Patent application title: METHODS FOR IDENTIFYING TARGETING DOMAINS AND METHOD AND COMPOSITIONS COMPRISING THE SAME
Inventors:
Anton Wellstein (Washington, DC, US)
Kevin J. Mcdonnell (Little Neck, NY, US)
Justinian R. Ngaiza (Centreville, VA, US)
Assignees:
GEORGETOWN UNIVERSITY
IPC8 Class: AA61K5110FI
USPC Class:
424 149
Class name: Drug, bio-affecting and body treating compositions radionuclide or intended radionuclide containing; adjuvant or carrier compositions; intermediate or preparatory compositions attached to antibody or antibody fragment or immunoglobulin; derivative
Publication date: 2012-08-02
Patent application number: 20120195829
Abstract:
The present invention is directed to a method of identifying tissue
targeting domains. In particular, the invention relates to methods for
identifying a polynucleotide encoding a targeting domain which directs
tumor cell localization to secondary sites, to methods of utilizing the
polynucleotide and corresponding polypeptide or fragments thereof and
compositions comprising the same.Claims:
1-37. (canceled)
38. A method of targeting delivery of a therapeutic agent or a detection moiety to the liver or lung of a subject comprising administering to the subject a targeting domain that is associated with liver metastasis or lung metastasis and is operably linked to the therapeutic agent or the detection moiety, thereby targeting delivery of the therapeutic agent or the detection moiety to the liver or lung of the subject.
39. The method of claim 38, wherein the targeting domain comprises a polypeptide sequence selected from the group consisting of: SEQ ID NO:10, SEQ ID NO:11, SEQ ID NO:12, SEQ ID NO:13, SEQ ID NO:14, SEQ ID NO:15, SEQ ID NO:16, SEQ ID NO:17, SEQ ID NO:18, SEQ ID NO:19, SEQ ID NO:20, SEQ ID NO:21, SEQ ID NO:22, SEQ ID NO:23, SEQ ID NO:24, SEQ ID NO:25, SEQ ID NO:26, SEQ ID NO:27, SEQ ID NO:28, SEQ ID NO:29, SEQ ID NO:31, SEQ ID NO:32, SEQ ID NO:33, SEQ ID NO:34, SEQ ID NO:35, and SEQ ID NO:36.
40. The method of claim 38, wherein the targeting domain is encoded by: (a) a polynucleotide selected from the group consisting of SEQ ID NO:1, SEQ ID NO:2, SEQ ID NO:3, SEQ ID NO:4, SEQ ID NO:5, SEQ ID NO:6, SEQ ID NO:7, SEQ ID NO:8, and SEQ ID NO:9; or (b) a polynucleotide that hybridizes under highly stringent conditions to a polynucleotide selected from the group consisting of SEQ ID NO:1, SEQ ID NO:2, SEQ ID NO:3, SEQ ID NO:4, SEQ ID NO:5, SEQ ID NO:6, SEQ ID NO:7, SEQ ID NO:8, and SEQ ID NO:9.
41. The method of claim 38, wherein the therapeutic agent is a radioisotope, a cytotoxic agent, a clotting factor, or a thrombolytic factor.
42. The method of claim 38, wherein the detection moiety is a radioisotope, a dye, a pigment, or a fluorescent molecule.
43. An antibody directed against a targeting domain of a polypeptide (a) comprising a polypeptide sequence selected from the group consisting of SEQ ID NO:10, SEQ ID NO:11, SEQ ID NO:12, SEQ ID NO:13, SEQ ID NO:14, SEQ ID NO:15, SEQ ID NO:16, SEQ ID NO:17, SEQ ID NO:18, SEQ ID NO:19, SEQ ID NO:20, SEQ ID NO:21, SEQ ID NO:22, SEQ ID NO:23, SEQ ID NO:24, SEQ ID NO:25, SEQ ID NO:26, SEQ ID NO:27, SEQ ID NO:28, SEQ ID NO:29, SEQ ID NO:31, SEQ ID NO:32, SEQ ID NO:33, SEQ ID NO:34, SEQ ID NO:35, and SEQ ID NO:36; (b) encoded by a polynucleotide selected from the group consisting of SEQ ID NO:1, SEQ ID NO:2, SEQ ID NO:3, SEQ ID NO:4, SEQ ID NO:5, SEQ ID NO:6, SEQ ID NO:7, SEQ ID NO:8, and SEQ ID NO:9; or (c) encoded by a polynucleotide that hybridizes under highly stringent conditions to a polynucleotide selected from the group consisting of SEQ ID NO:1, SEQ ID NO:2, SEQ ID NO:3, SEQ ID NO:4, SEQ ID NO:5, SEQ ID NO:6, SEQ ID NO:7, SEQ ID NO:8, and SEQ ID NO:9, wherein the targeting domain of the polypeptide is associated with metastasis.
44. The antibody of claim 43, wherein the antibody is a polyclonal antibody or a monoclonal antibody.
45. The antibody of claim 43, wherein the antibody is derived from a mouse or a human.
46. The antibody of claim 43, wherein the antibody is a chimeric antibody or a humanized antibody.
47. A method of treating metastasis in a subject in need of such treatment comprising administering to the subject the antibody of claim 43 coupled to a therapeutic agent in an amount effective to treat the metastasis.
48. The method of claim 43, wherein the therapeutic agent is a radioisotope, a cytotoxic agent, a clotting factor, or a thrombolytic factor.
49. The method of claim 43, wherein the metastasis is liver metastasis or lung metastasis.
50. A method of detecting metastasis in a subject having the metastasis comprising administering to the subject the antibody of claim 43 coupled to an imaging moiety and detecting the metastasis.
51. The method of claim 50, wherein the imaging moiety is a radioisotope, a dye, a pigment, or a fluorescent molecule.
52. The method of claim 50, wherein the metastasis is liver metastasis or lung metastasis.
53. A method of treating liver metastasis or lung metastasis in a subject in need of such treatment comprising administering to the subject a targeting domain associated with liver metastasis or lung metastasis, respectively, in an amount effective to treat the liver metastasis or lung metastasis, wherein the targeting domain is operably linked to a therapeutic agent.
54. The method of claim 53, wherein the therapeutic agent is a radioisotope, a cytotoxic agent, a clotting factor, or a thrombolytic factor.
55. A method of treating liver metastasis or lung metastasis in a subject in need of such treatment comprising administering to the subject a composition in an amount effective to treat the liver or lung metastasis wherein the composition comprises: (a) a targeting domain associated with the liver metastasis or lung metastasis and operably linked to a therapeutic agent; or (b) an antibody coupled to a therapeutic agent and directed against a targeting domain that is associated with the liver metastasis or lung metastasis.
56. The method of claim 55, wherein the therapeutic agent is a radioisotope, a cytotoxic agent, a clotting factor, or a thrombolytic factor.
57. The method of claim 55, wherein the antibody is a polyclonal antibody or a monoclonal antibody.
58. The method of claim 55, wherein the antibody is derived from a mouse or a human.
59. The method of claim 55, wherein the antibody is a chimeric antibody or a humanized antibody.
Description:
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001] This application claims the benefit of U.S. Provisional Application Ser. No. 60/345,520, filed Feb. 8, 2002, the disclosure of which is incorporated herein by reference in its entirety.
FIELD OF THE INVENTION
[0003] The present invention relates to the identification of tissue targeting domains. In particular, the invention relates to methods for identifying a polynucleotide encoding a targeting domain which directs tumor cell localization to secondary sites, to methods of utilizing the polynucleotide and corresponding polypeptide or fragments thereof and compositions comprising the same.
BACKGROUND OF THE INVENTION
[0004] Cancer is a leading cause of death in the United States. In 2001, of the over half a million deaths in the United States, one in every four deaths was likely attributable to cancer. If all cancers were diagnosed at a localized stage, the five year survival rate would be over 95%. Overall, metastasis is considered by many to be the deadliest aspect of cancer.
[0005] It has been clinically noted that particular primary tumors tend to metastasize to specific distant organs. For example, prostate cancer often metastasizes to the bone, breast cancer may metastasize to the liver, melanomas tend to spread to lymph nodes, ovarian cancer metastasizes to other areas of the body including the lungs, brain, lymph and bones. Once a cancer has spread, it becomes much more lethal. No longer is a simple surgical intervention (e.g., to remove the primary tumor) an effective form of treatment. In addition, "blunt instruments" (e.g., radiation treatment and chemotherapy) affect not only cancerous cells, but also normal tissues throughout the body.
[0006] Cancer metastasis involves a series of sequential steps. After the initial transforming event, growth of neoplastic cells must be progressive. Extensive vascularization or angiogeneis also must occur, which allows blood vessels to grow into the tumor mass, bringing nourishment and allowing increased tumor growth. As the vascularization increases and the tumor grows, the thin walled venules or anastomoses of the capillary network allow for the penetration of cancer cells. These cells then may detach from the tumor mass and enter the circulation in a process called "embolization." The majority of cancer cell aggregates that enter the circulation is destroyed, yet some of the aggregates migrate to distant capillary beds and begin a process called extravasation. During extravasation, tumor cells exit the capillary network, colonize a distant organ and create a secondary or metastatic tumor. This "homing," or metastasis, of particular types of tumor cells to specific "target" organs provides further evidence that organ-specific markers exist.
[0007] The extravasation process is thought to begin with a type of adherence to the vascular walls--either by (i) attachment to specific proteins on the endothelial surface of the vasculature or (ii) a non-specific type of adhesion to the homing molecule at the target organ. In either case, there exists the possibility for multiple molecules and mechanisms of adherence of both homing molecules from primary tumors of different origins and for target molecules at the site of secondary metastatic tumors in specific organs.
[0008] Dreyer and Hood formulated the "Area Code Hypothesis" in the study of embryology and tissue differentiation. J. Supramol Struct. 1977; 7(3-4); 531-559. This hypothesis is concerned with the structure, function and regulation of cell-surface molecules that mediate recognition during embryogenesis. Ruoslahti and Pasqualini, who applied the area code hypothesis, developed a method that involved putting random peptide sequences in a phage display library, which then were injected into mice (see e.g. U.S. Pat. Nos. 5,622,699 and 6,232,287). Such "in vivo phage display" led to the identification of several molecular motifs, which localized to specific organs. The goal which Ruoslahti and Pasqualini hoped to achieve was a method to specifically attack metastatic tumors using the identified motifs. However, the physiologic basis for this targeting remains unknown, and neither the native homing molecules on the metastatic cell or the target molecule at the site of the secondary tumor have been identified, with singular exceptions.
[0009] In addition, Pasqualini and Ruoslahti expressed skepticism that organs, which filter a high blood volume would be amenable to the procedure they described, due to their ability to non-specifically capture a large number of blood borne peptides. Nevertheless, the clinical observation that particular primary tumors do home in on target organs in spite of their small volume of blood flow, prior to colonizing organs with high volumes, such as the liver, kidney or lungs is well documented. If these mechanisms could be identified, powerful new ways to study and treat cancer would be available. There is, therefore, a well recognized need to identify molecules that allow the homing of cancer cells in vivo. In the same vein, there is a need for a mechanism to identify molecules at distant sites that are targeted by metastasizing cells.
SUMMARY OF THE INVENTION
[0010] The present invention overcomes the problems and disadvantages associated with current strategies and designs and provides new diagnostic and therapeutic methods relating to metastatic disease. The invention relates generally to a method of identifying a polynucleotide encoding a targeting domain which directs tumor cell localization to secondary site (e.g., metastasis), to the isolated polynucleotide and/or corresponding polypetide identified by the method, to methods of utilizing the polynucleotide and/or corresponding polypeptide in diagnostic and therapeutic applications and to compositions comprising the same.
[0011] One aspect of the invention provides a method of identifying a polynucleotide encoding a targeting domain associated with metastasis of tumor cells, the method comprising: (a) administering a phage displaying libraries comprising a collection of phages containing polynucleotides from, preferably, a primary tumor cell into a subject; (b) selecting phage that localize in a target organ or tissue; (c) collecting phage from the selected organ or tissue; (d) repeating steps (a) and (c) for one or more cycles; and (e) identifying one or more polynucleotides encoding a targeting domain or fragment thereof from selected phage that are associated with tumor cell metastasis. This method allows for the identification of polynucleotides and their expression products that are associated with metastasis and, preferably, those responsible for metastatic disease or organ targeting.
[0012] In one embodiment of the method, cDNA libraries from different primary tumors are packaged into T7 phage and injected in vivo into mice. After circulation, organs are extracted, a phage titer determined, and phage amplified in bacterial cells. This process of injection, organ removal and rescue and amplification of phage from the target organ (i.e. biopanning), is repeated multiple times and results in enrichment for phage possessing organ selectivity when compared to empty plasmid controls. The polynucleotide and/or expression product (e.g., polypeptide) of the phage exhibiting organ selectivity are characterized by well-known biochemical methods.
[0013] Another aspect of this invention is directed to isolated polynucleotides identified by the method described herein. In one embodiment, the polynucleotide hybridizes under stringent conditions to a polynucleotide comprising the sequence of SEQ ID NOS: 1, 2, 3, 4, 5, 6, 7, 8, or 9. In another embodiment, the polynucleotide comprises the sequence of SEQ ID NOS: 1, 2, 3, 4, 5, 6, 7, 8 or 9. In yet another embodiment the isolated polynucleotide encodes a polypeptide comprising the polypeptide sequence of SEQ ID NOS: 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, or 36. In yet another embodiment, the polynucleotide encodes a polypeptide comprising a targeting domain of the polypeptide of SEQ ID NOS: 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, or 36.
[0014] Another aspect of this invention is directed to isolated polypeptides identified by the method described herein. In one embodiment the polypeptide comprises the amino acid sequence of SEQ ID NOS: 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, or 36.
[0015] In yet another aspect of the invention, microarrays comprising the polynucleotides and/or polypeptides of the invention are provided.
[0016] Yet another aspect of the inventions relates to an antibody directed against the polypeptides of the invention.
[0017] In yet another aspect, polypeptides comprising the targeting domain coupled to a moiety (e.g., therapeutic or detection moiety) are provided.
[0018] Yet another aspect of this invention provides methods of prognosing and/or diagnosing metastatic disease in a subject. In one embodiment, the method comprises detecting the level of a polynucleotide encoding a polypeptide comprising the targeting domain in a sample obtained from a subject, wherein a higher level of the polynucleotide relative to a control sample (e.g., population controls or non-metastaic control sample) is indicative of metastatic disease. In another embodiment, the method comprises detecting the presence or absence of a polynucleotide encoding a polypeptide comprising the targeting domain in a sample obtained from the subject, wherein the presence of the polynucleotide is indicative of metastatic disease.
[0019] Yet another aspect of this invention provides methods of prognosing and/or diagnosing metastatic disease in a subject. In one embodiment, the method comprises detecting the level of a polypeptide comprising the targeting domain in a sample obtained from a subject, wherein a higher level of the polypeptide relative to a control sample (e.g., population controls or non-metastaic control sample) is indicative of metastatic disease. In another embodiment, the method comprises detecting the presence or absence of a polypeptide comprising a targeting domain in a sample obtained from the subject, wherein the presence of the polypeptide is indicative of metastatic disease.
[0020] Yet another aspect of the invention relates to detection of metastatic disease in a subject, such as a human utilizing antibodies coupled to a radiologic or other imaging molecules to detect metastasis in the subject.
[0021] A further aspect of the invention comprises methods of treating metastasis in a subject in need of such treatment. In some embodiments the method comprises administering to a subject in need of such treatment a targeting domain linked to a therapeutic agent in an amount effective to treat the metastasis, or an effective amount of a composition that inhibits the metastasis (e.g., collection of phage or phage expression products identified by the method herein; a targeting domain linked to a therapeutic agent and/or an antibody directed against a polypeptide comprising a targeting domain).
[0022] Yet another aspect of the invention provides methods of screening for candidate agents that inhibit the selectivity of the targeting domain.
[0023] Yet another aspect of the invention relates to kits and compositions for use in the methods described herein.
[0024] Other embodiments and advantages of the invention are set forth in part in the description, which follows, and in part, may be obvious from this description, or may be learned from the practice of the invention.
DESCRIPTION OF THE FIGURES
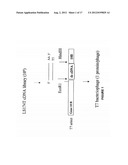
[0025] FIG. 1: is a diagrammatic representation of the preparation of the LS174T library showing the directional cloning of tumor cDNA into the T7 expression vector. cDNA, digested with EcoR1 and HindIII produced cDNA with and EcoR1 5' and HindIII 3' end. Ligation of these fragments into the T7 vector was through the corresponding EcoR1/HindIII vector arms so that inserts were in the sense orientation relative to the upstream expression signal.
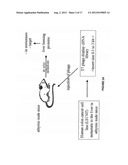
[0026] FIG. 2: (A and B) is a schematic representation of the experimental protocol. Mice were injected with the LS174T cDNA expression library over three successive rounds of biopanning using the liver-retained clones. (C) Phage titers after each round of biopanning are illustrated. Corresponding PCR gels created using T7 Up and Down arms as primers reveal band clarification after the second and third rounds of biopanning. (D) Organ distribution of injected phage clones after the fourth round of biopanning is illustrated.
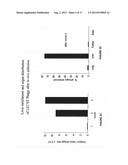
[0027] FIG. 3: (A) shows the degree of liver selectivity by individual phage clones injected into immunodeficient non-tumor bearing mice. Comparison is made between the portal vein and inferior vena cava. Titers of individual clones represented as a proportion of the phage numbers from kidneys of the respective animals are shown. (B) Using the two vascular beds with the largest volume, numbers for lung and liver binding (RCF of 1205LU) in each animal are presented together. Each bar represents the mean of three platings.
[0028] FIG. 4: shows the clone TD, frequency of clones in percent, and peptide insert length from the liver-selected library after the final round (four rounds total) of biopanning.
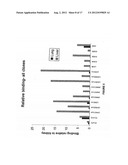
[0029] FIG. 5: shows the relative binding of all clones to lung and liver as compared to kidney.
[0030] FIG. 6: (A-C) shows the polynucleotide sequences for SEQ ID NOs: 1-9.
[0031] FIG. 7: (A-E) shows the polypeptide sequences for SEQ ID NOs: 10-36.
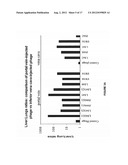
[0032] FIG. 8: is a comparison of patient cases with and without known metastasis indicating a highly significant increases in expression levels and frequency of the phage-display derived metastasis genes and of PTN for the cases with metastasis.
DETAILED DESCRIPTION OF THE INVENTION
[0033] The practice of the present invention will employ, unless otherwise indicated, conventional techniques of molecular biology (including recombinant techniques), microbiology, cell biology, biochemistry and immunology, which are within the skill of the art. Such techniques are explained fully in the literature, such as, Molecular Cloning: A Laboratory Manual, second edition (Sambrook et al., 1989) Cold Spring Harbor Press; Oligonucleotide Synthesis (M. J. Gait, ed., 1984); Methods in Molecular Biology, Humana Press; Cell Biology: A Laboratory Notebook (J. E. Cellis, ed., 1998) Academic Press; Animal Cell Culture (R. I. Freshney, ed., 1987); Introduction to Cell and Tissue Culture (J. P. Mather and P. E. Roberts, 1998) Plenum Press; Cell and Tissue Culture: Laboratory Procedures (A. Doyle, J. B. Griffiths, and D. G. Newell, eds., 1993-8) J. Wiley and Sons; Methods in Enzymology (Academic Press, Inc.); Handbook of Experimental Immunology (D. M. Weir and C. C. Blackwell, eds.); Gene Transfer Vectors for Mammalian Cells (J. M. Miller and M. P. Calos, eds., 1987); Current Protocols in Molecular Biology (F. M. Ausubel et al., eds., 1987); PCR: The Polymerase Chain Reaction, (Mullis et al., eds., 1994); Current Protocols in Immunology (J. E. Coligan et al., eds., 1991); Short Protocols in Molecular Biology (Wiley and Sons, 1999); Immunobiology (C. A. Janeway and P. Travers, 1997); Antibodies (P. Finch, 1997); Antibodies: a practical approach (D. Catty., ed., IRL, Press, 1988-1989); Monoclonal antibodies: a practical approach (P. Shepherd and C. Dean, eds., Oxford University Press, 2000); Using antibodies: a laboratory manual (E. Harlow and D. Lane (Cold Spring Harbor Laboratory Press, 1999); The Antibodies (M. Zanetti and J. D. Capra, eds., Harwood Academic Publishers, 1995); and Cancer: Principles and Practice of Oncology (V. T. DeVita et al., eds., J. B. Lippincott Company, 1993).
DEFINITIONS
[0034] As used herein, the singular form "a", "an", and "the" includes plural references unless indicated otherwise. For example, "a" polynucleotide includes one or more polynucleotides and "a targeting domain" means one or more targeting domains.
[0035] The term "targeting domain" or "homing domain" or "homing molecule" or "homing protein" generally, but not exclusively, refers to a polypeptide that selectively or preferentially targets a particular cell type and/or tissue. By way of example, a targeting domain directs tumor cell localization from a primary tumor (e.g., colon cancer) to secondary sites (e.g., liver, lung, marrow and/or lung).
[0036] The term "selectively targets" or "preferentially targets" (used interchangeably herein) is a term well understood in the art, and methods to determine such specific or preferential targeting are also well known in the art. A polypeptide is said to exhibit "selective" or "preferential" targeting if it reacts or associates more frequently, more rapidly, with greater duration and/or with greater affinity with a particular cell and/or tissue than it does with alternative cells and/or tissues. "Selectively targets" or "preferentially targets" does not necessarily require (although it can include) exclusive binding. By way of example, a polypeptide identified by the methods described herein selectively or preferentially targets a cell or tissue if it exhibits between about 3 to about 300 fold selectivity. Method for determining selective or preferential targeting are exemplified herein.
[0037] An "subject" may be any animal, preferably a vertebrate, most preferably a mammal. Examples, include, but are not limited to, rodents (e.g., mouse or rats), cats, dogs, rabbits, farm animals (e.g., pigs, horses, cows) or humans.
[0038] As embodied and broadly described herein, the present invention is directed to novel methods of identifying polynucleotides encoding targeting domains associated with metastatisis disease or disorders, the polynucleotides and polypeptides identified by the method. The invention is also directed to diagnostic and therapeutic compositions, kits and methods useful in the treatment, prevent and detection of metastatic disorders.
[0039] Method of Identifying Targeting Domains
[0040] The ability of tumors to metastasize is, at least in part, the result of genes whose products comprise a targeting domain which selectively directs a tumor cell from the primary tumor to secondary tissue metastatic sites. By way of example, the targeting domain may act as a receptor to a ligand on the surface of vessels in the specific target organs or as ligands to receptor proteins on the surface of vessels in those organs. This invention is based on the discovery of a method which identifies polynucleotides associated with metastasis and the targeting domain encoded by such polynucleotides. These polynucleotides and/or polypeptides, provide an understanding of the mechanism of action of metastatic tumors at a molecular level.
[0041] One embodiment of the invention is directed to libraries created from primary tumors that can be utilized for the identification of homing genes. As different homing proteins may have different targets in different tissues, another embodiment of the invention to directed to methods for identifying targeting molecules from different primary tissue types and their complimentary targets from different sites of metastasis.
[0042] Genes responsible for metastatic targeting of one or more organs, tissues and other areas of the body, can be identified by the current invention. Primary tumors that are known to metastasize to particular organs are selected for the production of a phage library. That library is then injected into an appropriate model. The model is preferably a mammal such as, for example, a human, a mouse or a rabbit, but may also be any other mammal. Alternatively, the method may utilize any animal, including non-mammals, that allows for the injection of phage library and demonstrates a significant response.
[0043] Human tumor libraries are often not available or may not yield the greatest response in a non-human model. Accordingly, the present invention comprises tumor libraries derived from animals that show a response in the model. With the successful completion of the human genome project, identification of homing and target genes in a non-human library for the identification of human homologues.
[0044] Accordingly, this invention provides a method of identifying a polynucleotide encoding a targeting domain associated with metastasis of tumor cells, the method comprising: (a) administering a phage displaying libraries comprising a collection of phages containing polynucleotides from a primary tumor cell into a subject; (b) selecting phage that localize in a target organ or tissue; (c) collecting phage from the selected organ or tissue; (d) repeating steps (a) and (c) for one or more cycles; and (e) identifying one or more polynucleotides encoding a targeting domain or fragment thereof from selected phage that are associated with tumor cell metastasis.
[0045] Any library may be used in the method described herein. Standard methods may be utilized to create the library or the library may be obtained from a commercial source. Examples of libraries that may be used in the method include, but are not limited to libraries created from primary tumors of lung, stomach, colon, rectum, prostate, pancreas, liver, leukemia, breast, uterus, ovary, melanoma, urinary tract, bladder, cervix, lymph, brain, nervous system or combinations thereof, peptide libraries, or libraries comprising molecules sharing common functional domains or sequence (e.g., kinases, cytokines, growth factors etc) or polynucleotides from any eukaryarotic cell. In one embodiment, a library used in the method is reused with the clones isolated from the first screen subtracted out from the library to minimize repetitive isolation of the same clone. Methods of creating subtraction libraries are well known in the art.
[0046] Any phage may be used to create the library. Preferably, the phage used in the creation of the library has one or more of the following characteristics: the ability to contain and relatively large polynucleotides, such as, for example, between about 300-3000 nucleotides and/or expresses the clone from the library at a low copy numbers, such as, for example, between about 0.1 copy to about 1 copy per phage. Such phage are commercially available (e.g., a T7Select vector using T7Select 1-1 phage). By way of example, a phage display library may comprise and express polynucleotides isolated from a primary tumor, such as, for example, colon cancer or from a cell line such as, for example, a colon cancer cell line (e.g., LS174T; American tissue culture collection, ATCC, Rockville, Md.). Preferably, the phage themselves (i.e. phage without a recombinant insert) have a low relative retention to target organs or cells. Retention, which may relate to direct binding, non-specific association, or active uptake, will cause phage to nonspecifically associate with target cells. By identifying and selecting only phage with low retentions by target cells, the highest selectivity can be achieved. Relative retention of phage to target tissue is preferably less than 50%, more preferably less than 10%, and still more preferably less than 1%.
[0047] The library is administered to any subject, preferably a mouse or other mammal. The animal may be a normal animal or an animal model of disease. Alternatively, the library may be contacted with in vitro systems or models. In an animal, such as for example, a mouse, a volume of between about 10 microliters to about 100 microliters containing between about 107 to about 1010 phage is administered to a mouse. Phage, based on the expression product displayed, target to selected organs, tissues or other areas of the body. Accordingly, the library is administered and allowed to circulate for a time to sufficient to allow binding to the target tissue and/or organ of the binding domains expressed in the library. The optimal circulation time will vary with the size/weight of the animal, volume and/or complexity of the library. By way of example, for a mouse circulation time may be preferably between about one minute to about ten minutes.
[0048] After sufficient circulation time the animal is euthanized and the target organs collected for analysis. The method described herein may be further enhanced by further comprising perusing the anesthetized animal with an isotonic salt solution with or without proteins (e.g., BSA) to minimize non-specific binding of phage. Examples of isotonic salt solutions include, but are not limited to phosphate buffer. Perfusion is continued, preferably until desanguination (e.g., little or no blood exits the vena cava, organs appear white in color.) By way of example, volumes of between about 1 to about 100, preferably about 20 times the volume of the animal may be used.
[0049] Any organ or tissue may be harvested for analysis. By way of example, but not limited to bone marrow, lung, skin, liver and/or brain. Generally the tissue or organ harvested will be selected based on the origin of the library. By way of example, metastasis in colon cancer is often to the liver, marrow, lung and/or bone marrow. If the library used in the method comprised polynucleotides from a primary colon cancer tumor or cell line, liver lung and/or bone marrow can be harvested
[0050] Phage are collected from the selected tissues and/or organs, amplified, if necessary, and injected into another animal. Through successive rounds of injection, selection, and amplification, a collection of phage can be isolated that are specific for the selection criteria. By way of example, between about two to about five rounds of injection, selection, and amplification may performed. These collections can be further selected or the polynucleotides from individual or groups of phage isolated and identified. Polynucleotides identified by these methods can be used for both diagnostic and therapeutic purposes. The polynucleotide expression products identified may be useful to distinguish metastatic from non-metastatic disease. Alternatively, the products may be useful in identifying new therapies for the treatment of metastatic and for the screening of promising pharmaceutical products.
[0051] The method described herein for identifying targeting domains may also be utilized to identify targeting domains in other diseases or disorders. By way of example, such diseases or disorders may include, but are not limited to, arteriosclerosis, coronary artery disease, stroke, diabetic vascular damage (e.g., kidney vascular damage) or retinopathy. Examples of animals models to be used in the methods described herein include, but are not limited to, cardiovascular diseases in pig, rat, rabbit arterial stenosis and vascularization. (e.g., Goodman and Gilman's: the Pharmaceutical Basis of Therapeutics Pergamon Press (1990)).
[0052] Polynucleotides
[0053] Another aspect of this invention is directed to isolated polynucleotides identified by the method described herein. The term polynucleotide is used broadly and refers to polymeric nucleotides of any length (e.g., oligonucleotides, genes, small inhibiting RNA etc). The polynucleotide of the invention may be, for example, linear, circular, supercoiled, single stranded, double stranded or branched. The nucleotides comprising the polynucleotide may be naturally occurring nucleotides or modified nucleotides. The polynucleotides illustrated in FIG. 6A-6B (SEQ ID NOS. 1-9) and/or their complement represent preferred embodiments of the invention. It is, however, understood by one skilled in the art that due to the degeneracy of the genetic code variations in the polynucleotide sequences shown will still result in a polynucleotide sequence capable of encoding a targeting domain. Such polynucleotide sequences are therefore functionally equivalent to the sequence set forth in FIG. 6A-6C and are intended to be encompassed within the present invention. Further, a person of skill in the art will understand that there are naturally occurring allelic variations of the polynucleotide sequences shown in FIG. 6A-6C these variations are also intended to be encompassed by the present invention.
[0054] In one embodiment the polynucleotide comprises the sequence of SEQ ID NOS: 1, 2, 3, 4, 5, 6, 7, 8, or 9. In yet another embodiment the isolated polynucleotide encodes a polypeptide comprising the polypeptide sequence of SEQ ID NOS: 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, or 36. In another embodiment, the polynucleotide encodes a polypeptide comprising a targeting domain of the polypeptide of SEQ ID NOS: 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, or 36.
[0055] This invention also relates to a polynucleotide that hybridizes under stringent conditions to a polynucleotide comprising the sequence of SEQ ID NOS: 1, 2, 3, 4, 5, 6, 7, 8, or 9. Hybridization reactions can be performed under conditions of different "stringency". Conditions that increase stringency of a hybridization reaction of widely known and published in the art. See, for example, Sambrook et al. (1989). Examples of relevant conditions include (in order of increasing stringency): incubation temperatures of 25° C., 37° C., 50° C. and 68° C.; buffer concentrations of 10×SSC, 6×SSC, 4×SSC, 1×SSC, 0.1×SSC (where SSC is 0.15 M NaCl and 15 mM citrate buffer) and their equivalents using other buffer systems; formamide concentrations of 0%, 25%, 50%, and 75%; incubation times from 5 minutes to 24 hours; 1, 2, or more washing steps; wash incubation times of 1, 2, or 15 minutes; and wash solutions of 6×SSC, 1×SSC, 0.1×SSC, or deionized water. In a preferred embodiment hybridization and wash conditions are done at high stringency. By way of example hybridization may be performed at 50% formamide and 4×SSC followed by washes of 2×SSC/formamide at 50° C. and with 1×SSC (see example).
[0056] Polypeptides
[0057] Another aspect of this invention is directed to isolated polypeptides identified by the methods described herein. The term polypeptide is used broadly herein to include peptide or protein or fragments thereof. Also intended to be encompassed are peptidomimetics, which include chemically modified peptides, peptide-like molecules containing normaturally occurring amino acids, peptoids and the like, have the selective binding of the targeting domains provided herein. ("Burger's Medicinal Chemistry and Drug Discovery" 5th ed., vols. 1 to 3 (ed. M. E. Wolff; Wiley Interscience 1995).
[0058] In one embodiment the polypeptide comprises the amino acid sequence of SEQ ID NOS: 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, or 36. This invention further includes polypeptides or analogs thereof having substantially the same function as the polypeptides of this invention. Such polypeptides include, but are not limited to, a substitution, addition or deletion mutant of the polypeptide. This invention also encompasses proteins or peptides that are substantially homologous to the polypeptides.
[0059] The term "analog" includes any polypeptide having an amino acid residue sequence substantially identical to at least one the polypeptide sequences specifically shown herein (FIG. 7A-7E) in which one or more residues have been conservatively substituted with a functionally similar residue and which displays the functional aspects of the polypeptides as described herein. Examples of conservative substitutions include the substitution of one non-polar (hydrophobic) residue such as isoleucine, valine, leucine or methionine for another, the substitution of one polar (hydrophilic) residue for another such as between arginine and lysine, between glutamine and asparagine, between glycine and serine, the substitution of one basic residue such as lysine, arginine or histidine for another, or the substitution of one acidic residue, such as aspartic acid or glutamic acid or another.
[0060] The phrase "conservative substitution" also includes the use of a chemically derivatized residue in place of a non-derivatized residue. "Chemical derivative" refers to a subject polypeptide having one or more residues chemically derivatized by reaction of a functional side group. Examples of such derivatized molecules include for example, those molecules in which free amino groups have been derivatized to form amine hydrochlorides, p-toluene sulfonyl groups, carbobenzoxy groups, t-butyloxycarbonyl groups, chloroacetyl groups or formyl groups. Free carboxyl groups may be derivatized to form salts, methyl and ethyl esters or other types of esters or hydrazides. Free hydroxyl groups may be derivatized to form O-acyl or O-alkyl derivatives. The imidazole nitrogen of histidine may be derivatized to form N-im-benzylhistidine. Also included as chemical derivatives are those proteins or peptides which contain one or more naturally-occurring amino acid derivatives of the twenty standard amino acids. For examples: 4-hydroxyproline may be substituted for proline; 5-hydroxylysine may be substituted for lysine; 3-methylhistidine may be substituted for histidine; homoserine may be substituted for serine; and ornithine may be substituted for lysine. Polypeptides of the present invention also include any polypeptide having one or more additions and/or deletions or residues relative to the sequence of a any one of the polypeptides whose sequences is described herein.
[0061] Methods of Prognosing and/or Diagnosing
[0062] level of metastasis may be correlated to the level of primary tumor growth leading to increased neoplastic embolism which then increases the level of neoplastic aggregates in the blood stream. A method of quantitatively measuring the RNA transcription product in the blood would allow an estimation of primary tumor growth and the metastatic potential of the tumor. In some cases, quantitative measurements can be made with a PCR or, alternatively, other methods to quantitatively measure transcription may be desirable. In many situations, different primary tumors metastasize to different organs.
[0063] The methods provided herein may be prognostic (e.g., detect subclinical metastasis, detection of subclinical metastasis in at risk patients, risk of metastasis) or diagnostic (e.g., detect metastasis, monitor disease progression or treatment). One embodiment, provides methods of prognosing and/or diagnosing metastatic disease in a subject. In one embodiment, the method comprises detecting the level of a polynucleotide encoding a polypeptide comprising the targeting domain in a sample obtained from a subject, wherein a higher level of the polynucleotide relative to a control sample (e.g., population controls or non-metastaic control sample) is indicative of metastatic disease. In another embodiment, the method comprises detecting the presence or absence of a polynucleotide encoding a polypeptide comprising the targeting domain in a sample obtained from the subject, wherein the presence of the polynucleotide is indicative of metastatic disease. Conventional methodology may be used to detect the polynucleotides in the method described herein. Examples include, but are not limited to, PCR analysis, RT-PCR, Northern analysis or microarrays as described herein below. Examples of a sample obtained from a subject include, but is not limited to, blood, biopsy sample, pathology sample, urine or cerebrospinal fluid.
[0064] Yet another aspect of this invention provides methods of prognosing, imaging and/or diagnosing metastatic disease in a subject. In one embodiment, the method comprises detecting the level of a polypeptide comprising the targeting domain in a sample obtained from a subject, wherein a higher level of the polypeptide relative to a control sample (e.g., population controls or non-metastaic control sample) is indicative of metastatic disease. In another embodiment, the method comprises detecting the presence or absence of a polypeptide comprising a targeting domain in a sample obtained from the subject, wherein the presence of the polypeptide is indicative of metastatic disease. Conventional methodology may be used to detect the polypeptides in the method described herein.
[0065] Examples include, but are not limited to, Western blot analysis or protein microarrays. Other methods of quantitative analysis of proteins include, for example, proteomics technologies such as isotope coded affinity tag reagents, MALDI TOF/TOF tandem mass spectrometry, and 2D-gel/mass spectrometry technologies. These technologies are commercially available from, for example, Large Scale Proteomics, Inc. (Germantown, Md.) and Oxford Glycosystems (Oxford UK). Methods for quantitatively measuring proteins such as ELISA analyses are well known. Kits for measuring levels of many proteins using ELISA assays are commercially available from many suppliers. In addition, methods for developing ELISA assays in the laboratory are well known. See for example Antibodies: A Laboratory Manual (Harlow and Lane Eds. Cold Spring Harbor Press). Antibodies for use in such ELISA methods either are commercially available or are prepared using well-known methods. Examples of a sample obtained from a subject include, but is not limited to, blood, biopsy sample, pathology sample, urine or cerebrospinal fluid.
[0066] Microarrays
[0067] In yet another aspect of the invention, microarrays comprising one or more of the polynucleotides and/or one or more of the polypeptides of the invention. Preferred polynucleotide sequences are shown in FIG. 6A-6C. preferred polypeptide sequences are shown in FIG. 7A-7C. Methods of making microarrays are known in the art. By way of example, one or more of the polynucleotide sequences described herein may comprise an array of polynucleotides attached to a support (e.g., dot blots on a nylon hybridization membrane Sambrook et al., or Ausubel et al) that is contacted with the nucleic acids isolated from, for example, a patient sample.
[0068] Microarrays may be a solid phase on the surface of which are immobilized a population of the nucleic acids of the invention. Microarrays can be generated in a number of ways. The polynucleotides can be attached to a solid support or surface, which may be made from, for example, glass, plastic (e.g., polypropylene, nylon), polyacrylamide, nitrocellulose, or other materials. Methods for attaching the nucleic acids to the surface of the solid phase include, but are not limited to, printing on glass plates (Schena et al, 1995, Science 270:467-470; DeRisi et al, 1996, Nature Genetics 14:457-460; Shalon et al., 1996, Genome Res. 6:639-645; and Schena et al., 1995, Proc. Natl. Acad. Sci. U.S.A. 93:10539-11286); or ink jet printer.
[0069] The microarrays can also be high-density oligonucleotide arrays. Techniques are known for producing arrays containing thousands of oligonucleotides complementary to defined sequences (see, Fodor et al., (1991) Science 251:767-773; Pease et al., (1994) Proc. Natl. Acad. Sci. U.S.A. 91:5022-5026; Lockhart et al., (1996) Nature Biotechnology 14:1675; U.S. Pat. Nos. 5,578,832; 5,556,752; and 5,510,270; Blanchard et al., Biosensors & Bioelectronics 11:687-690). Other methods for making microarrays may also be utilized (Maskos and Southern, (1992) Nuc. Acids. Res. 20:1679-1684; U.S. Pat. No. 6,136,592; WO 200054883; WO 200055363; WO 200053812; WO 200014273). The microarrays may be used as is or incorporated into a biochip, multiwell or other device.
[0070] Antibodies
[0071] The invention also provides antibodies which specifically bind one or more of the polypeptides of the invention. The antibodies can be monoclonal antibodies, polyclonal antibodies, antibody fragments (e.g., Fab, Fab', F(ab')2, Fv, Fc, etc.), chimeric antibodies, bispecific antibodies, heteroconjugate antibodies, single chain (ScFv), mutants thereof, fusion proteins comprising an antibody portion, humanized antibodies, and any other modified configuration of the immunoglobulin molecule that comprises an antigen recognition site of the required specificity, including glycosylation variants of antibodies, amino acid sequence variants of antibodies, and covalently modified antibodies: The antibodies may be murine, rat, human, or any other origin (including chimeric or humanized antibodies). The epitope(s) can be continuous or discontinuous. The antibodies may be made by any method known in the art and tested by the method described herein. In an alternative, antibodies may be made recombinantly and expressed using any method known in the art. In another alternative, antibodies may be made recombinantly by phage display technology. See, for example, U.S. Pat. Nos. 5,565,332; 5,580,717; 5,733,743; 6,265,150; and Winter et al., Annu. Rev. Immunol. 12:433-455 (1994). Alternatively, the phage display technology (McCafferty et al., Nature 348:552-553 (1990)) can be used to produce human antibodies and antibody fragments in vitro, from immunoglobulin variable (V) domain gene repertoires from unimmunized donors.
[0072] Yet another aspect of the invention relates to detection of metastatic disease in a subject, such as a human utilizing one or more antibodies described herein coupled to a radiologic (e.g., I125) or other imaging molecules (e.g., dyes, pigments or fluorescent molecules such as luciferase, fluoroscein or commercially available fluorescent molecules from quantum.com). The antibodies may be coupled to the radiologic or imaging molecule by methods known in the art.
[0073] Another embodiment relates to the polypeptides comprising the targeting domains described herein (e.g., FIG. 7A-7E) coupled to a moiety, such as a therapeutic moiety or a detection moiety. The moiety may be any molecule. Examples of therapeutic moieties include, but are not limited to, ricin, radioisotopes, or clotting or thrombolytic factors. Examples of a detection moiety include, but are not limited to, radioisotopes, dyes, pigments or fluorescent molecules such as luciferase, fluoroscein or commercially available fluorescent molecules from quantum.com. The polypeptide may be coupled to the radiologic or imaging molecule by methods known in the art and used to target delivery of the therapeutic or detection moiety to the liver.
[0074] Screening Methods
[0075] The methods of this invention can screen for a candidate agent that blocks, suppresses or reduces (including significantly) the binding of the targeting domains. Exemplary types of agents that may be screened for ability to inhibit one or more of the targeting domains described herein include, but are not limited to, antibodies, an anti-sense molecule directed to one or more polynucleotide sequences encoding the targeting domain, an NGF inhibitory compound, a structural analog, a dominant-negative mutation, immunoadhesin, small molecules having a molecular weight of 100 to 20,000 daltons, 500 to 15,000 daltons, or 1000 to 10,000 daltons. Libraries of small molecules are commercially available.
[0076] In many situations, different primary tumors metastasize to different organs. Another embodiment of the present invention is directed to libraries of particular primary tumors made, and to the methods disclosed herein that allow for the identification of genes specific for the site of each secondary metastasis. Conversely, since primary tumors of specific organs appear to express specific homing genes RT-PCR analysis of blood samples will allow the identification of such expression before clinical symptoms of the primary tumor present themselves. Thus the present method may allow the diagnosis of subclinical tumor genesis.
[0077] Methods of Treatment
[0078] A further aspect of the invention comprises methods of treating metastasis in a subject in need of such treatment. In some embodiments the method comprises administering to a subject in need of such treatment a targeting domain linked to a therapeutic agent in an amount effective to treat the metastasis, or an effective amount of a composition that inhibits the metastasis (e.g., collection of phage or phage expression products identified by the method herein; a targeting domain linked to a therapeutic agent and/or an antibody directed against a polypeptide comprising a targeting domain).
[0079] Yet another aspect of the invention relates to kits and compositions comprising the polynucleotides, polypeptides, antibodies or couple moieties described herein.
[0080] Methods of Inhibiting Gene Expression
[0081] The identification of genes that allow primary tumors to specifically target distant organs as sites of secondary metastasis provides new therapeutic methods of treatment. For example, the ability of tumor cells to induce transcription of an identified gene may be altered. Methods for down regulating genes are well known. It has been shown that antisense RNA introduced into a cell will bind to a complementary mRNA and thus inhibit the translation of that molecule. In a similar manner, antisense single stranded cDNA may be introduced into a cell with the same result. Further, co-suppression of genes by homologous transgenes may be effected because the ectopically integrated sequences impair the expression of the endogenous genes (Cogoni et al. Antonie van Leeuwenhoek, 1994; 65(3):205-9), and may also result in the transcription of antisense RNA (Hamada, W. and Spanu, P D; Mol. Gen. Genet. 1998). Methods of using short interfering RNA (RNAi) to specifically inhibit gene expression in eukaryotic cells have recently been described. Sec Tuschl et al., Nature 411:494-498 (2001). In all of the above methods, transfection of cells can be effected using adeno-viral or other viral vectors
[0082] In addition, stable triple-helical structures can be formed by bonding of oligodeoxyribonucleotides (ODNs) to polypurine tracts of double stranded DNA. (See, for example, Rininsland, Proc. Nat'l Acad. Sci. USA 94:5854-5859 (1997). Triplex formation can inhibit DNA replication by inhibition of transcription of elongation and is a very stable molecule.
[0083] Methods to Inhibit the Activity of Specific Proteins
[0084] While the present invention can be used to identify genes responsible for the homing and targeting of secondary tumors, the invention also recognized that it will allow the identification of the protein responsible for these phenomena. Thus, it is conceived that the present invention, by identifying that protein, will allow means of affecting gene products at the secondary metastatic site. Specifically, the site of metastasis may be targeted due to a surface protein found in the vascular walls of the endothelium at that site. It will then be possible to affect the expression of that gene down regulating it such that the metastatic tumor cells are not able to enter the endothelium and consequently will be unable to promote secondary tumor growth.
[0085] When a specific protein has been implicated in the metastatic ability of primary tumor cells its activity can be altered by several methods. First, specific antibodies may be used to bind the target protein thereby blocking its ability to attract secondary metastasis. In addition, antibodies against the homing protein may be used with a similar result. Such antibodies may be used to bind the protein thereby blocking its activity. Specific antibodies may be obtained though the use of conventional hybridoma technology or may be isolated from libraries commercially available from Dyax (Cambridge, Mass.), MorphoSys (Martinsried, Germany), Biosite (San Diego, Calif.) and Cambridge Antibody Technology (Cambridge, UK). In addition, identified proteins may act as cellular receptors. Identification of such receptors will allow the design of specific ligand antagonists which may affect the metastasis by either 1) binding to the receptor on the metastasizing tumor cell or 2) binding to the target of the metastatic cell in the vasculature at the site of the secondary tumor.
[0086] In addition, identification of metastatic proteins also allow for the design of drugs to specifically target both the primary tumor and the secondary tumor. For example, a protein on the surface of a tumor cell that allows it to home-in at a site of secondary metastasis will also allow the design of drugs that bind to that protein at the site of the primary tumor, as well as, to tumor cells which are immobilized in the blood. Similarly, identification of such proteins will allow the design of a drug or agent having an epitope similar to the identified gene product allowing the drug to home-in at the site of the metastases. Thus specific targeting of the primary and secondary tumors may be effected.
[0087] In addition, since the invention described herein allows for the identification of genes responsible for metastatic potential of primary tumors, another embodiment of the invention is directed to kits containing primers specific for those genes. Because metastatic tumor cells travel in the blood stream, use of such kits will only necessitate the drawing of blood from a patient and the use of PCR to perforin RT-PCR to identify clinically the presence of a tumor, as well as, its metastatic potential.
[0088] The following examples illustrate embodiments of the invention, but should not be viewed as limiting the scope of the invention.
EXAMPLES
Example 1
In Vivo Phage Display Identification of Metastatic Cancer Genes
Materials and Methods
[0089] Tumor cell lines: Colon cancer cell line (LS174T; American tissue culture collection, ATCC, Rockville, Md.), Melanoma cell line (1205LU; a gift from M. Heerlyn, Wistar Institute, Philadelphia).
[0090] Animal: Athymic nude mice were used for the in vivo selection studies.
[0091] Generation of cDNA phage libraries: cDNA libraries of the cancer cell lines LS174T and 1205LU were constructed by using the Orient Express directional random primer strategy (Novagen, Inc.; Darmstadt, DRG). The cDNA were then inserted into a T7Select vector using T7Select 1-1 (up to 1200 amino acids and approximately 1 copy/phage). The cDNA was inserted into the gene of capsid protein 10 and the fusion protein expressed on the surface of the phage capsid (FIG. 1). Using 300 nucleotides as the minimum size of cDNA to be inserted into each phage a library having a diversity of 1-4×106 was obtained. Once the vectors, which contain the inserts from the cDNA library, were prepared they were packaged into the T-7 phage and amplified in E. coli strain BL21 in preparation for biopanning.
[0092] In vivo phage display selection: 100 μl (108) of stock phage library was intravenously injected into mice (inferior vena cava for LS174T and portal vein for 1205LU) (FIGS. 2A and B). After a circulation time of 5 minutes mice were perfused via the heart and through the inferior vena cava until the perforate was clear of blood. Liver, lung, kidney and brain were extracted and stored at -80 degrees centigrade. The organ of metastasis (lung or liver) was used to measure the phage titer which was then amplified in E. coli in preparation for the subsequent round of biopanning. A total of three to four rounds of biopanning were conducted. The organ-selected library obtained on completion, was used to randomly select plaques for sequencing. Individual clones selected from the target organ, were isolated, amplified and intravenously injected into mice to determine their degree of organ selectivity.
[0093] FIG. 2C illustrates the rise in phage titer measured as a percent of phage injected, and quantified from the liver. After the 1st round of biopanning, phage titer retained in the liver was only 0.03% and this rose by almost three logarithms (orders of magnitude) by the third round. By the fourth round of biopanning (FIG. 2D), 97% of the total number of retained phage from the third round, were retained in the liver, compared to just over 2% in the lungs and well under 1% in both the kidneys and brain (FIG. 2D). Alternatively, 76% of the total number of phage injected from the third round, were retained in the liver, indicating that after four rounds of biopanning, clones were selected which predominantly favored the liver.
[0094] Identification of Clones: The clones selected by 4 rounds of biopanning were plated. Sixty plaques were selected, amplified by PCR using primers from T7 and the nucleotide sequence determined. The number of clones sequenced depended upon the degeneracy of the library with respect to each clone. The sequences were then analyzed and the identity of the gene obtained by using the BLAST (n) program. Translated sequences started at the 5' EcoR1 site (GAAT TC) at the 5' junction between the T-7 select vector and the tumor cell gene. Any one of the three frames in which translation occurred was used as the authentic frame for translation. In the case of known genes, all three frames were run through the BLAST (p) program to determine the correct frame for translation. For unknown genes, only frame translations, which were twenty amino acids or longer were used. The obtained amino acid sequences from various clones were grouped and analyzed by the CLUSTALW program (for multiple sequence alignment) in search of regions of homology among multiple clones.
[0095] Twenty five distinct clones were identified. Of the twenty five identified, seven were of unknown identity, fifteen were known to be either nuclear or cytoplasmic proteins, and one was associated with the cell membrane. Very surprisingly, none of the proteins identified were known to traverse the cell membrane. Of the twenty five different clones, LS42 was the most abundant being repeated seventeen times. A BLASTp search of the 151 amino acid insert shows that this peptide completely matched PA28 alpha subunit or IGUP 1-5111 from position 99 to 248. In the full length protein the first twenty four amino acids are indicated as the molecule's signal peptide allowing for secretion from the cytoplasm. Other than exhibiting 29% identity with β myosin heavy chain, this protein does not appear to be a member of a known family of proteins. However, it bears the cell adhesion motif RGD tripeptide. These results are both surprising and unexpected as a role for cell adhesion by PA28alpha subunit has never been reported.
[0096] Immunohistochemistry
[0097] To demonstrate that the retention of clones in the liver was not due to non-specific trapping but to direct binding to vascular cells, liver sections were probed with a T7 tag antibody and detected phage by immunohistochemistry. Localization of bacteriophage injected into mice was determined by immunohistochemistry analysis of brain, lung, liver and kidney tissue sections. After mice had been injected with bacteriophage and subsequently perfused, brain, lung, kidney and liver were removed and placed in 10% formaldehyde for 1-2 hours. Organs were then placed in 70% ethyl alcohol for at least two hours. Organs were embedded and sectioned and immunohistochemistry of tissue sections on glass slides was carried out. Briefly, embedding medium and formaldehyde were removed by pre-heating slides overnight at 55° C., followed by multiple treatments with xylene and ethanol. Sections were washed, blocked with 10% horse serum and after several more washes with phosphate buffered saline, (PBS), incubated with the primary antibody overnight. The following day sections were washed with PBS after which the biotinylated second antibody was added. Positive reactions were detected with avidin-biotin complex followed by incubation with DAB solution. Positive staining appeared as dark brown.
[0098] Northern Blot Analysis
[0099] Total RNA from cell lines was isolated with the RNA STAT-60 method (Tel-test, Friendswood, Tex.). RNA was separated and blotted as previously described (Fang et al., JBC, 1992, 267:25889-97). Blots were hybridized, washed and autoradiographed for 48 hrs with cDNA complementary to the gene which encodes for the 151 amino acid expression product for PA28a subunit. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as a loading control.
[0100] In Situ Hybridization
[0101] To demonstrate that clones thus far identified in the mouse are significant in humans, phage inserts were used to prepare probes which were then used to probe human tissue arrays of both normal and cancer tissues. These tissue samples included breast, prostate, colon, brain and lymphatic system and both primary and metastatic tissue. In situ hybridization's were carried out as previously described (Stiletto et al., 2000). Briefly, deparaffinized sections of fomalin-fixed tissues were treated at 37° C. for 10 minutes with proteinase K and then washed twice with SSC. Slides were incubated overnight with respective oligonucleotides, in hybridization solution (50% formamide, 4×SSC, 1×Denhardt's solution, 5 mg/ml heat denatured salmon sperm DNA, 2.5 mg/ml yeast tRNA, 10% dextran sulfate). Slides were washed with 2×SSC for 30 minutes at room temperature, with 2×SSC/formamide at 50° C. and with 1×SSC at room temperature for five minutes. Anti-digoxigenin-alkaline phosphatase conjugate was used for immunological detection of bound probes. In the breast cancer tissues, the results showed that the gene was strongly expressed in three cases, medium expression in four cases and little or no expression in five cases. Out of the six clones tested three, PA28a, Epsilon tubule chain, and CAT-292E10, showed positive staining in both the primary and metastatic tumor tissue, leaving the surrounding non cancerous tissue unstained.
[0102] Organ Homing by Individual Clones
[0103] After identification of the first twenty five clones from the liver-selective library, it was determine whether individual clones were capable of favoring the liver after being intravenously injected into mice. The nine most abundant clones were individually amplified and separately injected into no-tumor bearing mice (FIG. 4). The nucleic acid sequences for the nine clones are provided in FIG. 6A-C. Translations of the three reading frames for all nine clones is provided in FIG. 7A-E. To minimize the possibility that clones were retained in the liver because of direct blood flow from portal vein injections, a selection of clones were also injected, in separate mice, through the inferior vena cava, and organ distribution of retained phage estimated. FIG. 2D illustrates the organ distribution of phage injected into mice via the inferior vena cava or portal vein. To standardize the phage numbers among different experiments, phage titers were represented as a proportion of the kidney titers within each experiment. Phage titers in the kidneys were selected for comparison since they were not in the direct circulatory pathway of phage injected either via the inferior vena cava or portal vein. This new number was then used to calculate the liver to lung ratio which was finally used as a measure of liver selectivity. A ratio of one indicates that the clone was distributed equally between the lungs and liver. Injection of the control, wild type T7 phage via the inferior vena cava resulted in a liver to lung ratio of just over one. A slight preference for the liver (two fold) was seen when the route of injection was the portal vein (FIG. 3A).
[0104] Despite greater liver to lung ratios when the route of delivery was the portal vein as compared to the inferior vena cava, there was a clear preference for the liver in the clones tested (at least fourteen fold), in comparison to the numbers seen with the control, empty T7 phage (FIG. 5). While some of the repeat experiments for individual clones showed variable numbers (e.g. 29 fold and 270 fold for PA28a), all ratios were well above those seen for control phage. Thus, the clones selected for intravenous injection are mostly liver-selective. As further confirmation of the selectivity of these clones, the only clone injected individually whose sequence was outside the open reading frame, JN42, displayed a very weak level of selectivity (two fold), comparable to that of the control phage.
[0105] To demonstrate that the method described herein is generally applicable to all possible metastatic tissues, a cell line known to metastasize to organs besides the liver was tested. One of the tumor cell lines used was the human melanoma cell line 1205LU which predominantly metastasizes to the lungs. Using the same principal as that previously described for LS174T, a cDNA library was generated from 1205LU which was then spliced into the T7 phage. After biopanning this library in mice, individual clones were tested for their ability to preferentially home to the lungs (FIG. 3B). In spite of skepticism in the literature about being able to show selection in the lungs due to their high perfusion, a preference of the RFC2 clone for the lugs over the liver by 2.9 fold was shown (see Pasqualini and Ruoslahti Nature 38:1996, 364-366).
[0106] Identification of genes that predict and potentially drive metastasis in patients with gastrointestinal carcinoma can be a key concern for diagnostics and therapy.
[0107] Patients with different cancers as well as a secreted fibroblast growth factor binding protein (FGF-BP) that is upregulated early in the progression of colon cancer, i.e. dysplasia. We report the mRNA expression of five of the novel metastasis genes from phage display as well as PTN and FGFBP using a series of 39 tissue microarrays representing cancers of the pancreas (n=106), ampulla (n=54), bile duct (n=40), colon (n=37) and liver metastases from colon cancers (n=35). Each of these tumors was represented by several cores on the arrays (mean 4.8±2.6) and 708 cores of 22 different reference tissues were used as controls. Staining for mRNA was performed by in situ hybridization (ISH) with digoxigenin-labeled antisense mRNA probes.
[0108] Corresponding controls were performed with sense probes. Staining was evaluated without prior knowledge of the clinical data. Each core was classified according to staining intensity and frequency and tumor cases were classified by percentage of positively staining cancer cells into 6 groups from negative to highly positive.
[0109] Results show a distinct frequency and intensity of gene expression in most of the primary lesions (56.2%-92.2%) and very high expression in the liver metastases (69.09-100%). Expression of all of the genes was low in the pancreas non-adeno-carcinoma (25.8-27.3%) and in the according non-neoplastic reference tissues (0%-25%).
[0110] A subsequent comparison of patient cases with and without known metastasis showed highly significant increases in expression levels and frequency of the phage-display derived metastasis genes and of PTN for the cases with metastasis (FIG. 8). Tumors with known metastasis typically showed >75% positive tumor cells. Tumors with low or now expression (<25%) were typically without metastasis (all p-values <0.05). No significant correlations were found for FGF-BP. We conclude that genes discovered by phage display and PTN can serve to distinguish amongst GI cancers with different stage and outcome.
[0111] Other embodiments and uses of the invention will be apparent to those skilled in the art from consideration of the specification and practice of the invention disclosed herein. All references cited herein, including all publications, U.S. and foreign patents and patent applications, are specifically and entirely incorporated by reference. It is intended that the specification and examples be considered exemplary only with the true scope and spirit of the invention indicated by the following claims.
Sequence CWU
1
361497DNAHomo sapiensmisc_feature(24)..(24)n = A,T,C or G 1gaattcaagc
aaagtattta tctngactcg ccacactcca cgggaaagca atatgaaatg 60atctgctgca
gtgctctgag ccctaggatt catctttctt ttcaccgtag gtggcctgac 120tggcattgta
ttagcaaact catcactaga catcgtacta cacgacacgt actacgttgt 180agctcacttc
cactatgtcc tatcaatagg agctgtattt gccatcatag gaggcttcat 240tcattgattt
cccctattct caggctacac cctagaccaa acctacgcca aaatccattt 300cactatcata
ttcatcggcg taaatctaac tttcttccca caacactttc tcggcctaac 360cggaatgccc
cgacgttact cggactaccc cgatgcatac accacatgaa acatcctatc 420atctgtaggc
tcaagcttgc ggccgcactc gagtaactag tttacccctt ggggcctcta 480aacgggtctt
gaggggt 4972711DNAHomo
sapiensmisc_feature(1)..(711)n = A,T,C or G 2gaattcaagc gcacctttca
gaagctacac tagcaggaaa aaattccatc aagcaattnc 60attagtaatt tncnataatt
ncacaaaaga tncttgatct tacttgaagt atacatgagg 120ggaaagagcc ccctcagcag
gtgttcccgt tgcttacaga agcaaactaa aggacctaaa 180actggaggca agccaggatg
ccaaaaaggg ggaagagaaa tgataaagaa ccattcataa 240attccatgtc tacttcaaga
catttgtcta atgaccctta cataataagt attttaggga 300aaactaccac ccttttaaga
taaaagtaca atcttaaaag ctgtagttct caattatagt 360aatatttctt acttccagta
atatgtctca ataccttgga ctgctggatg tcaaaagaca 420atacctgggg gtcatctntg
agatctgaac aaatagagga attctctagg actgtatact 480ctctattttg gctttttgaa
tgaagtacag acaggcttct ctgctatcct ccaggcagtg 540taatagtcaa ggaaaagggc
aacagttttg gatcattcct tagacactaa tcagctgggg 600aaagagttca ttggnaaaag
tgtcctccca agaatggttt acaccaagca gagaggacat 660gtcactgaaa tggggaaagg
gaaacccccg tttccacagt cactgttagc a 7113532DNAHomo
sapiensmisc_feature(1)..(532)n = A,T,C or G 3gaattcangc gaaggatatg
canaagngat gtccacaaga gttcattgan cgctgaaatg 60aaactctttg ctcaacaatg
caaggaggta cnacatcctt atcatcacag cacttattcc 120aggtaaaaaa nctccanttt
tatttaatan nnnaatgant gngtcactga aggaaggttc 180agttgttnnn nctttacctg
cngangcngg cgnaaacntt gatancncct tggggcgaca 240anncnggtnt cagaggggna
attactcaca tcggcgagag gngagctggc cacggggggg 300gncactcagg ccngcaccct
ngattccaac nncatcaccn nacnnntgaa ggccatcanc 360ccngacaaag atacttttta
ttttgatgtg aaagangact ttcncnttgg tacnatnggt 420nntttccatn tcggnnctnn
ngngntnnaa ggtggtaaag gaanttttcc cccngncntn 480cnggnncccg tgnnccccac
cgtccngnnc ttcgnccgcc ccactnctnt nc 5324743DNAHomo
sapiensmisc_feature(1)..(743)n = A,T,C or G 4gaattcaagc anactttggg
gaaaaggagg ttcttaaaat cagtgtttcc cctttgtgca 60cttgtagaaa aaaaagaaan
accttctaga gctgatttga tggacaatgg ananagcnnn 120ccctgtgnnt atnataangg
aagctagctg ctctncggtc acctttgcnt agnannatac 180tttaacctgg cttttacagn
agtagtaact gccctccaac cgtcttaann gnnaatntcg 240gagccnattg cgcngtgntc
cacctacggc naatatttnc nccnaggagg atggntttcc 300cngccagtan tnccttngcn
ttnaacctca cgtgaccttc ttangcnatt cncgctcgcc 360gcaagangtc tttgttnttc
ccttctcgca ctccttntnt ntctnngngc cgtgncgncc 420ncttccttnc gctgaccngg
ctcgnnnctn nttgcncntt cagggngctc ttnccaagct 480cctcngggnt nntgcatttt
ttncncccng nntgncngcc ccnccgcccn gcncctgntt 540cagccttaca cttcnggcan
cggcctacan ggggataaan canncatttg tcncgggcgt 600ttacntnctc ccgtcccacc
atctnngcca tnttcnccnn gggnngtnct tttnctacct 660cccccccccn cncnctncan
tcntttaccn gttcgcgctc ctctntgcgt tcgngccncc 720ncgtcgcnct tttncnncnc
ttt 7435714DNAHomo
sapiensmisc_feature(1)..(714)n = A,T,C or G 5gaattcaagc ggaacgctca
cggactgtgt ggtaatgaga gatccaaaca ccaagcgctc 60caggggcttt gggtttgtca
catatgccac tgtggaggag gtggatgcag ctatgaatgc 120aaggccacac aaggtggatg
gaagagttgt ggaaccaaag agagctgtct ccagagaaga 180ttctcaaaga ccaggtgccc
acttaactgt gaaaaagata tttgttggtg gcattaaaga 240agacactgaa gaacatcacc
taagagatta ttttgaacag tatggaaaaa ttgaagtgat 300tgaaatcatg actgaccgag
gcagtggcaa gaaaaggggc tttgcctttg taacctttga 360cgaccatgac tccgtggata
agattgtcat tcagaaatac catactgtga atggccacaa 420ctgtgaagtt agaaaagccc
tgtcaaagca agagatggct agtgcttcat ccaaccaaag 480aggtcgaagt ggttctggaa
actttggtgg tggtcgtgga ggtggtttca gtgggaatga 540caacttcngt cctggaggaa
acttnnantg gtcctggtgg ctttggtggc aaccgtggtg 600gtggtggata tggtggcaat
ggggatggct ataatggatt tggtatgatg gaancaattt 660tggnggtggt ggaagctaca
ntgattttgg gaantaaaac aatcaatcct caaa 7146294DNAHomo
sapiensmisc_feature(1)..(294)n = A,T,C or G 6aaattcangc gttatcgtcc
tttcttccat tcttaacagt atgtgcccat ttgcaaaaca 60aaaatgctaa taatcagtaa
tagtcctata aaagatgtta actctgttta gtcattgact 120gatcttgctc taaccttaaa
attttgtgat tattgacctc tgttgcattt attctaaagc 180cccccgaagc ttgcggccgc
actcgagtaa ctagttaacc ccttggggcc tctaaacggg 240tcttgagggg taacttggnt
cctcgngggn ggnggcangc ttcggggggg tttg 2947705DNAHomo
sapiensmisc_feature(1)..(705)n = A,T,C or G 7gaattcaagc aagacaagga
tgaaaagaag aagggggagg atgaagacaa aggtcctccc 60tgtggcccag tgaactgcaa
tgaaaagatc gtggtccttc tgcagcgctt gaagcctgag 120atcaaggatg tcattgagca
gctcaacctg gtcaccacct ggttgcagct gcagatacct 180cggattgagg atggtaacaa
ttttggagtg gctgtccagg agaaggtgtt tgagctgatg 240accagcctcc acaccaagct
agaaggcttc cacactcaaa tctctaagta tttctctgag 300cgtggtgatg cagtgactaa
agcagccaag cagccccatg tgggtgatta tcggcagctg 360gtgcacgagc tggatgaggc
agagtaccgg gacatccggc tgatggtcat ggagatccgc 420aatgcttatg ctgtgttata
tgacatcatc ctgaagaact tcgagaagct caagaagccc 480aggggagaaa caaagggaat
gatctattga gagccctctc tcccattctg tgatgagtac 540agcaganacc ttcctgcttt
ttantgggga cccanatttt ccccaaactt gctctgttga 600gatttttccc tcaccttgcc
tctcangcac aataaatata nttataccac tgccaaagct 660tgcgggcgca ctccantaac
tagttaaccc cttggggcct ctaaa 7058657DNAHomo
sapiensmisc_feature(1)..(657)n = A,T,C or G 8gaattctcaa aatagttcag
aaaaatcatg tttcaaagta ctcacattct tccagatgag 60gaaaaaatgg tgaaggaaag
aaaaaggaaa ttgaaagaag tattaatcca aactttcaaa 120gaaaatcaac agtgtcaaaa
acggtatttc gctgcctggc acaagctgat tcttgatcat 180aggattaagc tggggaaagc
tgggaccctg tctgactgga agattcagct gaaggtcctg 240cgggcctgga gagactacac
aagattccag aagttggagc gggagactca agccttggaa 300aatgatctta gggaagaaaa
cagaaaacaa caactggcca ctgagtataa ccggaaacaa 360gttctccgac actgctttac
agaatggcag cattggcatg gcgccgagct cctgaagaga 420gagctggctc tcacaaaaga
ggaaactang aagaagatgg ntgcactgct gcaggcagca 480tcactgggga aactcagtgc
cantgggtta tcangcntca gtcnacctga gganggaaca 540gccntggtgg gtncnccant
naanaatggn cagganacng cnntgccccc ntttgtggga 600aaagcccccc ttgggnagca
ntgggtgnnt ntcnntcccc cccngggaag nacaana 6579420DNAHomo
sapiensmisc_feature(206)..(206)n = A,T,C or G 9gaattctgga aagttgggta
caactgtgaa gccaaagagt ctggttactt caagttctgg 60ggctttaaaa aagcagcata
agaagccctt tgatgcaatg aataacattg tggcaaattt 120gctcctcaac ctaacgaggg
aagcttgcgg ccgcactcga gtaactagtt aaccccttgg 180ggcctctaaa cgggtcttga
ggggtntact aagttactcg agtgcggccg caagcttccc 240tcgttaggtt gaggagcaaa
tttgccacaa tgttattcat tgcatcaaag ggcttcttat 300gctgcttttt taaagcccca
gaacttgaag taaccagact ctttggcttc acagttgtac 360ccaactttcc agaattcgga
tccccgagca tcacacctga ctggaatacg acagctncaa 42010160PRTHomo
sapiensVARIANT(1)..(1)Xaa = Any Amino Acid 10Xaa Asn Ser Ser Lys Val Phe
Ile Xaa Thr Arg His Thr Pro Arg Glu1 5 10
15Ser Asn Met Lys Ser Ala Ala Val Leu Ala Leu Gly Phe
Ile Phe Leu 20 25 30Phe Thr
Val Gly Gly Leu Thr Gly Ile Val Leu Ala Asn Ser Ser Leu 35
40 45Asp Ile Val Leu His Asp Thr Tyr Tyr Val
Val Ala His Phe His Tyr 50 55 60Val
Leu Ser Ile Gly Ala Val Phe Ala Ile Ile Gly Gly Phe Ile His65
70 75 80Phe Pro Leu Phe Ser Gly
Tyr Thr Leu Asp Gln Thr Tyr Ala Lys Ile 85
90 95His Phe Thr Ile Ile Phe Ile Gly Val Asn Leu Thr
Phe Phe Pro Gln 100 105 110His
Phe Leu Gly Leu Thr Gly Met Pro Arg Arg Tyr Ser Asp Tyr Pro 115
120 125Asp Ala Tyr Thr Thr Asn Ile Leu Ser
Ser Val Gly Ser Ser Leu Arg 130 135
140Pro His Ser Ser Asn Phe Thr Pro Trp Gly Leu Thr Gly Leu Glu Gly145
150 155 16011152PRTHomo
sapiensVARIANT(1)..(1)Xaa = Any Amino Acid 11Xaa Ile Gln Ala Lys Tyr Leu
Ser Xaa Leu Ala Thr Leu His Gly Lys1 5 10
15Ala Ile Asn Asp Leu Leu Gln Cys Ser Glu Pro Asp Ser
Ser Phe Phe 20 25 30Ser Pro
Val Ala Leu Ala Leu Tyr Gln Thr His His Thr Ser Tyr Tyr 35
40 45Thr Thr Arg Thr Thr Leu Leu Thr Ser Thr
Met Ser Tyr Gln Glu Leu 50 55 60Tyr
Leu Pro Ser Glu Ala Ser Phe Ile Asp Phe Pro Tyr Ser Gln Ala65
70 75 80Thr Pro Thr Lys Pro Thr
Pro Lys Ser Ile Ser Leu Ser Tyr Ser Ser 85
90 95Ala Ile Leu Ser Ser His Asn Thr Phe Ser Ala Pro
Glu Cys Pro Asp 100 105 110Val
Thr Arg Thr Thr Pro Met His Thr Pro His Glu Thr Ser Tyr His 115
120 125Leu Ala Gln Ala Cys Gly Arg Thr Arg
Val Thr Ser Leu Pro Leu Gly 130 135
140Ala Ser Lys Arg Val Leu Arg Gly145 15012163PRTHomo
sapiensVARIANT(8)..(8)Xaa = Any Amino Acid 12Glu Phe Lys Gln Ser Ile Tyr
Xaa Asp Ser Pro His Ser Thr Gly Lys1 5 10
15Gln Tyr Glu Met Ile Cys Cys Ser Ala Leu Ser Pro Arg
Ile His Leu 20 25 30Ser Phe
His Arg Arg Trp Pro Asp Trp His Cys Ile Ser Lys Leu Ile 35
40 45Thr Arg His Arg Thr Thr Arg His Val Leu
Arg Cys Ser Ser Leu Pro 50 55 60Leu
Cys Pro Ile Asn Arg Ser Cys Ile Cys His His Arg Arg Leu His65
70 75 80Ser Leu Ile Ser Pro Ile
Leu Arg Leu His Pro Arg Pro Asn Leu Arg 85
90 95Gln Asn Pro Phe His Tyr His Ile His Arg Arg Lys
Ser Asn Phe Leu 100 105 110Pro
Thr Thr Leu Ser Arg Pro Asn Arg Asn Ala Pro Thr Leu Leu Gly 115
120 125Leu Pro Arg Cys Ile His His Met Lys
His Pro Ile Ile Cys Arg Leu 130 135
140Lys Leu Ala Ala Ala Leu Glu Leu Val Tyr Pro Leu Gly Pro Leu Asn145
150 155 160Gly Ser
Gly13224PRTHomo sapiensVARIANT(1)..(224)Xaa = Any Amino Acid 13Xaa Asn
Ser Ser Ala Pro Phe Arg Ser Tyr Thr Ser Arg Lys Lys Phe1 5
10 15His Gln Ala Ile Xaa Leu Val Ile
Xaa Xaa Asn Xaa Thr Lys Asp Xaa 20 25
30Ser Tyr Leu Lys Tyr Thr Gly Glu Arg Ala Pro Ser Ala Gly Val
Pro 35 40 45Val Ala Tyr Arg Ser
Lys Leu Lys Asp Leu Lys Leu Glu Ala Ser Gln 50 55
60Asp Ala Lys Lys Gly Glu Glu Lys Arg Thr Ile His Lys Phe
His Val65 70 75 80Tyr
Phe Lys Thr Phe Val Pro Leu His Asn Lys Tyr Phe Arg Glu Asn
85 90 95Tyr His Pro Phe Lys Ile Lys
Val Gln Ser Lys Leu Phe Ser Ile Ile 100 105
110Val Ile Phe Leu Thr Ser Ser Asn Met Ser Gln Tyr Leu Gly
Leu Leu 115 120 125Asp Val Lys Arg
Gln Tyr Leu Gly Val Ile Xaa Glu Ile Thr Asn Arg 130
135 140Gly Ile Leu Asp Cys Ile Leu Ser Ile Leu Ala Phe
Met Lys Tyr Arg145 150 155
160Gln Ala Ser Leu Leu Ser Ser Arg Gln Cys Asn Ser Gln Gly Lys Gly
165 170 175Gln Gln Phe Trp Ile
Ile Pro Thr Leu Ile Ser Trp Gly Lys Ser Ser 180
185 190Leu Xaa Lys Val Ser Ser Gln Glu Trp Phe Thr Pro
Ser Arg Glu Asp 195 200 205Met Ser
Leu Lys Trp Gly Lys Gly Asn Pro Arg Phe His Ser His Cys 210
215 22014226PRTHomo sapiensVARIANT(1)..(226)Xaa =
Any Amino Acid 14Xaa Ile Gln Ala His Leu Ser Glu Ala Thr Leu Ala Gly Lys
Asn Ser1 5 10 15Ile Lys
Gln Xaa His Phe Xaa Ile Ile Xaa Gln Lys Xaa Leu Asp Leu 20
25 30Thr Ser Ile His Glu Gly Lys Glu Pro
Pro Gln Gln Val Phe Pro Leu 35 40
45Leu Thr Glu Ala Asn Arg Thr Asn Trp Arg Gln Ala Arg Met Pro Lys 50
55 60Arg Gly Lys Arg Asn Asp Lys Glu Pro
Phe Ile Asn Ser Met Ser Thr65 70 75
80Ser Arg His Leu Ser Asn Asp Pro Tyr Ile Ile Ser Ile Leu
Gly Lys 85 90 95Thr Thr
Thr Leu Leu Arg Lys Tyr Asn Leu Lys Ser Cys Ser Ser Gln 100
105 110Leu Tyr Phe Leu Leu Pro Val Ile Cys
Leu Asn Thr Leu Asp Cys Trp 115 120
125Met Ser Lys Asp Asn Thr Trp Gly Ser Ser Xaa Arg Ser Glu Gln Ile
130 135 140Glu Glu Phe Ser Arg Thr Val
Tyr Ser Leu Phe Trp Leu Phe Glu Ser145 150
155 160Thr Asp Arg Leu Leu Cys Tyr Pro Pro Gly Ser Val
Ile Val Lys Glu 165 170
175Lys Gly Asn Ser Phe Gly Ser Phe Leu Arg His Ser Ala Gly Glu Arg
180 185 190Val His Trp Xaa Lys Cys
Pro Pro Lys Asn Gly Leu His Gln Ala Glu 195 200
205Arg Thr Cys His Asn Gly Glu Arg Glu Thr Pro Val Ser Thr
Val Thr 210 215 220Val
Ser22515225PRTHomo sapiensVARIANT(1)..(225)Xaa = Any Amino Acid 15Glu Phe
Lys Arg Thr Phe Gln Lys Leu His Gln Glu Lys Ile Pro Ser1 5
10 15Ser Asn Xaa Ile Ser Asn Xaa Xaa
Xaa His Lys Arg Xaa Leu Ile Leu 20 25
30Leu Glu Val Tyr Met Arg Gly Lys Ser Pro Leu Ser Arg Cys Ser
Arg 35 40 45Cys Leu Gln Lys Gln
Thr Lys Gly Pro Lys Thr Gly Gly Lys Pro Gly 50 55
60Cys Gln Lys Gly Gly Arg Glu Met Ile Lys Asn His Ser Ile
Pro Cys65 70 75 80Leu
Leu Gln Asp Ile Cys Leu Met Thr Leu Thr Val Phe Gly Lys Leu
85 90 95Pro Pro Phe Asp Lys Ser Thr
Ile Leu Lys Ala Val Val Leu Asn Tyr 100 105
110Ser Asn Ile Ser Tyr Phe Gln Tyr Val Ser Ile Pro Trp Thr
Ala Gly 115 120 125Cys Gln Lys Thr
Ile Pro Gly Gly His Xaa Asp Leu Asn Lys Arg Asn 130
135 140Ser Leu Gly Leu Tyr Thr Leu Tyr Phe Gly Phe Leu
Asn Glu Val Gln145 150 155
160Thr Gly Phe Ser Ala Ile Leu Gln Ala Val Ser Arg Lys Arg Ala Thr
165 170 175Val Leu Asp His Ser
Leu Asp Thr Asn Gln Leu Gly Lys Glu Phe Ile 180
185 190Xaa Lys Ser Val Leu Pro Arg Met Val Tyr Thr Lys
Gln Arg Gly His 195 200 205Val Thr
Glu Met Gly Lys Gly Lys Pro Pro Phe Pro Gln Ser Leu Leu 210
215 220Ala22516174PRTHomo
sapiensVARIANT(1)..(174)Xaa = Any Amino Acid 16Xaa Asn Ser Xaa Glu Gly
Tyr Ala Xaa Xaa Met Ser Thr Arg Val His1 5
10 15Xaa Leu Lys Asn Ser Leu Leu Asn Asn Ala Arg Arg
Tyr Xaa Ile Leu 20 25 30Ile
Ile Thr Ala Leu Ile Pro Gly Lys Lys Xaa Pro Xaa Leu Phe Asn 35
40 45Xaa Xaa Met Xaa Xaa Ser Leu Lys Glu
Gly Ser Val Val Xaa Xaa Leu 50 55
60Pro Xaa Xaa Xaa Gly Xaa Asn Xaa Asp Xaa Xaa Leu Gly Arg Gln Xaa65
70 75 80Xaa Xaa Gln Arg Xaa
Asn Tyr Ser His Arg Arg Glu Xaa Ser Trp Pro 85
90 95Arg Gly Gly Xaa Leu Arg Xaa Ala Pro Xaa Ile
Pro Xaa Xaa Ser Xaa 100 105
110Xaa Xaa Arg Pro Ser Xaa Xaa Thr Lys Ile Leu Phe Ile Leu Met Lys
115 120 125Xaa Thr Phe Xaa Leu Val Xaa
Xaa Xaa Xaa Ser Xaa Ser Xaa Xaa Xaa 130 135
140Xaa Xaa Lys Val Val Lys Glu Xaa Phe Pro Xaa Xaa Xaa Xaa Xaa
Pro145 150 155 160Xaa Xaa
Pro Pro Ser Xaa Xaa Ser Xaa Ala Pro Xaa Xaa Xaa 165
17017173PRTHomo sapiensVARIANT(1)..(173)Xaa = Any Amino Acid
17Xaa Ile Xaa Ala Lys Asp Met Xaa Lys Xaa Cys Pro Gln Glu Phe Ile1
5 10 15Xaa Arg Asn Glu Thr Leu
Cys Ser Thr Met Gln Gly Gly Xaa Thr Ser 20 25
30Leu Ser Ser Gln His Leu Phe Gln Val Lys Xaa Leu Xaa
Phe Tyr Leu 35 40 45Ile Xaa Xaa
Xaa Xaa His Arg Lys Val Gln Leu Xaa Xaa Leu Tyr Leu 50
55 60Xaa Xaa Xaa Ala Xaa Xaa Leu Ile Xaa Pro Trp Gly
Asp Xaa Xaa Gly65 70 75
80Xaa Arg Gly Xaa Ile Thr His Ile Gly Glu Arg Xaa Ala Gly His Gly
85 90 95Gly Xaa His Ser Gly Xaa
His Pro Xaa Phe Gln Xaa His His Xaa Xaa 100
105 110Xaa Glu Gly His Xaa Pro Xaa Gln Arg Tyr Phe Leu
Phe Cys Glu Arg 115 120 125Xaa Leu
Xaa Xaa Trp Tyr Xaa Xaa Xaa Phe Pro Xaa Arg Xaa Xaa Xaa 130
135 140Xaa Xaa Arg Trp Arg Xaa Phe Ser Pro Xaa Xaa
Xaa Xaa Xaa Arg Xaa145 150 155
160Pro His Arg Xaa Xaa Leu Xaa Pro Pro His Xaa Xaa Xaa
165 17018174PRTHomo sapiensVARIANT(1)..(174)Xaa = Any
Amino Acid 18Glu Phe Xaa Arg Arg Ile Cys Xaa Xaa Asp Val His Lys Ser Ser
Leu1 5 10 15Xaa Ala Glu
Met Lys Leu Phe Ala Gln Gln Cys Lys Glu Val Xaa His 20
25 30Pro Tyr His His Ser Thr Tyr Ser Arg Lys
Xaa Ser Xaa Phe Ile Xaa 35 40
45Xaa Asn Xaa Xaa Val Thr Glu Gly Arg Phe Ser Cys Xaa Xaa Phe Thr 50
55 60Cys Xaa Xaa Xaa Xaa Lys Xaa Xaa Xaa
Leu Gly Ala Thr Xaa Xaa Xaa65 70 75
80Ser Glu Gly Xaa Leu Leu Thr Ser Ala Arg Xaa Glu Leu Ala
Thr Gly 85 90 95Gly Xaa
Thr Gln Ala Xaa Thr Xaa Asp Ser Asn Xaa Ile Thr Xaa Xaa 100
105 110Xaa Lys Ala Ile Xaa Xaa Asp Lys Asp
Thr Phe Tyr Phe Asp Val Lys 115 120
125Xaa Asp Phe Xaa Xaa Gly Xaa Xaa Gly Xaa Phe His Xaa Xaa Xaa Xaa
130 135 140Xaa Xaa Xaa Gly Gly Lys Gly
Xaa Phe Pro Xaa Xaa Xaa Xaa Xaa Pro145 150
155 160Val Xaa Pro Thr Val Xaa Xaa Phe Xaa Arg Pro Thr
Xaa Xaa 165 17019242PRTHomo
sapiensVARIANT(1)..(242)Xaa = Any Amino Acid 19Xaa Asn Ser Ser Xaa Leu
Trp Gly Lys Gly Gly Ser Asn Gln Cys Phe1 5
10 15Pro Phe Val His Leu Lys Lys Lys Lys Xaa Leu Leu
Glu Leu Ile Trp 20 25 30Thr
Met Xaa Xaa Xaa Xaa Pro Val Xaa Xaa Ile Xaa Glu Ala Ser Cys 35
40 45Ser Xaa Val Thr Phe Xaa Xaa Xaa Thr
Leu Thr Trp Leu Leu Gln Xaa 50 55
60Leu Pro Ser Asn Arg Leu Xaa Xaa Xaa Xaa Arg Ser Xaa Leu Arg Xaa65
70 75 80Xaa Pro Pro Thr Xaa
Asn Ile Xaa Xaa Xaa Glu Asp Xaa Phe Pro Xaa 85
90 95Gln Xaa Xaa Leu Xaa Xaa Xaa Pro His Val Thr
Phe Leu Xaa Xaa Xaa 100 105
110Arg Ser Pro Gln Xaa Val Phe Val Xaa Pro Phe Ser His Ser Xaa Xaa
115 120 125Xaa Xaa Xaa Pro Xaa Xaa Xaa
Leu Pro Xaa Ala Asp Xaa Ala Xaa Xaa 130 135
140Xaa Leu Xaa Xaa Gln Xaa Ala Leu Xaa Lys Leu Leu Xaa Xaa Xaa
Ala145 150 155 160Phe Phe
Xaa Pro Xaa Xaa Xaa Ala Xaa Pro Pro Xaa Xaa Xaa Phe Ser
165 170 175Leu Thr Leu Xaa Ala Xaa Ala
Tyr Xaa Gly Ile Xaa Xaa Xaa Phe Val 180 185
190Xaa Gly Val Tyr Xaa Leu Pro Ser His His Xaa Xaa His Xaa
Xaa Xaa 195 200 205Gly Xaa Xaa Phe
Xaa Tyr Leu Pro Pro Xaa Xaa Xaa Xaa Xaa Xaa Tyr 210
215 220Xaa Phe Ala Leu Leu Xaa Ala Phe Xaa Xaa Xaa Arg
Arg Xaa Phe Xaa225 230 235
240Xaa Leu20242PRTHomo sapiensVARIANT(1)..(242)Xaa = Any Amino Acid
20Xaa Ile Gln Ala Xaa Phe Gly Glu Lys Glu Val Leu Lys Ile Ser Val1
5 10 15Ser Pro Leu Cys Thr Cys
Arg Lys Lys Arg Xaa Thr Phe Ser Phe Asp 20 25
30Gly Gln Trp Xaa Xaa Xaa Xaa Leu Xaa Xaa Xaa Xaa Lys
Leu Ala Ala 35 40 45Xaa Arg Ser
Pro Leu Xaa Xaa Xaa Ile Leu Pro Gly Phe Tyr Xaa Ser 50
55 60Ser Asn Cys Pro Pro Thr Val Leu Xaa Xaa Asn Xaa
Gly Ala Xaa Cys65 70 75
80Xaa Val Xaa His Leu Arg Xaa Ile Phe Xaa Xaa Arg Arg Met Xaa Phe
85 90 95Xaa Ala Ser Xaa Xaa Xaa
Xaa Xaa Asn Leu Thr Pro Ser Xaa Xaa Ile 100
105 110Xaa Ala Arg Arg Lys Xaa Ser Leu Xaa Phe Pro Ser
Arg Thr Pro Xaa 115 120 125Xaa Ser
Xaa Xaa Arg Xaa Xaa Xaa Phe Leu Xaa Leu Thr Xaa Leu Xaa 130
135 140Xaa Xaa Cys Xaa Phe Arg Xaa Leu Xaa Pro Ser
Ser Xaa Gly Xaa Xaa145 150 155
160His Phe Xaa Xaa Xaa Xaa Xaa Xaa Pro Xaa Arg Xaa Xaa Pro Xaa Ser
165 170 175Ala Leu His Phe
Xaa Xaa Arg Pro Thr Xaa Gly Xaa Xaa His Leu Xaa 180
185 190Arg Ala Phe Xaa Xaa Ser Arg Pro Thr Ile Xaa
Ala Xaa Phe Xaa Xaa 195 200 205Xaa
Xaa Xaa Phe Xaa Thr Ser Pro Pro Xaa Xaa Xaa Xaa Xaa Phe Thr 210
215 220Xaa Ser Arg Ser Ser Xaa Arg Ser Xaa Xaa
Xaa Val Xaa Leu Xaa Xaa225 230 235
240Xaa Phe21244PRTHomo sapiensVARIANT(1)..(244)Xaa = Any Amino
Acid 21Glu Phe Lys Xaa Thr Leu Gly Lys Arg Arg Phe Leu Lys Ser Val Phe1
5 10 15Pro Leu Cys Ala Leu
Val Glu Lys Lys Glu Xaa Pro Ser Arg Ala Asp 20
25 30Leu Met Asp Asn Gly Xaa Ser Xaa Pro Cys Xaa Tyr
Xaa Xaa Gly Ser 35 40 45Leu Leu
Xaa Gly His Leu Cys Xaa Xaa Xaa Tyr Phe Asn Leu Ala Phe 50
55 60Thr Xaa Val Val Thr Ala Leu Gln Pro Ser Xaa
Xaa Xaa Ser Glu Xaa65 70 75
80Ile Ala Xaa Xaa Ser Thr Tyr Gly Xaa Tyr Xaa Xaa Xaa Gly Gly Trp
85 90 95Xaa Ser Xaa Pro Val
Xaa Pro Xaa Xaa Xaa Thr Ser Arg Asp Leu Leu 100
105 110Xaa Xaa Phe Xaa Leu Ala Ala Arg Xaa Leu Cys Xaa
Ser Leu Leu Ala 115 120 125Leu Leu
Xaa Xaa Xaa Xaa Ala Val Xaa Xaa Xaa Ser Xaa Arg Xaa Gly 130
135 140Ser Xaa Xaa Xaa Xaa Xaa Ser Gly Xaa Ser Xaa
Gln Ala Pro Xaa Xaa145 150 155
160Xaa Cys Ile Phe Xaa Xaa Xaa Xaa Xaa Xaa Pro Xaa Ala Xaa Xaa Leu
165 170 175Xaa Gln Pro Tyr
Thr Xaa Gly Xaa Gly Leu Xaa Gly Asp Lys Xaa Xaa 180
185 190Ile Cys Xaa Gly Arg Leu Xaa Xaa Pro Val Pro
Pro Ser Xaa Pro Xaa 195 200 205Xaa
Xaa Xaa Xaa Xaa Leu Xaa Leu Pro Pro Pro Pro Xaa Xaa Xaa Xaa 210
215 220Xaa Leu Xaa Val Arg Ala Pro Xaa Cys Val
Xaa Ala Xaa Xaa Ser Xaa225 230 235
240Phe Xaa Xaa Xaa22236PRTHomo sapiensVARIANT(1)..(236)Xaa = Any
Amino Acid 22Xaa Asn Ser Ser Gly Thr Leu Thr Asp Cys Val Val Met Arg Asp
Pro1 5 10 15Asn Thr Lys
Arg Ser Arg Gly Phe Gly Phe Val Thr Tyr Ala Thr Val 20
25 30Glu Glu Val Asp Ala Ala Met Asn Ala Arg
Pro His Lys Val Asp Gly 35 40
45Arg Val Val Glu Pro Lys Arg Ala Val Ser Arg Glu Asp Ser Gln Arg 50
55 60Pro Gly Ala His Leu Thr Val Lys Lys
Ile Phe Val Gly Gly Ile Lys65 70 75
80Glu Asp Thr Glu Glu His His Leu Arg Asp Tyr Phe Glu Gln
Tyr Gly 85 90 95Lys Ile
Glu Val Ile Glu Ile Met Thr Asp Arg Gly Ser Gly Lys Lys 100
105 110Arg Gly Phe Ala Phe Val Thr Phe Asp
Asp His Asp Ser Val Asp Lys 115 120
125Ile Val Ile Gln Lys Tyr His Thr Val Asn Gly His Asn Cys Glu Val
130 135 140Arg Lys Ala Leu Ser Lys Gln
Glu Met Ala Ser Ala Ser Ser Asn Gln145 150
155 160Arg Gly Arg Ser Gly Ser Gly Asn Phe Gly Gly Gly
Arg Gly Gly Gly 165 170
175Phe Ser Gly Asn Asp Asn Phe Xaa Pro Gly Gly Asn Xaa Xaa Trp Ser
180 185 190Trp Trp Leu Trp Trp Gln
Pro Trp Trp Trp Trp Ile Trp Trp Gln Trp 195 200
205Gly Trp Leu Trp Ile Trp Tyr Asp Gly Xaa Asn Phe Xaa Gly
Gly Gly 210 215 220Ser Tyr Xaa Asp Phe
Gly Xaa Asn Asn Gln Ser Ser225 230
23523228PRTHomo sapiensVARIANT(1)..(228)Xaa = Any Amino Acid 23Xaa Ile
Gln Ala Glu Arg Ser Arg Thr Val Trp Glu Ile Gln Thr Pro1 5
10 15Ser Ala Pro Gly Ala Leu Gly Leu
Ser His Met Pro Leu Trp Arg Arg 20 25
30Trp Met Gln Leu Met Gln Gly His Thr Arg Trp Met Glu Glu Leu
Trp 35 40 45Asn Gln Arg Glu Leu
Ser Pro Glu Lys Ile Leu Lys Asp Gln Val Pro 50 55
60Thr Leu Lys Arg Tyr Leu Leu Val Ala Leu Lys Lys Thr Leu
Lys Asn65 70 75 80Ile
Thr Glu Ile Ile Leu Asn Ser Met Glu Lys Leu Lys Leu Lys Ser
85 90 95Leu Thr Glu Ala Val Ala Arg
Lys Gly Ala Leu Pro Leu Pro Leu Thr 100 105
110Thr Met Thr Pro Trp Ile Arg Leu Ser Phe Arg Asn Thr Ile
Leu Met 115 120 125Ala Thr Thr Val
Lys Leu Glu Lys Pro Cys Gln Ser Lys Arg Trp Leu 130
135 140Val Leu His Pro Thr Lys Glu Val Glu Val Val Leu
Glu Thr Leu Val145 150 155
160Val Val Val Glu Val Val Ser Val Gly Met Thr Thr Xaa Val Leu Glu
165 170 175Glu Thr Xaa Xaa Gly
Pro Gly Gly Phe Gly Gly Asn Arg Gly Gly Gly 180
185 190Gly Tyr Gly Gly Asn Gly Asp Gly Tyr Asn Gly Phe
Gly Met Met Glu 195 200 205Xaa Ile
Leu Xaa Val Val Glu Ala Thr Xaa Ile Leu Gly Xaa Lys Thr 210
215 220Ile Asn Pro Gln22524223PRTHomo
sapiensVARIANT(1)..(223)Xaa = Any Amino Acid 24Glu Phe Lys Arg Asn Ala
His Gly Leu Cys Gly Asn Glu Arg Ser Lys1 5
10 15His Gln Ala Leu Gln Gly Leu Trp Val Cys His Ile
Cys His Cys Gly 20 25 30Gly
Gly Gly Cys Ser Tyr Glu Cys Lys Ala Thr Gln Gly Gly Trp Lys 35
40 45Ser Cys Gly Thr Lys Glu Ser Cys Leu
Gln Arg Arg Phe Ser Lys Thr 50 55
60Arg Cys Pro Leu Asn Cys Glu Lys Asp Ile Cys Trp Trp His Arg Arg65
70 75 80His Arg Thr Ser Pro
Lys Arg Leu Phe Thr Val Trp Lys Asn Ser Asp 85
90 95Asn His Asp Pro Arg Gln Trp Gln Glu Lys Gly
Leu Cys Leu Cys Asn 100 105
110Leu Arg Pro Leu Arg Gly Asp Cys His Ser Glu Ile Pro Tyr Cys Glu
115 120 125Trp Pro Gln Leu Ser Lys Ser
Pro Val Lys Ala Arg Asp Gly Cys Phe 130 135
140Ile Gln Pro Lys Arg Ser Lys Trp Phe Trp Lys Leu Trp Trp Trp
Ser145 150 155 160Trp Arg
Trp Phe Gln Trp Glu Gln Leu Xaa Ser Trp Arg Lys Leu Xaa
165 170 175Xaa Val Leu Val Ala Leu Val
Ala Thr Val Val Val Val Asp Met Val 180 185
190Ala Met Gly Met Ala Ile Met Asp Leu Val Trp Xaa Gln Phe
Trp Xaa 195 200 205Trp Trp Lys Leu
Xaa Phe Trp Glu Xaa Lys Gln Ser Ile Leu Lys 210 215
2202589PRTHomo sapiensVARIANT(1)..(89)Xaa = Any Amino Acid
25Xaa Asn Ser Xaa Val Ile Val Leu Ser Ser Ile Leu Asn Ser Met Cys1
5 10 15Pro Phe Ala Lys Gln Lys
Cys Ser Val Ile Val Leu Lys Met Leu Thr 20 25
30Leu Phe Ser His Leu Ile Leu Leu Pro Asn Phe Val Ile
Ile Asp Leu 35 40 45Cys Cys Ile
Tyr Ser Lys Ala Pro Arg Ser Leu Arg Pro His Ser Ser 50
55 60Asn Leu Thr Pro Trp Gly Leu Thr Gly Leu Glu Gly
Leu Xaa Ser Ser65 70 75
80Xaa Xaa Xaa Ala Xaa Phe Gly Gly Val 852692PRTHomo
sapiensVARIANT(1)..(92)Xaa = Any Amino Acid 26Xaa Ile Xaa Ala Leu Ser Ser
Phe Leu Pro Phe Leu Thr Val Cys Ala1 5 10
15His Leu Gln Asn Lys Asn Ala Asn Asn Gln Ser Tyr Lys
Arg Cys Leu 20 25 30Cys Leu
Val Ile Asp Ser Cys Ser Asn Leu Lys Ile Leu Leu Leu Thr 35
40 45Ser Val Ala Phe Ile Leu Lys Pro Pro Glu
Ala Cys Gly Arg Thr Arg 50 55 60Val
Thr Ser Pro Leu Gly Ala Ser Lys Arg Val Leu Arg Gly Asn Leu65
70 75 80Xaa Pro Xaa Gly Xaa Xaa
Xaa Ala Ser Gly Gly Phe 85 902792PRTHomo
sapiensVARIANT(1)..(92)Xaa = Any Amino Acid 27Lys Phe Xaa Arg Tyr Arg Pro
Phe Phe His Ser Gln Tyr Val Pro Ile1 5 10
15Cys Lys Thr Lys Met Leu Ile Ile Ser Asn Ser Pro Ile
Lys Asp Val 20 25 30Asn Ser
Val Ser Leu Thr Asp Leu Ala Leu Thr Leu Lys Phe Cys Asp 35
40 45Tyr Pro Leu Leu His Leu Phe Ser Pro Pro
Lys Leu Ala Ala Ala Leu 50 55 60Glu
Leu Val Asn Pro Leu Gly Pro Leu Asn Gly Ser Gly Val Thr Trp65
70 75 80Xaa Leu Xaa Xaa Xaa Gly
Xaa Leu Arg Gly Gly Leu 85 9028225PRTHomo
sapiensVARIANT(1)..(225)Xaa = Any Amino Acid 28Xaa Asn Ser Ser Lys Thr
Arg Met Lys Arg Arg Arg Gly Arg Met Lys1 5
10 15Thr Lys Val Leu Pro Val Ala Gln Thr Ala Met Lys
Arg Ser Trp Ser 20 25 30Phe
Cys Ser Ala Ser Leu Arg Ser Arg Met Ser Leu Ser Ser Ser Thr 35
40 45Trp Ser Pro Pro Gly Cys Ser Cys Arg
Tyr Leu Gly Leu Arg Met Val 50 55
60Thr Ile Leu Glu Trp Leu Ser Arg Arg Arg Cys Leu Ser Pro Ala Ser65
70 75 80Thr Pro Ser Lys Ala
Ser Thr Leu Lys Ser Leu Ser Ile Ser Leu Ser 85
90 95Val Val Met Gln Leu Lys Gln Pro Ser Ser Pro
Met Trp Val Ile Ile 100 105
110Gly Ser Trp Cys Thr Ser Trp Met Arg Gln Ser Thr Gly Thr Ser Gly
115 120 125Trp Ser Trp Arg Ser Ala Met
Leu Met Leu Cys Tyr Met Thr Ser Ser 130 135
140Arg Thr Ser Arg Ser Ser Arg Ser Pro Gly Glu Lys Gln Arg Glu
Ser145 150 155 160Ile Glu
Ser Pro Leu Ser His Ser Val Met Ser Thr Ala Xaa Thr Phe
165 170 175Leu Leu Phe Xaa Gly Asp Pro
Xaa Phe Pro Gln Thr Cys Ser Val Glu 180 185
190Ile Phe Pro Ser Pro Cys Leu Ser Xaa Thr Ile Asn Ile Xaa
Ile Pro 195 200 205Leu Pro Lys Leu
Ala Gly Ala Leu Xaa Leu Val Asn Pro Leu Gly Pro 210
215 220Leu22529215PRTHomo sapiensVARIANT(1)..(215)Xaa =
Any Amino Acid 29Xaa Ile Gln Ala Arg Gln Gly Lys Glu Glu Gly Gly Gly Arg
Gln Arg1 5 10 15Ser Ser
Leu Trp Pro Ser Glu Leu Gln Lys Asp Arg Gly Pro Ser Ala 20
25 30Ala Leu Glu Ala Asp Gln Gly Cys His
Ala Ala Gln Pro Gly His His 35 40
45Leu Val Ala Ala Ala Asp Thr Ser Asp Gly Trp Gln Phe Trp Ser Gly 50
55 60Cys Pro Gly Glu Gly Val Ala Asp Asp
Gln Pro Pro His Gln Ala Arg65 70 75
80Arg Leu Pro His Ser Asn Leu Val Phe Leu Ala Trp Cys Ser
Asp Ser 85 90 95Ser Gln
Ala Ala Pro Cys Gly Leu Ser Ala Ala Gly Ala Arg Ala Gly 100
105 110Gly Arg Val Pro Gly His Pro Ala Asp
Gly His Gly Asp Pro Gln Cys 115 120
125Leu Cys Cys Val Ile His His Pro Glu Glu Leu Arg Glu Ala Gln Glu
130 135 140Ala Gln Gly Arg Asn Lys Gly
Asn Asp Leu Leu Arg Ala Leu Ser Pro145 150
155 160Ile Leu Val Gln Gln Xaa Pro Ser Cys Phe Leu Xaa
Gly Thr Xaa Ile 165 170
175Phe Pro Lys Leu Ala Leu Leu Arg Phe Phe Pro His Leu Ala Ser Xaa
180 185 190Ala Gln Ile Xaa Leu Tyr
His Cys Gln Ser Leu Arg Ala His Ser Xaa 195 200
205Asn Leu Thr Pro Trp Gly Leu 210
21530232PRTHomo sapiensVARIANT(1)..(232) 30Glu Phe Lys Gln Asp Lys Asp
Glu Lys Lys Lys Gly Glu Asp Glu Asp1 5 10
15Lys Gly Pro Pro Cys Gly Pro Val Asn Cys Asn Glu Lys
Ile Val Val 20 25 30Leu Leu
Gln Arg Leu Lys Pro Glu Ile Lys Asp Val Ile Glu Gln Leu 35
40 45Asn Leu Val Thr Thr Trp Leu Gln Leu Gln
Ile Pro Arg Ile Glu Asp 50 55 60Gly
Asn Asn Phe Gly Val Ala Val Gln Glu Lys Val Phe Glu Leu Met65
70 75 80Thr Ser Leu His Thr Lys
Leu Glu Gly Phe His Thr Gln Ile Ser Lys 85
90 95Tyr Phe Ser Glu Arg Gly Asp Ala Val Thr Lys Ala
Ala Lys Gln Pro 100 105 110His
Val Gly Asp Tyr Arg Gln Leu Val His Glu Leu Asp Glu Ala Glu 115
120 125Tyr Arg Asp Ile Arg Leu Met Val Met
Glu Ile Arg Asn Ala Tyr Ala 130 135
140Val Leu Tyr Asp Ile Ile Leu Lys Asn Phe Glu Lys Leu Lys Lys Pro145
150 155 160Arg Gly Glu Thr
Lys Gly Met Ile Tyr Glu Pro Ser Leu Pro Phe Cys 165
170 175Asp Glu Tyr Ser Arg Xaa Leu Pro Ala Phe
Xaa Trp Gly Pro Xaa Phe 180 185
190Ser Pro Asn Leu Leu Cys Asp Phe Ser Leu Thr Leu Pro Leu Xaa His
195 200 205Asn Lys Tyr Xaa Tyr Thr Thr
Ala Lys Ala Cys Gly Arg Thr Pro Xaa 210 215
220Thr Ser Pro Leu Gly Ala Ser Lys225
23031214PRTHomo sapiensVARIANT(1)..(214)Xaa = Any Amino Acid 31Xaa Asn
Ser Gln Asn Ser Ser Glu Lys Ser Cys Phe Lys Val Leu Thr1 5
10 15Phe Phe Gln Met Arg Lys Lys Trp
Arg Lys Glu Lys Gly Asn Lys Lys 20 25
30Tyr Ser Lys Leu Ser Lys Lys Ile Asn Ser Val Lys Asn Gly Ile
Ser 35 40 45Leu Pro Gly Thr Ser
Phe Leu Ile Ile Gly Leu Ser Trp Gly Lys Leu 50 55
60Gly Pro Cys Leu Thr Gly Arg Phe Ser Arg Ser Cys Gly Pro
Gly Glu65 70 75 80Thr
Thr Gln Asp Ser Arg Ser Trp Ser Gly Arg Leu Lys Pro Trp Lys
85 90 95Met Ile Leu Gly Lys Lys Thr
Glu Asn Asn Asn Trp Pro Leu Ser Ile 100 105
110Thr Gly Asn Lys Phe Ser Asp Thr Ala Leu Gln Asn Gly Ser
Ile Gly 115 120 125Met Ala Pro Ser
Ser Arg Glu Ser Trp Leu Ser Gln Lys Arg Lys Leu 130
135 140Xaa Arg Arg Trp Xaa His Cys Cys Arg Gln His His
Trp Gly Asn Ser145 150 155
160Val Pro Xaa Gly Tyr Xaa Xaa Ser Val Xaa Leu Arg Xaa Glu Gln Xaa
165 170 175Trp Trp Xaa Xaa Xaa
Xaa Xaa Met Xaa Arg Xaa Xaa Xaa Cys Pro Xaa 180
185 190Leu Trp Glu Lys Pro Pro Leu Xaa Ser Xaa Gly Xaa
Xaa Xaa Xaa Pro 195 200 205Pro Xaa
Glu Xaa Gln Xaa 21032208PRTHomo sapiensVARIANT(1)..(208)Xaa = Any
Amino Acid 32Xaa Ile Leu Lys Ile Val Gln Lys Asn His Val Ser Lys Tyr Ser
His1 5 10 15Ser Ser Arg
Gly Lys Asn Gly Glu Gly Lys Lys Lys Glu Ile Glu Arg 20
25 30Ser Ile Asn Pro Asn Phe Gln Arg Lys Ser
Thr Val Ser Lys Thr Val 35 40
45Phe Arg Cys Leu Ala Gln Ala Asp Ser Ser Asp Ala Gly Glu Ser Trp 50
55 60Asp Pro Val Leu Glu Asp Ser Ala Glu
Gly Pro Ala Gly Leu Glu Arg65 70 75
80Leu His Lys Ile Pro Glu Val Gly Ala Gly Asp Ser Ser Leu
Gly Lys 85 90 95Ser Gly
Arg Lys Gln Lys Thr Thr Thr Gly His Val Pro Glu Thr Ser 100
105 110Ser Pro Thr Leu Leu Tyr Arg Met Ala
Ala Leu Ala Trp Arg Arg Ala 115 120
125Pro Glu Glu Arg Ala Gly Ser His Lys Arg Gly Asn Xaa Glu Glu Asp
130 135 140Xaa Cys Thr Ala Ala Gly Ser
Ile Thr Gly Glu Thr Gln Cys Xaa Trp145 150
155 160Val Ile Xaa Xaa Gln Xaa Thr Gly Xaa Asn Ser Xaa
Gly Gly Xaa Xaa 165 170
175Xaa Xaa Xaa Trp Xaa Gly Xaa Xaa Xaa Ala Pro Xaa Cys Gly Lys Ser
180 185 190Pro Pro Trp Xaa Ala Xaa
Gly Xaa Xaa Xaa Ser Pro Xaa Gly Lys Xaa 195 200
20533218PRTHomo sapiensVARIANT(1)..(218)Xaa = Any Amino Acid
33Glu Phe Ser Lys Phe Arg Lys Ile Met Phe Gln Ser Thr His Ile Leu1
5 10 15Pro Asp Glu Glu Lys Met
Val Lys Glu Arg Lys Arg Lys Leu Lys Glu 20 25
30Val Leu Ile Gln Thr Phe Lys Glu Asn Gln Gln Cys Gln
Lys Arg Tyr 35 40 45Phe Ala Ala
Trp His Lys Leu Ile Leu Asp His Arg Ile Lys Leu Gly 50
55 60Lys Ala Gly Thr Leu Ser Asp Trp Lys Ile Gln Leu
Lys Val Leu Arg65 70 75
80Ala Trp Arg Asp Tyr Thr Arg Phe Gln Lys Leu Glu Arg Glu Thr Gln
85 90 95Ala Leu Glu Asn Asp Leu
Arg Glu Glu Asn Arg Lys Gln Gln Leu Ala 100
105 110Thr Glu Tyr Asn Arg Lys Gln Val Leu Arg His Cys
Phe Thr Glu Trp 115 120 125Gln His
Trp His Gly Ala Glu Leu Leu Lys Arg Glu Leu Ala Leu Thr 130
135 140Lys Glu Glu Thr Xaa Lys Lys Met Xaa Ala Leu
Leu Gln Ala Ala Ser145 150 155
160Leu Gly Lys Leu Ser Ala Xaa Gly Leu Ser Xaa Xaa Ser Xaa Pro Glu
165 170 175Xaa Gly Thr Ala
Xaa Val Gly Xaa Pro Xaa Xaa Asn Xaa Gln Xaa Xaa 180
185 190Xaa Xaa Pro Xaa Phe Val Gly Lys Ala Pro Leu
Gly Xaa Xaa Trp Val 195 200 205Xaa
Xaa Xaa Pro Pro Xaa Gly Xaa Thr Xaa 210
21534135PRTHomo sapiensVARIANT(1)..(135)Xaa = Any Amino Acid 34Xaa Asn
Ser Gly Lys Leu Gly Thr Thr Val Lys Pro Lys Ser Leu Val1 5
10 15Thr Ser Ser Ser Gly Ala Leu Lys
Lys Gln His Lys Lys Pro Phe Asp 20 25
30Ala Met Asn Asn Ile Val Ala Asn Leu Leu Leu Asn Leu Thr Arg
Glu 35 40 45Ala Cys Gly Arg Thr
Arg Val Thr Ser Pro Leu Gly Ala Ser Lys Arg 50 55
60Val Leu Arg Gly Xaa Leu Ser Tyr Ser Ser Ala Ala Ala Ser
Phe Pro65 70 75 80Arg
Val Glu Glu Gln Ile Cys His Asn Val Ile His Cys Ile Lys Gly
85 90 95Leu Leu Met Leu Leu Phe Ser
Pro Arg Thr Ser Asn Gln Thr Leu Trp 100 105
110Leu His Ser Cys Thr Gln Leu Ser Arg Ile Arg Ile Pro Glu
His His 115 120 125Thr Leu Glu Tyr
Asp Ser Xaa 130 13535133PRTHomo
sapiensVARIANT(1)..(133)Xaa = Any Amino Acid 35Xaa Ile Leu Glu Ser Trp
Val Gln Leu Ser Gln Arg Val Trp Leu Leu1 5
10 15Gln Val Leu Gly Leu Lys Ser Ser Ile Arg Ser Pro
Leu Met Gln Ile 20 25 30Thr
Leu Trp Gln Ile Cys Ser Ser Thr Arg Gly Lys Leu Ala Ala Ala 35
40 45Leu Glu Leu Val Asn Pro Leu Gly Pro
Leu Asn Gly Ser Gly Xaa Tyr 50 55
60Val Thr Arg Val Arg Pro Gln Ala Ser Leu Val Arg Leu Arg Ser Lys65
70 75 80Phe Ala Thr Met Leu
Phe Ile Ala Ser Lys Gly Phe Leu Cys Cys Phe 85
90 95Phe Lys Ala Pro Glu Leu Glu Val Thr Arg Leu
Phe Gly Phe Thr Val 100 105
110Val Pro Asn Phe Pro Glu Phe Gly Ser Pro Ser Ile Thr Pro Asp Trp
115 120 125Asn Thr Thr Ala Xaa
13036129PRTHomo sapiensVARIANT(1)..(129)Xaa = Any Amino Acid 36Glu Phe
Trp Lys Val Gly Tyr Asn Cys Glu Ala Lys Glu Ser Gly Tyr1 5
10 15Phe Lys Phe Trp Gly Phe Lys Lys
Ala Ala Glu Ala Leu Cys Asn Glu 20 25
30His Cys Gly Lys Phe Ala Pro Gln Pro Asn Glu Gly Ser Leu Arg
Pro 35 40 45His Ser Ser Asn Leu
Thr Pro Trp Gly Leu Thr Gly Leu Glu Gly Xaa 50 55
60Thr Lys Leu Leu Glu Cys Gly Arg Lys Leu Pro Ser Leu Gly
Gly Ala65 70 75 80Asn
Leu Pro Gln Cys Tyr Ser Leu His Gln Arg Ala Ser Tyr Ala Ala
85 90 95Phe Leu Lys Pro Gln Asn Leu
Lys Pro Asp Ser Leu Ala Ser Gln Leu 100 105
110Tyr Pro Thr Phe Gln Asn Ser Asp Pro Arg Ala Ser His Leu
Thr Gly 115 120 125Ile
User Contributions:
Comment about this patent or add new information about this topic: