Patent application title: MOLECULAR SIGNATURE OF LIVER TUMOR GRADE AND USE TO EVALUATE PROGNOSIS AND THERAPEUTIC REGIMEN
Inventors:
Marie Annick Buendia (Le Perreux Sur Marne, FR)
Carolina Niell (Blanes (girona), ES)
Stefano Cairo (Longpont-Sur-Orge, FR)
Aurélien De Reynies (Paris, FR)
Aurélien De Reynies (Paris, FR)
Assignees:
INSTITUT PASTEUR
Centre National De La Recherche Scientifique (CNRS)
INSTITUT NATIONAL DE LA SANTE ET DE LA RECHERCHE MEDICALE (INSERM)
IPC8 Class: AC40B3004FI
USPC Class:
506009000
Class name:
Publication date: 2012-02-16
Patent application number: 20120040848
Abstract:
The present invention concerns a method to determine the gene expression
profile on a sample previously obtained from a patient diagnosed for a
liver tumor, comprising assaying the expression of a set of genes in this
sample and determining the gene expression profile (signature). In a
particular embodiment, said method enables to determine the grade of
liver tumor, such as hepatoblastoma (HB) or a hepatocellular carcinoma
(HCC). The invention is also directed to kits comprising a plurality of
pairs of primers or a plurality of probes specific for a set of genes, as
well as to solid support or composition comprising a set of probes
specific for a set of genes. These methods are useful to determine the
grade of a liver tumor in sample obtained from a patient, to determine
the risk of developing metastasis and/or to define the therapeutic
regimen to apply to a patient.Claims:
1. Method to determine the gene expression profile on a biological
sample, comprising: a. assaying the expression of a set of genes in a
sample previously obtained from a patient diagnosed for a liver tumor,
wherein said set comprises from 2 to 16 genes or consists of 2 to 16
genes, said 2 to 16 genes being chosen in the group consisting in the
alpha-fetoprotein (AFP), aldehyde dehydrogenase 2 (ALDH2), amyloid P
component serum (APCS), apolipoprotein C-IV (APOC4), aquaporin 9 (AQP9),
budding uninhibited by benzimidazoles 1 (BUB1), complement componant 1
(C1S), cytochrome p450 2E1 (CYP2E1), discs large homolog 7 (DLG7), dual
specificity phosphatase 9 (DUSP9), E2F5 transcription factor (E2F5),
growth hormone receptor (GHR), 4-hydroxyphenylpyruvase dioxygenase (DHP),
immunoglogulin superfamily member 1 (IGSF1), Notchless homolog 1 (NLE1)
and the ribosomal protein L10a (RPL10A) genes; and b. determining the
gene expression profile of said sample.
2. Method according to claim 1, which further comprises determining the grade of the liver tumor providing the sample, for example by comparing the obtained gene expression profile of said sample to the gene expression profile of a reference sample or to the gene expression profiles of a collection of reference samples or by applying a discretization method for classification.
3. Method according to claim 1 or 2, wherein the assay of the expression of said set of genes comprises a step of detecting nucleotide targets, wherein each nucleotide target is a product resulting from the expression of one of the genes in said set.
4. Method according to claim 2, wherein said nucleotide targets are mRNA.
5. Method according to any one of claims 1 to 4, wherein the assay of the expression of said set of genes comprises an amplification step, such as performed by qualitative polymerase chain reaction prior to a step of detecting the mRNA of each gene of said set.
6. Method according to any one of claims 1 to 5, wherein the assay of the expression of said set of genes comprises a hybridization step, such as one performed by hybridization on a solid or liquid support, especially on an array, prior to a step of detecting the mRNA of each gene of said set.
7. Method according to any one of claims 2 to 5, wherein said detected nucleotide targets are quantified with respect to at least one nucleotide target, expression product of an invariant gene, such as ACTG1, EFF1A1, PNN and RHOT2 genes.
8. Method according to any one of claims 1 to 7, wherein said liver tumor is a hepatoblastoma (HB) or a hepatocellular carcinoma (HCC).
9. Method according to any one of claims 2 to 8, wherein said method comprises, before step a., the preparation of said nucleotide targets from the sample.
10. Method according to any one of claims 1 to 9, wherein said set of genes comprises or consists in a set chosen in the group consisting of: (a) E2F5 and HPD genes; (b) APCS, BUB1, E2F5, GHR and HPD genes; (c) ALDH2, APCS, APOC4, BUB1, C1S, CYP2E1, E2F5, GHR and HPD genes; (d) ALDH2, APCS, APOC4, AQP9, BUB1, C1S, DUSP9, E2F5 and RPL10A genes; (e) ALDH2, APCS, APOC4, AQP9, C1S, CYP2E1, E2F5, GHR, IGSF1 and RPL10A genes; and (f) AFP, ALDH2, APCS, APOC4, AQP9, BUB1, C1S, CYP2E1, DLG7, DUSP9, E2F5, GHR, HPD, IGSF1, NLE1 and RPL10A genes.
11. Method enabling the determination of the tumor grade on a patient's biological sample, which comprises a classification of the tumor through discretization according the following steps: In a method according to any of claims 1 to 10, measuring the expression and especially the relative (normalized) expression of each gene in a set of genes defined as the signature of the tumor, for example by quantitative PCR thereby obtaining data as Ct or preferably Delta Ct in said biological sample wherein said set of genes is divided in two groups, a first group consisting of the proliferation-related genes and a second group consisting of the differentiation-related genes, comparing the values measured for each gene, to a cut-off value determined for each gene of the set of genes, and assigning a discretized value to each of said measured expression values with respect to said cut-off value, said discretized value being advantageously a "1" or a "2" and optionally a "1.5" value with respect to the cut-off value, determining the average of the discretized values for the genes, in each group of the set of genes, determining a score calculated as a ratio the average for the discretized values for the proliferating-related genes on the average for the discretized values for the differentiation-related genes, comparing the obtained score for the biological sample with one or more sample cut-off(s) value(s), wherein each cut-off value corresponds to a selected percentile, determining the tumor grade as C1 or C2, as a result of the classification of the biological sample with respect to said sample cut-off.
12. Method according to claim 11, wherein the relative expression determined for the profiled gene is obtained by normalizing with respect to the invariant RHOT2 gene.
13. Method according to claim 11, wherein the determination of the tumor grade on a biological sample comprises applying the following conditions: a) for a hepatoblastoma: the set of assayed genes for profiling is constituted of the 16 genes disclosed; the invariant gene (of reference) is RHOT2; the cut-offs value for each gene are: AFP: 3.96139596; ALDH2: 4.3590482; APCS: 4.4691582; APOC4: 2.03068712; AQP9: 3.38391456; BUB1: -1.41294708; C1S: 4.24839464; CYP2E1: 6.70659644; DLG7: -3.3912188; DUSP9: 2.07022648; E2F5: -0.72728656; GHR: -0.1505569200; HPD: 2.27655628; IGSF1: 0.1075015200; NLE: -0.02343571999; RPL10A: 6.19723876. the cut-off value for the sample is 0.91 and a sample with a score above 0.91 is classified into the C2 class and a sample with a score below 0.91 is classified into the C1 class. b) for a hepatocellular carcinoma: the set of assayed genes for profiling is constituted of the 16 genes disclosed; the invariant gene (of reference) is RHOT2; the cut-offs value for each gene is: TABLE-US-00033 Gene name Cut-off for Taqman Cut-off for SybrGreen AEP -1.2634010 -2.3753035 ALDH2 4.014143 5.314302 APCS 5.6142907 6.399079 APOC4 -0.7963158 4.656336 AQP9 4.2836011 5.446966 BUB1 -1.2736579 -3.634476 C1S 6.3514679 6.240002 CYP2E1 6.9562419 5.829384 DLG7 -2.335694 -4.614352 DUSP9 -7.979559 -1.8626715 E2F5 -0.4400218 -1.367846 GHR 1.0832632 1.169362 HPD 6.480328 6.736329 IGSF1 -4.8417785 7.6653982 NLE -1.6167268 -1.82226 RPL10A 6.2483056 5.731897
the cut-off value for the sample corresponding to the 67.sup.th percentile is 0.925 and the cut-off value corresponding to the 33.sup.th percentile is 0.66 and a sample with a score above 0.925 is classified into the C2 class and a sample with a score below 0.66 is classified into the C1 class.
14. Method according to claim 13 wherein in the case of a hepatocellular carcinoma, a sample with a score (initial score) between 0.66 and 0.925 is refined to obtain a modified score, the modified score being either "1" or "2" depending on the calculated average of the discretized values for the proliferation-related genes only, said average being discretized at a determined percentile (the 60.sup.th for example) and "1" is assigned if the sample has an average below the value at the percentile of reference and "2" is assigned if the sample has an average above the value at the percentile of reference.
15. Kit, suitable to carry out the method as defined in any one of claims 1 to 13, comprising a. a plurality of pairs of primers specific for a set of genes to be assayed, said set comprising 2 to 16 genes or consisting of 2 to 16 genes, said 2 to 16 genes being chosen in the group consisting of AFP, ALDH2, APCS, APOC4, AQP9, BUB1, C1S, CYP2E1, DLG7, DUSP9, E2F5, GHR, HPD, IGSF1, NLE1 and RPL10A genes; and b. optionally reagents necessary for the amplification of the nucleotide targets of these genes by said primers, and optionally reagents for detecting the amplification products.
16. Kit according to claim 14, wherein each primer is 10 to 30 bp in length and has at least 80% similarity with its complementary sequence in the nucleotide target, preferably 100%.
17. Kit according to claim 14 or 15, wherein said pairs of primers are chosen in the group consisting of: TABLE-US-00034 Forward primer gene (5'-3') Reverse primer (5'-3') AFP AACTATTGGCCTGTGGCGAG TCATCCACCACCAAGCTGC ALDH2 GTTTGGAGCCCAGTCACCCT GGGAGGAAGCTTGCATGATTC APCS GGCCAGGAATATGAACAAGCC CTTCTCCAGCGGTGTGATCA APOC4 GGAGCTGCTGGAGACAGTGG TTTGGATTCGAGGAACCAGG AQP9 GCTTCCTCCCTGGGACTGA CAACCAAAGGGCCCACTACA BUB1 ACCCCTGAAAAAGTGATGCCT TCATCCTGTTCCAAAAATCCG C1S TTGTTTGGTTCTGTCATCCGC TGGAACACATTTCGGCAGC CYP2E1 CAACCAAGAATTTCCTGATC AAGAAACAACTCCATGCGAGC CAG DLG7 GCAGGAAGAATGTGCTGAAA TCCAAGTCTTTGAGAAGGGCC CA DUSP9 CGGAGGCCATTGAGTTCATT ACCAGGTCATAGGCATCGTTG E2F5 CCATTCAGGCACCTTCTGGT ACGGGCTTAGATGAACTCGACT GHR CTTGGCACTGGCAGGATCA AGGTGAACGGCACTTGGTG HPD ATCTTCACCAAACCGGTGCA CCATGTTGGTGAGGTTACCCC IGSF1 CACTCACACTGAAAAACGCCC GGGTGGAGCAATTGAAAGTCA NLE1 ATGTGAAGGCCCAGAAGCTG GAGAACTTCGGGCCGTCTC RPL10A TATCCCCCACATGGACATCG TGCCTTATTTAAACCTGGGCC
and, a modified group of primers with respect to the above, wherein one or more primer(s) is modified, provided said primer(s) has at least 80% similarity with its non-modified version above.
18. A set of probes, suitable to carry out the method as defined in any one of claims 1 to 13, comprising a plurality of probes specific for a set of genes to assay, said set comprising or having from 2 to 16 genes, said 2 to 16 genes being chosen in the group consisting of AFP, ALDH2, APCS, APOC4, AQP9, BUB1, C1S, CYP2E1, DLG7, DUSP9, E2F5, GHR, HPD, IGSF1, NLE1 and RPL10A genes.
19. A set of probes according to claim 17, wherein said probes are 50 to 200 bp in length and have at least 80% similarity to the complementary sequence of the nucleotide target of the gene, preferably 100%.
20. A solid support, especially an array comprising a set of probes as defined in claims 17 or 18 linked to a support.
21. A composition comprising a set of probes as defined in claim 17 or 18, in solution.
22. A kit comprising a set of probes as defined in claim 17 or 18, a solid support as defined in claim 19 or a composition as defined in claim 20, and optionally reagents necessary for the hybridization of said nucleotide targets to said probes.
23. Set of probes, solid support, arrays, compositions or kits according to any one of claims 14 to 21, suitable for assaying a set of genes which comprises or consists in a set chosen in the group consisting of: (a) E2F5 and HPD genes; (b) APCS, BUB1, E2F5, GHR and HPD genes; (c) ALDH2, APCS, APOC4, BUB1, C1S, CYP2E1, E2F5, GHR and HPD genes; (d) ALDH2, APCS, APOC4, AQP9, BUB1, CIS, DUSP9, E2F5 and RPL10A genes; (e) ALDH2, APCS, APOC4, AQP9, C1S, CYP2E1, E2F5, GHR, IGSF1 and RPL10A genes; and (f) AFP, ALDH2, APCS, APOC4, AQP9, BUB1, C1S, CYP2E1, DLG7, DUSP9, E2F5, GHR, HPD, IGSF1, NLE1 and RPL10A genes.
24. Set of probes, solid support, arrays, compositions or kits according to any one of claims 14 to 23, wherein the invariant gene is the RHOT2 gene or the PNN gene.
25. Use of a set of probes, solid support, arrays, compositions or kits according to any one of claims 14 to 24, to determine the grade of a liver tumor in a sample obtained from a patient.
26. Use according to claim 25 or method of claim 11, wherein for a hepatoblastoma or for a hepatocellular carcinoma the cut-off value of the profiled genes are determined for the overexpressed proliferation-related genes at a percentile within the range of the 60.sup.th to the 80.sup.th percentile, especially at the 67.sup.th percentile and the cut-off value of the profiled genes are determined for the downregulated differentiation-related genes at a percentile within the range of the 30.sup.rd to 45.sup.th percentile, especially at the 33rd or 40.sup.th percentile and the cut-off value of the sample is determined within the same range of the 60.sup.th to the 80.sup.th percentile.
27. Use of a set of probes, arrays, compositions or kits according to any one of claims 14 to 21, to determine, in a patient, the risk of developing metastasis.
28. Use of a set of probes, arrays, compositions or kits according to any one of claims 14 to 21, to define the therapeutic regimen to apply to said patient.
Description:
[0001] The present invention relates to a method to in vitro determine
the grade of a liver tumor in a sample previously obtained from a
patient, using a molecular signature based on the expression of a set of
genes comprising at least 2, especially has or consist of 2 to 16 genes,
preferably a set of 16 genes. In a particular embodiment, the method
focuses on hepatoblastoma (HB) or hepatocellular carcinoma (HCC), in
adults or in children. The invention is also directed to sets of primers,
sets of probes, compositions, kits or arrays, comprising primers or
probes specific for a set of genes comprising at least 2 genes,
especially has or consists of 2 to 16 genes, preferably exactly 16 genes.
Said sets, kits and arrays are tools suitable to determine the grade of a
liver tumor in a patient.
[0002] The liver is a common site of metastases from a variety of organs such as lung, breast, colon and rectum. However, liver is also a site of different kinds of cancerous tumors that start in the liver (primary liver cancers). The most frequent is the Hepatocellular Carcinoma (HCC) (about 3 out of 4 primary liver cancers are this type) and is mainly diagnosed in adults. In the United States approximately 10,000 new patients are diagnosed with hepatocellular carcinoma each year. Less frequent liver tumours are cholangiocarcinoma (CC) in adults and hepatoblastoma (HB) in children.
[0003] The prognosis and treatment options associated with these different kinds of cancers is difficult to predict, and is dependent in particular on the stage of the cancer (such as the size of the tumor, whether it affects part or all of the liver, has spread to other places in the body or its aggressiveness). Therefore, it is important for clinicians and physicians to establish a classification of primary liver cancers (HCC or HB) to propose the most appropriate treatment and adopt the most appropriate surgery strategy. Some factors are currently used (degree of local invasion, histological types of cancer with specific grading, tumour markers and general status of the patient) but have been found to not be accurate and sufficient enough to ensure a correct classification.
[0004] As far as the HB is concerned, the PRETEXT (pre-treatment extent of disease) system designed by the International Childhood Liver Tumor Strategy Group (SIOPEL) is a non invasive technique commonly used by clinicians, to assess the extent of liver cancer, to determine the time of surgery and to adapt the treatment protocol. This system is based on the division of the liver in four parts and the determination of the number of liver sections that are free of tumor (Aronson et al. 2005; Journal of Clinical Oncology; 23 (6): 1245-1252). A revised staging system taking into account other criteria, such as caudate lobe involvement, extrahepatic abdominal disease, tumor focality, tumor rupture or intraperitoneal haemorrhage, distant metastases, lymph node metastases, portal vein involvement and involvement of the IVC (inferior vena cava) and/or hepatic veins, has been recently proposed (Roebuck; 2007; Pediatr Radiol; 37: 123-132). However, the PRETEXT system, even if reproducible and providing good prognostic value, is based on imaging and clinical symptoms, making this system dependent upon the technicians and clinicians. There is thus a need for a system, complementary to the PRETEXT system, based on genetic and molecular features of the liver tumors.
[0005] The present invention concerns a method or process of profiling gene expression for a set of genes, in a sample previously obtained from a patient diagnosed for a liver tumor. In a particular embodiment said method is designed to determine the grade of a liver tumor in a patient.
[0006] By "liver tumor" or "hepatic tumor", it is meant a tumor originating from the liver of a patient, which is a malignant tumor (comprising cancerous cells), as opposed to a benign tumor (non cancerous) which is explicitly excluded. Malignant liver tumors encompass two main kinds of tumors: hepatoblastoma (HB) or hepatocellular carcinoma (HCC). These two tumor types can be assayed for the presently reported molecular signature. However, the present method may also be used to assay malignant liver tumors which are classified as unspecified (non-HB, non-HCC).
[0007] The present method may be used to determine the grade of a liver tumor or several liver tumors of the same patient, depending on the extent of the liver cancer. For convenience, the expression "a liver tumor" will be used throughout the specification to possibly apply to "one or several liver tumor(s)". The term "neoplasm" may also be used as a synonymous of "tumor".
[0008] In a particular embodiment, the tumor whose grade has to be determined is located in the liver. The presence of the tumor(s) in the liver may be diagnosed by ultrasound scan, x-rays, blood test, CT scans (computerised tomography) and/or MRI scans (magnetic resonance imaging).
[0009] In a particular embodiment, the tumor, although originating from the liver, has extended to other tissues or has given rise to metastasis.
[0010] In a particular embodiment, the patient is a child i.e., a human host who is under 20 years of age according to the present application. Therefore, in a particular embodiment, the liver tumor is a paediatric HB or a paediatric HCC. In another embodiment, the liver tumor is an adult HCC.
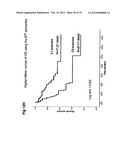
[0011] A grade is defined as a subclass of the liver tumor, corresponding to prognostic factors, such as tumor status, liver function and general health status. The present method of the invention allows or at least contributes to differentiating liver tumors having a good prognosis from tumors with a bad prognosis, in terms of evolution of the patient's disease. A good prognosis tumor is defined as a tumor with good survival probability for the patient (more than 80% survival at two years for HB and more than 50% survival at two years for HCC), low probability of metastases and good response to treatment for the patient. In contrast, a bad prognosis tumor is defined as a tumor with an advanced stage, such as one having vascular invasion or/and extrahepatic metastasis, and associated with a low survival probability for the patient (less than 50% survival in two years).
[0012] The method of the invention is carried out on a sample isolated from the patient who has previously been diagnosed for the tumor(s) and who, optionally, may have been treated by surgery. In a preferred embodiment, the sample is the liver tumor (tumoral tissue) or of one of the liver tumors identified by diagnosis imaging and obtained by surgery or a biopsy of this tumor. The tumor located in the liver tumor is called the primary tumor.
[0013] In another embodiment, the sample is not the liver tumor, but is representative of this tumor. By "representative", it is meant that the sample is regarded as having the same features as the primary tumors, when considering the gene expression profile assayed in the present invention. Therefore, the sample may also consist of metastatic cells (secondary tumors spread into different part(s) of the body) or of a biological fluid containing cancerous cells (such as blood).
[0014] The sample may be fixed, for example in formalin (formalin fixed). In addition or alternatively, the sample may be embedded in paraffin (paraffin-embedded) or equivalent products. In particular, the tested sample is a formalin-fixed, paraffin-embedded (FFPE) sample.
[0015] One advantage of the method of the present invention is that, despite the possible heterogeneity of some liver tumors (comprising epithelial tumor cells at different stages of liver differentiation within the same tumor), the assay has proved to be reproducible and efficient on liver tumor biopsies obtained from any part of the whole tumor. Therefore, there is no requirement for the isolation of cells presenting particular features except from the fact that they are obtained from a liver tumor or are representative thereof, to carry out the gene expression profile assay.
[0016] In a particular embodiment, the tumor originates from a patient having a Caucasian origin, in particular European, North American, Australian, New-Zealander or Afrikaners.
[0017] In a first step, the method or process of the invention comprises assaying the expression level of a set of genes in a sample, in order to get an expression profile thereof.
[0018] By "expression of a set of genes" (or "gene expression"), it is meant assaying, in particular detecting, the product or several products resulting from the expression of a gene, this product being in the form of a nucleic acid, especially RNA, mRNA, cDNA, polypeptide, protein or any other formats. In a particular embodiment, the assay of the gene expression profile comprises detecting a set of nucleotide targets, each nucleotide target corresponding to the expression product of a gene encompassed in the set.
[0019] The expression "nucleotide target" means a nucleic acid molecule whose expression must be measured, preferably quantitatively measured. By "expression measured", it is meant that the expression product(s), in particular the transcription product(s) of a gene, are measured. By "quantitative" it is meant that the method is used to determine the quantity or the number of copies of the expression products, in particular the transcription products or nucleotide targets, originally present in the sample. This must be opposed to the qualitative measurement, whose aim is to determine the presence or absence of said expression product(s) only.
[0020] A nucleotide target is in particular a RNA, and most particularly a total RNA. In a preferred embodiment, the nucleotide target is mRNA or transcripts. According to the methods used to measure the gene expression level, the mRNA initially present in the sample may be used to obtain cDNA or cRNA, which is then detected and possibly measured.
[0021] In an embodiment, the expression of the gene is assayed directly on the sample, in particular in the tumor. In an alternative embodiment, the expression products or the nucleotide targets are prepared from the sample, in particular are isolated or even purified. When the nucleotide targets are mRNA, a further step comprising or consisting in the retro-transcription of said mRNA into cDNA (complementary DNA) may also be performed prior to the step of detecting expression. Optionally, the cDNA may also be transcribed in vitro to provide cRNA.
[0022] During the step of preparation, and before assaying the expression, the expression product(s) or the nucleotide target(s) may be labelled, with isotopic (such as radioactive) or non isotopic (such as fluorescent, coloured, luminescent, affinity, enzymatic, magnetic, thermal or electrical) markers or labels.
[0023] It is noteworthy that steps carried out for assaying the gene expression must not alter the qualitative or the quantitative expression (number of copies) of the expression product(s) or of the nucleotide target(s), or must not interfere with the subsequent step comprising assaying the qualitative or the quantitative expression of said expression product(s) or nucleotide target(s).
[0024] The step of profiling gene expression comprises determining the expression of a set of genes. Such a set is defined as a group of genes that must be assayed for one test, and especially performed at the same time, on the same patient's sample. A set comprises at least 2 and has especially from 2 to 16 genes, said 2 to 16 genes being chosen from the 16 following genes: alpha-fetoprotein (AFP), aldehyde dehydrogenase 2 (ALDH2), amyloid P component serum (APCS), apolipoprotein C-IV (APOC4), aquaporin 9 (AQP9), budding uninhibited by benzimidazoles 1 (BUB1), complement componant 1 (C1S), cytochrome p450 2E1 (CYP2E1), discs large homolog 7 (DLG7), dual specificity phosphatase 9 (DUSP9), E2F5 transcription factor (E2F5), growth hormone receptor (GHR), 4-hydroxyphenylpyruvase dioxygenase (HPD), immunoglogulin superfamily member 1 (IGSF1), Notchless homolog 1 (NLE1) and the ribosomal protein L10a (RPL10A) genes.
[0025] A complete description of these 16 genes is given in Table 1. This table lists, from left to right, the symbol of the gene, the complete name of the gene, the number of the SEQ ID provided in the sequence listing, the Accession Number from the NCBI database on June 2008, the human chromosomal location and the reported function (when known).
[0026] A set of genes comprises at least 2 out the 16 genes of Table 1, and particularly at least or exactly 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14 or 15 out of the 16 genes of Table 1. In a particular embodiment, the set comprises or consists of the 16 genes of Table 1 i.e. the set of genes comprises or consists of AFP, ALDH2, APCS, APOC4, AQP9, BUB1, C1S, CYP2E1, DLG7, DUSP9, E2F5, GHR, HPD, IGSF1, NLE1 and RPL10A genes. Accordingly, unless otherwise stated when reference is made in the present application to a set of 2 to 16 genes of Table 1, it should be understood as similarly applying to any number of genes within said 2 to 16 range.
[0027] In other particular embodiments, the set of genes comprises or consists of one of the following sets: (a) the E2F5 and HPD genes, (b) the APCS, BUB1, E2F5, GHR and HPD genes, (c) the ALDH2, APCS, APOC4, BUB1, C1S, CYP2E1, E2F5, GHR and HPD genes, (d) the ALDH2, APCS, APOC4, AQP9, BUB1, C1S, DUSP9, E2F5 and RPL10A genes, or (e) the ALDH2, APCS, APOC4, AQP9, C1S, CYP2E1, E2F5, GHR, IGSF1 and RPL10A genes.
[0028] As indicated by the expression "comprises from 2 to 16 genes of Table 1", the set may, besides the specific genes of Table 1, contain additional genes not listed in Table 1. This means that the set must comprises from 2 to 16 genes of Table 1, i.e. 2 to 16 genes of Table 1 (in particular 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, or 16 genes), and optionally comprises one or more additional genes. Said set may also be restricted to said 2 to 16 genes of Table 1.
[0029] Additional genes may be selected for the difference of expression observed between the various grades of liver cancer, in particular between a tumor of good prognosis and a tumor of poor prognosis. TABLE-US-00001 TABLE 1 mRNA Accession Protein symbol Gene name SEQ ID No Location Function SEQ ID AFP alpha-fetoprotein 1 NM_001134 4q11-q13 plasma protein synthesized 2 by the fetal liver ALDH2 aldehyde dehydrogenase 2 3 NM_000690 12q24.2 liver enzyme involved in 4 family (mitochondrial) alcohol metabolism APCS amyloid P component, serum 5 NM_001639 1q21-q23 secreted glycoprotein 6 AP0C4 apolipoprotein C-IV 7 NM_001646 19q13.2 secreted liver protein 8 AQP9 aquaporin 9 9 NM_020980 15q22.1-22.2 water-selective membrane channel 10 BUB1 BUB1 budding uninhibited 11 AF043294 2q14 kinase involved in spindle 12 by benzimidazoles 1 homolog checkpoint (yeast) C1S complement component 1, s 13 M18767 12p13 component of the cleavage and 14 subcomponent polyadenylation specificity factor complex CYP2E1 cytochrome P450, family 2, 15 AF182276 10q24.3-qter cytochrome P450 family member subfamily E, polypeptide 1 involved in drug metabolism DLG7 discs, large homolog7 17 NM_014750 14q22.3 cell cycle regulator involved 18 (Drosophila) (DLGAP5) in kinetocore formation DUSP9 dual specificity phosphatase 9 19 NM_001395 Xq28 phosphatase involved in 20 regulation of MAP Kinases E2F5 E2F transcription factor 5, 21 U15642 8q21.2 transcription factor involved in cell 22 p130-binding cycle regulation GHR Growth hormone receptor 23 NM_000163 5p13-p12 transmembrane receptor for 24 growth hormone HPD 4-hydroxyphenylpyruvate 25 NM_002150 12q24-qter enzyme involved in amino-acid 26 dioxygenase degradation IGSF1 immunoglobulin superfamily, 27 NM_001555 Xq25 cell recognition and 28 member 1 regulation of cell behavior NLE1 notchless homolog 1 29 NM_08096 17q12 unknown 30 (Drosophila) RPL10A ribosomal protein L10a 31 NM_007104 6p21.3-p21.2 ribosomal protein of 60S subunit 32
[0030] The invention also relates to a set of genes comprising or consisting of the 16 genes of Table 1 (i.e., AFP, ALDH2, APCS, APOC4, AQP9, BUB1, C1S, CYP2E1, DLG7, DUSP9, E2F5, GHR, HPD, IGSF1, NLE1 and RPL10A genes), in which 1, 2, 3, 4 or 5 genes out of the 16 genes are substituted by a gene presenting the same features in terms of difference of expression between a tumor of a good prognosis and a tumor of poor prognosis.
[0031] In a particular embodiment, the number of genes of the set does not exceed 100, particularly 50, 30, 20, more particularly 16 and even more particularly is maximum 5, 6, 7, 8, 9 or 10.
[0032] When considering adding or substituting a gene or several genes to the disclosed set, the person skilled in the art will consider one or several of the following features: [0033] (a) the added gene(s) and/or the substituted gene(s) of Table 1 must present the same features in terms of difference of expression between a tumor of a good prognosis and a tumor of poor prognosis as the genes of Table 1 when taken as a whole. Thus, the expression of the added gene or of the substituted gene in a tumor of a good prognosis is either overexpressed or underexpressed of a factor of at least 2, preferably of at least 5, and more preferably of at least 10, as compared to its expression in a tumor of poor prognosis. [0034] (b) besides presenting the feature in a), the added gene and/or the substituted gene may also provide, in combination with the other genes of the set, discriminant results with respect to the grade of the liver tumors; this discrimination is reflected by the homogeneity of expression profile of this gene in the tumors of a good prognosis on the one hand, and the tumors of poor prognosis in the other hand; and [0035] (c) finally, besides features of a) and/or b), the added gene and/or the substituted gene is optionally chosen among genes that are involved in liver differentiation, in particular having a specific expression in fetal liver, or genes that are involved in proliferation, for example in mitosis or associated with ribosomes.
[0036] Examples of genes which can be added or may replace genes of the set may be identified in following Table 2. TABLE-US-00002 TABLE 2 list of genes according to p value. Gene mean mean ratio Parametric symbol rC1 rC2 rC2/rC1 p-value FDR Description IPO4 123.7 248.3 2.0 2.00E-07 0.00036 importin 4 CPSF1 467.8 1010.7 2.2 2.00E-07 0.00036 cleavage and polyadenylation specific factor 1, 160 kDa MCM4 25.8 90.7 3.5 1.10E-06 0.00115 MCM4 minichromosome maintenance deficient 4 (S. cerevisise) EIF3S3 1319 2601.2 2.0 1.20E-06 0.00119 eukaryotic translation initiation factor 3, subunit 3 gamma, 40 kDa NCL 1319 2655.6 2.0 1.30E-06 0.00122 nucleolin CDC25C 35.7 99.3 2.8 1.40E-06 0.00124 cell division cycle 25C CENPA 28.2 78.4 2.8 1.50E-06 0.00124 centromere protein A, 17 kDa KIF14 24.7 54.2 2.2 1.50E-06 0.00124 kinesin family member 14 IPW 145.7 397.6 2.7 1.90E-06 0.0015 imprinted in Prader-Willi syndrome KNTC2 26.8 65.1 2.4 2.20E-06 0.00157 kinetochore associated 2 TMEM48 264 71.7 2.7 2.30E-06 0.00157 transmembrane protein 48 BOP1 87.2 270.9 3.1 2.30E-06 0.00157 block of proliferation 1 EIF3S9 170 372.4 2.2 2.30E-06 0.00157 eukaryotic translation initiation factor 3, subunit 9 eta, 116 kDa PH-4 340.9 168.2 0.5 2.40E-06 0.00158 hypoxia-inducible factor prolyl 4- hydroxylase SMC4L1 151.5 359.3 2.4 2.50E-06 0.0016 SMC4 structural maintenance of chromosomes 4-like 1 (yeast) TTK 23.7 74.2 3.1 2.60E-06 0.00161 TTK protein kinase LAMA3 696 136.3 0.2 2.80E-06 0.00168 laminin, alpha 3 C10orf72 192.6 67.7 0.4 2.90E-06 0.00169 Chromosome 10 open reading frame 72 TPX2 73.4 401.5 5.5 3.10E-06 0.00171 TPX2, microtubule-associated, homolog (Xenopus laevis) MSH2 75.5 212.1 2.8 3.20E-06 0.00171 mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) DKC1 358.1 833.5 2.3 3.20E-06 0.00171 dyskeratosis congenita 1, dyskerin STK6 86.4 395.3 4.6 3.30E-06 0.00172 serine/threonine kinase 6 CCT6A 200.5 526.6 2.6 3.50E-06 0.00173 chaperonin containing TCP1, subunit 6A (zeta 1) SULT1C1 67.5 314.8 4.7 3.50E-06 0.00173 sulfotransferase family, cytosolic, 1C, member 1 ILF3 142.3 294.5 2.1 3.70E-06 0.00174 interleukin enhancer binding factor 3, 90 kDa IMPDH2 916.9 2385.6 2.6 3.70E-06 0.00174 IMP (inosine monophosphate) dehydrogenase 2 HIC2 63.4 208.8 3.3 3.90E-06 0.00179 hypermethylated in cancer 2 AFM 1310.3 237.4 0.2 4.10E-06 0.00184 afamin MCM7 187.3 465.3 2.5 4.30E-06 0.00189 MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) CNAP1 70.2 177.5 2.5 4.40E-06 0.00189 chromosome condensation-related SMC- associated protein 1 CBARA1 958 475 0.5 4.60E-06 0.00194 calcium binding atopy-related autoantigen 1 PLA2G4C 123.3 51.2 0.4 4.90E-06 0.00194 phospholipase A2, group IVC (cytosolic, calcium-independent) CPSF1 301.9 616 2.0 5.00E-06 0.00194 cleavage and polyadenylation specific factor 1, 160 kDa SNRPN 30.9 100.6 3.3 5.00E-06 0.00194 Small nuclear ribonucleoprotein polypeptide N RPL5 2754.8 4961 1.8 5.20E-06 0.00194 ribosomal protein L5 C1R 1446.5 366.4 0.3 5.30E-06 0.00194 complement component 1, r subcomponent C16orf34 630.4 1109.6 1.8 5.30E-06 0.00194 chromosome 16 open reading frame 34 PHB 309.3 915.1 3.0 5.30E-06 0.00194 prohibitin BZW2 387.4 946.4 2.4 5.40E-06 0.00194 basic leucine zipper and W2 domains 2 ALAS1 1075.8 466.5 0.4 5.50E-06 0.00194 aminolevulinate, delta-, synthase 1 FLJ20364 48.6 112.4 2.3 5.70E-06 0.00198 hypothetical protein FLJ20364 RANBP1 593.7 1168.1 2.0 5.90E-06 0.00201 RAN binding protein 1 SKB1 354.7 687.4 1.9 6.20E-06 0.00208 SKB1 homolog (S. pombe) ABHD6 402.2 196.9 0.5 6.50E-06 0.00213 abhydrolase domain containing 6 CCNB1 60.4 330 5.5 6.60E-06 0.00213 cyclin B1 NOL5A 246.9 716.2 2.9 7.00E-06 0.00213 nucleolar protein 5A (56 kDa with KKE/D repeat) RPL8 3805.7 7390.5 1.9 7.00E-06 0.00213 ribosomal protein L8 BLNK 211.1 39.8 0.2 7.10E-06 0.00213 B-cell linker BYSL 167.3 269.7 1.6 7.10E-06 0.00213 bystin-like UBE1L 247.6 142.3 0.6 7.20E-06 0.00213 ubiquitin-activating enzyme E1-like CHD7 118.6 312 2.6 7.40E-06 0.00215 chromodomain helicase DNA binding protein 7 DKFZp762E1 70.2 219.4 3.1 7.60E-06 0.00218 hypothetical protein DKFZp762E1312 312 (HJURP) NUP210 178.4 284.9 1.8 7.70E-06 0.00218 nucleoporin 210 kDa PLK1 72.8 185.2 2.5 7.90E-06 0.0022 polo-like kinase 1 (Drosophila) ENPEP 116.2 29.4 0.3 8.00E-06 0.0022 glutamyl aminopeptidase (aminopeptidase A) HCAP-G 17.7 57.8 3.3 8.40E-06 0.00228 chromosome condensation protein G UGT2B4 1117.8 246.7 0.2 9.20E-06 0.00245 UDP glucuronosyltransferase 2 family, polypeptide B4 C20orf27 129.7 245.3 1.9 9.30E-06 0.00245 chromosome 20 open reading frame 27 C6orf149 178.7 491.1 2.7 9.40E-06 0.00245 chromosome 6 open reading frame 149 (LYRM4) The Accession Numbers of the genes of Table 2. as found in NCBI database in June 2008, are the following: IPO4 (BC136759), CPSF1 (NM_013291), MCM4 (NM_005914.2; NM_182746.1; two accession numbers for the same gene correspond to 2 different isoforms of the gene), EIF3S3 (NM_003756.2), NCL (NM_005381.2), CDC25C (NM_001790.3), CENPA (NM_001809.3; NM_001042426.1), K1F14 (BC113742), IPW # (U12897), KNTC2 (AK313184), TMEM48 (NM_018087), BOP1 (NM_015201), EIF3S9 (NM_003751; NM_001037283). PH-4 (NM _177939), SMC4L1 (NM_005496; NM_001002800), TTK (AK315696), LAMA3 (NM_198129), C10orf72 (NM_001031746; NM_144984), TPX2 (NM_012112), MSH2 (NM_000251), DKC1 (NM_001363), STK6 (AY892410), CCT6A (NM_001762; # NM_001009186), SULT1C1 (AK313193), ILF3 (NM_012218; NM_004516), IMPDH2 (NM_000884), HIC2 (NM_015094), AFM (NM_001133), MCM7 (NM_005916; NM_182776), CNAP1(AK128354), CBARA1 (AK225695), PLA2G4C (NM_003706), CPSF1(NM_013291), SNRPN (BC000611), RPL5 (AK314720), C1R (NM_001733), C16orf34 (CH471112), PHB (AK312649), BZW2 (BC017794), ALAS1(AK312566), # FLJ20364 (NM_017785), RANBP1 (NM_002882), SKB1 (AF015913), ABHD6 (NM_020676), CCNB1 (NM_031966), NOL5A (NM_006392), RPL8 (NM_000973; NM_033301), BLNK (NM_013314; NM_001114094), BYSL (NM_004053), UBE1L(AY889910), CHD7 (NM_017780), DKFZp762E1312 (NM_018410), NUP210(NM_024923), PLK1(NM_005030), ENPEP(NM_001977), # HCAP-G(NM_022346), UGT2B4 (NM_021139), C20orf27 (NM_001039140) and C6orf149 (NM_020408).
[0037] In a particular embodiment of the invention, the set of genes of the invention is designed to determine the grade of hepatoblastoma, in particular paediatric hepatoblastoma. In another embodiment, the set of genes is designed to determine the grade of hepatocellular carcinoma, in particular paediatric HCC or adult HCC.
[0038] The expression of the genes of the set may be assayed by any conventional methods, in particular any conventional methods known to measure the quantitative expression of RNA, preferably mRNA.
[0039] The expression may be measured after carrying out an amplification process, such as by PCR, quantitative PCR (qPCR) or real-time PCR. Kits designed for measuring expression after an amplification step are disclosed below.
[0040] The expression may be measured using hybridization method, especially with a step of hybridizing on a solid support, especially an array, a macroarray or a microarray or in other conditions especially in solution. Arrays and kits of the invention, designed for measuring expression by hybridization method are disclosed below.
[0041] The expression of a gene may be assayed in two manners: [0042] to determine absolute gene expression that corresponds to the number of copies of the product of expression of a gene, in particular the number of copies of a nucleotide target, in the sample; and [0043] to determine the relative expression that corresponds to the number of copies of the product of expression of a gene, in particular the number of copies of a nucleotide target, in the sample over the number of copies of the expression product or the number of copies of a nucleotide target of a different gene (calculation also known as normalisation). This different gene is not one of the genes contained in the set to be assayed. This different gene is assayed on the same sample and at the same time as the genes of the set to be assayed, and is called an invariant gene or a normalizer. The invariant gene is generally selected for the fact that its expression is steady whatever the sample to be tested. The expression "steady whatever the sample" means that the expression of an invariant gene does not vary significantly between a normal liver cell and the corresponding tumor cell in a same patient and/or between different liver tumor samples in a same patient. In the present specification, a gene is defined as invariant when its absolute expression does not vary in function of the grade of the liver tumors, in particular does not vary in function of the grade of the HB or HCC tumor, and/or does not vary between liver tumor and normal liver cells.
[0044] In the present invention, the expression which is assayed is preferably the relative expression of each gene, calculated with reference to at least one (preferably 1, 2, 3 or 4) invariant gene(s). Invariant genes, suitable to perform the invention, are genes whose expression is constant whatever the grade of the liver tumors, such as for example ACTG1, EFF1A1, PNN and RHOT2 genes, whose features are summarized in Table 3. In a particular embodiment preferred, the relative expression is calculated with respect to at least the RHOT2 gene or with respect to the RHOT2 gene.
[0045] In another advantageous embodiment, the relative expression is calculated with respect to at least the PNN gene or with respect to the PNN gene. It may be calculated with respect to the RHOT2 and PNN genes.
[0046] The calculation of the absolute expression or of the relative expression of each gene of the set and of each invariant gene being assayed with the same method from the same sample, preferably at the same time, enables to determine for each sample a gene expression profile. TABLE-US-00003 TABLE 3 Features of invariant genes. ACTG1, EEF1A1, PNN and RHOT2 proteins are defined in SEQ ID NOs: 34, 36, 38 and 40 respectively. symbol Gene name SEQ 10* Accession No Location Function ACTG1 actin, gamma 1 33 NM_001614 17q25 cytoplasmic actin cytoskeleton in nonmuscle cells EEF1A1 eukaryotic translation 35 NM13 001 402 6q14.1 enzymatic delivery of elongation factor 1 aminoacyl tRNAs to alpha 1 the ribosome PNN pinin, desmosome 37 NM_002687 14q21.1 transcriptional associated protein corepressor, RNA splicing regulator RHOT2 ras homolog gene 39 NM_138769 16p13.3 Signaling by Rho family, member T2 GTPases, mitochondrial protein
[0047] An additional step of the method or process comprises the determination of the grade of said liver tumor, referring to the gene expression profile that has been assayed. In a particular embodiment of the invention, the method is designed to determine the grade of hepatoblastoma, in particular paediatric hepatoblastoma. In another embodiment, the method is designed to determine the grade of hepatocellular carcinoma, in particular paediatric HCC or adult HCC.
[0048] According to a particular embodiment of the invention, in the step of the method which is performed to determine the grade of the liver tumor, a gene expression profile or a signature (preferably obtained after normalization), which is thus specific for each sample, is compared to the gene expression profile of a reference sample or to the gene expression profiles of each sample of a collection of reference samples (individually tested) whose grade is known, so as to determine the grade of said liver tumor. This comparison step is carried out with at least one prediction algorithm. In a particular embodiment, the comparison step is carried out with 1, 2, 3, 4, 5 or 6 prediction algorithms chosen in the following prediction algorithms: Compound Covariate Predictor (CCP), Linear Discriminator Analysis (LDA), One Nearest Neighbor (1NN), Three Nearest Neighbor (3NN), Nearest Centroid (NC) and Support Vector Machine (SVM). These six algorithms are part of the "Biometric Research Branch (BRB) Tools" developed by the National Cancer Institut (NCI) and are available on http://linus.nci.nih.gov/BRB-ArrayTools.html. Equivalent algorithms may be used instead of or in addition to the above ones. Each algorithm classifies tumors within either of the two groups, defined as tumors with good prognosis (such as C1) or tumors with bad prognosis (such as C2); each group comprises the respective reference samples used for comparison, and one of these two groups also comprises the tumor to be classified.
[0049] Therefore, when 6 algorithms are used, the grade of a tumor sample may be assigned with certainty to the class of good prognosis or to the class of bad prognosis, when 5 or 6 of the above algorithms classified the tumor sample in the same group. In contrast, when less than 5 of the above algorithms classify a tumor sample in the same group, it provides an indication of the grade rather than a definite classification.
[0050] Reference samples which can be used for comparison with the gene expression profile of a tumor to be tested are one or several sample(s) representative for tumor with poor prognosis (such as C2), one or several sample(s) representative of tumor with good prognosis (such as C1), one or several sample(s) of a normal adult liver and/or one or several sample(s) of a fetal liver.
[0051] Table 4 lists the level of expression of each gene of Table 1 depending upon the status of the reference sample i.e., robust tumor with poor prognostic and robust tumor with good prognostic. Examples of methods to identify such robust tumors are provided in the examples. The present invention provides a new classification method in this respect, which is based on discretization of continuous values. TABLE-US-00004 TABLE 4 Level of expression of the genes of Table 1, with respect to the status of the robust tumors Nucleotide Expression status in robust tumor target with poor prognosis with good prognosis AFP overexpressed underexpressed ALDH2 underexpressed overexpressed APCS underexpressed ovorexpressed APOC4 underexpressed overexpressed AQP9 underexpressed overexpressed BUB1 overexpressed underexpressed C1S underexpressed overexpressed CYP2E1 underexpressed overexpressed DLG7 overexpressed underexpressed DUSP9 overexpressed underexpressed E2F5 overexpressed underexpressed GHR underexpressed overexpressed HPD underexpressed overexpressed IGSF1 overexpressed underexpressed NLE1 overexpressed underexpressed RPL10A overexpressed underexpressed
[0052] Reference samples usually correspond to so-called "robust tumor" for which all the marker genes providing the signature are expressed (either under expressed or overexpressed) as expected i.e., in accordance with the results disclosed in Table 5, when tested in similar conditions, as disclosed in the examples hereafter.
[0053] A robust tumor having an overexpression of one or several gene(s) selected among ALDH2, APCS, APOC4, AQP9, C1S, CYP2E1, GHR and HPD genes (these genes belong to the so-called group of differentiation-related genes), and/or an underexpression of one or several gene(s) selected among AFP, BUB1, DLG7, DUSP9, E2F5, IGSF1, NLE1 and RPL10A genes (these genes belong to the so-called group of proliferation-related genes), is an indicator of a robust liver tumor, in particular of a hepatoblastoma, with a good prognosis. A robust tumor having an overexpression of one or several gene(s) selected among AFP, BUB1, DLG7, DUSP9, E2F5, IGSF1, NLE1 and RPL10A genes, and/or an underexpression of one or several gene(s) among ALDH2, APCS, APOC4, AQP9, C1S, CYP2E1, GHR and HPD genes, is an indicator of a robust liver tumor, in particular of a hepatoblastoma, with a poor prognosis. In the present application, a gene is said "underexpressed" when its expression is lower than the expression of the same gene in the other tumor grade, and a gene is said "overexpressed" when its expression is higher than the expression of the same gene in the other tumor grade.
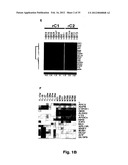
[0054] In a particular embodiment, Table 5 provides the gene expression profiles of the 16 genes of Table 1 in 13 samples of hepatoblastoma (HB) including 8 samples that have been previously identified as rC1 subtype and 5 samples that have been previously identified as rC2 subtype. This Table can therefore be used for comparison, to determine the gene expression profile of a HB tumor to be classified, with the robust tumors disclosed (constituting reference samples), for a set of genes as defined in the present application. Said comparison involves using the classification algorithms which are disclosed herein, for both the selected reference samples and the assayed sample. TABLE-US-00005 TABLE 5 ##STR1## Normalized qPCR data of 16 genes in 13 HB samples including 8 samples of the rC1 subtype and 5 samples of the rC2 subtype (in grey). The qpCR values have been obtained by measuring the expression of the 16 genes in 8 samples of the rC1 subtype and 5 samples of the rC2 subtype by the SYBR green method using the primers as disclosed in Table 6 below and in the conditions reported in the examples, and normalized by the ROTH2 gene (primers in Table 7).
[0055] The method of the present invention is also suitable to classify new tumor samples, and to use them as new reference samples. Therefore, the gene expression values of these new reference samples may be used in combination or in place of some of the values reported in Table 5.
[0056] In another embodiment of the invention, the step of determining the tumor grade comprises performing a method of discretization of continuous values of gene expression obtained on the set of genes the tested patients' samples. Discretization is generally defined as the process of transforming a continuous-valued variable into a discrete one by creating a set of contiguous intervals (or equivalently a set of cutpoints) that spans the range of the variable's values. Discretization has been disclosed for use in classification performance in Lustgarten J. L. et al, 2008.
[0057] The inventors have observed that discretization can be effective in determining liver tumor grade, especially for those tumors described in the present application, including Hepatoblastoma (HB) or Hepatocellular carcinoma (HCC).
[0058] The discretization method is especially disclosed in the examples where it is illustrated by using data obtained on tumor samples wherein these data are those obtained from profiling the 16 genes providing the large set of genes for expression profiling according to the invention. It is pointed out that the discretization method may however be carried out on a reduced number of profiled genes within this group of 16 genes, starting from a set consisting of 2 genes (or more genes) including one (or more) overexpressed proliferation-related genes chosen among AFP, BUB1, DLG7, DUSP9, E2F5, IGSF1, NLE1 and RPL10A and one down-regulated differentiation-related gene chosen among ALDH2, APCS, APOC4, AQP9, C1S, CYP2E1, GHR, HPD, said genes being thus classified as a result of gene profiles observed on robust tumors with poor prognosis (according to the classification in Table 4 above). In particular embodiments of the discretization method, the number of assayed gene for expression profiling is 2, 4, 6, 8, 10, 12, 14 or 16 and the same number of genes in each category (either the group of overexpressed proliferation-related genes or the group of downregulated differentiation-related gene) is used to perform the method.
[0059] The invention thus relates to a method enabling the determination of the tumor grade on a patient's sample, which comprises a classification of the tumor through discretization according to the following steps: [0060] measuring the expression and especially the relative (normalized) expression of each gene in a set of genes defined as the signature of the tumor, for example by quantitative PCR thereby obtaining data as Ct or preferably Delta Ct, wherein said set of genes is divided in two groups, a first group consisting of the proliferation-related genes and a second group consisting of the differentiation-related genes (as disclosed above), [0061] comparing the values measured for each gene, to a cut-off value determined for each gene of the set of genes, and assigning a discretized value to each of said measured values with respect to said cut-off value, said discretized value being advantageously a "1" or a "2" value assigned with respect to the cut-off value of the gene and optionally, if two cut-offs values are used for one gene, a further discretized value such as a "1.5" or another value between "1" or "2" may be assigned for the measured values which are intermediate between the cut-offs values, [0062] determining the average of the discretized values for the genes, in each group of the set of genes, [0063] determining the ratio of the average for the discretized values for the proliferation-related genes on the average for the discretized values for the differentiation-related genes, thereby obtaining a score for the sample, [0064] comparing the obtained score for the sample with one or more sample cut-off(s), wherein each cut-off has been assessed for a selected percentile, [0065] determining the tumor grade as C1 or C2, as a result of the classification of the sample with respect to said sample cut-off.
[0066] The above defined ratio of average values may be alternatively calculated as the ratio of the average for the discresized values for the differentiation-related genes on the average for the discretized values for the proliferation-related genes, to obtain a score. If this calculation made is adopted the cut-offs values are inversed, i.e., are calculated as 1/xxx.
[0067] In order to carry out the discretization method of the invention, the data obtained on the assayed genes for profiling a patient's sample are preferably normalized with respect to one or more invariant gene(s) of the present invention, in order to prevent detrimental impact on the results that may arise from possible inaccuracy in the quantification of initial nucleic acid, especially RNA, in the sample.
[0068] Normalization with respect to one invariant gene only, especially when said invariant gene is RHOT2 gene has proved to be relevant in the results obtained by the inventors. Similarly normalization with respect to PNN gene would be an advantageous possibility because the gene does also not vary in expression.
[0069] In order to design a discretization method for the determination of tumor grade of an individual sample of a patient, according to the invention, cut-offs values have to be determined to allow the determination of the tumor grade. The cut-offs values can be determined experimentally by carrying out the following steps on expression profiling results obtained on a determined number of tumor samples: [0070] defining a cut-off (threshold value) for each gene in the set of genes designed for the signature, said cut-off corresponding to the value of the absolute or preferably relative (i.e. normalized) expression of said gene at a selected percentile and said percentile being selected for each of two groups of genes defined in the set of genes. In order to do so, the set of profiled genes comprises the same number of genes within each of the 2 groups of genes consisting of the group of overexpressed proliferation-related genes encompassing AFP, BUB1, DLG7, DUSP9, E2F5, IGSF1, NLE1 and RPL10A and the group of down-regulated differentiation-related gene encompassing ALDH2, APCS, APOC4, AQP9, C15, CYP2E1, GHR, HPD (said groups being defined based on gene profiles on robust tumors with poor prognosis), [0071] in each tumor sample assigning to each expression value (especially normalized expression value) obtained for each expression profiled gene in the sample, a discretized value which is codified with respect to the cut-off value determined for the same gene and in line with the defined contiguous intervals of continuous values, e.g. a discretized value of "1" or "2" if two intervals (categories) are defined or a discretized value of "1", "1.5" (or another value between 1 and 2) or "2" if three intervals are defined, said assignment of discretized value being advantageously such that the "1" is assigned for expression values falling below the cut-off found for the differentiation-related genes and for expression values falling below the cut-off found for the proliferation-related genes, the "2" is assigned for expression values falling above the cut-off found for the differentiation-related genes and for expression values falling above the cut-off found for the proliferation-related genes, and optionally if a "1.5" is used it is assigned to values found between the cut-offs; [0072] on each tumor sample, determining in each group (proliferation-related genes group or differentiation-related genes group) the average value of said assigned discretized values of profiled genes of the set of profiled genes; [0073] determining a score for each sample, as the ratio between the average expression values of said genes in said two groups of genes in the set of profiled genes; [0074] determining on the basis of the obtained scores for all the tumor samples, one or more cut-off value(s) for the sample, corresponding to the respective value(s) at one or more (especially 2 or 3) percentile(s), wherein said percentile(s) is (are) either identical or different from the percentiles(s) selected for the genes. When the cut-offs values for each gene of the set of genes for profiling have been obtained for a sufficient number of relevant samples and the cut-off value for the sample is determined on the basis of the same samples, these cut-offs can be adopted as reference cut-offs for the user who will be carrying out the analysis of any further patient's tumor sample, especially for the purpose of determining the tumor grade in a patient's sample, if the analysis is performed in identical or similar conditions as the conditions which led to the establishment of the cut-offs values.
[0075] Therefore the invention provides cut-offs values as reference cut-offs, in order to carry out the determination of tumor grade in particular testing conditions as those disclosed below and in the examples.
[0076] In a particular embodiment of the method of discretization, the cut-off for each gene is the value corresponding to a determined percentile, which can be different for each of the considered two groups of genes (proliferation-related genes on the one hand and differentiation-related genes on the other hand). The selected percentile (or quantile) is determined with respect to the fraction of tumors (such as 1/3 or more) harbouring some chosen features such as overexpression of proliferation-related genes and/or dowregulation of differentiation-related genes, in the two groups of genes of the set of genes. Especially, when one intends to assign more weight to tumors displaying strong overexpression of proliferation-related genes and/or strong downregulation of differentiation-related genes, the cut-off corresponds to a high quantile (above the 50th, preferably the 60th, or even above the 65th, such as the 67th and for example within the range of 55th and 70th) for said proliferation-related genes and the cut-off corresponds to a low quantile (below the 50th, preferably equal to or below the 40th for example the 33rd, and for example within the range of between 20th and 40th) of the differentiation-related genes. The cut-off for each group of genes and the cut-off for the sample may be determined with respect to the same percentile(s) or may be determined with respect to different percentile.
[0077] According to a particular embodiment of the invention, for HB tumors, the percentile which is chosen for the overexpressed proliferation-related genes is the 67th and the percentile which is chosen for the downregulated differentiation-related genes is the 33rd. According to a particular embodiment of the invention, for HC tumors, the percentile which is chosen for the overexpressed proliferation-related genes is the 60th and the percentile which is chosen for the downregulated differentiation-related genes is the 40rd.
[0078] Each percentile (or cut-off value corresponding to the percentile) defines a cutpoint and the discretized values for each gene are either "1" or "2" below or above said percentile. The values "1" and "2" are distributed with respect to the percentiles so as to create the highest difference in the values of the calculated ratio for the most different tumor grades. This is illustrated in the examples for the selected percentiles.
[0079] It has been observed that in a preferred embodiment of the invention, the relative values of the profiled genes are determined by real-time PCR (qPCR).
[0080] Conditions to carry out the real-time PCR are disclosed herein, especially in the examples, as conditions applicable to analyzed samples.
[0081] PCR primers and probes suitable for the performance of RT-PCR are those disclosed herein for the various genes.
[0082] In a particular embodiment of the invention, the analysed tumor is a hepatoblastoma and its grade is determined by discretization as disclosed above and illustrated in the examples, taking into account that: [0083] the set of assayed genes for profiling is constituted of the 16 genes disclosed; [0084] the invariant gene (of reference) is RHOT2; [0085] the cut-offs value for each gene based on -dCt (minus delta Ct) measures) are: AFP: 3.96139596; ALDH2: 4.3590482; APCS: 4.4691582; APOC4: 2.03068712; AQP9: 3.38391456; BUB1: -1.41294708; C1S: 4.24839464; CYP2E1: 6.70659644; DLG7: -3.3912188; DUSP9: 2.07022648; E2F5: -0.72728656; GHR: -0.1505569200; HPD: 2.27655628; IGSF1: 0.1075015200; NLE: -0.02343571999; RPL10A: 6.19723876. [0086] the cut-off value for the sample is 0.91 (for the 67th) and optionally a further the cut-off value for the sample is 0.615 (for the 33rd). In such a case, a sample with a score above 0.91 is classified into the C2 class and a sample with a score below 0.91 is classified into the C1 class. The reference to the cut-off at 0.615 may be used to refine the results for values between both cut-offs.
[0087] In another embodiment of the invention, the tumor is an hepatocellular carcinoma and its grade is determined by discretization as disclosed above and illustrated in the examples, taking into account that: [0088] the set of assayed genes for profiling is constituted of the 16 genes disclosed; [0089] the invariant gene (of reference) is RHOT2;
[0090] the cut-offs value for each gene based on -dCt (minus delta Ct) measures) are: TABLE-US-00006 Gene name Cut-off for Taqman Cut-off for SybrGreen AFP -1.2634010 -2.3753035 ALDH2 4.014143 5.314302 APCS 5.6142907 6.399079 APQC4 -0.7963158 4.656336 AQP9 4.2836011 5.446966 BUB1 -1.2736579 -3.634476 C1S 6.3514679 6.240002 CYP2E1 6.9562419 5.829384 DLG7 -2.335694 -4.614352 DUSP9 -7.979559 -1.8626715 E2F5 -0.4400218 -1.367846 GHR 1.0832632 1.169362 HPD 6.7480328 6.736329 IGSF1 -4.8417785 7.6653982 NLE -1.6167268 -1.82226 RPL10A 6.2483056 5.731897
[0091] the cut-off value for the score of a sample based on the ration between the average of the discretized values of the "proliferation-related genes" on the "differentiation-related genes" are 0.66 determined as the 30th percentile of the score) and 0.925 (determined as the 67th percentile of the score) In such a case, a sample with a score above 0.925 is classified into the C2 class and a sample with a score below 0.66 is classified into the C1 class. The sample with a score (initial score) between 0.66 and 0.925 can be assigned to an intermediate class. It can alternatively be classified as C1 or C2 using a modified score corresponding to the average of the discretized values of the "proliferation-related genes". A new cut-off value is determined for said genes, which is the cut-off value for the modified score (in the present case it is 1.3). This cut-off can be determined via a percentile (here the 60th) of the distribution of the modified scores, using the samples of the intermediate class. A sample (initially classified in the intermediate class) with a modified score below 1.3 can be re-classified into the C1 class, and a sample with a modified score above 1.3 can be re-classified into the C2 class.
[0092] It is observed that the refinement of the results which are between the cut-offs of the samples is advantageous for hepatocellular carcinoma in order to increase the relevancy of the information on the tumor grade.
[0093] Generally said refinement of the classification of the intermediate results in the HCC is obtained by performing the following steps:
[0094] a modified score is determined which corresponds to the average of the discretized values of the "proliferation-related genes" only for the sample. A new cut-off value is determined for said genes, which is the cut-off value for the modified score (in the present case it is 1.3). This cut-off can be determined via a percentile (here the 60th) of the distribution of the modified scores, using the samples of the intermediate class. A sample (initially classified in the intermediate class) with a modified score below the "proliferation cut-off" (for example 1.3) can be re-classified into the C1 class, and a sample with a modified score above the "proliferation cut-off" (for example 1.3) can be re-classified into the C2 class.
[0095] From the 16 genes expressed in liver cells listed in Table 1, a set comprising from 2 to 16 genes (or more generally a set as defined herein) may be used to assay the grade of tumor cells in a tumor originating from the liver. The results obtained, after determining the expression of each of the genes of the set, are then treated for classification according to the steps disclosed herein. The invention relates to each and any combination of genes disclosed in Table 1, to provide a set comprising from 2 to 16 of these genes, in particular a set comprising or consisting of 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15 or 16 of these genes. In the designed set, one or many genes of Table 1 may be modified by substitution or by addition of one or several genes as explained above, which also enable to determine the grade of the liver tumor, when assayed in combination with the other genes.
[0096] In a preferred embodiment, the liver tumor is a paediatric HB, and the method or process of the invention enables to distinguish a first class, called C1, qualifying as a good prognosis tumor and a second class, called C2, qualifying as a poor prognosis tumor. The C1 grade is predominantly composed of fetal histotype cells (i.e., well differentiated and non proliferative cells). In contrast, the C2 grade presents cells other than the fetal histotype such as embryonic, atypic (crowded fetal), small cell undifferiantiated (SCUD) and/or macrotrabecular cells.
[0097] The present invention also relates to a kit suitable to determine the grade of a liver tumor from the sample obtained from a patient. This kit is appropriate to carry out the method or process described in the present application.
[0098] In a particular embodiment, the kit comprises a plurality of pairs of primers specific for a set of genes to be assayed, said set comprising from 2 to 16 genes, said 2 to 16 genes being chosen in the group consisting of AFP, ALDH2, APCS, APOC4, AQP9, BUB1, C1S, CYP2E1, DLG7, DUSP9, E2F5, GHR, HPD, IGSF1, NLE1 and RPL10A genes.
[0099] By "plurality", it is mean that the kit comprises at least as many pairs of primers as genes to enable assaying each selected gene, and in particular the nucleotide target of this gene. Accordingly, each gene and in particular its nucleotide target is specifically targeted by a least one of these pairs of primers. In a particular embodiment, the kit comprises the same number of pairs of primers as the number of genes to assay and each primer pair specifically targets one of the genes, and in particular the nucleotide targets of one of these genes, and does not hybridize with the other genes of the set.
[0100] The kits of the invention are defined to amplify the nucleotide targets of the sets of genes as described in the present invention. Therefore, the kit of the invention comprises from 2 to 16 pairs of primers which, when taken as a whole, are specific for said from 2 to 16 genes out of the 16 genes of Table 1. In particular, the kit comprises or consists of 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14 or 15 pairs of primers specific for 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14 or 15 out of the 16 genes of Table 1. In a particular embodiment, the kit comprises or consists of 16 pairs of primers specific for the 16 genes of Table 1 i.e., a primer pair specific for each of the following genes: AFP, ALDH2, APCS, APOC4, AQP9, BUB1, C1S, CYP2E1, DLG7, DUSP9, E2F5, GHR, HPD, IGSF1, NLE1 and RPL10A genes.
[0101] When the set of genes has been modified by the addition or substitution of at least one gene as described above, the kit is adapted to contain a pair of primers specific for each added or substituted gene(s). As indicated by the term "comprises", the kit may, besides the pairs of primers specific for the genes of Table 1, contain additional pair(s) of primers.
[0102] In a particular embodiment, the kit comprises at least one pair of primers (preferably one) for at least one invariant gene (preferably one or two) to be assayed for the determination of the expression profile of the genes, by comparison with the expression profile of the invariant gene.
[0103] The number of pairs of primers of the kit usually does not exceed 100, particularly 50, 30, 20, more particularly 16, and even more particularly is maximum 5, 6, 7, 8, 9 or 10.
[0104] In the kits of the invention, it is understood that, for each gene, at least one pair of primers and preferably exactly one pair, enabling to amplify the nucleotide targets of this gene, is present. When the kits provide several pairs of primers for the same gene, the gene expression level is measured by amplification with only one pair of primers. It is excluded that amplification may be performed using simultaneously several pairs of primers for the same gene.
[0105] As defined herein, a pair of primers consists of a forward polynucleotide and a backward polynucleotide, having the capacity to match its nucleotide target and to amplify, when appropriate conditions and reagents are brought, a nucleotide sequence framed by their complementary sequence, in the sequence of their nucleotide target.
[0106] The pairs of primers present in the kits of the invention are specific for a gene i.e., each pair of primers amplifies the nucleotide targets of one and only one gene among the set. Therefore, it is excluded that a pair of primers specific for a gene amplifies, in a exponential or even in a linear way, the nucleotide targets of another gene and/or other nucleic acids contained in sample. In this way, the sequence of a primer (whose pair is specific for a gene) is selected to be not found in a sequence found in another gene, is not complementary to a sequence found in this another gene and/or is not able to hybridize in amplification conditions as defined in the present application with the sequence of the nucleotide targets of this another gene.
[0107] In a particular embodiment, the forward and/or backward primer(s) may be labelled, either by isotopic (such as radioactive) or non isotopic (such as fluorescent, biotin, fluororochrome) methods. The label of the primer(s) leads to the labelling of the amplicon (product of amplification), since the primers are incorporated in the final product.
[0108] The design of a pair of primers is well known in the art and in particular may be carried out by reference to Sambrook et al. (Molecular Cloning, A laboratory Manual, Third Edition; chapter 8 and in particular pages 8.13 to 8.16). Various softwares are available to design pairs of primers, such as Oligo® or Primer3.
[0109] Therefore, each primer of the pair (forward and backward) has, independently from each other, the following features: [0110] their size is from 10 and 50 bp, preferably 15 to 30 bp; and [0111] they have the capacity to hybridize with the sequence of the nucleotide targets of a gene.
[0112] In a particular embodiment, when the pairs of primers are used in a simultaneous amplification reaction carried out on the sample, the various primers have the capacity to hybridize with their respective nucleotide targets at the same temperature and in the same conditions.
[0113] Conventional conditions for PCR amplification are well known in the art and in particular in Sambrook et al. An example of common conditions for amplification by PCR is dNTP (200 mM), MgCl2 (0.5-3 mM) and primers (100-200 nM).
[0114] In a particular embodiment, the sequence of the primer is 100% identical to one of the strands of the sequence of the nucleotide target to which it must hybridize with, i.e. is 100% complementary to the sequence of the nucleotide target to which it must hybridize. In another embodiment, the identity or complementarity is not 100%, but the similarity is at least 80%, at least 85%, at least 90% or at least 95% with its complementary sequence in the nucleotide target. In a particular embodiment, the primer differs from its counterpart in the sequence of the sequence of the nucleotide target by 1, 2, 3, 4 or 5 mutation(s) (deletion, insertion and/or substitution), preferably by 1, 2, 3, 4 or 5 nucleotide substitutions. In a particular embodiment, the mutations are not located in the last 5 nucleotides of the 3' end of the primer.
[0115] In a particular embodiment, the primer, which is not 100% identical or complementary, keeps the capacity to hybridize with the sequence of the nucleotide target, similarly to the primer that is 100% identical or 100% complementary with the sequence of the nucleotide target (in the hybridization conditions defined herein). In order to be specific, at least one of the primers (having at least 80% similarity as defined above) of the pair specific for a gene can not hybridize with the sequence found in the nucleotide targets of another gene of the set and of another gene of the sample.
[0116] In a particular embodiment, the pairs of primers used for amplifying a particular set of genes are designed, besides some or all of the features explained herein, in order that the amplification products (or amplicons) of each gene have approximately the same size. By "approximately" is meant that the difference of size between the longest amplicon and the shortest amplicon of the set is less than 30% (of the size of the longest amplicon), preferably less than 20%, more preferably less than 10%. As particular embodiments, the size of the amplicon is between 100 and 300 bp, such as about 100, 150, 200, 250 or 300 bp.
[0117] The nucleotide sequences of the 16 genes of Table 1 are provided in the Figures, and may be used to design specific pairs of primers for amplification, in view of the explanations above.
[0118] Examples of primers that may be used to measure the expression of the genes of Table 1, in particular to amplify the nucleotide targets of the genes of Table 1, are the primers having the sequence provided in Table 6 or variant primers having at least 80% similarity (or more as defined above) with the sequences defined in Table 6. TABLE-US-00007 TABLE 6 Sequence of forward and backward primers of the 16 genes defined in Table 1. These primers may be used in any real- time PCR, in particular the SYBR green technique, except for the Taqman ® protocol. Product size Target (bp) Forward primer (5'-3') Reverse primer (5'-3') AFP 151 AACTATTGGCCTGTGGCGAG TCATCCACCACCAAGCTGC ALDH2 151 GTTTGGAGCCCAGTCACCCT GGGAGGAAGCTTGCATGATTC APCS 151 GGCCAGGAATATGAACAAGCC CTTCTCCAGCGGTGTGATCA APOC4 151 GGAGCTGCTGGAGACAGTGG TTTGGATTCGAGGAACCAGG AQP9 151 GCTTCCTCCCTGGGACTGA CAACCAAAGGGCCCACTACA BUB1 152 ACCCCTGAAAAAGTGATGCCT TCATCCTGTTCCAAAAATCCG C1S 141 TTGTTTGGTTCTGTCATCCGC TGGAACACATTTCGGCAGC CYP2E1 151 CAACCAAGAATTTCCTGATCCAG AAGAAACAACTCCATGCGAGC DLG7 151 GCAGGAAGAATGTGCTGAAACA TCCAAGTCTTTGAGAAGGGCC DUSP9 151 CGGAGGCCATTGAGTTCATT ACCAGGTCATAGGCATCGTTG E2F5 151 CCATTCAGGCACCTTCTGGT ACGGGCTTAGATGAACTCGACT GHR 151 CTTGGCACTGGCAGGATCA AGGTGAACGGCACTTGGTG HPD 151 ATCTTCACCAAACCGGTGCA CCATGTTGGTGAGGTTACCCC IGSF1 152 CACTCACACTGAAAAACGCCC GGGTGGAGCAATTGAAAGTCA NLE1 151 ATGTGAAGGCCCAGAAGCTG GAGAACTTCGGGCCGTCTC RPL10A 151 TATCCCCCACATGGACATCG TGCCTTATTTAAACCTGGGCC
[0119] The kit of the invention may further comprise one or many pairs of primers specific for one or many invariant genes, in particular specific for ACTG1, EFF1A1, PNN and/or RHOT2 genes. The pair of primers specific for invariant gene(s) may be designed and selected as explained above for the pair of primers specific for the genes of the set of the invention. In a particular embodiment, the pairs of primers of the invariant genes are designed in order that their amplification product (or amplicon) has approximately the same size as the amplicon of the genes of the set to be assayed (the term approximately being defined as above, with respect to the longest amplicon of the set of genes). Examples of primers that may be used to amplify the particular invariant genes are primers having the sequence provided in Table 7 or primers having at least 80% similarity (or more as defined above) with the sequences defined in Table 7. TABLE-US-00008 TABLE 7 Sequence of forward and backward primers specific for the invariant genes defined in Table 3. These primers may be used in real-time PCR, in particular the SYBR green technique, except for the Taqman ® protocol. Product size Target (bp) Forward primer (5'-3') Reverse primer (5'-3') ACTG1 151 GATGGCCAGGTCATCACCAT ACAGGTCTTTGCGGATGTCC EFF1A1 151 TCACCCGTAAGGATGGCAAT CGGCCAACAGGAACAGTACC PNN 151 CCTTTCTGGTCCTGGTGGAG TGATTCTCTTCTGGTCCGACG RHOT2 151 CTGCGGACTATCTCTCCCCTC AAAAGGCTTTGCAGCTCCAC
[0120] The kits of the invention may also further comprise, in association with or independently of the pairs of primers specific for the invariant gene(s), reagents necessary for the amplification of the nucleotide targets of the sets of the invention and if any, of the nucleotide targets of the invariant genes.
[0121] The kits of the invention may also comprise probes as disclosed herein in the context of sets of probes, compositions and arrays. In particular, the kits also comprise the four dNTPs (nucleotides), amplification buffer, a polymerase (in particular a DNA polymerase, and more particularly a thermostable DNA polymerase) and/or salts necessary for the activity of the polymerase (such as Mg2+).
[0122] Finally, the kits may also comprise one or several control sample(s) i.e., at least one sample(s) representative of tumor with bad (i.e., poor) prognosis (in particular a HB C2 grade), at least one sample(s) representative of tumor with good prognosis (in particular a HB C1 grade), at least one sample of a normal adult liver and/or at least one sample of a fetal liver.
[0123] The kits may also comprise instructions to carry out the amplification step or the various steps of the method of the invention.
[0124] The invention is also directed to a set of probes suitable to determine the grade of a liver tumor from the sample obtained from a patient. This set of probes is appropriate to carry out the method or process described in the present invention. It may also be part of the kit.
[0125] This set of probes comprises a plurality of probes in particular from 2 to 16 probes, these 2 to 16 probes being specific for genes chosen in the group consisting of AFP, ALDH2, APCS, APOC4, AQP9, BUB1, C1S, CYP2E1, DLG7, DUSP9, E2F5, GHR, HPD, IGSF1, NLE1 and RPL10A genes.
[0126] By "plurality", it is mean that the set of probes comprises at least as many probes as genes to assay. In a particular embodiment, the array comprises the same number of probes as the number of genes to assay.
[0127] The probes of the sets of the invention are selected for their capacity to hybridize to the nucleotide targets of the sets of genes as described in the present invention. Therefore, the set of probes of the invention comprise from 2 to 16 probes specific for 2 to 16 genes out of the 16 genes of Table 1. In particular, the sets of probes comprise or consist of 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14 or 15 probes specific of 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14 or 15 out of the 16 genes of Table 1. In a particular embodiment, the sets of probes comprise or consist of 16 probes specific for the 16 genes of Table 1 i.e., a probe specific of each of the following genes: AFP, ALDH2, APCS, APOC4, AQP9, BUB1, C1S, CYP2E1, DLG7, DUSP9, E2F5, GHR, HPD, IGSF1, NLE1 and RPL10A genes.
[0128] The specificity of the probes is defined according to the same parameters as those applying to define specific primers.
[0129] When the set of genes has been modified by the addition or substitution of at least one gene as described above, the set of probes is adapted to contain a probe specific for the added or substituted gene(s). As indicated by the term "comprises", the set of probes may, besides the probes specific for the genes of Table 1, contain additional probe(s).
[0130] The number of probes of the set does usually not exceed 100, particularly 50, 30, 20, more particularly 16, and even more particularly is maximum 5, 6, 7, 8, 9 or 10.
[0131] In the set of probes of the invention, it is understood that for each gene corresponds at least one probe to which the nucleotide target of this gene hybridize to. The set of probes may comprise several probes for the same gene, either probes having the same sequence or probes having different sequences.
[0132] As defined herein, a probe is a polynucleotide, especially DNA, having the capacity to hybridize to the nucleotide target of a gene. Hybridization is usually carried out at a temperature ranging from 40 to 60° C. in hybridization buffer (see example of buffers below). These probes may be oligonucleotides, PCR products or cDNA vectors or purified inserts. The size of each probe is independently to each other from 15 and 1000 bp, preferably 100 to 500 bp or 15 to 500 bp, more preferably 50 to 200 bp or 15 to 100 bp. The design of probes is well known in the art and in particular may be carried out by reference to Sambrook et al. (Molecular Cloning, A laboratory Manual, Third Edition; chapters 9 and 10 and in particular pages 10.1 to 10.10).
[0133] The probes may be optionally labelled, either by isotopic (radioactive) or non isotopic (biotin, fluororochrome) methods. Methods to label probes are disclosed in Sambrook et al. (Molecular Cloning, A laboratory Manual, Third Edition; chapter 8 and in particular page 9.3). In a particular embodiment, the probes are modified to confer them different physicochemical properties (such as by methylation, ethylation). In another particular embodiment, the probes may be modified to add a functional group (such as a thiol group), and optionally immobilized on bead (preferably glass beads).
[0134] In a particular embodiment, the sequence of the probe is 100% identical to a part of one strand of the sequence of the nucleotide target to which it must hybridize, i.e. is 100% complementary to a part of the sequence of the nucleotide target to which it must hybridize. In another embodiment, the identity or complementarity is not 100% and the similarity is at least 80%, at least 85%, at least 90% or at least 95% with a part of the sequence of the nucleotide target. In a particular embodiment, the probe differs from a part of one strand of the sequence of the nucleotide target by 1 to 10 mutation(s) (deletion, insertion and/or substitution), preferably by 1 to 10 nucleotide substitutions. By "a part of", it is meant consecutive nucleotides of the nucleotide target, which correspond to the sequence of the probe.
[0135] In a particular embodiment, the probe, which is not 100% identical or complementary, keeps the capacity to hybridize, in particular to specifically hybridize, to the sequence of the nucleotide target, similarly to the probe which is 100% identical or 100% complementary with the sequence of the nucleotide target (in the hybridization conditions defined herein).
[0136] In a particular embodiment, the size of the probes used to assay a set of genes is approximately the same for all the probes. By "approximately" is meant that the difference of size between the longest probe and the shortest probe of the set is less than 30% (of the size of the longest probe), preferably less than 20%, more preferably less than 10%.
[0137] The set of probes of the invention may further comprise at least one (preferably one) probe specific for at least one invariant gene (preferably one or two), in particular specific for ACTG1, EFF1A1, PNN and/or RHOT2 genes. The probes specific for invariant gene(s) may be designed and selected as explained above for the probes specific for genes of the sets of the invention. In a particular embodiment, the probes specific of the invariant genes have approximately the same size as the probes specific of the genes of the set of be assayed (the term approximately being defined as above, with respect to the longest probes of the set of genes).
[0138] The invention is also directed to an array suitable to determine the grade of a liver tumor from the sample obtained from a patient. This array is appropriate to carry out the method or process described in the present application.
[0139] An array is defined as a solid support on which probes as defined above, are spotted or immobilized. The solid support may be porous or non-porous, and is usually glass slides, silica, nitrocellulose, acrylamide or nylon membranes or filters.
[0140] The arrays of the invention comprise a plurality of probes specific for a set of genes to be assayed. In particular, the array comprises, spotted on it, a set of probes as defined above.
[0141] The invention also relates to a composition comprising a set of probes as defined above in solution.
[0142] In a first embodiment, the probes (as defined above in the set of probes) may be modified to confer them different physicochemical properties (such as methylation, ethylation). The nucleotide targets (as defined herein and prepared from the sample) are linked to particles, preferably magnetic particles, for example covered with ITO (indium tin oxide) or polyimide. The solution of probes is then put in contact with the target nucleotides linked to the particles. The probe/target complexes are then detected, for example by mass spectrometry.
[0143] Alternatively, probes may be modified to add a functional group (such as a thiol group) and immobilized on beads (preferably glass beads). These probes immobilized on beads are put in contact with a sample comprising the nucleotide targets, and the probe/target complexes are detected, for example by capillary reaction.
[0144] The invention is also directed to kits comprising the sets of probes, the compositions or the arrays of the invention and preferably the primer pairs disclosed herein. These kits may also further comprise reagents necessary for the hybridization of the nucleotide targets of the sets of genes and/or of the invariant genes, to the probes (as such, in the compositions or on the arrays) and the washing of the array to remove unbound nucleotides targets.
[0145] In a particular embodiment, the kits also comprise reagents necessary for the hybridization, such as prehybridization buffer (for example containing 5×SSC, 0.1% SDS and 1% bovine serum albumin), hybridization buffer (for example containing 50% formamide, 10×SSC, and 0.2% SDS), low-stringency wash buffer (for example containing 1×SSC and 0.2% SDS) and/or high-stringency wash buffer (for example containing 0.1×SSC and 0.2% SDS).
[0146] The kits may also comprise one or several control sample(s) i.e., at least one sample(s) representative for tumor with poor prognosis, at least one sample(s) representative of tumor with good prognosis, at least one sample of a normal adult liver and/or at least one sample of a fetal liver. Alternatively, it may comprise the representation of a gene expression profile of such tumors.
[0147] Finally, the invention provides a kit as described above further comprising instructions to carry out the method or process of the invention.
[0148] The arrays and/or kits (either comprising pairs of primers or probes or arrays or compositions of the invention or all the components) according to the invention may be used in various aspects, in particular to determine the grade of a liver tumor from a patient, especially by the method disclosed in the present application.
[0149] The arrays and/or kits according to the invention are also useful to determine, depending upon the grade of the liver tumor, the risk for a patient to develop metastasis. Indeed, the classification of a liver tumor in the class with poor prognosis is highly associated with the risk of developing metastasis.
[0150] In another embodiment, the arrays and/or kits according to the invention are also useful to define, depending upon the grade of the liver tumor, the therapeutic regimen to apply to the patient.
[0151] The invention also relates to a support comprising the data identifying the gene expression profile obtained when carrying out the method of the invention.
BRIEF DESCRIPTION OF THE DRAWINGS
[0152] The colour version of the drawings as filed is available upon request to the European Patent Office.
[0153] FIG. 1. Identification of Two HB Subclasses by Expression Profiling.
[0154] (A) Schematic overview of the approach used to identify robust clusters of samples, including two tumor clusters (rC1 and rC2) and one non-tumor cluster (NL) (B) Expression profiles of 982 probe sets (824 genes) that discriminate rC1 and rC2 samples (p<0.001, two-sample t test). Data are plotted as a heatmap where red and green correspond to high and low expression in log2-transformed scale. (C) Molecular classification of 25 HB samples and status of CTNNB1 gene and β-catenin protein. C1 and C2 classification was based on rC1 and rC2 gene signature by using six different statistical predictive methods (CCP, LDA, 1NN, 3NN, NC and SVM) and the leave-one-out cross-validation. Black and gray squares indicate mutations of the CTNNB1 and AXIN1 genes. Immunohistochemical analysis of β-catenin in representative C1 and C2 cases is shown. (D) Expression of representative Wnt-related and β-catenin target genes (p<0.005, two-sample t test) in HB subclasses and non-tumor livers (NL). (E) Classification of hepatoblastoma by expression profile of a 16-gene signature. (F) Classification of normal human livers of children with HB (from 3 months to 6 years of age) (NT) or fetal livers at 17 to 35 weeks of gestation (FL) by expression profile of a 16-gene signature.
[0155] FIG. 2: Molecular HB subclasses are related to liver development stages. (A) Distinctive histologic and immunostaining patterns of HB subclasses C1 and C2. From top to bottom: numbers indicate the ratio of mixed epithelial-mesenchymal tumors and of tumors with predominant fetal histotype in C1 and C2 subtypes; hematoxylin and eosin (H&E) and immunostaining of Ki-67, AFP and GLUL in representative samples. Magnification, ×400. (B) Expression of selected markers of mature hepatocytes and hepatoblast/liver progenitors in HB subclasses and non-tumor livers.
[0156] FIG. 3: Validation of the 16-gene signature by qPCR in an independent set of 41 HBs. Expression profiles of the 16 genes forming the HB classifier are shown as a heatmap that indicates high (red) and low (green) expression according to log2-transformed scale. HB tumors, HB biopsies (b) and human fetal livers (FL) at different weeks (w) of gestation were assigned to class 1 or 2 by using the 16-gene expression profile, six different statistical predictive methods (CCP, LDA, 1NN, 3NN, NC and SVM) and leave-one-out cross-validation. Black boxes in the rows indicate from top to bottom: human fetal liver, mixed epithelial-mesenchymal histology, predominant fetal histotype, and β-catenin mutation.
[0157] FIG. 4: Gene expression of the 16 genes of the prognostic liver cancer signature assessed by qPCR is presented as box-plot. The boxes represent the 25-75 percentile range, the lines the 10-90 percentile range, and the horizontal bars the median values.
[0158] FIG. 5: Expression level of the 16 liver prognostic signature genes shown case by case in 46 hepatoblastomas and 8 normal livers. C1 tumors (green), C2 tumors (red) and normal liver (white).
[0159] FIG. 6. Correlation between molecular HB subtypes and clinical outcome in 61 patients. (A) Association of clinical and pathological data with HB classification in the complete set of 61 patients. Only significant correlations (Chi-square test) are shown. PRETEXT IV stage indicates tumorous involvement of all liver sections. (B) Kaplan-Meier plots of overall survival for 48 patients that received preoperative chemotherapy. Profiling via the 16-gene expression signature was used to define C1 and C2 subclasses in tumors resected after chemotherapy, and differences between survival curves were assessed with the log-rank test. (C) Overall survival of 17 HB patients for which pretreatment biopsies or primary surgery specimens were available. The signature was applied exclusively to tumor samples without prior therapy. (D) Multivariate analysis including 3 variables associated to patient's survival. The predominant histotype is defined as either fetal or other (including embryonal, crowed-fetal, macrotrabecular or SCUD types). Tumor stage is defined by PRETEXT stage (Perilongo et al., 2000) and/or distant metastasis at diagnosis and/or vascular invasion. HR, Hazard Ratio; CI, Confidence Interval.
[0160] FIG. 7: Clinical, pathological and genetic characteristics of 61 HB cases. SR: standard risk; HR: high risk according to SIOPEL criteria; NA: not available; PRETEXT: pre-treatment extent of disease according to SIOPEL; DOD: dead of disease; *: Vascular invasion was defined by radiological analysis; **: The predominant epithelial histotype variable categorized as "others" included embryonal, crowded fetal, macrotrabecular, and undifferentiated histotypes.
[0161] FIG. 8: Clinical, pathological and genetic characteristics of 66 HB samples; Tumor ID number indicates patient number. When more than one sample from the same patient was analyzed, the representative sample used for statistical analysis of clinical correlations is marked by an asterisk; b: biopsy. HB74F: fetal component of HB74; HB74e: embryonal component of HB74. Gender: M, male; F, female; Y, yes; N, no; NA, not available. Multifocality: S, solitary nodules; M, multiple nodules. Histology: E, epithelial; M, mixed; CF, crowded fetal; F, fetal; E, embryonal; M, macrotrabecular; PF, pure fetal; S, SCUD. PRETEXT β-catenin status: wt, wild-type; Δex3, in-frame deletion of part or all exon 3 sequence; FAP, familial polyposis kindred; AXIN1, Axin 1 nonsense mutation (R533stop, CGA to TGA).stage: I to IV according to SIOPEL (Aronson et al., 2005). Treatment protocol: S, standard risk; H, high risk according to SIOPEL. Outcome: A, alive free of disease; DOD, dead of disease; D, death unrelated to cancer; R, alive with recurrence of disease.
[0162] FIG. 9: Correlation between molecular HB subtypes and clinical outcome in 86 patients. (A) Association of clinical and pathological data with HB classification in the complete set of 86 patients. Only significant correlations (Chi-square test) are shown. PRETEXT IV stage indicates tumorous involvement of all liver sections. (B) Kaplan-Meier plots of overall survival for 73 patients that received preoperative chemotherapy. Profiling via the 16-gene expression signature was used to define C1 and C2 subclasses in tumors resected after chemotherapy, and differences between survival curves were assessed with the log-rank test. (C) Overall survival of 29 HB patients for which pretreatment biopsies or primary surgery specimens were available. The signature was applied exclusively to tumor samples without prior therapy. (D) Multivariate analysis including 3 variables associated to patient's survival. The predominant histotype is defined as either fetal or other (including embryonal, crowed-fetal, macrotrabecular or SCUD types). Tumor stage is defined by PRETEXT stage (Perilongo et al., 2000) and/or distant metastasis at diagnosis and/or vascular invasion. HR, Hazard Ratio; CI, Confidence Interval.
[0163] FIG. 10: Correlation between molecular HCC subtypes and clinical outcome in 64 patients. Kaplan-Meier estimates of overall survival in 64 HCC patients using molecular classification with 16 genes, with the unsupervised clustering (centroid) (A) or unsupervised clustering (average) (B).
[0164] FIG. 11: Analysis of the probability of overall survival (OS) of 85 hepatoblastoma patients using Kaplan-Meier estimates. Left pannel: cases were classified by the discretization method into 3 classes using as cut-offs the 33rd percentile and the 67th percentile. Middle pannel: cases were classified into 2 classes using the 33rd percentile. Right pannel: cases were classified into 2 classes using the 67th percentile.
[0165] FIG. 12: Analysis of the probability of overall survival (OS) or disease-free survival (DFS) of 113* HCC patients using Kaplan-Meier estimates and log-rank test. Among the total series of 114 patients, survival data were not available for one case.
Patients were treated either by partial hepatectomy (PH) or by orthotopic liver transplantation (OLT). Unless specified, the follow-up was closed at 146 months.
A: HCC cases were classified into 3 classes by the discretization method using as cut-offs the 33rd and the 67th percentiles.
B: 47 HCC cases previously classified into the intermediate class (33<p<67, see pannel A) were subdivided into 2 new subclasses using the 60th percentile of proliferation-related genes.
C: 92 HCC cases treated by partial hepatectomy (PH) were classified into 3 classes as in pannel A.
D: 21 HCC cases treated by orthotopic liver transplantation (OLT) were classified into 2 classes using as cut-off the 67th percentile.
E: HCC cases were classified into 2 classes using different combinations of scores as described in Table F.
F: HCC cases were classified into 2 classes using as cut-off the 33rd percentile.
G: HCC cases were classified into 2 classes using as cut-off the 50th percentile.
H: HCC cases were classified into 2 classes using as cut-off the 67th percentile.
I: 92 HCC cases treated by partial hepatectomy (PH) were classified into 2 classes using as cut-off the 33rd percentile.
J: 92 HCC cases treated by partial hepatectomy (PH) were classified into 2 classes using as cut-off the 50th percentile.
K: 92 HCC cases treated by partial hepatectomy (PH) were classified into 2 classes using as cut-off the 67th percentile.
L: Disease-free survival of 113 HCC cases after classification into 2 classes using as cut-off the 67th percentile. Follow-up was closed at 48 months. Data were not significant when the follow-up was closed at 146 months.
M: Disease-free survival of 92 HCC cases treated by PH, after classification into 2 classes using as cut-off the 67th percentile. Follow-up was closed at 48 months. Data were not significant when the follow-up was closed at 146 months.
[0166] FIG. 13: Analysis of the probability of overall survival (OS) or disease-free survival (DFS) HCC patients using Kaplan-Meier estimates and log-rank test.
EXAMPLES
Experimental Procedures
[0167] A. Patients and Tissue Samples.
[0168] Sixty-six tumor specimens and biopsies from 61 patients with hepatoblastoma were collected from different hospitals in France (52 cases), Italy (6 cases), United Kingdom (1 case), Switzerland (1 case) and Slovakia (1 case). Forty-eight patients received chemotherapy treatment prior to surgery, most being enrolled in clinical trials of the International Childhood Liver Tumour Strategy Group (SIOPEL) (Perilongo et al., 2000). Samples from fresh tumors avoiding fibrotic and necrotic areas and from adjacent non tumor livers were snap frozen at the time of surgery and stored at -80° C. FIG. 7 describes patient characteristics and clinicopathological parameters.
[0169] Patients were children with median age of 2 years, and male:female ratio of 1.5. The median follow-up was 32 months; during this period, 15 patients died from disease. The histology of all tumor specimens was centrally reviewed by expert pathologist according to previously described criteria (Finegold et al., 2007; Zimmermann, 2005). Twenty-five tumors were analyzed on oligonucleotide microarrays and 24 of them, for which DNA was available, were subjected to aCGH analysis, while a second set of 41 tumors was analyzed by qPCR (FIG. 8). No difference was observed in significant clinical and pathological data as well as in the percentage of cases carrying β-catenin mutation between the two sets. This study has been approved by the Ethics Committee of Institut Pasteur, and informed consent of the families was obtained at each Medical Center, in accordance with European Guidelines for biomedical research and with national laws in each country.
[0170] B. Oligonucleotide Microarrays and Gene Expression Data Analysis
[0171] Twenty-five HB samples and 4 non-tumor samples including a pool of livers from 3 males and a second from 3 females were analyzed using Affymetrix HG-U133A oligonucleotide arrays. Total RNA was prepared using FastPrep® system (Qbiogene, Strasbourg, France) and RNeasy mini Kit (Qiagen, Courtaboeuf, France). RNA quality was checked with the Agilent 2100 Bioanalyzer (Agilent Technologies, Palo Alto, Calif.). Microarray experiments were performed according to the manufacturer's instructions. Affymetrix microarray data were normalized using RMA method (Irizarry et al., 2003). Class discovery was done as described elsewhere (Lamant et al., 2007). Pathway and Gene Ontology enrichment analyses were performed using GSEA method (Subramanian et al., 2005) and hypergeometric tests. For supervised tests and class prediction, we used Biometric Research Branch (BRB) ArrayTools v3.2.2 software, developed by R. Simon and A. Peng. Permutations of the measurements are then used to estimate the FDR (the percentage of genes identified by chance). Additionally, mouse fetal livers at E18.5 and postnatal livers at 8 days of birth were profiled on Affymetrix MG-U74A, B v2 arrays. Data were processed and analyzed as aforementioned.
[0172] Except when indicated, transcriptome analysis was carried out using either an assortment of R system software packages (http://www.R-project.org, v2.3.0) including those of Bioconductor v1.8 (Gentleman et al., 2004) or original R code.
[0173] B.1. Normalization
[0174] Raw data from Affymetrix HG-U133A 2.0 GeneChip® microarrays were normalized in batch using robust multi-array average method (R package affy, v1.10.0) (Irizarry et al., 2003). Probe sets corresponding to control genes or having a "_x_" annotation were masked yielding a total of 19,787 probe sets available for further analyses.
[0175] B.2. Class Discovery
Step 1
Variance Test
[0176] The variance of each probe set across samples was tested and compared to the median variance of all the probe sets, using the model: ((n-1)×Var(probe set)/Varmed), where n refers to the number of samples. By using the same filtering tool of BRB ArrayTools software, the P-value for each probe set was obtained by comparison of this model to a percentile of Chi-square distribution with (n-1) degrees of freedom.
Robust Coefficient of Variation (rCV)
The rCV was calculated for each probe set as follows. After ordering the intensity values of n samples from min to max, we eliminated the min and max values and we calculated the coefficient of variation (CV) for the remaining values.
Unsupervised Probe Sets Selection
Unsupervised selection of probe set lists was based on the two following criteria:
(i) variance test at P<0.01,
(ii) rCV less than 10 and superior to a given rCV percentile. We used eight rCV percentile thresholds (60%; 70%; 80%; 90%; 95%; 97.5%; 99%; 99.5%), which yielded 8 probe set lists.
Step 2: Generation of a Series of 24 Dendrograms
[0177] Hierarchical clustering was performed by using the 8 rCV-ranked probe sets lists, 3 different linkage methods (average, complete and Ward's), and 1-Pearson correlation as a distance metric (package cluster v1.9.3). This analysis generated 24 dendrograms.
Step 3:
Stability Assessment
[0178] The intrinsic stability of each of the 24 dendrograms was assessed by comparing each dendrogram to the dendrograms obtained after data "perturbation" or "resampling" (100 iterations). Perturbation stands for the addition of random gaussian noise (μ=0, σ=1.5×median variance calculated from the data set) to the data matrix, and resampling for the random substitution of 5% of the samples by virtual sample's profiles, generated randomly. The comparison between dendrograms across all iterations yielded a mean `similarity score` (see below). The overall stability was assessed by calculating a mean similarity score, using all pairs of the 24 dendrograms.
Similarity Score
[0179] To compare two dendrograms, we compared the two partitions in k clusters (k=2 to 8) obtained from these two dendrograms. To compare a pair of partitions, we used a similarity measure, which corresponds to the symmetric difference distance (Robinson and Foulds, 1981).
Step 4: Identification of Robust Clusters
[0180] We identified groups in which any pair of samples was co-classified in at least 22 of the 24 partitions, and considered only groups made of 4 samples or more. Then, for any pair of these groups, we calculated the mean number of co-classification of any sample in the first group with any sample in the second group. We aggregated the groups for which this score was at least 18 (over the 24 partitions).
[0181] B.3. Supervised Tests
[0182] We compared gene expression between two classes of samples by using the Student's t test with random variance model option (BRB ArrayTools software, version 3.4.0a, developed by Dr. Richard Simon and Amy Peng Lam, http://linus.nci.nih.gov/BRB-ArrayTools.html). False Discovery Rates were assessed by using 1000 random permutations of labels (Monte Carlo approach).
[0183] B.4. Classification
[0184] To classify samples according to gene expression profile, we used the Class prediction tool of BRB ArrayTools software using all 6 following algorithms: Compound Covariate Predictor (CCP), Linear Discriminant Analysis (LDA), 1-Nearest Neighbor (1NN), 3-Nearest Neighbors (3NN), Nearest Centroid (NC) and Support Vector Machines (SVM). Each sample was classified according to the majority of the 6 algorithms. Samples classified as C2 by at least 3 algorithms were classified accordingly.
[0185] B.5. Gene Ontology and Pathway Analysis
[0186] We used a hypergeometric test to measure the association between a gene (probe set) list and a gene ontology term (GO term), as in GO stats R package (R. Gentleman). To this end, we mapped the gene list and the GO terms to non-redundant Entrez Gene identifiers by using the annotation file HG-U133_Plus--2.annot.csv (http://www.affymetrix.com, Dec. 14, 2006). GO terms and their relationships (parent/child) were downloaded from http://www.geneontology.org (version Dec. 31, 2006). The list of proteins associated to GO terms (table gene_association.goa_human) and mapping the Entrez Gene ids (table human.xrefs) were downloaded from ftp://ftp.ebi.ac.uk/pub/databases/GO/goa.
[0187] KEGG pathway annotation was done by Onto-tools software (http://vortex.cs.wayne.edu/ontoexpress/servlet/UserInfo). We designated a significance threshold of each hypergeometric test at P<0.001, and the condition that a GO term or pathway be represented by at least 3 Entrez Gene identifiers.
[0188] B.6. Gene Set Enrichment Analysis (gsea)
[0189] GSEA (Subramanian et al., 2005) was used to evaluate the correlation of a specific gene list with two different sample groups (phenotypes). Briefly, this method calculates an enrichment score after ranking all genes in the dataset based on their correlation with a chosen phenotype and identifying the rank positions of all the members of a defined gene set. We used the signal2noise ratio as a statistic to compare specific and random phenotypes in order to evaluate statistical differences.
[0190] C. Array-Based Comparative Genomic Hybridization (aCGH)
[0191] Genomic DNA from 24 HBs and 3 non-tumor liver samples was analyzed using aCGH chips designed by the CIT-CGH consortium. This array contains 3400 sequence-verified PAC/BAC clones spaced at approximately 1 Mb intervals, spotted in triplicate on Ultra Gaps slides (Corning Inc, Corning, N.Y.).
[0192] The aCGH chip was designed by CIT-CGH consortium (Olivier Delattre laboratory, Curie Institute, Paris; Charles Theillet laboratory, CRLC Val d'Aurelle, Montpellier; Stanislas du Manoir laboratory, IGBMC, Strasbourg and the company IntegraGen®). DNAs were labeled by the random priming method (Bioprime DNA labelling system; Invitrogen, Cergy-Pontoise, France) with cyanine-5 (Perkin-Elmer, Wellesley, Mass.). Using the same procedure, we labeled control DNAs with cyanine-3. After ethanol-precipitation with 210 μg of Human Cot-1 DNA (Invitrogen), resuspension in hybridization buffer (50% formamide), denaturation at 95° C. for 10 minutes and prehybridization at 37° C. for 90 minutes, probes were cohybridized on aCGH. The aCGH slides were previously preblocked with a buffer containing 2.6 mg succinic anhydride/118 ml N-methyl-2-pyrrolidinone/32 ml sodium tetraborate decahydrate, pH 8.0 (Sigma-Aldrich, Lyon, France). After washing, arrays were scanned using a 4000B scan (Axon, Union City, Calif.). Image analysis was performed with Genepix 5.1 software (Axon) and ratios of Cy5/Cy3 signals were determined. The aCGH data were normalized using lowess per block method (Dudoit et al., 2002). Comparison between groups was done using chi-square test or Fisher's exact test, as appropriate.
[0193] Status assignment (Gain/Loss) was performed using R package GLAD v1.6.0. Computation of recurrent minimal genomic alterations was done using slight modification of a previously described method (Rouveirol et al., 2006). For comparison between groups, we used the Fischer exact test. Complete aCGH data will be published elsewhere.
[0194] D. Mouse Microarray Analysis
[0195] Murine Genome Affymetrix U74v2 A and B arrays were used to investigate liver expression at embryonic day 18.5 (E18.5) and at 8 days after birth (PN8). Each time point consisted of a pool of livers from 3-5 animals analyzed in triplicate. Microarray experiments were performed according to the manufacturer's instructions.
[0196] Publicly available Affymetrix Mouse Genome (MG) 430 2.0 array liver expression data at embryonic time points E11.5, E12.5, E13.5, E14.5, and E16.5 days of gestation (Otu et al., 2007), were downloaded from the Gene Expression Omnibus (GEO) database (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE6998).
[0197] MG-U74v2, MG-430 2.0 and HG-133A 2.0 array intra- and cross-species probeset comparison was achieved by using the Affymetrix NetAffx analysis center and by choosing "Good Match" degree of specificity. Unification of sample replicates, multiple array data standardization and Heatmap visualization was done by using dCHIP v1.6 software. Comparison of fetal liver stages by supervised analysis was performed using BRB ArrayTools software as previously described, by classing E11.5 and E12.5 as "Early" and E14.5 and E16.5 as "Late" fetal liver stage. Supervised signature was applied to HB array data, and intensity cut-off=60 was chosen in order to remove probesets that did not reach such intensity level in at least one sample.
[0198] E. Quantitative PCR Analysis (qPCR)
[0199] For qPCR analysis, we used RNA from 52 tumor samples (including 11 samples analyzed on microarrays, see FIG. 8), and from 8 non-tumor livers and 5 human fetal livers (RNAs purchased from BioChain Institute, Hayward, Calif.).
[0200] RNA was extracted by using either Trizol, RNeasy kit (QIAGEN) or miRvana kit (Ambion), then quantified and quality-checked by Agilent technology. For each cDNA preparation, 1 μg of RNA was diluted at the final concentration of 100 ng/μl, and reverse transcribed with the Superscript RT kit (Invitrogen, Carlsbad, Calif.) following the manufacturer's protocol. Random primers (Promega, Charbonnieres-les-Bains, France) were added at the final concentration of 30 ng/μl and the final volume was 20 μl.
[0201] The cDNA was diluted 1:25, and 5 μl were used for each qPCR reaction. We added 5 μl of 2×Sybr Green Master mix (Applied Biosystems) and 0.3 μl of each specific primer (final concentration 300 nM). Each reaction was performed in triplicate. qPCR reactions were run on the Applied Biosystems 7900HT Fast Real-Time PCR System with a 384-well thermo-block, in the following conditions: 2 min at 50° C. to activate Uracil-N-glycosylase (UNG)-mediated erase of aspecific reaction; 10 min at 95° C. to activate the polymerase and inactivate the UNG; 40 cycles (15 sec at 95° C. denaturation step and 1 min at 60° C. annealing and extension); and final dissociation step to verify amplicon specificity.
[0202] The lists of primers used for qPCR are provided in Table 6 and Table 7 above.
[0203] F. Immunohistochemistry (IHC)
[0204] IHC was carried out as reported previously (Wei et al., 2000). For antigen retrieval at 95° C., we used 1 mM EDTA (pH 8) for β-catenin and Ki-67 IHC, and 10 mM citrate buffer (pH 6) for AFP and GLUL IHC. We used monoclonal antibodies against β-catenin and GLUL (Cat. Nos. 610154 and 610517; BD Biosciences, Le Pont de Claix, France) and Ki-67 (M7240, Dako, Trappes, France) and polyclonal antibody against AFP (N1501, Dako). Reactions were visualized using the ChemMate Dako Envision Detection kit (Dako) and diaminobenzidine. Subcellular distribution and quantitative evaluation of immunostaining in the different histotypes were assessed by examining at least ten random high-power fields.
[0205] G. Clinical Data Analysis
[0206] We used the Chi-square test for comparisons between groups. Survival curves were calculated according to the Kaplan-Meier method, using the log-rank test to assess differences between curves. Variables independently related to survival were determined by stepwise forward Cox regression analysis. Follow-up was closed at February 2007 or at time of death. Statistical analysis was done with SPSS software v10.0 (SPSS Inc., Chicago, Ill.).
[0207] H. Examples of Other Pairs of Primers and Probes for the 16 Genes of Table 1 and the 4 Invariant Genes (Table 3) that can be Used in the Taqman® Method. TABLE-US-00009 AFP forward primer: GCCAGTGCTGCACTTCTTCA AFP reverse primer: TGTTTCATCCACCACCAAGCT AFP probe: ATGCCAACAGGAGGCCATGCTTCA (for each polynucleotide, the sequence is given from 5' to 3') ALDH2 forward primer: TGCAGGATGGCATGACCAT ALDH2 reverse primer: TCTTGAACTTCAGGATCTGCATCA ALDH2 probe: CCAAGGAGGAGATCTTCGGGCCA APCS forward primer: AGCTGGGAGTCCTCATCAGGTA APCS reverse primer: CGCAGACCCTTTTTCACCAA APCS probe: TGCTGAATTTTGGATCAATGGGACACC APOC4 forward primer: TGAAGGAGCTGCTGGAGACA APOC4 reverse primer: CGGGCTCCAGAACCATTG APOC4 probe: TGGTGAACAGGACCAGAGACGGGTG AQP9 forward primer: GCCATCGGCCTCCTGATTA AQP9 reverse primer: GTTCATGGCACAGCCACTGT AQP9 probe: TGTCATTGCTTCCTCCCTGGGACTG BUB1 forward primer: ACATCTGGTTTTCAGTGTGTTGAGA BUB1 reverse primer: GTTGCAGCAACCCCAAAGTAA BUB1 probe: TCAGCAACAAACCATGGAACTACCA GATCG C1S forward primer: TCCCAATGACAAGACCAAATTCT C1S reverse primer: AGAGCCCATAGGTCCCACACT C1S probe: CGCAGCTGGCCTGGTGTCCTG CYP2E1 forward CATGAGATTCAGCGGTTCATCA primer: CYP2E1 reverse GGTGTCTCGGGTTGCTTCA primer: CYP2E1 probe: CCTCGTGCCCTCCAACCTGCC DLG7 forward primer: GCTGGAGAGGAGACATCAAGAAC DLG7 reverse primer: CCTGGTTGTAGAGGTGAAAAAGTAATC DLG7 probe: TGCCAGACACATTTCTTTTGGTGGTAA CC DUSP9 forward primer: GGCCTACCTCATGCAGAAGCT DUSP9 reverse primer: GGGAGATGTTAGACTTCTTCCTCTTG DUSP9 probe: CACCTCTCTCTCAACGATGCCTATGA CCTG E2F5 forward primer: CCTGTTCCCCCACCTGATG E2F5 reverse primer: TTTCTGTGGAGTCACTGGAGTCA E2F5 probe: CCTCACACAGCCTTCCTCCCAGTCC GHR forward primer: CCCAGGTGAGCGACATTACA GHR reverse primer: CATCCCTGCCTTATTCTTTTGG GHR probe: CAGCAGGTAGTGTGGTCCTTTCCCCG HPD forward primer: CCCACGCTCTTCCTGGAA HPD reverse primer: TTGCCGGCTCCAAAACC HPD probe: TCATCCAGGGCCACAACCACCA IGSF1 forward primer: GACCATTGCCCTTGAAGAGTGT IGSF1 reverse primer: GAGAGGTTGATGAAGGAGAATTGG IGSF1 probe: ACCAAGAAGGAGAACCAGGCACCCC NLE1 forward primer: TGCCTCCTTTGACAAGTCCAT NLE1 reverse primer: CGCGTAGGGAAGCCAGGTA NLE1 probe: TGGGATGGGAGGAGGGGCA RPL10A forward primer TCGGCCCAGGTTTAAATAAGG RPL10A reverse primer CCACTTTGGCCACCATGTTT RPL10A Taqman probe AGTTCCCTTCCCTGCTCACACACAACG ACTG1 forward primer: GGCGCCCAGCACCAT ACTG1 reverse primer: CCGATCCACACCGAGTACTTG ACTG1 probe: ATCAAGATCATCGCACCCCCAGAGG EEF1A1 forward GCGGTGGGTGTCATCAAAG primer: EEF1A11 reverse TGGGCAGACTTGGTGACCTT primer: EEF1A11 probe: AGTGGACAAGAAGGCTGCTGGAGCTG PNN forward primer: GAATTCCCGGTCCGACAGA PNN reverse primer: TTTCGGTCTCTTTCACTTCTTGAA PNN probe: AGAGGTCTATATCAGAGAGTAGTCGA TCAGGCAAAAGA RHQT2 forward primer: CCCAGCACCACGATCTTCAC RHOT2 reverse primer: CCAGAAGGAAGAGGGATGCA RHOT2 Taqman probe: CAGCTCGCCACCATGGCCG
Results
[0208] Identification of Two HB Subclasses by Gene Expression Profiling
[0209] For robust unsupervised classification, we generated and screened a series of 24 dendrograms to identify samples that co-clustered whatever the method and the gene list. We obtained two robust subgroups of tumors named robust Cluster 1 (rC1, n=8) and robust Cluster 2 (rC2, n=5) (FIG. 1A). Comparison of rC1 and rC2 expression profiles identified 824 genes (p<0.001, false discovery rate (FDR)=0.02) (FIG. 1B). KEGG pathway analysis pinpointed a strong enrichment of cell cycle related genes (p<10-11), most being up-regulated in rC2 tumors. These: genes were mainly assigned to GO categories including mitosis regulation, spindle checkpoint, nucleotide biosynthesis, RNA helicase activity, ribosome biogenesis, and translational regulation. Evidence that rC2 tumors were faster proliferating than rC1 tumors was further confirmed by Ki-67 immunostaining (see FIG. 2A).
[0210] The remaining tumors were classified into C1 (rC1-related) and C2 (rC2-related) subclasses by applying a predictive approach based on the rC1/rC2 gene signature and using robust samples as training set (FIG. 1C). Both groups exhibited similar, high rates of β-catenin mutations, and accordingly, immunohistochemistry (IHC) of β-catenin showed cytoplasmic and nuclear staining of the protein in the majority of HBs. However, β-catenin localization was predominantly membranous and cytoplasmic in C1 tumors, whereas it showed frequent loss of membrane anchoring and intense nuclear accumulation in C2 tumors (FIG. 1C).
[0211] We observed differential expression of a number of Wnt members and targets between subclasses. C2 tumors showed increased expression of MYCN, BIRC5 that encodes the anti-apoptotic factor Survivin, NPM1 (encoding nucleophosmin) and HDAC2. By contrast, most C1 tumors prominently expressed the Wnt antagonist DKK3, BMP4, and genes previously found to be activated in liver tumors carrying mutant β-catenin (Boyault et al., 2007; Renard et al., 2007; Stahl et al., 2005). Remarkably, most genes related to liver functions are expressed in the perivenous area of adult livers, such as GLUL, RHBG, and two members of the cytochrome p450 family: CYP2E1 and CYP1A1 (Benhamouche et al., 2006; Braeuning et al., 2006) (FIG. 1D).
[0212] Further evidence that the rC1 subclass was enriched in genes assigned to the hepatic perivenous program was provided by Gene Set Enrichment Analysis (GSEA), a computational method for assessing enrichment of a predefined gene list in one class as compared with another (Subramanian et al., 2005). Thus, Wnt/β-catenin signaling appears to activate different transcriptional programs in HB subtypes, likely reflecting different cellular contexts.
[0213] HB Subclasses Evoke Distinct Phases of Liver Development
[0214] Next, we sought to determine whether HB subclasses were associated with specific histological phenotypes. Mixed epithelial-mesenchymal tumors that represented 20% of cases were not significantly associated with C1 and C2 subclasses. By contrast, a tight association was found with the main epithelial component, which defines the cell type occupying more than 50% of tumor cross-sectional areas. Sixteen out of 18 C1 tumors displayed a predominant fetal phenotype, including 4 `pure fetal` cases, whereas all C2 tumors showed a more immature pattern, with prevailing embryonal or crowded-fetal histotypes associated with high proliferation (Finegold, 1994) (p<0.0001) (FIG. 2A). Further relationship between molecular subclasses and hepatic developmental stages was provided by the finding that a number of mature hepatocyte markers were markedly downregulated in C2 compared to C1 tumors (Tables 1 and 2). Conversely, C2 tumors showed strong overexpression (35-fold) of the oncofetal AFP gene associated to high protein levels in tumor cells by IHC (FIG. 2A) and in patients' sera (r=0.79, p<0.0001). C2 tumors also abundantly expressed hepatic progenitor markers such as KRT19 (encoding cytokeratin 19) and TACSTD1, also known as Ep-CAM (FIG. 2B).
[0215] To better define the relationships between HB subclasses and phases of hepatic differentiation, we first generated a liver development-related gene signature by making use of publicly available mouse fetal and adult liver data sets (Otu et al., 2007). When applied to HB samples, this signature was able to distinguish by hierarchical clustering two HB groups closely matching the C1/C2 classification. Next, we integrated HB gene expression data with the orthologous genes expressed in mouse livers at embryonic days (E) 11.5 to 18.5, and at 8 days of birth. In unsupervised clustering, most C2 tumors co-clustered with mouse livers at early stages of embryonic development (E11.5 and E12.5), whereas C1 tumors gathered with mouse livers at late fetal and postnatal stages. Together, these data comfort the notion that tumor cells in C2 and C1 subtypes are arrested at different points of the hepatic differentiation program.
[0216] Identification of a 16-Gene Signature as HB Classifier
[0217] To investigate the relevance of molecular HB classification in an independent set of tumors, we defined a HB classifier signature derived from the top list of genes differentially expressed between rC1 and rC2 clusters. After qPCR assessment, a list of 16 top genes at p≦10-7 was selected to form a class predictor (Table 1). Most of these genes show drastic variations in expression level during liver development, and among them, BUB1 and DLG7 have been repeatedly identified as hESC markers (Assou et al., 2007). The 16-gene expression profile was first investigated in rC1 and rC2 samples used as training set, and it predicted classification with 100% of accuracy in these samples, using either microarray or qPCR data. The robustness of this signature was confirmed by correct classification into C1 and C2 subclasses of all 13 remaining tumors analyzed by microarray (FIG. 1E). Expression profiles of fetal livers and normal liver for these 16-gene signature were also assayed (FIG. 1F). This signature was therefore employed to classify a new, independent set of 41 HB samples by qPCR (FIGS. 4 and 5 and Table 8), resulting in 21 tumors categorized as C1 and 20 tumors as C2 subtype (FIG. 3).
[0218] Extending our previous observation, C1/C2 classification in this new set of tumors was unrelated to CTNNB1 mutation rate. Using qPCR, we also confirmed enhanced expression in C2 tumors of liver progenitor markers such as AFP, Ep-CAM, and KRT19, as well as MYCN (FIG. 3). Moreover, while a similar percentage of C1 and C2 tumors displayed mesenchymal components, a predominant fetal histotype was found in 95% of tumors of the C1 subtype, whereas in 82% of C2 tumors, the major component displayed less differentiated patterns such as embryonal, crowded-fetal, macrotrabecular and SCUD types (p<0.0001) (FIG. 3). To further assess the association of HB subclasses with liver development, 5 human fetal livers at different weeks of gestation were included in the qPCR studies. In unsupervised clustering, fetal livers at late (>35 weeks) and earlier (17 to 26 weeks) developmental stages were classified as C1 and C2 respectively, further supporting that HB subclasses reflect maturation arrest at different developmental phases. TABLE-US-00010 TABLE 8 Gene expression of the prognostic signature for liver cancer by quantitative RT-PCR. C1 C2 NL Fold-change median min max median min max median min max C1/NL C2/NL C2/C1 C1/C2 AFP 0.4 0.0 33.3 30.7 0.0 456.1 0.2 0.0 8.8 2.3 38.1 16.5 0.1 ALDH2 87.1 13.2 356.7 15.0 2.2 74.4 240.4 151.6 387.6 0.3 0.1 0.2 5.2 APCS 61.6 1.1 338.9 1.9 0.0 276.2 158.6 92.7 509.5 0.2 0.0 0.1 19.8 APOC4 21.3 4.3 122.8 1.6 0.1 24.2 47.0 22.3 112.4 0.5 0.0 0.1 16.1 AQP9 60.6 8.0 540.6 2.5 0.1 90.1 46.6 38.0 72.7 1.3 0.1 0.1 18.9 BUB1 0.0 0.0 0.4 0.9 0.1 3.9 0.0 0.0 0.1 1.2 16.1 13.4 0.1 C1S 51.1 14.9 277.2 7.5 1.3 96.0 223.4 129.3 565.3 0.2 0.0 0.2 5.7 CYP2E1 583.2 97.7 3463.0 19.7 0.4 1504.0 1128.6 527.6 1697.0 0.7 0.0 0.0 51.6 DLG7 0.0 0.0 0.0 0.1 0.0 0.5 0.0 0.0 0.0 1.7 12.4 7.3 0.1 DUSP9 1.5 0.4 45.7 19.1 0.0 179.0 0.6 0.2 1.3 4.0 18.3 4.6 0.2 E2F5 0.2 0.0 2.0 1.1 0.1 11.7 0.1 0.0 0.5 1.8 6.5 3.5 0.3 GHR 5.2 0.0 54.0 0.5 0.0 2.4 35.2 20.8 54.5 0.1 0.0 0.1 8.6 HPD 22.9 0.9 182.0 1.2 0.1 23.8 111.5 62.6 165.7 0.2 0.0 0.1 14.0 IGSF1 0.1 0.0 1.7 1.7 0.0 19.8 0.1 0.0 0.1 2.2 22.4 10.2 0.1 NLE 0.4 0.1 4.8 0.8 0.3 5.1 0.4 0.2 0.8 1.2 2.2 1.8 0.5 RPL10A 73.3 12.0 230.4 98.2 11.9 432.8 86.9 54.1 159.9 0.8 1.1 1.5 0.7 NL, non-tumor liver; C1, good prognosis hepatoblastomas; C2, bad prognosis hepatoblastomas. Shown are the median values of 46 hepatoblastomas from 41 patients, the minimal and maximal values in each class, and the fold changes between classes. Data are presented in arbitrary units after normalization of the raw quantitative PCR values with genes (ACTG1, # EFF1A1, PNN and RHOT2) that presents highly similar values in all samples. Gene expression of the 16 genes are presented on FIGS. 4 and 5.
[0219] The 16-Gene Signature as a Strong Independent Prognostic Factor
[0220] In a First Set of 61 Patients
[0221] The clinical impact of HB molecular classification was addressed in a first set of 61 patients (FIGS. 7 and 8), comprising 37 (61%) C1 and 24 (39%) C2 cases. Besides strong association with predominant immature histotypes, HBs of the C2 subclass were tightly associated with features of advanced tumor stage, such as vascular invasion and extrahepatic metastasis (FIG. 6A). Accordingly, overall survival of these patients was markedly impaired. Kaplan-Meier estimates of overall survival probability at 2-years were 50% for patients with C2 tumors and 90% for patients with C1 tumors (p=0.0001, log rank test), and similar trends were seen for disease-free survival probabilities (data not shown). Next, we examined whether pre-operative chemotherapy treatment given to 48 patients could affect tumor classification. These cases were evenly distributed among HB subclasses, with no significant association with molecular classification. Of note, available pretreatment biopsies were assigned to the same subclass as matched resected tumors in 3 out of 4 cases (see FIG. 3; HB112 and HB112b have been both classified as C1 grade, and HB114 and HB114b have been both classified as C2 grade). We examined the performance of the 16-gene signature on the 48 tumors resected after chemotherapy, and found significant difference in outcome between patients with C1 and C2 type HBs (p=0.0021, log rank test) (FIG. 6B). Remarkably, Kaplan-Meier analysis confirmed C2 subclass as a poor prognostic group in 17 cases for which pre-treatment biopsies or primary surgery specimens were available (p=0.0318, log rank test) (FIG. 6C).
[0222] We further assessed the prognostic validity of the 16-gene signature for all patients in multivariate analysis, using a Cox proportional hazards model with pathological and clinical variables associated to patients' survival. This analysis identified the signature as an independent prognostic factor, with better performance than tumor stage defined by PRETEXT stage, vascular invasion and extrahepatic metastases (FIG. 6D). Thus, this signature demonstrated strong prognostic relevance when compared to current clinical criteria.
[0223] In a Second Set of 86 Patients
[0224] The clinical impact of HB molecular classification was addressed in a second set of patients (comprising the sample of the first set), comprising 53 (61%) C1 and 33 (39%) C2 cases. Besides strong association with predominant immature histotypes, HBs of the C2 subclass were tightly associated with features of advanced tumor stage, such as vascular invasion and extrahepatic metastasis (FIG. 9A). Accordingly, overall survival of these patients was markedly impaired. Kaplan-Meier estimates of overall survival probability at 2-years were 60% for patients with C2 tumors and 94% for patients with C1 tumors (p=0.00001, log rank test), and similar trends were seen for disease-free survival probabilities (Table 9). TABLE-US-00011 Table 9 N. of patients 61 C1+25 C2 = 86 P value Survival (all patients) Alive/Dead C1 50/3 <0.00001 C2 20/13 DFS (all Datients) DFS/others C1 48/5 <0.00001 C2 18/15 Survival (non-treated Patients) Alive/Dead C1 12/0 0.0164 C2 11/6 DES (non-treated patients) DES/others C1 12/0 0.0213 C2 12/6 Survival analysis (Kaplan Mejer, log rank test); DES: disease-free survival; Others: dead or alive with recurrent disease.
[0225] Next, we examined whether pre-operative chemotherapy treatment given to 73 patients could affect tumor classification. These cases were evenly distributed among HB subclasses, with no significant association with molecular classification. We examined the performance of the 16-gene signature on the 73 tumors resected after chemotherapy, and found significant difference in outcome between patients with C1 and C2 type HBs (p=0.0002, log rank test) (FIG. 9B). Remarkably, Kaplan-Meier analysis confirmed C2 subclass as a poor prognostic group in 29 cases for which pre-treatment biopsies or primary surgery specimens were available (p=0.0164, log rank test) (FIG. 9C).
[0226] We further assessed the prognostic validity of the 16-gene signature for all patients in multivariate analysis, using a Cox proportional hazards model with pathological and clinical variables associated to patients' survival. This analysis identified the signature as an independent prognostic factor, with better performance than tumor stage defined by PRETEXT stage, vascular invasion and extrahepatic metastases (FIG. 9D).
[0227] Finally, various clinical elements of 103 HB samples from 86 patients were compared with respect to their classification as C1 or C2 grade using the 16-gene signature (Table 10). TABLE-US-00012 TABLE 10 Clinical correlations. N. of patients 61 + 25 = 86 p-value (chi-square) Gender ns Chemotherapy treatment Yes/No C1 47/6 ns C2 26/7 Chemotherapy protocol STD/High C1 30/13 0.007 C2 9/16 TUMOR STAGE Early/Advanced C1 32/20 0.005 C2 10/23 Metastasis No/Yes C1 43/10 0.004 C2 17/16 Vascular Invasion No/Yes C1 36/15 0.005 C2 13/20 Advanced Pretext stage (IV) No/Yes C1 42/9 ns C2 24/7 Multifocality No/Yes C1 36/17 ns C2 18/14 Histology Ep/Mixed C1 31/21 ns C2 20/13 Main EDith ComD Fetal/NonFetal C1 48/4 <0.0001 C2 6/22 STD: standard risks (cisplatine) - High:high risk (cisplatine/doxorubicine, intensified treatment); Tumor stage (defined as Vasc. Inv and/or metastasis and/or PRETEXT stage IV); metastasis: extrahepatic metastasis (mainly lung); vascular invasion is determined by imagery; Pretext IV (involved an intrahepatic extent of the tumor to all hepatic sections); # multifocality (more than 2 tumor nodules); Ep: pure epithelial form - Mixed: mesenchymatous and epithelial mixed form; Fetal: well differentiated; non fetal: embryonic, atypic, SCUD and/or macrotrabecular cells.
[0228] The above results carried out on a first set of 61 patients, and on a second completed set of 86 patients, demonstrate that the 16-gene signature, identified in the present application, is a strong prognostic relevance when compared to current clinical criteria.
Discussion
[0229] The present application demonstrates that, using integrated molecular and genetic studies, hepatoblastoma encompass two major molecular subclasses of tumors that evoke early and late phases of prenatal liver development. Aberrant activation of the canonical Wnt pathway represented a seminal event in both tumor types, with cumulated mutation rates of β-catenin, APC and AXIN over 80%. However, depending on tumor differentiation stage, Wnt signaling activated distinct transcriptional programs involved in tumor growth and invasiveness or in liver metabolism. Further comparisons of immature, embryonal-type HBs with the bulk of more differentiated, fetal-type tumors revealed a tight correlation between stage of hepatic maturation arrest and clinical behavior, notably vascular invasion and metastatic spread, and patients' survival.
[0230] Molecular Hb Subclasses are Determined by Liver Differentiation Stages
[0231] In this study, expression-based classification of HB was achieved through a highly reliable statistical method combining different unsupervised hierarchical clustering approaches. This method led to the selection of two robust tumor subgroups, and this robustness was confirmed using a new, independent set of samples and 16 relevant genes discriminating these tumor subgroups. These results demonstrated that the most significant differences between HB subclasses can be ascribed to distinct hepatic differentiation stages, as defined by comparison with expression profiles of mouse livers at early (E11.5-E12.5) and late (E14.5-E18.5) embryonic stages. These studies also provide biological relevance to early histologic classification that distinguished fetal and embryonal cells as major HB components (Weinberg and Finegold, 1983). The C1 subclass recapitulates liver features at the latest stage of intrauterine life, both by expression profile and by mostly fetal morphologic patterns, while in the C2 subclass, transcriptional program and predominant embryonal histotype resemble earlier stages of liver development. Thus, despite frequent morphological heterogeneity in HB, these expression-based subclasses closely matched the histologic types found to be prevailing after microscopic examination of the entire tumor mass.
[0232] These results, showing that childhood liver tumors recapitulate programs of their developing counterpart, are in line with recent studies using cross-species comparisons. It has been demonstrated that clinically distinct medulloblastoma subtypes can be identified by their similarity with precise stages of murine cerebellar development (Kho et al., 2004). Evidence for conserved mechanisms between development and tumorigenesis was also obtained in Wilms' tumor, the embryonic kidney malignancy, which shares expression of sternness and imprinted genes with murine metanephric blastema (Dekel et al., 2006). It was noticed that HBs, like Wilms' tumors, exhibit robust overexpression of a number of paternally expressed genes like DLK1, IGF2, PEG3, and PEG10 that are involved in growth induction processes and downregulated with differentiation during development.
[0233] Previous studies using stem cell markers and markers of hepatocytic and biliary lineages have described differential patterns among HB components that reflect sequential stages of liver development (Schnater et al., 2003). The present data extent these observations, and indicate that immature C2-type tumor cells evoke hepatic cancer progenitor cells, with distinctive overexpression of highly relevant markers such as cytokeratin 19 and Ep-CAM (Roskams, 2006). Recently, embryonic stem/progenitor cells have been isolated from human fetal livers, either by enrichment of blast-like cells in primary hepatoblast cultures or by immunoselection of Ep-CAM-positive epithelial cells (Dan et al., 2006; Schmeizer et al., 2007). These cell lines have self-renewal capacity and can differentiate into mature hepatocytes and cholangiocytes, and one of them also gives rise to various mesenchymal lineages (Dan et al., 2006). Whether HBs arise from transformation of these cell types is presently unknown. As malignant mesenchymal derivatives are frequently admixed with epithelial tissues in HB, it is tempting to speculate that this tumor occurs from a multipotent progenitor harboring characteristics of mesenchymal-epithelial transitional cells. Moreover, since no significant differences in gene expression profiles was noted here between pure epithelial and mixed epithelial-mesenchymal HBs, tumor cells likely kept intrinsic capacities to undergo epithelial-mesenchymal transition.
[0234] A salient feature of immature HBs is the characteristic interplay of sternness and proliferation found in aggressive tumors (Glinsky et al., 2005). The C2-type expression profile was significantly enriched in hESC markers, including the mitotic cell cycle and spindle assembly checkpoint regulators cyclin B1, BUB1, BUB1B, and Aurora kinases. These mitotic kinases are centrosomal proteins that ensure proper spindle assembly and faithful chromosome segregation in mitosis. Overexpression of these kinases or other components of the spindle checkpoint induces centrosome amplification and defects in chromosome segregation leading to chromosome number instability and aneuploidy (Marumoto et al., 2005; Zhou et al., 1998). Non-disjunctional events are involved in developmental syndromes (Hassold and Hunt, 2001), and might be responsible for increased rate of chromosomal imbalances evidenced here in C2-type HBs.
[0235] Context-Dependent Transcriptional Programs Driven by Wnt Signalling
[0236] Mutational activation of β-catenin is a hallmark of HB, and accordingly, we found intracellular accumulation and nuclear localization of the protein in virtually all tumors, albeit with variable frequencies and intensities. Both immature and differentiated tumors overexpressed AXIN2 and DKK1, reflecting an attempt to activate a negative feedback loop aimed at limiting the Wnt signal. However, the two HB subtypes showed significant differences in β-catenin immunoexpression, illustrated by concomitant nuclear accumulation and decreased membranous localization of the protein in poorly differentiated, highly proliferative HBs. Heterogeneous distribution of nuclear β-catenin within colorectal tumors has been linked to different levels of Wnt signaling activity, resulting from differential combinations of autocrine and paracrine factors (Fodde and Brabletz, 2007). Similarly, nuclear β-catenin might be related to the absence of membranous E-cadherin in immature HBs, as we reported previously (Wei et al., 2000), and to cross-talks with growth-stimulating pathways in less differentiated cells. In this context, increased dosage of Wnt signaling might induce migratory and invasive phenotype.
[0237] Major differences between the two HB subtypes were found here in expression levels of Wnt targets involved in liver functions. Recent studies have demonstrated that Wnt/β-catenin signaling governs liver metabolic zonation by controlling positively the perivenous gene expression program and negatively the periportal program (Benhamouche et al., 2006). In our study, overexpression of hepatic perivenous markers such as GLUL was prominent in differentiated HBs, while genes encoding periportal functions like GLS2 were downregulated. This profile is highly similar to those of human and murine HCCs expressing mutant β-catenin (Boyault et al., 2007; Stahl et al., 2005), and corresponds to an hepatic signature of Wnt target genes. Accordingly, the zonation-related profile was lessened in poorly differentiated HBs, and mutant β-catenin was found to activate a different, muscle-related expression program in the pediatric Wilms' tumor (Zirn et al., 2006).
[0238] Clinical Implications
[0239] The clinical behavior of many human solid tumors has been related to their differentiation status and proliferative rate. We show that HB does not depart from this rule, with strong correlation of molecular subclasses linked to hepatic differentiation with clinical tumor stage and patient's outcome. This correlation was mainly determined by differences in invasive and metastatic phenotypes between the two subclasses, but not by differences in tumor localization or tumor extension across liver sections, which defines the preoperative staging (PRETEXT) utilized to evaluate tumor resectability (Perilongo et al., 2000). Major differences in expression profiles of the two molecular HB subtypes led us to elucidate a 16-gene signature that proved highly efficient in stratification of HBs as well as normal livers according to hepatic developmental stage. Most importantly, this classifier also discriminated aggressive tumors, exhibited powerful survival predictor capacities in pre-treatment biopsies and surgical specimens, and demonstrated strong prognostic relevance when confronted to current clinical criteria in multivariate analysis. Although immature HBs have been associated to worse clinical outcome as opposed to differentiated HBs (Weinberg and Finegold, 1983), frequent cellular heterogeneity has hampered the use of histopathologic criteria for defining risk groups, excepted for a minority of cases showing `pure fetal` or SCUD types. The expression signature afforded here enables direct appraisal of the global degree of tumor cell maturation, allowing to bypass these difficulties. Thus, it can improve the outcome prediction and clinical management of hepatoblastoma, by identifying cases with increased risk of developing metastasis, or conversely, by avoiding unnecessary over-treatment.
[0240] In conclusion, the present application identifies a 16-gene signature that distinguishes two HB subclasses and that is able to discriminate invasive and metastatic hepatoblastomas, and predicts prognosis with high accuracy. The identification of this expression signature with dual capacities may be used in recognizing liver developmental stage and in predicting disease outcome. This signature can be applied to improve clinical management of pediatric liver cancer and develop novel therapeutic strategies, and is therefore relevant for therapeutic targeting of tumor progenitor populations in liver cancer.
[0241] Analysis of 64 Hepatocellular Carcinoma (HCC) from 64 Patients
[0242] Real time RT-PCR (Taqman methodology) was performed on 67 HCC samples, as disclosed for HB samples above. The clinical characteristics of the 67 patients diagnosed with HCC as well as the features of the HCC samples are disclosed in Tables 11 and 12 below.
[0243] Amplification was carried out with primers of the 16-gene signature disclosed in Table 6. Data were normalized to the expression of the ROTH2 gene (primers disclosed in Table 7) and analyzed by the ΔCt method. Quantitative PCR data are disclosed in Table 13. TABLE-US-00013 TABLE 11 features of the HCC samples obtained from 67 patients (pages 60 to 62) Tumor follow-up tumor grade tumor differentiation tumor vascular invasion recurrence or Id length (years) (Edmonson) according to OMS size macro micro metastasis HC1 0.07 3 moderately differentiated 120 NA absent no recurrence HC10 0.95 4 moderately/poorly differentiated 75 absent absent no recurrence HC11 11.10 NA NA 15 absent absent no recurrence HC12 0.05 NA Well differentiated 60 NA NA no recurrence HC14 1.00 NA moderately/poorly differentiated 80 NA NA no recurrence HC15 1.22 3 moderately differentiated 60 present present no recurrence HC17 10.96 2 Well differentiated 100 absent absent no recurrence HC18 0.39 3 moderately differentiated 140 present present NA HC20 15.40 NA Well differentiated 40 NA NA no recurrence HC21 0.70 NA NA 100 NA NA NA HC22 11.50 NA Well differentiated 45 absent absent no recurrence HC23 11.93 2 Well differentiated 50 absent absent no recurrence HC25 15.87 2 Well differentiated 140 absent absent NA HC27 0.10 NA Well differentiated 15 absent absent no recurrence HC28 0.10 NA moderately differentiated 120 NA present no recurrence HC3 3.33 2 Well differentiated 60 absent absent recurrence HC30 11.78 3 moderately differentiated 16 NA NA no recurrence HC32 0.66 2 Well differentiated 60 absent NA no recurrence HC34 14.72 2 Well differentiated 140 absent absent recurrence HC37 0.20 NA moderately differentiated 35 present present non HC38 1.12 NA NA 50 absent NA recurrence HC4 11.48 2 Well differentiated 100 absent absent no recurrence HC41 7.44 2 Well differentiated 30 NA absent recurrence HC42 10.58 3 moderately differentiated 130 possible; present no recurrence non certain HC43 10.20 NA moderately differentiated 15 NA NA no recurrence HC52 0.25 3 moderately differentiated 110 absent absent no recurrence HC58 8.30 2 moderately differentiated 100 absent absent no recurrence HC6 1.25 2 Well differentiated 90 absent present recurrence HC64 5.25 3 moderately differentiated 40 absent absent recurrence HC66 8.93 2-3 Well to moderately differentiated 75 absent absent no recurrence HC7 1.50 2-3 Well differentiated 100 present present recurrence HC8 8.48 3 moderately differentiated 30 absent absent no recurrence HC9 0.02 3-4 moderately/poorly differentiated 100 present present no recurrence HC101 1.00 2-3 Well to moderately differentiated 35 present present no recurrence HC102 0.10 NA Poorly differentiated 200 present present no recurrence HC103 1.82 2-3 Well to moderately differentiated 55 absent present recurrence HC104 0.17 2-3 Well to moderately differentiated 160 Possible; present no recurrence non certain HC105 0.56 3 moderately differentiated 40 present present recurrence HC106 1.70 3 moderately differentiated 80 present present no recurrence HC107 1.75 2 Well differentiated 60 absent absent no recurrence HC108 1.62 3 moderately differentiated 26 absent present no recurrence HC109 1.00 1-2 Well to very well differentiated 30 absent absent no recurrence HC110 1.00 3 moderately differentiated 30 present present no recurrence HC111 0.60 3 moderately differentiated 40 present present no recurrence HC1112 1.48 2-3 Well to moderately differentiated 18 absent absent no recurrence HC113 1.00 2-3 Well to moderately differentiated 50 present present no recurrence HC114 0.44 2 Well differentiated 36 absent absent no recurrence HC119 0.75 1 Well differentiated 90 absent absent no recurrence HC120 0.69 3 moderately differentiated 140 absent absent no recurrence HC121 1.00 2-3 Well to moderately differentiated 28 absent absent no recurrence HC122 0.93 1 Very well differentiated 40 absent absent no recurrence HC123 0.90 3 moderately differentiated 26 absent present no recurrence HC124 0.82 2-3 Well to moderately differentiated 20 absent present no recurrence HC125 0.60 3 moderately differentiated 150 Possible; present no recurrence non certain HC126 0.75 2 Well differentiated 20 present present recurrence HC127 0.40 3 moderately differentiated 43 probable probable no recurrence HC128 0.52 3 moderately differentiated 62 absent absent no recurrence HC129 0.30 3 moderately differentiated 25 absent present no recurrence HC131 0.42 1-2 Well differentiated 130 present present recurrence HC132 0.25 2-3 Well to moderately differentiated 115 present present recurrence HC133 0.44 2 Well to moderately differentiated 110 absent present no recurrence HC134 0.10 3 moderately differentiated 30 absent present no recurrence HC135 0.14 3 moderately differentiated 38 absent Possible; no recurrence non certain HC136 0.26 2-3 Well to moderately differentiated 120 absent present no recurrence N.A: non available; macro: macrovacular invasion; micro: microvacular invasion
[0244] TABLE-US-00014 TABLE 12 features of the HCC samples obtained from 67 patients, and features of patients (pages 63 and 64) Chronic Other Tumor Score METAVIR viral Viral etiology etiolo- ID Activity Fibrosis hepatitis HBV HCV alcohol gies HC1 NA 4 no no no yes HC10 NA 4 yes yes no no HC11 NA NA yes yes yes no HC12 NA NA yes yes no no HC14 NA NA yes no yes yes HC15 3 3 no no no yes HC17 NA 3 yes yes no no HC18 2 4 no no no yes HC20 NA NA no no no yes HC21 NA NA no no no yes HC22 NA NA no no no yes HC23 NA 0 no no no no HC25 0 0 no no no no HC27 NA NA yes no yes no HC28 0 0 no no no no HC3 NA 4 yes no yes no HC30 NA 4 no no no yes HC32 NA 4 yes no yes no HC34 NA 0 no no no no HC37 NA NA no no no yes HC38 NA 4 yes no yes no HC4 NA 1 no no no no HC41 NA 4 yes no yes no HC42 2 1 yes yes no no HC43 NA NA yes no yes no HC52 NA 4 yes yes no no HC58 2 3 yes no yes no HC6 NA 1 no no no yes Hemochro HC64 2 2 yes no yes no HC66 NA 4 yes yes no yes HC7 2 3 no no no yes HC8 NA 4 yes no yes no HC9 1 3 no no no yes HC101 2 4 yes yes yes yes HC102 1 1 yes yes yes no HC103 3 4 yes yes no no HC104 0 1 no no no no HC105 2 4 yes no yes no HC106 1 4 yes yes no no HC107 0 0-1 no no no yes HC108 1 1 yes no yes no HC109 2 4 no no no yes NASH HC110 1 4 yes no yes yes HC111 1 4 no no no yes HC112 2 2 no no no no NASH HC113 1 4 yes no yes no HC114 2 3 no no no yes HC119 2 1 no no no no NASH HC120 2 3 yes yes no no HC121 2 4 yes no yes no HC122 0 1 no no no no HC123 2 4 yes no yes yes HC124 1 4 yes yes no no HC125 2 4 no no no yes NASH HC126 1 4 yes yes no no HC127 2 4 yes no yes no HC128 1 1 no no no no NASH HC129 2 4 no no no yes HC131 0 1 no no no no HC132 1 1 yes yes no no HC133 2 2 no no no yes HC134 2 3 yes no yes no HC135 1 2 yes yes no no HC136 0 1 no no no no N.A: non available; HBV: hepatitis B virus; HCV; hepatitis C virus; hemochro: hemochromatosis; NASH non alcoholic steatohepatitis.
[0245] TABLE-US-00015 TABLE 13 Quantitative PCR data of the 16-gene signature normalized to the expression of the ROTH2 gene (pages 65 to 68) HC1 HC3 HC4 HC6 HC7 HC8 HC9 HC10 HC11 AFP -2.212911 -3.865709 -7.6758115 -7.9469815 5.311541 2.0890815 -70483095 2.3869635 0.6488335 ALDH2 6.2372335 6.230074 2.186358 5.4231035 4.0446765 3.9297005 3.0017225 0.95212 5.958108 AP0C4 0.614689 0.95786 -1.608247 0.9614255 -3.550537 -0.6776965 -9.6721075 NA 1.076151 APCS 7.0721355 7.52919 5.845683 7.3704745 5.1967915 6.567126 -0.017488 -1.0272875 7.7638255 AQP9 6.047695 6.7334475 3.759528 7.006052 6.747103 3.1082155 3.7536735 1.3400495 6.122144 BUB1 -3.841505 -0.147459 -4.221132 -0.5252045 -0.299039 -1.214781 2.980029 -1.864677 -2.362454 C1S 8.163492 8.7963405 5.8997645 8.162856 4.062593 7.2991535 4.830331 2.639902 8.319293 CYP2E1 10.3093235 10.428074 7.1147515 10.1334265 11.024027 7.7910075 0.5825245 3.604805 9.575619 DLG7 -5.30317 -2.057513 -4.4226465 -1.6282005 -1.169221 -2.80866 1.3733475 NA -2.8432205 DUSP9 -11.616567 -8.8462855 -9.4268185 -10.22051 -6.6521625 -9.6946695 -9.5262655 NA NA E2F5 0.05328 -1.909804 -1.7432195 0.024339 -0.2833465 -0.0193165 0.711082 -1.344368 -0.736822 GHR 2.655512 2.069524 -2.0012965 1.887805 -1.7428205 2.342442 -2.3242195 -0.4900285 4.757848 HPD 9.449416 8.549803 9.415253 8.5958965 6.183977 5.329776 -0.011478 2.932809 9.029214 IGSF1 -6.46034 -7.249974 NA -7.1580385 -3.192514 -2.806768 -4.026769 NA -7.6390015 NLE1 -1.159417 -1.5801355 -3.1459935 0.6940375 -0.3919565 -1.579419 -0.80375 NA -1.9328755 RPL10A 6.6225235 6.0562915 4.4121905 6.8637555 7.1381125 6.2574845 6.3016635 9.1966395 7.379063 HC12 HC15 HC17 HC18 HC20 HC21 HC22 HC23 HC25 AFP -6.538312 6.14089 7.1950405 -6.856588 -0.65281 -4.3070475 -4.418018 -5.538438 -3.90298 ALDH2 4.6271565 4.5178635 2.6522585 1.840894 6.287083 2.175112 5.331214 5.853486 6.162477 AP0C4 -1.221393 -5.156026 -2.395651 -3.84764 3.2094885 -6.2591235 0.5455545 0.5708905 1.834891 APCS 6.942673 3.380102 4.5167035 4.916924 8.2117635 5.9159775 6.6835035 6.9009145 8.798759 AQP9 4.1878425 2.373344 2.8711295 3.6093495 7.354605 1.1452535 5.7992305 6.651868 8.758959 BUB1 -3.293346 0.8830545 1.0884485 -0.063545 -1.4635025 0.0802935 -2.173361 -2.5475915 -2.5679685 C1S 6.850023 7.1343975 6.035123 4.263272 8.471663 5.7190985 7.2514145 8.2212235 8.5606875 CYP2E1 7.284587 4.9390935 6.037085 5.811062 10.2536915 1.2878015 8.0876755 9.047509 10.814935 DLG7 -4.7199665 -0.1414205 0.666284 -1.512286 -2.1165725 -0.322455 -3.3904095 -3.848364 -3.34202 DUSP9 NA -4.4342765 -3.163581 -8.7756845 -9.6208445 -7.8162765 -10.827291 NA -7.1111525 E2F5 -2.4002515 1.399564 1.206766 -2.426129 -1.1944835 -0.0686475 -0.7133385 -1.4330655 0.049846 GHR 2.2402875 0.2426 -2.353691 -2.9035 4.5756335 0.71981 2.416651 3.7226655 1.9012935 HPD 9.656029 4.473096 0.6808655 5.7101575 10.6864405 4.0108195 9.8859985 9.583194 9.1845675 IGSF1 -7.466951 0.0722075 -6.0490105 -2.4248235 NA -2.954514 -5.6986975 -7.200325 NA NLE1 -1.64183 -0.321593 -0.386649 -1.3815525 -1.118745 -1.618369 -1.9449755 -1.823275 -1.770127 RPL10A 5.178571 6.8777395 7.068098 5.9464565 7.542193 6.309556 7.194012 5.9526365 7.4507165 HC26 HC27 HC28 HC30 HC32 HC34 HC37 HC38 HC41 AFP -5.69175 -0.626755 NA 6.4370325 0.0037145 -6.6945705 -1.3519745 4.053435 -2.7156435 ALDH2 5.0135775 5.6309605 1.913778 3.8476295 6.802666 5.11617 5.808058 4.596143 6.3503265 AP0C4 0.2581675 1.53158 -6.0251725 0.2797975 2.574347 0.5860455 -0.0768065 -0.129322 2.281983 APCS 7.2072275 7.2809855 1.0475505 7.1142435 7.500133 7.134934 6.755895 5.045701 5.612517 AQP9 3.8645965 5.4736555 0.9613895 5.0250435 7.530391 6.9427395 6.3416265 6.0302545 7.8444565 BUB1 0.545363 -0.8889165 -5.7426525 -0.190936 -5.1317805 -1.2674215 -2.4955985 0.321483 -0.587016 C1S 7.2351705 8.172076 4.910584 7.5279395 7.854502 7.719763 6.921051 6.101331 6.88808 CYP2E1 0.671071 8.6350095 3.6858305 7.5682115 9.4408715 8.545814 10.1686795 8.1123675 9.5090495 DLG7 -0.9710395 -2.3158215 NA -0.189092 -5.7080765 -2.339621 -2.6534895 -1.4386515 -1.840185 DUSP9 -8.5287915 -10.241011 NA -9.0027 -9.73163 -9.9728495 NA -5.2298755 -8.727439 E2F5 -1.1845665 -0.4045835 -4.334386 1.0623035 -0.054818 -1.4281575 -1.2212655 -0.037887 0.466649 GHR 1.964045 2.623084 -1.9788575 2.635437 2.0027475 1.563203 2.9415775 0.2025015 1.428749 HPD 7.6403735 9.597772 3.3142495 7.537 9.0015185 8.3685675 10.367265 7.547286 8.0015745 IGSF1 -5.4960635 -5.588995 NA -2.651022 NA -10.112616 -7.5570255 -0.680358 -7.243446 NLE1 -1.851733 -1.851285 -2.4559905 -1.2674865 -1.208576 -1.934745 -1.9881245 -2.1250395 -0.15624 RPL10A 5.9670715 7.6623025 5.521873 7.5046195 8.8437815 6.594006 6.901637 5.1574215 7.7043325 HC42 HC43 KC44 HC52 HC58 HC60 HC64 HC66 HC101 AFP -5.216493 -1.7983435 -0.564605 10.3337105 1.891958 7.624821 5.0266755 3.156328 -6.873135 ALDH2 4.4086495 5.457548 7.1344115 2.1920375 2.1172735 3.6860195 4.992107 3.8408415 4.339036 AP0C4 -0.627239 -0.7055185 0.499817 -8.124407 -11.8524 -0.545509 0.7860345 -0.6773785 -0.5787185 APCS 4.1054755 7.607914 7.567581 5.9818015 -4.1106695 8.100997 7.4148835 8.2106815 6.288568 AQP9 6.063786 4.7175855 6.058158 -0.4848805 -2.817265 6.8503395 7.0526325 6.2767975 4.6233735 BUB1 -2.224818 -2.8634735 -3.5668895 -1.2986035 1.9395175 -0.576028 -1.367463 -1.1272665 0.081457 C1S 6.3060565 7.9862115 8.547705 5.6337865 3.691331 8.167253 7.1364365 8.026875 7.321092 CYP2E1 9.1411555 8.760714 9.1133175 1.7693015 -4.3317445 9.1875325 9.682147 8.601088 5.806032 DLG7 -3.2531575 -4.2390495 -4.814388 -2.599359 0.1957495 -2.2644225 -2.386875 -2.7680135 -1.3084655 DUSP9 NA -10.525647 NA -3.8059605 -3.656912 -6.618755 -7.3184655 -11.5673955 -8.828389 E2F5 -0.3673235 -0.894345 -1.894272 0.4419525 0.804087 -0.432422 -0.2876185 -0.968982 -1.871516 GHR -1.2545195 3.2916395 4.5598275 -1.843696 -3.7242975 -1.4079225 0.349645 -1.2501855 0.1466275 HPD 8.2669835 8.997825 9.158005 2.481945 1.8257985 8.4643875 8.6027575 8.5231325 5.7252795 IGSF1 -2.899766 -5.5544715 -5.769786 2.254168 1.3471695 -0.7884805 -3.3382005 -9.185554 -4.1394545 NLE1 -0.9401045 -1.8422595 -2.0303285 -1.9474305 -1.209522 -1.9133155 -1.817699 -1.962008 -1.4546305 RPL10A 5.577659 5.480403 5.8488475 5.6154705 6.0601515 5.7041285 6.4617635 5.415169 6.144011 HC102 KC103 HC104 HC105 HC106 HC107 HC108 HC109 HC110 AFP -4.119697 1.6193685 5.5094265 2.3444245 -3.42054 -4.136209 -4.500336 -4.833024 -3.5240185 ALDH2 2.476355 3.889904 4.936239 4.239726 6.1642895 6.7443095 3.6076385 5.8617665 3.6707715 AP0C4 -5.453696 -0.54698 -0.5059805 -3.577778 -0.7836775 4.4534435 -2.478085 0.729565 -0256479 APCS -2.3952165 6.014572 5.624234 7.703333 7.8462545 9.2080655 7.275462 6.222909 5.043319 AQP9 0.0196725 7.151639 0.501258 4.2748785 5.85931 8.8878655 4.4353395 6.4504115 4.5999895 BUB1 -0.5553155 -2.086008 -1.311194 0.945674 -4.8909655 -1.7415115 -0.3807995 -2.2918285 -1.449943 C1S 5.939374 5.965432 6.716137 7.774455 8.060072 9.2061165 7.1031155 7.406001 6.9163195 CYP2E1 -2.8566735 8.266311 9.0888685 5.698899 9.9949555 9.3234825 3.889942 8.7101925 7.1 45766 DLG7 -2.1385165 -2.957914 -1.821739 -0.814912 -6.2678815 -1.357756 -2.2445545 -3.222524 -2.333076 DUSP9 -8.6628475 -12.521336 -5.396553 -5.4214725 -11.174152 -6.6136855 -8.0946735 -10.4709205 -11.616244 E2F5 0.830934 -1.8003215 -2.305498 2.0730715 -2.208171 2.78876 0.0923905 -1.9924345 -2.512512 GHR 0.947389 0.636723 1.6860905 0.682142 5.342392 2.935929 1.6363755 2.9233285 1.0803015 HPD 0.568809 6.717282 8.46781 2.288109 9.4440475 10.460972 2.9674235 7.8859205 8.1908235 IGSF1 -2.708733 -9.802921 0.1438735 -1.422332 -7.401009 NA -7.967992 -10.0122565 -8.1469415 NLE1 -1.1534675 -2.594702 -1.610158 -0.471391 -1.968983 -0.000835 -0.932052 -2.6102395 -2.3529485 RPL10A 5.283399 4.423835 6.21159 6.315756 5.769397 8.6686655 5.818028 5.541229 5.245476 HC111 HC112 HC113 HC114 HC119 HC120 HC121 HC122 HC123 AFP -1.883473 -2.8803905 1.208649 -5.4433695 1.0580855 -4.0065425 -4.254961 -2.3763095 0.821555 ALDH2 3.8304065 4.8726745 4.407016 4.7113965 6.159706 4.257398 4.556431 6.2844515 4.220769 AP0C4 -1.130067 -0.7777655 -2.366969 -0.833543 1.894453 -3.5241745 -2.167313 1.279577 -0.68167 APCS 5.976754 6.764675 5.197177 6.723142 9.375177 5.6838965 6.2688205 6.9942545 5.778659 AQP9 4.1657805 5.2735435 2.681192 4.445291 7.6266135 6.8239115 4.38702 6.8198535 6.410177 BUB1 0.621548 0.3135015 -3.4825665 -1.7431855 -0.797564 -0.0740105 -2.4486685 -6.0183915 -1.190323 C1S 6.278164 7.455794 6.338901 7.866014 9.1461175 8.5708615 8.118416 7.7653135 5.383781 CYP2E1 4.46942 2.5741475 6.443846 7.3429245 7.095824 7.6044515 7.765037 9.450349 8.528543 DLG7 -0.769283 -0.9196845 -4.5602875 -3.1500875 -1.712686 -1.9563135 -2.852561 -7.228946 -2.929576 DUSP9 -9.137462 -10.105965 -7.8299455 -11.804112 -9.106547 -5.8119685 -9.706684 -9.9054825 -11.584458 E2F5 1.045678 0.0373705 -2.82243 -0.0450475 -0.0248045 1.229768 -0.910943 -3.5033365 -0.646839 GHR 1.1576425 2.5391085 2.16232 2.5053965 3.7649595 3.196589 2.2774645 2.400201 -1.810364 HPD 7.245347 7.714358 6.685692 6.835254 9.220498 8.5127155 7.480725 8.7301975 4.7774665 IGSF1 -1.86965 -3.4428695 -2.045068 -5.1813245 -5.39017 -9.404196 -5.980435 -8.6480295 -5.1400615 NLE1 -1.012752 -1.119237 -2.156348 -1.3170345 -0.400823 -1.1096815 -1.758163 -2.2430545 -1.5951645 RPL10A 5.568205 6.1905075 5.8884625 5.795905 7.954231 6.4517175 6.4042545 5.199782 4.7323885 HC124 HC125 HC126 HC127 HC128 HC129 HC131 HC132 HC133 AFP 3.9525335 -4.806564 -5.899437 -0.0390765 5.8636305 -3.430757 -1.491189 5.4265205 -5.1621395 ALDH2 4.027289 4.5451465 5.02839 2.41699 5.085525 4.6298475 5.425994 3.105643 4.2462915 AP0C4 -0.0499065 2.6326775 0.407895 0.8680995 -0.626498 -1.863955 2.4702 -6.9974515 0.63156 APCS 5.391271 6.5321595 5.2838365 4.846116 5.087517 4.8448705 8.6617295 -3.2748865 7.145861 AQP9 4.463488 8.370224 3.6163545 1.8613935 4.3184915 2.870839 7.4772145 3.9244375 6.05182 BUB1 -1.592563 1.1627945 -2.6943025 -2.048769 -1.3297375 -2.3688215 -0.727709 0.2895395 -4.9277675 C1S 5.151686 8.4244055 7.1365955 6.3641695 6.828468 7.302922 7.525072 4.390082 7.3188145 CYP2E1 9.520436 9.426232 5.226091 6.1813065 7.4344035 2.692798 8.98645 7.0455735 8.1908895 DLG7 -2.03781 0.3286545 -3.944339 -2.96212 -2.6299155 -3.6405185 -1.461713 -1.5572645 -5.5447335 DUSP9 -8.81055 -9.3740615 -8.7174575 -8.672372 -8.499355 -7.0627455 -8.415907 -3.3843145 -8.022457 E2F5 0.574165 -0.028878 -3.271927 -2.162802 -4.393094 -0.470421 0.154573 1.9018925 -2.6341525 GHR 2.2369305 0.697866 1.824385 0.129431 1.9716885 2.332961 4.009655 1.7710325 2.2298335 HPD 7.832169 5.7813 1.865621 3.4481965 5.7052855 5.502918 8.960383 2.3653865 6.1281315 IGSF1 -1.4450915 -10.2234745 -7.659377 -3.1503205 -2.72995 -5.692623 -7.5832005 -1.947055 NA NLE1 -0.1499775 -0.405397 -2.033278 -2.205965 -1.949352 -1.683808 -1.5313675 0.2035885 -1.4173895 RPL10A 6.691521 7.1196575 5.389272 4.3385115 6.6181545 4.8697295 6.775249
6.7796075 5.762015 AFP HC134 HC135 HC136 ALDH2 2.8738695 -0.909107 -0.4105125 AP0C4 4.061101 2.7442165 6.0408575 APCS -0.1134065 -0.7630605 0.7390785 AQP9 7.5103485 0.959726 7.150737 BUB1 5.550642 4.0595615 5.996196 C1S 1.7425995 -1.2018365 -4.288554 CYP2E1 8.4609335 4.667223 8.243333 DLG7 7.859701 4.30592 9.042865 DUSP9 0.8148735 -2.250305 -5.5267715 E2F5 -4.96739 -5.794605 -10.9307725 GHR 3.1030595 0.986165 -2.4040865 HPD 1.3138565 -0.6955465 4.013948 IGSF1 7.231144 6.7262275 8.223611 NLE1 -0.3848995 -4.394354 -7.4962365 RPL10A 0.794433 -0.9780515 -2.426321 AFP 7.7140665 6.689595 5.5069335 NA: non available
[0246] Data were then analyzed by unsupervised clustering (dCHIP software) using 2 methods: average and centroid. Tumors were clustered into 2 groups, C1 and C2. Most of the samples have been attributed the same classification using the 2 methods, except for 6 samples (9%) that have been attributed a different classification (Table 15).
Clinical Parameters Associated to the C1 and C2 Molecular Subclasses
[0247] The clinico-pathological parameters of patients and tumors were compared between the two groups C1 and C2, using student's t test and Kaplan-Meier estimates. Since some data are not available for 3 patients, the following statistical studies were performed on 64 tumors.
[0248] Survival Analysis
[0249] There is a strong correlation of the molecular classification into C1 and C2 with patient's survival by using both classifications (Log rank: Centroid p=0.020 and Average p=0.024) (FIG. 10). In this figure, censored cases indicate the end of the follow-up (the last visit) for individual cases. Probability of survival at two years is 78% for C1 subclass and 39% for C2 subclass (the follow-up may be less than 2 years for some patients).
[0250] Association of HCC Classification with Clinical Variables
[0251] Table 14 shows the correlation between some clinical variable and the classification of the tumors. TABLE-US-00016 TABLE 14 Variable C1 C2 p-value Tumor grade >2 (Edmonson) 13/29 21/23 <0.0001 Moderately-poorly differentiated (OMS) 17/36 23/25 <0.0001 Macrovascular Invasion 6/30 9/21 0.074 Microvascular Invasion 13/32 15/22 0.043 Recurrence 7/36 5/25 ns (ns: non-significant)
[0252] TABLE-US-00017 TABLE 15 Classification of samples by unsupervised clustering (dCHIP software): average and centroid methods. Tumor ID average centroid comparison HC1 C1 C1 Same HC10 C2 C2 Same HC11 C1 C1 Same HC12 C1 C1 Same HC14 C1 C1 Same HC15 C2 C2 Same HC17 C2 C2 Same HC18 C2 C2 Same HC20 C1 C1 Same HC21 C2 C2 Same HC22 C1 C1 Same HC23 C1 C1 Same HC25 C1 C1 Same HC26 C1 C2 Different HC27 C1 C1 Same HC28 C2 C2 Same HC3 C1 C1 Same HC30 C2 C2 Same HC32 C1 C1 Same HC34 C1 C1 Same HC37 C1 C1 Same HC38 C2 C2 Same HC4 C1 C1 Same HC41 C1 C1 Same HC42 C2 C1 Different HC43 C1 C1 Same HC44 C1 C1 Same HC52 C2 C2 Same HC58 C2 C2 Same HC6 C1 C1 Same HC60 C2 C2 Same HC64 C2 C2 Same HC66 C1 C1 Same HC7 C2 C2 Same HC8 C2 C2 Same HC9 C2 C2 Same HC101 C1 C2 Different HC102 C2 C2 Same HC103 C1 C1 Same HC104 C2 C2 Same HC105 C2 C2 Same HC106 C1 C1 Same HC107 C1 C1 Same HC108 C1 C1 Same HC109 C1 C1 Same HC110 C1 C1 Same HC111 C2 C2 Same HC112 C1 C2 Different HC113 C2 C2 Same HC114 C1 C1 Same HC119 C1 C1 Same HC120 C1 C1 Same HC121 C1 C1 Same HC122 C1 C1 Same HC123 C2 C1 Different HC124 C2 C2 Same HC125 C1 C1 Same HC126 C1 C1 Same HC127 C2 C2 Same HC128 C2 C2 Same HC129 C1 C2 Different HC131 C1 C1 Same HC132 C2 C2 Same HC133 C1 C1 Same HC134 C2 C2 Same HC135 C2 C2 Same HC136 C1 C1 Same
[0253] In a second analysis, the global set of 64 tumors was analyzed independently of the C1/C2 classification, for parameters associated to survival. Results are presented in Table 16. TABLE-US-00018 TABLE 16 Variable Log rank Tumor grade >2 0.108 Mod-poor Duff. Degree 0.400 Macrovasc. mv. 0.004 Microvasc. mv. 0.026 recurrence ns Tumor size 2cm 0.397 Score METAVIR Activity ns Score METAVIR Fibrosis 0.038 <2 vs. ≧ 2 (variable 3) Chronic hepatitis 0.948 HBV 0.093 HCV 0.352 Alcohol 0.225 (ns: non-significant)
[0254] These results demonstrate that the methods and the signatures of the invention are able to determine the grade not only of HB tumors but also of HCC tumors. The inventors have shown that hierarchical clustering is an efficient method for classification of tumor grade especially for HB. For HCC, this method may be less sufficient (less robust) when the amplitude of variation of expression results of the genes is less important than for HB.
Classification of Hepatoblastomas and Hepatocellular Carcinomas Using the Method of Discretization of Continuous Values.
[0255] 85 hepatoblastomas (HBs) and 114 hepatocellular carcinomas (HCCs) including to the samples used in the above examples have been analyzed by quantitative PCR using the 16-gene signature and have been classified by the method of discretization of continuous values in order to determine their tumor grade.
Description of the Methodology for Classification
[0256] The inventors have designed a methodology for classification based on the principle of discretization of continuous values which refers to the process of converting continuous variables to "discretized" or nominal sets of values.
[0257] The major advantage of the discretization method relies on the definition of a cut-off for codification of each qPCR value (either by the Taqman or by the SybrGreen method), which provides an intrinsic score to directly classify an individual sample. There is hence no requirement to compare a sample to a large series of samples. In contrast, in other classification methods, the assigned subclass (such as C1 or C2 disclosed herein) is relative to the values obtained in a large number of cases. Moreover, the use of the average discretized values allows to tolerate missing values when analyzing the qPCR results (i.e. missed amplification of one of the genes for technical reasons).
[0258] Using the qPCR data of the 16 genes normalized to the reference RHOT2 gene (-deltaCt values), a cut-off (or threshold) has been defined for each gene. The -deltaCt values are converted into discrete values "1" or "2" depending on an assigned cut-off. In order to privilege the identification of samples that display strong overexpression of proliferation-related genes and/or strong downregulation of differentiation-related genes, the cut-offs have been defined as follows:
[0259] for the 8 proliferation-related genes (AFP, BUB1, DLG7, DUSP9, E2F5, IGSF1, NLE1, RPL10A), all -DeltaCts with a value above the 67th percentile have been assigned discretized value "2", otherwise the assigned value was "1".
[0260] for the 8 differentiation-related genes (ALDH2, APCS, APOC4, AQP9, C1S, CYP2E1, GHR, HPD), all -deltaCts with a value below the 33rd percentile have been assigned discretized value "1", otherwise the assigned value was "2".
Classification of 85 Hepatoblastomas (HB)
RNA Preparation and Quantitative PCR
[0261] RNA was extracted by using either Trizol, RNeasy kit (QIAGEN) or miRvana kit (Ambion), then quantified and quality-checked by Agilent technology.
[0262] For quantitative PCR analysis, the Sybr Green approach was used as described in point E. above. For each cDNA preparation, 1 μg of RNA was diluted at the final concentration of 100 ng/μl, and reverse transcribed with the Superscript RT kit (Invitrogen) following the manufacturer's protocol. Random primers were added at the final concentration of 30 ng/μl and the final volume was 20 μl. The cDNA was diluted 1:25, and 5 μl were used for each qPCR reaction. We added 5 μl of 2×Sybr Green Master mix (Applied Biosystems) and 0.3 μl of each specific primer (disclosed in point H. above) (final concentration 300 nM). Each reaction was performed in triplicate. qPCR reactions were run on the Applied Biosystems 7900HT Fast Real-Time PCR System with a 384-well thermo-block, and the conditions were the following:
[0263] 2 min at 50° C. to activate Uracil-N-glycosylase (UNG)-mediated erase of a specific reaction
[0264] 10 min at 95° C. to activate the polymerase and inactivate the UNG
[0265] 40 cycles:
[0266] 15 sec at 95° C. denaturation step
[0267] 1 min at 60° C. annealing and extension
[0268] Final dissociation step to verify amplicon specificity.
The normalized qPCR (deltaCt) values of the 85 HB samples are given in Table A.
Analysis of qPCR Data.
[0269] Assignment of a discretized value for the 8 proliferation-related genes ("AFP" "BUB1" "DLG7" "DUSP9" "E2F5" "IGSF1" "NLE" "RPL10A") was based on the 67th quantile (i.e. percentile), given that around 1/3 of HB cases overexpress proliferation genes, which is correlated with tumor aggressiveness and poor outcome. Assignment of a discretized value for the 8 differentiation-related genes ("ALDH2" "APCS" "APOC4" "AQP9" "C1S" "CYP2E1" "GHR" "HPD") was based on the 33rd quantile, given that around 1/3 of HB cases underexpress differentiation genes, which is correlated with tumor aggressiveness and poor outcome.
[0270] The cut-offs (or thresholds) selected for the -deltaCT value of each gene were determined after considering said chosen percentiles for each group of genes are as follows:
[0271] AFP: 3.96139596; ALDH2: 4.3590482; APCS: 4.4691582; APOC4: 2.03068712; AQP9: 3.38391456; BUB1: -1.41294708; C1S: 4.24839464; CYP2E1: 6.70659644; DLG7: -3.3912188; DUSP9: 2.07022648; E2F5: -0.72728656; GHR: -0.1505569200; HPD: 2.27655628; IGSF1: 0.1075015200; NLE: -0.02343571999; RPL10A: 6.19723876
[0272] For the sample, the relative expression value is determined for each gene of the set of profiled genes. Each value is compared to the cut-off for the corresponding gene and is then discretized as a result of its position with respect to said cut-off.
[0273] The next step consisted in assigning a discretized score to each sample as follows:
1--the average of the "discretized" values of the 8 proliferation-related genes was determined. The 8 proliferation-related genes are the following: AFP, BUB1, DLG7, DUSP9, E2F5, IGSF1, NLE, and RPL10A.
2--the average of the "discretized" values of the 8 differentiation-related genes was determined. The 8 differentiation-related genes are the following: ALDH2, APCS, APOC4, AQP9, C1S, CYP2E1, GHR, and HPD.
3--The score for each sample was determined as the ratio between the average of proliferation-related genes and the average of differentiation-related genes.
According to this calculation, a score of 2 is the maximal score for highly proliferating and poorly differentiated tumors, whereas well differentiated and slowly proliferating tumors will have a minimal score of 0.5.
[0274] Based on the scores assigned to the 85 HB samples analyzed, cut-offs were identified to separate the samples into relevant subclasses. Two different cut-offs that correspond to the 33rd (0.615), and 67th percentile (0.91) have been assessed, leading to the definition of either 2 or 3 subclasses. These data together with the clinical data of 85 HB cases are given in the Table B.
Statistical Analysis of Clinical Correlations
[0275] All statistical correlations were analyzed using the discrete classification into 2 subclasses with the 67th percentile (see 3rd column of the table given in Table B). TABLE-US-00019 Samples with Samples with p-values score <67th score >67th (chi- Characteristics percentile percentile square test) Previous C1/02 52/5 2/26 1.0739e-14 classification Gender Male/Female 28/29 7/21 0.03368 PRETEXT.stage 30/25 11/15 0.30367 I-II/III-IV Distant Metastasis 45/12 15/13 0.015808 No/Yes Vascular invasion 38/17 11/17 0.0090345 No/Yes Multifocality No/Yes 38/18 15/13 0.20088 Histology 34/22 16/22 0.75303 Epithelial/Mesenchymal β-catenin mutation 8/45 8/16 0.067697 No/Yes Main epithelial 49/7 5/21 2.33206e-9 component Fetal/Other* *Other = embryonal, macrotrabecular, crowded fetal
[0276] The best correlation of the discrete classification was observed with the previous classification into C1 and C2 classes, followed by the main epithelial histological component. The correlation with patients' survival is also excellent, as shown by using the Kaplan-Meier estimates and the log-rank test. Illustrative Kaplan-Meier curves are given in FIG. 11 for specific cancer-related survival, using different percentiles to classify the tumors.
[0277] In conclusion, this study shows that the discretization method allows to classify hepatoblastoma as efficiently as the previously described method.
[0278] A similar approach was therefore applied to the analysis of hepatocellular carcinoma.
Analysis of 114 Hepatocellular Carcinomas (HC)
RNA Preparation
[0279] RNA was extracted by using either Trizol, RNeasy kit (QIAGEN) or miRvana kit (Ambion), then quantified and quality-checked by Agilent technology.
[0280] For each cDNA preparation, 1 μg of RNA was diluted at the final concentration of 100 ng/μl, and reverse transcribed with the Superscript RT kit (Invitrogen) following the manufacturer's protocol. Random primers were added at the final concentration of 30 ng/μl and the final volume was 20 μl. The cDNA was diluted 1:25, and 5 μl were used for each qPCR reaction. We added 5 μl of 2×Sybr Green Master mix or the Taqman Master mix (Applied Biosystems) and specific primers (and probes when using Taqman chemistry) at the concentration indicated by the manufacturer. Each reaction was performed in triplicate. qPCR reactions were run on the Applied Biosystems 7900HT Fast Real-Time PCR System with a 384-well thermo-block, and the conditions were the following:
[0281] 2 min at 50° C. to activate Uracil-N-glycosylase (UNG)-mediated erase of aspecific reaction (omit if using the Taqman approach)
[0282] 10 min at 95° C. to activate the polymerase and inactivate the UNG
[0283] 40 cycles:
[0284] 15 sec at 95° C. denaturation step
[0285] 1 min at 60° C. annealing and extension
[0286] Final dissociation step to verify amplicon specificity (omit if using the Taqman approach)
Quantitative PCR
[0287] Real time RT-PCR was performed for 16 genes on 114 HCC samples using two different technologies:
[0288] Sybr Green as described above for hepatoblastoma (26 samples).
[0289] Taqman methodology (88 samples) using primers and probes designed and publicly released by Applied Biosystems company.
Examples
[0290] TABLE-US-00020 AFP forward primer: GCCAGTGCTGCACTTCTTCA AFP reverse primer: TGTTTCATCCACCACCAAGCT AFP Taqman probe: ATGCCAACAGGAGGCCATGCTTCA RHOT2 forward primer: CCCAGCACCACCATCTTGAC RHOT2 reverse primer: CCAGAAGGAAGAGGGATGCA RHOT2 Taqman probe: CAGCTCGCCACCATGGCCG
[0291] Each reaction was performed in triplicate for Sybr Green protocol and in duplicate for the taqman protocol. qPCR reactions were run on the Applied Biosystems 7900HT Fast Real-Time PCR System with a 384-well thermo-block.
Raw data for each gene were normalized to the expression of the ROTH2 gene, providing the deltaCt values that were then used for tumor classification into subclasses using the discretization method.
The normalized qPCR values (deltaCt) of the 16 genes in 26 HCC samples analyzed by the Sybr Green approach is given in Table C. The deltaCt values for 88 HCCs analyzed by the Taqman approach are given in Table D.
Analysis of qPCR Data.
[0292] The -deltaCt values for each gene in each sample was used. The cut-offs (or thresholds) selected for each gene using the Taqman method or the SybrGreen method are as follows: TABLE-US-00021 Table E of cut-offs for discretization values Gene name Cut-off for Taqman Cut-off for SybrGreen AFP -1.2634010 -2.3753035 ALDH2 4.014143 5.314302 APCS 5.6142907 6.399079 APOC4 -0.7963158 4.656336 AQP9 4.2836011 5.446966 BUB1 -1.2736579 -3.634476 C1S 6.3514679 6.240002 CYP2E1 6.9562419 5.829384 DLG7 -2.335694 -4.614352 DUSP9 -7.979559 -1.8626715 E2F5 -0.4400218 -1.367846 GHR 1.0832632 1.169362 HPD 6.7480328 6.736329 IGSF1 -4.8417785 7.6653982 NLE -1.6167268 -1.82226 RPL10A 6.2483056 5.731897
[0293] For the sample, the relative expression value is determined for each gene of the set of profiled genes. Each value is compared to the cut-off for the corresponding gene and is then discretized as a result of its position with respect to said cut-off.
[0294] The next step consisted in assigning a score to each sample as follows:
1--the average of the "discretized" values of the 8 proliferation-related genes was determined. The 8 proliferation-related genes are the following: AFP, BUB1, DLG7, DUSP9, E2F5, IGSF1, NLE, and RPL10A.
2--the average of the "discretized" values of the 8 differentiation-related genes was determined. The 8 differentiation-related genes are the following: ALDH2, APCS, APOC4, AQP9, C1S, CYP2E1, GHR, and HPD.
3--The score for each sample was determined as the ratio between the to average of proliferation-related genes and the average of differentiation-related genes.
According to this calculation, a score of 2 is the theoretical maximal score for highly proliferating and poorly differentiated tumors, whereas well differentiated and slowly proliferating tumors will have a theoretical minimal score of 0.5.
[0295] Based on the scores assigned to the 114 samples analyzed, cut-offs are identified to separate the samples into relevant subclasses. Three different cut-offs that correspond to the 30rd (0.66), 50th (0.8125) and 67th percentile (0.925) have been assessed, leading to 4 different classification methods. TABLE-US-00022 TABLE F of discretized values for 114 HCCs using 3 different thresholds and 4 combinations Method 1 3-class: (1): <q30 Method 2 Method 3 Method 4 (2): q30 2-class: 2-class: 2-class: Over- Follow- q67; (1): <q30 (1): <q67 (1): <q50; all.survi- up Sample score (3): >g67 (2): >q30 (2): >q67 (2): >q50 val (years) HC 001 0.6875 2 2 1 1 1 0.07 HC 003 0.6875 2 2 1 1 1 3.33 HC 004 0.7272727 2 2 1 1 0 11.48 HC 006 0.8125 2 2 1 2 1 1.25 HC 007 1.4545455 3 2 2 2 1 1.5 HC 008 1.0769231 3 2 2 2 1 8.48 HC 009 1.75 3 2 2 2 1 0.02 HC 010 1.5 3 2 2 2 1 0.95 HC 011 0.6428571 1 1 1 1 0 12.2 HC 012 0.5714286 1 1 1 1 1 0.05 HC 014 0.625 1 1 1 1 1 1 HC 015 1.6 3 2 2 2 1 1.22 HC 017 1.875 3 2 2 2 0 10.96 HC 018 1.5 3 2 2 2 1 0.39 HC 020 0.7857143 2 2 1 1 0 15.4 HC 021 1.5555556 3 2 2 2 1 0.7 HC 022 0.5625 1 1 1 1 0 11.5 HC 023 0.5 1 1 1 1 0 11.93 HC 025 0.7142857 2 2 1 1 1 15.87 HC 026 0.7142857 2 2 1 1 1 0.83 HC 027 0.8125 2 2 1 2 1 0.1 HC 028 1 3 2 2 2 1 0.1 HC 030 1 3 2 2 2 1 12.4 HC 032 0.7857143 2 2 1 1 1 0.66 HC 034 0.625 1 1 1 1 0 15.7 HC 037 0.5714286 1 1 1 1 1 0.2 HC 038 1.0769231 3 2 2 2 1 1.12 HC 041 0.8666667 2 2 1 2 1 7.44 HC 042 0.8791209 2 2 1 2 0 10.58 HC 043 0.5 1 1 1 1 0 10.9 HC 052 1.3333333 3 2 2 2 NA 0.25 HC 058 1.875 3 2 2 2 0 8.3 HC 060 1 3 2 2 2 NA NA HC 064 0.8666667 2 2 1 2 1 5.25 HC 066 0.7142857 2 2 1 1 0 8.93 HC 101 0.9230769 2 2 1 2 0 2.5 HC 102 1.625 3 2 2 2 0 0.1 HC 103 0.75 2 2 1 1 0 1.82 HC 104 0.8666667 2 2 1 2 0 2.1 HC 105 1.4545455 3 2 2 2 0 0.56 HC 106 0.5 1 1 1 1 0 2 HC 107 0.8571429 2 2 1 2 0 1.75 HC 108 1 3 2 2 2 0 1.62 HC 109 0.5 1 1 1 1 0 1.3 HC 110 0.6923077 2 2 1 1 0 1.95 HC 111 1.1818182 3 2 2 2 1 0.7 HC 112 0.8666667 2 2 1 2 0 1.48 HC 113 1.1 3 2 2 2 1 1 HC 114 0.6666667 2 2 1 1 0 0.44 HC 115 0.875 2 2 1 2 0 0.75 HC 116 0.9333333 3 2 2 2 0 0.69 HC 117 0.6 1 1 1 1 0 1.2 HC 118 0.5 1 1 1 1 0 0.93 HC 119 0.8461538 2 2 1 2 0 1.2 HC 120 1 3 2 2 2 0 0.82 HC 121 0.9285714 3 2 2 2 0 0.6 HC 122 0.6666667 2 2 1 1 0 0.75 HC 123 1 3 2 2 2 0 0.8 HC 124 0.7857143 2 2 1 1 0 0.52 HC 125 0.8181818 2 2 1 2 0 0.9 HC 126 0.8125 2 2 1 2 0 0.42 HC 127 1.6 3 2 2 2 0 0.25 HC 128 0.6095238 1 1 1 1 0 0.44 HC 129 1 3 2 2 2 1 0.15 HC 130 1.7777778 3 2 2 2 0 0.14 HC 131 0.5625 1 1 1 1 0 0.26 HC 137 1.2222222 3 2 2 2 0 5.67 HC 138 0.75 2 2 1 1 0 5.58 HC 139 1.3333333 3 2 2 2 0 6 HC 140 0.5714286 1 1 1 1 0 4.17 HC 141 0.6153846 1 1 1 1 0 5.08 HC 142 0.8888889 2 2 1 2 1 4.08 HC 143 1.375 3 2 2 2 0 2.83 HC 144 0.6153846 1 1 1 1 0 6 HC 145 0.8 2 2 1 1 0 5.58 HC 146 0.9 2 2 1 2 0 4.33 HC 147 0.6666667 2 2 1 1 0 3.83 HC 148 1.1 3 2 2 2 0 3.08 HC 149 1.2222222 3 2 2 2 1 3.42 HC 150 0.6666667 2 2 1 1 0 5.42 HC 151 0.6153846 1 1 1 1 0 2.25 HC 152 0.6428571 1 1 1 1 1 3.67 HC 153 0.6923077 2 2 1 1 1 4.83 HC 154 1.375 3 2 2 2 1 2.21 HC 155 0.8181818 2 2 1 2 0 4.1 HC 156 1.4 3 2 2 2 1 2.31 HC 157 1 3 2 2 2 1 3.59 HC 159 0.7272727 2 2 1 1 1 2.42 HC 161 0.6 1 1 1 1 0 4.47 HC 162 1.1111111 3 2 2 2 0 3.49 HC 163 0.6 1 1 1 1 1 2.21 HC 164 0.6428571 1 1 1 1 0 4.54 HC 165 0.6428571 1 1 1 1 0 4.72 HC 168 0.6 1 1 1 1 0 6 HC 169 0.6 1 1 1 1 1 2.78 HC 170 0.5625 1 1 1 1 0 5.29 HC 171 0.8181818 2 2 1 2 0 4.57 HC 172 0.8333333 2 2 1 2 0 3.9 HC 173 0.6428571 1 1 1 1 0 4.21 HC 176 0.6428571 1 1 1 1 0 4.57 HC 177 0.6666667 2 2 1 1 0 5.42 HC 178 0.7142857 2 2 1 1 0 2.5 HC 179 0.8181818 2 2 1 2 0 5.17 HC 180 0.8571429 2 2 1 2 1 3.58 HC 181 1 3 2 2 2 0 6.83 HC 182 0.5625 1 1 1 1 0 3.5 HC 183 0.7333333 2 2 1 1 1 4.08 HC 184 0.9230769 2 2 1 2 1 2.08 HC 185 0.7692308 2 2 1 1 0 2.25 HC 186 0.9285714 3 2 2 2 1 2.17 HC 187 0.6428571 1 1 1 1 0 7.67 HC 188 0.7142857 2 2 1 1 0 4.67 HC 189 0.8666667 2 2 1 2 1 3.25 HC 190 0.7619048 2 2 1 1 0 5.58
[0296] Samples were separated into the corresponding subgroups, and subsequent analysis was carried out using the 4 classification methods. Survival for each group was determined using the Kaplan-Meier estimates and the log-rank test.
Statistical Analysis of Clinical Correlations with the Subclasses for 114 HCCs
[0297] A complete table with all clinical and pathological data collected for 114 HCC patients is given in Table G. The different parameters are represented as follows: TABLE-US-00023 TABLE H Clinical and pathological parameters and molecular classification of 114 HB cases. Characteristics Etiology.sup.†* Alcohol 40 (36%) HCV 26 (23%) HBV 23 (20%) Hemochromatosis 6 (5%) NASH 6 (5%) Unknown 23 (20%) Treatment (SR, OLT) 93/21 Chronic viral hepatitist.sup.† 46 (41%) Liver cirrhosis.sup.† 44 (48%) Tumor characteristics Macrovascular invasion.sup.† 20 (25%) Microvascular invasion.sup.† 47 (50%) Mean tumor size, cm (range).sup.† 7.9 (1.5-22) Multifocality 46 (48%) Histology: Edmonson Tumor grade.sup.† (1/2/3/4) 7/35/47/5 OMS Tumor differentiation (W/M/P) 51/55/6 Classification with 16-genes by discretization 40th Percentile (C1/C2) 30/84 50th Percentile (C1/C2) 55/59 67th Percentile (C1/C2) 77/37 Mean follow-up, months (range) 43.6 (0.26-146) Tumor recurrence.sup.† 43 (40%) Alive/DOD.sup.† 75/38 Abbreviations: HCV, hepatitis C virus; HBV, hepatitis B virus; NASH, Nonalcoholic steatohepatitis; SR, surgical resection; OLT, orthotopic liver transplantation; W, well differentiated; M, moderately differentiated; P, poorly differentiated; NA, not available; DOD, dead of cancer. *12 cases have more than one etiological agent and data were not available for 2 Gases. .sup.†Data were not available for all cases. Percentages were deduced from available data.
[0298] In a second step, the intrinsic parameters of the tumors correlated with patients' survival were analyzed. In this series of tumors, only tumor grade (Edmonson) and vascular invasion were significantly correlated with survival. TABLE-US-00024 TABLE I Summary of the clinical variables associated to overall survival (Kaplan-Meier curves and log-rank test). This Table does not take into account the molecular classification N. N. patients Log Variable patients Log rank With PH rank Edmonson Tumor grade 94 0.028 73 0.032 (1-2/3-4) Tumor diff. OMS 111 0.406 90 0.647 (Well/Moderate-poorly duff.) High proliferation: >10 45 0.054 34 0.402 Mitosis in 10 fields 40× (N/Y) Macrovascular Invasion 79 0.001 59 0.010 (N/Y) Microvascular Invasion 92 0.007 72 0.050 (N/Y) Tumor size≧10 cm 113 0.298 92 0.314
Classification by Discretization of Continuous Values
[0299] The clinico-pathological parameters were compared between the tumor groups using student's t test and chi-square test. Survival was analyzed by using Kaplan-Meier curves and log rank test. A special attention was given to the classification with the 67th percentile. Follow-up was closed at 146 months for overall survival (OS) and at 48 months for disease-free survival (DFS). TABLE-US-00025 TABLE J Association of 16-gene classification by discretization with clinical and pathological data (chi-square test). Abbreviations: P33, 33rd percentile, P50 50th percentile and P67, 67th percentile. p-value P67 Variable P33 P50 P67 C1 C2 comments Edmonson Tumor 0.006 <0.001 <0.001 38/27 4/25 20 cases with grade: grade 1 missing values and 2 (well differentiated) vs. 3 and 4 (moderately and poorly diff.) Tumor 0.006 0.001 <0.001 45/32 6/29 2 cases with missing differentiation values OMS (Well/Moderate- versus poorly differentiated) High proliferation: 0.021 0.001 0.001 22/7 4/12 >10 mitosis in 10 fields 40× (N/Y) Macrovascular 0.097 0.033 0.008 44/8 16/12 The cases defined as Invasion (N/Y) possible are considered negative. Microvascular 0.071 0.001 0.009 37/26 9/21 The cases defined as Invasion (N/Y) possible are considered negative. Tumor size ns ns 0.015 57/20 19/18 Different cut-offs </≧10 cm assessed: 2, 3, 5 and 10 cm Multifocality (N/Y) ns ns ns 35/30 15/16 Macronodules of ns ns ns 24/9 12/4 regeneration Norm Liver A0F0- ns ns ns 48/17 27/7 A0F1 Cirrhosis AXF4 ns ns ns 31/29 17/15 (N/Y) Score METAVIR 0.053 0.044 ns 19/32 5/20 Activity >0 (N/Y) Score METAVIR ns 0.20 ns 31/20 15/10 Activity >1 (N/Y) Score METAVIR 0.041 ns ns 5/48 2/27 Fibrosis >0 (N/Y) Score METAVIR ns ns ns 19/35 7/22 Fibrosis >1 (N/Y) Score METAVIR ns ns ns 24/30 8/21 Fibrosis >2 (N/Y) Score METAVIR ns ns ns 26/28 15/14 Fibrosis >3 (N/Y) Chronic viral 0.047 ns ns 48/29 18/17 hepatitis (N/Y) HBV (N/Y) 0.075 ns ns 62/15 27/8 HCV (N/Y) ns ns ns 61/16 25/10 Alcohol (N/Y) ns ns ns 47/30 25/10 Recurrence (N/Y) ns ns ns 41/32 24/11 HCC034 and HCC030 censored Survival (N/Y) 0.050 0.023 0.031 56/21 19/17 HCC025 and HCC030 censored DFS (N/Y) ns ns ns 35/42 15/21 HCC025 and HCC030 censored
[0300] In conclusion, these data show significant correlations between molecular classification using the 3 methods and the following parameters: Tumor grade (Edmonson), tumor differentiation (OMS), proliferation rate, vascular invasion and survival. In contrast, the classifications were not correlated with etiological factors (viral hepatitis, alcohol, etc. . . . ), with the state of the disease in adjacent, non tumoral livers or with tumor recurrence.
The data suggest that classification using the 67th percentile seems to be the most adequate and is strongly recommended to classify HCCs.
Multivariate Analysis
[0301] To further determine the efficiency of the molecular classification using the 67th percentile, we performed multivariate analysis with the Cox regression test on two sets of patients for which all data were available:
[0302] 91 patients that received either surgical resection or orthoptic liver transplantation (OLT)
[0303] 71 patients that received surgical resection.
[0304] Different variables associated to survival in the clinical settings have been included in the multivariate analysis: 1) Edmonson grade, 2) microvascular invasion and 3) Molecular classification using the 67th percentile. TABLE-US-00026 TABLE K Multivariate test (Cox regression). N patients variable HR 95% CI p-value 91 Molec classsif (p67) 2.534 (1.214-5.289) 0.016 (surgical Edmonson Tumor grade 1.690 (0.747-3.823) 0.205 resections 1-2/3-4) and OLT) Microvascular Invasion 2.451 (1.105-5435) 0.024 (N/Y) 71 Molec classsif (p67) 2.646 (1.1156.278) 0.032 (only Edmonson Tumor grade (1- 2.697 (1.103-6.592) 0.026 surgical 213-4) resections) Microvascular Invasion 1.681 (0.648-4.359) 0.282 (N/Y)
Correlation of the Molecular Classifications with Survival
[0305] For overall survival (OS) and disease-free survival (DFS), we compared the efficiency of the 3 methods of discretization that separate the samples into 2 subclasses. Independent studies were made for patients that received surgical resection and for patients that received orthoptic liver transplantation (OLT). The ability of the 16-gene signature to discriminate between recurrent and non-recurrent tumors was also assessed. TABLE-US-00027 Table L Summary of survival analysis using Kaplan-Meier curves and log-rank test Analysis N. patients Classif. method Log rank OS 113 P33 0.037 113 P50 0.005 113 P67 0.002 DFS 113 P33 0.078 113 P50 0.019 113 P67 0.072 recurrence 108 p33* 0.134* 108 p50* 0.115* 108 P67 1.000 Analysis of 92 cases that received surgical resection OS 92 P33 0.032 92 P50 0.009 92 P67 0.013 DFS 92 P40 ns 92 P50 ns 92 P67 ns recurrence 88 P33 ns 88 P50 ns 88 P67 ns Abbreviations: OS, overall survival; DFS, disease free survival *There is a trend but it is not significant and it is lost in the P60 analysis
[0306] The different analyses are illustrated in the Kaplan-Meier plots shown in FIG. 12. The discretization method of classification showed the same efficiency in the analysis of tumors obtained either from surgical resection (also called partial hepatectomy, PH) or from orthotopic liver transplantation (OLT), showing that the clinical management of the tumor had no impact on the classification.
[0307] In conclusion, the method described herein is able to classify HCC cases according to tumor grade and patient's survival, and represents a powerful tool at diagnosis to stratify the tumors according to the prognosis, and for further clinical management of HCC. In particular, it may be an excellent tool for the decision of orthotopic liver transplantation, since the criteria used currently are limited and often poorly informative of the outcome.
Protocol for Applying the Method to a New Sample
[0308] The following protocol is designed according to the invention:
1--extract total RNA from the tumor specimen using well established technologies.
2--synthesize cDNA synthesis (suggested conditions: 1 μg RNA and 300 ng of random hexamers for a 20 μl-reaction)
[0309] 3--amplify the selected genes said genes being in equal number of each of the groups defined as overexpressed proliferation-related genes group and downregulated differentiation-related genes group (profiled genes within the group of 2 to 16 genes) and the reference gene (invariant gene) such as for example the RHOT2 gene 1:5 cDNA dilution, using either Taqman or SybrGreen qPCR technology.
4--determine the Delta Ct (DCt) value for each gene
5--compare the value with the threshold of reference (for HB or for HC) in order to assign a discretized value of "1" or "2".
[0310] 5--determine the average of discretized values in each group, i.e., for the selected proliferation-related genes (up to 8) separately for and the selected differentiation-related genes (up to 8) and determine the ratio of these 2 average values which is the score of the sample.
6--compare the result with the reference scores corresponding to the following cut-offs:
C1
[0311] |30rd=0.6667
[0312] |50th=0.8125
[0313] |67th=0.925
C2
Example
[0314] For patient X having an HC tumor a Taqman qPCR is performed.
[0315] Step one: assignment of discretized values to each selected gene among proliferation-related genes and differentiation-related genes.
Example
The DCt of AFP is -4.0523
The cut-off for AFP for qPCR using Taqman technology is -1.2634010 Given that -4.0523 is lower than the cut-off, the assigned discretized value is 2.
[0316] Step two: Determination of the average of discretized values for the 2 sets of 8 genes:
AFP=2; BUB1=1; DLG7=2; DUSP9=2; E2F5=2; IGSF1=1; NLE=2; RPL10A=1;
Average of Proliferation-Related Genes: (2+1+2+2+2+1+2+1)/8=1.625
ALDH2=1; APCS=1; APOC4=1; AQP9=1; C1S=2; CYP2E1=2; GHR=1; HPD=2;
Average of Differentiation-Related Genes: (1+1+1+1+2+2+1+2)/8=1.375
[0317] Step Three: calculate the ratio proliferation/differentiation score.
In this example: 1.625/1.375=1.18182
[0318] Step 4: compare the result with the reference scores:
C1
[0319] |30rd percentile=0.6667
[0320] |50th percentile=0.8125
[0321] |67th percentile=0.925
C2
Classification based on the value of the ratio=1.18182.
[0322] As the value is above the 67th percentile, the assigned class is C2. TABLE-US-00028 TABLE A id AFP ALDH2 APCS AP0C4 AQP9 BUB1 C1S CYP2E1 HB1 -7.684892 -4.592702 -0.660189 -2.651319 -4.194894 -1.068025 -1.394659 -3.334692 HB100 -7.682724 -3.849128 -0.372566 0.297278 -0.305738 0.65983 -2.572264 -7.352142 HB101 1.801478 -7.157316 -1.166513 -4.924476 -8.067838 6.222865 -5.284734 -11.757699 HB102 -7.761115 -5.696697 -1.044129 -2.374592 -3.447046 2.724363 -3.657616 -5.769417 HB103 2.908026 -2.580629 -2.748625 -2.55635 1.480624 3.891875 -2.819372 0.454623 HB106 0.294848 -7.534485 -1.424535 -5.377043 -7.886612 4.855797 -6.80698 -11.496242 HB107 0.719866 -6.546079 -9.18522 -3.425075 -6.189664 3.901806 -5.609115 -10.6711555 HB11 1.492805 -3.560021 -5.094387 -1.031623 -8.42849 2.086834 -6.166353 -9.043371 HB112 4.155252 -6.486961 -0.154814 -4.48155 -5.634596 3.762347 -7.88579 -8.960815 HB114 6.2971 -3.966456 5.02266 0.604275 3.037682 4.23408 -5.29691 -0.313326 HB118 0.318307 -4.311795 -5.146409 -3.787568 -5.428442 2.329959 -5.284827 -7.342423 HB121 -0.971033 -6.879043 -8.355819 -4.679393 -6.361435 2.329708 -6.559457 -8.87105 HB122 2.188721 -6.220957 -7.7399 -3.410743 -5.745306 3.309004 -6.327656 -8.906339 HB125 2.929931 -4.053616 -4.882212 -2.32494 -3.352398 5.067815 -4.255762 -7.887455 HB126 2.458273 -5.577951 -6.518289 -3.182407 -5.243351 5.270089 -5.814672 -8.188307 HB129 -4.930877 -2.124281 -0.744262 1.154663 -0.846572 0.421372 -2.925458 -4.708874 HB130 -4.86199 -1.139837 -1.398588 0.115559 -1.313951 1.669543 -2.37235 0.175598 HB131 5.545406 -1.714367 -1.045683 2.628822 1.903853 1.972112 -2.306818 0.069456 HB132 2.654369 -3.71955 -6.543987 -3.876868 -4.7099 4.043489 -4.801651 -7.725089 HB136 5.005516 -3.234557 -4.827283 2.471208 -0.502385 -1.945351 -4.324749 -4.844765 HB140 2.835457 -7.041546 -6.88604 -5.561912 -5.089682 4.140594 -6.023758 -10.477228 HB142 5.200474 -4.919616 2.416807 2.058522 -3.396171 1.380591 -5.965126 1.196438 HB145 3.58286 -5.186236 -5.18731 NA -5.118895 5.58416 -5.786933 -7.880334 HB146 -1.290056 -5.422341 -5.973879 -3.869993 -5.908024 0.982626 -4.124487 -8.751883 HB147 -9.442257 -3.655303 -0.362122 1.179633 -2.349782 -1.51351 -2.756099 0.30832 HB148 -3.566401 -5.382548 -6.721533 -2.380348 -6.951359 1.183916 -4.188648 -7.101147 HB150 2.356994 -5.56181 -5.496186 -4.45536 -5.603247 5.136577 -5.435261 -8.522001 HB153 -2.086302 -4.364035 -4.049735 -1.1908 -4.342186 2.437297 -6.055092 -7.522683 HB155 -1.951256 -5.140738 -7.17357 -0.801318 4.538929 4.038538 -5.939438 3.058475 HB156 -6.523604 -4.658012 -5.112322 -1.499462 -1.13031 1.970226 -4.763811 -8.138508 HB157 -8.747252 -3.193287 -0.914511 0.563787 -0.139273 0.648195 -3.089302 -2.404646 HB160 4.40621 -0.878277 -2.381785 -1.9527 0.770799 4.516203 -2.89522 1.197611 HB162 -1.127062 -5.142195 -6.564426 -2.432348 -5.179601 3.27157 -4.959578 -9.351464 HB165 -1.015428 -1.578048 -1.612095 -1.677494 1.921123 -0.416058 -4.579384 -0.458984 HB167 -7.323435 -5.692388 -6.461153 -2.470512 -4.912208 -0.369976 -4.949694 -10.583324 HB170 -0.980072 -5.786627 -7.265156 -3.690367 -5.952908 1.548967 -6.61768 -8.574004 HB171 2.310988 -5.687635 -7.127181 -3.794631 -5.898635 2.05689 -6.420469 -8.856566 HB172 4.547243 -0.385469 -1.804453 -1.833478 2.11442 4.373205 -3.929151 1.277285 HB173 1.889759 -5.184791 -4.471618 -2.235657 -5.743057 2.116789 -4.966413 -7.319851 HB175 -2.0436 -6.05152 -8.152949 -2.996302 -3.829205 3.036838 -5.151913 -9.108766 HB184 -6.561121 -2.895788 -5.35813 -1.653786 0.293844 -0.082754 -3.084271 -3.362889 HB20 4.752153 -4.811256 -5.712608 -2.133951 -5.361771 5.572378 -4.283688 -8.390209 HB28 -4.001793 -4.719296 -7.514733 -2.385516 -3.869707 0.599685 -5.187286 -9.373678 HB3 0.027392 -4.565046 -4.462833 -2.255273 -4.14636 4.676108 -5.373064 -6.610781 HB33 -7.497741 -3.066759 -5.881277 0.250334 0.950966 0.500246 -3.829096 -6.510795 HB39 -8.613403 -3.166427 3.421734 1.699859 -0.944463 -0.146929 -1.480822 -0.727464 HB48 -4.768603 -3.632136 -4.882397 -2.170561 -4.965403 1.366439 -3.944489 -9.061667 HB49 1.818606 -5.933777 -5.948111 -4.936781 -5.434931 4.576628 -5.318794 -9.381172 HB5 -2.282703 -6.147963 -7.059143 -4.107155 -7.593099 2.501017 -6.573836 -9.813634 HB54 1.132255 -4.844075 -5.655802 -2.937193 -4.595442 3.040468 -4.999207 -8.199672 HB59 1.334928 -6.792009 -7.221196 -5.590302 -6.300828 1.42553 -5.648808 -9.279234 HB6 -1.610623 -7.099329 -7.979286 -5.729452 -5.2647225 2.920021 -5.482511 -10.151809 HB60 -0.594337 -5.206398 -6.67766 -1.663871 -2.889326 3.97632 -5.504179 -6.743858 HB61 -5.058775 -6.113525 -5.991888 -3.527984 -5.387419 3.269827 -6.119246 -8.943929 HB62 -1.989342 -4.487171 -6.502588 -0.923844 -4.712471 3.449967 -4.22945 -7.087853 HB63 -0.891056 -4.153057 -5.680458 -2.637115 -5.710062 4.49543 -2.939154 -9.095241 HB65 3.025127 -4.346225 -5.338104 -1.175748 -1.226393 -0.613979 -5.196916 -4.645702 HB66 -1.861761 -4.166485 -5.897819 -2.09279 -3.003258 4.774807 -4.585607 -6.839392 HB68 -4.313608 -6.550704 -6.762513 -3.66757 -5.982654 4.060667 -5.956246 -8.393607 HB69 -1.820363 -9.245314333 -8.965648 -7.384871667 -9.430164667 -2.026701667 -8.961309 -12.31658 HB7 1.334084 -4.488213 -5.853708 -2.13753 -5.142938 4.894117 -4.082335 -8.118103 HB70 2.021391 -5.678476 -7.496267 -5.781771 -4.346458 2.174971 -7.066038 -8.392057 HB72 -11.99570467 3.978023333 -1.371737333 -2.543168667 -6.278723667 -5.504070333 -7.162789667 -8.103601333 HB73 -10.69629133 -8.263771333 -4.869197667 -2.900671333 -5.802080667 -5.324255333 -8.090371 -9.754354333 HB74F 3.831288 -7.73216 -4.940396 -6.3439 -6.355995 6.130615 -5.584023 -10.472842 HB75 0.474553 -6.309769 -2.777247 -4.334006 -6.807299 4.545387 -5.115577 -10.418948 HB77 2.915987 -5.645872 -6.698372 -2.284956 -5.392377 4.544876 -5.559466 -8.695429 HB78 -3.945686 -2.82555 -2.986284 -1.790335 -0.938738 4.523136 -2.620165 -5.945013 HB79 -0.781193 -5.652768 -5.454157 -3.953162 -5.051444 0.254305 -5.44242 -9.05667 HB8 -6.696169 -3.108913 0.498461 1.361801 -3.322642 0.055848 -0.348492 -1.877119 HB80 -8.8331005 -4.713883 -2.9124615 -2.810437 -0.838727 -0.7226515 -2.5925445 -5.408417 HB81 -4.851198667 -10.55296467 -10.55292033 -7.621321667 -10.19195633 -2.962795333 -10.17992067 -12.72629433 HB82 -1.942166 -5.620028 -5.739178 -3.972123 -6.520482 0.934055 -3.737063 -8.932744 HB83 -4.169107 -9.660034667 -9.382586667 -8.05219 -10.951863 -3.521245667 -10.12345167 -9.850559667 HB86 -6.283735 -5.287677 0.896101 -1.494853 -2.934412 -0.46896 -2.879366 -5.76077 HB89 2.996384 -7.323446 -7.464817 -5.120874 -5.856518 4.907738 -6.676481 -9.415603 HB9 -3.679937 -4.761778 -6.571455 -2.775269 -6.201772 2.209541 -3.895565 -8.86438 HB90 2.024206 -8.47846 -1.33932 -6.745716 -6.677122 5.899195 -8.114672 -10.459034 HB93 -4.610162 -5.583852 -5.277197 -1.990982 -2.698011 -1.085743 -4.488914 -3.388975 HB94 1.79868 -5.621254 -7.718202 -6.940586 -6.67335 3.551727 -6.54809 -8.572742 HB95 -0.444835 -5.745006 -8.404602 -5.637613 -6.396063 6.671045 -5.701559 -10.554918 HB96 -4.775396 -6.402052 -6.123253 -4.340961 -5.066688 3.365736 -6.521753 -9.090145 HB97 -6.841231 -6.21691 -6.275051 -3.638382 -3.617558 2.362203 -6.58495 -5.781372 HB98 -4.911783 -2.946932 6.478933 4.211147 0.395926 2.311268 -2.827802 0.584022 HB99 -4.551378 -1.14591 -5.549696 -1.796859 1.62906 2.600714 -2.483835 -3.848236 HB1 4.140368 -5.212318 -0.812424 1.207583 3.840983 -0.715134 -0.812792 -8.675945 HB100 4.399124 -5.749706 0.27698 1.907294 -0.113253 -2.800323 0.547899 -6.153046 HB101 7.086329 -0.641871 0.737702 -3.913751 -4.340259 7.086329 0.191689 -6.757648 HB102 7.380694 -4.303866 1.144778 0.2784 -0.284245 -2.545668 0.856607 -6.803817 HB103 5.997143 0.880421 3.697478 1.249386 -2.713306 1.392197 -0.453035 -4.535615 HB106 6.79755 -1.540745 0.77722 -4.155098 -5.747164 2.274385 0.291903 -6.637275 HB107 5.239962 -1.184244 3.145996 -1.891404 -4.433271 3.119114 -0.053334 -6.319917 HB11 3.688558 -1.412987 -0.179621 -0.149048 -1.897658 2.297186 -0.19686 -5.623341 HB112 6.035002 -2.179125 -0.998979 -3.575994 -4.671755 -0.776138 -2.252113 -7.8479 HB114 6.2971 2.615827 0.886564 0.002487 1.919397 2.50863 1.785623 -7.055851 HB118 3.935101 2.405105 2.275962 -0.451819 -4.812319 2.339813 0.486307 -5.904633 HB121 3.458157 -2.1882 1.247645 -1.155575 -5.938235 3.750147 1.867907 -5.131548 HB122 3.562777 1.229723 2.386559 -1.961029 -5.590919 2.406687 1.976893 -5.368023 HB125 5.700252 0.274642 2.864883 0.118717 -3.155289 2.138032 -0.470879 -3.478449 HB126 6.32602 0.274197 3.089709 -1.334371 -5.227705 2.726599 0.54385 -4.787822 HB129 4.474485 -3.829751 1.158283 3.025728 1.984295 -0.074354 1.326073 -5.682215 HB130 5.297728 -2.554008 2.251163 3.317556 0.885962 0.039307 1.389742 -4.829542 HB131 5.801168 2.269272 2.226921 1.235598 2.035452 5.621114 1.777334 -4.96776 HB132 8.18041 0.433104 4.507503 -0.157093 -2.441422 5.855213 2.895208 -3.579579 HB136 1.140686 0.10165 -2.336947 0.261203 0.124159 3.807218 -0.676358 -7.113232 HB140 9.015818 -0.401264 2.325356 -3.379816 -3.148068 3.156456 0.80129 -7.308986 HB142 6.203192 4.554631 3.03661 2.598877 4.150455 8.782461 1.428955 -6.630178 HB145 6.734264 1.908734 2.518779 -1.358174 -5.181668 4.610406 1.707345 -4.6775 HB146 0.991164 -0.681828 0.1227 -0.510651 -4.471483 0.777004 0.176935 -5.992209 HB147 -1.376061 -4.733546 -2.588397 1.772494 -1.944032 -2.698708 -0.565682 -7.527854 HB148 1.7033 -1.806502 -0.663069 -1.376372 -5.121145 -0.683001 -0.431826 -6.201895 HB150 5.800233 0.8436 2.758596 -1.181738 -5.492037 2.891937 0.439392 -4.69542 HB153 3.096912 -2.657862 0.449197 -0.480929 -4.261986 3.34336 1.423023 -5.963837 HB155 4.360922 -1.23259 0.752365 -3.062474 0.657144 -1.091013 0.911424 -5.964497 HB156 2.483547 -1.214228 0.687246 -1.107338 -3.806189 -1.181305 0.159847 -5.65452 HB157 0.181175 -4.1451 0.297747 1.940187 -3.850885 -1.38623 0.041349 -5.820536 HB160 6.224569 2.906158 4.403545 2.633949 -2.138569 3.355814 -0.100123 -4.568688 HB162 4.25017 -1.453283 1.117439 -0.163468 -4.733881 1.809885 -0.022627 -4.822098 HB165 -0.010488 1.837305 0.47467 -2.953007 -0.655058 -1.791164 -0.933062 -5.535221 HB167 0.509668 -1.707485 0.198742 0.269552 -4.442331 -1.197651 -0.240385 -5.755341 HB170 2.567207 1.148738 1.360144 -2.397242 -4.944439 2.424619 -0.463297 -5.539725 HB171 2.278353 1.67404 2.062277 -1.193735 -4.984552 2.19098 0.230044 -4.81411
HB172 6.060459 2.366999 3.689341 2.93017 -1.316921 2.571021 -0.153162 -3.812616 HB173 2.779999 1.921427 3.05205 -0.20919 -4.475376 0.418818 0.678606 -4.361307 HB175 4.414558 -1.623242 1.49 -0.662783 -4.684446 3.524049 1.78088 -5.173616 HB184 1.361379 -1.542307 -0.588812 1.814793 -2.048922 -0.326393 0.097971 -4.663763 HB20 9.423325 -0.34174 2.066057 -0.975735 -3.695854 4.361484 1.157495 -5.27136 HB28 1.922989 -2.304861 1.222545 -0.120436 -5.154703 -0.192738 1.819854 -5.824864 HB3 7.285685 0.65201 2.301029 -0.049158 0.117373 4.46221 1.743745 -6.911792 HB33 1.659659 -4.338262 -0.148233 1.134133 -4.625204 -2.34198 1.272614 -5.63922 HB39 2.485354 -4.927491 -1.241931 1.694781 -0.33289 -2.652634 -0.149609 -6.579218 HB48 1.583391 -3.620772 -0.089081 1.342382 -2.330218 0.686163 1.169838 -6.508074 HB49 5.652893 2.41148 3.776672 -1.220476 -5.746779 4.727596 2.190021 -4.286949 HB5 3.674234 -2.082424 0.98073 -1.943451 -6.561791 1.592167 0.449005 -6.230808 HB54 3.556268 3.982183 3.025795 -0.158057 -4.638333 3.623678 1.995039 -5.061096 HB59 5.127336 0.250753 3.459226 -2.269072 -4.727738 6.045093 1.466312 -6.48303 HB6 6.733353 -0.246309 3.812183 -2.459856 -3.728987 0.835057 2.205872 -7.208765 HB60 5.188517 2.869544 3.228365 -0.276338 -4.031974 2.026116 2.577353 -4.502382 HB61 5.827933 -5.51457 1.00606 -3.272672 -4.816797 -0.203871 0.753758 -6.140918 HB62 4.328277 0.708512 1.218963 1.021692 -3.265138 0.731519 2.223877 -5.334147 HB63 5.003075 -1.082094 0.951357 1.316553 2.000601 4.964996 1.31674 -6.741518 HB65 2.978487 -0.087486 -1.274388 0.080222 -2.417946 1.06702 -1.371523 -6.195428 HB66 8.039274 -0.423313 2.141981 -1.148424 -1.349111 -0.305017 1.586659 -5.393141 HB68 7.010986 -0.530541 2.520261 0.232431 -1.779051 -0.603113 2.342104 -4.959414 HB69 3.071106 -0.626059667 3.421015 -5.118794333 -6.824055667 11.819556 -0.603036 -2.847600667 HB7 8.076437 -0.833011 1.354912 -0.884629 -2.106592 2.978739 2.384133 -5.458546 HB70 4.083519 3.896364 2.616204 -3.614294 -6.063097 2.060379 1.506083 -4.669554 HB72 -1.688566667 -8.976227 -1.809694 -1.750672 -3.40203 -6.090071333 -2.505424 -5.054027 HB73 -2.068555667 -9.537516 -1.965151 -0.544775 -5.542041333 -7.013002667 -3.078154667 -5.580986333 HB74F 8.986048 0.497828 4.585503 -2.916191 -3.041943 7.759608 1.654283 -6.380865 HB75 7.231393 -2.411839 0.378995 -1.925637 -5.055106 2.61456 1.017432 -5.77539 HB77 9.66177 -0.139299 2.727198 -1.675013 -4.079932 2.793758 2.146337 -4.964228 HB78 5.293419 -0.185629 1.735594 0.020191 -3.984125 -2.010153 -0.114956 -3.94071 HB79 1.90306 1.145681 1.319285 -1.978228 -5.757335 0.01942 -0.194167 -5.016158 HB8 1.950257 -4.043236 -1.814636 2.280516 1.100353 0.314694 0.29834 -7.823095 HB80 2.660644 -4.9166885 -0.374031 0.675995 -0.4253495 -4.2048885 -0.8782055 -7.919531 HB81 2.155925333 -5.738363 0.932455333 -5.565798 -8.171378 -1.999123333 -2.092100667 -4.795482 HB82 1.47049 -3.938165 -0.549544 -1.023595 -3.267403 8.008069 0.067941 -7.635394 HB83 2.492243 -4.003930333 4.737920667 -4.561133333 -6.966227667 -0.028684333 -0.855054667 1.789090833 HB86 3.219092 -5.894534 -0.496662 0.35847 -0.121981 -2.31061 0.046472 -8.510995 HB89 8.255339 1.284916 3.638735 -2.665258 -5.177704 3.273649 1.279167 -5.898171 HB9 4.940411 -1.989636 0.700504 -0.698988 -3.255601 2.609339 1.300875 -6.54224 HB90 6.54891 1.104162 1.408459 -5.754423 -7.507485 4.45026 1.52717 -6.250036 HB93 3.902565 -7.483471 -0.488108 0.969648 -1.415501 -1.818147 -0.829773 -7.824402 HB94 8.669386 -1.132305 0.490788 8.498726 -6.819645 7.800646 -0.149162 -5.793072 HB95 6.921267 -1.620869 2.726241 -2.193777 -5.454765 1.364738 0.279802 -5.172451 HB96 6.685021 -0.591271 1.973021 -4.924202 -4.91283 1.722505 1.829525 -5.638435 HB97 6.474525 -5.800537 1.05047 -0.911789 -4.571465 -4.308964 -0.87035 -6.60257 HB98 6.837198 -2.065483 2.482301 1.17723 -0.98407 -0.701098 1.175939 -5.166874 HB99 6.353711 -4.201828 1.467552 1.703655 -0.109186 -0.822266 1.226265 -3.572067
[0323] TABLE-US-00029 TABLE B 67th percentile- percentile- related related previous 16 gene tumor score score score based clas- AFP at diagnosis Treatment PRETEXT ID (ratio) (2-classes) (3-classes) sification Gender Age months ng/mL treatment protocol stage HB122 0.5 1 1 C1 M 10 8000 Y H I HB126 0.5 1 1 C1 F 12 153840 Y S II HB145 0.5 1 1 C1 M 7 56000 Y S II HB150 0.5 1 1 C1 F 5 82000 Y S III HB175 0.5 1 1 C1 M 9 220000 Y S I HB20 0.5 1 1 C1 F 50 880 Y S II HB49 0.5 1 1 C1 F 15 11000 Y S II HB54 0.5 1 1 C1 M 10 180 N N I HB70 0.5 1 1 C1 F 42 812 Y S II HB77 0.5 1 1 C1 F 9 204000 Y S II HB89 0.5 1 1 C1 M 13 448 Y S I HB95 0.5 1 1 C1 M 28 1000000 Y H IV HB118 0.53333333 1 1 C1 M 17 14500 Y S NA HB132 0.53333333 1 1 C1 F 23 2078 Y NA III HB121 0.5625 1 1 C1 F 14 296000 Y S III HB140 0.5625 1 1 C1 M 3 22758 Y S II HB162 0.5625 1 1 C1 F 9 960000 Y S III HB171 0.5625 1 1 C1 F 17 300 Y S II HB173 0.5625 1 1 C1 F 27 66810 Y S I HB59 0.5625 1 1 C1 F 24 5643 Y S II HB6 0.5625 1 1 C1 M 24 320000 Y S II HB74F 0.5625 1 1 C1 M 96 325 N N I HB96 0.5625 1 1 C1 M 101 2265000 Y H IV HB60 0.57142857 1 1 C1 F 30 1990800 Y H II HB7 0.57142857 1 1 C1 M 33 45000 Y S I HB101 0.6 1 1 C1 M 42 67747 Y S III HB106 0.6 1 1 C1 F 11 320000 Y H IV HB90 0.6 1 1 C1 F 74 300 N N II HB62 0.61538462 1 2 C1 M 16 1708400 Y H IV HB107 0.625 1 2 C1 M 30 16000 Y H IV HB170 0.625 1 2 C1 M 20 123000 Y H III HB5 0.625 1 2 C1 M 84 300000 Y H III HB125 0.64285714 1 2 C1 F 15 360000 Y H IV HB75 0.66666667 1 2 C1 M 21 131000 Y S II HB9 0.66666667 1 2 C1 F 16 84000 Y NA III HB94 0.66666667 1 2 C1 M 29 1270 Y S I HB61 0.6875 1 2 C1 F 126 346000 Y NA IV HB69 0.6875 1 2 C1 M 25 1163 Y S I HB79 0.6875 1 2 C1 M 144 1200 Y S II HB3 0.69230769 1 2 C1 F 22 3192 Y S I HB66 0.69230769 1 2 C1 M 6 1000000 Y S III HB68 0.71428571 1 2 C1 F 11 119320 Y S III HB146 0.73333333 1 2 C1 F 11 NA N N NA HB155 0.75 1 2 C2 M 9 849500 Y S II HB63 0.75 1 2 C1 M 204 NA N N III HB11 0.76923077 1 2 C1 F 18 626100 Y H IV HB153 0.78571429 1 2 C1 F 27 1000000 Y H IV HB28 0.8125 1 2 C1 M 34 172500 Y NA II HB83 0.8125 1 2 C1 M 15 285 Y S II HB156 0.85714286 1 2 C2 F 2 468000 Y S III HB112 0.86666667 1 2 C1 M 36 725 Y S II HB82 0.86666667 1 2 C1 M 120 179000 N N II HB97 0.86666667 1 2 C1 F 42 700000 Y H IV HB81 0.875 1 2 C1 M 22 322197 Y H III HB103 0.9 1 2 C2 F 57 750000 Y H IV HB114 0.9 1 2 C2 F 21 8783 Y S II HB142 0.90909091 1 2 C2 F 48 605000 Y H III HB148 0.93333333 2 3 C1 M 17 200730 Y S II HB167 0.93333333 2 3 C2 M 34 1500000 Y H NA HB73 0.9375 2 3 C2 F 24 667786 Y H III HB131 1 2 3 C2 M 6 7511 Y H II HB65 1 2 3 C2 M 6 1740 N N III HB78 1 2 3 C1 M 126 376000 Y S II HB72 1.07142857 2 3 C2 F 16 1412000 Y S III HB48 1.07692308 2 3 C2 M 72 35558 Y H IV HB102 1.09090909 2 3 C2 M 41 1331000 N N II HB160 1.125 2 3 C2 M 45 342000 Y H II HB172 1.125 2 3 C2 M 50 64170 Y H II HB99 1.22222222 2 3 C2 M 72 277192 N N IV HB130 1.25 2 3 C2 F 19 1980000 Y H II HB98 1.25 2 3 C2 M 60 1285000 Y H III HB136 1.3 2 3 C2 M 6 31828 Y S III HB165 1.3 2 3 C2 M 13 18600 Y S II HB1 1.36363636 2 3 C2 F 43 3000 Y H IV HB93 1.36363636 2 3 C2 M 22 107000 Y S III HB129 1.375 2 3 C2 M 96 14000 N N I HB33 1.4 2 3 C2 M 12 765890 Y H IV HB100 1.44444444 2 3 C2 M 48 576000 N N III HB184 1.44444444 2 3 C2 M 41 912500 Y H IV HB157 1.55555556 2 3 C2 M 7 356000 Y H NA HB80 1.6 2 3 C2 M 180 37000 Y H III HB86 1.66666667 2 3 C2 M 0.08 74000 N N III HB8 1.75 2 3 C2 F 8 44610 Y NA II HB147 2 2 3 C2 F 9 2355000 Y S II HB39 2 2 3 C2 F 11 862067 Y S III Main Epi- thelial beta- tumor Distant Vascular Multi- Histol- com- catenin Follow-up Surgery Follow-up ID Metastasis invasion focality ogy ponent status (months) Outcome Type speOS (years) HB126 N N S Mx F mut 18 A R 0 1.5 HB145 N N S Mx F mut 17 A R 0 1.416666667 HB150 N N S Mx F mut 14 A R 0 1.166666667 HB175 N N M Mx F NA 6 A R 0 0.5 HB20 N N M Mx F mut 7 A R 0 0.583333333 HB49 N N S Ep F mut 42 A R 0 3.5 HB54 N N S Ep F mut 6 D R 0 0.5 HB70 N N S Ep PF mut 49 A R 0 4.083333333 HB77 N N S Ep PF mut 53 R R 0 4.416666667 HB89 N N S Ep F mut 37 A R 0 3.083333333 HB95 N N S Ep F mut 33 A R 0 2.75 HB118 Y Y M Mx F mut 32 A LT 0 2.666666667 HB132 N N S Mx F mut 121 A R 0 10.08333333 HB121 N N M Mx F mut 18 A R 0 1.5 HB140 N N S Mx F mut 22 A R 0 1.833333333 HB162 N N S Mx F mut 13 A R 0 1.083333333 HB171 N N S Ep F mut 9 A R 0 0.75 HB173 N N S Ep F NA 11 A R 0 0.916666667 HB59 N N S Ep PF mut 72 A R 0 6 HB6 N Y S Ep F mut 48 A R 0 4 HB74F N Y S Ep F mut 35 A R 0 2.916666667 HB96 N Y M Ep F mut 23 R LT 0 1.916666667 HB60 N Y S Ep F wt 63 A R 0 5.25 HB7 N Y S Mx F mut 46 A R 0 3.833333333 HB101 N N S Ep F mut 20 A R 0 1.666666667 HB106 N N S Mx F mut 25 A R 0 2.083333333 HB90 N N S Ep F mut 35 A R 0 2.916666667 HB62 N N S Mx F mut 69 A R 0 5.75 HB107 Y Y M Ep F mut 25 A LT 0 2.083333333 HB170 Y Y M Ep F wt (FAP) 15 A R 0 1.25 HB5 Y Y M Ep F mut 24 DOD R 1 2 HB125 Y N M Mx F mut 17 A LT 0 1.416666667 HB75 N Y S Mx F mut 41 A R 0 3.416666667 HB9 N N S Ep PF mut 91 A R 0 7.583333333 HB94 N N S Ep PF wt 29 A R 0 2.416666667 HB61 Y Y M Mx F mut 5 DOD R 1 0.416666667 HB69 N N S Ep PF wt 55 A R 0 4.583333333 HB79 N N M Ep M mut 39 A LT 0 3.25 HB3 N N S Ep F wt 55 A R 0 4.583333333 HB66 N N S Ep F mut 68 A R 0 5.666666667 HB68 N N S Mx E mut 52 A R 0 4.333333333 HB146 N NA S NA NA NA 1 D R 0 0.083333333 HB155 N N S Mx CF mut 8 A R 0 0.666666667 HB63 N Y M Mx F mut 96 A R 0 8 HB11 Y Y M Mx F mut 21 DOD R 1 1.75 HB153 Y N M Mx CF mut 8 A LT 0 0.666666667 HB28 N N S Ep F wt 120 A R 0 10 HB83 N N S Ep PF mut 53 A R 0 4.416666667 HB156 N N NA Ep F NA 6 A R 0 0.5 HB112 N N S Ep F wt 32 A R 0 2.666666667 HB82 N N S Ep F mut 63 A R 0 5.25 HB97 N Y M Ep F mut 30 A R 0 2.5 HB81 Y Y M Ep F mut 36 A R 0 3 HB103 Y Y M Ep M mut 9 DOD M 1 0.75 HB114 N N S Mx E mut 23 A P 0 1.916666667 HB142 Y Y S Ep NA mut 16 A R 0 1.333333333 HB148 N N S Mx F mut 11 A R 0 0.916666667 HB167 Y Y M Ep F mut 2 A R 0 0.166666667 HB73 Y Y S Ep E mut 16 DOD R 1 1.333333333 HB131 Y N S Ep E wt 1 DOD R 1 0.083333333 HB65 N N M Mx E wt 2 DOD R 1 0.166666667 HB78 N Y M Ep CF wt 32 A R 0 2.666666667 HB72 Y Y M Mx E mut 9.5 DOD R 1 0.791666667 HB48 N Y M Ep CF mut 9 DOD R 1 0.75 HB102 N N S Ep CF mut 4 D B 0 0.333333333 HB160 Y Y S Mx E NA 14 R R 0 1.166666667 HB172 Y Y M Mx F/E NA 10 A R 0 0.833333333 HB99 Y Y M Ep E mut 7 DOD B 1 0.583333333 HB130 Y N S Mx NA mut 62 A R 0 5.166666667 HB98 Y Y S Ep M wt (FAP) 30 A M 0 2.5 HB136 N N S Mx F wt 34 A R 0 2.833333333 HB165 N N M Mx F/E mut 4 A R 0 0.333333333 HB1 Y Y M Ep E wt (FAP) 12 DOD R 1 1 HB93 N Y M Mx E mut 33 A LT 0 2.75 HB129 N N S Mx E wt (FAP) 54 DOD R 1 4.5 HB33 N Y M Ep CF wt(AX1N1) 3.5 DOD R 1 0.291666667 HB100 N N S Ep F mut 20 A B 0 1.666666667 HB184 Y Y M Ep E NA 14 DOD LT 1 1.166666667 HB157 Y N M Ep CF mut 5 R LT 0 0.416666667 HB80 Y Y S Ep CF mut 14 DOD R 1 1.166666667 HB86 N Y S Ep E mut 57 A R 0 4.75 HB8 N Y S Ep E mut 135 A R 0 11.25 HB147 N N S Mx F NA 12 A R 0 1 HB39 N Y S Mx NA mut 66 A R 0 5.5
[0324] TABLE-US-00030 TABLE C Gene Name AFP ALDH2 APCS AP0C4 AQP9 BUB1 C1S CYP2E1 DLG7 HC161 2.079447 -5.920384 -6.086912 -7.366206 -7.320175 4.176845 -6.502865 9.12672475 5.322878 HC162 4.056751 -3.64102 -4.586098 -5.663246 -4.233021 3.559124 -4.64283 -4.136919 5.950173 HC163 3.323238 -6.086663 -6.399079 -4.052853 -6.010302 4.772507 -6.776158 -8.515956 5.551408 HC164 3.075226 -6.146711 -7.241796 -3.371322 -5.446966 3.634476 -7.462807 -5.829384 3.98399 HC165 2.685177 -7.0470725 -6.294538 -7.242275 -6.94561 4.029514 -5.926596 -3.033642 5.723743 HC168 1.501031 -6.016314 -6.696324 -5.130347 -5.64774 3.305894 -6.883263 -4.411302 4.362859 HC169 2.880925 -6.024682 -6.87168 -4.19185 -6.058572 4.09117 -6.767215 -8.63753 4.614352 HC170 2.3753035 -6.6226955 -8.3702955 -5.4072375 -5.6954625 5.5639145 -8.0538815 -9.7948605 6.6275145 HC171 3.001804 -2.573977 -4.213123 -4.040859 -4.992701 3.583809 -5.226561 1.25382 3.874142 HC172 1.164528 -5.314302 -6.094852 -4.127298 -3.890072 3.991173 -6.240002 2.279678 5.651484 HC173 4.694127 -6.373823 -5.51865 -6.056863 -6.314031 4.30288 -4.863168 -8.649852 5.564261 HC176 4.066485 -5.552505 -5.444218 -5.551191 -5.815727 6.073568 -5.850428 -9.402043 6.051409 HC177 2.692613 -5.43842 -3.091896 -4.656336 -5.907612 3.452047 -6.412596 -10.50172 4.083836 HC178 -0.554213 -5.646708 -7.296414 -4.588115 -5.579087 3.125179 -6.556397 -6.591304 4.755443 HC179 1.910595 -4.139932 -8.136252 -6.036987 -2.847761 3.895205 -4.943672 -5.283326 5.054346 HC180 3.212685 -5.831134 -7.519348 -5.962761 -6.611712 1.5179 -6.130592 -9.203789 2.22658 HC181 6.030393 -4.04397 -2.03808 -0.956533 -2.850753 5.430957 -4.712002 -2.555649 5.031845 HC182 3.376941 -7.072651 -7.74873 -5.2003 -5.445893 6.665657 -7.899793 -10.089271 7.487442 HC183 3.149578 -4.684626 -7.045155 -3.800078 -7.042931 2.40337 -6.412624 -9.657513 3.396236 HC184 -0.093476 5.985909 -7.203484 5.482853 -6.208594 1.558788 -6.347367 -9.658434 2.407985 HC185 1.405595 -4.748444 -5.89589 -3.780913 -2.802368 4.37289 -5.800822 -5.410746 4.6459 HC186 1.666457 -5.52819 -7.953401 -3.287374 -3.805233 1.040678 -7.309734 -6.699831 2.197157 HC187 3.652111 -4.151991 -7.459358 -6.247812 -5.346647 4.211928 -6.33068 -8.629261 4.520672 HC188 0.355562 -5.261937 -7.83848 -4.759525 -4.839348 5.111208 -7.787661 -4.575966 5.635841 HC189 1.239891 -4.501697 -8.737075 -6.152778 -6.402122 5.0291015 -6.951675 -5.450079 4.419359 HC190 3.306642 -4.365515 -7.399538 -4.721411 -6.178224 3.016906 -4.970499 -5.850237 9.264351 Gene Name DUSP9 E2F5 GHR HPD IGSF1 NLE RPL10A HC161 3.702615 1.025512 -0.817005 -7.653863 14.149408 5.1985405 -5.81852 HC162 1.738977 1.432598 -0.231753 -6.700146 14.781699 1.231146 -5.9665735 HC163 4.00436 1.072797 -2.746621 -6.213082 8.2477055 2.203781 -5.49725 HC164 4.25604 2.567639 -3.606813 -6.079645 12.649441 1.946926 -5.171041 HC165 1.788757 1.157215 -1.197022 -7.969042 14.270796 2.620134 -6.219366 HC168 5.625335 2.2963 -1.169362 -7.52548 8.041574 2.337152 -5.42627 HC169 3.838008 1.60884 -2.921191 -6.51064 8.136143 2.099644 -5.731897 HC170 1.8626715 1.6955475 -3.9034625 -7.4271305 7.756398 2.6917235 -5.8132855 HC171 5.349357 2.074272 -1.437519 -5.297939 6.325863 3.057537 -3.95361 HC172 5.592005 1.291773 -0.040049 -6.989866 6.998259 3.186024 -3.946432 HC173 4.718896 1.367846 -2.3934 -7.781412 9.1259525 1.82226 -4.957916 HC176 2.248373 2.709599 -3.2392 -7.594156 7.5288985 1.817325 -5.042318 HC177 -0.297108 2.149313 -2.166834 -7.847734 5.8240705 1.530536 -5.640103 HC178 4.943904 1.038474 -1.620902 -5.659262 5.416822 1.855914 -4.954215 HC179 1.464274 1.372578 -0.386778 -6.31274 7.244471 1.887378 -5.218281 HC180 0.161194 -0.215954 -0.371454 -6.978048 5.185486 1.004282 -6.187635 HC181 4.322323 2.990459 2.18165 -0.651095 4.292234 4.670446 -2.978533 HC182 2.395117 2.329727 -4.420263 -7.357922 7.932783 2.869667 -5.574881 HC183 3.7002215 -0.85541 0.078707 -7.143723 11.999761 0.63414 -6.105039 HC184 2.266351 6.244093 0.670045 -6.27671 6.935964 1.564672 -6.568913 HC185 1.811225 2.225761 -1.246884 -7.344763 10.1413645 1.39443 -5.015711 HC186 -2.717975 1.183123 -2.657936 -7.680597 8.921477 1.289946 -6.631908 HC187 -0.066629 -2.0378 1.078709 -8.251018 7.478678 1.655093 -5.763416 HC188 1.839584 0.638515 -1.989428 -6.736329 12.8628775 2.27923 -4.743699 HC189 6.509026 -0.7698 -2.238756 -8.600128 11.305903 -0.437812 -7.061492 HC190 0.70722 4.181534 -0.773062 -4.881306 2.422048 -5.53509
[0325] TABLE-US-00031 TABLE D Table of normalized qPCR data (deltaCt values) of 88 HCCs analyzed by the Taqman method Gene name AFP ALDH2 AP0C4 APCS AQP9 BUB1 C1S CYP2E1 HC 001 2.212911 -6.2372335 -0.614689 -7.0721355 -6.047695 3.841505 -8.163492 -10.3093235 HC 003 3.865709 -6.230074 -0.95786 -7.52919 -6.7334475 0.147459 -8.7963405 -10.428074 HC 004 7.6758115 -2.186358 1.608247 -5.845683 -3.759528 4.221132 -5.8997645 -7.1147515 HC 006 7.9469815 -5.4231035 -0.9614255 -7.3704745 -7.006052 0.5252045 -8.162856 -10.1334265 HC 007 -5.311541 -4.0446765 3.550537 -5.1967915 -6.747103 0.299039 -4.062593 -11.024027 HC 008 -2.0890815 -3.9297005 0.6776965 -6.567126 -3.1082155 1.214781 -7.2991535 -7.791007S HC 009 7.0483095 -3.0017225 9.6721075 0.017488 -3.7536735 -2.980029 -4.830331 -0.5825245 HC 010 -2.3869635 -0.95212 0 1.0272875 -1.3400495 1.864677 -2.639902 -3.604805 HC 011 -0.6488335 -5.958108 -1.076151 -7.7638255 -6.122144 2.362454 -8.319293 -9.575619 HC 012 6.538312 -4.6271565 1.221393 -6.942673 -4.1878425 3.293346 -6.850023 -7.284587 HC 014 2.987769 -5.194577 -1.3542145 -6.5396565 -6.8623455 1.363697 -6.8939375 -10.7465595 HC 015 -6.14089 -4.5178635 5.156026 -3.380102 -2.373344 -0.8830545 -7.1343975 -4.9390935 HC 017 -7.1950405 -2.6522585 2.395651 -4.5167035 -2.8711295 -1.0884485 -6.035123 -6.037085 HC 018 6.856588 -1.840894 3.84764 -4.916924 -3.6093495 0.063545 -4.263272 -5.811062 HC 020 0.65281 -6.287083 -3.2094885 -8.2117635 -7.354605 1.4635025 -8.471663 -10.2536915 HC 021 4.3070475 -2.175112 6.2591235 -5.9159775 -1.1452535 -0.0802935 -5.7190985 -1.2878015 HC 022 4.418018 -5.331214 -0.5455545 -6.6835035 -5.7992305 2.173361 -7.2514145 -8.0876755 HC23 5.538438 -5.853486 -0.5708905 -6.9009145 -6.651868 2.5475915 -8.2212235 -9.047509 HC 025 3.90298 -6.162477 -1.834891 -8.798759 -8.758959 2.5679685 -8.5606875 -10.814935 HC 026 5.69175 -5.0135775 -0.2581675 -7.2072275 -3.8645965 -0.545363 -7.2351705 -0.671071 HC 027 0.626755 -5.6309605 -1.53158 -7.2809855 -5.4736555 0.8889165 -8.172076 -8.6350095 HC 028 0 -1.913778 6.0251725 -1.0475505 -0.9613895 5.7426525 -4.910584 -3.6858305 HC 030 -6.4370325 -3.8476295 -0.2797975 -7.1142435 -5.0250435 0.190936 -7.5279395 -7.5682115 HC 032 -0.0037145 -6.802666 -2.574347 -7.500133 -7.530391 5.1317805 -7.854502 -9.4408715 HC 034 6.6945705 -5.11617 -0.5860455 -7.134934 -6.9427395 1.2674215 -7.719763 -8.545814 HC 037 1.3519745 -5.808058 0.0768065 -6.755895 -6.3416265 2.4955985 -6.921051 -10.1686795 HC 038 -4.053435 -4.596143 0.129322 -5.045701 -6.0302545 -0.321483 -6.101331 -8.1123675 HC 041 2.7156435 -6.3503265 -2.281983 -5.612517 -7.8444565 0.587016 -6.88808 -9.5090495 HC 042 5.216493 -4.4086495 0.627239 -4.1054755 -6.063786 2.224818 -6.3060565 -9.1411555 HC 043 1.7983435 -5.457548 0.7055185 -7.607914 -4.7175855 2.8634735 -7.9862115 -8.760714 HC 052 -10.3337105 -2.1920375 8.124407 -5.9818015 0.4848805 1.2986035 -5.6337865 -1.7693015 HC 058 -1.891958 -2.1172735 11.8524 4.1106695 2.817265 -1.9395175 -3.691331 4.3317445 HC 060 -7.624821 -3.6860195 0.545509 -8.100997 -6.8503395 0.576028 -8.167253 -9.1875325 HC 064 -5.0266755 -4.992107 -0.7860345 -7.4148835 -7.0526325 1.367463 -7.1364365 -9.682147 HC 066 -3.156328 -3.8408415 0.6773785 -8.2106815 -6.2767975 1.1272665 -8.026875 -8.601088 HC101 6.873135 -4.339036 0.5787185 -6.288568 -4.6233735 -0.081457 -7.321092 -5.806032 HC102 4.119697 -2.476355 5.453696 2.3952165 -0.0196725 0.5553155 -5.939374 2.8566735 HC103 -1.6193685 -3.889904 0.54698 -6.014572 -7.151639 2.086008 -5.965432 -8.266311 HC104 -5.5094265 -4.936239 0.5059805 -5.624234 -0.501258 1.311194 -6.716137 -9.0888685 HC105 -2.3444245 -4.239726 3.577778 -7.703333 -4.2748785 -0.945674 -7.774455 -5.698899 HC106 3.42054 -6.1642895 0.7836775 -7.8462545 -5.85931 4.8909655 -8.060072 -9.9949555 HC107 4.136209 -6.7443095 -4.4534435 -9.2080655 -8.8878655 1.7415115 -9.2061165 -9.3234825 HC108 4.500336 -3.6076385 2.478085 -7.275462 -4.4353395 0.3807995 -7.1031155 -3.889942 HC109 4.833024 -5.8617665 -0.729565 -6.222909 -6.4504115 2.2918285 -7.406001 -8.7101925 HC110 3.5240185 -3.6707715 0.256479 -5.043319 -4.5999895 1.449943 -6.9163195 -7.145766 HC111 1.883473 -3.8304065 1.130067 -5.976754 -4.1657805 -0.621548 -6.278164 -4.46942 HC112 2.8803905 -4.8726745 0.7777655 -6.764675 -5.2735435 -0.3135015 -7.455794 -2.5741475 HC113 -1.208649 -4.407016 2.366969 -5.197177 -2.681192 3.4825665 -6.338901 -6.443846 HC114 5.4433695 -4.7113965 0.833543 -6.723142 -4.445291 1.7431855 -7.866014 -7.3429245 HC119 -1.0580855 -6.159706 -1.894453 -9.375177 -7.6266135 0.797564 -9.1461175 -7.095824 HC120 4.0065425 -4.257398 3.5241745 -5.6838965 -6.8239115 0.0740105 -8.5708615 -7.6044515 HC121 4.254961 -4.556431 2.167313 -6.2688205 -4.38702 2.4486685 -8.118416 -7.765037 HC122 2.3763095 -6.2844515 -1.279577 -6.9942545 -6.8198535 6.0183915 -7.7653135 -9.450349 HC123 -0.821555 -4.220769 0.68167 -5.778659 -6.410177 1.190323 -5.383781 -8.528543 HC124 -3.9525335 -4.027289 0.0499065 -5.391271 -4.463488 1.592563 -5.151686 -9.520436 HC125 4.806564 -4.5451465 -2.6326775 -6.5321595 -8.370224 -1.1627945 -8.4244055 -9.426232 HC126 5.899437 -5.02839 -0.407895 -5.2838365 -3.6163545 2.6943025 -7.1365955 -5.226091 HC127 0.0390765 -2.41699 -0.8680995 -4.846116 -1.8613935 2.048769 -6.3641695 -6.1813065 HC128 -5.8636305 -5.085525 0.626498 -5.087517 -4.3184915 1.3297375 -6.828468 -7.4344035 HC129 3.430757 -4.6298475 1.863955 -4.8448705 -2.870839 2.3688215 -7.302922 -2.692798 HC131 1.491189 -5.425994 -2.4702 -8.6617295 -7.4772145 0.727709 -7.525072 -8.98645 HC132 -5.4265205 -3.105643 6.9974515 3.2748865 -3.9244375 -0.2895395 -4.390082 -7.0455735 HC133 5.1621395 -4.2462915 -0.63156 -7.145861 -6.05182 4.9277675 -7.3188145 -8.1908895 HC134 -2.8738695 -4.061101 0.1134065 -7.5103485 -5.550642 -1.7425995 -8.4609335 -7.859701 HC135 0.909107 -2.7442165 0.7630605 -0.959726 -4.0595615 1.2018365 -4.667223 -4.30592 HC136 0.4105125 -6.0408575 -0.7390785 -7.150737 -5.996196 4.288554 -8.243333 -9.042865 HC137 -4.378388 -3.2913795 3.209294 -4.421328 -0.5225755 4.2185175 -5.647363 -5.532515 HC138 2.4762965 -4.8248625 1.154563 -4.883388 -3.440722 3.408251 -6.459976 -7.2458685 HC139 2.7547595 -2.9782295 3.0252085 -5.3858735 -5.0157665 0.9503045 -6.0281485 -1.1920485 HC140 6.3489955 -4.644452 -1.006979 -2.1507335 -5.3387635 4.075603 -6.7373815 6.646618 HC141 2.4010865 -4.8883675 0.787009 -4.7365085 -4.1224775 4.2728925 -6.8664705 -2.6765195 HC142 4.5984525 -3.7946485 2.8271835 -4.9243665 -3.1411815 4.0713025 -6.3482925 2.654871 HC143 -4.0727165 -2.59764 1.855993 -4.8795135 -2.222047 1.6908025 -4.948264 -3.1057735 HC144 4.7344185 -4.3542505 -1.002913 -0.432856 -5.16696 2.510931 -5.3365195 -4.456082 HC145 8.5175565 -3.375805 0.8672075 -5.0765195 -4.091142 3.9700095 -6.960951 0.8009 HC146 5.741507 -3.5738745 1.2439275 -5.1950135 -3.4305425 2.9843625 -5.666896 -0.913546 HC147 6.0474775 -3.0470955 0.2246755 -5.6213855 -5.257189 2.7534355 -5.349428 -6.933909 HC148 -1.306432 -4.0108565 0.267747 -6.3544915 -3.1846315 1.1995135 -6.2066555 -4.1428355 HC149 -3.9190605 -3.3456535 2.735403 -1.9099995 -1.1810265 2.704253 -5.707004 -5.9300895 HC150 6.1556695 -2.9923905 -1.9485835 -5.821769 -6.3127705 2.452404 -4.984573 -7.3184395 HC151 5.5488065 -4.234966 1.372415 -5.8812085 -4.0297925 3.4239945 -7.2861515 -2.304461 HC152 4.917902 -3.97386 -4.005999 -6.5072455 -7.124415 2.5576145 -5.752235 -9.98327 HC153 5.6708455 -5.004032 -3.204075 -3.8195495 -6.2020215 1.9670395 -5.979251 -7.7421455 HC154 6.699114 -2.0392575 9.6136985 0.885791 -0.68511 1.755108 -0.7395055 2.544628 HC155 6.238831 -3.802053 2.0022335 -6.3105565 -2.974712 4.2276825 -7.058571 -4.1514335 HC156 -1.582839 -3.5688085 0.917505 -3.9333845 -4.163765 1.0763025 -4.6064345 -8.4802835 HC157 3.657864 -4.2315665 2.513598 -7.2096625 -4.573216 -0.284071 -5.856564 -7.9837885 HC159 3.4650565 -2.6801805 2.2596385 -4.0834345 -4.42904 3.44645 -5.923485 -7.778452 Gene name AFP ALDH2 AP0C4 APCS AQP9 BUB1 C1S CYP2E1 HC 001 5.30317 11.616567 -0.05328 -2.655512 -9.449416 6.46034 1.159417 -6.6225235 HC 003 2.057513 8.8462855 1.909804 -2.069524 -8.549803 7.249974 1.5801355 -6.0562915 HC 004 4.4226465 9.4268185 1.7432195 2.0012965 -9.415253 0 3.1459935 -4.4121905 HC 006 1.6282005 10.22051 -0.024339 -1.887805 -8.5958965 7.1580385 -0.6940375 -6.8637555 HC 007 1.169221 6.6521625 0.2833465 1.7428205 -6.183977 3.192514 0.3919565 -7.1381125 HC 008 2.80866 9.6946695 0.0193165 -2.342442 -5.329776 2.806768 1.579419 -6.2574845 HC 009 -1.3733475 9.5262655 -0.711082 2.3242195 0.011478 4.026769 0.80375 -6.3016635 HC 010 0 0 1.344368 0.4900285 -2.932809 0 0 -9.1966395 HC 011 2.8432205 0 0.736822 -4.757848 -9.029214 7.6390015 1.9328755 -7.379063 HC 012 4.7199665 0 2.4002515 -2.2402875 -9.656029 7.466951 1.64183 -5.178571 HC 014 3.3543285 7.7629895 1.5332515 -1.09511 -9.5837645 8.5836025 1.47219 -5.831244 HC 015 0.1414205 4.4342765 -1.399564 -0.2426 -4.473096 -0.0722075 0.321593 -6.8777395 HC 017 -0.666284 3.163581 -1.206766 2.353691 -0.6808655 6.0490105 0.386649 -7.068098 HC 018 1.512286 8.7756845 2.426129 2.9035 -5.7101575 2.4248235 1.3815525 -5.9464565 HC 020 2.1165725 9.6208445 1.1944835 -4.5756335 -10.6864405 0 1.118745 -7.542193 HC 021 0.322455 7.8162765 0.0686475 -0.71981 -4.0108195 2.954814 1.618369 -6.309556 HC 022 3.3904095 10.82729 10.7133385 -2.416651 -9.8859985 5.6986975 1.9449755 -7.194012 HC23 3.848364 0 1.4330655 -3.7226655 -9.583194 7.200325 1.823275 -5.9526365 HC 025 3.34202 7.1111525 -0.049846 -1.9012935 -9.1845675 0 1.770127 -7.4507165 HC 026 0.9710395 8.5287915 1.1845665 -1.964045 -7.6403735 5.4960635 1.851733 -5.9670715 HC 027 2.3158215 10.241011 0.4045835 -2.623084 -9.597772 5.588995 1.851285 -7.6623025 HC 028 0 0 4.334386 1.9788575 -3.3142495 0 2.4559905 -5.521873 HC 030 0.189092 9.0027 -1.0623035 -2.635437 -7.537 2.651022 1.2674865 -7.5046195 HC 032 5.7080765 9.73163 0.054818 -2.0027475 -9.0015185 0 1.208576 -8.8437815 HC 034 2.339621 9.9728495 1.4281575 -1.563203 -8.3685675 10.112616 1.934745 -6.594006 HC 037 2.6534895 0 1.2212655 -2.9415775 -10.367265 7.5570255 1.9881245 -6.901637 HC 038 1.4386515 5.2298755 0.037887 -0.2025015 -7.547286 0.680358 2.1250395 -5.1574215 HC 041 1.840185 8.727439 -0.466649 -1.428749 -8.0015745 7.243446 0.15624 -7.7043325 HC 042 3.2531575 0 0.3673235 1.2545195 -8.2669835 2.899766 0.9401045 -5.577659 HC 043 4.2390495 10.525647 0.894345 -3.2916395 -8.997825 5.5544715 1.8422595 -5.480403 HC 052 2.599359 3.8059605 -0.4419525 1.843696 -2.481945 -2.254168 1.9474305 -5.6154705
HC 058 -0.1957495 3.656912 -0.804087 3.7242975 -1.8257985 -1.3471695 1.209522 -6.0601515 HC 060 2.2644225 6.618755 0.432422 1.4079225 -8.4643875 0.7884805 1.9133155 -5.7041285 HC 064 2.386875 7.3184655 0.2876185 -0.349645 -8.6027575 3.3382005 1.817699 -6.4617635 HC 066 2.7680135 11.5673955 0.968982 1.2501855 -8.5231325 9.185554 1.962008 -5.415169 HC101 1.3084655 8.828389 1.871516 -0.1466275 -5.7252795 4.1394545 1.4546305 -6.144011 HC102 2.1385165 8.6628475 -0.830934 -0.947389 -0.568809 2.708733 1.1534675 -5.283399 HC103 2.957914 12.521336 1.8003215 -0.636723 -6.717282 9.802921 2.594702 -4.423835 HC104 1.821739 5.396553 2.305498 -1.6860905 -8.46781 -0.1438735 1.610158 -6.21159 HC105 0.814912 5.4214725 -2.0730715 -0.682142 -2.288109 1.422332 0.471391 -6.315756 HC106 6.2678815 11.174152 2.208171 -5.342392 -9.4440475 7.401009 1.968983 -5.769397 HC107 1.357756 6.6136855 -2.78876 -2.935929 -10.460972 0 0.000835 -8.6686655 HC105 2.2445545 8.0946735 -0.0923905 -1.6363755 -2.9674235 7.967992 0.932052 -5.818028 HC109 3.222524 10.4709205 1.9924345 -2.9233285 -7.8859205 10.0122565 2.6102395 -5.541229 HC110 2.333076 11.616244 2.512512 -1.0803015 -8.1908235 8.1469415 2.3529485 -5.245476 HC111 0.769283 9.137462 -1.045678 -1.1576425 -7.245347 1.86965 1.012752 -5.568205 HC112 0.9196845 10.105965 -0.0373705 -2.5391085 -7.714358 3.4428695 1.119237 -6.1905075 HC113 4.5602875 7.8299455 2.82243 -2.16232 -6.685692 2.045068 2.156348 -5.8884625 HC114 3.1500875 11.804112 0.0450475 -2.5053965 -6.835254 5.1813245 1.3170345 -5.795905 HC119 1.712686 9.106547 0.0248045 -3.7649595 -9.220498 5.39017 0.400823 -7.954231 HC120 1.9563135 5.8119685 -1.229768 -3.196589 -8.5127155 9.404196 1.1096815 -6.4517175 HC121 2.852561 9.706684 0.910943 -2.2774645 -7.480725 5.980435 1.758163 -6.4042545 HC122 7.228946 9.9054825 3.5033365 -2.400201 -8.7301975 8.6480295 2.2430545 -5.199782 HC123 2.929576 11.584458 0.646839 1.810364 -4.7774665 5.1400615 1.5951645 -4.7323885 HC124 2.03781 8.81055 -0.574165 -2.2369305 -7.832169 1.4450915 0.1499775 -6.691521 HC125 -0.3286545 9.3740615 0.028878 -0.697866 -5.7813 10.2234745 0.405397 -7.1196575 HC128 3.944339 8.7174575 3.271927 -1.824385 -1.865621 7.659377 2.033278 -5.389272 HC127 2.96212 8.672372 2.162602 -0.129431 -3.4481965 3.1503205 2.205965 -4.3385115 HC128 2.6299155 8.499355 4.393094 -1.9716885 -5.7052855 2.72995 1.949352 -6.6181545 HC129 3.6405185 7.0627455 0.470421 -2.332961 -5.502918 5.692623 1.683808 -4.8697295 HC131 1.461713 8.415907 -0.154573 -4.009655 -8.960383 7.5832005 1.5313675 -6.775249 HC132 1.5572645 3.3843145 -1.9018925 -1.7710325 -2.3653865 1.947055 -0.2035885 -6.7796075 HC133 5.5447335 8.022457 2.6341825 -2.2298335 -6.1281315 0 1.4173895 -5.762015 HC134 -0.8148735 4.96739 -3.1030595 -1.3138565 -7.231144 0.3848995 -0.794433 -7.7140665 HC135 2.250305 5.794605 -0.986165 0.6955465 -6.7262275 4.394354 0.9780515 -6.689595 HC136 5.5267715 10.9307725 2.4040865 -4.013948 -8.223611 7.4962365 2.426321 -5.5069335 HC137 5.2105355 4.767228 5.62451 -1.6355645 -5.8875425 1.0556075 3.7311615 -5.2271275 HC 138 5.028429 5.576937 4.1601375 -1.738341 -6.019837 7.169314 4.19882 -4.2322595 HC 139 2.940447 4.3133685 0.685194 1.632571 -4.6240035 3.333358 1.7913325 -6.6866335 HC 140 5.1767035 10.874029 2.488357 -3.1717235 -7.5439415 9.276635 5.0732625 -4.266519 HC 141 6.1148255 7.979559 2.66802 -1.687093 -7.2596615 #DIV/0! 3.5973445 -4.952551 HC 142 5.8031125 8.2104255 2.0983905 -1.5934495 -5.8074755 9.442329 3.4164995 4.6520795 HC 143 3.470906 3.981805 1.474377 0.695168 -2.049901 3.754627 3.058019 -4.7443975 HC 144 3.844786 10.7187705 3.540563 -1.6857605 -6.869217 11.9441575 4.417722 -4.817306 HC 145 5.482263 9.313039 2.112409 -1.525041 -6.669204 10.0458615 3.0082705 -5.7677005 HC 148 5.1824885 7.611916 2.8802325 -1.791636 -6.9831945 5.450716 3.884913 -4.427413 HC 147 4.5366875 9.358894 3.2373475 -2.0156545 -6.053345 8.7065355 3.732017 -4.317148 HC 148 2.490156 5.4985645 8.523611 -0.773246 -3.7206575 5.663583 3.295068 -6.0532135 HC 149 3.4454215 6.8563245 2.4724295 -0.9357605 -7.337568 -0.063395 4.267075 -5.7767065 HC 150 3.585447 7.980274 3.118546 0.5916635 -5.762837 9.1651835 2.811495 -5.7495535 HC 151 4.613043 8.9062765 2.2090065 -2.8000785 -7.251033 9.44137 3.5959505 -4.6972005 HC 152 4.17552 10.736246 4.56538 -1.578246 -8.106859 12.118351 2.6658355 -6.944767 HC 153 3.133394 7.298329 3.85894 -0.616143 -7.947464 11.674272 2.670245 -5.0796695 HC 154 3.2541115 3.139705 -0.3936805 -1.070278 -4.611328 1.5925535 2.2396475 -6.2090535 HC 155 5.7341595 6.4585135 2.4375015 -0.254649 -7.297162 10.0981895 3.3878795 -5.37231 HC 156 2.1302465 4.4056075 1.070339 0.42868 -6.890963 2.0124875 2.225275 -7.037827 HC 157 1.3778545 2.0950385 -0.56173 -0.8411435 -8.474893 7.2842685 1.6720135 -6.6310375 HC 159 5.727853 8.8523415 2.7886015 -1.0442865 -7.268645 8.8204775 2.861685 -5.4777465
[0326] TABLE-US-00032 TABLE G Date of follow- Date of 1st secondary date of tumor HC 000 tumor surgery or (PH) or Date of Date of up recur- recurrence or OLT after secondary grade identification transplantation (OLT) last visit death (years) rence metastasis hepatectomy OLT Edmondson HC 001 12/12/1996 PH 07/01/1997 0.07 N 3 HC 003 21/02/1997 PH 20/06/2000 3.33 Y 4/11/1998 N 2 HC 004 28/02/1997 PH 20/08/2008 11.48 N 2 HC 006 07/10/1996 PH 06/01/1998 1.25 N 28/11/1997 N 2 HC 007 02/07/1996 PH 31/12/1997 1.50 Y 4/11/1997 N 2-3 HC 008 05/06/1996 PH 24/01/2005 8.48 N 3 HC 009 28/08/1996 PH 05/09/1996 0.02 N 3-4 HC 010 10/10/1996 PH 20/09/1997 0.95 N 4 HC 011 10/10/1996 OLT 14/12/2008 12.20 N 2 HC 012 24/10/1995 OLT 14/11/1995 0.05 N 2 HC 014 10/06/1995 OLT 27/07/1995 1.00 N 3-4 HC 015 21/07/1995 PH 10/10/1996 1.22 Y 10/10/1996 N 3 HC 017 05/05/1997 PH 16/04/2008 10.96 N 2 HC 018 07/05/1997 PH 28/09/1997 0.39 NA 3 HC 020 13/05/1993 OLT 20/10/2008 15.40 N 2 HC 021 15/01/1992 PH 28/09/1992 0.70 Y 15/06/1992 N NA HC 022 15/03/1997 OLT 02/09/2008 11.50 N 2 HC 023 20/07/1995 PH 20/06/2007 11.93 N 2 HC 025 05/10/1992 PH 13/08/2008 15.87 N 2 HC 026 04/06/1993 OLT 18/04/1994 0.83 NA 2 HC 027 20/01/1993 OLT 15/02/1993 0.10 N 2 HC 028 16/02/1996 OLT 13/03/1996 0.10 N 3 HC 030 10/04/1996 PH 07/09/2008 12.40 Y 15/10/1996 Y 17/12/1993 3 HC 032 17/02/1993 PH 17/10/1993 0.66 N 2 HC 034 10/03/1993 PH 05/11/2008 15.70 Y 15/11/1995 Y 20/06/1996 2 HC 037 08/06/1997 OLT 13/08/1997 0.20 N 3 HC 038 16/07/1997 PH 28/08/1998 1.12 Y 1/01/1998 N NA HC 041 24/11/1997 PH 01/05/2005 7.44 Y 29/06/1999 Y 9/3/2000 2 2nd recurrence 15/1/2005 HC 042 05/11/1997 PH 03/06/2008 10.58 N 3 HC 043 19/11/1997 OLT 22/10/2008 10.90 N 3 HC 052 17/02/1999 PH 18/05/1999 PDV 0.25 N 3 HC 058 14/10/1999 PH 30/01/2008 8.30 N 2 HC 060 15/05/1925 PH NA NA HC 064 10/04/2000 PH 09/07/2005 5.25 Y 15/10/2001 N 3 HC 066 15/09/1999 PH 18/08/2008 8.93 N 2-3 HC 101 03/05/2006 0LT 27/10/2008 2.50 N 2-3 HC 102 12/07/2006 PH 18/08/2006 0.10 N 4 HC 103 16/08/2006 PH 11/06/2008 1.82 Y 15/1/2007 N 2-3 HC 104 20/09/2006 PH 05/11/2008 2.10 N 2-3 HC 105 11/12/2006 PH 04/07/2007 0.56 Y 15/04/2007 N 3 HC 106 22/01/2007 OLT 16/01/2009 2.00 Y 3 HC 107 25/01/2007 PH 23/10/2008 1.75 N 2 HC 108 12/02/2007 PH 24/09/2008 1.62 N 3 HC 109 19/02/2007 OLT 26/05/2008 1.30 N 2-3 HC 110 6/02/2007 OLT 04/02/2009 1.95 N 2-3 HC 111 07/03/2007 OLT 03/10/2007 0.70 N 2-3 HC 112 19/03/2007 PH 08/09/2008 1.48 N 2-3 HC 113 23/03/2007 OLT 15/03/2008 1.00 N 2-3 HC 114 03/04/2007 PH 11/09/2007 0.44 N 2 HC 115 01/08/2007 PH 29/04/2008 0.75 N 1 HC 116 09/08/2008 PH 18/04/2008 0.69 N 3 HC 117 25/10/2007 OLT 23/12/2008 1.20 N 2-3 HC 118 25/10/2007 PH 28/09/2008 0.93 N 1 HC 119 03/12/2007 OLT 08/01/2009 1.20 N 2-3 HC 120 18/12/2007 PH 14/10/2008 0.82 N Y 12/05/2008 2-3 HC 121 02/01/2008 PH 08/08/2008 0.60 N 3 HC 122 16/01/2008 PH 17/10/2008 0.75 Y 10/10/2008 N 2 HC 123 11/02/2008 OLT 01/12/2008 0.80 N 3 HC 124 20/02/2008 PH 26/08/2008 0.52 N 3 HC 125 22/02/2008 OLT 08/01/2009 0.90 N 3 HC 126 12/03/2008 PH 14/08/2008 0.42 Y 6/8/2008 N 1-2 HC 127 19/03/2008 PH 20/06/2008 0.25 Y 4/6/2008 N 2-3 HC 128 20/03/2008 PH 29/08/2008 0.44 N 2 HC 129 01/04/2008 0LT 31/05/2008 0.15 N 3 HC 130 07/04/2008 PH 27/05/2008 0.14 N 3 HC 131 10/04/2008 PH 15/07/2008 0.26 N 2-3 HC 137 19/07/2002 PH 31/03/2008 . 5.67 N . NA HC 138 25/04/2003 PH 03/12/2008 . 5.58 Y 03/10/2003 NA HC 139 15/05/2002 PH 09/05/2008 . 6.00 N . NA HC 140 03/06/2004 PH 5/08/2008 . 4.17 Y 30/06/2005 NA HC 141 06/02/2004 PH 12/03/2009 . 5.08 Y Dec. 2005 NA HC 142 14/05/2002 PH 21/06/2006 21/06/2006 4.08 Y 24/03/2006 NA HC 143 04/03/2002 PH 26/01/2007 . 2.83 Y 2005 NA HC 144 27/06/2002 PH 17/06/2008 . 6.00 Y 16/03/2004 NA HC 145 14/11/2002 PH 30/07/2008 . 5.58 Y 09/06/2005 NA HC 146 30/07/2004 PH 11/12/2008 . 4.33 Y June 2005 NA HC 147 23/11/2004 PH 22/09/2008 . 3.83 Y 12/06/2008 NA HC 148 12/09/2003 PH 15/10/2006 . 3.08 N NA HC 149 26/08/2003 PH 16/01/2007 16/01/2007 3.42 N NA HC 150 31/01/2003 PH 23/06/2008 . 5.42 N NA HC 151 10/12/2004 PH 15/03/2007 . 2.25 N NA HC 152 14/05/2003 PH 17/01/2007 17/01/2007 3.67 Y mars-09 NA HC 153 25/02/2003 PH 24/12/2007 24/12/2007 4.83 Y 06/05/2005 NA HC 154 06/09/2004 PH 23/11/2006 2.21 Y 01/01/2005 N 2-3 HC 155 18/10/2004 PH 09/12/2008 4.10 Y 18/10/2004 Y 31/05/2005 2 HC 156 03/02/2005 PH 28/05/2007 2.31 Y 15/06/2006 3 HC 157 24/02/2003 PH 26/10/2006 3.59 Y 15/08/2004 2 HC 159 16/10/2002 PH 18/03/2005 2.42 Y 03/05/2004 2 HC 161 20/08/2003 PH 06/02/2008 4.47 Y 2 HC 162 30/10/2003 PH 25/04/2007 3.49 N 3 HC 163 20/09/2004 PH 07/12/2006 2.21 Y 01/09/2006 N 3 HC 164 05/09/2002 PH 21/03/2007 4.54 N 1 HC 165 08/08/2003 PH 29/05/2008 4.72 N 2 HC 168 10/02/2003 PH 04/02/2009 6.00 Y 15/07/2004 Y 18/02/2008 2 HC 169 10/06/2002 PH 22/03/2005 22/03/2005 2.78 Y 15/03/2003 N 2 HC 170 14/03/2002 PH 28/06/2007 5.29 N 1 HC 171 25/03/2004 PH 17/10/2008 4.57 Y 15/11/2004 N 4 HC 172 10/01/2005 PH 25/11/2008 3.90 Y 25/11/2005 N 3 HC 173 18/12/2003 PH 03/03/2008 4.21 N 1 HC 176 13/03/2002 PH 05/10/2006 4.57 N 2 HC 177 29/10/2003 PH mars-09 5.42 Y 01/2009 2 HC 178 19/03/2003 PH 19/09/2005 2.50 N 2 HC 179 27/10/2000 PH 06/12/2005 5.17 Y 10/2002 2-3 HC 180 9/4/2002 PH 03/11/2005 03/11/2005 3.58 Y 05/2005 3 HC 181 27/05/2002 PH mars-09 6.83 Y 04/2008 2 HC 182 30/03/2004 PH October 3.50 N 1 2007 HC 183 21/07/2003 PH 02/09/2007 02/09/2007 4.08 Y July 2007 3 HC 184 18/01/2002 PH 08/02/2004 08/02/2004 2.08 Y April 2002 2 HC 185 19/11/2002 PH 03/03/2005 2.25 N 3 HC 186 31/08/2004 PH 06/11/2006 06/11/2006 2.17 N 3 HC 187 7/06/2001 PH fevr-09 7.67 Y March 2003 1 HC 188 29/07/2004 PH avr-09 4.67 Y July 2004 2 HC 189 30/04/2002 PH 13/08/2005 13/08/2005 3.25 Y January 2 2005 HC 190 29/07/2003 PH mars-09 5.58 N 3 number max of Macro- mitosis Ndules Nrmal Score Score Tumor vascular vascular per 10 of liver Cir- META- META- HC 000 tumor differenti- tumor invasion invasion fields × multiple regen- A0F0 or rhosis VIR VIR identification ation (OMS) size (mm) macro micro 40 Ndules eration A0F1 AXF4 Activity Fibrosis HC 001 moderately 120 N N NA N N Y NA 4 differentiated HC 003 well 60 N N NA N N Y NA 4 differentiated HC 004 well 100 N N NA N Y N 0 1 differentiated HC 006 well 90 N Y NA N Y N 0 1 differentiated HC 007 well 100 Y Y NA Y N N 2 3 differentiated HC 008 moderately 30 N N NA N N Y N 4 differentiated HC 009 Moderately 100 Y Y NA Y N N 1 3 poorly HC 010 moderately- 75 N N NA N N Y NA 4 poorly HC 011 well 15 N N NA Y N Y NA 4 differentiated HC 012 well 60 N N NA Y N Y NA 4 differentiated HC 014 Moderate 80 Y Y NA Y N Y NA 4 poor HC 015 moderately 60 Y Y NA Y N N 3 3 differentiated HC 017 well 100 N N NA N N N NA 3 differentiated HC 018 moderately 140 Y Y NA N N Y 2 4 differentiated HC 020 well 40 NA NA NA Y N Y NA 4 differentiated HC 021 NA 100 NA NA NA Y N Y NA 4 HC 022 well 45 N N NA Y N Y NA 4 differentiated HC 023 well 50 N N NA N Y N NA 0 differentiated HC 025 well 140 N N NA N Y N 0 0 differentiated HC 026 well 30 Y Y NA Y N Y NA 4 differentiated HC 027 well 15 N N NA Y Y N Y NA 4 differentiated HC 028 moderately 120 N Y NA Y Y N 0 0 differentiated HC 030 moderately 16 NA NA NA N N Y NA 4 differentiated HC 032 well 60 N NA NA Y N Y NA 4 differentiated HC 034 well 140 N N NA Y Y N NA 0 differentiated HC 037 moderately 35 Y Y NA Y Y N Y NA 4 differentiated HC 038 moderately 50 N N NA Y N Y NA 4 differentiated HC 041 well 30 N N NA N N Y NA 4 differentiated HC 042 moderately 130 prob- Y NA N N N 2 1 differentiated able HC 043 moderately 15 N N NA Y N Y N 4 differentiated HC 052 moderately 110 N Y NA Y N Y N 4 differentiated HC 058 moderately 100 N N NA N N N 2 3 differentiated HC 060 well 55 N N NA differentiated HC 064 moderately 40 N N NA N N N 2 2 differentiated HC 066 well 75 N N NA Y N Y NA 4 moderately HC 101 well 35 Y Y 18 Y Y N Y 2 4 moderately HC 102 Peu 200 Y Y 7 N N N N 1 1 differencie HC 103 well 55 N Y 8 N Y N Y 3 4 moderately HC 104 well 160 prob- Y 10 Y N Y N 0 1 moderately able HC 105 moderately 40 Y Y 20 Y Y N Y 2 4 differentiated HC 106 moderately 80 Y Y 32 Y N N Y 1 4 differentiated HC 107 well 60 N N 1 N N Y N 0 0-1 differentiated HC 108 moderately 26 N Y 18 N N N N 1 1 differentiated HC 109 well 30 N N <1 Y Y N Y 2 4 moderately HC 110 well 30 N Y 1a5 Y Y N Y 1 4 moderately HC 111 well 40 Y Y 45 Y Y N Y 1 4 moderately HC 112 well 18 N N 0 N N N N 2 2 moderately HC 113 well 50 Y Y 25 Y Y N Y 1 4 moderately HC 114 well 36 N N <1 N N N N 2 3 differentiated HC 115 well 90 N N 0 N N N N 2 1 differentiated HC 116 moderately 140 N N 12 N N N N 2 3 differentiated HC 117 well 28 N N 4 Y Y N Y 2 4 moderately HC 118 well 40 N N <1 N N Y N 0 1 differentiated HC 119 well 26 N Y 15 Y Y N Y 2 4 moderately HC 120 well 20 N Y 3 Y N N Y 1 4 moderately HC 121 moderately 150 prob- Y 8a30 Y Y N Y 2 4 differentiated able HC 122 well 20 Y Y 8 Y ? N Y 1 4
differentiated HC 123 moderately 43 prob- prob- 4 Y N N Y 2 4 differentiated able able HC 124 moderately 62 N N 4 N N N N 1 1 differentiated HC 125 moderately 33 N Y 2 Y N N Y 2 4 differentiated HC 126 well 130 Y Y 2 Y N Y N 0 1 differentiated HC 127 well 115 Y Y >100 N N N N 1 1 moderately HC 128 well 110 N Y 5 N N N N 2 2 moderately HC 129 moderately 30 N Y 40 Y N N N 2 3 differentiated HC 130 moderately 38 N prob- 12 N N N N 1 2 differentiated able HC 131 well 120 N Y 20a25 N N Y N 0 1 moderately HC 137 moderately 10 NA NA NA Y . . . differentiated HC 138 well 5.5 NA NA NA Y N . . differentiated HC 139 moderately 16 NA NA NA Y . . . differentiated HC 140 well 15 NA NA NA N N 0 1 differentiated HC 141 well 3.5 NA NA NA N N . . differentiated HC 142 well 8 NA NA NA Y . . . differentiated HC 143 well 3 NA NA NA N Y 1 4 differentiated HC 144 well 15 NA NA NA Y . . . differentiated HC 145 well 6 NA NA NA N . 0 3 differentiated HC 146 well 7.5 NA NA NA N N . 2 differentiated HC 147 moderately 15 NA NA NA N N 0 3 differentiated HC 148 moderately 21 NA NA NA Y N . . differentiated HC 149 NA 8 NA NA NA N N 0 0 HC 150 moderately 13 NA NA NA N . 0 3 differentiated HC 151 well 6.5 NA NA NA N Y 2 4 differentiated HC 152 well 3.5 NA NA NA N N 0 2 differentiated HC 153 well 5 NA NA NA N . 0 3 differentiated HC 154 well 45 Y Y 25 N N Y N 0 1 differentiated HC 155 well 24 N N 1 N N N Y 2 4 differentiated HC 156 moderately 70 N Y 16 Y N N Y 2 4 differentiated HC 157 well 140 Y Y 2 N N Y N 0 1 differentiated HC 159 well 35 N N NA N N N Y 2 4 differentiated HC 161 well 210 N Y 2 N N N N 1 1 differentiated HC 162 moderately 130 Y Y 77 N N Y N 0 0 differentiated HC 163 moderately 80 N Y 4 N N N Y 1 4 differentiated HC 164 well 90 N N 1 N N Y N 0 1 differentiated HC 165 well 30 N Y 4 N N N N 0 2 differentiated HC 168 well 25 N N 1 Y Y N Y 2 4 differentiated HC 169 well 35 N N NA N N N Y 2 4 differentiated HC 170 well 220 N N 0 N N Y N 0 0 differentiated HC 171 Peu 70 Y Y 10 Y N N N 1 2 differencie HC 172 moderately 40 N Y 28 N N N N 2 3 differentiated HC 173 well 40 N N 0 N N Y N 0 0 differentiated HC 176 well 75 N N NA N N Y N 0 0 differentiated HC 177 moderately 2.3 NA N NA Y A1 F4 differentiated HC 178 well 6.5 NA N NA Y A1 F4 differentiated HC 179 well-moder- 9 NA Y NA Y A2 F1 ate-poor HC 180 moderately 15 NA Y NA Y A2 F2 differentiated HC 181 well 3.5 NA Y NA Y A1 F4 moderately HC 182 well 11 NA N NA N F1 differentiated HC 183 well 8 NA Y NA N A1 F3 differentiated HC 184 well 6.5 NA N NA N F1 differentiated HC 185 moderately 3.5 NA N NA N A1 F4 differentiated HC 186 well 17 NA Y NA N F0 moderately HC 187 well 8 NA Y NA N F4 differentiated HC 188 well 13 NA N NA N F0 differentiated HC 189 well 22 NA Y NA Y F1 differentiated HC 190 moderately 15 NA N NA Y A1 F3 differentiated chronic HC 000 tumor viral Etiology Etiology identification hepatitis HBV HCV alcool Hemochromatos --NASH HC 001 N N N Y N N HC 003 Y N Y N N N HC 004 N N N N N N HC 006 N N N Y Y N HC 007 N N N Y N N HC 008 Y N Y N N N HC 009 N N N Y N N HC 010 Y Y N N N N HC 011 Y Y Y N N N HC 012 Y Y N N N N HC 014 Y N Y Y N N HC 015 N N N Y N N HC 017 Y Y N N N N HC 018 N N N Y N N HC 020 N N N Y N N HC 021 N N N Y N N HC 022 N N N Y N N HC 023 N N N N N N HC 025 N N N N N N HC 026 Y Y N N N N HC 027 Y N Y N N N HC 028 N N N N N N HC 030 N N N Y N N HC 032 Y N Y N N N HC 034 N N N N N N HC 037 N N N Y N N HC 038 Y N Y N N N HC 041 Y N Y N N N HC 042 Y Y N N N N HC 043 Y N Y N N N HC 052 Y Y N N N N HC 058 Y N Y N N N HC 060 HC 064 Y N Y N N N HC 066 Y Y N Y N N HC 101 Y Y Y Y N N HC 102 Y Y Y N N N HC 103 Y Y N N N N HC 104 N N N N N N HC 105 Y N Y N N N HC 106 Y Y N N N N HC 107 N N N Y N N HC 108 Y N Y N N N HC 109 N N N Y N Y HC 110 Y N Y Y N N HC 111 N N N Y N N HC 112 N N N N N Y HC 113 Y N Y N N N HC 114 N N N Y N N HC 115 N N N N N Y HC 116 Y Y N N N N HC 117 Y N Y N N N HC 118 N N N N N N HC 119 Y N Y Y N N HC 120 Y Y N N N N HC 121 N N N Y N Y HC 122 Y Y N N N N HC 123 Y N Y N N N HC 124 N N N N N Y HC 125 N N N Y N N HC 126 N N N N N N HC 127 Y Y N N N N HC 128 N N N Y N N HC 129 Y N Y N N N HC 130 Y Y N N N N HC 131 N N N N N N HC 137 N N N Y N N HC 138 N N N N N N HC 139 N N N N N N HC 140 N N N N N N HC 141 N N N N N N HC 142 N N N Y N N HC 143 N N N N Y N HC 144 N N N N N N HC 145 N N N Y N N HC 146 Y Y N Y N N HC 147 N N N N Y N HC 148 N N N N N N HC 149 N N N N N N HC 150 N N N Y N N HC 151 N N N N N Y HC 152 N N N N Y N HC 153 Y Y N Y N N HC 154 N N N N N N HC 155 N N N Y N N HC 156 Y N Y N N N HC 157 N N N N N N HC 159 N N N Y N N HC 161 N N N N N N HC 162 N N N N N N HC 163 N N N Y N N HC 164 N N N N N N HC 165 N N N N Y N HC 168 Y N Y N N N HC 169 N N N Y N N HC 170 N N N N N N HC 171 N N N Y N N HC 172 N N N N Y N HC 173 N N N N N N HC 176 N N N N N N HC 177 Y Y N N N N HC 178 N N N Y N N HC 179 Y N Y N N N HC 180 Y Y N N N N HC 181 N N N Y N N HC 182 Y N Y N N N HC 183 Y Y N N N N HC 184 N N N Y N N HC 185 Y N Y N N N HC 186 NA NA NA NA NA NA HC 187 N N N Y N N HC 188 N N N Y N N HC 189 N N N Y N N HC 190 Y Y N N N N
REFERENCES
[0327] Assou, S., Le Carrour, T., Tondeur, S., Strom, S., Gabelle, A., Marty, S., Nadal, L., Pantesco, V., Reme, T., Hugnot, J. P., et al. (2007). A meta-analysis of human embryonic stem cells transcriptome integrated into a web-based expression atlas. Stem Cells 25, 961-973. [0328] Boyault, S., Rickman, D. S., de Reynies, A., Balabaud, C., Rebouissou, S., Jeannot, E., Herault, A., Saric, J., Belghiti, J., Franco, D., et al. (2007). Transcriptome classification of HCC is related to gene alterations and to new therapeutic targets. Hepatology 45, 42-52. [0329] Finegold, M. J., Lopez-Terrada, D. H., Bowen, J., Washington, M. K., and Qualman, S. J. (2007). Protocol for the examination of specimens from pediatric patients with hepatoblastoma. Arch Pathol Lab Med 131, 520-529. [0330] Fodde, R., and Brabletz, T. (2007). Wnt/beta-catenin signaling in cancer sternness and malignant behavior. Curr Opin Cell Biol 19, 150-158. [0331] Glinsky, G. V., Berezovska, O., and Glinskii, A. B. (2005). Microarray analysis identifies a death-from-cancer signature predicting therapy failure in patients with multiple types of cancer. J Clin Invest 115, 1503-1521. [0332] Hirschman, B. A., Pollock, B. H., and Tomlinson, G. E. (2005). The spectrum of APC mutations in children with hepatoblastoma from familial adenomatous polyposis kindreds. J Pediatr 147, 263-266. [0333] Irizarry, R. A., Hobbs, B., Collin, F., Beazer-Barclay, Y. D., Antonellis, K. J., Scherf, U., and Speed, T. P. (2003). Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249-264. [0334] Lee, J. S., Heo, J., Libbrecht, L., Chu, I. S., Kaposi-Novak, P., Calvisi, D. F., Mikaelyan, A., Roberts, L. R., Demetris, A. J., Sun, Z., et al., (2006). A novel prognostic subtype of human hepatocellular carcinoma derived from hepatic progenitor cells. Nat Med 12, 410-416. [0335] McLin, V. A., Rankin, S. A., and Zorn, A. M. (2007). Repression of Wnt/β-catenin signaling in the anterior endoderm is essential for liver and pancreas development. Development 134, 2207-2217. [0336] Perilongo, G., Shafford, E., and Plaschkes, J. (2000). SIOPEL trials using preoperative chemotherapy in hepatoblastoma. Lancet Oncol 1, 94-100. [0337] Rowland, J. M. (2002). Hepatoblastoma: assessment of criteria for histologic classification. Med Pediatr Oncol 39, 478-483. [0338] Schnater, J. M., Kohler, S. E., Lamers, W. H., von Schweinitz, D., and Aronson, D. C. (2003). Where do we stand with hepatoblastoma? A review. Cancer 98, 668-678. [0339] Taniguchi, K., Roberts, L. R., Aderca, I. N., Dong, X., Qian, C., Murphy, L. M., Nagorney, D. M., Burgart, L. J., Roche, P. C., Smith, D. I., et al. (2002). Mutational spectrum of beta-catenin, AXIN1, and AXIN2 in hepatocellular carcinomas and hepatoblastomas. Oncogene 21, 4863-4871. [0340] Wei, Y., Fabre, M., Branchereau, S., Gauthier, F., Perilongo, G., and Buendia, M. A. (2000). Activation of beta-catenin in epithelial and mesenchymal hepatoblastomas. Oncogene 19, 498-504. [0341] Lustgarten, J. L. et al (2008)--Improving classification performance with discretization on biomedical datasets. AMIA 2008 Symposium Proceedings,
Sequence CWU
1
140 1 2014 DNA Homo sapiens CDS (31)..(1857) AFP mRNA 1 ccactgccaa
taacaaaata actagcaacc atg aag tgg gtg gaa tca att ttt 54
Met Lys Trp Val Glu Ser Ile Phe
1 5 tta att ttc cta cta aat ttt act gaa tcc aga
aca ctg cat aga aat 102 Leu Ile Phe Leu Leu Asn Phe Thr Glu Ser Arg
Thr Leu His Arg Asn 10 15 20 gaa
tat gga ata gct tcc ata ttg gat tct tac caa tgt act gca gag 150 Glu
Tyr Gly Ile Ala Ser Ile Leu Asp Ser Tyr Gln Cys Thr Ala Glu 25
30 35 40 ata agt tta gct gac ctg
gct acc ata ttt ttt gcc cag ttt gtt caa 198 Ile Ser Leu Ala Asp Leu
Ala Thr Ile Phe Phe Ala Gln Phe Val Gln 45
50 55 gaa gcc act tac aag gaa gta agc aaa atg gtg aaa
gat gca ttg act 246 Glu Ala Thr Tyr Lys Glu Val Ser Lys Met Val Lys
Asp Ala Leu Thr 60 65 70
gca att gag aaa ccc act gga gat gaa cag tct tca ggg tgt tta gaa 294
Ala Ile Glu Lys Pro Thr Gly Asp Glu Gln Ser Ser Gly Cys Leu Glu
75 80 85 aac cag cta cct gcc ttt ctg
gaa gaa ctt tgc cat gag aaa gaa att 342 Asn Gln Leu Pro Ala Phe Leu
Glu Glu Leu Cys His Glu Lys Glu Ile 90 95
100 ttg gag aag tac gga cat tca gac tgc tgc agc caa agt gaa gag gga
390 Leu Glu Lys Tyr Gly His Ser Asp Cys Cys Ser Gln Ser Glu Glu Gly
105 110 115 120 aga cat
aac tgt ttt ctt gca cac aaa aag ccc act cca gca tcg atc 438 Arg His
Asn Cys Phe Leu Ala His Lys Lys Pro Thr Pro Ala Ser Ile
125 130 135 cca ctt ttc caa gtt cca gaa
cct gtc aca agc tgt gaa gca tat gaa 486 Pro Leu Phe Gln Val Pro Glu
Pro Val Thr Ser Cys Glu Ala Tyr Glu 140 145
150 gaa gac agg gag aca ttc atg aac aaa ttc att tat gag ata
gca aga 534 Glu Asp Arg Glu Thr Phe Met Asn Lys Phe Ile Tyr Glu Ile
Ala Arg 155 160 165 agg cat ccc
ttc ctg tat gca cct aca att ctt ctt tgg gct gct cgc 582 Arg His Pro
Phe Leu Tyr Ala Pro Thr Ile Leu Leu Trp Ala Ala Arg 170
175 180 tat gac aaa ata att cca tct tgc tgc aaa gct gaa
aat gca gtt gaa 630 Tyr Asp Lys Ile Ile Pro Ser Cys Cys Lys Ala Glu
Asn Ala Val Glu 185 190 195
200 tgc ttc caa aca aag gca gca aca gtt aca aaa gaa tta aga gaa agc
678 Cys Phe Gln Thr Lys Ala Ala Thr Val Thr Lys Glu Leu Arg Glu Ser
205 210 215 agc ttg tta aat
caa cat gca tgt gca gta atg aaa aat ttt ggg acc 726 Ser Leu Leu Asn
Gln His Ala Cys Ala Val Met Lys Asn Phe Gly Thr 220
225 230 cga act ttc caa gcc ata act gtt act aaa ctg
agt cag aag ttt acc 774 Arg Thr Phe Gln Ala Ile Thr Val Thr Lys Leu
Ser Gln Lys Phe Thr 235 240 245
aaa gtt aat ttt act gaa atc cag aaa cta gtc ctg gat gtg gcc cat 822
Lys Val Asn Phe Thr Glu Ile Gln Lys Leu Val Leu Asp Val Ala His 250
255 260 gta cat gag cac tgt tgc aga gga gat
gtg ctg gat tgt ctg cag gat 870 Val His Glu His Cys Cys Arg Gly Asp
Val Leu Asp Cys Leu Gln Asp 265 270 275
280 ggg gaa aaa atc atg tcc tac ata tgt tct caa caa gac act
ctg tca 918 Gly Glu Lys Ile Met Ser Tyr Ile Cys Ser Gln Gln Asp Thr
Leu Ser 285 290 295 aac
aaa ata aca gaa tgc tgc aaa ctg acc acg ctg gaa cgt ggt caa 966 Asn
Lys Ile Thr Glu Cys Cys Lys Leu Thr Thr Leu Glu Arg Gly Gln
300 305 310 tgt ata att cat gca gaa aat
gat gaa aaa cct gaa ggt cta tct cca 1014 Cys Ile Ile His Ala Glu Asn
Asp Glu Lys Pro Glu Gly Leu Ser Pro 315 320
325 aat cta aac agg ttt tta gga gat aga gat ttt aac caa ttt tct
tca 1062 Asn Leu Asn Arg Phe Leu Gly Asp Arg Asp Phe Asn Gln Phe Ser
Ser 330 335 340 ggg gaa aaa aat atc
ttc ttg gca agt ttt gtt cat gaa tat tca aga 1110 Gly Glu Lys Asn Ile
Phe Leu Ala Ser Phe Val His Glu Tyr Ser Arg 345 350
355 360 aga cat cct cag ctt gct gtc tca gta att
cta aga gtt gct aaa gga 1158 Arg His Pro Gln Leu Ala Val Ser Val Ile
Leu Arg Val Ala Lys Gly 365 370
375 tac cag gag tta ttg gag aag tgt ttc cag act gaa aac cct ctt gaa
1206 Tyr Gln Glu Leu Leu Glu Lys Cys Phe Gln Thr Glu Asn Pro Leu Glu
380 385 390 tgc caa gat aaa
gga gaa gaa gaa tta cag aaa tac atc cag gag agc 1254 Cys Gln Asp Lys
Gly Glu Glu Glu Leu Gln Lys Tyr Ile Gln Glu Ser 395
400 405 caa gca ttg gca aag cga agc tgc ggc ctc ttc cag
aaa cta gga gaa 1302 Gln Ala Leu Ala Lys Arg Ser Cys Gly Leu Phe Gln
Lys Leu Gly Glu 410 415 420 tat tac
tta caa aat gcg ttt ctc gtt gct tac aca aag aaa gcc ccc 1350 Tyr Tyr
Leu Gln Asn Ala Phe Leu Val Ala Tyr Thr Lys Lys Ala Pro 425
430 435 440 cag ctg acc tcg tcg gag ctg
atg gcc atc acc aga aaa atg gca gcc 1398 Gln Leu Thr Ser Ser Glu Leu
Met Ala Ile Thr Arg Lys Met Ala Ala 445
450 455 aca gca gcc act tgt tgc caa ctc agt gag gac aaa
cta ttg gcc tgt 1446 Thr Ala Ala Thr Cys Cys Gln Leu Ser Glu Asp Lys
Leu Leu Ala Cys 460 465 470
ggc gag gga gcg gac att att atc gga cac tta tgt atc aga cat gaa 1494
Gly Glu Gly Ala Asp Ile Ile Ile Gly His Leu Cys Ile Arg His Glu
475 480 485 atg act cca gta aac cct ggt
gtt ggc cag tgc tgc act tct tca tat 1542 Met Thr Pro Val Asn Pro Gly
Val Gly Gln Cys Cys Thr Ser Ser Tyr 490 495
500 gcc aac agg agg cca tgc ttc agc agc ttg gtg gtg gat gaa aca tat
1590 Ala Asn Arg Arg Pro Cys Phe Ser Ser Leu Val Val Asp Glu Thr Tyr
505 510 515 520 gtc cct
cct gca ttc tct gat gac aag ttc att ttc cat aag gat ctg 1638 Val Pro
Pro Ala Phe Ser Asp Asp Lys Phe Ile Phe His Lys Asp Leu
525 530 535 tgc caa gct cag ggt gta gcg
ctg caa aca atg aag caa gag ttt ctc 1686 Cys Gln Ala Gln Gly Val Ala
Leu Gln Thr Met Lys Gln Glu Phe Leu 540 545
550 att aac ctt gtg aag caa aag cca caa ata aca gag gaa caa
ctt gag 1734 Ile Asn Leu Val Lys Gln Lys Pro Gln Ile Thr Glu Glu Gln
Leu Glu 555 560 565 gct gtc att
gca gat ttc tca ggc ctg ttg gag aaa tgc tgc caa ggc 1782 Ala Val Ile
Ala Asp Phe Ser Gly Leu Leu Glu Lys Cys Cys Gln Gly 570
575 580 cag gaa cag gaa gtc tgc ttt gct gaa gag gga caa
aaa ctg att tca 1830 Gln Glu Gln Glu Val Cys Phe Ala Glu Glu Gly Gln
Lys Leu Ile Ser 585 590 595
600 aaa act cgt gct gct ttg gga gtt taa attacttcag gggaagagaa
1877 Lys Thr Arg Ala Ala Leu Gly Val 605 gacaaaacga
gtctttcatt cggtgtgaac ttttctcttt aattttaact gatttaacac 1937 tttttgtgaa
ttaatgaaat gataaagact tttatgtgag atttccttat cacagaaata 1997 aaatatctcc
aaatgtt 2014 2 608 PRT
Homo sapiens 2 Met Lys Trp Val Glu Ser Ile Phe Leu Ile Phe Leu Leu Asn
Phe Thr 1 5 10 15 Glu
Ser Arg Thr Leu His Arg Asn Glu Tyr Gly Ile Ala Ser Ile Leu
20 25 30 Asp Ser Tyr Gln Cys Thr Ala
Glu Ile Ser Leu Ala Asp Leu Ala Thr 35 40
45 Ile Phe Phe Ala Gln Phe Val Gln Glu Ala Thr Tyr Lys Glu Val
Ser 50 55 60 Lys Met Val Lys Asp
Ala Leu Thr Ala Ile Glu Lys Pro Thr Gly Asp 65 70
75 80 Glu Gln Ser Ser Gly Cys Leu Glu Asn Gln
Leu Pro Ala Phe Leu Glu 85 90
95 Glu Leu Cys His Glu Lys Glu Ile Leu Glu Lys Tyr Gly His Ser Asp
100 105 110 Cys Cys Ser Gln
Ser Glu Glu Gly Arg His Asn Cys Phe Leu Ala His 115
120 125 Lys Lys Pro Thr Pro Ala Ser Ile Pro Leu Phe Gln
Val Pro Glu Pro 130 135 140 Val Thr
Ser Cys Glu Ala Tyr Glu Glu Asp Arg Glu Thr Phe Met Asn 145
150 155 160 Lys Phe Ile Tyr Glu Ile Ala
Arg Arg His Pro Phe Leu Tyr Ala Pro 165
170 175 Thr Ile Leu Leu Trp Ala Ala Arg Tyr Asp Lys Ile
Ile Pro Ser Cys 180 185 190
Cys Lys Ala Glu Asn Ala Val Glu Cys Phe Gln Thr Lys Ala Ala Thr
195 200 205 Val Thr Lys Glu Leu Arg Glu
Ser Ser Leu Leu Asn Gln His Ala Cys 210 215
220 Ala Val Met Lys Asn Phe Gly Thr Arg Thr Phe Gln Ala Ile Thr Val
225 230 235 240 Thr Lys
Leu Ser Gln Lys Phe Thr Lys Val Asn Phe Thr Glu Ile Gln
245 250 255 Lys Leu Val Leu Asp Val Ala
His Val His Glu His Cys Cys Arg Gly 260 265
270 Asp Val Leu Asp Cys Leu Gln Asp Gly Glu Lys Ile Met Ser
Tyr Ile 275 280 285 Cys Ser Gln
Gln Asp Thr Leu Ser Asn Lys Ile Thr Glu Cys Cys Lys 290
295 300 Leu Thr Thr Leu Glu Arg Gly Gln Cys Ile Ile His
Ala Glu Asn Asp 305 310 315
320 Glu Lys Pro Glu Gly Leu Ser Pro Asn Leu Asn Arg Phe Leu Gly Asp
325 330 335 Arg Asp Phe Asn
Gln Phe Ser Ser Gly Glu Lys Asn Ile Phe Leu Ala 340
345 350 Ser Phe Val His Glu Tyr Ser Arg Arg His Pro
Gln Leu Ala Val Ser 355 360 365
Val Ile Leu Arg Val Ala Lys Gly Tyr Gln Glu Leu Leu Glu Lys Cys 370
375 380 Phe Gln Thr Glu Asn Pro Leu Glu Cys
Gln Asp Lys Gly Glu Glu Glu 385 390 395
400 Leu Gln Lys Tyr Ile Gln Glu Ser Gln Ala Leu Ala Lys Arg
Ser Cys 405 410 415 Gly
Leu Phe Gln Lys Leu Gly Glu Tyr Tyr Leu Gln Asn Ala Phe Leu
420 425 430 Val Ala Tyr Thr Lys Lys Ala
Pro Gln Leu Thr Ser Ser Glu Leu Met 435 440
445 Ala Ile Thr Arg Lys Met Ala Ala Thr Ala Ala Thr Cys Cys Gln
Leu 450 455 460 Ser Glu Asp Lys Leu
Leu Ala Cys Gly Glu Gly Ala Asp Ile Ile Ile 465 470
475 480 Gly His Leu Cys Ile Arg His Glu Met Thr
Pro Val Asn Pro Gly Val 485 490
495 Gly Gln Cys Cys Thr Ser Ser Tyr Ala Asn Arg Arg Pro Cys Phe Ser
500 505 510 Ser Leu Val Val
Asp Glu Thr Tyr Val Pro Pro Ala Phe Ser Asp Asp 515
520 525 Lys Phe Ile Phe His Lys Asp Leu Cys Gln Ala Gln
Gly Val Ala Leu 530 535 540 Gln Thr
Met Lys Gln Glu Phe Leu Ile Asn Leu Val Lys Gln Lys Pro 545
550 555 560 Gln Ile Thr Glu Glu Gln Leu
Glu Ala Val Ile Ala Asp Phe Ser Gly 565
570 575 Leu Leu Glu Lys Cys Cys Gln Gly Gln Glu Gln Glu
Val Cys Phe Ala 580 585 590
Glu Glu Gly Gln Lys Leu Ile Ser Lys Thr Arg Ala Ala Leu Gly Val
595 600 605 3 2001 DNA Homo sapiens CDS
(45)..(1598) ALDH2 mRNA 3 ctagctctgc tctcggtccg ctcgctgtcc gctagcccgc
tgcg atg ttg cgc gct 56
Met Leu Arg Ala 1
gcc gcc cgc ttc ggg ccc cgc ctg ggc cgc cgc ctc ttg tca gcc gcc 104
Ala Ala Arg Phe Gly Pro Arg Leu Gly Arg Arg Leu Leu Ser Ala Ala 5
10 15 20 gcc acc cag gcc gtg
cct gcc ccc aac cag cag ccc gag gtc ttc tgc 152 Ala Thr Gln Ala Val
Pro Ala Pro Asn Gln Gln Pro Glu Val Phe Cys 25
30 35 aac cag att ttc ata aac aat gaa tgg cac gat
gcc gtc agc agg aaa 200 Asn Gln Ile Phe Ile Asn Asn Glu Trp His Asp
Ala Val Ser Arg Lys 40 45
50 aca ttc ccc acc gtc aat ccg tcc act gga gag gtc atc tgt cag gta
248 Thr Phe Pro Thr Val Asn Pro Ser Thr Gly Glu Val Ile Cys Gln Val
55 60 65 gct gaa ggg gac aag gaa gat
gtg gac aag gca gtg aag gcc gcc cgg 296 Ala Glu Gly Asp Lys Glu Asp
Val Asp Lys Ala Val Lys Ala Ala Arg 70 75
80 gcc gcc ttc cag ctg ggc tca cct tgg cgc cgc atg gac gca tca cac
344 Ala Ala Phe Gln Leu Gly Ser Pro Trp Arg Arg Met Asp Ala Ser His
85 90 95 100 agg ggc
cgg ctg ctg aac cgc ctg gcc gat ctg atc gag cgg gac cgg 392 Arg Gly
Arg Leu Leu Asn Arg Leu Ala Asp Leu Ile Glu Arg Asp Arg
105 110 115 acc tac ctg gcg gcc ttg gag
acc ctg gac aat ggc aag ccc tat gtc 440 Thr Tyr Leu Ala Ala Leu Glu
Thr Leu Asp Asn Gly Lys Pro Tyr Val 120 125
130 atc tcc tac ctg gtg gat ttg gac atg gtc ctc aaa tgt ctc
cgg tat 488 Ile Ser Tyr Leu Val Asp Leu Asp Met Val Leu Lys Cys Leu
Arg Tyr 135 140 145 tat gcc ggc
tgg gct gat aag tac cac ggg aaa acc atc ccc att gac 536 Tyr Ala Gly
Trp Ala Asp Lys Tyr His Gly Lys Thr Ile Pro Ile Asp 150
155 160 gga gac ttc ttc agc tac aca cgc cat gaa cct gtg
ggg gtg tgc ggg 584 Gly Asp Phe Phe Ser Tyr Thr Arg His Glu Pro Val
Gly Val Cys Gly 165 170 175
180 cag atc att ccg tgg aat ttc ccg ctc ctg atg caa gca tgg aag ctg
632 Gln Ile Ile Pro Trp Asn Phe Pro Leu Leu Met Gln Ala Trp Lys Leu
185 190 195 ggc cca gcc ttg
gca act gga aac gtg gtt gtg atg aag gta gct gag 680 Gly Pro Ala Leu
Ala Thr Gly Asn Val Val Val Met Lys Val Ala Glu 200
205 210 cag aca ccc ctc acc gcc ctc tat gtg gcc aac
ctg atc aag gag gct 728 Gln Thr Pro Leu Thr Ala Leu Tyr Val Ala Asn
Leu Ile Lys Glu Ala 215 220 225
ggc ttt ccc cct ggt gtg gtc aac att gtg cct gga ttt ggc ccc acg 776
Gly Phe Pro Pro Gly Val Val Asn Ile Val Pro Gly Phe Gly Pro Thr 230
235 240 gct ggg gcc gcc att gcc tcc cat gag
gat gtg gac aaa gtg gca ttc 824 Ala Gly Ala Ala Ile Ala Ser His Glu
Asp Val Asp Lys Val Ala Phe 245 250 255
260 aca ggc tcc act gag att ggc cgc gta atc cag gtt gct gct
ggg agc 872 Thr Gly Ser Thr Glu Ile Gly Arg Val Ile Gln Val Ala Ala
Gly Ser 265 270 275 agc
aac ctc aag aga gtg acc ttg gag ctg ggg ggg aag agc ccc aac 920 Ser
Asn Leu Lys Arg Val Thr Leu Glu Leu Gly Gly Lys Ser Pro Asn
280 285 290 atc atc atg tca gat gcc gat
atg gat tgg gcc gtg gaa cag gcc cac 968 Ile Ile Met Ser Asp Ala Asp
Met Asp Trp Ala Val Glu Gln Ala His 295 300
305 ttc gcc ctg ttc ttc aac cag ggc cag tgc tgc tgt gcc ggc tcc
cgg 1016 Phe Ala Leu Phe Phe Asn Gln Gly Gln Cys Cys Cys Ala Gly Ser
Arg 310 315 320 acc ttc gtg cag gag
gac atc tat gat gag ttt gtg gag cgg agc gtt 1064 Thr Phe Val Gln Glu
Asp Ile Tyr Asp Glu Phe Val Glu Arg Ser Val 325 330
335 340 gcc cgg gcc aag tct cgg gtg gtc ggg aac
ccc ttt gat agc aag acc 1112 Ala Arg Ala Lys Ser Arg Val Val Gly Asn
Pro Phe Asp Ser Lys Thr 345 350
355 gag cag ggg ccg cag gtg gat gaa act cag ttt aag aag atc ctc ggc
1160 Glu Gln Gly Pro Gln Val Asp Glu Thr Gln Phe Lys Lys Ile Leu Gly
360 365 370 tac atc aac acg
ggg aag caa gag ggg gcg aag ctg ctg tgt ggt ggg 1208 Tyr Ile Asn Thr
Gly Lys Gln Glu Gly Ala Lys Leu Leu Cys Gly Gly 375
380 385 ggc att gct gct gac cgt ggt tac ttc atc cag ccc
act gtg ttt gga 1256 Gly Ile Ala Ala Asp Arg Gly Tyr Phe Ile Gln Pro
Thr Val Phe Gly 390 395 400 gat gtg
cag gat ggc atg acc atc gcc aag gag gag atc ttc ggg cca 1304 Asp Val
Gln Asp Gly Met Thr Ile Ala Lys Glu Glu Ile Phe Gly Pro 405
410 415 420 gtg atg cag atc ctg aag ttc
aag acc ata gag gag gtt gtt ggg aga 1352 Val Met Gln Ile Leu Lys Phe
Lys Thr Ile Glu Glu Val Val Gly Arg 425
430 435 gcc aac aat tcc acg tac ggg ctg gcc gca gct gtc
ttc aca aag gat 1400 Ala Asn Asn Ser Thr Tyr Gly Leu Ala Ala Ala Val
Phe Thr Lys Asp 440 445 450
ttg gac aag gcc aat tac ctg tcc cag gcc ctc cag gcg ggc act gtg 1448
Leu Asp Lys Ala Asn Tyr Leu Ser Gln Ala Leu Gln Ala Gly Thr Val
455 460 465 tgg gtc aac tgc tat gat gtg
ttt gga gcc cag tca ccc ttt ggt ggc 1496 Trp Val Asn Cys Tyr Asp Val
Phe Gly Ala Gln Ser Pro Phe Gly Gly 470 475
480 tac aag atg tcg ggg agt ggc cgg gag ttg ggc gag tac ggg ctg cag
1544 Tyr Lys Met Ser Gly Ser Gly Arg Glu Leu Gly Glu Tyr Gly Leu Gln
485 490 495 500 gca tac
act gaa gtg aaa act gtc aca gtc aaa gtg cct cag aag aac 1592 Ala Tyr
Thr Glu Val Lys Thr Val Thr Val Lys Val Pro Gln Lys Asn
505 510 515 tca taa gaatcatgca agcttcctcc
ctcagccatt gatggaaagt tcagcaagat 1648 Ser cagcaacaaa accaagaaaa
atgatccttg cgtgctgaat atctgaaaag agaaattttt 1708 cctacaaaat ctcttgggtc
aagaaagttc tagaatttga attgataaac atggtgggtt 1768 ggctgagggt aagagtatat
gaggaacctt ttaaacgaca acaatactgc tagctttcag 1828 gatgattttt aaaaaataga
ttcaaatgtg ttatcctctc tctgaaacgc ttcctataac 1888 tcgagtttat aggggaagaa
aaagctattg tttacaatta tatcaccatt aaggcaactg 1948 ctacaccctg ctttgtattc
tgggctaaga ttcattaaaa actagctgct ctt 2001 4 517 PRT Homo sapiens
4 Met Leu Arg Ala Ala Ala Arg Phe Gly Pro Arg Leu Gly Arg Arg Leu 1
5 10 15 Leu Ser Ala Ala Ala
Thr Gln Ala Val Pro Ala Pro Asn Gln Gln Pro 20
25 30 Glu Val Phe Cys Asn Gln Ile Phe Ile Asn Asn Glu
Trp His Asp Ala 35 40 45 Val
Ser Arg Lys Thr Phe Pro Thr Val Asn Pro Ser Thr Gly Glu Val 50
55 60 Ile Cys Gln Val Ala Glu Gly Asp Lys Glu
Asp Val Asp Lys Ala Val 65 70 75
80 Lys Ala Ala Arg Ala Ala Phe Gln Leu Gly Ser Pro Trp Arg Arg
Met 85 90 95 Asp Ala
Ser His Arg Gly Arg Leu Leu Asn Arg Leu Ala Asp Leu Ile 100
105 110 Glu Arg Asp Arg Thr Tyr Leu Ala Ala
Leu Glu Thr Leu Asp Asn Gly 115 120
125 Lys Pro Tyr Val Ile Ser Tyr Leu Val Asp Leu Asp Met Val Leu Lys
130 135 140 Cys Leu Arg Tyr Tyr Ala Gly
Trp Ala Asp Lys Tyr His Gly Lys Thr 145 150
155 160 Ile Pro Ile Asp Gly Asp Phe Phe Ser Tyr Thr Arg
His Glu Pro Val 165 170
175 Gly Val Cys Gly Gln Ile Ile Pro Trp Asn Phe Pro Leu Leu Met Gln
180 185 190 Ala Trp Lys Leu Gly Pro
Ala Leu Ala Thr Gly Asn Val Val Val Met 195 200
205 Lys Val Ala Glu Gln Thr Pro Leu Thr Ala Leu Tyr Val Ala
Asn Leu 210 215 220 Ile Lys Glu Ala
Gly Phe Pro Pro Gly Val Val Asn Ile Val Pro Gly 225 230
235 240 Phe Gly Pro Thr Ala Gly Ala Ala Ile
Ala Ser His Glu Asp Val Asp 245 250
255 Lys Val Ala Phe Thr Gly Ser Thr Glu Ile Gly Arg Val Ile Gln
Val 260 265 270 Ala Ala Gly
Ser Ser Asn Leu Lys Arg Val Thr Leu Glu Leu Gly Gly 275
280 285 Lys Ser Pro Asn Ile Ile Met Ser Asp Ala Asp
Met Asp Trp Ala Val 290 295 300 Glu
Gln Ala His Phe Ala Leu Phe Phe Asn Gln Gly Gln Cys Cys Cys 305
310 315 320 Ala Gly Ser Arg Thr Phe
Val Gln Glu Asp Ile Tyr Asp Glu Phe Val 325
330 335 Glu Arg Ser Val Ala Arg Ala Lys Ser Arg Val Val
Gly Asn Pro Phe 340 345 350
Asp Ser Lys Thr Glu Gln Gly Pro Gln Val Asp Glu Thr Gln Phe Lys
355 360 365 Lys Ile Leu Gly Tyr Ile Asn
Thr Gly Lys Gln Glu Gly Ala Lys Leu 370 375
380 Leu Cys Gly Gly Gly Ile Ala Ala Asp Arg Gly Tyr Phe Ile Gln Pro
385 390 395 400 Thr Val
Phe Gly Asp Val Gln Asp Gly Met Thr Ile Ala Lys Glu Glu
405 410 415 Ile Phe Gly Pro Val Met Gln
Ile Leu Lys Phe Lys Thr Ile Glu Glu 420 425
430 Val Val Gly Arg Ala Asn Asn Ser Thr Tyr Gly Leu Ala Ala
Ala Val 435 440 445 Phe Thr Lys
Asp Leu Asp Lys Ala Asn Tyr Leu Ser Gln Ala Leu Gln 450
455 460 Ala Gly Thr Val Trp Val Asn Cys Tyr Asp Val Phe
Gly Ala Gln Ser 465 470 475
480 Pro Phe Gly Gly Tyr Lys Met Ser Gly Ser Gly Arg Glu Leu Gly Glu
485 490 495 Tyr Gly Leu Gln
Ala Tyr Thr Glu Val Lys Thr Val Thr Val Lys Val 500
505 510 Pro Gln Lys Asn Ser 515 5 928 DNA
Homo sapiens CDS (97)..(768) APCS mRNA 5 gggcatgaat atcagacgct
agggggacag ccactgtgtt gtctgctacc ctcatcctgg 60 tcactgcttc tgctataaca
gccctaggcc aggaat atg aac aag ccg ctg ctt 114
Met Asn Lys Pro Leu Leu
1 5 tgg atc tct gtc ctc acc agc ctc ctg gaa gcc ttt
gct cac aca gac 162 Trp Ile Ser Val Leu Thr Ser Leu Leu Glu Ala Phe
Ala His Thr Asp 10 15 20
ctc agt ggg aag gtg ttt gta ttt cct aga gaa tct gtt act gat cat 210
Leu Ser Gly Lys Val Phe Val Phe Pro Arg Glu Ser Val Thr Asp His
25 30 35 gta aac ttg atc aca ccg ctg
gag aag cct cta cag aac ttt acc ttg 258 Val Asn Leu Ile Thr Pro Leu
Glu Lys Pro Leu Gln Asn Phe Thr Leu 40 45
50 tgt ttt cga gcc tat agt gat ctc tct cgt gcc tac agc ctc ttc tcc
306 Cys Phe Arg Ala Tyr Ser Asp Leu Ser Arg Ala Tyr Ser Leu Phe Ser
55 60 65 70 tac aat
acc caa ggc agg gat aat gag cta cta gtt tat aaa gaa aga 354 Tyr Asn
Thr Gln Gly Arg Asp Asn Glu Leu Leu Val Tyr Lys Glu Arg
75 80 85 gtt gga gag tat agt cta tac
att gga aga cac aaa gtt aca tcc aaa 402 Val Gly Glu Tyr Ser Leu Tyr
Ile Gly Arg His Lys Val Thr Ser Lys 90 95
100 gtt atc gaa aag ttc ccg gct cca gtg cac atc tgt gtg agc
tgg gag 450 Val Ile Glu Lys Phe Pro Ala Pro Val His Ile Cys Val Ser
Trp Glu 105 110 115 tcc tca tca
ggt att gct gaa ttt tgg atc aat ggg aca cct ttg gtg 498 Ser Ser Ser
Gly Ile Ala Glu Phe Trp Ile Asn Gly Thr Pro Leu Val 120
125 130 aaa aag ggt ctg cga cag ggt tac ttt gta gaa gct
cag ccc aag att 546 Lys Lys Gly Leu Arg Gln Gly Tyr Phe Val Glu Ala
Gln Pro Lys Ile 135 140 145
150 gtc ctg ggg cag gaa cag gat tcc tat ggg ggc aag ttt gat agg agc
594 Val Leu Gly Gln Glu Gln Asp Ser Tyr Gly Gly Lys Phe Asp Arg Ser
155 160 165 cag tcc ttt gtg
gga gag att ggg gat ttg tac atg tgg gac tct gtg 642 Gln Ser Phe Val
Gly Glu Ile Gly Asp Leu Tyr Met Trp Asp Ser Val 170
175 180 ctg ccc cca gaa aat atc ctg tct gcc tat cag
ggt acc cct ctc cct 690 Leu Pro Pro Glu Asn Ile Leu Ser Ala Tyr Gln
Gly Thr Pro Leu Pro 185 190 195
gcc aat atc ctg gac tgg cag gct ctg aac tat gaa atc aga gga tat 738
Ala Asn Ile Leu Asp Trp Gln Ala Leu Asn Tyr Glu Ile Arg Gly Tyr 200
205 210 gtc atc atc aaa ccc ttg gtg tgg gtc
tga ggtcttgact caacgagagc 788 Val Ile Ile Lys Pro Leu Val Trp Val
215 220 acttgaaaat gaaatgactg tctaagagat ctggtcaaag
caactggata ctagatctta 848 catctgcagc tctttcttct ttgaatttcc tatctgtatg
tctgcctaat taaaaaaata 908 tatattgtat tatgctacct
928 6 223 PRT Homo sapiens 6 Met Asn Lys Pro Leu
Leu Trp Ile Ser Val Leu Thr Ser Leu Leu Glu 1 5
10 15 Ala Phe Ala His Thr Asp Leu Ser Gly Lys Val
Phe Val Phe Pro Arg 20 25
30 Glu Ser Val Thr Asp His Val Asn Leu Ile Thr Pro Leu Glu Lys Pro
35 40 45 Leu Gln Asn Phe Thr Leu Cys
Phe Arg Ala Tyr Ser Asp Leu Ser Arg 50 55
60 Ala Tyr Ser Leu Phe Ser Tyr Asn Thr Gln Gly Arg Asp Asn Glu Leu
65 70 75 80 Leu Val
Tyr Lys Glu Arg Val Gly Glu Tyr Ser Leu Tyr Ile Gly Arg
85 90 95 His Lys Val Thr Ser Lys Val
Ile Glu Lys Phe Pro Ala Pro Val His 100 105
110 Ile Cys Val Ser Trp Glu Ser Ser Ser Gly Ile Ala Glu Phe
Trp Ile 115 120 125 Asn Gly Thr
Pro Leu Val Lys Lys Gly Leu Arg Gln Gly Tyr Phe Val 130
135 140 Glu Ala Gln Pro Lys Ile Val Leu Gly Gln Glu Gln
Asp Ser Tyr Gly 145 150 155
160 Gly Lys Phe Asp Arg Ser Gln Ser Phe Val Gly Glu Ile Gly Asp Leu
165 170 175 Tyr Met Trp Asp
Ser Val Leu Pro Pro Glu Asn Ile Leu Ser Ala Tyr 180
185 190 Gln Gly Thr Pro Leu Pro Ala Asn Ile Leu Asp
Trp Gln Ala Leu Asn 195 200 205
Tyr Glu Ile Arg Gly Tyr Val Ile Ile Lys Pro Leu Val Trp Val 210
215 220 7 577 DNA Homo sapiens CDS (5)..(388)
APOC4 mRNA 7 agaa atg tcc ctc ctc aga aac agg ctc cag gcc ctg cct gcc
ctg tgc 49 Met Ser Leu Leu Arg Asn Arg Leu Gln Ala Leu Pro Ala
Leu Cys 1 5 10 15
ctc tgc gtg ctg gtc ctg gcc tgc att ggg gca tgc cag cca gag gcc 97
Leu Cys Val Leu Val Leu Ala Cys Ile Gly Ala Cys Gln Pro Glu Ala
20 25 30 cag gaa gga acc ctg agc
ccc cca cca aag cta aag atg agt cgc tgg 145 Gln Glu Gly Thr Leu Ser
Pro Pro Pro Lys Leu Lys Met Ser Arg Trp 35
40 45 agc ctg gtg agg ggc agg atg aag gag ctg ctg gag
aca gtg gtg aac 193 Ser Leu Val Arg Gly Arg Met Lys Glu Leu Leu Glu
Thr Val Val Asn 50 55 60 agg
acc aga gac ggg tgg caa tgg ttc tgg agc ccg agc acc ttc cgg 241 Arg
Thr Arg Asp Gly Trp Gln Trp Phe Trp Ser Pro Ser Thr Phe Arg 65
70 75 ggc ttc atg cag acc tac tat gac gac cac
ctg agg gac ctg ggt ccg 289 Gly Phe Met Gln Thr Tyr Tyr Asp Asp His
Leu Arg Asp Leu Gly Pro 80 85 90
95 ctc acc aag gcc tgg ttc ctc gaa tcc aaa gac agc ctc ttg aag
aag 337 Leu Thr Lys Ala Trp Phe Leu Glu Ser Lys Asp Ser Leu Leu Lys
Lys 100 105 110 acc cac
agc ctg tgc ccc agg ctt gtc tgt ggg gac aag gac cag ggt 385 Thr His
Ser Leu Cys Pro Arg Leu Val Cys Gly Asp Lys Asp Gln Gly 115
120 125 taa aatgttcata aaagccaggt
gtggttgtgg cgggtgcctg tagtcccagc 438 tactcaggag gctgaggtag
gatgatggct tgagcccagg agttcgagac cagcctgggc 498 aacacagcga gatctcttgg
gggtaaaaca aaaagaaaaa aaaaagttca tacttctcca 558 ataaataaag tctcacctg
577 8 127 PRT Homo sapiens
8 Met Ser Leu Leu Arg Asn Arg Leu Gln Ala Leu Pro Ala Leu Cys Leu 1
5 10 15 Cys Val Leu Val Leu
Ala Cys Ile Gly Ala Cys Gln Pro Glu Ala Gln 20
25 30 Glu Gly Thr Leu Ser Pro Pro Pro Lys Leu Lys Met
Ser Arg Trp Ser 35 40 45 Leu
Val Arg Gly Arg Met Lys Glu Leu Leu Glu Thr Val Val Asn Arg 50
55 60 Thr Arg Asp Gly Trp Gln Trp Phe Trp Ser
Pro Ser Thr Phe Arg Gly 65 70 75
80 Phe Met Gln Thr Tyr Tyr Asp Asp His Leu Arg Asp Leu Gly Pro
Leu 85 90 95 Thr Lys
Ala Trp Phe Leu Glu Ser Lys Asp Ser Leu Leu Lys Lys Thr 100
105 110 His Ser Leu Cys Pro Arg Leu Val Cys
Gly Asp Lys Asp Gln Gly 115 120
125 9 2801 DNA Homo sapiens CDS (186)..(1073) AQP9 mRNA 9 ccaccagaag
acgattaagc cacagcctct aattggaacg gcatttgtac agtcagagac 60 tcttaccaga
catctccagg aatctgtgag ccattgtcaa aacgtccatt ttcatctggc 120 tgtgaaagtg
aggaccacaa caggtaggta ttggtagaaa caggagtcct cagagaagcc 180 ccaag atg
cag cct gag gga gca gaa aag gga aaa agc ttc aag cag aga 230 Met
Gln Pro Glu Gly Ala Glu Lys Gly Lys Ser Phe Lys Gln Arg 1
5 10 15 ctg gtc ttg aag agc agc
tta gcg aaa gaa acc ctc tct gag ttc ttg 278 Leu Val Leu Lys Ser Ser
Leu Ala Lys Glu Thr Leu Ser Glu Phe Leu 20
25 30 ggc acg ttc atc ttg att gtc ctt gga tgt ggc tgt
gtt gcc caa gct 326 Gly Thr Phe Ile Leu Ile Val Leu Gly Cys Gly Cys
Val Ala Gln Ala 35 40 45
att ctc agt cga gga cgt ttt gga ggg gtc atc act atc aat gtt gga 374
Ile Leu Ser Arg Gly Arg Phe Gly Gly Val Ile Thr Ile Asn Val Gly
50 55 60 ttt tca atg gca gtt gca atg
gcc att tat gtg gct ggc ggt gtc tct 422 Phe Ser Met Ala Val Ala Met
Ala Ile Tyr Val Ala Gly Gly Val Ser 65 70
75 ggt ggt cac atc aac cca gct gtg tct tta gca atg tgt ctc ttt gga
470 Gly Gly His Ile Asn Pro Ala Val Ser Leu Ala Met Cys Leu Phe Gly
80 85 90 95 cgg atg
aaa tgg ttc aaa ttg cca ttt tat gtg gga gcc cag ttc ttg 518 Arg Met
Lys Trp Phe Lys Leu Pro Phe Tyr Val Gly Ala Gln Phe Leu
100 105 110 gga gcc ttt gtg ggg gct gca
acc gtc ttt ggc att tac tat gat gga 566 Gly Ala Phe Val Gly Ala Ala
Thr Val Phe Gly Ile Tyr Tyr Asp Gly 115 120
125 ctt atg tcc ttt gct ggt gga aaa ctg ctg atc gtg gga gaa
aat gca 614 Leu Met Ser Phe Ala Gly Gly Lys Leu Leu Ile Val Gly Glu
Asn Ala 130 135 140 aca gca cac
att ttt gca aca tac cca gct ccg tat cta tct ctg gcg 662 Thr Ala His
Ile Phe Ala Thr Tyr Pro Ala Pro Tyr Leu Ser Leu Ala 145
150 155 aac gca ttt gca gat caa gtg gtg gcc acc atg ata
ctc ctc ata atc 710 Asn Ala Phe Ala Asp Gln Val Val Ala Thr Met Ile
Leu Leu Ile Ile 160 165 170
175 gtc ttt gcc atc ttt gac tcc aga aac ttg gga gcc ccc aga ggc cta
758 Val Phe Ala Ile Phe Asp Ser Arg Asn Leu Gly Ala Pro Arg Gly Leu
180 185 190 gag ccc att gcc
atc ggc ctc ctg att att gtc att gct tcc tcc ctg 806 Glu Pro Ile Ala
Ile Gly Leu Leu Ile Ile Val Ile Ala Ser Ser Leu 195
200 205 gga ctg aac agt ggc tgt gcc atg aac cca gct
cga gac ctg agt ccc 854 Gly Leu Asn Ser Gly Cys Ala Met Asn Pro Ala
Arg Asp Leu Ser Pro 210 215 220
aga ctt ttc act gcc ttg gca ggc tgg ggg ttt gaa gtc ttc aga gct 902
Arg Leu Phe Thr Ala Leu Ala Gly Trp Gly Phe Glu Val Phe Arg Ala 225
230 235 gga aac aac ttc tgg tgg att cct gta
gtg ggc cct ttg gtt ggt gct 950 Gly Asn Asn Phe Trp Trp Ile Pro Val
Val Gly Pro Leu Val Gly Ala 240 245 250
255 gtc att gga ggc ctc atc tat gtt ctt gtc att gaa atc cac
cat cca 998 Val Ile Gly Gly Leu Ile Tyr Val Leu Val Ile Glu Ile His
His Pro 260 265 270 gag
cct gac tca gtc ttt aag aca gaa caa tct gag gac aaa cca gag 1046 Glu
Pro Asp Ser Val Phe Lys Thr Glu Gln Ser Glu Asp Lys Pro Glu
275 280 285 aaa tat gaa ctc agt gtc atc
atg tag tggcatgctc agctctggat 1093 Lys Tyr Glu Leu Ser Val Ile
Met 290 295 ttgcagtcag tttgggattc tcttcagaaa
gatggcatct aagtgtctgt gttcttgtaa 1153 gcctgaggtg gaatccaccc agttttgtct
gctagccata tgggacatct aattggaaaa 1213 gcatctgcat aaaagtttgg aaacaatgac
cacttctcta ccattgtccc ccacccccac 1273 cccccagaat aacgctgact gtcccctgaa
acagccttct ctcctgccct gtttatttca 1333 tcctcgatgg gaattcttgc taggtaagca
ctaataactc ggcatgaaat ggtgtcacca 1393 aaaccctttt cttcagtatc gacaaagatt
acattctgag taccaaccaa accctaaatt 1453 gaaagacaaa actatggttt cagtcaacat
attcatgaat tagggagcta atgggttaag 1513 cttccagttc ccgctatgct actggatttg
tataaatact gatattctcc aaacctagtg 1573 gtgtagggag caagagaatg cagctggaag
gcacaagggg aggacattgt ggcattcaga 1633 aactgcagga gacaagatga atttgagaag
ccaaatggaa tttttaatgg aaaccattta 1693 tcagattaat ctcttgctct cctgcatttt
agaggacacc aattaatttc ctggtcttta 1753 gtatataata acctaaaata ccattgtaac
ctcagtcatg aaaaatacat cactctgtct 1813 ttttagctca aatgtatttt cctaattgcc
cacttgagaa cagacatttg acaagttata 1873 tcaacgactg tgcttgtcca ttattttaca
catgccctag aagccaaaac tgaaagccac 1933 tggatcctgg tctagctgaa tcttcagagt
gggaggtctc caaaaagata ttaccttatt 1993 gggcttaaca attcacaagg cactttcaca
cccattatct aatttaatcc tcataatgac 2053 tatgtgaggc aaatgccaca ttgcccattt
ttcagataaa gaaacaaaat cttagggaag 2113 ataagttgag ttgtccaaga gcacactgaa
agttgaatgt tatctaatgc attcctctac 2173 ctttcagaag atcagtagct ggctgacaat
ctttgccaaa tcttccttgc tagccagaag 2233 tggaattggc agcttctaga atatgtacac
ctctggacaa aatgttcctc aatcttaaga 2293 tacaaagacc ctcattgtct gggtctattc
ccacacttac tgagtacaga tgaaggaaag 2353 tggtagcaat ttaatcataa ctttcatttg
ctgaaaaaca ttatgagaag gcctcccttc 2413 ctaagccacc tctggtcttg ctaagtcttg
atcttgcttc ctgccagcac caaacattac 2473 attcagggga tttcctctgg ctcagtcttt
tccccttgaa gttctctaat agatgttact 2533 tttgacaaaa gatcgcctat gagttacaag
caccagggga tgctctacat caagggatgc 2593 accttcagtc aaactgtcaa aaagcccaga
attcccaaag gcattaggtt tcccaactgc 2653 tttgtgctga tatcagaaca gcagaaatta
aatgtgaaat gtttctgatg acttatgttc 2713 tacaatctat ggacatacgg gatttttttt
tcttgctttg aagctacctg gatatttcct 2773 atttgaaata aaattgttcg gtcattgt
2801 10 295 PRT Homo sapiens 10 Met
Gln Pro Glu Gly Ala Glu Lys Gly Lys Ser Phe Lys Gln Arg Leu 1
5 10 15 Val Leu Lys Ser Ser Leu Ala
Lys Glu Thr Leu Ser Glu Phe Leu Gly 20 25
30 Thr Phe Ile Leu Ile Val Leu Gly Cys Gly Cys Val Ala Gln
Ala Ile 35 40 45 Leu Ser Arg
Gly Arg Phe Gly Gly Val Ile Thr Ile Asn Val Gly Phe 50
55 60 Ser Met Ala Val Ala Met Ala Ile Tyr Val Ala Gly
Gly Val Ser Gly 65 70 75
80 Gly His Ile Asn Pro Ala Val Ser Leu Ala Met Cys Leu Phe Gly Arg
85 90 95 Met Lys Trp Phe Lys
Leu Pro Phe Tyr Val Gly Ala Gln Phe Leu Gly 100
105 110 Ala Phe Val Gly Ala Ala Thr Val Phe Gly Ile Tyr
Tyr Asp Gly Leu 115 120 125 Met
Ser Phe Ala Gly Gly Lys Leu Leu Ile Val Gly Glu Asn Ala Thr 130
135 140 Ala His Ile Phe Ala Thr Tyr Pro Ala Pro
Tyr Leu Ser Leu Ala Asn 145 150 155
160 Ala Phe Ala Asp Gln Val Val Ala Thr Met Ile Leu Leu Ile Ile
Val 165 170 175 Phe Ala
Ile Phe Asp Ser Arg Asn Leu Gly Ala Pro Arg Gly Leu Glu 180
185 190 Pro Ile Ala Ile Gly Leu Leu Ile Ile
Val Ile Ala Ser Ser Leu Gly 195 200
205 Leu Asn Ser Gly Cys Ala Met Asn Pro Ala Arg Asp Leu Ser Pro Arg
210 215 220 Leu Phe Thr Ala Leu Ala Gly
Trp Gly Phe Glu Val Phe Arg Ala Gly 225 230
235 240 Asn Asn Phe Trp Trp Ile Pro Val Val Gly Pro Leu
Val Gly Ala Val 245 250
255 Ile Gly Gly Leu Ile Tyr Val Leu Val Ile Glu Ile His His Pro Glu
260 265 270 Pro Asp Ser Val Phe Lys
Thr Glu Gln Ser Glu Asp Lys Pro Glu Lys 275 280
285 Tyr Glu Leu Ser Val Ile Met 290 295
11 3445 DNA Homo sapiens CDS (58)..(3315) BUB1 mRNA 11 cggcggcttc
tagtttgcgg ttcaggtttg gccgctgccg gccagcgtcc tctggcc 57 atg gac acc
ccg gaa aat gtc ctt cag atg ctt gaa gcc cac atg cag 105 Met Asp Thr
Pro Glu Asn Val Leu Gln Met Leu Glu Ala His Met Gln 1 5
10 15 agc tac aag ggc aat gac cct ctt ggt
gaa tgg gaa aga tac ata cag 153 Ser Tyr Lys Gly Asn Asp Pro Leu Gly
Glu Trp Glu Arg Tyr Ile Gln 20 25
30 tgg gta gaa gag aat ttt cct gag aat aaa gaa tac ttg ata act tta
201 Trp Val Glu Glu Asn Phe Pro Glu Asn Lys Glu Tyr Leu Ile Thr Leu
35 40 45 cta gaa cat tta atg aag
gaa ttt tta gat aag aag aaa tac cac aat 249 Leu Glu His Leu Met Lys
Glu Phe Leu Asp Lys Lys Lys Tyr His Asn 50 55
60 gac cca aga ttc atc agt tat tgt tta aaa ttt gct gag tac aac
agt 297 Asp Pro Arg Phe Ile Ser Tyr Cys Leu Lys Phe Ala Glu Tyr Asn
Ser 65 70 75 80 gac
ctc cat caa ttt ttt gag ttt ctg tac aac cat ggg att gga acc 345 Asp
Leu His Gln Phe Phe Glu Phe Leu Tyr Asn His Gly Ile Gly Thr
85 90 95 ctg tca tcc cct ctg tac att
gcc tgg gcg ggg cat ctg gaa gcc caa 393 Leu Ser Ser Pro Leu Tyr Ile
Ala Trp Ala Gly His Leu Glu Ala Gln 100 105
110 gga gag ctg cag cat gcc agt gct gtc ctt cag aga gga att
caa aac 441 Gly Glu Leu Gln His Ala Ser Ala Val Leu Gln Arg Gly Ile
Gln Asn 115 120 125 cag gct gaa
ccc aga gag ttc ctg caa caa caa tac agg tta ttt cag 489 Gln Ala Glu
Pro Arg Glu Phe Leu Gln Gln Gln Tyr Arg Leu Phe Gln 130
135 140 aca cgc ctc act gaa acc cat ttg cca gct caa gct
aga acc tca gaa 537 Thr Arg Leu Thr Glu Thr His Leu Pro Ala Gln Ala
Arg Thr Ser Glu 145 150 155
160 cct ctg cat aat gtt cag gtt tta aat caa atg ata aca tca aaa tca
585 Pro Leu His Asn Val Gln Val Leu Asn Gln Met Ile Thr Ser Lys Ser
165 170 175 aat cca gga aat
aac atg gcc tgc att tct aag aat cag ggt tca gag 633 Asn Pro Gly Asn
Asn Met Ala Cys Ile Ser Lys Asn Gln Gly Ser Glu 180
185 190 ctt tct gga gtg ata tct tca gct tgt gat aaa
gag tca aat atg gaa 681 Leu Ser Gly Val Ile Ser Ser Ala Cys Asp Lys
Glu Ser Asn Met Glu 195 200 205
cga aga gtg atc acg att tct aaa tca gaa tat tct gtg cac tca tct 729
Arg Arg Val Ile Thr Ile Ser Lys Ser Glu Tyr Ser Val His Ser Ser 210
215 220 ttg gca tcc aaa gtt gat gtt gag cag
gtt gtt atg tat tgc aag gag 777 Leu Ala Ser Lys Val Asp Val Glu Gln
Val Val Met Tyr Cys Lys Glu 225 230 235
240 aag ctt att cgt ggg gaa tca gaa ttt tcc ttt gaa gaa ttg
aga gcc 825 Lys Leu Ile Arg Gly Glu Ser Glu Phe Ser Phe Glu Glu Leu
Arg Ala 245 250 255 cag
aaa tac aat caa cgg aga aag cat gag caa tgg gta aat gaa gac 873 Gln
Lys Tyr Asn Gln Arg Arg Lys His Glu Gln Trp Val Asn Glu Asp
260 265 270 aga cat tat atg aaa agg aaa
gaa gca aat gct ttt gaa gaa cag cta 921 Arg His Tyr Met Lys Arg Lys
Glu Ala Asn Ala Phe Glu Glu Gln Leu 275 280
285 tta aaa cag aaa atg gat gaa ctt cat aag aag ttg cat cag gtg
gtg 969 Leu Lys Gln Lys Met Asp Glu Leu His Lys Lys Leu His Gln Val
Val 290 295 300 gag aca tcc cat gag
gat ctg ccc gct tcc cag gaa agg tcc gag gtt 1017 Glu Thr Ser His Glu
Asp Leu Pro Ala Ser Gln Glu Arg Ser Glu Val 305 310
315 320 aat cca gca cgt atg ggg cca agt gta ggc
tcc cag cag gaa ctg aga 1065 Asn Pro Ala Arg Met Gly Pro Ser Val Gly
Ser Gln Gln Glu Leu Arg 325 330
335 gcg cca tgt ctt cca gta acc tat cag cag aca cca gtg aac atg gaa
1113 Ala Pro Cys Leu Pro Val Thr Tyr Gln Gln Thr Pro Val Asn Met Glu
340 345 350 aag aac cca aga
gag gca cct cct gtt gtt cct cct ttg gca aat gct 1161 Lys Asn Pro Arg
Glu Ala Pro Pro Val Val Pro Pro Leu Ala Asn Ala 355
360 365 att tct gca gct ttg gtg tcc cca gcc acc agc cag
agc att gct cct 1209 Ile Ser Ala Ala Leu Val Ser Pro Ala Thr Ser Gln
Ser Ile Ala Pro 370 375 380 cct gtt
cct ttg aaa gcc cag aca gta aca gac tcc atg ttt gca gtg 1257 Pro Val
Pro Leu Lys Ala Gln Thr Val Thr Asp Ser Met Phe Ala Val 385
390 395 400 gcc agc aaa gat gct gga tgt
gtg aat aag agt act cat gaa ttc aag 1305 Ala Ser Lys Asp Ala Gly Cys
Val Asn Lys Ser Thr His Glu Phe Lys 405
410 415 cca cag agt gga gca gag atc aaa gaa ggg tgt gaa
aca cat aag gtt 1353 Pro Gln Ser Gly Ala Glu Ile Lys Glu Gly Cys Glu
Thr His Lys Val 420 425 430
gcc aac aca agt tct ttt cac aca act cca aac aca tca ctg gga atg 1401
Ala Asn Thr Ser Ser Phe His Thr Thr Pro Asn Thr Ser Leu Gly Met
435 440 445 gtt cag gca acg cca tcc aaa
gtg cag cca tca ccc acc gtg cac aca 1449 Val Gln Ala Thr Pro Ser Lys
Val Gln Pro Ser Pro Thr Val His Thr 450 455
460 aaa gaa gca tta ggt ttc atc atg aat atg ttt cag gct cct aca ctt
1497 Lys Glu Ala Leu Gly Phe Ile Met Asn Met Phe Gln Ala Pro Thr Leu
465 470 475 480 cct gat
att tct gat gac aaa gat gaa tgg caa tct cta gat caa aat 1545 Pro Asp
Ile Ser Asp Asp Lys Asp Glu Trp Gln Ser Leu Asp Gln Asn
485 490 495 gaa gat gca ttt gaa gcc cag
ttt caa aaa aat gta agg tca tct ggg 1593 Glu Asp Ala Phe Glu Ala Gln
Phe Gln Lys Asn Val Arg Ser Ser Gly 500 505
510 gct tgg gga gtc aat aag atc atc tct tct ttg tca tct gct
ttt cat 1641 Ala Trp Gly Val Asn Lys Ile Ile Ser Ser Leu Ser Ser Ala
Phe His 515 520 525 gtg ttt gaa
gat gga aac aaa gaa aat tat gga tta cca cag cct aaa 1689 Val Phe Glu
Asp Gly Asn Lys Glu Asn Tyr Gly Leu Pro Gln Pro Lys 530
535 540 aat aaa ccc aca gga gcc agg acc ttt gga gaa cgc
tct gtc agc aga 1737 Asn Lys Pro Thr Gly Ala Arg Thr Phe Gly Glu Arg
Ser Val Ser Arg 545 550 555
560 ctt cct tca aaa cca aag gag gaa gtg cct cat gct gaa gag ttt ttg
1785 Leu Pro Ser Lys Pro Lys Glu Glu Val Pro His Ala Glu Glu Phe Leu
565 570 575 gat gac tca act
gta tgg ggt att cgc tgc aac aaa acc ctg gca ccc 1833 Asp Asp Ser Thr
Val Trp Gly Ile Arg Cys Asn Lys Thr Leu Ala Pro 580
585 590 agt cct aag agc cca gga gac ttc aca tct gct
gca caa ctt gcg tct 1881 Ser Pro Lys Ser Pro Gly Asp Phe Thr Ser Ala
Ala Gln Leu Ala Ser 595 600 605
aca cca ttc cac aag ctt cca gtg gag tca gtg cac att tta gaa gat 1929
Thr Pro Phe His Lys Leu Pro Val Glu Ser Val His Ile Leu Glu Asp 610
615 620 aaa gaa aat gtg gta gca aaa cag tgt
acc cag gcg act ttg gat tct 1977 Lys Glu Asn Val Val Ala Lys Gln Cys
Thr Gln Ala Thr Leu Asp Ser 625 630 635
640 tgt gag gaa aac atg gtg gtg cct tca agg gat gga aaa ttc
agt cca 2025 Cys Glu Glu Asn Met Val Val Pro Ser Arg Asp Gly Lys Phe
Ser Pro 645 650 655 att
caa gag aaa agc cca aaa cag gcc ttg tcg tct cac atg tat tca 2073 Ile
Gln Glu Lys Ser Pro Lys Gln Ala Leu Ser Ser His Met Tyr Ser
660 665 670 gca tcc tta ctt cgt ctg agc
cag cct gct gca ggt ggg gta ctt acc 2121 Ala Ser Leu Leu Arg Leu Ser
Gln Pro Ala Ala Gly Gly Val Leu Thr 675 680
685 tgt gag gca gag ttg ggc gtt gag gct tgc aga ctc aca gac act
gac 2169 Cys Glu Ala Glu Leu Gly Val Glu Ala Cys Arg Leu Thr Asp Thr
Asp 690 695 700 gct gcc att gca gaa
gat cca cca gat gct att gct ggg ctc caa gca 2217 Ala Ala Ile Ala Glu
Asp Pro Pro Asp Ala Ile Ala Gly Leu Gln Ala 705 710
715 720 gaa tgg atg cag atg agt tca ctt ggg act
gtt gat gct cca aac ttc 2265 Glu Trp Met Gln Met Ser Ser Leu Gly Thr
Val Asp Ala Pro Asn Phe 725 730
735 att gtt ggg aac cca tgg gat gat aag ctg att ttc aaa ctt tta tct
2313 Ile Val Gly Asn Pro Trp Asp Asp Lys Leu Ile Phe Lys Leu Leu Ser
740 745 750 ggg ctt tct aaa
cca gtg agt tcc tat cca aat act ttt gaa tgg caa 2361 Gly Leu Ser Lys
Pro Val Ser Ser Tyr Pro Asn Thr Phe Glu Trp Gln 755
760 765 tgt aaa ctt cca gcc atc aag ccc aag act gaa ttt
caa ttg ggt tct 2409 Cys Lys Leu Pro Ala Ile Lys Pro Lys Thr Glu Phe
Gln Leu Gly Ser 770 775 780 aag ctg
gtc tat gtc cat cac ctt ctt gga gaa gga gcc ttt gcc cag 2457 Lys Leu
Val Tyr Val His His Leu Leu Gly Glu Gly Ala Phe Ala Gln 785
790 795 800 gtg tac gaa gct acc cag gga
gat ctg aat gat gct aaa aat aaa cag 2505 Val Tyr Glu Ala Thr Gln Gly
Asp Leu Asn Asp Ala Lys Asn Lys Gln 805
810 815 aaa ttt gtt tta aag gtc caa aag cct gcc aac ccc
tgg gaa ttc tac 2553 Lys Phe Val Leu Lys Val Gln Lys Pro Ala Asn Pro
Trp Glu Phe Tyr 820 825 830
att ggg acc cag ttg atg gaa aga cta aag cca tct atg cag cac atg 2601
Ile Gly Thr Gln Leu Met Glu Arg Leu Lys Pro Ser Met Gln His Met
835 840 845 ttt atg aag ttc tat tct gcc
cac tta ttc cag aat ggc agt gta tta 2649 Phe Met Lys Phe Tyr Ser Ala
His Leu Phe Gln Asn Gly Ser Val Leu 850 855
860 gta gga gag ctc tac agc tat gga aca tta tta aat gcc att aac ctc
2697 Val Gly Glu Leu Tyr Ser Tyr Gly Thr Leu Leu Asn Ala Ile Asn Leu
865 870 875 880 tat aaa
aat acc cct gaa aaa gtg atg cct caa ggt ctt gtc atc tct 2745 Tyr Lys
Asn Thr Pro Glu Lys Val Met Pro Gln Gly Leu Val Ile Ser
885 890 895 ttt gct atg aga atg ctt tac
atg att gag caa gtg cat gac tgt gaa 2793 Phe Ala Met Arg Met Leu Tyr
Met Ile Glu Gln Val His Asp Cys Glu 900 905
910 atc att cat gga gac att aaa cca gac aat ttc ata ctt gga
aac gga 2841 Ile Ile His Gly Asp Ile Lys Pro Asp Asn Phe Ile Leu Gly
Asn Gly 915 920 925 ttt ttg gaa
cag gat gat gaa gat gat tta tct gct ggc ttg gca ctg 2889 Phe Leu Glu
Gln Asp Asp Glu Asp Asp Leu Ser Ala Gly Leu Ala Leu 930
935 940 att gac ctg ggt cag agt ata gat atg aaa ctt ttt
cca aaa gga act 2937 Ile Asp Leu Gly Gln Ser Ile Asp Met Lys Leu Phe
Pro Lys Gly Thr 945 950 955
960 ata ttc aca gca aag tgt gaa aca tct ggt ttt cag tgt gtt gag atg
2985 Ile Phe Thr Ala Lys Cys Glu Thr Ser Gly Phe Gln Cys Val Glu Met
965 970 975 ctc agc aac aaa
cca tgg aac tac cag atc gat tac ttt ggg gtt gct 3033 Leu Ser Asn Lys
Pro Trp Asn Tyr Gln Ile Asp Tyr Phe Gly Val Ala 980
985 990 gca aca gta tat tgc atg ctc ttt ggc act
tac atg aaa gtg aaa aat 3081 Ala Thr Val Tyr Cys Met Leu Phe Gly Thr
Tyr Met Lys Val Lys Asn 995 1000
1005 gaa gga gga gag tgt aag cct gaa ggt ctt ttt aga agg ctt cct
3126 Glu Gly Gly Glu Cys Lys Pro Glu Gly Leu Phe Arg Arg Leu Pro
1010 1015 1020 cat ttg gat atg tgg aat
gaa ttt ttt cat gtt atg ttg aat att 3171 His Leu Asp Met Trp Asn
Glu Phe Phe His Val Met Leu Asn Ile 1025 1030
1035 cca gat tgt cat cat ctt cca tct ttg gat ttg tta agg caa
aag 3216 Pro Asp Cys His His Leu Pro Ser Leu Asp Leu Leu Arg Gln
Lys 1040 1045 1050 ctg aag aaa gta
ttt caa caa cac tat act aac aag att agg gcc 3261 Leu Lys Lys Val
Phe Gln Gln His Tyr Thr Asn Lys Ile Arg Ala 1055
1060 1065 cta cgt aat agg cta att gta ctg ctc tta gaa
tgt aag cgt tca 3306 Leu Arg Asn Arg Leu Ile Val Leu Leu Leu Glu
Cys Lys Arg Ser 1070 1075 1080 cga
aaa taa aatttggata tagacagtcc ttaaaaatca cactgtaaat 3355 Arg
Lys 1085 atgaatctgc tcactttaaa cctgtttttt tttcatttat tgtttatgta
aatgtttgtt 3415 aaaaataaat cccatggaat atttccatgt
3445 12 1085 PRT Homo sapiens 12 Met Asp Thr Pro Glu Asn Val
Leu Gln Met Leu Glu Ala His Met Gln 1 5
10 15 Ser Tyr Lys Gly Asn Asp Pro Leu Gly Glu Trp Glu
Arg Tyr Ile Gln 20 25 30
Trp Val Glu Glu Asn Phe Pro Glu Asn Lys Glu Tyr Leu Ile Thr Leu
35 40 45 Leu Glu His Leu Met Lys Glu
Phe Leu Asp Lys Lys Lys Tyr His Asn 50 55
60 Asp Pro Arg Phe Ile Ser Tyr Cys Leu Lys Phe Ala Glu Tyr Asn Ser
65 70 75 80 Asp Leu
His Gln Phe Phe Glu Phe Leu Tyr Asn His Gly Ile Gly Thr
85 90 95 Leu Ser Ser Pro Leu Tyr Ile
Ala Trp Ala Gly His Leu Glu Ala Gln 100 105
110 Gly Glu Leu Gln His Ala Ser Ala Val Leu Gln Arg Gly Ile
Gln Asn 115 120 125 Gln Ala Glu
Pro Arg Glu Phe Leu Gln Gln Gln Tyr Arg Leu Phe Gln 130
135 140 Thr Arg Leu Thr Glu Thr His Leu Pro Ala Gln Ala
Arg Thr Ser Glu 145 150 155
160 Pro Leu His Asn Val Gln Val Leu Asn Gln Met Ile Thr Ser Lys Ser
165 170 175 Asn Pro Gly Asn
Asn Met Ala Cys Ile Ser Lys Asn Gln Gly Ser Glu 180
185 190 Leu Ser Gly Val Ile Ser Ser Ala Cys Asp Lys
Glu Ser Asn Met Glu 195 200 205
Arg Arg Val Ile Thr Ile Ser Lys Ser Glu Tyr Ser Val His Ser Ser 210
215 220 Leu Ala Ser Lys Val Asp Val Glu Gln
Val Val Met Tyr Cys Lys Glu 225 230 235
240 Lys Leu Ile Arg Gly Glu Ser Glu Phe Ser Phe Glu Glu Leu
Arg Ala 245 250 255 Gln
Lys Tyr Asn Gln Arg Arg Lys His Glu Gln Trp Val Asn Glu Asp
260 265 270 Arg His Tyr Met Lys Arg Lys
Glu Ala Asn Ala Phe Glu Glu Gln Leu 275 280
285 Leu Lys Gln Lys Met Asp Glu Leu His Lys Lys Leu His Gln Val
Val 290 295 300 Glu Thr Ser His Glu
Asp Leu Pro Ala Ser Gln Glu Arg Ser Glu Val 305 310
315 320 Asn Pro Ala Arg Met Gly Pro Ser Val Gly
Ser Gln Gln Glu Leu Arg 325 330
335 Ala Pro Cys Leu Pro Val Thr Tyr Gln Gln Thr Pro Val Asn Met Glu
340 345 350 Lys Asn Pro Arg
Glu Ala Pro Pro Val Val Pro Pro Leu Ala Asn Ala 355
360 365 Ile Ser Ala Ala Leu Val Ser Pro Ala Thr Ser Gln
Ser Ile Ala Pro 370 375 380 Pro Val
Pro Leu Lys Ala Gln Thr Val Thr Asp Ser Met Phe Ala Val 385
390 395 400 Ala Ser Lys Asp Ala Gly Cys
Val Asn Lys Ser Thr His Glu Phe Lys 405
410 415 Pro Gln Ser Gly Ala Glu Ile Lys Glu Gly Cys Glu
Thr His Lys Val 420 425 430
Ala Asn Thr Ser Ser Phe His Thr Thr Pro Asn Thr Ser Leu Gly Met
435 440 445 Val Gln Ala Thr Pro Ser Lys
Val Gln Pro Ser Pro Thr Val His Thr 450 455
460 Lys Glu Ala Leu Gly Phe Ile Met Asn Met Phe Gln Ala Pro Thr Leu
465 470 475 480 Pro Asp
Ile Ser Asp Asp Lys Asp Glu Trp Gln Ser Leu Asp Gln Asn
485 490 495 Glu Asp Ala Phe Glu Ala Gln
Phe Gln Lys Asn Val Arg Ser Ser Gly 500 505
510 Ala Trp Gly Val Asn Lys Ile Ile Ser Ser Leu Ser Ser Ala
Phe His 515 520 525 Val Phe Glu
Asp Gly Asn Lys Glu Asn Tyr Gly Leu Pro Gln Pro Lys 530
535 540 Asn Lys Pro Thr Gly Ala Arg Thr Phe Gly Glu Arg
Ser Val Ser Arg 545 550 555
560 Leu Pro Ser Lys Pro Lys Glu Glu Val Pro His Ala Glu Glu Phe Leu
565 570 575 Asp Asp Ser Thr
Val Trp Gly Ile Arg Cys Asn Lys Thr Leu Ala Pro 580
585 590 Ser Pro Lys Ser Pro Gly Asp Phe Thr Ser Ala
Ala Gln Leu Ala Ser 595 600 605
Thr Pro Phe His Lys Leu Pro Val Glu Ser Val His Ile Leu Glu Asp 610
615 620 Lys Glu Asn Val Val Ala Lys Gln Cys
Thr Gln Ala Thr Leu Asp Ser 625 630 635
640 Cys Glu Glu Asn Met Val Val Pro Ser Arg Asp Gly Lys Phe
Ser Pro 645 650 655 Ile
Gln Glu Lys Ser Pro Lys Gln Ala Leu Ser Ser His Met Tyr Ser
660 665 670 Ala Ser Leu Leu Arg Leu Ser
Gln Pro Ala Ala Gly Gly Val Leu Thr 675 680
685 Cys Glu Ala Glu Leu Gly Val Glu Ala Cys Arg Leu Thr Asp Thr
Asp 690 695 700 Ala Ala Ile Ala Glu
Asp Pro Pro Asp Ala Ile Ala Gly Leu Gln Ala 705 710
715 720 Glu Trp Met Gln Met Ser Ser Leu Gly Thr
Val Asp Ala Pro Asn Phe 725 730
735 Ile Val Gly Asn Pro Trp Asp Asp Lys Leu Ile Phe Lys Leu Leu Ser
740 745 750 Gly Leu Ser Lys
Pro Val Ser Ser Tyr Pro Asn Thr Phe Glu Trp Gln 755
760 765 Cys Lys Leu Pro Ala Ile Lys Pro Lys Thr Glu Phe
Gln Leu Gly Ser 770 775 780 Lys Leu
Val Tyr Val His His Leu Leu Gly Glu Gly Ala Phe Ala Gln 785
790 795 800 Val Tyr Glu Ala Thr Gln Gly
Asp Leu Asn Asp Ala Lys Asn Lys Gln 805
810 815 Lys Phe Val Leu Lys Val Gln Lys Pro Ala Asn Pro
Trp Glu Phe Tyr 820 825 830
Ile Gly Thr Gln Leu Met Glu Arg Leu Lys Pro Ser Met Gln His Met
835 840 845 Phe Met Lys Phe Tyr Ser Ala
His Leu Phe Gln Asn Gly Ser Val Leu 850 855
860 Val Gly Glu Leu Tyr Ser Tyr Gly Thr Leu Leu Asn Ala Ile Asn Leu
865 870 875 880 Tyr Lys
Asn Thr Pro Glu Lys Val Met Pro Gln Gly Leu Val Ile Ser
885 890 895 Phe Ala Met Arg Met Leu Tyr
Met Ile Glu Gln Val His Asp Cys Glu 900 905
910 Ile Ile His Gly Asp Ile Lys Pro Asp Asn Phe Ile Leu Gly
Asn Gly 915 920 925 Phe Leu Glu
Gln Asp Asp Glu Asp Asp Leu Ser Ala Gly Leu Ala Leu 930
935 940 Ile Asp Leu Gly Gln Ser Ile Asp Met Lys Leu Phe
Pro Lys Gly Thr 945 950 955
960 Ile Phe Thr Ala Lys Cys Glu Thr Ser Gly Phe Gln Cys Val Glu Met
965 970 975 Leu Ser Asn Lys
Pro Trp Asn Tyr Gln Ile Asp Tyr Phe Gly Val Ala 980
985 990 Ala Thr Val Tyr Cys Met Leu Phe Gly Thr
Tyr Met Lys Val Lys Asn 995 1000
1005 Glu Gly Gly Glu Cys Lys Pro Glu Gly Leu Phe Arg Arg Leu Pro
1010 1015 1020 His Leu Asp Met Trp Asn
Glu Phe Phe His Val Met Leu Asn Ile 1025 1030
1035 Pro Asp Cys His His Leu Pro Ser Leu Asp Leu Leu Arg Gln
Lys 1040 1045 1050 Leu Lys Lys Val
Phe Gln Gln His Tyr Thr Asn Lys Ile Arg Ala 1055
1060 1065 Leu Arg Asn Arg Leu Ile Val Leu Leu Leu Glu
Cys Lys Arg Ser 1070 1075 1080 Arg
Lys 1085 13 2748 DNA Homo sapiens CDS (304)..(2370) C1S mRNA 13
ggacagggag gctggccgga ggttcctgca gagggagcgt caaggccctg tgctgctgtc 60
cctgggggcc agaggggttg cccagcatgc ccactggcag gagagaggga actgacccac 120
ttgctcctac cagcttctga aggtgacact gagccccagg tgacgccgca ccaccaaaga 180
aggtgcttgt gtttgtcaga caaatacagc caggcctgcc accccttagg ctccaaagtc 240
cggaggtgca gaaagccagg accaagagac aggcagctca ccagggtgga caaatcgcca 300
gag atg tgg tgc att gtc ctg ttt tca ctt ttg gca tgg gtt tat gct 348
Met Trp Cys Ile Val Leu Phe Ser Leu Leu Ala Trp Val Tyr Ala 1
5 10 15 gag cct acc atg tat
ggg gag atc ctg tcc cct aac tat cct cag gca 396 Glu Pro Thr Met Tyr
Gly Glu Ile Leu Ser Pro Asn Tyr Pro Gln Ala 20
25 30 tat ccc agt gag gta gag aaa tct tgg gac ata
gaa gtt cct gaa ggg 444 Tyr Pro Ser Glu Val Glu Lys Ser Trp Asp Ile
Glu Val Pro Glu Gly 35 40
45 tat ggg att cac ctc tac ttc acc cat ctg gac att gag ctg tca gag
492 Tyr Gly Ile His Leu Tyr Phe Thr His Leu Asp Ile Glu Leu Ser Glu
50 55 60 aac tgt gcg tat gac tca gtg
cag ata atc tca gga gac act gaa gaa 540 Asn Cys Ala Tyr Asp Ser Val
Gln Ile Ile Ser Gly Asp Thr Glu Glu 65 70
75 ggg agg ctc tgt gga cag agg agc agt aac aat ccc cac tct cca att
588 Gly Arg Leu Cys Gly Gln Arg Ser Ser Asn Asn Pro His Ser Pro Ile
80 85 90 95 gtg gaa
gag ttc caa gtc cca tac aac aaa ctc cag gtg atc ttt aag 636 Val Glu
Glu Phe Gln Val Pro Tyr Asn Lys Leu Gln Val Ile Phe Lys
100 105 110 tca gac ttt tcc aat gaa gag
cgt ttt acg ggg ttt gct gca tac tat 684 Ser Asp Phe Ser Asn Glu Glu
Arg Phe Thr Gly Phe Ala Ala Tyr Tyr 115 120
125 gtt gcc aca gac ata aat gaa tgc aca gat ttt gta gat gtc
cct tgt 732 Val Ala Thr Asp Ile Asn Glu Cys Thr Asp Phe Val Asp Val
Pro Cys 130 135 140 agc cac ttc
tgc aac aat ttc att ggt ggt tac ttc tgc tcc tgc ccc 780 Ser His Phe
Cys Asn Asn Phe Ile Gly Gly Tyr Phe Cys Ser Cys Pro 145
150 155 ccg gaa tat ttc ctc cat gat gac atg aag aat tgc
gga gtt aat tgc 828 Pro Glu Tyr Phe Leu His Asp Asp Met Lys Asn Cys
Gly Val Asn Cys 160 165 170
175 agt ggg gat gta ttc act gca ctg att ggg gag att gca agt ccc aat
876 Ser Gly Asp Val Phe Thr Ala Leu Ile Gly Glu Ile Ala Ser Pro Asn
180 185 190 tat ccc aaa cca
tat cca gag aac tca agg tgt gaa tac cag atc cgg 924 Tyr Pro Lys Pro
Tyr Pro Glu Asn Ser Arg Cys Glu Tyr Gln Ile Arg 195
200 205 ttg gag aaa ggg ttc caa gtg gtg gtg acc ttg
cgg aga gaa gat ttt 972 Leu Glu Lys Gly Phe Gln Val Val Val Thr Leu
Arg Arg Glu Asp Phe 210 215 220
gat gtg gaa gca gct gac tca gcg gga aac tgc ctt gac agt tta gtt 1020
Asp Val Glu Ala Ala Asp Ser Ala Gly Asn Cys Leu Asp Ser Leu Val 225
230 235 ttt gtt gca gga gat cgg caa ttt ggt
cct tac tgt ggt cat gga ttc 1068 Phe Val Ala Gly Asp Arg Gln Phe Gly
Pro Tyr Cys Gly His Gly Phe 240 245 250
255 cct ggg cct cta aat att gaa acc aag agt aat gct ctt gat
atc atc 1116 Pro Gly Pro Leu Asn Ile Glu Thr Lys Ser Asn Ala Leu Asp
Ile Ile 260 265 270 ttc
caa act gat cta aca ggg caa aaa aag ggc tgg aaa ctt cgc tat 1164 Phe
Gln Thr Asp Leu Thr Gly Gln Lys Lys Gly Trp Lys Leu Arg Tyr
275 280 285 cat gga gat cca atg ccc tgc
cct aag gaa gac act ccc aat tct gtt 1212 His Gly Asp Pro Met Pro Cys
Pro Lys Glu Asp Thr Pro Asn Ser Val 290 295
300 tgg gag cct gcg aag gca aaa tat gtc ttt aga gat gtg gtg cag
ata 1260 Trp Glu Pro Ala Lys Ala Lys Tyr Val Phe Arg Asp Val Val Gln
Ile 305 310 315 acc tgt ctg gat ggg
ttt gaa gtt gtg gag gga cgt gtt ggt gca aca 1308 Thr Cys Leu Asp Gly
Phe Glu Val Val Glu Gly Arg Val Gly Ala Thr 320 325
330 335 tct ttc tat tcg act tgt caa agc aat gga
aag tgg agt aat tcc aaa 1356 Ser Phe Tyr Ser Thr Cys Gln Ser Asn Gly
Lys Trp Ser Asn Ser Lys 340 345
350 ctg aaa tgt caa cct gtg gac tgt ggc att cct gaa tcc att gag aat
1404 Leu Lys Cys Gln Pro Val Asp Cys Gly Ile Pro Glu Ser Ile Glu Asn
355 360 365 ggt aaa gtt gaa
gac cca gag agc act ttg ttt ggt tct gtc atc cgc 1452 Gly Lys Val Glu
Asp Pro Glu Ser Thr Leu Phe Gly Ser Val Ile Arg 370
375 380 tac act tgt gag gag cca tat tac tac atg gaa aat
gga gga ggt ggg 1500 Tyr Thr Cys Glu Glu Pro Tyr Tyr Tyr Met Glu Asn
Gly Gly Gly Gly 385 390 395 gag tat
cac tgt gct ggt aac ggg agc tgg gtg aat gag gtg ctg ggc 1548 Glu Tyr
His Cys Ala Gly Asn Gly Ser Trp Val Asn Glu Val Leu Gly 400
405 410 415 ccg gag ctg ccg aaa tgt gtt
cca gtc tgt gga gtc ccc aga gaa ccc 1596 Pro Glu Leu Pro Lys Cys Val
Pro Val Cys Gly Val Pro Arg Glu Pro 420
425 430 ttt gaa gaa aaa cag agg ata att gga gga tcc gat
gca gat att aaa 1644 Phe Glu Glu Lys Gln Arg Ile Ile Gly Gly Ser Asp
Ala Asp Ile Lys 435 440 445
aac ttc ccc tgg caa gtc ttc ttt gac aac cca tgg gct ggt gga gcg 1692
Asn Phe Pro Trp Gln Val Phe Phe Asp Asn Pro Trp Ala Gly Gly Ala
450 455 460 ctc att aat gag tac tgg gtg
ctg acg gct gct cat gtt gtg gag gga 1740 Leu Ile Asn Glu Tyr Trp Val
Leu Thr Ala Ala His Val Val Glu Gly 465 470
475 aac agg gag cca aca atg tat gtt ggg tcc acc tca gtg cag acc tca
1788 Asn Arg Glu Pro Thr Met Tyr Val Gly Ser Thr Ser Val Gln Thr Ser
480 485 490 495 cgg ctg
gca aaa tcc aag atg ctc act cct gag cat gtg ttt att cat 1836 Arg Leu
Ala Lys Ser Lys Met Leu Thr Pro Glu His Val Phe Ile His
500 505 510 ccg gga tgg aag ctg ctg gaa
gtc cca gaa gga cga acc aat ttt gat 1884 Pro Gly Trp Lys Leu Leu Glu
Val Pro Glu Gly Arg Thr Asn Phe Asp 515 520
525 aat gac att gca ctg gtg cgg ctg aaa gac cca gtg aaa atg
gga ccc 1932 Asn Asp Ile Ala Leu Val Arg Leu Lys Asp Pro Val Lys Met
Gly Pro 530 535 540 acc gtc tct
ccc atc tgc cta cca ggc acc tct tcc gac tac aac ctc 1980 Thr Val Ser
Pro Ile Cys Leu Pro Gly Thr Ser Ser Asp Tyr Asn Leu 545
550 555 atg gat ggg gac ctg gga ctg atc tca ggc tgg ggc
cga aca gag aag 2028 Met Asp Gly Asp Leu Gly Leu Ile Ser Gly Trp Gly
Arg Thr Glu Lys 560 565 570
575 aga gat cgt gct gtt cgc ctc aag gcg gca agg tta cct gta gct cct
2076 Arg Asp Arg Ala Val Arg Leu Lys Ala Ala Arg Leu Pro Val Ala Pro
580 585 590 tta aga aaa tgc
aaa gaa gtg aaa gtg gag aaa ccc aca gca gat gca 2124 Leu Arg Lys Cys
Lys Glu Val Lys Val Glu Lys Pro Thr Ala Asp Ala 595
600 605 gag gcc tat gtt ttc act cct aac atg atc tgt
gct gga gga gag aag 2172 Glu Ala Tyr Val Phe Thr Pro Asn Met Ile Cys
Ala Gly Gly Glu Lys 610 615 620
ggc atg gat agc tgt aaa ggg gac agt ggt ggg gcc ttt gct gta cag 2220
Gly Met Asp Ser Cys Lys Gly Asp Ser Gly Gly Ala Phe Ala Val Gln 625
630 635 gat ccc aat gac aag acc aaa ttc tac
gca gct ggc ctg gtg tcc tgg 2268 Asp Pro Asn Asp Lys Thr Lys Phe Tyr
Ala Ala Gly Leu Val Ser Trp 640 645 650
655 ggg ccc cag tgt ggg acc tat ggg ctc tac aca cgg gta aag
aac tat 2316 Gly Pro Gln Cys Gly Thr Tyr Gly Leu Tyr Thr Arg Val Lys
Asn Tyr 660 665 670 gtt
gac tgg ata atg aag act atg cag gaa aat agc acc ccc cgt gag 2364 Val
Asp Trp Ile Met Lys Thr Met Gln Glu Asn Ser Thr Pro Arg Glu
675 680 685 gac taa tccagataca tcccaccagc
ctctccaagg gtggtgacca atgcattacc 2420 Asp ttctgttcct tatgatattc
tcattatttc atcatgactg aaagaagaca cgagcgaatg 2480 atttaaatag aacttgattg
ttgagacgcc ttgctagagg tagagtttga tcatagaatt 2540 gtgctggtca tacatttgtg
gtctgactcc ttggggtcct ttccccggag tacctattgt 2600 agataacact atgggtgggg
cactcctttc ttgcactatt ccacagggat accttaattc 2660 tttgtttcct ctttacctgt
tcaaaattcc atttacttga tcattctcag tatccactgt 2720 ctatgtacaa taaaggatgt
ttataagc 2748 14 688 PRT Homo
sapiens 14 Met Trp Cys Ile Val Leu Phe Ser Leu Leu Ala Trp Val Tyr Ala
Glu 1 5 10 15 Pro Thr
Met Tyr Gly Glu Ile Leu Ser Pro Asn Tyr Pro Gln Ala Tyr 20
25 30 Pro Ser Glu Val Glu Lys Ser Trp Asp
Ile Glu Val Pro Glu Gly Tyr 35 40
45 Gly Ile His Leu Tyr Phe Thr His Leu Asp Ile Glu Leu Ser Glu Asn
50 55 60 Cys Ala Tyr Asp Ser Val Gln
Ile Ile Ser Gly Asp Thr Glu Glu Gly 65 70
75 80 Arg Leu Cys Gly Gln Arg Ser Ser Asn Asn Pro His
Ser Pro Ile Val 85 90
95 Glu Glu Phe Gln Val Pro Tyr Asn Lys Leu Gln Val Ile Phe Lys Ser
100 105 110 Asp Phe Ser Asn Glu Glu
Arg Phe Thr Gly Phe Ala Ala Tyr Tyr Val 115 120
125 Ala Thr Asp Ile Asn Glu Cys Thr Asp Phe Val Asp Val Pro
Cys Ser 130 135 140 His Phe Cys Asn
Asn Phe Ile Gly Gly Tyr Phe Cys Ser Cys Pro Pro 145 150
155 160 Glu Tyr Phe Leu His Asp Asp Met Lys
Asn Cys Gly Val Asn Cys Ser 165 170
175 Gly Asp Val Phe Thr Ala Leu Ile Gly Glu Ile Ala Ser Pro Asn
Tyr 180 185 190 Pro Lys Pro
Tyr Pro Glu Asn Ser Arg Cys Glu Tyr Gln Ile Arg Leu 195
200 205 Glu Lys Gly Phe Gln Val Val Val Thr Leu Arg
Arg Glu Asp Phe Asp 210 215 220 Val
Glu Ala Ala Asp Ser Ala Gly Asn Cys Leu Asp Ser Leu Val Phe 225
230 235 240 Val Ala Gly Asp Arg Gln
Phe Gly Pro Tyr Cys Gly His Gly Phe Pro 245
250 255 Gly Pro Leu Asn Ile Glu Thr Lys Ser Asn Ala Leu
Asp Ile Ile Phe 260 265 270
Gln Thr Asp Leu Thr Gly Gln Lys Lys Gly Trp Lys Leu Arg Tyr His
275 280 285 Gly Asp Pro Met Pro Cys Pro
Lys Glu Asp Thr Pro Asn Ser Val Trp 290 295
300 Glu Pro Ala Lys Ala Lys Tyr Val Phe Arg Asp Val Val Gln Ile Thr
305 310 315 320 Cys Leu
Asp Gly Phe Glu Val Val Glu Gly Arg Val Gly Ala Thr Ser
325 330 335 Phe Tyr Ser Thr Cys Gln Ser
Asn Gly Lys Trp Ser Asn Ser Lys Leu 340 345
350 Lys Cys Gln Pro Val Asp Cys Gly Ile Pro Glu Ser Ile Glu
Asn Gly 355 360 365 Lys Val Glu
Asp Pro Glu Ser Thr Leu Phe Gly Ser Val Ile Arg Tyr 370
375 380 Thr Cys Glu Glu Pro Tyr Tyr Tyr Met Glu Asn Gly
Gly Gly Gly Glu 385 390 395
400 Tyr His Cys Ala Gly Asn Gly Ser Trp Val Asn Glu Val Leu Gly Pro
405 410 415 Glu Leu Pro Lys
Cys Val Pro Val Cys Gly Val Pro Arg Glu Pro Phe 420
425 430 Glu Glu Lys Gln Arg Ile Ile Gly Gly Ser Asp
Ala Asp Ile Lys Asn 435 440 445
Phe Pro Trp Gln Val Phe Phe Asp Asn Pro Trp Ala Gly Gly Ala Leu 450
455 460 Ile Asn Glu Tyr Trp Val Leu Thr Ala
Ala His Val Val Glu Gly Asn 465 470 475
480 Arg Glu Pro Thr Met Tyr Val Gly Ser Thr Ser Val Gln Thr
Ser Arg 485 490 495 Leu
Ala Lys Ser Lys Met Leu Thr Pro Glu His Val Phe Ile His Pro
500 505 510 Gly Trp Lys Leu Leu Glu Val
Pro Glu Gly Arg Thr Asn Phe Asp Asn 515 520
525 Asp Ile Ala Leu Val Arg Leu Lys Asp Pro Val Lys Met Gly Pro
Thr 530 535 540 Val Ser Pro Ile Cys
Leu Pro Gly Thr Ser Ser Asp Tyr Asn Leu Met 545 550
555 560 Asp Gly Asp Leu Gly Leu Ile Ser Gly Trp
Gly Arg Thr Glu Lys Arg 565 570
575 Asp Arg Ala Val Arg Leu Lys Ala Ala Arg Leu Pro Val Ala Pro Leu
580 585 590 Arg Lys Cys Lys
Glu Val Lys Val Glu Lys Pro Thr Ala Asp Ala Glu 595
600 605 Ala Tyr Val Phe Thr Pro Asn Met Ile Cys Ala Gly
Gly Glu Lys Gly 610 615 620 Met Asp
Ser Cys Lys Gly Asp Ser Gly Gly Ala Phe Ala Val Gln Asp 625
630 635 640 Pro Asn Asp Lys Thr Lys Phe
Tyr Ala Ala Gly Leu Val Ser Trp Gly 645
650 655 Pro Gln Cys Gly Thr Tyr Gly Leu Tyr Thr Arg Val
Lys Asn Tyr Val 660 665 670
Asp Trp Ile Met Lys Thr Met Gln Glu Asn Ser Thr Pro Arg Glu Asp
675 680 685 15 1543 DNA Homo sapiens CDS
(10)..(1491) CYP2E1 mRNA 15 agcggcacc atg tct gcc ctc gga gtc acc gtg
gcc ctg ctg gtg tgg gcg 51 Met Ser Ala Leu Gly Val Thr Val
Ala Leu Leu Val Trp Ala 1 5 10
gcc ttc ctc ctg ctg gtg tcc atg tgg agg cag gtg cac agc agc tgg 99
Ala Phe Leu Leu Leu Val Ser Met Trp Arg Gln Val His Ser Ser Trp 15
20 25 30 aat ctg ccc cca ggc
cct ttc ccg ctt ccc atc atc ggg aac ctc ttc 147 Asn Leu Pro Pro Gly
Pro Phe Pro Leu Pro Ile Ile Gly Asn Leu Phe 35
40 45 cag ttg gaa ttg aag aat att ccc aag tcc ttc
acc cgg ttg gcc cag 195 Gln Leu Glu Leu Lys Asn Ile Pro Lys Ser Phe
Thr Arg Leu Ala Gln 50 55
60 cgc ttc ggg ccg gtg ttc acg ctg tac gtg ggc tcg cag cgc atg gtg
243 Arg Phe Gly Pro Val Phe Thr Leu Tyr Val Gly Ser Gln Arg Met Val
65 70 75 gtg atg cac ggc tac aag gcg
gtg aag gaa gcg ctg ctg gac tac aag 291 Val Met His Gly Tyr Lys Ala
Val Lys Glu Ala Leu Leu Asp Tyr Lys 80 85
90 gac gag ttc tcg ggc aga ggc gac ctc ccc gcg ttc cat gcg cac agg
339 Asp Glu Phe Ser Gly Arg Gly Asp Leu Pro Ala Phe His Ala His Arg
95 100 105 110 gac agg
gga atc att ttt aat aat gga cct acc tgg aag gac atc cgg 387 Asp Arg
Gly Ile Ile Phe Asn Asn Gly Pro Thr Trp Lys Asp Ile Arg
115 120 125 cgg ttt tcc ctg acc acc ctc
cgg aac tat ggg atg ggg aaa cag ggc 435 Arg Phe Ser Leu Thr Thr Leu
Arg Asn Tyr Gly Met Gly Lys Gln Gly 130 135
140 aat gag agc cgg atc cag agg gag gcc cac ttc ctg ctg gaa
gca ctc 483 Asn Glu Ser Arg Ile Gln Arg Glu Ala His Phe Leu Leu Glu
Ala Leu 145 150 155 agg aag acc
caa ggc cag cct ttc gac ccc acc ttc ctc atc ggc tgc 531 Arg Lys Thr
Gln Gly Gln Pro Phe Asp Pro Thr Phe Leu Ile Gly Cys 160
165 170 gcg ccc tgc aac gtc ata gcc gac atc ctc ttc cgc
aag cat ttt gac 579 Ala Pro Cys Asn Val Ile Ala Asp Ile Leu Phe Arg
Lys His Phe Asp 175 180 185
190 tac aat gat gag aag ttt cta agg ctg atg tat ttg ttt aat gag aac
627 Tyr Asn Asp Glu Lys Phe Leu Arg Leu Met Tyr Leu Phe Asn Glu Asn
195 200 205 ttc cac cta ctc
agc act ccc tgg ctc cag ctt tac aat aat ttt ccc 675 Phe His Leu Leu
Ser Thr Pro Trp Leu Gln Leu Tyr Asn Asn Phe Pro 210
215 220 agc ttt cta cac tac ttg cct gga agc cac aga
aaa gtc ata aaa aat 723 Ser Phe Leu His Tyr Leu Pro Gly Ser His Arg
Lys Val Ile Lys Asn 225 230 235
gtg gct gaa gta aaa gag tat gtg tct gaa agg gtg aag gag cac cat 771
Val Ala Glu Val Lys Glu Tyr Val Ser Glu Arg Val Lys Glu His His 240
245 250 caa tct ctg gac ccc aac tgt ccc cgg
gac ctc acc gac tgc ctg ctc 819 Gln Ser Leu Asp Pro Asn Cys Pro Arg
Asp Leu Thr Asp Cys Leu Leu 255 260 265
270 gtg gaa atg gag aag gaa aag cac agt gca gag cgc ttg tac
aca atg 867 Val Glu Met Glu Lys Glu Lys His Ser Ala Glu Arg Leu Tyr
Thr Met 275 280 285 gac
ggt atc acc gtg act gtg gcc gac ctg ttc ttt gcg ggg aca gag 915 Asp
Gly Ile Thr Val Thr Val Ala Asp Leu Phe Phe Ala Gly Thr Glu
290 295 300 acc acc agc aca act ctg aga
tat ggg ctc ctg att ctc atg aaa tac 963 Thr Thr Ser Thr Thr Leu Arg
Tyr Gly Leu Leu Ile Leu Met Lys Tyr 305 310
315 cct gag att gaa gag aag ctc cat gaa gaa att gac agg gtg att
ggg 1011 Pro Glu Ile Glu Glu Lys Leu His Glu Glu Ile Asp Arg Val Ile
Gly 320 325 330 cca agc cga atc cct
gcc atc aag gat agg caa gag atg ccc tac atg 1059 Pro Ser Arg Ile Pro
Ala Ile Lys Asp Arg Gln Glu Met Pro Tyr Met 335 340
345 350 gat gct gtg gtg cat gag att cag cgg ttc
atc acc ctc gtg ccc tcc 1107 Asp Ala Val Val His Glu Ile Gln Arg Phe
Ile Thr Leu Val Pro Ser 355 360
365 aac ctg ccc cat gaa gca acc cga gac acc att ttc aga gga tac ctc
1155 Asn Leu Pro His Glu Ala Thr Arg Asp Thr Ile Phe Arg Gly Tyr Leu
370 375 380 atc ccc aag ggc
aca gtc gta gtg cca act ctg gac tct gtt ttg tat 1203 Ile Pro Lys Gly
Thr Val Val Val Pro Thr Leu Asp Ser Val Leu Tyr 385
390 395 gac aac caa gaa ttt cct gat cca gaa aag ttt aag
cca gaa cac ttc 1251 Asp Asn Gln Glu Phe Pro Asp Pro Glu Lys Phe Lys
Pro Glu His Phe 400 405 410 ctg aat
gaa aat gga aag ttt aag tac agt gac tat ttc aag cca ttt 1299 Leu Asn
Glu Asn Gly Lys Phe Lys Tyr Ser Asp Tyr Phe Lys Pro Phe 415
420 425 430 tcc aca gga aaa cga gtg tgt
gct gga gaa ggc ctg gct cgc atg gag 1347 Ser Thr Gly Lys Arg Val Cys
Ala Gly Glu Gly Leu Ala Arg Met Glu 435
440 445 ttg ttt ctt ttg ttg tgt gcc att ttg cag cat ttt
aat ttg aag cct 1395 Leu Phe Leu Leu Leu Cys Ala Ile Leu Gln His Phe
Asn Leu Lys Pro 450 455 460
ctc gtt gac cca aag gat atc gac ctc agc cct ata cat att ggg ttt 1443
Leu Val Asp Pro Lys Asp Ile Asp Leu Ser Pro Ile His Ile Gly Phe
465 470 475 ggc tgt atc cca cca cgt tac
aaa ctc tgt gtc att ccc cgc tca tga 1491 Gly Cys Ile Pro Pro Arg Tyr
Lys Leu Cys Val Ile Pro Arg Ser 480 485
490 gtgtgtggag gacaccctga accccccgct ttcaaacaag ttttcaaatt gt
1543 16 493 PRT Homo sapiens 16 Met Ser Ala Leu Gly Val Thr Val Ala Leu
Leu Val Trp Ala Ala Phe 1 5 10
15 Leu Leu Leu Val Ser Met Trp Arg Gln Val His Ser Ser Trp Asn Leu
20 25 30 Pro Pro Gly Pro Phe
Pro Leu Pro Ile Ile Gly Asn Leu Phe Gln Leu 35
40 45 Glu Leu Lys Asn Ile Pro Lys Ser Phe Thr Arg Leu
Ala Gln Arg Phe 50 55 60 Gly Pro
Val Phe Thr Leu Tyr Val Gly Ser Gln Arg Met Val Val Met 65
70 75 80 His Gly Tyr Lys Ala Val Lys
Glu Ala Leu Leu Asp Tyr Lys Asp Glu 85
90 95 Phe Ser Gly Arg Gly Asp Leu Pro Ala Phe His Ala
His Arg Asp Arg 100 105 110
Gly Ile Ile Phe Asn Asn Gly Pro Thr Trp Lys Asp Ile Arg Arg Phe
115 120 125 Ser Leu Thr Thr Leu Arg Asn
Tyr Gly Met Gly Lys Gln Gly Asn Glu 130 135
140 Ser Arg Ile Gln Arg Glu Ala His Phe Leu Leu Glu Ala Leu Arg Lys
145 150 155 160 Thr Gln
Gly Gln Pro Phe Asp Pro Thr Phe Leu Ile Gly Cys Ala Pro
165 170 175 Cys Asn Val Ile Ala Asp Ile
Leu Phe Arg Lys His Phe Asp Tyr Asn 180 185
190 Asp Glu Lys Phe Leu Arg Leu Met Tyr Leu Phe Asn Glu Asn
Phe His 195 200 205 Leu Leu Ser
Thr Pro Trp Leu Gln Leu Tyr Asn Asn Phe Pro Ser Phe 210
215 220 Leu His Tyr Leu Pro Gly Ser His Arg Lys Val Ile
Lys Asn Val Ala 225 230 235
240 Glu Val Lys Glu Tyr Val Ser Glu Arg Val Lys Glu His His Gln Ser
245 250 255 Leu Asp Pro Asn
Cys Pro Arg Asp Leu Thr Asp Cys Leu Leu Val Glu 260
265 270 Met Glu Lys Glu Lys His Ser Ala Glu Arg Leu
Tyr Thr Met Asp Gly 275 280 285
Ile Thr Val Thr Val Ala Asp Leu Phe Phe Ala Gly Thr Glu Thr Thr 290
295 300 Ser Thr Thr Leu Arg Tyr Gly Leu Leu
Ile Leu Met Lys Tyr Pro Glu 305 310 315
320 Ile Glu Glu Lys Leu His Glu Glu Ile Asp Arg Val Ile Gly
Pro Ser 325 330 335 Arg
Ile Pro Ala Ile Lys Asp Arg Gln Glu Met Pro Tyr Met Asp Ala
340 345 350 Val Val His Glu Ile Gln Arg
Phe Ile Thr Leu Val Pro Ser Asn Leu 355 360
365 Pro His Glu Ala Thr Arg Asp Thr Ile Phe Arg Gly Tyr Leu Ile
Pro 370 375 380 Lys Gly Thr Val Val
Val Pro Thr Leu Asp Ser Val Leu Tyr Asp Asn 385 390
395 400 Gln Glu Phe Pro Asp Pro Glu Lys Phe Lys
Pro Glu His Phe Leu Asn 405 410
415 Glu Asn Gly Lys Phe Lys Tyr Ser Asp Tyr Phe Lys Pro Phe Ser Thr
420 425 430 Gly Lys Arg Val
Cys Ala Gly Glu Gly Leu Ala Arg Met Glu Leu Phe 435
440 445 Leu Leu Leu Cys Ala Ile Leu Gln His Phe Asn Leu
Lys Pro Leu Val 450 455 460 Asp Pro
Lys Asp Ile Asp Leu Ser Pro Ile His Ile Gly Phe Gly Cys 465
470 475 480 Ile Pro Pro Arg Tyr Lys Leu
Cys Val Ile Pro Arg Ser 485 490 17 2871
DNA Homo sapiens CDS (110)..(2650) DLG7 mRNA 17 ggaggctcgg gttgtgaggg
ttcctgcttc ggagtcggcg gtggtcgtcc agaccgagtg 60 ttctttactt tttgtttggt
tgaggtttca cgctagaagg tggctcagg atg tct tca 118
Met Ser Ser
1 tca cat ttt gcc agt cga cac agg aag gat ata agt
act gaa atg att 166 Ser His Phe Ala Ser Arg His Arg Lys Asp Ile Ser
Thr Glu Met Ile 5 10 15 aga act
aaa att gct cat agg aaa tca ctg tct cag aaa gaa aat aga 214 Arg Thr
Lys Ile Ala His Arg Lys Ser Leu Ser Gln Lys Glu Asn Arg 20
25 30 35 cat aag gaa tac gaa cga aat
aga cac ttt ggt ttg aaa gat gta aac 262 His Lys Glu Tyr Glu Arg Asn
Arg His Phe Gly Leu Lys Asp Val Asn 40
45 50 att cca acc ttg gaa ggt aga att ctt gtt gaa tta
gat gag aca tct 310 Ile Pro Thr Leu Glu Gly Arg Ile Leu Val Glu Leu
Asp Glu Thr Ser 55 60 65
caa ggg ctt gtt cca gaa aag acc aat gtt aag cca agg gca atg aaa 358
Gln Gly Leu Val Pro Glu Lys Thr Asn Val Lys Pro Arg Ala Met Lys
70 75 80 act att cta ggt gat caa cga
aaa cag atg ctc caa aaa tac aaa gaa 406 Thr Ile Leu Gly Asp Gln Arg
Lys Gln Met Leu Gln Lys Tyr Lys Glu 85 90
95 gaa aag caa ctt caa aaa ttg aaa gag cag aga gag aaa gct aaa cga
454 Glu Lys Gln Leu Gln Lys Leu Lys Glu Gln Arg Glu Lys Ala Lys Arg
100 105 110 115 gga ata
ttt aaa gtg ggt cgt tat aga cct gat atg cct tgt ttt ctt 502 Gly Ile
Phe Lys Val Gly Arg Tyr Arg Pro Asp Met Pro Cys Phe Leu
120 125 130 tta tca aac cag aat gct gtg
aaa gct gag cca aaa aag gct att cca 550 Leu Ser Asn Gln Asn Ala Val
Lys Ala Glu Pro Lys Lys Ala Ile Pro 135 140
145 tct tct gta cgg att aca agg tca aag gcc aaa gac caa atg
gag cag 598 Ser Ser Val Arg Ile Thr Arg Ser Lys Ala Lys Asp Gln Met
Glu Gln 150 155 160 act aag att
gat aac gag agt gat gtt cga gca atc cga cct ggt cca 646 Thr Lys Ile
Asp Asn Glu Ser Asp Val Arg Ala Ile Arg Pro Gly Pro 165
170 175 aga caa act tct gaa aag aaa gtg tca gac aaa gag
aaa aaa gtt gtg 694 Arg Gln Thr Ser Glu Lys Lys Val Ser Asp Lys Glu
Lys Lys Val Val 180 185 190
195 cag cct gta atg ccc acg tcg ttg aga atg act cga tca gct act caa
742 Gln Pro Val Met Pro Thr Ser Leu Arg Met Thr Arg Ser Ala Thr Gln
200 205 210 gca gca aag cag
gtt ccc aga aca gtc tca tct acc aca gca aga aag 790 Ala Ala Lys Gln
Val Pro Arg Thr Val Ser Ser Thr Thr Ala Arg Lys 215
220 225 cca gtc aca aga gct gct aat gaa aac gaa cca
gaa gga aag gtg cca 838 Pro Val Thr Arg Ala Ala Asn Glu Asn Glu Pro
Glu Gly Lys Val Pro 230 235 240
agt aaa gga aga cct gcc aaa aat gta gaa aca aaa ccc gac aag ggt 886
Ser Lys Gly Arg Pro Ala Lys Asn Val Glu Thr Lys Pro Asp Lys Gly 245
250 255 att tct tgt aaa gtc gat agt gaa gaa
aat act ttg aat tca caa act 934 Ile Ser Cys Lys Val Asp Ser Glu Glu
Asn Thr Leu Asn Ser Gln Thr 260 265 270
275 aat gca aca agt gga atg aat cca gat gga gtc tta tca aaa
atg gaa 982 Asn Ala Thr Ser Gly Met Asn Pro Asp Gly Val Leu Ser Lys
Met Glu 280 285 290 aac
tta cct gag ata aat act gca aaa ata aaa ggg aag aat tcc ttt 1030 Asn
Leu Pro Glu Ile Asn Thr Ala Lys Ile Lys Gly Lys Asn Ser Phe
295 300 305 gca cct aag gat ttt atg ttt
cag cca ctg gat ggt ctg aag acc tat 1078 Ala Pro Lys Asp Phe Met Phe
Gln Pro Leu Asp Gly Leu Lys Thr Tyr 310 315
320 caa gta aca cct atg act ccc aga agt gcc aat gct ttt ttg aca
ccc 1126 Gln Val Thr Pro Met Thr Pro Arg Ser Ala Asn Ala Phe Leu Thr
Pro 325 330 335 agt tac acc tgg act
cct tta aaa aca gaa gtt gat gag tct caa gca 1174 Ser Tyr Thr Trp Thr
Pro Leu Lys Thr Glu Val Asp Glu Ser Gln Ala 340 345
350 355 aca aaa gaa att ttg gca caa aaa tgt aaa
act tac tct acc aag aca 1222 Thr Lys Glu Ile Leu Ala Gln Lys Cys Lys
Thr Tyr Ser Thr Lys Thr 360 365
370 ata cag caa gat tca aat aaa ttg cca tgt cct ttg ggt cct cta act
1270 Ile Gln Gln Asp Ser Asn Lys Leu Pro Cys Pro Leu Gly Pro Leu Thr
375 380 385 gtt tgg cat gaa
gaa cat gtt tta aat aaa aat gaa gct act act aaa 1318 Val Trp His Glu
Glu His Val Leu Asn Lys Asn Glu Ala Thr Thr Lys 390
395 400 aat tta aat ggc ctt cca ata aaa gaa gtc cca tca
ctt gaa aga aat 1366 Asn Leu Asn Gly Leu Pro Ile Lys Glu Val Pro Ser
Leu Glu Arg Asn 405 410 415 gaa ggt
cga att gct cag ccc cac cat ggt gtg cca tat ttc aga aat 1414 Glu Gly
Arg Ile Ala Gln Pro His His Gly Val Pro Tyr Phe Arg Asn 420
425 430 435 atc ctc cag tca gaa act gag
aaa tta act tca cat tgc ttc gag tgg 1462 Ile Leu Gln Ser Glu Thr Glu
Lys Leu Thr Ser His Cys Phe Glu Trp 440
445 450 gac agg aaa ctt gaa ttg gac att cca gat gat gct
aaa gat ctt att 1510 Asp Arg Lys Leu Glu Leu Asp Ile Pro Asp Asp Ala
Lys Asp Leu Ile 455 460 465
cgc aca gca gtt ggt caa aca aga ctc ctt atg aag gaa agg ttt aaa 1558
Arg Thr Ala Val Gly Gln Thr Arg Leu Leu Met Lys Glu Arg Phe Lys
470 475 480 cag ttt gaa gga ctg gtt gat
gat tgt gaa tat aaa cga ggt ata aag 1606 Gln Phe Glu Gly Leu Val Asp
Asp Cys Glu Tyr Lys Arg Gly Ile Lys 485 490
495 gag act acc tgt aca gat ctg gat gga ttt tgg gat atg gtt agt ttt
1654 Glu Thr Thr Cys Thr Asp Leu Asp Gly Phe Trp Asp Met Val Ser Phe
500 505 510 515 cag ata
gaa gat gta atc cac aaa ttc aac aat ctg atc aaa ctt gag 1702 Gln Ile
Glu Asp Val Ile His Lys Phe Asn Asn Leu Ile Lys Leu Glu
520 525 530 gaa tct ggg tgg caa gtc aat
aat aat atg aat cat aat atg aac aaa 1750 Glu Ser Gly Trp Gln Val Asn
Asn Asn Met Asn His Asn Met Asn Lys 535 540
545 aat gtc ttt agg aaa aaa gtt gtc tca ggt ata gca agt aaa
cca aaa 1798 Asn Val Phe Arg Lys Lys Val Val Ser Gly Ile Ala Ser Lys
Pro Lys 550 555 560 cag gat gat
gct gga aga att gca gcg aga aat cgc cta gct gcc ata 1846 Gln Asp Asp
Ala Gly Arg Ile Ala Ala Arg Asn Arg Leu Ala Ala Ile 565
570 575 aaa aat gca atg aga gag aga att agg cag gaa gaa
tgt gct gaa aca 1894 Lys Asn Ala Met Arg Glu Arg Ile Arg Gln Glu Glu
Cys Ala Glu Thr 580 585 590
595 gca gtt tct gtg ata cca aag gaa gtt gat aaa ata gtg ttc gat gct
1942 Ala Val Ser Val Ile Pro Lys Glu Val Asp Lys Ile Val Phe Asp Ala
600 605 610 gga ttt ttc aga
gtt gaa agt cct gtt aaa tta ttc tca gga ctt tct 1990 Gly Phe Phe Arg
Val Glu Ser Pro Val Lys Leu Phe Ser Gly Leu Ser 615
620 625 gtc tct tct gaa ggc cct tct caa aga ctt gga
aca cct aag tct gtc 2038 Val Ser Ser Glu Gly Pro Ser Gln Arg Leu Gly
Thr Pro Lys Ser Val 630 635 640
aac aaa gct gta tct cag agt aga aat gag atg ggc att cca caa caa 2086
Asn Lys Ala Val Ser Gln Ser Arg Asn Glu Met Gly Ile Pro Gln Gln 645
650 655 act aca tca cca gaa aat gcc ggt cct
cag aat acg aaa agt gaa cat 2134 Thr Thr Ser Pro Glu Asn Ala Gly Pro
Gln Asn Thr Lys Ser Glu His 660 665 670
675 gtg aag aag act ttg ttt ttg agt att cct gaa agc agg agc
agc ata 2182 Val Lys Lys Thr Leu Phe Leu Ser Ile Pro Glu Ser Arg Ser
Ser Ile 680 685 690 gaa
gat gct cag tgt cct gga tta cca gat tta att gaa gaa aat cat 2230 Glu
Asp Ala Gln Cys Pro Gly Leu Pro Asp Leu Ile Glu Glu Asn His
695 700 705 gtt gta aat aag aca gac ttg
aag gtg gat tgt tta tcc agt gag aga 2278 Val Val Asn Lys Thr Asp Leu
Lys Val Asp Cys Leu Ser Ser Glu Arg 710 715
720 atg agt ttg cct ctt ctt gct ggt gga gta gca gat gat att aat
act 2326 Met Ser Leu Pro Leu Leu Ala Gly Gly Val Ala Asp Asp Ile Asn
Thr 725 730 735 aac aaa aaa gaa gga
att tca gat gtt gtg gaa gga atg gaa ctg aat 2374 Asn Lys Lys Glu Gly
Ile Ser Asp Val Val Glu Gly Met Glu Leu Asn 740 745
750 755 tct tca att aca tca cag gat gtt ttg atg
agt agc cct gaa aaa aat 2422 Ser Ser Ile Thr Ser Gln Asp Val Leu Met
Ser Ser Pro Glu Lys Asn 760 765
770 aca gct tca caa aat agc atc tta gaa gaa ggg gaa act aaa att tct
2470 Thr Ala Ser Gln Asn Ser Ile Leu Glu Glu Gly Glu Thr Lys Ile Ser
775 780 785 cag tca gaa cta
ttt gat aat aaa agt ctc act act gaa tgc cac ctt 2518 Gln Ser Glu Leu
Phe Asp Asn Lys Ser Leu Thr Thr Glu Cys His Leu 790
795 800 ctt gat tca cca ggt cta aac tgc agt aat cca ttt
act cag ctg gag 2566 Leu Asp Ser Pro Gly Leu Asn Cys Ser Asn Pro Phe
Thr Gln Leu Glu 805 810 815 agg aga
cat caa gaa cat gcc aga cac att tct ttt ggt ggt aac ctg 2614 Arg Arg
His Gln Glu His Ala Arg His Ile Ser Phe Gly Gly Asn Leu 820
825 830 835 att act ttt tca cct cta caa
cca gga gaa ttt tga atttaaaaat 2660 Ile Thr Phe Ser Pro Leu Gln
Pro Gly Glu Phe 840 845 aaatccaaac
attttccttc atattatcaa tgcttatata ttccttagac tattgaaatt 2720 ttggagaaaa
tgtatttgtg ttcacttcta tagcatataa tgttttaata ttctgtgttc 2780 atcaaagtgt
attttagata tactctttct caagggaagt ggggatattt tgtacatttt 2840 caacacagaa
taaaaaatgt actgtgcctt g 2871 18 846 PRT
Homo sapiens 18 Met Ser Ser Ser His Phe Ala Ser Arg His Arg Lys Asp Ile
Ser Thr 1 5 10 15 Glu
Met Ile Arg Thr Lys Ile Ala His Arg Lys Ser Leu Ser Gln Lys
20 25 30 Glu Asn Arg His Lys Glu Tyr
Glu Arg Asn Arg His Phe Gly Leu Lys 35 40
45 Asp Val Asn Ile Pro Thr Leu Glu Gly Arg Ile Leu Val Glu Leu
Asp 50 55 60 Glu Thr Ser Gln Gly
Leu Val Pro Glu Lys Thr Asn Val Lys Pro Arg 65 70
75 80 Ala Met Lys Thr Ile Leu Gly Asp Gln Arg
Lys Gln Met Leu Gln Lys 85 90
95 Tyr Lys Glu Glu Lys Gln Leu Gln Lys Leu Lys Glu Gln Arg Glu Lys
100 105 110 Ala Lys Arg Gly
Ile Phe Lys Val Gly Arg Tyr Arg Pro Asp Met Pro 115
120 125 Cys Phe Leu Leu Ser Asn Gln Asn Ala Val Lys Ala
Glu Pro Lys Lys 130 135 140 Ala Ile
Pro Ser Ser Val Arg Ile Thr Arg Ser Lys Ala Lys Asp Gln 145
150 155 160 Met Glu Gln Thr Lys Ile Asp
Asn Glu Ser Asp Val Arg Ala Ile Arg 165
170 175 Pro Gly Pro Arg Gln Thr Ser Glu Lys Lys Val Ser
Asp Lys Glu Lys 180 185 190
Lys Val Val Gln Pro Val Met Pro Thr Ser Leu Arg Met Thr Arg Ser
195 200 205 Ala Thr Gln Ala Ala Lys Gln
Val Pro Arg Thr Val Ser Ser Thr Thr 210 215
220 Ala Arg Lys Pro Val Thr Arg Ala Ala Asn Glu Asn Glu Pro Glu Gly
225 230 235 240 Lys Val
Pro Ser Lys Gly Arg Pro Ala Lys Asn Val Glu Thr Lys Pro
245 250 255 Asp Lys Gly Ile Ser Cys Lys
Val Asp Ser Glu Glu Asn Thr Leu Asn 260 265
270 Ser Gln Thr Asn Ala Thr Ser Gly Met Asn Pro Asp Gly Val
Leu Ser 275 280 285 Lys Met Glu
Asn Leu Pro Glu Ile Asn Thr Ala Lys Ile Lys Gly Lys 290
295 300 Asn Ser Phe Ala Pro Lys Asp Phe Met Phe Gln Pro
Leu Asp Gly Leu 305 310 315
320 Lys Thr Tyr Gln Val Thr Pro Met Thr Pro Arg Ser Ala Asn Ala Phe
325 330 335 Leu Thr Pro Ser
Tyr Thr Trp Thr Pro Leu Lys Thr Glu Val Asp Glu 340
345 350 Ser Gln Ala Thr Lys Glu Ile Leu Ala Gln Lys
Cys Lys Thr Tyr Ser 355 360 365
Thr Lys Thr Ile Gln Gln Asp Ser Asn Lys Leu Pro Cys Pro Leu Gly 370
375 380 Pro Leu Thr Val Trp His Glu Glu His
Val Leu Asn Lys Asn Glu Ala 385 390 395
400 Thr Thr Lys Asn Leu Asn Gly Leu Pro Ile Lys Glu Val Pro
Ser Leu 405 410 415 Glu
Arg Asn Glu Gly Arg Ile Ala Gln Pro His His Gly Val Pro Tyr
420 425 430 Phe Arg Asn Ile Leu Gln Ser
Glu Thr Glu Lys Leu Thr Ser His Cys 435 440
445 Phe Glu Trp Asp Arg Lys Leu Glu Leu Asp Ile Pro Asp Asp Ala
Lys 450 455 460 Asp Leu Ile Arg Thr
Ala Val Gly Gln Thr Arg Leu Leu Met Lys Glu 465 470
475 480 Arg Phe Lys Gln Phe Glu Gly Leu Val Asp
Asp Cys Glu Tyr Lys Arg 485 490
495 Gly Ile Lys Glu Thr Thr Cys Thr Asp Leu Asp Gly Phe Trp Asp Met
500 505 510 Val Ser Phe Gln
Ile Glu Asp Val Ile His Lys Phe Asn Asn Leu Ile 515
520 525 Lys Leu Glu Glu Ser Gly Trp Gln Val Asn Asn Asn
Met Asn His Asn 530 535 540 Met Asn
Lys Asn Val Phe Arg Lys Lys Val Val Ser Gly Ile Ala Ser 545
550 555 560 Lys Pro Lys Gln Asp Asp Ala
Gly Arg Ile Ala Ala Arg Asn Arg Leu 565
570 575 Ala Ala Ile Lys Asn Ala Met Arg Glu Arg Ile Arg
Gln Glu Glu Cys 580 585 590
Ala Glu Thr Ala Val Ser Val Ile Pro Lys Glu Val Asp Lys Ile Val
595 600 605 Phe Asp Ala Gly Phe Phe Arg
Val Glu Ser Pro Val Lys Leu Phe Ser 610 615
620 Gly Leu Ser Val Ser Ser Glu Gly Pro Ser Gln Arg Leu Gly Thr Pro
625 630 635 640 Lys Ser
Val Asn Lys Ala Val Ser Gln Ser Arg Asn Glu Met Gly Ile
645 650 655 Pro Gln Gln Thr Thr Ser Pro
Glu Asn Ala Gly Pro Gln Asn Thr Lys 660 665
670 Ser Glu His Val Lys Lys Thr Leu Phe Leu Ser Ile Pro Glu
Ser Arg 675 680 685 Ser Ser Ile
Glu Asp Ala Gln Cys Pro Gly Leu Pro Asp Leu Ile Glu 690
695 700 Glu Asn His Val Val Asn Lys Thr Asp Leu Lys Val
Asp Cys Leu Ser 705 710 715
720 Ser Glu Arg Met Ser Leu Pro Leu Leu Ala Gly Gly Val Ala Asp Asp
725 730 735 Ile Asn Thr Asn
Lys Lys Glu Gly Ile Ser Asp Val Val Glu Gly Met 740
745 750 Glu Leu Asn Ser Ser Ile Thr Ser Gln Asp Val
Leu Met Ser Ser Pro 755 760 765
Glu Lys Asn Thr Ala Ser Gln Asn Ser Ile Leu Glu Glu Gly Glu Thr 770
775 780 Lys Ile Ser Gln Ser Glu Leu Phe Asp
Asn Lys Ser Leu Thr Thr Glu 785 790 795
800 Cys His Leu Leu Asp Ser Pro Gly Leu Asn Cys Ser Asn Pro
Phe Thr 805 810 815 Gln
Leu Glu Arg Arg His Gln Glu His Ala Arg His Ile Ser Phe Gly
820 825 830 Gly Asn Leu Ile Thr Phe Ser
Pro Leu Gln Pro Gly Glu Phe 835 840
845 19 2284 DNA Homo sapiens CDS (114)..(1268) DUSP9 mRNA 19
cgcttcccgc cgcccgagct tcggaaactt cccggccgcg acgcagggaa ccggcgcgga 60
gaaccgagca gagcggagcg cccgtggtcc agcgtgtagg gagccgatcg ccc atg 116
Met
1 gag ggt ctg ggc cgc tcg
tgc ctg tgg ctg cgt cgg gag ctg tcg ccc 164 Glu Gly Leu Gly Arg Ser
Cys Leu Trp Leu Arg Arg Glu Leu Ser Pro 5
10 15 ccg cgg ccg cgg ctc ctg ctc ctg gac tgc cgc agc
cgc gag ctg tac 212 Pro Arg Pro Arg Leu Leu Leu Leu Asp Cys Arg Ser
Arg Glu Leu Tyr 20 25 30 gag
tcg gcg cgc atc ggt ggg gcg ctg agc gtg gcc ctg ccg gcg ctc 260 Glu
Ser Ala Arg Ile Gly Gly Ala Leu Ser Val Ala Leu Pro Ala Leu 35
40 45 ctg ctg cgc cgc ctg cgg agg ggc agc ctg
tcg gtg cgc gcg ctc ctg 308 Leu Leu Arg Arg Leu Arg Arg Gly Ser Leu
Ser Val Arg Ala Leu Leu 50 55 60
65 cct ggg ccg ccg ctg cag ccg ccc ccg cct gcc ccc gtg ctc ctg
tac 356 Pro Gly Pro Pro Leu Gln Pro Pro Pro Pro Ala Pro Val Leu Leu
Tyr 70 75 80 gac cag
ggc ggg ggc cgg cgc cgg cgc ggg gag gcc gag gcc gag gcc 404 Asp Gln
Gly Gly Gly Arg Arg Arg Arg Gly Glu Ala Glu Ala Glu Ala 85
90 95 gag gag tgg gag gcc gag tcg gtg ctg
ggc acc ctg ctg cag aag ctg 452 Glu Glu Trp Glu Ala Glu Ser Val Leu
Gly Thr Leu Leu Gln Lys Leu 100 105
110 cga gag gaa ggc tac ctg gcc tac tac ctc cag gga ggc ttc agc aga
500 Arg Glu Glu Gly Tyr Leu Ala Tyr Tyr Leu Gln Gly Gly Phe Ser Arg
115 120 125 ttc cag gcc gag tgc cct cac
ctg tgt gag acc agc ctt gct ggc cgt 548 Phe Gln Ala Glu Cys Pro His
Leu Cys Glu Thr Ser Leu Ala Gly Arg 130 135
140 145 gcc ggc tcc agc atg gcg ccg gtg ccc ggt cca gtg
ccc gtg gtg ggg 596 Ala Gly Ser Ser Met Ala Pro Val Pro Gly Pro Val
Pro Val Val Gly 150 155
160 ttg ggc agc ctg tgc ctg ggc tcc gac tgc tct gat gcg gaa tcc gag
644 Leu Gly Ser Leu Cys Leu Gly Ser Asp Cys Ser Asp Ala Glu Ser Glu
165 170 175 gct gac cgc gac tcc atg
agc tgt ggc ctg gat tcg gag ggt gcc aca 692 Ala Asp Arg Asp Ser Met
Ser Cys Gly Leu Asp Ser Glu Gly Ala Thr 180 185
190 ccc cca cca gtg ggg ctg cgg gca tcc ttc cct gtc cag atc
ctg ccc 740 Pro Pro Pro Val Gly Leu Arg Ala Ser Phe Pro Val Gln Ile
Leu Pro 195 200 205 aac ctc tat ctg
ggc agt gcc cgg gat tcc gcc aat ttg gag agc ctg 788 Asn Leu Tyr Leu
Gly Ser Ala Arg Asp Ser Ala Asn Leu Glu Ser Leu 210 215
220 225 gcc aaa ctg ggc atc cgc tac atc ctc
aat gtc acc ccc aac ctc cca 836 Ala Lys Leu Gly Ile Arg Tyr Ile Leu
Asn Val Thr Pro Asn Leu Pro 230 235
240 aac ttc ttc gag aag aat ggt gac ttt cac tac aag cag atc ccc
atc 884 Asn Phe Phe Glu Lys Asn Gly Asp Phe His Tyr Lys Gln Ile Pro
Ile 245 250 255 tcc gac cac
tgg agc cag aac ctg tcg cgg ttc ttt ccg gag gcc att 932 Ser Asp His
Trp Ser Gln Asn Leu Ser Arg Phe Phe Pro Glu Ala Ile 260
265 270 gag ttc att gat gag gcc ttg tcc cag aac tgc
ggg gtg ctc gtc cac 980 Glu Phe Ile Asp Glu Ala Leu Ser Gln Asn Cys
Gly Val Leu Val His 275 280 285 tgc
ttg gcg ggg gtc agc cgt tct gtc acc gtc act gtg gcc tac ctc 1028 Cys
Leu Ala Gly Val Ser Arg Ser Val Thr Val Thr Val Ala Tyr Leu 290
295 300 305 atg cag aag ctc cac ctc
tct ctc aac gat gcc tat gac ctg gtc aag 1076 Met Gln Lys Leu His Leu
Ser Leu Asn Asp Ala Tyr Asp Leu Val Lys 310
315 320 agg aag aag tct aac atc tcc ccc aac ttc aac ttc
atg ggg cag ttg 1124 Arg Lys Lys Ser Asn Ile Ser Pro Asn Phe Asn Phe
Met Gly Gln Leu 325 330 335
ctg gac ttt gag cgc agc ttg cgg ctg gag gag cgc cac tcg cag gag 1172
Leu Asp Phe Glu Arg Ser Leu Arg Leu Glu Glu Arg His Ser Gln Glu
340 345 350 cag ggc agt ggg ggg cag gca
tct gcg gcc tcc aac ccg ccc tcc ttc 1220 Gln Gly Ser Gly Gly Gln Ala
Ser Ala Ala Ser Asn Pro Pro Ser Phe 355 360
365 ttc acc acc ccc acc agt gat ggc gcc ttc gag ctg gcc ccc acc tag
1268 Phe Thr Thr Pro Thr Ser Asp Gly Ala Phe Glu Leu Ala Pro Thr 370
375 380 ggccccgtgg ccggcaggcc ggcccctgcc
ccacccccac ccacgggtgt ccctgcccac 1328 tcgtgtggca agggagggga gggcaggagg
gctcggcctg agcagggtgc tggggggaga 1388 gcgcaatacc tcacgcgggc tgccgtccta
atcaacgtgc ctatggcggg accacgctcg 1448 gagcctgcct cttctgcgac tgttactttt
tctttgcggg atgggggtgg gggttccctc 1508 tccaggtggt tgtccaggcc catgtcccgg
ccctgggtgc tcagccagct cggctaggcc 1568 ctgcgcctcc ctgcgcttcc cccttcagga
agggtgtgtg ccacctcgtt gcactggatc 1628 ccagtggctg cttgggggag aggcgtttgc
catcactggt gttgtcacct ccctgtttct 1688 ccaccaaggg cttgggcctc tcggggctgg
ggcctcccag gggatgggga cccagagtgc 1748 agtggccgcc cacatccatg gcctaggagc
tactgggcag gttcccggcc acacatctgg 1808 tgggctgttt tgtttttttt ttttcctctt
cccccagatg tcttgacggg atcactgggg 1868 ctctttgtga gtgagggtgg ccaaactacc
gccggaggag atggggtctc agagcgagag 1928 ctgcggaggg ggaggggaag aagaaggcct
cacttttgct gctgcggggc ccacacagcc 1988 gctgctactt tggggggtgg ggaaggggcc
aagctgcaga cacacacagt cattcatttc 2048 tgtccacacc cctgtgggtg gcgggtgtgc
gtgtgtgtgc ttgtgtgtgc gcacgtgtcg 2108 gcgctcacac acacatgcta gcccactgat
gcacccagcc cagggctggc agtctttgca 2168 gcgtggggcc gtctcaccct ggagcctgga
gaggatctat gcttgtttgt ttttgtaatc 2228 catatcatag ttgctttctt taattgttcc
ttctgaataa acagtttatt taagat 2284 20 384 PRT Homo sapiens 20 Met
Glu Gly Leu Gly Arg Ser Cys Leu Trp Leu Arg Arg Glu Leu Ser 1
5 10 15 Pro Pro Arg Pro Arg Leu Leu
Leu Leu Asp Cys Arg Ser Arg Glu Leu 20 25
30 Tyr Glu Ser Ala Arg Ile Gly Gly Ala Leu Ser Val Ala Leu
Pro Ala 35 40 45 Leu Leu Leu
Arg Arg Leu Arg Arg Gly Ser Leu Ser Val Arg Ala Leu 50
55 60 Leu Pro Gly Pro Pro Leu Gln Pro Pro Pro Pro Ala
Pro Val Leu Leu 65 70 75
80 Tyr Asp Gln Gly Gly Gly Arg Arg Arg Arg Gly Glu Ala Glu Ala Glu
85 90 95 Ala Glu Glu Trp Glu
Ala Glu Ser Val Leu Gly Thr Leu Leu Gln Lys 100
105 110 Leu Arg Glu Glu Gly Tyr Leu Ala Tyr Tyr Leu Gln
Gly Gly Phe Ser 115 120 125 Arg
Phe Gln Ala Glu Cys Pro His Leu Cys Glu Thr Ser Leu Ala Gly 130
135 140 Arg Ala Gly Ser Ser Met Ala Pro Val Pro
Gly Pro Val Pro Val Val 145 150 155
160 Gly Leu Gly Ser Leu Cys Leu Gly Ser Asp Cys Ser Asp Ala Glu
Ser 165 170 175 Glu Ala
Asp Arg Asp Ser Met Ser Cys Gly Leu Asp Ser Glu Gly Ala 180
185 190 Thr Pro Pro Pro Val Gly Leu Arg Ala
Ser Phe Pro Val Gln Ile Leu 195 200
205 Pro Asn Leu Tyr Leu Gly Ser Ala Arg Asp Ser Ala Asn Leu Glu Ser
210 215 220 Leu Ala Lys Leu Gly Ile Arg
Tyr Ile Leu Asn Val Thr Pro Asn Leu 225 230
235 240 Pro Asn Phe Phe Glu Lys Asn Gly Asp Phe His Tyr
Lys Gln Ile Pro 245 250
255 Ile Ser Asp His Trp Ser Gln Asn Leu Ser Arg Phe Phe Pro Glu Ala
260 265 270 Ile Glu Phe Ile Asp Glu
Ala Leu Ser Gln Asn Cys Gly Val Leu Val 275 280
285 His Cys Leu Ala Gly Val Ser Arg Ser Val Thr Val Thr Val
Ala Tyr 290 295 300 Leu Met Gln Lys
Leu His Leu Ser Leu Asn Asp Ala Tyr Asp Leu Val 305 310
315 320 Lys Arg Lys Lys Ser Asn Ile Ser Pro
Asn Phe Asn Phe Met Gly Gln 325 330
335 Leu Leu Asp Phe Glu Arg Ser Leu Arg Leu Glu Glu Arg His Ser
Gln 340 345 350 Glu Gln Gly
Ser Gly Gly Gln Ala Ser Ala Ala Ser Asn Pro Pro Ser 355
360 365 Phe Phe Thr Thr Pro Thr Ser Asp Gly Ala Phe
Glu Leu Ala Pro Thr 370 375 380 21
1724 DNA Homo sapiens CDS (31)..(1071) E2F5 mRNA 21 ggggcccgac
caccgcgggg ccgggacgcg atg gcg gcg gca gag ccc gcg agc 54
Met Ala Ala Ala Glu Pro Ala Ser
1 5 tcg ggc cag cag gcg ccg gca ggg cag ggg cag
ggc cag cgg ccg ccg 102 Ser Gly Gln Gln Ala Pro Ala Gly Gln Gly Gln
Gly Gln Arg Pro Pro 10 15 20 ccg
cag cct ccg cag gcg caa gcc ccg cag ccg ccc ccg ccg ccg cag 150 Pro
Gln Pro Pro Gln Ala Gln Ala Pro Gln Pro Pro Pro Pro Pro Gln 25
30 35 40 ctc ggg ggc gcc ggg ggc
ggc agc agc agg cac gag aag agc ctg ggg 198 Leu Gly Gly Ala Gly Gly
Gly Ser Ser Arg His Glu Lys Ser Leu Gly 45
50 55 ctg ctc act acc aag ttc gtg tcg ctg ctg cag gag
gcc aag gac ggc 246 Leu Leu Thr Thr Lys Phe Val Ser Leu Leu Gln Glu
Ala Lys Asp Gly 60 65 70
gtt ctg gat ctc aaa gcg gct gct gat act ttg gct gtg agg caa aaa 294
Val Leu Asp Leu Lys Ala Ala Ala Asp Thr Leu Ala Val Arg Gln Lys
75 80 85 agg aga att tat gat atc acc
aat gtc tta gag gga att gac ttg att 342 Arg Arg Ile Tyr Asp Ile Thr
Asn Val Leu Glu Gly Ile Asp Leu Ile 90 95
100 gaa aaa aag tca aaa aac agt atc cag tgg aaa ggt gta ggt gct ggc
390 Glu Lys Lys Ser Lys Asn Ser Ile Gln Trp Lys Gly Val Gly Ala Gly
105 110 115 120 tgt aat
act aaa gaa gtc ata gat aga tta aga tat ctt aaa gct gaa 438 Cys Asn
Thr Lys Glu Val Ile Asp Arg Leu Arg Tyr Leu Lys Ala Glu
125 130 135 att gaa gat cta gaa ctg aag
gaa aga gaa ctt gat cag cag aag ttg 486 Ile Glu Asp Leu Glu Leu Lys
Glu Arg Glu Leu Asp Gln Gln Lys Leu 140 145
150 tgg cta cag caa agc atc aaa aat gtg atg gac gat tcc att
aat aat 534 Trp Leu Gln Gln Ser Ile Lys Asn Val Met Asp Asp Ser Ile
Asn Asn 155 160 165 aga ttt tcc
tat gta act cat gaa gac atc tgt aat tgc ttt aat ggt 582 Arg Phe Ser
Tyr Val Thr His Glu Asp Ile Cys Asn Cys Phe Asn Gly 170
175 180 gat aca ctt ttg gcc att cag gca cct tct ggt aca
caa ctg gag gta 630 Asp Thr Leu Leu Ala Ile Gln Ala Pro Ser Gly Thr
Gln Leu Glu Val 185 190 195
200 ccc att cca gaa atg ggt cag aat gga caa aag aaa tac cag atc aat
678 Pro Ile Pro Glu Met Gly Gln Asn Gly Gln Lys Lys Tyr Gln Ile Asn
205 210 215 cta aag agt cat
tca gga cct atc cat gtg ctg ctt ata aat aaa gag 726 Leu Lys Ser His
Ser Gly Pro Ile His Val Leu Leu Ile Asn Lys Glu 220
225 230 tcg agt tca tct aag ccc gtg gtt ttt cct gtt
ccc cca cct gat gac 774 Ser Ser Ser Ser Lys Pro Val Val Phe Pro Val
Pro Pro Pro Asp Asp 235 240 245
ctc aca cag cct tcc tcc cag tcc ttg act cca gtg act cca cag aaa 822
Leu Thr Gln Pro Ser Ser Gln Ser Leu Thr Pro Val Thr Pro Gln Lys 250
255 260 tcc agc atg gca act caa aat ctg cct
gag caa cat gtc tct gaa aga 870 Ser Ser Met Ala Thr Gln Asn Leu Pro
Glu Gln His Val Ser Glu Arg 265 270 275
280 agc cag gct ctg cag cag aca tca gct aca gat ata tct tca
gca gga 918 Ser Gln Ala Leu Gln Gln Thr Ser Ala Thr Asp Ile Ser Ser
Ala Gly 285 290 295 tct
att agt gga gat atc att gat gag tta atg tct tct gac gtg ttt 966 Ser
Ile Ser Gly Asp Ile Ile Asp Glu Leu Met Ser Ser Asp Val Phe
300 305 310 cct ctc tta agg ctt tct cct
acc ccg gca gat gac tac aac ttt aat 1014 Pro Leu Leu Arg Leu Ser Pro
Thr Pro Ala Asp Asp Tyr Asn Phe Asn 315 320
325 tta gat gat aac gaa gga gtt tgt gat ctg ttt gat gtc cag ata
cta 1062 Leu Asp Asp Asn Glu Gly Val Cys Asp Leu Phe Asp Val Gln Ile
Leu 330 335 340 aat tat tag
attccatgga aacttgggac tgttatctac ctctaactgt 1111 Asn Tyr 345
gtaacatttt agacttctta ataacctaaa tatttaaaat aatgaatgta acaccttttt 1171
tagttcactg attctgaagt gttcttccct aatactttct ttacttcaca aaacttcaac 1231
cataaaaaca aagggctctg attgctttag gggataagtg atttaatatc cacaaacgtc 1291
cccactccca aaagtaacta tattctggat ttcaactttt cttctaattg tgaatccttc 1351
tgttttttct tcttaaggag gaaagttaaa ggacactaca ggtcatcaaa aacaagttgg 1411
ccaaggactc attacttgtc ttatattttt actgccacta aactgcctgt atttctgtat 1471
gtccttctat ccaaacagac gttcactgcc acttgtaaag tgaaggatgt aaacgaggat 1531
atataactgt ttcagtgaac agattttgtg aagtgccttc tgttttagca ctttaagttt 1591
atcacatttt gttgacttct gacattccac tttcctaggt tataggaaag atctgtttat 1651
gtagtttgtt tttaaaatgt gccaatgcct gtacattaac aagattttta aaaataaaat 1711
tgtataaaac att 1724
22 346 PRT Homo sapiens 22 Met Ala Ala Ala Glu Pro Ala Ser Ser Gly Gln
Gln Ala Pro Ala Gly 1 5 10
15 Gln Gly Gln Gly Gln Arg Pro Pro Pro Gln Pro Pro Gln Ala Gln Ala
20 25 30 Pro Gln Pro Pro Pro Pro
Pro Gln Leu Gly Gly Ala Gly Gly Gly Ser 35 40
45 Ser Arg His Glu Lys Ser Leu Gly Leu Leu Thr Thr Lys Phe
Val Ser 50 55 60 Leu Leu Gln Glu
Ala Lys Asp Gly Val Leu Asp Leu Lys Ala Ala Ala 65 70
75 80 Asp Thr Leu Ala Val Arg Gln Lys Arg
Arg Ile Tyr Asp Ile Thr Asn 85 90
95 Val Leu Glu Gly Ile Asp Leu Ile Glu Lys Lys Ser Lys Asn Ser
Ile 100 105 110 Gln Trp Lys
Gly Val Gly Ala Gly Cys Asn Thr Lys Glu Val Ile Asp 115
120 125 Arg Leu Arg Tyr Leu Lys Ala Glu Ile Glu Asp
Leu Glu Leu Lys Glu 130 135 140 Arg
Glu Leu Asp Gln Gln Lys Leu Trp Leu Gln Gln Ser Ile Lys Asn 145
150 155 160 Val Met Asp Asp Ser Ile
Asn Asn Arg Phe Ser Tyr Val Thr His Glu 165
170 175 Asp Ile Cys Asn Cys Phe Asn Gly Asp Thr Leu Leu
Ala Ile Gln Ala 180 185 190
Pro Ser Gly Thr Gln Leu Glu Val Pro Ile Pro Glu Met Gly Gln Asn
195 200 205 Gly Gln Lys Lys Tyr Gln Ile
Asn Leu Lys Ser His Ser Gly Pro Ile 210 215
220 His Val Leu Leu Ile Asn Lys Glu Ser Ser Ser Ser Lys Pro Val Val
225 230 235 240 Phe Pro
Val Pro Pro Pro Asp Asp Leu Thr Gln Pro Ser Ser Gln Ser
245 250 255 Leu Thr Pro Val Thr Pro Gln
Lys Ser Ser Met Ala Thr Gln Asn Leu 260 265
270 Pro Glu Gln His Val Ser Glu Arg Ser Gln Ala Leu Gln Gln
Thr Ser 275 280 285 Ala Thr Asp
Ile Ser Ser Ala Gly Ser Ile Ser Gly Asp Ile Ile Asp 290
295 300 Glu Leu Met Ser Ser Asp Val Phe Pro Leu Leu Arg
Leu Ser Pro Thr 305 310 315
320 Pro Ala Asp Asp Tyr Asn Phe Asn Leu Asp Asp Asn Glu Gly Val Cys
325 330 335 Asp Leu Phe Asp
Val Gln Ile Leu Asn Tyr 340 345 23 4413 DNA
Homo sapiens CDS (44)..(1960) GHR mRNA 23 ccgcgctctc tgatcagagg
cgaagctcgg aggtcctaca ggt atg gat ctc tgg 55
Met Asp Leu Trp
1 cag ctg ctg ttg acc ttg gca ctg gca gga tca agt gat gct
ttt tct 103 Gln Leu Leu Leu Thr Leu Ala Leu Ala Gly Ser Ser Asp Ala
Phe Ser 5 10 15 20
gga agt gag gcc aca gca gct atc ctt agc aga gca ccc tgg agt ctg 151
Gly Ser Glu Ala Thr Ala Ala Ile Leu Ser Arg Ala Pro Trp Ser Leu
25 30 35 caa agt gtt aat cca ggc
cta aag aca aat tct tct aag gag cct aaa 199 Gln Ser Val Asn Pro Gly
Leu Lys Thr Asn Ser Ser Lys Glu Pro Lys 40
45 50 ttc acc aag tgc cgt tca cct gag cga gag act ttt
tca tgc cac tgg 247 Phe Thr Lys Cys Arg Ser Pro Glu Arg Glu Thr Phe
Ser Cys His Trp 55 60 65 aca
gat gag gtt cat cat ggt aca aag aac cta gga ccc ata cag ctg 295 Thr
Asp Glu Val His His Gly Thr Lys Asn Leu Gly Pro Ile Gln Leu 70
75 80 ttc tat acc aga agg aac act caa gaa tgg
act caa gaa tgg aaa gaa 343 Phe Tyr Thr Arg Arg Asn Thr Gln Glu Trp
Thr Gln Glu Trp Lys Glu 85 90 95
100 tgc cct gat tat gtt tct gct ggg gaa aac agc tgt tac ttt aat
tca 391 Cys Pro Asp Tyr Val Ser Ala Gly Glu Asn Ser Cys Tyr Phe Asn
Ser 105 110 115 tcg ttt
acc tcc atc tgg ata cct tat tgt atc aag cta act agc aat 439 Ser Phe
Thr Ser Ile Trp Ile Pro Tyr Cys Ile Lys Leu Thr Ser Asn 120
125 130 ggt ggt aca gtg gat gaa aag tgt ttc
tct gtt gat gaa ata gtg caa 487 Gly Gly Thr Val Asp Glu Lys Cys Phe
Ser Val Asp Glu Ile Val Gln 135 140
145 cca gat cca ccc att gcc ctc aac tgg act tta ctg aac gtc agt tta
535 Pro Asp Pro Pro Ile Ala Leu Asn Trp Thr Leu Leu Asn Val Ser Leu
150 155 160 act ggg att cat gca gat atc
caa gtg aga tgg gaa gca cca cgc aat 583 Thr Gly Ile His Ala Asp Ile
Gln Val Arg Trp Glu Ala Pro Arg Asn 165 170
175 180 gca gat att cag aaa gga tgg atg gtt ctg gag tat
gaa ctt caa tac 631 Ala Asp Ile Gln Lys Gly Trp Met Val Leu Glu Tyr
Glu Leu Gln Tyr 185 190
195 aaa gaa gta aat gaa act aaa tgg aaa atg atg gac cct ata ttg aca
679 Lys Glu Val Asn Glu Thr Lys Trp Lys Met Met Asp Pro Ile Leu Thr
200 205 210 aca tca gtt cca gtg tac
tca ttg aaa gtg gat aag gaa tat gaa gtg 727 Thr Ser Val Pro Val Tyr
Ser Leu Lys Val Asp Lys Glu Tyr Glu Val 215 220
225 cgt gtg aga tcc aaa caa cga aac tct gga aat tat ggc gag
ttc agt 775 Arg Val Arg Ser Lys Gln Arg Asn Ser Gly Asn Tyr Gly Glu
Phe Ser 230 235 240 gag gtg ctc tat
gta aca ctt cct cag atg agc caa ttt aca tgt gaa 823 Glu Val Leu Tyr
Val Thr Leu Pro Gln Met Ser Gln Phe Thr Cys Glu 245 250
255 260 gaa gat ttc tac ttt cca tgg ctc tta
att att atc ttt gga ata ttt 871 Glu Asp Phe Tyr Phe Pro Trp Leu Leu
Ile Ile Ile Phe Gly Ile Phe 265 270
275 ggg cta aca gtg atg cta ttt gta ttc tta ttt tct aaa cag caa
agg 919 Gly Leu Thr Val Met Leu Phe Val Phe Leu Phe Ser Lys Gln Gln
Arg 280 285 290 att aaa atg
ctg att ctg ccc cca gtt cca gtt cca aag att aaa gga 967 Ile Lys Met
Leu Ile Leu Pro Pro Val Pro Val Pro Lys Ile Lys Gly 295
300 305 atc gat cca gat ctc ctc aag gaa gga aaa tta
gag gag gtg aac aca 1015 Ile Asp Pro Asp Leu Leu Lys Glu Gly Lys Leu
Glu Glu Val Asn Thr 310 315 320 atc
tta gcc att cat gat agc tat aaa ccc gaa ttc cac agt gat gac 1063 Ile
Leu Ala Ile His Asp Ser Tyr Lys Pro Glu Phe His Ser Asp Asp 325
330 335 340 tct tgg gtt gaa ttt att
gag cta gat att gat gag cca gat gaa aag 1111 Ser Trp Val Glu Phe Ile
Glu Leu Asp Ile Asp Glu Pro Asp Glu Lys 345
350 355 act gag gaa tca gac aca gac aga ctt cta agc agt
gac cat gag aaa 1159 Thr Glu Glu Ser Asp Thr Asp Arg Leu Leu Ser Ser
Asp His Glu Lys 360 365 370
tca cat agt aac cta ggg gtg aag gat ggc gac tct gga cgt acc agc 1207
Ser His Ser Asn Leu Gly Val Lys Asp Gly Asp Ser Gly Arg Thr Ser
375 380 385 tgt tgt gaa cct gac att ctg
gag act gat ttc aat gcc aat gac ata 1255 Cys Cys Glu Pro Asp Ile Leu
Glu Thr Asp Phe Asn Ala Asn Asp Ile 390 395
400 cat gag ggt acc tca gag gtt gct cag cca cag agg tta aaa ggg gaa
1303 His Glu Gly Thr Ser Glu Val Ala Gln Pro Gln Arg Leu Lys Gly Glu
405 410 415 420 gca gat
ctc tta tgc ctt gac cag aag aat caa aat aac tca cct tat 1351 Ala Asp
Leu Leu Cys Leu Asp Gln Lys Asn Gln Asn Asn Ser Pro Tyr
425 430 435 cat gat gct tgc cct gct act
cag cag ccc agt gtt atc caa gca gag 1399 His Asp Ala Cys Pro Ala Thr
Gln Gln Pro Ser Val Ile Gln Ala Glu 440 445
450 aaa aac aaa cca caa cca ctt cct act gaa gga gct gag tca
act cac 1447 Lys Asn Lys Pro Gln Pro Leu Pro Thr Glu Gly Ala Glu Ser
Thr His 455 460 465 caa gct gcc
cat att cag cta agc aat cca agt tca ctg tca aac atc 1495 Gln Ala Ala
His Ile Gln Leu Ser Asn Pro Ser Ser Leu Ser Asn Ile 470
475 480 gac ttt tat gcc cag gtg agc gac att aca cca gca
ggt agt gtg gtc 1543 Asp Phe Tyr Ala Gln Val Ser Asp Ile Thr Pro Ala
Gly Ser Val Val 485 490 495
500 ctt tcc ccg ggc caa aag aat aag gca ggg atg tcc caa tgt gac atg
1591 Leu Ser Pro Gly Gln Lys Asn Lys Ala Gly Met Ser Gln Cys Asp Met
505 510 515 cac ccg gaa atg
gtc tca ctc tgc caa gaa aac ttc ctt atg gac aat 1639 His Pro Glu Met
Val Ser Leu Cys Gln Glu Asn Phe Leu Met Asp Asn 520
525 530 gcc tac ttc tgt gag gca gat gcc aaa aag tgc
atc cct gtg gct cct 1687 Ala Tyr Phe Cys Glu Ala Asp Ala Lys Lys Cys
Ile Pro Val Ala Pro 535 540 545
cac atc aag gtt gaa tca cac ata cag cca agc tta aac caa gag gac 1735
His Ile Lys Val Glu Ser His Ile Gln Pro Ser Leu Asn Gln Glu Asp 550
555 560 att tac atc acc aca gaa agc ctt acc
act gct gct ggg agg cct ggg 1783 Ile Tyr Ile Thr Thr Glu Ser Leu Thr
Thr Ala Ala Gly Arg Pro Gly 565 570 575
580 aca gga gaa cat gtt cca ggt tct gag atg cct gtc cca gac
tat acc 1831 Thr Gly Glu His Val Pro Gly Ser Glu Met Pro Val Pro Asp
Tyr Thr 585 590 595 tcc
att cat ata gta cag tcc cca cag ggc ctc ata ctc aat gcg act 1879 Ser
Ile His Ile Val Gln Ser Pro Gln Gly Leu Ile Leu Asn Ala Thr
600 605 610 gcc ttg ccc ttg cct gac aaa
gag ttt ctc tca tca tgt ggc tat gtg 1927 Ala Leu Pro Leu Pro Asp Lys
Glu Phe Leu Ser Ser Cys Gly Tyr Val 615 620
625 agc aca gac caa ctg aac aaa atc atg cct tag cctttctttg
gtttcccaag 1980 Ser Thr Asp Gln Leu Asn Lys Ile Met Pro 630
635 agctacgtat ttaatagcaa agaattgact ggggcaataa cgtttaagcc
aaaacaatgt 2040 ttaaaccttt tttgggggag tgacaggatg gggtatggat tctaaaatgc
cttttcccaa 2100 aatgttgaaa tatgatgtta aaaaaataag aagaatgctt aatcagatag
atattcctat 2160 tgtgcaatgt aaatatttta aagaattgtg tcagactgtt tagtagcagt
gattgtctta 2220 atattgtggg tgttaatttt tgatactaag cattgaatgg ctatgttttt
aatgtatagt 2280 aaatcacgct ttttgaaaaa gcgaaaaaat caggtggctt ttgcggttca
ggaaaattga 2340 atgcaaacca tagcacaggc taattttttg ttgtttctta aataagaaac
ttttttattt 2400 aaaaaactaa aaactagagg tgagaaattt aaactataag caagaaggca
aaaatagttt 2460 ggatatgtaa aacatttatt ttgacataaa gttgataaag attttttaat
aatttagact 2520 tcaagcatgg ctattttata ttacactaca cactgtgtac tgcagttggt
atgacccctc 2580 taaggagtgt agcaactaca gtctaaagct ggtttaatgt tttggccaat
gcacctaaag 2640 aaaaacaaac tcgtttttta caaagccctt ttatacctcc ccagactcct
tcaacaattc 2700 taaaatgatt gtagtaatct gcattattgg aatataattg ttttatctga
atttttaaac 2760 aagtatttgt taatttagaa aactttaaag cgtttgcaca gatcaactta
ccaggcacca 2820 aaagaagtaa aagcaaaaaa gaaaaccttt cttcaccaaa tcttggttga
tgccaaaaaa 2880 aaatacatgc taagagaagt agaaatcata gctggttcac actgaccaag
atacttaagt 2940 gctgcaattg cacgcggagt gagtttttta gtgcgtgcag atggtgagag
ataagatcta 3000 tagcctctgc agcggaatct gttcacaccc aacttggttt tgctacataa
ttatccagga 3060 agggaataag gtacaagaag cattttgtaa gttgaagcaa atcgaatgaa
attaactggg 3120 taatgaaaca aagagttcaa gaaataagtt tttgtttcac agcctataac
cagacacata 3180 ctcatttttc atgataatga acagaacata gacagaagaa acaaggtttt
cagtccccac 3240 agataactga aaattattta aaccgctaaa agaaactttc tttctcacta
aatcttttat 3300 aggatttatt taaaatagca aaagaagaag tttcatcatt ttttacttcc
tctctgagtg 3360 gactggcctc aaagcaagca ttcagaagaa aaagaagcaa cctcagtaat
ttagaaatca 3420 ttttgcaatc ccttaatatc ctaaacatca ttcatttttg ttgttgttgt
tgttgttgag 3480 acagagtctc gctctgtcgc caggctagag tgcggtggcg cgatcttgac
tcactgcaat 3540 ctccacctcc cacaggttca ggcgattccc gtgcctcagc ctcctgagta
gctgggacta 3600 caggcacgca ccaccatgcc aggctaattt ttttgtattt tagcagagac
ggggtttcac 3660 catgttggcc aggatggtct cgatctcctg acctcgtgat ccacccgact
cggcctccca 3720 aagtgctggg attacaggtg taagccaccg tgcccagccc taaacatcat
tcttgagagc 3780 attgggatat ctcctgaaaa ggtttatgaa aaagaagaat ctcatctcag
tgaagaatac 3840 ttctcatttt ttaaaaaagc ttaaaacttt gaagttagct ttaacttaaa
tagtatttcc 3900 catttatcgc agaccttttt taggaagcaa gcttaatggc tgataatttt
aaattctctc 3960 tcttgcagga aggactatga aaagctagaa ttgagtgttt aaagttcaac
atgttatttg 4020 taatagatgt ttgatagatt ttctgctact ttgctgctat ggttttctcc
aagagctaca 4080 taatttagtt tcatataaag tatcatcagt gtagaaccta attcaattca
aagctgtgtg 4140 tttggaagac tatcttacta tttcacaaca gcctgacaac atttctatag
ccaaaaatag 4200 ctaaatacct caatcagtct cagaatgtca ttttggtact ttggtggcca
cataagccat 4260 tattcactag tatgactagt tgtgtctggc agtttatatt taactctctt
tatgtctgtg 4320 gattttttcc ttcaaagttt aataaattta ttttcttgga ttcctgatag
tgtgcttctg 4380 ttatcaaaca ccaacataaa aatgatctaa acc
4413 24 638 PRT Homo sapiens 24 Met Asp Leu Trp Gln Leu Leu
Leu Thr Leu Ala Leu Ala Gly Ser Ser 1 5
10 15 Asp Ala Phe Ser Gly Ser Glu Ala Thr Ala Ala Ile
Leu Ser Arg Ala 20 25 30
Pro Trp Ser Leu Gln Ser Val Asn Pro Gly Leu Lys Thr Asn Ser Ser
35 40 45 Lys Glu Pro Lys Phe Thr Lys
Cys Arg Ser Pro Glu Arg Glu Thr Phe 50 55
60 Ser Cys His Trp Thr Asp Glu Val His His Gly Thr Lys Asn Leu Gly
65 70 75 80 Pro Ile
Gln Leu Phe Tyr Thr Arg Arg Asn Thr Gln Glu Trp Thr Gln
85 90 95 Glu Trp Lys Glu Cys Pro Asp
Tyr Val Ser Ala Gly Glu Asn Ser Cys 100 105
110 Tyr Phe Asn Ser Ser Phe Thr Ser Ile Trp Ile Pro Tyr Cys
Ile Lys 115 120 125 Leu Thr Ser
Asn Gly Gly Thr Val Asp Glu Lys Cys Phe Ser Val Asp 130
135 140 Glu Ile Val Gln Pro Asp Pro Pro Ile Ala Leu Asn
Trp Thr Leu Leu 145 150 155
160 Asn Val Ser Leu Thr Gly Ile His Ala Asp Ile Gln Val Arg Trp Glu
165 170 175 Ala Pro Arg Asn
Ala Asp Ile Gln Lys Gly Trp Met Val Leu Glu Tyr 180
185 190 Glu Leu Gln Tyr Lys Glu Val Asn Glu Thr Lys
Trp Lys Met Met Asp 195 200 205
Pro Ile Leu Thr Thr Ser Val Pro Val Tyr Ser Leu Lys Val Asp Lys 210
215 220 Glu Tyr Glu Val Arg Val Arg Ser Lys
Gln Arg Asn Ser Gly Asn Tyr 225 230 235
240 Gly Glu Phe Ser Glu Val Leu Tyr Val Thr Leu Pro Gln Met
Ser Gln 245 250 255 Phe
Thr Cys Glu Glu Asp Phe Tyr Phe Pro Trp Leu Leu Ile Ile Ile
260 265 270 Phe Gly Ile Phe Gly Leu Thr
Val Met Leu Phe Val Phe Leu Phe Ser 275 280
285 Lys Gln Gln Arg Ile Lys Met Leu Ile Leu Pro Pro Val Pro Val
Pro 290 295 300 Lys Ile Lys Gly Ile
Asp Pro Asp Leu Leu Lys Glu Gly Lys Leu Glu 305 310
315 320 Glu Val Asn Thr Ile Leu Ala Ile His Asp
Ser Tyr Lys Pro Glu Phe 325 330
335 His Ser Asp Asp Ser Trp Val Glu Phe Ile Glu Leu Asp Ile Asp Glu
340 345 350 Pro Asp Glu Lys
Thr Glu Glu Ser Asp Thr Asp Arg Leu Leu Ser Ser 355
360 365 Asp His Glu Lys Ser His Ser Asn Leu Gly Val Lys
Asp Gly Asp Ser 370 375 380 Gly Arg
Thr Ser Cys Cys Glu Pro Asp Ile Leu Glu Thr Asp Phe Asn 385
390 395 400 Ala Asn Asp Ile His Glu Gly
Thr Ser Glu Val Ala Gln Pro Gln Arg 405
410 415 Leu Lys Gly Glu Ala Asp Leu Leu Cys Leu Asp Gln
Lys Asn Gln Asn 420 425 430
Asn Ser Pro Tyr His Asp Ala Cys Pro Ala Thr Gln Gln Pro Ser Val
435 440 445 Ile Gln Ala Glu Lys Asn Lys
Pro Gln Pro Leu Pro Thr Glu Gly Ala 450 455
460 Glu Ser Thr His Gln Ala Ala His Ile Gln Leu Ser Asn Pro Ser Ser
465 470 475 480 Leu Ser
Asn Ile Asp Phe Tyr Ala Gln Val Ser Asp Ile Thr Pro Ala
485 490 495 Gly Ser Val Val Leu Ser Pro
Gly Gln Lys Asn Lys Ala Gly Met Ser 500 505
510 Gln Cys Asp Met His Pro Glu Met Val Ser Leu Cys Gln Glu
Asn Phe 515 520 525 Leu Met Asp
Asn Ala Tyr Phe Cys Glu Ala Asp Ala Lys Lys Cys Ile 530
535 540 Pro Val Ala Pro His Ile Lys Val Glu Ser His Ile
Gln Pro Ser Leu 545 550 555
560 Asn Gln Glu Asp Ile Tyr Ile Thr Thr Glu Ser Leu Thr Thr Ala Ala
565 570 575 Gly Arg Pro Gly
Thr Gly Glu His Val Pro Gly Ser Glu Met Pro Val 580
585 590 Pro Asp Tyr Thr Ser Ile His Ile Val Gln Ser
Pro Gln Gly Leu Ile 595 600 605
Leu Asn Ala Thr Ala Leu Pro Leu Pro Asp Lys Glu Phe Leu Ser Ser 610
615 620 Cys Gly Tyr Val Ser Thr Asp Gln Leu
Asn Lys Ile Met Pro 625 630 635 25 1403
DNA Homo sapiens CDS (23)..(1204) HPD mRNA 25 ggaggtttga ctaagatcaa tc
atg acg act tac agt gac aaa ggg gca aag 52
Met Thr Thr Tyr Ser Asp Lys Gly Ala Lys 1
5 10 cct gag aga ggc cga ttc ctc cac ttc cac tct
gtg acc ttc tgg gtt 100 Pro Glu Arg Gly Arg Phe Leu His Phe His Ser
Val Thr Phe Trp Val 15 20
25 ggc aac gcc aag cag gcc acg tca ttc tac tgc agc aag atg ggc ttt
148 Gly Asn Ala Lys Gln Ala Thr Ser Phe Tyr Cys Ser Lys Met Gly Phe
30 35 40 gaa cct cta gcc tac agg
ggc ctg gag acc ggt tcc cgg gag gtg gtc 196 Glu Pro Leu Ala Tyr Arg
Gly Leu Glu Thr Gly Ser Arg Glu Val Val 45 50
55 agc cat gta atc aaa caa ggg aag att gtg ttt gtc ctc tcc
tca gcg 244 Ser His Val Ile Lys Gln Gly Lys Ile Val Phe Val Leu Ser
Ser Ala 60 65 70 ctc aac ccc tgg
aac aaa gag atg ggc gat cac ctg gtg aaa cac ggt 292 Leu Asn Pro Trp
Asn Lys Glu Met Gly Asp His Leu Val Lys His Gly 75 80
85 90 gac gga gtg aag gac att gcg ttc gag
gtg gaa gat tgt gac tac atc 340 Asp Gly Val Lys Asp Ile Ala Phe Glu
Val Glu Asp Cys Asp Tyr Ile 95 100
105 gtg cag aaa gca cgg gaa cgg ggc gcc aaa atc atg cgg gag ccc
tgg 388 Val Gln Lys Ala Arg Glu Arg Gly Ala Lys Ile Met Arg Glu Pro
Trp 110 115 120 gta gag caa
gac aag ttt ggg aag gtg aag ttt gct gtg ctg cag acg 436 Val Glu Gln
Asp Lys Phe Gly Lys Val Lys Phe Ala Val Leu Gln Thr 125
130 135 tat ggg gac acc aca cac acc ctg gtg gag aag
atg aac tac atc ggc 484 Tyr Gly Asp Thr Thr His Thr Leu Val Glu Lys
Met Asn Tyr Ile Gly 140 145 150 caa
ttc ttg cct gga tat gag gcc cca gcg ttc atg gac ccc cta ctt 532 Gln
Phe Leu Pro Gly Tyr Glu Ala Pro Ala Phe Met Asp Pro Leu Leu 155
160 165 170 cct aaa ctg ccc aaa tgc
agt ctg gag atg atc gac cac att gtg gga 580 Pro Lys Leu Pro Lys Cys
Ser Leu Glu Met Ile Asp His Ile Val Gly 175
180 185 aac cag cct gat cag gag atg gtg tcc gcc tcc gaa
tgg tac ctg aaa 628 Asn Gln Pro Asp Gln Glu Met Val Ser Ala Ser Glu
Trp Tyr Leu Lys 190 195 200
aac ctg cag ttc cac cgc ttc tgg tcc gtg gat gac acg cag gtg cac 676
Asn Leu Gln Phe His Arg Phe Trp Ser Val Asp Asp Thr Gln Val His
205 210 215 acg gaa tat agc tct ctg cga
tcc att gtg gtg gcc aac tat gaa gag 724 Thr Glu Tyr Ser Ser Leu Arg
Ser Ile Val Val Ala Asn Tyr Glu Glu 220 225
230 tcc atc aag atg ccc atc aat gag cca gcg cct ggc aag aag aag tcc
772 Ser Ile Lys Met Pro Ile Asn Glu Pro Ala Pro Gly Lys Lys Lys Ser
235 240 245 250 cag atc
cag gaa tat gtg gac tat aac ggg ggc gct ggg gtc cag cac 820 Gln Ile
Gln Glu Tyr Val Asp Tyr Asn Gly Gly Ala Gly Val Gln His
255 260 265 atc gct ctc aag acc gaa gac
atc atc aca gcg att cgc cac ttg aga 868 Ile Ala Leu Lys Thr Glu Asp
Ile Ile Thr Ala Ile Arg His Leu Arg 270 275
280 gag aga ggc ctg gag ttc tta tct gtt ccc tcc acg tac tac
aaa caa 916 Glu Arg Gly Leu Glu Phe Leu Ser Val Pro Ser Thr Tyr Tyr
Lys Gln 285 290 295 ctg cgg gag
aag ctg aag acg gcc aag atc aag gtg aag gag aac att 964 Leu Arg Glu
Lys Leu Lys Thr Ala Lys Ile Lys Val Lys Glu Asn Ile 300
305 310 gat gcc ctg gag gag ctg aaa atc ctg gtg gac tac
gac gag aaa ggc 1012 Asp Ala Leu Glu Glu Leu Lys Ile Leu Val Asp Tyr
Asp Glu Lys Gly 315 320 325
330 tac ctc ctg cag atc ttc acc aaa ccg gtg cag gac cgg ccc acg ctc
1060 Tyr Leu Leu Gln Ile Phe Thr Lys Pro Val Gln Asp Arg Pro Thr Leu
335 340 345 ttc ctg gaa gtc
atc cag cgc cac aac cac cag ggt ttt gga gcc ggc 1108 Phe Leu Glu Val
Ile Gln Arg His Asn His Gln Gly Phe Gly Ala Gly 350
355 360 aac ttc aac tca ctg ttc aag gct ttc gag gag
gag cag aac ctg cgg 1156 Asn Phe Asn Ser Leu Phe Lys Ala Phe Glu Glu
Glu Gln Asn Leu Arg 365 370 375
ggt aac ctc acc aac atg gag acc aat ggg gtg gtg ccc ggc atg taa 1204
Gly Asn Leu Thr Asn Met Glu Thr Asn Gly Val Val Pro Gly Met 380
385 390 gccccgccca ccccacggag gccacagcca
cacagccacg ccccctgatt ctggaactcg 1264 cccaacttcc ctactggctg ctccccttgg
gtcccgccca ccagcggact cggcccccaa 1324 ggctccgccc acactgacca cgcccctcgg
cggggccgcc ctctgctcca gccctcccga 1384 ttaaagcgtg ccccggtcc
1403 26 393 PRT Homo sapiens 26 Met
Thr Thr Tyr Ser Asp Lys Gly Ala Lys Pro Glu Arg Gly Arg Phe 1
5 10 15 Leu His Phe His Ser Val Thr
Phe Trp Val Gly Asn Ala Lys Gln Ala 20 25
30 Thr Ser Phe Tyr Cys Ser Lys Met Gly Phe Glu Pro Leu Ala
Tyr Arg 35 40 45 Gly Leu Glu
Thr Gly Ser Arg Glu Val Val Ser His Val Ile Lys Gln 50
55 60 Gly Lys Ile Val Phe Val Leu Ser Ser Ala Leu Asn
Pro Trp Asn Lys 65 70 75
80 Glu Met Gly Asp His Leu Val Lys His Gly Asp Gly Val Lys Asp Ile
85 90 95 Ala Phe Glu Val Glu
Asp Cys Asp Tyr Ile Val Gln Lys Ala Arg Glu 100
105 110 Arg Gly Ala Lys Ile Met Arg Glu Pro Trp Val Glu
Gln Asp Lys Phe 115 120 125 Gly
Lys Val Lys Phe Ala Val Leu Gln Thr Tyr Gly Asp Thr Thr His 130
135 140 Thr Leu Val Glu Lys Met Asn Tyr Ile Gly
Gln Phe Leu Pro Gly Tyr 145 150 155
160 Glu Ala Pro Ala Phe Met Asp Pro Leu Leu Pro Lys Leu Pro Lys
Cys 165 170 175 Ser Leu
Glu Met Ile Asp His Ile Val Gly Asn Gln Pro Asp Gln Glu 180
185 190 Met Val Ser Ala Ser Glu Trp Tyr Leu
Lys Asn Leu Gln Phe His Arg 195 200
205 Phe Trp Ser Val Asp Asp Thr Gln Val His Thr Glu Tyr Ser Ser Leu
210 215 220 Arg Ser Ile Val Val Ala Asn
Tyr Glu Glu Ser Ile Lys Met Pro Ile 225 230
235 240 Asn Glu Pro Ala Pro Gly Lys Lys Lys Ser Gln Ile
Gln Glu Tyr Val 245 250
255 Asp Tyr Asn Gly Gly Ala Gly Val Gln His Ile Ala Leu Lys Thr Glu
260 265 270 Asp Ile Ile Thr Ala Ile
Arg His Leu Arg Glu Arg Gly Leu Glu Phe 275 280
285 Leu Ser Val Pro Ser Thr Tyr Tyr Lys Gln Leu Arg Glu Lys
Leu Lys 290 295 300 Thr Ala Lys Ile
Lys Val Lys Glu Asn Ile Asp Ala Leu Glu Glu Leu 305 310
315 320 Lys Ile Leu Val Asp Tyr Asp Glu Lys
Gly Tyr Leu Leu Gln Ile Phe 325 330
335 Thr Lys Pro Val Gln Asp Arg Pro Thr Leu Phe Leu Glu Val Ile
Gln 340 345 350 Arg His Asn
His Gln Gly Phe Gly Ala Gly Asn Phe Asn Ser Leu Phe 355
360 365 Lys Ala Phe Glu Glu Glu Gln Asn Leu Arg Gly
Asn Leu Thr Asn Met 370 375 380 Glu
Thr Asn Gly Val Val Pro Gly Met 385 390 27 5413 DNA Homo
sapiens CDS (1145)..(5128) IGSF1 mRNA 27 ctccctgacc ccttgcgctt
cccgagtgag gcaatgcctc gccctgcttc ggctcgcgga 60 cggtgcgcgc acccactgac
ctgcgcccac tgtctggcac tccctagtga gatgaacccg 120 gtacctcaga tggaaatgca
gaaatcaccc gtcttctgcg tcgctcacgc tgggagctgt 180 agatcgcagc tgttcctatt
cggccatctt ggctcctcca ttctgctttt aagatacaag 240 tgatgccttc aaatttgagc
tcagtttctt atggtcgttt aggagcatga gggtttctgc 300 aataatctac cagtgaaatg
atgatcaaga tcttggaaag cactctactg gtcaagcaga 360 atagttcctg tcatctccat
taagaatcag cttattggcc gggcgcagtg gctcatgcct 420 gtaatcccag cactttggga
ggctgaggcg ggtggatcac gaggtcagga gatcgagacc 480 atcctgtcta acaaggaatc
ctttctggca ctcaagagtt ttcaattcaa tcctgatgaa 540 tgaatcatta tttgatacca
gcctaagtga agactctgcc tggtgacttt tgcaaggtgg 600 tttctggctg caacatggga
agatttggga tcagtgagat ggaagagatt tttctggcta 660 ctagcgttcc taagaaagag
ctttagtgtt tatccagcta ttggtatgca gacaactata 720 cccttatgag aaacactagc
agagaagaag agttcttgac tgggtctggg ccatgaaact 780 gaccggataa aagtgggaga
agaagagatg ttgaaggagg agtttgctgt tttctcagtc 840 tcctgagaaa ttccagatat
cttctttcta cacacatcct ggagagggga aagaaagcct 900 ctctctgtca cccaggctgg
aatgcactgg tgccatctcc actcactgca gcttcaacct 960 cctgggttca agtgatcctc
ctgcctctca gccttccgag tagctgggac tacagtttgc 1020 tgcatctgga ggagctcact
ggagaatctc caacatcgga gcgggccttc aactaccatc 1080 ccaccacctg ctgaggagaa
aaattcttca agactcagag cacacagcca gcaccagagg 1140 cccc atg acc ctg gac
aga cca ggg gag ggg gcc acc atg ctg aag aca 1189 Met Thr Leu Asp
Arg Pro Gly Glu Gly Ala Thr Met Leu Lys Thr 1 5
10 15 ttc act gtt ttg ctc ttt tgc att ctg atg
gac cct caa cca gag ttg 1237 Phe Thr Val Leu Leu Phe Cys Ile Leu Met
Asp Pro Gln Pro Glu Leu 20 25
30 tgg ata gag tcc aac tac ccc cag gcc cct tgg gag aac atc acg ctt
1285 Trp Ile Glu Ser Asn Tyr Pro Gln Ala Pro Trp Glu Asn Ile Thr Leu
35 40 45 tgg tgc cga agc ccc
tct cgg ata tca agc aag ttc ctg ctg ctg aag 1333 Trp Cys Arg Ser Pro
Ser Arg Ile Ser Ser Lys Phe Leu Leu Leu Lys 50
55 60 gat aag aca cag atg acc tgg atc cgc cct tcc cac
aag acc ttc caa 1381 Asp Lys Thr Gln Met Thr Trp Ile Arg Pro Ser His
Lys Thr Phe Gln 65 70 75 gtt tca
ttc ctt ata ggt gcc ctt act gag tcc aat gca ggt ctt tac 1429 Val Ser
Phe Leu Ile Gly Ala Leu Thr Glu Ser Asn Ala Gly Leu Tyr 80
85 90 95 cgg tgc tgc tac tgg aag gag
aca ggc tgg tca aag ccc agt aaa gtt 1477 Arg Cys Cys Tyr Trp Lys Glu
Thr Gly Trp Ser Lys Pro Ser Lys Val 100
105 110 cta gag ttg gag gca cca ggc caa ctg ccc aag ccc
atc ttc tgg att 1525 Leu Glu Leu Glu Ala Pro Gly Gln Leu Pro Lys Pro
Ile Phe Trp Ile 115 120 125
cag gct gag acc ccc gct ctt cct ggg tgt aat gtt aac atc ctc tgc 1573
Gln Ala Glu Thr Pro Ala Leu Pro Gly Cys Asn Val Asn Ile Leu Cys
130 135 140 cat ggc tgg ctg cag gat ttg
gta ttc atg ctg ttt aaa gag gga tat 1621 His Gly Trp Leu Gln Asp Leu
Val Phe Met Leu Phe Lys Glu Gly Tyr 145 150
155 gca gag cct gtg gat tac caa gtc cca act ggg aca atg gcc ata ttc
1669 Ala Glu Pro Val Asp Tyr Gln Val Pro Thr Gly Thr Met Ala Ile Phe
160 165 170 175 tcc att
gac aac ctg aca cct gag gat gaa ggg gtt tac atc tgc cgc 1717 Ser Ile
Asp Asn Leu Thr Pro Glu Asp Glu Gly Val Tyr Ile Cys Arg
180 185 190 act cat atc cag atg ctc ccc
acc ctg tgg tca gag ccc agc aac ccc 1765 Thr His Ile Gln Met Leu Pro
Thr Leu Trp Ser Glu Pro Ser Asn Pro 195 200
205 ctg aag ctg gtt gta gca gga ctc tac ccc aaa cca act ttg
aca gcc 1813 Leu Lys Leu Val Val Ala Gly Leu Tyr Pro Lys Pro Thr Leu
Thr Ala 210 215 220 cat cct ggg
ccc atc atg gca cct gga gaa agc ctg aat ctc agg tgc 1861 His Pro Gly
Pro Ile Met Ala Pro Gly Glu Ser Leu Asn Leu Arg Cys 225
230 235 caa ggg cca atc tat gga atg acc ttt gct cta atg
agg gtt gaa gac 1909 Gln Gly Pro Ile Tyr Gly Met Thr Phe Ala Leu Met
Arg Val Glu Asp 240 245 250
255 ttg gag aag tcc ttt tac cac aag aag aca ata aaa aat gag gca aat
1957 Leu Glu Lys Ser Phe Tyr His Lys Lys Thr Ile Lys Asn Glu Ala Asn
260 265 270 ttc ttc ttc cag
tct ttg aag atc caa gat act gga cat tac ctc tgt 2005 Phe Phe Phe Gln
Ser Leu Lys Ile Gln Asp Thr Gly His Tyr Leu Cys 275
280 285 ttt tac tat gac gca tca tat aga ggt tca ctc
ctt agt gat gtc ctg 2053 Phe Tyr Tyr Asp Ala Ser Tyr Arg Gly Ser Leu
Leu Ser Asp Val Leu 290 295 300
aaa atc tgg gtg act gac act ttc ccc aag acc tgg cta ctt gct cgg 2101
Lys Ile Trp Val Thr Asp Thr Phe Pro Lys Thr Trp Leu Leu Ala Arg 305
310 315 ccc agt gct gtg gtc caa atg ggt cag
aat gtg agc cta cgg tgt cga 2149 Pro Ser Ala Val Val Gln Met Gly Gln
Asn Val Ser Leu Arg Cys Arg 320 325 330
335 gga cca gtg gat gga gtg ggt ctt gca ctc tat aag aaa gga
gaa gac 2197 Gly Pro Val Asp Gly Val Gly Leu Ala Leu Tyr Lys Lys Gly
Glu Asp 340 345 350 aaa
cca ctt caa ttt ttg gat gcc acc agc atc gat gac aac aca tca 2245 Lys
Pro Leu Gln Phe Leu Asp Ala Thr Ser Ile Asp Asp Asn Thr Ser
355 360 365 ttc ttc ctc aac aat gta acc
tac agt gat act ggc atc tat agc tgc 2293 Phe Phe Leu Asn Asn Val Thr
Tyr Ser Asp Thr Gly Ile Tyr Ser Cys 370 375
380 cac tat ctt ctc acc tgg aag acc tcc att agg atg cca tca cac
aac 2341 His Tyr Leu Leu Thr Trp Lys Thr Ser Ile Arg Met Pro Ser His
Asn 385 390 395 act gtg gag ctt atg
gtt gta gat aag ccc ccc aaa ccc tcc ctg tca 2389 Thr Val Glu Leu Met
Val Val Asp Lys Pro Pro Lys Pro Ser Leu Ser 400 405
410 415 gct tgg cca agc act gtg ttc aag cta gga
aag gcc atc acc ctt cag 2437 Ala Trp Pro Ser Thr Val Phe Lys Leu Gly
Lys Ala Ile Thr Leu Gln 420 425
430 tgc cga gta tct cat cca gta ctg gaa ttt tct ctg gaa tgg gaa gaa
2485 Cys Arg Val Ser His Pro Val Leu Glu Phe Ser Leu Glu Trp Glu Glu
435 440 445 aga gaa aca ttc
caa aaa ttc tca gta aac gga gac ttc atc atc agt 2533 Arg Glu Thr Phe
Gln Lys Phe Ser Val Asn Gly Asp Phe Ile Ile Ser 450
455 460 aat gtt gac ggg aaa ggc aca ggg acc tac agt tgc
agc tat cgc gta 2581 Asn Val Asp Gly Lys Gly Thr Gly Thr Tyr Ser Cys
Ser Tyr Arg Val 465 470 475 gag aca
cat cct aac atc tgg tca cat cgc agt gag ccc ctg aag ctg 2629 Glu Thr
His Pro Asn Ile Trp Ser His Arg Ser Glu Pro Leu Lys Leu 480
485 490 495 atg ggg cca gca ggc tat ctc
acc tgg aat tac gtt ctg aat gaa gct 2677 Met Gly Pro Ala Gly Tyr Leu
Thr Trp Asn Tyr Val Leu Asn Glu Ala 500
505 510 atc agg ttg tct cta atc atg cag ctt gtt gcc ttg
ctg ttg gta gtg 2725 Ile Arg Leu Ser Leu Ile Met Gln Leu Val Ala Leu
Leu Leu Val Val 515 520 525
ctg tgg ata agg tgg aag tgt cgg aga ctc aga atc aga gaa gcc tgg 2773
Leu Trp Ile Arg Trp Lys Cys Arg Arg Leu Arg Ile Arg Glu Ala Trp
530 535 540 ttg ctg gga aca gct caa ggg
gtc acc atg ctc ttc ata gtc acg gcc 2821 Leu Leu Gly Thr Ala Gln Gly
Val Thr Met Leu Phe Ile Val Thr Ala 545 550
555 ctt ctc tgc tgt gga ctg tgc aat ggg gta ttg ata gaa gag act gaa
2869 Leu Leu Cys Cys Gly Leu Cys Asn Gly Val Leu Ile Glu Glu Thr Glu
560 565 570 575 ata gtc
atg cca acc cct aag cct gag ctg tgg gca gag acc aac ttt 2917 Ile Val
Met Pro Thr Pro Lys Pro Glu Leu Trp Ala Glu Thr Asn Phe
580 585 590 cct ctg gcc ccg tgg aag aac
tta acc ctc tgg tgc aga agc cct tct 2965 Pro Leu Ala Pro Trp Lys Asn
Leu Thr Leu Trp Cys Arg Ser Pro Ser 595 600
605 ggc tca act aag gag ttt gtg ttg ctg aag gat ggg acc ggg
tgg atc 3013 Gly Ser Thr Lys Glu Phe Val Leu Leu Lys Asp Gly Thr Gly
Trp Ile 610 615 620 gcc act cgc
ccg gcc tca gag cag gtc cgg gct gcc ttc ccc ctt ggc 3061 Ala Thr Arg
Pro Ala Ser Glu Gln Val Arg Ala Ala Phe Pro Leu Gly 625
630 635 gcc ctg acc cag agc cac acc ggg agc tac cac tgc
cat tca tgg gag 3109 Ala Leu Thr Gln Ser His Thr Gly Ser Tyr His Cys
His Ser Trp Glu 640 645 650
655 gag atg gct gta tcg gag ccc agt gag gca ctt gag ctg gtg ggg aca
3157 Glu Met Ala Val Ser Glu Pro Ser Glu Ala Leu Glu Leu Val Gly Thr
660 665 670 gac atc ctc ccc
aaa cct gtc att tct gct tcc ccc aca atc cgg ggc 3205 Asp Ile Leu Pro
Lys Pro Val Ile Ser Ala Ser Pro Thr Ile Arg Gly 675
680 685 cag gaa cta caa ctc cgg tgc aaa gga tgg ctg
gca ggc atg ggg ttt 3253 Gln Glu Leu Gln Leu Arg Cys Lys Gly Trp Leu
Ala Gly Met Gly Phe 690 695 700
gct ctg tat aag gag gga gag caa gaa cct gtc cag caa ctt ggt gct 3301
Ala Leu Tyr Lys Glu Gly Glu Gln Glu Pro Val Gln Gln Leu Gly Ala 705
710 715 gtt gga aga gaa gcc ttc ttt aca atc
cag aga atg gag gat aaa gac 3349 Val Gly Arg Glu Ala Phe Phe Thr Ile
Gln Arg Met Glu Asp Lys Asp 720 725 730
735 gaa ggc aat tac agc tgc cgc act cac act gaa aaa cgc ccc
ttc aag 3397 Glu Gly Asn Tyr Ser Cys Arg Thr His Thr Glu Lys Arg Pro
Phe Lys 740 745 750 tgg
tct gag ccc agt gag ccg ctg gag ctt gtc ata aaa gaa atg tac 3445 Trp
Ser Glu Pro Ser Glu Pro Leu Glu Leu Val Ile Lys Glu Met Tyr
755 760 765 cct aag ccc ttc ttc aag aca
tgg gcc agc cct gtg gtc acc cct ggt 3493 Pro Lys Pro Phe Phe Lys Thr
Trp Ala Ser Pro Val Val Thr Pro Gly 770 775
780 gcc cga gtg act ttc aat tgc tcc acc ccc cac cag cat atg agc
ttt 3541 Ala Arg Val Thr Phe Asn Cys Ser Thr Pro His Gln His Met Ser
Phe 785 790 795 att ctt tac aaa gat
gga agt gaa ata gca tcc agt gac agg tcc tgg 3589 Ile Leu Tyr Lys Asp
Gly Ser Glu Ile Ala Ser Ser Asp Arg Ser Trp 800 805
810 815 gca agt ccg ggg gcc agt gca gct cac ttt
cta atc att tcg gtg ggc 3637 Ala Ser Pro Gly Ala Ser Ala Ala His Phe
Leu Ile Ile Ser Val Gly 820 825
830 att ggt gat gga ggg aat tac agc tgc cga tat tat gac ttt tct atc
3685 Ile Gly Asp Gly Gly Asn Tyr Ser Cys Arg Tyr Tyr Asp Phe Ser Ile
835 840 845 tgg tct gag ccc
agc gac cct gtg gag ctc gtg gtg aca gaa ttc tac 3733 Trp Ser Glu Pro
Ser Asp Pro Val Glu Leu Val Val Thr Glu Phe Tyr 850
855 860 ccc aaa ccc act ctc ctg gca cag cca ggt cct gtg
gtg ttt cct ggg 3781 Pro Lys Pro Thr Leu Leu Ala Gln Pro Gly Pro Val
Val Phe Pro Gly 865 870 875 aag agt
gtg atc ctg cgc tgc caa ggg act ttc cag ggc atg agg ttc 3829 Lys Ser
Val Ile Leu Arg Cys Gln Gly Thr Phe Gln Gly Met Arg Phe 880
885 890 895 gcc ctc ttg cag gag gga gcc
cat gtt ccc tta cag ttt cgg agt gtc 3877 Ala Leu Leu Gln Glu Gly Ala
His Val Pro Leu Gln Phe Arg Ser Val 900
905 910 tca ggg aac tca gct gac ttc ctt ctc cac act gtt
gga gca gag gac 3925 Ser Gly Asn Ser Ala Asp Phe Leu Leu His Thr Val
Gly Ala Glu Asp 915 920 925
tct ggg aac tat agc tgt atc tac tat gag aca acc atg tca aac agg 3973
Ser Gly Asn Tyr Ser Cys Ile Tyr Tyr Glu Thr Thr Met Ser Asn Arg
930 935 940 ggg tca tat ctc agt atg ccc
ctt atg atc tgg gtg act gac aca ttc 4021 Gly Ser Tyr Leu Ser Met Pro
Leu Met Ile Trp Val Thr Asp Thr Phe 945 950
955 cct aag cca tgg ttg ttt gct gag ccc agt tct gtg gtt ccc atg ggg
4069 Pro Lys Pro Trp Leu Phe Ala Glu Pro Ser Ser Val Val Pro Met Gly
960 965 970 975 cag aat
gtt act ctc tgg tgc cga ggg ccg gtc cat gga gta gga tac 4117 Gln Asn
Val Thr Leu Trp Cys Arg Gly Pro Val His Gly Val Gly Tyr
980 985 990 att ctg cac aaa gaa gga gaa
gcc act tca atg cag ctc tgg gga tcc 4165 Ile Leu His Lys Glu Gly Glu
Ala Thr Ser Met Gln Leu Trp Gly Ser 995
1000 1005 acc agt aat gac ggg gca ttc ccc atc acc aat
ata tct ggt act 4210 Thr Ser Asn Asp Gly Ala Phe Pro Ile Thr Asn
Ile Ser Gly Thr 1010 1015 1020
agc atg ggg cgt tac agc tgc tgc tac cac cct gac tgg acc agt 4255
Ser Met Gly Arg Tyr Ser Cys Cys Tyr His Pro Asp Trp Thr Ser
1025 1030 1035 tct atc aag ata caa cct
agc aac acc ctg gaa ctc cta gtc aca 4300 Ser Ile Lys Ile Gln Pro
Ser Asn Thr Leu Glu Leu Leu Val Thr 1040 1045
1050 ggc tta ctc ccc aaa ccc agc cta tta gcc cag cct ggt
ccc atg 4345 Gly Leu Leu Pro Lys Pro Ser Leu Leu Ala Gln Pro Gly
Pro Met 1055 1060 1065 gtg gcc
cct ggc gaa aat atg act ctt cag tgt caa ggg gaa ctg 4390 Val Ala
Pro Gly Glu Asn Met Thr Leu Gln Cys Gln Gly Glu Leu 1070
1075 1080 cca gac tca aca ttt gtc ctg ttg aag
gag ggg gct cag gag cct 4435 Pro Asp Ser Thr Phe Val Leu Leu Lys
Glu Gly Ala Gln Glu Pro 1085 1090
1095 tta gag caa cag agg cca agt ggg tac agg gct gac ttc tgg atg
4480 Leu Glu Gln Gln Arg Pro Ser Gly Tyr Arg Ala Asp Phe Trp Met
1100 1105 1110 cca gca gtg aga ggt gaa
gac tct ggg atc tat agc tgt gtt tat 4525 Pro Ala Val Arg Gly Glu
Asp Ser Gly Ile Tyr Ser Cys Val Tyr 1115 1120
1125 tat ttg gac tct act ccc ttt gca gct tca aat cac agt
gac tcc 4570 Tyr Leu Asp Ser Thr Pro Phe Ala Ala Ser Asn His Ser
Asp Ser 1130 1135 1140 ctg gag
atc tgg gtg act gat aag ccc cct aaa ccc tct ctg tca 4615 Leu Glu
Ile Trp Val Thr Asp Lys Pro Pro Lys Pro Ser Leu Ser 1145
1150 1155 gcc tgg ccc agc acc atg ttc aag tta
ggg aag gac atc acc ctt 4660 Ala Trp Pro Ser Thr Met Phe Lys Leu
Gly Lys Asp Ile Thr Leu 1160 1165
1170 cag tgc cga gga ccc ctg cca ggt gtt gaa ttt gtc cta gaa cat
4705 Gln Cys Arg Gly Pro Leu Pro Gly Val Glu Phe Val Leu Glu His
1175 1180 1185 gat gga gaa gaa gca cct
cag cag ttt tca gag gat gga gac ttt 4750 Asp Gly Glu Glu Ala Pro
Gln Gln Phe Ser Glu Asp Gly Asp Phe 1190 1195
1200 gtc atc aac aac gta gaa gga aaa ggc att gga aac tac
agc tgc 4795 Val Ile Asn Asn Val Glu Gly Lys Gly Ile Gly Asn Tyr
Ser Cys 1205 1210 1215 agc tac
cgc ctc cag gcc tac cct gat atc tgg tca gag cct agt 4840 Ser Tyr
Arg Leu Gln Ala Tyr Pro Asp Ile Trp Ser Glu Pro Ser 1220
1225 1230 gat ccc ctg gag ctg gtg ggg gca gca
ggg cct gtt gct cag gag 4885 Asp Pro Leu Glu Leu Val Gly Ala Ala
Gly Pro Val Ala Gln Glu 1235 1240
1245 tgc act gta ggg aac att gtc cga agt agc cta atc gtg gtg gtt
4930 Cys Thr Val Gly Asn Ile Val Arg Ser Ser Leu Ile Val Val Val
1250 1255 1260 gtt gta gcc ttg ggg gta
gtg cta gcc ata gag tgg aag aag tgg 4975 Val Val Ala Leu Gly Val
Val Leu Ala Ile Glu Trp Lys Lys Trp 1265 1270
1275 cct cga ctg cga acc aga ggc tca gag aca gac gga aga
gac cag 5020 Pro Arg Leu Arg Thr Arg Gly Ser Glu Thr Asp Gly Arg
Asp Gln 1280 1285 1290 acc att
gcc ctt gaa gag tgt aac caa gaa gga gaa cca ggc acc 5065 Thr Ile
Ala Leu Glu Glu Cys Asn Gln Glu Gly Glu Pro Gly Thr 1295
1300 1305 cct gcc aat tct cct tca tca acc tct
cag aga atc tct gtg gaa 5110 Pro Ala Asn Ser Pro Ser Ser Thr Ser
Gln Arg Ile Ser Val Glu 1310 1315
1320 ctg ccc gtt cca ata taa taatctcctc ctttacaaga gctttcctct
5158 Leu Pro Val Pro Ile 1325 cctctctctt gctctcagag acctataaat
ccaaccagtt accctgcaag tcagccccat 5218 ctgctgttcc ttggtctcta atcacctgag
ctgggtaaag gggattctgg gagttgagag 5278 ctctgccagg gtgagatgtt tcctgaagag
aggttcccca cccctgtaac tcctcactgt 5338 actgatttac tggcgcatga aattctatta
aaaatgcatt cttctgaata aaaagagtat 5398 tcactattta acttc
5413 28 1327 PRT Homo sapiens 28 Met
Thr Leu Asp Arg Pro Gly Glu Gly Ala Thr Met Leu Lys Thr Phe 1
5 10 15 Thr Val Leu Leu Phe Cys Ile
Leu Met Asp Pro Gln Pro Glu Leu Trp 20 25
30 Ile Glu Ser Asn Tyr Pro Gln Ala Pro Trp Glu Asn Ile Thr
Leu Trp 35 40 45 Cys Arg Ser
Pro Ser Arg Ile Ser Ser Lys Phe Leu Leu Leu Lys Asp 50
55 60 Lys Thr Gln Met Thr Trp Ile Arg Pro Ser His Lys
Thr Phe Gln Val 65 70 75
80 Ser Phe Leu Ile Gly Ala Leu Thr Glu Ser Asn Ala Gly Leu Tyr Arg
85 90 95 Cys Cys Tyr Trp Lys
Glu Thr Gly Trp Ser Lys Pro Ser Lys Val Leu 100
105 110 Glu Leu Glu Ala Pro Gly Gln Leu Pro Lys Pro Ile
Phe Trp Ile Gln 115 120 125 Ala
Glu Thr Pro Ala Leu Pro Gly Cys Asn Val Asn Ile Leu Cys His 130
135 140 Gly Trp Leu Gln Asp Leu Val Phe Met Leu
Phe Lys Glu Gly Tyr Ala 145 150 155
160 Glu Pro Val Asp Tyr Gln Val Pro Thr Gly Thr Met Ala Ile Phe
Ser 165 170 175 Ile Asp
Asn Leu Thr Pro Glu Asp Glu Gly Val Tyr Ile Cys Arg Thr 180
185 190 His Ile Gln Met Leu Pro Thr Leu Trp
Ser Glu Pro Ser Asn Pro Leu 195 200
205 Lys Leu Val Val Ala Gly Leu Tyr Pro Lys Pro Thr Leu Thr Ala His
210 215 220 Pro Gly Pro Ile Met Ala Pro
Gly Glu Ser Leu Asn Leu Arg Cys Gln 225 230
235 240 Gly Pro Ile Tyr Gly Met Thr Phe Ala Leu Met Arg
Val Glu Asp Leu 245 250
255 Glu Lys Ser Phe Tyr His Lys Lys Thr Ile Lys Asn Glu Ala Asn Phe
260 265 270 Phe Phe Gln Ser Leu Lys
Ile Gln Asp Thr Gly His Tyr Leu Cys Phe 275 280
285 Tyr Tyr Asp Ala Ser Tyr Arg Gly Ser Leu Leu Ser Asp Val
Leu Lys 290 295 300 Ile Trp Val Thr
Asp Thr Phe Pro Lys Thr Trp Leu Leu Ala Arg Pro 305 310
315 320 Ser Ala Val Val Gln Met Gly Gln Asn
Val Ser Leu Arg Cys Arg Gly 325 330
335 Pro Val Asp Gly Val Gly Leu Ala Leu Tyr Lys Lys Gly Glu Asp
Lys 340 345 350 Pro Leu Gln
Phe Leu Asp Ala Thr Ser Ile Asp Asp Asn Thr Ser Phe 355
360 365 Phe Leu Asn Asn Val Thr Tyr Ser Asp Thr Gly
Ile Tyr Ser Cys His 370 375 380 Tyr
Leu Leu Thr Trp Lys Thr Ser Ile Arg Met Pro Ser His Asn Thr 385
390 395 400 Val Glu Leu Met Val Val
Asp Lys Pro Pro Lys Pro Ser Leu Ser Ala 405
410 415 Trp Pro Ser Thr Val Phe Lys Leu Gly Lys Ala Ile
Thr Leu Gln Cys 420 425 430
Arg Val Ser His Pro Val Leu Glu Phe Ser Leu Glu Trp Glu Glu Arg
435 440 445 Glu Thr Phe Gln Lys Phe Ser
Val Asn Gly Asp Phe Ile Ile Ser Asn 450 455
460 Val Asp Gly Lys Gly Thr Gly Thr Tyr Ser Cys Ser Tyr Arg Val Glu
465 470 475 480 Thr His
Pro Asn Ile Trp Ser His Arg Ser Glu Pro Leu Lys Leu Met
485 490 495 Gly Pro Ala Gly Tyr Leu Thr
Trp Asn Tyr Val Leu Asn Glu Ala Ile 500 505
510 Arg Leu Ser Leu Ile Met Gln Leu Val Ala Leu Leu Leu Val
Val Leu 515 520 525 Trp Ile Arg
Trp Lys Cys Arg Arg Leu Arg Ile Arg Glu Ala Trp Leu 530
535 540 Leu Gly Thr Ala Gln Gly Val Thr Met Leu Phe Ile
Val Thr Ala Leu 545 550 555
560 Leu Cys Cys Gly Leu Cys Asn Gly Val Leu Ile Glu Glu Thr Glu Ile
565 570 575 Val Met Pro Thr
Pro Lys Pro Glu Leu Trp Ala Glu Thr Asn Phe Pro 580
585 590 Leu Ala Pro Trp Lys Asn Leu Thr Leu Trp Cys
Arg Ser Pro Ser Gly 595 600 605
Ser Thr Lys Glu Phe Val Leu Leu Lys Asp Gly Thr Gly Trp Ile Ala 610
615 620 Thr Arg Pro Ala Ser Glu Gln Val Arg
Ala Ala Phe Pro Leu Gly Ala 625 630 635
640 Leu Thr Gln Ser His Thr Gly Ser Tyr His Cys His Ser Trp
Glu Glu 645 650 655 Met
Ala Val Ser Glu Pro Ser Glu Ala Leu Glu Leu Val Gly Thr Asp
660 665 670 Ile Leu Pro Lys Pro Val Ile
Ser Ala Ser Pro Thr Ile Arg Gly Gln 675 680
685 Glu Leu Gln Leu Arg Cys Lys Gly Trp Leu Ala Gly Met Gly Phe
Ala 690 695 700 Leu Tyr Lys Glu Gly
Glu Gln Glu Pro Val Gln Gln Leu Gly Ala Val 705 710
715 720 Gly Arg Glu Ala Phe Phe Thr Ile Gln Arg
Met Glu Asp Lys Asp Glu 725 730
735 Gly Asn Tyr Ser Cys Arg Thr His Thr Glu Lys Arg Pro Phe Lys Trp
740 745 750 Ser Glu Pro Ser
Glu Pro Leu Glu Leu Val Ile Lys Glu Met Tyr Pro 755
760 765 Lys Pro Phe Phe Lys Thr Trp Ala Ser Pro Val Val
Thr Pro Gly Ala 770 775 780 Arg Val
Thr Phe Asn Cys Ser Thr Pro His Gln His Met Ser Phe Ile 785
790 795 800 Leu Tyr Lys Asp Gly Ser Glu
Ile Ala Ser Ser Asp Arg Ser Trp Ala 805
810 815 Ser Pro Gly Ala Ser Ala Ala His Phe Leu Ile Ile
Ser Val Gly Ile 820 825 830
Gly Asp Gly Gly Asn Tyr Ser Cys Arg Tyr Tyr Asp Phe Ser Ile Trp
835 840 845 Ser Glu Pro Ser Asp Pro Val
Glu Leu Val Val Thr Glu Phe Tyr Pro 850 855
860 Lys Pro Thr Leu Leu Ala Gln Pro Gly Pro Val Val Phe Pro Gly Lys
865 870 875 880 Ser Val
Ile Leu Arg Cys Gln Gly Thr Phe Gln Gly Met Arg Phe Ala
885 890 895 Leu Leu Gln Glu Gly Ala His
Val Pro Leu Gln Phe Arg Ser Val Ser 900 905
910 Gly Asn Ser Ala Asp Phe Leu Leu His Thr Val Gly Ala Glu
Asp Ser 915 920 925 Gly Asn Tyr
Ser Cys Ile Tyr Tyr Glu Thr Thr Met Ser Asn Arg Gly 930
935 940 Ser Tyr Leu Ser Met Pro Leu Met Ile Trp Val Thr
Asp Thr Phe Pro 945 950 955
960 Lys Pro Trp Leu Phe Ala Glu Pro Ser Ser Val Val Pro Met Gly Gln
965 970 975 Asn Val Thr Leu
Trp Cys Arg Gly Pro Val His Gly Val Gly Tyr Ile 980
985 990 Leu His Lys Glu Gly Glu Ala Thr Ser Met
Gln Leu Trp Gly Ser Thr 995 1000
1005 Ser Asn Asp Gly Ala Phe Pro Ile Thr Asn Ile Ser Gly Thr Ser
1010 1015 1020 Met Gly Arg Tyr Ser Cys
Cys Tyr His Pro Asp Trp Thr Ser Ser 1025 1030
1035 Ile Lys Ile Gln Pro Ser Asn Thr Leu Glu Leu Leu Val Thr
Gly 1040 1045 1050 Leu Leu Pro Lys
Pro Ser Leu Leu Ala Gln Pro Gly Pro Met Val 1055
1060 1065 Ala Pro Gly Glu Asn Met Thr Leu Gln Cys Gln
Gly Glu Leu Pro 1070 1075 1080 Asp
Ser Thr Phe Val Leu Leu Lys Glu Gly Ala Gln Glu Pro Leu 1085
1090 1095 Glu Gln Gln Arg Pro Ser Gly Tyr Arg
Ala Asp Phe Trp Met Pro 1100 1105
1110 Ala Val Arg Gly Glu Asp Ser Gly Ile Tyr Ser Cys Val Tyr Tyr
1115 1120 1125 Leu Asp Ser Thr Pro Phe
Ala Ala Ser Asn His Ser Asp Ser Leu 1130 1135
1140 Glu Ile Trp Val Thr Asp Lys Pro Pro Lys Pro Ser Leu Ser
Ala 1145 1150 1155 Trp Pro Ser Thr
Met Phe Lys Leu Gly Lys Asp Ile Thr Leu Gln 1160
1165 1170 Cys Arg Gly Pro Leu Pro Gly Val Glu Phe Val
Leu Glu His Asp 1175 1180 1185 Gly
Glu Glu Ala Pro Gln Gln Phe Ser Glu Asp Gly Asp Phe Val 1190
1195 1200 Ile Asn Asn Val Glu Gly Lys Gly Ile
Gly Asn Tyr Ser Cys Ser 1205 1210
1215 Tyr Arg Leu Gln Ala Tyr Pro Asp Ile Trp Ser Glu Pro Ser Asp
1220 1225 1230 Pro Leu Glu Leu Val Gly
Ala Ala Gly Pro Val Ala Gln Glu Cys 1235 1240
1245 Thr Val Gly Asn Ile Val Arg Ser Ser Leu Ile Val Val Val
Val 1250 1255 1260 Val Ala Leu Gly
Val Val Leu Ala Ile Glu Trp Lys Lys Trp Pro 1265
1270 1275 Arg Leu Arg Thr Arg Gly Ser Glu Thr Asp Gly
Arg Asp Gln Thr 1280 1285 1290 Ile
Ala Leu Glu Glu Cys Asn Gln Glu Gly Glu Pro Gly Thr Pro 1295
1300 1305 Ala Asn Ser Pro Ser Ser Thr Ser Gln
Arg Ile Ser Val Glu Leu 1310 1315
1320 Pro Val Pro Ile 1325 29 2557 DNA Homo sapiens CDS (14)..(1471)
NLE1 mRNA 29 gtggggacgc agg atg gcg gca gca gtg ccg gac gag gcg gtg gcg
cgc 49 Met Ala Ala Ala Val Pro Asp Glu Ala Val Ala
Arg 1 5 10 gat gtg cag cgg
ttg cta gtg cag ttc cag gat gag ggc ggg cag ctg 97 Asp Val Gln Arg
Leu Leu Val Gln Phe Gln Asp Glu Gly Gly Gln Leu 15
20 25 ctg ggt tcc ccg ttc gac gtg ccc gtg gac atc acc
ccg gac agg ctg 145 Leu Gly Ser Pro Phe Asp Val Pro Val Asp Ile Thr
Pro Asp Arg Leu 30 35 40 cag ctc
gtg tgc aac gcg cta ctg gcc cag gag gat ccc ctg cca ctg 193 Gln Leu
Val Cys Asn Ala Leu Leu Ala Gln Glu Asp Pro Leu Pro Leu 45
50 55 60 gct ttc ttt gtc cac gat gct
gag atc gtc tcc tca ctg ggg aag acg 241 Ala Phe Phe Val His Asp Ala
Glu Ile Val Ser Ser Leu Gly Lys Thr 65
70 75 ttg gag tcc cag gca gtg gag aca gag aag gtc cta
gac atc atc tac 289 Leu Glu Ser Gln Ala Val Glu Thr Glu Lys Val Leu
Asp Ile Ile Tyr 80 85 90
cag cca cag gct atc ttc aga gtc cgg gct gtg act cgc tgc acc agc 337
Gln Pro Gln Ala Ile Phe Arg Val Arg Ala Val Thr Arg Cys Thr Ser
95 100 105 tcc ttg gag ggt cac agt gag
gca gtc att tct gtg gcc ttc agc cct 385 Ser Leu Glu Gly His Ser Glu
Ala Val Ile Ser Val Ala Phe Ser Pro 110 115
120 acg gga aag tac ctg gcc agt ggc tct gga gac acc acc gtg cgc ttc
433 Thr Gly Lys Tyr Leu Ala Ser Gly Ser Gly Asp Thr Thr Val Arg Phe
125 130 135 140 tgg gat
ctc agc aca gag aca cca cat ttc aca tgc aag gga cac aga 481 Trp Asp
Leu Ser Thr Glu Thr Pro His Phe Thr Cys Lys Gly His Arg
145 150 155 cac tgg gtc ctt agt ata tcc
tgg tct cca gat ggc agg aag ctg gcc 529 His Trp Val Leu Ser Ile Ser
Trp Ser Pro Asp Gly Arg Lys Leu Ala 160 165
170 tca ggc tgc aag aat ggc cag att ctc ctc tgg gac cca agc
aca ggg 577 Ser Gly Cys Lys Asn Gly Gln Ile Leu Leu Trp Asp Pro Ser
Thr Gly 175 180 185 aag cag gtg
ggc agg acc ctc gct ggc cac agc aag tgg atc aca ggc 625 Lys Gln Val
Gly Arg Thr Leu Ala Gly His Ser Lys Trp Ile Thr Gly 190
195 200 ctg agc tgg gag ccc ctc cat gcg aac cct gag tgc
cgc tat gtg gcc 673 Leu Ser Trp Glu Pro Leu His Ala Asn Pro Glu Cys
Arg Tyr Val Ala 205 210 215
220 agc agc tcc aag gat ggc agt gtg cgg atc tgg gac aca act gca ggc
721 Ser Ser Ser Lys Asp Gly Ser Val Arg Ile Trp Asp Thr Thr Ala Gly
225 230 235 cgc tgt gag cgc
atc ctc acc ggg cac acc cag tcg gtc acc tgt ctc 769 Arg Cys Glu Arg
Ile Leu Thr Gly His Thr Gln Ser Val Thr Cys Leu 240
245 250 cgg tgg gga ggg gac ggg ctt ctc tac tct gcc
tcc cag gac cgc acc 817 Arg Trp Gly Gly Asp Gly Leu Leu Tyr Ser Ala
Ser Gln Asp Arg Thr 255 260 265
atc aaa gtc tgg aga gct cat gac ggt gtg ctg tgc cgg act ctg caa 865
Ile Lys Val Trp Arg Ala His Asp Gly Val Leu Cys Arg Thr Leu Gln 270
275 280 ggc cac ggc cac tgg gtg aac acc atg
gcc ctc agc act gac tat gcc 913 Gly His Gly His Trp Val Asn Thr Met
Ala Leu Ser Thr Asp Tyr Ala 285 290 295
300 ctg cgc act ggg gcc ttt gaa cct gct gag gcc tca gtt aat
ccc caa 961 Leu Arg Thr Gly Ala Phe Glu Pro Ala Glu Ala Ser Val Asn
Pro Gln 305 310 315 gac
ctc caa gga tcc ttg cag gag ttg aag gag agg gct ctg agc cga 1009 Asp
Leu Gln Gly Ser Leu Gln Glu Leu Lys Glu Arg Ala Leu Ser Arg
320 325 330 tac aac ctc gtg cgg ggc cag
ggt cca gag agg ctg gtg tct ggc tcc 1057 Tyr Asn Leu Val Arg Gly Gln
Gly Pro Glu Arg Leu Val Ser Gly Ser 335 340
345 gac gac ttc acc tta ttc ctg tgg tcc cca gca gag gac aaa aag
cct 1105 Asp Asp Phe Thr Leu Phe Leu Trp Ser Pro Ala Glu Asp Lys Lys
Pro 350 355 360 ctc act cgg atg aca
gga cac caa gct ctc atc aac cag gtg ctc ttc 1153 Leu Thr Arg Met Thr
Gly His Gln Ala Leu Ile Asn Gln Val Leu Phe 365 370
375 380 tct cct gac tcc cgc atc gtg gct agt gcc
tcc ttt gac aag tcc atc 1201 Ser Pro Asp Ser Arg Ile Val Ala Ser Ala
Ser Phe Asp Lys Ser Ile 385 390
395 aag ctg tgg gat ggc agg acg ggc aag tac ctg gct tcc cta cgc ggc
1249 Lys Leu Trp Asp Gly Arg Thr Gly Lys Tyr Leu Ala Ser Leu Arg Gly
400 405 410 cac gtg gct gcc
gtg tac cag att gcg tgg tca gct gac agt cgg ctc 1297 His Val Ala Ala
Val Tyr Gln Ile Ala Trp Ser Ala Asp Ser Arg Leu 415
420 425 ctg gtc agc ggc agc agt gac agc aca ctg aag gtg
tgg gat gtg aag 1345 Leu Val Ser Gly Ser Ser Asp Ser Thr Leu Lys Val
Trp Asp Val Lys 430 435 440 gcc cag
aag ctg gcc atg gac ctg ccc ggc cac gcg gat gag gta tat 1393 Ala Gln
Lys Leu Ala Met Asp Leu Pro Gly His Ala Asp Glu Val Tyr 445
450 455 460 gct gtt gac tgg agt cca gat
ggc cag aga gtg gca agt ggt ggg aag 1441 Ala Val Asp Trp Ser Pro Asp
Gly Gln Arg Val Ala Ser Gly Gly Lys 465
470 475 gac aaa tgc ctc cgg ata tgg agg aga tga
gacggcccga agttctctct 1491 Asp Lys Cys Leu Arg Ile Trp Arg Arg
480 485 gacccccacc tcgactcggc ctctgccagc
tgccttccct gccagagaac aaaggctgag 1551 atggcagtgc acacaccctc cccaccagtg
gggacctgag aatgcgtgtg gcctgctgtc 1611 ctcgatagac cggaatgggg ttttcccaca
gatccccgcc tgtggcacac cccagagcca 1671 gaaatcgaag gtcacaggaa gttgtcactg
aacttggccc gtgtctgcta ctctgtacct 1731 tgctggtaca gacaggggtg gtgggcagcc
aggctctatg agtgggcccc tagtgtcagc 1791 tctgtacagg gtcagatccc aggttctatg
accaaataag taacttaagt tttgtgtgtt 1851 gggttctaat tccttgtcct agaatcccca
tgactcaatc aaggactgtg ctaaatgaga 1911 ttgtccagcc cccgcccttg cactggacta
cgccaaaacc acactgacca ggcacttgcc 1971 ttccctctct tcccccgtgt tggtaagaga
gaggccagtt gtgatagtgg ccaaggagaa 2031 tctagggctg tattgttgtc cactgcagta
ggcaccggcc acatgtgact gctggcatga 2091 aatagaagtg cagttcctcc atcgcactgg
gtaaggcctc cagtattgga cagcacacag 2151 aaaggttttc atcatcaaga gagttctgct
ggtcagccct gctccagggg atgcctctgc 2211 cttcgcatag cacactgctt gaggccctgc
caggcaccaa gcactgccct gggcccatgg 2271 gatagagcgg ggaaggtgat ggctcttcca
gaggattccc tcagatgggg aggcagcagt 2331 atgagctctg agcagaagtg ggtattgttg
atacagagga agttctttgc cacgagaact 2391 ttcaagcagt gaaaggaatt cccatcagga
ctcagacccc aggccgagat cttgccctga 2451 atgtaccctg cctctgcttt ctcctgcatc
ccatgctaag cagggtcatg gtctgaacta 2511 ctcagattgg atttccaaac catccttgta
taaactgctc agaact 2557 30 485 PRT Homo sapiens 30 Met
Ala Ala Ala Val Pro Asp Glu Ala Val Ala Arg Asp Val Gln Arg 1
5 10 15 Leu Leu Val Gln Phe Gln Asp
Glu Gly Gly Gln Leu Leu Gly Ser Pro 20 25
30 Phe Asp Val Pro Val Asp Ile Thr Pro Asp Arg Leu Gln Leu
Val Cys 35 40 45 Asn Ala Leu
Leu Ala Gln Glu Asp Pro Leu Pro Leu Ala Phe Phe Val 50
55 60 His Asp Ala Glu Ile Val Ser Ser Leu Gly Lys Thr
Leu Glu Ser Gln 65 70 75
80 Ala Val Glu Thr Glu Lys Val Leu Asp Ile Ile Tyr Gln Pro Gln Ala
85 90 95 Ile Phe Arg Val Arg
Ala Val Thr Arg Cys Thr Ser Ser Leu Glu Gly 100
105 110 His Ser Glu Ala Val Ile Ser Val Ala Phe Ser Pro
Thr Gly Lys Tyr 115 120 125 Leu
Ala Ser Gly Ser Gly Asp Thr Thr Val Arg Phe Trp Asp Leu Ser 130
135 140 Thr Glu Thr Pro His Phe Thr Cys Lys Gly
His Arg His Trp Val Leu 145 150 155
160 Ser Ile Ser Trp Ser Pro Asp Gly Arg Lys Leu Ala Ser Gly Cys
Lys 165 170 175 Asn Gly
Gln Ile Leu Leu Trp Asp Pro Ser Thr Gly Lys Gln Val Gly 180
185 190 Arg Thr Leu Ala Gly His Ser Lys Trp
Ile Thr Gly Leu Ser Trp Glu 195 200
205 Pro Leu His Ala Asn Pro Glu Cys Arg Tyr Val Ala Ser Ser Ser Lys
210 215 220 Asp Gly Ser Val Arg Ile Trp
Asp Thr Thr Ala Gly Arg Cys Glu Arg 225 230
235 240 Ile Leu Thr Gly His Thr Gln Ser Val Thr Cys Leu
Arg Trp Gly Gly 245 250
255 Asp Gly Leu Leu Tyr Ser Ala Ser Gln Asp Arg Thr Ile Lys Val Trp
260 265 270 Arg Ala His Asp Gly Val
Leu Cys Arg Thr Leu Gln Gly His Gly His 275 280
285 Trp Val Asn Thr Met Ala Leu Ser Thr Asp Tyr Ala Leu Arg
Thr Gly 290 295 300 Ala Phe Glu Pro
Ala Glu Ala Ser Val Asn Pro Gln Asp Leu Gln Gly 305 310
315 320 Ser Leu Gln Glu Leu Lys Glu Arg Ala
Leu Ser Arg Tyr Asn Leu Val 325 330
335 Arg Gly Gln Gly Pro Glu Arg Leu Val Ser Gly Ser Asp Asp Phe
Thr 340 345 350 Leu Phe Leu
Trp Ser Pro Ala Glu Asp Lys Lys Pro Leu Thr Arg Met 355
360 365 Thr Gly His Gln Ala Leu Ile Asn Gln Val Leu
Phe Ser Pro Asp Ser 370 375 380 Arg
Ile Val Ala Ser Ala Ser Phe Asp Lys Ser Ile Lys Leu Trp Asp 385
390 395 400 Gly Arg Thr Gly Lys Tyr
Leu Ala Ser Leu Arg Gly His Val Ala Ala 405
410 415 Val Tyr Gln Ile Ala Trp Ser Ala Asp Ser Arg Leu
Leu Val Ser Gly 420 425 430
Ser Ser Asp Ser Thr Leu Lys Val Trp Asp Val Lys Ala Gln Lys Leu
435 440 445 Ala Met Asp Leu Pro Gly His
Ala Asp Glu Val Tyr Ala Val Asp Trp 450 455
460 Ser Pro Asp Gly Gln Arg Val Ala Ser Gly Gly Lys Asp Lys Cys Leu
465 470 475 480 Arg Ile
Trp Arg Arg 485 31 704 DNA Homo sapiens CDS (23)..(676)
RPL10A mRNA 31 ggttagcgcg gcgtgagaag cc atg agc agc aaa gtc tct cgc gac
acc ctg 52 Met Ser Ser Lys Val Ser Arg Asp
Thr Leu 1 5 10
tac gag gcg gtg cgg gaa gtc ctg cac ggg aac cag cgc aag cgc cgc 100
Tyr Glu Ala Val Arg Glu Val Leu His Gly Asn Gln Arg Lys Arg Arg
15 20 25 aag ttc ctg gag acg gtg
gag ttg cag atc agc ttg aag aac tat gat 148 Lys Phe Leu Glu Thr Val
Glu Leu Gln Ile Ser Leu Lys Asn Tyr Asp 30
35 40 ccc cag aag gac aag cgc ttc tcg ggc acc gtc agg
ctt aag tcc act 196 Pro Gln Lys Asp Lys Arg Phe Ser Gly Thr Val Arg
Leu Lys Ser Thr 45 50 55 ccc
cgc cct aag ttc tct gtg tgt gtc ctg ggg gac cag cag cac tgt 244 Pro
Arg Pro Lys Phe Ser Val Cys Val Leu Gly Asp Gln Gln His Cys 60
65 70 gac gag gct aag gcc gtg gat atc ccc cac
atg gac atc gag gcg ctg 292 Asp Glu Ala Lys Ala Val Asp Ile Pro His
Met Asp Ile Glu Ala Leu 75 80 85
90 aaa aaa ctc aac aag aat aaa aaa ctg gtc aag aag ctg gcc aag
aag 340 Lys Lys Leu Asn Lys Asn Lys Lys Leu Val Lys Lys Leu Ala Lys
Lys 95 100 105 tat gat
gcg ttt ttg gcc tca gag tct ctg atc aag cag att cca cga 388 Tyr Asp
Ala Phe Leu Ala Ser Glu Ser Leu Ile Lys Gln Ile Pro Arg 110
115 120 atc ctc ggc cca ggt tta aat aag gca
gga aag ttc cct tcc ctg ctc 436 Ile Leu Gly Pro Gly Leu Asn Lys Ala
Gly Lys Phe Pro Ser Leu Leu 125 130
135 aca cac aac gaa aac atg gtg gcc aaa gtg gat gag gtg aag tcc aca
484 Thr His Asn Glu Asn Met Val Ala Lys Val Asp Glu Val Lys Ser Thr
140 145 150 atc aag ttc caa atg aag aag
gtg tta tgt ctg gct gta gct gtt ggt 532 Ile Lys Phe Gln Met Lys Lys
Val Leu Cys Leu Ala Val Ala Val Gly 155 160
165 170 cac gtg aag atg aca gac gat gag ctt gtg tat aac
att cac ctg gct 580 His Val Lys Met Thr Asp Asp Glu Leu Val Tyr Asn
Ile His Leu Ala 175 180
185 gtc aac ttc ttg gtg tca ttg ctc aag aaa aac tgg cag aat gtc cgg
628 Val Asn Phe Leu Val Ser Leu Leu Lys Lys Asn Trp Gln Asn Val Arg
190 195 200 gcc tta tat atc aag agc
acc atg ggc aag ccc cag cgc cta tat taa 676 Ala Leu Tyr Ile Lys Ser
Thr Met Gly Lys Pro Gln Arg Leu Tyr 205 210
215 ggcacatttg aataaattct attaccag
704 32 217 PRT Homo sapiens 32 Met Ser Ser Lys Val Ser Arg Asp
Thr Leu Tyr Glu Ala Val Arg Glu 1 5 10
15 Val Leu His Gly Asn Gln Arg Lys Arg Arg Lys Phe Leu Glu
Thr Val 20 25 30 Glu Leu
Gln Ile Ser Leu Lys Asn Tyr Asp Pro Gln Lys Asp Lys Arg 35
40 45 Phe Ser Gly Thr Val Arg Leu Lys Ser Thr
Pro Arg Pro Lys Phe Ser 50 55 60
Val Cys Val Leu Gly Asp Gln Gln His Cys Asp Glu Ala Lys Ala Val 65
70 75 80 Asp Ile Pro His Met
Asp Ile Glu Ala Leu Lys Lys Leu Asn Lys Asn 85
90 95 Lys Lys Leu Val Lys Lys Leu Ala Lys Lys Tyr
Asp Ala Phe Leu Ala 100 105
110 Ser Glu Ser Leu Ile Lys Gln Ile Pro Arg Ile Leu Gly Pro Gly Leu
115 120 125 Asn Lys Ala Gly Lys Phe Pro
Ser Leu Leu Thr His Asn Glu Asn Met 130 135
140 Val Ala Lys Val Asp Glu Val Lys Ser Thr Ile Lys Phe Gln Met Lys
145 150 155 160 Lys Val
Leu Cys Leu Ala Val Ala Val Gly His Val Lys Met Thr Asp
165 170 175 Asp Glu Leu Val Tyr Asn Ile
His Leu Ala Val Asn Phe Leu Val Ser 180 185
190 Leu Leu Lys Lys Asn Trp Gln Asn Val Arg Ala Leu Tyr Ile
Lys Ser 195 200 205 Thr Met Gly
Lys Pro Gln Arg Leu Tyr 210 215 33 1919 DNA Homo
sapiens CDS (75)..(1202) ACTG1 mRNA 33 gtctcagtcg ccgctgccag ctctcgcact
ctgttcttcc gccgctccgc cgtcgcgttt 60 ctctgccggt cgca atg gaa gaa gag
atc gcc gcg ctg gtc att gac aat 110 Met Glu Glu Glu
Ile Ala Ala Leu Val Ile Asp Asn 1 5
10 ggc tcc ggc atg tgc aaa gct ggt ttt gct ggg gac gac gct ccc
cga 158 Gly Ser Gly Met Cys Lys Ala Gly Phe Ala Gly Asp Asp Ala Pro
Arg 15 20 25 gcc gtg ttt cct
tcc atc gtc ggg cgc ccc aga cac cag ggc gtc atg 206 Ala Val Phe Pro
Ser Ile Val Gly Arg Pro Arg His Gln Gly Val Met 30
35 40 gtg ggc atg ggc cag aag gac tcc tac gtg ggc gac
gag gcc cag agc 254 Val Gly Met Gly Gln Lys Asp Ser Tyr Val Gly Asp
Glu Ala Gln Ser 45 50 55
60 aag cgt ggc atc ctg acc ctg aag tac ccc att gag cat ggc atc gtc
302 Lys Arg Gly Ile Leu Thr Leu Lys Tyr Pro Ile Glu His Gly Ile Val
65 70 75 acc aac tgg gac gac
atg gag aag atc tgg cac cac acc ttc tac aac 350 Thr Asn Trp Asp Asp
Met Glu Lys Ile Trp His His Thr Phe Tyr Asn 80
85 90 gag ctg cgc gtg gcc ccg gag gag cac cca gtg ctg
ctg acc gag gcc 398 Glu Leu Arg Val Ala Pro Glu Glu His Pro Val Leu
Leu Thr Glu Ala 95 100 105 ccc
ctg aac ccc aag gcc aac aga gag aag atg act cag att atg ttt 446 Pro
Leu Asn Pro Lys Ala Asn Arg Glu Lys Met Thr Gln Ile Met Phe 110
115 120 gag acc ttc aac acc ccg gcc atg tac gtg
gcc atc cag gcc gtg ctg 494 Glu Thr Phe Asn Thr Pro Ala Met Tyr Val
Ala Ile Gln Ala Val Leu 125 130 135
140 tcc ctc tac gcc tct ggg cgc acc act ggc att gtc atg gac tct
gga 542 Ser Leu Tyr Ala Ser Gly Arg Thr Thr Gly Ile Val Met Asp Ser
Gly 145 150 155 gac ggg
gtc acc cac acg gtg ccc atc tac gag ggc tac gcc ctc ccc 590 Asp Gly
Val Thr His Thr Val Pro Ile Tyr Glu Gly Tyr Ala Leu Pro 160
165 170 cac gcc atc ctg cgt ctg gac ctg gct
ggc cgg gac ctg acc gac tac 638 His Ala Ile Leu Arg Leu Asp Leu Ala
Gly Arg Asp Leu Thr Asp Tyr 175 180
185 ctc atg aag atc ctc act gag cga ggc tac agc ttc acc acc acg gcc
686 Leu Met Lys Ile Leu Thr Glu Arg Gly Tyr Ser Phe Thr Thr Thr Ala
190 195 200 gag cgg gaa atc gtg cgc gac
atc aag gag aag ctg tgc tac gtc gcc 734 Glu Arg Glu Ile Val Arg Asp
Ile Lys Glu Lys Leu Cys Tyr Val Ala 205 210
215 220 ctg gac ttc gag cag gag atg gcc acc gcc gca tcc
tcc tct tct ctg 782 Leu Asp Phe Glu Gln Glu Met Ala Thr Ala Ala Ser
Ser Ser Ser Leu 225 230
235 gag aag agc tac gag ctg ccc gat ggc cag gtc atc acc att ggc aat
830 Glu Lys Ser Tyr Glu Leu Pro Asp Gly Gln Val Ile Thr Ile Gly Asn
240 245 250 gag cgg ttc cgg tgt ccg
gag gcg ctg ttc cag cct tcc ttc ctg ggt 878 Glu Arg Phe Arg Cys Pro
Glu Ala Leu Phe Gln Pro Ser Phe Leu Gly 255 260
265 atg gaa tct tgc ggc atc cac gag acc acc ttc aac tcc atc
atg aag 926 Met Glu Ser Cys Gly Ile His Glu Thr Thr Phe Asn Ser Ile
Met Lys 270 275 280 tgt gac gtg gac
atc cgc aaa gac ctg tac gcc aac acg gtg ctg tcg 974 Cys Asp Val Asp
Ile Arg Lys Asp Leu Tyr Ala Asn Thr Val Leu Ser 285 290
295 300 ggc ggc acc acc atg tac ccg ggc att
gcc gac agg atg cag aag gag 1022 Gly Gly Thr Thr Met Tyr Pro Gly Ile
Ala Asp Arg Met Gln Lys Glu 305 310
315 atc acc gcc ctg gcg ccc agc acc atg aag atc aag atc atc gca
ccc 1070 Ile Thr Ala Leu Ala Pro Ser Thr Met Lys Ile Lys Ile Ile Ala
Pro 320 325 330 cca gag cgc
aag tac tcg gtg tgg atc ggt ggc tcc atc ctg gcc tca 1118 Pro Glu Arg
Lys Tyr Ser Val Trp Ile Gly Gly Ser Ile Leu Ala Ser 335
340 345 ctg tcc acc ttc cag cag atg tgg att agc aag
cag gag tac gac gag 1166 Leu Ser Thr Phe Gln Gln Met Trp Ile Ser Lys
Gln Glu Tyr Asp Glu 350 355 360 tcg
ggc ccc tcc atc gtc cac cgc aaa tgc ttc taa acggactcag 1212 Ser
Gly Pro Ser Ile Val His Arg Lys Cys Phe 365 370
375 cagatgcgta gcatttgctg catgggttaa ttgagaatag aaatttgccc
ctggcaaatg 1272 cacacacctc atgctagcct cacgaaactg gaataagcct tcgaaaagaa
attgtccttg 1332 aagcttgtat ctgatatcag cactggattg tagaacttgt tgctgatttt
gaccttgtat 1392 tgaagttaac tgttcccctt ggtatttgtt taataccctg tacatatctt
tgagttcaac 1452 ctttagtacg tgtggcttgg tcacttcgtg gctaaggtaa gaacgtgctt
gtggaagaca 1512 agtctgtggc ttggtgagtc tgtgtggcca gcagcctctg atctgtgcag
ggtattaacg 1572 tgtcagggct gagtgttctg ggatttctct agaggctggc aagaaccagt
tgttttgtct 1632 tgcgggtctg tcagggttgg aaagtccaag ccgtaggacc cagtttcctt
tcttagctga 1692 tgtctttggc cagaacaccg tgggctgtta cttgctttga gttggaagcg
gtttgcattt 1752 acgcctgtaa atgtattcat tcttaattta tgtaaggttt tttttgtacg
caattctcga 1812 ttctttgaag agatgacaac aaattttggt tttctactgt tatgtgagaa
cattaggccc 1872 cagcaacacg tcattgtgta aggaaaaata aaagtgctgc cgtaacc
1919 34 375 PRT Homo sapiens 34 Met Glu Glu Glu Ile Ala Ala
Leu Val Ile Asp Asn Gly Ser Gly Met 1 5
10 15 Cys Lys Ala Gly Phe Ala Gly Asp Asp Ala Pro Arg
Ala Val Phe Pro 20 25 30
Ser Ile Val Gly Arg Pro Arg His Gln Gly Val Met Val Gly Met Gly
35 40 45 Gln Lys Asp Ser Tyr Val Gly
Asp Glu Ala Gln Ser Lys Arg Gly Ile 50 55
60 Leu Thr Leu Lys Tyr Pro Ile Glu His Gly Ile Val Thr Asn Trp Asp
65 70 75 80 Asp Met
Glu Lys Ile Trp His His Thr Phe Tyr Asn Glu Leu Arg Val
85 90 95 Ala Pro Glu Glu His Pro Val
Leu Leu Thr Glu Ala Pro Leu Asn Pro 100 105
110 Lys Ala Asn Arg Glu Lys Met Thr Gln Ile Met Phe Glu Thr
Phe Asn 115 120 125 Thr Pro Ala
Met Tyr Val Ala Ile Gln Ala Val Leu Ser Leu Tyr Ala 130
135 140 Ser Gly Arg Thr Thr Gly Ile Val Met Asp Ser Gly
Asp Gly Val Thr 145 150 155
160 His Thr Val Pro Ile Tyr Glu Gly Tyr Ala Leu Pro His Ala Ile Leu
165 170 175 Arg Leu Asp Leu
Ala Gly Arg Asp Leu Thr Asp Tyr Leu Met Lys Ile 180
185 190 Leu Thr Glu Arg Gly Tyr Ser Phe Thr Thr Thr
Ala Glu Arg Glu Ile 195 200 205
Val Arg Asp Ile Lys Glu Lys Leu Cys Tyr Val Ala Leu Asp Phe Glu 210
215 220 Gln Glu Met Ala Thr Ala Ala Ser Ser
Ser Ser Leu Glu Lys Ser Tyr 225 230 235
240 Glu Leu Pro Asp Gly Gln Val Ile Thr Ile Gly Asn Glu Arg
Phe Arg 245 250 255 Cys
Pro Glu Ala Leu Phe Gln Pro Ser Phe Leu Gly Met Glu Ser Cys
260 265 270 Gly Ile His Glu Thr Thr Phe
Asn Ser Ile Met Lys Cys Asp Val Asp 275 280
285 Ile Arg Lys Asp Leu Tyr Ala Asn Thr Val Leu Ser Gly Gly Thr
Thr 290 295 300 Met Tyr Pro Gly Ile
Ala Asp Arg Met Gln Lys Glu Ile Thr Ala Leu 305 310
315 320 Ala Pro Ser Thr Met Lys Ile Lys Ile Ile
Ala Pro Pro Glu Arg Lys 325 330
335 Tyr Ser Val Trp Ile Gly Gly Ser Ile Leu Ala Ser Leu Ser Thr Phe
340 345 350 Gln Gln Met Trp
Ile Ser Lys Gln Glu Tyr Asp Glu Ser Gly Pro Ser 355
360 365 Ile Val His Arg Lys Cys Phe 370
375 35 3528 DNA Homo sapiens CDS (64)..(1452) EEF1A1 mRNA 35
ctttttcgca acgggtttgc cgccagaaca caggtgtcgt gaaaactacc cctaaaagcc 60
aaa atg gga aag gaa aag act cat atc aac att gtc gtc att gga cac 108
Met Gly Lys Glu Lys Thr His Ile Asn Ile Val Val Ile Gly His 1
5 10 15 gta gat tcg ggc aag
tcc acc act act ggc cat ctg atc tat aaa tgc 156 Val Asp Ser Gly Lys
Ser Thr Thr Thr Gly His Leu Ile Tyr Lys Cys 20
25 30 ggt ggc atc gac aaa aga acc att gaa aaa ttt
gag aag gag gct gct 204 Gly Gly Ile Asp Lys Arg Thr Ile Glu Lys Phe
Glu Lys Glu Ala Ala 35 40
45 gag atg gga aag ggc tcc ttc aag tat gcc tgg gtc ttg gat aaa ctg
252 Glu Met Gly Lys Gly Ser Phe Lys Tyr Ala Trp Val Leu Asp Lys Leu
50 55 60 aaa gct gag cgt gaa cgt ggt
atc acc att gat atc tcc ttg tgg aaa 300 Lys Ala Glu Arg Glu Arg Gly
Ile Thr Ile Asp Ile Ser Leu Trp Lys 65 70
75 ttt gag acc agc aag tac tat gtg act atc att gat gcc cca gga cac
348 Phe Glu Thr Ser Lys Tyr Tyr Val Thr Ile Ile Asp Ala Pro Gly His
80 85 90 95 aga gac
ttt atc aaa aac atg att aca ggg aca tct cag gct gac tgt 396 Arg Asp
Phe Ile Lys Asn Met Ile Thr Gly Thr Ser Gln Ala Asp Cys
100 105 110 gct gtc ctg att gtt gct gct
ggt gtt ggt gaa ttt gaa gct ggt atc 444 Ala Val Leu Ile Val Ala Ala
Gly Val Gly Glu Phe Glu Ala Gly Ile 115 120
125 tcc aag aat ggg cag acc cga gag cat gcc ctt ctg gct tac
aca ctg 492 Ser Lys Asn Gly Gln Thr Arg Glu His Ala Leu Leu Ala Tyr
Thr Leu 130 135 140 ggt gtg aaa
caa cta att gtc ggt gtt aac aaa atg gat tcc act gag 540 Gly Val Lys
Gln Leu Ile Val Gly Val Asn Lys Met Asp Ser Thr Glu 145
150 155 cca ccc tac agc cag aag aga tat gag gaa att gtt
aag gaa gtc agc 588 Pro Pro Tyr Ser Gln Lys Arg Tyr Glu Glu Ile Val
Lys Glu Val Ser 160 165 170
175 act tac att aag aaa att ggc tac aac ccc gac aca gta gca ttt gtg
636 Thr Tyr Ile Lys Lys Ile Gly Tyr Asn Pro Asp Thr Val Ala Phe Val
180 185 190 cca att tct ggt
tgg aat ggt gac aac atg ctg gag cca agt gct aac 684 Pro Ile Ser Gly
Trp Asn Gly Asp Asn Met Leu Glu Pro Ser Ala Asn 195
200 205 atg cct tgg ttc aag gga tgg aaa gtc acc cgt
aag gat ggc aat gcc 732 Met Pro Trp Phe Lys Gly Trp Lys Val Thr Arg
Lys Asp Gly Asn Ala 210 215 220
agt gga acc acg ctg ctt gag gct ctg gac tgc atc cta cca cca act 780
Ser Gly Thr Thr Leu Leu Glu Ala Leu Asp Cys Ile Leu Pro Pro Thr 225
230 235 cgt cca act gac aag ccc ttg cgc ctg
cct ctc cag gat gtc tac aaa 828 Arg Pro Thr Asp Lys Pro Leu Arg Leu
Pro Leu Gln Asp Val Tyr Lys 240 245 250
255 att ggt ggt att ggt act gtt cct gtt ggc cga gtg gag act
ggt gtt 876 Ile Gly Gly Ile Gly Thr Val Pro Val Gly Arg Val Glu Thr
Gly Val 260 265 270 ctc
aaa ccc ggt atg gtg gtc acc ttt gct cca gtc aac gtt aca acg 924 Leu
Lys Pro Gly Met Val Val Thr Phe Ala Pro Val Asn Val Thr Thr
275 280 285 gaa gta aaa tct gtc gaa atg
cac cat gaa gct ttg agt gaa gct ctt 972 Glu Val Lys Ser Val Glu Met
His His Glu Ala Leu Ser Glu Ala Leu 290 295
300 cct ggg gac aat gtg ggc ttc aat gtc aag aat gtg tct gtc aag
gat 1020 Pro Gly Asp Asn Val Gly Phe Asn Val Lys Asn Val Ser Val Lys
Asp 305 310 315 gtt cgt cgt ggc aac
gtt gct ggt gac agc aaa aat gac cca cca atg 1068 Val Arg Arg Gly Asn
Val Ala Gly Asp Ser Lys Asn Asp Pro Pro Met 320 325
330 335 gaa gca gct ggc ttc act gct cag gtg att
atc ctg aac cat cca ggc 1116 Glu Ala Ala Gly Phe Thr Ala Gln Val Ile
Ile Leu Asn His Pro Gly 340 345
350 caa ata agc gcc ggc tat gcc cct gta ttg gat tgc cac acg gct cac
1164 Gln Ile Ser Ala Gly Tyr Ala Pro Val Leu Asp Cys His Thr Ala His
355 360 365 att gca tgc aag
ttt gct gag ctg aag gaa aag att gat cgc cgt tct 1212 Ile Ala Cys Lys
Phe Ala Glu Leu Lys Glu Lys Ile Asp Arg Arg Ser 370
375 380 ggt aaa aag ctg gaa gat ggc cct aaa ttc ttg aag
tct ggt gat gct 1260 Gly Lys Lys Leu Glu Asp Gly Pro Lys Phe Leu Lys
Ser Gly Asp Ala 385 390 395 gcc att
gtt gat atg gtt cct ggc aag ccc atg tgt gtt gag agc ttc 1308 Ala Ile
Val Asp Met Val Pro Gly Lys Pro Met Cys Val Glu Ser Phe 400
405 410 415 tca gac tat cca cct ttg ggt
cgc ttt gct gtt cgt gat atg aga cag 1356 Ser Asp Tyr Pro Pro Leu Gly
Arg Phe Ala Val Arg Asp Met Arg Gln 420
425 430 aca gtt gcg gtg ggt gtc atc aaa gca gtg gac aag
aag gct gct gga 1404 Thr Val Ala Val Gly Val Ile Lys Ala Val Asp Lys
Lys Ala Ala Gly 435 440 445
gct ggc aag gtc acc aag tct gcc cag aaa gct cag aag gct aaa tga 1452
Ala Gly Lys Val Thr Lys Ser Ala Gln Lys Ala Gln Lys Ala Lys 450
455 460 atattatccc taatacctgc caccccactc
ttaatcagtg gtggaagaac ggtctcagaa 1512 ctgtttgttt caattggcca tttaagttta
gtagtaaaag actggttaat gataacaatg 1572 catcgtaaaa ccttcagaag gaaaggagaa
tgttttgtgg accactttgg ttttcttttt 1632 tgcgtgtggc agttttaagt tattagtttt
taaaatcagt actttttaat ggaaacaact 1692 tgaccaaaaa tttgtcacag aattttgaga
cccattaaaa aagttaaatg agaaacctgt 1752 gtgttccttt ggtcaacacc gagacattta
ggtgaaagac atctaattct ggttttacga 1812 atctggaaac ttcttgaaaa tgtaattctt
gagttaacac ttctgggtgg agaatagggt 1872 tgttttcccc ccacataatt ggaaggggaa
ggaatatcat ttaaagctat gggagggttg 1932 ctttgattac aacactggag agaaatgcag
catgttgctg attgcctgtc actaaaacag 1992 gccaaaaact gagtccttgt gttgcataga
aagcttcatg ttgctaaacc aatgttaagt 2052 gaatctttgg aaacaaaatg tttccaaatt
actgggatgt gcatgttgaa acgtgggtta 2112 aaatgactgg gcagtgaaag ttgactattt
gccatgacat aagaaataag tgtagtggct 2172 agtgtacacc ctatgagtgg aagggtccat
tttgaagtca gtggagtaag ctttatgcca 2232 gtttgatggt ttcacaagtt ctattgagtg
ctattcagaa taggaacaag gttctaatag 2292 aaaaagatgg caatttgaag tagctataaa
attagactaa tctacattgc ttttctcctg 2352 cagagtctaa taccttttat gctttgataa
ttagcagttt gtctacttgg tcactaggaa 2412 tgaaactaca tggtaatagg cttaacaggt
gtaatagccc acttactcct gaatctttaa 2472 gcatttgtgc atttgaaaaa tgcttttcgc
gatcttcctg ctgggattac aggcatgagc 2532 cactgtgcct gacctcccat atgtaaaagt
gtctaaaggt ttttttttgg ttataaaagg 2592 aaaatttttg cttaagtttg aaggataggt
aaaattaaag gacatgcttt ctgtttgtgt 2652 gatggttttt aaaaattttt tttaagatgg
agttcttgtt gcccaggcta gaatgcaatg 2712 gcaaaatctc actgcaatct cctcctcctg
ggttcaagca attctcctac ttcagcctcc 2772 caagtagctg ggattacagg catgtgctaa
tttggtgttt ttaatagaga tgaggttttt 2832 ccatgttggt caggctggtc tcaaactcct
gaccttaggt gatcgcctcg gcctcctaaa 2892 gtgctggaat tacaggcatg agccaccatg
cctggccagg acatgtgttc ttaaggacat 2952 gctaagcagg agttaaagca gcccaagaga
taaggcctct taaagtgact ggcaatgtgt 3012 attgctcaag attcaaaggt acttgaattg
gccatagaca agtctgtaat gaagtgttat 3072 cgttttccct catctgagtc tgaattagat
aaaatgcctt cccatcagcc agtgctctga 3132 ggtatcaagt ctaaattgaa ctagagattt
ttgtccttag tttctttgct atctaatgtt 3192 tacacaagta aatagtctaa gatttgctgg
atgacagaaa aaacaggtaa ggcctttaat 3252 agatggccaa tagatgccct gataatgaaa
gttgacacct gtaagattta ccagtagaga 3312 attcttgaca tgcaaggaag caagatttaa
ctgaaaaatt gttcccactg gaagcaggaa 3372 tgagtcagtt tacttgcata tactgagatt
gagattaact tcctgtgaaa cccagtgtct 3432 tagacaactg tggcttgagc accacctgct
ggtattcatt acaaacttgc tcactacaat 3492 aaatgaattt taagctttaa aaaaaaaaaa
aaaaaa 3528 36 462 PRT Homo sapiens 36 Met
Gly Lys Glu Lys Thr His Ile Asn Ile Val Val Ile Gly His Val 1
5 10 15 Asp Ser Gly Lys Ser Thr Thr
Thr Gly His Leu Ile Tyr Lys Cys Gly 20 25
30 Gly Ile Asp Lys Arg Thr Ile Glu Lys Phe Glu Lys Glu Ala
Ala Glu 35 40 45 Met Gly Lys
Gly Ser Phe Lys Tyr Ala Trp Val Leu Asp Lys Leu Lys 50
55 60 Ala Glu Arg Glu Arg Gly Ile Thr Ile Asp Ile Ser
Leu Trp Lys Phe 65 70 75
80 Glu Thr Ser Lys Tyr Tyr Val Thr Ile Ile Asp Ala Pro Gly His Arg
85 90 95 Asp Phe Ile Lys Asn
Met Ile Thr Gly Thr Ser Gln Ala Asp Cys Ala 100
105 110 Val Leu Ile Val Ala Ala Gly Val Gly Glu Phe Glu
Ala Gly Ile Ser 115 120 125 Lys
Asn Gly Gln Thr Arg Glu His Ala Leu Leu Ala Tyr Thr Leu Gly 130
135 140 Val Lys Gln Leu Ile Val Gly Val Asn Lys
Met Asp Ser Thr Glu Pro 145 150 155
160 Pro Tyr Ser Gln Lys Arg Tyr Glu Glu Ile Val Lys Glu Val Ser
Thr 165 170 175 Tyr Ile
Lys Lys Ile Gly Tyr Asn Pro Asp Thr Val Ala Phe Val Pro 180
185 190 Ile Ser Gly Trp Asn Gly Asp Asn Met
Leu Glu Pro Ser Ala Asn Met 195 200
205 Pro Trp Phe Lys Gly Trp Lys Val Thr Arg Lys Asp Gly Asn Ala Ser
210 215 220 Gly Thr Thr Leu Leu Glu Ala
Leu Asp Cys Ile Leu Pro Pro Thr Arg 225 230
235 240 Pro Thr Asp Lys Pro Leu Arg Leu Pro Leu Gln Asp
Val Tyr Lys Ile 245 250
255 Gly Gly Ile Gly Thr Val Pro Val Gly Arg Val Glu Thr Gly Val Leu
260 265 270 Lys Pro Gly Met Val Val
Thr Phe Ala Pro Val Asn Val Thr Thr Glu 275 280
285 Val Lys Ser Val Glu Met His His Glu Ala Leu Ser Glu Ala
Leu Pro 290 295 300 Gly Asp Asn Val
Gly Phe Asn Val Lys Asn Val Ser Val Lys Asp Val 305 310
315 320 Arg Arg Gly Asn Val Ala Gly Asp Ser
Lys Asn Asp Pro Pro Met Glu 325 330
335 Ala Ala Gly Phe Thr Ala Gln Val Ile Ile Leu Asn His Pro Gly
Gln 340 345 350 Ile Ser Ala
Gly Tyr Ala Pro Val Leu Asp Cys His Thr Ala His Ile 355
360 365 Ala Cys Lys Phe Ala Glu Leu Lys Glu Lys Ile
Asp Arg Arg Ser Gly 370 375 380 Lys
Lys Leu Glu Asp Gly Pro Lys Phe Leu Lys Ser Gly Asp Ala Ala 385
390 395 400 Ile Val Asp Met Val Pro
Gly Lys Pro Met Cys Val Glu Ser Phe Ser 405
410 415 Asp Tyr Pro Pro Leu Gly Arg Phe Ala Val Arg Asp
Met Arg Gln Thr 420 425 430
Val Ala Val Gly Val Ile Lys Ala Val Asp Lys Lys Ala Ala Gly Ala
435 440 445 Gly Lys Val Thr Lys Ser Ala
Gln Lys Ala Gln Lys Ala Lys 450 455
460 37 3606 DNA Homo sapiens CDS (98)..(2251) PNN mRNA 37 attggctgag
cccggctgtc agtcctttcg cgcctcggcg gcgcggcata gcccggctcg 60 gcctgtaaag
cagtctcaag cctgccgcag ggagaag atg gcg gtc gcc gtg aga 115
Met Ala Val Ala Val Arg
1 5 act ttg cag gaa cag ctg gaa aag gcc
aaa gag agt ctt aag aac gtg 163 Thr Leu Gln Glu Gln Leu Glu Lys Ala
Lys Glu Ser Leu Lys Asn Val 10 15
20 gat gag aac att cgc aag ctc acc ggg cgg gat ccg aat gac gtg agg
211 Asp Glu Asn Ile Arg Lys Leu Thr Gly Arg Asp Pro Asn Asp Val Arg
25 30 35 ccc atc caa gcc aga ttg
ctg gcc ctt tct ggt cct ggt gga ggt aga 259 Pro Ile Gln Ala Arg Leu
Leu Ala Leu Ser Gly Pro Gly Gly Gly Arg 40 45
50 gga cgt ggt agt tta tta ctg agg cgt gga ttc tca gat agt gga
gga 307 Gly Arg Gly Ser Leu Leu Leu Arg Arg Gly Phe Ser Asp Ser Gly
Gly 55 60 65 70 gga
ccc cca gcc aaa cag aga gac ctt gaa ggg gca gtc agt agg ctg 355 Gly
Pro Pro Ala Lys Gln Arg Asp Leu Glu Gly Ala Val Ser Arg Leu
75 80 85 ggc ggg gag cgt cgg acc aga
aga gaa tca cgc cag gaa agc gac ccg 403 Gly Gly Glu Arg Arg Thr Arg
Arg Glu Ser Arg Gln Glu Ser Asp Pro 90 95
100 gag gat gat gat gtt aaa aag cca gca ttg cag tct tca gtt
gta gct 451 Glu Asp Asp Asp Val Lys Lys Pro Ala Leu Gln Ser Ser Val
Val Ala 105 110 115 acc tcc aaa
gag cgc aca cgt aga gac ctt atc cag gat caa aat atg 499 Thr Ser Lys
Glu Arg Thr Arg Arg Asp Leu Ile Gln Asp Gln Asn Met 120
125 130 gat gaa aag gga aag caa agg aac cgg cga ata ttt
ggc ttg ttg atg 547 Asp Glu Lys Gly Lys Gln Arg Asn Arg Arg Ile Phe
Gly Leu Leu Met 135 140 145
150 ggt acc ctt caa aaa ttt aaa caa gaa tcc act gtt gct act gaa agg
595 Gly Thr Leu Gln Lys Phe Lys Gln Glu Ser Thr Val Ala Thr Glu Arg
155 160 165 caa aag cgg cgc
cag gaa att gaa caa aaa ctt gaa gtt cag gca gaa 643 Gln Lys Arg Arg
Gln Glu Ile Glu Gln Lys Leu Glu Val Gln Ala Glu 170
175 180 gaa gag aga aag cag gtt gaa aat gaa agg aga
gaa ctg ttt gaa gag 691 Glu Glu Arg Lys Gln Val Glu Asn Glu Arg Arg
Glu Leu Phe Glu Glu 185 190 195
agg cgt gct aaa cag aca gaa ctg cgg ctt ttg gaa cag aaa gtt gag 739
Arg Arg Ala Lys Gln Thr Glu Leu Arg Leu Leu Glu Gln Lys Val Glu 200
205 210 ctt gcg cag ctg caa gaa gaa tgg aat
gaa cat aat gcc aaa ata att 787 Leu Ala Gln Leu Gln Glu Glu Trp Asn
Glu His Asn Ala Lys Ile Ile 215 220 225
230 aaa tat ata aga act aag aca aag ccc cat ttg ttt tat att
cct gga 835 Lys Tyr Ile Arg Thr Lys Thr Lys Pro His Leu Phe Tyr Ile
Pro Gly 235 240 245 aga
atg tgt cca gct acc caa aaa cta ata gaa gag tca cag aga aaa 883 Arg
Met Cys Pro Ala Thr Gln Lys Leu Ile Glu Glu Ser Gln Arg Lys
250 255 260 atg aac gct tta ttt gaa ggt
aga cgc atc gaa ttt gca gaa caa ata 931 Met Asn Ala Leu Phe Glu Gly
Arg Arg Ile Glu Phe Ala Glu Gln Ile 265 270
275 aat aaa atg gag gct agg cct aga aga caa tca atg aag gaa aaa
gag 979 Asn Lys Met Glu Ala Arg Pro Arg Arg Gln Ser Met Lys Glu Lys
Glu 280 285 290 cat cag gtg gtg cgt
aat gaa gaa cag aag gcg gaa caa gaa gag ggt 1027 His Gln Val Val Arg
Asn Glu Glu Gln Lys Ala Glu Gln Glu Glu Gly 295 300
305 310 aag gtg gct cag cga gag gaa gag ttg gag
gag aca ggt aat cag cac 1075 Lys Val Ala Gln Arg Glu Glu Glu Leu Glu
Glu Thr Gly Asn Gln His 315 320
325 aat gat gta gaa ata gag gaa gca gga gag gaa gag gaa aag gaa ata
1123 Asn Asp Val Glu Ile Glu Glu Ala Gly Glu Glu Glu Glu Lys Glu Ile
330 335 340 gcg att gtt cat
agt gat gca gag aaa gaa cag gag gag gaa gaa caa 1171 Ala Ile Val His
Ser Asp Ala Glu Lys Glu Gln Glu Glu Glu Glu Gln 345
350 355 aaa cag gaa atg gag gtt aag atg gag gag gaa act
gag gta agg gaa 1219 Lys Gln Glu Met Glu Val Lys Met Glu Glu Glu Thr
Glu Val Arg Glu 360 365 370 agt gag
aag cag cag gat agt cag cct gaa gaa gtt atg gat gtg cta 1267 Ser Glu
Lys Gln Gln Asp Ser Gln Pro Glu Glu Val Met Asp Val Leu 375
380 385 390 gag atg gtt gag aat gtc aaa
cat gta att gct gac cag gag gta atg 1315 Glu Met Val Glu Asn Val Lys
His Val Ile Ala Asp Gln Glu Val Met 395
400 405 gaa act aat cga gtt gaa agt gta gaa cct tca gaa
aat gaa gct agc 1363 Glu Thr Asn Arg Val Glu Ser Val Glu Pro Ser Glu
Asn Glu Ala Ser 410 415 420
aaa gaa ttg gaa cca gaa atg gaa ttt gaa att gag cca gat aaa gaa 1411
Lys Glu Leu Glu Pro Glu Met Glu Phe Glu Ile Glu Pro Asp Lys Glu
425 430 435 tgt aaa tcc ctt tct cct ggg
aaa gag aat gtc agt gct tta gac atg 1459 Cys Lys Ser Leu Ser Pro Gly
Lys Glu Asn Val Ser Ala Leu Asp Met 440 445
450 gaa aag gag tct gag gaa aaa gaa gaa aaa gaa tct gag ccc caa cct
1507 Glu Lys Glu Ser Glu Glu Lys Glu Glu Lys Glu Ser Glu Pro Gln Pro
455 460 465 470 gag cct
gtg gct caa cct cag cct cag tct cag ccc cag ctt cag ctt 1555 Glu Pro
Val Ala Gln Pro Gln Pro Gln Ser Gln Pro Gln Leu Gln Leu
475 480 485 caa tcc cag tcc caa cca gta
ctc cag tcc cag cct ccc tct cag cct 1603 Gln Ser Gln Ser Gln Pro Val
Leu Gln Ser Gln Pro Pro Ser Gln Pro 490 495
500 gag gat ttg tca tta gct gtt tta cag cca aca ccc caa gtt
act cag 1651 Glu Asp Leu Ser Leu Ala Val Leu Gln Pro Thr Pro Gln Val
Thr Gln 505 510 515 gag caa ggg
cat tta cta cct gag agg aag gat ttt cct gta gag tct 1699 Glu Gln Gly
His Leu Leu Pro Glu Arg Lys Asp Phe Pro Val Glu Ser 520
525 530 gta aaa ctc act gag gta cca gta gag cca gtc ttg
aca gta cat cca 1747 Val Lys Leu Thr Glu Val Pro Val Glu Pro Val Leu
Thr Val His Pro 535 540 545
550 gag agc aag agc aaa acc aaa act agg agc aga agt aga ggt cga gct
1795 Glu Ser Lys Ser Lys Thr Lys Thr Arg Ser Arg Ser Arg Gly Arg Ala
555 560 565 aga aat aaa aca
agc aag agt aga agt cga agc agt agc agt agc agt 1843 Arg Asn Lys Thr
Ser Lys Ser Arg Ser Arg Ser Ser Ser Ser Ser Ser 570
575 580 tct agt agc agt tca acc agt agc agc agt gga
agt agt tcc agc agt 1891 Ser Ser Ser Ser Ser Thr Ser Ser Ser Ser Gly
Ser Ser Ser Ser Ser 585 590 595
gga agt agt agc agt cgc agt agt tcc agt agc agc tcc agt aca agt 1939
Gly Ser Ser Ser Ser Arg Ser Ser Ser Ser Ser Ser Ser Ser Thr Ser 600
605 610 ggc agc agc agc aga gat agt agc agt
agc act agt agt agt agt gag 1987 Gly Ser Ser Ser Arg Asp Ser Ser Ser
Ser Thr Ser Ser Ser Ser Glu 615 620 625
630 agt aga agt cgg agt agg ggc cgg gga cat aat aga gat aga
aag cac 2035 Ser Arg Ser Arg Ser Arg Gly Arg Gly His Asn Arg Asp Arg
Lys His 635 640 645 aga
agg agc gtg gat cgg aag aga agg gat act tca gga cta gaa aga 2083 Arg
Arg Ser Val Asp Arg Lys Arg Arg Asp Thr Ser Gly Leu Glu Arg
650 655 660 agt cac aaa tct tca aaa ggt
ggt agt agt aga gat aca aaa gga tca 2131 Ser His Lys Ser Ser Lys Gly
Gly Ser Ser Arg Asp Thr Lys Gly Ser 665 670
675 aag gat aag aat tcc cgg tcc gac aga aag agg tct ata tca gag
agt 2179 Lys Asp Lys Asn Ser Arg Ser Asp Arg Lys Arg Ser Ile Ser Glu
Ser 680 685 690 agt cga tca ggc aaa
aga tct tca aga agt gaa aga gac cga aaa tca 2227 Ser Arg Ser Gly Lys
Arg Ser Ser Arg Ser Glu Arg Asp Arg Lys Ser 695 700
705 710 gac agg aaa gac aaa agg cgt taa
tggaagaagc caggctttct tagccattct 2281 Asp Arg Lys Asp Lys Arg Arg
715 ttgcagcaga agatttcttg ataaaaaagg attacctttc cttgtaaaga
ggatgctgcc 2341 ttaagaattg catgttgtaa aaaatctttt tggaaaatac agactgtttg
tttaccagac 2401 attcttgtac tttttgcata attttgtaag agttatttat caaaattatg
tgaggttcca 2461 aaatatgtaa aaatgataat aataaaaaaa gattaacatc ccttgtcatc
ttttttaaat 2521 atcctatact cttcagtaag aatctgtata ttttaatagg caaatcttta
agtctgttcc 2581 cttctaattc tgtatcatac attgcttttg tagaaataaa tgtgtgttta
tttcattatt 2641 tttgggatgt cctcgttgac acttgtataa taaatatcct ctttatcatt
ttcagctttt 2701 aacactagat actgcacgtg attagaatgt ttttgaaggt ttcctcgttt
ttatttgcct 2761 tggacagttt ttagttgtca gagtcagagc tttgcagctt tgagggggaa
cagttctctt 2821 taaaatcatt tgctattttc tattctccct tgttatttta atctaagcat
tttcccccgt 2881 ttctcatatt ttaaccatat gttcggtaga taactaaaca gtatgatctg
gttggcattt 2941 tcttcctgat gatgggagcg tcattctttt gtcttcatgg ttacttgtgt
gatataacat 3001 acatctgtta aagaaaatca cttctttcta ggggagggag gtagaaaagt
atctttcaaa 3061 cttggttttt gagtttgtgt cttgtcttaa ctttgtgttg gctctaactt
aaacatgctg 3121 atatgtgttt tcaagaattt tgtttaagga agtattgtat ggaagtccac
aaaatgaagg 3181 aagttcatct aggttttaat atgtaagcaa gataacacac aagtgtacca
agtgattatt 3241 aactttgttg ttttacaaat ttgtatgaac ttggagtatc tgttggccat
tactatacat 3301 gtgcaaataa atgtggctta gacttgtgtg actgcttaag actagtactt
gtattaactt 3361 tctgatgctt tatacaagag agcacttaaa ttgcatcctt ctttgagttt
aacactgtag 3421 ctttagcctt gactttgaat attcattttg ccttcccttg acaagtaaat
ggttacagtg 3481 aaaatgtgaa gaagtatttc cagttagttt ttgtgtgtat ataactcact
tgtgtaagtg 3541 gatgtgttaa aagatctctt tttaaaaatc cttttattgg agtaattatt
aaaacagtaa 3601 atgca
3606 38 717 PRT Homo sapiens 38 Met Ala Val Ala Val Arg Thr
Leu Gln Glu Gln Leu Glu Lys Ala Lys 1 5
10 15 Glu Ser Leu Lys Asn Val Asp Glu Asn Ile Arg Lys
Leu Thr Gly Arg 20 25 30
Asp Pro Asn Asp Val Arg Pro Ile Gln Ala Arg Leu Leu Ala Leu Ser
35 40 45 Gly Pro Gly Gly Gly Arg Gly
Arg Gly Ser Leu Leu Leu Arg Arg Gly 50 55
60 Phe Ser Asp Ser Gly Gly Gly Pro Pro Ala Lys Gln Arg Asp Leu Glu
65 70 75 80 Gly Ala
Val Ser Arg Leu Gly Gly Glu Arg Arg Thr Arg Arg Glu Ser
85 90 95 Arg Gln Glu Ser Asp Pro Glu
Asp Asp Asp Val Lys Lys Pro Ala Leu 100 105
110 Gln Ser Ser Val Val Ala Thr Ser Lys Glu Arg Thr Arg Arg
Asp Leu 115 120 125 Ile Gln Asp
Gln Asn Met Asp Glu Lys Gly Lys Gln Arg Asn Arg Arg 130
135 140 Ile Phe Gly Leu Leu Met Gly Thr Leu Gln Lys Phe
Lys Gln Glu Ser 145 150 155
160 Thr Val Ala Thr Glu Arg Gln Lys Arg Arg Gln Glu Ile Glu Gln Lys
165 170 175 Leu Glu Val Gln
Ala Glu Glu Glu Arg Lys Gln Val Glu Asn Glu Arg 180
185 190 Arg Glu Leu Phe Glu Glu Arg Arg Ala Lys Gln
Thr Glu Leu Arg Leu 195 200 205
Leu Glu Gln Lys Val Glu Leu Ala Gln Leu Gln Glu Glu Trp Asn Glu 210
215 220 His Asn Ala Lys Ile Ile Lys Tyr Ile
Arg Thr Lys Thr Lys Pro His 225 230 235
240 Leu Phe Tyr Ile Pro Gly Arg Met Cys Pro Ala Thr Gln Lys
Leu Ile 245 250 255 Glu
Glu Ser Gln Arg Lys Met Asn Ala Leu Phe Glu Gly Arg Arg Ile
260 265 270 Glu Phe Ala Glu Gln Ile Asn
Lys Met Glu Ala Arg Pro Arg Arg Gln 275 280
285 Ser Met Lys Glu Lys Glu His Gln Val Val Arg Asn Glu Glu Gln
Lys 290 295 300 Ala Glu Gln Glu Glu
Gly Lys Val Ala Gln Arg Glu Glu Glu Leu Glu 305 310
315 320 Glu Thr Gly Asn Gln His Asn Asp Val Glu
Ile Glu Glu Ala Gly Glu 325 330
335 Glu Glu Glu Lys Glu Ile Ala Ile Val His Ser Asp Ala Glu Lys Glu
340 345 350 Gln Glu Glu Glu
Glu Gln Lys Gln Glu Met Glu Val Lys Met Glu Glu 355
360 365 Glu Thr Glu Val Arg Glu Ser Glu Lys Gln Gln Asp
Ser Gln Pro Glu 370 375 380 Glu Val
Met Asp Val Leu Glu Met Val Glu Asn Val Lys His Val Ile 385
390 395 400 Ala Asp Gln Glu Val Met Glu
Thr Asn Arg Val Glu Ser Val Glu Pro 405
410 415 Ser Glu Asn Glu Ala Ser Lys Glu Leu Glu Pro Glu
Met Glu Phe Glu 420 425 430
Ile Glu Pro Asp Lys Glu Cys Lys Ser Leu Ser Pro Gly Lys Glu Asn
435 440 445 Val Ser Ala Leu Asp Met Glu
Lys Glu Ser Glu Glu Lys Glu Glu Lys 450 455
460 Glu Ser Glu Pro Gln Pro Glu Pro Val Ala Gln Pro Gln Pro Gln Ser
465 470 475 480 Gln Pro
Gln Leu Gln Leu Gln Ser Gln Ser Gln Pro Val Leu Gln Ser
485 490 495 Gln Pro Pro Ser Gln Pro Glu
Asp Leu Ser Leu Ala Val Leu Gln Pro 500 505
510 Thr Pro Gln Val Thr Gln Glu Gln Gly His Leu Leu Pro Glu
Arg Lys 515 520 525 Asp Phe Pro
Val Glu Ser Val Lys Leu Thr Glu Val Pro Val Glu Pro 530
535 540 Val Leu Thr Val His Pro Glu Ser Lys Ser Lys Thr
Lys Thr Arg Ser 545 550 555
560 Arg Ser Arg Gly Arg Ala Arg Asn Lys Thr Ser Lys Ser Arg Ser Arg
565 570 575 Ser Ser Ser Ser
Ser Ser Ser Ser Ser Ser Ser Thr Ser Ser Ser Ser 580
585 590 Gly Ser Ser Ser Ser Ser Gly Ser Ser Ser Ser
Arg Ser Ser Ser Ser 595 600 605
Ser Ser Ser Ser Thr Ser Gly Ser Ser Ser Arg Asp Ser Ser Ser Ser 610
615 620 Thr Ser Ser Ser Ser Glu Ser Arg Ser
Arg Ser Arg Gly Arg Gly His 625 630 635
640 Asn Arg Asp Arg Lys His Arg Arg Ser Val Asp Arg Lys Arg
Arg Asp 645 650 655 Thr
Ser Gly Leu Glu Arg Ser His Lys Ser Ser Lys Gly Gly Ser Ser
660 665 670 Arg Asp Thr Lys Gly Ser Lys
Asp Lys Asn Ser Arg Ser Asp Arg Lys 675 680
685 Arg Ser Ile Ser Glu Ser Ser Arg Ser Gly Lys Arg Ser Ser Arg
Ser 690 695 700 Glu Arg Asp Arg Lys
Ser Asp Arg Lys Asp Lys Arg Arg 705 710
715 39 2509 DNA Homo sapiens CDS (68)..(1924) RHOT2 mRNA 39 gagcgaaagg
cttgaggacc aggtcggggc cgggttccgg gtcggggagc ggctccgggc 60 ggcagct atg
agg cgg gac gtg cgc atc ctg tta ctg ggc gag gcc cag 109 Met
Arg Arg Asp Val Arg Ile Leu Leu Leu Gly Glu Ala Gln 1
5 10 gtg ggg aag acg tcg ctg atc ctg tcc ctg gtg ggc
gag gag ttc ccc 157 Val Gly Lys Thr Ser Leu Ile Leu Ser Leu Val Gly
Glu Glu Phe Pro 15 20 25
30 gag gag gtc cct ccc cgc gcg gag gag atc acg atc ccc gcg gac gtc
205 Glu Glu Val Pro Pro Arg Ala Glu Glu Ile Thr Ile Pro Ala Asp Val
35 40 45 acc ccg gag aag gtg
ccc acc cac atc gtg gac tac tca gaa gcc gag 253 Thr Pro Glu Lys Val
Pro Thr His Ile Val Asp Tyr Ser Glu Ala Glu 50
55 60 cag acg gac gag gag ctg cgg gag gag atc cac aag
gca aac gtg gtg 301 Gln Thr Asp Glu Glu Leu Arg Glu Glu Ile His Lys
Ala Asn Val Val 65 70 75 tgt
gtg gtg tat gac gtc tct gag gag gcc acc att gag aag att cga 349 Cys
Val Val Tyr Asp Val Ser Glu Glu Ala Thr Ile Glu Lys Ile Arg 80
85 90 act aag tgg atc cca ctg gtg aat ggg ggg
acc acg cag ggg ccc agg 397 Thr Lys Trp Ile Pro Leu Val Asn Gly Gly
Thr Thr Gln Gly Pro Arg 95 100 105
110 gtg ccc atc atc cta gtg ggc aac aag tca gac ctg cgg tcg ggg
agc 445 Val Pro Ile Ile Leu Val Gly Asn Lys Ser Asp Leu Arg Ser Gly
Ser 115 120 125 tcc atg
gag gcc gtg ctc ccc atc atg agc cag ttt ccc gag att gag 493 Ser Met
Glu Ala Val Leu Pro Ile Met Ser Gln Phe Pro Glu Ile Glu 130
135 140 acc tgc gtg gag tgt tcg gcc aag aac
ctg agg aac atc tca gag ctg 541 Thr Cys Val Glu Cys Ser Ala Lys Asn
Leu Arg Asn Ile Ser Glu Leu 145 150
155 ttc tac tac gcc cag aag gcc gtc ctg cat ccc aca gcc ccc ctc tat
589 Phe Tyr Tyr Ala Gln Lys Ala Val Leu His Pro Thr Ala Pro Leu Tyr
160 165 170 gac cct gag gcc aag cag ttg
agg ccc gcg tgc gcc cag gcg ctg acg 637 Asp Pro Glu Ala Lys Gln Leu
Arg Pro Ala Cys Ala Gln Ala Leu Thr 175 180
185 190 cgc atc ttc agg ctc tca gat cag gac ctg gac cag
gcg ctc agt gac 685 Arg Ile Phe Arg Leu Ser Asp Gln Asp Leu Asp Gln
Ala Leu Ser Asp 195 200
205 gaa gag ctc aac gct ttc cag aaa tcc tgc ttt ggg cac ccc ctg gcc
733 Glu Glu Leu Asn Ala Phe Gln Lys Ser Cys Phe Gly His Pro Leu Ala
210 215 220 ccg cag gcc ctg gag gac
gtg aag acg gtg gtg tgc agg aac gtg gcg 781 Pro Gln Ala Leu Glu Asp
Val Lys Thr Val Val Cys Arg Asn Val Ala 225 230
235 ggc ggc gtg cgg gag gac cgg ctg acc ctg gat ggt ttc ctc
ttc ctg 829 Gly Gly Val Arg Glu Asp Arg Leu Thr Leu Asp Gly Phe Leu
Phe Leu 240 245 250 aac acg ctc ttc
atc cag cgc ggc cgg cac gag acc acc tgg acc atc 877 Asn Thr Leu Phe
Ile Gln Arg Gly Arg His Glu Thr Thr Trp Thr Ile 255 260
265 270 ctg cgg cgc ttc ggc tac agc gat gcc
ctg gag ctg act gcg gac tat 925 Leu Arg Arg Phe Gly Tyr Ser Asp Ala
Leu Glu Leu Thr Ala Asp Tyr 275 280
285 ctc tcc cct ctg atc cac gtg ccc ccc ggc tgc agc acg gag ctc
aac 973 Leu Ser Pro Leu Ile His Val Pro Pro Gly Cys Ser Thr Glu Leu
Asn 290 295 300 cac ctt ggc
tac cag ttt gtg cag aga gtg ttt gag aag cac gac cag 1021 His Leu Gly
Tyr Gln Phe Val Gln Arg Val Phe Glu Lys His Asp Gln 305
310 315 gac cgc gac ggc gcc ctc tcg ccc gtg gag ctg
caa agc ctt ttc agt 1069 Asp Arg Asp Gly Ala Leu Ser Pro Val Glu Leu
Gln Ser Leu Phe Ser 320 325 330 gtg
ttc cca gca gcg ccc tgg ggc ccc gag ctc cca cgc aca gtc cgc 1117 Val
Phe Pro Ala Ala Pro Trp Gly Pro Glu Leu Pro Arg Thr Val Arg 335
340 345 350 aca gag gcc ggc cgg ttg
ccc ctg cac gga tac ctc tgc cag tgg acc 1165 Thr Glu Ala Gly Arg Leu
Pro Leu His Gly Tyr Leu Cys Gln Trp Thr 355
360 365 ctg gtg acc tac ctg gac gtc cgg agc tgc ctt gga
cac cta ggc tac 1213 Leu Val Thr Tyr Leu Asp Val Arg Ser Cys Leu Gly
His Leu Gly Tyr 370 375 380
ctg ggc tac ccc acc ctc tgt gag cag gac cag gcc cat gcc atc aca 1261
Leu Gly Tyr Pro Thr Leu Cys Glu Gln Asp Gln Ala His Ala Ile Thr
385 390 395 gtc act cgt gag aag agg ctg
gac cag gag aag gga cag acg cag cgg 1309 Val Thr Arg Glu Lys Arg Leu
Asp Gln Glu Lys Gly Gln Thr Gln Arg 400 405
410 agc gtc ctc ctg tgc aag gtg gta ggg gcc cgt gga gtg ggc aag tct
1357 Ser Val Leu Leu Cys Lys Val Val Gly Ala Arg Gly Val Gly Lys Ser
415 420 425 430 gcc ttc
ctg cag gct ttt ctc ggc cgc ggc ctg ggg cac cag gac acg 1405 Ala Phe
Leu Gln Ala Phe Leu Gly Arg Gly Leu Gly His Gln Asp Thr
435 440 445 agg gag cag cct ccc ggc tac
gcc atc gac acg gtg cag gtc aat gga 1453 Arg Glu Gln Pro Pro Gly Tyr
Ala Ile Asp Thr Val Gln Val Asn Gly 450 455
460 cag gag aag tac ttg atc ctc tgt gag gtg ggc aca gat ggt
ctg ctg 1501 Gln Glu Lys Tyr Leu Ile Leu Cys Glu Val Gly Thr Asp Gly
Leu Leu 465 470 475 gcc aca tcg
ctg gac gcc acc tgt gac gtt gcc tgc ttg atg ttt gat 1549 Ala Thr Ser
Leu Asp Ala Thr Cys Asp Val Ala Cys Leu Met Phe Asp 480
485 490 ggc agt gac cca aag tcc ttt gca cat tgt gcc agc
gtc tac aag cac 1597 Gly Ser Asp Pro Lys Ser Phe Ala His Cys Ala Ser
Val Tyr Lys His 495 500 505
510 cat tac atg gac ggg cag acc ccc tgc ctc ttt gtc tcc tcc aag gcc
1645 His Tyr Met Asp Gly Gln Thr Pro Cys Leu Phe Val Ser Ser Lys Ala
515 520 525 gac ctg ccc gaa
ggt gtc gcg gtg tct ggc cca tca ccg gcc gag ttt 1693 Asp Leu Pro Glu
Gly Val Ala Val Ser Gly Pro Ser Pro Ala Glu Phe 530
535 540 tgc cgc aag cac cgg cta ccc gct ccc gtg ccg
ttc tcc tgt gct ggc 1741 Cys Arg Lys His Arg Leu Pro Ala Pro Val Pro
Phe Ser Cys Ala Gly 545 550 555
cca gcc gag ccc agc acc acc atc ttc acc cag ctc gcc acc atg gcc 1789
Pro Ala Glu Pro Ser Thr Thr Ile Phe Thr Gln Leu Ala Thr Met Ala 560
565 570 gcc ttc cca cat ttg gtc cac gca gag
ctg cat ccc tct tcc ttc tgg 1837 Ala Phe Pro His Leu Val His Ala Glu
Leu His Pro Ser Ser Phe Trp 575 580 585
590 ctc cgg ggg ctg ctg ggg gtt gtc ggg gcc gcc gtg gcc gca
gtc ctc 1885 Leu Arg Gly Leu Leu Gly Val Val Gly Ala Ala Val Ala Ala
Val Leu 595 600 605 agc
ttc tca ctc tac agg gtc ctg gtg aag agc cag tga ggcccctggt 1934 Ser
Phe Ser Leu Tyr Arg Val Leu Val Lys Ser Gln 610
615 acccaagccc cctcccctga cctgggtgtg cctcgctgct ggggctctgc aggggcagca
1994 cagctggggt gcaggccagg ctgccactcc gggaacgcct ttgcgccggg actttttgtt
2054 tctgaaggca gtcgatctgc agcggggcct tatgctgcca tgcactgccc tggctcctgc
2114 cggaccccca gggtgggccg tggcaggtgg ctgagcagga gctcccaagt gccggccacc
2174 gctgtcaggg attgcccacc cctgggcatc atgtgtgtgg ggccggggag cacaggtgtg
2234 ggagctggtg accccagacc cagaattctc agggctctac ccccctttcc tggtcctagg
2294 tggccagtgg gtatgaggag ggctggaagg cagagctttg ggccaaaagc aggcgttggg
2354 gggtcccccc tcaagtttgg agccgtttcc gtggttgtag cagaggaccg gaggttgggt
2414 tcctgattaa acttcactgt gtgttttcta tctcggatcc cagtctctga agacaacttg
2474 ctttgattca acctaaaaaa aaaaaaaaaa aaaaa
2509 40 618 PRT Homo sapiens 40 Met Arg Arg Asp Val Arg Ile Leu Leu Leu
Gly Glu Ala Gln Val Gly 1 5 10
15 Lys Thr Ser Leu Ile Leu Ser Leu Val Gly Glu Glu Phe Pro Glu Glu
20 25 30 Val Pro Pro Arg Ala
Glu Glu Ile Thr Ile Pro Ala Asp Val Thr Pro 35
40 45 Glu Lys Val Pro Thr His Ile Val Asp Tyr Ser Glu
Ala Glu Gln Thr 50 55 60 Asp Glu
Glu Leu Arg Glu Glu Ile His Lys Ala Asn Val Val Cys Val 65
70 75 80 Val Tyr Asp Val Ser Glu Glu
Ala Thr Ile Glu Lys Ile Arg Thr Lys 85
90 95 Trp Ile Pro Leu Val Asn Gly Gly Thr Thr Gln Gly
Pro Arg Val Pro 100 105 110
Ile Ile Leu Val Gly Asn Lys Ser Asp Leu Arg Ser Gly Ser Ser Met
115 120 125 Glu Ala Val Leu Pro Ile Met
Ser Gln Phe Pro Glu Ile Glu Thr Cys 130 135
140 Val Glu Cys Ser Ala Lys Asn Leu Arg Asn Ile Ser Glu Leu Phe Tyr
145 150 155 160 Tyr Ala
Gln Lys Ala Val Leu His Pro Thr Ala Pro Leu Tyr Asp Pro
165 170 175 Glu Ala Lys Gln Leu Arg Pro
Ala Cys Ala Gln Ala Leu Thr Arg Ile 180 185
190 Phe Arg Leu Ser Asp Gln Asp Leu Asp Gln Ala Leu Ser Asp
Glu Glu 195 200 205 Leu Asn Ala
Phe Gln Lys Ser Cys Phe Gly His Pro Leu Ala Pro Gln 210
215 220 Ala Leu Glu Asp Val Lys Thr Val Val Cys Arg Asn
Val Ala Gly Gly 225 230 235
240 Val Arg Glu Asp Arg Leu Thr Leu Asp Gly Phe Leu Phe Leu Asn Thr
245 250 255 Leu Phe Ile Gln
Arg Gly Arg His Glu Thr Thr Trp Thr Ile Leu Arg 260
265 270 Arg Phe Gly Tyr Ser Asp Ala Leu Glu Leu Thr
Ala Asp Tyr Leu Ser 275 280 285
Pro Leu Ile His Val Pro Pro Gly Cys Ser Thr Glu Leu Asn His Leu 290
295 300 Gly Tyr Gln Phe Val Gln Arg Val Phe
Glu Lys His Asp Gln Asp Arg 305 310 315
320 Asp Gly Ala Leu Ser Pro Val Glu Leu Gln Ser Leu Phe Ser
Val Phe 325 330 335 Pro
Ala Ala Pro Trp Gly Pro Glu Leu Pro Arg Thr Val Arg Thr Glu
340 345 350 Ala Gly Arg Leu Pro Leu His
Gly Tyr Leu Cys Gln Trp Thr Leu Val 355 360
365 Thr Tyr Leu Asp Val Arg Ser Cys Leu Gly His Leu Gly Tyr Leu
Gly 370 375 380 Tyr Pro Thr Leu Cys
Glu Gln Asp Gln Ala His Ala Ile Thr Val Thr 385 390
395 400 Arg Glu Lys Arg Leu Asp Gln Glu Lys Gly
Gln Thr Gln Arg Ser Val 405 410
415 Leu Leu Cys Lys Val Val Gly Ala Arg Gly Val Gly Lys Ser Ala Phe
420 425 430 Leu Gln Ala Phe
Leu Gly Arg Gly Leu Gly His Gln Asp Thr Arg Glu 435
440 445 Gln Pro Pro Gly Tyr Ala Ile Asp Thr Val Gln Val
Asn Gly Gln Glu 450 455 460 Lys Tyr
Leu Ile Leu Cys Glu Val Gly Thr Asp Gly Leu Leu Ala Thr 465
470 475 480 Ser Leu Asp Ala Thr Cys Asp
Val Ala Cys Leu Met Phe Asp Gly Ser 485
490 495 Asp Pro Lys Ser Phe Ala His Cys Ala Ser Val Tyr
Lys His His Tyr 500 505 510
Met Asp Gly Gln Thr Pro Cys Leu Phe Val Ser Ser Lys Ala Asp Leu
515 520 525 Pro Glu Gly Val Ala Val Ser
Gly Pro Ser Pro Ala Glu Phe Cys Arg 530 535
540 Lys His Arg Leu Pro Ala Pro Val Pro Phe Ser Cys Ala Gly Pro Ala
545 550 555 560 Glu Pro
Ser Thr Thr Ile Phe Thr Gln Leu Ala Thr Met Ala Ala Phe
565 570 575 Pro His Leu Val His Ala Glu
Leu His Pro Ser Ser Phe Trp Leu Arg 580 585
590 Gly Leu Leu Gly Val Val Gly Ala Ala Val Ala Ala Val Leu
Ser Phe 595 600 605 Ser Leu Tyr
Arg Val Leu Val Lys Ser Gln 610 615 41 20 DNA
Artificial AFP Forward primer (5'-3') 41 aactattggc ctgtggcgag
20 42 19 DNA Artificial AFP Reverse
primer (5'-3') 42 tcatccacca ccaagctgc
19 43 20 DNA Artificial ALDH2 Forward primer (5'-3') 43
gtttggagcc cagtcaccct 20
44 21 DNA Artificial ALDH2 Reverse primer (5'-3') 44 gggaggaagc
ttgcatgatt c 21 45 21 DNA
Artificial APCS Forward primer (5'-3') 45 ggccaggaat atgaacaagc c
21 46 20 DNA Artificial APCS
Reverse primer (5'-3') 46 cttctccagc ggtgtgatca
20 47 20 DNA Artificial APOC4 Forward primer
(5'-3') 47 ggagctgctg gagacagtgg
20 48 20 DNA Artificial APOC4 Reverse primer (5'-3') 48
tttggattcg aggaaccagg 20
49 19 DNA Artificial AQP9 Forward primer (5'-3') 49 gcttcctccc tgggactga
19 50 20 DNA Artificial
AQP9 Reverse primer (5'-3') 50 caaccaaagg gcccactaca
20 51 21 DNA Artificial BUB1 Forward primer
(5'-3') 51 acccctgaaa aagtgatgcc t
21 52 21 DNA Artificial BUB1 Reverse primer (5'-3') 52
tcatcctgtt ccaaaaatcc g 21
53 21 DNA Artificial C1S Forward primer (5'-3') 53 ttgtttggtt ctgtcatccg
c 21 54 19 DNA Artificial
C1S Reverse primer (5'-3') 54 tggaacacat ttcggcagc
19 55 23 DNA Artificial CYP2E1 Forward primer
(5'-3') 55 caaccaagaa tttcctgatc cag
23 56 21 DNA Artificial CYP2E1 Reverse primer (5'-3') 56
aagaaacaac tccatgcgag c 21
57 22 DNA Artificial DLG7 Forward primer (5'-3') 57 gcaggaagaa
tgtgctgaaa ca 22 58 21 DNA
Artificial DLG7 Reverse primer (5'-3') 58 tccaagtctt tgagaagggc c
21 59 20 DNA Artificial DUSP9
Forward primer (5'-3') 59 cggaggccat tgagttcatt
20 60 21 DNA Artificial DUSP9 Reverse primer
(5'-3') 60 accaggtcat aggcatcgtt g
21 61 20 DNA Artificial E2F5 Forward primer (5'-3') 61
ccattcaggc accttctggt 20
62 22 DNA Artificial E2F5 Reverse primer (5'-3') 62 acgggcttag
atgaactcga ct 22 63 19 DNA
Artificial GHR Forward primer (5'-3') 63 cttggcactg gcaggatca
19 64 19 DNA Artificial GHR Reverse
primer (5'-3') 64 aggtgaacgg cacttggtg
19 65 20 DNA Artificial HPD Forward primer (5'-3') 65
atcttcacca aaccggtgca 20
66 21 DNA Artificial HPD Reverse primer (5'-3') 66 ccatgttggt gaggttaccc
c 21 67 21 DNA Artificial
IGSF1 Forward primer (5'-3') 67 cactcacact gaaaaacgcc c
21 68 21 DNA Artificial IGSF1 Reverse primer
(5'-3') 68 gggtggagca attgaaagtc a
21 69 20 DNA Artificial NLE1 Forward primer (5'-3') 69
atgtgaaggc ccagaagctg 20
70 19 DNA Artificial NLE1 Reverse primer (5'-3') 70 gagaacttcg ggccgtctc
19 71 20 DNA Artificial
RPL10A Forward primer (5'-3') 71 tatcccccac atggacatcg
20 72 21 DNA Artificial RPL10A Reverse
primer (5'-3') 72 tgccttattt aaacctgggc c
21 73 20 DNA Artificial ACTG1 Forward primer (5'-3') 73
gatggccagg tcatcaccat 20
74 20 DNA Artificial ACTG1 Reverse primer (5'-3') 74 acaggtcttt
gcggatgtcc 20 75 20 DNA
Artificial EFF1A1 Forward primer (5'-3') 75 tcacccgtaa ggatggcaat
20 76 20 DNA Artificial EFF1A1
Reverse primer (5'-3') 76 cggccaacag gaacagtacc
20 77 20 DNA Artificial PNN Forward primer (5'-3')
77 cctttctggt cctggtggag
20 78 21 DNA Artificial PNN Reverse primer (5'-3') 78 tgattctctt
ctggtccgac g 21 79 21 DNA
Artificial RHOT2 Forward primer (5'-3') 79 ctgcggacta tctctcccct c
21 80 20 DNA Artificial RHOT2
Reverse primer (5'-3') 80 aaaaggcttt gcagctccac
20 81 20 DNA Artificial AFP Forward primer 81
gccagtgctg cacttcttca 20
82 21 DNA Artificial AFP Reverse primer 82 tgtttcatcc accaccaagc t
21 83 24 DNA Artificial AFP probe
83 atgccaacag gaggccatgc ttca
24 84 19 DNA Artificial ALDH2 Forward primer 84 tgcaggatgg catgaccat
19 85 24 DNA Artificial
ALDH2 Reverse primer 85 tcttgaactt caggatctgc atca
24 86 23 DNA Artificial ALDH2 probe 86 ccaaggagga
gatcttcggg cca 23 87 22 DNA
Artificial APCS Forward primer 87 agctgggagt cctcatcagg ta
22 88 20 DNA Artificial APCS Reverse
primer 88 cgcagaccct ttttcaccaa
20 89 27 DNA Artificial APCS probe 89 tgctgaattt tggatcaatg
ggacacc 27 90 20 DNA Artificial
APOC4 Forward primer 90 tgaaggagct gctggagaca
20 91 18 DNA Artificial APOC4 Reverse primer 91
cgggctccag aaccattg 18
92 25 DNA Artificial APOC4 probe 92 tggtgaacag gaccagagac gggtg
25 93 19 DNA Artificial AQP9 Forward
primer 93 gccatcggcc tcctgatta
19 94 20 DNA Artificial AQP9 Reverse primer 94 gttcatggca
cagccactgt 20 95 25 DNA
Artificial AQP9 probe 95 tgtcattgct tcctccctgg gactg
25 96 25 DNA Artificial BUB1 Forward primer 96
acatctggtt ttcagtgtgt tgaga 25
97 21 DNA Artificial BUB1 Reverse primer 97 gttgcagcaa ccccaaagta a
21 98 30 DNA Artificial BUB1
probe 98 tcagcaacaa accatggaac taccagatcg
30 99 23 DNA Artificial C1S Forward primer 99 tcccaatgac
aagaccaaat tct 23 100 21 DNA
Artificial C1S Reverse primer 100 agagcccata ggtcccacac t
21 101 21 DNA Artificial C1S probe 101
cgcagctggc ctggtgtcct g 21
102 22 DNA Artificial CYP2E1 Forward primer 102 catgagattc agcggttcat ca
22 103 19 DNA Artificial
CYP2E1 Reverse primer 103 ggtgtctcgg gttgcttca
19 104 21 DNA Artificial CYP2E1 probe 104
cctcgtgccc tccaacctgc c 21
105 23 DNA Artificial DLG7 Forward primer 105 gctggagagg agacatcaag aac
23 106 27 DNA Artificial DLG7
Reverse primer 106 cctggttgta gaggtgaaaa agtaatc
27 107 29 DNA Artificial DLG7 probe 107 tgccagacac
atttcttttg gtggtaacc 29 108 21 DNA
Artificial DUSP9 Forward primer 108 ggcctacctc atgcagaagc t
21 109 26 DNA Artificial DUSP9 Reverse
primer 109 gggagatgtt agacttcttc ctcttg
26 110 30 DNA Artificial DUSP9 probe 110 cacctctctc tcaacgatgc
ctatgacctg 30 111 19 DNA Artificial
E2F5 Forward primer 111 cctgttcccc cacctgatg
19 112 23 DNA Artificial E2F5 Reverse primer 112
tttctgtgga gtcactggag tca 23
113 25 DNA Artificial E2F5 probe 113 cctcacacag ccttcctccc agtcc
25 114 20 DNA Artificial GHR Forward
primer 114 cccaggtgag cgacattaca
20 115 22 DNA Artificial GHR Reverse primer 115 catccctgcc
ttattctttt gg 22 116 26 DNA
Artificial GHR probe 116 cagcaggtag tgtggtcctt tccccg
26 117 18 DNA Artificial HPD Forward primer 117
cccacgctct tcctggaa 18
118 17 DNA Artificial HPD Reverse primer 118 ttgccggctc caaaacc
17 119 22 DNA Artificial HPD
probe 119 tcatccagcg ccacaaccac ca
22 120 22 DNA Artificial IGSF1 Forward primer 120 gaccattgcc
cttgaagagt gt 22 121 24 DNA
Artificial IGSF1 Reverse primer 121 gagaggttga tgaaggagaa ttgg
24 122 25 DNA Artificial IGSF1 probe
122 accaagaagg agaaccaggc acccc
25 123 21 DNA Artificial NLE1 Forward primer 123 tgcctccttt gacaagtcca t
21 124 19 DNA Artificial
NLE1 Reverse primer 124 cgcgtaggga agccaggta
19 125 19 DNA Artificial NLE1 probe 125 tgggatggca
ggacgggca 19 126 21 DNA
Artificial RPL10A Forward primer 126 tcggcccagg tttaaataag g
21 127 20 DNA Artificial RPL10A Reverse
primer 127 ccactttggc caccatgttt
20 128 27 DNA Artificial RPL10A probe 128 agttcccttc cctgctcaca
cacaacg 27 129 15 DNA Artificial
ACTG1 Forward primer 129 ggcgcccagc accat
15 130 21 DNA Artificial ACTG1 Reverse primer 130
ccgatccaca ccgagtactt g 21
131 25 DNA Artificial ACTG1 probe 131 atcaagatca tcgcaccccc agagc
25 132 19 DNA Artificial EFF1A1
Forward primer 132 gcggtgggtg tcatcaaag
19 133 20 DNA Artificial EFF1A1 Reverse primer 133
tgggcagact tggtgacctt 20
134 26 DNA Artificial EFF1A1 probe 134 agtggacaag aaggctgctg gagctg
26 135 19 DNA Artificial PNN Forward
primer 135 gaattcccgg tccgacaga
19 136 24 DNA Artificial PNN Reverse primer 136 tttcggtctc
tttcacttct tgaa 24 137 38 DNA
Artificial PNN probe 137 agaggtctat atcagagagt agtcgatcag gcaaaaga
38 138 20 DNA Artificial RHOT2 Forward Primer 138
cccagcacca ccatcttcac 20
139 20 DNA Artificial RHOT2 Reverse primer 139 ccagaaggaa gagggatgca
20 140 19 DNA Artificial
RHOT2 probe 140 cagctcgcca ccatggccg
19
User Contributions:
Comment about this patent or add new information about this topic:
| People who visited this patent also read: | |
| Patent application number | Title |
|---|---|
| 20120039794 | HYDROGEN PROCESS |
| 20120039779 | METHOD FOR FEEDING REDUCING AGENT INTO AN EXHAUST GAS SYSTEM AND CORRESPONDING EXHAUST GAS SYSTEM AND MOTOR VEHICLE |
| 20120039778 | Method and Apparatus for Controlling Acid Gas Emissions from Cement Plants |
| 20120039769 | DISK-BASED FLUID SAMPLE COLLECTION DEVICE |
| 20120039768 | NOZZLE ASSEMBLY FOR MOUNTING IN MECHANICAL OR ELECTRONIC PIPETTE |