Patent application title: Vascularization Inhibitors
Inventors:
Tadamitsu Kishimoto (Osaka, JP)
Takashi Nagasawa (Osaka, JP)
Kazunobu Tachibana (Osaka, JP)
IPC8 Class: AG01N3350FI
USPC Class:
435 611
Class name: Measuring or testing process involving enzymes or micro-organisms; composition or test strip therefore; processes of forming such composition or test strip involving nucleic acid nucleic acid based assay involving a hybridization step with a nucleic acid probe, involving a single nucleotide polymorphism (snp), involving pharmacogenetics, involving genotyping, involving haplotyping, or involving detection of dna methylation gene expression
Publication date: 2013-09-12
Patent application number: 20130236889
Abstract:
This invention provides a therapeutic agent for inhibiting
neovascularization, a therapeutic agent for a solid cancer, a therapeutic
agent for a disease pathologically caused by neovascularization, and a
therapeutic agent for repairing a tissue comprising as the effective
ingredient, a substance that potentiates the action of CXCR4.
Based on the finding that vascularization is suppressed in CXCR4 knockout
mice, it becomes possible to prepare a therapeutic agent for suppressing
vascularization, a therapeutic agent for a solid cancer, a therapeutic
agent for a disease pathologically caused by neovascularization, each of
which comprises as the effective ingredient, a substance that inhibits
the action of CXCR4, as well as to prepare a therapeutic agent for
repairing a tissue comprising as the effective ingredient, a substance
that potentiates the action of CXCR4. Methods for treatment are made
possible that use these therapeutic agents.Claims:
1-27. (canceled)
28. A method for screening a substance that inhibits vascularization, treats a solid cancer or treats a disease pathologically caused by neovascularization, based on the inhibition of the action due to CXCR4, which comprises a step of identifying a substance that inhibits the action due to CXCR4 as a substance that inhibits vascularization, treats a solid cancer or treats a disease pathologically caused by neovascularization.
29. The method according to claim 28, which comprises a step of identifying a substance that inhibits (1) the binding between SDF-1 and CXCR4, (2) the signaling from CXCR4 to nuclei, (3) the expression of CXCR4, or (4) the expression of SDF-1 as a substance that inhibits vascularization, treats a solid cancer or treats a disease pathologically caused by neovascularization.
30. A method for evaluating vascularization inhibitory activity, anti-solid tumor activity or anti-disease pathologically caused by neovascularization activity of a substance, based on the inhibition of the action due to CXCR4, which comprises a step of measuring the inhibitory activity of the action due to CXCR4.
31. The method according to claim 30, which comprises a step of measuring the inhibitory activity of (1) the binding between SDF-1 and CXCR4, (2) the signaling from CXCR4 to nuclei, (3) the expression of CXCR4, or (4) the expression of SDF-1.
Description:
TECHNICAL FIELD
[0001] This invention relates to a novel vascularization inhibitor, an anti-solid cancer agent, and a therapeutic agent for a disease pathologically caused by neovascularization, each comprising a CXCR4 inhibitor as the effective ingredient. Further, the invention relates to a tissue-repairing agent comprising a CXCR4 potentiator as the effective ingredient.
BACKGROUND ART
[0002] In the past, it has been known that when tumor cells invade out of the blood vessels, vascular endothelial cells rupture. It is also known that neovascularization is deeply involved in the proliferation and migration of cancer and that tumor cells produce and release a variety of neovascularization factors. Especially, neovascularization is considered crucial to the proliferation of solid tumors.
[0003] Therefore, a substance that inhibits the neovascularization has the potential to be an anticancer agent that is provided with a novel mode of action. For this reason, several types of neovascularization inhibitory substances such as steroids and metabolic products of microorganisms have already been tested for use. ("Manual for Studies on Cancer Invasion and Metastasis," The Cancer Metastasis Society Ed., Kinhodo Publisher, 159-182 (1994)). However, it is strongly desired that novel neovascularization inhibitory substances with the action of more effectively inhibiting the proliferation and metastasis of cancers be discovered.
DISCLOSURE OF THE INVENTION
[0004] This invention provides a vascularization inhibitor, an anti-solid cancer agent, or a therapeutic agent for a disease pathologically caused by neovascularization, each comprising an inhibitor of a chemokine receptor with the action of more effectively inhibiting the proliferation, invasion and metastasis of a cancer. Further, an object of the invention is to provide a tissue-repairing agent comprising as the effective ingredient, a potentiator of chemokine receptors.
[0005] Specifically, the present inventors pursued extensive research in order to solve the above-identified problems; and as a result, they have discovered that when knockout mice lacking in pre-B-cell growth stimulating factor/stromal-cell derived factor (hereafter referred to as "PBSF/SDF-1" or "SDF-1") which is a CXC chemokine as well as in CXCR4 which is a chemokine receptor are created, the vascularization in the mice is suppressed, and namely, suppression of CXCR4 results in the suppression of vascularization. Such finding means that the chemokine receptor CXCR4 is essential for neovascularization.
[0006] Neovascularization of living tissues generally occurs through remodeling of the preexisting vascular system when they grow to perform their specific functions during development. Analyses of the mutant mice have determined that the molecules required by early vascular systems are largely receptor tyrosine kinases and their ligands. (Risau, w. Nature 386, 671-674 (1997); Folkman, J. & D'Amore, P. A. Cell 87, 1158-1155 (1996); and Lindahl, P., et al., science 277, 242-245 (1997)). However, substances responsible for vascularization during organogenesis have not yet been identified, because most of these mice die during early gestation before development of their tissues.
[0007] The structure of chemokine receptor CXCR4 according to this invention has already been known. (Bleul, C. C. et al., Nature 382, 829-883 (1996); Oberlin, E. et al., Nature 382, 888-835 (1996); and Nagasawa, T. et al., Proc. Natl. Acad. Sci. USA 93, 14726-14729 (1996)). CXCR4 is a seven-transmembrane-spanning G-protein-coupled protein and a receptor for PBSF/SDF-1 which is a CXC chemokine. The aforementioned factor is thought to be responsible for B-cell lymphopoiesis, bone marrow myelopoiesis and cardiac ventricular septum formation (Nagasawa, T. et al., Nature 382, 685-688 (1996)). CXCR4 also functions as a co-receptor for T-cell-line-tropic HIV-1 (Feng, Y. et al., Science 272, 872-877 (1996)). CXCR4 has further been reported to be expressed in cultured endothelial cells (Volin, M. V. et al., Biochem. Biophys. Res. Commun. 242, 46-53 (1998)).
[0008] In addition, the present inventors have discovered that the above-mentioned CXCR4 is expressed in developing vascular endothelial cells, and that mice lacking CXCR4 or its ligand PBSF/SDF-1 show defective formation of the large vessels being supplied to the gastrointestinal tract. Such finding means that the CXCR4 and PBSF/SDF-1 signaling systems are essential for the formation of median arteriovein supplying nutrient to the gastrointestinal tract. Furthermore, the present inventors have found that mice lacking CXCR4 are apt to die in utero just as seen in mice lacking PGSF/SDF-1. Such finding suggests that CXCR4 is the most critical, primary physiological receptor for PBSF/SDF-1.
[0009] Based on the foregoing observations by the present inventors, it is contemplated that substances capable of inhibiting the action due to CXCR4 may inhibit vascularization and thus can be effective anticancer agents, since the vascularization is essential for the maintenance and enlargement of cancerous tissues.
[0010] It is likewise contemplated that substances capable of inhibiting CXCR4 can be therapeutic agents for the treatment of diseases involving neovascularization.
[0011] It is further contemplated that promotion of the action due to CXCR4 accelerates vascularization and thus can be a remedy for a disease where the vascularization is desired.
[0012] More specifically, as will be summarized below, this invention provides a vascularization inhibitor, an anti-solid tumor agent, or a therapeutic agent for a disease pathologically caused by neovascularization, each comprising as the effective ingredient, a substance that inhibits the action due to CXCR4. The invention also provides a tissue-repairing agent or the like comprising as the effective ingredient, a substance that potentiates the action due to CXCR4.
[0013] That is, this invention provides a vascularization inhibitor comprising a CXCR4 inhibitor as the effective ingredient.
[0014] Also, this invention provides an anti-solid cancer agent comprising a CXCR4 inhibitor as the effective ingredient.
[0015] Further, this invention provides a therapeutic agent for a disease pathologically caused by neovascularization, comprising a CXCR4 inhibitor as the effective ingredient.
[0016] Still further, the invention provides a tissue-repairing agent comprising a CXCR4 potentiator as the effective ingredient.
[0017] Because the formation of median or large arterioveins is essential for the maintenance and enlargement of a cancer tissue that exceeds a certain size, the vascularization inhibitor of this invention blocks the CXCR4 or PBSF/SDF-1 signaling system, thus suppressing the maintenance and enlargement of the cancer tissue.
[0018] The finding obtained in this invention suggests the possibility that the CXCR4 and PBSF/SDF-1 signaling systems contribute to the universal vascularization. Therefore, in diseases of which particular kind of cancer or neovascularization is the major pathological cause, it is likely that CXCR4 or PBSF/SDF-1 is deeply involved in the pathological cause; in this case, there is the possibility that these diseases can be suppressed by blocking CXCR4 or PBSF/SDF-1 individually or concurrently with other molecules.
[0019] In the present specification and the drawings, the abbreviations for bases or amino acids are those following the IUPAC-IUB Commission on Biochemistry Nomenclature or those based on what is customary in the art. Illustrated below are their examples. Where amino acids are meant and there may be their optical isomers, they represent L-forms unless otherwise indicated.
DNA: deoxyribonucleic acid cDNA: complementary deoxyribonucleic acid A: adenine T: thymine G: guanine C: cytosine RNA: ribonucleic acid mRNA: messenger ribonucleic acid G or Gly: glycine A or Ala: alanine V or Val: valine L or Leu: leucine I or Ile: isoleucine S or Ser: serine T or Thr: threonine C or Cys: cysteine M or Met: methionine E or Glu: glutamic acid D or Asp: aspartic acid K or Lys: lysine R or Arg: arginine H or His: histidine F or Phe: phenylalanine Y or Tyr: tyrosine W or Trp: tryptophan P or Pro: proline N or Asn: asparagine Q or Gln: glutamine BSA: bovine serum albumin FBS: fetal bovine serum PBS: phosphate buffer saline SDS: sodium dodecyl sulfate
BRIEF DESCRIPTION OF THE DRAWINGS
[0020] FIG. 1 is a graph showing a targeting strategy for the CXCR4 gene. In the figure, there are shown the CXCR4 wild-type allele at the top, a targeting vector in the middle, and a predicted mutant allele at the bottom. The coding regions of the genes are indicated by black boxes. Empty boxes indicate the 5'- and 3'-untranslated regions. Dotted lines indicate homologous fragments used in the targeting vector. Probe A is an external probe for Southern hybridization. Restriction sites are E (EcoRI), Sh (SphI), and X (XhoI), respectively.
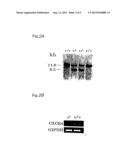
[0021] FIG. 2A is a photograph showing the Southern blot analysis of tail DNAs from wild-type (+/+) and heterozygous mutant (+/-) mice. The EcoRI-EcoRI fragments from the 11.8-kb wild-type and the 8.2-kb targeted allele which were identified by probe A are shown in the figure.
[0022] FIG. 2B is a photograph showing the RT-PCR amplification analysis of CXCR4 expression. Total RNAs were prepared from E18.5 wild-type and homozygous mutant embryos, and amplified with CXCR4-specific primers. The RT-PCR amplification employed G3PDH mRNA, which was universally expressed, as a control for the presence of any amplifiable RNA.
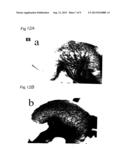
[0023] FIG. 3 is a photograph showing defects of gastrointestinal blood vessels at the mesentery and mid-gut loop region in a wild-type CXCR4.sup.-/- embryo at E13.5, resulting from immunohistostaining of the mesentery and intestine with anti-PECAM-1 antibody. Arrow indicates a large branch of superior mesenteric artery or superior mesenteric vein being supplied to the small intestine in the wild-type mesentery. "du" represents duodenum; "p," the proximal part of mid-gut loop; and "dm," the distal part of mid-gut loop.
[0024] FIG. 4 is a photograph showing defects of gastrointestinal blood vessels at the cross-sections of mesentery in the wild-type CXCR4.sup.-/- embryo at E13.5, resulting from immunohistostaining of the mesentery and intestine with the anti-PECAM-1 antibody. "a" represents artery, and "v" vein.
[0025] FIG. 5 is a photograph showing defects of gastrointestinal blood vessels in the jejunum in a wild-type CXCR4.sup.-/- embryo at E17.5, resulting from immunohistostaining of the mesentery and intestine with the anti-PECAM-1 antibody. Arrow indicates a large branch of superior mesenteric artery or superior mesenteric vein being supplied to the small intestine in the wild-type mesentery.
[0026] FIG. 6 is a photograph showing defects of gastrointestinal blood vessels at the more distal part of the jejunum in the wild-type CXCR4.sup.-/- embryo at E17.5, resulting from immunohistostaining of the mesentery and intestine with the anti-PECAM-1 antibody. Arrow indicates a large branch of superior mesenteric artery or superior mesenteric vein being supplied to the small intestine in the wild-type mesentery.
[0027] FIG. 7 is a photograph showing defects of gastrointestinal blood vessels at the mesentery and mid-gut loop regions in a mutant CXCR4.sup.-/- embryo at E13.5, resulting from immunohistostaining of the mesentery and intestine of mutant with the anti-PECAM-1 antibody. "p" represents the proximal part of mid-gut loop; and "dm," the distal part of mid-gut loop.
[0028] FIG. 8 is a photograph showing defects of gastrointestinal blood vessels at the cross-sections of mesentery in the mutant CXCR4.sup.-/- embryo at E13.5, resulting from immunohistostaining of the mesentery and intestine of the mutant with the anti-PECAM-1 antibody.
[0029] FIG. 9 is a photograph showing defects of gastrointestinal blood vessels in the jejunum in a mutant CXCR4.sup.-/- embryo at E17.5, resulting from immunohistostaining of the mesentery and intestine of the mutant with the anti-PECAM-1 antibody.
[0030] FIG. 10 is a photograph showing defects of gastrointestinal blood vessels at a more distal part of the jejunum in the mutant CXCR4.sup.-/- embryo at 17.5, resulting from immunohistostaining of the mesentery and intestine of the mutant with the anti-PECAM-1 antibody.
[0031] FIG. 11 is a photograph showing defects of gastrointestinal blood vessels, which are a haemorrhagic lesion of the unstained intestine of a mutant mouse, an E16.5 mutant CXCR4.sup.-/- embryo, resulting from immunohistostaining of the mesentery and intestine of the mutant with the anti-PECAM-1 antibody.
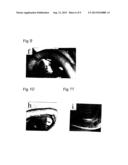
[0032] FIG. 12A is a photograph showing the result of immunohistostaining the stomach of an E13.5 wild-type with the anti-PECAM-1 antibody. Arrow indicates a large vessel only seen in the wild-type.
[0033] FIG. 12B is a photograph showing the result of immunohistostaining the stomach of an E13.5 mutant with the anti-PECAM-1 antibody.
[0034] FIG. 12C is a photograph showing the result of immunohistostaining the stomach of an E15.5 wild-type with the anti-PECAM-1 antibody. Inset in the photograph shows haematoxylin and eosin-stained sections of large vessels in the wall of stained stomach at E15.5. Arrow indicates a large vessel only seen in the wild-type.
[0035] FIG. 12D is a photograph showing the result of immunohistostaining the stomach of an E15.5 mutant with the anti-PECAM-1 antibody.
[0036] FIG. 13A is a photograph showing CXCR4 and PBSF/SDF-1 expression in a gastrointestinal tract tissue through in situ hybridization. Serial sections of the wild-type mesentery connecting to the mid-gut loop were stained with haematoxylin and eosin. "m" represents mesentery, "i" intestine, "a" superior mesenteric artery, and "v" superior mesenteric vein.
[0037] FIG. 13B is a photograph showing CXCR4 and PBSF/SDF-1 expression in a gastrointestinal tract tissue through in situ hybridization. Hybridization was done with a CXCR4-specific probe. Arrows indicate the stained endothelial cells of the mesenteric vessels.
[0038] FIG. 13C is a photograph showing CXCR4 and PBSF/SDF-1 expression in a gastrointestinal tract tissue through in situ hybridization. Hybridization was done with a PBSF/SDF-1-specific probe. PBSF/SDF-1 was expressed in mesenchymal cells surrounding the endothelial cells in the mesentery.
[0039] FIG. 13D is a photograph showing CXCR4 and PBSF/SDF-1 expression in the gastrointestinal tract tissue through in situ hybridization. Serial sections of the wild-type mesentery connecting to the mid-gut loop were stained with haematoxylin and eosin. FIG. 13D is an enlargement of blood vessels arising from the superior mesenteric artery shown in FIG. 13A, where strong expression of CXCR4 was observed in the vascular endothelial cells.
[0040] FIG. 13E is a photograph showing CXCR4 and PBSF/SDF-1 expression in the gastrointestinal tract tissue through in situ hybridization. Hybridization was done with the CXCR4-specific probe. FIG. 13E is an enlargement of blood vessels arising from the superior mesenteric artery shown in FIG. 13B, where strong expression of CXCR4 was observed in the vascular endothelial cells. Arrow indicates the stained endothelial cells of the mesenteric vessels.
[0041] FIG. 13F is a photograph showing CXCR4 and PBSF/SDF-1 expression in a gastrointestinal tract tissue through in situ hybridization. It is a section of an E18.5 wild-type embryonic bone marrow, showing CXCR4 expression in hematopoietic cells but no expression in spindle-shaped stroma cells.
BEST MODE FOR CARRYING OUT THE INVENTION
[0042] The vascularization inhibitor, the anti-solid cancer agent, or the therapeutic agent for a disease pathologically caused by neovascularization according to this invention comprises as the effective ingredient, a substance that inhibits the action of CXCR4 which is a chemokine receptor. On the other hand, the tissue-repairing agent according to the invention comprises as the effective ingredient, a substance that potentiates the action of CXCR4.
[0043] The amino acid sequence of CXCR4 has already been known. Specifically, the amino acid sequence of human CXCR4 and the amino acid sequence of murine CXCR4 are set forth in SEQ ID NOs: 1 and 3, respectively. The base sequence of human CXCR4 and the base sequence of murine CXCR4 are set forth in SEQ ID NO: 2 (base positions 1-1056) and SEQ ID NO: 4 (base positions 1-1077), respectively.
[0044] Also, the amino acid sequence of SDF-1, which is a ligand binding to CXCR4, has already been known. There are two types of SDF-1 differing in the length of amino acid sequence, i.e., SDF-1-α and SDF-1-β. Specifically, the amino acid sequence of human SDF-1-α is set forth in SEQ ID NO: 5 and its base sequence in SEQ ID NO: 6 (base positions 474-740). Human SDF-1-β (SEQ ID No: 9) is derived from human SDF-1-α by appending four amino acid residues, Arg Phe Lys Met, to a C-terminus thereof.
[0045] The amino acid sequence of murine SDF-1-α is set forth in SEQ ID NO: 7 and its base sequence in SEQ ID NO: 8 (base positions 82-348). Murine SDF-1-β (SEQ ID No: 10) is derived from murine SDF-1-α by appending four amino acid residues, Arg Leu Lys Met, to a C-terminus thereof. For human and murine SDF-1's, the sequence of from the 1st amino acid (Met) to the 21st amino acid (Gly) is a signal sequence.
[0046] CXC chemokines that have hitherto been known include, in addition to PBSF/SDF-1 mentioned above, IL-8 (Yoshimura., T. et al., Proc. Natl. Acad. Sci. U.S.A., 84, 9233-9237 (1987)), NAP-2 (Walz. A., et al., Biochem. Biophys. Res. Comun., 159, 969-975 (1989)), NAP-4, GRO α (Richmondo, A. et al., J. Cell. Biochem., 36, 185-198 (1988)), GRO β (Haskill, S. et al., Proc. Natl. Acad. Sci. U.S.A., 87, 77732-7736 (1990)), GRO γ (Haskill, S. et al., ibid. (1990)), GCP-2 (Proost, P. et al., J. Immunol., 150, 1000-1010 (1993)), ENA-78 (Wayz, A. et al., J. Exp. Med., 174, 1355-1362 (1991)), PF-4 (Deuel, T. F. et al., Proc. Natl. Acad. Sci. U.S.A. 74, 2256-2258 (1977)), and IP-10 (Dewald, B. et al., Immunol. Lett., 32, 81-84 (1992)).
[0047] There are no particular limitations to substances that inhibit the action due to CXCR4 that can be used in this invention; and they may be substances that inhibit the action due to CXCR4 with the result of inhibition of neovascularization.
[0048] Specifically mentioned are: (1) a substance based on inhibition of the binding itself between the ligand (SDF-1) and the receptor (CXCR4); (2) a substance based on inhibition of the signaling from CXCR4 to nuclei; (3) a substance that inhibits the expression of CXCR4 itself; and (4) a substance that inhibits the expression of SDF-1 itself.
[0049] (1) For the substance that inhibits the binding itself between SDF-1 and CXCR4, there are a substance that inhibits SDF-1 and a substance that inhibits CXCR4.
[0050] More specifically, the substance that inhibits SDF-1 is classified into a substance that inhibits CXCR4 in antagonistic competition with SDF-1 and a substance that inhibits SDF-1 from binding to CXCR4 by binding to SDF-1. For the substance that inhibits CXCR4 in antagonistic competition with SDF-1, there are concretely mentioned a protein having a SDF-1-like structure, a fused protein of the foregoing protein with another peptide or polypeptide, a low molecular weight compound having a structure similar to a partial peptide of SDF-1 or a binding site of SDF-1, and the like.
[0051] For the substance that inhibits SDF-1 from binding to CXCR4 by binding to SDF-1, there are concretely mentioned an anti-SDF-1 antibody, a fragment thereof having possessing binding activity, a fused protein possessing binding activity to SDF-1, a substance that induces a structural change in SDF-1, a low molecular weight compound that binds to the CXCR4-binding site of SDF-1, and the like.
[0052] More specifically, the substance that inhibits CXCR4 is classified into a substance that inhibits CXCR4 in antagonistic competition with CXCR4 for binding to SDF-1 and a substance that inhibits SDF-1 from binding to CXCR4 by binding to CXCR4. For the substance that inhibits CXCR4 in antagonistic competition with CXCR4 for binding to SDF-1, there are concretely mentioned a soluble CXCR4 that antagonizes CXCR4 in inhibition, a protein having a CXCR4-like structure, a fused protein of the foregoing protein with another peptide or polypeptide, a low molecular weight compound having a structure similar to a partial peptide of CXCR4 or a binding site of CXCR4, and the like.
[0053] For the substance that inhibits SDF-1 from binding to CXCR4 by binding to CXCR4, there are concretely mentioned an anti-CXCR4 antibody, a fragment thereof possessing its binding activity, a fused protein possessing binding activity to CXCR4, a substance that induces a structural change in SDF-1, a low molecular weight compound that binds to the SDF-1-binding site, and the like.
[0054] Examples of the substance that inhibits the binding itself between CXCR4 and SDF-1 include T22 (T. Murakami, et al., J. Exp. Med., 186, 1389-1393 (1997)), ALX40-4C (J. Exp. Med., 186, 1395-1400 (1997)), AMD3100 (J. Exp. Med., 186, 1383-1388 (1997); Nat. Med., 4, 72-77 (1998)), and the like. As to the methods for preparation of these substances, they can, for example, be done by the method as described in J. Exp. Med., 186, 1189-1191 (1997) with any possible modifications.
[0055] (2) There is no particular limitation to the substance based on inhibition of the signaling from CXCR4 to nuclei insofar as it is a substance having such action. For the substance based on inhibition of the signaling from CXCR4 to nuclei, there are mentioned inhibitors of the signaling system existing downstream of a G protein-coupled protein, such as an MAK cascade inhibitor, a phospholipase C (PLC) inhibitor and a kinase inhibitor for PI3 kinase.
[0056] (3) For the substance that inhibits the expression of CXCR4 itself, there are mentioned a substance that apparently makes CXCR4 disappear on cells and a substance that inhibits the expression of CXCR4 itself. A specific example of the substance that apparently makes CXCR4 disappear on cells is a substance that induces down-regulation of CXCR4. The "induction of down-regulation of CXCR4" specifically means such a function that it acts on the cell membrane to alter mobility thereof and thereby to make CXCR4 disappear from the cell membrane. For example, dexamethasone is mentioned as a substance possessing the function.
[0057] For the substance that inhibits the expression of CXCR4 itself, there are concretely mentioned an antigene, an antisense (antisense oligonucleotide and antisense RNA expressed by antisense vector), a ribozyme, and a substance that inhibit the expression control site of CXCR4 such as a promoter or an enhancer.
[0058] From the examples which will be described later, it has become evident that when a vector containing a part of the CXCR4 gene is used to cause the deficiency of CXCR4, vascularization is suppressed. Therefore, the inhibition of CXCR4 by the antigene, antisense, or ribozyme of CXCR4 will suppress vascularization.
[0059] Antisense oligonucleotides that can preferably be used in this invention include CXCR4 genes, SDF-1 genes against CXCR4, nucleotides (DNAs or RNAs) selectively hybridizable to the genes of substances that are involved in the signaling system based on CXCR4, and derivatives thereof (such as antisense oligonucleotides). This invention, for example, encompasses antisense oligonucleotides that hybridize to any site of the base sequence of human CXCR4 gene as set forth in SEQ ID NO: 2.
[0060] Preferably, the antisense oligonucleotide is an antisense oligonucleotide to at least 20 consecutive nucleotides within the base sequence set forth in SEQ ID NO: 2. More preferably, the antisense oligonucleotide is the at least 20 consecutive nucleotides containing a translation initiation codon.
[0061] As used herein, "antisense oligonucleotide" is not only one having nucleotides that correspond to the nucleotides constituting the predetermined region of DNA or RNA and that are all complementary thereto, but also may allow one or more mismatches of nucleotide to be present therein insofar as the oligonucleotide and the DNA or the RNA are able to selectively and stably hybridize to the base sequence set forth in SEQ ID NO: 2. By "selectively and stably hybridize" is meant those having at least 70%, preferably at least 80%, more preferably at least 90%, most preferably 95% or greater homology of base sequence in the nucleotide sequence region of at least 20, and preferably 30 consecutive nucleotides. In the present specification, "homology" indicates "identity."
[0062] When the oligonucleotide derivative used in this invention is a deoxyribonucleotide, the structure of each derivative is represented by formula 1:
##STR00001##
[0063] In the formula, X may independently be any of oxygen (O), sulfur (S), a lower alkyl group, a primary amine and a secondary amine. Y may independently be either oxygen (O) or sulfur (S). B is selected from adenine, guanine, thymine, or cytosine, and is principally an oligonucleotide complementary to DNA or RNA of the human CXCR4 gene. R is independently hydrogen (H), a dimethoxytrytyl group or a lower alkyl group. n is from 7 to 28.
[0064] Preferable oligonucleotide derivatives are not limited to oligonucleotides that have not been modified, but may be modified oligonucleotides, as will be illustrated below. These modified forms include lower alkyl phosphonate derivatives of such types as methyl phosphonate or ethyl phosphonate, phosphorothioate derivatives, phosphoroamidates, and the like.
##STR00002##
[0065] These oligonucleotide derivatives can be obtained by standard methods as described below. The oligonucleotides of formula (I) wherein X and Y are both O may readily be prepared with a commercial DNA synthesizer such as one available from Applied Biosystems; for their preparation method, solid phase synthesis employing phosphoroamidites, solid phase synthesis employing hydrogen phosphonates and the like can be used to obtain them.
[0066] The phosphoric triester modified forms wherein X is a lower alkoxy group can be obtained by standard methods: for example, oligonucleotides obtained by chemical synthesis are treated with a DMF/methanol/2,6-lutidine solution of tosyl chloride. The alkyl phosphonate modified forms wherein X is an alkyl group can be obtained according standard methods, for example, by using phosphoamidites. The phosphorothioate modified forms wherein X is S can be obtained according standard methods, for example, by solid phase synthesis using sulfur, or alternatively by solid phase synthesis using tetraethyl thiuram disulfide. The phosphorodithioate modified forms wherein X and Y are both S can be obtained according to solid phase synthesis, for example, by converting bisamidites into thioamidites and allowing sulfur to act on the thioamidites. The phosphoroamidate modified forms wherein X is a primary or secondary amine can be obtained according to solid phase synthesis, for example, by treating hydrogenphosphonates with a primary or secondary amine; alternatively, they can be obtained by oxidizing amidites with tert-butyl hydrogen peroxide.
[0067] Purification and the assurance of purity may be carried out by high speed liquid chromatography or polyacrylamide gel electrophoresis. The confirmation of molecular weights may be carried out by electrospray ionization mass spectrometry or fast atom bombardment-mass spectrometry.
[0068] The antisense oligonucleotide derivatives used in this invention act on a CXCR4 receptor or a ligand thereof, as well as on cells producing a signaling substance based on CXCR4, and bind to DNA or RNA encoding the peptide, thereby inhibiting its transcription or translation, or promoting the decomposition of mRNA; as a result, they possess an inhibitory effect on the action due to CXCR4 by suppressing the expression of the peptide.
[0069] Consequently, the antisense oligonucleotides used in this invention have utility in inhibiting neovascularization. Neovascularization inhibitors comprising the antisense oligonucleotides of this invention are useful as therapeutic agents for cancers, particularly solid cancers.
[0070] Preparation of CXCR4 antisense vectors may follow the methods that are commonly used. Specifically, cDNA encoding CXCR4 is linked to an AAV vector (adeno-associated virus vector), MLV vector (murine leukemic virus vector), HIV-vector, or the like in the antisense direction. By "antisense direction" is meant linking to the 3'-side of the cDNA to be introduced in the downstream of the promoter. The antisense RNAs synthesized from cDNAs contained in these vectors constitutively suppress the expression of CXCR4 in hosts.
[0071] Antisense DNAs or RNAs can be introduced into cells by using means such as the liposome method, the HVJ liposome method, or the positively charged liposome method. The introduction of a CXCR4 antisense DNA or RNA allows for the constitutive inhibition of expression of CXCR4.
[0072] (4) For the substance that inhibits the expression of SDF-1 itself against CXCR4, there are mentioned an antisense substance that inhibits the expression of SDF-1 and a substance that inhibits the expression control site such as a promoter.
[0073] The antibodies described above that can be used in this invention, such as anti-SDF-1 antibodies and anti-CXCR4 antibodies can be prepared as polyclonal or monoclonal antibodies using techniques that are known in the art. Especially, monoclonal antibodies derived from mammals are preferred as the antibodies that are used in the invention. The monoclonal antibodies derived from mammals include those produced by hybridomas and those produced by the hosts that have been transformed with expression vectors containing antibody genes by techniques in genetic engineering. These antibodies are those which possess the above-mentioned properties.
[0074] Antibody-producing hybridomas can principally be prepared using techniques known in the art in the following manner. That is, a desired antigen is used as the sensitizing antigen to carry out immunization according to a conventional immunization method; and the resulting immunized cells are fused with parent cells known in the art by a conventional cell fusion method, and monoclonal antibody-producing cells are subjected to cloning by a conventional cloning method.
[0075] The mammals to be immunized with the sensitizing antigen are not particularly limited, but should preferably be selected in consideration of their compatibility with the parent cells to be used in the cell fusion. Generally, animals belonging to the rodent family, such as mouse, rat and hamster are used. Immunization of animals with the sensitizing antigen is carried out by a method known in the art.
[0076] For the mammalian myeloma cells used as the other parent cells for fusion with the immunocytes, a variety of cell lines already known in the art may appropriately be employed. The cell fusion between the immunocytes and myeloma cells may be carried out basically according to a conventional method, such as the method of Milstein et al (Kohler, G. and Milstein, C., Methods Enzymol., 73, 3-46 (1981)) with any possible modifications.
[0077] The obtained hybridomas are selected by culturing in a common selection medium, such as HAT medium (medium containing hypoxanthine, aminopterin and thymidine). Culturing in the HAT medium is continued for a sufficient time to allow killing of all the cells other than the targeted hybridomas (the non-fused cells), which is usually from a few days to a few weeks. The usual limiting dilution method is then performed for screening and cloning of hybridomas producing the target antibodies. Antibodies may be acquired from the thus-obtained hybridomas following a method that is commonly employed.
[0078] In addition to acquisition of the hybridomas by immunizing an animal other than human with the antigen as described above, human lymphocytes are sensitized in vitro with a desired antigen protein or antigen-expressing cell, and the sensitized B lymphocytes are fused with human myeloma cells, such as U266, whereby desired humanized antibodies possessing the binding activity toward the desired antigen or antigen-expressing cell may be obtained. Furthermore, the desired humanized antibodies may be acquired according to the aforementioned method by administering the antigen or the antigen-expressing cell to a transgenic animal having a reparatory of human antibody genes.
[0079] An antigen gene is cloned as a monoclonal antibody from the hybridoma and incorporated into a suitable vector; and this is introduced into a host, and the gene manipulation technology is used to produce a recombinant antibody, which may then be used in this invention. See, for example, Carl, A. K. Borrenbaeck, James, W. Larrick, Therapeutic Monoclonal Antibodies, published in the United Kingdom by Macmillan Publishers Ltd. 1990.
[0080] In this invention, recombinant antibodies that have been artificially modified for the purpose of lowering heterogeneous antigenicity against humans, such as chimeric antibodies (European Patent Publication EP125023) and humanized antibodies (European Patent Publication EP125023), may be used. These antibodies can be produced by known methods.
[0081] The chimeric antibody comprises the variable region of an antibody derived from a mammal other than human and the constant region (C region) derived from a human antibody. The humanized antibody comprises a complementarity-determining region of an antibody derived from a mammal other than human, a framework region (FR) derived from a human antibody and a C region. They are useful as the effective ingredients in this invention because of diminished antigenicity in the human body.
[0082] The antibodies used in this invention may be fragments of antibody or modified substances thereof insofar as they can desirably be used in the invention. The fragments of antibody, for example, include Fab, F(ab')2, Fv, and single chain Fv (scFv) obtainable by linking Fv from H chain and Fv from L chain via a suitable linker. Specifically, an antibody is treated with an enzyme such as papain or pepsin to produce antibody fragments. Alternatively, genes encoding these antibody fragments are constructed and are introduced into an expression vector, after which they are expressed in suitable host cells.
[0083] The phage library method can be utilized to obtain the antibodies that are used in this invention. (Marks, C. et al., The New England Journal Medicine 335, 730-733). For example, a cDNA library is acquired from human B cells that comprises a human antibody V region, such as the gene encoding scFv. This cDNA library is introduced into a phage vector such as the M13 phage surface presenting vector, and this is allowed to infect E. coli. The cDNA library is expressed in E. coli, and the antibody V region is produced on the cell surfaces. If selection is made on a plate coated with the desired antigen based on antigen-binding activity, a gene encoding the desired antibody can be obtained.
[0084] Antibodies possessing stronger binding activity to antigens may be obtained by the chain shuffling method with application of the phage library method. (Akamatsu, Y. and Tsurushita, N. Medical Immunology 27, 273-286 (1994)). Specifically, one member of the V region of an antibody gene that has been separated (such as VH) is fixed; and a new library is constructed from the mixture of the one member and the other member prepared from B cells (such as VL). Clones that bind to the antigen more strongly than do the others may be separated from the library.
[0085] It is also possible to obtain antibodies possessing stronger binding activity to antigens by introducing artificial mutations into the amino acid sequences of the antibodies. (Akamatsu, Y. and Tsurushita, N. Medical Immunology 27, 273-286 (1994)). More specifically, the mutation is introduced into a gene encoding the cloned antibody V region, and this gene is expressed by the phage library method described above. Thereby, it becomes possible to obtain a gene encoding the antibody that possesses stronger binding activity to antigen.
[0086] These fragments of antibody can be produced by hosts after their genes are acquired and expressed in the same manner as described previously above. As used in the present specification, "antibody" encompasses these fragments of antibody.
[0087] Modified forms of antibody can employ antibodies that are bound to various molecules such as polyethylene glycol (PEG). As used in the present specification, the "antibody" encompasses these modified forms of antibody. Acquisition of these modified forms of antibody can be done by subjecting the obtained antibodies to chemical modification.
[0088] These methods have already been established in the art. The antibodies that are expressed and produced as described above can be separated from the host cells intracellularly or extracellularly, and can be purified to homogeneity according to methods that are commonly used. Concentration measurement can be carried out by the measurement of absorbance, ELISA or the like.
[0089] For the CXCR4 inhibitor used in this invention, there is mentioned a protein having an SDF-1- or CXCR4-like structure (structure-resembling protein). This substance is one that possesses binding activity to SDF-1 or CXCR4 and that does not transmit its biological activity. That is, it blocks signaling by SDF-1, because it binds to CXCR4 in a competitive manner with SDF-1, but does not transmit the biological activity of SDF-1.
[0090] SDF-1 structure-resembling proteins may be prepared by introducing mutations into the amino acid sequence of SDF-1 through substitution of amino acid residues thereof. For SDF-1 on which the SDF-1 structure-resembling proteins are based, its source does not matter; however, it is preferably human SDF-1 in consideration of its antigenicity or the like. Specifically, the amino acid sequence of SDF-1 is used to predict its secondary structure by using a molecular modeling program known in the art such as WHATIF (Vriend et al., J. Mol. Graphics. 8, 52-56 (1990)); and further the influence on the whole, of the amino acid residue to be substituted is evaluated to carry out the preparation.
[0091] After a suitable amino acid residue to be substituted has been determined, a vector containing the base sequence that encodes human SDF-1 gene is used as a template, and the introduction of mutation is carried out by the PCR method (polymerase chain reaction), which is commonly done, so that the amino acid may be substituted. This allows a gene encoding the SDF-1 structure-resembling protein to be obtained. This gene is incorporated into a suitable expression vector as appropriate, and the SDF-1 structure-resembling protein can be obtained according to the methods for the expression, production and purification of the recombinant antibodies as described previously. SDF-1 of which the N-terminus has been deleted is known as an SDF-1 structure-resembling protein (EMBO J. 16, 6996-7007 (1997)).
[0092] The SDF-1 partial peptide or the CXCR4 partial peptide that is used in this invention is a substance that possesses binding activity to CXCR4 or to SDF-1 and that does not transmit the biological activity of SDF-1. That is, the SDF-1 partial peptide or the CXCR4 partial peptide specifically inhibits SDF-1 from binding to CXCR4, because each binds to CXCR4 or to SDF-1 and traps either of them.
[0093] Consequently, they block signaling by SDF-1, because they do not transmit the biological activity of SDF-1.
[0094] The SDF-1 partial peptide or the CXCR4 partial peptide is a peptide comprising a part or the whole of the amino acid sequence of the region, which is responsible for the binding between SDF-1 and CXCR4, in the amino acid sequence of SDF-1 or of CXCR4. Such a peptide usually comprises 10-80 amino acid residues, preferably 20-50 amino acid residues, and more preferably 20-40 amino acid residues.
[0095] The SDF-1 partial peptide or the CXCR4 partial peptide can be prepared by identifying in the amino acid sequence of SDF-1 or of CXCR4, the region that is responsible for the binding between SDF-1 and CXCR4 and by preparing a part or the whole of the amino acid sequence of said region according to a method that is commonly known, such as a technique in genetic engineering or peptide synthesis.
[0096] To prepare the SDF-1 partial peptide or the CXCR4 partial peptide by the technique in genetic engineering, a DNA sequence encoding the desired peptide is incorporated into an expression vector, and the methods for expression, production and purification of the recombinant antibodies as described previously are followed with any possible modifications, thus enabling the preparation.
[0097] To prepare the SDF-1 partial peptide or the CXCR4 partial peptide by peptide synthesis, methods that are commonly used in the peptide synthesis, such as the solid phase synthesis or the liquid phase synthesis, can be employed. Concretely, the method as described in "Development of Drugs, Peptide Synthesis Vol. 14, Ed. by Nariaki Yajima, Hirokawa Publisher (1991) may be followed with any possible modifications. For the solid phase synthesis, the following method is, for example, employed: an amino acid corresponding to the C-terminus of the peptide to be synthesized is allowed to bind to a support insoluble in an organic solvent; and the reaction wherein an amino acid protected with suitable protecting groups at its α-amino group and side-chain functional group is condensed to the foregoing amino acid one amino acid at a time in the direction of from the C-terminus to the N-terminus and the reaction for detaching the protecting group for the α-amino group of the amino acid or peptide bound to the resin are alternately repeated to elongate the peptide chain. The solid phase peptide synthesis is largely classified into the Boc method and the Fmoc method, depending on the kind of protecting groups to be used.
[0098] After the objective peptide is synthesized in this manner, deprotection reaction is done and the peptide is cleaved from the Support for the peptide chain. In the cleavage reaction from the peptide chain, the Boc method can usually employ hydrogen fluoride or trifluoromethanesulfonic acid and the Fmoc method can usually employ TFA. In the Boc method, the protected peptide resin mentioned above is, for example, treated in hydrogen fluoride in the presence of anisole. Subsequently, deprotection of the protecting group and cleavage from the support recovers the peptide.
[0099] The product is lyophilized to yield a crude peptide. In the Fmoc method, deprotection reaction and cleavage reaction from the support for the peptide chain can also be carried out in TFA using manipulations similar to those mentioned above.
[0100] The resulting crude peptide can be separated and purified by being applied on HPLC. The elution may then be carried out under optimum conditions with a solvent of the water-acetonitrile system that is usually used to purify proteins. The fractions corresponding to the peaks of the obtained profile of chromatography are fractionally separated and lyophilized. The peptide fractions thus purified are identified by molecular weight analysis through mass spectrometry, amino acid composition analysis, amino acid sequencing, or the like.
[0101] For the SDF-1 partial peptide or the CXCR4 partial peptide that is used in this invention, its sequence does not matter insofar as each binds to CXCR4 or to SDF-1 and possess no signaling activity. The amino acid sequences that are already known can be used both for the SDF-1 partial peptide and the CXCR4 partial peptide. For example, when the ligand is SDF-1, the amino acid sequences set forth in SEQ ID NO: (human) and SEQ ID NO: 7 (mouse) are usable.
[0102] There is no particular limitation to the substance that potentiates the action due to CXCR4 that can be used in this invention. For the substance that potentiates SDF-1, there are mentioned SDF-1 itself, an agonist of SDF-1, and a potentiator of SDF-1 expression. Furthermore, for the substance that potentiates a CXCR4 receptor, there are mentioned CXCR4 itself, an agonist of CXCR4, and a potentiator of CXCR4 expression.
[0103] As explained above, use of the vascularization inhibitor according to this invention comprising the CXCR4 inhibitor as the effective ingredient allows for the inhibition of vascularization; therefore, it will exert an antitumor effect (inhibition of neovascularization) on solid cancer in addition to antitumor effects on angiosarcoma (cancer of blood vessels themselves) and Kaposi's sarcoma. It will also exert therapeutic effects against diseases pathologically caused by neovascularization, such as chronic articular rheumatism, psoriasis, and diabetic retinopathy.
[0104] If the therapeutic agent for a disease pathologically caused by neovascularization which comprises the substance that potentiates the action of CXCR4 is used, it will be possible to promote neovascularization. The use will exert therapeutic effects on myocardial infarction and diseases involving neovascularization after surgery, such as wound healing, repairing and remodeling of bones, repairing of cartilage, growth of hair, myocardial infarction, brain infarction, and brain trauma.
[0105] For the neovascularization inhibition and promotion test methods that can be used in this invention, a neovascularization assay may be employed. There is no particular limitation to this assay, and a method that is commonly known can preferably be used. ("Research Manual for the Invasion and Metastasis of Cancers" the Cancer Metastasis Study Group Ed., Kinhodo, 159-182, (1994)). Specifically, among others, there are mentioned (I) the method for measuring cleavage of the spaces between vascular endothelial cells (i.e., effect on the vascular endothelial cells) which is based on the finding that the vascular endothelial cells rupture when tumor cells invade out of the blood vessels (from the permeability of FITC-dextran); (II) the cornea method known as an in vivo measurement method for identifying a candidate factor that exerts the function of a neovascularization-inducing factor in vivo; (IV) CAM method (chickembryochorioallantoic membrane); (V) the dorsal subcutanea method for measuring the quantity of induced blood vessel by the naked eyes; and (VI) the method for determining the formation of lumen by vascular endothelial cells.
[0106] To confirm the antitumor effect, there may be mentioned in vivo experiments using a transplant model or transplant metastasis model and in vitro experiments with cancer cells. Specifically, the method described in "Research Manual for the Invasion and Metastasis of Cancers" the Cancer Metastasis Study Group Ed., Kinhodo, 7-158 (1994) may be used.
[0107] The vascularization inhibitor, anti-sold cancer agent, therapeutic agent, tissue-repairing agent, or the like according to this invention may be systemically (by oral route) or locally administered. For example, intravenous injection (such as intravenous drip infusion), intramuscular injection, intraperitoneal injection, or subcutaneous injection may be selected: an appropriate method of administration may be selected depending on the age or the severity of the subject.
[0108] The effective unit dose is chosen within the range of from 0.09 mg to 100 mg per kg of body weight. Alternatively, a dose of 1-1000 mg, preferably a dose of 5-50 mg for the subject may be chosen.
[0109] The vascularization inhibitor, anti-sold cancer agent, therapeutic agent, tissue-repairing agent, or the like according to this invention may together contain pharmaceutically acceptable carriers or additives, depending on the route of administration. Examples of such carriers and additives include water, organic solvents that are pharmaceutically acceptable, collagen, poly(vinyl alcohol), poly(vinylpyrrolidone), carboxyvinylpolymer, sodium carboxymethyl cellulose, sodium polyacrylate, sodium arginate, water-soluble dextran, sodium carboxymethyl starch, pectin, methylcellulose, xanthan gum, gum arabic, casein, gelatin, agar, diglycerin, propylene glycol, poly(ethylene glycol), vaseline, paraffin, stearyl alcohol, stearic acid, human serum albumin (HSA), mannitol, sorbitol, lactose, surfactants that are accepted as drug additives, etc.
[0110] The additives to be used may appropriately (or in combination) be selected among those mentioned above depending on the dosage form, but are not limited thereto.
[0111] This invention will be hereinbelow illustrated in greater details by way of examples; however, the invention is in no way limited by those examples.
EXAMPLES
Creation and Analysis of Mice Completely Lacking CXCR4
[0112] Genome DNA containing the CXCR4 locus was isolated from a murine cell line 129DNA library (STRATAGENE).
[0113] A 1.1-kb genomic fragment containing the 5'-coding region of exon 2 was replaced by the neomycin resistant gene, and the herpes simplex thymidine kinase gene was linked to the 5'-terminus.
[0114] The targeting vector was introduced into the cells on day 14.1 of embryogenesis (referred to as "E14.1" hereafter) by electroporation, and a homologous recombination was selected by the use of G418 and ganciclovir and identified by PCR.
[0115] The structure of the mutant locus and the presence of a single insert in the ES cell colony were confirmed by Southern hybridization. According to the method as described in Nagasawa, T. et al., Nature 382, 685-688 (1996), the mutant ES cell colony was used to create a mutant mouse by the injection of undifferentiated embryonic cells.
[0116] For Southern hybridization, tail DNA was digested with EcoRI, transferred to a nylon membrane, and hybridized to 550-bp probe A with the 5'-homology region.
[0117] RT-PCR was performed for 40 cycles, using 3 μg of total RNA isolated from the fetal liver of E18.5 embryos as a starting material according to the standard method. A 630-bp PCR product was amplified using CXCR4-specific primers, i.e., forward primer (SEQ ID NO: 11) and reverse primer (SEQ ID NO: 12). Histological analysis and flow cytometry analysis were carried out substantially according to the method as described in Nagasawa, T. et al., Nature 382, 685-688 (1996).
[0118] Immunohistostaining substantially followed the method of Adachi, S., Yoshida, H., Kataoka, H. Nishikawa, S.- I. Int. Immunol. 9, 507-514. The sections of embryos and organs were fixed with 4% paraformaldehyde, dehydrated with methanol, decolorized with 30% hydrogen peroxide in methanol, and hydrated again.
[0119] After incubating in PBSMT (PBS containing 1% skim milk powder and 0.3% v/v TritonX-100), the sample was incubated with diluted anti-PECAM antibody (1:250) (PharMingen) in PBSMT at 4° C. overnight. Then, the sample was washed with PBSMT, and incubated with diluted horseradish peroxidase labeled anti-rat Ig antibody (1:500) (Biosource) in PBST at 4° C. overnight.
[0120] Subsequently, the sample was thoroughly washed. The embryo was incubated in PBS containing 250 μg/ml diaminobenzidine (Dojin Chemicals) and 0.08% NiCl2 for 30 min. Hydrogen peroxide was added to the sample to provide a final concentration of 0.01%, after which peroxidase staining was carried out. The reaction was quenched after about 30 min.
[0121] According to the method as described in Nagasawa, T. et al., Nature 382, 685-688 (1996), a fragment of murine CXCR4 or PBSF/SDF-1 cDNA was used as a probe to carry out antisense transcription.
[0122] Mice lacking CXCR4 were created to determine the physiological function of CXCR4. Specifically, a targeting vector was constructed, such that most of exon 2 within the CXCR4 gene which contained all transmembrane-spanning regions critical to the receptor function had been deleted and the deleted portion would be replaced by the neomycin resistant gene (neo). This would result in that after homologous recombination, the complete CXCR4 gene would be substantially deleted.
[0123] FIG. 1 is a graph showing a targeting strategy for the CXCR4 gene (denoted as top, middle, and bottom). There are shown the CXCR4 wild-type allele at the top, a targeting vector in the middle, and a predicted mutant allele at the bottom. The coding regions of the genes are indicated by black boxes. Empty boxes indicate the 5'- and 3'-untranslated regions. Dotted lines indicate homologous fragments used in the targeting vector. Probe A is an external probe for Southern hybridization. Here, restriction sites are E (EcoRI), Sh (SphI), and X (XhoI), respectively.
[0124] FIG. 2A is a photograph showing the Southern blot analysis of tail DNAs from wild-type (+/+) and heterozygous mutant (+/-) mice. The EcoRI-EcoRI fragments from the 11.8-kb wild-type and the 8.2-kb targeted allele which were identified by probe A are shown in the figure. FIG. 2B is a photograph showing the RT-PCR amplification analysis of CXCR4 expression. Total RNAs were prepared from E18.5 wild-type and homozygous mutant embryos, and amplified with CXCR4-specific primers. The RT-PCR amplification employed G3PDH mRNA, which was universally expressed, as a control for the presence of any amplifiable RNA.
[0125] Mice with a CXCR4.sup.+/- heterozygous mutation were created. The mice were healthy and fertile. CXCR4.sup.-/- homozygous mutant embryos were present at the expected ratios until E15.5 of embryogenesis. However, about half of the CXCR4.sup.-/- embryos were dead at E18.5 and CXCR4.sup.-/- neonates died within an hour similarly to mice lacking PBSF/SDF-1 as previously reported (Nagasawa, T. et al., Nature 382, 685-688 (1996)).
[0126] To elucidate the functions of CXCR4 during embryogenesis, expression of CXCR4 in developing embryos was examined by in situ hybridization. High levels of CXCR4 transcripts were detected in the endothelium of developing blood vessels during embryogenesis.
[0127] Based on this finding, the effect of CXCR4 gene deficiency on vascularization was investigated. On visual inspection, vitellin and umbilical vessels were normal. Histological examination of E18.5 CXCR4.sup.-/- embryos demonstrated the presence of the major blood vessels, including aorta, vena cava, carotid artery, jugular vein, coeliac artery and superior mesenteric artery and superior mesenteric vein. To visualize the vascular system of organs, whole-mount preparations of wild-type and mutant embryos were then immunostained with an anti-PECAM-1 antibody. It is known that PECAM-1 is specifically and stably expressed in all endothelial cells during embryonic period. (Vecchi, A. et al. Eur. J. cell Biol. 63, 247-255 (1994); Baldwin, H. S. et al., Development 120, 2539-2558 (1994)).
[0128] Consequently, in the gastrointestinal tract, including stomach, intestine and mesentery, a highly branched homogeneous vascular network was observed in both the wild-type and the mutant by E11.5. The formation of large and small vessels through remodeling in the mesentery connecting to mid-gut loop was observed at around E12.5. As FIG. 3 shows, many large branches of superior mesenteric artery and vein supplying nutrient to the intestine were formed in wild-type embryos at E13.5. On the other hand, these large branches were not present in the mesenteries of CXCR4.sup.-/- embryos at E13.5; instead, only small vessels were formed. Histological analysis by microscopy revealed superior mesenteric arteries and veins in the mesenteries of wild-type embryos at E13.5. This showed that the branched vessels were paired between the artery and the vein (FIG. 4). In contrast, most of the vessels in CXCR4.sup.-/- embryos were not paired, but were single, as can be seen in FIG. 8. However, the superior mesenteric arteries and veins within the mesenteries of the wild-type embryos were normal. In E17.5 wild-type embryos, the large mesenteric vessels split into many branches and reach the intestine (FIGS. 5 and 6). However, in the CXCR4.sup.-/- embryos such vessels corresponding to the large mesenteric vessels were substantially absent (FIGS. 6 and 9). Also, a few large vessels with aberrant branching were observed in the mutant mesenteries (FIGS. 6 and 9). In most of E16.5 mutant embryos multiple haemorrhagic lesions were observed in their small intestines because of such defective vascular system. This pathogenesis is believed to be the result of aberration of the circulatory system governing the intestine (FIG. 11).
[0129] The above-mentioned results have demonstrated that CXCR4 is essential for the normal vascularization of the small intestine: the mechanism is believed to be due to that CXCR4 is involved in the branching and/or remodeling of the mesenteric vessels.
[0130] In the stomach, large vessels branching out from mesenchymal vessels along the lesser curvature were formed and distributed to the entire ventral and dorsal surfaces in the wild-type embryos by E13.5 (FIGS. 12A and 12C). Histological analysis revealed that these vessels were paired between the artery and the vein in the E15.5 wild-type mice as the inset in FIG. 12 C shows. However, the corresponding vessels were not found in the mutant embryos (FIGS. 12B and 12D). Formation of the network of small vessels that surrounds the stomach seemed to be normal in the mutant embryos (FIG. 12D).
[0131] Histological analysis of the stomachs and intestines of E18.5 mutant embryos detected no obvious abnormalities in organogenesis. For example, the smooth muscle layers (both the outer and inner layers) of the gastrointestinal tract of mutant mice seemed to be normal in the longitudinal and vertical directions.
[0132] FIGS. 3-6 are photographs showing defects of the gastrointestinal vessels in the CXCR4.sup.-/- embryos, and also photographs showing immunohistostaining of mesenteries and intestines of the wild-types with the anti-PECAN antibody. FIG. 3 shows the mesentry and mid-gut loop regions at E13.5. FIG. 5 shows a jejunum at E17.5. FIG. 6 shows a jejunum at E17.5. FIG. 7 shows a cross-section of the stained mesentry at E13.5. The arrows in FIGS. 3, 5, and 6 indicate large branches of superior mesenteric artery or superior mesenteric vein supplying nutrient to the small intestine, in wild-type mesentries.
[0133] Similarly, FIGS. 7-11 are photographs showing defects of the gastrointestinal vessels in the CXCR4.sup.-/- embryos, and also photographs showing immunohistostaining of mesenteries and intestines of the mutants with the anti-PECAM antibody. FIG. 7 shows the mesentry and mid-gut loop regions at E13.5. FIG. 9 shows a jejunum at E17.5. FIG. 10 shows a jejunum at E17.5. FIG. 7 shows a cross-section of the stained mesentry at E13.5. FIG. 11 shows the haemorrhagic lesions of unstained intestine of a mutant mouse at E16.5. The arrows in FIGS. 9 and 10 indicate large vessels displaying aberrant running and/or branching in the mutant mice.
[0134] FIGS. 12A-12D are photographs showing the results of immunohistostaining stomachs with the anti-PECAM antibody. FIGS. 12A and 12B show E13.5; FIGS. 12C and 12D show E15.5; FIGS. 12A and 12C show the wild types; and FIGS. 12B and 12D show the mutants. The inset in the photograph of FIG. 12C shows a haematoxylin-and-eosin-stained section of large vessels in the wall of stained stomach at E15.5. The arrows in FIGS. 12A and 12C indicate large vessels only observed in the mutant. "du" represents duodenum; "p" represents the proximal part of mid-gut loop; "dm" represents the distal part of mid-gut loop; "a" represents artery; and "v" represents vein.
[0135] These findings indicate that the abnormalities in vascularization in the CXCR4.sup.-/- mice are not a secondary outcome of those in the gastrointestinal tracts themselves. Abnormalities similar to those in vascularization were also observed in the mice lacking PBSF/SDF-1.
[0136] In situ hybridization analysis indicated that CXCR4 transcripts were expressed in the endothelial cells of blood vessels in the mesentery and in the wall of the intestine and stomach in the E12.5 wild-type embryos (FIGS. 13B and 13E). Particularly, strong expression was observed in the endothelial cells of branches arising from the superior mesenteric arteries (FIGS. 13B and 13E). In contrast, PBSF/SDF-1 was expressed at high levels in mesenchymal cells surrounding the endothelial cells of the mesentry, but not in the endothelial cells or the wall of intestine and stomach (FIG. 13C).
[0137] FIGS. 13A-13F are photographs showing an analysis of CXCR4 and PBSF/SDF-1 expression in the gastrointestinal tract through in situ hybridization. Serial sections of the wild-type mesentery connecting to the mid-gut loop were used; one piece was stained with haematoxylin and eosin (FIGS. 13A and 13D); another piece was hybridized to the CXCR4-specific probe (FIGS. 13B and 13E); an additional piece was hybridized to the PBSF/SDF-1-specific probe (FIG. 13E). FIGS. 13D and 13E are enlargements of branched vessels arising from the superior mesenteric artery shown in FIGS. 13A and 13B, indicating strong expression of CXCR4 in endothelial cells of the vessel. The arrows and arrowheads in FIGS. 13B and 13E indicate the endothelial cells of mesenteric blood vessels with observed CXCR4 expression. PBSF/SDF-1 is expressed in the mesenchymal cells surrounding endothelial cells in the mesentery (FIG. 13E). FIG. 13F is a cross-section of bone marrow of the E18.5 wild-type embryo, showing CXCR4 expression in haemotopoietic cells but not in spindle-shaped stromal cells. "m" represents mesentry; "i" represents intestine; "a" represents superior mesenteric artery; and "v" represents superior mesenteric vein.
[0138] The expression patterns obtained indicate that PBSF/SDF-1 produced by mesenchymal cells acts on CXCR4 on endothelial cells; and this strongly suggests the presence of the paracrine signal by a cytokine that plays a extremely important role in the mesenteric mesenchyme. Thus, the phenotype that lacks large vessels in the stomachs of CXCR4.sup.-/- and PBSF/SDF-1.sup.-/- may result from abnormalities in the vascular branching and/or remodelling in the mesenchyme along the lesser curvature of the stomach.
[0139] To examine the vascular systems of other organs, a whole-mount yolk sac, brain and heart were stained with an anti-PECAM-1 monoclonal antibody. There was no obvious difference between CXCR4.sup.-/- and PBSF/SDF-1.sup.-/- and the wild-type in the formation of large and small vessels in the whole-mount yolk sac (E12.5, E14.5), the head region (E11.5) and the heart (E12.5-E14.5).
[0140] In summary, the aforementioned experimental results have shown that CXCR4 and PBSF/SDF-1 are essential for the formation of a mature vascular system which is supplied to the gastrointestinal tract by acting on the endothelial cells of blood vessels and regulating vascular branching and/or remodelling.
[0141] A flow cytometric analysis revealed that the number of B-cell progenitors in fetal livers of CXCR4.sup.-/- mice was severely reduced. Histological analysis did not detect myelocytes and their progenitors in the medullary cavity. In addition, defects of the membranous portion of the cardiac ventricular septum were found in the hearts of E18.5 mutant mice. These abnormalities are very similar to the phenotype found in the mice lacking PBSF/SDF-1, supporting the thinking that CXCR4 is a primary physiological receptor for PBSF/SDF-1.
[0142] In the E18.5 wild-type embryonic bone marrow as determined by in situ hybridization, CXCR4 transcripts were expressed in hematopoietic cells, while they were not expressed in spindle-shaped stroma cells where the expression of PBSF/SDF-1 transcripts had been observed (FIG. 3D). The results of these expression patterns mean the presence of paracrine signaling in bone marrow.
[0143] Analyses of receptor tyrosine kinases (RTKs), such as Flk-1 and Tie-2, and their ligands, such as VEGF, angiopoietin-1 and PDGF-B that have hitherto been reported, using mutant mice have indicated that they play an important role in development of the vascular system, and that many of them are required for the very early stage of vascularization in genesis as well as for vascularization in all parts of body, including the york sac and the extra-embryonic vasculature. (Shalaby, F. et al., Nature 376, 62-66 (1995); Fong, G.- H., Rossant, J., Gertsenstein, M. & Breitman, M. L., Nature 376, 66-70 (1995); Dumount, D. H. et al., Genes Dev 8, 1897-1909 (1994); Sato, T. N. et al., Nature, 376, 70-74 (1995); Carmeliet, P. et al., Nature, 880, 435-439 (1996); Ferrara, N. et al., Nature, 380, 439-442 (1996); and Suri, C. et al., Cell 87, 1171-1180 (1996)).
[0144] In contrast, the functions of CXCR4 and PBSF/SDF-1 operate at later stages of genesis and are organ-specific. Tie-2 and its ligand, angiopoietin-1, are thought to be necessary for the branching and/or remodelling in the early vascular system. (Sato, T. N. et al., Nature 376, 70-74 (1995); Suri, C. et al., Cell 87, 1171-1180 (1996)). Their roles in the formation of the mature vascular system in the gastrointestinal tract are not clear. Obvious abnormalities found in the York sac vascular system of Tie-2 or angiopoietin-1.sup.-/- mice were not noted in the CXCR4 or PBSF/SDF-1.sup.-/- mice. CXC chemokines, such as PF4 (Maione, T. E. et al., Science 247, 77-79 (1990)), IL-8 (Koch, A. E. et al., Science 258, 1798-1801 (1992)), IP-10 (Luster, A. D. et al., J. Exp. Med. 182, 219-231 (1995)), and Groβ (Cao, Y. H., et al., J. J. Exp. Med. 182, 2069-2077 (1995)), have been reported to be neovascularization regulators. However, expression of their receptors in endothelial cells and their physiological roles have not yet been elucidated. Although coagulation factor V (CuI, J., et al., Nature, 384, 66-68 (1996)) and tissue factor (Carmeliet, P. et al., Nature 883, 75-78 (1996)) have been shown to be essential for the york sac vascular system, it is not clear as to what kind of acceptor mediates these factors.
[0145] With the background mentioned above, it can be said that this invention have demonstrated the presence of a novel signaling system--chemokines and a seven-transmembrane spanning, G-protein-coupled receptors--essential for vascularization.
[0146] It has recently been shown that mice lacking the α-subunit of the heterotrimeric GTP-binding protein Gα13 have abnormalities, such as no formation of a york-sac vascular system and enlargement of embryonic small vessels. Although the phenotypes are different from those of mice lacking CXCR4, it is necessary to examine the possibility of CXCR4's coupling to Gα13.
[0147] It is known that CXCR4 and CCR5 are essential co-receptors when T-cell line tropic and macrophage-tropic HIV-1 strains infect host cells, respectively. (Feng, Y., et al., Science 272, 872-877 (1996); Fauci, A. S., Nature 884, 529-584 (1996)). Between the two, the people homozygous for CCR5 deletion have been discovered; and they are resistant to HIV-1 infection and have no obvious health problems. (Liu, R et al., Cell 86, 367-377 (1996); Samson, M. et al., Nature 382, 722-725 (1996); and Dean, M. et al., Science 273, 1856-1861).
[0148] Concerning the other CXCR4, it has, however, been strongly suggested that homologous CXCR4 deletion is unlikely to occur in humans since mice lacking CXCR4 are apt to die in utero. There still remains the possibility that additional genetic factors or homologous viable mutations may exist in long-term survivors who are resistant to T cell-line-tropic HIV-1.
INDUSTRIAL APPLICABILITY
[0149] Based on the finding of this invention that vascularization is suppressed in CXCR4 knockout mice, a vascularization inhibitor comprising as the effective ingredient, a substance that inhibits the action of CXCR4 can be prepared: since vascularization is essential for the maintenance and enlargement of cancerous tissues, the finding is utilized in the preparation of an anti-solid cancer agent and a therapeutic agent for a disease pathologically caused by neovascularization, which comprises as the effective ingredient a substance that inhibits the action of CXCR4, as well as in the preparation of a tissue-repairing agent comprising as the effective ingredient, a substance that potentiates the action of CXCR4.
Sequence CWU
1
1
121352PRTHomo sapiens 1Met Glu Gly Ile Ser Ile Tyr Thr Ser Asp Asn Tyr Thr
Glu Glu Met 1 5 10 15
Gly Ser Gly Asp Tyr Asp Ser Met Lys Glu Pro Cys Phe Arg Glu Glu
20 25 30 Asn Ala Asn Phe
Asn Lys Ile Phe Leu Pro Thr Ile Tyr Ser Ile Ile 35
40 45 Phe Leu Thr Gly Ile Val Gly Asn Gly
Leu Val Ile Leu Val Met Gly 50 55
60 Tyr Gln Lys Lys Leu Arg Ser Met Thr Asp Lys Tyr Arg
Leu His Leu 65 70 75
80 Ser Val Ala Asp Leu Leu Phe Val Ile Thr Leu Pro Phe Trp Ala Val
85 90 95 Asp Ala Val Ala
Asn Trp Tyr Phe Gly Asn Phe Leu Cys Lys Ala Val 100
105 110 His Val Ile Tyr Thr Val Asn Leu Tyr
Ser Ser Val Leu Ile Leu Ala 115 120
125 Phe Ile Ser Leu Asp Arg Tyr Leu Ala Ile Val His Ala Thr
Asn Ser 130 135 140
Gln Arg Pro Arg Lys Leu Leu Ala Glu Lys Val Val Tyr Val Gly Val 145
150 155 160 Trp Ile Pro Ala Leu
Leu Leu Thr Ile Pro Asp Phe Ile Phe Ala Asn 165
170 175 Val Ser Glu Ala Asp Asp Arg Tyr Ile Cys
Asp Arg Phe Tyr Pro Asn 180 185
190 Asp Leu Trp Val Val Val Phe Gln Phe Gln His Ile Met Val Gly
Leu 195 200 205 Ile
Leu Pro Gly Ile Val Ile Leu Ser Cys Tyr Cys Ile Ile Ile Ser 210
215 220 Lys Leu Ser His Ser Lys
Gly His Gln Lys Arg Lys Ala Leu Lys Thr 225 230
235 240 Thr Val Ile Leu Ile Leu Ala Phe Phe Ala Cys
Trp Leu Pro Tyr Tyr 245 250
255 Ile Gly Ile Ser Ile Asp Ser Phe Ile Leu Leu Glu Ile Ile Lys Gln
260 265 270 Gly Cys
Glu Phe Glu Asn Thr Val His Lys Trp Ile Ser Ile Thr Glu 275
280 285 Ala Leu Ala Phe Phe His Cys
Cys Leu Asn Pro Ile Leu Tyr Ala Phe 290 295
300 Leu Gly Ala Lys Phe Lys Thr Ser Ala Gln His Ala
Leu Thr Ser Val 305 310 315
320 Ser Arg Gly Ser Ser Leu Lys Ile Leu Ser Lys Gly Lys Arg Gly Gly
325 330 335 His Ser Ser
Val Ser Thr Glu Ser Glu Ser Ser Ser Phe His Ser Ser 340
345 350 21588DNAHomo
sapiensCDS(1)..(1059) 2atg gag ggg atc agt ata tac act tca gat aac tac
acc gag gaa atg 48Met Glu Gly Ile Ser Ile Tyr Thr Ser Asp Asn Tyr
Thr Glu Glu Met 1 5 10
15 ggc tca ggg gac tat gac tcc atg aag gaa ccc tgt
ttc cgt gaa gaa 96Gly Ser Gly Asp Tyr Asp Ser Met Lys Glu Pro Cys
Phe Arg Glu Glu 20 25
30 aat gct aat ttc aat aaa atc ttc ctg ccc acc atc
tac tcc atc atc 144Asn Ala Asn Phe Asn Lys Ile Phe Leu Pro Thr Ile
Tyr Ser Ile Ile 35 40
45 ttc tta act ggc att gtg ggc aat gga ttg gtc atc
ctg gtc atg ggt 192Phe Leu Thr Gly Ile Val Gly Asn Gly Leu Val Ile
Leu Val Met Gly 50 55 60
tac cag aag aaa ctg aga agc atg acg gac aag tac
agg ctg cac ctg 240Tyr Gln Lys Lys Leu Arg Ser Met Thr Asp Lys Tyr
Arg Leu His Leu 65 70 75
80 tca gtg gcc gac ctc ctc ttt gtc atc acg ctt ccc
ttc tgg gca gtt 288Ser Val Ala Asp Leu Leu Phe Val Ile Thr Leu Pro
Phe Trp Ala Val 85 90
95 gat gcc gtg gca aac tgg tac ttt ggg aac ttc cta
tgc aag gca gtc 336Asp Ala Val Ala Asn Trp Tyr Phe Gly Asn Phe Leu
Cys Lys Ala Val 100 105
110 cat gtc atc tac aca gtc aac ctc tac agc agt gtc
ctc atc ctg gcc 384His Val Ile Tyr Thr Val Asn Leu Tyr Ser Ser Val
Leu Ile Leu Ala 115 120
125 ttc atc agt ctg gac cgc tac ctg gcc atc gtc cac
gcc acc aac agt 432Phe Ile Ser Leu Asp Arg Tyr Leu Ala Ile Val His
Ala Thr Asn Ser 130 135 140
cag agg cca agg aag ctg ttg gct gaa aag gtg gtc
tat gtt ggc gtc 480Gln Arg Pro Arg Lys Leu Leu Ala Glu Lys Val Val
Tyr Val Gly Val 145 150 155
160 tgg atc cct gcc ctc ctg ctg act att ccc gac ttc
atc ttt gcc aac 528Trp Ile Pro Ala Leu Leu Leu Thr Ile Pro Asp Phe
Ile Phe Ala Asn 165 170
175 gtc agt gag gca gat gac aga tat atc tgt gac cgc
ttc tac ccc aat 576Val Ser Glu Ala Asp Asp Arg Tyr Ile Cys Asp Arg
Phe Tyr Pro Asn 180 185
190 gac ttg tgg gtg gtt gtg ttc cag ttt cag cac atc
atg gtt ggc ctt 624Asp Leu Trp Val Val Val Phe Gln Phe Gln His Ile
Met Val Gly Leu 195 200
205 atc ctg cct ggt att gtc atc ctg tcc tgc tat tgc
att atc atc tcc 672Ile Leu Pro Gly Ile Val Ile Leu Ser Cys Tyr Cys
Ile Ile Ile Ser 210 215 220
aag ctg tca cac tcc aag ggc cac cag aag cgc aag
gcc ctc aag acc 720Lys Leu Ser His Ser Lys Gly His Gln Lys Arg Lys
Ala Leu Lys Thr 225 230 235
240 aca gtc atc ctc atc ctg gct ttc ttc gcc tgt tgg
ctg cct tac tac 768Thr Val Ile Leu Ile Leu Ala Phe Phe Ala Cys Trp
Leu Pro Tyr Tyr 245 250
255 att ggg atc agc atc gac tcc ttc atc ctc ctg gaa
atc atc aag caa 816Ile Gly Ile Ser Ile Asp Ser Phe Ile Leu Leu Glu
Ile Ile Lys Gln 260 265
270 ggg tgt gag ttt gag aac act gtg cac aag tgg att
tcc atc acc gag 864Gly Cys Glu Phe Glu Asn Thr Val His Lys Trp Ile
Ser Ile Thr Glu 275 280
285 gcc cta gct ttc ttc cac tgt tgt ctg aac ccc atc
ctc tat gct ttc 912Ala Leu Ala Phe Phe His Cys Cys Leu Asn Pro Ile
Leu Tyr Ala Phe 290 295 300
ctt gga gcc aaa ttt aaa acc tct gcc cag cac gca
ctc acc tct gtg 960Leu Gly Ala Lys Phe Lys Thr Ser Ala Gln His Ala
Leu Thr Ser Val 305 310 315
320 agc aga ggg tcc agc ctc aag atc ctc tcc aaa gga
aag cga ggt gga 1008Ser Arg Gly Ser Ser Leu Lys Ile Leu Ser Lys Gly
Lys Arg Gly Gly 325 330
335 cat tca tct gtt tcc act gag tct gag tct tca agt
ttt cac tcc agc 1056His Ser Ser Val Ser Thr Glu Ser Glu Ser Ser Ser
Phe His Ser Ser 340 345
350 taa cacagatgta aaagactttt ttttatacga taaataactt
ttttttaagt 1109tacacatttt tcagatataa aagactgacc aatattgtac
agtttttatt gcttgttgga 1169tttttgtctt gtgtttcttt agtttttgtg aagtttaatt
gacttattta tataaatttt 1229ttttgtttca tattgatgtg tgtctaggca ggacctgtgg
ccaagttctt agttgctgta 1289tgtctcgtgg taggactgta gaaaagggaa ctgaacattc
cagagcgtgt agttaatcac 1349gtaaagctag aaatgatccc cagctgttta tgcatagata
atctctccat tcccgtggaa 1409cgtttttcct gttcttaaga cgtgattttg ctgtagaaga
tggcacttat aaccaaagcc 1469caaagtggta tagaaatgct ggtttttcag ttttcaggag
tgggttgatt tcagcaccta 1529cagtgtacag tcttgtatta agttgttaat aaaagtacat
gttaaactta aaaaaaaaa 15883359PRTMus musculus 3Met Glu Pro Ile Ser Val
Ser Ile Tyr Thr Ser Asp Asn Tyr Ser Glu 1 5
10 15 Glu Val Gly Ser Gly Asp Tyr Asp Ser Asn Lys
Glu Pro Cys Phe Arg 20 25
30 Asp Glu Asn Val His Phe Asn Arg Ile Phe Leu Pro Thr Ile Tyr
Phe 35 40 45 Ile
Ile Phe Leu Thr Gly Ile Val Gly Asn Gly Leu Val Ile Leu Val 50
55 60 Met Gly Tyr Gln Lys Lys
Leu Arg Ser Met Thr Asp Lys Tyr Arg Leu 65 70
75 80 His Leu Ser Val Ala Asp Leu Leu Phe Val Ile
Thr Leu Pro Phe Trp 85 90
95 Ala Val Asp Ala Met Ala Asp Trp Tyr Phe Gly Lys Phe Leu Cys Lys
100 105 110 Ala Val
His Ile Ile Tyr Thr Val Asn Leu Tyr Ser Ser Val Leu Ile 115
120 125 Leu Ala Phe Ile Ser Leu Asp
Arg Tyr Leu Ala Ile Val His Ala Thr 130 135
140 Asn Ser Gln Arg Pro Arg Lys Leu Leu Ala Glu Lys
Ala Val Tyr Val 145 150 155
160 Gly Val Trp Ile Pro Ala Leu Leu Leu Thr Ile Pro Asp Phe Ile Phe
165 170 175 Ala Asp Val
Ser Gln Gly Asp Ile Ser Gln Gly Asp Asp Arg Tyr Ile 180
185 190 Cys Asp Arg Leu Tyr Pro Asp Ser
Leu Trp Met Val Val Phe Gln Phe 195 200
205 Gln His Ile Met Val Gly Leu Ile Leu Pro Gly Ile Val
Ile Leu Ser 210 215 220
Cys Tyr Cys Ile Ile Ile Ser Lys Leu Ser His Ser Lys Gly His Gln 225
230 235 240 Lys Arg Lys Ala
Leu Lys Thr Thr Val Ile Leu Ile Leu Ala Phe Phe 245
250 255 Ala Cys Trp Leu Pro Tyr Tyr Val Gly
Ile Ser Ile Asp Ser Phe Ile 260 265
270 Leu Leu Gly Val Ile Lys Gln Gly Cys Asp Phe Glu Ser Ile
Val His 275 280 285
Lys Trp Ile Ser Ile Thr Glu Ala Leu Ala Phe Phe His Cys Cys Leu 290
295 300 Asn Pro Ile Leu Tyr
Ala Phe Leu Gly Ala Lys Phe Lys Ser Ser Ala 305 310
315 320 Gln His Ala Leu Asn Ser Met Ser Arg Gly
Ser Ser Leu Lys Ile Leu 325 330
335 Ser Lys Gly Lys Arg Gly Gly His Ser Ser Val Ser Thr Glu Ser
Glu 340 345 350 Ser
Ser Ser Phe His Ser Ser 355 41758DNAMus
musculusCDS(1)..(1080) 4atg gaa ccg atc agt gtg agt ata tac act tct gat
aac tac tct gaa 48Met Glu Pro Ile Ser Val Ser Ile Tyr Thr Ser Asp
Asn Tyr Ser Glu 1 5 10
15 gaa gtg ggg tct gga gac tat gac tcc aac aag gaa
ccc tgc ttc cgg 96Glu Val Gly Ser Gly Asp Tyr Asp Ser Asn Lys Glu
Pro Cys Phe Arg 20 25
30 gat gaa aac gtc cat ttc aat agg atc ttc ctg ccc
acc atc tac ttc 144Asp Glu Asn Val His Phe Asn Arg Ile Phe Leu Pro
Thr Ile Tyr Phe 35 40
45 atc atc ttc ttg act ggc ata gtc ggc aat gga ttg
gtg atc ctg gtc 192Ile Ile Phe Leu Thr Gly Ile Val Gly Asn Gly Leu
Val Ile Leu Val 50 55 60
atg ggt tac cag aag aag cta agg agc atg acg gac
aag tac cgg ctg 240Met Gly Tyr Gln Lys Lys Leu Arg Ser Met Thr Asp
Lys Tyr Arg Leu 65 70 75
80 cac ctg tca gtg gct gac ctc ctc ttt gtc atc aca
ctc ccc ttc tgg 288His Leu Ser Val Ala Asp Leu Leu Phe Val Ile Thr
Leu Pro Phe Trp 85 90
95 gca gtt gat gcc atg gct gac tgg tac ttt ggg aaa
ttt ttg tgt aag 336Ala Val Asp Ala Met Ala Asp Trp Tyr Phe Gly Lys
Phe Leu Cys Lys 100 105
110 gct gtc cat atc atc tac act gtc aac ctc tac agc
agc gtt ctc atc 384Ala Val His Ile Ile Tyr Thr Val Asn Leu Tyr Ser
Ser Val Leu Ile 115 120
125 ctg gcc ttc atc agc ctg gac cgg tac ctc gcc att
gtc cac gcc acc 432Leu Ala Phe Ile Ser Leu Asp Arg Tyr Leu Ala Ile
Val His Ala Thr 130 135 140
aac agt caa agg cca agg aaa ctg ctg gct gaa aag
gca gtc tat gtg 480Asn Ser Gln Arg Pro Arg Lys Leu Leu Ala Glu Lys
Ala Val Tyr Val 145 150 155
160 ggc gtc tgg atc cca gcc ctc ctc ctg act ata cct
gac ttc atc ttt 528Gly Val Trp Ile Pro Ala Leu Leu Leu Thr Ile Pro
Asp Phe Ile Phe 165 170
175 gcc gac gtc agc cag ggg gac atc agt cag ggg gat
gac agg tac atc 576Ala Asp Val Ser Gln Gly Asp Ile Ser Gln Gly Asp
Asp Arg Tyr Ile 180 185
190 tgt gac cgc ctt tac ccc gat agc ctg tgg atg gtg
gtg ttt caa ttc 624Cys Asp Arg Leu Tyr Pro Asp Ser Leu Trp Met Val
Val Phe Gln Phe 195 200
205 cag cat ata atg gtg ggt ctc atc ctg ccc ggc atc
gtc atc ctc tcc 672Gln His Ile Met Val Gly Leu Ile Leu Pro Gly Ile
Val Ile Leu Ser 210 215 220
tgt tac tgc atc atc atc tct aag ctg tca cac tcc
aag ggc cac cag 720Cys Tyr Cys Ile Ile Ile Ser Lys Leu Ser His Ser
Lys Gly His Gln 225 230 235
240 aag cgc aag gcc ctc aag acg aca gtc atc ctc atc
cta gct ttc ttt 768Lys Arg Lys Ala Leu Lys Thr Thr Val Ile Leu Ile
Leu Ala Phe Phe 245 250
255 gcc tgc tgg ctg cca tat tat gtg ggg atc agc atc
gac tcc ttc atc 816Ala Cys Trp Leu Pro Tyr Tyr Val Gly Ile Ser Ile
Asp Ser Phe Ile 260 265
270 ctt ttg gga gtc atc aag caa gga tgt gac ttc gag
agc att gtg cac 864Leu Leu Gly Val Ile Lys Gln Gly Cys Asp Phe Glu
Ser Ile Val His 275 280
285 aag tgg atc tcc atc aca gag gcc ctc gcc ttc ttc
cac tgt tgc ctg 912Lys Trp Ile Ser Ile Thr Glu Ala Leu Ala Phe Phe
His Cys Cys Leu 290 295 300
aac ccc atc ctc tat gcc ttc ctc ggg gcc aag ttc
aaa agc tct gcc 960Asn Pro Ile Leu Tyr Ala Phe Leu Gly Ala Lys Phe
Lys Ser Ser Ala 305 310 315
320 cag cat gca ctc aac tcc atg agc aga ggc tcc agc
ctc aag atc ctt 1008Gln His Ala Leu Asn Ser Met Ser Arg Gly Ser Ser
Leu Lys Ile Leu 325 330
335 tcc aaa gga aag cgg ggt gga cac tct tcc gtc tcc
acg gag tca gaa 1056Ser Lys Gly Lys Arg Gly Gly His Ser Ser Val Ser
Thr Glu Ser Glu 340 345
350 tcc tcc agt ttt cac tcc agc taa cccttatgca
aagacttata taatatatat 1110Ser Ser Ser Phe His Ser Ser
355
atatatatga taaagaactt ttttatgtta cacattttcc
agatataaga gactgaccag 1170tcttgtacag tttttttttt tttttaattg actgttggga
gtttatgttc ctctagtttt 1230tgtgaggttt gacttaattt atataaatat tgttttttgt
ttgtttcatg tgaatgagcg 1290tctaggcagg acctgtggcc aagttcttag tagctgttta
tctgtgtgta ggactgtaga 1350actgtagagg aagaaactga acattccaga atgtgtggta
aattgaataa agctagccgt 1410gatcctcagc tgttgctgca taatctcttc attccgagga
gcaccccacc cccaccccca 1470cccccacccc attcttaaat tgtttggtta tgctgtgtga
tggtttgttt ggtttttttt 1530tgttgttgtt gttgtttttt ttttctgtaa aagatggcac
ttaaaaccaa agcctgaaat 1590ggtggtagaa atgctggggt tttttttgtt tgtttgtttt
ttcagttttc aagagtagat 1650tgacttcagt ccctacaaat gtacagtctt gtattacatt
gttaataaaa gtcaatgata 1710aacttaaaaa aaaaaaaaaa aaaaaaaaaa aaaaaaaaaa
aaaaaaaa 1758589PRTArtificial SequenceLigand peptide -
Human SDF-1-alpha 5Met Asn Ala Lys Val Val Val Val Leu Val Leu Val Leu
Thr Ala Leu 1 5 10 15
Cys Leu Ser Asp Gly Lys Pro Val Ser Leu Ser Tyr Arg Cys Pro Cys
20 25 30 Arg Phe Phe Glu
Ser His Val Ala Arg Ala Asn Val Lys His Leu Lys 35
40 45 Ile Leu Asn Thr Pro Asn Cys Ala Leu
Gln Ile Val Ala Arg Leu Lys 50 55
60 Asn Asn Asn Arg Gln Val Cys Ile Asp Pro Lys Leu Lys
Trp Ile Gln 65 70 75
80 Glu Tyr Leu Glu Lys Ala Leu Asn Lys 85
62244DNAHomo sapiensCDS(471)..(743) 6gcacgggaca ggccgggcca cacccaccgg
ggcgagctcg gagggcggcg ctctgggcgg 60agggcccggc ggctcggccc agggcgcgtt
acctcgtcgc cggggccgga gagggcgggc 120ggaggcacgg ggcctggagg cgccaggcgg
aggatgcggg cgacacggtg gcggcggcga 180ccgcgcgacc gggcgggcgg gcgggcaggg
gcgagcggag ggagggagcg gactgcggca 240ggatctgtcg aggaaaaatc ttgcggccgg
cgattccccg ccttttaagc gcagcctgca 300ctccccccac cccacgcagg ggcgggcctt
ccccaacgcg ggcgcccact ggccgccgcg 360cgccgctccc ctccagctcg cctgcgcctc
tcactctccg tcagccgcat tgcccgctcg 420gcgtccggcc cccgacccgc gctcgtccgc
ccgcccgccc gcccgcccgc gcc atg 476
Ala Met
1 aac gcc aag gtc gtg gtc gtg ctg
gtc ctc gtg ctg acc gcg ctc tgc 524Asn Ala Lys Val Val Val Val Leu
Val Leu Val Leu Thr Ala Leu Cys 5 10
15 ctc agc gac ggg aag ccc gtc agc
ctg agc tac aga tgc cca tgc cga 572Leu Ser Asp Gly Lys Pro Val Ser
Leu Ser Tyr Arg Cys Pro Cys Arg 20 25
30 ttc ttc gaa agc cat gtt gcc aga
gcc aac gtc aag cat ctc aaa att 620Phe Phe Glu Ser His Val Ala Arg
Ala Asn Val Lys His Leu Lys Ile 35 40
45 50 ctc aac act cca aac tgt gcc ctt
cag att gta gcc cgg ctg aag aac 668Leu Asn Thr Pro Asn Cys Ala Leu
Gln Ile Val Ala Arg Leu Lys Asn 55
60 65 aac aac aga caa gtg tgc att gac
ccg aag cta aag tgg att cag gag 716Asn Asn Arg Gln Val Cys Ile Asp
Pro Lys Leu Lys Trp Ile Gln Glu 70
75 80 tac ctg gag aaa gct tta aac aag
taa gcacaacagc caaaaaggac 763Tyr Leu Glu Lys Ala Leu Asn Lys
85 90
tttccgctag acccactcga ggaaaactaa
aaccttgtga gagatgaaag ggcaaagacg 823tgggggaggg ggccttaacc atgaggacca
ggtgtgtgtg tggggtgggc acattgatct 883gggatcgggc ctgaggtttg ccagcattta
gaccctgcat ttatagcata cggtatgata 943ttgcagctta tattcatcca tgccctgtac
ctgtgcacgt tggaattttt attactgggg 1003tttttctaag aaagaaattg tattatcaac
agcattttca agcagttagt tccttcatga 1063tcatcacaat catcatcatt ctcattctca
ttttttaaat caacgagtac ttcaagatct 1123gaatttggct tgtttggagc atctcctctg
ctcccctggg gagtctgggc acagtcaggt 1183ggtggcttaa cagggagctg gaaaaagtgt
cctttcttca gacactgagg ctcccgcagc 1243agcgcccctc ccaagaggaa ggcctctgtg
gcactcagat accgactggg gctgggcgcc 1303gccactgcct tcacctcctc tttcaacctc
agtgattggc tctgtgggct ccatgtagaa 1363gccactatta ctgggactgt gctcagagac
ccctctccca gctattccta ctctctcccc 1423gactccgaga gcatgcatta atcttgcttc
tgcttctcat ttctgtagcc tgatcagcgc 1483cgcaccagcc gggaagaggg tgattgctgg
ggctcgtgcc ctgcatccct ctcctcccag 1543ggcctgcccc acagctcggg ccctctgtga
gatccgtctt tggcctcctc cagaatggag 1603ctggccctct cctggggatg tgtaatggtc
cccctgctta cccgcaaaag acaagtcttt 1663acagaatcaa atgcaatttt aaatctgaga
gctcgctttg agtgactggg ttttgtgatt 1723gcctctgaag cctatgtatg ccatggaggc
actaacaaac tctgaggttt ccgaaatcag 1783aagcgaaaaa atcagtgaat aaaccatcat
cttgccacta ccccctcctg aagccacagc 1843agggtttcag gttccaatca gaactgttgg
caaggtgaca tttccatgca taaatgcgat 1903ccacagaagg tcctggtggt atttgtaact
ttttgcaagg cattttttta tatatatttt 1963tgtgcacatt tttttttacg tttctttaga
aaacaaatgt atttcaaaat atatttatag 2023tcgaacaatt catatatttg aagtggagcc
atatgaatgt cagtagttta tacttctcta 2083ttatctcaaa ctactggcaa tttgtaaaga
aatatatatg atatataaat gtgattgcag 2143cttttcaatg ttagccacag tgtatttttt
cacttgtact aaaattgtat caaatgtgac 2203attatatgca ctagcaataa aatgctaatt
gtttcatggt a 2244789PRTArtificial SequenceLigand
peptide - Murine SDF-1-alpha 7Met Asp Ala Lys Val Val Ala Val Leu Ala Leu
Val Leu Ala Ala Leu 1 5 10
15 Cys Ile Ser Asp Gly Lys Pro Val Ser Leu Ser Tyr Arg Cys Pro Cys
20 25 30 Arg Phe
Phe Glu Ser His Ile Ala Arg Ala Asn Val Lys His Leu Lys 35
40 45 Ile Leu Asn Thr Pro Asn Cys
Ala Leu Gln Ile Val Ala Arg Leu Lys 50 55
60 Asn Asn Asn Arg Gln Val Cys Ile Asp Pro Lys Leu
Lys Trp Ile Gln 65 70 75
80 Glu Tyr Leu Glu Lys Ala Leu Asn Lys 85
81781DNAMus musculusCDS(82)..(351) 8gaccactttc cctctcggtc cacctcggtg
tcctcttgct gtccagctct gcagcctccg 60gcgcgccctc ccgcccacgc c atg gac
gcc aag gtc gtc gcc gtg ctg gcc 111 Met Asp
Ala Lys Val Val Ala Val Leu Ala 1
5 10 ctg gtg ctg gcc gcg ctc tgc atc
agt gac ggt aaa cca gtc agc ctg 159Leu Val Leu Ala Ala Leu Cys Ile
Ser Asp Gly Lys Pro Val Ser Leu 15
20 25 agc tac cga tgc ccc tgc cgg ttc
ttc gag agc cac atc gcc aga gcc 207Ser Tyr Arg Cys Pro Cys Arg Phe
Phe Glu Ser His Ile Ala Arg Ala 30
35 40 aac gtc aag cat ctg aaa atc ctc
aac act cca aac tgt gcc ctt cag 255Asn Val Lys His Leu Lys Ile Leu
Asn Thr Pro Asn Cys Ala Leu Gln 45 50
55 att gtt gca cgg ctg aag aac aac
aac aga caa gtg tgc att gac ccg 303Ile Val Ala Arg Leu Lys Asn Asn
Asn Arg Gln Val Cys Ile Asp Pro 60 65
70 aaa tta aag tgg atc caa gag tac
ctg gag aaa gct tta aac aag taa 351Lys Leu Lys Trp Ile Gln Glu Tyr
Leu Glu Lys Ala Leu Asn Lys 75 80
85 gcacaacagc ccaaaggact
ttccagtaga cccccgagga aggctgacat ccgtgggaga 411tgcaagggca gtggtgggga
ggagggcctg aaccctggcc aggatggccg gcgggacagc 471actgactggg gtcatgctaa
ggtttgccag cataaagaca ctccgccata gcatatggta 531cgatattgca gcttatattc
atccctgccc tcgcccgtgc acaatggagc ttttataact 591ggggtttttc taaggaattg
tattacccta accagttagc ttcatcccca ttctcctcat 651cctcatcttc attttaaaaa
gcagtgatta cttcaagggc tgtattcagt ttgctttgga 711gcttctcttt gccctggggc
ctctgggcac agttatagac ggtggctttg cagggagccc 771tagagagaaa ccttccacca
gagcagagtc cgaggaacgc tgcagggctt gtcctgcagg 831gggcgctcct cgacagatgc
cttgtcctga gtcaacacaa gatccggcag agggaggctc 891ctttatccag ttcagtgcca
gggtcgggaa gcttccttta gaagtgatcc ctgaagctgt 951gctcagagac cctttcctag
ccgttcctgc tctctgcttg cctccaaacg catgcttcat 1011ctgacttccg cttctcacct
ctgtagcctg acggaccaat gctgcaatgg aagggaggag 1071agtgatgtgg ggtgccccct
ccctctcttc cctttgcttt cctctcactt gggccctttg 1131tgagattttt ctttggcctc
ctgtagaatg gagccagacc atcctggata atgtgagaac 1191atgcctagat ttacccacaa
aacacaagtc tgagaattaa tcataaacgg aagtttaaat 1251gaggatttgg accttggtaa
ttgtccctga gtcctatata tttcaacagt ggctctatgg 1311gctctgatcg aatatcagtg
atgaaaataa taataataat aataataacg aataagccag 1371aatcttgcca tgaagccaca
gtggggattc tgggttccaa tcagaaatgg agacaagata 1431aaacttgcat acattcttat
gatcacagac ggccctggtg gtttttggta actatttaca 1491aggcattttt ttacatatat
ttttgtgcac tttttatgtt tctttggaag acaaatgtat 1551ttcagaatat atttgtagtc
aattcatata tttgaagtgg agccatagta atgccagtag 1611atatctctat gatcttgagc
tactggcaac ttgtaaagaa atatatatga catataaatg 1671tattgtagct ttccggtgtc
agccacggtg tatttttcca cttggaatga aattgtatca 1731actgtgacat tatatgcact
agcaataaaa tgctaattgt ttcatgctgt 178194PRTArtificial
SequenceAdded peptide to derive Human SDF-1-beta 9Arg Phe Lys Met 1
104PRTArtificial SequenceAdded peptide to derive Murine
SDF-1-beta 10Arg Leu Lys Met 1 1127DNAArtificial
SequenceCXCR4-specific forward primer 11tagcggccgc gttgccatgg aaccgat
271227DNAArtificial
SequenceCXCR4-specific reverse primer 12gcgtcgactt tgcataaggg ttagctg
27
User Contributions:
Comment about this patent or add new information about this topic:
| People who visited this patent also read: | |
| Patent application number | Title |
|---|---|
| 20140318410 | LOCOMOTIVE WITH VARIABLE POWER MODULES |
| 20140318409 | CARRIAGE ASSEMBLY FOR AN AIRPLANE TRANSPORTING SYSTEM ON AN AIRPORT APRON |
| 20140318408 | Drive Apparatus for a Magnetic Levitation Transport System |
| 20140318407 | MACHINE AND METHOD FOR RAILWAY TRACK MAINTENANCE, FOR TRACK LEVELLING, ALIGNMENT, COMPACTION AND STABILISATION, CAPABLE OF OPERATING WITHOUT INTERRUPTING THE FORWARD MOVEMENT THEREOF |
| 20140318406 | BULLET |