Patent application title: Methods to Treat Cancer with 10-propargyl-10-deazaaminopterin and Methods for Assessing Cancer for Increased Sensitivity to 10-propargyl-10-deazaaminopterin
Inventors:
Owen A. O'Connor (Scarsdale, NY, US)
Francis Sirotnak (Hampton Bays, NY, US)
Assignees:
SLOAN-KETTERING INSTITUTE FOR CANCER RESEARCH
IPC8 Class: AA61K314985FI
USPC Class:
514249
Class name: Hetero ring is six-membered consisting of two nitrogens and four carbon atoms (e.g., pyridazines, etc.) polycyclo ring system having a 1,2- or 1,4-diazine as one of the cyclos 1,4-diazine as one of the cyclos
Publication date: 2010-07-01
Patent application number: 20100168118
Claims:
1. A method of selecting a patient for treatment of a cancer with
10-propargyl-10-deazaminopterin, the method comprising the steps of:(a)
obtaining a sample of the patient's cancer tissue;(b) determining the
expression level of at least one selected polypeptide expressed by the
sample;(c) obtaining a reference expression level for the at least one
selected polypeptide for a cancer having sensitivity to
10-propargyl-10-deazaminopterin;(d) comparing the expression data for the
at least one selected polypeptide of step (b) with the reference
expression for the at least one selected polypeptide of step (c), wherein
a match of the sample expression of the at least one selected polypeptide
to the reference expression of the at least one selected polypeptide
indicates the patient's cancer has greater sensitivity to
10-propargyl-10-deazaminopterin, wherein the at least one selected
polypeptide is selected from the group consisting of reduced folate
carrier-1 enzyme (RFC-1), dihydrofolate reductase (DHFR),
folylpoly-gamma-glutamate synthetase (FPGS), thymidylate synthase (TS),
γ-glutamyl hydrolase (GGH), and glycinamide ribonucleotide
formyltransferase (GARFT); and(e) selecting the patient for treatment
10-propargyl-10-deazaminopterin when the expression of the at least one
sample polypeptide and the at least one reference polypeptide matches.
2. The method of claim 1, wherein the reference cancer is a T-cell lymphoma, a NSCLC, or a multiple myeloma.
3. The method of claim 1, wherein the at least one selected polypeptide is RFC-1.
4. The method of claim 1, wherein the at least one selected polypeptide is TS.
5. The method of claim 1, wherein the at least one selected polypeptide is DHFR.
6. The method of claim 1, wherein a match is defined as the at least one selected polypeptide having an expression level of at least 50% of the reference's at least one selected polypeptide expression level.
7. The method of claim 1, wherein the patient's cancer is lymphoma, multiple myeloma, or NSCLC.
8. The method of claim 7, wherein the lymphoma is a T-cell lymphoma selected from the group consisting of lymphoblastic lymphomas in which the malignancy occurs in primitive lymphoid progenitors from the thymus; mature or peripheral T-cell neoplasms, including T-cell prolymphocytic leukemia, T-cell granular lymphocytic leukemia, aggressive NK-cell leukemia, cutaneous T-cell lymphoma (Mycosis fungoides/Sezary syndrome), anaplastic large cell lymphoma, T-cell type, enteropathy-type T-cell lymphoma, Adult T-cell leukemia/lymphoma including those associated with HTLV-1, and angioimmunoblastic T-cell lymphoma, and subcutaneous panniculitic T-cell lymphoma; and peripheral T-cell lymphomas that initially involve a lymph node paracortex.
9. A method for assessing sensitivity of a patient's cancer to treatment with 10-propargyl-10-deazaminopterin comprising the steps of:(a) obtaining a sample of the patient's cancer tissue;(b) determining the expression level of at least one selected polypeptide expressed by the sample;(c) obtaining a reference expression level for the at least one selected polypeptide for a least one cancer having sensitivity to 10-propargyl-10-deazaminopterin;(d) comparing the expression data for the at least one selected polypeptide of step (b) with the reference expression for the at least one selected polypeptide of step (c), wherein a match of the sample expression level of the at least one selected polypeptide to the reference expression level of the at least one selected polypeptide indicates the patient's cancer has greater sensitivity to 10-propargyl-10-deazaminopterin, wherein the at least one selected polypeptide is selected from the group consisting of reduced folate carrier-1 enzyme (RFC-1), dihydrofolate reductase (DHFR), folylpoly-gamma-glutamate synthetase (FPGS), thymidylate synthase (TS), γ-glutamyl hydrolase (GGH), and glycinamide ribonucleotide formyltransferase (GARFT); and(e) generating a report of the sensitivity of the sample to 10-propargyl-10-deazaminopterin.
10. The method of claim 9, wherein the reference cancer is a T-cell lymphoma, NSCLC, or multiple myeloma.
11. The method of claim 9, wherein the at least one selected polypeptide is RFC-1.
12. The method of claim 9, wherein the at least one selected polypeptide is TS.
13. The method of claim 9, wherein the at least one selected polypeptide is DHFR.
14. The method of claim 9, wherein a match is defined as the at least one selected polypeptide having an expression level of at least 50% of the reference's at least one selected polypeptide expression level.
15. The method of claim 9, wherein the patient's cancer is lymphoma, NSCLC, or multiple myeloma.
16. The method of claim 15, wherein the lymphoma is a T-cell lymphoma selected from the group consisting of lymphoblastic lymphomas in which the malignancy occurs in primitive lymphoid progenitors from the thymus; mature or peripheral T-cell neoplasms, including T-cell prolymphocytic leukemia, T-cell granular lymphocytic leukemia, aggressive NK-cell leukemia, cutaneous T-cell lymphoma (Mycosis fungoides/Sezary syndrome), anaplastic large cell lymphoma, T-cell type, enteropathy-type T-cell lymphoma, Adult T-cell leukemia/lymphoma including those associated with HTLV-1, and angioimmunoblastic T-cell lymphoma, and subcutaneous panniculitic T-cell lymphoma; and peripheral T-cell lymphomas that initially involve a lymph node paracortex.
17. A method for the treatment of multiple myeloma comprising administering to a patient diagnosed with having multiple myeloma a pharmaceutically acceptable composition comprising a therapeutically effective amount of 10-propargyl-10-deazaminopterin.
18. The method of claim 17, wherein the 10-propargyl-10-deazaminopterin is substantially free of 10-deazaminopterin.
19. The method of claim 17, wherein the 10-propargyl-10-deazaminopterin is administered in an amount of from about 30 to about 275 mg/m2 per dose.
20. A method to modulate the expression of a polypeptide in a patient's cancer comprising administering to the patient an effective amount of 10-propargyl-10-deazaminopterin, wherein the polypeptide is selected from the group consisting of reduced folate carrier-1 enzyme (RFC-1), dihydrofolate reductase (DHFR), folylpoly-gamma-glutamate synthetase (FPGS), thymidylate synthase (TS), γ-glutamyl hydrolase (GGH), and glycinamide ribonucleotide formyltransferase (GARFT).
21. The method of claim 20, wherein the patient has lymphoma, multiple myeloma, or NSCLC.
22. The method of claim 20, wherein the modulation is down-regulation and the polypeptide is TS or DHFR.
23. The method of claim 20, wherein the 10-propargyl-10-deazaminopterin is substantially free of 10-deazaminopterin.
24. The method of claim 20, wherein the 10-propargyl-10-deazaminopterin is administered in an amount of from about 30 to about 275 mg/m2 per dose.
25. The method of claim 20, wherein the polypeptide is RFC-1.
26. A kit for assessing sensitivity of a patient's cancer to treatment with 10-propargyl-10-deazaminopterin comprising at least two sets of selected polypeptide RNA-specific primers wherein each set of specific primers produces double stranded DNA complementary to at least one selected polypeptides, wherein each first primers of said sets contains a sequence which can selectively hybridize to RNA, cDNA or an EST complementary to one of the selected polypeptides to create an extension product and each said second primers of said sets is capable of selectively hybridizing to said extension product, wherein the at least one selected polypeptide is selected from the group consisting of reduced folate carrier-1 enzyme (RFC-1), dihydrofolate reductase (DHFR), folylpoly-gamma-glutamate synthetase (FPGS), thymidylate synthase (TS), γ-glutamyl hydrolase (GGH), and glycinamide ribonucleotide formyltransferase (GARFT).
27. The kit of claim 26, wherein the at least one selected peptide is RFC-1.
Description:
STATEMENT OF RELATED APPLICATIONS
[0001]The present application claims the benefit of U.S. Provisional Application No. 60/869,528, filed Dcember 11, 2006, which application is incorporated herein by reference in its entirety. The present invention claims priority from and is a continuation in part application of pending U.S. Ser. No. 11/568,254, filed Oct. 24, 2006, which is a 371 of PCT/US2005/019170 filed on May 31, 2005, claiming priority to U.S. Ser. No. 60/521,593, filed on May 30, 2004; each of which are incorporated by reference herein in their entireties.
TECHNICAL FIELD
[0002]The present invention relates to methods to treat cancer with 10-propargyl-10-deazaminopterin and methods for assessing cancers and selecting patients for treatment based for increased sensitivity to 10-propargyl-10-deazaminopterin.
BACKGROUND OF THE INVENTION
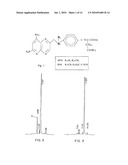
[0003]10-Propargyl-10-deazaminopterin (herein "PDX" or "10-propargyl-10dAM" or "pralatrexate") is a member of a large class of compounds which have been tested and in some cases found useful in the treatment of tumors. This compound, which has the structure shown in FIG. 1, was disclosed by DeGraw et al., "Synthesis and Antitumor Activity of 10-Propargyl-10-deazaminopterin." J. Medical Chem. 36: 2228-2231 (1993) and shown to act as an inhibitor of growth in the murine L1210 cell line and to a lesser extent of the enzyme dihydrofolate reductase ("DHFR"). In addition, some results were presented for the antitumor properties of the compound using the E0771 murine mammary tumor model. These data were equivocal because of the small number of mice used in the test (3 per dosage), the absence of any standard deviation information which would quantify the reliability of the data, and the fact that the highest dose used was in fact toxic to the mice. Nevertheless, assuming these data have some predictive value for the efficacy of a drug in treating human tumors, it would at best predict a drug which, at equivalent levels of tolerance, had properties comparable to or perhaps slightly better than methotrexate.
[0004]PCT Publication No. WO98/02163, discloses the surprising observation that more highly purified 10-propargyl-10-dAM compositions when tested in a xenograft model for their efficacy against human tumors have now been shown to be far superior to methotrexate ("MTX") and are even superior to edatrexate ("ETX"), a more recent clinical candidate. Moreover, 10-propargyl-10dAM showed a surprising ability to cure tumors such that there was no evidence of tumor growth several weeks after the cessation of therapy. Thus, highly purified composition containing 10-propargyl-10dAM. can be used in accordance with the invention to treat tumors, including both solid tumors and leukemias. The composition is illustrated for use in treatment of human mammary tumors and human lung cancer.
[0005]Subsequent studies with 10-propargyl-10-dAM have shown that it is useful on its own and in combinations with other therapeutic agents. For example, Sirotnak et al., Clinical Cancer Research Vol. 6, 3705-3712 (2000) reports that co-administration of 10-propargyl-10-dAM and probenecid, an inhibitor of a cMOAT/MRP-like plasma membrane ATPase greatly enhances the efficacy of 10-propargyl-10-dAM against human solid tumors in vivo. 10-propargyl-10-dAM and combinations of 10-propargyl-10-dAM with platinum based chemotherapeutic agents have been shown to be effective against mesothelioma. (Khokar, et al., Clin. Cancer Res. 7: 3199-3205 (2001).
[0006]Another subsequent study showed that 10-propargyl-10-dAM has particular utility in the treatment of T-cell lymphomas, even with patients with drug resistant T-cell lymphomas, disclosed in U.S. Patent Publication No. 2005/0267117, which is incorporated by reference herein in its entirety. Other studies have shown a method for assessing sensitivity of a lymphoma to treatment with 10-propargyl-10-dAM by determining the amount of reduced folate carrier-1 enzyme (RFC-1) expressed by the sample, wherein a higher level of expressed RFC-1 is indicative of greater sensitivity to 10-propargyl-10-dAM, disclosed in PCT Publication No. WO 2005/117892, which is incorporated by reference herein in its entirety.
[0007]However, a need still remains in the art for determining which other cancers for which 10-propargyl-10-dAM has particular utility in treating, and also for methods for selecting patients for treatment with 10-propargyl-10-dAM, as well as methods for assessing sensitivity of a cancer including lymphoma to 10-propargyl-10-dAM. These and other needs are addressed by the present invention.
[0008]The term "lymphomas" refers to a variety of disease states, including Non-Hodgkins Lymphoma (NHL); diffuse large B-cell lymphoma (DLBCL); follicular lymphoma (FL); Hodgkin's Disease; Burkitt's Lymphoma; cutaneous T-cell lymphoma; primary central nervous system lymphoma, and lymphomatous metastases. In most cases, lymphoma is characterized by the presence of cancerous B-cells. However, in T-cell lymphomas, the disease state is characterized by cancerous T-lymphocytes.
[0009]All references cited herein, both supra and infra, are hereby incorporated by reference herein in their entireties.
SUMMARY OF THE INVENTION
[0010]The present invention relates to a method for assessing the sensitivity of a patient's cancer to treatment with 10-propargyl-10-deazaminopterin and a method for selecting a patient for treatment of cancer with 10-propargyl-10-deazaminopterin, by determining the amount of a selected polypeptide expressed by the cancer and comparing the amount with the amount of the selected polypeptide expressed by a reference cancer, wherein the polypeptide includes a member of folate pathways within cells and may include at least one of reduced folate carrier-1 enzyme (RFC-1), dihydrofolate reductase (DHFR), folylpoly-gamma-glutamate synthetase (FPGS), thymidylate synthase (TS), γ-glutamyl hydrolase (GGH), and glycinamide ribonucleotide formyltransferase (GARFT). The present invention also relates to the use of 10-propargyl-10-deazaminopterin in the treatment of multiple myeloma.
BRIEF DESCRIPTION OF THE DRAWINGS
[0011]FIG. 1 shows the structure of 10-propargyl-10-dAM and methotrexate;
[0012]FIG. 2 shows an HPLC of an impure 10-propargyl-10-dAM preparation prepared in accordance with the prior art;
[0013]FIG. 3 shows an HPLC of a highly purified 10-PROPARGYL-10-DAM preparation in accordance with the invention;
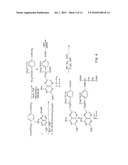
[0014]FIG. 4 shows a synthetic scheme useful in preparing the compound in accordance with the invention;
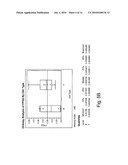
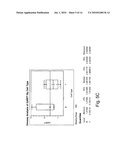
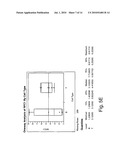
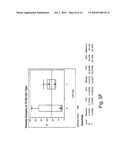
[0015]FIG. 5(a)-(f) shows a comparison of relative expression of selected folate pathway genes in B- and T-cell lymphoma cell lines; and
[0016]FIG. 6(a)-(f) shows a comparison of relative gene expression of selected folate pathway genes during treatment of B-cell (RL) and T-cell (HT) lymphoma cells with 10-propargyl-10-dAM or MTX.
DETAILED DESCRIPTION OF THE INVENTION
[0017]The present invention relates to a method for assessing the sensitivity of a patient's cancer to treatment with 10-propargyl-10-deazaminopterin and a method for selecting a patient for treatment of cancer with 10-propargyl-10-deazaminopterin, by determining the amount of a selected polypeptide expressed by the cancer and comparing the amount with the amount of the selected polypeptide expressed by a reference cancer, wherein the polypeptide includes a member of folate pathways within cells and may include at least one of reduced folate carrier-1 enzyme (RFC-1), dihydrofolate reductase (DHFR), folylpoly-gamma-glutamate synthetase (FPGS), thymidylate synthase (TS), γ-glutamyl hydrolase (GGH), and glycinamide ribonucleotide formyltransferase (GARFT). The present invention also relates to the use of 10-propargyl-10-deazaminopterin in the treatment of multiple myeloma.
[0018]As used in the specification and claims of this application, the term "lymphomas" refers to Non-Hodgkins Lymphoma (NHL); diffuse large B-cell lymphoma (DLBCL); follicular lymphoma (FL); Hodgkin's Disease; Burkitt's Lymphoma; cutaneous T-cell lymphoma; primary central nervous system lymphoma, and lymphomatous metastases. In one embodiment of the present invention, this application relates to the use of 10-propargyl-10-deazaminopterin in the treatment of T-cell lymphoma.
[0019]T-cell lymphomas are lymphomas in which the T cells of the patient are determined to be cancerous. T-cell lymphomas encompass a variety of conditions including without limitation: (a) lymphoblastic lymphomas in which the malignancy occurs in primitive lymphoid progenitors from the thymus; (b) mature or peripheral T-cell neoplasms, including T-cell prolymphocytic leukemia, T-cell granular lymphocytic leukemia, aggressive NK-cell leukemia, cutaneous T-cell lymphoma (Mycosis fungoides/Sezary syndrome), anaplastic large cell lymphoma, T-cell type, enteropathy-type T-cell lymphoma, Adult T-cell leukemia/lymphoma including those associated with HTLV-1, and angioimmunoblastic T-cell lymphoma, and subcutaneous panniculitic T-cell lymphoma; and (c) peripheral T-cell lymphomas that initially involve a lymph node paracortex and never grow into a true follicular pattern.
[0020]In one embodiment, the present invention includes a method for the treatment of multiple myeloma comprising administering to a patient diagnosed with having multiple myeloma a pharmaceutically acceptable composition comprising a therapeutically effective amount of 10-propargyl-10-deazaminopterin. In one embodiment, the 10-propargyl-10-deazaminopterin is substantially free of 10-deazaminopterin.
[0021]In one embodiment of the invention, the composition comprises "highly purified" 10-propargyl-10-dAM. As used in the specification and claims hereof, compositions which are "highly purified" contain 10-propargyl-10-dAM substantially free of other folic acid derivatives, particularly 10-deazaminopterin, which can interfere with the antitumor activity of the 10-propargyl-10-dAM. A composition within the scope of the invention may include carriers or excipients for formulating the 10-propargyl-10-dAM into a suitable dosage unit form for therapeutic use, as well as additional, non-folate therapeutic agents.
[0022]10-propargyl-10-dAM can be synthesized using the method disclosed in the DeGraw paper, supra or in Example 7 of U.S. Pat. No. 5,354,751, which is incorporated herein by reference. HPLC evaluation of the product prepared by this method shows the presence of a substantial amount (about 4.6%) of an impurity A (FIG. 2) which has a retention time consistent with 10-deazaminopterin. Thus, if this synthetic approach is employed further purification is necessary beyond that disclosed in the DeGraw et al. paper. Such purification can be carried out by additional HPLC or crystallization to remove the 10-deazaminopterin and other folic acid derivatives which may be present.
[0023]For use in the present invention, 10-propargyl-10-dAM is advantageously formulated as part of a pharmaceutical preparation. The specific dosage form will depend on the method of administration, but may include tablets, capsules, oral liquids, and injectable solutions for intravenous, intramuscular or intraperitoneal administration. One suitable dosing schedule involves the administration of 150 mg/m2 every two weeks. Lower doses may of course be indicated depending on the tolerance of an individual patient, or if more frequent administration were adopted. For example, doses on the order of 40 to 120 mg/m2 of body surface area/day are appropriate. Dosages of 30 mg/m2 weekly for 3 weeks followed by a one week rest, 30 mg/m2 weekly×6 weeks followed by a one week rest, or gradually increasing doses of 10-propargyl-10-dAM on the weekly×6 week schedule are also suitable. Higher doses could be utilized if less frequent administration were used. Thus, in a general sense, dosages of 30 to 275 mg/m2 are suitably used with various dosing schedules, for example 135 to 275 mg/m2 for biweekly dosages, and 30 to 150 mg/m2 for weekly dosages. The determination of suitable dosages using protocols similar to those described in U.S. Pat. No. 6,323,205, which is incorporated herein by reference, is within the skill in the art. In one embodiment, the 10-propargyl-10-deazaminopterin is administered in an amount of from about 30 to about 275 mg/m2 per dose. Methods of the present invention also include administration of 10-propargyl-10-deazaminopterin weekly; administration of 10-propargyl-10-deazaminopterin in a dose of about 30 mg/m2; administration of 10-propargyl-10-deazaminopterin in an amount of from about 30 to about 150 mg/m2 per dose; administration of 10-propargyl-10-deazaminopterin biweekly; and/or administering 10-propargyl-10-deazaminopterin in a dosage amount of about 135 to about 275 mg/m2.
[0024]10-propargyl-10-dAM may be used in combinations with other cytotoxic and antitumor compounds, including vinca alkaloids such as vinblastine, navelbine, and vindesine; probenicid, nucleotide analogs such as gemcitabine, 5-fluorouracil, and cytarabine; alkylating agents such as cyclophosphamide or ifosfamide; cisplatin or carboplatin; leucovorin; taxanes such a paclitaxel or docetaxel; anti-CD20 monoclonal antibodies, with or without radioisotopes, and antibiotics such as doxorubicin and mitomycin. Combinations of 10-propargyl-10-dAM with several of these other antitumor agents or with growth factor inhibitors and anti-angiogenic agents may also be used.
[0025]10-propargyl-10-dAM and other agents may be concurrently administered or utilized in combination as part of a common treatment regimen, in which the 10-propargyl-10-dAM and the other agent(s) are administered at different times. For example, the other agent may be administered before, immediately afterward or after a period of time (for example 24 hours) relative to the 10-propargyl-10-dAM administration. Thus, for purposes of this application, the term administering refers generally to concurrent administration or to sequential administration of the drugs and in either order in a parallel treatment regimen with or without a separation in time between the drugs unless otherwise specified.
[0026]10-propargyl-10-dAM is suitably used in combination with folic acid and vitamin B12 supplementation to reduce the side effects of the treatment. For example, patients may be treated with folic acid (1 mg/m2 daily starting 1 week prior to treatment with 10-propargyl-10-dAM, or alternatively 1 mg perioral (p.o.) daily not based on body surface area (BSA)); and B12 (1 mg/m2 monthly, or alternatively given intramuscularly (I.M.) every 8-10 weeks as 1 mg (not based on BSA), or alternatively p.o. daily 1 mg (not based on BSA)).
[0027]One embodiment of the present invention includes a method for assessing the sensitivity of a patient's cancer to treatment with 10-propargyl-10-deazaminopterin. This method includes the following steps, in any order. One step includes obtaining a sample of the patient's cancer tissue. Another step includes determining the amount of at least one selected polypeptide expressed by the sample. Another step includes obtaining a reference expression level for the at least one selected polypeptide for a cancer having sensitivity to 10-propargyl-10-deazaminopterin. Another step includes comparing the expression data for the at least one selected polypeptide with the reference expression for the at least one selected polypeptide. A match of the sample expression level of the at least one selected polypeptide to the reference expression level of the at least one selected polypeptide indicates the patient's cancer has increased sensitivity to 10-propargyl-10-deazaminopterin. Another step includes generating a report of the sensitivity of the sample to 10-propargyl-10-deazaminopterin. A report may be, without limitation, an oral report, a printed report, or an electronically transmitted report. Selected polypeptides include enzymes of any folate pathway in the cell and includes the polypeptides reduced folate carrier-1 enzyme (RFC-1), dihydrofolate reductase (DHFR), thymidylate synthase (TS), γ-glutamyl hydrolase (GGH) (also known as folypolyglutamate hydrolase (FPGH)), folylpoly-gamma-glutamate synthetase (FPGS), and glycinamide ribonucleotide formyltransferase (GARFT). Selected polypeptides are also variously referred to herein as "biomarkers of the invention."
[0028]Several proteins are implicated in the metabolism of folic acid and for the targets of anti-folates such as 10-propargyl-10-dAM and MTX in tumor cells. In most tumor cells, the protein encoded by RFC-1 mediates internalization of folate analogs. Once inside the cell, these analogs either bind dihydrofolate reductase (DHFR), thereby depleting intracellular reduced folate pools needed for purine and thymidine biosynthesis, or will be metabolized to a polyglutamate prior to binding to DHFR. Polyglutamylation is catalyzed by FPGS. FPGH (also known as GGH) mediates cleavage and clearance of these intracellular polyglutamated anti-folates. TS and GARFT are also involved in folate metabolism as "recycling" enzymes (thus directly affecting pools of nucleotides available for DNA synthesis). Without intending to be bound by a specific mechanism, it is believed that this correlation between RFC-1 expression levels and 10-propargyl-10-dAM sensitivity is a reflection of increased transport of 10-propargyl-10-dAM into tumor cells. Without being bound by theory, it is believed that alterations in other folate pathway enzymes discussed herein also correlate with 10-propargyl-10-dAM sensitivity; such as, for example, reduced DHFR levels correlating with a decrease in the amount of intracellular drug required to inhibit this enzyme, reduced GARFT and TS potentially reducing the pools of available nucleotides, increased FPGS increasing the rate of polyglutamylation of 10-propargyl-10-dAM and resulting in increased retention within the cell to facilitate ongoing activity against DHFR.
[0029]In another embodiment of the present invention, a method of selecting a patient for treatment of a cancer with 10-propargyl-10-deazaminopterin is provided. The method includes the following steps, in any order. One step includes obtaining a sample of the patient's cancer tissue. Another step includes determining the amount of at least one selected polypeptide expressed by the sample. Another step includes obtaining a reference expression level for the at least one selected polypeptide for a cancer having sensitivity to 10-propargyl-10-deazaminopterin. Another step includes comparing the expression data for the at least one selected polypeptide with the reference expression for the at least one selected polypeptide. A match of the sample expression level of the at least one selected polypeptide to the reference expression level of the at least one selected polypeptide indicates the patient's cancer has increased sensitivity to 10-propargyl-10-deazaminopterin. Selected polypeptides include reduced folate carrier-1 enzyme (RFC-1), dihydrofolate reductase (DHFR), thymidylate synthase (TS), γ-glutamyl hydrolase (GGH) (also known as folypolyglutamate hydrolase (FPGH)), folylpoly-gamma-glutamate synthetase (FPGS), and glycinamide ribonucleotide formyltransferase (GARFT). Another step includes selecting the patient for treatment 10-propargyl-10-deazaminopterin when the expression of the sample protein and the reference protein matches.
[0030]As used herein, the terms "protein" and "polypeptide" and "proteinaceous agent" are used interchangeably to refer to a chain of amino acids linked together by peptide bonds which optionally can comprise natural or non-natural amino acids. Optionally, the protein or peptide can comprise other molecules in addition to amino acids. Said chain can be of any length. Polypeptides of the present invention include enzymes related to folate pathways in cells including the selected polypeptides, including reduced folate carrier-1 enzyme (RFC-1), dihydrofolate reductase (DHFR), thymidylate synthase (TS), γ-glutamyl hydrolase (GGH) (also known as folypolyglutamate hydrolase (FPGH)), folylpoly-gamma-glutamate synthetase (FPGS), and glycinamide ribonucleotide formyltransferase (GARFT). The accession numbers and SEQ ID NOs of the selected polypeptides are as follows:
TABLE-US-00001 GenBank Poly- Accession SEQ ID peptide Full name Number NOs DHFR dihydrofolate reductase NM_000791 4, 5, 6 FPGS folylpolyglutamate synthetase M98045 13, 14, 15 GARFT glycinamide ribonucleotide X54199 16, 17, 18 transformylase GGH gamma-glutamyl hydrolase NM_003878 10, 11, 12 RFC-1 reduced folate carrier, member 1 NM_194255.1 1, 2, 3 TS Thymidylate synthase NM_001071 7, 8, 9
[0031]As used herein, nucleotide sequences of the gene products of the above identified selected polypeptides include, but are not limited to, the cDNA, genome-derived DNA and synthetic or semi-synthetic DNA or RNA. The full length gene nucleotide sequence of RFC-1 is contained in SEQ. ID. NO: 1; the full length gene nucleotide sequence of DHFR is contained in SEQ. ID. NO: 4, the full length gene nucleotide sequence of TS is contained in SEQ. ID. NO: 7, the full length gene nucleotide sequence of GGH is contained in SEQ. ID. NO: 10, the full length gene nucleotide sequence of FPGH is contained in SEQ. ID. NO: 13, and the full length gene nucleotide sequence of GARFT is contained in SEQ. ID. NO: 16.
[0032]The term "polynucleotide" is used to mean a polymeric form of nucleotides of any length, which contain deoxyribonucleotides, ribonucleotides, and/or their analogs. The terms "polynucleotide" and "nucleotide" as used herein are used interchangeably. Polynucleotides can have any three-dimensional structure, and can perform any function, known or unknown. The term "polynucleotide" includes double-stranded, single-stranded, and triple-helical molecules. Unless otherwise specified or required, any embodiment of the invention described herein that is a polynucleotide encompasses both the double-stranded form and each of two complementary single-stranded forms known or predicted to make up the double stranded form.
[0033]The following are non-limiting examples of polynucleotides: a gene or gene fragment, exons, introns, mRNA, tRNA, rRNA, ribozymes, cDNA, recombinant polynucleotides, branched polynucleotides, plasmids, vectors, isolated DNA of any sequence, isolated RNA of any sequence, nucleic acid probes, and primers. A polynucleotide can be comprised of modified nucleotides, such as methylated nucleotides and nucleotide analogs. Analogs of purines and pyrimidines are known in the art, and include, but are not limited to, aziridinylcytosine, 4-acetylcytosine, 5-fluorouracil, 5-bromouracil, 5-carboxymethylaminomethyl-2-thiouracil, 5-carboxymethyl-aminomethyluracil, inosine, N6-isopentenyladenine, 1-methyladenine, 1-methylpseudouracil, 1-methylguanine, 1-methylinosine, 2,2-dimethylguanine, 2-methyladenine, 2-methylguanine, 3-methylcytosine, 5-methylcytosine, pseudouracil, 5-pentynyluracil and 2,6-diaminopurine. The use of uracil as a substitute for thymine in a deoxyribonucleic acid is also considered an analogous form of pyrimidine.
[0034]A "fragment" (also called a "region") of a polynucleotide (i.e., a polynucleotide encoding a sarp) is a polynucleotide comprised of at least 9 contiguous nucleotides of the novel genes. Preferred fragments are comprised of a region encoding at least 5 contiguous amino acid residues, more preferably, at least 10 contiguous amino acid residues, and even more preferably at least 15 contiguous amino acid residues.
[0035]The term "recombinant" polynucleotide as used herein intends a polynucleotide of genomic, cDNA, semisynthetic, or synthetic in origin which, by virtue of its origin or manipulation: is not associated with all or a portion of a polynucleotide with which it is associated in nature; is linked to a polynucleotide other than that to which it is linked in nature; or does not occur in nature.
[0036]As used herein, reference to a selected gene product, protein or polypeptide in the present invention, including RFC-1 (SEQ ID NO:3), DHFR (SEQ ID NO:6), TS (SEQ ID NO:9), GGH, (also known as FPGH) (SEQ ID NO:12), FPGS (SEQ ID NO:15), and GARFT (SEQ ID NO:18), includes full-length proteins, fusion proteins, or any fragment or homologue of such a protein. The amino acid sequence for RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT from human are described herein as exemplary folate metabolism associated polypeptides and proteins. In addition, and by way of example, a "human RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT protein" refers to a RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT protein (generally including a homologue of a naturally occurring RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT protein) from a human (Homo sapiens) or to a RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT protein that has been otherwise produced from the knowledge of the structure (e.g., sequence) and perhaps the function of a naturally occurring RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT protein from Homo sapiens. In other words, a human RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT protein includes any RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT protein that has substantially similar structure and function of a naturally occurring RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT protein from Homo sapiens or that is a biologically active (i.e., has biological activity) homologue of a naturally occurring RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT protein from Homo sapiens as described in detail herein. As such, a human RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT protein can include purified, partially purified, recombinant, mutated/modified and synthetic proteins. According to the present invention, the terms "modification" and "mutation" can be used interchangeably, particularly with regard to the modifications/mutations to the amino acid sequence of RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT (or nucleic acid sequences) described herein.
[0037]As used herein, the term "homologue" is used to refer to a protein or peptide which differs from a naturally occurring protein or peptide (i.e., the "prototype" or "wild-type" protein) by minor modifications to the naturally occurring protein or peptide, but which maintains the basic protein and side chain structure of the naturally occurring form. Such changes include, but are not limited to: changes in one or a few amino acid side chains; changes one or a few amino acids, including deletions (e.g., a truncated version of the protein or peptide) insertions and/or substitutions; changes in stereochemistry of one or a few atoms; and/or minor derivatizations, including but not limited to: methylation, glycosylation, phosphorylation, acetylation, myristoylation, prenylation, palmitation, amidation and/or addition of glycosylphosphatidyl inositol. A homologue can have either enhanced, decreased, or substantially similar properties as compared to the naturally occurring protein or peptide. A homologue can include an agonist of a protein or an antagonist of a protein.
[0038]Homologues can be the result of natural allelic variation or natural mutation. A naturally occurring allelic variant of a nucleic acid encoding a protein is a gene that occurs at essentially the same locus (or loci) in the genome as the gene which encodes such protein, but which, due to natural variations caused by, for example, mutation or recombination, has a similar but not identical sequence. Allelic variants typically encode proteins having similar activity to that of the protein encoded by the gene to which they are being compared. One class of allelic variants can encode the same protein but have different nucleic acid sequences due to the degeneracy of the genetic code. Allelic variants can also comprise alterations in the 5' or 3' untranslated regions of the gene (e.g., in regulatory control regions). Allelic variants are well known to those skilled in the art.
[0039]Homologues can be produced using techniques known in the art for the production of proteins including, but not limited to, direct modifications to the isolated, naturally occurring protein, direct protein synthesis, or modifications to the nucleic acid sequence encoding the protein using, for example, classic or recombinant DNA techniques to effect random or targeted mutagenesis.
[0040]According to the present invention, an isolated RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT protein, including a biologically active homologue or fragment thereof, has at least one characteristic of biological activity of activity a wild-type, or naturally occurring RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT protein (which can vary depending on whether the homologue or fragment is an agonist, antagonist, or mimic of RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT, and the isoform of RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT).
[0041]Homologues of RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT, including peptide and non-peptide agonists and antagonists of RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT (analogues), can be products of drug design or selection and can be produced using various methods known in the art. Such homologues can be referred to as mimetics.
[0042]In one embodiment, a RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT homologue comprises, consists essentially of, or consists of, an amino acid sequence that is at least about 45%, or at least about 50%, or at least about 55%, or at least about 60%, or at least about 65%, or at least about 70%, or at least about 75%, or at least about 80%, or at least about 85%, or at least about 90%, or at least about 95% identical, or at least about 95% identical, or at least about 96% identical, or at least about 97% identical, or at least about 98% identical, or at least about 99% identical (or any percent identity between 45% and 99%, in whole integer increments), to a naturally occurring RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT amino acid sequence. A homologue of RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT differs from a reference (e.g., wild-type) RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT and therefore is less than 100% identical to the reference RFC-1, DHFR, TS, GGH, (also known as FPGH), FPGS, and GARFT at the amino acid level.
[0043]As used herein, unless otherwise specified, reference to a percent (%) identity refers to an evaluation of homology which is performed using: (1) a BLAST 2.0 Basic BLAST homology search using blastp for amino acid searches and blastn for nucleic acid searches with standard default parameters, wherein the query sequence is filtered for low complexity regions by default (described in Altschul, S. F., Madden, T. L., Sch{umlaut over (aa)}ffer, A. A., Zhang, J., Zhang, Z., Miller, W. & Lipman, D. J. (1997) "Gapped BLAST and PSI-BLAST: a new generation of protein database search programs." Nucleic Acids Res. 25:3389-3402, incorporated herein by reference in its entirety); (2) a BLAST 2 alignment (using the parameters described below); (3) and/or PSI-BLAST with the standard default parameters (Position-Specific Iterated BLAST. It is noted that due to some differences in the standard parameters between BLAST 2.0 Basic BLAST and BLAST 2, two specific sequences might be recognized as having significant homology using the BLAST 2 program, whereas a search performed in BLAST 2.0 Basic BLAST using one of the sequences as the query sequence may not identify the second sequence in the top matches. In addition, PSI-BLAST provides an automated, easy-to-use version of a "profile" search, which is a sensitive way to look for sequence homologues. The program first performs a gapped BLAST database search. The PSI-BLAST program uses the information from any significant alignments returned to construct a position-specific score matrix, which replaces the query sequence for the next round of database searching. Therefore, it is to be understood that percent identity can be determined by using any one of these programs.
[0044]The term, "primer", as used herein refers to an oligonucleotide, whether occurring naturally as in a purified restriction digest or produced synthetically, which is capable of acting as a point of initiation of synthesis when placed under conditions in which synthesis of a primer extension product, which is complementary to a nucleic acid strand, is induced, i.e., in the presence of nucleotides and an inducing agent such as a DNA polymerase and at a suitable temperature and pH. The primer may be either single-stranded or double-stranded and must be sufficiently long to prime the synthesis of the desired extension product in the presence of the inducing agent. The exact length of the primer will depend upon many factors, including temperature, source of primer and the method used. For example, for diagnostic applications, depending on the complexity of the target sequence, the oligonucleotide primer typically contains 15-25 or more nucleotides, although it may contain fewer nucleotides. The factors involved in determining the appropriate length of primer are readily known to one of ordinary skill in the art. In general, the design and selection of primers embodied by the instant invention is according to methods that are standard and well known in the art, see Dieffenbach, C. W., Lowe, T. M. J., Dveksler, G. S. (1995) General Concepts for PCR Primer Design. In: PCR Primer, A Laboratory Manual (Eds. Dieffenbach, C. W, and Dveksler, G. S.) Cold Spring Harbor Laboratory Press, New York, 133-155; Innis, M. A., and Gelfand, D. H. (1990) Optimization of PCRs. In: PCR protocols, A Guide to Methods and Applications (Eds. Innis, M. A., Gelfand, D. H., Sninsky, J. J., and White, T. J.) Academic Press, San Diego, 3-12; Sharrocks, A. D. (1994) The design of primers for PCR. In: PCR Technology, Current Innovations (Eds. Griffin, H. G., and Griffin, A. M, Ed.) CRC Press, London, 5-11.
[0045]As used herein, the terms "RNA portion" and "a portion thereof" in context of RNA products of a biomarker of the invention refer to an RNA transcript comprising a nucleic acid sequence of at least 6, at least 9, at least 15, at least 18, at least 21, at least 24, at least 30, at least 60, at least 90, at least 99, or at least 108, or more nucleotides of a RNA product of a biomarker of the invention.
[0046]Obtaining a sample of the patient's cancer tissue may be done by any methods known in the art. Bone marrow or lymph node biopsies and analysis of peripheral blood samples for cytogenetic and/or immunologic analysis is standard practice. Frozen tissue specimens may be obtained as well. As used herein a "sample" can be from any organism and can further include, but is not limited to, peripheral blood, plasma, urine, saliva, gastric secretion, feces, bone marrow specimens, primary tumors, metastatic tissue, embedded tissue sections, frozen tissue sections, cell preparations, cytological preparations, exfoliate samples (e.g., sputum), fine needle aspirations, amino cells, fresh tissue, dry tissue, and cultured cells or tissue. It is further contemplated that the biological sample of this invention can also be whole cells or cell organelles (e.g., nuclei). The sample can be unfixed or fixed according to standard protocols widely available in the art.
[0047]In some embodiments of the present invention, peripheral blood is drawn, or alternatively, if desired, leukocytes may be isolated by differential gradient separation, using, for example, ficoll-hypaque or sucrose gradient solutions for cell separations, followed by ammonium chloride or hypotonic lysis of remaining contaminating erythrocytes ("Cell Biology: A Laboratory Handbook", Volumes I-III Cellis, J. E., ed. (1994); "Current Protocols in Immunology" Volumes I-III Coligan J. E., ed. (1994); Stites et al. (eds)). Bone marrow and lymph node biopsies may be processed by collagenase/dispase treatment of the biopsy material, or by homogenization in order to obtain single cell suspensions ("Cell Biology: A Laboratory Handbook", Volumes I-III Cellis, J. E., ed. (1994); "Current Protocols in Immunology" Volumes I-III Coligan J. E., ed. (1994); Stites et al. (eds)).
[0048]The sample can be from a subject or a patient. As utilized herein, the "subject" or "patient" of the methods described herein can be any animal. In a preferred embodiment, the animal of the present invention is a human. In addition, determination of expression patterns is also contemplated for non-human animals which can include, but are not limited to, cats, dogs, birds, horses, cows, goats, sheep, guinea pigs, hamsters, gerbils, mice and rabbits.
[0049]The term "cancer," or "reference cancer", when used herein refers to or describes the pathological condition, preferably in a mammalian subject, that is typically characterized by unregulated cell growth. Non-limiting cancer types include carcinoma (e.g., adenocarcinoma), sarcoma, myeloma, leukemia, and lymphoma, and mixed types of cancers, such as adenosquamous carcinoma, mixed mesodermal tumor, carcinosarcoma, and teratocarcinoma. Representative cancers include, but are not limited to, bladder cancer, lung cancer, including NSCLC. Non-small cell lung cancer (NSCLC) is the most common type of lung cancer. It usually grows and spreads more slowly than small cell lung cancer. There are three forms of NSCLC: Adenocarcinomas are often found in an outer area of the lung. Squamous cell carcinomas are usually found in the center of the lung by an air tube (bronchus). Large cell carcinomas can occur in any part of the lung. Other cancers include breast cancer, colon cancer, rectal cancer, endometrial cancer, ovarian cancer; head and neck cancer, prostate cancer, and melanoma. Specifically included are AIDS-related cancers (e.g., Kaposi's Sarcoma, AIDS-related lymphoma), bone cancers (e.g., osteosarcoma, malignant fibrous histiocytoma of bone, Ewing's Sarcoma, and related cancers), and hematologic/blood cancers (e.g., adult acute lymphoblastic leukemia, childhood acute lymphoblastic leukemia, adult acute myeloid leukemia, childhood acute myeloid leukemia, chronic lymphocytic leukemia, chronic myelogenous leukemia, hairy cell leukemia, adult Hodgkin's disease, childhood Hodgkin's disease, Hodgkin's disease during pregnancy, adult non-Hodgkin's lymphoma, childhood non-Hodgkin's lymphoma, non-Hodgkin's lymphoma during pregnancy, primary central nervous system lymphoma, Waldenstrom's macroglobulinemia, multiple myeloma/plasmacell neoplasm, myelodysplastic syndrome, and myeloproliferative disorders), as well as lymphoblastic lymphomas in which the malignancy occurs in primitive lymphoid progenitors from the thymus; mature or peripheral T-cell neoplasms, including T-cell prolymphocytic leukemia, T-cell granular lymphocytic leukemia, aggressive NK-cell leukemia, cutaneous T-cell lymphoma (Mycosis fungoides/Sezary syndrome), anaplastic large cell lymphoma, T-cell type, enteropathy-type T-cell lymphoma, Adult T-cell leukemia/lymphoma including those associated with HTLV-1, and angioimmunoblastic T-cell lymphoma, and subcutaneous panniculitic T-cell lymphoma; and peripheral T-cell lymphomas that initially involve a lymph node paracortex and never grow into a true follicular pattern.
[0050]Also included are brain cancers (e.g., adult brain tumor, childhood brain stem glioma, childhood cerebellar astrocytoma, childhood cerebral astrocytoma, childhood ependymoma, childhood medulloblastoma, supratentorial primitive neuroectodermal and pineal, and childhood visual pathway and hypothalamic glioma), digestive/gastrointestinal cancers (e.g., anal cancer, extrahepatic bile duct cancer, gastrointestinal carcinoid tumor, colon cancer, esophageal cancer, gallbladder cancer, adult primary liver cancer, childhood liver cancer, pancreatic cancer, rectal cancer, small intestine cancer, and gastric cancer), musculoskeletal cancers (e.g., childhood rhabdomyosarcoma, adult soft tissue sarcoma, childhood soft tissue sarcoma, and uterine sarcoma), and endocrine cancers (e.g., adrenocortical carcinoma, gastrointestinal carcinoid tumor, islet cell carcinoma (endocrine pancreas), parathyroid cancer, pheochromocytoma, pituitary tumor, and thyroid cancer).
[0051]Also included are neurologic cancers (e.g., neuroblastoma, pituitary tumor, and primary central nervous system lymphoma), eye cancers (e.g., intraocular melanoma and retinoblastoma), genitourinary cancers (e.g., bladder cancer, kidney (renal cell) cancer, penile cancer, transitional cell renal pelvis and ureter cancer, testicular cancer, urethral cancer, Wilms' tumor and other childhood kidney tumors), respiratory/thoracic cancers (e.g., non-small cell lung cancer, small cell lung cancer, malignant mesothelioma, and malignant thymoma), germ cell cancers (e.g., childhood extracranial germ cell tumor and extragonadal germ cell tumor), skin cancers (e.g., melanoma, and merkel cell carcinoma), gynecologic cancers (e.g., cervical cancer, endometrial cancer, gestational trophoblastic tumor, ovarian epithelial cancer, ovarian germ cell tumor, ovarian low malignant potential tumor, uterine sarcoma, vaginal cancer, and vulvar cancer), and unknown primary cancers.
[0052]In one embodiment, the cancer is a lymphoma. In another embodiment, the cancer is a T-cell lymphoma. In yet another embodiment, the cancer is a multiple myeloma. In one embodiment, the sample and the reference cancer are both the same cancer sub-type, i.e., the sample cancer is derived from the same type of cell as the reference cancer. In another embodiment, the reference cancer is any one of or a combination of a cancer or cancerous cell line derived from a T-cell lymphoma or a multiple myeloma, such as, for example, lymphoblastic lymphomas in which the malignancy occurs in primitive lymphoid progenitors from the thymus; mature or peripheral T-cell neoplasms, including T-cell prolymphocytic leukemia, T-cell granular lymphocytic leukemia, aggressive NK-cell leukemia, cutaneous T-cell lymphoma (Mycosis fungoides/Sezary syndrome), anaplastic large cell lymphoma, T-cell type, enteropathy-type T-cell lymphoma, Adult T-cell leukemia/lymphoma including those associated with HTLV-1, and angioimmunoblastic T-cell lymphoma, and subcutaneous panniculitic T-cell lymphoma; and peripheral T-cell lymphomas that initially involve a lymph node paracortex.
[0053]In another embodiment of the present invention, the reference cancer or cancerous cell line is a reference cancer or cancerous cell line which is known to have a greater sensitivity to 10-propargyl-10-dAM. The term, "greater sensitivity," includes those cancers that are known or are found to have an enhanced response to 10-propargyl-10-dAM as compared to MTX. Increased sensitivity may be determined by those of skill in the art and may include assessment of effects seen in cell lines derived from that cancer and/or type of cancer, in animal models, such as mouse subcutaneous transplantation models, and therapeutic indicators such as remission or other indicia of reduced tumor burden in patients, such as increased apoptosis, decreased tumor volume, growth inhibition, and other indicia known to those in the art. An enhanced response can include differential effects seen at equivalent doses of, serum concentrations of, or other indicia of equivalence between, MDX and 10-propargyl-10-dAM.
[0054]The selected polypeptides may be quantitated and/or relative amounts determined by any method known in the art for quantitating and/or determining relative amounts of expression levels. The term, "quantitate" or "quantitation" also includes determination of relative amounts of a polypeptide or its transcript. Quantitating transcript RNA or portions thereof of a selected polypeptide is one such method. RNA may be extracted from biological samples via a number of standard techniques (see Current Protocols in Molecular Biology" Volumes I-III Ausubel, R. M., ed. (1994); Ausubel et al., "Current Protocols in Molecular Biology", John Wiley and Sons, Baltimore, Md. (1989)). Guanidium-based methods for cell lysis enabling RNA isolation, with subsequent cesium chloride step gradients for separation of the RNA from other cellular macromolecules, followed by RNA precipitation and resuspension, is an older, less commonly employed method of RNA isolation (Glisin, Ve. et al (1973) Biochemistry 13: 2633). Alternatively, RNA may be isolated in a single step procedure (U.S. Pat. No. 4,843,155, and Puissant, C. and Houdebine L. M. (1990) Biotechniques 8: 148-149). Single step procedures include the use of Guanidium isothiocyanate for RNA extraction, and subsequent phenol/chloroform/isoamyl alcohol extractions facilitating the separation of total RNA from other cellular proteins and DNA. Commercially available single-step formulations based on the above-cited principles may be employed, including, for example, the use of the TRIZOL reagent (Life Technologies, Gaithersburg, Md.).
[0055]According to further features of preferred embodiments of the present invention, monitoring selected polypeptide RNA/gene expression is via a number of standard techniques well described in the art, any of which can be employed to evaluate selected polypeptide expression. These assays comprise Northern blot and dot blot analysis, primer extension, RNase protection, RT-PCR, in-situ hybridization and chip hybridization. Specific selected polypeptide RNA sequences can be readily detected by hybridization of labeled probes to blotted RNA preparations extracted as above. In Northern blot analysis, fractionated RNA is subjected to denaturing agarose gel electrophoresis, which prevents RNA from assuming secondary structures that might inhibit size based separation. RNA is then transferred by capillary transfer to a nylon or nitrocellulose membrane support and may be probed with a labeled oligonucleotide probe complementary to the selected polypeptide sequence (Alwine, et al. (1977). Proc. Natl. Acad. Sci. USA 74: 5350-5354 and Current Protocols in Molecular Biology" Volumes I-III Ausubel, R. M., ed. (1994); Ausubel et al., "Current Protocols in Molecular Biology", John Wiley and Sons, Baltimore, Md. (1989)).
[0056]Alternatively, unfractionated RNA may be immobilized on a nylon or nitrocellulose membrane, and similarly probed for selected polypeptide-specific expression, by Slot/Dot blot analysis. RNA slot/dot blots can be prepared by hand, or alternatively constructed using a manifold apparatus, which facilitates comparing hybridization signals by densitometry scanning (Chomczynski P. (1992) Anal. Biochem. 201: 134-139). Primer extension is an additional means whereby quantification of the RNA may be accomplished. Primer extension provides an additional benefit in mapping the 5' terminus of a particular RNA, by extending a primer using the enzyme reverse transcriptase. In this case, the primer is an oligonucleotide (or restriction fragment) complementary to a portion of the selected polypeptide mRNA. The primer is end-labeled, and is allowed to hybridize to template selected polypeptide mRNA. Once hybridized, the primer is extended by addition of reverse transcriptase, and incorporation of unlabeled dexoynucleotides to for a single-stranded DNA complementary to template selected polypeptide mRNA. DNA is then analyzed on a sequencing gel, with the length of extended primer serving to map the 5' position of the mRNA, and the yield of extended product reflecting the abundance of RNA in the sample (Jones et al (1985) Cell 42: 559-572 and Micrendorf R. C. And Pfeffer, D. (1987). Methods Enzymol. 152: 563-566).
[0057]RNase protection assays provide a highly sensitive means of quantifying selected polypeptide RNA, even in low abundance. In protection assays, sequence-specific hybridization of ribonucleotide probes complementary to selected polypeptide RNA, with high specific activity are generated, and hybridized to sample RNA. Hybridization reactions are then treated with ribonuclease to remove free probe, leaving intact fragments of annealed probe hybridized to homologous selected polypeptide sequences in sample RNA. Fragments are then analyzed by electrophoresis on a sequencing gel, when appropriately-sized probe fragments are visualized (Zinn K. et al (1983) Cell 34: 865-879 and Melton S. A., et al (1984). Nucl. Acids Res. 12: 7035-7056).
[0058]RT-PCR is another means by which selected polypeptide expression is verified. RT-PCR is a particularly useful method for detecting rare transcripts, or transcripts in low abundance. RT-PCR employs the use of the enzyme reverse transcriptase to prepare cDNA from RNA samples, using deoxynucleotide primers complementary to the selected polypeptide mRNA. Once the cDNA is generated, it is amplified through the polymerase chain reaction, by the addition of deoxynucleotides and a DNA polymerase that functions at high temperatures. Through repetitive cycles of primer annealing, incorporation of deoxynucleotides facilitating cDNA extension, followed by strand denaturation, amplification of the desired sequence occurs, yielding an appropriately sized fragment that may be detected by agarose gel electrophoresis. Optimal reverse transcription, hybridization, and amplification conditions will vary depending upon the sequence composition and length(s) of the primers and target(s) employed, and the experimental method selected by the practitioner. Various guidelines may be used to select appropriate primer sequences and hybridization conditions (see, e.g., Sambrook et al., 1989, Molecular Cloning, A Laboratory Manual, (Volumes 1-3) Cold Spring Harbor Press, N.Y.; and Ausubel et al., 1989, Current Protocols in Molecular Biology, Green Publishing Associates and Wiley Interscience, N.Y.).
[0059]In-situ hybridization provides another tool for the detection and localization of cell/tissue specific selected polypeptide RNA expression. Labeled anti-sense RNA probes are hybridized to mRNAs in cells singly, or in processed tissue slices, which are immobilized on microscope glass slides (In Situ Hybridization: Medical Applications (eds. G. R. Coulton and J. de Belleroche), Kluwer Academic Publishers, Boston (1992); In Situ Hybridization: In Neurobiology; Advances in Methodology (eds. J. H. Eberwine, K. L. Valentino, and J. D. Barchas), Oxford University Press Inc., England (1994); and In Situ Hybridization: A Practical Approach (ed. D. G. Wilkinson), Oxford University Press Inc., England (1992)). Numerous non-isotopic systems have been developed to visualize labeled DNA probes including; a) fluorescence-based direct detection methods, b) the use of digoxigenin- and biotin-labeled DNA probes coupled with fluorescence detection methods, and c) the use of digoxigenin- and biotin-labeled DNA probes coupled with antibody-enzyme detection methods. When fluorescence-labeled anti-sense RNA probes are hybridized to cellular RNA, the hybridized probes can be viewed directly using a fluorescence microscope. Direct fluorochrome-labeling of the nucleic acid probes eliminate the need for multi-layer detection procedures (e.g., antibody-based-systems), which allows fast processing and also reduces non-specific background signals, hence providing a versatile and highly sensitive means of identifying selected polypeptide gene expression.
[0060]Chip hybridization utilizes selected polypeptide-specific oligonucleotides attached to a solid substrate, which may consist of a particulate solid phase such as nylon filters, glass slides or silicon chips [Schena et al. (1995) Science 270:467-470] designed as a microarray. Microarrays are known in the art and consist of a surface to which probes that correspond in sequence to gene products (such as cDNAs) can be specifically hybridized or bound at a known position for the detection of selected polypeptide gene expression. Quantification of the hybridization complexes is well known in the art and may be achieved by any one of several approaches. These approaches are generally based on the detection of a label or marker, such as any radioactive, fluorescent, biological or enzymatic tags or labels of standard use in the art. A label can be applied to either the oligonucleotide probes or the RNA derived from the biological sample.
[0061]In general, mRNA quantification is preferably effected alongside a calibration curve so as to enable accurate mRNA determination. Furthermore, quantifying transcript(s) originating from a biological sample is preferably effected by comparison to a normal sample, which sample is characterized by normal expression pattern of the examined transcript(s).
[0062]Selected polypeptide expression may also be evaluated at the level of protein expression, either by demonstration of the presence of the protein, or by its activity, with activity herein referring to the enzymatic activity of the selected polypeptide enzyme. Methods for monitoring specific polypeptide protein expression include the following methods discussed below. Anti-selected polypeptide-antibodies for use in selected polypeptide-specific protein detection are readily generated by methods known in the art and include both polyclonal and monoclonal antibodies. The antibodies preferably bind to both native and denatured selected polypeptides and may be detected by several well-known assays in the art, including ELISA, RIA, light emission immunoassays, Western blot analysis, immunofluorescence assays, immunohistochemistry and FACS analysis.
[0063]Enzyme linked immunosorbant (ELISA) assays and radioimmunoassays (RIA) follow similar principles for detection of specific antigens, in this case, selected polypeptides. In RIA a selected polypeptide-specific antibody is radioactively labeled, typically with 125I. In ELISA assays a selected polypeptide-specific antibody is chemically linked to an enzyme. Selected polypeptide-specific capturing antibody is immobilized onto a solid support. Unlabelled specimens, e.g., protein extracts from biopsy or blood samples are then incubated with the immobilized antibody under conditions where non-specific binding is blocked, and unbound antibody and/or protein removed by washing. Bound selected polypeptide is detected by a second selected polypeptide-specific labeled antibody. Antibody binding is measured directly in RIA by measuring radioactivity, while in ELISA binding is detected by a reaction converting a colorless substrate into a colored reaction product, as a function of linked-enzyme activity. Changes can thus readily be detected by spectrophotometry (Janeway C. A. et al (1997). "Immunbiology" 3rd Edition, Current Biology Ltd., Garland Publishing Inc.; "Cell Biology: A Laboratory Handbook", Volumes I-III Cellis, J. E., ed. (1994); "Current Protocols in Immunology" Volumes I-III Coligan J. E., ed. (1994); Stites et al. (eds)). Both assays therefore provide a means of quantification of selected polypeptide protein content in a biological sample.
[0064]Selected polypeptide protein expression may also be detected via light emission immunoassays. Much like ELISA and RIA, in light emission immunoassays the biological sample/protein extract to be tested is immobilized on a solid support, and probed with a specific label, labeled anti- selected polypeptide antibody. The label, in turn, is luminescent, and emits light upon binding, as an indication of specific recognition. Luminescent labels include substances that emit light upon activation by electromagnetic radiation, electro chemical excitation, or chemical activation and may include fluorescent and phosphorescent substances, scintillators, and chemiluminescent substances. The label can be a part of a catalytic reaction system such as enzymes, enzyme fragments, enzyme substrates, enzyme inhibitors, coenzymes, or catalysts; part of a chromogen system such as fluorophores, dyes, chemiluminescers, luminescers, or sensitizers; a dispersible particle that can be non-magnetic or magnetic, a solid support, a liposome, a ligand, a receptor, a hapten radioactive isotope, and so forth (U.S. Pat. Nos. 6,410,696, U.S. Pat. No. 4,652,533 and European Patent Application No. 0,345,776), and provide an additional, highly sensitive method for detection of selected polypeptide protein expression.
[0065]Western blot analysis is another means of assessing selected polypeptide content in a biological sample. Protein extracts from biological samples of, for example, hematopoietic cells, are solubilized in a denaturing ionizing environment, and aliquots are applied to polyacrylamide gel matrixes. Proteins separate based on molecular size properties as they migrate toward the anode. Antigens are then transferred to nitrocellulose, PVDF or nylon membranes, followed by membrane blocking to minimize non-specific binding. Membranes are probed with antibodies directly coupled to a detectable moiety, or are subsequently probed with a secondary antibody containing the detectable moiety. Typically the enzymes horseradish peroxidase or alkaline phosphatase are coupled to the antibodies, and chromogenic or luminescent substrates are used to visualize activity (Harlow E. et al (1998) Immunoblotting. In Antibodies: A Laboratory Manual, pp. 471-510 CSH Laboratory, cold Spring Harbor, N.Y. and Bronstein I. Et al. (1992) Biotechniques 12: 748-753). Unlike RIA, ELISA, light emission immunoassays and immunoblotting, which quantify selected polypeptide content in whole samples, immunofluorescence/immunocytochemistry may be used to detect proteins in a cell-specific manner, though quantification is compromised.
[0066]In some steps of the methods of the present invention, the level of expression of the RNA and/or protein products of one or more biomarkers of the invention, as measured by the amount or level of RNA or protein, is compared to see if the level of expression "matches." The term "match" indicates that the level of expression of mRNA, and/or one or more spliced variants of mRNA of the biomarker in the sample is compared with the level of expression of the same one or more biomarkers of the invention as measured by the amount or level of RNA, including mRNA and/or one or more spliced variants of mRNA in a reference sample, and is determined to be similar, for example, by one of skill in the art and/or in accordance with the discussion hereinbelow. A "match" can also include a measurement of the protein, or one or more protein variants encoded by the biomarker of the invention in the sample as compared with the amount or level of protein expression, including one or more protein variants of the biomarker or biomarkers of the invention in the reference sample. A match may be determined by comparing a first population of samples as compared with a second population of samples or a single sample to a reference using either a ratio of the level of expression or using p-value. When using p-value, a nucleic acid transcript including hnRNA and mRNA is identified as being differentially expressed as between a first and second population when the p-value is less than 0.1, less than 0.05, less than 0.01, less than 0.005, less than 0.001 etc. A "match" indicating that the level of expression of the biomarker or selected polypeptide of the sample or population of samples is similar to the level of expression of the biomarker or selected polypeptide of the reference may be determined by one of skill in the art, and includes a level of expression in the sample that is at least about 20%, at least about 25%, at least about 30%, at least about 35%, at least about 40%, at least about 45%, at least about 50%, at least about 55%, at least about 60%, at least about 65%, at least about 70%, at least about 75%, at least about 80%, at least about 85%, at least about 90%, at least about 98%, at least about 100%, at least about 110%, at least about 120%, at least about 130%, at least about 140%, at least about 150%, at least about 160%, at least about 170%, at least about 180%, at least about 200%, at least about three fold, at least about four fold, at least about five fold, at least about ten fold, of the reference.
[0067]In another embodiment, the present invention includes a method to modulate the expression of a selected polypeptide in a patient's cancer comprising administering to a patient an effective amount of 10-propargyl-10-deazaminopterin, wherein the selected polypcptide is selected from the group consisting reduced folate carrier-1 enzyme (RFC-1), dihydrofolate reductase (DHFR), thymidylate synthase (TS), γ-glutamyl hydrolase (GGH) (also known as folypolyglutamate hydrolase (FPGH)), folylpoly-gamma-glutamate synthetase (FPGS), and glycinamide ribonucleotide formyltransferase (GARFT). A patient's cancer can include any cancer. In one embodiment, the patient's cancer is a lymphoma; in one embodiment, the patient's cancer is a T-cell lymphoma; in another embodiment, the patient's cancer is multiple myeloma. In another embodiment, the patient's cancer is a NSCLC. The modulation can occur in vitro and/or in vivo.
[0068]Modulation includes both up-regulation and down-regulation. As used herein, the term "up regulated" or "increased level of expression" in the context of this invention refers to a sequence corresponding to a gene which is expressed wherein the measure of the quantity of the sequence demonstrates an increased level of expression of the gene in the patient as compared to prior to administration of 10-propargyl-10-deazaminopterin, and can be observed at any point in treatment with 10-propargyl-10-deazaminopterin. An "increased level of expression" according to the present invention, is an increase in expression of at least 1%, 2%, 3%, 4%, 5%, 6%, 7%, 8%, 9%, 10% or more, for example, 20%, 30%, 40%, or 50%, 60%, 70%, 80%, 90% or more, or greater than 1-fold, up to 2-fold, 3-fold, 4-fold, 5-fold, 10-fold, 50-fold, 100-fold or more. In one embodiment, the polypeptide that is up-regulated is RFC-1 and the cancer is a T-cell lymphoma, multiple myeloma, or a NSCLC.
[0069]"Down regulation" or "decreased level of expression" in the context of this invention refers to a sequence corresponding to a gene which is expressed wherein the measure of the quantity of the sequence demonstrates a decreased level of expression of the gene in the patient as compared to prior to administration of 10-propargyl-10-deazaminopterin, and can be observed at any point in treatment with 10-propargyl-10-deazaminopterin. A "decreased level of expression" according to the present invention, is a decrease in expression of at least 1%, 2%, 3%, 4%, 5%, 6%, 7%, 8%, 9%, 10% or more, for example, 20%, 30%, 40%, or 50%, 60%, 70%, 80%, 90% or more, or greater than 1-fold, up to 2-fold, 3-fold, 4-fold, 5-fold, 10-fold, 50-fold, 100-fold or more. In one embodiment, the 10-propargyl-10-deazaminopterin is substantially free of 10-deazaminopterin. In one embodiment, the modulation is down-regulation, and the downregulation of expression of polypeptide is TS and/or DFHR, and the cancer is NSCLC, T-cell lymphoma, or multiple myeloma.
[0070]In one embodiment, kits are provided for measuring a RNA product of a biomarker of the invention which comprise materials and reagents that are necessary for measuring the expression of the RNA product. For example, a microarray or RT-PCR kit may be used and contain only those reagents and materials necessary for measuring the levels of RNA products of any number of up to at least 1, at least 2, at least 3, at least 4, at least 5, at least 6, all or any combination of the biomarkers of the invention. Alternatively, in some embodiments, the kits can comprise materials and reagents that are not limited to those required to measure the levels of RNA products of any number of up to 1, 2, 3, 4, 5, 6, all or any combination of the biomarkers of the invention. For example, a microarray kit may contain reagents and materials necessary for measuring the levels of RNA products any number of up to 1, 2, 3, 4, 5, 6, all or any combination of the biomarkers of the invention, in addition to reagents and materials necessary for measuring the levels of the RNA products of any number of up to at least 1, at least 2, at least 3, at least 4, at least 5, at least 6, or more genes other than the biomarkers of the invention. In a specific embodiment, a microarray or RT-PCR kit contains reagents and materials necessary for measuring the levels of RNA products of any number of up to at least 1, at least 2, at least 3, at least 4, at least 5, at least 6, all or any combination of the biomarkers of the invention, and any number of up to 1, 2, 3, 4, 5, 10 or more genes that are not biomarkers of the invention.
[0071]For nucleic acid microarray kits, the kits generally comprise probes attached to a support surface. The probes may be labeled with a detectable label. In a specific embodiment, the probes are specific for the 5' region, the 3' region, the internal coding region, an exon(s), an intron(s), an exon junction(s), or an exon-intron junction(s), of any number of up to 1, 2, 3, 4, 5, 6, all or any combination of the biomarkers of the invention. The microarray kits may comprise instructions for performing the assay and methods for interpreting and analyzing the data resulting from the performance of the assay. The kits may also comprise hybridization reagents and/or reagents necessary for detecting a signal produced when a probe hybridizes to a target nucleic acid sequence. Generally, the materials and reagents for the microarray kits are in one or more containers. Each component of the kit is generally in its own a suitable container.
[0072]For RT-PCR kits, the kits generally comprise pre-selected primers specific for particular RNA products (e.g., an exon(s), an intron(s), an exon junction(s), and an exon-intron junction(s)) of any number of up to 1, 2, 3, 4, 5, 6, all or any combination of the biomarkers of the invention. The RT-PCR kits may also comprise enzymes suitable for reverse transcribing and/or amplifying nucleic acids (e.g., polymerases such as Taq), and deoxynucleotides and buffers needed for the reaction mixture for reverse transcription and amplification. The RT-PCR kits may also comprise probes specific for any number of up to 1, 2, 3, 4, 5, 6, all or any combination of the biomarkers of the invention. The probes may or may not be labeled with a detectable label (e.g., a fluorescent label). Each component of the RT-PCR kit is generally in its own suitable container. Thus, these kits generally comprise distinct containers suitable for each individual reagent, enzyme, primer and probe. Further, the RT-PCR kits may comprise instructions for performing the assay and methods for interpreting and analyzing the data resulting from the performance of the assay.
[0073]For antibody based kits, the kit can comprise, for example: (1) a first antibody (which may or may not be attached to a support) which binds to protein of interest (e.g., a protein product of any number of up to 1, 2, 3, 4, 5, 6, all or any combination of the biomarkers of the invention); and, optionally, (2) a second, different antibody which binds to either the protein, or the first antibody and is conjugated to a detectable label (e.g., a fluorescent label, radioactive isotope or enzyme). The antibody-based kits may also comprise beads for conducting an immunoprecipitation. Each component of the antibody-based kits is generally in its own suitable container. Thus, these kits generally comprise distinct containers suitable for each antibody. Further, the antibody-based kits may comprise instructions for performing the assay and methods for interpreting and analyzing the data resulting from the performance of the assay.
[0074]Additional objects, advantages, and novel features of the present invention will become apparent to one ordinarily skilled in the art upon examination of the following examples, which are not intended to be limiting. Additionally, each of the various embodiments and aspects of the present invention as delineated hereinabove and as claimed in the claims section below finds experimental support in the following examples.
EXAMPLES
[0075]The following examples are provided for illustrative purposes only and are not intended to limit the scope of the invention.
Example 1
[0076]FIG. 4 shows a synthetic scheme useful in preparing 10-propargyl-10-dAM in accordance with the invention. A mixture of 60% NaH in oil dispersion (1.06 g, 26.5 mmol) in 18 mL of sieve-dried THF was cooled to 0° C. The cold mixture was treated with a solution of homoterephthalic acid dimethyl ester (5.0 g, 24 mmol. compound 1 in FIG. 4) in dry THF (7 mL), and the mixture was stirred for 1 hour at 0° C. Propargyl bromide (26.4 mmol) was added, and the mixture was stirred at 00° C. for an additional 1 hour, and then at room temperature for 16 hours. The resulting mixture was treated with 2.4 mL of 50% acetic acid and then poured into 240 mL of water. The mixture was extracted with ether (2×150 mL). The ether extracts were combined, dried over Na2SO4, and concentrated to an orange-yellow oil. Chromatography on silica gel (600 mL of 230-400 mesh) with elution by cyclohexane-EtOAc (8:1) gave the product α-propargylhomoterephthalic acid dimethyl ester (compound 2) as a white solid (4.66) which appeared by TLC (cyclohexane-EtOAc, 3:1) to be homogeneous. Mass spectral data on this product, however, showed it to be a mixture of the desired product 2, and the dipropargylated compound. No starting material 1 was detected. HPLC shows the ratio of mono- to dipropargylated products to be about 3:1. Since the dipropargylated product, unlike compound 1, cannot produce an unwanted coproduct in the next step of the reaction, this material was suitable for conversion to compound 3. Absence of starting compound 1 in the product used to proceed in the synthesis is very important in order to avoid the sequential formation of 10-dAM during the transformations lading to the final product, because complete removal from 10-dAM from 10-propargyl-1-dAM is very difficult.
[0077]A mixture was formed by combining 0.36 g of a 60% NaH (9 mmol) in oil dispersion with 10 mL of dry DMF and cooled to 0-5° C. The cold mixture was treated drop-wise with a solution of the product of the first reaction (compound 2) (2.94 g, 12 mmol) in 10 mL dry DMF and then stirred at 0° C. for 30 minutes. After cooling to -25° C., a solution of 2,4,diamino-6-(bromomethyl)-pteridine hydrobromide-0.2 2-propanol (1.00 g, 2.9 mmol) in 10 mL dry DMF was added drop-wise while the temperature was maintained near -25° C. The temperature of the stirred mixture was allowed to rise to -10° C. over a period of 2 hours. After an additional 2 hours at -10° C., the temperature was allowed to rise to 20° C., stirring at room temperature was continued for 2 hours longer. The reaction was then adjusted to pH 7 by addition of solid CO2, After concentration in vacuo to remove solvent, the residue was stirred with diethyl ether and the ether insoluble material was collected, washed with water, and dried in vacuo to give 1.49 g of a crude product. This crude product was dissolved in CHCl3-MeOH (10:1) for application to a silica gel column. Elution by the same solvent system afforded 10-propargyl-10-carbomethoxy-4-deoxy-4-a-mino-10-deazapteroic acid methyl ester (compound 3) which was homogenous to TLC in 40% yield (485 mg).
[0078]A stirred suspension of compound 3 (400 mg, 0.95 mmol) in 2-methoxyethanol (5 mL) was treated with water (5 mL) and then 10% sodium hydroxide solution (3.9 mL). The mixture was stirred as room temperature for 4 hours, during which time solution occurred. The solution was adjusted to pH 8 with acetic acid and concentrated under high vacuum. The resulting residue was dissolved in 15 mL of water and acidified to pH 5.5-5.8 resulting in formation of a precipitate. The precipitate was collected, washed with water and dried in vacuo to recover 340 mg of compound 4 (91% yield). HPLC analysis indicated a product purity of 90%.
[0079]Compound 4 (330 mg) was decarboxylated by heating in 15 mL DMSO at 115-120° C. for 10 minutes. A test by HPLC after 10 minutes confirmed that the conversion was essentially complete. DMSO was removed by distillation in vacuo (bath at 40° C.). The residue was stirred with 0.5 N NaOH to give a clear solution, Acidification to pH 5.0 with 1 N HCl gave 10-propargyl-4-deoxy-4-amino-10-deazapteroic acid (compound 5) as a yellow solid in 70% yield. HPLC indicated product purity at this stage as 90%.
[0080]Compound 5 (225 mg, 0.65 mmol) was coupled with dimethyl L-glutamate hydrochloride (137 mg, 0.65 mmol) using BOP reagent (benzotriazole-1-yloxytris(dimethylamino) phosphonium hexafluorophosphate (287 mg, 0.65 mmol, Aldrich Chemical Co.) in DMF (10 mL) containing triethylamine (148 mg, 1.46 mmol). The mixture was stirred for 3 hours at 20-25° C. and then evaporated to dryness. The residue was stirred with water, and the water-insoluble crude product was collected and dried in vacuo. The crude product (350 mg) was purified by silica gel chromatography with elution by CHCl3-MeOH (10:1) containing triethylamine (0.25% by volume) to recover 165 mg of 10-propargyl-10-deazaminopterin dimethyl ester (compound 6, 50% yield) which was homogeneous to TLC(CHCl3-MeOH 5:1).
[0081]Compound 6 (165 mg, 0.326 mmol) was suspended in 10 mL stirred Me0H to which 0.72 mL (0.72 meq) 1N NaOH was added. Stirring at room temperature was continued until solution occurred after a few hours. The solution was kept at 20-25°. for 8 hours, then diluted with 10 mL water. Evaporation under reduced pressure removed the methanol, and the concentrated aqueous solution was left at 20-25° C. for another 24 hours. HPLC then showed the ester hydrolysis to be complete. The clear aqueous solution was acidified with acetic acid to pH 4.0 to precipitate 10-propargyl-10-deazaminopterin as a pale yellow solid, The collected, water washed and dried in vacuo product weighed 122 mg (79% yield). Assay by elemental analysis, proton NMR and mass spectroscopy were entirely consistent with the assigned structure. HPLC analysis indicated purity of 98% and established the product to be free of 10-deazaminopterin.
[0082]FIG. 3 shows an HPLC of a highly purified preparation consisting essentially of 10-propargyl-10-dAM in accordance with the invention prepared using the method described in Example 1. In this case, the amount of 10-propargyl-10-dAM (as determined by HPLC peak area) approaches 98%, and the peak corresponding to 10-deazaminopterin is not detected by the processing software although there is a minor baseline ripple in this area.
Example 2
[0083]This example describes testing of 10-propargyl-10-dAM and MTX for cytotoxicity against human lymphoma cell lines.
[0084]10-propargyl-10-dAM preparation prepared in accordance with Example 1 and an MTX preparation were tested for cytotoxicity against a panel of five human lymphoma cell lines. Experiments were performed as described previously. (Sirotnak et al., Cancer Chemother. Pharmacol. 12: 18-25 (1984). In brief, 2.5 to 5×103 cells were plated per well in 96-well flat bottom plates. Drug was added in a 0.9% NaCl solution (pH 7.0) over a range of concentrations, and cells were continuously exposed to drug for 5 days. Colorimetric dye (XTT or Alamar blue) was added for an addition period of time (XTT dye, 6 hours, Alamar blue, 24 hours). Each plate was then read on an automated plate reader at 590 nm. The percentage of inhibition was calculated as growth of cells exposed to drug divided by growth of controls (cells incubated with media only). IC50 values were determined as the drug concentrations at which cell growth was inhibited 50% as compared to controls. Experiments were repeated as least three times. Experiments were also conducted with continuous drug exposures lasting for 3 and 4 days with results similar to the 5 day results. The results of the study with 5 day drug exposures is summarized in Table 1. As shown, in every instance, the IC50 of 10-propargyl-10-dAM was substantially lower than the IC50 for MTX, indicating greater potency and/or the ability to use lower and therefore less toxic amounts to achieve the same efficacy.
TABLE-US-00002 TABLE 1 Relative growth inhibition in vitro Cell Line Lymphoma Type IC50 PDX (nM) IC50 MTX (nM) Hs445 Hodgkin's disease 1.6 ± 0.8 32 ± 2.2 HT Diffuse large B-cell 2.0 ± 0.4 35 ± 5.0 Raji Burkitt's 2.0 ± 0.3 16 ± 0.8 RL Transformed follicular 23.0 ± 2.0 210 ± 40.sup. SKI-DLCL-1 Diffuse large B-cell 5.1 ± 0.1 48 ± 2.5
Example 3
[0085]This example describes the effects of 10-propargyl-10-dAM used in a Phase I/II study on T-cell lymphomas.
[0086]In this study, patients with aggressive lymphoma were enrolled, including three patients with drug-resistant T-cell lymphoma. The following case summaries have been obtained. Each of these patients was also treated with folic acid (1 mg/m2 daily starting 1 week prior to treatment with 10-propargyl-10-dAM) and Vitamin B12 (1 mg/mg/m2 monthly) supplementation.
[0087]Patient 1 had a diagnosis of Peripheral T-cell Lymphoma, Stage 1V. Demographics: 48 Year old male; Prior Treatment: CHOP×4 cycles (July 2002-November 2002)--refractory, ICE×2 cycles (December 2002)--refractory, Campath (March 2003-June 2003)--mixed response; Pre-Treatment Staging: Extensive disease cutaneous disease. Treatment on Study: 10-propargyl-10-dAM 135 mg/m2 times.1 dose. Toxicitics observed wcrc Grade 3 stomatitis; neutropenia grade 3; sepsis; Response: Essentially complete remission by PET scan. Comment: This patient ultimately died after developing a bacteremia and sepsis from open skin lesions with Gram positive bacteria.
[0088]Patient 2 had a diagnosis of Lymphoblastic Lymphoma, Precursor T-cell, Stage IV. Demographics: 65 year old female; Prior Treatment: L20--Complex combination chemotherapy since May 2002, administered over two years. Has received MTX from May 2002 through February 2004. Relapsed December 2004. Pre-Treatment Staging: Extensive widespread relapse. Treatment on Study: 10-propargyl-10-dAM 30 mg/m2 2×3 weeks every 4 weeks. Completed 3 cycles to date. Toxicities: None Response: Complete remission by PET and CT. Comment: Patient with essentially methotrexate resistant disease with extensive sinus based disease which began resolving after one dose of 10-propargyl-10-dAM.
[0089]Patient 3 had a diagnosis of HTLV Associated T-cell Lymphoma. Demographics: 38 Year old male; Prior Treatment: EPOCH--infusional combination chemotherapy October 2003 to February 2004. Pre-Treatment Staging: Left axillary disease. Treatment on Study: 10-propargyl-10-dAM 30 mg/m2 weekly×3 every 4 weeks×2 cycles; Toxicities: None. Response: Complete remission. Comment: Complete disappearance of clinically evident disease by the end of the first cycle, very well tolerated, no toxicity.
[0090]Patient 4 had a diagnosis of Panniculitic T-cell Lymphoma. Demographics: 25 Year old male; Prior Treatment: Ontak (refractory), September 2002-November 2002; Targretin and IFNα January 2003-October 2003 (durable partial remission); CHOP April 2004-June 2004; ICE June 2004, CyPen July 2004-August 2004, Targretin/MTX September 2004 to February 2005; Treatment on Study: 10-propargyl-10-dAM 30 mg/m2 weekly×4. Response: Clinical complete remission by PET; Toxicities: None; Comment: healing subcutaneous lesions, too numerous to count, large ulcerative granulating lesion.
[0091]This Example shows that the 4 patients with T-cell lymphoma treated with 10-propargyl-10-dAM in this example have all met criteria for complete remission, even based on the sensitive PET imaging techniques. Interestingly, the patient treated at 135 mg/m2 received only a single dose of drug with a dramatic response to therapy, while the others had received only small modest doses on a weekly schedule.
Example 4
[0092]This example describes the effects of 10-propargyl-10-dAM on lymphoma growth in vivo.
[0093]Subcutaneous transplantation models in NOD/SCID mice were generated using three established human lymphoma cell line representative of aggressive transformed FL (RL) and de novo extranodal DLBCL (HT; SKI-DLCL-I) histologies. Methods are described in Wang et al., Leukemia and Lymphoma, 2003, Vol. 44 (6), pages 1027-1035 and Rots et al. Leukemia 14:2166-2175 (2000). Six to eight week old non-obese diabetic severe combined immunodeficient (NOD/SCID) mice (Jackson Laboratories, Bar Harbor, Me.) were sub-lethally irradiated with three cGy from a gamma source and inoculated with 10×106 lymphoma cells via a subcutaneous route. When tumor volumes approached 100 mm3, mice were divided into three groups, averaging 3-8 mice per group. Mice were treated with normal saline or the maximum tolerated dose (MTD) of MTX (40 mg/kg) or 10-propargyl-10-dAM (60 mg/kg) via an intraperitoneal route twice weekly for two weeks (four total doses). The MTD of each drug has been previously shown to result in less than 10% weight loss and no toxic deaths in nude mice. (Sirotnak, et al., Cancer Chemother. Pharmacol. 12: 26-30 (1984); Sirotnak et al., Cancer Chemother. Pharmacol. 42: 313-318 (1998). Engraftment rates in this experiment ranged from 80 to 90%. Palpable tumors formed under the skin approximately 7-10 days after inoculation and were readily measurable with calipers. Mice with subcutaneous lymphoma growths survived an average of 40-50 days after inoculation. Treatment results in lymphoma xenografted mice are summarized in Tables 2-4. As shown in Table 2 and 3, in the RL (transformed FL) and SKI-DLCL-I xenografts, 10-propargyl-10-dAM treatment resulted in much greater inhibition of lymphoma growth than MTX. These tumors were only minimally sensitive to MTX treatment with small reductions in growth an no regressions. 10-propargyl-10-dAM treatment, however, decreased tumor volumes by at least 50% from initial volumes and induced tumor regressions in 57% (5 of 9 mice) and 30% (3 of 10 mice) of RL and SKI-DLCL-I, respectively.
TABLE-US-00003 TABLE 2 Treatment of human RL (transformed follicular) non-Hodgkin's lymphoma xenografts. Avg Avg Change in Weight Tumor Tumor Tumor Complete Dose Change Diameter Volume Regression Regression Agent (mg/kg) (%) (mm ± SE) (mm ± SE) (%) (no/total) Control -- +15.9 12.5 ± 1.3 +1228 ± 238 -- 0/7 MTX 40 -14.8 10.9 ± 0.5 +619 ± 108 -- 0/12 PDX 60 -11.1 2.7 ± 1.1 -46 ± 34 56 5/9
TABLE-US-00004 TABLE 3 Treatment of human SKI-DLCL-1 (de novo diffuse large B-cell) non-Hodgkin's lymphoma xenografts Avg Avg Change in Weight Tumor Tumor Tumor Complete Dose Change Diameter Volume Regression Regression Agent (mg/kg) (%) (mm ± SE) (mm ± SE) (%) (no/total) Control -- +4.9 12 ± 0.3 +786 ± 64 -- 0/8 MTX 40 -1.9 9.5 ± 0.4 +299 ± 58 -- 0/10 PDX 60 -1.2 3.5 ± 0.7 -81 ± 16 54 3/10
[0094]As shown in Table 4, even more significant results were achieved using 10-propargyl-10-dAM in the treatment of HT xenografts. Although MTX treatment resulted in modest growth inhibition as compared to controls, there was no tumor regression in these animals. In contrast, 10-propargyl-10-dAM administration resulted in complete tumor regression in 89% (8 of 9) of the mice with an average tumor regression of 99%. At the nadir of tumor regression, HL xenograft mice treated with 10-propargyl-10-dAM had an average tumor diameter of 0.5 mm, as opposed to 11.2 mm for the control and 8.7 mm for MDX-treated mice.
TABLE-US-00005 TABLE 4 Treatment of human HT (diffuse large B-cell) non-Hodgkin's lymphoma xenografts. Avg Avg Change in Weight Tumor Tumor Tumor Complete Dose Change Diameter Volume Regression Regression Agent (mg/kg) (%) (mm ± SE) (mm ± SE) (%) (no/total) Control -- +13.2 11.2 ± 1.3 641 ± 252 -- 0/8 MTX 40 -9.8 8.7 ± 2.0 +300 ± 225 -- 0/7 PDX 60 -8.9 0.5 ± 0.3 -95 ± 0.8 99 8/9
Example 5
[0095]This example describes relative expression of selected genes in B- and T-cell lymphoma cell lines.
[0096]Several proteins are implicated in the metabolism of folic acid and targets for anti-folates in tumor cells. In most tumor cells, the protein encoded by RFC-1 mediates internalization of folate analogs. Once inside the cell, these analogs either bind dihydrofolate reductase (DHFR), thereby depleting intracellular reduced folate pools needed for purine and thymidine biosynthesis, or will be metabolized to a polyglutamate prior to binding to DHFR. Polyglutamylation is catalyzed by FPGS. FPGH mediates cleavage and clearance of these intracellular polyglutamated anti-folates. Using quantitative RT-PCR techniques, expression levels were determined in RL, HT and SKI-DCBCL-I cell lines for RFC-1, FPGS and FPGH using primers and methods as described in Wang et al., Leukemia and Lymphoma, 2003, Vol. 44 (6), pages 1027-1035 and Rots et al. Leukemia 14:2166-2175 (2000). The results of these determinations are summarized in Table 5. As shown, the HT cell line, which was most sensitive to 10-propargyl-10-dAM, also had the greatest levels of RFC-1 expression both on an absolute level and relative to FPGS while the levels of RFC-1 for SKI-DCBCL-I and RL, which had similar sensitivity to 10-propargyl-10-dAM, are similar to one another. Without intending to be bound by a specific mechanism, it is believed that this correlation between RFC-1 expression levels and 10-propargyl-10-dAM sensitivity is a reflection of increased transport of 10-propargyl-10-dAM into tumor cells.
TABLE-US-00006 TABLE 5 Relative levels of RFC-1, FPGS, and FPGH mRNA gene expression in lymphoma cell lines as determined by real-time RT-PCR. Cell RFC-1 (n = 5) FPGS (n = 5) FPGH (n = 5) HT 0.96 ± 0.2 4.92 ± 0.6 1.06 ± 0.2 SKI- 0.30 ± 0.04 6.84 ± 0.6 1.08 ± 0.10 DCBCL-I RL 0.41 ± 0.1 7.28 ± 0.8 0.58 ± 0.08
Example 6
[0097]This example describes relative expression of selected genes in B- and T-cell lymphoma cell lines.
[0098]Using quantitative RT-PCR techniques, expression levels were determined in RL and HT cell lines for DHFR, GARFT, GGH, TS, RFC-1, and FPGS using primers and methods described in Wang et al., Leukemia and Lymphoma, 2003, Vol. 44 (6), pages 1027-1035 and Rots et al. Leukemia 14:2166-2175 (2000). See FIG. 5(a)-(f). Results show that in particular, median expression of RFC-1 and TS trended higher in the T-cell lymphoma cell line than in the B-cell lymphoma cell line. Together with the data showing greater sensitivity of T-cell lymphoma lines to 10-propargyl-10-dAM treatment than B-cell lymphoma lines (including Example 3), these results suggest that T-cell lymphomas' observed greater susceptibility to treatment with 10-propargyl-10-dAM is related to enzymes of the folate pathway and includes differential expression of these enzymes in T-cell lymphomas. Such proteins include DHFR, GARFT, GGH, TS, RFC-1, FPGS, and particularly RFC-1 and TS.
Example 7
[0099]This example describes a comparison of relative gene expression of selected folate pathway genes during treatment of B-cell (RL) and T-cell (HT) lymphoma cells with 10-propargyl-10-dAM and MTX.
[0100]A 10-propargyl-10-dAM preparation prepared in accordance with Example 1 and an MTX preparation were tested for cytotoxicity against a representative B-cell human lymphoma cell lines (RL) and T-cell (HT) human lymphoma cell line. Experiments were performed as described previously. (Sirotnak et al., Cancer Chemother. Pharmacol. 12: 18-25 (1984). In brief, 2.5 to 5×103 cells were plated per well in 96-well flat bottom plates. Drug was added in a 0.9% NaCl solution (pH 7.0) with either 1 nM or 50 nM of 10-propargyl-10-dAM and MTX and gene expression of selected folate pathway enzymes was quantified by RT-PCR as discussed previously at time 0, 1 hour post-treatment, and 12 hours post treatment. Relative enzyme expression levels were determined for reduced folate carrier-1 enzyme (RFC-1), dihydrofolate reductase (DHFR), thymidylate synthase (TS), γ-glutamyl hydrolase (GGH) (also known as folypolyglutamate hydrolase (FPGH)), folylpoly-gamma-glutamate synthetase (FPGS), and glycinamide ribonucleotide formyltransferase (GARFT) was measured at time 0, at 1 hour, and at 12 hours post-treatment. See FIGS. 6(a)-(f). Ct refers to "cycle threshold" and refers to the number of PCR cycles required to generate enough product for the specific antibody to the target. Actin is used as an internal reference. The expression of the selected polypeptide is expressed as relative to actin.
[0101]Results show that RFC-1 expression is several-fold (approximately 7-fold) higher for T-cell (H9) than for B-cell (RL) cell lines and that treatment with low-dose 10-propargyl-10-dAM causes a rapid induction of RFC-1. Results also show that DHFR expression is similar in T-cell (H9) and B-cell (RL) cell lines, and is upregulated, followed by down-regulation, upon treatment of MTX and 10-propargyl-10-dAM. Results show that FPGS expression is similar in T-cell (H9) and B-cell (RL) cell lines, and low dose 10-propargyl-10-dAM produces a rapid two fold induction of FPGS. Results also show that TS expression is seven fold higher in T-cell (H9) than B-cell (RL) cell lines, with upregulation followed by downregulation upon treatment with MTX and 10-propargyl-10-dAM. Further, the results show that GARFT expression is two fold higher in T-cell (H9) than B-cell (RL) and is downregulated upon treatment with 10-propargyl-10-dAM and MTX; and that GGH expression is similar in T-cell (H9) and B-cell (RL) cell lines, with upregulation and downregulation of expression seen upon treatment with 10-propargyl-10-dAM and MTX.
Example 8
[0102]This example describes relative expression of selected genes in B- and T-cell lymphoma cell lines and in a multiple myeloma cell line.
[0103]Using quantitative RT-PCR techniques, expression levels were determined in a number of cell lines for DHFR, GARFT, GGH, TS, RFC-1, and FPGS using primers described in Rots et al. Leukemia 14:2166-2175 (2000), including SKI (B-cell lymphoma line), H9 (T-cell lymphoma line), CCL119 (T-cell lymphoma line), TIB152 (T-cell lymphoma line), RL (B-cell lymphoma line), HPB ALL (T-cell lymphoma line), JJN3 (multiple myeloma cell line), P121chikawa (T-cell lymphoma cell line), Lyl (B-cell lymphoma line), and CUTLL1 (T-cell lymphoma line). See Table 6. Results show that in particular, median expression of RFC-1 and TS in particular trended higher in the multiple myeloma cell line JJN3. Together with the data showing greater sensitivity of T-cell lymphoma lines to 10-propargyl-10-dAM treatment than B-cell lymphoma lines (including Example 3) and higher RFC-1 expression, these results show that multiple myeloma may have greater susceptibility to treatment with 10-propargyl-10-dAM, and that biomarkers for such sensitivity may include DHFR, GARFT, GGH, TS, RFC-1, AND FPGS, and RFC-1 and TS.
TABLE-US-00007 TABLE 6 PDX Cell Line Untreated RT-PCR A. Allos Sample COM- Actin Number MENTS RGI Accession Ct DHFR Ct DeltaCt A SKI AGR-06- 28.07 21.71 22.84 5.23 0002291 B H9 AGR-06- 30.48 8.72 23.94 6.54 0002292 C CCL119 AGR-06- 29.69 9.33 23.24 6.45 0002293 D TIB 152 AGR-06- 30.33 16.06 24.66 5.66 0002294 E RL AGR-06- 27.29 12.04 21.22 6.08 0002295 F HPB ALL AGR-06- 29.71 17.64 24.18 5.53 0002296 G JJN3 AGR-06- 30.60 32.52 25.96 4.64 0002297 H P12lchikawa AGR-06- 30.07 16.39 24.44 5.63 0002298 I Ly1 AGR-06- 31.77 10.30 25.47 6.30 0002299 J CUTLL1 AGR-06- 32.67 10.19 26.35 6.32 0002300 B. Allos Sample COM- Actin Number MENTS RGI Accession Ct FPGS Ct DeltaCt A SKI AGR-06-0002291 29.83 0.44 22.84 6.99 B H9 AGR-06-0002292 31.24 0.35 23.94 7.31 C CCL119 AGR-06-0002293 30.97 0.26 23.24 7.72 D TIB 152 AGR-06-0002294 31.30 0.55 24.66 6.64 E RL AGR-06-0002295 29.16 0.22 21.22 7.95 F HPB ALL AGR-06-0002296 31.34 0.39 24.18 7.16 G JJN3 AGR-06-0002297 33.09 0.40 25.96 7.13 H P12lchikawa AGR-06-0002298 31.89 0.32 24.44 7.45 I Ly1 AGR-06-0002299 33.57 0.20 25.47 8.10 J CUTLL1 AGR-06-0002300 33.56 0.37 26.35 7.21 C. Allos Sample RGI Act in Number COMMENTS Accession Ct GARFT Ct DeltaCt A SKI AGR-06- 29.26 1.55 22.84 6.42 0002291 B H9 AGR-06- 29.97 2.02 23.94 6.04 0002292 C CCL119 AGR-06- 30.18 1.09 23.24 6.93 0002293 D TIB 152 AGR-06- 30.83 1.84 24.66 6.17 0002294 E RL AGR-06- 27.92 1.27 21.22 6.70 0002295 F HPB ALL AGR-06- 30.72 1.43 24.18 6.54 0002296 G JJN3 AGR-06- 31.46 2.93 25.96 5.50 0002297 H P12lchikawa AGR-06- 31.14 1.28 24.44 6.70 0002298 I Ly1 AGR-06- 31.83 1.62 25.47 6.36 0002299 J CUTLL1 AGR-06- 33.73 0.79 26.35 7.38 0002300 D. Allos Sample COM- Actin Number MENTS RGI Accession Ct GGH Ct DeltaCt A SKI AGR-06-0002291 30.27 0.82 22.69 7.58 B H9 AGR-06-0002292 31.81 0.61 23.80 8.01 C CCL119 AGR-06-0002293 31.02 0.66 23.14 7.89 D TIB 152 AGR-06-0002294 32.15 0.81 24.54 7.61 E RL AGR-06-0002295 29.21 0.61 21.20 8.00 F HPB ALL AGR-06-0002296 31.86 0.73 24.11 7.75 G JJN3 AGR-06-0002297 28.68 23.16 25.92 2.76 H P12lchikawa AGR-06-0002298 32.79 0.48 24.45 8.34 I Ly1 AGR-06-0002299 33.39 0.61 25.39 8.00 J CUTLL1 AGR-06-0002300 34.15 0.67 26.28 7.87 E. Allos Sample COM- Actin Number MENTS RGI Accession Ct RFC1 Ct DeltaCt A SKI AGR-06-0002291 30.75 1.91 22.69 8.07 B H9 AGR-06-0002292 30.15 6.25 23.80 6.35 C CCL119 AGR-06-0002293 30.49 3.12 23.14 7.36 D TIB 152 AGR-06-0002294 31.45 4.27 24.54 6.90 E RL AGR-06-0002295 29.90 1.24 21.20 8.69 F HPB ALL AGR-06-0002296 31.19 3.79 24.11 7.08 G JJN3 AGR-06-0002297 31.77 8.82 25.92 5.86 H P12lchikawa AGR-06-0002298 31.26 4.56 24.45 6.81 I Ly1 AGR-06-0002299 32.25 4.40 25.39 6.86 J CUTLL1 AGR-06-0002300 32.70 6.01 26.28 6.41 F. Allos Sample COM- Actin Number MENTS RGI Accession Ct TS Ct DeltaCt A SKI AGR-06-0002291 28.48 5.26 22.69 5.80 B H9 AGR-06-0002292 28.09 14.94 23.80 4.29 C CCL119 AGR-06-0002293 27.23 17.13 23.14 4.10 D TIB 152 AGR-06-0002294 27.56 36.18 24.54 3.02 E RL AGR-06-0002295 27.49 3.75 21.20 6.29 F HPB ALL AGR-06-0002296 27.64 25.37 24.11 3.53 G JJN3 AGR-06-0002297 28.29 56.71 25.92 2.37 H P12lchikawa AGR-06-0002298 27.95 25.97 24.45 3.50 I Ly1 AGR-06-0002299 30.59 7.97 25.39 5.20 J CUTLL1 AGR-06-0002300 29.70 27.50 26.28 3.41
[0104]The foregoing discussion of the invention has been presented for purposes of illustration and description. The foregoing is not intended to limit the invention to the form or forms disclosed herein. Although the description of the invention has included description of one or more embodiments and certain variations and modifications, other variations and modifications are within the scope of the invention, e.g., as may be within the skill and knowledge of those in the art, after understanding the present disclosure. It is intended to obtain rights which include alternative embodiments to the extent permitted, including alternate, interchangeable and/or equivalent structures, functions, ranges or steps to those claimed, whether or not such alternate, interchangeable and/or equivalent structures, functions, ranges or steps are disclosed herein, and without intending to publicly dedicate any patentable subject matter.
Sequence CWU
1
1812839DNAHomo sapiens 1gtagtcccgg agtccgcgtg cgcggggccg ggtccgggag
ccccagggca gccgccccgc 60cgagtcgcag gcacagcgtc accttcgtcc cctccggagc
tgcacgtggc ctgagcagga 120tggtgccctc cagcccagcg gtggagaagc aggtgcccgt
ggaacctggg cctgaccccg 180agctccggtc ctggcggcac ctcgtgtgct acctttgctt
ctacggcttc atggcgcaga 240tacggccagg ggagagcttc atcaccccct acctcctggg
gcccgacaag aacttcacgc 300gggagcaggt cacgaacgag atcacgccgg tgctgtcgta
ctcctacctg gccgtgctgg 360tgcccgtgtt cctgctcacc gactacctgc gctacacgcc
ggtgctgctg ctgcaggggc 420tcagcttcgt gtcggtgtgg ctgctgctgc tgctgggcca
ctcggtggcg cacatgcagc 480tcatggagct cttctacagc gtcaccatgg ccgcgcgcat
cgcctattcc tcctacatct 540tctctctcgt gcggcccgcg cgctaccagc gtgtggccgg
ctactcgcgc gctgcggtgc 600tgctgggcgt gttcaccagc tccgtgctgg gccagctgct
ggtcactgtg ggccgagtct 660ccttctccac gctcaactac atctcgctgg ccttcctcac
cttcagcgtg gtcctcgccc 720tcttcctgaa gcgccccaag cgcagcctct tcttcaaccg
cgacgaccgg gggcggtgcg 780aaacctcggc ttcggagctg gagcgcatga atcctggccc
aggcgggaag ctgggacacg 840ccctgcgggt ggcctgtggg gactcagtgc tggcgcggat
gctgcgggag ctgggggaca 900gcctgcggcg gccgcagctg cgcctgtggt ccctctggtg
ggtcttcaac tcggccggct 960actacctggt ggtctactac gtgcacatcc tgtggaacga
ggtggacccc accaccaaca 1020gtgcgcgggt ctacaacggc gcggcagatg ctgcctccac
gctgctgggc gccatcacgt 1080ccttcgccgc gggcttcgtg aagatccgct gggcgcgctg
gtccaagctg ctcatcgcgg 1140gcgtcacggc cacgcaggcg gggctggtct tccttctggc
gcacacgcgc cacccgagca 1200gcatctggct gtgctatgcg gccttcgtgc tgttccgcgg
ctcctaccag ttcctcgtgc 1260ccatcgccac ctttcagatt gcatcttctc tgtctaaaga
gctctgtgcc ctggtcttcg 1320gggtcaacac gttctttgcc accatcgtca agaccatcat
cactttcatt gtctcggacg 1380tgcggggcct gggcctcccg gtccgcaagc agttccagtt
atactccgtg tacttcctga 1440tcctgtccat catctacttc ttgggggcca tgctggatgg
cctgcggcac tgccagcggg 1500gccaccaccc gcggcagccc ccggcccagg gcctgaggag
tgccgcggag gagaaggcag 1560cacaggcact gagcgtgcag gacaagggcc tcggaggcct
gcagccagcc cagagcccgc 1620cgctttcccc agaagacagc ctgggggctg tggggccagc
ctccctggag cagagacaga 1680gcgacccata cctggcccag gccccggccc cgcaggcagc
tgaattcctg agcccagtga 1740caaccccttc cccctgcact ctgtgctccg cccaagcctc
aggccctgag gctgcagatg 1800agacttgtcc ccagctggct gtccatcctc ctggtgtcag
caagctgggt ttgcagtgtc 1860ttccaagcga cggtgttcag aatgtgaacc agtgactctc
gggcgcccct gtggtaactt 1920tgcaggcggc cctcagtgca tccccacgac ccctgcctcg
agggccgcct gccttagcaa 1980tgggggcctc cgcttatcct gctagcaggc cccctaggat
tccccctgcc ctgtgccgca 2040ctctggcggt ggccacagcg tgctggcgac actcagggca
gctgcctggc catgctgtcc 2100ctgcactgtg ccccgcgggc tttgttgctg gaagaggtgg
gtggtgggct tctgcgtcca 2160ccaggcctca ctggctcatg ccccttgggg ggcttgagac
aaatcctttc tgccccccag 2220ggctagtgaa gtggcctctt ggataccagc tcaggggaca
ctggccccac aggagttgtg 2280agccctctag ggcagggtgg gagccgggac cctcaggtgt
agctgagctg tgacattgct 2340ggtcatcctt ggtgctcttg cttttttgaa agatgctttt
ttttttttta actgacgtag 2400aatgaagaac tgcatgtggc ttctctgtct ctgtggaaaa
gccatctcag gttggcggca 2460gacacattgt catcagaggg gagcagcggc tctggtcctc
ggagctggtt cctctctccc 2520accctaaggg cagccctcca tggtcctgtc tgtccttctg
aagtgtgtcc atcctgacct 2580gcgggtcctc agctgctccc acacttgtgc cagcccggag
gggactggtc ccggtcaccg 2640cggacgtgct ggccttggta tgtgccaggc ttgcctgggc
tgggcagcct tgggggggct 2700gcctttgtgg tgggcgctgg ggaagtacgt cccagcggcc
tcagggtcta aggagcgcta 2760gtgccttgcc cacaggtgcg ggaccatctg atgtgatgtg
aatactcttc ccacatacat 2820taaacacact taagtgaga
283922839DNAHomo sapiensCDS(120)..(1895)
2gtagtcccgg agtccgcgtg cgcggggccg ggtccgggag ccccagggca gccgccccgc
60cgagtcgcag gcacagcgtc accttcgtcc cctccggagc tgcacgtggc ctgagcagg
119atg gtg ccc tcc agc cca gcg gtg gag aag cag gtg ccc gtg gaa cct
167Met Val Pro Ser Ser Pro Ala Val Glu Lys Gln Val Pro Val Glu Pro1
5 10 15ggg cct gac ccc gag ctc
cgg tcc tgg cgg cac ctc gtg tgc tac ctt 215Gly Pro Asp Pro Glu Leu
Arg Ser Trp Arg His Leu Val Cys Tyr Leu 20 25
30tgc ttc tac ggc ttc atg gcg cag ata cgg cca ggg gag
agc ttc atc 263Cys Phe Tyr Gly Phe Met Ala Gln Ile Arg Pro Gly Glu
Ser Phe Ile 35 40 45acc ccc tac
ctc ctg ggg ccc gac aag aac ttc acg cgg gag cag gtc 311Thr Pro Tyr
Leu Leu Gly Pro Asp Lys Asn Phe Thr Arg Glu Gln Val 50
55 60acg aac gag atc acg ccg gtg ctg tcg tac tcc tac
ctg gcc gtg ctg 359Thr Asn Glu Ile Thr Pro Val Leu Ser Tyr Ser Tyr
Leu Ala Val Leu65 70 75
80gtg ccc gtg ttc ctg ctc acc gac tac ctg cgc tac acg ccg gtg ctg
407Val Pro Val Phe Leu Leu Thr Asp Tyr Leu Arg Tyr Thr Pro Val Leu
85 90 95ctg ctg cag ggg ctc agc
ttc gtg tcg gtg tgg ctg ctg ctg ctg ctg 455Leu Leu Gln Gly Leu Ser
Phe Val Ser Val Trp Leu Leu Leu Leu Leu 100
105 110ggc cac tcg gtg gcg cac atg cag ctc atg gag ctc
ttc tac agc gtc 503Gly His Ser Val Ala His Met Gln Leu Met Glu Leu
Phe Tyr Ser Val 115 120 125acc atg
gcc gcg cgc atc gcc tat tcc tcc tac atc ttc tct ctc gtg 551Thr Met
Ala Ala Arg Ile Ala Tyr Ser Ser Tyr Ile Phe Ser Leu Val 130
135 140cgg ccc gcg cgc tac cag cgt gtg gcc ggc tac
tcg cgc gct gcg gtg 599Arg Pro Ala Arg Tyr Gln Arg Val Ala Gly Tyr
Ser Arg Ala Ala Val145 150 155
160ctg ctg ggc gtg ttc acc agc tcc gtg ctg ggc cag ctg ctg gtc act
647Leu Leu Gly Val Phe Thr Ser Ser Val Leu Gly Gln Leu Leu Val Thr
165 170 175gtg ggc cga gtc tcc
ttc tcc acg ctc aac tac atc tcg ctg gcc ttc 695Val Gly Arg Val Ser
Phe Ser Thr Leu Asn Tyr Ile Ser Leu Ala Phe 180
185 190ctc acc ttc agc gtg gtc ctc gcc ctc ttc ctg aag
cgc ccc aag cgc 743Leu Thr Phe Ser Val Val Leu Ala Leu Phe Leu Lys
Arg Pro Lys Arg 195 200 205agc ctc
ttc ttc aac cgc gac gac cgg ggg cgg tgc gaa acc tcg gct 791Ser Leu
Phe Phe Asn Arg Asp Asp Arg Gly Arg Cys Glu Thr Ser Ala 210
215 220tcg gag ctg gag cgc atg aat cct ggc cca ggc
ggg aag ctg gga cac 839Ser Glu Leu Glu Arg Met Asn Pro Gly Pro Gly
Gly Lys Leu Gly His225 230 235
240gcc ctg cgg gtg gcc tgt ggg gac tca gtg ctg gcg cgg atg ctg cgg
887Ala Leu Arg Val Ala Cys Gly Asp Ser Val Leu Ala Arg Met Leu Arg
245 250 255gag ctg ggg gac agc
ctg cgg cgg ccg cag ctg cgc ctg tgg tcc ctc 935Glu Leu Gly Asp Ser
Leu Arg Arg Pro Gln Leu Arg Leu Trp Ser Leu 260
265 270tgg tgg gtc ttc aac tcg gcc ggc tac tac ctg gtg
gtc tac tac gtg 983Trp Trp Val Phe Asn Ser Ala Gly Tyr Tyr Leu Val
Val Tyr Tyr Val 275 280 285cac atc
ctg tgg aac gag gtg gac ccc acc acc aac agt gcg cgg gtc 1031His Ile
Leu Trp Asn Glu Val Asp Pro Thr Thr Asn Ser Ala Arg Val 290
295 300tac aac ggc gcg gca gat gct gcc tcc acg ctg
ctg ggc gcc atc acg 1079Tyr Asn Gly Ala Ala Asp Ala Ala Ser Thr Leu
Leu Gly Ala Ile Thr305 310 315
320tcc ttc gcc gcg ggc ttc gtg aag atc cgc tgg gcg cgc tgg tcc aag
1127Ser Phe Ala Ala Gly Phe Val Lys Ile Arg Trp Ala Arg Trp Ser Lys
325 330 335ctg ctc atc gcg ggc
gtc acg gcc acg cag gcg ggg ctg gtc ttc ctt 1175Leu Leu Ile Ala Gly
Val Thr Ala Thr Gln Ala Gly Leu Val Phe Leu 340
345 350ctg gcg cac acg cgc cac ccg agc agc atc tgg ctg
tgc tat gcg gcc 1223Leu Ala His Thr Arg His Pro Ser Ser Ile Trp Leu
Cys Tyr Ala Ala 355 360 365ttc gtg
ctg ttc cgc ggc tcc tac cag ttc ctc gtg ccc atc gcc acc 1271Phe Val
Leu Phe Arg Gly Ser Tyr Gln Phe Leu Val Pro Ile Ala Thr 370
375 380ttt cag att gca tct tct ctg tct aaa gag ctc
tgt gcc ctg gtc ttc 1319Phe Gln Ile Ala Ser Ser Leu Ser Lys Glu Leu
Cys Ala Leu Val Phe385 390 395
400ggg gtc aac acg ttc ttt gcc acc atc gtc aag acc atc atc act ttc
1367Gly Val Asn Thr Phe Phe Ala Thr Ile Val Lys Thr Ile Ile Thr Phe
405 410 415att gtc tcg gac gtg
cgg ggc ctg ggc ctc ccg gtc cgc aag cag ttc 1415Ile Val Ser Asp Val
Arg Gly Leu Gly Leu Pro Val Arg Lys Gln Phe 420
425 430cag tta tac tcc gtg tac ttc ctg atc ctg tcc atc
atc tac ttc ttg 1463Gln Leu Tyr Ser Val Tyr Phe Leu Ile Leu Ser Ile
Ile Tyr Phe Leu 435 440 445ggg gcc
atg ctg gat ggc ctg cgg cac tgc cag cgg ggc cac cac ccg 1511Gly Ala
Met Leu Asp Gly Leu Arg His Cys Gln Arg Gly His His Pro 450
455 460cgg cag ccc ccg gcc cag ggc ctg agg agt gcc
gcg gag gag aag gca 1559Arg Gln Pro Pro Ala Gln Gly Leu Arg Ser Ala
Ala Glu Glu Lys Ala465 470 475
480gca cag gca ctg agc gtg cag gac aag ggc ctc gga ggc ctg cag cca
1607Ala Gln Ala Leu Ser Val Gln Asp Lys Gly Leu Gly Gly Leu Gln Pro
485 490 495gcc cag agc ccg ccg
ctt tcc cca gaa gac agc ctg ggg gct gtg ggg 1655Ala Gln Ser Pro Pro
Leu Ser Pro Glu Asp Ser Leu Gly Ala Val Gly 500
505 510cca gcc tcc ctg gag cag aga cag agc gac cca tac
ctg gcc cag gcc 1703Pro Ala Ser Leu Glu Gln Arg Gln Ser Asp Pro Tyr
Leu Ala Gln Ala 515 520 525ccg gcc
ccg cag gca gct gaa ttc ctg agc cca gtg aca acc cct tcc 1751Pro Ala
Pro Gln Ala Ala Glu Phe Leu Ser Pro Val Thr Thr Pro Ser 530
535 540ccc tgc act ctg tgc tcc gcc caa gcc tca ggc
cct gag gct gca gat 1799Pro Cys Thr Leu Cys Ser Ala Gln Ala Ser Gly
Pro Glu Ala Ala Asp545 550 555
560gag act tgt ccc cag ctg gct gtc cat cct cct ggt gtc agc aag ctg
1847Glu Thr Cys Pro Gln Leu Ala Val His Pro Pro Gly Val Ser Lys Leu
565 570 575ggt ttg cag tgt ctt
cca agc gac ggt gtt cag aat gtg aac cag tga 1895Gly Leu Gln Cys Leu
Pro Ser Asp Gly Val Gln Asn Val Asn Gln 580
585 590ctctcgggcg cccctgtggt aactttgcag gcggccctca
gtgcatcccc acgacccctg 1955cctcgagggc cgcctgcctt agcaatgggg gcctccgctt
atcctgctag caggccccct 2015aggattcccc ctgccctgtg ccgcactctg gcggtggcca
cagcgtgctg gcgacactca 2075gggcagctgc ctggccatgc tgtccctgca ctgtgccccg
cgggctttgt tgctggaaga 2135ggtgggtggt gggcttctgc gtccaccagg cctcactggc
tcatgcccct tggggggctt 2195gagacaaatc ctttctgccc cccagggcta gtgaagtggc
ctcttggata ccagctcagg 2255ggacactggc cccacaggag ttgtgagccc tctagggcag
ggtgggagcc gggaccctca 2315ggtgtagctg agctgtgaca ttgctggtca tccttggtgc
tcttgctttt ttgaaagatg 2375cttttttttt ttttaactga cgtagaatga agaactgcat
gtggcttctc tgtctctgtg 2435gaaaagccat ctcaggttgg cggcagacac attgtcatca
gaggggagca gcggctctgg 2495tcctcggagc tggttcctct ctcccaccct aagggcagcc
ctccatggtc ctgtctgtcc 2555ttctgaagtg tgtccatcct gacctgcggg tcctcagctg
ctcccacact tgtgccagcc 2615cggaggggac tggtcccggt caccgcggac gtgctggcct
tggtatgtgc caggcttgcc 2675tgggctgggc agccttgggg gggctgcctt tgtggtgggc
gctggggaag tacgtcccag 2735cggcctcagg gtctaaggag cgctagtgcc ttgcccacag
gtgcgggacc atctgatgtg 2795atgtgaatac tcttcccaca tacattaaac acacttaagt
gaga 28393591PRTHomo sapiens 3Met Val Pro Ser Ser Pro
Ala Val Glu Lys Gln Val Pro Val Glu Pro1 5
10 15Gly Pro Asp Pro Glu Leu Arg Ser Trp Arg His Leu
Val Cys Tyr Leu 20 25 30Cys
Phe Tyr Gly Phe Met Ala Gln Ile Arg Pro Gly Glu Ser Phe Ile 35
40 45Thr Pro Tyr Leu Leu Gly Pro Asp Lys
Asn Phe Thr Arg Glu Gln Val 50 55
60Thr Asn Glu Ile Thr Pro Val Leu Ser Tyr Ser Tyr Leu Ala Val Leu65
70 75 80Val Pro Val Phe Leu
Leu Thr Asp Tyr Leu Arg Tyr Thr Pro Val Leu 85
90 95Leu Leu Gln Gly Leu Ser Phe Val Ser Val Trp
Leu Leu Leu Leu Leu 100 105
110Gly His Ser Val Ala His Met Gln Leu Met Glu Leu Phe Tyr Ser Val
115 120 125Thr Met Ala Ala Arg Ile Ala
Tyr Ser Ser Tyr Ile Phe Ser Leu Val 130 135
140Arg Pro Ala Arg Tyr Gln Arg Val Ala Gly Tyr Ser Arg Ala Ala
Val145 150 155 160Leu Leu
Gly Val Phe Thr Ser Ser Val Leu Gly Gln Leu Leu Val Thr
165 170 175Val Gly Arg Val Ser Phe Ser
Thr Leu Asn Tyr Ile Ser Leu Ala Phe 180 185
190Leu Thr Phe Ser Val Val Leu Ala Leu Phe Leu Lys Arg Pro
Lys Arg 195 200 205Ser Leu Phe Phe
Asn Arg Asp Asp Arg Gly Arg Cys Glu Thr Ser Ala 210
215 220Ser Glu Leu Glu Arg Met Asn Pro Gly Pro Gly Gly
Lys Leu Gly His225 230 235
240Ala Leu Arg Val Ala Cys Gly Asp Ser Val Leu Ala Arg Met Leu Arg
245 250 255Glu Leu Gly Asp Ser
Leu Arg Arg Pro Gln Leu Arg Leu Trp Ser Leu 260
265 270Trp Trp Val Phe Asn Ser Ala Gly Tyr Tyr Leu Val
Val Tyr Tyr Val 275 280 285His Ile
Leu Trp Asn Glu Val Asp Pro Thr Thr Asn Ser Ala Arg Val 290
295 300Tyr Asn Gly Ala Ala Asp Ala Ala Ser Thr Leu
Leu Gly Ala Ile Thr305 310 315
320Ser Phe Ala Ala Gly Phe Val Lys Ile Arg Trp Ala Arg Trp Ser Lys
325 330 335Leu Leu Ile Ala
Gly Val Thr Ala Thr Gln Ala Gly Leu Val Phe Leu 340
345 350Leu Ala His Thr Arg His Pro Ser Ser Ile Trp
Leu Cys Tyr Ala Ala 355 360 365Phe
Val Leu Phe Arg Gly Ser Tyr Gln Phe Leu Val Pro Ile Ala Thr 370
375 380Phe Gln Ile Ala Ser Ser Leu Ser Lys Glu
Leu Cys Ala Leu Val Phe385 390 395
400Gly Val Asn Thr Phe Phe Ala Thr Ile Val Lys Thr Ile Ile Thr
Phe 405 410 415Ile Val Ser
Asp Val Arg Gly Leu Gly Leu Pro Val Arg Lys Gln Phe 420
425 430Gln Leu Tyr Ser Val Tyr Phe Leu Ile Leu
Ser Ile Ile Tyr Phe Leu 435 440
445Gly Ala Met Leu Asp Gly Leu Arg His Cys Gln Arg Gly His His Pro 450
455 460Arg Gln Pro Pro Ala Gln Gly Leu
Arg Ser Ala Ala Glu Glu Lys Ala465 470
475 480Ala Gln Ala Leu Ser Val Gln Asp Lys Gly Leu Gly
Gly Leu Gln Pro 485 490
495Ala Gln Ser Pro Pro Leu Ser Pro Glu Asp Ser Leu Gly Ala Val Gly
500 505 510Pro Ala Ser Leu Glu Gln
Arg Gln Ser Asp Pro Tyr Leu Ala Gln Ala 515 520
525Pro Ala Pro Gln Ala Ala Glu Phe Leu Ser Pro Val Thr Thr
Pro Ser 530 535 540Pro Cys Thr Leu Cys
Ser Ala Gln Ala Ser Gly Pro Glu Ala Ala Asp545 550
555 560Glu Thr Cys Pro Gln Leu Ala Val His Pro
Pro Gly Val Ser Lys Leu 565 570
575Gly Leu Gln Cys Leu Pro Ser Asp Gly Val Gln Asn Val Asn Gln
580 585 59043932DNAHomo sapiens
4tcccagacag aacctactat gtgcggcggc agctggggcg ggaaggcggg agctgggggc
60gctgggggcg ctgcggccgc tgcggccgct gcagccgctg cagcgccagg gtccacctgg
120tcggctgcac ctgtggagga ggaggtggat ttcaggcttc ccgtagactg gaagaatcgg
180ctcaaaaccg cttgcctcgc aggggctgag ctggaggcag cgaggccgcc cgacgcaggc
240ttccggcgag acatggcagg gcaaggatgg cagcccggcg gcagggcctg gcgaggagcg
300cgagcccgcg gccgcagttc ccaggcgtct gcgggcgcga gcacgccgcg accctgcgtg
360cgccggggcg ggggggcggg gcctcgcctg cacaaatggg gacgaggggg gcggggcggc
420cacaatttcg cgccaaactt gaccgcgcgt tctgctgtaa cgagcgggct cggaggtcct
480cccgctgctg tcatggttgg ttcgctaaac tgcatcgtcg ctgtgtccca gaacatgggc
540atcggcaaga acggggacct gccctggcca ccgctcagga atgaattcag atatttccag
600agaatgacca caacctcttc agtagaaggt aaacagaatc tggtgattat gggtaagaag
660acctggttct ccattcctga gaagaatcga cctttaaagg gtagaattaa tttagttctc
720agcagagaac tcaaggaacc tccacaagga gctcattttc tttccagaag tctagatgat
780gccttaaaac ttactgaaca accagaatta gcaaataaag tagacatggt ctggatagtt
840ggtggcagtt ctgtttataa ggaagccatg aatcacccag gccatcttaa actatttgtg
900acaaggatca tgcaagactt tgaaagtgac acgttttttc cagaaattga tttggagaaa
960tataaacttc tgccagaata cccaggtgtt ctctctgatg tccaggagga gaaaggcatt
1020aagtacaaat ttgaagtata tgagaagaat gattaatatg aaggtgtttt ctagtttaag
1080ttgttccccc tccctctgaa aaaagtatgt atttttacat tagaaaaggt tttttgttga
1140ctttagatct ataattattt ctaagcaact agtttttatt ccccactact cttgtctcta
1200tcagatacca tttatgagac attcttgcta taactaagtg cttctccaag accccaactg
1260agtccccagc acctgctaca gtgagctgcc attccacacc catcacatgt ggcactcttg
1320ccagtccttg acattgtcgg gcttttcaca tgttggtaat atttattaaa gatgaagatc
1380cacataccct tcaactgagc agtttcacta gtggaaatac caaaagcttc ctacgtgtat
1440atccagaggt ttgtagataa atgttgccac cttgtttgta acagtgaaaa attgaaaaca
1500acctggaagt ccagtgatgg gaaaatgagt atgtttctgt cttagattgg ggaacccaaa
1560gcagattgca agactgaaat ttcagtgaaa gcagtgtatt tgctaggtca taccagaaat
1620catcaattga ggtacggaga aactgaactg agaaggtaag aaaagcaatt taaagtcagc
1680gagcaggttc tcattgataa caagctccat actgctgaga tacagggaaa tggagggggg
1740aaagctggag tattgatccc gcccccctcc ttggttgtca gctccctgtc ctgtgtgtgg
1800gcggaacata gtccagctgc tctatagcaa gtctcaggtg tttgcagtaa gaagctgctg
1860gcatgcacgg gaacagtgaa tgccaaacac ttaaagcaat tcgatgttta agtatgtaag
1920ttcttttttt tttagacagc gtttcgctct tgttgcccag gctagcatgc aatggtgtga
1980cctcggctta ctgcaacctc cgccttccca gattcaagcg attctcctgc ctcaggctcc
2040caagtagcta ggaccaggtg cgcgccacca cgcccggcta atttttgtat tttgtatttt
2100tagtagagat ggggtttcac catgttggtc aggctagtct cgaactcgtg accgcaagcg
2160attcacccac ctcagcctcc caaagtgctg ggattaccgg cttgagccac cacacccggc
2220acatcttcat tctttttatg tagtaaaaag tataaggcca cacatggttt atttgaagta
2280ttttataatt taaaaaaata cagaagcagg aaaaccaatt ataagttcaa gtgagggatg
2340atggttgctt gaaccaaagg gttgcatgta gtaagaaatt gtgatttaag atatatttta
2400aagttataag tagcaggata ttctgatgga gtttgacttt ggttttgggc ccagggagtt
2460tcagatgcct ttgagaaatg aatgaagtag agagaaaata aaagaaaaac cagccaggca
2520cagtggctca cacctgtaat cccagcgctt tgggaggcta aggcaggcag atcacttgag
2580accagcttgg gcaacatggc aaagccccat ctctacaaaa aacacaaaaa ttagctgggc
2640attgtggcgc acacctgtat tcccatctag tcaggaagct gagatggaag aattaattga
2700gcccacgagt tcaaggctgc agtgagtcgt gattgtgcca ctgcactcca gccggggtga
2760cagaagagac cttgtctcga aaaggaatct gaaaacaatg gaaccatgcc ttcataattc
2820tagaaagtta ttttcaactg ataaatctat attcacccaa ataatcaagg gtgaaggtaa
2880aataatacat ttttagacaa gcaaagactc aggggttacc tccatgtgcc ctttttaggg
2940aagctgttgg agaaaatact ccagcaaaat gaaggagtac acaaaccaga gaatgacatg
3000aatccagcaa ataggatcca acacaggcaa tattccagct atggagctag ctttaaaaag
3060gaacagtaaa aatattaatc ggttagctgg gtggaatggc ccatgcctgt agtcccagct
3120actcaggagg ctcagcagca ggacgacttg agcccaagag ttccagacca gcctggccac
3180cttagtgaga tcccttctct taaaaataat aacttattgc cagatttggg gcatttggaa
3240agaagttcat tgaagataaa gcaaaagtaa aaaaaaaaaa aaaaaaaaca aggggaaagg
3300gttggttagg caatcattct agggcagaaa gaagtacagg ataggaagag cataatacac
3360tgtttttctc aacaaggagc agtatgtaca cagtcataat gatgtgactg cttagcccct
3420aaatatggta actactctgg gacaatatgg gaggaaaagt gaagattgtg atggtgtaag
3480agctaaatcc tcatctgtca tatccagaaa tcactatata atatataata atgaaatgac
3540taagttatgt gaggaaaaaa acagaagaca ttgctaaaag agttaaaagt cattgctctg
3600gagaattagg agggatgggg caggggactg ttaggatgca ttataaactg aaaagccttt
3660ttaaaatttt atgtattaat atatgcattc acttgaaaaa ctaaaaaaaa acaataattt
3720ggaaaaaccc atgaaggtaa ctaacggaag gaaaaactaa gagaatgaaa agtatttgcc
3780tctggaaaga acaactggca ggactgttgt tttcattgta agacttttgg agccatttaa
3840ttgtacttaa ccattttcat ctatttcttt aataagaaca attccatctt aataaagagt
3900tacacttgtt aataagtaaa aaaaaaaaaa aa
393253932DNAHomo sapiensCDS(493)..(1056) 5tcccagacag aacctactat
gtgcggcggc agctggggcg ggaaggcggg agctgggggc 60gctgggggcg ctgcggccgc
tgcggccgct gcagccgctg cagcgccagg gtccacctgg 120tcggctgcac ctgtggagga
ggaggtggat ttcaggcttc ccgtagactg gaagaatcgg 180ctcaaaaccg cttgcctcgc
aggggctgag ctggaggcag cgaggccgcc cgacgcaggc 240ttccggcgag acatggcagg
gcaaggatgg cagcccggcg gcagggcctg gcgaggagcg 300cgagcccgcg gccgcagttc
ccaggcgtct gcgggcgcga gcacgccgcg accctgcgtg 360cgccggggcg ggggggcggg
gcctcgcctg cacaaatggg gacgaggggg gcggggcggc 420cacaatttcg cgccaaactt
gaccgcgcgt tctgctgtaa cgagcgggct cggaggtcct 480cccgctgctg tc atg gtt
ggt tcg cta aac tgc atc gtc gct gtg tcc cag 531 Met Val
Gly Ser Leu Asn Cys Ile Val Ala Val Ser Gln 1
5 10aac atg ggc atc ggc aag aac ggg gac ctg ccc tgg cca
ccg ctc agg 579Asn Met Gly Ile Gly Lys Asn Gly Asp Leu Pro Trp Pro
Pro Leu Arg 15 20 25aat gaa ttc aga
tat ttc cag aga atg acc aca acc tct tca gta gaa 627Asn Glu Phe Arg
Tyr Phe Gln Arg Met Thr Thr Thr Ser Ser Val Glu30 35
40 45ggt aaa cag aat ctg gtg att atg ggt
aag aag acc tgg ttc tcc att 675Gly Lys Gln Asn Leu Val Ile Met Gly
Lys Lys Thr Trp Phe Ser Ile 50 55
60cct gag aag aat cga cct tta aag ggt aga att aat tta gtt ctc
agc 723Pro Glu Lys Asn Arg Pro Leu Lys Gly Arg Ile Asn Leu Val Leu
Ser 65 70 75aga gaa ctc aag
gaa cct cca caa gga gct cat ttt ctt tcc aga agt 771Arg Glu Leu Lys
Glu Pro Pro Gln Gly Ala His Phe Leu Ser Arg Ser 80
85 90cta gat gat gcc tta aaa ctt act gaa caa cca gaa
tta gca aat aaa 819Leu Asp Asp Ala Leu Lys Leu Thr Glu Gln Pro Glu
Leu Ala Asn Lys 95 100 105gta gac atg
gtc tgg ata gtt ggt ggc agt tct gtt tat aag gaa gcc 867Val Asp Met
Val Trp Ile Val Gly Gly Ser Ser Val Tyr Lys Glu Ala110
115 120 125atg aat cac cca ggc cat ctt
aaa cta ttt gtg aca agg atc atg caa 915Met Asn His Pro Gly His Leu
Lys Leu Phe Val Thr Arg Ile Met Gln 130
135 140gac ttt gaa agt gac acg ttt ttt cca gaa att gat
ttg gag aaa tat 963Asp Phe Glu Ser Asp Thr Phe Phe Pro Glu Ile Asp
Leu Glu Lys Tyr 145 150 155aaa
ctt ctg cca gaa tac cca ggt gtt ctc tct gat gtc cag gag gag 1011Lys
Leu Leu Pro Glu Tyr Pro Gly Val Leu Ser Asp Val Gln Glu Glu 160
165 170aaa ggc att aag tac aaa ttt gaa gta
tat gag aag aat gat taa 1056Lys Gly Ile Lys Tyr Lys Phe Glu Val
Tyr Glu Lys Asn Asp 175 180
185tatgaaggtg ttttctagtt taagttgttc cccctccctc tgaaaaaagt atgtattttt
1116acattagaaa aggttttttg ttgactttag atctataatt atttctaagc aactagtttt
1176tattccccac tactcttgtc tctatcagat accatttatg agacattctt gctataacta
1236agtgcttctc caagacccca actgagtccc cagcacctgc tacagtgagc tgccattcca
1296cacccatcac atgtggcact cttgccagtc cttgacattg tcgggctttt cacatgttgg
1356taatatttat taaagatgaa gatccacata cccttcaact gagcagtttc actagtggaa
1416ataccaaaag cttcctacgt gtatatccag aggtttgtag ataaatgttg ccaccttgtt
1476tgtaacagtg aaaaattgaa aacaacctgg aagtccagtg atgggaaaat gagtatgttt
1536ctgtcttaga ttggggaacc caaagcagat tgcaagactg aaatttcagt gaaagcagtg
1596tatttgctag gtcataccag aaatcatcaa ttgaggtacg gagaaactga actgagaagg
1656taagaaaagc aatttaaagt cagcgagcag gttctcattg ataacaagct ccatactgct
1716gagatacagg gaaatggagg ggggaaagct ggagtattga tcccgccccc ctccttggtt
1776gtcagctccc tgtcctgtgt gtgggcggaa catagtccag ctgctctata gcaagtctca
1836ggtgtttgca gtaagaagct gctggcatgc acgggaacag tgaatgccaa acacttaaag
1896caattcgatg tttaagtatg taagttcttt tttttttaga cagcgtttcg ctcttgttgc
1956ccaggctagc atgcaatggt gtgacctcgg cttactgcaa cctccgcctt cccagattca
2016agcgattctc ctgcctcagg ctcccaagta gctaggacca ggtgcgcgcc accacgcccg
2076gctaattttt gtattttgta tttttagtag agatggggtt tcaccatgtt ggtcaggcta
2136gtctcgaact cgtgaccgca agcgattcac ccacctcagc ctcccaaagt gctgggatta
2196ccggcttgag ccaccacacc cggcacatct tcattctttt tatgtagtaa aaagtataag
2256gccacacatg gtttatttga agtattttat aatttaaaaa aatacagaag caggaaaacc
2316aattataagt tcaagtgagg gatgatggtt gcttgaacca aagggttgca tgtagtaaga
2376aattgtgatt taagatatat tttaaagtta taagtagcag gatattctga tggagtttga
2436ctttggtttt gggcccaggg agtttcagat gcctttgaga aatgaatgaa gtagagagaa
2496aataaaagaa aaaccagcca ggcacagtgg ctcacacctg taatcccagc gctttgggag
2556gctaaggcag gcagatcact tgagaccagc ttgggcaaca tggcaaagcc ccatctctac
2616aaaaaacaca aaaattagct gggcattgtg gcgcacacct gtattcccat ctagtcagga
2676agctgagatg gaagaattaa ttgagcccac gagttcaagg ctgcagtgag tcgtgattgt
2736gccactgcac tccagccggg gtgacagaag agaccttgtc tcgaaaagga atctgaaaac
2796aatggaacca tgccttcata attctagaaa gttattttca actgataaat ctatattcac
2856ccaaataatc aagggtgaag gtaaaataat acatttttag acaagcaaag actcaggggt
2916tacctccatg tgcccttttt agggaagctg ttggagaaaa tactccagca aaatgaagga
2976gtacacaaac cagagaatga catgaatcca gcaaatagga tccaacacag gcaatattcc
3036agctatggag ctagctttaa aaaggaacag taaaaatatt aatcggttag ctgggtggaa
3096tggcccatgc ctgtagtccc agctactcag gaggctcagc agcaggacga cttgagccca
3156agagttccag accagcctgg ccaccttagt gagatccctt ctcttaaaaa taataactta
3216ttgccagatt tggggcattt ggaaagaagt tcattgaaga taaagcaaaa gtaaaaaaaa
3276aaaaaaaaaa aacaagggga aagggttggt taggcaatca ttctagggca gaaagaagta
3336caggatagga agagcataat acactgtttt tctcaacaag gagcagtatg tacacagtca
3396taatgatgtg actgcttagc ccctaaatat ggtaactact ctgggacaat atgggaggaa
3456aagtgaagat tgtgatggtg taagagctaa atcctcatct gtcatatcca gaaatcacta
3516tataatatat aataatgaaa tgactaagtt atgtgaggaa aaaaacagaa gacattgcta
3576aaagagttaa aagtcattgc tctggagaat taggagggat ggggcagggg actgttagga
3636tgcattataa actgaaaagc ctttttaaaa ttttatgtat taatatatgc attcacttga
3696aaaactaaaa aaaaacaata atttggaaaa acccatgaag gtaactaacg gaaggaaaaa
3756ctaagagaat gaaaagtatt tgcctctgga aagaacaact ggcaggactg ttgttttcat
3816tgtaagactt ttggagccat ttaattgtac ttaaccattt tcatctattt ctttaataag
3876aacaattcca tcttaataaa gagttacact tgttaataag taaaaaaaaa aaaaaa
39326187PRTHomo sapiens 6Met Val Gly Ser Leu Asn Cys Ile Val Ala Val Ser
Gln Asn Met Gly1 5 10
15Ile Gly Lys Asn Gly Asp Leu Pro Trp Pro Pro Leu Arg Asn Glu Phe
20 25 30Arg Tyr Phe Gln Arg Met Thr
Thr Thr Ser Ser Val Glu Gly Lys Gln 35 40
45Asn Leu Val Ile Met Gly Lys Lys Thr Trp Phe Ser Ile Pro Glu
Lys 50 55 60Asn Arg Pro Leu Lys Gly
Arg Ile Asn Leu Val Leu Ser Arg Glu Leu65 70
75 80Lys Glu Pro Pro Gln Gly Ala His Phe Leu Ser
Arg Ser Leu Asp Asp 85 90
95Ala Leu Lys Leu Thr Glu Gln Pro Glu Leu Ala Asn Lys Val Asp Met
100 105 110Val Trp Ile Val Gly Gly
Ser Ser Val Tyr Lys Glu Ala Met Asn His 115 120
125Pro Gly His Leu Lys Leu Phe Val Thr Arg Ile Met Gln Asp
Phe Glu 130 135 140Ser Asp Thr Phe Phe
Pro Glu Ile Asp Leu Glu Lys Tyr Lys Leu Leu145 150
155 160Pro Glu Tyr Pro Gly Val Leu Ser Asp Val
Gln Glu Glu Lys Gly Ile 165 170
175Lys Tyr Lys Phe Glu Val Tyr Glu Lys Asn Asp 180
18571536DNAHomo sapiens 7gggggggggg ggaccacttg gcctgcctcc
gtcccgccgc gccacttggc ctgcctccgt 60cccgccgcgc cacttcgcct gcctccgtcc
cccgcccgcc gcgccatgcc tgtggccggc 120tcggagctgc cgcgccggcc cttgcccccc
gccgcacagg agcgggacgc cgagccgcgt 180ccgccgcacg gggagctgca gtacctgggg
cagatccaac acatcctccg ctgcggcgtc 240aggaaggacg accgcacggg caccggcacc
ctgtcggtat tcggcatgca ggcgcgctac 300agcctgagag atgaattccc tctgctgaca
accaaacgtg tgttctggaa gggtgttttg 360gaggagttgc tgtggtttat caagggatcc
acaaatgcta aagagctgtc ttccaaggga 420gtgaaaatct gggatgccaa tggatcccga
gactttttgg acagcctggg attctccacc 480agagaagaag gggacttggg cccagtttat
ggcttccagt ggaggcattt tggggcagaa 540tacagagata tggaatcaga ttattcagga
cagggagttg accaactgca aagagtgatt 600gacaccatca aaaccaaccc tgacgacaga
agaatcatca tgtgcgcttg gaatccaaga 660gatcttcctc tgatggcgct gcctccatgc
catgccctct gccagttcta tgtggtgaac 720agtgagctgt cctgccagct gtaccagaga
tcgggagaca tgggcctcgg tgtgcctttc 780aacatcgcca gctacgccct gctcacgtac
atgattgcgc acatcacggg cctgaagcca 840ggtgacttta tacacacttt gggagatgca
catatttacc tgaatcacat cgagccactg 900aaaattcagc ttcagcgaga acccagacct
ttcccaaagc tcaggattct tcgaaaagtt 960gagaaaattg atgacttcaa agctgaagac
tttcagattg aagggtacaa tccgcatcca 1020actattaaaa tggaaatggc tgtttagggt
gctttcaaag gagcttgaag gatattgtca 1080gtctttaggg gttgggctgg atgccgaggt
aaaagttctt tttgctctaa aagaaaaagg 1140aactaggtca aaaatctgtc cgtgacctat
cagttattaa tttttaagga tgttgccact 1200ggcaaatgta actgtgccag ttctttccat
aataaaaggc tttgagttaa ctcactgagg 1260gtatctgaca atgctgaggt tatgaacaaa
gtgaggagaa tgaaatgtat gtgctcttag 1320caaaaacatg tatgtgcatt tcaatcccac
gtacttataa agaaggttgg tgaatttcac 1380aagctatttt tggaatattt ttagaatatt
ttaagaattt cacaagctat tccctcaaat 1440ctgagggagc tgagtaacac catcgatcat
gatgtagagt gtggttatga actttatagt 1500tgttttatat gttgctataa taaagaagtg
ttctgc 153681536DNAHomo
sapiensCDS(106)..(1047) 8gggggggggg ggaccacttg gcctgcctcc gtcccgccgc
gccacttggc ctgcctccgt 60cccgccgcgc cacttcgcct gcctccgtcc cccgcccgcc
gcgcc atg cct gtg gcc 117
Met Pro Val Ala 1ggc
tcg gag ctg ccg cgc cgg ccc ttg ccc ccc gcc gca cag gag cgg 165Gly
Ser Glu Leu Pro Arg Arg Pro Leu Pro Pro Ala Ala Gln Glu Arg5
10 15 20gac gcc gag ccg cgt ccg
ccg cac ggg gag ctg cag tac ctg ggg cag 213Asp Ala Glu Pro Arg Pro
Pro His Gly Glu Leu Gln Tyr Leu Gly Gln 25
30 35atc caa cac atc ctc cgc tgc ggc gtc agg aag gac
gac cgc acg ggc 261Ile Gln His Ile Leu Arg Cys Gly Val Arg Lys Asp
Asp Arg Thr Gly 40 45 50acc
ggc acc ctg tcg gta ttc ggc atg cag gcg cgc tac agc ctg aga 309Thr
Gly Thr Leu Ser Val Phe Gly Met Gln Ala Arg Tyr Ser Leu Arg 55
60 65gat gaa ttc cct ctg ctg aca acc aaa
cgt gtg ttc tgg aag ggt gtt 357Asp Glu Phe Pro Leu Leu Thr Thr Lys
Arg Val Phe Trp Lys Gly Val 70 75
80ttg gag gag ttg ctg tgg ttt atc aag gga tcc aca aat gct aaa gag
405Leu Glu Glu Leu Leu Trp Phe Ile Lys Gly Ser Thr Asn Ala Lys Glu85
90 95 100ctg tct tcc aag
gga gtg aaa atc tgg gat gcc aat gga tcc cga gac 453Leu Ser Ser Lys
Gly Val Lys Ile Trp Asp Ala Asn Gly Ser Arg Asp 105
110 115ttt ttg gac agc ctg gga ttc tcc acc aga
gaa gaa ggg gac ttg ggc 501Phe Leu Asp Ser Leu Gly Phe Ser Thr Arg
Glu Glu Gly Asp Leu Gly 120 125
130cca gtt tat ggc ttc cag tgg agg cat ttt ggg gca gaa tac aga gat
549Pro Val Tyr Gly Phe Gln Trp Arg His Phe Gly Ala Glu Tyr Arg Asp
135 140 145atg gaa tca gat tat tca gga
cag gga gtt gac caa ctg caa aga gtg 597Met Glu Ser Asp Tyr Ser Gly
Gln Gly Val Asp Gln Leu Gln Arg Val 150 155
160att gac acc atc aaa acc aac cct gac gac aga aga atc atc atg tgc
645Ile Asp Thr Ile Lys Thr Asn Pro Asp Asp Arg Arg Ile Ile Met Cys165
170 175 180gct tgg aat cca
aga gat ctt cct ctg atg gcg ctg cct cca tgc cat 693Ala Trp Asn Pro
Arg Asp Leu Pro Leu Met Ala Leu Pro Pro Cys His 185
190 195gcc ctc tgc cag ttc tat gtg gtg aac agt
gag ctg tcc tgc cag ctg 741Ala Leu Cys Gln Phe Tyr Val Val Asn Ser
Glu Leu Ser Cys Gln Leu 200 205
210tac cag aga tcg gga gac atg ggc ctc ggt gtg cct ttc aac atc gcc
789Tyr Gln Arg Ser Gly Asp Met Gly Leu Gly Val Pro Phe Asn Ile Ala
215 220 225agc tac gcc ctg ctc acg tac
atg att gcg cac atc acg ggc ctg aag 837Ser Tyr Ala Leu Leu Thr Tyr
Met Ile Ala His Ile Thr Gly Leu Lys 230 235
240cca ggt gac ttt ata cac act ttg gga gat gca cat att tac ctg aat
885Pro Gly Asp Phe Ile His Thr Leu Gly Asp Ala His Ile Tyr Leu Asn245
250 255 260cac atc gag cca
ctg aaa att cag ctt cag cga gaa ccc aga cct ttc 933His Ile Glu Pro
Leu Lys Ile Gln Leu Gln Arg Glu Pro Arg Pro Phe 265
270 275cca aag ctc agg att ctt cga aaa gtt gag
aaa att gat gac ttc aaa 981Pro Lys Leu Arg Ile Leu Arg Lys Val Glu
Lys Ile Asp Asp Phe Lys 280 285
290gct gaa gac ttt cag att gaa ggg tac aat ccg cat cca act att aaa
1029Ala Glu Asp Phe Gln Ile Glu Gly Tyr Asn Pro His Pro Thr Ile Lys
295 300 305atg gaa atg gct gtt tag
ggtgctttca aaggagcttg aaggatattg 1077Met Glu Met Ala Val
310tcagtcttta ggggttgggc tggatgccga ggtaaaagtt ctttttgctc taaaagaaaa
1137aggaactagg tcaaaaatct gtccgtgacc tatcagttat taatttttaa ggatgttgcc
1197actggcaaat gtaactgtgc cagttctttc cataataaaa ggctttgagt taactcactg
1257agggtatctg acaatgctga ggttatgaac aaagtgagga gaatgaaatg tatgtgctct
1317tagcaaaaac atgtatgtgc atttcaatcc cacgtactta taaagaaggt tggtgaattt
1377cacaagctat ttttggaata tttttagaat attttaagaa tttcacaagc tattccctca
1437aatctgaggg agctgagtaa caccatcgat catgatgtag agtgtggtta tgaactttat
1497agttgtttta tatgttgcta taataaagaa gtgttctgc
15369313PRTHomo sapiens 9Met Pro Val Ala Gly Ser Glu Leu Pro Arg Arg Pro
Leu Pro Pro Ala1 5 10
15Ala Gln Glu Arg Asp Ala Glu Pro Arg Pro Pro His Gly Glu Leu Gln
20 25 30Tyr Leu Gly Gln Ile Gln His
Ile Leu Arg Cys Gly Val Arg Lys Asp 35 40
45Asp Arg Thr Gly Thr Gly Thr Leu Ser Val Phe Gly Met Gln Ala
Arg 50 55 60Tyr Ser Leu Arg Asp Glu
Phe Pro Leu Leu Thr Thr Lys Arg Val Phe65 70
75 80Trp Lys Gly Val Leu Glu Glu Leu Leu Trp Phe
Ile Lys Gly Ser Thr 85 90
95Asn Ala Lys Glu Leu Ser Ser Lys Gly Val Lys Ile Trp Asp Ala Asn
100 105 110Gly Ser Arg Asp Phe Leu
Asp Ser Leu Gly Phe Ser Thr Arg Glu Glu 115 120
125Gly Asp Leu Gly Pro Val Tyr Gly Phe Gln Trp Arg His Phe
Gly Ala 130 135 140Glu Tyr Arg Asp Met
Glu Ser Asp Tyr Ser Gly Gln Gly Val Asp Gln145 150
155 160Leu Gln Arg Val Ile Asp Thr Ile Lys Thr
Asn Pro Asp Asp Arg Arg 165 170
175Ile Ile Met Cys Ala Trp Asn Pro Arg Asp Leu Pro Leu Met Ala Leu
180 185 190Pro Pro Cys His Ala
Leu Cys Gln Phe Tyr Val Val Asn Ser Glu Leu 195
200 205Ser Cys Gln Leu Tyr Gln Arg Ser Gly Asp Met Gly
Leu Gly Val Pro 210 215 220Phe Asn Ile
Ala Ser Tyr Ala Leu Leu Thr Tyr Met Ile Ala His Ile225
230 235 240Thr Gly Leu Lys Pro Gly Asp
Phe Ile His Thr Leu Gly Asp Ala His 245
250 255Ile Tyr Leu Asn His Ile Glu Pro Leu Lys Ile Gln
Leu Gln Arg Glu 260 265 270Pro
Arg Pro Phe Pro Lys Leu Arg Ile Leu Arg Lys Val Glu Lys Ile 275
280 285Asp Asp Phe Lys Ala Glu Asp Phe Gln
Ile Glu Gly Tyr Asn Pro His 290 295
300Pro Thr Ile Lys Met Glu Met Ala Val305
310101280DNAHomo sapiens 10tgccgcagcc cccgcccgcc cgcagagctt ttgaaaggcg
gcgggaggcg gcgagcgcca 60tggccagtcc gggctgcctg ctgtgcgtgc tgggcctgct
actctgcggg gcggcgagcc 120tcgagctgtc tagaccccac ggcgacaccg ccaagaagcc
catcatcgga atattaatgc 180aaaaatgccg taataaagtc atgaaaaact atggaagata
ctatattgct gcgtcctatg 240taaagtactt ggagtctgca ggtgcgagag ttgtaccagt
aaggctggat cttacagaga 300aagactatga aatacttttc aaatctatta atggaatcct
tttccctgga ggaagtgttg 360acctcagacg ctcagattat gctaaagtgg ccaaaatatt
ttataacttg tccatacaga 420gttttgatga tggagactat tttcctgtgt ggggcacatg
ccttggattt gaagagcttt 480cactgctgat tagtggagag tgcttattaa ctgccacaga
tactgttgac gtggcaatgc 540cgctgaactt cactggaggt caattgcaca gcagaatgtt
ccagaatttt cctactgagt 600tgttgctgtc attagcagta gaacctctga ctgccaattt
ccataagtgg agcctctccg 660tgaagaattt tacaatgaat gaaaagttaa agaagttttt
caatgtctta actacaaata 720cagatggcaa gattgagttt atttcaacaa tggaaggata
taagtatcca gtatatggtg 780tccagtggca tccagagaaa gcaccttatg agtggaagaa
tttggatggc atttcccatg 840cacctaatgc tgtgaaaacc gcattttatt tagcagagtt
ttttgttaat gaagctcgga 900aaaacaacca tcattttaaa tctgaatctg aagaggagaa
agcattgatt tatcagttca 960gtccaattta tactggaaat atttcttcat ttcagcaatg
ttacatattt gattgaaagt 1020cttcaatttg ttaacagagc aaatttgaat aattccatga
ttaaactgtt agaataactt 1080gctactcatg gcaagattag gaagtcacag attcttttct
ataatgtgcc tggctctgat 1140tcttcattat gtatgtgact atttatataa cattagataa
ttaaatagtg agacataaat 1200agagtgcttt ttcatggaaa agccttctta tatctgaaga
ttgaaaaata aatttactga 1260aatacaaaaa aaaaaaaaaa
1280111280DNAHomo sapiensCDS(60)..(1016)
11tgccgcagcc cccgcccgcc cgcagagctt ttgaaaggcg gcgggaggcg gcgagcgcc
59atg gcc agt ccg ggc tgc ctg ctg tgc gtg ctg ggc ctg cta ctc tgc
107Met Ala Ser Pro Gly Cys Leu Leu Cys Val Leu Gly Leu Leu Leu Cys1
5 10 15ggg gcg gcg agc ctc gag
ctg tct aga ccc cac ggc gac acc gcc aag 155Gly Ala Ala Ser Leu Glu
Leu Ser Arg Pro His Gly Asp Thr Ala Lys 20 25
30aag ccc atc atc gga ata tta atg caa aaa tgc cgt aat
aaa gtc atg 203Lys Pro Ile Ile Gly Ile Leu Met Gln Lys Cys Arg Asn
Lys Val Met 35 40 45aaa aac tat
gga aga tac tat att gct gcg tcc tat gta aag tac ttg 251Lys Asn Tyr
Gly Arg Tyr Tyr Ile Ala Ala Ser Tyr Val Lys Tyr Leu 50
55 60gag tct gca ggt gcg aga gtt gta cca gta agg ctg
gat ctt aca gag 299Glu Ser Ala Gly Ala Arg Val Val Pro Val Arg Leu
Asp Leu Thr Glu65 70 75
80aaa gac tat gaa ata ctt ttc aaa tct att aat gga atc ctt ttc cct
347Lys Asp Tyr Glu Ile Leu Phe Lys Ser Ile Asn Gly Ile Leu Phe Pro
85 90 95gga gga agt gtt gac ctc
aga cgc tca gat tat gct aaa gtg gcc aaa 395Gly Gly Ser Val Asp Leu
Arg Arg Ser Asp Tyr Ala Lys Val Ala Lys 100
105 110ata ttt tat aac ttg tcc ata cag agt ttt gat gat
gga gac tat ttt 443Ile Phe Tyr Asn Leu Ser Ile Gln Ser Phe Asp Asp
Gly Asp Tyr Phe 115 120 125cct gtg
tgg ggc aca tgc ctt gga ttt gaa gag ctt tca ctg ctg att 491Pro Val
Trp Gly Thr Cys Leu Gly Phe Glu Glu Leu Ser Leu Leu Ile 130
135 140agt gga gag tgc tta tta act gcc aca gat act
gtt gac gtg gca atg 539Ser Gly Glu Cys Leu Leu Thr Ala Thr Asp Thr
Val Asp Val Ala Met145 150 155
160ccg ctg aac ttc act gga ggt caa ttg cac agc aga atg ttc cag aat
587Pro Leu Asn Phe Thr Gly Gly Gln Leu His Ser Arg Met Phe Gln Asn
165 170 175ttt cct act gag ttg
ttg ctg tca tta gca gta gaa cct ctg act gcc 635Phe Pro Thr Glu Leu
Leu Leu Ser Leu Ala Val Glu Pro Leu Thr Ala 180
185 190aat ttc cat aag tgg agc ctc tcc gtg aag aat ttt
aca atg aat gaa 683Asn Phe His Lys Trp Ser Leu Ser Val Lys Asn Phe
Thr Met Asn Glu 195 200 205aag tta
aag aag ttt ttc aat gtc tta act aca aat aca gat ggc aag 731Lys Leu
Lys Lys Phe Phe Asn Val Leu Thr Thr Asn Thr Asp Gly Lys 210
215 220att gag ttt att tca aca atg gaa gga tat aag
tat cca gta tat ggt 779Ile Glu Phe Ile Ser Thr Met Glu Gly Tyr Lys
Tyr Pro Val Tyr Gly225 230 235
240gtc cag tgg cat cca gag aaa gca cct tat gag tgg aag aat ttg gat
827Val Gln Trp His Pro Glu Lys Ala Pro Tyr Glu Trp Lys Asn Leu Asp
245 250 255ggc att tcc cat gca
cct aat gct gtg aaa acc gca ttt tat tta gca 875Gly Ile Ser His Ala
Pro Asn Ala Val Lys Thr Ala Phe Tyr Leu Ala 260
265 270gag ttt ttt gtt aat gaa gct cgg aaa aac aac cat
cat ttt aaa tct 923Glu Phe Phe Val Asn Glu Ala Arg Lys Asn Asn His
His Phe Lys Ser 275 280 285gaa tct
gaa gag gag aaa gca ttg att tat cag ttc agt cca att tat 971Glu Ser
Glu Glu Glu Lys Ala Leu Ile Tyr Gln Phe Ser Pro Ile Tyr 290
295 300act gga aat att tct tca ttt cag caa tgt tac
ata ttt gat tga 1016Thr Gly Asn Ile Ser Ser Phe Gln Gln Cys Tyr
Ile Phe Asp305 310 315aagtcttcaa
tttgttaaca gagcaaattt gaataattcc atgattaaac tgttagaata 1076acttgctact
catggcaaga ttaggaagtc acagattctt ttctataatg tgcctggctc 1136tgattcttca
ttatgtatgt gactatttat ataacattag ataattaaat agtgagacat 1196aaatagagtg
ctttttcatg gaaaagcctt cttatatctg aagattgaaa aataaattta 1256ctgaaataca
aaaaaaaaaa aaaa 128012318PRTHomo
sapiens 12Met Ala Ser Pro Gly Cys Leu Leu Cys Val Leu Gly Leu Leu Leu
Cys1 5 10 15Gly Ala Ala
Ser Leu Glu Leu Ser Arg Pro His Gly Asp Thr Ala Lys 20
25 30Lys Pro Ile Ile Gly Ile Leu Met Gln Lys
Cys Arg Asn Lys Val Met 35 40
45Lys Asn Tyr Gly Arg Tyr Tyr Ile Ala Ala Ser Tyr Val Lys Tyr Leu 50
55 60Glu Ser Ala Gly Ala Arg Val Val Pro
Val Arg Leu Asp Leu Thr Glu65 70 75
80Lys Asp Tyr Glu Ile Leu Phe Lys Ser Ile Asn Gly Ile Leu
Phe Pro 85 90 95Gly Gly
Ser Val Asp Leu Arg Arg Ser Asp Tyr Ala Lys Val Ala Lys 100
105 110Ile Phe Tyr Asn Leu Ser Ile Gln Ser
Phe Asp Asp Gly Asp Tyr Phe 115 120
125Pro Val Trp Gly Thr Cys Leu Gly Phe Glu Glu Leu Ser Leu Leu Ile
130 135 140Ser Gly Glu Cys Leu Leu Thr
Ala Thr Asp Thr Val Asp Val Ala Met145 150
155 160Pro Leu Asn Phe Thr Gly Gly Gln Leu His Ser Arg
Met Phe Gln Asn 165 170
175Phe Pro Thr Glu Leu Leu Leu Ser Leu Ala Val Glu Pro Leu Thr Ala
180 185 190Asn Phe His Lys Trp Ser
Leu Ser Val Lys Asn Phe Thr Met Asn Glu 195 200
205Lys Leu Lys Lys Phe Phe Asn Val Leu Thr Thr Asn Thr Asp
Gly Lys 210 215 220Ile Glu Phe Ile Ser
Thr Met Glu Gly Tyr Lys Tyr Pro Val Tyr Gly225 230
235 240Val Gln Trp His Pro Glu Lys Ala Pro Tyr
Glu Trp Lys Asn Leu Asp 245 250
255Gly Ile Ser His Ala Pro Asn Ala Val Lys Thr Ala Phe Tyr Leu Ala
260 265 270Glu Phe Phe Val Asn
Glu Ala Arg Lys Asn Asn His His Phe Lys Ser 275
280 285Glu Ser Glu Glu Glu Lys Ala Leu Ile Tyr Gln Phe
Ser Pro Ile Tyr 290 295 300Thr Gly Asn
Ile Ser Ser Phe Gln Gln Cys Tyr Ile Phe Asp305 310
315132158DNAHomo sapiens 13gcgcggcata acgacccagg tcgcggcgcg
gcggggcttg agcgcgtggc cggtgccgca 60ggagccgagc atggagtacc aggatgccgt
gcgcatgctc aataccctgc agaccaatgc 120cggctacctg gagcaggtga agcgccagcg
gggtgaccct cagacacagt tggaagccat 180ggaactgtac ctggcacgga gtgggctgca
ggtggaggac ttggaccggc tgaacatcat 240ccacgtcact gggacgaagg ggaagggctc
cacctgtgcc ttcacggaat gtatcctccg 300aagctatggc ctgaagacgg gattctttag
ctctccccac ctggtgcagg ttcgggagcg 360gatccgcatc aatgggcagc ccatcagtcc
tgagctcttc accaagtact tctggcgcct 420ctaccaccgg ctggaggaga ccaaggatgg
cagctgtgtc tccatgcccc cctacttccg 480cttcctgaca ctcatggcct tccacgtctt
cctccaagag aaggtggacc tggcagtggt 540ggaggtgggc attggcgggg cttatgactg
caccaacatc atcaggaagc ctgtggtgtg 600cggagtctcc tctcttggca tcgaccacac
cagcctcctg ggggatacgg tggagaagat 660cgcatggcag aaagggggca tctttaagca
aggtgtccct gccttcactg tgctccaacc 720tgaaggtccc ctggcagtgc tgagggaccg
agcccagcag atctcatgtc ctctatacct 780gtgtccgatg ctggaggccc tcgaggaagg
ggggccgccg ctgaccctgg gcctggaggg 840ggagcaccag cggtccaacg ccgccttggc
cttgcagctg gcccactgct ggctgcagcg 900gcaggaccgc catggtgctg gggagccaaa
ggcatccagg ccagggctcc tgtggcagct 960gcccctggca cctgtgttcc agcccacatc
ccacatgcgg ctcgggcttc ggaacacgga 1020gtggccgggc cggacgcagg tgctgcggcg
cgggcccctc acctggtacc tggacggtgc 1080gcacaccgcc agcagcgcgc aggcctgcgt
gcgctggttc cgccaggcgc tgcagggccg 1140cgagaggccg agcggtggcc ccgaggttcg
agtcttgctc ttcaatgcta ccggggaccg 1200ggacccggcg gccctgctga agctgctgca
gccctgccag tttgactatg ccgtcttctg 1260ccctaacctg acagaggtgt catccacagg
caacgcagac caacagaact tcacagtgac 1320actggaccag gtcctgctcc gctgcctgga
acaccagcag cactggaacc acctggacga 1380agagcaggcc agcccggacc tctggagtgc
ccccagccca gagcccggtg ggtccgcatc 1440cctgcttctg gcgccccacc caccccacac
ctgcagtgcc agctccctcg tcttcagctg 1500catttcacat gccttgcaat ggatcagcca
aggccgagac cccatcttcc agccacctag 1560tcccccaaag ggcctcctca cccaccctgt
ggctcacagt ggggccagca tactccgtga 1620ggctgctgcc atccatgtgc tagtcactgg
cagcctgcac ctggtgggtg gtgtcctgaa 1680gctgctggag cccgcactgt cccagtagcc
aaggcccggg gttggaggtg ggagcttccc 1740acacctgcct gcgttctccc catgaactta
catactaggt gccttttgtt tttggctttc 1800ctggttctgt ctagactggc ctaggggcca
gggctttggg atgggaggcc gggagaggat 1860gtctttttta aggctctgtg ccttggtctc
tccttcctct tggctgagat agcagagggg 1920ctccccgggt ctctcactgt tgcagtggcc
tggccgttca gcctgtctcc cccaacaccc 1980cgcctgcctc ctggctcagg cccagcttat
tgtgtgcgct gcctggccag gccctgggtc 2040ttgccatgtg ctgggtggta gatttcctcc
tcccagtgcc ttctgggaag ggagagggcc 2100tctgcctggg acactgcggg acagagggtg
gctggagtga attaaagcct ttgttttt 2158142158DNAHomo
sapiensCDS(71)..(1708) 14gcgcggcata acgacccagg tcgcggcgcg gcggggcttg
agcgcgtggc cggtgccgca 60ggagccgagc atg gag tac cag gat gcc gtg cgc
atg ctc aat acc ctg 109 Met Glu Tyr Gln Asp Ala Val Arg
Met Leu Asn Thr Leu 1 5 10cag
acc aat gcc ggc tac ctg gag cag gtg aag cgc cag cgg ggt gac 157Gln
Thr Asn Ala Gly Tyr Leu Glu Gln Val Lys Arg Gln Arg Gly Asp 15
20 25cct cag aca cag ttg gaa gcc atg gaa ctg
tac ctg gca cgg agt ggg 205Pro Gln Thr Gln Leu Glu Ala Met Glu Leu
Tyr Leu Ala Arg Ser Gly30 35 40
45ctg cag gtg gag gac ttg gac cgg ctg aac atc atc cac gtc act
ggg 253Leu Gln Val Glu Asp Leu Asp Arg Leu Asn Ile Ile His Val Thr
Gly 50 55 60acg aag ggg
aag ggc tcc acc tgt gcc ttc acg gaa tgt atc ctc cga 301Thr Lys Gly
Lys Gly Ser Thr Cys Ala Phe Thr Glu Cys Ile Leu Arg 65
70 75agc tat ggc ctg aag acg gga ttc ttt agc
tct ccc cac ctg gtg cag 349Ser Tyr Gly Leu Lys Thr Gly Phe Phe Ser
Ser Pro His Leu Val Gln 80 85
90gtt cgg gag cgg atc cgc atc aat ggg cag ccc atc agt cct gag ctc
397Val Arg Glu Arg Ile Arg Ile Asn Gly Gln Pro Ile Ser Pro Glu Leu 95
100 105ttc acc aag tac ttc tgg cgc ctc
tac cac cgg ctg gag gag acc aag 445Phe Thr Lys Tyr Phe Trp Arg Leu
Tyr His Arg Leu Glu Glu Thr Lys110 115
120 125gat ggc agc tgt gtc tcc atg ccc ccc tac ttc cgc
ttc ctg aca ctc 493Asp Gly Ser Cys Val Ser Met Pro Pro Tyr Phe Arg
Phe Leu Thr Leu 130 135
140atg gcc ttc cac gtc ttc ctc caa gag aag gtg gac ctg gca gtg gtg
541Met Ala Phe His Val Phe Leu Gln Glu Lys Val Asp Leu Ala Val Val
145 150 155gag gtg ggc att ggc ggg
gct tat gac tgc acc aac atc atc agg aag 589Glu Val Gly Ile Gly Gly
Ala Tyr Asp Cys Thr Asn Ile Ile Arg Lys 160 165
170cct gtg gtg tgc gga gtc tcc tct ctt ggc atc gac cac acc
agc ctc 637Pro Val Val Cys Gly Val Ser Ser Leu Gly Ile Asp His Thr
Ser Leu 175 180 185ctg ggg gat acg gtg
gag aag atc gca tgg cag aaa ggg ggc atc ttt 685Leu Gly Asp Thr Val
Glu Lys Ile Ala Trp Gln Lys Gly Gly Ile Phe190 195
200 205aag caa ggt gtc cct gcc ttc act gtg ctc
caa cct gaa ggt ccc ctg 733Lys Gln Gly Val Pro Ala Phe Thr Val Leu
Gln Pro Glu Gly Pro Leu 210 215
220gca gtg ctg agg gac cga gcc cag cag atc tca tgt cct cta tac ctg
781Ala Val Leu Arg Asp Arg Ala Gln Gln Ile Ser Cys Pro Leu Tyr Leu
225 230 235tgt ccg atg ctg gag gcc
ctc gag gaa ggg ggg ccg ccg ctg acc ctg 829Cys Pro Met Leu Glu Ala
Leu Glu Glu Gly Gly Pro Pro Leu Thr Leu 240 245
250ggc ctg gag ggg gag cac cag cgg tcc aac gcc gcc ttg gcc
ttg cag 877Gly Leu Glu Gly Glu His Gln Arg Ser Asn Ala Ala Leu Ala
Leu Gln 255 260 265ctg gcc cac tgc tgg
ctg cag cgg cag gac cgc cat ggt gct ggg gag 925Leu Ala His Cys Trp
Leu Gln Arg Gln Asp Arg His Gly Ala Gly Glu270 275
280 285cca aag gca tcc agg cca ggg ctc ctg tgg
cag ctg ccc ctg gca cct 973Pro Lys Ala Ser Arg Pro Gly Leu Leu Trp
Gln Leu Pro Leu Ala Pro 290 295
300gtg ttc cag ccc aca tcc cac atg cgg ctc ggg ctt cgg aac acg gag
1021Val Phe Gln Pro Thr Ser His Met Arg Leu Gly Leu Arg Asn Thr Glu
305 310 315tgg ccg ggc cgg acg cag
gtg ctg cgg cgc ggg ccc ctc acc tgg tac 1069Trp Pro Gly Arg Thr Gln
Val Leu Arg Arg Gly Pro Leu Thr Trp Tyr 320 325
330ctg gac ggt gcg cac acc gcc agc agc gcg cag gcc tgc gtg
cgc tgg 1117Leu Asp Gly Ala His Thr Ala Ser Ser Ala Gln Ala Cys Val
Arg Trp 335 340 345ttc cgc cag gcg ctg
cag ggc cgc gag agg ccg agc ggt ggc ccc gag 1165Phe Arg Gln Ala Leu
Gln Gly Arg Glu Arg Pro Ser Gly Gly Pro Glu350 355
360 365gtt cga gtc ttg ctc ttc aat gct acc ggg
gac cgg gac ccg gcg gcc 1213Val Arg Val Leu Leu Phe Asn Ala Thr Gly
Asp Arg Asp Pro Ala Ala 370 375
380ctg ctg aag ctg ctg cag ccc tgc cag ttt gac tat gcc gtc ttc tgc
1261Leu Leu Lys Leu Leu Gln Pro Cys Gln Phe Asp Tyr Ala Val Phe Cys
385 390 395cct aac ctg aca gag gtg
tca tcc aca ggc aac gca gac caa cag aac 1309Pro Asn Leu Thr Glu Val
Ser Ser Thr Gly Asn Ala Asp Gln Gln Asn 400 405
410ttc aca gtg aca ctg gac cag gtc ctg ctc cgc tgc ctg gaa
cac cag 1357Phe Thr Val Thr Leu Asp Gln Val Leu Leu Arg Cys Leu Glu
His Gln 415 420 425cag cac tgg aac cac
ctg gac gaa gag cag gcc agc ccg gac ctc tgg 1405Gln His Trp Asn His
Leu Asp Glu Glu Gln Ala Ser Pro Asp Leu Trp430 435
440 445agt gcc ccc agc cca gag ccc ggt ggg tcc
gca tcc ctg ctt ctg gcg 1453Ser Ala Pro Ser Pro Glu Pro Gly Gly Ser
Ala Ser Leu Leu Leu Ala 450 455
460ccc cac cca ccc cac acc tgc agt gcc agc tcc ctc gtc ttc agc tgc
1501Pro His Pro Pro His Thr Cys Ser Ala Ser Ser Leu Val Phe Ser Cys
465 470 475att tca cat gcc ttg caa
tgg atc agc caa ggc cga gac ccc atc ttc 1549Ile Ser His Ala Leu Gln
Trp Ile Ser Gln Gly Arg Asp Pro Ile Phe 480 485
490cag cca cct agt ccc cca aag ggc ctc ctc acc cac cct gtg
gct cac 1597Gln Pro Pro Ser Pro Pro Lys Gly Leu Leu Thr His Pro Val
Ala His 495 500 505agt ggg gcc agc ata
ctc cgt gag gct gct gcc atc cat gtg cta gtc 1645Ser Gly Ala Ser Ile
Leu Arg Glu Ala Ala Ala Ile His Val Leu Val510 515
520 525act ggc agc ctg cac ctg gtg ggt ggt gtc
ctg aag ctg ctg gag ccc 1693Thr Gly Ser Leu His Leu Val Gly Gly Val
Leu Lys Leu Leu Glu Pro 530 535
540gca ctg tcc cag tag ccaaggcccg gggttggagg tgggagcttc ccacacctgc
1748Ala Leu Ser Gln 545ctgcgttctc cccatgaact tacatactag
gtgccttttg tttttggctt tcctggttct 1808gtctagactg gcctaggggc cagggctttg
ggatgggagg ccgggagagg atgtcttttt 1868taaggctctg tgccttggtc tctccttcct
cttggctgag atagcagagg ggctccccgg 1928gtctctcact gttgcagtgg cctggccgtt
cagcctgtct cccccaacac cccgcctgcc 1988tcctggctca ggcccagctt attgtgtgcg
ctgcctggcc aggccctggg tcttgccatg 2048tgctgggtgg tagatttcct cctcccagtg
ccttctggga agggagaggg cctctgcctg 2108ggacactgcg ggacagaggg tggctggagt
gaattaaagc ctttgttttt 215815545PRTHomo sapiens 15Met Glu Tyr
Gln Asp Ala Val Arg Met Leu Asn Thr Leu Gln Thr Asn1 5
10 15Ala Gly Tyr Leu Glu Gln Val Lys Arg
Gln Arg Gly Asp Pro Gln Thr 20 25
30Gln Leu Glu Ala Met Glu Leu Tyr Leu Ala Arg Ser Gly Leu Gln Val
35 40 45Glu Asp Leu Asp Arg Leu Asn
Ile Ile His Val Thr Gly Thr Lys Gly 50 55
60Lys Gly Ser Thr Cys Ala Phe Thr Glu Cys Ile Leu Arg Ser Tyr Gly65
70 75 80Leu Lys Thr Gly
Phe Phe Ser Ser Pro His Leu Val Gln Val Arg Glu 85
90 95Arg Ile Arg Ile Asn Gly Gln Pro Ile Ser
Pro Glu Leu Phe Thr Lys 100 105
110Tyr Phe Trp Arg Leu Tyr His Arg Leu Glu Glu Thr Lys Asp Gly Ser
115 120 125Cys Val Ser Met Pro Pro Tyr
Phe Arg Phe Leu Thr Leu Met Ala Phe 130 135
140His Val Phe Leu Gln Glu Lys Val Asp Leu Ala Val Val Glu Val
Gly145 150 155 160Ile Gly
Gly Ala Tyr Asp Cys Thr Asn Ile Ile Arg Lys Pro Val Val
165 170 175Cys Gly Val Ser Ser Leu Gly
Ile Asp His Thr Ser Leu Leu Gly Asp 180 185
190Thr Val Glu Lys Ile Ala Trp Gln Lys Gly Gly Ile Phe Lys
Gln Gly 195 200 205Val Pro Ala Phe
Thr Val Leu Gln Pro Glu Gly Pro Leu Ala Val Leu 210
215 220Arg Asp Arg Ala Gln Gln Ile Ser Cys Pro Leu Tyr
Leu Cys Pro Met225 230 235
240Leu Glu Ala Leu Glu Glu Gly Gly Pro Pro Leu Thr Leu Gly Leu Glu
245 250 255Gly Glu His Gln Arg
Ser Asn Ala Ala Leu Ala Leu Gln Leu Ala His 260
265 270Cys Trp Leu Gln Arg Gln Asp Arg His Gly Ala Gly
Glu Pro Lys Ala 275 280 285Ser Arg
Pro Gly Leu Leu Trp Gln Leu Pro Leu Ala Pro Val Phe Gln 290
295 300Pro Thr Ser His Met Arg Leu Gly Leu Arg Asn
Thr Glu Trp Pro Gly305 310 315
320Arg Thr Gln Val Leu Arg Arg Gly Pro Leu Thr Trp Tyr Leu Asp Gly
325 330 335Ala His Thr Ala
Ser Ser Ala Gln Ala Cys Val Arg Trp Phe Arg Gln 340
345 350Ala Leu Gln Gly Arg Glu Arg Pro Ser Gly Gly
Pro Glu Val Arg Val 355 360 365Leu
Leu Phe Asn Ala Thr Gly Asp Arg Asp Pro Ala Ala Leu Leu Lys 370
375 380Leu Leu Gln Pro Cys Gln Phe Asp Tyr Ala
Val Phe Cys Pro Asn Leu385 390 395
400Thr Glu Val Ser Ser Thr Gly Asn Ala Asp Gln Gln Asn Phe Thr
Val 405 410 415Thr Leu Asp
Gln Val Leu Leu Arg Cys Leu Glu His Gln Gln His Trp 420
425 430Asn His Leu Asp Glu Glu Gln Ala Ser Pro
Asp Leu Trp Ser Ala Pro 435 440
445Ser Pro Glu Pro Gly Gly Ser Ala Ser Leu Leu Leu Ala Pro His Pro 450
455 460Pro His Thr Cys Ser Ala Ser Ser
Leu Val Phe Ser Cys Ile Ser His465 470
475 480Ala Leu Gln Trp Ile Ser Gln Gly Arg Asp Pro Ile
Phe Gln Pro Pro 485 490
495Ser Pro Pro Lys Gly Leu Leu Thr His Pro Val Ala His Ser Gly Ala
500 505 510Ser Ile Leu Arg Glu Ala
Ala Ala Ile His Val Leu Val Thr Gly Ser 515 520
525Leu His Leu Val Gly Gly Val Leu Lys Leu Leu Glu Pro Ala
Leu Ser 530 535 540Gln545163291DNAHomo
sapiens 16accgggcaag cgggaaccag gtggccaccc ggtgtcggtt tcattttcct
ttggaatttc 60tgctttacag acagaacaat ggcagcccga gtacttataa ttggcagtgg
aggaagggaa 120catacgctgg cctggaaact tgcacagtct catcatgtca aacaagtgtt
ggttgcccca 180ggaaacgcag gcactgcctg ctctgaaaag atttcaaata ccgccatctc
aatcagtgac 240cacactgccc ttgctcaatt ctgcaaagag aagaaaattg aatttgtagt
tgttggacca 300gaagcacctc tggctgctgg gattgttggg aacctgaggt ctgcaggagt
gcaatgcttt 360ggcccaacag cagaagcggc tcagttagag tccagcaaaa ggtttgccaa
agagtttatg 420gacagacatg gaatcccaac cgcacaatgg aaggctttca ccaaacctga
agaagcctgc 480agcttcattt tgagtgcaga cttccctgct ttggttgtga aggccagtgg
tcttgcagct 540ggaaaagggg tgattgttgc aaagagcaaa gaagaggcct gcaaagctgt
acaagagatc 600atgcaggaga aagcctttgg ggcagctgga gaaacaattg tcattgaaga
acttcttgac 660ggagaagagg tgtcgtgtct gtgtttcact gatggcaaga ctgtggcccc
catgccccca 720gcacaggacc ataagcgatt actggaggga gatggtggcc ctaacacagg
gggaatggga 780gcctattgtc cagcccctca ggtttctaat gatctattac taaaaattaa
agatactgtt 840cttcagagga cagtggatgg catgcagcaa gagggtactc catatacagg
tattctctat 900gctggaataa tgctgaccaa gaatggccca aaagttctag agtttaattg
ccgttttggt 960gatccagagt gccaagtaat cctcccactt cttaaaagtg atctttatga
agtgattcag 1020tccaccttag atggactgct ctgcacatct ctgcctgttt ggctagaaaa
ccacaccgcc 1080ctaactgttg tcatggcaag taaaggttat cctggagact acaccaaggg
tgtagagata 1140acagggtttc ctgaggctca agctctagga ctggaggtgt tccatgcagg
cactgccctc 1200aaaaatggca aagtagtaac tcatgggggt agagttcttg cagtcacagc
catccgggaa 1260aatctcatat cagcccttga ggaagccaag aaaggactag ctgctataaa
gtttgaggga 1320gcaatttata ggaaagacgt cggctttcgt gccatagctt tcctccagca
gcccaggagt 1380ttgacttaca aggaatctgg agtagatatc gcagctggaa atatgctggt
caagaaaatt 1440cagcctttag caaaagccac ttccagatca ggctgtaaag ttgatcttgg
aggttttgct 1500ggtctttttg atttaaaagc agctggtttc aaagatcccc ttctggcctc
tggaacagat 1560ggcgttggaa ctaaactaaa gattgcccag ctatgcaata aacatgatac
cattggtcaa 1620gatttggtag caatgtgtgt taatgatatt ctggcacaag gagcagagcc
cctcttcttc 1680cttgattact tttcctgtgg aaaacttgac ctcagtgtaa ctgaagctgt
tgttgctgga 1740attgctaaag cttgtggaaa agctggatgt gctctccttg gaggtgaaac
agcagaaatg 1800cctgacatgt atccccctgg agagtatgac ctagctgggt ttgccgttgg
tgccatggag 1860cgagatcaga aactccctca cctggaaaga atcactgagg gtgatgttgt
tgttggaata 1920gcttcatctg gtcttcatag caatggattt agccttgtga ggaaaatcgt
tgcaaaatct 1980tccctccagt actcctctcc agcacctgat ggttgtggtg accagacttt
aggggactta 2040cttctcacgc ctaccagaat ctacagccat tcactgttac ctgtcctacg
ttcaggacat 2100gtcaaagcct ttgcccatat tactggtgga ggattactag agaacatccc
cagagtcctc 2160cctgagaaac ttggggtaga tttagatgcc cagacctgga ggatccccag
ggttttctca 2220tggttgcagc aggaaggaca cctctctgag gaagagatgg ccagaacatt
taactgtggg 2280gttggcgctg tccttgtggt atcaaaggag cagacagagc agattctgag
ggatatccag 2340cagcacaagg aagaagcctg ggtgattggc agtgtggttg cacgagctga
aggttcccca 2400cgtgtgaaag tcaagaatct gattgaaagc atgcaaataa atgggtcagt
gttgaagaat 2460ggctccctga caaatcattt ctcttttgaa aaaaaaaagg ccagagtggc
tgtcttaata 2520tctggaacag gatcgaacct gcaagcactt atagacagta ctcgggaacc
aaatagctct 2580gcacaaattg atattgttat ctccaacaaa gccgcagtag ctgggttaga
taaagcggaa 2640agagctggta ttcccactag agtaattaat cataaactgt ataaaaatcg
tgtagaattt 2700gacagtgcaa ttgacctagt ccttgaagag ttctccatag acatagtctg
tcttgcagga 2760ttcatgagaa ttctttctgg cccctttgtc caaaagtgga atggaaaaat
gctcaatatc 2820cacccatcct tgctcccttc ttttaagggt tcaaatgccc atgagcaagc
cctggaaacc 2880ggagtcacag ttactgggtg cactgtacac tttgtagctg aagatgtgga
tgctggacag 2940attattttgc aagaagctgt tcccgtgaag aggggtgata ctgtcgcaac
tctttctgaa 3000agagtaaaat tagcagaaca taaaatattt cctgcagccc ttcagctggt
ggccagtgga 3060actgtacagc ttggagaaaa tggcaagatc tgttgggtta aagaggaatg
aagcctttta 3120attcagaaat ggggccagtt tagaaagaat tatttgctgt ttgcatggtg
gttttttatc 3180atggacttgg cccaaaagaa aaactgctaa aagacaaaaa agacctcacc
cttacttcat 3240ctattttttt aataaataga gactcactaa aaaaaaaaaa aaaaaaaaaa a
3291173291DNAHomo sapiensCDS(79)..(3111) 17accgggcaag
cgggaaccag gtggccaccc ggtgtcggtt tcattttcct ttggaatttc 60tgctttacag
acagaaca atg gca gcc cga gta ctt ata att ggc agt gga 111
Met Ala Ala Arg Val Leu Ile Ile Gly Ser Gly 1
5 10gga agg gaa cat acg ctg gcc tgg aaa ctt
gca cag tct cat cat gtc 159Gly Arg Glu His Thr Leu Ala Trp Lys Leu
Ala Gln Ser His His Val 15 20
25aaa caa gtg ttg gtt gcc cca gga aac gca ggc act gcc tgc tct gaa
207Lys Gln Val Leu Val Ala Pro Gly Asn Ala Gly Thr Ala Cys Ser Glu
30 35 40aag att tca aat acc gcc atc tca
atc agt gac cac act gcc ctt gct 255Lys Ile Ser Asn Thr Ala Ile Ser
Ile Ser Asp His Thr Ala Leu Ala 45 50
55caa ttc tgc aaa gag aag aaa att gaa ttt gta gtt gtt gga cca gaa
303Gln Phe Cys Lys Glu Lys Lys Ile Glu Phe Val Val Val Gly Pro Glu60
65 70 75gca cct ctg gct gct
ggg att gtt ggg aac ctg agg tct gca gga gtg 351Ala Pro Leu Ala Ala
Gly Ile Val Gly Asn Leu Arg Ser Ala Gly Val 80
85 90caa tgc ttt ggc cca aca gca gaa gcg gct cag
tta gag tcc agc aaa 399Gln Cys Phe Gly Pro Thr Ala Glu Ala Ala Gln
Leu Glu Ser Ser Lys 95 100
105agg ttt gcc aaa gag ttt atg gac aga cat gga atc cca acc gca caa
447Arg Phe Ala Lys Glu Phe Met Asp Arg His Gly Ile Pro Thr Ala Gln
110 115 120tgg aag gct ttc acc aaa cct
gaa gaa gcc tgc agc ttc att ttg agt 495Trp Lys Ala Phe Thr Lys Pro
Glu Glu Ala Cys Ser Phe Ile Leu Ser 125 130
135gca gac ttc cct gct ttg gtt gtg aag gcc agt ggt ctt gca gct gga
543Ala Asp Phe Pro Ala Leu Val Val Lys Ala Ser Gly Leu Ala Ala Gly140
145 150 155aaa ggg gtg att
gtt gca aag agc aaa gaa gag gcc tgc aaa gct gta 591Lys Gly Val Ile
Val Ala Lys Ser Lys Glu Glu Ala Cys Lys Ala Val 160
165 170caa gag atc atg cag gag aaa gcc ttt ggg
gca gct gga gaa aca att 639Gln Glu Ile Met Gln Glu Lys Ala Phe Gly
Ala Ala Gly Glu Thr Ile 175 180
185gtc att gaa gaa ctt ctt gac gga gaa gag gtg tcg tgt ctg tgt ttc
687Val Ile Glu Glu Leu Leu Asp Gly Glu Glu Val Ser Cys Leu Cys Phe
190 195 200act gat ggc aag act gtg gcc
ccc atg ccc cca gca cag gac cat aag 735Thr Asp Gly Lys Thr Val Ala
Pro Met Pro Pro Ala Gln Asp His Lys 205 210
215cga tta ctg gag gga gat ggt ggc cct aac aca ggg gga atg gga gcc
783Arg Leu Leu Glu Gly Asp Gly Gly Pro Asn Thr Gly Gly Met Gly Ala220
225 230 235tat tgt cca gcc
cct cag gtt tct aat gat cta tta cta aaa att aaa 831Tyr Cys Pro Ala
Pro Gln Val Ser Asn Asp Leu Leu Leu Lys Ile Lys 240
245 250gat act gtt ctt cag agg aca gtg gat ggc
atg cag caa gag ggt act 879Asp Thr Val Leu Gln Arg Thr Val Asp Gly
Met Gln Gln Glu Gly Thr 255 260
265cca tat aca ggt att ctc tat gct gga ata atg ctg acc aag aat ggc
927Pro Tyr Thr Gly Ile Leu Tyr Ala Gly Ile Met Leu Thr Lys Asn Gly
270 275 280cca aaa gtt cta gag ttt aat
tgc cgt ttt ggt gat cca gag tgc caa 975Pro Lys Val Leu Glu Phe Asn
Cys Arg Phe Gly Asp Pro Glu Cys Gln 285 290
295gta atc ctc cca ctt ctt aaa agt gat ctt tat gaa gtg att cag tcc
1023Val Ile Leu Pro Leu Leu Lys Ser Asp Leu Tyr Glu Val Ile Gln Ser300
305 310 315acc tta gat gga
ctg ctc tgc aca tct ctg cct gtt tgg cta gaa aac 1071Thr Leu Asp Gly
Leu Leu Cys Thr Ser Leu Pro Val Trp Leu Glu Asn 320
325 330cac acc gcc cta act gtt gtc atg gca agt
aaa ggt tat cct gga gac 1119His Thr Ala Leu Thr Val Val Met Ala Ser
Lys Gly Tyr Pro Gly Asp 335 340
345tac acc aag ggt gta gag ata aca ggg ttt cct gag gct caa gct cta
1167Tyr Thr Lys Gly Val Glu Ile Thr Gly Phe Pro Glu Ala Gln Ala Leu
350 355 360gga ctg gag gtg ttc cat gca
ggc act gcc ctc aaa aat ggc aaa gta 1215Gly Leu Glu Val Phe His Ala
Gly Thr Ala Leu Lys Asn Gly Lys Val 365 370
375gta act cat ggg ggt aga gtt ctt gca gtc aca gcc atc cgg gaa aat
1263Val Thr His Gly Gly Arg Val Leu Ala Val Thr Ala Ile Arg Glu Asn380
385 390 395ctc ata tca gcc
ctt gag gaa gcc aag aaa gga cta gct gct ata aag 1311Leu Ile Ser Ala
Leu Glu Glu Ala Lys Lys Gly Leu Ala Ala Ile Lys 400
405 410ttt gag gga gca att tat agg aaa gac gtc
ggc ttt cgt gcc ata gct 1359Phe Glu Gly Ala Ile Tyr Arg Lys Asp Val
Gly Phe Arg Ala Ile Ala 415 420
425ttc ctc cag cag ccc agg agt ttg act tac aag gaa tct gga gta gat
1407Phe Leu Gln Gln Pro Arg Ser Leu Thr Tyr Lys Glu Ser Gly Val Asp
430 435 440atc gca gct gga aat atg ctg
gtc aag aaa att cag cct tta gca aaa 1455Ile Ala Ala Gly Asn Met Leu
Val Lys Lys Ile Gln Pro Leu Ala Lys 445 450
455gcc act tcc aga tca ggc tgt aaa gtt gat ctt gga ggt ttt gct ggt
1503Ala Thr Ser Arg Ser Gly Cys Lys Val Asp Leu Gly Gly Phe Ala Gly460
465 470 475ctt ttt gat tta
aaa gca gct ggt ttc aaa gat ccc ctt ctg gcc tct 1551Leu Phe Asp Leu
Lys Ala Ala Gly Phe Lys Asp Pro Leu Leu Ala Ser 480
485 490gga aca gat ggc gtt gga act aaa cta aag
att gcc cag cta tgc aat 1599Gly Thr Asp Gly Val Gly Thr Lys Leu Lys
Ile Ala Gln Leu Cys Asn 495 500
505aaa cat gat acc att ggt caa gat ttg gta gca atg tgt gtt aat gat
1647Lys His Asp Thr Ile Gly Gln Asp Leu Val Ala Met Cys Val Asn Asp
510 515 520att ctg gca caa gga gca gag
ccc ctc ttc ttc ctt gat tac ttt tcc 1695Ile Leu Ala Gln Gly Ala Glu
Pro Leu Phe Phe Leu Asp Tyr Phe Ser 525 530
535tgt gga aaa ctt gac ctc agt gta act gaa gct gtt gtt gct gga att
1743Cys Gly Lys Leu Asp Leu Ser Val Thr Glu Ala Val Val Ala Gly Ile540
545 550 555gct aaa gct tgt
gga aaa gct gga tgt gct ctc ctt gga ggt gaa aca 1791Ala Lys Ala Cys
Gly Lys Ala Gly Cys Ala Leu Leu Gly Gly Glu Thr 560
565 570gca gaa atg cct gac atg tat ccc cct gga
gag tat gac cta gct ggg 1839Ala Glu Met Pro Asp Met Tyr Pro Pro Gly
Glu Tyr Asp Leu Ala Gly 575 580
585ttt gcc gtt ggt gcc atg gag cga gat cag aaa ctc cct cac ctg gaa
1887Phe Ala Val Gly Ala Met Glu Arg Asp Gln Lys Leu Pro His Leu Glu
590 595 600aga atc act gag ggt gat gtt
gtt gtt gga ata gct tca tct ggt ctt 1935Arg Ile Thr Glu Gly Asp Val
Val Val Gly Ile Ala Ser Ser Gly Leu 605 610
615cat agc aat gga ttt agc ctt gtg agg aaa atc gtt gca aaa tct tcc
1983His Ser Asn Gly Phe Ser Leu Val Arg Lys Ile Val Ala Lys Ser Ser620
625 630 635ctc cag tac tcc
tct cca gca cct gat ggt tgt ggt gac cag act tta 2031Leu Gln Tyr Ser
Ser Pro Ala Pro Asp Gly Cys Gly Asp Gln Thr Leu 640
645 650ggg gac tta ctt ctc acg cct acc aga atc
tac agc cat tca ctg tta 2079Gly Asp Leu Leu Leu Thr Pro Thr Arg Ile
Tyr Ser His Ser Leu Leu 655 660
665cct gtc cta cgt tca gga cat gtc aaa gcc ttt gcc cat att act ggt
2127Pro Val Leu Arg Ser Gly His Val Lys Ala Phe Ala His Ile Thr Gly
670 675 680gga gga tta cta gag aac atc
ccc aga gtc ctc cct gag aaa ctt ggg 2175Gly Gly Leu Leu Glu Asn Ile
Pro Arg Val Leu Pro Glu Lys Leu Gly 685 690
695gta gat tta gat gcc cag acc tgg agg atc ccc agg gtt ttc tca tgg
2223Val Asp Leu Asp Ala Gln Thr Trp Arg Ile Pro Arg Val Phe Ser Trp700
705 710 715ttg cag cag gaa
gga cac ctc tct gag gaa gag atg gcc aga aca ttt 2271Leu Gln Gln Glu
Gly His Leu Ser Glu Glu Glu Met Ala Arg Thr Phe 720
725 730aac tgt ggg gtt ggc gct gtc ctt gtg gta
tca aag gag cag aca gag 2319Asn Cys Gly Val Gly Ala Val Leu Val Val
Ser Lys Glu Gln Thr Glu 735 740
745cag att ctg agg gat atc cag cag cac aag gaa gaa gcc tgg gtg att
2367Gln Ile Leu Arg Asp Ile Gln Gln His Lys Glu Glu Ala Trp Val Ile
750 755 760ggc agt gtg gtt gca cga gct
gaa ggt tcc cca cgt gtg aaa gtc aag 2415Gly Ser Val Val Ala Arg Ala
Glu Gly Ser Pro Arg Val Lys Val Lys 765 770
775aat ctg att gaa agc atg caa ata aat ggg tca gtg ttg aag aat ggc
2463Asn Leu Ile Glu Ser Met Gln Ile Asn Gly Ser Val Leu Lys Asn Gly780
785 790 795tcc ctg aca aat
cat ttc tct ttt gaa aaa aaa aag gcc aga gtg gct 2511Ser Leu Thr Asn
His Phe Ser Phe Glu Lys Lys Lys Ala Arg Val Ala 800
805 810gtc tta ata tct gga aca gga tcg aac ctg
caa gca ctt ata gac agt 2559Val Leu Ile Ser Gly Thr Gly Ser Asn Leu
Gln Ala Leu Ile Asp Ser 815 820
825act cgg gaa cca aat agc tct gca caa att gat att gtt atc tcc aac
2607Thr Arg Glu Pro Asn Ser Ser Ala Gln Ile Asp Ile Val Ile Ser Asn
830 835 840aaa gcc gca gta gct ggg tta
gat aaa gcg gaa aga gct ggt att ccc 2655Lys Ala Ala Val Ala Gly Leu
Asp Lys Ala Glu Arg Ala Gly Ile Pro 845 850
855act aga gta att aat cat aaa ctg tat aaa aat cgt gta gaa ttt gac
2703Thr Arg Val Ile Asn His Lys Leu Tyr Lys Asn Arg Val Glu Phe Asp860
865 870 875agt gca att gac
cta gtc ctt gaa gag ttc tcc ata gac ata gtc tgt 2751Ser Ala Ile Asp
Leu Val Leu Glu Glu Phe Ser Ile Asp Ile Val Cys 880
885 890ctt gca gga ttc atg aga att ctt tct ggc
ccc ttt gtc caa aag tgg 2799Leu Ala Gly Phe Met Arg Ile Leu Ser Gly
Pro Phe Val Gln Lys Trp 895 900
905aat gga aaa atg ctc aat atc cac cca tcc ttg ctc cct tct ttt aag
2847Asn Gly Lys Met Leu Asn Ile His Pro Ser Leu Leu Pro Ser Phe Lys
910 915 920ggt tca aat gcc cat gag caa
gcc ctg gaa acc gga gtc aca gtt act 2895Gly Ser Asn Ala His Glu Gln
Ala Leu Glu Thr Gly Val Thr Val Thr 925 930
935ggg tgc act gta cac ttt gta gct gaa gat gtg gat gct gga cag att
2943Gly Cys Thr Val His Phe Val Ala Glu Asp Val Asp Ala Gly Gln Ile940
945 950 955att ttg caa gaa
gct gtt ccc gtg aag agg ggt gat act gtc gca act 2991Ile Leu Gln Glu
Ala Val Pro Val Lys Arg Gly Asp Thr Val Ala Thr 960
965 970ctt tct gaa aga gta aaa tta gca gaa cat
aaa ata ttt cct gca gcc 3039Leu Ser Glu Arg Val Lys Leu Ala Glu His
Lys Ile Phe Pro Ala Ala 975 980
985ctt cag ctg gtg gcc agt gga act gta cag ctt gga gaa aat ggc aag
3087Leu Gln Leu Val Ala Ser Gly Thr Val Gln Leu Gly Glu Asn Gly Lys
990 995 1000atc tgt tgg gtt aaa gag
gaa tga agccttttaa ttcagaaatg 3131Ile Cys Trp Val Lys Glu
Glu 1005 1010gggccagttt agaaagaatt atttgctgtt
tgcatggtgg ttttttatca tggacttggc 3191ccaaaagaaa aactgctaaa agacaaaaaa
gacctcaccc ttacttcatc tattttttta 3251ataaatagag actcactaaa aaaaaaaaaa
aaaaaaaaaa 3291181010PRTHomo sapiens 18Met Ala
Ala Arg Val Leu Ile Ile Gly Ser Gly Gly Arg Glu His Thr1 5
10 15Leu Ala Trp Lys Leu Ala Gln Ser
His His Val Lys Gln Val Leu Val 20 25
30Ala Pro Gly Asn Ala Gly Thr Ala Cys Ser Glu Lys Ile Ser Asn
Thr 35 40 45Ala Ile Ser Ile Ser
Asp His Thr Ala Leu Ala Gln Phe Cys Lys Glu 50 55
60Lys Lys Ile Glu Phe Val Val Val Gly Pro Glu Ala Pro Leu
Ala Ala65 70 75 80Gly
Ile Val Gly Asn Leu Arg Ser Ala Gly Val Gln Cys Phe Gly Pro
85 90 95Thr Ala Glu Ala Ala Gln Leu
Glu Ser Ser Lys Arg Phe Ala Lys Glu 100 105
110Phe Met Asp Arg His Gly Ile Pro Thr Ala Gln Trp Lys Ala
Phe Thr 115 120 125Lys Pro Glu Glu
Ala Cys Ser Phe Ile Leu Ser Ala Asp Phe Pro Ala 130
135 140Leu Val Val Lys Ala Ser Gly Leu Ala Ala Gly Lys
Gly Val Ile Val145 150 155
160Ala Lys Ser Lys Glu Glu Ala Cys Lys Ala Val Gln Glu Ile Met Gln
165 170 175Glu Lys Ala Phe Gly
Ala Ala Gly Glu Thr Ile Val Ile Glu Glu Leu 180
185 190Leu Asp Gly Glu Glu Val Ser Cys Leu Cys Phe Thr
Asp Gly Lys Thr 195 200 205Val Ala
Pro Met Pro Pro Ala Gln Asp His Lys Arg Leu Leu Glu Gly 210
215 220Asp Gly Gly Pro Asn Thr Gly Gly Met Gly Ala
Tyr Cys Pro Ala Pro225 230 235
240Gln Val Ser Asn Asp Leu Leu Leu Lys Ile Lys Asp Thr Val Leu Gln
245 250 255Arg Thr Val Asp
Gly Met Gln Gln Glu Gly Thr Pro Tyr Thr Gly Ile 260
265 270Leu Tyr Ala Gly Ile Met Leu Thr Lys Asn Gly
Pro Lys Val Leu Glu 275 280 285Phe
Asn Cys Arg Phe Gly Asp Pro Glu Cys Gln Val Ile Leu Pro Leu 290
295 300Leu Lys Ser Asp Leu Tyr Glu Val Ile Gln
Ser Thr Leu Asp Gly Leu305 310 315
320Leu Cys Thr Ser Leu Pro Val Trp Leu Glu Asn His Thr Ala Leu
Thr 325 330 335Val Val Met
Ala Ser Lys Gly Tyr Pro Gly Asp Tyr Thr Lys Gly Val 340
345 350Glu Ile Thr Gly Phe Pro Glu Ala Gln Ala
Leu Gly Leu Glu Val Phe 355 360
365His Ala Gly Thr Ala Leu Lys Asn Gly Lys Val Val Thr His Gly Gly 370
375 380Arg Val Leu Ala Val Thr Ala Ile
Arg Glu Asn Leu Ile Ser Ala Leu385 390
395 400Glu Glu Ala Lys Lys Gly Leu Ala Ala Ile Lys Phe
Glu Gly Ala Ile 405 410
415Tyr Arg Lys Asp Val Gly Phe Arg Ala Ile Ala Phe Leu Gln Gln Pro
420 425 430Arg Ser Leu Thr Tyr Lys
Glu Ser Gly Val Asp Ile Ala Ala Gly Asn 435 440
445Met Leu Val Lys Lys Ile Gln Pro Leu Ala Lys Ala Thr Ser
Arg Ser 450 455 460Gly Cys Lys Val Asp
Leu Gly Gly Phe Ala Gly Leu Phe Asp Leu Lys465 470
475 480Ala Ala Gly Phe Lys Asp Pro Leu Leu Ala
Ser Gly Thr Asp Gly Val 485 490
495Gly Thr Lys Leu Lys Ile Ala Gln Leu Cys Asn Lys His Asp Thr Ile
500 505 510Gly Gln Asp Leu Val
Ala Met Cys Val Asn Asp Ile Leu Ala Gln Gly 515
520 525Ala Glu Pro Leu Phe Phe Leu Asp Tyr Phe Ser Cys
Gly Lys Leu Asp 530 535 540Leu Ser Val
Thr Glu Ala Val Val Ala Gly Ile Ala Lys Ala Cys Gly545
550 555 560Lys Ala Gly Cys Ala Leu Leu
Gly Gly Glu Thr Ala Glu Met Pro Asp 565
570 575Met Tyr Pro Pro Gly Glu Tyr Asp Leu Ala Gly Phe
Ala Val Gly Ala 580 585 590Met
Glu Arg Asp Gln Lys Leu Pro His Leu Glu Arg Ile Thr Glu Gly 595
600 605Asp Val Val Val Gly Ile Ala Ser Ser
Gly Leu His Ser Asn Gly Phe 610 615
620Ser Leu Val Arg Lys Ile Val Ala Lys Ser Ser Leu Gln Tyr Ser Ser625
630 635 640Pro Ala Pro Asp
Gly Cys Gly Asp Gln Thr Leu Gly Asp Leu Leu Leu 645
650 655Thr Pro Thr Arg Ile Tyr Ser His Ser Leu
Leu Pro Val Leu Arg Ser 660 665
670Gly His Val Lys Ala Phe Ala His Ile Thr Gly Gly Gly Leu Leu Glu
675 680 685Asn Ile Pro Arg Val Leu Pro
Glu Lys Leu Gly Val Asp Leu Asp Ala 690 695
700Gln Thr Trp Arg Ile Pro Arg Val Phe Ser Trp Leu Gln Gln Glu
Gly705 710 715 720His Leu
Ser Glu Glu Glu Met Ala Arg Thr Phe Asn Cys Gly Val Gly
725 730 735Ala Val Leu Val Val Ser Lys
Glu Gln Thr Glu Gln Ile Leu Arg Asp 740 745
750Ile Gln Gln His Lys Glu Glu Ala Trp Val Ile Gly Ser Val
Val Ala 755 760 765Arg Ala Glu Gly
Ser Pro Arg Val Lys Val Lys Asn Leu Ile Glu Ser 770
775 780Met Gln Ile Asn Gly Ser Val Leu Lys Asn Gly Ser
Leu Thr Asn His785 790 795
800Phe Ser Phe Glu Lys Lys Lys Ala Arg Val Ala Val Leu Ile Ser Gly
805 810 815Thr Gly Ser Asn Leu
Gln Ala Leu Ile Asp Ser Thr Arg Glu Pro Asn 820
825 830Ser Ser Ala Gln Ile Asp Ile Val Ile Ser Asn Lys
Ala Ala Val Ala 835 840 845Gly Leu
Asp Lys Ala Glu Arg Ala Gly Ile Pro Thr Arg Val Ile Asn 850
855 860His Lys Leu Tyr Lys Asn Arg Val Glu Phe Asp
Ser Ala Ile Asp Leu865 870 875
880Val Leu Glu Glu Phe Ser Ile Asp Ile Val Cys Leu Ala Gly Phe Met
885 890 895Arg Ile Leu Ser
Gly Pro Phe Val Gln Lys Trp Asn Gly Lys Met Leu 900
905 910Asn Ile His Pro Ser Leu Leu Pro Ser Phe Lys
Gly Ser Asn Ala His 915 920 925Glu
Gln Ala Leu Glu Thr Gly Val Thr Val Thr Gly Cys Thr Val His 930
935 940Phe Val Ala Glu Asp Val Asp Ala Gly Gln
Ile Ile Leu Gln Glu Ala945 950 955
960Val Pro Val Lys Arg Gly Asp Thr Val Ala Thr Leu Ser Glu Arg
Val 965 970 975Lys Leu Ala
Glu His Lys Ile Phe Pro Ala Ala Leu Gln Leu Val Ala 980
985 990Ser Gly Thr Val Gln Leu Gly Glu Asn Gly
Lys Ile Cys Trp Val Lys 995 1000
1005Glu Glu 1010
User Contributions:
Comment about this patent or add new information about this topic: