Patent application title: DETECTING CANCER WITH ANTI-CCL25 AND ANTI-CCR9 ANTIBODIES
Inventors:
James W. Lillard (Smyrna, GA, US)
Rajesh Singh (Atlanta, GA, US)
Rajesh Singh (Atlanta, GA, US)
Shailesh Singh (Powder Springs, GA, US)
Shailesh Singh (Powder Springs, GA, US)
Assignees:
MOREHOUSE SCHOOL OF MEDICINE
IPC8 Class: AG01N33574FI
USPC Class:
435 612
Class name: Measuring or testing process involving enzymes or micro-organisms; composition or test strip therefore; processes of forming such composition or test strip involving nucleic acid with significant amplification step (e.g., polymerase chain reaction (pcr), etc.)
Publication date: 2012-04-26
Patent application number: 20120100554
Abstract:
Methods for detecting cancer in a subject are disclosed. The method
includes detecting the level of expression of one or more cancer markers
in a biological sample obtained from the subject; and comparing the level
of expression of the one or more cancer markers in the biological sample
to a normal level of expression of the one or more cancer markers. The
one or more cancer markers comprise CCL25 or CCR9 or both CCL25 and CCR9.Claims:
1. A method for detecting the presence of cancer in a subject,
comprising: detecting the level of expression of one or more cancer
markers in a biological sample obtained from said subject; and comparing
the level of expression of said one or more cancer markers in said
biological sample to a normal level of expression of said one or more
cancer markers, wherein a higher than normal level of expression of said
one or more cancer markers in said biological sample is indicative of the
presence of cancer in said subject, wherein said normal level of
expression of said one or more cancer markers is a predetermined value or
is obtained from a control sample of known normal non-cancerous cells of
the same origin or type as said biological sample, and wherein said
cancer is blastoma, carcinoma, leukemia, lymphoma, melanoma, myeloma,
sarcoma or germ cell tumor, and wherein said one or more cancer markers
comprises CCL25 or CCR9 or both CCL25 and CCR9.
2. The method of claim 1, wherein said one or more cancer markers further comprises CXCL13 or CXCR5 or both CXCL13 and CXCR5.
3. The method of claim 2, wherein said one or more cancer markers further comprises CXCL16 or CXCR6 or both CXCL16 and CXCR6.
4. The method of claim 1, wherein said one or more cancer markers further comprises CXCL16 or CXCR6 or both CXCL16 and CXCR6.
5. The method of claim 1, wherein said one or more cancer markers further comprises one or more cancer markers selected from the group consisting of CXCL13, CXCR5, CXCL16 CXCR6, CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CCL27, CXCL1, CXCL2, CXCL3, CXCL7, CXCL8, CXCL12, CX3CL1, CCR2, CCR7, CCR8, CCR10, CXCR1, CXCR2, CXCR4, CXCR7 and CX3CR1.
6. The method of claim 1, wherein said cancer is carcinoma.
7. The method of claim 6, wherein said carcinoma is breast cancer and wherein one or more cancer markers further comprises one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CXCR4, CXCR5, CXCR6 CX3CR1, HER2, RBM3 and CEA.
8. The method of claim 6, wherein said carcinoma is prostate cancer and wherein one or more cancer markers further comprises one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CXCR4, CXCR5, CXCR6 and CX3CR1.
9. The method of claim 6, wherein said carcinoma is brain cancer, pituitary cancer or bone cancer and wherein one or more cancer markers further comprises one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CXCR4, CXCR5, CXCR6 and CX3CR1.
10. The method of claim 6, wherein said carcinoma is colorectal cancer and wherein one or more cancer markers further comprises one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CXCR4, CXCR5, CXCR6, CX3CR1, seprase polypeptide, anti-p53, osteopontin, and ferritin.
11. The method of claim 6, wherein said carcinoma is ovarian cancer and wherein one or more cancer markers further comprises one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CXCR4, CXCR5, CXCR6, CX3CR1, cancer antigen 125 (CA-125), HE-4, OVX-1 macrophage colony stimulating factor (M-CSF) and lysophosphatidyl choline.
12. The method of claim 6, wherein said carcinoma is lung cancer and wherein one or more cancer markers further comprises one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CCL25, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CCR9, CXCR4, CXCR5, CXCR6, CX3CR1, kinesin family member 4A (KIF4A), Neural pentraxin I (NPTX1), fibroblast growth factor receptor 1 oncogene partner (FGFR1OP) protein and CEA.
13. The method of claim 6, wherein said carcinoma is pancreatic cancer or gastric cancer and wherein one or more cancer markers further comprises one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CCL25, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CCR9, CXCR4, CXCR5, CXCR6, CX3CR1 and CEA.
14. The method of claim 1, wherein said biological sample is plasma, saliva or urine.
15. A method for assessing the prognosis of a subject with a cancer, comprising: determining the expression level of one or more cancer markers in a biological sample from said subject, and comparing the level of expression of said one or more cancer markers in said biological sample to a control level of expression of said one or more cancer markers, wherein a higher level of expression of said one or more cancer markers in the biological sample relative to said control level indicates that the prognosis of said subject is poor, wherein a lower or similar level of expression of said one or more cancer markers in said biological sample relative to said control level indicates that the prognosis of said subject is good, wherein a poor prognosis indicates that said cancer is of an aggressive or invasive type, wherein said cancer is blastoma, carcinoma, leukemia, lymphoma, melanoma, myeloma, sarcoma or germ cell tumor, and wherein said one or more cancer markers comprise CCL25 or CCR9 or both CCL25 and CCR9.
16. The method of claim 15, wherein said one or more cancer markers further comprises (1) CXCL13 or CXCR5 or both CXCL13 and CXCR5, and/or (2) CXCL16 or CXCR6 or both CXCL16 and CXCR6.
17. The method of claim 15, wherein said one or more cancer markers further comprises one or more cancer markers selected from the group consisting of CXCL13, CXCR5, CXCL16 CXCR6, CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CCL27, CXCL1, CXCL2, CXCL3, CXCL7, CXCL8, CXCL12, CX3CL1, CCR2, CCR7, CCR8, CCR10, CXCR1, CXCR2, CXCR4, CXCR7 and CX3CR1.
18. The method of claim 15, wherein said biological sample is plasma, saliva or urine.
19. A method for monitoring the course of cancer treatment in a subject, comprising: determining the expression levels of one or more cancer markers in one or more biological samples obtained from said subject during or after said treatment, and comparing the level of expression of said one or more cancer markers in said one or more biological samples to a control level of expression of said one or more cancer markers, wherein said control level of said one or more cancer markers is a pre-treatment level of said one or more cancer markers in said subject or a predetermined reference level, wherein said treatment is deemed efficacious if said one or more cancer markers in said one or more biological samples is similar to or lower than said control level, wherein said cancer is blastoma, carcinoma, leukemia, lymphoma, melanoma, myeloma, sarcoma or germ cell tumor, and wherein said one or more cancer markers comprise CCL25 or CCR9 or both CCL25 and CCR9.
20. The method of claim 19, wherein said one or more cancer markers further comprises (1) CXCL13 or CXCR5 or both CXCL13 and CXCR5, and/or (2) CXCL16 or CXCR6 or both CXCL16 and CXCR6.
21. The method of claim 19, wherein said one or more cancer markers further comprises one or more cancer markers selected from the group consisting of CXCL13, CXCR5, CXCL16 CXCR6, CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CCL27, CXCL1, CXCL2, CXCL3, CXCL7, CXCL8, CXCL12, CX3CL1, CCR2, CCR7, CCR8, CCR10, CXCR1, CXCR2, CXCR4, CXCR7 and CX3CR1.
22. The method of claim 19, wherein said biological sample is plasma, saliva or urine.
23. A kit for detecting cancer, comprising: reagents for determining expression of CCL25 and/or CCR9 in a biological sample; and instructions for how to use said reagents, wherein said reagents comprise an anti-CCL25 antibody, an anti-CCR9 antibody, or both.
24. The kit of claim 23, further comprising: (1) reagents for determining expression of CXCL13 and/or CXCR5 in a biological sample; and instructions for how to use said reagents, wherein said reagents comprise an anti-CXCL13 antibody, an anti-CXCR5 antibody, or both.
25. The kit of claim 24, further comprising: reagents for determining expression of CXCL16 and/or CXCR6 in a biological sample; and instructions for how to use said reagents, wherein said reagents comprise an anti-CXCL16 antibody, an anti-CXCR6 antibody, or both.
26. The kit of claim 23, further comprising: reagents for determining expression of CXCL16 and/or CXCR6 in a biological sample; and instructions for how to use said reagents, wherein said reagents comprise an anti-CXCL16 antibody, an anti-CXCR6 antibody, or both.
Description:
[0001] This application is a Continuation-In-Part of U.S. patent
application Ser. No. 13/313,705, filed on Dec. 7, 2011, which is a
Continuation-In-Part of U.S. patent application Ser. No. 13/248,904,
filed on Sep. 29, 2011, which is a Continuation-In-Part of U.S. patent
application Ser. No. 13/233,769, filed on Sep. 15, 2011, which is a
Continuation-In-Part of U.S. patent application Ser. No. 12/967,273,
filed Dec. 14, 2010, which is a continuation of U.S. patent application
Ser. No. 10/712,398, filed on Nov. 14, 2003, now U.S. Pat. No. 7,919,083,
which claims priority of U.S. Provisional Patent Application No.
60/426,347, filed Nov. 15, 2002. This application is also a
Continuation-In-Part of U.S. patent application Ser. No. 13/312,343,
filed on Dec. 6, 2011, which is a Continuation-In-Part of U.S. patent
application Ser. No. 13/233,769, filed Sep. 15, 2011, which is a
Continuation-In-Part of U.S. patent application Ser. No. 12/967,273,
filed Dec. 14, 2010, which is a Continuation of U.S. patent application
Ser. No. 10/712,398, filed on Nov. 14, 2003, now U.S. Pat. No. 7,919,083,
which claims priority of U.S. Provisional Patent Application No.
60/426,347, filed Nov. 15, 2002. The entirety of all of the
aforementioned applications is incorporated herein by reference.
FIELD
[0002] This application generally relates to detection of cancer. In particular, the application relates to a method for detecting cancer using anti-chemokine and/or anti-chemokine receptor antibodies.
BACKGROUND
[0003] Cancer is one of the leading cause of death in the United States. Most cancer starts in just a single neoplastic cell. The neoplastic cell proliferate to form a local "tumor." A tumor simply means a swelling; it is not necessarily cancerous. A tumor which only grows in its place or origin, and cannot spread distantly, is a benign tumor and is not cancer. However, a tumor which has the capacity to spread (whether it actually does or not) is called a malignant tumor or cancer. A cancer may spread via the blood or lymphatic system to regional lymph nodes and to distant sites via a process called metastasis. A metastasized cancer is more difficult to treat because it now spreads into many different tissues and organs. It has been demonstrated that early treatment increase survival in many types of cancers, such as breast cancer, colon cancer, ovarian cancer and prostate cancer.
[0004] Chemokines are a superfamily of small, cytokine-like proteins that are resistant to hydrolysis, promote neovascularization or endothelial cell growth inhibition, induce cytoskeletal rearrangement, activate or inactivate lymphocytes, and mediate chemotaxis through interactions with G-protein coupled receptors. Chemokines can mediate the growth and migration of host cells that express their receptors.
[0005] Chemokine (C-C motif) ligand 25 (CCL25), also known as Thymus-Expressed Chemokine (TECK), is a small cytokine belonging to the CC chemokine family. CCL25 is chemotactic for thymocytes, macrophages, and dendritic cells. CCL25 elicits its effects by binding the chemokine receptors CCR9 and is believed to play a role in the development of T-cells. Human CCL25 is produced as a protein precursor containing 151 amino acids. The gene for CCL25 (scya25) is located on human chromosome 19.
[0006] Chemokine (C-C motif) receptor 9 (CCR9), also known as GPR 9-6, is very highly expressed in thymus (on both immature and mature T-cells) while low in lymph nodes and spleen. CCR9 is also abundant in the gut, with its expression associated with T cells of the intestine. To note, the chemokine binding protein D6 had previously been referred to as CCR9, but this molecule is a scavenger receptor not a true (signaling) chemokine receptor.
SUMMARY
[0007] One aspect of the present application relates to a method for detecting cancer in a subject. The method comprises detecting the level of expression of one or more cancer markers in a biological sample obtained from the subject; and comparing the level of expression of the one or more cancer markers in the biological sample to a normal level of expression of the one or more cancer markers, wherein a higher than normal level of expression of said one or more cancer markers in the biological sample is indicative of the presence of cancer in the subject, wherein the normal level of expression of the one or more cancer markers is a predetermined value or is obtained from a control sample of known normal non-cancerous cells of the same origin or type as the biological sample, wherein the cancer is blastoma, carcinoma, leukemia, lymphoma, melanoma, myeloma, sarcoma or germ cell tumor and wherein the one or more cancer markers comprise CCL25 or CCR9 or both CCL25 and CCR9.
[0008] Another aspect of the present application relates to a method for detecting cancer in a subject. The method comprises detecting the level of expression of one or more cancer markers in a biological sample obtained from the subject; and comparing the level of expression of the one or more cancer markers in the biological sample to a normal level of expression of the one or more cancer markers, wherein a higher than normal level of expression of said one or more cancer markers in the biological sample is indicative of the presence of cancer in the subject, wherein the normal level of expression of the one or more cancer markers is a predetermined value or is obtained from a control sample of known normal non-cancerous cells of the same origin or type as the biological sample, wherein the cancer is blastoma, carcinoma, leukemia, lymphoma, melanoma, myeloma, sarcoma or germ cell tumor and wherein the one or more cancer markers comprise: (1) one or more cancer markers selected from the group consisting of CCL25 and CCL9; and (2) one or more cancer markers selected from the group consisting of CXCL13 and CXCR5 and/or one or more cancer markers selected from the group consisting of CXCL16 and CXCR6.
[0009] Another aspect of the present application relates to a method for assessing the prognosis of a subject with a cancer. The method comprises determining the expression level of one or more cancer markers in a biological sample from the subject, and comparing the level of expression of the one or more cancer markers in the biological sample to a control level of expression of the one or more cancer markers, wherein a higher level of expression of the one or more cancer markers in the biological sample relative to the control level indicates that the prognosis of the subject is poor, and wherein a lower or similar level of expression of the one or more cancer markers in the biological sample relative to the control level indicates that the prognosis of the subject is good, wherein a poor prognosis indicates that the cancer is of an aggressive or invasive type, wherein the blastoma, carcinoma, leukemia, lymphoma, melanoma, myeloma, sarcoma or germ cell tumor and wherein the one or more cancer markers comprise CCL25 or CCR9 or both CCL25 and CCR9.
[0010] Another aspect of the present application relates to a method for assessing the prognosis of a subject with a cancer. The method comprises determining the expression level of one or more cancer markers in a biological sample from the subject, and comparing the level of expression of the one or more cancer markers in the biological sample to a control level of expression of the one or more cancer markers, wherein a higher level of expression of the one or more cancer markers in the biological sample relative to the control level indicates that the prognosis of the subject is poor, and wherein a lower or similar level of expression of the one or more cancer markers in the biological sample relative to the control level indicates that the prognosis of the subject is good, wherein a poor prognosis indicates that the cancer is of an aggressive or invasive type, wherein the cancer is blastoma, carcinoma, leukemia, lymphoma, melanoma, myeloma, sarcoma or germ cell tumor and wherein the one or more cancer markers comprise: (1) one or more cancer markers selected from the group consisting of CCL25 and CCL9; and (2) one or more cancer markers selected from the group consisting of CXCL13 and CXCR5 and/or one or more cancer markers selected from the group consisting of CXCL16 and CXCR6.
[0011] Another aspect of the present application relates to a method for monitoring the course of cancer treatment in a subject. The method comprises determining the expression levels of one or more cancer markers in one or more biological samples obtained from the subject during or after the treatment, and comparing the level of expression of the one or more cancer markers in the one or more biological samples to a control level of expression of the one or more cancer markers, wherein the control level of the one or more cancer markers is a pre-treatment level of the one or more cancer markers in the subject or a predetermined reference level, wherein the treatment is deemed efficacious if the one or more cancer markers in the one or more biological samples is similar to or lower than the control level, wherein the cancer is blastoma, carcinoma, leukemia, lymphoma, melanoma, myeloma, sarcoma or germ cell tumor and wherein the one or more cancer markers comprise CCL25 or CCR9 or both CCL25 and CCR9.
[0012] Another aspect of the present application relates to a method for monitoring the course of cancer treatment in a subject. The method comprises determining the expression levels of one or more cancer markers in one or more biological samples obtained from the subject during or after the treatment, and comparing the level of expression of the one or more cancer markers in the one or more biological samples to a control level of expression of the one or more cancer markers, wherein the control level of the one or more cancer markers is a pre-treatment level of the one or more cancer markers in the subject or a predetermined reference level, wherein the treatment is deemed efficacious if the one or more cancer markers in the one or more biological samples is similar to or lower than the control level, wherein the cancer is blastoma, carcinoma, leukemia, lymphoma, melanoma, myeloma, sarcoma or germ cell tumor and wherein the one or more cancer markers comprise: (1) one or more cancer markers selected from the group consisting of CCL25 and CCL9; and (2) one or more cancer markers selected from the group consisting of CXCL13 and CXCR5 and/or one or more cancer markers selected from the group consisting of CXCL16 and CXCR6.
[0013] Another aspect of the present application relates to a kit for detecting cancer. The kit comprises reagents for determining expression of CCL25 and/or CCR9 in a biological sample; and instructions for how to use the reagents, wherein the reagents comprise an anti-CCL25 antibody, an anti-CCR9 antibody, or both.
BRIEF DESCRIPTION OF THE DRAWINGS
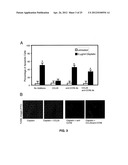
[0014] FIG. 1 shows CCL25 expression by breast cancer tissue.
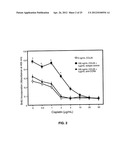
[0015] FIG. 2 shows that CCL25 inhibits cisplatin-induced reductions in breast cancer cell line growth.
[0016] FIG. 3 shows that CCL25 protects breast cancer cells from cisplatin-induced apoptosis.
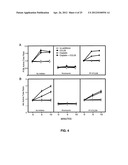
[0017] FIGS. 4A-B show PI3K and Akt activation by CCL25-CCR9 interactions in a breast cancer cell line.
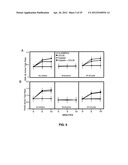
[0018] FIGS. 5A-B show GSK-3β and FKHR phosphorylation following CCL25 treatment of a breast cancer cell line.
[0019] FIG. 6 shows CCR9 and CCL25 expression by ovarian cancer tissues.
[0020] FIGS. 7A-B show an analysis of CCL25 expression by ovarian cancer tissues.
[0021] FIGS. 8A-B show an analysis of CCR9 expression by ovarian cancer tissues.
[0022] FIGS. 9A-B show CCR9 and CCL25 expression by ovarian cancer cell lines.
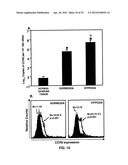
[0023] FIGS. 10A-B show hypoxia-regulated CCR9 mRNA and surface protein expression by ovarian cancer cells.
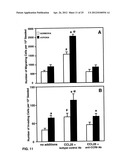
[0024] FIGS. 11A-B show hypoxia-mediated and CCL25-mediated migration and invasion of SKOV-3 cells.
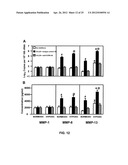
[0025] FIGS. 12A-B show CCL25-induced collagenase expression by SKOV-3 cells.
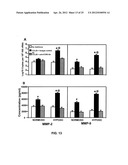
[0026] FIGS. 13A-B show CCL25-induced gelatinase expression by SKOV-3 cells.
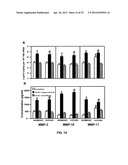
[0027] FIGS. 14A-B show CCL25-induced stromelysin expression by SKOV-3 cells.
[0028] FIG. 15 shows CCR9 expression by prostate cancer cells.
[0029] FIGS. 16A-D show CCR9 expression by prostate tissue.
[0030] FIGS. 17A-D show CCL25 expression by prostate cancer tissue.
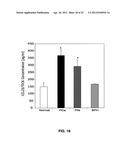
[0031] FIG. 18 shows serum CCL25 levels in normal healthy donors or patients with prostatic disease.
[0032] FIGS. 19A-C show CCL25 expression by mouse bone marrow cells.
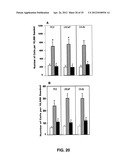
[0033] FIGS. 20A-B show CCR9-mediated prostate cancer cell migration and invasion.
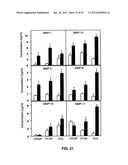
[0034] FIG. 21 shows CCL25-induced active MMP expression by prostate cancer cell lines.
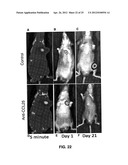
[0035] FIGS. 22A-F show inhibition of bone metastasis of PC3 prostate cancer cell line by CCR9 knockdown.
[0036] FIG. 23 shows serum CCL25 levels in lung cancer patients.
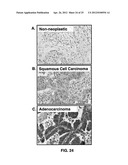
[0037] FIGS. 24A-C show CCR9 expression by non-neoplastic lung and lung cancer tissues.
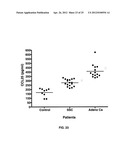
[0038] FIGS. 25A-C show CCR9-CCL25 expression by colon cancer tissues.
DETAILED DESCRIPTION
[0039] The following detailed description is presented to enable any person skilled in the art to make and use the application. For purposes of explanation, specific nomenclature is set forth to provide a thorough understanding of the present invention. However, it will be apparent to one skilled in the art that these specific details are not required to practice the invention. Descriptions of specific applications are provided only as representative examples. The present invention is not intended to be limited to the embodiments shown, but is to be accorded the widest possible scope consistent with the principles and features disclosed herein.
[0040] Unless otherwise defined, scientific and technical terms used in connection with the present invention shall have the meanings that are commonly understood by those of ordinary skill in the art. Further, unless otherwise required by context, singular terms shall include pluralities and plural terms shall include the singular.
[0041] As used herein, the following terms shall have the following meanings:
[0042] As used herein, the teen "antibody" refers to immunoglobulin molecules and immunologically active portions of immunoglobulin (Ig) molecules, i.e., molecules that contain an antigen binding site that specifically binds (immunoreacts with) an antigen. The term "antibody" is used in the broadest sense and specifically covers monoclonal antibodies (including full length monoclonal antibodies), polyclonal antibodies, multispecific antibodies (e.g., bispecific antibodies), and antibody fragments so long as they exhibit the desired biological activity. By "specifically bind" or "immunoreacts with" is meant that the antibody reacts with one or more antigenic determinants of the desired antigen and does not react (i.e., bind) with other polypeptides or binds at much lower affinity with other polypeptides. The term "antibody" also includes antibody fragments that comprise a portion of a full length antibody, generally the antigen binding or variable region thereof. Examples of antibody fragments include Fab, Fab', F(ab')2, and Fv fragments; diabodies; linear antibodies; single-chain antibody (scFv) molecules; and multispecific antibodies formed from antibody fragments. In certain embodiments of the application, it may be desirable to use an antibody fragment, rather than an intact antibody, to increase tumor penetration, for example. In this case, it may be desirable to use an antibody fragment that has been modified by any means known in the art in order to increase its serum half life.
[0043] The term "monoclonal antibody" as used herein refers to an antibody obtained from a population of substantially homogeneous antibodies, i.e., the individual antibodies comprising the population are identical except for possible naturally occurring mutations that may be present in minor amounts. The monoclonal antibodies herein specifically include "chimeric" antibodies in which a portion of the heavy and/or light chain is identical with or homologous to corresponding sequences in antibodies derived from a particular species or belonging to a particular antibody class or subclass, while the remainder of the chain(s) is identical with or homologous to corresponding sequences in antibodies derived from another species or belonging to another antibody class or subclass, as well as fragments of such antibodies, so long as they exhibit the desired biological activity (U.S. Pat. No. 4,816,567; and Morrison et al., Proc. Natl. Acad. Sci. USA 81:6851-6855 (1984)).
[0044] "Humanized" forms of non-human antibodies are chimeric antibodies which contain minimal sequence derived from non-human immunoglobulin. For the most part, humanized antibodies are human immunoglobulins (recipient antibody) in which residues from a hypervariable region of the recipient are replaced by residues from a hypervariable region of a non-human species (donor antibody) such as mouse, rat, rabbit or nonhuman primate having the desired specificity, affinity, and/or capacity. Methods for making humanized and other chimeric antibodies are known in the art.
[0045] "Bispecific antibodies" are antibodies that have binding specificities for at least two different antigens. In the present case, one of the binding specificities is for CXCL16 or CXCR6. The second binding target is any other antigen, and advantageously is a cell-surface protein or receptor or receptor subunit. Methods for making bispecific antibodies are known in the art.
[0046] The use of "heteroconjugate antibodies" is also within the scope of the present application. Heteroconjugate antibodies are composed of two covalently joined antibodies. Such antibodies have, for example, been proposed to target immune system cells to unwanted cells (U.S. Pat. No. 4,676,980). It is contemplated that the antibodies can be prepared in vitro using known methods in synthetic protein chemistry, including those involving crosslinking agents.
[0047] The teen "tumor" as used herein refers to a neoplasm or a solid lesion formed by an abnormal growth of cells. A tumor can be benign, pre-malignant or malignant.
[0048] A "primary tumor" is a tumor appearing at a first site within the subject and can be distinguished from a "metastatic tumor" which appears in the body of the subject at a remote site from the primary tumor.
[0049] The term "cancer," as used herein, refer to or describe the physiological condition in mammals that is typically characterized by unregulated cell growth. Exemplary cancers include: carcinoma, melanoma, sarcoma, lymphoma, leukemia, germ cell tumor, and blastoma. More particular examples of such cancers include squamous cell cancer (e.g., epithelial squamous cell cancer), lung cancer including small-cell lung cancer, non-small cell lung cancer, adenocarcinoma of the lung and squamous carcinoma of the lung, cancer of the peritoneum, hepatocellular cancer, gastric or stomach cancer including gastrointestinal cancer, pancreatic cancer, glioblastoma, cervical cancer, ovarian cancer, liver cancer, bladder cancer, cancer of the urinary tract, hepatoma, breast cancer, colon cancer, rectal cancer, colorectal cancer, endometrial or uterine carcinoma, salivary gland carcinoma, kidney or renal cancer, prostate cancer, vulval cancer, thyroid cancer, hepatic carcinoma, anal carcinoma, penile carcinoma, melanoma, multiple myeloma and B-cell lymphoma, brain, as well as head and neck cancer, and associated metastases.
[0050] The term "carcinoma" as used herein refers to an invasive malignant tumor consisting of transformed epithelial cells or transformed cells of unknown histogenesis, but which possess specific molecular or histological characteristics that are associated with epithelial cells, such as the production of cytokeratins or intercellular bridges. Exemplary carcinomas of the present application include ovarian cancer, vaginal cancer, cervical cancer, uterine cancer, prostate cancer, anal cancer, rectal cancer, colon cancer, stomach cancer, pancreatic cancer, insulinoma, adenocarcinoma, adenosquamous carcinoma, neuroendocrine tumor, breast cancer, lung cancer, esophageal cancer, oral cancer, brain cancer, medulloblastoma, neuroectodermal tumor, glioma, pituitary cancer, and bone cancer.
[0051] The term "lymphoma" as used herein is a cancer of lymphatic cells of the immune system. Lymphomas typically present as a solid tumor. Exemplary lymphomas include: small lymphocytic lymphoma, lymphoplasmacytic lymphoma, Waldenstrom macroglobulinemia, splenic marginal zone lymphoma, plasmacytoma, extranodal marginal zone B cell lymphoma, MALT lymphoma, nodal marginal zone B cell lymphoma (NMZL), follicular lymphoma, mantle cell lymphoma, diffuse large B cell lymphoma, mediastinal (thymic) large B cell lymphoma, intravascular large B cell lymphoma, primary effusion lymphoma, Burkitt lymphoma, B cell chronic lymphocytic lymphoma, classical Hodgkin lymphoma, nodular lymphocyte-predominant Hodgkin lymphoma, adult T cell lymphoma, nasal type extranodal NK/T cell lymphoma, enteropathy-type T cell lymphoma, hepatosplenic T cell lymphoma, blastic NK cell lymphoma, mycosis fungoide, Sezary syndrome, primary cutaneous CD30-positive T cell lymphoproliferative disorders, primary cutaneous anaplastic large cell lymphoma, lymphomatoid papulosis, angioimmunoblastic T cell lymphoma, unspecified peripheral T cell lymphoma, and anaplastic large cell lymphoma. Exemplary forms of classical Hodgkin lymphoma including: nodular sclerosis, mixed cellularity, lymphocyte-rich, and lymphocyte-depleted or not depleted.
[0052] The term "sarcoma" as used herein is a cancer that arises from transformed cells in one of a number of tissues that develop from embryonic mesoderm. Thus, sarcomas include tumors of bone, cartilage, fat, muscle, vascular, and hematopoietic tissues. For example, osteosarcoma arises from bone, chondrosarcoma arises from cartilage, liposarcoma arises from fat, and leiomyosarcoma arises from smooth muscle. Exemplary sarcomas include: Askin's tumor, botryodies, chondrosarcoma, Ewing's-PNET, malignant Hemangioendothelioma, malignant Schwannoma, osteosarcoma, soft tissue sarcomas. Subclases of soft tissue sarcomas include: alveolar soft part sarcoma, angiosarcoma, cystosarcoma phyllodes, dermatofibrosarcomadesmoid tumor, desmoplastic small round cell tumor, epithelioid sarcomaextraskeletal chondrosarcoma, extraskeletal osteosarcoma, fibrosarcoma, hemangiopericytoma, hemangiosarcoma, Kaposi's sarcoma, leiomyosarcoma, liposarcoma, lymphangiosarcomal, lymphosarcoma, malignant fibrous histiocytoma, neurofibrosarcoma, rhabdomyosarcoma, and synovial sarcoma.
[0053] The term "leukemia" as used herein is a cancer of the blood or bone marrow characterized by an abnormal increase of white blood cells. Leukemia is a broad term covering a spectrum of diseases. In turn, it is part of the even broader group of diseases called hematological neoplasms. Leukemia is subdivided into a variety of large groups; the first division is between acute and chronic forms of leukemia. Acute leukemia is characterized by a rapid increase in the numbers of immature blood cells. Crowding due to such cells makes the bone marrow unable to produce healthy blood cells. Chronic leukemia is characterized by the excessive build up of relatively mature, but still abnormal, white blood cells. Typically taking months or years to progress, the cells are produced at a much higher rate than normal cells, resulting in many abnormal white blood cells in the blood. Leukemia is also subdivided by the blood cells affected. This split divides leukemias into lymphoblastic or lymphocytic leukemias and myeloid or myelogenous leukemias. In lymphoblastic or lymphocytic leukemias, the cancerous change takes place in a type of marrow cell that normally goes on to form lymphocytes. In myeloid or myelogenous leukemias, the cancerous change takes place in a type of marrow cell that normally goes on to form red blood cells, some other types of white cells, and platelets. Combining these two classifications provides a total of four main categories. Within each of these four main categories, there are typically several subcategories. There are also rare types outside of this classification scheme. Exemplary leukemias include: acute lymphoblastic leukemia (ALL), chronic lymphocytic leukemia (CLL), acute myelogenous leukemia (AML), chronic myelogenous leukemia (CML), hairy cell leukemia (HCL), T-cell prolymphocytic leukemia, large granular lymphocytic leukemia, juvenile myelomonocytic leukemia, B-cell prolymphocytic leukemia, Burkitt leukemia, and adult T-cell leukemia.
[0054] The term "melanoma" as used herein is a cancer or malignant tumor of melanocytes. Melanocytes are cells that produce the dark pigment, melanin, which is responsible for the color of skin. They predominantly occur in skin, but are also found in other parts of the body, including the bowel and the eye. Melanoma is divided into the following stereotypes and subtypes: lentigo maligna, lentigo maligna melanoma, superficial spreading melanoma, acral lentiginous melanoma, mucosal melanoma, nodular melanoma, polypoid melanoma, desmoplastic melanoma, amelanotic melanoma, soft-tissue melanoma, melanoma with small nevus-like cells, melanoma with features of a Spitz nevus, and uveal melanoma.
[0055] The term "germ cell tumor (GCT)" as used herein is a neoplasm derived from germ cells. Germ cell tumors can be cancerous or non-cancerous tumors. Germ cells normally occur inside the gonads (ovary and testis). Germ cell tumors that originate outside the gonads may be birth defects resulting from errors during development of the embryo. Germ cell tumors are broadly divided in two classes: germinomatous or seminomatous and nongerminomatous or nonseminomatous germ cell tumors. Exemplary germinomatous or seminomatous germ cell tumors include: germinoma, dysgerminoma, and seminoma. Exemplary nongerminomatous or nonseminomatous germ cell tumors include: Embryonal carcinoma, endodermal sinus tumor or yolk sac tumor (EST, YST), choriocarcinoma, mature teratoma, dermoid cyst, immature teratoma, teratoma with malignant transformation, polyembryoma, gonadoblastoma, and mixed GCT.
[0056] The teen "metastasis" as used herein refers to the spread of a cancer or carcinoma from one organ or part to another non-adjacent organ or part.
[0057] The term "biological sample" refers to a sample of biological material obtained from a mammal subject, preferably a human subject, including a tissue, a tissue sample, a cell sample, a tumor sample, a stool sample, and a biological fluid, e.g., blood, plasma, serum, saliva, urine, cerebral or spinal fluid, lymph liquid and a nipple aspirate. A biological sample may be obtained in the form of, e.g., a tissue biopsy, such as, an aspiration biopsy, a brush biopsy, a surface biopsy, a needle biopsy, a punch biopsy, an excision biopsy, an open biopsy, an incision biopsy and an endoscopic biopsy. In one embodiment, the biological sample is a blood, serum or plasma sample. In another embodiment, the biological sample is a saliva sample. In yet another embodiment, the biological sample is a urine sample.
[0058] An "isolate" of a biological sample (e.g., an isolate of a tissue or tumor sample) refers to a material or composition (e.g., a biological material or composition) which has been separated, derived, extracted, purified or isolated from the sample and preferably is substantially free of undesirable compositions and/or impurities or contaminants associated with the biological sample.
[0059] A "tissue sample" includes a portion, piece, part, segment, or fraction of a tissue which is obtained or removed from an intact tissue of a subject, preferably a human subject.
[0060] A "tumor sample" includes to a portion, piece, part, segment, or fraction of a tumor, for example, a tumor which is obtained or removed from a subject (e.g., removed or extracted from a tissue of a subject), preferably a human subject. A tumor sample may be obtained from a primary tumor or a metastatic tumor.
[0061] A "mammal" for purposes of treatment refers to any animal classified as a mammal, including humans, non-human primates, domestic and farm animals, and zoo, sports, or pet animals, such as dogs, horses, cats, cows, etc. Preferably, the mammal is human.
[0062] The term "increased level" refers to a level that is higher than a normal or control level customarily defined or used in the relevant art. For example, an increased level of immunostaining in a tissue is a level of immunostaining that would be considered higher than the level of immunostaining in a control tissue by a person of ordinary skill in the art.
[0063] Ranges may be expressed herein as from "about" one particular value, and/or to "about" another particular value. When such a range is expressed, another embodiment includes from the one particular value and/or to the other particular value. Similarly, when values are expressed as approximations, by use of the antecedent "about," it will be understood that the particular value forms another embodiment. It will be further understood that the endpoints of each of the ranges are significant both in relation to the other endpoint, and independently of the other endpoint. It is also understood that there are a number of values disclosed herein, and that each value is also herein disclosed as "about" that particular value in addition to the value itself. For example, if the value "10" is disclosed, then "about 10" is also disclosed. It is also understood that when a value is disclosed that "less than or equal to" the value, "greater than or equal to the value" and possible ranges between values are also disclosed, as appropriately understood by the skilled artisan. For example, if the value "10" is disclosed the "less than or equal to 10" as well as "greater than or equal to 10" is also disclosed. As used herein, the term "antibody" refers to immunoglobulin molecules and immunologically active portions of immunoglobulin (Ig) molecules, i.e., molecules that contain an antigen binding site that specifically binds (immunoreacts with) an antigen.
Method for Detecting Cancer by Measuring CCL25 and/or CCR9 Expression or Activity
[0064] CCL25 is a ligand for the CCR9 chemokine receptor. Both the chemokine and the receptor appear to play a role in the regulation of metastasis and invasion of cancer. Both CCL25 and CCR9 are locally up-regulated in multiple carcinoma tissue types compared to normal tissues, including ovarian, lung, breast, prostate, colon, bone and pancreatic cancers. CCL25 levels are also increased in the serum of patients with those cancers. Additionally, soluble CCL25 chemokine enhances both in vivo and in vitro proliferation and migration of cancer cells.
[0065] CCR9 is a member of the chemokine receptor family of G protein coupled receptors (GPCRs) that may have a diverse role in cancer cell survival that presumably supports protection against chemotherapeutic drugs. We show that interaction of CCR9 with CCL25 modulates matrix metalloproteinase (MMP) expression and enhances the migration and invasive potential of carcinoma cells. This suggests that CCR9-CCL25 interaction contributes to carcinoma cell migration and invasion. Accordingly, blocking this axis has the potential to inhibit carcinoma cell metastasis).
[0066] One aspect of the present application relates to a method for detecting the presence of cancer in a subject, comprising: detecting the level of expression of one or more cancer markers in a biological sample obtained from said subject; and comparing the level of expression of said one or more cancer markers in said biological sample to a normal level of expression of said one or more cancer markers, wherein a higher than normal level of expression of said one or more cancer markers in said biological sample is indicative of the presence of cancer in said subject, wherein said normal level of expression of said one or more cancer markers is a predetermined value or is obtained from a control sample of known normal non-cancerous cells of the same origin or type as said biological sample, and wherein said cancer is blastoma, carcinoma, leukemia, lymphoma, melanoma, myeloma, sarcoma or germ cell tumor and wherein said one or more cancer markers comprises CCL25 or CCR9 or both CCL25 and CCR9.
[0067] In one embodiment, said one or more cancer markers comprise (1) CCL25 or CCR9 or both CCL25 and CCR9 and (2) CXCL13 or CXCR5 or both CXCL13 and CXCR5. In another embodiment, said one or more cancer markers comprise (1) CCL25 or CCR9 or both CCL25 and CCR9 and (2) CXCL16 or CXCR6 or both CXCL16 and CXCR6.
[0068] In another embodiment, said one or more cancer markers comprise (1) CCL25 or CCR9 or both CCL25 and CCR9, (2) CXCL13 or CXCR5 or both CXCL13 and CXCR5, and (3) CXCL16 or CXCR6 or both CXCL16 and CXCR6. In another embodiment, said one or more cancer markers comprise (1) CCL25 or CCR9 or both CCL25 and CCR9, and/or (2) CXCL13 or CXCR5 or both CXCL13 and CXCR5, and/or (3) CXCL16 or CXCR6 or both CXCL16 and CXCR6' and (4) one or more other cancer markers.
[0069] In still another embodiment, said one or more other cancer markers comprise (1) CCL25 or CCR9 or both CCL25 and CCR9, and (2) one or more cancer markers selected from the group consisting of CXCL1, CXCL2, CXCL3, CXCL4, CXCL6, CXCL7, CXCL8, CXCL9, CXCL10, CXCL11, CXCL12, CXCL13, CXCL14, CXCL15, CXCL16, CXCR1, CXCR2, CXCR3, CXCR4, CXCR5, CXCR6, CXCR7, CCL1, CCL2, CCL3, CCL4, CCL5, CCL6, CCL7, CCL8, CCL9, CCL10, CCL11, CCL12, CCL13, CCL14, CCL15, CCL16, CCL17, CCL18, CCL19, CCL20, CCL21, CCL22, CCL24, CCL25, CCL25-1, CCL25-2, CCL27, CCL28, CCR1, CCR2, CCR3, CCR4, CCR5, CCR6, CCR7, CCR8, CCR9, CCR10, CCR11, XCL1, XCL2, XCR1, CX3CR1, CX3CL1, HER2, RNA binding motif 3 ("RBM3"), carcinoembryonic Antigen (CEA), prostate specific antigen (PSA), chromgranin A (CGA), dehydroepiandrosterone (DHEA), neuron-specific enolase (NSE), prostatic acid phosphatase (PAP), prolactin, B7-H3, seprase polypeptide, anti-p53, osteopontin, ferritin, lysophosphatidyl choline, kinesin family member 4A (KIF4A), Neural pentraxin I (NPTX1) and fibroblast growth factor receptor 1 oncogene partner (FGFR1OP) protein.
[0070] In another embodiment, said cancer is breast cancer and wherein the one or more cancer markers comprise (1) CCL25 or CCR9 or both CCL25 and CCR9, and (2) one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CXCR4, CXCR5, CXCR6 CX3CR1, HER2, RBM3 and CEA.
[0071] In another embodiment, said carcinoma is prostate cancer and wherein the one or more cancer markers comprise (1) CCL25 or CCR9 or both CCL25 and CCR9, and (2) one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CXCR4, CXCR5, CXCR6, CX3CR1, PSA, CEA, CGA, DHEA, NSE, PAP, prolactin and B7-H3.
[0072] In another further embodiment, said carcinoma is colorectal cancer and wherein the one or more cancer markers comprise (1) CCL25 or CCR9 or both CCL25 and CCR9, and (2) one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CXCR4, CXCR5, CXCR6, CX3CR1, seprase polypeptide, anti-p53, osteopontin, and ferritin.
[0073] In another further embodiment, said carcinoma is ovarian cancer and wherein the one or more cancer markers comprise (1) CCL25 or CCR9 or both CCL25 and CCR9, and (2) one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CXCR4, CXCR5, CXCR6, CX3CR1, cancer antigen 125 (CA-125), HE-4, OVX-1 macrophage colony stimulating factor (M-CSF) and lysophosphatidyl choline.
[0074] In another further embodiment, said carcinoma is lung cancer and wherein the one or more cancer markers comprise (1) CCL25 or CCR9 or both CCL25 and CCR9, and (2) one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CCL25, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CCR9, CXCR4, CXCR5, CXCR6, CX3CR1, kinesin family member 4A (KIF4A), Neural pentraxin I (NPTX1), fibroblast growth factor receptor 1 oncogene partner (FGFR1OP) protein and CEA.
[0075] In another further embodiment, said carcinoma is pancreatic cancer or gastric cancer and wherein the one or more cancer markers comprise (1) CCL25 or CCR9 or both CCL25 and CCR9, and (2) one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CXCR4, CXCR5, CXCR6, CX3CR1 and CEA.
[0076] In another further embodiment, the carcinoma is brain cancer, pituitary cancer or bone cancer and wherein one or more cancer markers further comprises one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CXCR4, CXCR5, CXCR6 and CX3CR1.
[0077] In some other embodiments, the biological sample is a plasma sample, a saliva sample or a urine sample.
[0078] In the context of the present application, the term "detecting" is intended to encompass predictions and likelihood analysis. The present method is intended to be used clinically in making decisions concerning treatment modalities, including therapeutic intervention, diagnostic criteria such as disease stages, and disease monitoring and surveillance for cancer. According to the present application, an intermediate result for examining the condition of a subject may be provided. Such intermediate result may be combined with additional information to assist a doctor, nurse, or other practitioner to diagnose that a subject suffers from the disease. Alternatively, the present application may be used to detect cancerous cells in a subject-derived tissue, and provide a doctor with useful information to diagnose that the subject suffers from the disease. The subject is preferably a human, but may also include other mammals such as non-human primate, mouse, rat, dog, cat, horse, and cow.
Method for Assessing the Prognosis of a Subject Having Cancer
[0079] The present method for detecting cancer may also be applied for assessing the prognosis of a patient with the cancer by comparing the expression level of one or more cancer markers in a patient-derived biological sample with that of a reference sample.
[0080] Therefore, another aspect of the present application relates to a method for assessing the prognosis of a subject with a cancer, comprising: determining the expression level of one or more cancer markers in a biological sample from said subject, and comparing the level of expression of said one or more cancer markers in said biological sample to a control level of expression of said one or more cancer markers, wherein a higher level of expression of said one or more cancer markers in the biological sample relative to said control level indicates that the prognosis of said subject is poor, wherein a lower or similar level of expression of said one or more cancer markers in said biological sample relative to said control level indicates that the prognosis of said subject is good, wherein a poor prognosis indicates that said cancer is of an aggressive or invasive type, wherein said cancer is blastoma, carcinoma, leukemia, lymphoma, melanoma, myeloma, sarcoma or germ cell tumor, and wherein said one or more cancer markers comprise CCL25 or CCR9 or both CCL25 and CCR9.
[0081] In one embodiment, said one or more cancer markers further comprises CXCL13 or CXCR5 or both CXCL13 and CXCR5. In another embodiment, said one or more cancer markers further comprises CXCL16 or CXCR6 or both CXCL16 and CXCR6.
[0082] In another embodiment, said one or more cancer markers further comprises (1) CXCL13 or CXCR5 or both CXCL13 and CXCR5, and (2) CXCL16 or CXCR6 or both CXCL16 and CXCR6.
[0083] In still another embodiment, said one or more cancer markers further comprises one or more cancer markers selected from the group consisting of CXCL13, CXCR5, CXCL16, CXCR6, CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CCL27, CXCL1, CXCL2, CXCL3, CXCL5, CXCL6, CXCL7, CXCL8, CXCL12, CX3CL1, CCR2, CCR7, CCR8, CCR10, CXCR1, CXCR2, CXCR4, CXCR7 and CX3CR1.
[0084] Alternatively, the level of one or more cancer markers in the biological sample may be measured over a spectrum of disease stages to assess the prognosis of the patient. An increase in the expression level of one or more cancer markers as compared to a normal control level indicates less favorable prognosis. A similarity in the expression level of one or more cancer markers as compared to a normal control level indicates a more favorable prognosis of the patient.
[0085] In some other embodiments, the biological sample is a plasma sample, a saliva sample or a urine sample.
Method for Monitoring the Course of Cancer Treatment
[0086] In certain embodiments, the level(s) of one or more cancer markers is used to monitor the course of treatment of cancer. In this method, a test biological sample is provided from a subject undergoing treatment for cancer. Preferably, multiple test biological samples are obtained from the subject at various time points before, during or after the treatment. The expression level of the cancer marker in the post-treatment sample may then be compared with the level of the cancer marker in the pre-treatment sample or, alternatively, with a reference sample (e.g., a normal control level). For example, if the post-treatment marker level is lower than the pre-treatment marker level, one can conclude that the treatment was efficacious. Likewise, if the post-treatment marker level is similar to, or the same as, the normal control marker level, one can also conclude that the treatment was efficacious.
[0087] An "efficacious" treatment is one that leads to a reduction in the level of a cancer marker or a decrease in size, prevalence or metastatic potential of cancer in a subject. When a treatment is applied prophylactically, "efficacious" means that the treatment retards or prevents occurrence of cancer or alleviates a clinical symptom of cancer. The assessment of cancer can be made using standard clinical protocols. Furthermore, the efficaciousness of a treatment can be determined in association with any known method for diagnosing or treating cancer. For example, cancer is routinely diagnosed histopathologically or by identifying symptomatic anomalies such as weight loss and loss of appetite.
[0088] Accordingly, another aspect of the present application relates to a method for monitoring the course of cancer treatment in a subject, comprising: determining the expression levels of one or more cancer markers in one or more biological samples obtained from said subject during or after said treatment, and comparing the level of expression of said one or more cancer markers in said one or more biological samples to a control level of expression of said one or more cancer markers, wherein said control level of said one or more cancer markers is a pre-treatment level of said one or more cancer markers in said subject or a predetermined reference level, wherein said treatment is deemed efficacious if said one or more cancer markers in said one or more biological samples is similar to or lower than said control level, wherein said cancer is blastoma, carcinoma, leukemia, lymphoma, melanoma, myeloma, sarcoma or germ cell tumor, and wherein said one or more cancer markers comprise CCL25 or CCR9 or both CCL25 and CCR9.
[0089] In one embodiment, said one or more cancer markers further comprises CXCL13 or CXCR5 or both CXCL13 and CXCR5. In another embodiment, said one or more cancer markers further comprises CXCL16 or CXCR6 or both CXCL16 and CXCR6.
[0090] In another embodiment, said one or more cancer markers further comprises (1) CXCL13 or CXCR5 or both CXCL13 and CXCR5 and (2) CXCL16 or CXCR6 or both CXCL16 and CXCR6.
[0091] In still another embodiment, said one or more cancer markers further comprises one or more cancer markers selected from the group consisting of CXCL13, CXCR5, CXCL16, CXCR6, CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CCL27, CXCL1, CXCL2, CXCL3, CXCL5, CXCL6, CXCL7, CXCL8, CXCL12, CX3CL1, CCR2, CCR7, CCR8, CCR10, CXCR1, CXCR2, CXCR4, CXCR7 and CX3CR1.
Cancer Markers
[0092] The term "cancer marker" as used herein, refers to or describes a polypeptide or a polynuceotide whose expression level, alone or in combination with other polypeptides or a polynuceotides, is correlated with cancer or prognosis of cancer. The correlation may relate to either an increased or decreased expression of the polypeptide or a polynuceotide. For example, the expression of the polypeptide or a polynuceotide may be indicative of cancer, or lack of expression of the polypeptide or a polynuceotide may be correlated with poor prognosis in a cancer patient.
[0093] The teen "expression level of a cancer marker" may be measured at the transcription level, in which case the presence and/or the amount of a polynucleotide is determined, or at the translation level, in which case the presence and/or the amount of a polypeptide is determined. Cancer marker expression may be characterized using any suitable method.
[0094] Examples of the cancer marker include CCL25, CCR9 and other chemokines and chemokine receptors such as CXCL1, CXCL2, CXCL3, CXCL4, CXCL6, CXCL7, CXCL8, CXCL9, CXCL10, CXCL11, CXCL12, CXCL 13, CXCL14, CXCL15, CXCL16, CXCR1, CXCR2, CXCR3, CXCR4, CXCR5, CXCR6, CXCR7, CCL1, CCL2, CCL3, CCL4, CCL5, CCL6, CCL7, CCL8, CCL9, CCL10, CCL11, CCL12, CCL13, CCL14, CCL15, CCL16, CCL17, CCL18, CCL19, CCL20, CCL21, CCL22, CCL24, CCL27, CCL28, CCR1, CCR2, CCR3, CCR4, CCR5, CCR6, CCR7, CCR8, CCR10, CCR11, XCL1, XCL2, XCR1, CX3CR1, CX3CL1, RNA binding motif 3 ("RBM3"), carcinoembryonic Antigen (CEA), prostate specific antigen (PSA), chromgranin A (CGA), dehydroepiandrosterone (DHEA), neuron-specific enolase (NSE), prostatic acid phosphatase (PAP), prolactin, B7-H3, seprase polypeptide, anti-p53, osteopontin, ferritin, lysophosphatidyl choline, kinesin family member 4A (KIF4A), Neural pentraxin I (NPTX1) and fibroblast growth factor receptor 1 oncogene partner (FGFR1OP) protein.
[0095] In one embodiment, the cancer markers used in the present application are selected from a melanoma marker panel that includes CCL25, CCR9, CXCL13, CXCR5, CXCL16, CXCR6, CCL27, CXCL1, CXCL2, CXCL3, CXCL5, CXCL6, CXCL7, CXCL8, CXCL12, CX3CL1, CCR10, CXCR1, CXCR2, CXCR4, and CX3CR1. The markers in the melanoma panel may be used for detecting melanoma or predicting the prognosis of a subject having melanoma.
[0096] In one embodiment, the cancer markers described above are selected from a carcinoma marker panel that includes CCL25, CCR9, CXCL13, CXCR5, CCL1, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CXCL16, CCR7, CCR8, CXCR4, CXCR6 and CX3CR1. The markers in the carcinoma panel may be used for detecting carcinoma or predicting the prognosis of a subject having carcinoma.
[0097] In another embodiment, the cancer markers described above are selected from a breast cancer marker panel that includes CCL25, CCR9, CXCL13, CXCR5, CCL1, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CXCL16, CCR7, CCR8, CXCR4, CXCR6, CX3CR1, HER2, RNA binding motif 3 ("RBM3") and carcinoembryonic Antigen (CEA). The markers in the breast cancer panel may be used for detecting breast cancer or predicting the prognosis of a subject having breast cancer.
[0098] In another embodiment, the cancer markers described above are selected from a prostate cancer marker panel that includes CCL25, CCR9, CXCL13, CXCR5, CXCL16, CXCR6, CCL1, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CCR7, CCR8, CXCR4, CX3CR1, PSA, CEA, CGA, DHEA, NSE, PAP, prolactin and B7-H3. The markers in the breast cancer panel may be used for detecting prostate cancer or predicting the prognosis of a subject having prostate cancer.
[0099] In another embodiment, the one or more cancer markers described above are selected from a colonrectal cancer marker panel that includes CCL25, CCR9, CXCL13, CXCR5, CXCL16, CXCR6, CCL1, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CCR7, CCR8, CXCR4, CX3CR1, seprase polypeptide, anti-p53, osteopontin, and ferritin. The markers in the colonrectal cancer panel may be used for detecting colonrectal cancer or predicting the prognosis of a subject having colonrectal cancer.
[0100] In another embodiment, the cancer markers described above are selected from an ovarian cancer marker panel that includes CCL25, CCR9, CXCL13, CXCR5, CXCL16, CXCR6, CCL1, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CCR7, CCR8, CXCR4, CX3CR1, cancer antigen 125 (CA-125), HE-4, OVX-1 macrophage colony stimulating factor (M-CSF) and lysophosphatidyl choline. The markers in the ovarian cancer panel may be used for detecting ovarian cancer or predicting the prognosis of a subject having ovarian cancer.
[0101] In another embodiment, the cancer markers described above are selected from a lung cancer marker panel that includes CCL25, CCR9, CXCL13, CXCR5, CXCL16, CXCR6, CCL1, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CCR7, CCR8, CXCR4, CX3CR1, kinesin family member 4A (KIF4A), Neural pentraxin I (NPTX1), fibroblast growth factor receptor 1 oncogene partner (FGFR1OP) protein and CEA. The markers in the lung cancer panel may be used for detectsing lung cancer or predicting the prognosis of a subject having lung cancer.
[0102] In another embodiment, the one or more cancer markers described above are selected from a pancreatic cancer or gastric marker panel that includes CCL25, CCR9, CXCL13, CXCR5, CXCL16, CXCR6, CCL1, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CCR7, CCR8, CXCR4, CX3CR1 and CEA. The markers in the pancreatic cancer panel may be used for detecting pancreatic or gastric cancer or predicting the prognosis of a subject having pancreatic cancer.
[0103] In another embodiment, the one or more cancer markers described above are selected from a brain cancer, pituitary cancer, bone cancer, pancratic cancer or gastric marker panel that comprises one or more cancer markers selected from the group consisting of CCL1, CCL2, CCL4, CCL17, CCL19, CCL21, CCL22, CXCL12, CXCL13, CXCL16, CX3CL1, CCR2, CCR7, CCR8, CXCR4, CXCR5, CXCR6 and CX3CR1.
Detection Methods
[0104] The expression of the cancer marker(s) can be determined at the transcription level (i.e., the amount of mRNA) or the translation level (i.e., the amount of protein). In certain embodiments, expression of the cancer marker(s) is determined at the mRNA level by quantitative RT-PCR, Northern blot or other methods known to a person of ordinary skill in the art. In other embodiments, the expression of the cancer marker(s) is determined at the protein level by ELISA, Western blot or other types of immuno-detection methods using anti-cancer marker antibodies, such as anti-CCL25 and anti-CCR9 antibodies, anti-CXCL13 and anti-CXCR5 antibodies, and anti-CXCL16 and anti-CXCR6 antibodies.
[0105] In certain embodiments, the anti-CCL25 and/or anti-CCR9 antibodies include antibodies that bind specifically to a CCL25 peptide or a CCR9 peptide. Examples of the CCL25 peptides include, but are not limited to, peptides consisting of, or comprising, one or more sequences selected from the group consisting of LAYHYPIGWAVL (SEQ ID NO:116), KRHRKVCGNPKSREVQRAMKLLDARNKVFAKLHH (SEQ ID NO:117), FEDCCLAYHYPIGWAVLRRA (SEQ ID NO:118), IQEVSGSCNLPAAIFYLPKRHRKVCGN (SEQ ID NO:119), AMKLLDAR (SEQ ID NO:120), KVFAKLHHN (SEQ ID NO:121), QAGPHAVKKL (SEQ ID NO:122), FYLPKRHRKVCGNP (SEQ ID NO:123) YLPKRHRKVCGNPK (SEQ ID NO:124), LPKRHRKVCGNPKS (SEQ ID NO:125), PKRHRKVCGNPKSR (SEQ ID NO:126), CGNPKSREVQRAMK (SEQ ID NO:127), GNPKSREVQRAMKL (SEQ ID NO:128), KFSNPISSSKRNVS (SEQ ID NO:129), PKSREV (SEQ ID NO:130), LHHNTQT (SEQ ID NO:131) and SSSKRN (SEQ ID NO:132). Examples of the CCR9 peptides include, but are not limited to, peptides consisting of, or comprising, one or more sequences selected from the group consisting of QFASHFLPP (SEQ ID NO:133), AAADQWKFQ (SEQ ID NO:134), TFMCKVVNSM (SEQ ID NO:135), IAICTMVYPS (SEQ ID NO:136) and VQTIDAYAMFISNCAVSTNIDICFQ (SEQ ID NO:137).
[0106] In other embodiments, the anti-CXCL13 and/or anti-CXCR5 antibodies include antibodies that bind specifically to a CXCL13 peptide or a CXCR5 peptide. Examples of the CXCL13 peptides include, but are not limited to, peptides consisting of, or comprising, one or more sequences selected from the group consisting of RSSSTLPVPVFKRKIP (SEQ ID NO:45), PRGNGCPRKEIIVWKK (SEQ ID NO:46), LPRGNGCPRKEIIVWK (SEQ ID NO:47), QILPRGNGCPRKEIIV (SEQ ID NO:48), ILPRGNGCPRKEIIVW (SEQ ID NO:49), RIQILPRGNGCPRKEI (SEQ ID NO:50), RGNGCPRKEIIVWKKN (SEQ ID NO:51), KRSSSTLPVPVFKRKI (SEQ ID NO:52), IQILPRGNGCPRKEII (SEQ ID NO:53), DRIQILPRGNGCPRKE (SEQ ID NO:54), RKRSSSTLPVPVFKRK (SEQ ID NO:55), RCRCVQESSVFIPRRF (SEQ ID NO:56), GNGCPRKEIIVWKKNK (SEQ ID NO:57), CVQESSVFIPRRFIDR (SEQ ID NO:58), IDRIQILPRGNGCPRK (SEQ ID NO:59), LRCRCVQESSVFIPRR (SEQ ID NO:60), FIDRIQILPRGNGCPR (SEQ ID NO:61), RCVQESSVFIPRRFID (SEQ ID NO:62), CRCVQESSVFIPRRFI (SEQ ID NO:63), QESSVFIPRRFIDRIQ (SEQ ID NO:64), RFIDRIQILPRGNGCP (SEQ ID NO:65), VQESSVFIPRRFIDRI (SEQ ID NO:66), ESSVFIPRRFIDRIQI (SEQ ID NO:67), SLRCRCVQESSVFIPR (SEQ ID NO:68), NGCPRKEIIVWKKNKS (SEQ ID NO:69), PQAEWIQRMMEVLRKR (SEQ ID NO:70), RRFIDRIQILPRGNGC (SEQ ID NO:71), LRKRSSSTLPVPVFKR (SEQ ID NO:72), VQESSVFIPRR (SEQ ID NO:73, EWIQRMMEVLRKRSSSTLPVPVFKRK (SEQ ID NO:74), KKNK (SEQ ID NO:75), RKRSSS (SEQ ID NO:76), RGNGCP (SEQ ID NO:77), VYYTSLRCRCVQESSVFIPRR (SEQ ID NO:78), DRIQILP (SEQ ID NO:79), RKEIIVW (SEQ ID NO:80) and KSIVCVDPQ (SEQ ID NO:81). Examples of the CXCR5 peptides include, but are not limited to, peptides consisting of, or comprising, one or more sequences selected from the group consisting of TSLVENHLCPATE (SEQ ID NO:82), EGSVGWVLGTFLCKT (SEQ ID NO:83), LPRCTFS (SEQ ID NO:84), LARLKAVDNT (SEQ ID NO:85) and MASFKAVFVP (SEQ ID NO:86).
[0107] The anti-CXCL16 and/or anti-CXCR6 antibodies include antibodies that bind specifically to a CXCL16 peptide or a CXCR6 peptide. Examples of the CXCL16 peptides include, but are not limited to, peptides consisting of, or comprising, one or more sequences selected from the group consisting of AAGPEAGENQKQPEKN (SEQ ID NO:87), SQASEGASSDIHTPAQ (SEQ ID NO:88), STLQSTQRPTLPVGSL (SEQ ID NO:89), SWSVCGGNKDPWVQEL (SEQ ID NO:90), GPTARTSATVPVLCLL (SEQ ID NO:91), SGIVAHQKHLLPTSPP (SEQ ID NO:92), RLRKHL (SEQ ID NO:93), LQSTQRP (SEQ ID NO:94), SSDKELTRPNETT (SEQ ID NO:95), AGENQKQPEKNA (SEQ ID NO:96), NEGSVT (SEQ ID NO:97), ISSDSPPSV (SEQ ID NO:98), CGGNKDPW (SEQ ID NO:99), LLPTSPPISQASEGASSDIHT (SEQ ID NO:100), STQRPTLPVGSLSSDKELTRPNETTIHT (SEQ ID NO:101), SLAAGPEAGENQKQPEKNAGPTARTSA (SEQ ID NO:102), TGSCYCGKR (SEQ ID NO:103), DSPPSVQ (SEQ ID NO:104), RKHLRAYHRCLYYTRFQLLSWSVCGG (SEQ ID NO:105), WVQELMSCLDLKECGHAYSGIVAHQKHLLPTSPPISQ (SEQ ID NO:106), SDIHTPAQMLLSTLQ (SEQ ID NO:107), RPTLPVGSL (SEQ ID NO:108), TAGHSLAAG (SEQ ID NO:109), GKRISSDSPPSVQ (SEQ ID NO:110) and KDPWVQELMSCLDLKECGHAYSGIVAHQKH (SEQ ID NO:111). Examples of the CXCR6 peptides include, but are not limited to, peptides consisting of, or comprising, one or more sequences selected from the group consisting of HQDFLQFSKV (SEQ ID NO:112), AGIHEWVFGQVMCK (SEQ ID NO:113), PQIIYGNVFNLDKLICGYHDEAI (SEQ ID NO:114) and YYAMTSFHYTIMVTEA (SEQ ID NO:115).
[0108] In one embodiment, the antibody is conjugated to a solid support. By "solid support" is meant a non-aqueous matrix to which an antibody of the present application can adhere or attach. Examples of solid phases encompassed herein include those formed partially or entirely of glass (e.g., controlled pore glass), polysaccharides (e.g., agarose), polyacrylamides, silicones, and plastics such as polystyrene, polypropylene and polyvinyl alcohol.
Enzyme-Linked Immunosorbent Assay (ELISA)
[0109] In certain embodiments, the cancer markers are detected using enzyme-linked immunosorbent assay (ELISA) which is typically carried out using antibody coated assay plate or wells. Commonly used ELISA assay employs either a sandwich immunoassay or a competitive binding immunoassay.
[0110] Briefly, a sandwich immunoassay is a method using two antibodies, which bind to different sites on the antigen or ligand. The primary antibody, which is highly specific for the antigen, is attached to a solid surface. The antigen is then added followed by addition of a second antibody referred to as the detection antibody. The detection antibody binds the antigen to a different epitope than the primary antibody. As a result the antigen is `sandwiched` between the two antibodies. The antibody binding affinity for the antigen is usually the main determinant of immunoassay sensitivity. As the antigen concentration increases the amount of detection antibody increases leading to a higher measured response. The standard curve of a sandwich-binding assay has a positive slope. To quantify the extent of binding different reporters can be used. Typically an enzyme is attached to the secondary antibody which must be generated in a different species than primary antibodies (i.e. if the primary antibody is a rabbit antibody than the secondary antibody would be an anti-rabbit from goat, chicken, etc., but not rabbit). The substrate for the enzyme is added to the reaction that forms a colorimetric readout as the detection signal. The signal generated is proportional to the amount of target antigen present in the sample.
[0111] The antibody linked reporter used to measure the binding event determines the detection mode. A spectrophotometric plate reader may be used for colorimetric detection. Several types of reporters have been recently developed in order to increase sensitivity in an immunoassay. For example, chemiluminescent substrates have been developed which further amplify the signal and can be read on a luminescent plate reader. Also, a fluorescent readout where the enzyme step of the assay is replaced with a fluorophor tagged antibody is becoming quite popular. This readout is then measured using a fluorescent plate reader.
[0112] A competitive binding assay is based upon the competition of labeled and unlabeled ligand for a limited number of antibody binding sites. Competitive inhibition assays are often used to measure small analytes. These assays are also used when a matched pair of antibodies to the analyte does not exist. Only one antibody is used in a competitive binding ELISA. This is due to the steric hindrance that occurs if two antibodies would attempt to bind to a very small molecule. A fixed amount of labeled ligand (tracer) and a variable amount of unlabeled ligand are incubated with the antibody. According to law of mass action the amount of labeled ligand is a function of the total concentration of labeled and unlabeled ligand. As the concentration of unlabeled ligand is increased, less labeled ligand can bind to the antibody and the measured response decreases. Thus the lower the signal, the more unlabeled analyte there is in the sample. The standard curve of a competitive binding assay has a negative slope.
Microbeads
[0113] In certain other embodiments, the cancer markers are detected using antibody coated microbeads. In some embodiments, the microbeads are magnetic beads. In other embodiments, the beads are internally color-coded with fluorescent dyes and the surface of the bead is tagged with an anti-cancer marker antibody (e.g., an anti-CCL25 or anti-CCR9 antibody) that can bind a cancer marker in a test sample. The cancer marker, in turn, is either directly labeled with a fluorescent tag or indirectly labeled with an anti-marker antibody conjugated to a fluorescent tag. Hence, there are two sources of color, one from the bead and the other from the fluorescent tag. Alternatively, the beads can be internally coded by different sizes.
[0114] By using a blend of different fluorescent intensities from the two dyes, as well as beads of different sizes, the assay can measure up to hundreds of different cancer markers. During the assay, a mixture containing the color/size-coded beads, fluorescence labeled anti-marker antibodies, and the sample are combined and injected into an instrument that uses precision fluidics to align the beads. The beads then pass through a laser and, on the basis of their color or size, either get sorted or measured for color intensity, which is processed into quantitative data for each reaction.
[0115] When samples are directly labeled with fluorophores, the system can read and quantitate only fluorescence on beads without removing unbound fluorophores in solution. The assays can be multiplexed by differentiating various colored or sized beads. Real time measurement is achievable when a sample is directly required for unlabeled samples. Standard assay steps include incubation of a sample with anti-marker antibody coated beads, incubation with biotin or fluorophore-labeled secondary antibody, and detection of fluorescence signals. Fluorescent signals can be developed on bead (by adding streptavidin-fluorophore conjugates for biotinylated secondary antibody) and read out by a bead analyzer. Depending on the anti-marker immobilized on the bead surface, a bead-based immunoassay can be a sandwich type or a competitive type immunoassay.
Test Stick
[0116] In some other embodiments, the cancer markers in a liquid biosample are detected using a test stick. The test stick typically contains a fluid impermeable housing and a fluid permeable "stick" having one or more detection zones. In one embodiment, each detection zone contains a dried binding reagent that binds to a cancer marker in a biosample. In another embodiment, the dried binding reagent is a labeled binding reagent. In another embodiment, the test stick may further comprise a control zone to indicate that the assay test has been carried out satisfactorily, namely the reagents were present in the test stick and that they become mobilized during running the test and have been transported along the flow path. The control zone can also indicate that the reagents within the device are capable of immunochemical interactions, confirming the chemical integrity of the device. This is important when considering the storage and shipment of the device under desiccated conditions within a certain temperature range. The control zone is typically positioned downstream from the detection zone(s) and may, for example, comprise an immobilized binding reagent for a labeled binding reagent. The labeled binding reagent may be present in a mobilizable form upstream from the control zone and detection zone. The labeled binding reagent may be the same or different to the labeled binding reagent for the cancer marker.
[0117] In one embodiment, the test stick comprise a porous sample receiver in fluid connection with and upstream from one or more flow-paths. The porous sample receiver may be common to all assays. Thus a fluid sample applied to the common sample application region of the device is able to travel along the one or more flow-paths to the respective detection zones. The porous sample receiver may be provided within a housing or may at least partially extend out of said housing and may serve for example to collect a body fluid. The porous sample receiver may also act as a fluid reservoir. The porous sample receiving member can be made from any bibulous, porous or fibrous material capable of absorbing liquid rapidly. The porosity of the material can be unidirectional (i.e. with pores or fibers running wholly or predominantly parallel to an axis of the member) or multidirectional (omnidirectional, so that the member has an amorphous sponge-like structure). Porous plastics material, such as polypropylene, polyethylene (preferably of very high molecular weight), polyvinylidene fluoride, ethylene vinylacetate, acrylonitrile and polytetrafluoro-ethylene can be used. Other suitable materials include glass-fiber.
[0118] If desired, an absorbent "sink" can be provided at the distal end of the carrier material. The absorbent sink may comprise, for example, Whatman 3MM chromatography paper, and should provide sufficient absorptive capacity to allow any unbound labeled binding reagent to wash out of the detection zone(s). As an alternative to such a sink it can be sufficient to have a length of porous solid phase material which extends beyond the detection zone(s).
[0119] Following the application of a binding reagent to a detection zone, the remainder of the porous solid phase material may be treated to block any remaining binding sites. Blocking can be achieved by treatment for example with protein (e.g. bovine serum albumin or milk protein), or with polyvinyl alcohol or ethanolamine, or combinations thereof. To assist the free mobility of the labeled binding reagent when the porous carrier is moistened with the sample, the porous carrier may further comprise a sugar such as sucrose or lactose and/or other substances, such as polyvinyl alcohol (PVA) or polyvinyl pyrrolidone (PVP). Such material may be deposited, for example, as an aqueous solution in the region to which the labeled binding reagent is to be applied. Such materials could be applied to the porous carrier as a first application followed by the application of the label; alternatively, such materials could be mixed with the label and applied to the porous carrier or combinations of both. Such material may be deposited upstream from or at the labeled binding reagent.
[0120] Alternatively, the porous carrier may not be blocked at the point of manufacture; instead the means for blocking the porous carrier are included in a material upstream from the porous carrier. On wetting the test strip, the means for blocking the porous carrier are mobilized and the blocking means flow into and through the porous carrier, blocking as the flow progresses. The blocking means include proteins such as BSA and casein as well as polymers such as PVP, PVA as well as sugars and detergents such as Triton-X100. The blocking means could be present in the macroporous carrier material.
[0121] The dried binding reagents may be provided on a porous carrier material provided upstream from a porous carrier material comprising the detection zone. The upstream porous carrier material may be macroporous. The macroporous carrier material should be low or non-protein-binding, or should be easily blockable by means of reagents such as BSA or PVA, to minimize non-specific binding and to facilitate free movement of the labeled reagent after the macroporous body has become moistened with the liquid sample. The macroporous carrier material can be pre-treated with a surface active agent or solvent, if necessary, to render it more hydrophilic and to promote rapid uptake of the liquid sample. Suitable materials for a macroporous carrier include plastic materials such as polyethylene and polypropylene, or other materials such as paper or glass-fiber. In the case that the labeled binding reagent is labeled with a detectable particle, the macroporous body may have a pore size at least ten times greater than the maximum particle size of the particle label. Larger pore sizes give better release of the labeled reagent. As an alternative to a macroporous carrier, the labeled binding reagent may be provided on a non-porous substrate provided upstream from the detection zone, said non-porous substrate forming part of the flow-path. In another embodiment, the test stick may further comprise a sample receiving member for receiving the fluid sample. The sample receiving member may extend from the housing.
[0122] The housing may be constructed of a fluid impermeable material. The housing will also desirably exclude ambient light. The housing will be considered to substantially exclude ambient light if less than 10%, preferably less than 5%, and most preferably less than 1%, of the visible light incident upon the exterior of the device penetrates to the interior of the device. A light-impermeable synthetic plastics material such as polycarbonate, ABS, polystyrene, polystyrol, high density polyethylene, or polypropylene containing an appropriate light-blocking pigment is a suitable choice for use in fabrication of the housing. An aperture may be provided on the exterior of the housing which communicates with the assay provided within the interior space within the housing. Alternatively, the aperture may serve to allow a porous sample receiver to extend from the housing to a position external from the housing.
Microarray
[0123] In other embodiments, the cancer markers are detected by a protein microarray containing immobilized cancer marker-specific antibodies on its surface. The microarray can be used in a "sandwich" assay in which the antibody on the microarray captures a cancer marker in the test sample and the captured marker is detected by a labeled secondary antibody that specifically binds to the captured marker. In a preferred embodiment, the secondary antibody is biotinylated or enzyme-labeled. The detection is achieved by subsequent incubation with a streptavidin-fluorophore conjugate (for fluorescence detection) or an enzyme substrate (for colorimetric detection).
[0124] Typically, a microarray assay contains multiple incubation steps, including incubation with the samples and incubation with various reagents (e.g., primary antibodies, secondary antibodies, reporting reagents, etc.). Repeated washes are also needed between the incubation steps. In one embodiment, the microarray assays is performed in a fast assay mode that requires only one or two incubations. It is also conceivable that the formation of a detectable immune complex (e.g., a captured cancer marker/anti-marker antibody/label complex) may be achieved in a single incubation step by exposing the protein microarray to a mixture of the sample and all the necessary reagents. In one embodiment, the primary and secondary antibodies are the same antibody.
[0125] In another embodiment, the protein microarray provides a competitive immunoassay. Briefly, a microarray comprising immobilized anti-marker antibodies is incubated with a test sample in the presence of a labeled cancer marker standard. The labeled cancer marker competes with the unlabeled cancer marker in the test sample for the binding to the immobilized antigen-specific antibody. In such a competitive setting, an increased concentration of the specific cancer marker in the test sample would lead to a decreased binding of the labeled cancer marker standard to the immobilized antibody and hence a reduced signal intensity from the label.
[0126] The microarray can be processed in manual, semi-automatic or automatic modes. Manual mode refers to manual operations for all assay steps including reagent and sample delivery onto microarrays, sample incubation and microarray washing. Semi-automatic modes refer to manual operation for sample and reagent delivery onto microarray, while incubation and washing steps operate automatically. In an automatic mode, three steps (sample/reagent delivery, incubation and washing) can be controlled by a computer or an integrated breadboard unit with a keypad. For example, the microarray can be processed with a ProteinArray Workstation (PerkinElmer Life Sciences, Boston, Mass.) or Assay 1200®. Workstation (Zyomyx, Hayward, Calif.). Scanners by fluorescence, colorimetric and chemiluminescence, can be used to detect microarray signals and capture microarray images. Quantitation of microarray-based assays can also be achieved by other means, such as mass spectrometry and surface plasma resonance. Captured microarray images can be analyzed by stand-alone image analysis software or with image acquisition and analysis software package. For example, quantification of an antigen microarray can be achieved with a fluorescent PMT-based scanner--ScanArray 3000 (General Scanning, Watertown, Mass.) or colorimetric CCD-based scanner--VisionSpot (Allied Biotech, Ijamsville, Md.). Typically, the image analysis would include data acquisition and preparation of assay report with separate software packages. To speed up the whole assay process from capturing an image to generating an assay report, all the analytical steps including image capture, image analysis, and report generation, can be confined in and/or controlled by one software package. Such an unified control system would provide the image analysis and the generation of assay report in a user-friendly manner.
Implantable Biosensors
[0127] In other embodiments, the cancer markers are detected using implantable biosensors. Biosensors are electronic devices that produce electronic signals as the result of biological interactions. In one embodiment, the biosensors use antibodies, receptors, nucleic acids, or other members of a binding pair to bind with a cancer marker, which is typically the other member of the binding pair. Biosensors may be used with a blood sample to determine the presence of a cancer marker without the need for sample preparation and/or separation steps typically required for the automated immunoassay systems.
[0128] In one embodiment, the sensor is a nanoscale device. The sensor system includes a biological recognition element attached to a nanowire and a detector that is capable of determining a property associated with the nanowire. The biological recognition element is one member of a binding pair (e.g., a receptor of the cancer marker or an anti-cancer marker antibody) where the cancer marker being measured is the other member of the binding pair. Preferably, the nanowire sensor includes a semiconductor nanowire with an exterior surface formed thereon to form a gate electrode and a first end in electrical contact with a conductor to form a source electrode and a second end in contact with a conductor to form a drain electrode. In one embodiment the sensor is a field effect transistor comprising a substrate formed of an insulating material, a source electrode, a drain electrode and a semiconductor nanowire disposed there between with a biological recognition element attached on a surface of the nanowire. When a binding event occurs between the biological recognition element and its specific binding partner, a detectable change is caused in a current-voltage characteristic of the field effect transistor.
[0129] In another embodiment, the sensor system includes an array of sensors. One or more of the sensors in the array is associated with a protective member that prevents the associated sensor from interacting with the surrounding environment. At a selected time, the protective member may be disabled, thereby allowing the sensor to begin operating to interact with the surrounding fluid or tissue so that the biological recognition element can interact with the other member of its binding pair if that pair member is present.
[0130] In another embodiment, the protective member is formed of a conductive material that can oxidize, is biocompatible, bio-absorbable, and that may be dissolved in solution such as blood upon application of an electric potential. For example, a sensor may be farmed within a well of a substrate that is capped by a conductive material such as a biocompatible metal or an electrically-erodible polymer. In another embodiment, the protective member is formed using a material that dissolves over a predetermined period of time.
Mass Spectrometry
[0131] In other embodiments, the cancer markers are detected using mass spectrometry (MS) such as MALDI/TOF (time-of-flight), SELDI/TOF, liquid chromatography-mass spectrometry (LC-MS), gas chromatography-mass spectrometry (GC-MS), high performance liquid chromatography-mass spectrometry (HPLC-MS), capillary electrophoresis-mass spectrometry, nuclear magnetic resonance spectrometry, or tandem mass spectrometry (e.g., MS/MS, MS/MS/MS, ESI-MS/MS, etc.).
[0132] Mass spectrometry methods are well known in the art and have been used to quantify and/or identify biomolecules, such as proteins. Further, mass spectrometric techniques have been developed that permit at least partial de novo sequencing of isolated proteins. In certain embodiments, a gas phase ion spectrophotometer is used. In other embodiments, laser-desorption/ionization mass spectrometry is used to analyze the sample. Modem laser desorption/ionization mass spectrometry ("LDI-MS") can be practiced in two main variations: matrix assisted laser desorption/ionization ("MALDI") mass spectrometry and surface-enhanced laser desorption/ionization ("SELDI"). In MALDI, the analyte is mixed with a solution containing a matrix, and a drop of the liquid is placed on the surface of a substrate. The matrix solution then co-crystallizes with the biological molecules. The substrate is inserted into the mass spectrometer. Laser energy is directed to the substrate surface where it desorbs and ionizes the biological molecules without significantly fragmenting them. In SELDI, the substrate surface is modified so that it is an active participant in the desorption process. In one embodiment, the surface is derivatized with adsorbent and/or capture reagents that selectively bind the protein of interest. In another embodiment, the surface is derivatized with energy absorbing molecules that are not desorbed when struck with the laser. In another embodiment, the surface is derivatized with molecules that bind the protein of interest and that contain a photolytic bond that is broken upon application of the laser. In each of these methods, the derivatizing agent generally is localized to a specific location on the substrate surface where the sample is applied. The two methods can be combined by, for example, using a SELDI affinity surface to capture an analyte and adding matrix-containing liquid to the captured analyte to provide the energy absorbing material.
[0133] Detection of the presence of a cancer marker will typically involve detection of signal intensity. This, in turn, can reflect the quantity and character of a polypeptide bound to the substrate. For example, in certain embodiments, the signal strength of peak values from spectra of a first sample and a second sample can be compared (e.g., visually, by computer analysis etc.), to determine the relative amounts of particular biomolecules. Software programs such as the Biomarker Wizard program (Ciphergen Biosystems, Inc., Fremont, Calif.) can be used to aid in analyzing mass spectra. The mass spectrometers and their techniques are well known to those of skill in the art.
[0134] A person skilled in the art understands that any of the components of a mass spectrometer (e.g., desorption source, mass analyzer, detect, etc.) and varied sample preparations can be combined with other suitable components or preparations described herein, or to those known in the art. For example, in some embodiments a control sample may contain heavy atoms (e.g. 13C) thereby permitting the test sample to be mixed with the known control sample in the same mass spectrometry run.
[0135] In one preferred embodiment, a laser desorption time-of-flight (TOF) mass spectrometer is used. In laser desorption mass spectrometry, a substrate with a bound marker is introduced into an inlet system. The marker is desorbed and ionized into the gas phase by laser from the ionization source. The ions generated are collected by an ion optic assembly, and then in a time-of-flight mass analyzer, ions are accelerated through a short high voltage field and let drift into a high vacuum chamber. At the far end of the high vacuum chamber, the accelerated ions strike a sensitive detector surface at a different time. Since the time-of-flight is a function of the mass of the ions, the elapsed time between ion formation and ion detector impact can be used to identify the presence or absence of molecules of specific mass to charge ratio.
[0136] In some embodiments the relative amounts of one or more cancer markers present in a first or second sample is determined, in part, by executing an algorithm with a computer. The algorithm identifies at least one peak value in the first mass spectrum and the second mass spectrum. The algorithm then compares the signal strength of the peak value of the first mass spectrum to the signal strength of the peak value of the second mass spectrum of the mass spectrum. The relative signal strengths are an indication of the amount of the cancer marker that is present in the first and second samples. A standard containing a known amount of a cancer marker can be analyzed as the second sample to better quantify the amount of the biomolecule present in the first sample. In certain embodiments, the identity of the cancer markers in the first and second sample can also be determined.
Determination of Standard Value, Specificity and Sensitivity
[0137] In the present application, the standard expression level of a cancer marker, such as the blood concentration of CCL25, can be determined statistically. For example, the blood concentration of CCL25 in healthy individuals can be measured to determine the standard blood concentration of CCL25 statistically. When a statistically sufficient population can be gathered, a value in the range of twice or three times the standard deviation (S.D.) from the mean value is often used as the standard value. Therefore, values corresponding to the mean value+2×S.D. or mean value+3×S.D. may be used as standard values. The standard values set as described theoretically comprise 90% and 99.7% of healthy individuals, respectively.
[0138] Alternatively, standard values can also be set based on the actual expression level (e.g., blood concentration of CCL25) in cancer patients. Generally, standard values set this way minimize the percentage of false positives, and are selected from a range of values satisfying conditions that can maximize detection sensitivity. Herein, the percentage of false positives refers to a percentage, among healthy individuals, of patients whose blood concentration of CCL25 is judged to be higher than a standard value. On the contrary, the percentage, among healthy individuals, of patients whose blood concentration of CCL25 is judged to be lower than a standard value indicates specificity. That is, the sum of the false positive percentage and the specificity is always 1. The detection sensitivity refers to the percentage of patients whose blood concentration of CCL25 is judged to be higher than a standard value, among all cancer patients within a population of individuals for whom the presence of cancer has been determined.
[0139] As used herein, the teem "test sensitivity" is the ability of a screening test to identify true disease, also characterized by being a test with high sensitivity has few false negatives, additionally a test independent of disease prevalence. The test sensitivity is calculated as true positive tests per total affected patients tested, expressed as a percentage.
[0140] The term "Test Specificity" is a screening test which is correctly negative in the absence of disease, has high specificity and few false positives, is independent of disease prevalence. The test specificity is calculated as true negative tests per unaffected individuals tested, expressed as a percentage.
[0141] The term "PPV" (Positive Predictive Value) is the percent of patients with positive test having disease, and thus assesses reliability of positive test. Calculation:
PPV=(True positive)/(True+False positives). 1.
[0142] The term "NPV" (Negative Predictive Value) refers to patients with negative test that do not have disease, and assesses reliability of negative test. Calculation:
NPV=(True negative)/(true and false negatives). 2.
[0143] As the relationship shown above indicates, each of the values for sensitivity, specificity, positive predictive value, and negative predictive value, which are indexes for evaluating the detection accuracy, varies depending on the standard value for judging the level of the blood concentration of CCL25.
[0144] A standard value is usually set such that the false positive ratio is low and the sensitivity is high. However, as also apparent from the relationship shown above, there is a trade-off between the false positive ratio and sensitivity. That is, if the standard value is decreased, the detection sensitivity increases. However, since the false positive ratio also increases, it is difficult to satisfy the conditions to have a "low false positive ratio". Considering this situation, for example, values that give the following predicted results may be selected as the preferable standard values in the present application: (1) standard values for which the false positive ratio is 50% or less (that is, standard values for which the specificity is not less than 50%) and (2) standard values for which the sensitivity is not less than 20%.
[0145] The standard values can be set using receiver operating characteristic (ROC) curve. An ROC curve is a graph that shows the detection sensitivity on the vertical axis and the false positive ratio (that is, "1--specificity") on the horizontal axis. A ROC curve can be obtained by plotting the changes in the sensitivity and the false positive ratio, which were obtained after continuously varying the standard value for determining the high/low degree of the blood concentration of a cancer marker, such as CCL25.
[0146] The "standard value" for obtaining the ROC curve is a value temporarily used for the statistical analyses. The "standard value" for obtaining the ROC curve can generally be continuously varied within a range that allows to cover all selectable standard values. For example, the standard value can be varied between the smallest and largest measured blood CCL25 values in an analyzed population.
[0147] Based on the obtained ROC curve, a preferable standard value to be used in the present application can be selected from a range that satisfies the above-mentioned conditions. Alternatively, a standard value can be selected based on a ROC curve produced by varying the standard values from a range that comprises most of the measured blood CCL25.
Kits for Detecting Cancer
[0148] Another aspect of the present application relates to a kit for detecting cancer, comprising: reagents for determining expression of CCL25 and/or CCR9 in a biological sample; and instructions for how to use said reagents, wherein said reagents comprise an anti-CCL25 antibody, an anti-CCR9 antibody, or both.
[0149] In a particular embodiment, said kit further comprises reagents for determining expression of CXCL13 and/or CXCR5 in a biological sample; and instructions for how to use said reagents, wherein said reagents comprise an anti-CXCL13 antibody, an anti-CXCR5 antibody, or both. In a further particular embodiment, said kit further comprises reagents for determining expression of CXCL16 and/or CXCR6 in a biological sample; and instructions for how to use said reagents, wherein said reagents comprise an anti-CXCL16 antibody, an anti-CXCR6 antibody, or both.
[0150] In another particular embodiment, said kit further comprises reagents for determining expression of CXCL16 and/or CXCR6 in a biological sample; and instructions for how to use said reagents, wherein said reagents comprise an anti-CXCL16 antibody, an anti-CXCR6 antibody, or both.
Example 1
In Vitro Analysis of CCL25 and CCR9 Expression and Activity in Various Carcinomas
[0151] As shown in FIG. 1, CCL25 is expressed by breast cancer tissue. Breast cancer tissue was stained with isotype control or anti-CCL25 antibodies. Magenta color shows CCL25 staining. An Aperio ScanScope CS system with a 40× objective captured digital images. A representative case of breast cancer indicated and immuno-intensity of CCL25.
[0152] FIG. 2 demonstrates CCL25 inhibition of cisplatin-induced reductions in breast cancer cell line growth is demonstrated. MDA-MB-231 cells were cultured with 0 or 100 ng/ml of CCL25 plus isotype control or anti-CCR9 Ab for 24 hours, along with increasing concentrations of cisplatin. Cell proliferation was determined by BrdU incorporation and assays were repeated 3 times and performed in triplicate. Asterisks indicate statistical significant differences (p<0.01) between CCL25-treated and untreated BrCa cells.
[0153] FIGS. 3A-B show that CCL25 protects breast cancer cells from cisplatin-induced apoptosis. MDA-MB-231 cells were cultured for 24 hours with 5 mg/ml of cisplatin alone or with 0 or 100 ng/ml CCL25 plus 1 mg/ml of anti-human CCR9 or isotype controls (A). Cells were harvested and stained with annexin V and propidium iodide (PI). Analysis by flow cytometry of the stained cells distinguished apoptotic (annexin V positive) cells from viable (no fluorescence) and necrotic (PI positive) cells. Asterisks indicate statistical significant differences (p<0.01) between CCL25-treated and untreated breast cancer cells. MDA-MB-231 cell line was cultured for 24 hours with 5 mg/ml cisplatin or with 0 or 100 ng/ml of CCL25 plus 1 mg/ml or anti-human CCR9 or isotype control Abs (B). Detection of apoptotic cells was carried out using the terminal deoxynucleotidyl transferase-mediated dUTP nick-end labeling (TUNEL) method. Apoptotic cells exhibited nuclear green fluorescence with a standard fluorescence filter set (520±20 nm). Asterisks indicate statistical significant differences (p<0.01) between cisplatin CCL25-treated and untreated breast cancer cell line.
[0154] FIGS. 4A-B show PI3K and Akt activation by CCL25-CCR9 interactions in a breast cancer cell line. MDA-MB-231 cells were tested for their ability to activate PI3K and Akt following treatment with CCL25, cisplatin and specific kinase inhibitors (wortmannin and PF-573,228). In situ total and phosphorylated PI3K and Akt levels were quantified by Fast Activated Cell-based ELISA before (0 minutes) or after (5 or 10 minutes) CCL25 stimulation in the presence of cisplatin and kinase inhibitors. The ratio ±SEM of active (phosphorylated) to total PI3K (A) or Akt (B) are presented in from 3 separate experiments performed in triplicate. Asterisks indicate statistical differences between untreated and CCL25-treated cells and CCL25+cisplatin-treated cells.
[0155] FIGS. 5A-B show GSK-3β and FKHR phosphorylation following CCL25 treatment of a breast cancer cell line. MDA-MB-231 cells were tested for their ability to phosphorylate GSK-3β and FKHR following treatment with CCL25, cisplatin and specific-kinase inhibitors (wortmannin and PF-573,228). In situ total and phosphorylated GSK-3β and FKHR levels were quantified by Fast Activated Cell-based ELISA before (0 minutes) or after (5 or 10 minutes) CCL25 stimulation in the presence of cisplatin and kinase inhibitors. The ratio of phosphorylated to total GSK-3β (A) or FKHR (B) are presented in ±SE from 3 separate experiments performed in triplicate. Asterisks indicate statistical differences (p<0.01) between untreated and CCL25-treated cells and CCL25+cisplatin-treated cells.
[0156] FIG. 6 shows CCR9 and CCL25 expression by ovarian cancer tissues. Ovarian cancer tissues from non-neoplastic (n=8), serous adenocarcinoma (n=9), serous papillary cystadenoma (n=1), endometrioid adenocarcinoma (n=5), mucinous adenocarcinoma (n=2), Cystadenoma (n=3), mucinous boderline adenocarcinoma (n=1), clear cell carcinoma (n=5), granulosa cell tumor (n=3), dysgerminoma (n=3), transitional cell carcinoma (n=3), Brenner tumor (n=1), yolk sac tumor (n=4), adenocarcinoma (n=1) and fibroma (n=2) were stained with isotype control or anti-CCR9 and CCL25 antibodies. Brown (DAB) color shows CCR9 staining and Magenta color show CCL25. An Aperio ScanScope CS system with a 40× objective captured digital images of each slide. Representative cases show immunointensities of CCR9 and CCL25.
[0157] FIGS. 7A-B show an analysis of CCL25 expression by ovarian cancer tissues. CCL25 expression were analyzed and presented by modified box plot (A). Lower, middle and upper lines, respectively, in the box represent the first quartile (Q1), Median (Q2) and third quartile (Q3). Upper and lower whiskers represent the median ±1.5 (Q3-Q1). Significant differences from non-neoplastic are indicated in the lower panel. The table (B) shows respective p values or significant differences between non-neoplastic tissue (NN) and serous adenocarcinoma (SA), endometrioid adenocarcinoma (EC), mucinous adenocarcinoma (MA), cystadenoma (C), mucinous boderline adenocarcinoma (MBA), clear cell carcinoma (CCC), granulosa cell tumor (GCT), dysgerminoma (D), transitional cell carcinoma (TCC), Brenner tumor (BT), yolk sac tumor (YST), adenocarcinoma (A), and fibroma (F).
[0158] FIGS. 8A-B show an analysis of CCR9 expression by ovarian cancer tissues. CCR9 expression was analyzed and presented by modified box plot (A). Lower, middle and upper lines, respectively, in the box represent the first quartile (Q1), Median (Q2) and third quartile (Q3). Upper and lower whiskers represent the median ±1.5 (Q3-Q1 significant differences from non-neoplastic are indicated in the lower panel. The table (B) shows respective p values or significant differences between non-neoplastic tissue (NN) and serous adenocarcinoma (SA), endometrioid adenocarcinoma (EC), mucinous adenocarcinoma (MA), cystadenoma (C), mucinous boderline adenocarcinoma (MBA), clear cell carcinoma (CCC), granulosa cell tumor (GCT), dysgerminoma (D), transitional cell carcinoma (TCC), Brenner tumor (BT), yolk sac tumor (YST), adenocarcinoma (A), and fibroma (F).
[0159] FIGS. 9A-B show CCR9 and CCL25 expression by ovarian cancer cell lines. Ovarian cancer cells were stained with fluorescein (FITC)-conjugated anti-CCR9 or FITC-conjugated isotype control antibody and analyzed by FACS (A). Ovarian cancer cells were stained with FITC-conjugated anti-CCR9, intracellular CCL25 was stained with phycoerythrin (PE)-conjugated anti-CCL25 antibody and nuclei were stained with Draq-5 (B). Merged data show the expression of CCR9 on the surface and CCL25 expression in the nucleus.
[0160] FIGS. 10A-B show hypoxia-regulated CCR9 mRNA and surface protein expression by ovarian cancer cells. Total RNA was isolated from SKOV-3 cell line under normoxic and hypoxic conditions or from normal primary ovary tissue. Quantitative RT-PCR analysis of CCR9 mRNA expression was performed in triplicate. The copies of transcripts are expressed relative to actual copies of 18S rRNA+SE (A). SKOV-3 cells under normoxia and hypoxia were stained with PE-conjugated isotype control antibody (Ab) (solid histogram) or PE-conjugated anti-CCR9 monoclonal Ab (open histogram) and quantified by flow cytometry (B). The mean fluorescent intensities of PE-positive cells are shown. Symbols indicate statistical significant (p<0.01) differences in CCR9 expression between normal tissue or isotype control and OvCa cells (@) or between normoxic and hypoxic cells (*).
[0161] FIGS. 11A-B show hypoxia-mediated and CCL25-mediated migration and invasion of SKOV-3 cells. SKOV-3 cells were tested for their ability to migrate toward chemotactic gradients of CCL25 (A). Cells were co-cultured with 1.0 μg/ml mouse anti-CCR9 antibody (Ab) or isotype control Ab during migration assays using 100 ng/ml of CCL25 under normoxic or hypoxic conditions. Also, SKOV-3 cells were tested for their ability to invade or translocate cross Matrigel® matrix in response to 100 ng/ml of CCL25 under hypoxic or normoxic conditions (B). Cells were co-cultured with 1.0 μg/ml monoclonal antibodies against CCR9 during invasion assays using 100 ng/ml of CCL25 under normoxic or hypoxic conditions. The number of cells (+SE) that migrated or invaded is shown with symbols that indicate significant (p<0.01) differences between CCL25-treated and untreated normoxic cells (#), CCL25-treated and untreated hypoxic cells (@), or similarly treated normoxic and hypoxic cells (*).
[0162] FIGS. 12A-B show CCL25-induced collagenase expression by SKOV-3 cells. Cells were tested for their ability to express collagenases (MMP-1, MMP-8, and MMP-13) mRNA and active protein. SKOV-3 cells were cultured for 24 hours alone, with 100 ng/ml of CCL25+1 μg/ml of isotype control antibody (Ab), or CCL25+1 μg/ml of mouse anti-CCR9 Ab under normoxic or hypoxic conditions. Total RNA was isolated and quantitative RT-PCR analysis was performed for mRNA expression of collagenases and transcript copies are presented relative to actual copies of 18S rRNA (A). Active collagenases were quantified by Fluorokine and Biotrak assays in conditioned media (B). Symbols indicate significant (p<0.01) differences between CCL25-treated and untreated normoxic cells (#), CCL25-treated and untreated hypoxic cells (@), or similarly treated normoxic and hypoxic cells (*).
[0163] FIGS. 13A-B show CCL25-induced gelatinase expression by SKOV-3 cells. Cells were tested for their ability to express gelatinases (MMP-2 and MMP-9) mRNA and active protein. SKOV-3 cells were cultured for 24 hours alone, with 100 ng/ml of CCL25+1 μg/ml of isotype control antibody (Ab), or CCL25+1 μg/ml of mouse anti-CCR9 Ab under normoxic or hypoxic conditions. Total RNA was isolated and quantitative RT-PCR analysis was performed for mRNA expression of gelatinases and transcript copies are presented relative to actual copies of 18S rRNA (A). Active gelatinases in conditioned media were quantified by Fluorokine and Biotrak assays (B). Symbols indicate significant (p<0.01) differences between CCL25-treated and untreated normoxic cells (#), CCL25-treated and untreated hypoxic cells (@), or similarly treated normoxic and hypoxic cells (*).
[0164] FIGS. 14A-B show CCL25-induced stromelysin expression by SKOV-3 cells. Cells were tested for their ability to express stromelysins (MMP-3, MMP-10, and MMP-11) mRNA and active protein. SKOV-3 cells were cultured for 24 hours alone, with 100 ng/ml of CCL25+1 μg/ml of isotype control antibody (Ab), or CCL25+1 μg/ml of mouse anti-CCR9 Ab under normoxic or hypoxic conditions. Total RNA was isolated and quantitative RT-PCR analysis was performed for mRNA expression of stromelysins and transcript copies are presented relative to actual copies of 18S rRNA (A). Active stromelysins were quantified by Fluorokine and Biotrak assays in conditioned media (B). Symbols indicate significant (p<0.01) differences between CCL25-treated and untreated normoxic cells (#), CCL25-treated and untreated hypoxic cells (@), or similarly treated normoxic and hypoxic cells (*).
[0165] FIG. 15 shows CCR9 expression by prostate cancer cell lines. Prostate cancer cell lines (C4-2B, LNCaP, and PC3) and normal prostate cells (RWPE-1) were stained with FITC-conjugated anti-human CCR9 (green) and 7AAD (nuclear stain; red). Positively stained cells were imaged and quantified by Amnis ImageStream. Panels on the right show the mean fluorescence intensity of CCR9 staining.
[0166] FIGS. 16A-D show CCR9 expression by prostate tissue. Tissue microarrays (TMA) were obtained from the National Institutes of Health (NIH), National Cancer Institute (NCI) and the University of Alabama at Birmingham and stained for CCR9. Aperio Scan Scope system with a 40× objective captured digital images of each slide. Representative cases of prostate cancer (CaP)(A), matched benign prostate tissue (MB)(B) and negative controls are indicated and intensities of CCR9 for all tissues scanned and analyzed were quantified using ImageScope software (v. 6.25). FIG. 27D shows the CCR9 immunointensity between MB, benign prostatic hyperplasia (BPH), and prostate cancer (PCa). Asterisks indicate significant (p<0.01) differences in CCR9 immunointensity between MB, BPH, and PCa tissue.
[0167] FIGS. 17A-D show CCL25 expression by prostate cancer tissue. Neuroendocrine differentiation of endocrine-paracrine cell phenotypes frequently occurs in prostatic malignancies and has potential prognostic and therapeutic implications. Paracrine cell phenotypes can be considered to be an androgen-insensitive, post-mitotic subpopulation in the prostate and prostate cancer. FIG. 17A demonstrates the expression of CCL25 in paracrine pattern within prostate interepithelial neoplasia. The double-headed arrow points to multiple paracrine cells producing CCL25 (red); brown arrow points cells expressing CCR9 (Brown). FIG. 17B shown cell stained red for CCL25. Brown arrow points the cell NSE. FIGS. 17A and C are higher magnifications of FIGS. 17D and B, respectively.
[0168] FIG. 18 shows serum CCL25 levels in normal healthy donors or patients with prostatic disease. ELISA was used to quantify CCL25 in serum from normal healthy donors, prostate cancer (PCa), prostate interepithelial neoplasia (PIN), and benign prostate hyperplasia (BPH). Asterisks indicate significant differences (p<0.05) of CCL25 levels compared to normal healthy donors.
[0169] FIGS. 19A-C shows CCL25 expression by mouse bone marrow cells. Bone marrow cells from non-tumor bearing (A) and tumor-bearing (B) mice were aspirated and stained with FITC-conjugated anti-CCL25 antibody. Positively stained cells (C) were quantified by Amnis ImageStream. Image-based analysis was performed using IDEAS-software and indicated a 1.6 fold increase in CCL25 expression by bone marrow cells after prostate tumor challenge.
[0170] FIGS. 20A-B show CCR9-mediated prostate cancer cell migration (A) and invasion (B). LNCaP, PC3, and C4-2b cells were tested for their ability to migrate to no additions (open bar), 100 ng/mL of CCL25 (hashed bar), or 100 ng/mL of CCL25+1 μg/mL anti-CCL25 antibody (solid bar). The number of cells (±SEM) that migrated and invaded in response to CCL25 from the initial 104 cells used to seed the migration and invasion chamber, show migration was CCL25 dependent and inhibited by anti-CCL25 antibody blockade. Asterisks indicate significant differences (p<0.01) between no additions and CCL25-treated cells.]
[0171] FIG. 21 shows CCL25-induced active matrix metalloproteinase (MMP) expression by LNCaP, PC3, and C4-2b prostate cancer cell lines. Cells were cultured for 24 hours without (open boxes) or with 100 ng/mL CCL25 (solid boxes). MMP-1, MMP-2, MMP-13, MMP-9, MMP-10, and MMP-11 protein levels, in cultured supernatants, were determined by MMP activity assays. Asterisks show a significant (P<0.05) increase or decrease in MMP secretion by a CCL25-treated cell line compared with the untreated cell line.
[0172] FIGS. 22A-F show inhibition of bone metastasis of PC3 prostate cancer cell line by CCR9 knockdown. Mice were challenged with a luciferase- and doxycyclene-inducible CCR9-specific shRNA-expressing PC3 cell line (A, D). Mice were challenged with this cell line by intracardiac injection. Subsequently, mice received no additions or doxycycline (0.2 mg/mL) in drinking for 21 days. Metastasis and tumor growth was monitored every week by Caliper Xenogen 100 in vivo imaging system. There were no changes 24 hours post challenge (B, E), but three weeks after challenge significantly less CCR9 knockdown PC3 (F) cells grew as bone metastases than compared to CCR9-positive PC3 cells (C).
[0173] FIG. 23 shows serum CCL25 levels in lung cancer patients. CCL25 ELISAs were performed to quantify CCL25 levels in serum from patients diagnosed with adenocarcinoma (Adeno Ca; n=14), squamous cell carcinoma (SSC; n=17), and normal healthy donors (control; n=9). ELISAs were capable of detecting >5 pg/mL of CCL25. Solid circles indicate individual serum CCL25 levels and lines show median concentrations of each group. Asterisks indicate significant differences (p<0.01) between controls and groups with lung cancer.
[0174] FIGS. 24A-D show CCR9 expression by non-neoplastic lung and lung cancer tissues. Lung tissues from non-neoplastic (n=8)(A), adenocarcinoma (n=54)(B), and squamous cell carcinoma (n=24)(C) were stained with isotype control or anti-CCR9 antibodies. Brown (DAB) color show CCR9 staining. Aperio ScanScope CS system with a 40× objective captured digital images of each slide.
[0175] FIGS. 25A-D show CCR9-CCL25 expression by colon cancer tissues. Colon tissues from non-neoplastic (n=8) and adenocarcinoma (n=16) were stained with isotype control (A), anti-CCR9 (B) or anti-CCL25 (C) antibodies. Brown (DAB) stain indicates CCR9 positivity and magenta stain show CCL25 positivity. Aperio ScanScope CS system with a 40× objective captured digital images.
Example 2
Detecting Chemokine Expression Levels with Real Time-PCR Analysis Primer Design
[0176] Messenger RNA sequences for CXCL1, CXCL2, CXCL3, CXCL4, CXCL5, CXCL6, CXCL7, CXCL8, CXCL9, CXCL10, CXCL11, CXCL12, CXCL13, CXCL14, CXCL15, CXCL16, CXCR1, CXCR2, CXCR3, CXCR4, CXCR5, CXCR5a, CXCR5b, CXCR6, CXCR7, CCL1, CCL2, CCL3, CCL4, CCL5, CCL6, CCL7, CCL8, CCL9, CCL10, CCL11, CCL12, CCL13, CCL14, CCL15, CCL16, CCL17, CCL18, CCL19, CCL20, CCL21, CCL22, CCL24, CCL25, CCL25-1, CCL25-2, CCL27, CCL28, CCR1, CCR2, CCR3, CCR4, CCR5, CCR6, CCR7, CCR8, CCR9, CCR10, CCR11, XCL1, XCL2, XCR1, CX3CR1, or CX3CL1 were obtained from the NIH-NCBI gene bank database. Primers were designed using the BeaconJ 2.0 computer program. Thermodynamic analysis of the primers was conducted using computer programs: Primer PremierJ and MIT Primer 3. The resulting primer sets were compared against the entire human genome to confirm specificity.
Real Time PCR Analysis
[0177] Cancer cell lines (ATCC, Rockville, Md.) were cultured in RMPI-1640 containing 10% fetal calf serum supplemented with non-essential amino acids, L-glutamate, and sodium pyruvate (complete media). Primary tumor and normal-paired matched tissues were obtained from clinical isolates (Clinomics Biosciences, Frederick, Md. and UAB Tissue Procurement, Birmingham, Ala.). Messenger RNA (mRNA) was isolated from 106 cells using TriReagent (Molecular Research Center, Cincinnati, Ohio) according to manufacturer's protocols. Potential genomic DNA contamination was removed from these samples by treatment with 10 U/Fl of RNase free DNase (Invitrogen, San Diego, Calif.) for 15 minutes at 37° C. RNA was then precipitated and resuspended in RNA Secure (Ambion, Austin, Tex.). The cDNA was generated by reverse transcribing approximately 2 μg of total RNA using Taqman7 reverse transcription reagents (Applied Biosystems, Foster City, Calif.) according to manufacturer's protocols. Subsequently, cDNAs were amplified with specific human cDNA primers, to CXCL1, CXCL2, CXCL3, CXCL4, CXCL5, CXCL6, CXCL7, CXCL8, CXCL9, CXCL10, CXCL11, CXCL12, CXCL13, CXCL14, CXCL15, CXCL16, CXCR1, CXCR2, CXCR3, CXCR4, CXCR5, CXCR5a, CXCR5b, CXCR6, CXCR7, CCL1, CCL2, CCL3, CCL4, CCL5, CCL6, CCL7, CCL8, CCL9, CCL10, CCL11, CCL12, CCL13, CCL14, CCL15, CCL16, CCL17, CCL18, CCL19, CCL20, CCL21, CCL22, CCL24, CCL25, CCL25-1, CCL25-2, CCL27, CCL28, CCR1, CCR2, CCR3, CCR4, CCR5, CCR6, CCR7, CCR8, CCR9, CCR10, CCR11, XCL1, XCL2, XCR1, CX3CR1 or CX3CL1, using SYBR7 Green PCR master mix reagents (Applied Biosystems) according to manufacturer's protocol. The level of copies of mRNA of these targets were evaluated by real-time PCR analysis using the BioRad Icycler and software (Hercules, Calif.).
[0178] The RT-PCR products obtained using CXCL1-, CXCL2-, CXCL3-, CXCL4-, CXCL5-, CXCL6-, CXCL7-, CXCL8-, CXCL9-, CXCL10-, CXCL11-, CXCL12-, CXCL13-, CXCL14-, CXCL15-, CXCL16-, CXCR1-, CXCR2-, CXCR3-, CXCR4-, CXCR5-, CXCR5a-, CXCR5b-, CXCR6-, CXCR7-, CCL1-, CCL2-, CCL3-, CCL4-, CCL5-, CCL6-, CCL7-, CCL8-, CCL9-, CCL10-, CCL11-, CCL12-, CCL13-, CCL14-, CCL15-, CCL16-, CCL17-, CCL18-, CCL19-, CCL20-, CCL21-, CCL22-, CCL24-, CCL25-, CCL25-1-, CCL25-2-, CCL27-, CCL28-, CCR1-, CCR2-, CCR3-, CCR4-, CCR5-, CCR6-, CCR7-, CCR8-, CCR9-, CCR10-, CCR11-, XCL1-, XCL2-, XCR1-, CX3CR1-, or CX3CL1-specific primer sets did not cross react with other gene targets due to exclusion of primers that annealed to host sequences (NIH-NCBI Genebank). The primers produced different size amplicon products relative the polymorphisms that resulted in CXCR5a versus CXCR5b and CCL25, CCL25-1, versus CCL25-2. To this end, RT-PCR analysis of adenoma, carcinoma, leukemia, lymphoma, melanoma, and/or myeloma cell lines and tumor tissue revealed that chemokines and chemokine receptors were differentially expressed by cancer cells.
Example 3
Anti-Chemokine and Anti-Chemokine Receptor Antibodies Inhibit Tumor Cell Growth In Vitro and In Vivo
Anti-Sera Preparation
[0179] 15 amino acid peptides from CXCR1, CXCR2, CXCL1, CXCL2, CXCL3, CXCL5, CXCL6, CXCL7, CXCL8, CXCL12, CXCR5a, CXCR5b, CXCL13, CXCR6, CXCL16, CCL16, CCL25, CCL25-1, CCL25-2, CCR9, CX3CR1, and CX3CL1 (SEQ ID NOS:1-21) were synthesized (Sigma Genosys, The Woodlands, Tex.) and conjugated to hen egg lysozyme (Pierce, Rockford, Ill.) to generate the antigen for subsequent immunizations for anti-sera preparation or monoclonal antibody generation. The endotoxin levels of chemokine peptide conjugates were quantified by the chromogenic Limulus amebocyte lysate assay (Cape Cod, Inc., Falmouth, Miss.) and shown to be <5 EU/mg. 100 μg of the antigen was used as the immunogen together with complete Freund's adjuvant Ribi Adjuvant system (RAS) for the first immunization in a final volume of 1.0 ml. This mixture was administered in 100 ml aliquots on two sites of the back of the rabbit subcutaneously and 400 ml intramuscularly in each hind leg muscle. Three to four weeks later, rabbits received 100 μg of the antigen in addition to incomplete Freund's adjuvant for 3 subsequent immunizations. Anti-sera were collected when anti-CXCR1, -CXCR2, -CXCL1, -CXCL2, -CXCL3, -CXCL5, -CXCL6-CXCL7, -CXCL8, -CXCL12, -CXCR5a, -CXCR5b, -CXCL13, -CXCR6, -CXCL16, -CCL16, -CCL25, -CCL25-1, -CCL25-2, -CCR9, -CX3CR1, and -CX3CL1 antibody titers reached 1:1,000,000. Subsequently, normal or anti-sera were heat-inactivated and diluted 1:50 in PBS.
Monoclonal Antibody Preparation
[0180] 15 amino acid peptides from CXCR1, CXCR2, CXCL1, CXCL2, CXCL3, CXCL5, CXCL6, CXCL7, CXCL8, CXCL12, CXCR5a, CXCR5b, CXCL13, CXCR6, CXCL16, CCL16, CCL25, CCL25-1, CCL25-2, CCR9, CX3CR1, and CX3CL1 were synthesized (Sigma Genosys) and conjugated to hen egg lysozyme (Pierce) to generate the "antigen" for subsequent immunizations for anti-sera preparation or monoclonal antibody generation. The endotoxin levels of chemokine peptide conjugates were quantified by the chromogenic Limulus amebocyte lysate assay (Cape Cod, Inc., Falmouth, Miss.) and shown to be <5 EU/mg. 100 μg of the antigen was used as the immunogen together with complete Freund's adjuvant Ribi Adjuvant system (RAS) for the first immunization in a final volume of 200 μl. This mixture was subcutaneously administered in 100 μl aliquots at two sites of the back of a rat, mouse, or immunoglobulin-humanized mouse. Two weeks later, animals received 100 μg of the antigen in addition to incomplete Freund's adjuvant for 3 subsequent immunizations. Serum were collected and when anti-CXCR1, -CXCR2, -CXCL1, -CXCL2, -CXCL3, -CXCL5, -CXCL6-CXCL7, -CXCL8, -CXCL12, -CXCR5a, -CXCR5b, -CXCL13, -CXCR6, -CXCL16, -CCL16, -CCL25, -CCL25-1, -CCL25-2, -CCR9, -CX3CR1, or -CX3CL1 antibody titers reached 1:2,000,000, hosts were sacrificed and splenocytes were isolated for hybridoma generation. Briefly, B cells from the spleen or lymph nodes of immunized hosts were fused with immortal myeloma cell lines (e.g., YB2/0). Hybridomas were next isolated after selective culturing conditions (i.e., HAT-supplemented media) and limiting dilution methods of hybridoma cloning. Cells that produce antibodies with the desired specificity were selected using ELISA. Hybridomas from normal rats or mice were humanized with molecular biological techniques in common use. After cloning a high affinity and prolific hybridoma, antibodies were isolated from ascites or culture supernatants and adjusted to a titer of 1:2,000,000 and diluted 1:50 in PBS.
Anti-Sera or Monoclonal Antibody Treatment
[0181] Immunodeficient nude NIH-III mice (8 to 12 weeks old, Charles River Laboratory, Wilmington, Mass.), which lack T, B, and NK cells, received 1×106 cancer cells, subcutaneously, for the establishment of a tumor. The established solid tumor was then removed from the host for immediate implantation or stored in liquid nitrogen for later implantation. Freshly isolated or liquid nitrogen frozen tumor tissue (1 g) were surgically implanted in the intestinal adipose tissue for the generation of tumor. Once the xenografted tumor growth reached 5 mm in size, the NIH-III mice received 200 μl intraperitoneal injections of either anti-sera or monoclonal antibodies every three days and the tumor was monitored for progression or regression of growth.
Data Analysis
[0182] SigmaStat 2000 (Chicago, Ill.) software was used to analyze and confine the statistical significance of data. The data were subsequently analyzed by the Student's t-test, using a two-factor, unpaired test. In this analysis, treated samples were compared to untreated controls. The significance level was set at p<0.05.
In Vitro Growth Studies
[0183] The adenoma, carcinoma, leukemia, lymphoma, melanoma, and/or myeloma cell lines were grown in complete media in the presence or absence of antibodies specific for CXCR1, CXCR2, CXCL1, CXCL2, CXCL3, CXCL5, CXCL6 CXCL7, CXCL8, CXCR4, CXCL12, CXCR5a, CXCR5b, CXCL13, CXCR6, CXCL16, CCL16, CCR9, CCL25, CCL25-1, CCL25-2, CX3CR1, or CX3CL1. The growth of cancer cell lines expressing CXCR1 and/or CXCR2 were inhibited by antibodies to CXCR1, CXCR2, CXCL1, CXCL2, CXCL3, CXCL5, CXCL6, CXCL7, or CXCL8. Similarly, the growth of cancer cell lines expressing CXCR4 were inhibited by antibodies to CXCR4 or CXCL12. The growth of cancer cell lines expressing CXCR5a or CXCR5a were inhibited by antibodies to CXCR5a, CXCR5b, or CXCL13. The proliferation of cancer cell lines expressing CXCR6 were inhibited by antibodies to CXCR6 or CXCL16. The growth of cancer cell lines expressing CCR9 were inhibited by antibodies to CCR9, CCL25, CCL25-1, or CCL25-2. The propagation of cancer cell lines expressing CX3CR1 were inhibited by antibodies to CX3CR1 or CXC3L1. Of interest, antibodies against the soluble ligands, CXCL1, CXCL2, CXCL3, CXCL5, CXCL6, CXCL7, CXCL8, CXCL12, CXCL13, CXCL16, CCL16, CCL25, CCL25-1, CCL25-2, or CX3CL1, were more effective at growth inhibition that those directed against the membrane receptors.
In Vitro Angiogenesis Studies
[0184] Microvascular endothelial cells (Cell Systems, Kirkland, Wash.) were grown according to supplier's protocols and allowed to form microvascular venules in an in vitro assay for angiogenesis (BD-Biocoat, Hercules, Calif.), in the presence or absence of antibodies specific for CXCR1, CXCR2, CXCL1, CXCL2, CXCL3, CXCL5, CXCL6, CXCL7, CXCL8, CXCR4, CXCL12, CXCR5a, CXCR5b, CXCL13, CXCR6, CXCL16, CCL16, CCR9, CCL25, CCL25-1, CCL25-2, CX3CR1, or CX3CL1. The angiogenesis was inhibited by antibodies against CXCR1, CXCR2, CXCL1, CXCL2, CXCL3, CXCL5, CXCL6, CXCL7, CXCL8, CXCR4, CXCL12, CXCR6 or CXCL16.
In Vivo Growth Studies
[0185] Cancer cell lines or primary tumor tissue were adoptively transferred into NIH-III mice and allowed to form the xenograft tumor of interest. Antibodies directed against CXCR1, CXCR2, CXCL1, CXCL2, CXCL3, CXCL5, CXCL6, CXCL7, CXCL8, CXCR4, CXCL12, CXCR5a, CXCR5b, CXCL13, CXCR6, CXCL16, CCL16, CCR9, CCL25, CCL25-1, CCL25-2; CX3CR1, or CX3CL1 differentially affected the progression and regression of tumor size. In certain cases, antibodies directed towards CXCR1, CXCR2, CXCL1, CXCL2, CXCL3, CXCL5, CXCL6, CXCL7, CXCL8, CXCR4, CXCL12, CXCR6 or CXCL16 effectively lead to both regression and impeding progression of tumor growth. Antibodies directed against CXCR4, CXCL12, CXCR5a, CXCR5b, CXCL13, CCL16, CCR9, CCL25, CCL25-1, CCL25-2, CX3CR1, or CX3CL1 were effective at inhibiting the progression of tumor size.
[0186] The protein sequences of the chemokines used herein are recorded in NIH-NCBI GenBank as: (1) CXCR1 (ACCESSION# NP 000625), SEQ ID NO:1, (2) CXCR2(ACCESSION# NP 001548), SEQ ID NO:2, (3) CXCL1 (ACCESSION# NP 001502), SEQ ID NO:3, (4) CXCL2 (ACCESSION# NP 002080), SEQ ID NO:4, (5) CXCL3 (ACCESSION# NP 002081), SEQ ID NO:5, (6) CXCL5 (ACCESSION# NP 002985), SEQ ID NO:6, (7) CXCL6 (ACCESSION# NP 002984), SEQ ID NO:7, (8) CXCL7 (ACCESSION# NP 002695), SEQ ID NO:8, (9) CXCL8 (IL-8, ACCESSION# NP 000575), SEQ ID NO:9, (10) CXCR4 (ACCESSION# NP 003458), SEQ ID NO:10, (11) CXCL12 (ACCESSION# NP 000600), SEQ ID NO:11, (12) CXCR5A (ACCESSION# NP 116743), SEQ ID NO:12, (13) CXCR5B (ACCESSION# NP 001707), SEQ ID NO:13, (14) CXCL13 (ACCESSION# NP 006410), SEQ ID NO:14, (15) CXCR6 (ACCESSION# NP 006555), SEQ ID NO:15, (16) CXCL16 (ACCESSION# NP 071342), SEQ ID NO:16, (17) CCL16 (ACCESSION# NP 004581), SEQ ID NO:17, (18) CCL25 (ACCESSION# NP-005615.2), SEQ ID NO:18, (19) CCL25-1 (ACCESSION# NP 005615), SEQ ID NO:19, (20) CCL25-2 (ACCESSION# NP 683686), SEQ ID NO:20, (21) CX3CR1 (ACCESSION# NP 001328), SEQ ID NO:21, and (22) CX3CL1 (ACCESSION# NP 002987), SEQ ID NO:22.
[0187] The cDNA sequences are known and are available in NIH-NCBI GenBank under the following accession numbers: (23) CXCR1 (ACCESSION# NM 000634), SEQ ID NO:23, (24) CXCR2(ACCESSION# NM 001557), SEQ ID NO:24, (25) CXCL1 (ACCESSION# NM 001511), SEQ ID NO:25, (26) CXCL2 (ACCESSION# NM 002089), SEQ ID NO:26, (27) CXCL3 (ACCESSION# NM 002090), SEQ ID NO:27, (28) CXCL5 (ACCESSION# NM 002994), SEQ ID NO:28, (29) CXCL6 (ACCESSION# NM 002993), SEQ ID NO:29, (30) CXCL7 (ACCESSION# NM 002704), SEQ ID NO:30, (31) CXCL8 (IL-8, ACCESSION# NM 000584), SEQ ID NO:31, (32) CXCR4 (ACCESSION# NM 003467), SEQ ID NO:32, (33) CXCL12 (ACCESSION# NM 000609), SEQ ID NO:33, (34) CXCR5A (ACCESSION# NM 032966), SEQ ID NO:34, (35) CXCR5B (ACCESSION# NM 001716), SEQ ID NO:35, (36) CXCL13 (ACCESSION# NM 006419), SEQ ID NO:36, (37) CXCR6 (ACCESSION# NM 006564), SEQ ID NO:37, (38) CXCL16 (ACCESSION# NM 022059), SEQ ID NO:38, (39) CCL16 (ACCESSION# NM 004590), SEQ ID NO:39, (40) CCL25 (ACCESSION# NM 005624.3), SEQ ID NO:40, (41) CCL25-1 (ACCESSION# NM 005624), SEQ ID NO:41, (42) CCL25-2 (ACCESSION# NM 148888), SEQ ID NO:42, (43) CX3CR1 (ACCESSION# NM 001337), SEQ ID NO:43, and (44) CX3CL1 (ACCESSION# NM 002996), SEQ ID NO:44.
[0188] As shown in the table below, the particular chemokines which are most which any tumor expresses may vary. The methods of the present application may be customized for a particular patient, depending on the chemokines over-expressed by the patient's own tumor. It is possible to identify the particular chemokines which are over-expressed in the tumor using methods of the application and administer antibodies against that over-expressed chemokine. The tailoring of treatment for the cancer patient is novel, and is a particularly valuable aspect of the application.
[0189] TABLE 1 indicates the differing amounts of particular chemokines over-expressed in particular tumors that were studied.
TABLE-US-00001 TABLE 1 Chemokine, Chemokine Receptor and Cancer Association (dependent on stage of disease). Cancer Chemokine Chemokine Receptor Carcinoma CCL1, CCL2, CCL4, CCL17, CCR2, CCR7, CCR8, CCL19, CCL21, CCL22, CCL25 CCR9 CXCL12, CXCL13, CXCL16 CXCR4, CXCR5, CX3CL1 CXCR6 CX3CR1 Leukemia CCL1, CCL4, CCL17, CCL19, CCR7, CCR8, CCR9 CCL21, CCL22, CCL25 CXCL12 CXCR4, CXCR7 Lymphoma CXCL12, CXCL13 CXCR4, CXCR5 Melanoma CCL25, CCL27 CCR9, CCR10 CXCL1, CXCL2, CXCL3, CXCL5, CXCR1, CXCR2, CXCL6, CXCL7, CXCL8, CXCR4, CXCR5, CXCL12, CXCL13, CXCL16 CXCR6, CXCR7 CX3CL1 CX3CR1 Sarcoma CCL1, CCL3, CCL4, CCL5, CCL7, CCR3, CCR5, CCR8 CCL8, CCL11, CCL13, CCL17, CCL22, CCL24 CXCL12 CXCR4, CXCR7 CX3CL1 CX3CR1
Example 4
CCR9-CCL25 Induced Anti-Apoptotic and/or Survival Signal Involved in PCa Chemo Resistance
[0190] LNCaP (hormone responsive, wild type p53 expression), PC3 (hormone refractory, p53 null), and DU145 (hormone refractory, p53 mutated) cell lines are grown with or without CCL25 and with or without doxorubicin (1 μM/2 μM/4 μM), etoposide (20 μM/40 μM), estramustine (4 μM/10 μM), or docetaxel (10 nM/20 nM/40 nM) for 4, 8, 12, and 24 hours. Expression and activation of cell survival, pro- and anti-apoptotic signals (Akt, Src, CamKII, FAK, FKHR, FOXO, CREB, NF-κB, Myc, Fos, Jun Apaf1, Bax, Bcl2, BclXL, BaK, Bad, Bik, Bim, TP53, Caspase-3, -6, -8, -9, survivin, vitronectin, β-Catenin) and molecules responsible for drug resistance or metabolism (Twist-1, Snail-1, Glutathione-S-transferase-π (GST-π), p53, topoisomerase I, IIα, IIβ, and ABC drug transporters) are accessed by real-time PCR and Western blot. Briefly, after treatment of cells, changes in the gene expression is tested using real-time PCR. Activation of signaling molecules is also be tested by phosphorylation specific antibody (i.e., Western blot analysis). To further confirm the role of the activated signaling molecules, following CCL25 treatment, expression or activity of the candidate molecules is inhibited using chemical inhibitors or siRNAs and target genes are analyzed by real-time PCR and Western blot analysis. Subsequently, the response of treated cells to chemotherapeutic drugs is evaluated by Vybrant apoptosis assay (Molecular probes) kit.
RNA Isolation and Real-Time PCR
[0191] Total RNA is isolated by TRIZOL® (Invitrogen) method and quantified by UV spectrophotometry. Quality of RNA is analyzed by electrophoresis. The cDNA synthesis is completed using the ISCRIPT® cDNA synthesis kit (BioRad) as described by the manufacturer. Real-time PCR is performed using IQ® SYBR green supermix (BioRad) as described by manufacturer and specific primers designed against FAK, FKHR, FOXO, Apaf1, Bax, Bcl2, BclXL, BaK, Bad, Bid, XIAP, Bik, Bim, TP53, cytochrome C, Caspase-3, -6, -8, -9, survivin, lamin, CamKII, vitronectin, β-Catenin, cadherins, Twist-1, Snail-1, CREB, NF-κB, Myc, Fos, Jun, β-actin and GAPDH. The results are calculated by delta delta Ct to quantify fold changes in mRNAs compared to untreated groups.
Western Blotting
[0192] Cells are harvested and resuspended in lysis buffer to extract total protein. Lysis buffer contains 50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 1% Triton X-100, 1% deoxycholate, 0.1% SDS, 5 mM EDTA supplemented with protease inhibitors, 1 mM phenylmethylsulphonylfluoride, 1 mM benzamidine, 10 μg/mL soybean trypsin inhibitor, 50 μg/mL leupeptin, 1 μg/mL pepstatin and 20 μg/mL aprotinin. Cell lysates are stored on ice for 30 min, centrifuged (14000×g) for 20 min at 4° C., and supernatant is used for Western blot analysis of genes demonstrating significant modulation in mRNA level. Similarly, phosphor-specific antibodies are used to test changes in the level of phophorylation of Akt1/2/3, mTOR, FAK, FKHR, FOXO, and GSK-3β. Moreover, activation of caspases and PARP, following cleavage are evaluated using specific antibodies. The results obtained after chemiluminescent detection of protein bands by ECL plus reagent (Pharmecia) on X-ray film is normalized to β-actin and/or GAPDH using Image J image analysis software (NIH).
Detection of Cytochrome C Release
[0193] Cells are collected and washed in PBS, and resuspended in extraction buffer containing 220 mM mannitol, 68 mM sucrose, 50 mM PIPES-KOH, pH 7.4, 50 mM KCl, 5 mM EGTA, 2 mM MgCl2, 1 mM DTT, and protease inhibitors. After 30 min incubation on ice, cells are homogenized using Glass-Teflon homogenizer and homogenates will be spun at 14,000 g for 15 min. Cytosolic extracts are used for Western blot analysis using anti-cytochrome C monoclonal antibody (PharMingen).
siRNA Transfection, Chemical Inhibitor, and Apoptosis Detection
[0194] Prostate cancer cell lines are transfected with gene specific and nonspecific control siRNAs (Dharmacon) using LipofectAMINE 2000 (Invitrogen). Optimum gene knock-down time and siRNA concentration are confirmed by western blot analysis and further evaluated for cell survival following drug treatment with or without CXCL16, control antibody, and/or anti-CXCR6 antibody. The detection of changes in live, apoptotic, and necrotic cells is evaluated as follows: cell survival is tested by Vybrant apoptosis as described by the manufacturer (Molecular probe), using FACScan flow cytometer and CellQuest® software (BD Pharmingen). Change in down-stream gene expression after gene knockdown is tested using real-time PCR and Western blotting.
[0195] Cells treated with CCL25 show enhanced expression of cell survival and drug transporter proteins which show differences in their expression pattern in hormone responsive and non responsive cells. Anti-CCL25 Abs effectively reverse the effect of CCL25 in PCa cells. Doxorubicin, estramustine, etoposide and docetaxel induce apoptosis in PCa cells without CCL25 treatment (or CCR9 blockade).
Example 5
CCR9-CCL25 Induced Changes in ABC Drug Transporters
[0196] LNCaP, PC3, and DU145 cells are grown with or without CCL25, anti-CCL25 antibody, control antibody, and/or anti-CCR9 antibodies along with or without doxorubicin, estramustine, etoposide or docetaxel for 4, 8, 12 or 16 hours as described earlier. After treatment, changes in the ABC transporter and Twist-1 mRNA expression are quantified by real-time PCR, as described above, using specific primers directed for ABC and Twist-1 cDNA. The genes demonstrating significant alterations in mRNA expression are further tested by Western blot analysis. Nuclear extracts from treated cells are evaluated by chromatin immuno-precipitation (ChIP) assay to determine whether the transcriptional factors induced by CXCL16 bind the promoter region of ABC transporters and Twist-1.
Chromatin Immuno Precipitation (ChIP)
[0197] The results from Example 4 provide information about the genes that are regulated as well as those that may modulate transcription factors activated by CCR9-CCL25 interaction. Based on these results, target transcription factors and genes are selected. Specific PCR primers are designed against the promoter region of these genes containing the binding sites of transcription factors. PCR primer are used to amplify the DNA being precipitated along with transcription factors. Cells are harvested by trypsinization in the presence of 20 mM butyrate. 50,000 cells are re-suspended in 500 μl PBS/butyrate. Proteins and DNA are cross-linked with 1% formaldehyde for 8 min at room temperature and cross-linking is stopped with 125 mM glycine for 5 min. Cells are centrifuged at 470 g in a swing-out rotor with soft deceleration settings for 10 min at 4° C. and washed twice in 0.5 ml ice-cold PBS/butyrate by vortexing followed by centrifugation. Cells are lysed by addition of lysis buffer (50 mM Tris-HCl, pH 8, 10 mM EDTA, 1% SDS, protease inhibitor cocktail (Sigma-Aldrich), 1 mM PMSF, 20 mM butyrate, vortexing and subsequent centrifugation. This procedure is known to produce chromatin fragments of 500 bp. The sonicated lysate is diluted 8-fold in RIPA buffer containing a protease inhibitor cocktail, 1 mM PMSF, and 20 mM butyrate (RIPA ChIP buffer). RIPA ChIP buffer (330 μl) is added to the pellet and mixed by vortexing. Immunoprecipitation and washes of the ChIP material is accomplished by the use of antibody-directed against specific transcription factors. Chromatin is aliquoted into tubes containing antibody-bead complexes. Input sample is placed in a tube for phenol-chloroform isoamyl alcohol isolation. The immunoprecipitated material is washed three times and transferred into a new tube while in TE. DNA elution in 1% SDS, cross-link reversal and proteinase K digestion is carried out in a single step for 2 hrs at 68° C. DNA is extracted with phenol-chloroform isoamylalcohol, and ethanol-precipitation in presence of acrylamide carrier (Sigma-Aldrich) and dissolved in TE. Immunoprecipitated DNA from 3-4 independent ChIPs is analyzed by real time PCR. Real-time PCR data is expressed as percent (±SD) precipitated (antibody-bound) DNA relative to input DNA, in three independent replicate ChIP assays.
[0198] Phosphorylation and activation of transcription factors such as CREB, Fos, Jun, and NFkB via CCR9-CCL25 signaling subsequently leads to increases in expression of ABC transporters and Twist-1. Decreases in gene expression are observed if negative regulatory elements are present in the same promoter. Since hormone-dependent and refractory PCa cells have differences in the expression of these intracellular signaling molecules, they show variations in genes to be modulated by hormone dependent and refractory conditions. The modulation in gene expression shows differences with drug treatment in presence of CCL25 and in absence of CCL25 treatment.
Example 6
In Vivo Evaluation of CCL25-Directed Therapy
[0199] Male nude mice are subcutaneously challenged by luciferase expressing androgen responsive (LNCaP-Luc) and non-responsive (PC3-Luc) cells. Tumor development is measured non-invasively using in vivo imaging system. After establishment of a measurable tumor, mice are divided into treatment (A, B, C, D and E) and control groups (F, G, H, I, J and K). Group "A" receives CCL25 neutralizing antibodies (12.5 mg/kg/day) every alternate day and controls (group F) receive isotype control antibodies (12.5 mg/kg/day). Group "B," "C," "D" and "E" receive CCL25 neutralizing antibodies (12.5 mg/kg/day) with intraperitoneal injection of doxorubicin (5 mg/kg/day on days 1 to 3 followed by administration on days 15 to 17), intravenous injection of etoposide (10 mg/kg/day; on day 1, 5, 9, 14, 19 and 24), intravenous injection of estramustine (4 mg/kg/day on day 1-5 and day 26-31), or intraperitoneal injection of docetaxel (8 mg/kg/day twice a week for 4 weeks), respectively. Controls for these treatment groups ("G," "H," "I" and "J," respectively) receive theses drugs using similar concentration and injection protocol with isotype control antibodies (12.5 mg/kg/day). Group "K" receives PBS and serves as placebo. Tumor progression and regression in treatment and controls are evaluated by non-invasive in vivo imaging. The tumor from treated groups and untreated control groups is excised and evaluated for the changes in the cell survival and drug resistance proteins by immunohistochemistry. In the context used herein, the term "CCL25 neutralizing antibodies" means anti-CCL25 antibodies and/or anti-CCR9 antibodies.
Statistics (Significance) and Sample Size
[0200] Sample size (or power) calculations are relevant to the design of preliminary studies and determining the requirements for proposed experiments. To interpret our results, significance tests and statistical analysis are also critical. The traditional α-value, i.e., p=0.01, is used to evaluate the statistical significance of this study. The proposed experiment will require a minimum of 10 mice per group. The data is expressed as the mean±SEM and compared using a two-tailed paired (or unpaired) student's t-test for normally distributed samples or an unpaired Mann Whitney U test as a non-parametric test for samples not normally distributed. The results are analyzed using SYSTAT (Systat software Inc.) statistical program. Single-factor and two-factor variance ANOVA analyses are used to evaluate groups and subgroups, respectively. Hence, results are considered statistically significant if p values are <0.05.
Animals
[0201] Six to eight week old male nude mice are subcutaneously injected with PCa cells. Briefly, 5×106 Luciferase expressing PC3 cells are resuspended in 100 μl of sterile PBS and injected into the flanks of nude mice under isoflurane anesthesia. Luciferase expressing LNCaP cells (5×106 cell) are mixed with 50% Matrigel (Becton Dickinson) and injected in the flanks of nude mice under isoflurane anesthesia.
Analysis of In Vivo Tumor Growth
[0202] Tumor bearing nude mice receive 150 mg/kg D-Luciferin (Xenogen) by intra-peritoneal injection Using 25×5/8'' gauge needle 15 minutes before imaging. The mice are imaged using the IVIS100 in vivo imaging system and results expressed in photons/sec/cm2/sr. Tumor volume is measured by use of calipers and calculated by the formula (Larger diameter)×(smaller diameter)2×0.5.
Cell Survival, Apoptotic and Drug Resistant Gene Expression Analysis
[0203] Tumors from all groups are excised three days after completion of treatment protocols. Tumors are fixed in 4% PFA and embedded in paraffin. Paraffin sections (thickness 7 μm) are mounted on glass slides, deparaffinized and re-hydrated (Xylene for 5 min; absolute, 95% and 70% ethanol for 1 min each). The rehydrated sections are used for peroxidase based immunohistochemical staining for drug transporters, PI3K, Akt, FAK, FKHR, FOXO, Apaf1, Bax, Bcl2, BclXL, BaK, Bad, Bid, XIAP, Bik, Bim, TP53, Cytochrome C, Caspase-3, -6, -8, -9, survivin, lamin, CamKII, vitronectin, β-Catenin, cadherins, Twist-1, CREB, NF-κB, Myc, Fos, Jun, CCR9 and CCL25. After staining, slides are scanned and analyzed by the Aperio scanscope (Aperio) system.
[0204] CCL25 neutralization leads to decreased cell survival in response to drugs, thus reduction of tumor volume. However, the response also varies among the tumors formed by hormone sensitive (LNCaP) and hormone refractory (PC3 cells). Further, chemotherapeutic drugs have lower efficacy in the tumors with a functional CCR9-CCL25 axis, which may enhance the expression of ABC proteins known to transport these drugs out of the cell.
[0205] The above description is for the purpose of teaching the person of ordinary skill in the art how to practice the present invention, and is not intended to detail all those obvious modifications and variations of it that will become apparent to the skilled worker upon reading the description. It is intended, however, that all such obvious modifications and variations be included within the scope of the present invention, which is defined by the following claims. The claims are intended to cover the components and steps in any sequence that is effective to meet the objectives there intended, unless the context specifically indicates the contrary. All the references cited in the specification are herein incorporated by reference in their entirely.
Sequence CWU
1
1371350PRTHomo sapiens 1Met Ser Asn Ile Thr Asp Pro Gln Met Trp Asp Phe
Asp Asp Leu Asn1 5 10
15Phe Thr Gly Met Pro Pro Ala Asp Glu Asp Tyr Ser Pro Cys Met Leu
20 25 30Glu Thr Glu Thr Leu Asn Lys
Tyr Val Val Ile Ile Ala Tyr Ala Leu 35 40
45Val Phe Leu Leu Ser Leu Leu Gly Asn Ser Leu Val Met Leu Val Ile
50 55 60Leu Tyr Ser Arg Val Gly Arg
Ser Val Thr Asp Val Tyr Leu Leu Asn65 70
75 80Leu Ala Leu Ala Asp Leu Leu Phe Ala Leu Thr Leu
Pro Ile Trp Ala 85 90
95Ala Ser Lys Val Asn Gly Trp Ile Phe Gly Thr Phe Leu Cys Lys Val
100 105 110Val Ser Leu Leu Lys Glu Val
Asn Phe Tyr Ser Gly Ile Leu Leu Leu 115 120
125Ala Cys Ile Ser Val Asp Arg Tyr Leu Ala Ile Val His Ala Thr
Arg 130 135 140Thr Leu Thr Gln Lys Arg
His Leu Val Lys Phe Val Cys Leu Gly Cys145 150
155 160Trp Gly Leu Ser Met Asn Leu Ser Leu Pro Phe
Phe Leu Phe Arg Gln 165 170
175Ala Tyr His Pro Asn Asn Ser Ser Pro Val Cys Tyr Glu Val Leu Gly
180 185 190Asn Asp Thr Ala Lys Trp
Arg Met Val Leu Arg Ile Leu Pro His Thr 195 200
205Phe Gly Phe Ile Val Pro Leu Phe Val Met Leu Phe Cys Tyr
Gly Phe 210 215 220Thr Leu Arg Thr Leu
Phe Lys Ala His Met Gly Gln Lys His Arg Ala225 230
235 240Met Arg Val Ile Phe Ala Val Val Leu Ile
Phe Leu Leu Cys Trp Leu 245 250
255Pro Tyr Asn Leu Val Leu Leu Ala Asp Thr Leu Met Arg Thr Gln Val
260 265 270Ile Gln Glu Ser Cys
Glu Arg Arg Asn Asn Ile Gly Arg Ala Leu Asp 275
280 285Ala Thr Glu Ile Leu Gly Phe Leu His Ser Cys Leu
Asn Pro Ile Ile 290 295 300Tyr Ala Phe
Ile Gly Gln Asn Phe Arg His Gly Phe Leu Lys Ile Leu305
310 315 320Ala Met His Gly Leu Val Ser
Lys Glu Phe Leu Ala Arg His Arg Val 325
330 335Thr Ser Tyr Thr Ser Ser Ser Val Asn Val Ser Ser
Asn Leu 340 345 3502360PRTHomo
sapiens 2Met Glu Asp Phe Asn Met Glu Ser Asp Ser Phe Glu Asp Phe Trp Lys1
5 10 15Gly Glu Asp Leu
Ser Asn Tyr Ser Tyr Ser Ser Thr Leu Pro Pro Phe 20
25 30Leu Leu Asp Ala Ala Pro Cys Glu Pro Glu Ser
Leu Glu Ile Asn Lys 35 40 45Tyr
Phe Val Val Ile Ile Tyr Ala Leu Val Phe Leu Leu Ser Leu Leu 50
55 60Gly Asn Ser Leu Val Met Leu Val Ile Leu
Tyr Ser Arg Val Gly Arg65 70 75
80Ser Val Thr Asp Val Tyr Leu Leu Asn Leu Ala Leu Ala Asp Leu
Leu 85 90 95Phe Ala Leu
Thr Leu Pro Ile Trp Ala Ala Ser Lys Val Asn Gly Trp 100
105 110Ile Phe Gly Thr Phe Leu Cys Lys Val Val
Ser Leu Leu Lys Glu Val 115 120
125Asn Phe Tyr Ser Gly Ile Leu Leu Leu Ala Cys Ile Ser Val Asp Arg 130
135 140Tyr Leu Ala Ile Val His Ala Thr
Arg Thr Leu Thr Gln Lys Arg Tyr145 150
155 160Leu Val Lys Phe Ile Cys Leu Ser Ile Trp Gly Leu
Ser Leu Leu Leu 165 170
175Ala Leu Pro Val Leu Leu Phe Arg Arg Thr Val Tyr Ser Ser Asn Val
180 185 190Ser Pro Ala Cys Tyr Glu
Asp Met Gly Asn Asn Thr Ala Asn Trp Arg 195 200
205Met Leu Leu Arg Ile Leu Pro Gln Ser Phe Gly Phe Ile Val
Pro Leu 210 215 220Leu Ile Met Leu Phe
Cys Tyr Gly Phe Thr Leu Arg Thr Leu Phe Lys225 230
235 240Ala His Met Gly Gln Lys His Arg Ala Met
Arg Val Ile Phe Ala Val 245 250
255Val Leu Ile Phe Leu Leu Cys Trp Leu Pro Tyr Asn Leu Val Leu Leu
260 265 270Ala Asp Thr Leu Met
Arg Thr Gln Val Ile Gln Glu Thr Cys Glu Arg 275
280 285Arg Asn His Ile Asp Arg Ala Leu Asp Ala Thr Glu
Ile Leu Gly Ile 290 295 300Leu His Ser
Cys Leu Asn Pro Leu Ile Tyr Ala Phe Ile Gly Gln Lys305
310 315 320Phe Arg His Gly Leu Leu Lys
Ile Leu Ala Ile His Gly Leu Ile Ser 325
330 335Lys Asp Ser Leu Pro Lys Asp Ser Arg Pro Ser Phe
Val Gly Ser Ser 340 345 350Ser
Gly His Thr Ser Thr Thr Leu 355 3603107PRTHomo
sapiens 3Met Ala Arg Ala Ala Leu Ser Ala Ala Pro Ser Asn Pro Arg Leu Leu1
5 10 15Arg Val Ala Leu
Leu Leu Leu Leu Leu Val Ala Ala Gly Arg Arg Ala 20
25 30Ala Gly Ala Ser Val Ala Thr Glu Leu Arg Cys
Gln Cys Leu Gln Thr 35 40 45Leu
Gln Gly Ile His Pro Lys Asn Ile Gln Ser Val Asn Val Lys Ser 50
55 60Pro Gly Pro His Cys Ala Gln Thr Glu Val
Ile Ala Thr Leu Lys Asn65 70 75
80Gly Arg Lys Ala Cys Leu Asn Pro Ala Ser Pro Ile Val Lys Lys
Ile 85 90 95Ile Glu Lys
Met Leu Asn Ser Asp Lys Ser Asn 100
1054107PRTHomo sapiens 4Met Ala Arg Ala Thr Leu Ser Ala Ala Pro Ser Asn
Pro Arg Leu Leu1 5 10
15Arg Val Ala Leu Leu Leu Leu Leu Leu Val Ala Ala Ser Arg Arg Ala
20 25 30Ala Gly Ala Pro Leu Ala Thr
Glu Leu Arg Cys Gln Cys Leu Gln Thr 35 40
45Leu Gln Gly Ile His Leu Lys Asn Ile Gln Ser Val Lys Val Lys
Ser 50 55 60Pro Gly Pro His Cys Ala
Gln Thr Glu Val Ile Ala Thr Leu Lys Asn65 70
75 80Gly Gln Lys Ala Cys Leu Asn Pro Ala Ser Pro
Met Val Lys Lys Ile 85 90
95Ile Glu Lys Met Leu Lys Asn Gly Lys Ser Asn 100
1055107PRTHomo sapiens 5Met Ala His Ala Thr Leu Ser Ala Ala Pro Ser
Asn Pro Arg Leu Leu1 5 10
15Arg Val Ala Leu Leu Leu Leu Leu Leu Val Ala Ala Ser Arg Arg Ala
20 25 30Ala Gly Ala Ser Val Val Thr
Glu Leu Arg Cys Gln Cys Leu Gln Thr 35 40
45Leu Gln Gly Ile His Leu Lys Asn Ile Gln Ser Val Asn Val Arg
Ser 50 55 60Pro Gly Pro His Cys Ala
Gln Thr Glu Val Ile Ala Thr Leu Lys Asn65 70
75 80Gly Lys Lys Ala Cys Leu Asn Pro Ala Ser Pro
Met Val Gln Lys Ile 85 90
95Ile Glu Lys Ile Leu Asn Lys Gly Ser Thr Asn 100
1056114PRTHomo sapiens 6Met Ser Leu Leu Ser Ser Arg Ala Ala Arg Val
Pro Gly Pro Ser Ser1 5 10
15Ser Leu Cys Ala Leu Leu Val Leu Leu Leu Leu Leu Thr Gln Pro Gly
20 25 30Pro Ile Ala Ser Ala Gly Pro
Ala Ala Ala Val Leu Arg Glu Leu Arg 35 40
45Cys Val Cys Leu Gln Thr Thr Gln Gly Val His Pro Lys Met Ile
Ser 50 55 60Asn Leu Gln Val Phe Ala
Ile Gly Pro Gln Cys Ser Lys Val Glu Val65 70
75 80Val Ala Ser Leu Lys Asn Gly Lys Glu Ile Cys
Leu Asp Pro Glu Ala 85 90
95Pro Phe Leu Lys Lys Val Ile Gln Lys Ile Leu Asp Gly Gly Asn Lys
100 105 110Glu Asn 7114PRTHomo
sapiens 7Met Ser Leu Pro Ser Ser Arg Ala Ala Arg Val Pro Gly Pro Ser Gly1
5 10 15Ser Leu Cys Ala
Leu Leu Ala Leu Leu Leu Leu Leu Thr Pro Pro Gly 20
25 30Pro Leu Ala Ser Ala Gly Pro Val Ser Ala Val
Leu Thr Glu Leu Arg 35 40 45Cys
Thr Cys Leu Arg Val Thr Leu Arg Val Asn Pro Lys Thr Ile Gly 50
55 60Lys Leu Gln Val Phe Pro Ala Gly Pro Gln
Cys Ser Lys Val Glu Val65 70 75
80Val Ala Ser Leu Lys Asn Gly Lys Gln Val Cys Leu Asp Pro Glu
Ala 85 90 95Pro Phe Leu
Lys Lys Val Ile Gln Lys Ile Leu Asp Ser Gly Asn Lys 100
105 110Lys Asn 8128PRTHomo sapiens 8Met Ser Leu
Arg Leu Asp Thr Thr Pro Ser Cys Asn Ser Ala Arg Pro1 5
10 15Leu His Ala Leu Gln Val Leu Leu Leu
Leu Ser Leu Leu Leu Thr Ala 20 25
30Leu Ala Ser Ser Thr Lys Gly Gln Thr Lys Arg Asn Leu Ala Lys Gly
35 40 45Lys Glu Glu Ser Leu Asp Ser
Asp Leu Tyr Ala Glu Leu Arg Cys Met 50 55
60Cys Ile Lys Thr Thr Ser Gly Ile His Pro Lys Asn Ile Gln Ser Leu65
70 75 80Glu Val Ile Gly
Lys Gly Thr His Cys Asn Gln Val Glu Val Ile Ala 85
90 95Thr Leu Lys Asp Gly Arg Lys Ile Cys Leu
Asp Pro Asp Ala Pro Arg 100 105
110Ile Lys Lys Ile Val Gln Lys Lys Leu Ala Gly Asp Glu Ser Ala Asp
115 120 125999PRTHomo sapiens 9Met Thr
Ser Lys Leu Ala Val Ala Leu Leu Ala Ala Phe Leu Ile Ser1 5
10 15Ala Ala Leu Cys Glu Gly Ala Val
Leu Pro Arg Ser Ala Lys Glu Leu 20 25
30Arg Cys Gln Cys Ile Lys Thr Tyr Ser Lys Pro Phe His Pro Lys
Phe 35 40 45Ile Lys Glu Leu Arg
Val Ile Glu Ser Gly Pro His Cys Ala Asn Thr 50 55
60Glu Ile Ile Val Lys Leu Ser Asp Gly Arg Glu Leu Cys Leu
Asp Pro65 70 75 80Lys
Glu Asn Trp Val Gln Arg Val Val Glu Lys Phe Leu Lys Arg Ala
85 90 95Glu Asn Ser10352PRTHomo
sapiens 10Met Glu Gly Ile Ser Ile Tyr Thr Ser Asp Asn Tyr Thr Glu Glu
Met1 5 10 15Gly Ser Gly
Asp Tyr Asp Ser Met Lys Glu Pro Cys Phe Arg Glu Glu 20
25 30Asn Ala Asn Phe Asn Lys Ile Phe Leu Pro
Thr Ile Tyr Ser Ile Ile 35 40
45Phe Leu Thr Gly Ile Val Gly Asn Gly Leu Val Ile Leu Val Met Gly 50
55 60Tyr Gln Lys Lys Leu Arg Ser Met Thr
Asp Lys Tyr Arg Leu His Leu65 70 75
80Ser Val Ala Asp Leu Leu Phe Val Ile Thr Leu Pro Phe Trp
Ala Val 85 90 95Asp Ala
Val Ala Asn Trp Tyr Phe Gly Asn Phe Leu Cys Lys Ala Val 100
105 110His Val Ile Tyr Thr Val Asn Leu Tyr
Ser Ser Val Leu Ile Leu Ala 115 120
125Phe Ile Ser Leu Asp Arg Tyr Leu Ala Ile Val His Ala Thr Asn Ser
130 135 140Gln Arg Pro Arg Lys Leu Leu
Ala Glu Lys Val Val Tyr Val Gly Val145 150
155 160Trp Ile Pro Ala Leu Leu Leu Thr Ile Pro Asp Phe
Ile Phe Ala Asn 165 170
175Val Ser Glu Ala Asp Asp Arg Tyr Ile Cys Asp Arg Phe Tyr Pro Asn
180 185 190Asp Leu Trp Val Val Val
Phe Gln Phe Gln His Ile Met Val Gly Leu 195 200
205Ile Leu Pro Gly Ile Val Ile Leu Ser Cys Tyr Cys Ile Ile
Ile Ser 210 215 220Lys Leu Ser His Ser
Lys Gly His Gln Lys Arg Lys Ala Leu Lys Thr225 230
235 240Thr Val Ile Leu Ile Leu Ala Phe Phe Ala
Cys Trp Leu Pro Tyr Tyr 245 250
255Ile Gly Ile Ser Ile Asp Ser Phe Ile Leu Leu Glu Ile Ile Lys Gln
260 265 270Gly Cys Glu Phe Glu
Asn Thr Val His Lys Trp Ile Ser Ile Thr Glu 275
280 285Ala Leu Ala Phe Phe His Cys Cys Leu Asn Pro Ile
Leu Tyr Ala Phe 290 295 300Leu Gly Ala
Lys Phe Lys Thr Ser Ala Gln His Ala Leu Thr Ser Val305
310 315 320Ser Arg Gly Ser Ser Leu Lys
Ile Leu Ser Lys Gly Lys Arg Gly Gly 325
330 335His Ser Ser Val Ser Thr Glu Ser Glu Ser Ser Ser
Phe His Ser Ser 340 345
3501193PRTHomo sapiens 11Met Asn Ala Lys Val Val Val Val Leu Val Leu Val
Leu Thr Ala Leu1 5 10 15
Cys Leu Ser Asp Gly Lys Pro Val Ser Leu Ser Tyr Arg Cys Pro Cys
20 25 30 Arg Phe Phe Glu Ser His Val
Ala Arg Ala Asn Val Lys His Leu Lys 35 40
45Ile Leu Asn Thr Pro Asn Cys Ala Leu Gln Ile Val Ala Arg Leu
Lys 50 55 60Asn Asn Asn Arg Gln Val
Cys Ile Asp Pro Lys Leu Lys Trp Ile Gln65 70
75 80Glu Tyr Leu Glu Lys Ala Leu Asn Lys Arg Phe
Lys Met 85 9012327PRTHomo sapiens 12Met
Ala Ser Phe Lys Ala Val Phe Val Pro Val Ala Tyr Ser Leu Ile1
5 10 15Phe Leu Leu Gly Val Ile Gly
Asn Val Leu Val Leu Val Ile Leu Glu 20 25
30Arg His Arg Gln Thr Arg Ser Ser Thr Glu Thr Phe Leu Phe
His Leu 35 40 45Ala Val Ala Asp
Leu Leu Leu Val Phe Ile Leu Pro Phe Ala Val Ala 50 55
60Glu Gly Ser Val Gly Trp Val Leu Gly Thr Phe Leu Cys
Lys Thr Val65 70 75
80Ile Ala Leu His Lys Val Asn Phe Tyr Cys Ser Ser Leu Leu Leu Ala
85 90 95Cys Ile Ala Val Asp Arg
Tyr Leu Ala Ile Val His Ala Val His Ala 100
105 110Tyr Arg His Arg Arg Leu Leu Ser Ile His Ile Thr
Cys Gly Thr Ile 115 120 125Trp Leu
Val Gly Phe Leu Leu Ala Leu Pro Glu Ile Leu Phe Ala Lys 130
135 140Val Ser Gln Gly His His Asn Asn Ser Leu Pro
Arg Cys Thr Phe Ser145 150 155
160Gln Glu Asn Gln Ala Glu Thr His Ala Trp Phe Thr Ser Arg Phe Leu
165 170 175Tyr His Val Ala
Gly Phe Leu Leu Pro Met Leu Val Met Gly Trp Cys 180
185 190Tyr Val Gly Val Val His Arg Leu Arg Gln Ala
Gln Arg Arg Pro Gln 195 200 205Arg
Gln Lys Ala Val Arg Val Ala Ile Leu Val Thr Ser Ile Phe Phe 210
215 220Leu Cys Trp Ser Pro Tyr His Ile Val Ile
Phe Leu Asp Thr Leu Ala225 230 235
240Arg Leu Lys Ala Val Asp Asn Thr Cys Lys Leu Asn Gly Ser Leu
Pro 245 250 255Val Ala Ile
Thr Met Cys Glu Phe Leu Gly Leu Ala His Cys Cys Leu 260
265 270Asn Pro Met Leu Tyr Thr Phe Ala Gly Val
Lys Phe Arg Ser Asp Leu 275 280
285Ser Arg Leu Leu Thr Lys Leu Gly Cys Thr Gly Pro Ala Ser Leu Cys 290
295 300Gln Leu Phe Pro Ser Trp Arg Arg
Ser Ser Leu Ser Glu Ser Glu Asn305 310
315 320Ala Thr Ser Leu Thr Thr Phe
32513372PRTHomo sapiens 13Met Asn Tyr Pro Leu Thr Leu Glu Met Asp Leu Glu
Asn Leu Glu Asp1 5 10
15Leu Phe Trp Glu Leu Asp Arg Leu Asp Asn Tyr Asn Asp Thr Ser Leu
20 25 30Val Glu Asn His Leu Cys Pro
Ala Thr Glu Gly Pro Leu Met Ala Ser 35 40
45Phe Lys Ala Val Phe Val Pro Val Ala Tyr Ser Leu Ile Phe Leu
Leu 50 55 60Gly Val Ile Gly Asn Val
Leu Val Leu Val Ile Leu Glu Arg His Arg65 70
75 80Gln Thr Arg Ser Ser Thr Glu Thr Phe Leu Phe
His Leu Ala Val Ala 85 90
95Asp Leu Leu Leu Val Phe Ile Leu Pro Phe Ala Val Ala Glu Gly Ser
100 105 110Val Gly Trp Val Leu Gly
Thr Phe Leu Cys Lys Thr Val Ile Ala Leu 115 120
125His Lys Val Asn Phe Tyr Cys Ser Ser Leu Leu Leu Ala Cys
Ile Ala 130 135 140Val Asp Arg Tyr Leu
Ala Ile Val His Ala Val His Ala Tyr Arg His145 150
155 160Arg Arg Leu Leu Ser Ile His Ile Thr Cys
Gly Thr Ile Trp Leu Val 165 170
175Gly Phe Leu Leu Ala Leu Pro Glu Ile Leu Phe Ala Lys Val Ser Gln
180 185 190Gly His His Asn Asn
Ser Leu Pro Arg Cys Thr Phe Ser Gln Glu Asn 195
200 205Gln Ala Glu Thr His Ala Trp Phe Thr Ser Arg Phe
Leu Tyr His Val 210 215 220Ala Gly Phe
Leu Leu Pro Met Leu Val Met Gly Trp Cys Tyr Val Gly225
230 235 240Val Val His Arg Leu Arg Gln
Ala Gln Arg Arg Pro Gln Arg Gln Lys 245
250 255Ala Val Arg Val Ala Ile Leu Val Thr Ser Ile Phe
Phe Leu Cys Trp 260 265 270Ser
Pro Tyr His Ile Val Ile Phe Leu Asp Thr Leu Ala Arg Leu Lys 275
280 285Ala Val Asp Asn Thr Cys Lys Leu Asn
Gly Ser Leu Pro Val Ala Ile 290 295
300Thr Met Cys Glu Phe Leu Gly Leu Ala His Cys Cys Leu Asn Pro Met305
310 315 320Leu Tyr Thr Phe
Ala Gly Val Lys Phe Arg Ser Asp Leu Ser Arg Leu 325
330 335Leu Thr Lys Leu Gly Cys Thr Gly Pro Ala
Ser Leu Cys Gln Leu Phe 340 345
350Pro Ser Trp Arg Arg Ser Ser Leu Ser Glu Ser Glu Asn Ala Thr Ser
355 360 365Leu Thr Thr Phe
37014109PRTHomo sapiens 14Met Lys Phe Ile Ser Thr Ser Leu Leu Leu Met Leu
Leu Val Ser Ser1 5 10
15Leu Ser Pro Val Gln Gly Val Leu Glu Val Tyr Tyr Thr Ser Leu Arg
20 25 30Cys Arg Cys Val Gln Glu Ser
Ser Val Phe Ile Pro Arg Arg Phe Ile 35 40
45Asp Arg Ile Gln Ile Leu Pro Arg Gly Asn Gly Cys Pro Arg Lys
Glu 50 55 60Ile Ile Val Trp Lys Lys
Asn Lys Ser Ile Val Cys Val Asp Pro Gln65 70
75 80Ala Glu Trp Ile Gln Arg Met Met Glu Val Leu
Arg Lys Arg Ser Ser 85 90
95Ser Thr Leu Pro Val Pro Val Phe Lys Arg Lys Ile Pro 100
10515342PRTHomo sapiens 15Met Ala Glu His Asp Tyr His Glu Asp
Tyr Gly Phe Ser Ser Phe Asn1 5 10
15Asp Ser Ser Gln Glu Glu His Gln Asp Phe Leu Gln Phe Ser Lys
Val 20 25 30Phe Leu Pro Cys
Met Tyr Leu Val Val Phe Val Cys Gly Leu Val Gly 35
40 45Asn Ser Leu Val Leu Val Ile Ser Ile Phe Tyr His
Lys Leu Gln Ser 50 55 60Leu Thr Asp
Val Phe Leu Val Asn Leu Pro Leu Ala Asp Leu Val Phe65 70
75 80Val Cys Thr Leu Pro Phe Trp Ala
Tyr Ala Gly Ile His Glu Trp Val 85 90
95Phe Gly Gln Val Met Cys Lys Ser Leu Leu Gly Ile Tyr Thr
Ile Asn 100 105 110Phe Tyr Thr
Ser Met Leu Ile Leu Thr Cys Ile Thr Val Asp Arg Phe 115
120 125Ile Val Val Val Lys Ala Thr Lys Ala Tyr Asn
Gln Gln Ala Lys Arg 130 135 140Met Thr
Trp Gly Lys Val Thr Ser Leu Leu Ile Trp Val Ile Ser Leu145
150 155 160Leu Val Ser Leu Pro Gln Ile
Ile Tyr Gly Asn Val Phe Asn Leu Asp 165
170 175Lys Leu Ile Cys Gly Tyr His Asp Glu Ala Ile Ser
Thr Val Val Leu 180 185 190Ala
Thr Gln Met Thr Leu Gly Phe Phe Leu Pro Leu Leu Thr Met Ile 195
200 205Val Cys Tyr Ser Val Ile Ile Lys Thr
Leu Leu His Ala Gly Gly Phe 210 215
220Gln Lys His Arg Ser Leu Lys Ile Ile Phe Leu Val Met Ala Val Phe225
230 235 240Leu Leu Thr Gln
Met Pro Phe Asn Leu Met Lys Phe Ile Arg Ser Thr 245
250 255His Trp Glu Tyr Tyr Ala Met Thr Ser Phe
His Tyr Thr Ile Met Val 260 265
270Thr Glu Ala Ile Ala Tyr Leu Arg Ala Cys Leu Asn Pro Val Leu Tyr
275 280 285Ala Phe Val Ser Leu Lys Phe
Arg Lys Asn Phe Trp Lys Leu Val Lys 290 295
300Asp Ile Gly Cys Leu Pro Tyr Leu Gly Val Ser His Gln Trp Lys
Ser305 310 315 320Ser Glu
Asp Asn Ser Lys Thr Phe Ser Ala Ser His Asn Val Glu Ala
325 330 335Thr Ser Met Phe Gln Leu
34016120PRTHomo sapiens 16Met Lys Val Ser Glu Ala Ala Leu Ser Leu Leu
Val Leu Ile Leu Ile1 5 10
15Ile Thr Ser Ala Ser Arg Ser Gln Pro Lys Val Pro Glu Trp Val Asn
20 25 30Thr Pro Ser Thr Cys Cys Leu
Lys Tyr Tyr Glu Lys Val Leu Pro Arg 35 40
45Arg Leu Val Val Gly Tyr Arg Lys Ala Leu Asn Cys His Leu Pro
Ala 50 55 60Ile Ile Phe Val Thr Lys
Arg Asn Arg Glu Val Cys Thr Asn Pro Asn65 70
75 80Asp Asp Trp Val Gln Glu Tyr Ile Lys Asp Pro
Asn Leu Pro Leu Leu 85 90
95Pro Thr Arg Asn Leu Ser Thr Val Lys Ile Ile Thr Ala Lys Asn Gly
100 105 110Gln Pro Gln Leu Leu Asn
Ser Gln 115 12017120PRTHomo sapiens 17Met Lys Val
Ser Glu Ala Ala Leu Ser Leu Leu Val Leu Ile Leu Ile1 5
10 15Ile Thr Ser Ala Ser Arg Ser Gln Pro
Lys Val Pro Glu Trp Val Asn 20 25
30Thr Pro Ser Thr Cys Cys Leu Lys Tyr Tyr Glu Lys Val Leu Pro Arg
35 40 45Arg Leu Val Val Gly Tyr Arg
Lys Ala Leu Asn Cys His Leu Pro Ala 50 55
60Ile Ile Phe Val Thr Lys Arg Asn Arg Glu Val Cys Thr Asn Pro Asn65
70 75 80Asp Asp Trp Val
Gln Glu Tyr Ile Lys Asp Pro Asn Leu Pro Leu Leu 85
90 95Pro Thr Arg Asn Leu Ser Thr Val Lys Ile
Ile Thr Ala Lys Asn Gly 100 105
110Gln Pro Gln Leu Leu Asn Ser Gln 115
12018150PRTHomo sapiens 18Met Asn Leu Trp Leu Leu Ala Cys Leu Val Ala Gly
Phe Leu Gly Ala1 5 10
15Trp Ala Pro Ala Val His Thr Gln Gly Val Phe Glu Asp Cys Cys Leu
20 25 30Ala Tyr His Tyr Pro Ile Gly
Trp Ala Val Leu Arg Arg Ala Trp Thr 35 40
45Tyr Arg Ile Gln Glu Val Ser Gly Ser Cys Asn Leu Pro Ala Ala
Ile 50 55 60Phe Tyr Leu Pro Lys Arg
His Arg Lys Val Cys Gly Asn Pro Lys Ser65 70
75 80Arg Glu Val Gln Arg Ala Met Lys Leu Leu Asp
Ala Arg Asn Lys Val 85 90
95Phe Ala Lys Leu His His Asn Thr Gln Thr Phe Gln Ala Gly Pro His
100 105 110Ala Val Lys Lys Leu Ser
Ser Gly Asn Ser Lys Leu Ser Ser Ser Lys 115 120
125Phe Ser Asn Pro Ile Ser Ser Ser Lys Arg Asn Val Ser Leu
Leu Ile 130 135 140Ser Ala Asn Ser Gly
Leu145 15019150PRTHomo sapiens 19Met Asn Leu Trp Leu Leu
Ala Cys Leu Val Ala Gly Phe Leu Gly Ala1 5
10 15Trp Ala Pro Ala Val His Thr Gln Gly Val Phe Glu
Asp Cys Cys Leu 20 25 30Ala
Tyr His Tyr Pro Ile Gly Trp Ala Val Leu Arg Arg Ala Trp Thr 35
40 45Tyr Arg Ile Gln Glu Val Ser Gly Ser
Cys Asn Leu Pro Ala Ala Ile 50 55
60Phe Tyr Leu Pro Lys Arg His Arg Lys Val Cys Gly Asn Pro Lys Ser65
70 75 80Arg Glu Val Gln Arg
Ala Met Lys Leu Leu Asp Ala Arg Asn Lys Val 85
90 95Phe Ala Lys Leu His His Asn Thr Gln Thr Phe
Gln Ala Gly Pro His 100 105
110Ala Val Lys Lys Leu Ser Ser Gly Asn Ser Lys Leu Ser Ser Ser Lys
115 120 125Phe Ser Asn Pro Ile Ser Ser
Ser Lys Arg Asn Val Ser Leu Leu Ile 130 135
140Ser Ala Asn Ser Gly Leu145 1502084PRTHomo sapiens
20Met Asn Leu Trp Leu Leu Ala Cys Leu Val Ala Gly Phe Leu Gly Ala1
5 10 15Trp Ala Pro Ala Val His
Thr Gln Gly Val Phe Glu Asp Cys Cys Leu 20 25
30Ala Tyr His Tyr Pro Ile Gly Trp Ala Val Leu Arg Arg
Ala Trp Thr 35 40 45Tyr Arg Ile
Gln Glu Val Ser Gly Ser Cys Asn Leu Pro Ala Ala Ile 50
55 60Arg Pro Ser Cys Cys Lys Glu Val Glu Phe Trp Lys
Leu Gln Val Ile65 70 75
80Ile Val Gln Val21355PRTHomo sapiens 21Met Asp Gln Phe Pro Glu Ser Val
Thr Glu Asn Phe Glu Tyr Asp Asp1 5 10
15Leu Ala Glu Ala Cys Tyr Ile Gly Asp Ile Val Val Phe Gly
Thr Val 20 25 30Phe Leu Ser
Ile Phe Tyr Ser Val Ile Phe Ala Ile Gly Leu Val Gly 35
40 45Asn Leu Leu Val Val Phe Ala Leu Thr Asn Ser
Lys Lys Pro Lys Ser 50 55 60Val Thr
Asp Ile Tyr Leu Leu Asn Leu Ala Leu Ser Asp Leu Leu Phe65
70 75 80Val Ala Thr Leu Pro Phe Trp
Thr His Tyr Leu Ile Asn Glu Lys Gly 85 90
95Leu His Asn Ala Met Cys Lys Phe Thr Thr Ala Phe Phe
Phe Ile Gly 100 105 110Phe Phe
Gly Ser Ile Phe Phe Ile Thr Val Ile Ser Ile Asp Arg Tyr 115
120 125Leu Ala Ile Val Leu Ala Ala Asn Ser Met
Asn Asn Arg Thr Val Gln 130 135 140His
Gly Val Thr Ile Ser Leu Gly Val Trp Ala Ala Ala Ile Leu Val145
150 155 160Ala Ala Pro Gln Phe Met
Phe Thr Lys Gln Lys Glu Asn Glu Cys Leu 165
170 175Gly Asp Tyr Pro Glu Val Leu Gln Glu Ile Trp Pro
Val Leu Arg Asn 180 185 190Val
Glu Thr Asn Phe Leu Gly Phe Leu Leu Pro Leu Leu Ile Met Ser 195
200 205Tyr Cys Tyr Phe Arg Ile Ile Gln Thr
Leu Phe Ser Cys Lys Asn His 210 215
220Lys Lys Ala Lys Ala Ile Lys Leu Ile Leu Leu Val Val Ile Val Phe225
230 235 240Phe Leu Phe Trp
Thr Pro Tyr Asn Val Met Ile Phe Leu Glu Thr Leu 245
250 255Lys Leu Tyr Asp Phe Phe Pro Ser Cys Asp
Met Arg Lys Asp Leu Arg 260 265
270Leu Ala Leu Ser Val Thr Glu Thr Val Ala Phe Ser His Cys Cys Leu
275 280 285Asn Pro Leu Ile Tyr Ala Phe
Ala Gly Glu Lys Phe Arg Arg Tyr Leu 290 295
300Tyr His Leu Tyr Gly Lys Cys Leu Ala Val Leu Cys Gly Arg Ser
Val305 310 315 320His Val
Asp Phe Ser Ser Ser Glu Ser Gln Arg Ser Arg His Gly Ser
325 330 335Val Leu Ser Ser Asn Phe Thr
Tyr His Thr Ser Asp Gly Asp Ala Leu 340 345
350Leu Leu Leu 35522397PRTHomo sapiens 22Met Ala Pro
Ile Ser Leu Ser Trp Leu Leu Arg Leu Ala Thr Phe Cys1 5
10 15His Leu Thr Val Leu Leu Ala Gly Gln
His His Gly Val Thr Lys Cys 20 25
30Asn Ile Thr Cys Ser Lys Met Thr Ser Lys Ile Pro Val Ala Leu Leu
35 40 45Ile His Tyr Gln Gln Asn Gln
Ala Ser Cys Gly Lys Arg Ala Ile Ile 50 55
60Leu Glu Thr Arg Gln His Arg Leu Phe Cys Ala Asp Pro Lys Glu Gln65
70 75 80Trp Val Lys Asp
Ala Met Gln His Leu Asp Arg Gln Ala Ala Ala Leu 85
90 95Thr Arg Asn Gly Gly Thr Phe Glu Lys Gln
Ile Gly Glu Val Lys Pro 100 105
110Arg Thr Thr Pro Ala Ala Gly Gly Met Asp Glu Ser Val Val Leu Glu
115 120 125Pro Glu Ala Thr Gly Glu Ser
Ser Ser Leu Glu Pro Thr Pro Ser Ser 130 135
140Gln Glu Ala Gln Arg Ala Leu Gly Thr Ser Pro Glu Leu Pro Thr
Gly145 150 155 160Val Thr
Gly Ser Ser Gly Thr Arg Leu Pro Pro Thr Pro Lys Ala Gln
165 170 175Asp Gly Gly Pro Val Gly Thr
Glu Leu Phe Arg Val Pro Pro Val Ser 180 185
190Thr Ala Ala Thr Trp Gln Ser Ser Ala Pro His Gln Pro Gly
Pro Ser 195 200 205Leu Trp Ala Glu
Ala Lys Thr Ser Glu Ala Pro Ser Thr Gln Asp Pro 210
215 220Ser Thr Gln Ala Ser Thr Ala Ser Ser Pro Ala Pro
Glu Glu Asn Ala225 230 235
240Pro Ser Glu Gly Gln Arg Val Trp Gly Gln Gly Gln Ser Pro Arg Pro
245 250 255Glu Asn Ser Leu Glu
Arg Glu Glu Met Gly Pro Val Pro Ala His Thr 260
265 270Asp Ala Phe Gln Asp Trp Gly Pro Gly Ser Met Ala
His Val Ser Val 275 280 285Val Pro
Val Ser Ser Glu Gly Thr Pro Ser Arg Glu Pro Val Ala Ser 290
295 300Gly Ser Trp Thr Pro Lys Ala Glu Glu Pro Ile
His Ala Thr Met Asp305 310 315
320Pro Gln Arg Leu Gly Val Leu Ile Thr Pro Val Pro Asp Ala Gln Ala
325 330 335Ala Thr Arg Arg
Gln Ala Val Gly Leu Leu Ala Phe Leu Gly Leu Leu 340
345 350Phe Cys Leu Gly Val Ala Met Phe Thr Tyr Gln
Ser Leu Gln Gly Cys 355 360 365Pro
Arg Lys Met Ala Gly Glu Met Ala Glu Gly Leu Arg Tyr Ile Pro 370
375 380Arg Ser Cys Gly Ser Asn Ser Tyr Val Leu
Val Pro Val385 390 395232502DNAHomo
sapiens 23tattcatcaa gtgccctcta gctgttaagt cactctgatc tctgactgca
gctcctactg 60ttggacacac ctggccggtg cttcagttag atcaaaccat tgctgaaact
gaagaggaca 120tgtcaaatat tacagatcca cagatgtggg attttgatga tctaaatttc
actggcatgc 180cacctgcaga tgaagattac agcccctgta tgctagaaac tgagacactc
aacaagtatg 240ttgtgatcat cgcctatgcc ctagtgttcc tgctgagcct gctgggaaac
tccctggtga 300tgctggtcat cttatacagc agggtcggcc gctccgtcac tgatgtctac
ctgctgaacc 360tggccttggc cgacctactc tttgccctga ccttgcccat ctgggccgcc
tccaaggtga 420atggctggat ttttggcaca ttcctgtgca aggtggtctc actcctgaag
gaagtcaact 480tctacagtgg catcctgctg ttggcctgca tcagtgtgga ccgttacctg
gccattgtcc 540atgccacacg cacactgacc cagaagcgtc acttggtcaa gtttgtttgt
cttggctgct 600ggggactgtc tatgaatctg tccctgccct tcttcctttt ccgccaggct
taccatccaa 660acaattccag tccagtttgc tatgaggtcc tgggaaatga cacagcaaaa
tggcggatgg 720tgttgcggat cctgcctcac acctttggct tcatcgtgcc gctgtttgtc
atgctgttct 780gctatggatt caccctgcgt acactgttta aggcccacat ggggcagaag
caccgagcca 840tgagggtcat ctttgctgtc gtcctcatct tcctgctttg ctggctgccc
tacaacctgg 900tcctgctggc agacaccctc atgaggaccc aggtgatcca ggagagctgt
gagcgccgca 960acaacatcgg ccgggccctg gatgccactg agattctggg atttctccat
agctgcctca 1020accccatcat ctacgccttc atcggccaaa attttcgcca tggattcctc
aagatcctgg 1080ctatgcatgg cctggtcagc aaggagttct tggcacgtca tcgtgttacc
tcctacactt 1140cttcgtctgt caatgtctct tccaacctct gaaaaccatc gatgaaggaa
tatctcttct 1200cagaaggaaa gaataaccaa caccctgagg ttgtgtgtgg aaggtgatct
ggctctggac 1260aggcactatc tgggttttgg ggggacgcta taggatgtgg ggaagttagg
aactggtgtc 1320ttcaggggcc acaccaacct tctgaggagc tgttgaggta cctccaagga
ccggcctttg 1380cacctccatg gaaacgaagc accatcattc ccgttgaacg tcacatcttt
aacccactaa 1440ctggctaatt agcatggcca catctgagcc ccgaatctga cattagatga
gagaacaggg 1500ctgaagctgt gtcctcatga gggctggatg ctctcgttga ccctcacagg
agcatctcct 1560caactctgag tgttaagcgt tgagccacca agctggtggc tctgtgtgct
ctgatccgag 1620ctcagggggg tggttttccc atctcaggtg tgttgcagtg tctgctggag
acattgaggc 1680aggcactgcc aaaacatcaa cctgccagct ggccttgtga ggagctggaa
acacatgttc 1740cccttggggg tggtggatga acaaagagaa agagggtttg gaagccagat
ctatgccaca 1800agaaccccct ttacccccat gaccaacatc gcagacacat gtgctggcca
cctgctgagc 1860cccaagtgga acgagacaag cagcccttag cccttcccct ctgcagcttc
caggctggcg 1920tgcagcatca gcatccctag aaagccatgt gcagccacca gtccattggg
caggcagatg 1980ttcctaataa agcttctgtt ccgtgcttgt ccctgtggaa gtatcttggt
tgtgacagag 2040tcaagggtgt gtgcagcatt gttggctgtt cctgcagtag aatgggggca
gcacctccta 2100agaaggcacc tctctgggtt gaagggcagt gttccctggg gctttaactc
ctgctagaac 2160agtctcttga ggcacagaaa ctcctgttca tgcccatacc cctggccaag
gaagatccct 2220ttgtccacaa gtaaaaggaa atgctcctcc agggagtctc agcttcaccc
tgaggtgagc 2280atcatcttct gggttaggcc ttgcctaggc atagccctgc ctcaagctat
gtgagctcac 2340cagtccctcc ccaaatgctt tccatgagtt gcagtttttt cctagtctgt
tttccctcct 2400tggagacagg gccctgtcgg tttattcact gtatgtcctt ggtgcctgga
gcctactaaa 2460tgctcaataa ataatgatca caggaaaaaa aaaaaaaaaa aa
2502242880DNAHomo sapiens 24aggttcaaaa cattcagaga cagaaggtgg
atagacaaat ctccaccttc agactggtag 60gctcctccag aagccatcag acaggaagat
gtgaaaatcc ccagcactca tcccagaatc 120actaagtggc acctgtcctg ggccaaagtc
ccaggacaga cctcattgtt cctctgtggg 180aatacctccc caggagggca tcctggattt
cccccttgca acccaggtca gaagtttcat 240cgtcaaggtt gtttcatctt ttttttcctg
tctaacagct ctgactacca cccaaccttg 300aggcacagtg aagacatcgg tggccactcc
aataacagca ggtcacagct gctcttctgg 360aggtgtccta caggtgaaaa gcccagcgac
ccagtcagga tttaagttta cctcaaaaat 420ggaagatttt aacatggaga gtgacagctt
tgaagatttc tggaaaggtg aagatcttag 480taattacagt tacagctcta ccctgccccc
ttttctacta gatgccgccc catgtgaacc 540agaatccctg gaaatcaaca agtattttgt
ggtcattatc tatgccctgg tattcctgct 600gagcctgctg ggaaactccc tcgtgatgct
ggtcatctta tacagcaggg tcggccgctc 660cgtcactgat gtctacctgc tgaacctagc
cttggccgac ctactctttg ccctgacctt 720gcccatctgg gccgcctcca aggtgaatgg
ctggattttt ggcacattcc tgtgcaaggt 780ggtctcactc ctgaaggaag tcaacttcta
tagtggcatc ctgctactgg cctgcatcag 840tgtggaccgt tacctggcca ttgtccatgc
cacacgcaca ctgacccaga agcgctactt 900ggtcaaattc atatgtctca gcatctgggg
tctgtccttg ctcctggccc tgcctgtctt 960acttttccga aggaccgtct actcatccaa
tgttagccca gcctgctatg aggacatggg 1020caacaataca gcaaactggc ggatgctgtt
acggatcctg ccccagtcct ttggcttcat 1080cgtgccactg ctgatcatgc tgttctgcta
cggattcacc ctgcgtacgc tgtttaaggc 1140ccacatgggg cagaagcacc gggccatgcg
ggtcatcttt gctgtcgtcc tcatcttcct 1200gctctgctgg ctgccctaca acctggtcct
gctggcagac accctcatga ggacccaggt 1260gatccaggag acctgtgagc gccgcaatca
catcgaccgg gctctggatg ccaccgagat 1320tctgggcatc cttcacagct gcctcaaccc
cctcatctac gccttcattg gccagaagtt 1380tcgccatgga ctcctcaaga ttctagctat
acatggcttg atcagcaagg actccctgcc 1440caaagacagc aggccttcct ttgttggctc
ttcttcaggg cacacttcca ctactctcta 1500agacctcctg cctaagtgca gccccgtggg
gttcctccct tctcttcaca gtcacattcc 1560aagcctcatg tccactggtt cttcttggtc
tcagtgtcaa tgcagccccc attgtggtca 1620caggaagtag aggaggccac gttcttacta
gtttcccttg catggtttag aaagcttgcc 1680ctggtgcctc accccttgcc ataattacta
tgtcatttgc tggagctctg cccatcctgc 1740ccctgagccc atggcactct atgttctaag
aagtgaaaat ctacactcca gtgagacagc 1800tctgcatact cattaggatg gctagtatca
aaagaaagaa aatcaggctg gccaacgggg 1860tgaaaccctg tctctactaa aaatacaaaa
aaaaaaaaaa attagccggg cgtggtggtg 1920agtgcctgta atcacagcta cttgggaggc
tgagatggga gaatcacttg aacccgggag 1980gcagaggttg cagtgagccg agattgtgcc
cctgcactcc agcctgagcg acagtgagac 2040tctgtctcag tccatgaaga tgtagaggag
aaactggaac tctcgagcgt tgctgggggg 2100gattgtaaaa tggtgtgacc actgcagaag
acagtatggc agctttcctc aaaacttcag 2160acatagaatt aacacatgat cctgcaattc
cacttatagg aattgaccca caagaaatga 2220aagcagggac ttgaacccat atttgtacac
caatattcat agcagcttat tcacaagacc 2280caaaaggcag aagcaaccca aatgttcatc
aatgaatgaa tgaatggcta agcaaaatgt 2340gatatgtacc taacgaagta tccttcagcc
tgaaagagga atgaagtact catacatgtt 2400acaacacgga cgaaccttga aaactttatg
ctaagtgaaa taagccagac atcaacagat 2460aaatagttta tgattccacc tacatgaggt
actgagagtg aacaaattta cagagacaga 2520aagcagaaca gtgattacca gggactgagg
ggaggggagc atgggaagtg acggtttaat 2580gggcacaggg tttatgttta ggatgttgaa
aaagttctgc agataaacag tagtgatagt 2640tgtaccgcaa tgtgacttaa tgccactaaa
ttgacactta aaaatggttt aaatggtcaa 2700ttttgttatg tatattttat atcaatttaa
aaaaaaacct gagccccaaa aggtatttta 2760atcaccaagg ctgattaaac caaggctaga
accacctgcc tatatttttt gttaaatgat 2820ttcattcaat atcttttttt taataaacca
tttttacttg ggtgtttata aaaaaaaaaa 2880251119DNAHomo sapiens 25cacagagccc
gggccgcagg cacctcctcg ccagctcttc cgctcctctc acagccgcca 60gacccgcctg
ctgagcccca tggcccgcgc tgctctctcc gccgccccca gcaatccccg 120gctcctgcga
gtggcactgc tgctcctgct cctggtagcc gctggccggc gcgcagcagg 180agcgtccgtg
gccactgaac tgcgctgcca gtgcttgcag accctgcagg gaattcaccc 240caagaacatc
caaagtgtga acgtgaagtc ccccggaccc cactgcgccc aaaccgaagt 300catagccaca
ctcaagaatg ggcggaaagc ttgcctcaat cctgcatccc ccatagttaa 360gaaaatcatc
gaaaagatgc tgaacagtga caaatccaac tgaccagaag ggaggaggaa 420gctcactggt
ggctgttcct gaaggaggcc ctgcccttat aggaacagaa gaggaaagag 480agacacagct
gcagaggcca cctggattgt gcctaatgtg tttgagcatc gcttaggaga 540agtcttctat
ttatttattt attcattagt tttgaagatt ctatgttaat attttaggtg 600taaaataatt
aagggtatga ttaactctac ctgcacactg tcctattata ttcattcttt 660ttgaaatgtc
aaccccaagt tagttcaatc tggattcata tttaatttga aggtagaatg 720ttttcaaatg
ttctccagtc attatgttaa tatttctgag gagcctgcaa catgccagcc 780actgtgatag
aggctggcgg atccaagcaa atggccaatg agatcattgt gaaggcaggg 840gaatgtatgt
gcacatctgt tttgtaactg tttagatgaa tgtcagttgt tatttattga 900aatgatttca
cagtgtgtgg tcaacatttc tcatgttgaa actttaagaa ctaaaatgtt 960ctaaatatcc
cttggacatt ttatgtcttt cttgtaaggc atactgcctt gtttaatggt 1020agttttacag
tgtttctggc ttagaacaaa ggggcttaat tattgatgtt ttcatagaga 1080atataaaaat
aaagcactta tagaaaaaaa aaaaaaaaa
1119261234DNAHomo sapiens 26gagctccggg aatttccctg gcccgggact ccgggctttc
cagccccaac catgcataaa 60aggggttcgc cgttctcgga gagccacaga gcccgggcca
caggcagctc cttgccagct 120ctcctcctcg cacagccgct cgaaccgcct gctgagcccc
atggcccgcg ccacgctctc 180cgccgccccc agcaatcccc ggctcctgcg ggtggcgctg
ctgctcctgc tcctggtggc 240cgccagccgg cgcgcagcag gagcgcccct ggccactgaa
ctgcgctgcc agtgcttgca 300gaccctgcag ggaattcacc tcaagaacat ccaaagtgtg
aaggtgaagt cccccggacc 360ccactgcgcc caaaccgaag tcatagccac actcaagaat
gggcagaaag cttgtctcaa 420ccccgcatcg cccatggtta agaaaatcat cgaaaagatg
ctgaaaaatg gcaaatccaa 480ctgaccagaa ggaaggagga agcttattgg tggctgttcc
tgaaggaggc cctgccctta 540caggaacaga agaggaaaga gagacacagc tgcagaggcc
acctggattg cgcctaatgt 600gtttgagcat cacttaggag aagtcttcta tttatttatt
tatttattta tttgtttgtt 660ttagaagatt ctatgttaat attttatgtg taaaataagg
ttatgattga atctacttgc 720acactctccc attatattta ttgtttattt taggtcaaac
ccaagttagt tcaatcctga 780ttcatattta atttgaagat agaaggtttg cagatattct
ctagtcattt gttaatattt 840cttcgtgatg acatatcaca tgtcagccac tgtgatagag
gctgaggaat ccaagaaaat 900ggccagtgag atcaatgtga cggcagggaa atgtatgtgt
gtctattttg taactgtaaa 960gatgaatgtc agttgttatt tattgaaatg atttcacagt
gtgtggtcaa catttctcat 1020gttgaagctt taagaactaa aatgttctaa atatcccttg
gacattttat gtctttcttg 1080taaggcatac tgccttgttt aatgttaatt atgcagtgtt
tccctctgtg ttagagcaga 1140gaggtttcga tatttattga tgttttcaca aagaacagga
aaataaaata tttaaaaata 1200taaaaaaaaa aaaaaaaaaa aaaaaaaaaa aaaa
1234271166DNAHomo sapiens 27gctccgggaa tttccctggc
ccggccgctc cgggctttcc agtctcaacc atgcataaaa 60agggttcgcc gatcttgggg
agccacacag cccgggtcgc aggcacctcc ccgccagctc 120tcccgcttct cgcacagctt
cccgacgcgt ctgctgagcc ccatggccca cgccacgctc 180tccgccgccc ccagcaatcc
ccggctcctg cgggtggcgc tgctgctcct gctcctggtg 240gccgccagcc ggcgcgcagc
aggagcgtcc gtggtcactg aactgcgctg ccagtgcttg 300cagacactgc agggaattca
cctcaagaac atccaaagtg tgaatgtaag gtcccccgga 360ccccactgcg cccaaaccga
agtcatagcc acactcaaga atgggaagaa agcttgtctc 420aaccccgcat cccccatggt
tcagaaaatc atcgaaaaga tactgaacaa ggggagcacc 480aactgacagg agagaagtaa
gaagcttatc agcgtatcat tgacacttcc tgcagggtgg 540tccctgccct taccagagct
gaaaatgaaa aagagaacag cagctttcta gggacagctg 600gaaaggactt aatgtgtttg
actatttctt acgagggttc tacttattta tgtatttatt 660tttgaaagct tgtattttaa
tattttacat gctgttattt aaagatgtga gtgtgtttca 720tcaaacatag ctcagtcctg
attatttaat tggaatatga tgggttttaa atgtgtcatt 780aaactaatat ttagtgggag
accataatgt gtcagccacc ttgataaatg acagggtggg 840gaactggagg gtggggggat
tgaaatgcaa gcaattagtg gatcactgtt agggtaaggg 900aatgtatgta cacatctatt
ttttatactt tttttttaaa aaaagaatgt cagttgttat 960ttattcaaat tatctcacat
tatgtgttca acatttttat gctgaagttt cccttagaca 1020ttttatgtct tgcttgtagg
gcataatgcc ttgtttaatg tccattctgc agcgtttctc 1080tttcccttgg aaaagagaat
ttatcattac tgttacattt gtacaaatga catgataata 1140aaagttttat gaaaaaaaaa
aaaaaa 1166282475DNAHomo sapiens
28gtgcagaagg cacgaggaag ccacagtgct ccggatcctc caatcttcgc tcctccaatc
60tccgctcctc cacccagttc aggaacccgc gaccgctcgc agcgctctct tgaccactat
120gagcctcctg tccagccgcg cggcccgtgt ccccggtcct tcgagctcct tgtgcgcgct
180gttggtgctg ctgctgctgc tgacgcagcc agggcccatc gccagcgctg gtcctgccgc
240tgctgtgttg agagagctgc gttgcgtttg tttacagacc acgcaaggag ttcatcccaa
300aatgatcagt aatctgcaag tgttcgccat aggcccacag tgctccaagg tggaagtggt
360agcctccctg aagaacggga aggaaatttg tcttgatcca gaagcccctt ttctaaagaa
420agtcatccag aaaattttgg acggtggaaa caaggaaaac tgattaagag aaatgagcac
480gcatggaaaa gtttcccagt cttcagcaga gaagttttct ggaggtctct gaacccaggg
540aagacaagaa ggaaagattt tgttgttgtt tgtttatttg tttttccagt agttagcttt
600cttcctggat tcctcacttt gaagagtgtg aggaaaacct atgtttgccg cttaagcttt
660cagctcagct aatgaagtgt ttagcatagt acctctgcta tttgctgtta ttttatctgc
720tatgctattg aagttttggc aattgactat agtgtgagcc aggaatcact ggctgttaat
780ctttcaaagt gtcttgaatt gtaggtgact attatatttc caagaaatat tccttaagat
840attaactgag aaggctgtgg atttaatgtg gaaatgatgt ttcataagaa ttctgttgat
900ggaaatacac tgttatcttc acttttataa gaaataggaa atattttaat gtttcttggg
960gaatatgtta gagaatttcc ttactcttga ttgtgggata ctatttaatt atttcacttt
1020agaaagctga gtgtttcaca ccttatctat gtagaatata tttccttatt cagaatttct
1080aaaagtttaa gttctatgag ggctaatatc ttatcttcct ataattttag acattcttta
1140tctttttagt atggcaaact gccatcattt acttttaaac tttgatttta tatgctattt
1200attaagtatt ttattaggag taccataatt ctggtagcta aatatatatt ttagatagat
1260gaagaagcta gaaaacaggc aaattcctga ctgctagttt atatagaaat gtattctttt
1320agtttttaaa gtaaaggcaa acttaacaat gacttgtact ctgaaagttt tggaaacgta
1380ttcaaacaat ttgaatataa atttatcatt tagttataaa aatatatagc gacatcctcg
1440aggccctagc atttctcctt ggatagggga ccagagagag cttggaatgt taaaaacaaa
1500acaaaacaaa aaaaaacaag gagaagttgt ccaagggatg tcaatttttt atccctctgt
1560atgggttaga ttttccaaaa tcataatttg aagaaggcca gcatttatgg tagaatatat
1620aattatatat aaggtggcca cgctggggca agttccctcc ccactcacag ctttggcccc
1680tttcacagag tagaacctgg gttagaggat tgcagaagac gagcggcagc ggggagggca
1740gggaagatgc ctgtcgggtt tttagcacag ttcatttcac tgggattttg aagcatttct
1800gtctgaatgt aaagcctgtt ctagtcctgg tgggacacac tggggttggg ggtgggggaa
1860gatgcggtaa tgaaaccggt tagtcagtgt tgtcttaata tccttgataa tgctgtaaag
1920tttattttta caaatatttc tgtttaagct atttcacctt tgtttggaaa tccttccctt
1980ttaaagagaa aatgtgacac ttgtgaaaag gcttgtagga aagctcctcc ctttttttct
2040ttaaaccttt aaatgacaaa cctaggtaat taatggttgt gaatttctat ttttgctttg
2100tttttaatga acatttgtct ttcagaatag gattctgtga taatatttaa atggcaaaaa
2160caaaacataa ttttgtgcaa ttaacaaagc tactgcaaga aaaataaaac atttcttggt
2220aaaaacgtat gtatttatat attatatatt tatatataat atatattata tatttagcat
2280tgctgagctt tttagatgcc tattgtgtat cttttaaagg ttttgaccat tttgttatga
2340gtaattacat atatattaca ttcactatat taaaattgta cttttttact atgtgtctca
2400ttggttcata gtctttattt tgtcctttga ataaacatta aaagatttct aaacttcaaa
2460aaaaaaaaaa aaaaa
2475291677DNAHomo sapiens 29accccttctt tccacactgc cccctgagtt cagggaattt
ccccagcatc ccaaagcttg 60agtttcctgc cagtcgggag ggatgaatgc agataaaggg
agtgcagaag gcacgaggaa 120accaaagtgc tctgtatcct ccagtctccg cgcctccacc
cagctcagga acccgcgaac 180cctctcttga ccactatgag cctcccgtcc agccgcgcgg
cccgtgtccc gggtccttcg 240ggctccttgt gcgcgctgct cgcgctgctg ctcctgctga
cgccgccggg gcccctcgcc 300agcgctggtc ctgtctctgc tgtgctgaca gagctgcgtt
gcacttgttt acgcgttacg 360ctgagagtaa accccaaaac gattggtaaa ctgcaggtgt
tccccgcagg cccgcagtgc 420tccaaggtgg aagtggtagc ctccctgaag aacgggaagc
aagtttgtct ggacccggaa 480gccccttttc taaagaaagt catccagaaa attttggaca
gtggaaacaa gaaaaactga 540gtaacaaaaa agaccatgca tcataaaatt gcccagtctt
cagcggagca gttttctgga 600gatccctgga cccagtaaga ataagaagga agggttggtt
tttttccatt ttctacatgg 660attccctact ttgaagagtg tgggggaaag cctacgcttc
tccctgaagt ttacagctca 720gctaatgaag tactaatata gtatttccac tatttactgt
tattttacct gataagttat 780tgaacccttt ggcaattgac catattgtga gcaaagaatc
actggttatt agtctttcaa 840tgaatattga attgaagata actattgtat ttctatcata
cattccttaa agtcttaccg 900aaaaggctgt ggatttcgta tggaaataat gttttattag
tgtgctgttg agggaggtat 960cctgttgttc ttactcactc ttctcataaa ataggaaata
ttttagttct gtttcttggg 1020gaatatgtta ctctttaccc taggatgcta tttaagttgt
actgtattag aacactgggt 1080gtgtcatacc gttatctgtg cagaatatat ttccttattc
agaatttcta aaaatttaag 1140ttctgtaagg gctaatatat tctcttccta tggttttaga
cgtttgatgt cttcttagta 1200tggcataatg tcatgattta ctcattaaac tttgattttg
tatgctattt tttcactata 1260ggatgactat aattctggtc actaaatata cactttagat
agatgaagaa gcccaaaaac 1320agataaattc ctgattgcta atttacatag aaatgtattc
tcttggtttt ttaaataaaa 1380gcaaaattaa caatgatctg tgctctgaaa gttttgaaaa
tatatttgaa caatttgaat 1440ataaattcat catttagtcc tcaaaatata tatagcattg
ctaagatttt cagatatcta 1500ttgtggatct tttaaaggtt ttgaccattt tgttatgagg
aattatacat gtatcacatt 1560cactatatta aaattgcact tttatttttt cctgtgtgtc
atgttggttt ttggtacttg 1620tattgtcatt tggagaaaca ataaaagatt tctaaaccaa
aaaaaaaaaa aaaaaaa 1677301307DNAHomo sapiens 30acttatctgc agacttgtag
gcagcaactc accctcactc agaggtcttc tggttctgga 60aacaactcta gctcagcctt
ctccaccatg agcctcagac ttgataccac cccttcctgt 120aacagtgcga gaccacttca
tgccttgcag gtgctgctgc ttctgtcatt gctgctgact 180gctctggctt cctccaccaa
aggacaaact aagagaaact tggcgaaagg caaagaggaa 240agtctagaca gtgacttgta
tgctgaactc cgctgcatgt gtataaagac aacctctgga 300attcatccca aaaacatcca
aagtttggaa gtgatcggga aaggaaccca ttgcaaccaa 360gtcgaagtga tagccacact
gaaggatggg aggaaaatct gcctggaccc agatgctccc 420agaatcaaga aaattgtaca
gaaaaaattg gcaggtgatg aatctgctga ttaatttgtt 480ctgtttctgc caaacttctt
taactcccag gaagggtaga attttgaaac cttgattttc 540tagagttctc atttattcag
gatacctatt cttactgtat taaaatttgg atatgtgttt 600cattctgtct caaaaatcac
attttattct gagaaggttg gttaaaagat ggcagaaaga 660agatgaaaat aaataagcct
ggtttcaacc ctctaattct tgcctaaaca ttggactgta 720ctttgcattt ttttctttaa
aaatttctat tctaacacaa cttggttgat ttttcctggt 780ctactttatg gttattagac
atactcatgg gtattattag atttcataat ggtcaatgat 840aataggaatt acatggagcc
caacagagaa tatttgctca atacattttt gttaatatat 900ttaggaactt aatggagtct
ctcagtgtct tagtcctagg atgtcttatt taaaatactc 960cctgaaagtt tattctgatg
tttattttag ccatcaaaca ctaaaataat aaattggtga 1020atatgaatct tataaactgt
ggttagctgg tttaaagtga atatatttgc cactagtaga 1080acaaaaatag atgatgaaaa
tgaattaaca tatctacata gttataattc tatcattaga 1140atgagcctta taaataagta
caatatagga cttcaacctt actagactcc taattctaaa 1200ttctactttt ttcatcaaca
gaactttcat tcatttttta aaccctaaaa cttataccca 1260cactattctt acaaaaatat
tcacatgaaa taaaaatttg ctattga 1307311718DNAHomo sapiens
31gagggtgcat aagttctcta gtagggtgat gatataaaaa gccaccggag cactccataa
60ggcacaaact ttcagagaca gcagagcaca caagcttcta ggacaagagc caggaagaaa
120ccaccggaag gaaccatctc actgtgtgta aacatgactt ccaagctggc cgtggctctc
180ttggcagcct tcctgatttc tgcagctctg tgtgaaggtg cagttttgcc aaggagtgct
240aaagaactta gatgtcagtg cataaagaca tactccaaac ctttccaccc caaatttatc
300aaagaactga gagtgattga gagtggacca cactgcgcca acacagaaat tattgtaaag
360ctttctgatg gaagagagct ctgtctggac cccaaggaaa actgggtgca gagggttgtg
420gagaagtttt tgaagagggc tgagaattca taaaaaaatt cattctctgt ggtatccaag
480aatcagtgaa gatgccagtg aaacttcaag caaatctact tcaacacttc atgtattgtg
540tgggtctgtt gtagggttgc cagatgcaat acaagattcc tggttaaatt tgaatttcag
600taaacaatga atagtttttc attgtaccat gaaatatcca gaacatactt atatgtaaag
660tattatttat ttgaatctac aaaaaacaac aaataatttt taaatataag gattttccta
720gatattgcac gggagaatat acaaatagca aaattgaggc caagggccaa gagaatatcc
780gaactttaat ttcaggaatt gaatgggttt gctagaatgt gatatttgaa gcatcacata
840aaaatgatgg gacaataaat tttgccataa agtcaaattt agctggaaat cctggatttt
900tttctgttaa atctggcaac cctagtctgc tagccaggat ccacaagtcc ttgttccact
960gtgccttggt ttctccttta tttctaagtg gaaaaagtat tagccaccat cttacctcac
1020agtgatgttg tgaggacatg tggaagcact ttaagttttt tcatcataac ataaattatt
1080ttcaagtgta acttattaac ctatttatta tttatgtatt tatttaagca tcaaatattt
1140gtgcaagaat ttggaaaaat agaagatgaa tcattgattg aatagttata aagatgttat
1200agtaaattta ttttatttta gatattaaat gatgttttat tagataaatt tcaatcaggg
1260tttttagatt aaacaaacaa acaattgggt acccagttaa attttcattt cagataaaca
1320acaaataatt ttttagtata agtacattat tgtttatctg aaattttaat tgaactaaca
1380atcctagttt gatactccca gtcttgtcat tgccagctgt gttggtagtg ctgtgttgaa
1440ttacggaata atgagttaga actattaaaa cagccaaaac tccacagtca atattagtaa
1500tttcttgctg gttgaaactt gtttattatg tacaaataga ttcttataat attatttaaa
1560tgactgcatt tttaaataca aggctttata tttttaactt taagatgttt ttatgtgctc
1620tccaaatttt ttttactgtt tctgattgta tggaaatata aaagtaaata tgaaacattt
1680aaaatataat ttgttgtcaa agtaaaaaaa aaaaaaaa
1718321691DNAHomo sapiens 32aacttcagtt tgttggctgc ggcagcaggt agcaaagtga
cgccgagggc ctgagtgctc 60cagtagccac cgcatctgga gaaccagcgg ttaccatgga
ggggatcagt atatacactt 120cagataacta caccgaggaa atgggctcag gggactatga
ctccatgaag gaaccctgtt 180tccgtgaaga aaatgctaat ttcaataaaa tcttcctgcc
caccatctac tccatcatct 240tcttaactgg cattgtgggc aatggattgg tcatcctggt
catgggttac cagaagaaac 300tgagaagcat gacggacaag tacaggctgc acctgtcagt
ggccgacctc ctctttgtca 360tcacgcttcc cttctgggca gttgatgccg tggcaaactg
gtactttggg aacttcctat 420gcaaggcagt ccatgtcatc tacacagtca acctctacag
cagtgtcctc atcctggcct 480tcatcagtct ggaccgctac ctggccatcg tccacgccac
caacagtcag aggccaagga 540agctgttggc tgaaaaggtg gtctatgttg gcgtctggat
ccctgccctc ctgctgacta 600ttcccgactt catctttgcc aacgtcagtg aggcagatga
cagatatatc tgtgaccgct 660tctaccccaa tgacttgtgg gtggttgtgt tccagtttca
gcacatcatg gttggcctta 720tcctgcctgg tattgtcatc ctgtcctgct attgcattat
catctccaag ctgtcacact 780ccaagggcca ccagaagcgc aaggccctca agaccacagt
catcctcatc ctggctttct 840tcgcctgttg gctgccttac tacattggga tcagcatcga
ctccttcatc ctcctggaaa 900tcatcaagca agggtgtgag tttgagaaca ctgtgcacaa
gtggatttcc atcaccgagg 960ccctagcttt cttccactgt tgtctgaacc ccatcctcta
tgctttcctt ggagccaaat 1020ttaaaacctc tgcccagcac gcactcacct ctgtgagcag
agggtccagc ctcaagatcc 1080tctccaaagg aaagcgaggt ggacattcat ctgtttccac
tgagtctgag tcttcaagtt 1140ttcactccag ctaacacaga tgtaaaagac ttttttttat
acgataaata actttttttt 1200aagttacaca tttttcagat ataaaagact gaccaatatt
gtacagtttt tattgcttgt 1260tggatttttg tcttgtgttt ctttagtttt tgtgaagttt
aattgactta tttatataaa 1320ttttttttgt ttcatattga tgtgtgtcta ggcaggacct
gtggccaagt tcttagttgc 1380tgtatgtctc gtggtaggac tgtagaaaag ggaactgaac
attccagagc gtgtagtgaa 1440tcacgtaaag ctagaaatga tccccagctg tttatgcata
gataatctct ccattcccgt 1500ggaacgtttt tcctgttctt aagacgtgat tttgctgtag
aagatggcac ttataaccaa 1560agcccaaagt ggtatagaaa tgctggtttt tcagttttca
ggagtgggtt gatttcagca 1620cctacagtgt acagtcttgt attaagttgt taataaaagt
acatgttaaa cttaaaaaaa 1680aaaaaaaaaa a
1691333545DNAHomo sapiens 33gccgcacttt cactctccgt
cagccgcatt gcccgctcgg cgtccggccc ccgacccgcg 60ctcgtccgcc cgcccgcccg
cccgcccgcg ccatgaacgc caaggtcgtg gtcgtgctgg 120tcctcgtgct gaccgcgctc
tgcctcagcg acgggaagcc cgtcagcctg agctacagat 180gcccatgccg attcttcgaa
agccatgttg ccagagccaa cgtcaagcat ctcaaaattc 240tcaacactcc aaactgtgcc
cttcagattg tagcccggct gaagaacaac aacagacaag 300tgtgcattga cccgaagcta
aagtggattc aggagtacct ggagaaagct ttaaacaaga 360ggttcaagat gtgagagggt
cagacgcctg aggaaccctt acagtaggag cccagctctg 420aaaccagtgt tagggaaggg
cctgccacag cctcccctgc cagggcaggg ccccaggcat 480tgccaagggc tttgttttgc
acactttgcc atattttcac catttgatta tgtagcaaaa 540tacatgacat ttatttttca
tttagtttga ttattcagtg tcactggcga cacgtagcag 600cttagactaa ggccattatt
gtacttgcct tattagagtg tctttccacg gagccactcc 660tctgactcag ggctcctggg
ttttgtattc tctgagctgt gcaggtgggg agactgggct 720gagggagcct ggccccatgg
tcagccctag ggtggagagc caccaagagg gacgcctggg 780ggtgccagga ccagtcaacc
tgggcaaagc ctagtgaagg cttctctctg tgggatggga 840tggtggaggg ccacatggga
ggctcacccc cttctccatc cacatgggag ccgggtctgc 900ctcttctggg agggcagcag
ggctaccctg agctgaggca gcagtgtgag gccagggcag 960agtgagaccc agccctcatc
ccgagcacct ccacatcctc cacgttctgc tcatcattct 1020ctgtctcatc catcatcatg
tgtgtccacg actgtctcca tggccccgca aaaggactct 1080caggaccaaa gctttcatgt
aaactgtgca ccaagcagga aatgaaaatg tcttgtgtta 1140cctgaaaaca ctgtgcacat
ctgtgtcttg tttggaatat tgtccattgt ccaatcctat 1200gtttttgttc aaagccagcg
tcctcctctg tgaccaatgt cttgatgcat gcactgttcc 1260ccctgtgcag ccgctgagcg
aggagatgct ccttgggccc tttgagtgca gtcctgatca 1320gagccgtggt cctttggggt
gaactacctt ggttccccca ctgatcacaa aaacatggtg 1380ggtccatggg cagagcccaa
gggaattcgg tgtgcaccag ggttgacccc agaggattgc 1440tgccccatca gtgctccctc
acatgtcagt accttcaaac tagggccaag cccagcactg 1500cttgaggaaa acaagcattc
acaacttgtt tttggttttt aaaacccagt ccacaaaata 1560accaatcctg gacatgaaga
ttctttccca attcacatct aacctcatct tcttcaccat 1620ttggcaatgc catcatctcc
tgccttcctc ctgggccctc tctgctctgc gtgtcacctg 1680tgcttcgggc ccttcccaca
ggacatttct ctaagagaac aatgtgctat gtgaagagta 1740agtcaacctg cctgacattt
ggagtgttcc ccttccactg agggcagtcg atagagctgt 1800attaagccac ttaaaatgtt
cacttttgac aaaggcaagc acttgtgggt ttttgttttg 1860tttttcattc agtcttacga
atacttttgc cctttgatta aagactccag ttaaaaaaaa 1920ttttaatgaa gaaagtggaa
aacaaggaag tcaaagcaag gaaactatgt aacatgtagg 1980aagtaggaag taaattatag
tgatgtaatc ttgaattgta actgttcttg aatttaataa 2040tctgtagggt aattagtaac
atgtgttaag tattttcata agtatttcaa attggagctt 2100catggcagaa ggcaaaccca
tcaacaaaaa ttgtccctta aacaaaaatt aaaatcctca 2160atccagctat gttatattga
aaaaatagag cctgagggat ctttactagt tataaagata 2220cagaactctt tcaaaacctt
ttgaaattaa cctctcacta taccagtata attgagtttt 2280cagtggggca gtcattatcc
aggtaatcca agatatttta aaatctgtca cgtagaactt 2340ggatgtacct gcccccaatc
catgaaccaa gaccattgaa ttcttggttg aggaaacaaa 2400catgacccta aatcttgact
acagtcagga aaggaatcat ttctatttct cctccatggg 2460agaaaataga taagagtaga
aactgcaggg aaaattattt gcataacaat tcctctacta 2520acaatcagct ccttcctgga
gactgcccag ctaaagcaat atgcatttaa atacagtctt 2580ccatttgcaa gggaaaagtc
tcttgtaatc cgaatctctt tttgctttcg aactgctagt 2640caagtgcgtc cacgagctgt
ttactaggga tccctcatct gtccctccgg gacctggtgc 2700tgcctctacc tgacactccc
ttgggctccc tgtaacctct tcagaggccc tcgctgccag 2760ctctgtatca ggacccagag
gaaggggcca gaggctcgtt gactggctgt gtgttgggat 2820tgagtctgtg ccacgtgttt
gtgctgtggt gtgtccccct ctgtccaggc actgagatac 2880cagcgaggag gctccagagg
gcactctgct tgttattaga gattacctcc tgagaaaaaa 2940ggttccgctt ggagcagagg
ggctgaatag cagaaggttg cacctccccc aaccttagat 3000gttctaagtc tttccattgg
atctcattgg acccttccat ggtgtgatcg tctgactggt 3060gttatcaccg tgggctccct
gactgggagt tgatcgcctt tcccaggtgc tacacccttt 3120tccagctgga tgagaatttg
agtgctctga tccctctaca gagcttccct gactcattct 3180gaaggagccc cattcctggg
aaatattccc tagaaacttc caaatcccct aagcagacca 3240ctgataaaac catgtagaaa
atttgttatt ttgcaacctc gctggactct cagtctctga 3300gcagtgaatg attcagtgtt
aaatgtgatg aatactgtat tttgtattgt ttcaattgca 3360tctcccagat aatgtgaaaa
tggtccagga gaaggccaat tcctatacgc agcgtgcttt 3420aaaaaataaa taagaaacaa
ctctttgaga aacaacaatt tctactttga agtcatacca 3480atgaaaaaat gtatatgcac
ttataatttt cctaataaag ttctgtactc aaatgtagcc 3540accaa
3545342896DNAHomo sapiens
34ccactctaag gaatgcggtc cctttgacag gcgaaaaact gaagttggaa aagacaaagt
60gatttgttca aaattgaaat ttgaaacttg acatttggtc agtgggccct atgtaggaaa
120aaacctccaa gagagctagg gttcctctca gagaggaaag acaggtcctt aggtcctcac
180cctcccgtct ccttgccctt gcagttctgg gaactggaca gattggacaa ctataacgac
240acctccctgg tggaaaatca tctctgccct gccacagagg ggcccctcat ggcctccttc
300aaggccgtgt tcgtgcccgt ggcctacagc ctcatcttcc tcctgggcgt gatcggcaac
360gtcctggtgc tggtgatcct ggagcggcac cggcagacac gcagttccac ggagaccttc
420ctgttccacc tggccgtggc cgacctcctg ctggtcttca tcttgccctt tgccgtggcc
480gagggctctg tgggctgggt cctggggacc ttcctctgca aaactgtgat tgccctgcac
540aaagtcaact tctactgcag cagcctgctc ctggcctgca tcgccgtgga ccgctacctg
600gccattgtcc acgccgtcca tgcctaccgc caccgccgcc tcctctccat ccacatcacc
660tgtgggacca tctggctggt gggcttcctc cttgccttgc cagagattct cttcgccaaa
720gtcagccaag gccatcacaa caactccctg ccacgttgca ccttctccca agagaaccaa
780gcagaaacgc atgcctggtt cacctcccga ttcctctacc atgtggcggg attcctgctg
840cccatgctgg tgatgggctg gtgctacgtg ggggtagtgc acaggttgcg ccaggcccag
900cggcgccctc agcggcagaa ggcagtcagg gtggccatcc tggtgacaag catcttcttc
960ctctgctggt caccctacca catcgtcatc ttcctggaca ccctggcgag gctgaaggcc
1020gtggacaata cctgcaagct gaatggctct ctccccgtgg ccatcaccat gtgtgagttc
1080ctgggcctgg cccactgctg cctcaacccc atgctctaca ctttcgccgg cgtgaagttc
1140cgcagtgacc tgtcgcggct cctgacgaag ctgggctgta ccggccctgc ctccctgtgc
1200cagctcttcc ctagctggcg caggagcagt ctctctgagt cagagaatgc cacctctctc
1260accacgttct aggtcccagt gtcccctttt attgctgctt ttccttgggg caggcagtga
1320tgctggatgc tccttccaac aggagctggg atcctaaggg ctcaccgtgg ctaagagtgt
1380cctaggagta tcctcatttg gggtagctag aggaaccaac ccccatttct agaacatccc
1440tgccagctct tctgccggcc ctggggctag gctggagccc agggagcgga aagcagctca
1500aaggcacagt gaaggctgtc cttacccatc tgcacccccc tgggctgaga gaacctcacg
1560cacctcccat cctaatcatc caatgctcaa gaaacaactt ctacttctgc ccttgccaac
1620ggagagcgcc tgcccctccc agaacacact ccatcagctt aggggctgct gacctccaca
1680gcttcccctc tctcctcctg cccacctgtc aaacaaagcc agaagctgag caccagggga
1740tgagtggagg ttaaggctga ggaaaggcca gctggcagca gagtgtggcc ttcggacaac
1800tcagtcccta aaaacacaga cattctgcca ggcccccaag cctgcagtca tcttgaccaa
1860gcaggaagct cagactggtt gagttcaggt agctgcccct ggctctgacc gaaacagcgc
1920tgggtccacc ccatgtcacc ggatcctggg tggtctgcag gcagggctga ctctaggtgc
1980ccttggaggc cagccagtga cctgaggaag cgtgaaggcc gagaagcaag aaagaaaccc
2040gacagaggga agaaaagagc tttcttcccg aaccccaagg agggagatgg atcaatcaaa
2100cccggcggtc ccctccgcca ggcgagatgg ggtggggtgg agaactccta gggtggctgg
2160gtccagggga tgggaggttg tgggcattga tggggaagga ggctggcttg tcccctcctc
2220actcccttcc cataagctat agacccgagg aaactcagag tcggaacgga gaaaggtgga
2280ctggaagggg cccgtgggag tcatctcaac catcccctcc gtggcatcac cttaggcagg
2340gaagtgtaag aaacacactg aggcagggaa gtccccaggc cccaggaagc cgtgccctgc
2400ccccgtgagg atgtcactca gatggaaccg caggaagctg ctccgtgctt gtttgctcac
2460ctggggtgtg ggaggcccgt ccggcagttc tgggtgctcc ctaccacctc cccagccttt
2520gatcaggtgg ggagtcaggg acccctgccc ttgtcccact caagccaagc agccaagctc
2580cttgggaggc cccactgggg aaataacagc tgtggctcac gtgagagtgt cttcacggca
2640ggacaacgag gaagccctaa gacgtccctt ttttctctga gtatctcctc gcaagctggg
2700taatcgatgg gggagtctga agcagatgca aagaggcaag aggctggatt ttgaattttc
2760tttttaataa aaaggcacct ataaaacagg tcaatacagt acaggcagca cagagacccc
2820cggaacaagc ctaaaaattg tttcaaaata aaaaccaaga agatgtcttc acatattgta
2880aaaaaaaaaa aaaaaa
2896352919DNAHomo sapiens 35aaaaaaaaaa agtgatgagt tgtgaggcag gtcgcggccc
tactgcctca ggagacgatg 60cgcagctcat ttgcttaaat ttgcagctga cggctgccac
ctctctagag gcacctggcg 120gggagcctct caacataaga cagtgaccag tctggtgact
cacagccggc acagccatga 180actacccgct aacgctggaa atggacctcg agaacctgga
ggacctgttc tgggaactgg 240acagattgga caactataac gacacctccc tggtggaaaa
tcatctctgc cctgccacag 300aggggcccct catggcctcc ttcaaggccg tgttcgtgcc
cgtggcctac agcctcatct 360tcctcctggg cgtgatcggc aacgtcctgg tgctggtgat
cctggagcgg caccggcaga 420cacgcagttc cacggagacc ttcctgttcc acctggccgt
ggccgacctc ctgctggtct 480tcatcttgcc ctttgccgtg gccgagggct ctgtgggctg
ggtcctgggg accttcctct 540gcaaaactgt gattgccctg cacaaagtca acttctactg
cagcagcctg ctcctggcct 600gcatcgccgt ggaccgctac ctggccattg tccacgccgt
ccatgcctac cgccaccgcc 660gcctcctctc catccacatc acctgtggga ccatctggct
ggtgggcttc ctccttgcct 720tgccagagat tctcttcgcc aaagtcagcc aaggccatca
caacaactcc ctgccacgtt 780gcaccttctc ccaagagaac caagcagaaa cgcatgcctg
gttcacctcc cgattcctct 840accatgtggc gggattcctg ctgcccatgc tggtgatggg
ctggtgctac gtgggggtag 900tgcacaggtt gcgccaggcc cagcggcgcc ctcagcggca
gaaggcagtc agggtggcca 960tcctggtgac aagcatcttc ttcctctgct ggtcacccta
ccacatcgtc atcttcctgg 1020acaccctggc gaggctgaag gccgtggaca atacctgcaa
gctgaatggc tctctccccg 1080tggccatcac catgtgtgag ttcctgggcc tggcccactg
ctgcctcaac cccatgctct 1140acactttcgc cggcgtgaag ttccgcagtg acctgtcgcg
gctcctgacg aagctgggct 1200gtaccggccc tgcctccctg tgccagctct tccctagctg
gcgcaggagc agtctctctg 1260agtcagagaa tgccacctct ctcaccacgt tctaggtccc
agtgtcccct tttattgctg 1320cttttccttg gggcaggcag tgatgctgga tgctccttcc
aacaggagct gggatcctaa 1380gggctcaccg tggctaagag tgtcctagga gtatcctcat
ttggggtagc tagaggaacc 1440aacccccatt tctagaacat ccctgccagc tcttctgccg
gccctggggc taggctggag 1500cccagggagc ggaaagcagc tcaaaggcac agtgaaggct
gtccttaccc atctgcaccc 1560ccctgggctg agagaacctc acgcacctcc catcctaatc
atccaatgct caagaaacaa 1620cttctacttc tgcccttgcc aacggagagc gcctgcccct
cccagaacac actccatcag 1680cttaggggct gctgacctcc acagcttccc ctctctcctc
ctgcccacct gtcaaacaaa 1740gccagaagct gagcaccagg ggatgagtgg aggttaaggc
tgaggaaagg ccagctggca 1800gcagagtgtg gccttcggac aactcagtcc ctaaaaacac
agacattctg ccaggccccc 1860aagcctgcag tcatcttgac caagcaggaa gctcagactg
gttgagttca ggtagctgcc 1920cctggctctg accgaaacag cgctgggtcc accccatgtc
accggatcct gggtggtctg 1980caggcagggc tgactctagg tgcccttgga ggccagccag
tgacctgagg aagcgtgaag 2040gccgagaagc aagaaagaaa cccgacagag ggaagaaaag
agctttcttc ccgaacccca 2100aggagggaga tggatcaatc aaacccggcg gtcccctccg
ccaggcgaga tggggtgggg 2160tggagaactc ctagggtggc tgggtccagg ggatgggagg
ttgtgggcat tgatggggaa 2220ggaggctggc ttgtcccctc ctcactccct tcccataagc
tatagacccg aggaaactca 2280gagtcggaac ggagaaaggt ggactggaag gggcccgtgg
gagtcatctc aaccatcccc 2340tccgtggcat caccttaggc agggaagtgt aagaaacaca
ctgaggcagg gaagtcccca 2400ggccccagga agccgtgccc tgcccccgtg aggatgtcac
tcagatggaa ccgcaggaag 2460ctgctccgtg cttgtttgct cacctggggt gtgggaggcc
cgtccggcag ttctgggtgc 2520tccctaccac ctccccagcc tttgatcagg tggggagtca
gggacccctg cccttgtccc 2580actcaagcca agcagccaag ctccttggga ggccccactg
gggaaataac agctgtggct 2640cacgtgagag tgtcttcacg gcaggacaac gaggaagccc
taagacgtcc cttttttctc 2700tgagtatctc ctcgcaagct gggtaatcga tgggggagtc
tgaagcagat gcaaagaggc 2760aagaggctgg attttgaatt ttctttttaa taaaaaggca
cctataaaac aggtcaatac 2820agtacaggca gcacagagac ccccggaaca agcctaaaaa
ttgtttcaaa ataaaaacca 2880agaagatgtc ttcacatatt gtaaaaaaaa aaaaaaaaa
2919361219DNAHomo sapiens 36gagaagatgt ttgaaaaaac
tgactctgct aatgagcctg gactcagagc tcaagtctga 60actctacctc cagacagaat
gaagttcatc tcgacatctc tgcttctcat gctgctggtc 120agcagcctct ctccagtcca
aggtgttctg gaggtctatt acacaagctt gaggtgtaga 180tgtgtccaag agagctcagt
ctttatccct agacgcttca ttgatcgaat tcaaatcttg 240ccccgtggga atggttgtcc
aagaaaagaa atcatagtct ggaagaagaa caagtcaatt 300gtgtgtgtgg accctcaagc
tgaatggata caaagaatga tggaagtatt gagaaaaaga 360agttcttcaa ctctaccagt
tccagtgttt aagagaaaga ttccctgatg ctgatatttc 420cactaagaac acctgcattc
ttcccttatc cctgctctgg attttagttt tgtgcttagt 480taaatctttt ccaggaaaaa
gaacttcccc atacaaataa gcatgagact atgtaaaaat 540aaccttgcag aagctgatgg
ggcaaactca agcttcttca ctcacagcac cctatataca 600cttggagttt gcattcttat
tcatcaggga ggaaagtttc tttgaaaata gttattcagt 660tataagtaat acaggattat
tttgattata tacttgttgt ttaatgttta aaatttctta 720gaaaacaatg gaatgagaat
ttaagcctca aatttgaaca tgtggcttga attaagaaga 780aaattatggc atatattaaa
agcaggcttc tatgaaagac tcaaaaagct gcctgggagg 840cagatggaac ttgagcctgt
caagaggcaa aggaatccat gtagtagata tcctctgctt 900aaaaactcac tacggaggag
aattaagtcc tacttttaaa gaatttcttt ataaaattta 960ctgtctaaga ttaatagcat
tcgaagatcc ccagacttca tagaatactc agggaaagca 1020tttaaagggt gatgtacaca
tgtatccttt cacacatttg ccttgacaaa cttctttcac 1080tcacatcttt ttcactgact
ttttttgtgg ggggcggggc cggggggact ctggtatcta 1140attctttaat gattcctata
aatctaatga cattcaataa agttgagcaa acattttact 1200taaaaaaaaa aaaaaaaaa
1219371953DNAHomo sapiens
37gcagaccttg cttcatgagc aagctcatct ctggaacaaa ctggcaaagc atctctgctg
60gtgttcatca gaacagacac catggcagag catgattacc atgaagacta tgggttcagc
120agtttcaatg acagcagcca ggaggagcat caagacttcc tgcagttcag caaggtcttt
180ctgccctgca tgtacctggt ggtgtttgtc tgtggtctgg tggggaactc tctggtgctg
240gtcatatcca tcttctacca taagttgcag agcctgacgg atgtgttcct ggtgaaccta
300cccctggctg acctggtgtt tgtctgcact ctgcccttct gggcctatgc aggcatccat
360gaatgggtgt ttggccaggt catgtgcaag agcctactgg gcatctacac tattaacttc
420tacacgtcca tgctcatcct cacctgcatc actgtggatc gtttcattgt agtggttaag
480gccaccaagg cctacaacca gcaagccaag aggatgacct ggggcaaggt caccagcttg
540ctcatctggg tgatatccct gctggtttcc ttgccccaaa ttatctatgg caatgtcttt
600aatctcgaca agctcatatg tggttaccat gacgaggcaa tttccactgt ggttcttgcc
660acccagatga cactggggtt cttcttgcca ctgctcacca tgattgtctg ctattcagtc
720ataatcaaaa cactgcttca tgctggaggc ttccagaagc acagatctct aaagatcatc
780ttcctggtga tggctgtgtt cctgctgacc cagatgccct tcaacctcat gaagttcatc
840cgcagcacac actgggaata ctatgccatg accagctttc actacaccat catggtgaca
900gaggccatcg catacctgag ggcctgcctt aaccctgtgc tctatgcctt tgtcagcctg
960aagtttcgaa agaacttctg gaaacttgtg aaggacattg gttgcctccc ttaccttggg
1020gtctcacatc aatggaaatc ttctgaggac aattccaaga ctttttctgc ctcccacaat
1080gtggaggcca ccagcatgtt ccagttatag gccttgccag ggtttcgaga agctgctctg
1140gaatttgcaa gtcatggctg tgccctcttg atgtggtgag gcaggctttg tttatagctt
1200gcgcattctc atggagaagt tatcagacac tctggctggt ttggaatgct tcttctcagg
1260catgaacatg tactgttctc ttcttgaaca ctcatgctga aagcccaagt agggggtcta
1320aaatttttaa ggactttcct tcctccatct ccaagaatgc tgaaaccaag ggggatgaca
1380tgtgactcct atgatctcag gttctccttg attgggactg gggctgaagg ttgaagaggt
1440gagcacggcc aacaaagctg ttgatggtag gtggcacact gggtgcccaa gctcagaagg
1500ctcttctgac tactgggcaa agagtgtaga tcagagcagc agtgaaaaca agtgctggca
1560ccaccaggca cctcacagaa atgagatcag gctctgcctc accttggggc ttgacttttg
1620tataggtaga tgttcagatt gctttgatta atccagaata actagcacca gggactatga
1680atgggcaaaa ctgaattata agaggctgat aattccagtg gtccatggaa tgcttgaaaa
1740atgtgcaaaa cagcgtttaa gactgtaatg aatctaagca gcatttctga agtggactct
1800ttggtggctt tgcattttaa aaatgaaatt ttccaatgtc tgccacacaa acgtatgtaa
1860atgtatatac ccacacacat acacacatat gtcatatatt actagcatat gagtttcata
1920gctaagaaat aaaactgtta aagtctccaa act
1953382344DNAHomo sapiens 38ggtgcgtccg cgggtggctg ccccgcaggt gcgcgcggcc
ggggctggcg gcgactctct 60ccaccgggcc gcccgggagg ctcatgcagc gcggctgggt
cccgcggcgc ccggatcggg 120gaagtgaaag tgcctcggag gaggagggcc ggtccggcag
tgcagccgcc tcacaggtcg 180gcggacgggc caggcgggcg gcctcctgaa ccgaaccgaa
tcggctcctc gggccgtcgt 240cctcccgccc ctcctcgccc gccgccggag ttttctttcg
gtttcttcca agattcctgg 300ccttccctcg acggagccgg gcccagtgcg ggggcgcagg
gcgcgggagc tccacctcct 360cggctttccc tgcgtccaga ggctggcatg gcgcgggccg
agtactgagc gcacggtcgg 420ggcacagcag ggccgggggg tgcagctggc tcgcgcctcc
tctccggccg ccgtctcctc 480cggtccccgg cgaaagccat tgagacacca gctggacgtc
acgcgccgga gcatgtctgg 540gagtcagagc gaggtggctc catccccgca gagtccgcgg
agccccgaga tgggacggga 600cttgcggccc gggtcccgcg tgctcctgct cctgcttctg
ctcctgctgg tgtacctgac 660tcagccaggc aatggcaacg agggcagcgt cactggaagt
tgttattgtg gtaaaagaat 720ttcttccgac tccccgccat cggttcagtt catgaatcgt
ctccggaaac acctgagagc 780ttaccatcgg tgtctatact acacgaggtt ccagctcctt
tcctggagcg tgtgtggggg 840caacaaggac ccatgggttc aggaattgat gagctgtctt
gatctcaaag aatgtggaca 900tgcttactcg gggattgtgg cccaccagaa gcatttactt
cctaccagcc ccccaatttc 960tcaggcctca gagggggcat cttcagatat ccacacccct
gcccagatgc tcctgtccac 1020cttgcagtcc actcagcgcc ccaccctccc agtaggatca
ctgtcctcgg acaaagagct 1080cactcgtccc aatgaaacca ccattcacac tgcgggccac
agtctggcag ctgggcctga 1140ggctggggag aaccagaagc agccggaaaa aaatgctggt
cccacagcca ggacatcagc 1200cacagtgcca gtcctgtgcc tcctggccat catcttcatc
ctcaccgcag ccctttccta 1260tgtgctgtgc aagaggagga gggggcagtc accgcagtcc
tctccagatc tgccggttca 1320ttatatacct gtggcacctg actctaatac ctgagccaag
aatggaagct tgtgaggaga 1380cggactctat gttgcccagg ctgttatgga actcctgagt
caagtgatcc tcccaccttg 1440gcctctgaag gtgcgaggat tataggcgtc acctaccaca
tccagcctac acgtatttgt 1500taatatctaa cataggacta accagccact gccctctctt
aggcccctca tttaaaaacg 1560gttatactat aaaatctgct tttcacactg ggtgataata
acttggacaa attctatgtg 1620tattttgttt tgttttgctt tgctttgttt tgagacggag
tctcgctctg tcatccaggc 1680tggagtgcag tggcatgatc tcggctcact gcaaccccca
tctcccaggt tcaagcgatt 1740ctcctgcctc ctcctgagta gctgggacta caggtgctca
ccaccacacc cggctaattt 1800tttgtatttt tagtagagac ggggtttcac catgttgacc
aggctggtct cgaactcctg 1860acctggtgat ctgcccaccc aggcctccca aagtgctggg
attaaaggtg tgagccacca 1920tgcctggccc tatgtgtgtt ttttaactac taaaaattat
ttttgtaatg attgagtctt 1980ctttatggaa acaactggcc tcagcccttg cgcccttact
gtgattcctg gcttcatttt 2040ttgctgatgg ttccccctcg tcccaaatct ctctcccagt
acaccagttg ttcctccccc 2100acctcagccc tctcctgcat cctcctgtac ccgcaacgaa
ggcctgggct ttcccaccct 2160ccctccttag caggtgccgt gctgggacac catacgggtt
ggtttcacct cctcagtccc 2220ttgcctaccc cagtgagagt ctgatcttgt ttttattgtt
attgctttta ttattattgc 2280ttttattatc attaaaactc tagttcttgt tttgtctctc
cgaaaaaaaa aaaaaaaaaa 2340aaaa
2344391497DNAHomo sapiens 39cggaccacca gcaacagaca
acatcttcat tcggctctcc ctgaagctgt actgcctcgc 60tgagaggatg aaggtctccg
aggctgccct gtctctcctt gtcctcatcc ttatcattac 120ttcggcttct cgcagccagc
caaaagttcc tgagtgggtg aacaccccat ccacctgctg 180cctgaagtat tatgagaaag
tgttgccaag gagactagtg gtgggataca gaaaggccct 240caactgtcac ctgccagcaa
tcatcttcgt caccaagagg aaccgagaag tctgcaccaa 300ccccaatgac gactgggtcc
aagagtacat caaggatccc aacctacctt tgctgcctac 360caggaacttg tccacggtta
aaattattac agcaaagaat ggtcaacccc agctcctcaa 420ctcccagtga tgaccaggct
ttagtggaag cccttgttta cagaagagag gggtaaacct 480atgaaaacag gggaagcctt
attaggctga aactagccag tcacattgag agaagcagaa 540caatgatcaa aataaaggag
aagtatttcg aatattttct caatcttagg aggaaatacc 600aaagttaagg gacgtgggca
gaggtacgct cttttatttt tatatttata tttttatttt 660tttgagatag ggtcttactc
tgtcacccag gctggagtgc agtggtgtga tcttggctca 720cttgatcttg gctcactgta
acctccacct cccaggctca agtgatcctc ccaccccagc 780ctcctgagta gctgggacta
caggcttgcg ccaccacacc tggctaattt ttgtattttt 840ggtagagacg ggattctacc
atgttgccca ggctggtctc aaactcgtgt gcccaagcaa 900tccacctgcc tcagccttcc
aaaagtgctg ggattacagg cgtgagccac cacatccggc 960cagtgcactc ttaatacaca
gaaaaatata tttcacatcc ttctcctgct ctctttcaat 1020tcctcacttc acaccagtac
acaagccatt ctaaatactt agccagtttc cagccttcca 1080gatgatcttt gccctctggg
tcttgaccca ttaagagccc catagaactc ttgatttttc 1140ctgtccatct ttatggattt
ttctggatct atattttctt caattattct ttcattttat 1200aatgcaactt tttcatagga
agtccggatg ggaatattca cattaatcat ttttgcagag 1260actttgctag atcctctcat
attttgtctt cctcagggtg gcaggggtac agagagtgcc 1320tgattggaaa aaaaaaaaaa
agagagagag agagaagaag aagaagaaga gacacaaatc 1380tctacctccc atgttaagct
ttgcaggaca gggaaagaaa gggtatgaga cacggctagg 1440ggtaaactct tagtccaaaa
cccaagcatg caataaataa aactccctta tttgaca 1497401002DNAHomo sapiens
40agatgggaca gcttggccta cagcccggcg ggcatcagct cccttgaccc agtggatatc
60ggtggccccg ttattcgtcc aggtgcccag ggaggaggac ccgcctgcag catgaacctg
120tggctcctgg cctgcctggt ggccggcttc ctgggagcct gggcccccgc tgtccacacc
180caaggtgtct ttgaggactg ctgcctggcc taccactacc ccattgggtg ggctgtgctc
240cggcgcgcct ggacttaccg gatccaggag gtgagcggga gctgcaatct gcctgctgcg
300atattctacc tccccaagag acacaggaag gtgtgtggga accccaaaag cagggaggtg
360cagagagcca tgaagctcct ggatgctcga aataaggttt ttgcaaagct ccaccacaac
420acgcagacct tccaagcagg ccctcatgct gtaaagaagt tgagttctgg aaactccaag
480ttatcatcgt ccaagtttag caatcccatc agcagcagta agaggaatgt ctccctcctg
540atatcagcta attcaggact gtgagccggc tcatttctgg gctccatcgg cacaggaggg
600gccggatctt tctccgataa aaccgtcgcc ctacagaccc agctgtcccc acgcctctgt
660cttttgggtc aagtcttaat ccctgcacct gagttggtcc tccctctgca cccccaccac
720ctcctgcccg tctggcaact ggaaagaggg agttggcctg attttaagcc ttttgccgct
780ccggggacca gcagcaatcc tgggcagcca gtggctcttg tagagaagac ttaggatacc
840tctctcactt tctgtttctt gccgtccacc ccgggccatg ccagtgtgtc cctctgggtc
900cctccaaaac tctggtcagt tcaaggatgc ccctcccagg ctatgctttt ctataacttt
960taaataaacc ttggggggtg atggagtcat tcctgcctgt ta
1002411002DNAHomo sapiens 41agatgggaca gcttggccta cagcccggcg ggcatcagct
cccttgaccc agtggatatc 60ggtggccccg ttattcgtcc aggtgcccag ggaggaggac
ccgcctgcag catgaacctg 120tggctcctgg cctgcctggt ggccggcttc ctgggagcct
gggcccccgc tgtccacacc 180caaggtgtct ttgaggactg ctgcctggcc taccactacc
ccattgggtg ggctgtgctc 240cggcgcgcct ggacttaccg gatccaggag gtgagcggga
gctgcaatct gcctgctgcg 300atattctacc tccccaagag acacaggaag gtgtgtggga
accccaaaag cagggaggtg 360cagagagcca tgaagctcct ggatgctcga aataaggttt
ttgcaaagct ccaccacaac 420acgcagacct tccaagcagg ccctcatgct gtaaagaagt
tgagttctgg aaactccaag 480ttatcatcgt ccaagtttag caatcccatc agcagcagta
agaggaatgt ctccctcctg 540atatcagcta attcaggact gtgagccggc tcatttctgg
gctccatcgg cacaggaggg 600gccggatctt tctccgataa aaccgtcgcc ctacagaccc
agctgtcccc acgcctctgt 660cttttgggtc aagtcttaat ccctgcacct gagttggtcc
tccctctgca cccccaccac 720ctcctgcccg tctggcaact ggaaagaggg agttggcctg
attttaagcc ttttgccgct 780ccggggacca gcagcaatcc tgggcagcca gtggctcttg
tagagaagac ttaggatacc 840tctctcactt tctgtttctt gccgtccacc ccgggccatg
ccagtgtgtc cctctgggtc 900cctccaaaac tctggtcagt tcaaggatgc ccctcccagg
ctatgctttt ctataacttt 960taaataaacc ttggggggtg atggagtcat tcctgcctgt
ta 100242744DNAHomo sapiens 42atgaacctgt ggctcctggc
ctgcctggtg gccggcttcc tgggagcctg ggcccccgct 60gtccacaccc aaggtgtctt
tgaggactgc tgcctggcct accactaccc cattgggtgg 120gctgtgctcc ggcgcgcctg
gacttaccgg atccaggagg tgagcgggag ctgcaatctg 180cctgctgcga tcaggccctc
atgctgtaaa gaagttgagt tctggaaact ccaagttatc 240atcgtccaag tttagcaatc
ccatcagcag cagtaagagg aatgtctccc tcctgatatc 300agctaattca ggactgtgag
ccggctcatt tctgggctcc atcggcacag gaggggccgg 360atctttctcc gataaaaccg
tcgccctaca gacccagctg tccccacgcc tctgtctttt 420gggtcaagtc ttaatccctg
cacctgagtt ggtcctccct ctgcaccccc accacctcct 480gcccgtctgg caactggaaa
gagggagttg gcctgatttt aagccttttg ccgctccggg 540gaccagcagc aatcctgggc
agccagtggc tcttgtagag aagacttagg atacctctct 600cactttctgt ttcttgccgt
ccaccccggg ccatgccagt gtgtccctct gggtccctcc 660aaaactctgg tcagttcaag
gatgcccctc ccaggctatg cttttctata acttttaaat 720aaaccttggg gggtgatgga
gtca 744433108DNAHomo sapiens
43gaaatactcg tctctggtaa agtctgagca ggacagggtg gctgactggc agatccagag
60gttcccttgg cagtccacgc caggccttca ccatggatca gttccctgaa tcagtgacag
120aaaactttga gtacgatgat ttggctgagg cctgttatat tggggacatc gtggtctttg
180ggactgtgtt cctgtccata ttctactccg tcatctttgc cattggcctg gtgggaaatt
240tgttggtagt gtttgccctc accaacagca agaagcccaa gagtgtcacc gacatttacc
300tcctgaacct ggccttgtct gatctgctgt ttgtagccac tttgcccttc tggactcact
360atttgataaa tgaaaagggc ctccacaatg ccatgtgcaa attcactacc gccttcttct
420tcatcggctt ttttggaagc atattcttca tcaccgtcat cagcattgat aggtacctgg
480ccatcgtcct ggccgccaac tccatgaaca accggaccgt gcagcatggc gtcaccatca
540gcctaggcgt ctgggcagca gccattttgg tggcagcacc ccagttcatg ttcacaaagc
600agaaagaaaa tgaatgcctt ggtgactacc ccgaggtcct ccaggaaatc tggcccgtgc
660tccgcaatgt ggaaacaaat tttcttggct tcctactccc cctgctcatt atgagttatt
720gctacttcag aatcatccag acgctgtttt cctgcaagaa ccacaagaaa gccaaagcca
780ttaaactgat ccttctggtg gtcatcgtgt ttttcctctt ctggacaccc tacaacgtta
840tgattttcct ggagacgctt aagctctatg acttctttcc cagttgtgac atgaggaagg
900atctgaggct ggccctcagt gtgactgaga cggttgcatt tagccattgt tgcctgaatc
960ctctcatcta tgcatttgct ggggagaagt tcagaagata cctttaccac ctgtatggga
1020aatgcctggc tgtcctgtgt gggcgctcag tccacgttga tttctcctca tctgaatcac
1080aaaggagcag gcatggaagt gttctgagca gcaattttac ttaccacacg agtgatggag
1140atgcattgct ccttctctga agggaatccc aaagccttgt gtctacagag aacctggagt
1200tcctgaacct gatgctgact agtgaggaaa gatttttgtt gttatttctt acaggcacaa
1260aatgatggac ccaatgcaca caaaacaacc ctagagtgtt gttgagaatt gtgctcaaaa
1320tttgaagaat gaacaaattg aactctttga atgacaaaga gtagacattt ctcttactgc
1380aaatgtcatc agaacttttt ggtttgcaga tgacaaaaat tcaactcaga ctagtttagt
1440taaatgaggg tggtgaatat tgttcatatt gtggcacaag caaaagggtg tctgagccct
1500caaagtgagg ggaaaccagg gcctgagcca agctagaatt ccctctctct gactctcaaa
1560tcttttagtc attatagatc ccccagactt tacatgacac agctttatca ccagagaggg
1620actgacaccc atgtttctct ggccccaagg gcaaaattcc cagggaagtg ctctgatagg
1680ccaagtttgt atcaggtgcc catccctgga aggtgctgtt atccatgggg aagggatata
1740taagatggaa gcttccagtc caatctcatg gagaagcaga aatacatatt tccaagaagt
1800tggatgggtg ggtactattc tgattacaca aaacaaatgc cacacatcac ccttaccatg
1860tgcctgatcc agcctctccc ctgattacac cagcctcgtc ttcattaagc cctcttccat
1920catgtcccca aacctgcaag ggctccccac tgcctactgc atcgagtcaa aactcaaatg
1980cttggcttct catacgtcca ccatggggtc ctaccaatag attccccatt gcctcctcct
2040tcccaaagga ctccacccat cctatcagcc tgtctcttcc atatgacctc atgcatctcc
2100acctgctccc aggccagtaa gggaaataga aaaaccctgc ccccaaataa gaagggatgg
2160attccaaccc caactccagt agcttgggac aaatcaagct tcagtttcct ggtctgtaga
2220agagggataa ggtacctttc acatagagat catcctttcc agcatgagga actagccacc
2280aactcttgca ggtctcaacc cttttgtctg cctcttagac ttctgctttc cacacctggc
2340actgctgtgc tgtgcccaag ttgtggtgct gacaaagctt ggaagagcct gcaggtgctg
2400ctgcgtggca tagcccagac acagaagagg ctggttctta cgatggcacc cagtgagcac
2460tcccaagtct acagagtgat agccttccgt aacccaactc tcctggactg ccttgaatat
2520cccctcccag tcaccttgtg gcaagcccct gcccatctgg gaaaataccc catcattcat
2580gctactgcca acctggggag ccagggctat gggagcagct tttttttccc ccctagaaac
2640gtttggaaca atctaaaagt ttaaagctcg aaaacaattg taataatgct aaagaaaaag
2700tcatccaatc taaccacatc aatattgtca ttcctgtatt cacccgtcca gaccttgttc
2760acactctcac atgtttagag ttgcaatcgt aatgtacaga tggttttata atctgatttg
2820ttttcctctt aacgttagac cacaaatagt gctcgctttc tatgtagttt ggtaattatc
2880attttagaag actctaccag actgtgtatt cattgaagtc agatgtggta actgttaaat
2940tgctgtgtat ctgatagctc tttggcagtc tatatgtttg tataatgaat gagagaataa
3000gtcatgttcc ttcaagatca tgtaccccaa tttacttgcc attactcaat tgataaacat
3060ttaacttgtt tccaatgttt agcaaataca tattttatag aacttcca
3108443304DNAHomo sapiens 44ctgagctctg ccgcctggct ctagccgcct gcctggcccc
cgccgggact cttgcccacc 60ctcagccatg gctccgatat ctctgtcgtg gctgctccgc
ttggccacct tctgccatct 120gactgtcctg ctggctggac agcaccacgg tgtgacgaaa
tgcaacatca cgtgcagcaa 180gatgacatca aagatacctg tagctttgct catccactat
caacagaacc aggcatcatg 240cggcaaacgc gcaatcatct tggagacgag acagcacagg
ctgttctgtg ccgacccgaa 300ggagcaatgg gtcaaggacg cgatgcagca tctggaccgc
caggctgctg ccctaactcg 360aaatggcggc accttcgaga agcagatcgg cgaggtgaag
cccaggacca cccctgccgc 420cgggggaatg gacgagtctg tggtcctgga gcccgaagcc
acaggcgaaa gcagtagcct 480ggagccgact ccttcttccc aggaagcaca gagggccctg
gggacctccc cagagctgcc 540gacgggcgtg actggttcct cagggaccag gctccccccg
acgccaaagg ctcaggatgg 600agggcctgtg ggcacggagc ttttccgagt gcctcccgtc
tccactgccg ccacgtggca 660gagttctgct ccccaccaac ctgggcccag cctctgggct
gaggcaaaga cctctgaggc 720cccgtccacc caggacccct ccacccaggc ctccactgcg
tcctccccag ccccagagga 780gaatgctccg tctgaaggcc agcgtgtgtg gggtcaggga
cagagcccca ggccagagaa 840ctctctggag cgggaggaga tgggtcccgt gccagcgcac
acggatgcct tccaggactg 900ggggcctggc agcatggccc acgtctctgt ggtccctgtc
tcctcagaag ggacccccag 960cagggagcca gtggcttcag gcagctggac ccctaaggct
gaggaaccca tccatgccac 1020catggacccc cagaggctgg gcgtccttat cactcctgtc
cctgacgccc aggctgccac 1080ccggaggcag gcggtggggc tgctggcctt ccttggcctc
ctcttctgcc tgggggtggc 1140catgttcacc taccagagcc tccagggctg ccctcgaaag
atggcaggag agatggcgga 1200gggccttcgc tacatccccc ggagctgtgg tagtaattca
tatgtcctgg tgcccgtgtg 1260aactcctctg gcctgtgtct agttgtttga ttcagacagc
tgcctgggat ccctcatcct 1320catacccacc cccacccaag ggcctggcct gagctgggat
gattggaggg gggaggtggg 1380atcctccagg tgcacaagct ccaagctccc aggcattccc
caggaggcca gccttgacca 1440ttctccacct tccagggaca gagggggtgg cctcccaact
caccccagcc ccaaaactct 1500cctctgctgc tggctggtta gaggttccct ttgacgccat
cccagcccca atgaacaatt 1560atttattaaa tgcccagccc cttctgaccc atgctgccct
gtgagtacta cagtcctccc 1620atctcacaca tgagcatcag gccaggccct ctgcccactc
cctgcaacct gattgtgtct 1680cttggtcctg ctgcagttgc cagtcacccc ggccacctgc
ggtgctatct cccccagccc 1740catcctctgt acagagccca cgcccccact ggtgacatgt
cttttcttgc atgaggctag 1800tgtggtgttt cctggcactg cttccagtga ggctctgccc
ttggttaggc attgtgggaa 1860ggggagataa gggtatctgg tgactttcct ctttggtcta
cactgtgctg agtctgaagg 1920ctgggttctg atcctagttc caccatcaag ccaccaacat
actcccatct gtgaaaggaa 1980agagggaggt aaggaatacc tgtccccctg acaacactca
ttgacctgag gcccttctct 2040ccagcccctg gatgcagcct cacagtcctt accagcagag
caccttagac agtccctgcc 2100aatggactaa cttgtctttg gaccctgagg cccagagggc
ctgcaaggga gtgagttgat 2160agcacagacc ctgccctgtg ggcccccaaa tggaaatggg
cagagcagag accatccctg 2220aaggccccgc ccaggcttag tcactgagac agcccgggct
ctgcctccca tcacccgcta 2280agagggaggg agggctccag acacatgtcc aagaagccca
ggaaaggctc caggagcagc 2340cacattcctg atgcttcttc agagactcct gcaggcagcc
aggccacaag acccttgtgg 2400tcccacccca cacacgccag attctttcct gaggctgggc
tcccttccca cctctctcac 2460tccttgaaaa cactgttctc tgccctccaa gaccttctcc
ttcacctttg tccccaccgc 2520agacaggacc agggatttcc atgatgtttt ccatgagtcc
cctgtttgtt tctgaaaggg 2580acgctacccg ggaagggggc tgggacatgg gaaaggggaa
gttgtaggca taaagtcagg 2640ggttcccttt tttggctgct gaaggctcga gcatgcctgg
atggggctgc accggctggc 2700ctggcccctc agggtccctg gtggcagctc acctctccct
tggattgtcc ccgacccttg 2760ccgtctacct gaggggcctc ttatgggctg ggttctaccc
aggtgctagg aacactcctt 2820cacagatggg tgcttggagg aaggaaaccc agctctggtc
catagagagc aagacgctgt 2880gctgccctgc ccacctggcc tctgcactcc cctgctgggt
gtggcgcagc atattcagga 2940agctcagggc ctggctcagg tggggtcact ctggcagctc
agagagggtg ggagtgggtc 3000caatgcactt tgttctggct cttccaggct gggagagcct
ttcaggggtg ggacaccctg 3060tgatggggcc ctgcctcctt tgtgaggaag ccgctggggc
cagttggtcc cccttccatg 3120gactttgtta gtttctccaa gcaggacatg gacaaggatg
atctaggaag actttggaaa 3180gagtaggaag actttggaaa gacttttcca accctcatca
ccaacgtctg tgccattttg 3240tattttacta ataaaattta aaagtcttgt gaaaaaaaaa
aaaaaaaaaa aaaaaaaaaa 3300aaaa
33044516PRTHomo sapiens 45Arg Ser Ser Ser Thr Leu
Pro Val Pro Val Phe Lys Arg Lys Ile Pro1 5
10 154616PRTHomo sapiens 46Pro Arg Gly Asn Gly Cys Pro
Arg Lys Glu Ile Ile Val Trp Lys Lys1 5 10
154716PRTHomo sapiens 47Leu Pro Arg Gly Asn Gly Cys Pro
Arg Lys Glu Ile Ile Val Trp Lys1 5 10
154816PRTHomo sapiens 48Gln Ile Leu Pro Arg Gly Asn Gly Cys
Pro Arg Lys Glu Ile Ile Val1 5 10
154916PRTHomo sapiens 49Ile Leu Pro Arg Gly Asn Gly Cys Pro Arg
Lys Glu Ile Ile Val Trp1 5 10
155016PRTHomo sapiens 50Arg Ile Gln Ile Leu Pro Arg Gly Asn Gly Cys
Pro Arg Lys Glu Ile1 5 10
155116PRTHomo sapiens 51Arg Gly Asn Gly Cys Pro Arg Lys Glu Ile Ile Val
Trp Lys Lys Asn1 5 10
155216PRTHomo sapiens 52Lys Arg Ser Ser Ser Thr Leu Pro Val Pro Val Phe
Lys Arg Lys Ile1 5 10
155316PRTHomo sapiens 53Ile Gln Ile Leu Pro Arg Gly Asn Gly Cys Pro Arg
Lys Glu Ile Ile1 5 10
155416PRTHomo sapiens 54Asp Arg Ile Gln Ile Leu Pro Arg Gly Asn Gly Cys
Pro Arg Lys Glu1 5 10
155516PRTHomo sapiens 55Arg Lys Arg Ser Ser Ser Thr Leu Pro Val Pro Val
Phe Lys Arg Lys1 5 10
155616PRTHomo sapiens 56Arg Cys Arg Cys Val Gln Glu Ser Ser Val Phe Ile
Pro Arg Arg Phe1 5 10
155716PRTHomo sapiens 57Gly Asn Gly Cys Pro Arg Lys Glu Ile Ile Val Trp
Lys Lys Asn Lys1 5 10
155816PRTHomo sapiens 58Cys Val Gln Glu Ser Ser Val Phe Ile Pro Arg Arg
Phe Ile Asp Arg1 5 10
155916PRTHomo sapiens 59Ile Asp Arg Ile Gln Ile Leu Pro Arg Gly Asn Gly
Cys Pro Arg Lys1 5 10
156016PRTHomo sapiens 60Leu Arg Cys Arg Cys Val Gln Glu Ser Ser Val Phe
Ile Pro Arg Arg1 5 10
156116PRTHomo sapiens 61Phe Ile Asp Arg Ile Gln Ile Leu Pro Arg Gly Asn
Gly Cys Pro Arg1 5 10
156216PRTHomo sapiens 62Arg Cys Val Gln Glu Ser Ser Val Phe Ile Pro Arg
Arg Phe Ile Asp1 5 10
156316PRTHomo sapiens 63Cys Arg Cys Val Gln Glu Ser Ser Val Phe Ile Pro
Arg Arg Phe Ile1 5 10
156416PRTHomo sapiens 64Gln Glu Ser Ser Val Phe Ile Pro Arg Arg Phe Ile
Asp Arg Ile Gln1 5 10
156516PRTHomo sapiens 65Arg Phe Ile Asp Arg Ile Gln Ile Leu Pro Arg Gly
Asn Gly Cys Pro1 5 10
156616PRTHomo sapiens 66Val Gln Glu Ser Ser Val Phe Ile Pro Arg Arg Phe
Ile Asp Arg Ile1 5 10
156716PRTHomo sapiens 67Glu Ser Ser Val Phe Ile Pro Arg Arg Phe Ile Asp
Arg Ile Gln Ile1 5 10
156816PRTHomo sapiens 68Ser Leu Arg Cys Arg Cys Val Gln Glu Ser Ser Val
Phe Ile Pro Arg1 5 10
156916PRTHomo sapiens 69Asn Gly Cys Pro Arg Lys Glu Ile Ile Val Trp Lys
Lys Asn Lys Ser1 5 10
157016PRTHomo sapiens 70Pro Gln Ala Glu Trp Ile Gln Arg Met Met Glu Val
Leu Arg Lys Arg1 5 10
157116PRTHomo sapiens 71Arg Arg Phe Ile Asp Arg Ile Gln Ile Leu Pro Arg
Gly Asn Gly Cys1 5 10
157216PRTHomo sapiens 72Leu Arg Lys Arg Ser Ser Ser Thr Leu Pro Val Pro
Val Phe Lys Arg1 5 10
157311PRTHomo sapiens 73Val Gln Glu Ser Ser Val Phe Ile Pro Arg Arg1
5 107426PRTHomo sapiens 74Glu Trp Ile Gln Arg
Met Met Glu Val Leu Arg Lys Arg Ser Ser Ser1 5
10 15Thr Leu Pro Val Pro Val Phe Lys Arg Lys
20 25754PRTHomo sapiens 75Lys Lys Asn Lys1766PRTHomo
sapiens 76Arg Lys Arg Ser Ser Ser1 5776PRTHomo sapiens
77Arg Gly Asn Gly Cys Pro1 57821PRTHomo sapiens 78Val Tyr
Tyr Thr Ser Leu Arg Cys Arg Cys Val Gln Glu Ser Ser Val1 5
10 15Phe Ile Pro Arg Arg
20797PRTHomo sapiens 79Asp Arg Ile Gln Ile Leu Pro1
5807PRTHomo sapiens 80Arg Lys Glu Ile Ile Val Trp1
5819PRTHomo sapiens 81Lys Ser Ile Val Cys Val Asp Pro Gln1
58213PRTHomo sapiens 82Thr Ser Leu Val Glu Asn His Leu Cys Pro Ala Thr
Glu1 5 108315PRTHomo sapiens 83Glu Gly
Ser Val Gly Trp Val Leu Gly Thr Phe Leu Cys Lys Thr1 5
10 15847PRTHomo sapiens 84Leu Pro Arg Cys
Thr Phe Ser1 58510PRTHomo sapiens 85Leu Ala Arg Leu Lys Ala
Val Asp Asn Thr1 5 108610PRTHomo sapiens
86Met Ala Ser Phe Lys Ala Val Phe Val Pro1 5
108716PRTHomo sapiens 87Ala Ala Gly Pro Glu Ala Gly Glu Asn Gln Lys
Gln Pro Glu Lys Asn1 5 10
158816PRTHomo sapiens 88Ser Gln Ala Ser Glu Gly Ala Ser Ser Asp Ile His
Thr Pro Ala Gln1 5 10
158916PRTHomo sapiens 89Ser Thr Leu Gln Ser Thr Gln Arg Pro Thr Leu Pro
Val Gly Ser Leu1 5 10
159016PRTHomo sapiens 90Ser Trp Ser Val Cys Gly Gly Asn Lys Asp Pro Trp
Val Gln Glu Leu1 5 10
159116PRTHomo sapiens 91Gly Pro Thr Ala Arg Thr Ser Ala Thr Val Pro Val
Leu Cys Leu Leu1 5 10
159216PRTHomo sapiens 92Ser Gly Ile Val Ala His Gln Lys His Leu Leu Pro
Thr Ser Pro Pro1 5 10
15936PRTHomo sapiens 93Arg Leu Arg Lys His Leu1 5947PRTHomo
sapiens 94Leu Gln Ser Thr Gln Arg Pro1 59513PRTHomo sapiens
95Ser Ser Asp Lys Glu Leu Thr Arg Pro Asn Glu Thr Thr1 5
109612PRTHomo sapiens 96Ala Gly Glu Asn Gln Lys Gln Pro
Glu Lys Asn Ala1 5 10976PRTHomo sapiens
97Asn Glu Gly Ser Val Thr1 5989PRTHomo sapiens 98Ile Ser
Ser Asp Ser Pro Pro Ser Val1 5998PRTHomo sapiens 99Cys Gly
Gly Asn Lys Asp Pro Trp1 510021PRTHomo sapiens 100Leu Leu
Pro Thr Ser Pro Pro Ile Ser Gln Ala Ser Glu Gly Ala Ser1 5
10 15Ser Asp Ile His Thr
2010128PRTHomo sapiens 101Ser Thr Gln Arg Pro Thr Leu Pro Val Gly Ser Leu
Ser Ser Asp Lys1 5 10
15Glu Leu Thr Arg Pro Asn Glu Thr Thr Ile His Thr 20
2510227PRTHomo sapiens 102Ser Leu Ala Ala Gly Pro Glu Ala Gly Glu
Asn Gln Lys Gln Pro Glu1 5 10
15Lys Asn Ala Gly Pro Thr Ala Arg Thr Ser Ala 20
251039PRTHomo sapiens 103Thr Gly Ser Cys Tyr Cys Gly Lys Arg1
51047PRTHomo sapiens 104Asp Ser Pro Pro Ser Val Gln1
510526PRTHomo sapiens 105Arg Lys His Leu Arg Ala Tyr His Arg Cys Leu
Tyr Tyr Thr Arg Phe1 5 10
15Gln Leu Leu Ser Trp Ser Val Cys Gly Gly 20
2510637PRTHomo sapiens 106Trp Val Gln Glu Leu Met Ser Cys Leu Asp Leu Lys
Glu Cys Gly His1 5 10
15Ala Tyr Ser Gly Ile Val Ala His Gln Lys His Leu Leu Pro Thr Ser
20 25 30Pro Pro Ile Ser Gln
3510715PRTHomo sapiens 107Ser Asp Ile His Thr Pro Ala Gln Met Leu Leu Ser
Thr Leu Gln1 5 10
151089PRTHomo sapiens 108Arg Pro Thr Leu Pro Val Gly Ser Leu1
51099PRTHomo sapiens 109Thr Ala Gly His Ser Leu Ala Ala Gly1
511013PRTHomo sapiens 110Gly Lys Arg Ile Ser Ser Asp Ser Pro Pro Ser
Val Gln1 5 1011130PRTHomo sapiens 111Lys
Asp Pro Trp Val Gln Glu Leu Met Ser Cys Leu Asp Leu Lys Glu1
5 10 15Cys Gly His Ala Tyr Ser Gly
Ile Val Ala His Gln Lys His 20 25
3011210PRTHomo sapiens 112His Gln Asp Phe Leu Gln Phe Ser Lys Val1
5 1011314PRTHomo sapiens 113Ala Gly Ile His
Glu Trp Val Phe Gly Gln Val Met Cys Lys1 5
1011423PRTHomo sapiens 114Pro Gln Ile Ile Tyr Gly Asn Val Phe Asn Leu
Asp Lys Leu Ile Cys1 5 10
15Gly Tyr His Asp Glu Ala Ile 2011516PRTHomo sapiens 115Tyr
Tyr Ala Met Thr Ser Phe His Tyr Thr Ile Met Val Thr Glu Ala1
5 10 1511612PRTHomo sapiens 116Leu
Ala Tyr His Tyr Pro Ile Gly Trp Ala Val Leu1 5
1011734PRTHomo sapiens 117Lys Arg His Arg Lys Val Cys Gly Asn Pro
Lys Ser Arg Glu Val Gln1 5 10
15Arg Ala Met Lys Leu Leu Asp Ala Arg Asn Lys Val Phe Ala Lys Leu
20 25 30His His11820PRTHomo
sapiens 118Phe Glu Asp Cys Cys Leu Ala Tyr His Tyr Pro Ile Gly Trp Ala
Val1 5 10 15Leu Arg Arg
Ala 2011927PRTHomo sapiens 119Ile Gln Glu Val Ser Gly Ser Cys
Asn Leu Pro Ala Ala Ile Phe Tyr1 5 10
15Leu Pro Lys Arg His Arg Lys Val Cys Gly Asn 20
251208PRTHomo sapiens 120Ala Met Lys Leu Leu Asp Ala Arg1
51219PRTHomo sapiens 121Lys Val Phe Ala Lys Leu His His
Asn1 512210PRTHomo sapiens 122Gln Ala Gly Pro His Ala Val
Lys Lys Leu1 5 1012314PRTHomo sapiens
123Phe Tyr Leu Pro Lys Arg His Arg Lys Val Cys Gly Asn Pro1
5 1012414PRTHomo sapiens 124Tyr Leu Pro Lys Arg His Arg
Lys Val Cys Gly Asn Pro Lys1 5
1012514PRTHomo sapiens 125Leu Pro Lys Arg His Arg Lys Val Cys Gly Asn Pro
Lys Ser1 5 1012614PRTHomo sapiens 126Pro
Lys Arg His Arg Lys Val Cys Gly Asn Pro Lys Ser Arg1 5
1012714PRTHomo sapiens 127Cys Gly Asn Pro Lys Ser Arg Glu
Val Gln Arg Ala Met Lys1 5 1012814PRTHomo
sapiens 128Gly Asn Pro Lys Ser Arg Glu Val Gln Arg Ala Met Lys Leu1
5 1012914PRTHomo sapiens 129Lys Phe Ser Asn Pro
Ile Ser Ser Ser Lys Arg Asn Val Ser1 5
101306PRTHomo sapiens 130Pro Lys Ser Arg Glu Val1
51317PRTHomo sapiens 131Leu His His Asn Thr Gln Thr1
51326PRTHomo sapiens 132Ser Ser Ser Lys Arg Asn1
51339PRTHomo sapiens 133Gln Phe Ala Ser His Phe Leu Pro Pro1
51349PRTHomo sapiens 134Ala Ala Ala Asp Gln Trp Lys Phe Gln1
513510PRTHomo sapiens 135Thr Phe Met Cys Lys Val Val Asn Ser Met1
5 1013610PRTHomo sapiens 136Ile Ala Ile Cys Thr
Met Val Tyr Pro Ser1 5 1013725PRTHomo
sapiens 137Val Gln Thr Ile Asp Ala Tyr Ala Met Phe Ile Ser Asn Cys Ala
Val1 5 10 15Ser Thr Asn
Ile Asp Ile Cys Phe Gln 20 25
User Contributions:
Comment about this patent or add new information about this topic:
| People who visited this patent also read: | |
| Patent application number | Title |
|---|---|
| 20130026735 | TRAILER TONGUE CONNECTION UNIT |
| 20130026734 | SCOOTER WITH FOLDING DEVICE AND FOLDING DEVICE |
| 20130026733 | VEHICLE BODY FRAME |
| 20130026732 | DOUBLE BRIDGE BICYCLE FORK |
| 20130026731 | WHEEL SPEED SENSOR MOUNTING STRUCTURE |