gly Phe Leu Glu 65
70 75 80 Gly Gly Trp Glu Gly
Met Ile Ala Gly Trp His Gly Tyr Thr Ser His 85
90 95 Gly Ala His Gly Val Ala Val Ala Ala Asp
Leu Lys Ser Thr Gln Glu 100 105
110 Ala Ile Asn Lys Ile Thr Lys Asn Leu Asn Ser Leu Ser Glu Leu
Glu 115 120 125 Gly
Gly Asp Pro Glu Trp Asp Arg Glu Ile Asn Asn Tyr Thr Ser Ile 130
135 140 Ile Tyr Ser Leu Ile Glu
Glu Ser Gln Asn Gln Gln Glu Asn Gly Thr 145 150
155 160 Gly Gly Gly Ser Gly Ile Val Gln Gln Gln Asn
Asn Leu Leu Arg Ala 165 170
175 Ile Glu Ala Gln Gln His Leu Leu Gln Leu Thr Val Trp Gly Ile Lys
180 185 190 Gln Leu
Gln Gly Ser Gln Ile Glu Leu Ala Val Leu Leu Ser Asn Glu 195
200 205 Gly Ile Ile Asn Ser Glu Asp
Glu His Leu Leu Ala Leu Glu Arg Lys 210 215
220 Leu Lys Lys Met Leu Gly Pro Ser Ala Val Glu Ile
Gly Asn Gly Cys 225 230 235
240 Phe Glu Thr Lys His Lys Cys Asn Gln Thr Cys Leu Asp Arg Ile Ala
245 250 255 Ala Gly Thr
Phe Asp Ala Gly Glu Phe Ser Leu Pro Thr Phe Asp Ser 260
265 270 Leu Asn Ile Thr Ser Gly Gly Asp
Ile Ile Lys Leu Leu Asn Glu Gln 275 280
285 Val Asn Lys Glu Met Gln Ser Ser Asn Leu Tyr Met Ser
Met Ser Ser 290 295 300
Trp Cys Tyr Thr His Ser Leu Asp Gly Ala Gly Leu Phe Leu Phe Asp 305
310 315 320 His Ala Ala Glu
Glu Tyr Glu His Ala Lys Lys Leu Ile Ile Phe Leu 325
330 335 Asn Glu Asn Asn Val Pro Val Gln Leu
Thr Ser Ile Ser Ala Pro Glu 340 345
350 His Lys Phe Glu Gly Leu Thr Gln Ile Phe Gln Lys Ala Tyr
Glu His 355 360 365
Glu Gln His Ile Ser Glu Ser Ile Asn Asn Ile Val Asp His Ala Ile 370
375 380 Lys Ser Lys Asp His
Ala Thr Phe Asn Phe Leu Gln Trp Tyr Val Ala 385 390
395 400 Glu Gln His Glu Glu Glu Val Leu Phe Lys
Asp Ile Leu Asp Lys Ile 405 410
415 Glu Leu Ile Gly Asn Glu Asn His Gly Leu Tyr Leu Ala Asp Gln
Tyr 420 425 430 Val
Lys Gly Ile Ala Lys Ser Arg Lys Ser Gly Ser 435
440 1261332DNAArtificial SequenceSynthetic 126ggatccgctc
ttcctgctct tggcgatgcc cttcacgtac tggtcggcca ggtacaggcc 60gtggttctcg
ttgccgatca gctcgatctt gtccaggatg tccttgaaca gcacctcctc 120ctcgtgctgc
tcggccacgt accactgcag gaagttgaag gtggcgtggt ccttgctctt 180gatggcgtgg
tccacgatgt tgttgatgct ctcgctgatg tgctgctcgt gctcgtaggc 240cttctggaag
atctgggtca ggccctcgaa cttgtgctcg ggggcgctga tgctggtcag 300ctgcacgggc
acgttgttct cgttcaggaa gatgatcagc ttcttggcgt gctcgtactc 360ctcggcggcg
tggtcgaaca ggaacaggcc ggcgccgtcc aggctgtggg tgtagcacca 420gctgctcatg
ctcatgtaca ggttgctgct ctgcatctcc ttgttcacct gctcgttcag 480cagcttgatg
atgtcgcctc cggatgtgat attcaggcta tcgaaggtgg gcagagaaaa 540ctcgccggca
tcaaatgttc cggcggcaat tctatccagg caggtctggt tgcacttgtg 600ctttgtctca
aagcatccat tgccgatttc cacggcgcta ggtcccagca tcttcttcag 660ctttctttcc
agggccagca gatgttcgtc ctcgctattg atgatgccct cgttagacag 720cagcacagcc
agttcaatct ggctcccctg cagctgcttg atgccccaca ctgtcagctg 780cagcaggtgc
tgctgagctt caatggctcg cagcagattg ttctgctgct gcacgattcc 840ggagcctccc
ccagtcccat tctcctgctg gttctggctt tcctcaatca gactgtagat 900gatagatgtg
tagttgttga tttcgcgatc ccactcgggg tcgcctcctt ccagctcgct 960caggctgttc
aggttcttgg tgatcttgtt gatggcttcc tgtgtagatt tcagatcagc 1020agccactgcc
acgccatgtg cgccatgaga tgtgtagcca tgccatccag caatcattcc 1080ctcccatccg
ccttccagaa agccggcgat agctccaaaa aagcctctct ctttcagcag 1140cttggctgga
ggtctatact ttgtgccatt ggccagcttc agtcctgatc ccagagggat 1200cacgccggtc
acattcactt cgccctgtgt agctgttttc acgacgtgag ggctattgct 1260gctggtgatg
ccggtacaga ttctatcggc gttgcttgtg accaccatca gcagcacgat 1320gatggccttc
at
13321271332DNAArtificial SequenceSynthetic 127atgaaggcca tcatcgtgct
gctgatggtg gtgaccagca acgccgatag aatctgcacc 60ggcatcacca gcagcaatag
cccccatgtg gtgaaaacag ccacccaggg cgaagtgaat 120gtgacaggcg tgatccctct
gggatcagga ctgaagctgg ccaatggcac caagtacaga 180cctcccgcca agctgctgaa
agagagaggc ttctttggcg ccattgccgg atttctggaa 240ggcggctggg agggaatgat
tgccggctgg cacggctata catctcatgg ggcccatggc 300gtggctgtgg ccgccgatct
gaagtctacc caggaagcca tcaacaagat caccaagaac 360ctgaacagcc tgagcgagct
ggaaggaggc gaccccgagt gggatcgcga aatcaacaac 420tacacatcta tcatctacag
tctgattgag gaaagccaga accagcagga gaatgggact 480gggggaggct ccggaatcgt
gcagcagcag aacaatctgc tgcgagccat tgaagctcag 540cagcacctgc tgcagctgac
agtgtggggc atcaagcagc tgcaggggtc ccagattgaa 600ctggccgtgc tgctgtccaa
cgagggcatc atcaacagcg aggatgaaca cctgctggcc 660ctggaacgga agctgaagaa
gatgctgggc ccttctgccg tggagatcgg caacggctgc 720ttcgagacaa agcacaagtg
caaccagacc tgcctggata gaatcgccgc tggcaccttc 780aatgccggcg agttcagcct
gcctaccttc gacagcctga atatcacctc cggaggcgac 840atcatcaagc tgctgaacga
gcaggtgaac aaggagatgc agagcagcaa cctgtacatg 900agcatgagca gctggtgcta
cacccacagc ctggacggcg ccggcctgtt cctgttcgac 960cacgccgccg aggagtacga
gcacgccaag aagctgatca tcttcctgaa cgagaacaac 1020gtgcccgtgc agctgaccag
catcagcgcc cccgagcaca agttcgaggg cctgacccag 1080atcttccaga aggcctacga
gcacgagcag cacatcagcg agagcatcaa caacatcgtg 1140gaccacgcca tcaagagcaa
ggaccacgcc accttcaact tcctgcagtg gtacgtggcc 1200gagcagcacg aggaggaggt
gctgttcaag gacatcctgg acaagatcga gctgatcggc 1260aacgagaacc acggcctgta
cctggccgac cagtacgtga agggcatcgc caagagcagg 1320aagagcggat cc
1332128444PRTArtificial
SequenceSynthetic 128Met Lys Ala Ile Ile Val Leu Leu Met Val Val Thr Ser
Asn Ala Asp 1 5 10 15
Arg Ile Cys Thr Gly Ile Thr Ser Ser Asn Ser Pro His Val Val Lys
20 25 30 Thr Ala Thr Gln
Gly Glu Val Asn Val Thr Gly Val Ile Pro Leu Gly 35
40 45 Ser Gly Leu Lys Leu Ala Asn Gly Thr
Lys Tyr Arg Pro Pro Ala Lys 50 55
60 Leu Leu Lys Glu Arg Gly Phe Phe Gly Ala Ile Ala Gly
Phe Leu Glu 65 70 75
80 Gly Gly Trp Glu Gly Met Ile Ala Gly Trp His Gly Tyr Thr Ser His
85 90 95 Gly Ala His Gly
Val Ala Val Ala Ala Asp Leu Lys Ser Thr Gln Glu 100
105 110 Ala Ile Asn Lys Ile Thr Lys Asn Leu
Asn Ser Leu Ser Glu Leu Glu 115 120
125 Gly Gly Asp Pro Glu Trp Asp Arg Glu Ile Asn Asn Tyr Thr
Ser Ile 130 135 140
Ile Tyr Ser Leu Ile Glu Glu Ser Gln Asn Gln Gln Glu Asn Gly Thr 145
150 155 160 Gly Gly Gly Ser Gly
Ile Val Gln Gln Gln Asn Asn Leu Leu Arg Ala 165
170 175 Ile Glu Ala Gln Gln His Leu Leu Gln Leu
Thr Val Trp Gly Ile Lys 180 185
190 Gln Leu Gln Gly Ser Gln Ile Glu Leu Ala Val Leu Leu Ser Asn
Glu 195 200 205 Gly
Ile Ile Asn Ser Glu Asp Glu His Leu Leu Ala Leu Glu Arg Lys 210
215 220 Leu Lys Lys Met Leu Gly
Pro Ser Ala Val Glu Ile Gly Asn Gly Cys 225 230
235 240 Phe Glu Thr Lys His Lys Cys Asn Gln Thr Cys
Leu Asp Arg Ile Ala 245 250
255 Ala Gly Thr Phe Asn Ala Gly Glu Phe Ser Leu Pro Thr Phe Asp Ser
260 265 270 Leu Asn
Ile Thr Ser Gly Gly Asp Ile Ile Lys Leu Leu Asn Glu Gln 275
280 285 Val Asn Lys Glu Met Gln Ser
Ser Asn Leu Tyr Met Ser Met Ser Ser 290 295
300 Trp Cys Tyr Thr His Ser Leu Asp Gly Ala Gly Leu
Phe Leu Phe Asp 305 310 315
320 His Ala Ala Glu Glu Tyr Glu His Ala Lys Lys Leu Ile Ile Phe Leu
325 330 335 Asn Glu Asn
Asn Val Pro Val Gln Leu Thr Ser Ile Ser Ala Pro Glu 340
345 350 His Lys Phe Glu Gly Leu Thr Gln
Ile Phe Gln Lys Ala Tyr Glu His 355 360
365 Glu Gln His Ile Ser Glu Ser Ile Asn Asn Ile Val Asp
His Ala Ile 370 375 380
Lys Ser Lys Asp His Ala Thr Phe Asn Phe Leu Gln Trp Tyr Val Ala 385
390 395 400 Glu Gln His Glu
Glu Glu Val Leu Phe Lys Asp Ile Leu Asp Lys Ile 405
410 415 Glu Leu Ile Gly Asn Glu Asn His Gly
Leu Tyr Leu Ala Asp Gln Tyr 420 425
430 Val Lys Gly Ile Ala Lys Ser Arg Lys Ser Gly Ser
435 440 1291332DNAArtificial
SequenceSynthetic 129ggatccgctc ttcctgctct tggcgatgcc cttcacgtac
tggtcggcca ggtacaggcc 60gtggttctcg ttgccgatca gctcgatctt gtccaggatg
tccttgaaca gcacctcctc 120ctcgtgctgc tcggccacgt accactgcag gaagttgaag
gtggcgtggt ccttgctctt 180gatggcgtgg tccacgatgt tgttgatgct ctcgctgatg
tgctgctcgt gctcgtaggc 240cttctggaag atctgggtca ggccctcgaa cttgtgctcg
ggggcgctga tgctggtcag 300ctgcacgggc acgttgttct cgttcaggaa gatgatcagc
ttcttggcgt gctcgtactc 360ctcggcggcg tggtcgaaca ggaacaggcc ggcgccgtcc
aggctgtggg tgtagcacca 420gctgctcatg ctcatgtaca ggttgctgct ctgcatctcc
ttgttcacct gctcgttcag 480cagcttgatg atgtcgcctc cggaggtgat attcaggctg
tcgaaggtag gcaggctgaa 540ctcgccggca ttgaaggtgc cagcggcgat tctatccagg
caggtctggt tgcacttgtg 600ctttgtctcg aagcagccgt tgccgatctc cacggcagaa
gggcccagca tcttcttcag 660cttccgttcc agggccagca ggtgttcatc ctcgctgttg
atgatgccct cgttggacag 720cagcacggcc agttcaatct gggacccctg cagctgcttg
atgccccaca ctgtcagctg 780cagcaggtgc tgctgagctt caatggctcg cagcagattg
ttctgctgct gcacgattcc 840ggagcctccc ccagtcccat tctcctgctg gttctggctt
tcctcaatca gactgtagat 900gatagatgtg tagttgttga tttcgcgatc ccactcgggg
tcgcctcctt ccagctcgct 960caggctgttc aggttcttgg tgatcttgtt gatggcttcc
tgggtagact tcagatcggc 1020ggccacagcc acgccatggg ccccatgaga tgtatagccg
tgccagccgg caatcattcc 1080ctcccagccg ccttccagaa atccggcaat ggcgccaaag
aagcctctct ctttcagcag 1140cttggcggga ggtctgtact tggtgccatt ggccagcttc
agtcctgatc ccagagggat 1200cacgcctgtc acattcactt cgccctgggt ggctgttttc
accacatggg ggctattgct 1260gctggtgatg ccggtgcaga ttctatcggc gttgctggtc
accaccatca gcagcacgat 1320gatggccttc at
1332
User Contributions:
Comment about this patent or add new information about this topic: