Patent application title: Method of classifying antibody, method of identifying antigen, method of obtaining antibody or antibody set, method of constructing antibody panel and antibody or antibody set and use of the same
Inventors:
Atsushi Sugioka (Nagoya-Shi, JP)
Mototaka Sugiura (Toyoake-Shi, JP)
Yasushi Akahori (Nagoya-Shi, JP)
Nobuhiro Hayashi (Toyoake-Shi, JP)
Akihiko Takasaki (Nagoya-Shi, JP)
Miwa Morita (Toyoake-Shi, JP)
Gene Kurosawa (Nagoya-Shi, JP)
Mariko Sumitomo (Nagoya-Shi, JP)
Susumu Tsutsumi (Nagoya-Shi, JP)
Keiko Ogawa (Toyoake-Shi, JP)
Kazuki Matsuda (Nagoya-Shi, JP)
Chiho Muramatsu (Toyoake-Shi, JP)
Noriko Satou (Toyoake-Shi, JP)
Masachika Azuma (Nagoya-Shi, JP)
Yoshinori Ukai (Nagoya-Shi, JP)
Kazuhiro Suzuki (Toyota-Shi, JP)
Yoshikazu Kurosawa (Nagoya-Shi, JP)
Miho Tanaka (Nagoya-Shi, JP)
Mamoru Shiraishi (Nagoya-Shi, JP)
Assignees:
Institute for Antibodies Co., Ltd.
IPC8 Class: AC40B3004FI
USPC Class:
506 9
Class name: Combinatorial chemistry technology: method, library, apparatus method of screening a library by measuring the ability to specifically bind a target molecule (e.g., antibody-antigen binding, receptor-ligand binding, etc.)
Publication date: 2009-08-13
Patent application number: 20090203538
Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
Patent application title: Method of classifying antibody, method of identifying antigen, method of obtaining antibody or antibody set, method of constructing antibody panel and antibody or antibody set and use of the same
Inventors:
Atsushi Sugioka
Mototaka Sugiura
Yasushi Akahori
Nobuhiro Hayashi
Akihiko Takasaki
Miwa Morita
Gene Kurosawa
Mariko Sumitomo
Susumu Tsutsumi
Keiko Ogawa
Kazuki Matsuda
Chiho Muramatsu
Noriko Satou
Masachika Azuma
Yoshinori Ukai
Kazuhiro Suzuki
Yoshikazu Kurosawa
Miho Tanaka
Mamoru Shiraishi
Agents:
EDWARDS ANGELL PALMER & DODGE LLP
Assignees:
Institute for Antibodies Co., Ltd.
Origin: BOSTON, MA US
IPC8 Class: AC40B3004FI
USPC Class:
506 9
Abstract:
It is intended to provide a method whereby a plural number of antibodies
against cell surface antigens are quickly classified and to provide a
method whereby antigens of the thus classified antibodies are quickly
identified. Further, it is intended to provide a method of promoting the
utilization of the useful data obtained by the above methods.
Furthermore, it is intended to provide an antibody which is effective in
treating or diagnosing cancer. Namely, a method of classifying antibodies
which comprises: (1) the step of preparing a plural number of antibodies
respectively recognizing cell surface antigens; (2) the step of bringing
each of these antibodies into contact with a cell of the same species;
(3) the step of analyzing each of the cells having been treated in the
step (2) by flow cytometry and thus obtaining data indicating the
reactivity of each antibody with its cell surface antigen; and (4) the
step of comparing the thus obtained data and classifying the individual
antibodies depending on the similarity. A method of identifying antigens
which further comprises: (5) the step of selecting one to several
antibodies from each antibody group formed in the step (4) and
identifying antigens thereof; and (6) on the assumption that antigens of
the antibodies belonging to a single antibody group are the same or
highly related to one another, making relations between the antigens
having been identified in the step (5) and the antibody groups to thereby
identify the antigens. An antibody against HER1, an antibody against
HER2, an antibody against CD46, an antibody against ITGA3, an antibody
against ICAM1, an antibody against ALCAM, an antibody against CD147, an
antibody against C1qR, an antibody against CD44, an antibody against
CD73, an antibody against EpCAM and an antibody against HGFR, each
obtained by using the above methods.Claims:
1. A method of classifying antibody including the following steps:(1)
preparing a plurality of antibodies recognizing cell surface antigen;(2)
bringing each of the antibodies into contact with cells of the same
kinds;(3) analyzing each cell after step (2) by flow cytometry so as to
obtain data showing reactivity between the antibody and the cell surface;
and(4) comparing the obtained data and classifying antibodies based on
the similarity of the data.
2. The method of classifying antibody according to claim 1, wherein the cell surface antigen is an intact cell surface antigen.
3. The classifying method according to claim 1, wherein the cell surface antigen is a cell surface antigen of a cancer cell.
4. The classifying method according to claim 1, wherein the plurality of antibodies recognize cell surface antigen are composed of an assembly of antibodies derived from antibody clones selected as being capable of recognizing a cell surface antigen, from an antibody library.
5. The classifying method according to claim 4, wherein the antibody library is a phage antibody library.
6. The classifying method according to claim 1, wherein the antibody is an antibody to which a label material is bound or fused.
7. The classifying method according to claim 1, wherein the antibody does not include a label material and the method includes a step of labeling the antibody bound to the cell after step (2).
8. The classifying method according to claim 1, wherein the cell is an established cell line.
9. The classifying method according to claim 1, wherein the cell is an established cancer cell line.
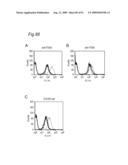
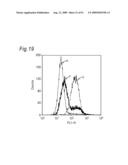
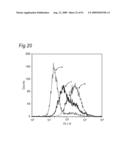
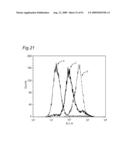
10. The classifying method according to claim 1, wherein the data are shown in a histogram showing a relationship between a binding amount of antibodies and a number of cells, and the similarity of the data is determined by comparing the shapes of the histograms.
11. The classifying method according to claim 1, wherein the data are shown in a histogram showing a relationship between a binding amount of antibodies and a number of cells, and the similarity of the data is determined based on one or more values selected from the group consisting of a median value, a mode, a maximum value, a range, a standard deviation, a kurtosis and a skewness of the histogram.
12. The classifying method according to claim 11, wherein the similarity of the data is determined based on the values of the median value, the mode, and the kurtosis and a skewness of the histogram.
13. The classifying method according to claim 10, wherein the binding amount of antibody is shown by a fluorescence intensity.
14. The classifying method according to claim 1, wherein in step (4), a plurality of antibodies having the identical or high similar data are classified into one antibody group.
15. The classifying method according to claim 1, wherein two or more kinds of cells are prepared and each kind of cell is subjected to steps (2) to (4).
16. The classifying method according to claim 15, wherein a plurality of antibodies having the identical or high similar data with respect to two or more kinds of cells in the cells are classified into one antibody group.
17. The classifying method according to claim 1, wherein an antibody that has been determined to have a low reactivity with respect to the cell surface antigen during classification or after classification is excluded.
18. The classifying method according to claim 1, wherein classification results of antibodies are displayed as a panel.
19. The classifying method according to claim 1, wherein after step (4), the following steps are carried out:(i) associating the classified antibodies to a combination of n pieces of parameters including a first parameter, a second parameter, . . . , and an n-th parameter (wherein, n represents an integer of 2 or more, each parameter has two or more parameter values and the same parameter value is given to two or more antibodies in each parameter);(ii) with respect to each parameter, preparing antibody mixtures of the antibodies having the same parameter value;(iii) examining a reactivity of each of the antibody mixtures with a target antigen by an enzyme linked immunosorbent assay (ELISA) so as to specify the antibody mixture which shows reactivity;(iv) specifying a combination of a parameter name and a parameter value that are common to the antibody group contained in the specified antibody mixture;(v) selecting an antibody corresponding to the combination specified in the step (iv) in terms of all parameters among the antibodies subjected to step (i); and(vi) classifying the selected antibodies into one antibody group.
20. The classifying method according to claim 19, wherein the steps (i) to (v) are repeated several times under the conditions in which the combination of parameters is different in each trial; an antibody in which results of all trials are not contradictory is selected; and the antibody is subjected to the step (vi).
21. The classifying method according to claim 19, further including the following steps between the step (v) and the step (vi);(v-1) newly associating the classified antibodies selected in step (v) with a combination of n pieces of parameters in a same manner as in the step (i);(v-2) with respect to each parameter, preparing the antibody mixture of antibodies having the same parameter value for each parameter;(v-3) examining a reactivity of each of the antibody mixtures with a target antigen by an enzyme linked immunosorbent assay (ELISA) so as to specify the antibody mixture showing the reactivity;(v-4) determining a combination of a parameter name and a parameter value that are common to the antibody group contained in the specified antibody mixture; and(v-5) selecting an antibody having the combination specified in the step (v-4) in terms of all parameters among the antibodies subjected to the step (v-1).
22. The classifying method according to claim 21, wherein the steps (v-1) to (v-4) are repeated twice or more.
23. The classifying method according to claim 19, wherein n is 3.
24. The classifying method according to claim 19, wherein two or more kinds of target antigens are prepared and the steps (iii) to (vi) are carried out by using each target antigen.
25. The classifying method according to claim 19, wherein the target antigen is an antigen selected from the group consisting of HER1, HER2, CD46, ITGA3, ICAM1, ALCAM, CD147, IgSF4, BCAM, C1qR, CD44, CD73, LAR, EpCAM and HGFR.
26. An identifying method of an antigen including the following steps:(1) preparing a plurality of antibodies recognizing cell surface antigen;(2) bringing each of the antibodies into contact with cells of the same kind;(3) analyzing each cell after step (2) by flow cytometry so as to obtain data showing the reactivity between the antibody and the cell surface;(4) comparing the obtained data and classifying antibodies based on the similarity of the data;(5) selecting one or several antibodies from each antibody group formed in the step (4) and identifying an antigen thereof; and(6) associating the antigens identified in the step (5) with an antibody group, based on the estimation that antigens to antibodies belonging to the same antibody group are identical or have high relationship.
27. The identification method according to claim 26, wherein in the step (5), one antibody is selected from each antibody group.
28. The identification method according to claim 26, wherein in the step (5), from the results of a flow cytometry analysis, an antibody that is determined to have a high reactivity with respect to an antigen is selected.
29. The identification method according to claim 26, wherein in the step (5), the identification of an antigen is carried out by one or more methods selected from the group consisting of an immunoprecipitation test, Western blotting, affinity chromatography, proteomics techniques (electrophoresis, mass spectrometry, genome data base retrieve, and analysis by bioinformatics), and an expression analysis of corresponding gene.
30. The identification method according to claim 26, further including a step of examining a reactivity between an antigen identified in the step (5) and an antibody belonging to an antibody group with which the antigen is associated in the step (6) so as to confirm that the estimation is correct.
31. The identification method according to claim 26, wherein an identification result of antigen is displayed as a panel.
32. The identification method according to claim 31, wherein the panel is any of the following (a) to (c):(a) a panel displaying a plurality of antibodies showing identical or high similar data in the flow cytometry analysis in the step (3) as one antibody group in which each antibody group is associated with its antigen;(b) a panel displaying a plurality of antibodies showing identical or high similar data in the flow cytometry analysis in the step (3) as one antibody group in which each antibody group is associated with a cell expressing a cell surface antigen recognized by the each antibody group; and(c) a panel displaying a plurality of antibodies showing identical or high similar data in the flow cytometry analysis in the step (3) as one antibody group in which each antibody group, its antigen and a cell expressing a cell surface antigen recognized by the antibody group are associated with each other.
33. A method of obtaining an antibody having a relationship with respect to a certain disease, the method comprising the following steps:(1) selecting one or two or more of antibody groups from the plurality of antibody groups classified by the classifying method according to claim 1;(2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and(3) selecting an antibody in the antibody group, to which an antibody having a specific reactivity to any of diseases belongs, as a useful antibody.
34. A method of obtaining an antibody having a relationship with respect to a certain disease, the method comprising the following steps:(1) selecting one or two or more of antibody groups from the plurality of antibody groups classified by the classifying method according to claim 19;(2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and(3) selecting an antibody in the antibody group, to which an antibody having a specific reactivity to any of diseases belongs, as a useful antibody.
35. A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:(1) selecting one or two or more of antibody groups from the plurality of antibody groups classified by the classifying method according to claim 1;(2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and(3') selecting a disease to which two or more antibodies show a specific reactivity, then selecting antibodies from the antibody group, to which the antibody having a specific reactivity to the disease belongs, and combining the selected antibodies.
36. A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:(1) selecting two or more antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to claim 1;(2) with respect to two kinds or more diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and(3) selecting antibodies from the antibody group, to which the antibody having a specific reactivity to any of disease belongs, and combining the selected antibodies.
37. A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:(1) selecting two or more antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to claim 1;(2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and(3) selecting an antibody from the antibody group to which the antibody having a specific reactivity to any of diseases belongs, and an antibody belonging to other antibody group whose antigen is common to that of the antibody group, and combining the selected antibodies.
38. A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:(1) selecting two or more antibody groups recognizing the common antigen from the plurality of antibody groups classified by the classifying method according to claim 1;(2) with respect to one kind or two or more kinds of pathologic conditions, examining a reactivity between an antibody in each of the selected antibody groups and a pathologic condition; and(3) connecting information about the reactivity and then combining the antibodies in the antibody groups.
39. A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:(1) selecting one or two or more antibody groups from the plurality of antibody groups classified by the classifying method according to claim 19;(2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and(3') selecting a disease to which two or more antibodies show a specific reactivity, then selecting antibodies from an antibody group which the antibodies showing a specific reactivity to the disease belong to, and combining the selected antibodies.
40. A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:(1) selecting two or more antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to claim 19;(2) with respect to two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease in two or more kinds of diseases; and(3) selecting antibodies from the antibody group to which the antibody having a specific reactivity to any of diseases belong, and combining the selected antibodies.
41. A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:(1) selecting two or more antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to claim 19;(2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and(3) selecting an antibody from the antibody group to which the antibody having a specific reactivity to any of disease belongs, and an antibody belonging to other antibody group whose antigen is common to that of the antibody group, and combining the selected antibodies.
42. A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:(1) selecting two or more antibody groups recognizing the common antigen from the plurality of antibody groups classified by the classifying method according to claim 19;(2) with respect to one kind or two or more kinds of pathologic conditions, examining a reactivity between an antibody in each of the selected antibody groups and a pathologic condition; and(3) associating information about the reactivity and then combining the antibodies in the antibody groups.
43. The obtaining method according to claim 33, wherein the disease is selected from the group consisting of: kidney cancer, hepatic cell carcinoma, gallbladder and liver cancer, alveolar cell carcinoma, lung squamous cell cancer, pulmonary adenocarcinoma, pancreas cancer, adenocarcinoma, and ovarian cancer.
44. The obtaining method according to claim 33, wherein in the step (2), the reactivity is examined by one or more methods selected from the group consisting of an immunostaining procedure, an immunoprecipitation method, a flow cytometry analysis, cell ELISA, an intermolecular interactive analysis between a disease-related molecule (disease causative gene product and the like) and an antibody, and application test to a disease model cell (or animal).
45. An isolated antibody obtained by the method according to claim 33.
46. An antibody set obtained by the method described in claim 35.
47. A production method of a panel displaying a relationship between an antibody and a disease, the method comprising the following steps:(1) selecting one or two or more of antibody groups from the plurality of antibody groups classified by the classifying method according to claim 1;(2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and(3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
48. A production method of a panel displaying a relationship between an antibody and a disease, the method comprising the following steps:(1) selecting two or more of antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to claim 1;(2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and(3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
49. A production method of a panel displaying a relationship between an antibody and a pathologic condition, the method comprising the following steps:(1) selecting two or more of antibody groups recognizing a common antigen from the plurality of antibody groups classified by the classifying method according to claim 1;(2) with respect to one kind or two or more kinds of pathologic condition, examining a reactivity between an antibody in each of the selected antibody groups and a certain pathologic condition of disease; and(3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
50. A production method of a panel displaying a relationship between an antibody and a disease, the method comprising the following steps:(1) selecting one or two or more of antibody groups from the plurality of antibody groups classified by the classifying method according to claim 19;(2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and(3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
51. A production method of a panel displaying a relationship between an antibody and a disease, the method comprising the following steps:(1) selecting two or more of antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to claim 19;(2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and(3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
52. A production method of a panel displaying a relationship between an antibody and a pathologic condition, the method comprising the following steps:(1) selecting two or more of antibody groups recognizing a common antigen from the plurality of antibody groups classified by the classifying method according to claim 19;(2) with respect to one kind or two or more kinds of pathologic condition, examining a reactivity between an antibody in each of the selected antibody groups and a certain pathologic condition of disease; and(3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
53. A panel produced by the method according to claim 47.
54. (canceled)
55. A method of testing a disease in which a cell surface antigen is an indicator, the method comprising the following steps:(1) preparing a cell or a tissue separated from a subject;(2) examining a reactivity between the cell or the tissue and each antibody displayed on the panel according to claim 53; and(3) collating the results in the step (2) with the panel.
56. A method of selecting an optimum treatment method for a certain disease, the method comprising the following steps:(1) preparing a cell or a tissue separated from a subject;(2) examining a reactivity between the cell or the tissue and each antibody displayed on the panel according to claim 53;(3) collating the results in the step (2) with the panel, and(4) selecting an effective antibody according to the results of collating.
57. The method according to claim 56, wherein the effective antibody is an antibody showing a specific reactivity in the step (2) or an antibody equivalent thereto.
58. The method according to claim 56, wherein the certain disease is a disease in which a cell surface antigen selected from the group consisting of HER1, HER2, CD46, ITGA3, ICAM1, ALCAM, CD147, IgSF4, BCAM, C1qR, CD44, CD73, LAR, EpCAM and HGFR is an indicator.
59. The method according to claim 56, wherein the panel displays two or more antibodies selected from the group consisting of 048-006 antibody, 057-091 antibody, 059-152 antibody, 048-040 antibody, 054-101 antibody, 055-147 antibody, 059-173 antibody, 067-149 antibody, 067-176 antibody, 015-126 antibody, 015-044 antibody, 015-102 antibody, 015-136 antibody, 015-143 antibody, 015-209 antibody, 039-016 antibody, 053-216 antibody, 075-024 antibody, 075-110 antibody, 086-032 antibody, 086-035 antibody, 086-036 antibody, 086-061 antibody, 086-138 antibody, 086-182 antibody, 035-224 antibody, 045-011 antibody, 051-144 antibody, 052-053 antibody, 052-073 antibody, 053-049 antibody, 3172-120 antibody, 066-069 antibody, 015-003 antibody, 064-002 antibody, 064-006 antibody, 064-012a antibody, 064-012b antibody, 064-014 antibody, 064-054 antibody, 064-085 antibody, 064-093 antibody, 064-116 antibody, 065-183 antibody, 067-142 antibody, 068-007 antibody, 052-033 antibody, 053-042 antibody, 053-051 antibody, 053-059 antibody, 053-085 antibody, 035-234 antibody, 040-107 antibody, 041-118 antibody, 066-174 antibody, 083-040 antibody, 029-143 antibody, 045-134 antibody, 062-101 antibody, 062-109 antibody, 084-103 antibody, 052-274 antibody, 029-067 antibody, 083-131 antibody, 059-053 antibody, 064-003 antibody, 067-213 antibody, 067-153 antibody, 067-126 antibody, 067-133 antibody, 067-287 antibody, 064-044 antibody, 065-030 antibody, 065-358 antibody, 066-019 antibody, 079-085 antibody, 067-024 antibody and 076-048 antibody.
60. A method of selecting an optimum treatment method of a certain disease, the method comprising the following steps:(1) preparing a panel displaying a reactivity between one or more antibodies selected from the group consisting of 048-006 antibody, 015-126 antibody, 067-133 antibody, 064-044 antibody, 076-048 antibody and 059-053 antibody, and a clinical cancer tissue of one or more diseases selected from the group consisting of squamous carcinoma, adenosquamous carcinoma, alveolar adenocarcinoma, adenocarcinoma, and large cell carcinoma, and a cell or tissue separated from a subject;(2) examining a reactivity between the cell or the tissue and each antibody displayed on the panel;(3) collating the results in the step (2) with the panel, and(4) selecting an effective antibody according to the results of collating.
61. The method according to claim 60, wherein the effective antibody is an antibody showing a specific reactivity in the step (2) or an antibody equivalent thereto.
62. The method according to claim 60, wherein the certain disease is a disease selected from the group consisting of squamous carcinoma, adenosquamous carcinoma, alveolar adenocarcinoma, adenocarcinoma, and large cell carcinoma.
63-85. (canceled)
Description:
CROSS REFERENCE TO RELATED APPLICATIONS
[0001]This application is a continuation-in-part of international application No. PCT/JP2007/063689, filed Jul. 9, 2007, which claims priority to Japanese applications No. 2006-189872, filed Jul. 10, 2007 and No. 2007-058458, filed Mar. 8, 2008. The contents of these three applications are hereby incorporated by reference in their entirety.
FIELD OF THE INVENTION
[0002]The present invention relates to a method of classifying a plurality of antibodies, a method of identifying antigen, a panel displaying characteristics of an antibody, and the like, as well as an antibody related to a disease and a use thereof.
BACKGROUND OF THE INVENTION
[0003]Success of Herceptin to breast cancer (see, non-patent document 1) and Rituxan (non-patent document 2) to malignant lymphoma B shows that an antibody is effective as a therapeutic agent to a cancer. Certain antibodies exhibit an ADCC effect (non-patent document 3) and/or a CDC effect (non-patent document 4) by forming a complex with an antigen molecule existing on the cell membrane and the effects kill a target cell (cell expressing an antigen). The ADCC effect or the CDC effect may cause apoptosis. Such an effect of an antibody is specific to an antigen. That is to say, an antibody acts on cells expressing an antigen which the antibody recognizes regardless of whether the cells are cancer cells or normal cells. Therefore, the success in development of antibody therapeutic agents to cancers is dependent on discovery of antigens expressing in a cancer-specific manner and recognized by an antibody so as to cause the ADCC effect or the CDC effect. An antibody against to such an antigen is a promising candidate of a therapeutic agent capable of reliably killing target cancer cells while minimizing the influence (side effect) on normal cells.
[0004]In antibody drug development, it is essential to obtain antibodies that recognize "intact state" target cancer antigens existing on the surface of a cell membrane. However, since the target cancer antigen is membrane protein, it has been difficult to obtain an antibody against even known cancer antigen. In order to solve these problems, present inventors have produced a huge human antibody library including as many as 100 billion independent clones and established a comprehensive acquisition method for antibodies to proteins (cell surface antigens) existing on the surface of the cell membrane of cancer cells and tissues by using the library (patent documents 1 to 3).
[Patent document 1] WO01/062907[Patent document 2] WO2001/096401[Patent document 3] Japanese Patent Unexamined Publication No. 2005-185281[Non-patent document 1] Mass R, et al.: The Concordance Between the Clinical Trials Assay (CTA) and Fluorescence in Situ Hybridization (FISH) in the Herceptin Pivotal Trials.: Proc Am Soci Clin Oncol 19, 75a, 2000[Non-patent document 2] Berinstein N L, Grillo-Lopez A J, White C A, Bence-Bruckler I, Maloney D, Czuczman M, et al. Association of serum Rituximab (IDEC-C2B8) concentration and anti-tumor response in the treatment of recurrent low-grade or follicular non-Hodgkin's lymphoma. Annals of Oncology 1998, 9:995-1001.[Non-patent document 3] Bruggemann M., Williams G. T., Bindon C. I., Clark M. R., Walker M. R., Jefferis R., Waldmann H., Neuberger M. S. (1987). Comparison of the effector functions of human immunoglobulins using a matched set of chimeric antibodies. J. Exp. Med., 166, 1351-1361.[Non-patent document 4] Loos M. (1982). The classical complement pathway: mechanism of activation of the first component by antigen-antibody complexes. Prog. Allergy, 30, 135-192. Mol. Immunol. 1982 May; 19 (5): 651-7.
SUMMARY OF THE INVENTION
[0005]Currently, the present inventors can comprehensively obtain antibodies to cell surface antigens. As the next step, it is necessary to identify an antibody to each antibody and to screen useful antibodies. However, it will take a much labor and time and considerably high cost to individually identify an antigen for the comprehensively obtained antibodies.
[0006]Furthermore, the comprehensively obtained antibodies may include unnecessary antibodies from the viewpoint that they do not have sufficient affinity and reactivity, or they have substantially the same as the other antibodies. Therefore, method for efficiently screening useful antibodies has been demanded.
[0007]On the other hand, the comprehensively obtained antibodies may include antibodies such as candidates of diagnostic agents and therapeutic agents, which are extremely important from the medical viewpoint.
[0008]Under such circumstances, the present invention aims at the effective use of comprehensively obtained antibodies to cell surface antigens in medical fields and research fields, and has an object to provide a useful method therefor. That is to say, the present invention has an object to provide a method of classifying a plurality of antibodies to cell surface antigens rapidly. Also, the present invention has another object to provide a method of rapidly identifying an antigen for the antibody. Furthermore, the present invention has a further object to provide a method of promoting to use useful information obtained by such methods. The present invention has a yet further object to provide an antibody effective for treatment and diagnosis of cancers.
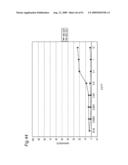
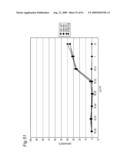
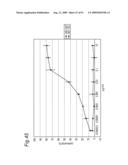
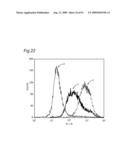
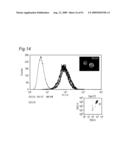
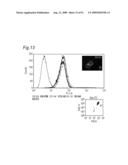
[0009]In view of the above-mentioned objects, the present inventors carry out an analysis of an antibody by the following approach: preparing cell lines that are expected to express cell surface antigens for the obtained antibodies; allowing each antibody to react with the cell lines; and carrying out the flow cytometry analysis.
[0010]The present inventors focus on the histogram of the results of the flow cytometry analysis and classify the antibodies based on the similarity so as to obtain a plurality of antibodies groups. Then, it is confirmed that antigens to antibodies belonging to the same antibody group are common. This fact means that it is possible to determine antigens for all antibodies by selecting the respective antibody in each antibody group and identifying the antigen of the representative antibody. Thus, the present inventors have succeeded in finding a method for identifying antigens comprehensively and rapidly. On the other hand, the present inventors carry out classification of antibodies and identification of an antigen according to the above-mentioned technique and consider the reactivity between each antibody group and clinical samples so as to search for clinically applicable antibodies. As a result, the present inventors have succeeded in finding a novel antibody specific to certain kinds of cancers. Furthermore, they have reached the findings that information obtained by using a clinical sample (relationship between the antibody and disease) is extremely useful for establishing methods for diagnosis and treatment.
[0011]The present invention provides, for example, a method of classifying antibody, and the like, mentioned below based on the above-mentioned results and findings.
<Method of Classifying Antibody>
[0012][1] A method of classifying antibody including the following steps:
[0013](1) preparing a plurality of antibodies recognizing cell surface antigen;
[0014](2) bringing each of the antibodies into contact with cells of the same kinds;
[0015](3) analyzing each cell after step (2) by flow cytometry so as to obtain data showing reactivity between the antibody and the cell surface; and
[0016](4) comparing the obtained data and classifying antibodies based on the similarity of the data.
[2] The method of classifying antibody according to [1], wherein the cell surface antigen is an intact cell surface antigen.[3] The classifying method according to [1] or [2], wherein the cell surface antigen is a cell surface antigen of a cancer cell.[4] The classifying method according to [1], wherein the plurality of antibodies recognize cell surface antigen are composed of an assembly of antibodies derived from antibody clones selected as being capable of recognizing a cell surface antigen, from an antibody library.[5] The classifying method according to [4], wherein the antibody library is a phage antibody library.[6] The classifying method according to [1], wherein the antibody is an antibody to which a label material is bound or fused.[7] The classifying method according to [1], wherein the antibody does not include a label material and the method includes a step of labeling the antibody bound to the cell after step (2).[8] The classifying method according to [1], wherein the cell is an established cell line.[9] The classifying method according to [1], wherein the cell is an established cancer cell line.[10] The classifying method according to [1], wherein the data are shown in a histogram showing a relationship between a binding amount of antibodies and a number of cells, and the similarity of the data is determined by comparing the shapes of the histograms.[11] The classifying method according to [1], wherein the data are shown in a histogram showing a relationship between a binding amount of antibodies and a number of cells, and the similarity of the data is determined based on one or more values selected from the group consisting of a median value, a mode, a maximum value, a range, a standard deviation, a kurtosis and a skewness of the histogram.[12] The classifying method according to [11], wherein the similarity of the data is determined based on the values of the median value, the mode, and the kurtosis and a skewness of the histogram.[13] The classifying method according to [10] or [11], wherein the binding amount of antibody is shown by a fluorescence intensity.[14] The classifying method according to [1], wherein in step (4), a plurality of antibodies having the identical or high similar data are classified into one antibody group.[15] The classifying method according to [1], wherein two or more kinds of cells are prepared and each kind of cell is subjected to steps (2) to (4).[16] The classifying method according to [15], wherein a plurality of antibodies having the identical or high similar data with respect to two or more kinds of cells in the cells are classified into one antibody group.[17] The classifying method according to [1], wherein an antibody that has been determined to have a low reactivity with respect to the cell surface antigen during classification or after classification is excluded.[18] The classifying method according to [1], wherein classification results of antibodies are displayed as a panel.[19] The classifying method according to any of [α] to [18], wherein after step (4), the following steps are carried out:
[0017](i) associating the classified antibodies to a combination of n pieces of parameters including a first parameter, a second parameter, . . . , and an n-th parameter (wherein, n represents an integer of 2 or more, each parameter has two or more parameter values and the same parameter value is given to two or more antibodies in each parameter);
[0018](ii) with respect to each parameter, preparing antibody mixtures of the antibodies having the same parameter value;
[0019](iii) examining a reactivity of each of the antibody mixtures with a target antigen by an enzyme linked immunosorbent assay (ELISA) so as to specify the antibody mixture which shows reactivity;
[0020](iv) specifying a combination of a parameter name and a parameter value that are common to the antibody group contained in the specified antibody mixture;
[0021](v) selecting an antibody corresponding to the combination specified in the step (iv) in terms of all parameters among the antibodies subjected to step (i); and
[0022](vi) classifying the selected antibodies into one antibody group.
[20] The classifying method according to [19], wherein the steps (i) to (v) are repeated several times under the conditions in which the combination of parameters is different in each trial; an antibody in which results of all trials are not contradictory is selected; and the antibody is subjected to the step (vi).[21] The classifying method according to [19], further including the following steps between the step (v) and the step (vi);
[0023](v-1) newly associating the classified antibodies selected in step (v) with a combination of n pieces of parameters in a same manner as in the step (i);
[0024](v-2) with respect to each parameter, preparing the antibody mixture of antibodies having the same parameter value for each parameter;
[0025](v-3) examining a reactivity of each of the antibody mixtures with a target antigen by an enzyme linked immunosorbent assay (ELISA) so as to specify the antibody mixture showing the reactivity;
[0026](v-4) determining a combination of a parameter name and a parameter value that are common to the antibody group contained in the specified antibody mixture; and
[0027](v-5) selecting an antibody having the combination specified in the step (v-4) in terms of all parameters among the antibodies subjected to the step (v-1).
[22] The classifying method according to [21], wherein the steps (v-1) to (v-4) are repeated twice or more.[23] The classifying method according to any of [19] to [22], wherein n is 3.[24] The classifying method according to any of [19] to [23], wherein two or more kinds of target antigens are prepared and the steps (iii) to (vi) are carried out by using each target antigen.[25] The classifying method according to any of [19] to [24], wherein the target antigen is an antigen selected from the group consisting of HER1, HER2, CD46, ITGA3, ICAM1, ALCAM, CD147, IgSF4, BCAM, C1qR, CD44, CD73, LAR, EpCAM and HGFR.
<Identifying Method of Antigen>
[0028][26] An identifying method of an antigen including the following steps:
[0029](1) preparing a plurality of antibodies recognizing cell surface antigen;
[0030](2) bringing each of the antibodies into contact with cells of the same kind;
[0031](3) analyzing each cell after step (2) by flow cytometry so as to obtain data showing the reactivity between the antibody and the cell surface;
[0032](4) comparing the obtained data and classifying antibodies based on the similarity of the data;
[0033](5) selecting one or several antibodies from each antibody group formed in the step (4) and identifying an antigen thereof; and
[0034](6) associating the antigens identified in the step (5) with an antibody group, based on the estimation that antigens to antibodies belonging to the same antibody group are identical or have high relationship, and.
[27] The identification method according to [26], wherein in the step (5), one antibody is selected from each antibody group.[28] The identification method according to [26], wherein in the step (5), from the results of a flow cytometry analysis, an antibody that is determined to have a high reactivity with respect to an antigen is selected.[29] The identification method according to [26], wherein in the step (5), the identification of an antigen is carried out by one or more methods selected from the group consisting of an immunoprecipitation test, Western blotting, affinity chromatography, proteomics techniques (electrophoresis, mass spectrometry, genome data base retrieve, and analysis by bioinformatics), and an expression analysis of corresponding gene.[30] The identification method according to [26], further including a step of examining a reactivity between an antigen identified in the step (5) and an antibody belonging to an antibody group with which the antigen is associated in the step (6) so as to confirm that the estimation is correct.[31] The identification method according to [26], wherein an identification result of antigen is displayed as a panel.[32] The identification method according to [31], wherein the panel is any of the following (a) to (c):
[0035](a) a panel displaying a plurality of antibodies showing identical or high similar data in the flow cytometry analysis in the step (3) as one antibody group in which each antibody group is associated with its antigen;
[0036](b) a panel displaying a plurality of antibodies showing identical or high similar data in the flow cytometry analysis in the step (3) as one antibody group in which each antibody group is associated with a cell expressing a cell surface antigen recognized by the each antibody group; and
[0037](c) a panel displaying a plurality of antibodies showing identical or high similar data in the flow cytometry analysis in the step (3) as one antibody group in which each antibody group, its antigen and a cell expressing a cell surface antigen recognized by the antibody group are associated with each other.
<Method of Obtaining Antibody or Antibody Set, Antibody or Antibody Set to be Obtained>
[0038][33] A method of obtaining an antibody having a relationship with respect to a certain disease, the method comprising the following steps:
[0039](1) selecting one or two or more of antibody groups from the plurality of antibody groups classified by the classifying method according to [1];
[0040](2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0041](3) selecting an antibody in the antibody group, to which an antibody having a specific reactivity to any of diseases belongs, as a useful antibody.
[34] A method of obtaining an antibody having a relationship with respect to a certain disease, the method comprising the following steps:
[0042](1) selecting one or two or more of antibody groups from the plurality of antibody groups classified by the classifying method according to [19];
[0043](2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0044](3) selecting an antibody in the antibody group, to which an antibody having a specific reactivity to any of diseases belongs, as a useful antibody.
[35] A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:
[0045](1) selecting one or two or more of antibody groups from the plurality of antibody groups classified by the classifying method according to [1];
[0046](2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0047](3') selecting a disease to which two or more antibodies show a specific reactivity, then selecting antibodies from the antibody group, to which the antibody having a specific reactivity to the disease belongs, and combining the selected antibodies.
[36] A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:
[0048](1) selecting two or more antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to [1];
[0049](2) with respect to two kinds or more diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0050](3) selecting antibodies from the antibody group, to which the antibody having a specific reactivity to any of disease belongs, and combining the selected antibodies.
[37] A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:
[0051](1) selecting two or more antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to [1];
[0052](2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0053](3) selecting an antibody from the antibody group to which the antibody having a specific reactivity to any of diseases belongs, and an antibody belonging to other antibody group whose antigen is common to that of the antibody group, and combining the selected antibodies.
[38] A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:
[0054](1) selecting two or more antibody groups recognizing the common antigen from the plurality of antibody groups classified by the classifying method according to [1];
[0055](2) with respect to one kind or two or more kinds of pathologic conditions, examining a reactivity between an antibody in each of the selected antibody groups and a pathologic condition; and
[0056](3) connecting information about the reactivity and then combining the antibodies in the antibody groups.
[39] A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:
[0057](1) selecting one or two or more antibody groups from the plurality of antibody groups classified by the classifying method according to [19];
[0058](2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0059](3') selecting a disease to which two or more antibodies show a specific reactivity, then selecting antibodies from an antibody group which the antibodies showing a specific reactivity to the disease belong to, and combining the selected antibodies.
[40] A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:
[0060](1) selecting two or more antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to [19];
[0061](2) with respect to two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease in two or more kinds of diseases; and
[0062](3) selecting antibodies from the antibody group to which the antibody having a specific reactivity to any of diseases belong, and combining the selected antibodies.
[41] A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:
[0063](1) selecting two or more antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to [19];
[0064](2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0065](3) selecting an antibody from the antibody group to which the antibody having a specific reactivity to any of disease belongs, and an antibody belonging to other antibody group whose antigen is common to that of the antibody group, and combining the selected antibodies.
[42] A method of obtaining an antibody set having a relationship with respect to a certain disease, the method comprising the following steps:
[0066](1) selecting two or more antibody groups recognizing the common antigen from the plurality of antibody groups classified by the classifying method according to [19];
[0067](2) with respect to one kind or two or more kinds of pathologic conditions, examining a reactivity between an antibody in each of the selected antibody groups and a pathologic condition; and
[0068](3) associating information about the reactivity and then combining the antibodies in the antibody groups.
[43] The obtaining method according any of [33] to [42], wherein the disease is selected from the group consisting of kidney cancer, hepatic cell carcinoma, gallbladder and liver cancer, alveolar cell carcinoma, lung squamous cell cancer, pulmonary adenocarcinoma, pancreas cancer, adenocarcinoma, and ovarian cancer.[44] The obtaining method according any of [33] to [42], wherein in the step (2), the reactivity is examined by one or more methods selected from the group consisting of an immunostaining procedure, an immunoprecipitation method, a flow cytometry analysis, cell ELISA, an intermolecular interactive analysis between a disease-related molecule (disease causative gene product and the like) and an antibody, and application test to a disease model cell (or animal).[45] An isolated antibody obtained by the method according to [33] or [34].[46] An antibody set obtained by the method described in any of [35] to [42].
<Production Method of Panel, Panel, and Combination of Antibody or Antibody Set and Panel>
[0069][47] A production method of a panel displaying a relationship between an antibody and a disease, the method comprising the following steps:
[0070](1) selecting one or two or more of antibody groups from the plurality of antibody groups classified by the classifying method according to [1];
[0071](2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0072](3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
[48] A production method of a panel displaying a relationship between an antibody and a disease, the method comprising the following steps:
[0073](1) selecting two or more of antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to [1];
[0074](2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0075](3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
[49] A production method of a panel displaying a relationship between an antibody and a pathologic condition, the method comprising the following steps:
[0076](1) selecting two or more of antibody groups recognizing a common antigen from the plurality of antibody groups classified by the classifying method according to [1];
[0077](2) with respect to one kind or two or more kinds of pathologic condition, examining a reactivity between an antibody in each of the selected antibody groups and a certain pathologic condition of disease; and
[0078](3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
[50] A production method of a panel displaying a relationship between an antibody and a disease, the method comprising the following steps:
[0079](1) selecting one or two or more of antibody groups from the plurality of antibody groups classified by the classifying method according to [19];
[0080](2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0081](3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
[51] A production method of a panel displaying a relationship between an antibody and a disease, the method comprising the following steps:
[0082](1) selecting two or more of antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to [19];
[0083](2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0084](3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
[52] A production method of a panel displaying a relationship between an antibody and a pathologic condition, the method comprising the following steps:
[0085](1) selecting two or more of antibody groups recognizing a common antigen from the plurality of antibody groups classified by the classifying method according to [19];
[0086](2) with respect to one kind or two or more kinds of pathologic condition, examining a reactivity between an antibody in each of the selected antibody groups and a certain pathologic condition of disease; and
[0087](3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
[53] A panel produced by the method according to any of [47] to [52].[54] A combination of an antibody or an antibody set and a panel selected from the group consisting of the following (a) to (d);
[0088](a) a combination of the isolated antibody obtained by the method according to [33] and the panel produced by the method according to [47];
[0089](b) a combination of the antibody set obtained by the method according to [35] and the panel produced by the method according to [47];
[0090](c) a combination of the antibody set obtained by the method according to [36] and the panel produced by the method according to [48];
[0091](d) a combination of the antibody set obtained by the method according to [37] and the panel produced by the method according to [48];
[0092](e) a combination of the antibody set obtained by the method according to [38] and the panel produced by the method according to [49];
[0093](f) an isolated antibody obtained by the method according to [34] and the panel produced by the method according to [50];
[0094](g) a combination of the antibody set obtained by the method according to [39] and the panel produced by the method according to [50];
[0095](h) a combination of the antibody set obtained by the method according to [40] and the panel produced by the method according to [51];
[0096](i) a combination of the antibody set obtained by the method according to [41] and the panel produced by the method according to [51]; and
[0097](j) a combination of the antibody set obtained by the method according to [42] and the panel produced by the method according to [52].
[55] A method of testing a disease in which a cell surface antigen is an indicator, the method comprising the following steps:
[0098](1) preparing a cell or a tissue separated from a subject;
[0099](2) examining a reactivity between the cell or the tissue and each antibody displayed on the panel according to [53]; and
[0100](3) collating the results in the step (2) with the panel.
<Method of Selecting Optimum Treatment Method>
[0101][56] A method of selecting an optimum treatment method for a certain disease, the method comprising the following steps:
[0102](1) preparing a cell or a tissue separated from a subject;
[0103](2) examining a reactivity between the cell or the tissue and each antibody displayed on the panel according to [53];
[0104](3) collating the results in the step (2) with the panel, and
[0105](4) selecting an effective antibody according to the results of collating.
[57] The method according to [56], wherein the effective antibody is an antibody showing a specific reactivity in the step (2) or an antibody equivalent thereto.[58] The method according to [56] or [57], wherein the certain disease is a disease in which a cell surface antigen selected from the group consisting of HER1, HER2, CD46, ITGA3, ICAM1, ALCAM, CD147, IgSF4, BCAM, C1qR, CD44, CD73, LAR, EpCAM and HGFR is an indicator.[59] The method according to any of [56] to [58], wherein the panel displays two or more antibodies selected from the group consisting of 048-006 antibody, 057-091 antibody, 059-152 antibody, 048-040 antibody, 054-101 antibody, 055-147 antibody, 059-173 antibody, 067-149 antibody, 067-176 antibody, 015-126 antibody, 015-044 antibody, 015-102 antibody, 015-136 antibody, 015-143 antibody, 015-209 antibody, 039-016 antibody, 053-216 antibody, 075-024 antibody, 075-110 antibody, 086-032 antibody, 086-035 antibody, 086-036 antibody, 086-061 antibody, 086-138 antibody, 086-182 antibody, 035-224 antibody, 045-011 antibody, 051-144 antibody, 052-053 antibody, 052-073 antibody, 053-049 antibody, 3172-120 antibody, 066-069 antibody, 015-003 antibody, 064-002 antibody, 064-006 antibody, 064-012a antibody, 064-012b antibody, 064-014 antibody, 064-054 antibody, 064-085 antibody, 064-093 antibody, 064-116 antibody, 065-183 antibody, 067-142 antibody, 068-007 antibody, 052-033 antibody, 053-042 antibody, 053-051 antibody, 053-059 antibody, 053-085 antibody, 035-234 antibody, 040-107 antibody, 041-118 antibody, 066-174 antibody, 083-040 antibody, 029-143 antibody, 045-134 antibody, 062-101 antibody, 062-109 antibody, 084-103 antibody, 052-274 antibody, 029-067 antibody, 083-131 antibody, 059-053 antibody, 064-003 antibody, 067-213 antibody, 067-153 antibody, 067-126 antibody, 067-133 antibody, 067-287 antibody, 064-044 antibody, 065-030 antibody, 065-358 antibody, 066-019 antibody, 079-085 antibody, 067-024 antibody and 076-048 antibody.[60] A method of selecting an optimum treatment method of a certain disease, the method comprising the following steps:
[0106](1) preparing a panel displaying a reactivity between one or more antibodies selected from the group consisting of 048-006 antibody, 015-126 antibody, 067-133 antibody, 064-044 antibody, 076-048 antibody and 059-053 antibody, and a clinical cancer tissue of one or more diseases selected from the group consisting of squamous carcinoma, adenosquamous carcinoma, alveolar adenocarcinoma, adenocarcinoma, and large cell carcinoma, and a cell or tissue separated from a subject;
[0107](2) examining a reactivity between the cell or the tissue and each antibody displayed on the panel;
[0108](3) collating the results in the step (2) with the panel, and
[0109](4) selecting an effective antibody according to the results of collating.
[61] The method according to [60], wherein the effective antibody is an antibody showing a specific reactivity in the step (2) or an antibody equivalent thereto.[62] The method according to [60] or [61], wherein the certain disease is a disease selected from the group consisting of squamous carcinoma, adenosquamous carcinoma, alveolar adenocarcinoma, adenocarcinoma, and large cell carcinoma.
<Isolated Antibody>
[0110][63] An isolated antibody having affinity to HER1, comprising:
[0111]a heavy chain variable region CDR3 and a light chain variable region CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3 and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (1) to (3);
[0112]heavy chain variable regions CDR2 and CDR3 and light chain variable regions CDR2 and CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (4) to (6);
[0113]heavy chain variable regions CDR1 to CDR3 and light chain variable regions CDR1 to CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (7) to (9) and (13) to (18); or
[0114]a heavy chain variable region and a light chain variable region specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region and SEQ ID NO showing an amino acid sequence of a light chain variable region) selected from the group consisting of the following (10) to (12) and (19) to (24);
(1) SEQ ID NO: 4 and SEQ ID NO: 8
(2) SEQ ID NO: 12 and SEQ ID NO: 16
(3) SEQ ID NO: 20 and SEQ ID NO: 24
(4) SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 7, and SEQ ID NO: 8
(5) SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 15, and SEQ ID NO: 16
(6) SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 23, and SEQ ID NO: 24
(7) SEQ ID NO: 2, SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 6, SEQ ID NO: 7, and SEQ ID NO: 8
(8) SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 14, SEQ ID NO: 15, and SEQ ID NO: 16
(9) SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 22, SEQ ID NO: 23, and SEQ ID NO: 24
(10) SEQ ID NO: 1, and SEQ ID NO: 5
(11) SEQ ID NO: 9, and SEQ ID NO: 13
(12) SEQ ID NO: 17, and SEQ ID NO: 21
(13) SEQ ID NO: 484 (VH CDR1), SEQ ID NO: 485 (VH CDR2), SEQ ID NO: 486 (VH CDR3), SEQ ID NO: 488 (VL CDR1), SEQ ID NO: 489 (VL CDR2), and SEQ ID NO: 490 (VL CDR3)
(14) SEQ ID NO: 492 (VH CDR1), SEQ ID NO: 493 (VH CDR2), SEQ ID NO: 494 (VH CDR3), SEQ ID NO: 496 (VL CDR1), SEQ ID NO: 497 (VL CDR2), and SEQ ID NO: 498 (VL CDR3)
(15) SEQ ID NO: 500 (VH CDR1), SEQ ID NO: 501 (VH CDR2), SEQ ID NO: 502 (VH CDR3), SEQ ID NO: 504 (VL CDR1), SEQ ID NO: 505 (VL CDR2), and SEQ ID NO: 506 (VL CDR3)
(16) SEQ ID NO: 508 (VH CDR1), SEQ ID NO: 509 (VH CDR2), SEQ ID NO: 510 (VH CDR3), SEQ ID NO: 512 (VL CDR1), SEQ ID NO: 513 (VL CDR2), and SEQ ID NO: 514 (VL CDR3)
(17) SEQ ID NO: 516 (VH CDR1), SEQ ID NO: 517 (VH CDR2), SEQ ID NO: 518 (VH CDR3), SEQ ID NO: 520 (VL CDR1), SEQ ID NO: 521 (VL CDR2), and SEQ ID NO: 522 (VL CDR3)
(18) SEQ ID NO: 524 (VH CDR1), SEQ ID NO: 525 (VH CDR2), SEQ ID NO: 526 (VH CDR3), SEQ ID NO: 528 (VL CDR1), SEQ ID NO: 529 (VL CDR2), and SEQ ID NO: 530 (VL CDR3)
(19) SEQ ID NO: 483 (VH), and SEQ ID NO: 487 (VL)
(20) SEQ ID NO: 491 (VH), and SEQ ID NO: 495 (VL)
(21) SEQ ID NO: 499 (VH), and SEQ ID NO: 503 (VL)
(22) SEQ ID NO: 507 (VH), and SEQ ID NO: 511 (VL)
(23) SEQ ID NO: 515 (VH), and SEQ ID NO: 519 (VL), and
(24) SEQ ID NO: 523 (VH), and SEQ ID NO: 527 (VL)
[0115][64] An isolated antibody having affinity to HER2, comprising:
[0116]a heavy chain variable region CDR3 and a light chain variable region CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3 and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) shown in the following (1);
[0117]heavy chain variable regions CDR2 and CDR3 and light chain variable regions CDR2 and CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) shown in the following (2);
[0118]heavy chain variable regions CDR1 to CDR3 and light chain variable regions CDR1 to CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (3) and (5) to (19); or
[0119]a heavy chain variable region and a light chain variable region specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region and SEQ ID NO showing an amino acid sequence of a light chain variable region) selected from the group consisting of the following (4) and (20) to (34);
(1) SEQ ID NO: 28, and SEQ ID NO: 32
(2) SEQ ID NO: 27, SEQ ID NO: 28, SEQ ID NO: 31, and SEQ ID NO: 32
(3) SEQ ID NO: 26, SEQ ID NO: 27, SEQ ID NO: 28, SEQ ID NO: 30, SEQ ID NO: 31, and SEQ ID NO:32
(4) SEQ ID NO: 25, and SEQ ID NO: 29
(5) SEQ ID NO: 532 (VH CDR1), SEQ ID NO: 533 (VH CDR2), SEQ ID NO: 534 (VH CDR3), SEQ ID NO: 536 (VL CDR1), SEQ ID NO: 537 (VL CDR2), and SEQ ID NO: 538 (VL CDR3)
(6) SEQ ID NO: 540 (VH CDR1), SEQ ID NO: 541 (VH CDR2), SEQ ID NO: 542 (VH CDR3), SEQ ID NO: 544 (VL CDR1), SEQ ID NO: 545 (VL CDR2), and SEQ ID NO: 546 (VL CDR3)
(7) SEQ ID NO: 548 (VH CDR1), SEQ ID NO: 549 (VH CDR2), SEQ ID NO: 550 (VH CDR3), SEQ ID NO: 552 (VL CDR1), SEQ ID NO: 553 (VL CDR2), and SEQ ID NO: 554 (VL CDR3)
(8) SEQ ID NO: 556 (VH CDR1), SEQ ID NO: 557 (VH CDR2), SEQ ID NO: 558 (VH CDR3), SEQ ID NO: 560 (VL CDR1), SEQ ID NO: 561 (VL CDR2), and SEQ ID NO: 562 (VL CDR3)
(9) SEQ ID NO: 564 (VH CDR1), SEQ ID NO: 565 (VH CDR2), SEQ ID NO: 566 (VH CDR3), SEQ ID NO: 568 (VL CDR1), SEQ ID NO: 569 (VL CDR2), and SEQ ID NO: 570 (VL CDR3)
(10) SEQ ID NO: 572 (VH CDR1), SEQ ID NO: 573 (VH CDR2), SEQ ID NO: 574 (VH CDR3), SEQ ID NO: 576 (VL CDR1), SEQ ID NO: 577 (VL CDR2), and SEQ ID NO: 578 (VL CDR3)
(11) SEQ ID NO: 580 (VH CDR1), SEQ ID NO: 581 (VH CDR2), SEQ ID NO: 582 (VH CDR3), SEQ ID NO: 584 (VL CDR1), SEQ ID NO: 585 (VL CDR2), and SEQ ID NO: 586 (VL CDR3)
(12) SEQ ID NO: 588 (VH CDR1), SEQ ID NO: 589 (VH CDR2), SEQ ID NO: 590 (VH CDR3), SEQ ID NO: 592 (VL CDR1), SEQ ID NO: 593 (VL CDR2), and SEQ ID NO: 594 (VL CDR3)
(13) SEQ ID NO: 596 (VH CDR1), SEQ ID NO: 597 (VH CDR2), SEQ ID NO: 598 (VH CDR3), SEQ ID NO: 600 (VL CDR1), SEQ ID NO: 601 (VL CDR2), and SEQ ID NO: 602 (VL CDR3)
(14) SEQ ID NO: 604 (VH CDR1), SEQ ID NO: 605 (VH CDR2), SEQ ID NO: 606 (VH CDR3), SEQ ID NO:608 (VL CDR1), SEQ ID NO:609 (VL CDR2), and SEQ ID NO: 610 (VL CDR3)
(15) SEQ ID NO: 612 (VH CDR1), SEQ ID NO: 613 (VH CDR2), SEQ ID NO: 614 (VH CDR3), SEQ ID NO: 616 (VL CDR1), SEQ ID NO: 617 (VL CDR2), and SEQ ID NO: 618 (VL CDR3)
(16) SEQ ID NO: 620 (VH CDR1), SEQ ID NO: 621 (VH CDR2), SEQ ID NO: 622 (VH CDR3), SEQ ID NO: 624 (VL CDR1), SEQ ID NO: 625 (VL CDR2), and SEQ ID NO: 626 (VL CDR3)
(17) SEQ ID NO: 628 (VH CDR1), SEQ ID NO: 629 (VH CDR2), SEQ ID NO: 630 (VH CDR3), SEQ ID NO: 632 (VL CDR1), SEQ ID NO: 633 (VL CDR2), and SEQ ID NO: 634 (VL CDR3)
(18) SEQ ID NO: 636 (VH CDR1), SEQ ID NO: 637 (VH CDR2), SEQ ID NO: 638 (VH CDR3), SEQ ID NO: 640 (VL CDR1), SEQ ID NO: 641 (VL CDR2), and SEQ ID NO: 642 (VL CDR3)
(19) SEQ ID NO: 644 (VH CDR1), SEQ ID NO: 645 (VH CDR2), SEQ ID NO: 646 (VH CDR3), SEQ ID NO: 648 (VL CDR1), SEQ ID NO: 649 (VL CDR2), and SEQ ID NO: 650 (VL CDR3)
(20) SEQ ID NO: 531 (VH), and SEQ ID NO: 535 (VL)
(21) SEQ ID NO: 539 (VH), and SEQ ID NO: 543 (VL)
(22) SEQ ID NO: 547 (VH), and SEQ ID NO: 551 (VL)
(23) SEQ ID NO: 555 (VH), and SEQ ID NO: 559 (VL)
(24) SEQ ID NO: 563 (VH), and SEQ ID NO: 567 (VL)
(25) SEQ ID NO: 571 (VH), and SEQ ID NO: 575 (VL)
(26) SEQ ID NO: 579 (VH), and SEQ ID NO: 583 (VL)
(27) SEQ ID NO: 587 (VH), and SEQ ID NO: 591 (VL)
(28) SEQ ID NO: 595 (VH), and SEQ ID NO: 599 (VL)
(29) SEQ ID NO: 603 (VH), and SEQ ID NO: 607 (VL)
(30) SEQ ID NO: 611 (VH), and SEQ ID NO: 615 (VL)
(31) SEQ ID NO: 619 (VH), and SEQ ID NO: 623 (VL)
(32) SEQ ID NO: 627 (VH), and SEQ ID NO: 631 (VL)
(33) SEQ ID NO: 635 (VH), and SEQ ID NO: 639 (VL), and
(34) SEQ ID NO: 643 (VH), and SEQ ID NO: 647 (VL)
[0120][65] An isolated antibody having affinity to CD46 antigen, comprising:
[0121]a heavy chain variable region CDR3 and a light chain variable region CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3 and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (1) to (7);
[0122]heavy chain variable regions CDR2 and CDR3 and light chain variable regions CDR2 and CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (8) to (14);
[0123]heavy chain variable regions CDR1 to CDR3 and light chain variable regions CDR1 to CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (15) to (22); or
[0124]a heavy chain variable region and a light chain variable region specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region and SEQ ID NO showing an amino acid sequence of a light chain variable region) selected from the group consisting of the following (23) to (30);
(1) SEQ ID NO: 36, and SEQ ID NO: 40
(2) SEQ ID NO: 44, and SEQ ID NO: 48
(3) SEQ ID NO: 52, and SEQ ID NO: 56
(4) SEQ ID NO: 60, and SEQ ID NO: 64
(5) SEQ ID NO: 68, and SEQ ID NO: 72
(6) SEQ ID NO: 76, and SEQ ID NO: 80
(7) SEQ ID NO: 84, and SEQ ID NO: 88
(8) SEQ ID NO: 35, SEQ ID NO: 36, SEQ ID NO: 39, and SEQ ID NO: 40
(9) SEQ ID NO: 43, SEQ ID NO: 44, SEQ ID NO: 47, and SEQ ID NO: 48
(10) SEQ ID NO: 51, SEQ ID NO: 52, SEQ ID NO: 55, and SEQ ID NO: 56
(1) SEQ ID NO: 59, SEQ ID NO: 60, SEQ ID NO: 63, and SEQ ID NO: 64
(12) SEQ ID NO: 67, SEQ ID NO: 68, SEQ ID NO: 71, and SEQ ID NO: 72
(13) SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 79, and SEQ ID NO: 80
(14) SEQ ID NO: 83, SEQ ID NO: 84, SEQ ID NO: 87, and SEQ ID NO: 88
(15) SEQ ID NO: 34, SEQ ID NO: 35, SEQ ID NO: 36, SEQ ID NO: 38, SEQ ID NO: 39, and SEQ ID NO: 40
(16) SEQ ID NO: 42, SEQ ID NO: 43, SEQ ID NO: 44, SEQ ID NO: 46, SEQ ID NO: 47, and SEQ ID NO: 48
(17) SEQ ID NO: 50, SEQ ID NO: 51, SEQ ID NO: 52, SEQ ID NO: 54, SEQ ID NO: 55, and SEQ ID NO: 56
(18) SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, SEQ ID NO: 62, SEQ ID NO: 63, and SEQ ID NO: 64
(19) SEQ ID NO: 66, SEQ ID NO: 67, SEQ ID NO: 68, SEQ ID NO: 70, SEQ ID NO: 71, and SEQ ID NO: 72
(20) SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 78, SEQ ID NO: 79, and SEQ ID NO: 80
(21) SEQ ID NO: 82, SEQ ID NO: 83, SEQ ID NO: 84, SEQ ID NO: 86, SEQ ID NO: 87, and SEQ ID NO: 88
(22) SEQ ID NO: 756 (VH CDR1), SEQ ID NO: 757 (VH CDR2), SEQ ID NO: 758 (VH CDR3), SEQ ID NO: 760 (VL CDR1), SEQ ID NO: 761 (VL CDR2), and SEQ ID NO: 762 (VL CDR3)
(23) SEQ ID NO: 33, and SEQ ID NO: 37
(24) SEQ ID NO: 41, and SEQ ID NO: 45
(25) SEQ ID NO: 49, and SEQ ID NO: 53
(26) SEQ ID NO: 57, and SEQ ID NO: 61
(27) SEQ ID NO: 65, and SEQ ID NO: 69
(28) SEQ ID NO: 73, and SEQ ID NO: 77
(29) SEQ ID NO: 81, and SEQ ID NO: 85
(30) SEQ ID NO: 755 (VH), and SEQ ID NO: 759 (VL)
[0125][66] An isolated antibody having affinity to ITAG3, comprising:
[0126]a heavy chain variable region CDR3 and a light chain variable region CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3 and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) shown in the following (1);
[0127]heavy chain variable regions CDR2 and CDR3 and light chain variable regions CDR2 and CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) shown in the following (2);
[0128]heavy chain variable regions CDR1 to CDR3 and light chain variable regions CDR1 to CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (3) and (5) to (16); or
[0129]a heavy chain variable region and a light chain variable region specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region and SEQ ID NO showing an amino acid sequence of a light chain variable region) selected from the group consisting of the following (4) and (17) to (28);
(1) SEQ ID NO: 92, and SEQ ID NO: 96
(2) SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 95, and SEQ ID NO: 96
(3) SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 94, and SEQ ID NO: 95,
(4) SEQ ID NO: 89, and SEQ ID NO: 93
(5) SEQ ID NO: 676 (VH CDR1), SEQ ID NO: 677 (VH CDR2), SEQ ID NO: 678 (VH CDR3), SEQ ID NO: 680 (VL CDR1), SEQ ID NO: 681 (VL CDR2), and SEQ ID NO: 682 (VL CDR3)
(6) SEQ ID NO: 684 (VH CDR1), SEQ ID NO: 685 (VH CDR2), SEQ ID NO: 686 (VH CDR3), SEQ ID NO: 688 (VL CDR1), SEQ ID NO: 689 (VL CDR2), and SEQ ID NO: 690 (VL CDR3)
(7) SEQ ID NO: 692 (VH CDR1), SEQ ID NO: 693 (VH CDR2), SEQ ID NO: 694 (VH CDR3), SEQ ID NO: 696 (VL CDR1), SEQ ID NO: 697 (VL CDR2), and SEQ ID NO: 698 (VL CDR3)
(8) SEQ ID NO: 700 (VH CDR1), SEQ ID NO: 701 (VH CDR2), SEQ ID NO: 702 (VH CDR3), SEQ ID NO: 704 (VL CDR1), SEQ ID NO: 705 (VL CDR2), and SEQ ID NO: 706 (VL CDR3)
(9) SEQ ID NO: 708 (VH CDR1), SEQ ID NO: 709 (VH CDR2), SEQ ID NO: 710 (VH CDR3), SEQ ID NO: 712 (VL CDR1), SEQ ID NO: 713 (VL CDR2), and SEQ ID NO: 714 (VL CDR3)
(10) SEQ ID NO: 716 (VH CDR1), SEQ ID NO: 717 (VH CDR2), SEQ ID NO: 718 (VH CDR3), SEQ ID NO: 720 (VL CDR1), SEQ ID NO: 721 (VL CDR2), and SEQ ID NO: 722 (VL CDR3)
(11) SEQ ID NO: 724 (VH CDR1), SEQ ID NO: 725 (VH CDR2), SEQ ID NO: 726 (VH CDR3), SEQ ID NO: 728 (VL CDR1), SEQ ID NO: 729 (VL CDR2), and SEQ ID NO: 730 (VL CDR3)
(12) SEQ ID NO: 732 (VH CDR1), SEQ ID NO: 733 (VH CDR2), SEQ ID NO: 734 (VH CDR3), SEQ ID NO: 736 (VL CDR1), SEQ ID NO: 737 (VL CDR2), and SEQ ID NO: 738 (VL CDR3)
(13) SEQ ID NO: 740 (VH CDR1), SEQ ID NO: 741 (VH CDR2), SEQ ID NO: 742 (VH CDR3), SEQ ID NO: 744 (VL CDR1), SEQ ID NO: 745 (VL CDR2), and SEQ ID NO: 746 (VL CDR3)
(14) SEQ ID NO: 748 (VH CDR1), SEQ ID NO: 749 (VH CDR2), SEQ ID NO: 750 (VH CDR3), SEQ ID NO: 752 (VL CDR1), SEQ ID NO: 753 (VL CDR2), and SEQ ID NO: 754 (VL CDR3)
(15) SEQ ID NO: 764 (VH CDR1), SEQ ID NO: 765 (VH CDR2), SEQ ID NO: 766 (VH CDR3), SEQ ID NO: 768 (VL CDR1), SEQ ID NO: 769 (VL CDR2), and SEQ ID NO: 770 (VL CDR3)
(16) SEQ ID NO: 772 (VH CDR1), SEQ ID NO: 773 (VH CDR2), SEQ ID NO: 774 (VH CDR3), SEQ ID NO: 776 (VL CDR1), SEQ ID NO: 777 (VL CDR2), and SEQ ID NO: 778 (VL CDR3)
(17) SEQ ID NO: 675 (VH), and SEQ ID NO: 679 (VL)
(18) SEQ ID NO: 683 (VH), and SEQ ID NO: 687 (VL)
(19) SEQ ID NO: 691 (VH), and SEQ ID NO: 695 (VL)
(20) SEQ ID NO: 699 (VH), and SEQ ID NO: 703 (VL)
(21) SEQ ID NO: 707 (VH), and SEQ ID NO: 711 (VL)
(22) SEQ ID NO: 715 (VH), and SEQ ID NO: 719 (VL)
(23) SEQ ID NO: 723 (VH), and SEQ ID NO: 727 (VL)
(24) SEQ ID NO: 731 (VH), and SEQ ID NO: 735 (VL)
(25) SEQ ID NO: 739 (VH), and SEQ ID NO: 743 (VL)
(26) SEQ ID NO: 747 (VH), and SEQ ID NO: 751 (VL)
(27) SEQ ID NO: 763 (VH), and SEQ ID NO: 767 (VL), and
(28) SEQ ID NO: 771 (VH), and SEQ ID NO: 775 (VL)
[0130][67] An isolated antibody having affinity to ICAM1, comprising:
[0131]a heavy chain variable region CDR3 and a light chain variable region CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3 and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (1) to (5);
[0132]heavy chain variable regions CDR2 and CDR3 and light chain variable regions CDR2 and CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (6) to (10);
[0133]heavy chain variable regions CDR1 to CDR3 and light chain variable regions CDR1 to CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (11) to (15); or
[0134]a heavy chain variable region and a light chain variable region specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region and SEQ ID NO showing an amino acid sequence of a light chain variable region) selected from the group consisting of the following (16) to (20);
(1) SEQ ID NO: 100, and SEQ ID NO: 104
(2) SEQ ID NO: 108, and SEQ ID NO: 112
(3) SEQ ID NO: 116, and SEQ ID NO: 120
(4) SEQ ID NO: 124, and SEQ ID NO: 128
(5) SEQ ID NO: 132, and SEQ ID NO: 136
(6) SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 103, and SEQ ID NO: 104
(7) SEQ ID NO: 107, SEQ ID NO: 108, SEQ ID NO: 111, and SEQ ID NO: 112
(8) SEQ ID NO: 115, SEQ ID NO: 116, SEQ ID NO: 119, and SEQ ID NO: 120
(9) SEQ ID NO: 123, SEQ ID NO: 124, SEQ ID NO: 127, and SEQ ID NO: 128
(10) SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 135, SEQ ID NO: 136
(11) SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 102, SEQ ID NO: 103, and SEQ ID NO: 104
(12) SEQ ID NO: 106, SEQ ID NO: 107, SEQ ID NO: 108, SEQ ID NO: 110, SEQ ID NO: 111, and SEQ ID NO: 112
(13) SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 116, SEQ ID NO: 118, SEQ ID NO: 119, and SEQ ID NO: 120
(14) SEQ ID NO: 122, SEQ ID NO: 123, SEQ ID NO: 124, SEQ ID NO: 126, SEQ ID NO: 127, and SEQ ID NO: 128
(15) SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 134, SEQ ID NO: 135, and SEQ ID NO: 136
(16) SEQ ID NO: 97, and SEQ ID NO: 101
(17) SEQ ID NO: 105, and SEQ ID NO: 109
(18) SEQ ID NO: 113, and SEQ ID NO: 117
(19) SEQ ID NO: 121, and SEQ ID NO: 125
(20) SEQ ID NO: 129, and SEQ ID NO: 133
[0135][68] An isolated antibody having affinity to ALCAM, comprising:
[0136]a heavy chain variable region CDR3 and a light chain variable region CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3 and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (1) to (5);
[0137]heavy chain variable regions CDR2 and CDR3 and light chain variable regions CDR2 and CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (6) to (10);
[0138]heavy chain variable regions CDR1 to CDR3 and light chain variable regions CDR1 to CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (11) to (15) and (21) to (28); or
[0139]a heavy chain variable region and a light chain variable region specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region and SEQ ID NO showing an amino acid sequence of a light chain variable region) selected from the group consisting of the following (16) to (20) and (29) to (36);
(1) SEQ ID NO: 140, and SEQ ID NO: 144
(2) SEQ ID NO: 148, and SEQ ID NO: 152
(3) SEQ ID NO: 156, and SEQ ID NO: 160
(4) SEQ ID NO: 164, and SEQ ID NO: 168
(5) SEQ ID NO: 172, and SEQ ID NO: 176
(6) SEQ ID NO: 139, SEQ ID NO: 140, SEQ ID NO: 143, and SEQ ID NO: 144
(7) SEQ ID NO: 147, SEQ ID NO: 148, SEQ ID NO: 151, and SEQ ID NO: 152
(8) SEQ ID NO: 155, SEQ ID NO: 156, SEQ ID NO: 159, and SEQ ID NO: 160
(9) SEQ ID NO: 163, SEQ ID NO: 164, SEQ ID NO: 167, and SEQ ID NO: 168
(10) SEQ ID NO: 171, SEQ ID NO: 172, SEQ ID NO: 175, and SEQ ID NO: 176
(1) SEQ ID NO: 138, SEQ ID NO: 139, SEQ ID NO: 140, SEQ ID NO: 142, SEQ ID NO: 143, and SEQ ID NO: 144
(12) SEQ ID NO: 146, SEQ ID NO: 147, SEQ ID NO: 148, SEQ ID NO: 150, SEQ ID NO: 151, and SEQ ID NO: 152
(13) SEQ ID NO: 154, SEQ ID NO: 155, SEQ ID NO: 156, SEQ ID NO: 158, SEQ ID NO: 159, and SEQ ID NO: 160
(14) SEQ ID NO: 162, SEQ ID NO: 163, SEQ ID NO: 164, SEQ ID NO: 166, SEQ ID NO: 167, and SEQ ID NO: 168
(15) SEQ ID NO: 170, SEQ ID NO: 171, SEQ ID NO: 172, SEQ ID NO: 174, SEQ ID NO: 175, and SEQ ID NO: 176
(16) SEQ ID NO: 137, and SEQ ID NO: 141
(17) SEQ ID NO: 145, and SEQ ID NO: 149
(18) SEQ ID NO: 153, and SEQ ID NO: 157
(19) SEQ ID NO: 161, SEQ ID NO: 165
(20) SEQ ID NO: 169, and SEQ ID NO: 173
(21) SEQ ID NO: 780 (VH CDR1), SEQ ID NO: 781 (VH CDR2), SEQ ID NO 782 (VH CDR3), SEQ ID NO: 784 (VL CDR1), SEQ ID NO: 785 (VL CDR2), and SEQ ID NO: 786 (VL CDR3)
(22) SEQ ID NO: 788 (VH CDR1), SEQ ID NO: 789 (VH CDR2), SEQ ID NO: 790 (VH CDR3), SEQ ID NO: 792 (VL CDR1), SEQ ID NO: 793 (VL CDR2), and SEQ ID NO: 794 (VL CDR3)
(23) SEQ ID NO: 796 (VH CDR1), SEQ ID NO: 797 (VH CDR2), SEQ ID NO: 798 (VH CDR3), SEQ ID NO: 800 (VL CDR1), SEQ ID NO: 801 (VL CDR2), and SEQ ID NO: 802 (VL CDR3)
(24) SEQ ID NO: 804 (VH CDR1), SEQ ID NO: 805 (VH CDR2), SEQ ID NO: 806 (VH CDR3), SEQ ID NO: 808 (VL CDR1), SEQ ID NO: 809 (VL CDR2), and SEQ ID NO: 810 (VL CDR3)
(25) SEQ ID NO: 812 (VH CDR1), SEQ ID NO: 813 (VH CDR2), SEQ ID NO: 814 (VH CDR3), SEQ ID NO: 816 (VL CDR1), SEQ ID NO: 817 (VL CDR2), and SEQ ID NO: 818 (VL CDR3)
(26) SEQ ID NO: 820 (VH CDR1), SEQ ID NO: 821 (VH CDR2), SEQ ID NO: 822 (VH CDR3), SEQ ID NO: 824 (VL CDR1), SEQ ID NO: 825 (VL CDR2), and SEQ ID NO: 826 (VL CDR3)
(27) SEQ ID NO: 828 (VH CDR1), SEQ ID NO: 829 (VH CDR2), SEQ ID NO: 830 (VH CDR3), SEQ ID NO: 832 (VL CDR1), SEQ ID NO: 833 (VL CDR2), and SEQ ID NO: 834 (VL CDR3)
(28) SEQ ID NO: 836 (VH CDR1), SEQ ID NO: 837 (VH CDR2), SEQ ID NO: 838 (VH CDR3), SEQ ID NO: 840 (VL CDR1), SEQ ID NO: 841 (VL CDR2), and SEQ ID NO: 842 (VL CDR3)
(29) SEQ ID NO: 779 (VH), and SEQ ID NO: 783 (VL)
(30) SEQ ID NO: 787 (VH), and SEQ ID NO: 791 (VL)
(31) SEQ ID NO: 795 (VH), and SEQ ID NO: 799 (VL)
(32) SEQ ID NO: 803 (VH), and SEQ ID NO: 807 (VL)
(33) SEQ ID NO: 811 (VH), and SEQ ID NO: 815 (VL)
(34) SEQ ID NO: 819 (VH), and SEQ ID NO: 823 (VL)
(35) SEQ ID NO: 827 (VH), and SEQ ID NO: 831 (VL), and
(36) SEQ ID NO: 835 (VH), and SEQ ID NO: 839 (VL)
[0140][69] An isolated antibody having affinity to CD147 antigen, comprising:
[0141]a heavy chain variable region CDR3 and a light chain variable region CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3 and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) shown in the following (1);
[0142]heavy chain variable regions CDR2 and CDR3 and light chain variable regions CDR2 and CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) shown in the following (2);
[0143]heavy chain variable regions CDR1 to CDR3 and light chain variable regions
[0144]CDR1 to CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) shown in the following (3); or
[0145]a heavy chain variable region and a light chain variable region specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region and SEQ ID NO showing an amino acid sequence of a light chain variable region) shown in the following (4);
(1) SEQ ID NO: 180, and SEQ ID NO: 184
(2) SEQ ID NO: 179, SEQ ID NO: 180, SEQ ID NO: 183, and SEQ ID NO: 184
(3) SEQ ID NO: 178, SEQ ID NO: 179, SEQ ID NO: 180, SEQ ID NO: 182, SEQ ID NO: 183, and SEQ ID NO: 184, and
(4) SEQ ID NO: 177, and SEQ ID NO: 181
[0146][70] An isolated antibody having affinity to C1qR, comprising:
[0147]heavy chain variable regions CDR1 to CDR3 and light chain variable regions CDR1 to CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) shown in the following (1); or
[0148]a heavy chain variable region and a light chain variable region specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region and SEQ ID NO showing an amino acid sequence of a light chain variable region) shown in the following (2);
(1) SEQ ID NO: (VH CDR1) 452, SEQ ID NO: 453 (VH CDR2), SEQ ID NO: 454 (VH CDR3), SEQ ID NO: (VL CDR1) 456, SEQ ID NO: 457 (VL CDR2), and SEQ ID NO: 458 (VL CDR3), and
(2) SEQ ID NO: 451 (VH), and SEQ ID NO: 455 (VL)
[0149][71] An isolated antibody having affinity to CD44, comprising:
[0150]heavy chain variable regions CDR1 to CDR3 and light chain variable regions CDR1 to CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ. ID NO showing an amino acid sequence of a light chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) shown in the following (1); or
[0151]a heavy chain variable region and a light chain variable region specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region and SEQ ID NO showing an amino acid sequence of a light chain variable region) shown in the following (2);
(1) SEQ ID NO: 460 (VH CDR1), SEQ ID NO: 461 (VH CDR2), SEQ ID NO: 462 (VH CDR3), SEQ ID NO: 464 (VL CDR1), SEQ ID NO: 465 (VL CDR2), and SEQ ID NO: 466 (VL CDR3), and
(2) SEQ ID NO: 459 (VH), and SEQ ID NO: 463 (VL)
[0152][72] An isolated antibody having affinity to CD73, comprising:
[0153]heavy chain variable regions CDR1 to CDR3 and light chain variable regions CDR1 to CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR1; SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) shown in the following (1); or
[0154]a heavy chain variable region and a light chain variable region specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region and SEQ ID NO showing an amino acid sequence of a light chain variable region) shown in the following (2);
(1) SEQ ID NO: 468 (VH CDR1), SEQ ID NO: 469 (VH CDR2), SEQ ID NO: 470 (VH CDR3), SEQ ID NO: 472 (VL CDR1), SEQ ID NO: 473 (VL CDR2), and SEQ ID NO: 474 (VL CDR3), and
(2) SEQ ID NO: 467 (VH), and SEQ ID NO: 471 (VL)
[0155][73] An isolated antibody having affinity to EpCAM, comprising:
[0156]heavy chain variable regions CDR1 to CDR3 and light chain variable regions CDR1 to CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) shown in the following (1); or
[0157]a heavy chain variable region and a light chain variable region specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region and SEQ ID NO showing an amino acid sequence of a light chain variable region) shown in the following (2);
(1) SEQ ID NO: 476 (VH CDR1), SEQ ID NO: 477 (VH CDR2), SEQ ID NO: 478 (VH CDR3), SEQ ID NO: 480 (VL CDR1), SEQ ID NO: 481 (VL CDR2), and SEQ ID NO: 482 (VL CDR3), and
(2) SEQ ID NO: 475 (VH), and SEQ ID NO: 479 (VL)
[0158][74] An isolated antibody having affinity to HGFR, comprising:
[0159]heavy chain variable regions CDR1 to CDR3 and light chain variable regions CDR1 to CDR3 specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR2, SEQ ID NO showing an amino acid sequence of a heavy chain variable region CDR3, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR1, SEQ ID NO showing an amino acid sequence of a light chain variable region CDR2, and SEQ ID NO showing an amino acid sequence of a light chain variable region CDR3) selected from the group consisting of the following (1) to (3); or
[0160]a heavy chain variable region and a light chain variable region specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region and SEQ ID NO showing an amino acid sequence of a light chain variable region) selected from the group consisting of the following (4) to (6);
(1) SEQ ID NO: 652 (VH CDR1), SEQ ID NO: 653 (VH CDR2), SEQ ID NO: 654 (VH CDR3), SEQ ID NO: 656 (VL CDR1), SEQ ID NO: 657 (VL CDR2), and SEQ ID NO: 658 (VL CDR3)
(2) SEQ ID NO: 660 (VH CDR1), SEQ ID NO: 661 (VH CDR2), SEQ ID NO: 662 (VH CDR3), SEQ ID NO: 664 (VL CDR1), SEQ ID NO: 665 (VL CDR2), and SEQ ID NO: 666 (VL CDR3)
(3) SEQ ID NO: 668 (VH CDR1), SEQ ID NO: 669 (VH CDR2), SEQ ID NO: 670 (VH CDR3), SEQ ID NO: 672 (VL CDR1), SEQ ID NO: 673 (VL CDR2), and SEQ ID NO: 674 (VL CDR3)
(4) SEQ ID NO: 651 (VH), and SEQ ID NO: 655 (VL)
(5) SEQ ID NO: 659 (VH), and SEQ ID NO: 663 (VL), and
(6) SEQ ID NO: 667 (VH), and SEQ ID NO: 671 (VL)
[0161][75] An isolated antibody having affinity to LAR, comprising:
[0162]a heavy chain variable region and a light chain variable region specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region and SEQ ID NO showing an amino acid sequence of a light chain variable region) selected from the group consisting of the following (1) to (5);
(1) SEQ ID NO: 944 (VH), and SEQ ID NO: 945 (VL)
(2) SEQ ID NO: 946 (VH), and SEQ ID NO: 947 (VL)
(3) SEQ ID NO: 948 (VH), and SEQ ID NO: 949 (VL)
(4) SEQ ID NO: 950 (VH), and SEQ ID NO: 951 (VL), and
(5) SEQ ID NO: 952 (VH), and SEQ ID NO: 953 (VL)
[0163][76] An isolated antibody having affinity to BCAM, comprising:
[0164]a heavy chain variable region and a light chain variable region specified by a combination of SEQ ID NOs (SEQ ID NO showing an amino acid sequence of a heavy chain variable region and SEQ ID NO showing an amino acid sequence of a light chain variable region) shown in the group consisting of the following (1);
(1) SEQ ID NO: 954 (VH), and SEQ ID NO: 955 (VL)
<Isolated Nucleic Acid Molecule, Vector, and the Like>
[0165][77] An isolated nucleic acid molecule, which encodes the heavy chain variable region and/or the light chain variable region of the antibody according to any of [63] to [76].[78] A vector including the nucleic acid molecule according to [77] in a form capable of being expressed.[79] A transformant into which the nucleic acid molecule according to [77] is introduced.[80] A cancer therapeutic agent comprising the antibody according to any of [63] to [76] as an effective ingredient.[81] A reagent for examining or studying cancer comprising the antibody according to any of [63] to [76].
<Examination Method>
[0166][82] A method for examining gallbladder and liver cancer or pancreas cancer, the method comprising the following steps:
[0167](1) preparing subject cells or tissues separated from a living body; and
[0168](2) detecting a CD46 antigen in the subject cells or tissues.
[83] A method for examining gallbladder and liver cancer or pancreas cancer, the method comprising the following steps:
[0169](1) preparing subject cells or tissues separated from a living body; and
[0170](2) detecting ITGA3 in the subject cells or tissues.
[84] A method for examining kidney cancer, hepatic cell carcinoma or gallbladder and liver cancer, the method comprising the following steps:
[0171](1) preparing subject cells or tissues separated from a living body; and
[0172](2) detecting ALCAM in the subject cells or tissues.
[85] A method for examining kidney cancer, the method comprising the following steps:
[0173](1) preparing subject cells or tissues separated from a living body; and
[0174](2) detecting a CD147 antigen in the subject cells or tissues.
BRIEF DESCRIPTION OF THE DRAWINGS
[0175]FIG. 1 shows one example of a method of obtaining an antibody or an antibody set related to a certain disease.
[0176]FIG. 2 shows another example of a method of obtaining an antibody set related to a certain disease.
[0177]FIG. 3 shows a further example of a method of obtaining an antibody set related to a certain disease.
[0178]FIG. 4 shows a yet further example of a method of obtaining an antibody set related to a certain disease.
[0179]FIG. 5 is a schematic view showing a vector used for producing an scFv antibody gene library.
[0180]FIG. 6 is a schematic view showing a structure of pscFvCA9-E8VHdVLd.
[0181]FIG. 7-1 shows a base sequence (SEQ ID NO: 401) of an insert part of pscFvCA9-E8VHdVLd and an amino acid sequence (SEQ ID NO: 402) encoded by the base sequence.
[0182]FIG. 7-2 shows a part continuing to FIG. 7-1.
[0183]FIG. 8-1 shows a base sequence (SEQ ID NO: 405) of an insert of pscFvCA-E8VHd and a restriction enzyme site and an amino acid sequence (SEQ ID NO: 406).
[0184]FIG. 8-2 shows a part continuing to FIG. 8-1.
[0185]FIG. 9 shows a process of screening of an antibody clone specific to liver cancer cell.
[0186]FIG. 10 shows an FCM reactivity (representative example) of an antibody clone, showing histogram (right) and cell fluorescence cytology image (left) showing the reactivity between an antibody clones 035-011 and 041-101 and undifferentiated malignant liver cancer cell line HLF.
[0187]FIG. 11 shows an FCM reactivity (representative example) of an antibody clone, showing histogram (right) and cell fluorescence cytology image (left) showing the reactivity between an antibody clones 041-129, 045-134 and 052-042 and undifferentiated malignant liver cancer cell line HLF.
[0188]FIG. 12 shows histograms obtained by FCM of seven kinds of antibodies, which are overwritten onto each other. This shows that each histogram has a unique shape.
[0189]FIG. 13 shows histograms obtained by FCM of seven kinds of antibodies, which are overwritten onto each other. This shows that all the histograms have high similarity to each other.
[0190]FIG. 14 shows histograms obtained by FCM of four kinds of antibodies, which are overwritten onto each other. This shows that all the histograms have high similarity to each other.
[0191]FIG. 15 shows histograms obtained by FCM of two kinds of antibodies, which are overwritten onto each other. This shows that two histograms have high similarity to each other.
[0192]FIG. 16 shows histograms obtained by FCM of three kinds of antibodies in various cells, which are overwritten onto each other. This shows that even when any cells are used, these antibodies provide histograms having a high similarity to each other.
[0193]FIG. 17 shows a method for classifying the antibody group into groups based on the results of the FCM analysis.
[0194]FIG. 18 is a table showing a classification of a plurality of antibody clones based on the results of the FCM analysis. Each reference mark in Table is shown by a shift amount from the histogram (reference histogram) provided by the negative control antibody. Double circle mark represents that the shift amount is 20 times or more (the peak value of the is 20 times or more of the reference histogram); "o" (circle mark) represents that the shift amount is 10 times or more; "Δ" (triangle mark) represents that the shift amount is 3 times or more; and "x" represents that the shift amount is less than 3, respectively (an oblique line means no data is obtained).
[0195]FIG. 19 shows the results of RNAi in which CD147 is a subject antigen. Gray color (a); cells that have not subjected to RNAi are stained with an anti-influenza antibody YA14 cp3 as a primary antibody; Green color (b); cells that have not subjected to RNAi are stained with 059-053 cp3 as a primary antibody; Red color (c); cells that have subjected to RNAi are stained with 059-053 cp3 as a primary antibody.
[0196]FIG. 20 shows the results of RNAi in which CD166 is a subject antigen. Gray color (a); cells that have not subjected to RNAi are stained with an anti-influenza antibody YA14 cp3 as a primary antibody; Green color (b); cells that have not subjected to RNAi are stained with 035-234 cp3 as a primary antibody; Red color (c); cells that have subjected to RNAi are stained with 035-234 cp3 as a primary antibody.
[0197]FIG. 21 shows the results of RNAi in which HER1 is a subject antigen. Gray color (a); cells that have not subjected to RNAi are stained with an anti-influenza antibody YA14 cp3 as a primary antibody; Green color (b); cells that have not subjected to RNAi are stained with 048-006 cp3 as a primary antibody; Red color (c); cells that have subjected to RNAi are stained with 048-006 cp3 as a primary antibody.
[0198]FIG. 22 shows the results of RNAi in which HER2 is a subject antigen. Gray color (a); cells that have not subjected to RNAi are stained with an anti-influenza antibody YA14 cp3 as a primary antibody; Green color (b); cells that have not subjected to RNAi are stained with 015-126 cp3 as a primary antibody; Red color (c); cells that have subjected to RNAi are stained with 015-126 cp3 as a primary antibody.
[0199]FIG. 23 shows the results of RNAi in which IgSF4 is a subject antigen. Gray color (a); cells that have not subjected to RNAI are stained with an anti-influenza antibody YA14 cp3 as a primary antibody; light blue color (b); cells that have not subjected to RNAi are stained with 035-273 cp3 as a primary antibody; orange color (c); cells that have subjected to RNAi are stained with 035-273 cp3 as a primary antibody.
[0200]FIG. 24 shows A: an EGF binding inhibitory activity (using A431 cells) of 048-006 antibody and 059-152 antibody; B: an EGF binding inhibitory activity of 048-006 antibody (using low concentration range, A431 cells), and C: an EGF binding inhibitory activity of 048-006 antibody (using low concentration range, A431 cells).
[0201]FIG. 25 shows A: HER1 phosphorylation signal inhibitory activity of 048-006 antibody and 059-152 antibody (results of Western blotting). Lane 1; antibody is not added, lane 2; HR1-007 added (10 μg/ml), lane 3; 048-006 antibody added (10 μg/ml), lane 4; 048-006 antibody added (10 μg/ml), lane 5; 059-152 antibody added (10 μg/ml), lane 6; and 059-152 antibody added (10 μg/ml). Upper part shows the results of Western blotting by using anti-phosphorylation tyrosine antibody (mouse monoclonal antibody). Lower part shows the results of Western blotting by using anti-β actin antibody (rabbit polyclonal antibody). B: HER1 phosphorylation signal inhibitory activity of a 048-006 antibody (low concentration range). Lane 1; not treated, lane 2; antibody is not added, lane 3; HR1-007 is added (1 μg/ml), lane 4; 048-006 antibody added (0.5 μg/ml), lane 5; 048-006 antibody added (0.1 μg/ml), lane 6; and 048-006 antibody added (0.05 μg/ml). After incubation with an antibody for 30 minutes, Her1 was added. Upper part shows the results of Western blotting by using anti-phosphorylation tyrosine antibody (mouse monoclonal antibody). Lower part shows the results of Western blotting by using anti-β actin antibody (rabbit polyclonal antibody). C: Comparison of HER1 phosphorylation signal inhibition effects of 048-006 antibody, 059-152 antibody and ERBITUX (using A-431 cells. Lane 1; HR1-007, lane 2; 048-006 antibody, lane 3; 059-152 antibody, lane 4; ERBITUX, lane 5; antibody is not added (EGF (+)), lane 6; antibody is not added (EGF (-)). D: Comparison of HER1 phosphorylation signal inhibition effects of 048-006 antibody, 059-152 antibody and ERBITUX (using CCF-RC1 cells). Lane 1; HR1-007, lane 2; 048-006 antibody, lane 3; 059-152 antibody, lane 4; ERBITUX, lane 5; antibody is not added (EGF (+)), lane 6; antibody is not added (EGF (-)). E: Comparison of HER1 phosphorylation signal inhibition effects of 048-006 antibody and 059-152 antibody clone and ERBITUX (using Caki-1 cells). Lane 1; HR1-007, lane 2; 048-006 antibody, lane 3; 059-152 antibody, lane 4; ERBITUX, lane 5; antibody is not added (EGF (+)), lane 6; antibody is not added (EGF (-)).
[0202]FIG. 26 shows a result of BIACORE experiment. Fixation method: CM5 chip of BIAcore is used and NHS is used so as to fix a partial sequence of HER1 to sensor. 048-006 antibody is allowed to flow at the above-mentioned concentration to observe signals.
[0203]FIG. 27 shows a result of an ADCC activity test. An antibody to be used: anti-ITGA3 antibody, a target culture cell: HLF.
[0204]FIG. 28 shows a result of an ADCC activity test. An antibody to be used: anti-HER1 antibody, a target culture cell: A-431.
[0205]FIG. 29 shows a result of an ADCC activity test. An antibody to be used: anti-HER1 antibody, a target culture cell: A549.
[0206]FIG. 30 shows a result of an ADCC activity test. An antibody to be used: anti-HER1 antibody, a target culture cell: ACHN.
[0207]FIG. 31 shows a result of an ADCC activity test. An antibody to be used: anti-HER1 antibody, a target culture cell: CCF-RC-1.
[0208]FIG. 32 shows a result of an ADCC activity test. An antibody to be used: anti-HER1 antibody, a target culture cell: NCI-H1373.
[0209]FIG. 33 shows a result of an ADCC activity test. An antibody to be used: anti-HER1 antibody, a target culture cell: SK-OV-3.
[0210]FIG. 34 shows a result of an ADCC activity test. An antibody to be used: anti-HER2 antibody, a target culture cell: BT-474.
[0211]FIG. 35 shows a result of an ADCC activity test. (a) An antibody to be used: anti-ALCAM antibody, 066-174 whose, a target culture cell: NCI-H1373. (b) An antibody to be used: anti-ALCAM antibody, 066-174, target culture cell: CW2. (c) An antibody to be used: anti-ALCAM antibody, 066-174, target culture cell: NCI-H441.
[0212]FIG. 36 shows a result of an ADCC activity test. (a) An antibody to be used: anti-ALCAM antibody, 035-234, target culture cell: CW2. (b) An antibody to be used: anti-ALCAM antibody, 035-234, target culture cell: NCI-H441.
[0213]FIG. 37 shows a result of an ADCC activity test. (a) An antibody to be used: anti-ICAM1 antibody, 053-051, target culture cell: NCI-H441. (b) An antibody to be used: anti-ICAM1 antibody, 053-051, target culture cell: HepG2.
[0214]FIG. 38 shows a result of an ADCC activity test. (a) An antibody to be used: anti-ICAM1 antibody, 053-059, target culture cell: NCI-H441. (b) An antibody to be used: anti-ICAM1 antibody, 053-059, target culture cell: HepG2.
[0215]FIG. 39 shows a result of an ADCC activity test. (a) An antibody to be used: anti-ICAM1 antibody, 053-085, target culture cell: NCI-H441. (b) An antibody to be used: anti-ICAM1 antibody, 053-085, target culture cell: HepG2.
[0216]FIG. 40 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-HER1 antibody, 048-006 antibody or 059-152 antibody,
[0217]target culture cell: CCF-RC-1.
[0218]FIG. 41 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-HER1 antibody, 048-006 antibody or 059-152 antibody, target culture cell: NCI-H1373.
[0219]FIG. 42 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-HER1 antibody, 048-006 antibody or 059-152 antibody,
[0220]target culture cell: A-431.
[0221]FIG. 43 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-ALCAM antibody, 041-118 antibody, target culture cell: NCI-H1373.
[0222]FIG. 44 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-EpCAM antibody, 067-153 antibody, target culture cell: MKN-45.
[0223]FIG. 45 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-EpCAM antibody, 067-153 antibody, target culture cell: HT-29.
[0224]FIG. 46 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-EpCAM antibody, 067-153 antibody, target culture cell: NCI-H1373.
[0225]FIG. 47 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-HGFR antibody, 067-133 antibody, target culture cell: NCI-H1373.
[0226]FIG. 48 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-HER1 antibody, 055-147 antibody or 059-173 antibody, target culture cell: CCF-RC1.
[0227]FIG. 49 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-HER1 antibody. 048-006 antibody, 059-152 antibody, 055-147 antibody or 059-173 antibody, target culture cell: HT-29.
[0228]FIG. 50 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-HER1 antibody, 048-006 antibody, 055-147 antibody or 059-173 antibody, target culture cell: A431.
[0229]FIG. 51 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-HER1 antibody, 048-006 antibody or 059-152 antibody, target culture cell: ACHN.
[0230]FIG. 52 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-ALCAM antibody, 035-234 antibody or 066-174 antibody,
[0231]target culture cell: NCI-H1373.
[0232]FIG. 53 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-ALCAM antibody, 035-234 antibody or 066-174 antibody, target culture cell: SKOv3.
[0233]FIG. 54 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-ALCAM antibody, 035-234 antibody or 066-174 antibody, target culture cell: CW-2.
[0234]FIG. 55 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-ALCAM antibody, 041-118 antibody, target culture cell: EBC-1.
[0235]FIG. 56 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-ALCAM antibody, 080-040 antibody, target culture cell: NCI-H1373.
[0236]FIG. 57 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-ICAM1 antibody, 053-042 antibody, target culture cell: NCI-H1373.
[0237]FIG. 58 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-ICAM1 antibody, 053-051 antibody, 053-059 antibody or 053-085 antibody, target culture cell: NCI-H1373.
[0238]FIG. 59 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-EpCAM antibody, 067-153 antibody, target culture cell: EBC-1.
[0239]FIG. 60 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-HGFR antibody 067-133 antibody, target culture cell: MKN-45.
[0240]FIG. 61 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-HGFR antibody 067-133 antibody, target culture cell: EBC-1.
[0241]FIG. 62 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-ITGA3 antibody, 015-003 antibody, target culture cell: ACHN.
[0242]FIG. 63 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-CD147 antibody, 059-053 antibody, target culture cell: CCF-RC1.
[0243]FIG. 64 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-CD147 antibody, 059-053 antibody, target culture cell: ACHN.
[0244]FIG. 65 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-PTP-LAR antibody, 064-044 antibody or 079-085 antibody, target culture cell: PC-14.
[0245]FIG. 66 shows antibody dosage dependence of the ADCC activity. An antibody to be used: anti-CD44 antibody. 064-003 antibody, target culture cell: PC-14.
[0246]FIG. 67 shows a result of a cell proliferation inhibition test. An antibody to be used: anti-HER1 antibody (048-006), target subjected cultured cell: A-431.
[0247]FIG. 68 shows a result of a cell proliferation inhibition test. An antibody to be used: anti-HER1 antibody (048-006), target subjected cultured cell: ACHN.
[0248]FIG. 69 shows a result of a cell proliferation inhibition test. An antibody to be used: anti-HER1 antibody (048-006), target subjected cultured cell: NCI-H1373.
[0249]FIG. 70 shows a result of a cell proliferation inhibition test. An antibody to be used: anti-HER1 antibody (048-006), target subjected cultured cell: SK-OV-3.
[0250]FIG. 71 shows a result of a cell proliferation inhibition test. An antibody to be used: anti-HER2 antibody (015-126), target subjected cultured cell: BT-474.
[0251]FIG. 72 shows a result of an antitumor experiment using mouse. An antibody to be used: anti-HER1 antibody (048-006), subject transplant cell: human lung cancer cell H1373 cell.
[0252]FIG. 73 shows a result of an antitumor experiment using mouse. An antibody to be used: anti-HER1 antibody (048-006), subject transplant cell: epidermoid tumor A-431.
[0253]FIG. 74 shows a result of an antitumor experiment using mouse. An antibody to be used: anti-HER1 antibody (048-006), subject transplant cell: epidermoid tumor A-431.
[0254]FIG. 75 shows a result of an antitumor experiment using mouse. An antibody to be used: anti-HER1 antibody (059-152), subject transplant cell: epidermoid tumor A-431.
[0255]FIG. 76 is a table showing culture conditions of cell lines to be used in experiments.
[0256]FIG. 77 is a conceptual diagram of three-dimensional ELISA, showing how each mixture antibody is prepared.
[0257]FIG. 78 is a conceptual diagram of three-dimensional ELISA, showing a procedure of specifying an antibody clone.
[0258]FIG. 79 shows a result of ELISA using a plate mixed antibody (antigen is CD147).
[0259]FIG. 80 shows a result of ELISA using a row mixed antibody (antigen is CD147).
[0260]FIG. 81 shows a result of ELISA using a column mixed antibody (antigen is CD147).
[0261]FIG. 82 shows a result of ELISA using a plate mixed antibody (antigen is HER1).
[0262]FIG. 83 shows a result of ELISA using a row mixed antibody (antigen is HER1).
[0263]FIG. 84 shows a result of ELISA using a column mixed antibody (antigen is HER1).
[0264]FIG. 85 shows a result of ELISA using a selected antibody clone (antigen is HER1).
[0265]FIG. 86 shows a RNAi effect on SKOv-3 cells. A: anit-ITGA3 antibody, B: anit-ITGB1 antibody, C: 015-003 cp3 antibody. Broken line: no RNAi, solid line: ITGA3 RNAi, light-colored solid line: ITGB1 RNAi, gray: and no primary antibody.
[0266]FIG. 87 shows a correspondence between a tissue that has been diagnosed to be specific in immunostaining using a clinical cancer specimen and each antibody clone.
[0267]FIG. 88 shows a reactivity of a clinical cancer specimen and each antibody clone. + represents positive to the immunostaining; ± represents weakly positive to the immunostaining; and - represents negative to the immunostaining.
DETAILED DESCRIPTION OF THE INVENTION
Terms
[0268]For convenience, certain terms employed in the specification are collected herein.
[0269]In the specification, the terms "comprise/include" and "comprising/including" are used to include the meaning of "consisting of:" Therefore, for example, "a product (or method) comprising/including a plurality of elements (members)" necessarily includes also the terms "a product (or method) consisting of a plurality of elements (members)"
[0270]The term "disease" herein is used interchangeably with the terms meaning that some function failure occurs, for example, illness and sickness. Furthermore, unless otherwise noted, in this specification, this term is used to encompass the words meaning the condition (state) of disease such as condition, pathologic condition, symptom, and state of health. That is to say, the term "disease" is used interchangeably with the terms such as condition and pathologic condition.
[0271]The term "isolated" used herein means a state in which it is taken out from the original environment (for example, a natural environment in the case of a natural material), that is to say, means a state that is a different state from the original existing state by an artificial manipulation.
[0272]An "isolated antibody" does not include an antibody in a state in which it is natural state and no external manipulation (artificial manipulation) is given. It does not include an antibody produced in the individual body and remaining therein. An isolated antibody is typically present in a state in which other kinds of antibodies are not contaminated, that is, present singly (as an assembly of the same kinds of antibodies). In the case of an "isolated" state of the CDR region, in addition to the state which is present singly, a state which is present together with the other regions of the antibody is included. That is, the term "isolated CDR" includes not only a CDR that is present singly but also a CDR that is present as a part of an isolated antibody is included.
[0273]"HER1" is also referred to as erbB1, c-erbB-1, EGFR (Epidermal Growth Factor Receptor), or v-erbB. Originally, a gene corresponding to a cancer gene erbB found in the retrovirus that infects chicken and causes carcinogenesis (erythroleukemia) on the genome is isolated. And this gene is determined to be a receptor of EGF. By the way, EGF (Epidermal Growth Factor) as a ligand was found as a factor for promoting the cleavage of the eyelids of newly born mouse and development of an incisor in an extracted solution of the mouse submaxillary gland in 1962, and has been studied widely as cell proliferation, differentiation and survival factors. EGF is a peptide composed of 53 amino acids and has a characteristic structure including three disulfide loops formed of six cysteine residues. Thereafter, this structure has been found in a large number of proteins and is referred to as EGF-like domain. The EGF family has one or more EGF-like domains and directly binds to a receptor type tyrosine kinase EGF receptor (EGFR) family (another name: ErbB family) so as to activate this.
[0274]On the other hand, currently, four kinds of receptor ErbB families has been found and they are called EGFR (ErbB-1), ErbB-2, ErbB-3, and ErbB-4. ErbB-1 and ErbB-2 overexpress in various human tumors and are involved in the deterioration of the prognosis or survival rate. Furthermore, stimuli of these receptors are involved in cell proliferation and in turn involved in several processes related to progress, infiltration, and metastasis of tumor. To date, a phosphorylation inhibiting agent specific to EGFR have been approved as a therapeutic agent for lung cancer. They are found to highly express in many cancers. Cetuximab (ERBITUX, which is mouse/human chimeric antibody) has been developed by ImClone Systems and already marketed. ERBITUX inhibits the initial process of activation of the information transmission passage by the phosphorylation of dimerized-EGFR when it binds to a receptor of EGF as a ligand. Note here that the amino acid sequence of HER1 is shown in SEQ ID NO: 369.
[0275]"HER2" is also referred to as erbB-2, c-erbB-2, or neu. HER2 belongs to a receptor type tyrosine kinase family and its over-expression and gene amplification in the breast cancer, ovarian cancer, stomach cancer, and the like, have been reported. HER2 is a molecule that was found in 1985 when DNA containing a region of gene similar to EGFR was amplified (gene amplification) in the brain tumor and breast cancer derived from glia cells was observed. HER2 has low shedding level and is thought to be very effective as a target molecule in treating cancers. In many institutions, the monoclonal antibody (MoAb) showing effects of promoting or suppressing the tumor proliferation has been produced. MoAb showing a tumor proliferation suppressing effect is used for clinical test as a simple substance of the antibody or in combination with anti-cancer drugs such as cisplatin, and its efficacy has been reported. The EGFR family includes four kinds, but only EGFR (HER1) and HER4 have both the ligand binding sites and tyrosine phosphorylation enzymatic activity sites. HER2 does not have the ligand binding site. Instead using a ligand, HER2 has a structure that is activated from the first in terms of dimer formation ability. Incidentally, HER3 lacks the tyrosine phosphorylation activity. Therefore, HER2-HER3 hetero-dimer is a functional molecule. Genentech isolated 11 kinds of mouse monoclonal antibodies to HER2 in 1989. Among them, 4D5 was made into a humanized antibody and succeeded in developing Trastuzumab (Herceptin). Note here that the amino acid sequence of HER2 is shown in SEQ ID NO: 370.
[0276]"CD46 antigen" is an O-type sugar chain bonded non-disulfide bonded dimer protein having a molecule weight of 56 to 66 kDa, which is also referred to as MCP (Membrane Co-factor Protein), gp45-70, HuLY-m5, measles virus receptor, MIC10, TLX-B antigen, TRA2, trophoblast leucocyte common antigen, and trophoblast-lymphocyte cross-reactive antigen. This molecule binds to C3b or C4b and is known as Membrane Co-factor Protein (MCP) that is a co-factor for promoting the degradation by serine protease or I factor in plasma. It is also a receptor of the surface protein of measles virus agglutinin and Streptococcus group A. It has been reported that it is expressed in the thymus gland cells, T lymphocyte, B lymphocyte, monocyte, granulocyte, NK cells, platelet, endothelial cell, epithelium cells and fibroblast but does not express in the erythrocyte. On the assumption that only cells inducing the production of antibody to cancer specific antigen abnormally expressing in carcinogenicity and escaping from the attack of cancer tissue by complement (complement-dependent cytotoxicity, CDC) may actually grow into cancer, the expression of molecule group having an effect of inhibiting the complement has been analyzed in detail. There have been many reports about the abnormal expression of CD46 in cancer cells, however, few evidence showing that the production of antibody against antigen specific to cancer cells are induced. An amino acid sequence of CD46 antigen is shown in SEQ ID NO: 371.
[0277]"ITGA3 (integrin alpha 3)" is also referred to as alpha 3 beta 1 Epiligrin Receptor, alpha 3 beta 1 Integrin, Epiligrin Receptor, CD49c, VLA-3, Gap b3, Galactoprotein b3, or Laminin-5 Receptor in which integrin α3 chain having a molecular weight of 150 kDa and integrin β1 chain (CD29 molecule) having a molecular weight of 130 kDa are bonded to each other non-covalently to form a VLA-3 complex (α3β1 or CD49c/CD29). It is known as a receptor of laminin, collagen, fibronectin, invasion and epiligrin. Integrin is a hetero dimer molecule composed of α chain and β chain. Twenty four types of α chains and nine types of β chain form a variety of molecule groups by various combination and selective splicing. The extracellular domain binds to the extracellular matrix (for example, collagen, fibronectin, laminin). The side of cytoplasm is bonded to actin filament via talin, filanin, and α-actinin. It functions as an adhesive molecule and further functions as an important role as information transmission molecule. Above all, α3β1 molecule is associated with a tetraspanin molecule C151. Note here that the amino acid sequence of ITGA3 is shown in SEQ ID NO: 372.
[0278]The "ICAM1 (Intercellular adhesion molecule-1)" is also referred to Intercellular Adhesion Molecule 1 or CD54 Antigen and is transmembrane glycoprotein having seven binding sites of the N-bonding sugar chain. The molecular weight is 90 kDa. ICAM belongs to Ig-superfamily and is known to be mainly involved with adhesion of leukocyte. It also mediates T lymphocyte adhesion to an antigen presenting cell (APC) and is involved with the interaction between T cell and T cell or between T cell and B cell. It also involved with the adhesion to endothelial cell in which monocyte, lymphocyte, and neutrophil are activated. ICAM is bonded to integrin of LFA-1 (CD11a/CD18) and Mac-1 (CD11b/CD18). Furthermore, it also is a receptor of rhinovirus. It is expressed on various kinds of activated cells in addition to the endothelial cells. For example, it is expressed on the monocyte. On B- and T-lymphocytes, thymus gland cells, dendritic cells, endothelial cells, fibroblast, keratinocyte, chondrocyte and epithelium cells, expression is enhanced. The characteristics required to obtain during the cancerization process of epithelium cells include capability of invading into cells, and furthermore migrating and being fixed in metastasis. Therefore, it is thought that the expression of adhesion factor contributes to carcinogenesis. The adhesion factor is roughly classified into five groups, i.e., selectin (E-, P-, and L-), molecules (Ig-superfamily) having an immunoglobulin-like domain, integrin, Cadherin, and CD44. In cancerization, it is recognized that the expression of E Cadherin is suppressed. Abnormal expression in some cancer cases has been reported. Note here that the amino acid sequence of ICAM1 is shown in SEQ ID NO: 373.
[0279]"ALCAM (Activaed leukocyte cell adhesion molecule)" is transmembrane protein that is also referred to as CD166 antigen, KG-CAM, CD6 Ligand, and Neurolin. ALCAM is an immunoglobulin superfamily molecule including ten N-bonding type sugar chain added sites. ALCAM has a molecular weight of 100 to 105 kDa and is composed of five extracellular Ig-like domains and the intracellular terminus having 32 amino acid, and short transmembrane region. ALCAM is one of the adhesive molecules, is present on the activated leukocyte and is identified as a ligand molecule to CD6 molecule (which functions as a signal receptor in T cells). ALCAM also functions as an adhesion factor in homophylic (ALCAM-ALCAM) or heterophylic (ALCAM-CD6) interaction. It is suggested that ALCAM can form oligomer at intercellular adhesion site via three C2-like domains near the membrane. The distribution of ALCAM is not restricted by cell strains and ALCAM is expressed in various types of cells such as hematopoietic cells, endothelial cells, epithelium cells of the thymic cortex and thymic medulla, mesenchymal cell of the bone marrow, fibroblast, liver cells, and the like. In the peripheral blood, it is weakly expressed in activated T- and B-cells, monocyte, circulated dendritic cells, and granulocyte. Although ALCAM shows wide dispersion of tissues, the expression of ALCAM is generally limited to cell populations involved in proliferation or migration. In the thymus gland, since ALCAM is expressed in CD6+thymus gland cells, and thymus gland epithelium cells, its interaction with CD6 molecule is thought to play a role in the differentiation of T cells. In addition, it is suggested that ALCAM adhesive molecules are involved in the fetal blood formation, differentiation of angioblastic cells, and capillary angiogenesis. The roles of ALCAM in cancerization is variously assumed (e.g., controlling of MMP activation, causing internalization and recycling, functioning as a substrate of ADM17 and ADAM10 (abbreviation of a disintegrin and metalloprotease), protecting from apoptosis and autophasy), however, no decisive roles have not reported. The interaction of ALCAM-CD6 is thought to be carried out in the both direction. The amino acid sequence of ALCAM is shown in SEQ ID NO: 374.
[0280]"CD147 antigen" is membrane glycoprotein belonging to an immunoglobulin superfamily and is also referred to as BSG, TCSF (Tumor cell-derived collagenase stimulatory factor), 5F7 protein, OK blood group protein, basigin protein, collagenase stimulatory factor protein, EMMPRIN (Extracellular matrix metalloproteinase Inducer), M6 activation antigen, human leukocyte activation antigen M6, or the like. D147 antigen has two aspects. One is observed when it functions on the cell surface, it exhibits the activation of MMP-1, 2, 3 (matrix metalloprotease) and the lectin activity recognizing oligomannose as membrane glycoprotein having two Ig domains. The activation of MMP receives much attention in cancers (which is also known as EMMPRIN in Europe and America). That is to say, CD147 antigen expressing in cancer cells activates MMP expressing in the surrounding fibroblast and contributes to the infiltration of cancers. On the other hand, the activation of oligomannose lectin is especially important in the interaction of nerve cells and indicated to have a relationship with respect to neurite outgrowth. The second aspect is a function in cells. CD147 antigen forms a homo dimer. It is reported that this formation needs N-terminal Ig domain and does not need addition of sugar chain. CD147 has the following interesting reports: integrin α3β1 and CD147 form a complex, and in this case, TM4SF (tetraspanin) molecule does not join the complex. In cancerization, the production of D147 changes anchorage-dependent growth to independent growth, which is promoted by the production of hyaluronic acid (hyaluronam). It is interesting that the receptor of hyaluronic acid includes CD44 and RHAMM. CD147 induces the production of MMP, and a part of CD147 is solubilized due to the effect of the MMP. CD147 acts on integrin so as to change the structure of cells. CD147 affects the angiogenesis. Furthermore, mass expression-cell proliferation of CD147 and Cyclophin A has been found.
[0281]The amino acid sequence of the CD147 antigen is shown in SEQ ID NO: 375.
[0282]"IgSF4" is an abbreviation of immunogloblin superfamily member 4 and is also referred to as BL2, ST17, NECL2, TSLC1, IGSF4A, SYNCAM, and sTSLC-1. IgSF4 has homology of NCAM (neural cell adhesion molecule) and amino acid sequence. IgSF4 is thought to be expressed from human 11-chromosome, 11q23.2. It has been reported that IgSF4 expressed as a suppression gene in a lung cancer specific manner and that IgSF4 is involved in the nerve adhesion in the brain (Biederer T et al. Science. 2002 Aug. 30; 297 (5586): 1525-31). The sequence information of IgSF4 is recorded in a NCBI-PUBMED database (Accession No. NM--01433, Definition: Homo sapiens immunoglobulin superfamily, member 4 (IGSF4), mRNA). As to the relationship with respect to the carcinogenesis, as shown by the name TSLC1 (tumor suppressor in lung cancer 1), it receives attention as a tumor suppressor gene. However, IgSF4 shows high expression in 100% adult T cell leukemia (ATL) cells and it is suggested that IgSF4 may work as oncogene. The amino acid sequence of IgSF4 is shown in SEQ ID NO: 376.
[0283]"C1qR" is a complement receptor encoding a type I membrane protein. This protein functions as a receptor for complement protein C1q, mannose binding lictin, and lung surfactant protein A. Two or more polypeptides of 70 kDa are bonded by disulfide bonding so as to form C1qR. Removing an immune complex is an important function of the complement and the C1q receptor is a functional receptor that is bonded to a collagen portion of C1q thereby linking the immune complex to phagocyte. It is suggested that C1qR forms complex with CD43. The amino acid sequence of C1 qR is shown in SEQ ID NO: 446.
[0284]"CD44" is a transmembrane protein belonging to a hyaladherin family, which is cell surface glycoprotein related to cellular interaction, cell adhesion and cell migration. It is a hyaluronic acid receptor. It is thought that a wide variety of the structural and functional isoforms of proteins by the selective splicing or post-translation modification of this molecule may be involved in tumor metastasis. The CD44 molecule is expressed in almost all the cells and tissues. However, in general, it is not expressed in the platelet, liver cell, cardiac muscle, uridiferous tubule epithelium, testis, and skin. The amino acid sequence of CD44 is shown in SEQ ID NO: 447.
[0285]"CD73" is also referred to as 5-prime-ribonucleotide phosphohydrolase and transforms purine 5-prime mononucleotides into nucleosides at the neutral pH. The enzyme mediates glycosylphosphatidyl inositol to the surface of the outside of the plasma membrane and is bonded to the surface of the outside of the plasma membrane. CD73 is a homodimer composed of two 70 kDA subunits. CD73 is used as a marker of the lymphocyte differentiation. It has been known that the deletion of this gene is related to various immune defective diseases. The amino acid sequence of CD73 is shown in SEQ ID NO: 448.
[0286]"EpCAM" has 22 or more names as to only the number of names used and cited several times in research paper. This antigen exists on genome 2p21. This antigen is a protein having a full length of 314aa, and 34920 Da. In the documents in which this molecule is examined at the mRNA level, it is detected in healthy human individuals, 100% in the peripheral blood (PB) level and 40% in the bone marrow (BM) level. It has been reported that it can be detected in large intestine but cannot detected in the liver, prostate, and lung. In cancer cell line, in the relationship with respect to p53, the methylation of EpCAM is lost due to the mutation or deletion of p53 and the amplification is induced. The amino acid sequence of EpCAM is shown in SEQ ID NO: 449.
[0287]The first Met gene discovered as a search product of oncogene using NIH3T3 gene is HGFR (Hepatocyte growth factor receptor). HGF is also referred to as a scatter factor and is utterly independently isolated as a molecule having an extremely different apparent function. Similar to HER1 and PDGF, HGFR is a receptor having a ligand binding domain outside the cells and has a tyrosine phosphorylation enzymatic activity site at the cytoplasm side, however, the function is extremely different. In general, when the cell proliferation factor or a differentiation induction factor is bonded to a receptor so as to cause the phosphorylation of protein, it finally activates the transcription factor and expresses a certain gene set by way of some of the limited information transmission pathway (Ras/MAP kinase pathway, and the like). In this case, the type of the cell response is finally determined by transcription factor. Thus, when the cancerization may activate some of the proliferation factors-receptor, it is thought that changes other than cancerization are not likely to occur in the cells. Currently, as to the cancerization, the phenomenon called epithelial-mesenchymal transition (EMT) receives much attention and the factor plays a core role in the phenomenon. In such examples, since a large number of molecules cooperatively function, detail analysis is needed. The amino acid sequence of HGFR is shown in SEQ ID NO: 450.
[0288]LAR (Leukocyte common Antigen-Related) belongs to a PTP (protein tyrosine phosphatase) family. The PTPs are known to be molecules to modulate the process in the various aspects of the cancerization, division cycle, differentiation, cell growth, and the like. The structure thereof includes an extracellular region, mono-transmembrane region, and two tandem catalyzing domain in the cytoplasm (homolog of protein tyrosine phosphatase). The extracellular region has a structure similar to nerve cell adhesion factor, which includes three Ig-like domains and nine non-Ig like domains (homolog of NCAM). The function of this molecule is involved in the cell adhesion in the formation adherents junctions in the epithelium. Note here that it is confirmed that this molecule is highly expressed in insulin sensitive mast cells, and insulin resistant cells. Therefore, it is suggested that it is related to insulin. Furthermore, it is reported that anti-LAR antibody has an insulin receptor inhibitory activity of the insulin receptor forced expressing body (Knock-down of LAR protein tyrosine phosphatase induces insulin resistance: Mander A, Hodgkinson C P, Sale G J.: FEBS Lett. 2005 Jun. 6; 579 (14): 3024-8.).
[0289]Furthermore, LAR is expressed on the membrane of all the leukocytes and is referred to as protein tyrosine phosphatase receptor type F (PTPRF) and protein sequence (SEQ ID NO: 941) thereof is registered as TDHULK in Protein sequence database of the Protein Information Resource (PIR).
[0290]BCAM (basal cell adhesion molecule) (Lutheran blood group) is referred to as CD239 antigen and its protein sequence is registered as Q86VC7 (UniProtKB/Swiss-Prot) and 13800 (PIR) (SEQ ID NO: 942). It produces a selective splicing product from a single gene in the chromosome 19q13.2-q13.3. It is a glycoprotein having an immunoglobulin-like domain. It is a mono-transmembrane type and expressed widely. Its expression in the pancreas is high and its expression in the brain is low. The BCAM antigen is modulated excessively in certain cells, thus inducing the malignant alteration of cancers. Also, it is shown that it is overexpressed in the living body with ovarian cancers.
[0291]In the present invention, "liver cancer" is intended to be widely interpreted and it includes liver carcinoma and liver sarcoma. Furthermore, the term "cancer" in the present invention is interchangeably with "tumor." Furthermore, in the stages before the pathological diagnosis is not established, that is, before whether the tumor is benign or malignant has not been determined, the term may include benign tumor, benign-malignant borderline lesion, and malignant tumor collectively.
[0292]Cancers are called under the name of the organs in which the cancers are developed or the name of development body tissue. Main examples include tongue cancer, gingival cancer, pharynx cancer, maxillary cancer, laryngeal cancer, salivary gland cancer, esophageal cancer, stomach cancer, small intestinal cancer, large bowel cancer, rectum cancer, liver cancer, biliary tract cancer, gallbladder cancer, pancreas cancer, lung cancer, breast cancer, thyroid gland cancer, adrenal gland cancer, hypophyseal tumor, pinealoma, uterine cancer, ovarian cancer, vaginal cancer, urinary bladder cancer, kidney cancer, prostate cancer, urethral cancer, retinoblastoma, conjunctival cancer, gliocystoma, glioblastoma, skin cancer, leukemia, malignant lymphoma, testicular tumor, osteosarcoma, rhabdomyoblastoma, leiomyosarcoma, blood vessel sarcoma, liposarcoma, chondrosarcoma, Ewing's sarcoma, and the like. Furthermore, depending upon the characteristics of the sites of the organs of development, cancers are subclassified into, for example, upper, middle, and lower pharynx cancers, upper, middle, and lower esophageal cancers, gastric cardia cancer, gastropyloric cancer, cervical cancer, cancer of uterine body, and the like. These cancers are included in the "cancers" of the present invention but the cancers are not limited to these alone.
[0293]In the specification, if necessary, the following abbreviations (in parentheses) are used according to the practice.
[0294]Heavy chain (H chain), light chain (L chain), heavy chain variable region (VH), light chain variable region (VL), complementarity determining region (CDR), first complementarity determining region (CDR1), second complementarity determining region (CDR2), third complementarity determining region (CDR3), first complementarity determining region of heavy chain (VH CDR1), second complementarity determining region of heavy chain (VH CDR2), third complementarity determining region of heavy chain (VH CDR3), first complementarity determining region of light chain (VL CDR1), second complementarity determining region of light chain (VL CDR2), third complementarity determining region of light chain (VL CDR3)
[0295]The first aspect of the present invention relates to a method of classifying antibody. The classifying method of the present invention includes the following steps.
[0296](1) preparing a plurality of antibodies recognizing cell surface antigen;
[0297](2) bringing each of the antibodies into contact with cells of the same kinds;
[0298](3) analyzing each cell after step (2) by flow cytometry so as to obtain data showing reactivity between the antibody and the cell surface; and
[0299](4) comparing the obtained data and classifying antibodies based on the similarity of the data.
Step (1)
[0300]In the classifying method of the present invention, firstly, a plurality of antibodies recognizing cell surface antigen are prepared. For convenience of explanation, the antibody classified by the classifying method of the present invention is also referred to as a "sample antibody."
[0301]In the present invention, the "cell surface antigen" is a molecule in which at least a part thereof exists outside the cell and which forms an antigenic determinant on the surface of the cell. For example, protein such as transmembrane type protein having a cell membrane transmembrane domain and an extracellular domain and GPI anchor type protein, which are linked to cell membrane via glycolipid and the like and existing on the surface of the extracellular surface, can form such an antigenic determinant. The cell surface antigen can be formed by a simple protein (basically, constituent includes only amino acids), a conjugated protein (constituent other than amino acid are contained. For example, glycoprotein and lipoprotein), or a modified protein (a protein modified by, for example, phosphorylation, acetylation, and methylation), and the like. Furthermore, two or more same types or different types of molecules may cooperatively form an antigen determinant.
[0302]The "cell surface antigen" of the present invention is not particularly limited to animal cells and may include cell surface antigens of plant cells, microorganism cells, and the like. Preferably, "cell surface antigen" of the present invention is the cell surface antigen of animal cells. It is known that the animal cells have various cell surface antigens. The "animal cells" herein include mammalian cells and non-mammalian cells, but preferably mammalian cells. Above all, human cells are preferable.
[0303]Preferably, a plurality of antibodies recognizing the intact cell surface antigen are prepared. The "intact state" means that the original state is maintained. It has the same meaning that "not denatured state."
[0304]The "antibody recognizing cell surface antigen" represents an antibody recognizing and binding the cell surface antigen with highly specific recognition mechanism between the antigen and the antibody. The origins, types, classes, forms and the like, of antibodies are not particularly limited. Therefore, the "antibody" in the present invention includes an antibody of non-human animals such as mouse and rat, a chimeric antibody in which a part of the region is substituted with that of other animal (including human), a humanized antibody, and human antibody. Preferably, human antibody or human type antibody (humanized antibody) are used. Antibody fragments such as Fab, Fab', F(ab')2, scFv, and dsFv antibody may be used. An antibody for treatment application includes an antibody in which VH and VL (Fv region) are converted into IgG type is included.
[0305]An antibody recognizing a cell surface antigen can be prepared by, for example, bringing an antibody library into contact with the cell surface antigens and recovering the antibodies bound to the cell surface antigens. One of such preparation methods is a method reported by the present inventors before (Japanese Patent Unexamined Publication No. 2005-185281). This method makes it possible to select an antibody clone recognizing intact cell surface antigen from the phage antibody library. The present invention can preferably use the antibody assembly derived from each antibody clone. The "assembly derived from each antibody clone" herein includes the selected antibody clone itself, or the product prepared by using the gene. The latter example includes an antibody in which genes of the selected antibody clone is transformed by an appropriate host (for example, E. coli) and the host is expressed, or an antibody to which further genetic engineering modification is added in the host or by the use of the host and then the modified antibody is expressed.
[0306]The above-mentioned publication discloses as the antibody having a human Fv region, scFv-CL-cp3 antibody (an antibody in which a phage protein cpIII is fused to scFV via the light chain constant region), scFv-CL-pp antibody (an: antibody in which two proteins A are fused to scFV via the light chain constant region), scFv-CL-pp-Avi antibody (an antibody in which avidin is fused to scFv-CL-pp antibody), scFv-CL-Avi antibody (an antibody in which avidin is fused to scFV via the light chain constant region), scFv-CL-pp-Avi or antibody obtained by biotining scFv-CL-Avi antibody (an antibody in which biotin is bonded to an avidin part), and the like. The present invention can preferably use any of these types of antibodies. These antibodies having a human Fv region are very useful in providing an antibody for treatment (production of an antibody for treatment can be proceeded advantageously).
[0307]Note here that the contents disclosed Japanese Patent Unexamined Publication No. 2005-185281 are herein incorporated by reference in its entity.
[0308]A combination of separately prepared antibodies may be used as the "plurality of antibodies recognizing cell surface antigen" in the present invention. In this case, the preparation method of each antibody may be the same as or different from each other.
[0309]An antibody in which a label material has been bound or fused in advance (which is collectively referred to as "labeled sample antibody") may be used. The former example can include an antibody labeled with fluorescence pigment. The latter example can include an antibody in which fluorescence proteins (fluorescence protein fused antibody) such as GFP (Green Fluorescent Protein) and RFP (Red Fluorescent Protein) have been fused. Such fluorescence protein fused antibody can be prepared easily by using genetic engineering technique.
Step (2)
[0310]Next, the sample antibodies are brought into contact with cells of the same kinds, respectively. That is to say, cells to be used are determined, and then the cells are brought into contact with the sample antibody for each sample antibody. The sample antibody recognizing the surface antigen of the cells to be used binds to the cell surface. The binding amount of the sample antibody is dependent upon the expression amount of the cell surface antigen recognized by the antibody.
[0311]Cells that are brought into contact with the sample antibody are not particularly limited and may be arbitrarily selected from animal cells, plant cells, microorganism cells, and the like. For example, in one preferable embodiment, cells derived from a patent having a certain disease (or having a certain pathologic condition) are used. The "certain disease" includes various kinds of cancers, for example. The tissues or organs from which the cells are derived are not particularly limited. An example of the certain disease include kidney cancer, hepatic cell carcinoma, gallbladder and liver cancer, alveolar cell carcinoma, lung squamous cell cancer, pulmonary adenocarcinoma, pancreas cancer, adenocarcinoma, ovarian cancer, and the like.
[0312]Cells forming a highly uniform cell population are preferably used. It is preferable because such cells can provide easier or simpler data, facilitates the comparison of data and provides more reliable comparison results in the below-mentioned flow cytometry analysis. The typical example of such cells is established cell line (cell line). Preferable examples include established cancer cell line such as liver cancer cell line HepG2, undifferentiated liver cancer cell line HLF, liver cancer cell line OCTH, intrahepatic bile duct cancer cell line RBE, pancreatic cancer cell line PANC-1, pancreas cancer cell line MIA-Paca2, kidney cancer cell line CCFRC1, kidney cancer cell line Caki-1, kidney cancer cell line ACHN, kidney cancer cell line 293T, ovarian cancer cell line KF28, ovarian cancer cell line SKOv3, ovarian cancer cell line KF-28, ovarian cancer cell line RMG-1, ovarian cancer cell line RMG-2, breast cancer cell line BT474, vulvar mucosa epithelium cell line A431, stomach cancer cell line SNU-5, stomach cancer cell line MKN45, stomach cancer cell line NCI-N87, cancer cell line RERF-LC-AI, pulmonary adenocarcinoma cell line PCI4, lung cancer cell line NCI-H441, lung squamous cell canceEBC1, pulmonary adenocarcinoma cell line H1373, pulmonary adenocarcinoma cell line A549, pulmonary adenocarcinoma cell line Calu-3, pulmonary adenocarcinoma cell line PC14, large bowel cancer cell line CaCo2, large bowel cancer cell line CW2, hamster ovarian cancer cell line CHO, and the like. Note here that cells whose uniformity is improved by culture operation is one of the most preferable cells.
[0313]Each sample antibody is brought into contact with cells in an appropriate solution. At this time, it is preferable that the conditions are set so that the properties of the sample antibody are not affected and cells are not damaged. For example, cells and the sample antibodies are co-existed in the culture solution suitable for the existence and proliferation of the cells, in the phosphoric acid buffer and citric acid buffer, in physiologic saline, or in a solution in which BSA for suppressing non-specific adsorption is added, at room temperature to low temperatures (for example, 0° C. to 25° C., preferably 4° C. to 15° C.), for 20 minutes to 3 hours. During this time, the solution may be stirred.
[0314]The conditions under which each sample antibody and cells are brought into contact with each other are made to be uniform in order to obtain highly reliable data.
[0315]After contacting operation mentioned above, labeling is carried out if necessary (other than the case when a labeled sample antibody is used). The "labeling" herein denotes labeling the sample antibody bound to the surface of the cells. For example, labeling can be carried out by reacting (contacting) an antibody having a specific binding ability to the sample antibody to which a label material has been bound (antibody to be detected) with cells after the contacting operation. Instead of directly binding an antibody to be detected to the sample antibody, other antibodies and the like may be interposed therebetween. Thus, various labeling techniques can be employed and a person skilled in the art can select an appropriate technique. In the flow cytometry analysis, in general, fluorescent dye is used as a label material. Fluorescent dye such as Alexa488, AMCA, Cascade Blue (registered trademark), FITC, PerCPTM, CyTM3, Texas Red (registered trademark), CyTM5, APC, TRITC, and the like, can be used.
Step (3)
[0316]Subsequently, cells after subjecting to the step (2) are analyzed by flow cytometry so as to obtain data showing the reactivity between the antibody and the cell surface. That is to say, cells after subjecting the contacting operation to the sample antibody are subjected to the flow cytometry analysis, and the binding property to the sample antibody is examined. Preferably, as the data showing the "reactivity" herein, histogram showing the relationship between the antibody binding amount and the number of cells is used. That is to say, one-parameter histogram in which the antibody binding amount is used as a parameter is used. The one-parameter histogram is one display method in the flow cytometry. The one-parameter histogram is generally shown in a graph in which X-axis represents one indicator (parameter) and Y-axis represents the number of cells. For the device used for the flow cytometry analysis, for example, devices from BECKMAN COULTER, Japan Becton, Dickinson and Company, and the like can be used in the present invention. The operation may be carried out according to the basic operation and analysis conditions attached to the device. Furthermore, many research paper and documents about the flow cytometry analysis are published. See, for example, Cao T M, et al. Cancer. 2001 Jun. 15; 91 (12): 2205-13., Storek K J, et al. Blood 97: 3380-3389, WEIR'S HANDBOOK OF EXPERIMENTAL IMMUNOLOGY Vol. II <Blackwell Science>, Little MT and R. Storb Nature Reviews Cancer 2002 2: 231-238.
[0317]Typical procedure of the flow cytometry analysis is described below. The sample antibody and cells are reacted with each other, then reacted with antibody to be detected labeled with fluorescent dye, so that cells are labeled with fluorescence. The amount of sample antibody to be bound varies depending upon the amount of antigen existing on the surface of the cells. As a result, the amount of fluorescent label of the cells becomes different. Therefore, by measuring the fluorescence intensity, the affinity between the antigen existing on the surface of the cell and the ample antibody and the amount of antigen can be estimated. In general, prior to the detection of the fluorescence intensity, forward scatter light (FSC) and side scatter light (SSC) are measured and gated, so that the fluorescence intensity of only the target cell population is measured. Specifically, for example, the forward scatter light and the side scatter light are shown in X-axis and Y-axis, respectively. The cell population (when established cell lines or cultured cells are used, the cell population becomes extremely uniform) that are assumed to be living cells from the data obtained by dot plot expansion are gated, and the fluorescence intensity within the gate is measured. The measurement result is shown in a form of, for example, histogram. Note here that the terms related to the histogram obtained in the flow cytometry analysis are mentioned below.
[0318]The "number of samples" denotes number of data and generally represented by n. The "total" denotes a total of data and generally represented by T. "Mean value" denotes an average of data and is calculated by dividing the total by the number of samples. The mean value is susceptible to abnormal data. The "median value" is a value located in the middle when the data are aligned in ascending numeric order. When the number of data is odd number, the average of two middle values is defined as a median value. The median value is less susceptible to abnormal data as compared with the mean value and shows the characteristics of the population more accurately. The "mode" denotes a value whose frequency is maximum in the data. In the case of the flow cytometry analysis, the mode is the same as a peak value. The mode is less susceptible to abnormal data as compared with the mean value. The "maximum value" is a maximum value of data and generally represented by Max.
[0319]The "range value" is difference between the maximum value and the minimum value and generally called range and referred to as R. The "dispersion" is a value showing the degree of variation of data. The larger the dispersion is, the larger the variation is. In general, it is referred to as V. The dispersion is obtained by dividing the sum of squares deviation by the number of samples (in the case of sample survey, divided by (number of samples -1)). The "standard deviation" denotes square root of the dispersion and is generally referred to as u. The "coefficient of variation" is a value obtained by dividing the standard deviation by an average value and is generally referred to as CV. Since the standard deviation does not clearly shows the degree of variation of data, the standard deviation is normalized by dividing it by the average value. In the flow cytometry analysis, it is frequently used as a value showing the resolving power of the device. The "kurtosis" is one of the indicators representing the distribution in the population and generally is referred to as H. The distribution in which the kurtosis is 0 is defined as normal distribution. When the kurtosis is larger than 0, the distribution has sharper apex than the normal distribution. When the kurtosis is smaller than 0, the distribution becomes more flatness than the normal distribution. The "skewness" denotes a value showing the left-right symmetry of the population and generally is referred to as G. When the skewness is 0, distribution becomes left-right symmetric. When the skewness is larger than 0, the distribution distorts in the right direction. When the skewness is smaller than 0, the distribution distorts in the left direction.
Step (4)
[0320]Next, the obtained data are compared and sample antibodies are classified based on the similarity of the obtained data. Herein, "based on the similarity" means that the similarly of data are used as a criterion of classification. An example of criterion (classification criterion) based on the similarity of data is shown below.
[0321](a) A plurality of antibodies having the identical or highly similar data are classified into one antibody group. Specifically, for example, plurality of antibodies having extremely similar histogram is defined as one group when the shape of the histogram showing the distribution of cells is determined by the kurtosis, skewness and the like.
[0322](b) An antibody providing specific data forms one antibody group by itself.
[0323](c) An antibody having a low reactivity with respect to the antigen is excluded
[0324](the antibody does not belong to any groups).
[0325]In the present invention, each antibody is classified by one or two or more criteria selected from the above-mentioned classification criteria (a) to (c).
[0326]The similarity of data can be determined based on the parameter specifying the data. However, the specific determination method is dependent upon the types of data. In the case where data are represented by numeric values, it is possible to determine the similarity based on the degree of similarity of numeric values (for example, when 1, 2, and 5 are given as data, it is determined that the similarity between 1 and 2 has high similarity).
[0327]Furthermore, when a histogram is given as data, it is possible to determine the similarity of data based on the shape of the histogram. As a result of the investigation by the present inventors, it is determined that the shape of the histogram in the flow cytometry analysis is highly dependent upon the kinds of the antigen. In other words, when the antigens to be recognized are the same, regardless of the kinds of antibodies, it is determined that the histogram having an identity or high similarity can be obtained. Base on this fact, in one embodiment of the present invention, by comparing the shapes of the histogram showing the results of the flow cytometry analysis, the similarity of data is determined. Specifically, the similarity of data can be determined by comparison by visual observation or by comparison of one or two or more of parameters specifying the histogram. The parameters herein can employ one or more values selected from the group consisting of median value, mode, maximum value, range, standard deviation, kurtosis, and skewness of the histogram. Preferably, determination is carried out in terms of two or more values, furthermore preferably three or more values, and yet furthermore preferably four or more values. By increasing parameters to be used in determination, the determination accuracy can be improved. Among these parameters, it is said to be advantageous that the median value, mode, or kurtosis that are parameters deeply related to the shapes of the histogram are employed for carrying out the determination at high accuracy. Preferably, a combination of two or more of these parameters is used. Specifically, for example, the similarity of the histogram may be determined based on the median value, mode, and kurtosis.
[0328]When two data to be compared have similar values in terms of employed parameters, the similarity between the two data is determined to be high. When the difference between two values (100×(A-B)/A (%) when the two values are A, B (A≧B)) is within 10%, preferably within 5%, and furthermore preferably within 3%, the two values are determined to be similar.
[0329]In one embodiment of the present invention, when or after the sample antibodies are classified, sample antibodies having a low reactivity to the cell surface antigen are removed. Thereby, an antibody group including highly useful sample antibodies can be formed. The degree of the reactivity of the antibody can be determined by using the results of the flow cytometry analysis. Specifically, the mode (peak value) of the histogram obtained with respect to the sample antibody to be determined and the mode (that is to say, the maximum mode in the group) of the histogram obtained with respect to the sample antibody having the maximum reactivity in the antibody group to which the sample antibody belongs. As a result, when the former is 1/2 or less of the latter, preferably 1/5 or less, furthermore preferably 1/10, it is determined that the sample antibody to be determined has low reactivity.
[0330]In one embodiment of the present invention, the reactivity of each sample antibody is examined in two or more kinds of cells and the sample antibodies are classified by using the results. That is to say, two or more kinds of cells are prepared and by using the prepared cells, steps (2) to (4) are carried out.
[0331]The expression amount, distribution, and the like of the cell surface antigens are dependent upon the kinds of cells. Therefore, two antibodies having high similarity in data obtained by using certain cells, that is, two antibodies having the common antigens should provide data having high similarity when the other cells are used. Thus, when the two antibodies to be compared provide data with high similarity with respect to more than two kinds of cells, the probability that the antibodies have the common antigens is extremely high. Furthermore, when such results are obtained, it can be easily determined that the two antibodies have the common antigens. Thus, the use of two kinds or more cells can make it accurate and easy to determine the identity of antigens.
[0332]In one preferable embodiment of the present invention, sample antibodies having identical or highly similar data with respect to at least two kinds of cells are classified into one antibody group.
[0333]Furthermore, by observing the classification results of the case where two or more kinds of cells are used, kinds or amount of antigens to be expressed can be compared between the cells. Therefore, more useful information can be provided in studying the properties of these cells.
[0334]In one embodiment of the present invention, a classification result is displayed as a panel. The "panel" in this specification is a product in which a plurality of elements (for example, antigen, antibody, antibody group, cell, name of disease, name of pathologic condition), are displayed in the form of tables or drawings, in which the elements are associated with each other, on media such as a display and paper. Each element is represented by general name, abbreviation, alias, or symbol or code representing thereof, and the like. The panel of the present invention shows the relationship with respect to two kinds or more of elements.
[0335]The term "associating to" in the present invention means that two or more elements are linked. Therefore, in the tabular format panel showing the association between an antigen and an antibody group, for example, both elements are displayed in adjacent to each other, or both elements are displayed in the same cells, or both elements are linked by a line or something, so that it can be understood that the both elements form a pair.
[0336]In the panel herein, typically, antibody groups are displayed in a way in which they are associated with each other for each antigen (or for each antigen having high association) expressed by the cells that have been subjected to the flow cytometry analysis. Therefore, this panel makes it possible to access antibodies useful for studying surface antigens of the cells. Thus, the panel itself of the present invention has a great value. A panel formed by using two kinds or more of cells makes it possible to understand the presence, expression amount, and the like, of antigens expressing between cells. Such a panel has further higher values.
[0337]In the panel of the present invention, antibodies may be arranged regularly in accordance with the reactivity to antigens. Thus, the difference in the reactivity between antibodies can be made obvious.
[0338]According to the classifying method in the present invention, a plurality of antibodies recognizing the same antigen (or antigens having high association) are associated with each other. In other words, for each antigen (or for each antigen having high association), antibody assembly (antibody group) recognizing the antigen can be obtained. These antibody groups are useful for studying cell surface antigen and have high usability. Furthermore, according to the classifying method of the present invention, a large number of antibodies can be classified rapidly for each antigen (or for each antigen having high association). That is to say, the classifying method of the present invention is useful for classification of a large number of antibodies and allows comprehensive classification of antibodies. The term "highly associated" or "having high association" used for antigen means that two or more antigens have a close association in a living body, for example, the antigens are not the same molecules but exhibit one function cooperatively (for example, two antigens are bound so as to form one complex functionally).
[0339]According to the classifying method of the present invention, typically, plurality of antibodies are associated with each other for each antigen (or for each antigen having high association). Therefore, in studying certain antigens, a plurality of antibodies can be used or suitable antibodies can be selectively used if necessary, which leads to better results or significant findings and means that studying can be proceeded advantageously.
[0340]On the other hand, by executing the classifying method of the present invention, it is possible to understand the expression amount of distribution of cell surface antigens (antigen are unknown) in certain cells (that is, cells that are brought into contact with the sample antibody). Thus, the classifying method of the present invention provides useful information on the properties of the certain cells and is useful for studying the cells (in particular, the surface antigens).
[0341]Note here that when antigens to all the sample antibodies are unknown, antigens to which each antibody group is associated are not identified. On the other hand, when some identified antigens are contained in a part of the sample antibodies, an antigen to which the antibody group containing the antibody becomes an identified antigen. Thus, it is also possible to associate an antibody group with the identified antigen.
[0342]According to the above-mentioned classifying method, antibodies are classified based on the reactivity between the antigens and certain cell surfaces and the antibody groups are formed. Therefore, antibodies belonging to the same antibody group have the same (or highly similar) reactivity to the surface of cells used for classification. However, it is not necessarily ensured that all the antibodies belonging to the same antibody group can recognize the same antigens. Even if the recognizing antigen is the same, the reactivity to cells expressing antigens on the cell surface may be different. Furthermore, the opposite case may occur (even if the recognizing antigen is different, the reactivity to cells expressing antigens on the cell surface may be the same, for example, one of the complex may be recognized).
[0343]Therefore, in order to form an antibody for each recognizing antigen, one embodiment of the present invention carries out the following steps (i) to (vi) after the step (4).
[0344](i) associating the classified antibodies with a combination of n pieces of parameters including a first parameter, a second parameter, . . . , and an n-th parameter (wherein, n represents an integer of 2 or more, each parameter has two or more parameter values and the same parameter value is given to two or more antibodies in each parameter);
[0345](ii) with respect to each parameter, preparing an antibody mixture of the antibodies having the same parameter value;
[0346](iii) examining a reactivity of each of the antibody mixtures with a target antigen by an enzyme linked immunosorbent assay (ELISA) so as to specify the antibody mixture which shows reactivity;
[0347](iv) specifying a combination of a parameter name and a parameter value that are common to the antibody group contained in the specified antibody mixture;
[0348](v) selecting an antibody corresponding to the combination specified in the step (iv) in terms of all parameters among the antibodies subjected to step (i); and
[0349](vi) classifying the selected antibodies into one antibody group.
[0350]According to the classifying method of this embodiment, an antibody group can be formed for each antigen to be recognized. That is to say, antibody groups having various individualities recognizing the same antigen can be obtained. Furthermore, the combination of the plurality of parameters is associated with each antigen and then an antibody mixture is prepared according to a predetermined regulation. Then, based on the results of ELISA (Enzyme-Linked immunosorbent assay) using the antibody mixture, an antibody recognizing a target antigen is determined. By this unique technique, antibodies can be classified rapidly and efficiently. Furthermore, at the same time when the antibodies are classified, as to at least a part of the antibodies, an antigen is identified. That is to say, the classifying method of this embodiment is a method of rapidly and efficiently obtaining an antibody whose antigen has been identified, which dramatically promote the increase in the number of antibodies whose antigens have been identified. On the other hand, the classification results show the presence form or expression from on the cell surface used in flow cytometry analysis, which provides extremely useful information for study and development of the application of antibody (for example, treatment of cancer). Furthermore, when the presence of a certain antigen is clarified based on the classification results, it is possible to obtain an unknown antigen (for example, complex counterpart) that is thought to be possible to exist in a form of a complex with the antigen. That is to say, the classifying method of this embodiment efficiently functions as determining a novel antigen or novel molecule complex. Hereinafter, each step is described in detail. For convenience of explanation, the classifying method of this embodiment is also referred to as "n dimensional ELISA method."
Step (i)
[0351]In this step, a combination of n pieces of parameters consisting of the first parameter, the second parameter, . . . , and the n-th parameter are associated with antibodies classified by the preceding steps (steps (1) to (4)). Thus, each antibody has n-dimensional address (a parameter value of the first parameter, a parameter value of the second parameter, . . . , and a parameter value of the n-th parameter).
[0352]In general, association is carried out with respect to all the antibodies that have been classified in the preceding steps, although the association is not limited to this. That is to say, the association may be carried out only a part of the antibodies that has been classified in the preceding steps. In this case, a part of antibodies are excluded from the antibodies to be classified.
[0353]Herein, "n" is an integer of two or more. That is to say, to each antibody, two or more combinations of parameters are associated. The number of "n" does not have an upper limit. When the number of "n" is too large, operations in the subsequent steps (for example, preparation of an antibody mixture, specification of an antibody mixture showing the reactivity) may be excessively complicated. Therefore, "n" is preferably three to five.
[0354]On the other hand, each parameter is made to have two or more parameter values and the same parameter values of each parameter are made to be provided to two or more kinds of antibodies. Specifically, parameter values of the first parameter may be 1, 2, 3 and 4, and each parameter value is provided to five kinds of antibodies, respectively. The number of the parameter values is set for each parameter. Furthermore, similar to the number of parameters, the number of the parameter values does not have an upper limit. In order to make the analysis in the following steps (iv) and (v) be efficient and improve the accuracy thereof, it is preferable that the kinds of antibodies contained in each antibody mixture are not excessively large number. Therefore, each parameter value may be set so that the kinds of antibodies contained in each antibody mixture is preferably 200 or less, and furthermore preferably, 100 or less. Specifically, for example, the number of the parameter values can be set to between 2 and 100. Note here that the kind of antibodies contained in each antibody mixture is dependent upon the setting of the parameter, and may not be equal between antibody mixtures.
Step (ii)
[0355]In this step, an antibody mixture, in which antibodies having the same parameter value are mixed, is prepared. The antibody mixture is prepared for each parameter. For example, when the values of the first parameter is 1, 2, 3 and 4, an antibody mixture mixing antibodies to which 1 is given as the first parameter, an antibody mixture mixing antibodies to which 2 is given as the first parameter, an antibody mixture mixing antibodies to which 3 is given as the first parameter, and an antibody mixture mixing antibodies to which 4 is given as the first parameter are prepared. By the same procedure, as to the remaining parameters, antibody mixtures are prepared. Thus, antibody mixtures in the same number as the total number of the number of the first parameter, the number of the second parameter, and the number of the n-th parameter are prepared.
[0356]In general, an antibody mixture, in which all antibodies having the same parameter values are mixed, are prepared although the antibody mixture is not limited to this. An antibody mixture may be prepared by selecting a part of all antibodies having the same parameter values and mixing thereof. Thus, the selection of antibodies may be carried out in this stage.
[0357]It is preferable that an antibody mixture is prepared so that all antibodies are contained in equal amount and the amount of each antibody (that is, concentration for each antibody) is equal between antibody mixtures. Adjusting the amount of antibodies in this way facilitates the specification of the antibody mixture based on the reactivity in the following ELISA.
Step (iii)
[0358]In this step, the reactivity between each of the antibody mixtures and the target antigen is examined by ELISA so as to specify the antibody mixture showing the reactivity. When at least one of the antibodies recognizing the target antigen is contained in the antibody to be used for preparing the antibody mixture, a plurality of antibody mixtures shows the reactivity. On the other hand, when the antibody recognizing the target antigen is not contained, any of the antibody mixtures will not show reactivity. In this case, the operation is terminated without continuing the following operations.
[0359]The target antigen herein nay include HER1, HER2, CD46, ITGA3, ICAM1, ALCAM, CD147, IgSF4, BCAM, CIqR, CD44, CD73, LAR, EpCAM, HGFR, and the like. The target antigen can be arbitrarily selected. The antigen determined by the below-mentioned identification methods (step (5) and (6)) may be used as the target antigen herein.
Step (iv)
[0360]In this step, a combination of a parameter name and a parameter value that are common to the antibody group contained in the specified antibody mixture is specified. In the present invention, the combination specified herein is referred to as "positive combination." Specifically, the positive combination is specified like (first parameter, parameter value a1), (second parameter, parameter value a2), . . . , (the n-th parameter, parameter value an). When a plurality of antibody mixtures having the different degree of reactivity are recognized in the step (iii), similarly, specification may be carried out for each level of the reactivity. For example, the middle level of positive combination may be specified like (first parameter, parameter value a1), (second parameter, parameter value a2), . . . , (the n-th parameter, parameter value an); and the high level of positive combination may be specified like (first parameter, parameter value b1), (second parameter, parameter value b2), . . . , (the n-th parameter, parameter value bn).
Step (v)
[0361]In this step, antibodies corresponding to the combination specified in step (iv) as to all parameters are selected from the antibody subjected to step (i). That is to say, antibodies in which all parameters are positive combination are selected. For example, when (first parameter, parameter value a1), (second parameter, parameter value a2), . . . , (the n-th parameter, parameter value an) are specified as the positive combination, antibodies having (parameter value a1, parameter value a2, . . . , parameter value an) is selected.
Step (vi)
[0362]In this step, the selected antibodies are classified into one antibody group. Thus, an antibody group showing the reactivity to the target antigen can be made into one group. In other words, an antibody group whose antigen is determined can be obtained. Note here that when only one antibody is selected in the step (v), this only one antibody makes one an antibody group.
[0363]When two or more kinds of target antigens are prepared and the above-mentioned steps (iii) to (vi) are carried out by using each target antigen, two or more antibody groups recognizing different antigens can be obtained.
[0364]In one embodiment of the present invention, the steps (i) to (v) are tried a plurality of times under the conditions in which the combination of parameters is changed every trial. For example, in the first trial, analysis is carried out in which four parameter combinations composed of numeric values (for example, antibody 1 (001, 001, 001, 001), antibody 2 (002, 002, 002, 002), . . . ) are associated with each antibody. In the second trial, analysis is carried out in which three parameter combinations composed of alphabets (for example, antibody 1 (ααα, ααα, ααα), antibody 2 (βββ, βββ, βββ, βββ), . . . ) are associated with each antibody. Note here that each trial is carried out so that the antibody group formed in each trial is not completely identical. The "antibody group is completely identical" means that the numbers of groups are the same and the kinds of antibodies contained in each group are the same over the all groups.
[0365]After a plurality of times of trials, antibodies in which the results in all trials are not contradictory and which show the binding positive reaction to the target antigen are selected. Then, the step (vi) is carried out by using the selected antibody (a plurality of antibodies).
[0366]When trials are carried out at a plurality of times and only an antibody that provides not-contradictory (that is, consistent) results are selected, an antibody having a target antigen reactivity (intended antibody) can be efficiently obtained.
[0367]The number of times of trial in the steps (i) to (v) is not particularly limited. It may be arbitrarily set by considering the number of antibodies to be treated, the number of "positive combinations" that is anticipated at one trial. For example, the number of times of trial can be twice to five times.
[0368]In a further embodiment of the present invention, the following steps are carried out between the step (v) and the step (vi).
[0369](v-1) newly associating the classified antibodies selected in step (v) with a combination of n pieces of parameters in a same manner as in the step (i);
[0370](v-2) with respect to each parameter, preparing the antibody mixture of antibodies having the same parameter value;
[0371](v-3) examining a reactivity of each of the antibody mixtures with a target antigen by an enzyme linked immunosorbent assay (ELISA) so as to specify the antibody mixture showing the reactivity;
[0372](v-4) determining a combination of a parameter name and a parameter value that are common to the antibody group contained in the specified antibody mixture; and
[0373](v-5) selecting an antibody having the combination specified in the step (v-4) in terms of all parameters among the antibodies subjected to the step (v-1).
[0374]Note here that the steps (v-1) to (v-4) are repeated twice or more, if necessary. In this embodiment, a combination of parameters is newly associated with antibodies selected in one trial. Then, the selection of antibody is carried out again. By repeating trials, the intended antibody is narrowed. Thus, classification accuracy is improved.
[0375]Herein, with reference to FIGS. 77 and 78, the principle of the n-dimensional ELISA method is described more particularly. FIGS. 77 and 78 are conceptual diagrams in a case where n is 3 (three dimensional ELISA method). In this example, a general-purposed 96-well microwell plate is used. Firstly, plates in the number necessary to the number of antibody clones are prepared. In this example, the number of antibody clones is made to be 4,800 and 50 plates (4,800 well in total) are prepared.
[0376]Next, the antibody clone is placed in the well sequentially and the antibody clones are arranged in the plate. Thus, each antibody clone is associated with an address consisting of a plate number (first parameter), a plate row name (second parameter), and a plate column number (third parameter). For example, the address of the antibody clone in the first plate, row A and first column in a well becomes (1, A, 1).
[0377]Subsequently, a mixture of antibody clones having the same plate number (referred to as a plate mixed antibody), a mixture of antibody clones having the same plate row name (referred to as a row mixed antibody), and a mixture of antibody clones having the same plate column number (referred to as a column mixed antibody) are prepared, respectively (FIG. 77). The number of the respective mixed antibodies are 50 (first plate mixed antibody to fifth plate mixed antibody), 8 (row A mixed antibody to row H mixed antibody), and 12 (first column mixed antibody to twelfth column mixed antibody), sequentially.
[0378]The mixed antibodies prepared as mentioned above are placed in wells in a newly prepared 96-well microwell plate sequentially, and the mixed antibodies are aligned in the plate. In this example, in the plate, the first to seventh columns are assigned to the plate mixed antibody, the eighth column is assigned to the row mixed antibody, and the ninth to tenth columns are assigned to the column mixed antibody (upper part of FIG. 78). The thus obtained plates are used and ELISA method is carried out. Then, by examining the well showing the reactivity, the address of the intended antibody clone (antibody clone showing the reactivity to the target antigen) is specified. In this example, a well in which the plate mixed antibody of the third plate is placed, a well in which the row mixed antibody of the row E is placed, and a well in which the column mixed antibody of the third column show the reactivity, (3, E, 3) is specified as an address of the intended antibody (lower part of FIG. 78). Finally, antibody clone to which the specified address is associated with is obtained as the intended antibody.
[0379]The second aspect of the present invention provides an identifying method of an antigen to each antibody classified in the classifying method of the present invention. In the identification method of the present invention, following the above-mentioned steps (1) to (4) in the classifying method of the present invention, the below-mentioned steps are carried out.
[0380](5) selecting one or several antibodies from each antibody group formed in the step (4) and identifying an antigen thereof, and
[0381](6) associating the antigens identified in the step (5) with an antibody group, on the estimation that antigens to antibodies belonging to the same antibody group are identical or have high relationship, and
Step (5)
[0382]In this step, antibodies to be identified are selected. The criteria of selection are not particularly limited, and antibodies that are judged to have high reactivity with respect to antigen from the results of the flow cytometry analysis may be selected. This is because when such an antibody is used, the identification operation using the antigen antibody reaction can be carried out advantageously.
[0383]The number of antibody to be selected is typically one, but the number is not necessarily limited to one. If necessary, several antibodies (for example, two or three antibodies) are selected. When a plurality of antibodies are selected from one antibody group, the identification results of antibodies can be compared with each other, and thereby the reliability of the identification results can be improved. On the other hand, when the identification operation is carried out by selecting a more than necessary number of antibodies, excessive workload is applied. As a result, the effect that is originally intended by the present invention is decreased. Then, it is preferable that the number of antibodies to be selected is small. Specifically, the number is preferably five or less, further preferably three or less, and the most preferably two or less. In order to maximize the effect of the present invention, the number of antibody to be selected from each antibody group is one.
[0384]Identification of an antigen to an selected antibody (hereinafter, referred to as "selected antibody") can be carried out by using a method such as mass spectrometry, immunoprecipitation test, Western blotting, affinity chromatography, RNAi, proteomics techniques (analysis by electrophoresis, mass spectrometry, genome data base retrieve, and bioinformatics), and analysis of expression of corresponding gene. Among them, a method by the proteomics technique based on the mass spectrometry is suitable for identification of unknown antigen and preferable for the identification method employed in the present invention. Note here that these methods are not exclusive to each other and two or more of them can be used if necessary.
[0385]The mass spectrometry is a method of determining the mass of samples by separating ions generated from samples such as protein and peptide according to mass/electric charge (m/z), and measuring the intensity thereof. Since soft ionization methods such as an ESI method (Electro Spray Ionization) and an MALDI method (Matrix Assisted Laser Deporption Ionization) are developed, the mass spectrometry is widely used for analyzing living body sample such as protein and peptide.
[0386]A mass spectrometer is generally composed of ion source, mass spectrometer, and detector. According to sample types and analysis purposes, various mass spectrometers are commercially available. For identification of protein or peptide, MS/MS (Mass spectrometry/mass spectrometry) by a tandem mass spectrometry such as ESI Q-TOF MS, MALDI-TOF MS, and the like are used. A measurement method combining liquid chromatography and mass spectrometer (LC-MAS (liquid chromatography/Electro Spray Ionization mass spectrometer), LC-MS/MS, etc.), and the like, can be also used.
[0387]In the tandem mass spectrometer, two mass spectrometers are linked in series in which ions generated in the ion source are separated in the first mass spectrometer (MS 1) and allowed to pass through only a single ion peak. Then, inactive gas particles are allowed to collide with the ions so as to be degraded into product ions. This product ion is analyzed by the second mass spectrometer (MS 2). According to the combination of the first mass spectrometer (MS 1) and the second mass spectrometer (MS 2), tandem mass spectrometers such as Q-TOF, TOF-TOF, Q-Q, and Q-IT (Iontrap) are present. Like Q-TOF (a tandem mass spectrometer in which Quadrupole mass spectrometer: Q-MS and TOF mass spectrometer (Time-of-flight mass spectrometer: TOF-MS are linked in series), hybrid type tandem mass spectrometer composed of two different kinds of mass spectrometers is excellent in MS/MS measurement ability and suitable for identifying the amino acid sequence of protein and peptide.
[0388]In order to identify the amino acid sequence from the results of the mass spectrometer, a PMF method (peptide mass fingerprinting method) of carrying out genome data search by using experiment results, MS/MS ion search method and the like, are used. Furthermore, de novo sequencing method of determining the amino acid sequence by mathematical operation from the MS/MS spectrum without carrying out genome data search may be used.
[0389]On the other hand, an immunoprecipitation test, Western blotting technique, affinity chromatography, RNAi, and the like, are effective method when a selected antibody is anticipated to recognize the known antigen. These methods can examine the reactivity between the selected antibody and well-known antigen. That is to say, in the immunoprecipitation test, it is examined whether or not the selected antibody and certain known antigen form an immunoprecipitate. When an immunoprecipitate is formed, the known antigen is determined to be the antigen of the selected antibody. On the other hand, in the Western blotting technique, it is examined whether or not the selected antibody can recognize an antigen protein transferred to a PVDF membrane etc. Furthermore, in the affinity chromatography, the adsorption property of the selective antibody to a column supporting a certain known antigen is examined. The presence or the degree of adsorption property is determined. Herein, as the known antigen, commercially available antigens, or antigens expressed from a gene and purified can be used. Furthermore, operations of the immunoprecipitation test, Western blotting technique, affinity chromatography, and the like, can be carried out in the usual manner. In the investigation in RNAi, RNAi of the known antigen is allowed to act on forcedly expressed cells or cells to which an antibody is reacted. It is determined that the subject antibody recognizes the subject antigen when the staining property FCM or the degree of cell immunostaining is reduced.
Step (6)
[0390]In the identification method of the present invention, following the step (5), it is assumed that antigens to each antigen belonging to the same antibody group are identical or have high association. According to the assumption, the antigens identified in the step (5) are associated with an antibody group. Thus, all antibodies belonging to the same antibody group are associated with one antigen.
[0391]In one embodiment of the present invention, the above assumption (estimation as to the association of antigen) is verified. That is to say, in this embodiment, the reactivity between the antigen identified in the step (5) and the antibody belonging to the antibody group with which the antigen is associated in the step (6) is examined so as to confirm that the above assumption is correct. Specifically, firstly, antibodies are selected from the antibody group that needs verification. Preferably, all the antibodies are selected, and the reactivity thereof is verified. Next, the reactivity of each antibody to the identified antigen (hereinafter, referred to as "identified antigen") is examined by using the immunoprecipitation test or ELISA (including cell ELISA), and RNAi. For example, in the immunoprecipitation test, by reacting the antibody to an solution or an extracted solution of cells that express the identified antigen, then, proteins recovered as the immunoprecipitates are detected by, for example, electrophoresis. Thereby, the reactivity of each antibody to the identified antigen can be confirmed. On the other hand, in ELISA, for example, by a series of operations including preparation of well in which an identified antigen is fixed, addition of antibody, addition of labeled antibody, and measurement amount of labeled antibodies, the reactivity of each antibody with respect to the identified antigen can be confirmed. Furthermore, also by examining the binding property to cells forcedly expressing the identified antigen, the reactivity of each antigen to the identified antigen can be confirmed. In the verification by RNAi, by allowing the known RNAi to act on cells forcedly expression the identified antigen or subjected cells showing the antibody reaction. When, the staining property of the subjected antibody in FCM and cell immunostaining is reduced, it is recognized that the subjected antigen is recognized.
[0392]Furthermore, when disease-related molecules (disease causative gene products, etc.) can be obtained in same forms such as purified protein or recombinant protein, the intermolecular interaction between such molecules and the antibodies can be examined in vitro (classical methods using fluorescence spectroscopy, gel filtration, and ultracentrifugation; a method using surface plasmon resonance phenomenon; a method using quartz-crystal resonator microbalance, and the like) or in vivo (monomolecular tracing method, fluorescence resonance energy metastasis (fluorescence resonance energy transfer: FRET) observation method, and the like).
[0393]When specific reactivity is observed between the identified antigen and each antibody, it is judged that the above assumption is correct.
[0394]In one embodiment of the present invention, identification results are displayed on a panel. Specifically, the panel is any of the following (a) to (c).
[0395](a) a panel displaying as one antibody group a plurality of antibodies providing data identical to or similar to each other in the flow cytometry analysis in the step (3) in which each antibody group is associated with its antigen;
[0396](b) a panel displaying as one antibody group a plurality of antibodies providing data identical to or similar to each other in the flow cytometry analysis in the step (3) in which each antibody in the antibody group is associated with a cell expressing a cell surface antigen recognized by the each antibody group; and
[0397](c) a panel displaying as one antibody group a plurality of antibodies providing histogramidentical to or similar to each other in the flow cytometry analysis in the step (3) in which each antibody group, its antigen and a cell expressing a cell surface antigen recognized by the antibody are associated with each other.
[0398]The above-mentioned panels are useful for studying identified antigens, and for studying or classifying certain cells displayed on the panel.
[0399]The panel (a) displays the relationship between each antigen to the antibody group. Therefore, it is useful in searching an antibody to a certain antigen. The panel (a) can be formed by displaying by the use of diagrams or tabular formats the association between each antibody group and the antigen by using identification results by steps (5) and (6) of the present invention in which a plurality of antibodies providing identical or highly similar data in the flow cytometry analysis in the step (3) of the present invention are defined as one group.
[0400]The panel (b) shows the association between the antibody group and cells. Therefore, it is useful in searching an antibody to a certain cell surface antigen. Furthermore, when the panel displays the association between the antibody group and a plurality of cells, useful information on the distribution of cell surface antigen can be provided. The panel (b) can be formed by displaying by the use of diagrams or tabular formats the association between each antibody group and cells expression the cell surface antigen recognized thereby by using identification results by steps (5) and (6) of the present invention in which a plurality of antibodies providing identical or highly similar data in the flow cytometry analysis in the step (3) of the present invention are defined as one group.
[0401]The panel (c) combines the panel (a) and the panel (b). This panel shows that the kinds or distribution state of a cell surface antigen expressed by certain cells and allows easy and rapid search of antibodies to the antigens of interest. The panel (c) can be formed by displaying by the use of diagrams or tabular formats the association between each antibody group and cells expression the cell surface antigen recognized by the antigen and each antibody group by using identification results by steps (5) and (6) of the present invention in which a plurality of antibodies providing identical or highly similar histogram in the flow cytometry analysis in the step (3) of the present invention are defined as one group.
[0402]In the identification method of the present invention, identification of antigen with respect to only a part of the antibodies in the antibody group, and as to the other antibodies, antigens are determined by estimation. Therefore, as compared with the case where identification operation is carried out for each antibody, necessary labor and time can be radically reduced. In other words, according to the identification method of the present invention, antigen of each antibody can be determined rapidly and easily. Note here that as shown in the below-mentioned Examples, as far as the present inventors have investigated, error in estimation has not been confirmed. The reliability of this method has been confirmed.
[0403]On the other hand, according to the identification method of the present invention, it is possible to understand the kinds of surface antigens expressed by certain cells. Furthermore, information on the expression amount can be obtained. When the classification of antibodies is carried out by using two kinds or more cells, information on the distribution state of the cell surface antigens can be obtained. Thus, the identification method of the present invention brings useful information as to the cell surface antigen.
[0404]As a result, according to the identification method of the present invention, it is possible to obtain an assembly of antibodies capable of recognizing antigens for each identified antigen (or for each of the plurality of antigens having high association). These antibody groups are useful for study of the cell surface antigens, classification and diagnosis of diseases, and the like. These antibody groups are expected to be applied to the field of treatment.
[0405]The present invention further provides an application of information obtained by the classifying method or the identification method of the present invention. As one of the applications, the third aspect of the present invention relates to a method of obtaining an antibody or an antibody set having a association with respect to a certain disease. The method of obtaining the antibody of the present invention (the first embodiment of the third aspect) includes the following steps.
[0406](1) selecting one or two or more of antibody groups from the plurality of antibody groups classified by the classifying method according to the present invention;
[0407](2) with respect to one kind or two or more kinds of diseases examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0408](3) selecting an antibody in the antibody group, to which an antibody having a specific reactivity to any of diseases belongs, as a useful antibody.
[0409]On the other hand, a method of obtaining an antibody set of the present invention (the second embodiment of the third aspect) includes the step (3') instead of the step (3):
[0410](3') selecting diseases to which two or more antibodies show a specific reactivity, then selecting antibodies from the antibody group, to which the antibody having a specific reactivity to the disease belongs, and combining the selected antibodies.
[0411]Hereinafter, the detail of each step is described with reference to FIG. 1. For convenience of explanation, in FIG. 1, it is assumed that the antibody groups 1 to 5 are obtained by the classifying method of the present invention and three antibodies belong to each antibody group. Furthermore, in this example, it is assumed that antigens to each antibody group have been already identified.
[0412]Firstly, in the step (1), focused antibody group (antibody groups 1, 3, and 5) are selected (FIG. 1, (1)). As in this example, two or more antibody groups may be selected.
[0413]Next, in the step (2), the reactivity between an antibody to each of the selected antibody groups and a certain disease is examined. Specifically, a sample (cells or tissues) derived from a patient having a certain disease is prepared, and then, the reactivity of each antibody to the sample is examined (FIG. 1, (2)). Two or more antibodies from each of the selected antibody groups are selected, and thereby the reactivity of them may be examined. The "certain disease" herein is not particularly limited but it may include various kinds of cancers, for example, kidney cancer, hepatic cell carcinoma, gallbladder and liver cancer, alveolar cell carcinoma, lung squamous cell cancer, pulmonary adenocarcinoma, pancreas cancer, adenocarcinoma, or ovarian cancer. In the example shown in FIG. 1, the reactivity with respect to two kinds or more of diseases are examined simultaneously. However, the examination is not limited to this alone. The reactivity to one disease may be examined. Furthermore, the reactivity with respect to a certain pathologic condition in the certain disease may be examined.
[0414]The reactivity with respect to the samples derived from a patient can be detected and evaluated by using an immunohistochemical staining technique, an immunoprecipitation method, flow cytometry analysis, cell ELISA and the like. These methods are not exclusive to each other and therefore two or more of these methods can be used if necessary. Among them, it is preferable to employ the immunohistochemical staining technique. The immunohistochemical staining technique permits rapid and sensitive detection. Furthermore, its operation is relatively simple.
[0415]In the immunohistochemical staining technique, tissues collected from a patient and an antibody are brought into contact with each other, and then, specifically bonded antibodies are detected. Concretely, the method of the present invention can be carried out according to the following immunohistochemical staining technique.
[0416]The immunohistochemical staining of living tissue is generally carried out by the following procedures (a) to 0). Note here that the immunohistochemical staining of living tissue can be referred to as various documents and publications (for example, "Enzyme-labeled Antibody Method" 3rd revised edition, K. Watanabe and K. Nakane (ed), Gakusai Kikaku).
(a) Immobilization--Paraffin Embedding Method Tissue surgically collected from a living body is immobilized in formalin, paraformaldehyde, absolute ethyl alcohol, and the like, and then embedded in paraffin. In general, it is dehydrated with alcohol, treated with xylene and embedded in paraffin. The paraffin embedded specimen is cut into a desired thickness (for example, 3 to 5 μm thick) and extended on a slide glass. Instead of the paraffin embedding specimen, an alcohol immobilized specimen, a dry sealed specimen, a frozen specimen, and the like may be used.
(b) Deparaffinization
[0417]In general, treatment is carried out with xylene, alcohol, and purified water sequentially in this order.
(c) Pretreatment (Antigen Activation)
[0418]If necessary, for antigen activation, for example, enzyme treatment, heat treatment and/or pressurization treatment are carried out.
(d) Removal of Endogeneous Peroxidase
[0419]When peroxidase is used as a labeling material for staining, endogeneous peroxidase activation is removed by carrying out with hydrogen peroxide solution.
(e) Non-Specific Reaction Inhibition
[0420]Non-specific reaction is inhibited by treating a section with bovine serum albumin solution (for example, 1% solution) for several minutes to several tens of minutes. Note here that this process may be omitted when the following primary antibody reaction is carried out by using an antibody solution impregnated with bovine serum albumin.
(f) Primary Antibody Reaction
[0421]An antibody diluted to an appropriate concentration is dropped on the slide glass and allowed to react for ten minutes to several hours. After reaction, the reacted produce is washed with an appropriate buffer solution such as phosphate buffer.
(g) Addition of Labeling Reagent
[0422]As the label material, peroxidase is frequently used. Secondary antibody bonded to peroxidase is dropped on the section and then allowed to react for ten minutes to several hours. After reaction, the reacted product is washed with an appropriate buffer solution such as phosphate buffer.
(h) Color Reaction
[0423]DAB (3,3'-diaminobenzidine) is dissolved in Tris buffer. Then, hydrogen peroxide solution is added. The thus prepared coloring solution is impregnated into a section for several minutes (for example, five minutes) so as to color the section. After coloring, the section is sufficiently washed with tapped water so as to remove DAB.
(i) Nuclear Staining
[0424]The section is subjected to nuclear staining by reacting it with Mayer hematoxylin for several seconds to several tens seconds. It was washed with flowing water for saddening (in general, for several minutes).
(j) Dehydration, Clearing, Encapsulation
[0425]The section is dehydrated with alcohol, clearing treated with xylene, and finally encapsulated with synthesized resin, glycerine, rubber syrup, and the like.
[0426]An antibody that is recognized to have specific reactivity to any of diseases can detect a cell surface antigen characterizing the disease with high sensitivity. Such an antibody is expected to be used as a diagnosis or treatment antibody of the disease. Then, in the step (3), an antibody of the antibody group including such an antibody is selected (FIG. 1 (3)). As a result, in this example, as to disease A, an antibody (antibody 1-1, 1-2 or 1-3) of the antibody group 1 and an antibody of the antibody group 3 (antibody 3-1, 3-2 or 3-3) are selected. As to disease B, an antibody (antibody 5-1, 5-2 or 5-3) of the antibody group 5 is selected. In this way, a specific antibody for a certain diseases can be obtained.
[0427]In the step (3'), a disease in which two or more antibodies show the specific reactivity is selected, and then, each antibody is selected from the antibody group to which the antibody showing the specific reactivity to the disease belongs, is selected, and the selective antibodies are combined (FIG. 1, (3')). That is to say, in this example, the disease A is selected and the antibodies of antibody groups 1 and 3, which are antibody groups to which the antibody showing the specific reactivity to the disease A belongs, are combined. Thus, the antibody set showing specific to a certain disease is obtained.
[0428]Herein, by comparing the specificities (cross reactivity) of the antibodies in the antibody group, an antibody having the most excellent property may be selected (in this example, antibody 1-2, antibody 3-3, and antibody 5-3 are selected. See, FIG. 1, (4)). By adding this step, more useful antibody or antibody set can be obtained.
[0429]Furthermore, an antibody set may be structured by combining an arbitrary antibody that does not have reactivity to the diseases with the antibodies selected as the antibodies showing the reactivity to a certain disease (in this example, for example, the antibody 4-1 is combined to an antibody of the antibody group 1 and antibody of antibody group 3). By using such an antibody set, detail characterization of the disease can be possible.
[0430]According to the obtaining method of the present invention, an antibody (or antibody set) to a disease-specific antigen can be obtained. The antibody (or antibody set), which are as it is or to which necessary modification is added, is useful for study, classifying, diagnosing and treating the disease or the pathologic condition. Thus, this method provides an extremely useful tool in the field of medicine.
[0431]The third embodiment of this aspect provides the obtaining method of antibody set including the following steps.
[0432](1) selecting two or more antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to the present invention;
[0433](2) with respect to two kinds or more of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0434](3) selecting antibodies from the antibody group, to which the antibody having a specific reactivity to any of disease belongs, and combining the selected antibodies.
[0435]Hereinafter, the detail of each step is described with reference to FIG. 2. For convenience of explanation, in FIG. 2, it is assumed that the antibody groups 1 to 6 are obtained by the classifying method of the present invention and three antibodies belong to each antibody group. The antigens (antigen A) in the antibody groups 1 to 3 are common. Similarly, the antigens (antigen B) in the antibody groups 4 and 5 are also common.
[0436]In the step (1) of this embodiment, two or more antibody groups recognizing different antigens (antibody groups 1, 4, and 6) are selected (see, FIG. 2 (1)). In the following step (2), the reactivity between the antibodies (antibodies 1-1, 4-1, and 6-1) in each of the selected antibody groups and certain diseases (diseases A to D) are examined (FIG. 2, (2)). In the step (3), antibodies in the antibody groups to which the antibody belong showing specific reactivity to any of diseases are combined. That is to say, in this example, an antibody of antibody group 1 to which an antibody 1-1 showing specific reactivity to disease A and an antibody of antibody group 4 to which an antibody 4-1 showing specific reactivity to disease B are combined to form an antibody set (FIG. 2, (3)). Thus, an antibody set (the antibody 1-1 and the antibody 4-1) including an antibody specific to disease A and an antibody specific to disease B is obtained. This antibody set is useful for detecting, for example, disease A or disease B and this antibody is a reagent effective to the discrimination of the diseases A and B.
[0437]Note here that by comparing the specificity (cross reactivity) and the like between the antibodies in the antibody group, an antibody having the most excellent property may be selected (In this example, the antibody 1-2 and the antibody 4-3 are selected. FIG. 2, (4)). By adding this step, it is possible to obtain a more useful antibody set.
[0438]As a result of carrying out the classifying method and the identification method of the present invention, assuming that a plurality of antibodies groups recognizing the same antigen are obtained, the fourth embodiment of this aspect provides a obtaining method of an antibody set including the following steps.
[0439](1) selecting two or more antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to the present invention;
[0440](2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0441](3) selecting an antibody from the antibody group to which the antibody having a specific reactivity to any of disease belongs, and an antibody belonging to other antibody group whose antigen is common to that of the antibody group, and combining the selected antibodies.
[0442]Hereinafter, the detail of each step is described with reference to FIG. 3. For convenience of explanation, in FIG. 3, it is assumed that the antibody groups 1 to 6 are obtained by the classifying method of the present invention and three antibodies belong to each antibody group. The antigens (antigen A) in the antibody groups 1 to 3 are common. Similarly, the antigens (antigen B) in the antibody groups 4 and 5 are also common.
[0443]In the step (1) of this embodiment, two or more antibody groups recognizing different antigens (antibody groups 1, 4, and 6) are selected (see, FIG. 3 (1)). In the following step (2), the reactivity between the antibodies (antibodies 1-1, 4-1, and 6-1) in each of the selected antibody groups- and certain diseases (diseases A to D) are examined (FIG. 3, (2)). In the step (3), an antibody of the antibody group to which an antibody showing the specific reactivity to any of diseases and an antibody belonging to other antibody group whose antigen is common to the group are selected, respectively. The selected antibodies are combined so as to form an antibody set (FIG. 3, (3)). That is to say, in this example, an antibody in antibody group 1 to which antibody 1-1 belongs showing specific reactivity to disease A and an antibody of the antibody groups 2 and 3 whose antigens are common are combined. Thus, an antibody set specific to the disease A is obtained. Similarly, an antibody in antibody group 4 to which antibody 4-1 belongs showing specific reactivity to disease B and an antibody of the antibody group 5 whose antigen is common to that of antibody group 4. Thus, an antibody set specific to the disease B is obtained. As shown in this example, "another antibody group" herein is not particularly one but a plurality antibody groups may be present.
[0444]Herein, even in the case of cancers of the same organ, depending upon patients, the pathologic condition (grade of malignancy) may be largely different. The difference in such pathologic conditions is thought to be involved to the expression forms of the specific antigens. On the other hand, the antibody sets obtained in this embodiment are not different in the level recognized by an antigen but include antibodies that are different in the level of epitope. That is to say, this is an antibody set including a plurality of antibodies that are different in the epitope to be recognized. Such an antibody set permits multilateral detection or evaluation of expression forms of antigen. For example, such an antibody set is useful for detection of certain pathologic conditions in, for example, cancers, or a determination of the grade of malignancy.
[0445]Note here that by comparing the specificity (cross reactivity) and the like in the antibodies in the antibody group, an antibody having the most excellent property may be finally selected (FIG. 3, (4)). By adding this step, it is possible to obtain a more useful antibody set.
[0446]As a result of carrying out the classifying method and the identification method of the present invention, assuming that a plurality of antibodies groups recognizing the same antigen are obtained, the fifth embodiment of this aspect provides a obtaining method of an antibody set including the following steps.
[0447](1) selecting two or more antibody groups recognizing the same antigen from the plurality of antibody groups classified by the classifying method according to the present invention;
[0448](2) with respect to one kind or two or more kinds of pathologic conditions, examining a reactivity between an antibody in each of the selected antibody groups and a pathologic condition; and
[0449](3) associating information about the reactivity and then combining the antibodies in the antibody groups.
[0450]Hereinafter, the detail of each step is described with reference to FIG. 4. For convenience of explanation, in FIG. 4, it is assumed that the antibody groups 1 to 6 are obtained by the classifying method of the present invention and three antibodies belong to each antibody group. The antigens (antigen A) in the antibody groups 1 to 3 are common. Similarly, the antigens (antigen B) in the antibody groups 4 and 5 are also common.
[0451]In the step (1) of this embodiment, two or more antibody groups recognizing common antigen (antibody groups 1 to 3) are selected (see, FIG. 4 (1)). In the following step (2), the reactivity between the antibodies (antibodies 1-1, 2-1, and 3-1) in each of the selected antibody groups and certain various diseases are examined (FIG. 4, (2)). Specifically, as to various pathologic conditions of certain disease, samples (cells or tissue) derived from a patient are prepared, and the reactivity between the samples and each antibody is examined. In the step (3), the obtained reactivity is associated with each other (FIG. 4, (2), right column), and then antibodies of each of the selected antibody groups (antibody groups 1 to 3) are combined so as to form an antibody set (FIG. 4, (3)). Thus, antibody sets specific to the certain pathologic condition of certain disease is obtained (in this example, an antibody set specific to pathologic condition of disease A including antibodies of the antibody groups 1 to 3 is obtained). The antibody set obtained in this embodiment is typically not different in the level of an antigen but include antibodies that are different in the level of epitope. Therefore, similar to the antibody set according to the above-mentioned embodiment, for example, the antibody set is useful detecting the certain pathologic condition in, for example, cancer, or a determination of the grade of malignancy. Note here that it is preferable that an antibody set is constructed by excluding antibodies showing no specific reactivity with respect to any pathologic conditions.
[0452]By comparing the specificity (cross reactivity) and the like in the antibodies in the antibody group, an antibody having the most excellent property may be finally selected (in this example, antibodies 1-2, 2-1 and 3-3 are selected, FIG. 4, (4)). By adding this step, it is possible to obtain a more useful antibody set.
[0453]A further aspect of the present invention provides a production method of a panel displaying a association between an antibody and a disease (or pathologic condition). In the first embodiment of this aspect, the following steps are carried out.
[0454](1) selecting one or two or more of antibody groups from the plurality of antibody groups classified by the classifying method according to the present invention;
[0455](2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0456](3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
[0457]When one antibody group is selected in the step (1), as to one antibody or a plurality of antibodies whose antigen is common, a panel displaying the association with respect to a certain disease can be obtained. In the latter case, as to a plurality of antibodies whose antigen is common antigen, from the viewpoint of the association with respect to the disease, difference or points of difference (one caused by the cross reactivity and the like) can be read out. That is to say, the panel gives an important suggestion as to the property of the antibody. On the other hand, when two or more antibody groups are selected in the step (1), as to a plurality of antibodies whose antigen is different (however, when several antibodies from each antibody group in the step (1), antibodies whose antigen is common is contaminated), a panel displaying the association with respect to the certain disease is obtained. This panel gives information on the antibody group useful for study, classification and diagnosis. The panel itself has a great value. Form this panel, the association between a plurality of antigen and disease can be read out. That is to say, the panel gives an important suggestion as to the association between each antigen and disease.
[0458]Herein, in the step (2), it is preferable to examine the reactivity of the antibody as to two or more diseases. Thus, a panel displaying the association (linkage) between each antibody and two or more diseases can be obtained. The panel displays more pieces of information and further displays the association between diseases. Suggestion that is useful and important for study, classification and diagnosis of the diseases can be obtained.
[0459]In the second embodiment of this aspect, the following steps are carried out.
[0460](1) selecting two or more of antibody groups recognizing different antigens from the plurality of antibody groups classified by the classifying method according to the present invention;
[0461](2) with respect to one kind or two or more kinds of diseases, examining a reactivity between an antibody in each of the selected antibody groups and a certain disease; and
[0462](3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
[0463]In this embodiment, a panel displaying the association between a plurality of antibodies whose antigen is different and a certain disease can be obtained. This panel gives information on an antibody group useful for study, classification and diagnosis for a disease and the panel itself has a great value. Form this panel, the association (linkage) between a plurality of antigens and disease can be read out. That is to say, the panel gives important suggestions as to the association between each antigen and disease as well as the association between antigens.
[0464]Herein, in the step (2), it is preferable to examine the reactivity of the antibody as to two or more diseases. Thus, a panel displaying the association between each antibody and two or more diseases can be obtained. The panel displays more pieces of information and further displays the association between diseases. Suggestion that is useful and important for study, classification and diagnosis of the diseases can be obtained.
[0465]In the third embodiment of this aspect, the following steps are carried out.
[0466](1) selecting two or more of antibody groups recognizing a common antigen from the plurality of antibody groups classified by the classifying method according to the present invention;
[0467](2) with respect to one kind or two or more kinds of pathologic condition, examining a reactivity between an antibody in each of the selected antibody groups and a certain pathologic condition of disease; and
[0468](3) associating the results of the step (2) with each antibody and displaying by using a drawing or a tabular format.
[0469]In this embodiment, as to a plurality of antibodies whose antigen is common, a panel displaying the association with respect to a pathologic condition of a certain disease can be obtained. This panel gives information on antibody group that is useful for study of each pathologic condition, study of difference between pathologic conditions, classification of pathologic conditions, or diagnosis on the level of the pathologic condition. The panel itself has a great value.
[0470]Herein, in the step (2), it is preferable to examine the reactivity of the antibody as to two or more pathologic conditions. Thus, a panel displaying the association between each antibody and two or more pathologic conditions can be obtained. This panel displays not only more pieces of information but also the association between the pathologic conditions. Suggestion that is useful and important to study, classification and diagnosis of each pathologic condition can be obtained.
[0471]Note here that the first embodiment of this aspect corresponds to the first and second embodiments of the third aspect. Similarly, the third aspect of the second embodiment corresponds to the third and fourth embodiments of the third aspect, respectively. Therefore, as to the matters that are not specifically noted in this aspect, the explanation of the corresponding third aspect is employed. In the panel of the present invention, the term "association between antibody and disease (or pathologic condition)" is displayed by characters showing subject diseases (or pathologic conditions) are positive or negative to the antibody (for example, "to positive," "to negative," "positive," and "negative") or marks (for example, "o," "x," "P," and "N") etc. The display is not limited to two-stage display and, display may be carried out in four stages, for example, strongly positive, moderate positive, weak positive, and negative.
[0472]The number of antibodies displayed in one panel is not particularly limited. For example, the number is 1 to 1000, preferably 2 to 100, and further preferably 5 to 59.
[0473]Furthermore, in addition to the association between an antibody and a certain disease (or pathologic condition), an antigen to each antibody may be shown. The combination of the panel of this aspect and the antibody (or antibody set) obtained in the above-mentioned obtaining method of the present invention becomes an effective tool for study, classification and diagnosis of diseases, pathologic conditions, or the like. That is to say, according to the combination, both information, i.e., an antibody (or an antibody set) specific to a disease or a pathologic condition and the association between the antibody (or the antibody set) and the disease or the pathologic condition can be obtained simultaneously.
[0474]The present invention further relates to a method of testing a disease in which a cell surface antigen is an indicator, the method comprising the following steps.
[0475](1) preparing a cell or a tissue separated from a subject;
[0476](2) examining a reactivity between the cell or the tissue and each antibody displayed on the panel (panel displaying the association between an antibody and a disease (or a pathologic condition)) according to the present invention; and
[0477](3) collating the results in the step (2) with the panel.
[0478]According to the testing method of the present invention, as to a disease or a pathologic condition to be tested (hereinafter, referred to as "diseased to be tested"), information about the presence of contraction of a subject, contraction risk, pathologic conditions, and the like, can be obtained. That is to say, the testing method of the present invention is effective means for diagnosing the subjected disease. Furthermore, when the testing method of the present invention is carried out along with the treatment, the therapeutic effect can be evaluated based on the testing results. Thus, the testing method of the present invention may be used for monitoring the therapeutic effect.
[0479]In the step (1), cells or tissue separated from a subject (that is, a living body) (hereinafter, referred to as "subject cell, and the like") are prepared. The term "separated from a subject" means a state in which a part of cells or tissue of a subject is extracted and completely isolated form a subject as a living body. A person who needs information about a disease to be tested is a subject. A subject may be a patient of a disease to be tested or may be an apparent healthy person. The "apparent healthy person" means a person who has not recognized to be a patient of a disease to be tested prior to the application of the testing method of the present invention.
[0480]In the step (2), the reactivity between the subject cells and the like and each antibody displayed on the panel of the present invention is examined. That is to say, by using an immunologic procedure (for example, immunohistochemical staining technique), whether or not the tested cells express an antigen recognized by each antibody is examined. According to the immunologic procedure, in general, information on the expression amount of antigens can be obtained. Therefore, in addition to the presence of expression antigen, the expression amount may be also examined. An example of the immunologic procedure includes ELISA method, radioimmunoassay, flow cytometry analysis, immunoprecipitation method, immune-blotting, and the like.
[0481]In the step (3), the results of the step (2) (reactivity of each antibody) is collated with the panel of the present invention. The panel of the present invention displays the association between each antibody and a disease or a pathologic condition. Therefore, this step clarifies the association between the tested cells etc. and the disease via the reactivity with respect to each antibody.
[0482]A further application of the above-mentioned panel also includes the following method of the present invention, that is, the optimum method of treating certain diseases, which includes the following steps.
[0483](1) preparing a cell or a tissue separated from a subject;
[0484](2) examining a reactivity between the cell or the tissue and each antibody displayed on the panel (a panel displaying the association between the antibody and disease (or pathologic condition)) according to the present invention;
[0485](3) collating the results in the step (2) with the panel, and
[0486](4) selecting an effective antibody according to the results of collating.
[0487]In the selection method of the present invention, similar to the above-mentioned testing method, after the steps (1) to (3) are carried out, according to the collation results, an effective antibody is selected (the step (4)). As the effective antibody, typically, an antibody showing a specific reactivity in the step (2) is selected. An antibody equivalent to the antibody showing a specific reactivity in the step (2) may be also selected. The "equivalent antibody" means an antibody having equivalent properties (reactivity or activity) to the reference antibody. An example of the equivalent antibody may be an antibody in which the sequence of the heavy chain variable region and the sequence of the light chain variable region are not substantially different from that of the reference antibody (completely identical, or slightly different so that the reactivity or activity is not affected). Another example of the equivalent antibody may be an antibody in which no difference is observed in all of the sequence of each CDR constituting heavy chain variable region and the sequence of each CDR constituting light chain variable region when it is compared with the reference antibody.
[0488]Diseases to which the selection method of the present invention is applied is a disease in which cell surface antigen selected from the group consisting of HER1, HER2, CD46, ITGA3, ICAM1, ALCAM, CD147, IgSF4, BCAM, C1qR, CD44, CD73, LAR, EpCAM and HGFR is an indicator. That is to say, for selecting optimum treatment methods suitable for various diseases characterized by the expression of the cell surface antigen, the present invention can be used. According to the present invention, optimum treatment method suitable for each patient can be selected. Thus, tailor-made medicine can be realized.
[0489]It is preferable that the panel used in the selection method of the present invention displays two or more antibodies selected from the group consisting of 048-006 antibody, 057-091 antibody, 059-152 antibody, 048-040 antibody, 054-101 antibody, 055-147 antibody, 059-173 antibody, 067-149 antibody, 067-176 antibody, 015-126 antibody, 015-044 antibody, 015-102 antibody, 015-136 antibody, 015-143 antibody, 015-209 antibody, 039-016 antibody, 053-216 antibody, 075-024 antibody, 075-110 antibody, 086-032 antibody, 086-035 antibody, 086-036 antibody, 086-061 antibody, 086-138 antibody, 086-182 antibody, 035-224 antibody, 045-011 antibody, 051-144 antibody, 052-053 antibody, 052-073 antibody, 053-049 antibody, 3172-120 antibody, 066-069 antibody, 015-003 antibody, 064-002 antibody, 064-006 antibody, 064-012a antibody, 064-012b antibody, 064-014 antibody, 064-054 antibody, 064-085 antibody, 064-093 antibody, 064-116 antibody, 065-183 antibody, 067-142 antibody, 068-007 antibody, 052-033 antibody, 053-042 antibody, 053-051 antibody, 053-059 antibody, 053-085 antibody, 035-234 antibody, 040-107 antibody, 041-118 antibody, 066-174 antibody, 083-040 antibody, 029-143 antibody, 045-134 antibody, 062-101 antibody, 062-109 antibody, 084-103 antibody, 052-274 antibody, 029-067 antibody, 083-131 antibody, 059-053 antibody, 064-003 antibody, 067-213 antibody, 067-153 antibody, 067-126 antibody, 067-133 antibody, 067-287 antibody, 064-044 antibody, 065-030 antibody, 065-358 antibody, 066-019 antibody, 079-085 antibody, 067-024 antibody, and 076-048 antibody.
[0490]In one embodiment of the selecting method of the present invention, the following steps are carried out.
[0491](1) preparing a panel displaying a reactivity between one or more antibodies selected from the group consisting of 048-006 antibody, 015-126 antibody, 067-133 antibody, 064-044 antibody, 076-048 antibody and 059-053 antibody, and a clinical cancer tissue of one or more diseases selected from the group consisting of squamous carcinoma, adenosquamous carcinoma, alveolar adenocarcinoma, adenocarcinoma, and large cell carcinoma, and a cell or tissue separated from a subject;
[0492](2) examining reactivity between the cell or the tissue and each antibody displayed on the panel;
[0493](3) collating the results in the step (2) with the panel, and
[0494](4) selecting an effective antibody according to the results of collating. In the step (1) of this embodiment, a panel displaying the reactivity between an antibody successfully obtained by the present inventor and clinical cancer tissue of a certain disease is prepared. In addition, cells or tissue separated from a subject are prepared. The step (2) or later are carried out similar to the above-mentioned embodiments. Note here that, a specific example of the panel to be used in this embodiment is a panel shown in FIG. 69.
[0495]Also in this embodiment, an antibody showing the specific reactivity in the step (2) or the equivalent antibody thereto is selected as an effective antibody. The selection method of this embodiment is preferred for selecting the suitable treatment method of squamous carcinoma, adenosquamous carcinoma, alveolar adenocarcinoma, adenocarcinoma, or large cell carcinoma.
[0496]As a further aspect of the present invention provides an isolated antibody (or an antibody set) obtained in the above-mentioned obtaining method of an antibody (or an obtaining method of an antibody set). As shown in the below-mentioned Examples, the present inventors have succeeded in actually obtaining by the method of the present invention, an antibody relevant to HER1, an antibody relevant to HER2, an antibody relevant to CD46, an antibody relevant to ITGA3, an antibody relevant to ICAM1, an antibody relevant to ALCAM, an antibody relevant to CD147, an antibody relevant to C1qR, an antibody relevant to CD44, an antibody relevant to CD73, an antibody relevant to EpCAM, an antibody relevant to HGFR, an antibody relevant to LAR, and an antibody relevant to BCAM. Furthermore, in the current testing method, it is possible to obtain an antibody capable of recognizing two clinical specimen s that are determined to have the same disease (pathologic condition). With this antibody, a certain disease can be newly classified based on the expression state of an antigen and further such a disease can be examined.
[0497]A further aspect of the present invention provides an antibody successfully obtained by the present inventors and the application thereof. As shown in the below-mentioned Examples, the present inventors succeeded in obtaining nine kinds of antibodies to HER1 (048-006 antibody, 057-091 antibody, 059-152 antibody, 048-040 antibody, 054-101 antibody, 055-147 antibody, 059-173 antibody, 067-149 antibody, and 067-176 antibody), 16 kinds of antibodies to HER2 (015-126 antibody, 015-044 antibody, 015-102 antibody, 015-136 antibody, 015-143 antibody, 015-209 antibody, 039-016 antibody, 053-216 antibody, 075-024 antibody, 075-110 antibody, 086-032 antibody, 086-035 antibody, 086-036 antibody, 086-061 antibody, 086-138 antibody, and 086-182 antibody), eight kinds of antibodies to CD46 (035-224 antibody, 045-011 antibody, 051-144 antibody, 052-053 antibody, 052-073 antibody, 053-049 antibody, 3172-120 antibody, and 066-069 antibody), 13 kinds of antibodies to ITGA3 (015-003 antibody, 064-002 antibody, 064-006 antibody, 064-012a antibody, 064-012b antibody, 064-014 antibody, 064-054 antibody, 064-085 antibody, 064-093 antibody, 064-116 antibody, 065-183 antibody, 067-142 antibody, and 068-007 antibody), five kinds of antibodies to ICAM1 (052-033 antibody, 053-042 antibody, 053-051 antibody, 053-059 antibody, and 053-085 antibody), 13 kinds of antibodies to ALCAM (035-234 antibody, 040-107 antibody, 041-118 antibody, 066-174 antibody, 083-040 antibody, 029-143 antibody, 045-134 antibody, 062-101 antibody, 062-109 antibody, 084-103 antibody, 052-274 antibody, 029-067 antibody, and 083-131 antibody), one kind of antibody to CD147 antibody (059-053 antibody), one kind of antibody to C1qR (070-016 antibody), one kind of antibody to CD44 (064-003 antibody), one kind of antibody to CD73 (067-213 antibody), one kind of antibody to EpCAM (067-153 antibody), three kinds of antibodies to HGFR (067-126 antibody, 067-133 antibody, and 067-287 antibody), five kinds of antibodies to LAR (064-044 antibody, 065-030 antibody, 065-358 antibody, 066-019 antibody, and 079-085 antibody), and one kind of antibody to BCAM (067-024 antibody). Since these antibodies are recognize an extracellular domain of antigen in a state in which it is expressed on the surface of the cell membrane, they are useful for staining cells and tissues, and the like. As a result of analysis of sequences of each antibody, the following sequence information is obtained. Note here that, following to the antibody name, the amino acid sequence of the heavy chain variable region; the amino acid sequence of the heavy chain CDR1; the amino acid sequence of the heavy chain CDR2; the amino acid sequence of the heavy chain CDR3; the amino acid sequence of the light chain variable region; the amino acid sequence of the light chain CDR1; the amino acid sequence of the light chain CDR2; and the amino acid sequence of the light chain CDR3 are described sequentially in this order.
1. Antibody to HER1
[0498]A plurality of antibodies clones are obtained. Among them, antibodies having the same amino acid sequence are included. As to the below-mentioned nine kinds of antibody clones, the sequences are analyzed.
[0499]048-006 antibody: SEQ ID NO: 1 (VH); SEQ ID NO: 2 (VH CDR1); SEQ ID NO: 3 (VH CDR2); SEQ ID NO: 4 (VH CDR3); SEQ ID NO: 5 (VL); SEQ ID NO: 6 (VL CDR1); SEQ ID NO: 7 (VL CDR2); SEQ ID NO: 8 (VL CDR3)
[0500]057-091 antibody: SEQ ID NO: 9 (VH); SEQ ID NO: 10 (VH CDR1); SEQ ID NO: 11 (VH CDR2); SEQ ID NO: 12 (VH CDR3); SEQ ID NO: 13 (VL); SEQ ID NO: 14 (VL CDR1); SEQ ID NO: 15 (VL CDR2); SEQ ID NO: 16 (VL CDR3)
[0501]059-152 antibody: SEQ ID NO: 17 (VH); SEQ ID NO: 18 (VH CDR1); SEQ ID NO: 19 (VH CDR2); SEQ ID NO: 20 (VH CDR3); SEQ ID NO: 21 (VL); SEQ ID NO: 22 (VL CDR1); SEQ ID NO: 23 (VL CDR2); SEQ ID NO: 24 (VL CDR3)
[0502]048-040 antibody: SEQ ID NO: 483 (VH); SEQ ID NO: 484 (VH CDR1); SEQ ID NO: 485 (VH CDR2); SEQ ID NO: 486 (VH CDR3); SEQ ID NO: 487 (VL); SEQ ID NO: 488 (VL CDR1); SEQ ID NO: 489 (VL CDR2); SEQ ID NO: 490 (VL CDR3)
[0503]054-101 antibody: SEQ ID NO: 491 (VH); SEQ ID NO: 492 (VH CDR1); SEQ ID NO: 493 (VH CDR2); SEQ ID NO: 494 (VH CDR3); SEQ ID NO: 495 (VL); SEQ ID NO: 496 (VL CDR1); SEQ ID NO: 497 (VL CDR2); SEQ ID NO: 498 (VL CDR3)
[0504]055-147 antibody: SEQ ID NO: 499 (VH); SEQ ID NO: 500 (VH CDR1); SEQ ID NO: 501 (VH CDR2); SEQ ID NO: 502 (VH CDR3); SEQ ID NO: 503 (VL); SEQ ID NO: 504 (VL CDR1); SEQ ID NO: 505 (VL CDR2); SEQ ID NO: 506 (VL CDR3)
[0505]059-173 antibody: SEQ ID NO: 507 (VH); SEQ ID NO: 508 (VH CDR1); SEQ ID NO: 509 (VH CDR2); SEQ ID NO: 510 (VH CDR3); SEQ ID NO: 511 (VL); SEQ ID NO: 512 (VL CDR1); SEQ ID NO: 513 (VL CDR2); SEQ ID NO: 514 (VL CDR3)
[0506]067-149 antibody: SEQ ID NO: 515 (VH); SEQ ID NO: 516 (VH CDR1); SEQ ID NO: 517 (VH CDR2); SEQ ID NO: 518 (VH CDR3); SEQ ID NO: 519 (VL); SEQ ID NO: 520 (VL CDR1); SEQ ID NO: 521 (VL CDR2); SEQ ID NO: 522 (VL CDR3)
[0507]067-176 antibody: SEQ ID NO: 523 (VH); SEQ ID NO: 524 (VH CDR1); SEQ ID NO: 525 (VH CDR2); SEQ ID NO: 526 (VH CDR3); SEQ ID NO: 527 (VL); SEQ ID NO: 528 (VL CDR1); SEQ ID NO: 529 (VL CDR2); SEQ ID NO: 530 (VL CDR3)
[0508]As mentioned in the below-mentioned Examples, the relationships between these antibodies and pancreatic cancer cell line PANC-1, kidney cancer cell line CCFRC1, kidney cancer cell line Caki-1, ovarian cancer cell line KF28, stomach cancer cell line SNU-5, lung squamous cell carcinoma line RERF-LC-AI, ovarian cancer cell line RMG-1, undifferentiated hepatic cell carcinoma cancer cell line HLF, ovarian cancer cell line SKOv3, pulmonary adenocarcinoma cell line PC14, kidney cancer cell line ACHN, lung squamous cell carcinoma line EBC1, vulva mucosal epithelial cell line A431, pulmonary adenocarcinoma cell line H1373, hepatic cell carcinoma cell line HepG2, and kidney cancer clinical specimen established cell line (as to the above mention, based on the results of the cell line staining), as well as the relationships between these antibodies and kidney cancer, hepatic cell carcinoma, gallbladder and liver cancer, lung squamous cell cancer, pulmonary adenocarcinoma, and pancreas cancer (as to the above mention, based on the results of the tissue staining) are experimentally confirmed.
2. Antibody to HER2
[0509]A plurality of antibodies clones are obtained. Among them, antibodies having the same amino acid sequence are included. As to the below-mentioned 16 kinds of antibody clones, the sequences are analyzed.
[0510]015-126 antibody SEQ ID NO: 25 (VH); SEQ ID NO: 26 (VH CDR1); SEQ ID NO: 27 (VH CDR2); SEQ ID NO: 28 (VH CDR3); SEQ ID NO: 29 (VL); SEQ ID NO: 30 (VL CDR1); SEQ ID NO: 31 (VL CDR2); SEQ ID NO: 32 (VL CDR3)
[0511]015-044 antibody SEQ ID NO: 531 (VH); SEQ ID NO: 532 (VH CDR1); SEQ ID NO: 533 (VH CDR2); SEQ ID NO: 534 (VH CDR3); SEQ ID NO: 535 (VL); SEQ ID NO: 536 (VL CDR1); SEQ ID NO: 537 (VL CDR2); SEQ ID NO: 538 (VL CDR3)
[0512]015-102 antibody SEQ ID NO: 539 (VH); SEQ ID NO: 540 (VH CDR1); SEQ ID NO: 541 (VH CDR2); SEQ ID NO: 542 (VH CDR3); SEQ ID NO: 543 (VL); SEQ ID NO: 544 (VL CDR1); SEQ ID NO: 545 (VL CDR2); SEQ ID NO: 546 (VL CDR3)
[0513]015-136 antibody SEQ ID NO: 547 (VH); SEQ ID NO: 548 (VH CDR1); SEQ ID NO: 549 (VH CDR2); SEQ ID NO: 550 (VH CDR3); SEQ ID NO: 551 (VL); SEQ ID NO: 552 (VL CDR1); SEQ ID NO: 553 (VL CDR2); SEQ ID NO: 554 (VL CDR3)
[0514]015-143 antibody SEQ ID NO: 555 (VH); SEQ ID NO: 556 (VH CDR1); SEQ ID NO: 557 (VH CDR2); SEQ ID NO: 558 (VH CDR3); SEQ ID NO: 559 (VL); SEQ ID NO: 560 (VL CDR1); SEQ ID NO: 561 (VL CDR2); SEQ ID NO: 562 (VL CDR3)
[0515]015-209 antibody SEQ ID NO: 563 (VH); SEQ ID NO: 564 (VH CDR1); SEQ ID NO: 565 (VH CDR2); SEQ ID NO: 566 (VH CDR3); SEQ ID NO: 567 (VL); SEQ ID NO: 568 (VL CDR1); SEQ ID NO: 569 (VL CDR2); SEQ ID NO: 570 (VL CDR3)
[0516]039-016 antibody SEQ ID NO: 571 (VH); SEQ ID NO: 572 (VH CDR1); SEQ ID NO: 573 (VH CDR2); SEQ ID NO: 574 (VH CDR3); SEQ ID NO: 575 (VL); SEQ ID NO: 576 (VL CDR1); SEQ ID NO: 577 (VL CDR2); SEQ ID NO: 578 (VL CDR3)
[0517]053-216 antibody SEQ ID NO: 579 (VH); SEQ ID NO: 580 (VH CDR1); SEQ ID NO: 581 (VH CDR2); SEQ ID NO: 582 (VH CDR3); SEQ ID NO: 583 (VL); SEQ ID NO: 584 (VL CDR1); SEQ ID NO: 585 (VL CDR2); SEQ ID NO: 586 (VL CDR3)
[0518]075-024 antibody SEQ ID NO: 587 (VH); SEQ ID NO: 588 (VH CDR1); SEQ ID NO: 589 (VH CDR2); SEQ ID NO: 590 (VH CDR3); SEQ ID NO: 591 (VL); SEQ ID NO: 592 (VL CDR1); SEQ ID NO: 593 (VL CDR2); SEQ ID NO: 594 (VL CDR3)
[0519]075-110 antibody SEQ ID NO: 595 (VH); SEQ ID NO: 596 (VH CDR1); SEQ ID NO: 597 (VH CDR2); SEQ ID NO: 598 (VH CDR3); SEQ ID NO: 599 (VL); SEQ ID NO: 600 (VL CDR1); SEQ ID NO: 601 (VL CDR2); SEQ ID NO: 602 (VL CDR3)
[0520]086-032 antibody SEQ ID NO: 603 (VH); SEQ ID NO: 604 (VH CDR1); SEQ ID NO: 605 (VH CDR2); SEQ ID NO: 606 (VH CDR3); SEQ ID NO: 607 (VL); SEQ ID NO: 608 (VL CDR1); SEQ ID NO: 609 (VL CDR2); SEQ ID NO: 610 (VL CDR3)
[0521]086-035 antibody SEQ ID NO: 611 (VH); SEQ ID NO: 612 (VH CDR1); SEQ ID NO: 613 (VH CDR2); SEQ ID NO: 614 (VH CDR3); SEQ ID NO: 615 (VL); SEQ ID NO: 616 (VL CDR1); SEQ ID NO: 617 (VL CDR2); SEQ ID NO: 618 (VL CDR3)
[0522]086-036 antibody SEQ ID NO: 619 (VH); SEQ ID NO: 620 (VH CDR1); SEQ ID NO: 621 (VH CDR2); SEQ ID NO: 622 (VH CDR3); SEQ ID NO: 623 (VL); SEQ ID NO: 624 (VL CDR1); SEQ ID NO: 625 (VL CDR2); SEQ ID NO: 626 (VL CDR3)
[0523]086-061 antibody SEQ ID NO: 627 (VH); SEQ ID NO: 628 (VH CDR1); SEQ ID NO: 629 (VH CDR2); SEQ ID NO: 630 (VH CDR3); SEQ ID NO: 631 (VL); SEQ ID NO: 632 (VL CDR1); SEQ ID NO: 633 (VL CDR2); SEQ ID NO: 634 (VL CDR3)
[0524]086-138 antibody SEQ ID NO: 635 (VH); SEQ ID NO: 636 (VH CDR1); SEQ ID NO: 637 (VH CDR2); SEQ ID NO: 638 (VH CDR3); SEQ ID NO: 639 (VL); SEQ ID NO: 640 (VL CDR1); SEQ ID NO: 641 (VL CDR2); SEQ ID NO: 642 (VL CDR3)
[0525]086-182 antibody SEQ ID NO: 643 (VH); SEQ ID NO: 644 (VH CDR1); SEQ ID NO: 645 (VH CDR2); SEQ ID NO: 646 (VH CDR3); SEQ ID NO: 647 (VL); SEQ ID NO: 648 (VL CDR1); SEQ ID NO: 649 (VL CDR2); SEQ ID NO: 650 (VL CDR3)
[0526]As mentioned in the below-mentioned Examples, the relationships between these antibodies and pulmonary adenocarcinoma cell line Calu-3, ovarian cancer cell line SKOv3, and breast cancer cell line BT474 (based on the results of the cell line staining) are experimentally confirmed.
3. Antibody to CD46
[0527]A plurality of antibodies clones are obtained. Among them, antibodies having the same amino acid sequence are included. Finally 87 kinds of antibody clones are identified. As to the below-mentioned eight kinds of antibody clones, the sequences are analyzed.
[0528]035-224 antibody SEQ ID NO: 33 (VH); SEQ ID NO: 34 (VH CDR1); SEQ ID NO: 35 (VH CDR2); SEQ ID NO: 36 (VH CDR3); SEQ ID NO: 37 (VL); SEQ ID NO: 38 (VL CDR1); SEQ ID NO: 39 (VL CDR2); SEQ ID NO: 40 (VL CDR3)
[0529]045-011 antibody SEQ ID NO: 41 (VH); SEQ ID NO: 42 (VH CDR1); SEQ ID NO: 43 (VH CDR2); SEQ ID NO: 44 (VH CDR3); SEQ ID NO: 45 (VL); SEQ ID NO: 46 (VL CDR1); SEQ ID NO: 47 (VL CDR2); SEQ ID NO: 48 (VL CDR3)
[0530]051-144 antibody SEQ ID NO: 49 (VH); SEQ ID NO: 50 (VH CDR1); SEQ ID NO: 51 (VH CDR2); SEQ ID NO: 52 (VH CDR3); SEQ ID NO: 53 (VL); SEQ ID NO: 54 (VL CDR1); SEQ ID NO: 55 (VL CDR2); SEQ ID NO: 56 (VL CDR3)
[0531]052-053 antibody SEQ ID NO: 57 (VH); SEQ ID NO: 58 (VH CDR1); SEQ ID NO: 59 (VH CDR2); SEQ ID NO: 60 (VH CDR3); SEQ ID NO: 61 (VL); SEQ ID NO: 62 (VL CDR1); SEQ ID NO: 63 (VL CDR2); SEQ ID NO: 64 (VL CDR3)
[0532]052-073 antibody SEQ ID NO: 65 (VH); SEQ ID NO: 66 (VH CDR1); SEQ ID NO: 67 (VH CDR2); SEQ ID NO: 68 (VH CDR3); SEQ ID NO: 69 (VL); SEQ ID NO: 70 (VL CDR1); SEQ ID NO: 71 (VL CDR2); SEQ ID NO: 72 (VL CDR3)
[0533]053-049 antibody SEQ ID NO: 73 (VH); SEQ ID NO: 74 (VH CDR1); SEQ ID NO: 75 (VH CDR2); SEQ ID NO: 76 (VH CDR3); SEQ ID NO: 77 (VL); SEQ ID NO: 78 (VL CDR1); SEQ ID NO: 79 (VL CDR2); SEQ ID NO: 80 (VL CDR3)
[0534]3172-120 antibody SEQ ID NO: 81 (VH); SEQ ID NO: 82 (VH CDR1); SEQ ID NO: 83 (VH CDR2); SEQ ID NO: 84 (VH CDR3); SEQ ID NO: 85 (VL); SEQ ID NO: 86 (VL CDR1); SEQ ID NO: 87 (VL CDR2); SEQ ID NO: 88 (VL CDR3)
[0535]066-069 antibody SEQ ID NO: 755 (VH); SEQ ID NO: 756 (VH CDR1); SEQ ID NO: 757 (VH CDR2); SEQ ID NO: 758 (VH CDR3); SEQ ID NO: 759 (VL); SEQ ID NO: 760 (VL CDR1); SEQ ID NO: 761 (VL CDR2); SEQ ID NO: 762 (VL CDR3)
[0536]As mentioned in the below-mentioned Examples, the relationships between these antibodies and large bowel cancer cell line CaCo2, stomach cancer cell line MKN45, undifferentiated hepatic cell carcinoma cell line HLF, liver cancer cell line HepG2, intrahepatic bile duct cell cancer cell line RBE, pancreas cancer cell line PANC1, kidney cancer cell line CCFRC1, kidney cancer cell line Caki-1, lung cancer cell line NCI-H441, lung squamous cell cancer EBC1, stomach cancer cell line NCI-N87, stomach cancer cell line SNU-5, lung squamous cell carcinoma line RERF-LC-AI, hepatic cell carcinoma clinical specimen s, breast cancer cell line BT474, kidney cancer cell line 293T, pulmonary adenocarcinoma cell line PC14, kidney cancer cell line ACHN, and pulmonary adenocarcinoma cell line H1373 (as to the above mention, based on the results of the cell line staining), as well as the relationships between these kidney cancer, hepatic cell carcinoma, gallbladder and liver cancer, pulmonary adenocarcinoma, and pancreas cancer (as to the above mention, based on the results of the tissue staining) are experimentally confirmed.
4. Antibody to ITGA3
[0537]A plurality of antibodies clones are obtained. Among them, antibodies having the same amino acid sequence are included. As to the below-mentioned 13 kinds of antibody clones, the sequences are analyzed.
[0538]015-003 antibody SEQ ID NO: 89 (VH); SEQ ID NO: 90 (VH CDR1); SEQ ID NO: 91 (VH CDR2); SEQ ID NO: 92 (VH CDR3); SEQ ID NO: 93 (VL); SEQ ID NO: 94 (VL CDR1); SEQ ID NO: 95 (VL CDR2); SEQ ID NO: 96 (VL CDR3)
[0539]064-002 antibody SEQ ID NO: 675 (VH); SEQ ID NO: 676 (VH CDR1); SEQ ID NO: 677 (VH CDR2); SEQ ID NO: 678 (VH CDR3); SEQ ID NO: 679 (VL); SEQ ID NO: 680 (VL CDR1); SEQ ID NO: 681 (VL CDR2); SEQ ID NO: 682 (VL CDR3)
[0540]064-006 antibody SEQ ID NO: 683 (VH); SEQ ID NO: 684 (VH CDR1); SEQ ID NO: 685 (VH CDR2); SEQ ID NO: 686 (VH CDR3); SEQ ID NO: 687 (VL); SEQ ID NO: 688 (VL CDR1); SEQ ID NO: 689 (VL CDR2); SEQ ID NO: 690 (VL CDR3)
[0541]064-012a antibody SEQ ID NO: 691 (VH); SEQ ID NO: 692 (VH CDR1); SEQ ID NO: 693 (VH CDR2); SEQ ID NO: 694 (VH CDR3); SEQ ID NO: 695 (VL); SEQ ID NO: 696 (VL CDR1); SEQ ID NO: 697 (VL CDR2); SEQ ID NO: 698 (VL CDR3)
[0542]064-012b antibody SEQ ID NO: 699 (VH); SEQ ID NO: 700 (VH CDR1); SEQ ID NO: 701 (VH CDR2); SEQ ID NO: 702 (VH CDR3); SEQ ID NO: 703 (VL); SEQ ID NO: 704 (VL CDR1); SEQ ID NO: 705 (VL CDR2); SEQ ID NO: 706 (VL CDR3)
[0543]064-014 antibody SEQ ID NO: 707 (VH); SEQ ID NO: 708 (VH CDR1); SEQ ID NO: 709 (VH CDR2); SEQ ID NO: 710 (VH CDR3); SEQ ID NO: 711 (VL); SEQ ID NO: 712 (VL CDR1); SEQ ID NO: 713 (VL CDR2); SEQ ID NO: 714 (VL CDR3)
[0544]064-054 antibody SEQ ID NO: 715 (VH); SEQ ID NO: 716 (VH CDR1); SEQ ID NO: 717 (VH CDR2); SEQ ID NO: 718 (VH CDR3); SEQ ID NO: 719 (VL); SEQ ID NO: 720 (VL CDR1); SEQ ID NO: 721 (VL CDR2); SEQ ID NO: 722 (VL CDR3)
[0545]064-085 antibody SEQ ID NO: 723 (VH); SEQ ID NO: 724 (VH CDR1); SEQ ID NO: 725 (VH CDR2); SEQ ID NO: 726 (VH CDR3); SEQ ID NO: 727 (VL); SEQ ID NO: 728 (VL CDR1); SEQ ID NO: 729 (VL CDR2); SEQ ID NO: 730 (VL CDR3)
[0546]064-093 antibody SEQ ID NO: 731 (VH); SEQ ID NO: 732 (VH CDR1); SEQ ID NO: 733 (VH CDR2); SEQ ID NO: 734 (VH CDR3); SEQ ID NO: 735 (VL); SEQ ID NO: 736 (VL CDR1); SEQ ID NO: 737 (VL CDR2); SEQ ID NO: 738 (VL CDR3)
[0547]064-116 antibody SEQ ID NO: 739 (VH); SEQ ID NO: 740 (VH CDR1); SEQ ID NO: 741 (VH CDR2); SEQ ID NO: 742 (VH CDR3); SEQ ID NO: 743 (VL); SEQ ID NO: 744 (VL CDR1); SEQ ID NO: 745 (VL CDR2); SEQ ID NO: 746 (VL CDR3)
[0548]065-183 antibody SEQ ID NO: 747 (VH); SEQ ID NO: 748 (VH CDR1); SEQ ID NO: 749 (VH CDR2); SEQ ID NO: 750 (VH CDR3); SEQ ID NO: 751 (VL); SEQ ID NO: 752 (VL CDR1); SEQ ID NO: 753 (VL CDR2); SEQ ID NO: 754 (VL CDR3)
[0549]067-142 antibody SEQ ID NO: 763 (VH); SEQ ID NO: 764 (VH CDR1); SEQ ID NO: 765 (VH CDR2); SEQ ID NO: 766 (VH CDR3); SEQ ID NO: 767 (VL); SEQ ID NO: 768 (VL CDR1); SEQ ID NO: 769 (VL CDR2); SEQ ID NO: 770 (VL CDR3)
[0550]068-007 antibody SEQ ID NO: 771 (VH); SEQ ID NO: 772 (VH CDR1); SEQ ID NO: 773 (VH CDR2); SEQ ID NO: 774 (VH CDR3); SEQ ID NO: 775 (VL); SEQ ID NO: 776 (VL CDR1); SEQ ID NO: 777 (VL CDR2); SEQ ID NO: 778 (VL CDR3)
[0551]As mentioned in the below-mentioned Examples, the relationships between these antibodies and undifferentiated hepatic cell carcinoma cell line HLF, ovarian cancer cell line SKOv3, kidney cancer cell line ACHN, kidney cancer cell line Caki-1, pulmonary adenocarcinoma cell line H1373, lung squamous cell cancer EBC1, vulva mucosal epithelial cell line A431, breast cancer cell line BT474, pulmonary adenocarcinoma cell line PC14, kidney cancer cell line CCFRC1, hepatic cell carcinoma cell line OCTH, intrahepatic bile duct cell cancer RBE, pancreas cancer cell line PANC-1, pancreas cancer cell line MIA-Paca2, pulmonary adenocarcinoma cell line A549, pulmonary adenocarcinoma cell line NCI-N441, lung squamous cell carcinoma line Calu-3, lung squamous cell carcinoma line RERF-LC-AI, stomach cancer cell line SNU5, stomach cancer cell line MKN45, stomach cancer cell line NCI-N87, large bowel cancer cell line CW2, ovarian cancer cell line SKOv3, ovarian cancer cell line KF-28, ovarian cancer cell line RMG-1, and ovarian cancer cell line RMG-2 (as to the above mention, based on the results of the cell line staining), as well as the relationships between these antibodies and gallbladder and liver cancer and pancreas cancer (as to the above mention, based on the results of the tissue staining) are experimentally confirmed.
5. Antibody to ICAM1
[0552]A plurality of antibodies clones are obtained. Among them, antibodies having the same amino acid sequence are included. Finally, 22 kinds of antibody clones are identified. As to the below-mentioned five kinds of antibody clones, the sequences are analyzed.
[0553]052-033 antibody SEQ ID NO: 97 (VH); SEQ ID NO: 98 (VH CDR1); SEQ ID NO: 99 (VH CDR2); SEQ ID NO: 100 (VH CDR3); SEQ ID NO: 101 (VL); SEQ ID NO: 102 (VL CDR1); SEQ ID NO: 103 (VL CDR2); SEQ ID NO: 104 (VL CDR3)
[0554]053-042 antibody SEQ ID NO: 105 (VH); SEQ ID NO: 106 (VH CDR1); SEQ ID NO: 107 (VH CDR2); SEQ ID NO: 108 (VH CDR3); SEQ ID NO: 109 (VL); SEQ ID NO: 110 (VL CDR1); SEQ ID NO: 111 (VL CDR2); SEQ ID NO: 112 (VL CDR3)
[0555]053-051 antibody SEQ ID NO: 113 (VH); SEQ ID NO: 114 (VH CDR1); SEQ ID NO: 115 (VH CDR2); SEQ ID NO: 116 (VH CDR3); SEQ ID NO: 117 (VL); SEQ ID NO: 118 (VL CDR1); SEQ ID NO: 119 (VL CDR2); SEQ ID NO: 120 (VL CDR3)
[0556]053-059 antibody SEQ ID NO: 121 (VH); SEQ ID NO: 122 (VH CDR1); SEQ ID NO: 123 (VH CDR2); SEQ ID NO: 124 (VH CDR3); SEQ ID NO: 125 (VL); SEQ ID NO: 126 (VL CDR1); SEQ ID NO: 127 (VL CDR2); SEQ ID NO: 128 (VL CDR3)
[0557]053-085 antibody SEQ ID NO: 129 (VH); SEQ ID NO: 130 (VH CDR1); SEQ ID NO: 131 (VH CDR2); SEQ ID NO: 132 (VH CDR3); SEQ ID NO: 133 (VL); SEQ ID NO: 134 (VL CDR1); SEQ ID NO: 135 (VL CDR2); SEQ ID NO: 136 (VL CDR3)
[0558]As mentioned in the below-mentioned Examples, the relationships between these antibodies and liver cancer cell line HepG2, pulmonary adenocarcinoma cell line PC14, and cell line established from kidney clinical specimen (as to the above mention, based on the results of the cell line staining), as well as the relationships between these antibodies and hepatic cell carcinoma (as to the above mention, based on the results of the tissue staining) are experimentally confirmed.
6. Antibody to ALCAM
[0559]A plurality of antibodies clones are obtained. Among them, antibodies having the same amino acid sequence are included. As to the below-mentioned 13 kinds of antibody clones, the sequences are analyzed.
[0560]035-234 antibody SEQ ID NO: 137 (VH); SEQ ID NO: 138 (VH CDR1); SEQ ID NO: 139 (VH CDR2); SEQ ID NO: 140 (VH CDR3); SEQ ID NO: 141 (VL); SEQ ID NO: 142 (VL CDR1); SEQ ID NO: 143 (VL CDR2); SEQ ID NO: 144 (VL CDR3)
[0561]040-107 antibody SEQ ID NO: 145 (VH); SEQ ID NO: 146 (VH CDR1); SEQ ID NO: 147 (VH CDR2); SEQ ID NO: 148 (VH CDR3); SEQ ID NO: 149 (VL); SEQ ID NO: 150 (VL CDR1); SEQ ID NO: 151 (VL CDR2); SEQ ID NO: 152 (VL CDR3)
[0562]041-118 antibody SEQ ID NO: 153 (VH); SEQ ID NO: 154 (VH CDR1); SEQ ID NO: 155 (VH CDR2); SEQ ID NO: 156 (VH CDR3); SEQ ID NO: 157 (VL); SEQ ID NO: 158 (VL CDR1); SEQ ID NO: 159 (VL CDR2); SEQ ID NO: 160 (VL CDR3)
[0563]066-174 antibody SEQ ID NO: 161 (VH); SEQ ID NO: 162 (VH CDR1); SEQ ID NO: 163 (VH CDR2); SEQ ID NO: 164 (VH CDR3); SEQ ID NO: 165 (VL); SEQ ID NO: 166 (VL CDR1); SEQ ID NO: 167 (VL CDR2); SEQ ID NO: 168 (VL CDR3)
[0564]083-040 antibody SEQ ID NO: 169 (VH); SEQ ID NO: 170 (VH CDR1); SEQ ID NO: 171 (VH CDR2); SEQ ID NO: 172 (VH CDR3); SEQ ID NO: 173 (VL); SEQ ID NO: 174 (VL CDR1); SEQ ID NO: 175 (VL CDR2); SEQ ID NO: 176 (VL CDR3)
[0565]029-143 antibody SEQ ID NO: 779 (VH); SEQ ID NO: 780 (VH CDR1); SEQ ID NO: 781 (VH CDR2); SEQ ID NO 782 (VH CDR3); SEQ ID NO: 783 (VL); SEQ ID NO: 784 (VL CDR1); SEQ ID NO: 785 (VL CDR2); SEQ ID NO: 786 (VL CDR3)
[0566]045-134 antibody SEQ ID NO: 787 (VH); SEQ ID NO: 788 (VH CDR1); SEQ ID NO: 789 (VH CDR2); SEQ ID NO: 790 (VH CDR3); SEQ ID NO: 791 (VL); SEQ ID NO: 792 (VL CDR1); SEQ ID NO: 793 (VL CDR2); SEQ ID NO: 794 (VL CDR3)
[0567]062-101 antibody SEQ ID NO: 795 (VH); SEQ ID NO: 796 (VH CDR1); SEQ ID NO: 797 (VH CDR2); SEQ ID NO: 798 (VH CDR3); SEQ ID NO: 799 (VL); SEQ ID NO: 800 (VL CDR1); SEQ ID NO: 801 (VL CDR2); SEQ ID NO: 802 (VL CDR3)
[0568]062-109 antibody SEQ ID NO: 803 (VH); SEQ ID NO: 804 (VH CDR1); SEQ ID NO: 805 (VH CDR2); SEQ ID NO: 806 (VH CDR3); SEQ ID NO: 807 (VL); SEQ ID NO: 808 (VL CDR1); SEQ ID NO: 809 (VL CDR2); SEQ ID NO: 810 (VL CDR3)
[0569]084-103 antibody SEQ ID NO: 811 (VH); SEQ ID NO: 812 (VH CDR1); SEQ ID NO: 813 (VH CDR2); SEQ ID NO: 814 (VH CDR3); SEQ ID NO: 815 (VL); SEQ ID NO: 816 (VL CDR1); SEQ ID NO: 817 (VL CDR2); SEQ ID NO: 818 (VL CDR3)
[0570]052-274 antibody SEQ ID NO: 819 (VH); SEQ ID NO: 820 (VH CDR1); SEQ ID NO: 821 (VH CDR2); SEQ ID NO: 822 (VH CDR3); SEQ ID NO: 823 (VL); SEQ ID NO: 824 (VL CDR1); SEQ ID NO: 825 (VL CDR2); SEQ ID NO: 826 (VL CDR3)
[0571]029-067 antibody SEQ ID NO: 827 (VH); SEQ ID NO: 828 (VH CDR1); SEQ ID NO: 829 (VH CDR2); SEQ ID NO: 830 (VH CDR3); SEQ ID NO: 831 (VL); SEQ ID NO: 832 (VL CDR1); SEQ ID NO: 833 (VL CDR2); SEQ ID NO: 834 (VL CDR3)
[0572]083-131 antibody SEQ ID NO: 835 (VH); SEQ ID NO: 836 (VH CDR1); SEQ ID NO: 837 (VH CDR2); SEQ ID NO: 838 (VH CDR3); SEQ ID NO: 839 (VL); SEQ ID NO: 840 (VL CDR1); SEQ ID NO: 841 (VL CDR2); SEQ ID NO: 842 (VL CDR3)
[0573]As mentioned in the below-mentioned Examples, the relationships between these antibodies and liver cancer cell line (HepG2, OCTH, Hep3B, and HLF), kidney cancer cell line (Caki-1, CCFRC1, ACHN, 293T, and cell line established from the clinical specimen), lung cancer cell line (PC14, NCI-H441, EB. C-1, RERF-LC-AI, A549, and H1373), ovarian cancer cell line (SKOv3, KF-28, RMG1, and RMG2), stomach cancer cell line (NCI-N87), large bowel cancer cell line (CW2), breast cancer cell line (BT474), acute myelocytic leukemia AML clinical specimen, and hamster ovarian cancer cell line CHO (as to the above mention, based on the results of the cell line staining), as well as the relationships between these antibodies and kidney cancer, hepatic cell carcinoma, gallbladder and liver cancer, lung squamous cell cancer, alveolar cell carcinoma, and adenocarcinoma (as to the above mention, based on the results of the tissue staining) are experimentally confirmed.
7. Antibody to CD147
[0574]A plurality of antibodies clones are obtained. Among them, antibodies having the same amino acid sequence are included. As to the below-mentioned one kind of antibody clone, the sequence is analyzed.
[0575]059-053 antibody SEQ ID NO: 177 (VH); SEQ ID NO: 178 (VH CDR1); SEQ ID NO: 179 (VH CDR2); SEQ ID NO: 180 (VH CDR3); SEQ ID NO: 181 (VL); SEQ ID NO: 182 (VL CDR1); SEQ ID NO: 183 (VL CDR2); SEQ ID NO: 184 (VL CDR3)
[0576]As mentioned in the below-mentioned Examples, the relationships between this antibody the and liver cancer cell line HepG2, kidney cancer cell line CCFRC1, kidney cancer cell line ACHN, kidney cancer cell line Caki-1, pulmonary adenocarcinoma PC14, and cell line established from kidney cancer clinical specimen (as to the above mention, based on the results of the cell line staining), as well as the relationships between these antibodies and kidney cancer (as to the above mention, based on the results of the tissue staining) are experimentally confirmed.
8. Antibody to C1qR
[0577]A plurality of antibodies clones are obtained. Among them, antibodies having the same amino acid sequence are included. As to the below-mentioned one kind of antibody clone, the sequence is analyzed.
[0578]070-016 antibody SEQ ID NO: 451 (VH); SEQ ID NO: (VH CDR1) 452; SEQ ID NO: 453 (VH CDR2); SEQ ID NO: 454 (yH CDR3); SEQ ID NO: 455 (VL); SEQ ID NO: (VL CDR1) 456; SEQ ID NO: 457 (VL CDR2); SEQ ID NO: 458 (VL CDR3)
[0579]The relationship between this antibody and leukemia is experimentally confirmed. That is to say, in cell line staining using this antibody, leukemia AML cell line Nohno 1 and leukemia AML clinical specimen shows a strong positive property (MFI=20 or more). Furthermore, in the process of growing the leukemia cell line, this antibody is added to the growing temperature, rapid aggregation of cancer cells can be confirmed. Moreover, the antibody amount necessary to cause these phenomena is relatively low concentration.
9. Antibody to CD44
[0580]A plurality of antibodies clones are obtained. Among them, antibodies having the same amino acid sequence are included. As to the below-mentioned one kind of antibody clone, the sequence is analyzed.
[0581]064-003 antibody SEQ ID NO: 459 (VH); SEQ ID NO: 460 (VH CDR1); SEQ ID NO: 461 (VH CDR2); SEQ ID NO: 462 (VH CDR3); SEQ ID NO: 463 (VL); SEQ ID NO: 464 (VL CDR1); SEQ ID NO: 465 (VL CDR2); SEQ ID NO: 466 (VL CDR3)
[0582]The relationships between this antibody and liver cancer, lung cancer, ovarian cancer, and stomach cancer are experimentally confirmed. That is to say, in the cell staining using this antibody, hepatic cell carcinoma HLF, pulmonary adenocarcinoma cell line PC14, pulmonary adenocarcinoma cell line NCI-H1373, and ovary adenocarcinoma cell line SKOv3 show the strong positive property (MFI=20 or more), and epidermoid cancer cell line A431 and lung squamous cell cancer EBC1 show the weak positive property (MFI=3 or more). Furthermore, in immunostaining using this antibody, a case in which a pulmonary adenocarcinoma clinical specimen shows cancer specific stained image is observed, and cancer portions of alveolar cell carcinoma and lung squamous cell cancer show the weak positive property.
10. Antibody to CD73
[0583]A plurality of antibodies clones are obtained. Among them, antibodies having the same amino acid sequence are included. As to the below-mentioned one kind of antibody clone, the sequence is analyzed.
[0584]067-213 antibody SEQ ID NO: 467 (VH); SEQ ID NO: 468 (VH CDR1); SEQ ID NO: 469 (VH CDR2); SEQ ID NO: 470 (VH CDR3); SEQ ID NO: 471 (VL); SEQ ID NO: 472 (VL CDR1); SEQ ID NO: 473 (VL CDR2); SEQ ID NO: 474 (VL CDR3)
[0585]The relationships between this antibody and liver cancer, lung cancer, and ovarian cancer are experimentally confirmed. That is to say, in the cell staining using this antibody, pulmonary adenocarcinoma cell line NCI-H1373, and lung squamous cell cancer EBC1 show the strong positive property (MFI=20 or more), and liver cancer cell line HLF, ovary adenocarcinoma cell line SKOv3, and pulmonary adenocarcinoma cell line PC14 show the weak positive property (MFI=3 or more). Furthermore, in immunostaining using this antibody, a cancer-specific stained image is obtained in a pulmonary adenocarcinoma clinical specimen and a stained image showing the weak positive property to a cancer portion is obtained in lung squamous cell cancer.
11. Antibody to EpCAM
[0586]A plurality of antibodies clones are obtained. Among them, antibodies having the same amino acid sequence are included. As to the below-mentioned one kind of antibody clone, the sequence is analyzed.
[0587]067-153 antibody SEQ ID NO: 475 (VH); SEQ ID NO: 476 (VH CDR1); SEQ ID NO: 477 (VH CDR2); SEQ ID NO: 478 (VH CDR3); SEQ ID NO: 479 (VL); SEQ ID NO: 480 (VL CDR1); SEQ ID NO: 481 (VL CDR2); SEQ ID NO: 482 (VL CDR3)
[0588]The relationships between this antibody and liver cancer, lung cancer, ovarian cancer, stomach cancer, and large bowel cancer are experimentally confirmed. That is to say, in the cell staining using this antibody, pulmonary adenocarcinoma cell line NCI-H1373 and lung squamous cell carcinoma line LK-2 show the strong positive property (MFI=20 or more); lung squamous cell cancer EBC1 and pulmonary adenocarcinoma cell line PCI4 show the positive property (MFI=10 or more); and ovary adenocarcinoma cell line SKOv3 shows the weak positive property (MFI=3 or more). Furthermore, in immunostaining using this antibody, an extremely excellent cancer-specific stained image is obtained in each clinical specimen of large bowel cancer, pulmonary adenocarcinoma, lung squamous cell cancer, stomach cancer. A stained image having a weak cancer specific positive property is obtained in a part of hepatic cell carcinoma clinical specimens.
12. Antibody to HGFR
[0589]A plurality of antibodies clones are obtained. Among them, antibodies having the same amino acid sequence are included. Finally 87 kinds of antibody clones are identified. As to the below-mentioned three kinds of antibody clones, the sequences are analyzed.
[0590]067-126 antibody SEQ ID NO: 651 (VH); SEQ ID NO: 652 (VH CDR1); SEQ ID NO: 653 (VH CDR2); SEQ ID NO: 654 (VH CDR3); SEQ ID NO: 655 (VL); SEQ ID NO: 656 (VL CDR1); SEQ ID NO: 657 (VL CDR2); SEQ ID NO: 658 (VL CDR3)
[0591]067-133 antibody SEQ ID NO: 659 (VH); SEQ ID NO: 660 (VH CDR1); SEQ ID NO: 661 (VH CDR2); SEQ ID NO: 662 (VH CDR3); SEQ ID NO: 663 (VL); SEQ ID NO: 664 (VL CDR1); SEQ ID NO: 665 (VL CDR2); SEQ ID NO: 666 (VL CDR3)
[0592]067-287 antibody SEQ ID NO: 667 (VH); SEQ ID NO: 668 (VH CDR1); SEQ ID NO: 669 (VH CDR2); SEQ ID NO: 670 (VH CDR3); SEQ ID NO: 671 (VL); SEQ ID NO: 672 (VL CDR1); SEQ ID NO: 673 (VL CDR2); SEQ ID NO: 674 (VL CDR3)
[0593]The relationships between this antibody and lung cancer, liver cancer, ovarian cancer, large bowel cancer, and stomach cancer are experimentally confirmed. That is to say, in cell line staining using this antibody, lung squamous cell cancer EBC1 shows a strong positive property (MFI=20 or more); alveolar adenocarcinoma NCI-H1373 shows the positive property (MFI=10 or more); and epidermoid cancer cell line A431, ovary adenocarcinoma cell line SKOv3, pulmonary adenocarcinoma cell line PC14, and hepatic cell carcinoma HLF show the weak positive property (MFI=3 or more). Furthermore, in immunostaining using this antibody, a weak positive property to cancer portion in a part of lung squamous cell cancer clinical specimen is obtained.
13. Antibody to LAR
[0594]A plurality of antibodies clones are obtained. Among them, antibodies having the same amino acid sequence are included. As to the below-mentioned five kinds of antibody clones, the sequence is analyzed. [0595]064-044 antibody SEQ ID NO: 944 (VH); and SEQ ID NO: 945 (VL) 065-030 antibody SEQ ID NO: 946 (VH); and SEQ ID NO: 947 (VL) [0596]065-358 antibody SEQ ID NO: 948 (VH); and SEQ ID NO: 949 (VL) [0597]066-019 antibody SEQ ID NO: 950 (VH); and SEQ ID NO: 951 (VL) [0598]079-085 antibody SEQ ID NO: 952 (VH); and SEQ ID NO: 953 (VL)
[0599]In the immunostaining using these antibodies, a positive property is observed in a cancer portion in a part of the lung cancer clinical specimens.
14. Antibody to BCAM
[0600]A plurality of antibodies clones are obtained. Among them, antibodies having the same amino acid sequence are included. As to the below-mentioned one kind of antibody clone, the sequence is analyzed. [0601]067-024 antibody SEQ ID NO: 954 (VH); and SEQ ID NO: 955 (VL)
[0602]In the immunostaining using these antibodies, a positive property is observed in a cancer portion in a part of the clinical specimens of lung cancer, liver cancer, and kidney cancer.
[0603]The first embodiment of this aspect provides an isolated antibody having a specific binding property to HER1. The antibody of this form includes the heavy chain variable region CDR3 and the light chain variable region CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the group consisting of the following (1) to (3). Preferably, it includes the heavy chain variable regions CDR2 and CDR3 and the light chain variable regions CDR2 and CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the group consisting of the following (4) to (6). Furthermore, preferably, it includes the heavy chain variable regions CDR1 to CDR3 and the light chain variable regions CDR1 to CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2; and SEQ ID NO showing the amino acid sequence of the light chain variable region. CDR3) selected from the group consisting of the following (7) to (9) and (13) to (18). The most preferably, it includes the heavy chain variable region and the light chain variable region specified by the combination of SEQ ID NOs (SEQ ID NO showing the heavy chain variable region and SEQ ID NO showing the light chain variable region) selected from the group consisting of the following (10) to (12) and (19) to (24).
(Combination of CDR3)
(1) SEQ ID NO: 4, SEQ ID NO: 8
(2) SEQ ID NO: 12, SEQ ID NO: 16
(3) SEQ ID NO: 20, SEQ ID NO: 24
(Combination of CDR2 and CDR3)
(4) SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 7, SEQ ID NO: 8
(5) SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 15, SEQ ID NO: 16
(6) SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 23, SEQ ID NO: 24
(Combination of CDR1 to CDR3)
(7) SEQ ID NO: 2, SEQ ID NO: 3, SEQ ID NO: 4, SEQ ID NO: 6 SEQ ID NO: 7, SEQ ID NO: 8
(8) SEQ ID NO: 10, SEQ ID NO: 11, SEQ ID NO: 12, SEQ ID NO: 14, SEQ ID NO: 15, SEQ ID NO: 16
(9) SEQ ID NO: 18, SEQ ID NO: 19, SEQ ID NO: 20, SEQ ID NO: 22, SEQ ID NO: 23, SEQ ID NO: 24
(13) SEQ ID NO: 484 (VH CDR1), SEQ ID NO: 485 (VH CDR2), SEQ ID NO: 486 (VH CDR3), SEQ ID NO: 488 (VL CDR1), SEQ ID NO: 489 (VL CDR2), SEQ ID NO: 490 (VL CDR3)
(14) SEQ ID NO: 492 (VH CDR1), SEQ ID NO: 493 (VH CDR2), SEQ ID NO: 494 (VH CDR3), SEQ ID NO: 496 (VL CDR1), SEQ ID NO: 497 (VL CDR2), SEQ ID NO: 498 (VL CDR3)
(15), SEQ ID NO: 500 (VH CDR1), SEQ ID NO: 501 (VH CDR2), SEQ ID NO: 502 (VH CDR3), SEQ ID NO: 504 (VL CDR1), SEQ ID NO: 505 (VL CDR2), SEQ ID NO: 506 (VL CDR3)
(16) SEQ ID NO: 508 (VH CDR1), SEQ ID NO: 509 (VH CDR2), SEQ ID NO: 510 (VH CDR3), SEQ ID NO: 512 (VL CDR1), SEQ ID NO: 513 (VL CDR2), SEQ ID NO: 514 (VL CDR3)
(17) SEQ ID NO: 516 (VH CDR1), SEQ ID NO: 517 (VH CDR2), SEQ ID NO: 518 (VH CDR3), SEQ ID NO: 520 (VL CDR1), SEQ ID NO: 521 (VL CDR2), SEQ ID NO: 522 (VL CDR3)
(18) SEQ ID NO: 524 (VH CDR1), SEQ ID NO: 525 (VH CDR2), SEQ ID NO: 526 (VH CDR3), SEQ ID NO: 528 (VL CDR1), SEQ ID NO: 529 (VL CDR2), SEQ ID NO: 530 (VL CDR3)
(Combination of Heavy Chain Variable Region and Light Chain Variable Region)
(10) SEQ ID NO: 1, SEQ ID NO: 5
(11) SEQ ID NO: 9, SEQ ID NO: 13
(12) SEQ ID NO: 17, SEQ ID NO: 21
(19) SEQ ID NO: 483 (VH), SEQ ID NO: 487 (VL)
(20) SEQ ID NO: 491 (VH), SEQ ID NO: 495 (VL)
(21) SEQ ID NO: 499 (VH), SEQ ID NO: 503 (VL)
(22) SEQ ID NO: 507 (VH), SEQ ID NO: 511 (VL)
(23) SEQ ID NO: 515 (VH), SEQ ID NO: 519 (VL)
(24) SEQ ID NO: 523 (VH), SEQ ID NO: 527 (VL)
[0604]Note here that (1), (4), (7), and (10) correspond to 048-006 antibody; (2), (5), (8), and (11) correspond to 057-091 antibody; (3), (6), (9), and (12) correspond to 059-152 antibody; (13) and (19) correspond to 048-040 antibody; (14) and (20) correspond to 054-101 antibody; (15) and (21) correspond to 055-147 antibody; (16) and (22) correspond to 059-173 antibody; (17) and (23) correspond to 067-149 antibody; as well as (18) and (24) correspond to 067-176 antibody. Therefore, the antibody of the present invention is expected to have high specificity to HER1.
[0605]The second embodiment of this aspect provides an isolated antibody having a specific binding property to HER2. The antibody of this form includes the heavy chain variable region CDR3 and the light chain variable region CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) shown in the following (1). Preferably, it includes the heavy chain variable regions CDR2 and CDR3 and the light chain variable regions CDR2 and CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the following (2). Furthermore, preferably, it includes the heavy chain variable regions CDR1 to CDR3 and the light chain variable regions CDR1 to CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the group consisting of the following (3) and (5) to (19). The most preferably, it includes the heavy chain variable region and the light chain variable region specified by the combination of SEQ ID NOs (SEQ ID NO showing the heavy chain variable region and SEQ ID NO showing the light chain variable region) selected from the group consisting of the following (4) and (20) to (34).
(Combination of CDR3)
(1) SEQ ID NO: 28, SEQ ID NO: 32
(Combination of CDR2 and CDR3)
(2) SEQ ID NO: 27, SEQ ID NO: 28, SEQ ID NO: 31, SEQ ID NO: 32
(Combination of CDR1 to CDR3)
(3) SEQ ID NO: 26, SEQ ID NO: 27, SEQ ID NO: 28, SEQ ID NO: 30, SEQ ID NO: 31, SEQ ID NO: 32
(5) SEQ ID NO: 532 (VH CDR1), SEQ ID NO: 533 (VH CDR2), SEQ ID NO: 534 (VH CDR3), SEQ ID NO: 536 (VL CDR1), SEQ ID NO: 537 (VL CDR2), SEQ ID NO: 538 (VL CDR3)
(6) SEQ ID NO: 540 (VH CDR1), SEQ ID NO: 541 (VH CDR2), SEQ ID NO: 542 (VH CDR3), SEQ ID NO: 544 (VL CDR1), SEQ ID NO: 545 (VL CDR2), SEQ ID NO: 546 (VL CDR3)
(7) SEQ ID NO: 548 (VH CDR1), SEQ ID NO: 549 (VH CDR2), SEQ ID NO: 550 (VH CDR3), SEQ ID NO: 552 (VL CDR1), SEQ ID NO: 553 (VL CDR2), SEQ ID NO: 554 (VL CDR3)
(8) SEQ ID NO: 556 (VH CDR1), SEQ ID NO: 557 (VH CDR2), SEQ ID NO: 558 (VH CDR3), SEQ ID NO: 560 (VL CDR1), SEQ ID NO: 561 (VL CDR2), SEQ ID NO: 562 (VL CDR3)
(9) SEQ ID NO: 564 (VH CDR1), SEQ ID NO: 565 (VH CDR2), SEQ ID NO: 566 (VH CDR3), SEQ ID NO: 568 (VL CDR1), SEQ ID NO: 569 (VL CDR2), SEQ ID NO: 570 (VL CDR3)
(10) SEQ ID NO: 572 (VH CDR1), SEQ ID NO: 573 (VH CDR1), SEQ ID NO: 574 (VH CDR3), SEQ ID NO: 576 (VL CDR1), SEQ ID NO: 577 (VL CDR2), SEQ ID NO: 578 (VL CDR3)
(11) SEQ ID NO: 580 (VH CDR1), SEQ ID NO: 581 (VH CDR2), SEQ ID NO: 582 (VH CDR3), SEQ ID NO: 584 (VL CDR1), SEQ ID NO: 585 (VL CDR2), SEQ ID NO: 586 (VL CDR3)
(12) SEQ ID NO: 588 (VH CDR1), SEQ ID NO: 589 (VH CDR2), SEQ ID NO: 590 (VH CDR3), SEQ ID NO: 592 (VL CDR1), SEQ ID NO: 593 (VL CDR2), SEQ ID NO: 594 (VL CDR3)
(13) SEQ ID NO; 596 (VH CDR1), SEQ ID NO: 597 (VH CDR2), SEQ ID NO: 598 (VH CDR3), SEQ ID NO: 600 (VL CDR1), SEQ ID NO: 601 (VL CDR2), SEQ ID NO: 602 (VL CDR3)
(14) SEQ ID NO: 604 (VH CDR1), SEQ ID NO: 605 (VH CDR2), SEQ ID NO: 606 (VH CDR3), SEQ ID NO: 608 (VL CDR1), SEQ ID NO: 609 (VL CDR2), SEQ ID NO: 610 (VL CDR3)
(15) SEQ ID NO: 612 (VH CDR1), SEQ ID NO: 613 (VH CDR2), SEQ ID NO: 614 (VH CDR3), SEQ ID NO: 616 (VL CDR1), SEQ ID NO: 617 (VL CDR2), SEQ ID NO: 618 (VL CDR3)
(16) SEQ ID NO: 620 (VH CDR1), SEQ ID NO: 621 (VH CDR2), SEQ ID NO: 622 (VH CDR3), SEQ ID NO: 624 (VL CDR1), SEQ ID NO: 625 (VL CDR2), SEQ ID NO: 626 (VL CDR3)
(17) SEQ ID NO: 628 (VH CDR1), SEQ ID NO: 629 (VH CDR2), SEQ ID NO: 630 (VH CDR3), SEQ ID NO: 632 (VL CDR1), SEQ ID NO: 633 (VL CDR2), SEQ ID NO: 634 (VL CDR3)
(18) SEQ ID NO: 636 (VH CDR1), SEQ ID NO: 637 (VH CDR2), SEQ ID NO: 638 (VH CDR3), SEQ ID NO: 640 (VL CDR1), SEQ ID NO: 641 (VL CDR2), SEQ ID NO: 642 (VL CDR3)
(19) SEQ ID NO: 644 (VH CDR1), SEQ ID NO: 645 (VH CDR2), SEQ ID NO: 646 (VH CDR3), SEQ ID NO: 648 (VL CDR1), SEQ ID NO: 649 (VL CDR2), SEQ ID NO: 650 (VL CDR3)
(Combination of Heavy Chain Variable Region and Light Chain Variable Region)
(4) SEQ ID NO: 25, SEQ ID NO: 29
(20) SEQ ID NO: 531 (VH), SEQ ID NO: 535 (VL)
(21) SEQ ID NO: 539 (VH), SEQ ID NO: 543 (VL)
(22) SEQ ID NO: 547 (VH), SEQ ID NO: 551 (VL)
(23) SEQ ID NO: 555 (VH), SEQ ID NO: 559 (VL)
(24) SEQ ID NO: 563 (VH), SEQ ID NO: 567 (VL)
(25) SEQ ID NO: 571 (VH), SEQ ID NO: 575 (VL)
(26) SEQ ID NO: 579 (VH), SEQ ID NO: 583 (VL)
(27) SEQ ID NO: 587 (VH), SEQ ID NO: 591 (VL)
(28) SEQ ID NO: 595 (VH), SEQ ID NO: 599 (VL)
(29) SEQ ID NO: 603 (VH), SEQ ID NO: 607 (VL)
(30) SEQ ID NO: 611 (VH), SEQ ID NO: 615 (VL)
(31) SEQ ID NO: 619 (VH), SEQ ID NO: 623 (VL)
(32) SEQ ID NO: 627 (VH), SEQ ID NO: 631 (VL)
(33) SEQ ID NO: 635 (VH), SEQ ID NO: 639 (VL)
(34) SEQ ID NO: 643 (VH), SEQ ID NO: 647 (VL)
[0606]Note here that (1) to (4) correspond to 015-126 antibody; (5) and (20) correspond to 015-044 antibody; (6) and (21) correspond to 015-102 antibody; (7) and (22) correspond to 015-136 antibody; (8) and (23) correspond to 015-143 antibody; (9) and (24) correspond to 015-209 antibody; (10) and (25) correspond to 039-016 antibody; (11) and (26) correspond to 053-216 antibody; (12) and (27) correspond to 075-024 antibody; (13) and (28) correspond to 075-110 antibody; (14), (29) correspond to 086-032 antibody; (15) and (30) correspond to 086-035 antibody; (16) and (31) correspond to 086-036 antibody; (17) and (32) correspond to 086-061 antibody; (18) and (33) correspond to 086-138 antibody; as well as (19) and (34) correspond to 086-182 antibody. Therefore, the antibody of the present invention is expected to have high specificity to HER2.
[0607]The third embodiment of this aspect provides an isolated antibody having a specific binding property to CD46 antigen. The antibody of this form includes the heavy chain variable region CDR3 and the light chain variable region CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the following the group consisting of (1) to (7). Preferably, it includes the heavy chain variable regions CDR2 and CDR3 and the light chain variable regions CDR2 and CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the following the group consisting of (8) to (14). Furthermore preferably, it includes the heavy chain variable regions CDR1 to CDR3 and the light chain variable regions CDR1 to CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR 1, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the following the group consisting of (15) to (21). The most preferably, it includes the heavy chain variable region and the light chain variable region specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region and SEQ ID NO showing the amino acid sequence of the light chain variable region) selected from the following the group consisting of (22) to (28).
(Combination of CDR3)
(1) SEQ ID NO: 36, SEQ ID NO: 40
(2) SEQ ID NO: 44, SEQ ID NO: 48
(3) SEQ ID NO: 52, SEQ ID NO: 56
(4) SEQ ID NO: 60, SEQ ID NO: 64
(5) SEQ ID NO: 68, SEQ ID NO: 72
(6) SEQ ID NO: 76, SEQ ID NO: 80
(7) SEQ ID NO: 84, SEQ ID NO: 88
(Combination of CDR2 and CDR3)
(8) SEQ ID NO: 35, SEQ ID NO: 36, SEQ ID NO: 39, SEQ ID NO: 40
(9) SEQ ID NO: 43, SEQ ID NO: 44, SEQ ID NO: 47, SEQ ID NO: 48
(10) SEQ ID NO: 51, SEQ ID NO: 52, SEQ ID NO: 55, SEQ ID NO: 56
(11) SEQ ID NO: 59, SEQ ID NO: 60, SEQ ID NO: 63, SEQ ID NO: 64
(12) SEQ ID NO: 67, SEQ ID NO: 68, SEQ ID NO: 71, SEQ ID NO: 72
(13) SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 79, SEQ ID NO: 80
(14) SEQ ID NO: 83, SEQ ID NO: 84, SEQ ID NO: 87, SEQ ID NO: 88
(Combination of CDR1 to CDR3)
(15) SEQ ID NO: 34, SEQ ID NO: 35, SEQ ID NO: 36, SEQ ID NO: 38, SEQ ID NO: 39, SEQ ID NO: 40
(16) SEQ ID NO: 42, SEQ ID NO: 43, SEQ ID NO: 44, SEQ ID NO: 46, SEQ ID NO: 47, SEQ ID NO: 48
(17) SEQ ID NO: 50, SEQ ID NO: 51, SEQ ID NO: 52, SEQ ID NO: 54, SEQ ID NO: 55, SEQ ID NO: 56
(18) SEQ ID NO: 58, SEQ ID NO: 59, SEQ ID NO: 60, SEQ ID NO: 62, SEQ ID NO: 63, SEQ ID NO: 64
(19) SEQ ID NO: 66, SEQ ID NO: 67, SEQ ID NO: 68, SEQ ID NO: 70, SEQ ID NO: 71, SEQ ID NO: 72
(20) SEQ ID NO: 74, SEQ ID NO: 75, SEQ ID NO: 76, SEQ ID NO: 78, SEQ ID NO: 79, SEQ ID NO: 80
(21) SEQ ID NO: 82, SEQ ID NO: 83, SEQ ID NO: 84, SEQ ID NO: 86, SEQ ID NO: 87, SEQ ID NO: 88
(22) SEQ ID NO: 756 (VH CDR1), SEQ ID NO: 757 (VH CDR2), SEQ ID NO: 758 (VH CDR3), SEQ ID NO: 760 (VL CDR1), SEQ ID NO: 761 (VL CDR2), SEQ ID NO: 762 (VL CDR3)
(Combination of Heavy Chain Variable Region and Light Chain Variable Region)
(23) SEQ ID NO: 33, SEQ ID NO: 37
(24) SEQ ID NO: 41, SEQ ID NO: 45
(25) SEQ ID NO: 49, SEQ ID NO: 53
(26) SEQ ID NO: 57, SEQ ID NO: 61
(27) SEQ ID NO: 65, SEQ ID NO: 69
(28) SEQ ID NO: 73, SEQ ID NO: 77
(29) SEQ ID NO: 81, SEQ ID NO: 85
(30) SEQ ID NO: 755 (VH), SEQ ID NO: 759 (VL)
[0608]Note here that (1), (8), (15) and (23) correspond to 035-224 antibody; (2), (9), (16), and (24) correspond to 045-011 antibody; (3), (10), (17), and (25) correspond to 051-144 antibody; (4), (11), (18), and (26) correspond to 052-053 antibody; (5), (12), (19), and (27) correspond to 052-073 antibody; (6), (13), (20), and (28) correspond to 053-049 antibody; (7), (14), (21), and (29) correspond to 3172-120 antibody; as well as (22) and (30) correspond to 066-069 antibody. Therefore, the antibody of the present invention is expected to have high specificity to a CD46 antigen.
[0609]The fourth embodiment of this aspect provides an isolated antibody having a specific binding property to ITGA3. The antibody of this form includes the heavy chain variable region CDR3 and the light chain variable region CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) shown in the following (1). Preferably, it includes the heavy chain variable regions CDR2 and CDR3 and the light chain variable regions CDR2 and CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) shown in the following (2). Furthermore, preferably, it includes the heavy chain variable regions CDR1 to CDR3 and the light chain variable regions CDR1 to CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the group consisting of the following (3) and (5) to (17). The most preferably, it includes the heavy chain variable region and the light chain variable region specified by the combination of SEQ ID NOs (SEQ ID NO showing the heavy chain variable region and SEQ ID NO showing the light chain variable region) selected from the group consisting of the following (4) and (18) to (30).
(Combination of CDR3)
(1) SEQ ID NO: 92, SEQ ID NO: 96
(Combination of CDR2 and CDR3)
(2) SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 95, SEQ ID NO: 96
(Combination of CDR 1 to CDR3)
(3) SEQ ID NO: 90, SEQ ID NO: 91, SEQ ID NO: 92, SEQ ID NO: 94, SEQ ID NO: 95, SEQ ID NO: 96
(5) SEQ ID NO: 676 (VH CDR1), SEQ ID NO: 677 (VH CDR2), SEQ ID NO: 678 (VH CDR3), SEQ ID NO: 680 (VL CDR1), SEQ ID NO: 681 (VL CDR2), SEQ ID NO: 682 (VL CDR3)
(6) SEQ ID NO: 684 (VH CDR1), SEQ ID NO: 685 (VH CDR2), SEQ ID NO: 686 (VH CDR3), SEQ ID NO: 688 (VL CDR1), SEQ ID NO: 689 (VL CDR2), SEQ ID NO: 690 (VL CDR3)
(7) SEQ ID NO: 692 (VH CDR1), SEQ ID NO: 693 (VH CDR2), SEQ ID NO: 694 (VH CDR3), SEQ ID NO: 696 (VL CDR1), SEQ ID NO: 697 (VL CDR2), SEQ ID NO: 698 (VL CDR3)
(8) SEQ ID NO: 700 (VH CDR1), SEQ ID NO: 701 (VH CDR2), SEQ ID NO: 702 (VH CDR3), SEQ ID NO: 704 (VL CDR1), SEQ ID NO: 705 (VL CDR2), SEQ ID NO: 706 (VL CDR3)
(9) SEQ ID NO: 708 (VH CDR1), SEQ ID NO: 709 (VH CDR2), SEQ ID NO: 710 (VH CDR3), SEQ ID NO: 712 (VL CDR1), SEQ ID NO: 713 (VL CDR2), SEQ ID NO: 714 (VL CDR3)
(10) SEQ ID NO: 716 (VH CDR1), SEQ ID NO: 717 (VH CDR2), SEQ ID NO: 718 (VH CDR3), SEQ ID NO: 720 (VL CDR1), SEQ ID NO: 721 (VL CDR2), SEQ ID NO: 722 (VL CDR3)
(11) SEQ ID NO: 724 (VH CDR1), SEQ ID NO: 725 (VH CDR2), SEQ ID NO: 726 (VH CDR3), SEQ ID NO: 728 (VL CDR1), SEQ ID NO: 729 (VL CDR2), SEQ ID NO: 730 (VL CDR3)
(12) SEQ ID NO: 732 (VH CDR1), SEQ ID NO: 733 (VH CDR2), SEQ ID NO: 734 (VH CDR3), SEQ ID NO: 736 (VL CDR1), SEQ ID NO: 737 (VL CDR2), SEQ ID NO: 738 (VL CDR3)
(13) SEQ ID NO: 740 (VH CDR1), SEQ ID NO: 741 (VH CDR2), SEQ ID NO: 742 (VH CDR3), SEQ ID NO: 744 (VL CDR1), SEQ ID NO: 745 (VL CDR2), SEQ ID NO: 746 (VL CDR3)
(14) SEQ ID NO: 748 (VH CDR1), SEQ ID NO: 749 (VH CDR2), SEQ ID NO: 750 (VH CDR3), SEQ ID NO: 752 (VL CDR1), SEQ ID NO: 753 (VL CDR2), SEQ ID NO: 754 (VL CDR3)
(15) SEQ ID NO: 764 (VH CDR1), SEQ ID NO: 765 (VH CDR2), SEQ ID NO: 766 (VH CDR3), SEQ ID NO: 768 (VL CDR1), SEQ ID NO: 769 (VL CDR2), SEQ ID NO: 770 (VL CDR3)
(16) SEQ ID NO: 772 (VH CDR1), SEQ ID NO: 773 (VH CDR2), SEQ ID NO: 774 (VH CDR3), SEQ ID NO: 776 (VL CDR1), SEQ ID NO: 777 (VL CDR2), SEQ ID NO: 778 (VL CDR3)
(Combination of Heavy Chain Variable Region and Light Chain Variable Region)
(4) SEQ ID NO: 89, SEQ ID NO: 93
(17) SEQ ID NO: 675 (VH), SEQ ID NO: 679 (VL)
(18) SEQ ID NO: 683 (VH), SEQ ID NO: 687 (VL)
(19) SEQ ID NO: 691 (VH), SEQ ID NO: 695 (VL)
(20) SEQ ID NO: 699 (VH), SEQ ID NO: 703 (VL)
(21) SEQ ID NO: 707 (VH), SEQ ID NO: 711 (VL)
(22) SEQ ID NO: 715 (VH), SEQ ID NO: 719 (VL)
(23) SEQ ID NO: 723 (VH), SEQ ID NO: 727 (VL)
(24) SEQ ID NO: 731 (VH), SEQ ID NO: 735 (VL)
(25) SEQ ID NO: 739 (VH), SEQ ID NO: 743 (VL)
(26) SEQ ID NO: 747 (VH), SEQ ID NO: 751 (VL)
(27) SEQ ID NO: 763 (VH), SEQ ID NO: 767 (VL)
(28) SEQ ID NO: 771 (VH), SEQ ID NO. 775 (VL)
[0610]Note here that (1) to (4) correspond to 015-003 antibody; (5) and (17) correspond to 064-002 antibody; (6) and (18) correspond to 064-006 antibody; (7) and (19) correspond to 064-012a antibody; (8) and (20) correspond to 064-012b antibody; (9) and (21) correspond to 064-014 antibody; (10) and (22) correspond to 064-054 antibody; (11) and (23) correspond to 064-085 antibody; (12) and (24) correspond to 064-093 antibody; (13) and (25) correspond to 064-116 antibody; (14) and (26) correspond to 065-183 antibody; (15) and (27) correspond to 067-142 antibody; as well as (16) and (28) correspond to 068-007 antibody. Therefore, the antibody of the present invention is expected to have high specificity to ITGA3.
[0611]The fifth embodiment of this aspect provides an isolated antibody having a specific binding property to ICAM1. The antibody of this form includes the heavy chain variable region CDR3 and the light chain variable region CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the following the group consisting of (1) to (5). Preferably, it includes the heavy chain variable regions CDR2 and CDR3 and the light chain variable regions CDR2 and CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the following the group consisting of (6) to (10). Furthermore preferably, it includes the heavy chain variable regions CDR1 to CDR3 and the light chain variable regions CDR1 to CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the following the group consisting of (11) to (15). The most preferably, it includes the heavy chain variable region and the light chain variable region specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region and SEQ ID NO showing the amino acid sequence of the light chain variable region) selected from the following the group consisting of (16) to (20).
(Combination of CDR3)
(1) SEQ ID NO: 100, SEQ ID NO: 104
(2) SEQ ID NO: 108, SEQ ID NO: 112
(3) SEQ ID NO: 116, SEQ ID NO: 120
(4) SEQ ID NO: 124, SEQ ID NO: 128
(5) SEQ ID NO: 132, SEQ ID NO: 136
(Combination of CDR2 and CDR3)
(6) SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 103, SEQ ID NO: 104
(7) SEQ ID NO: 107, SEQ ID NO: 108, SEQ ID NO: 111, SEQ ID NO: 112
(8) SEQ ID NO: 115, SEQ ID NO: 116, SEQ ID NO: 119, SEQ ID NO: 120
(9) SEQ ID NO: 123, SEQ ID NO: 124, SEQ ID NO: 127, SEQ ID NO: 128
(10) SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 135, SEQ ID NO: 136
(Combination of CDR1 to CDR3)
(11) SEQ ID NO: 98, SEQ ID NO: 99, SEQ ID NO: 100, SEQ ID NO: 102, SEQ ID NO: 103, SEQ ID NO: 104
(12) SEQ ID NO: 106, SEQ ID NO: 107, SEQ ID NO: 108, SEQ ID NO: 110, SEQ ID NO: 111, SEQ ID NO: 112
(13) SEQ ID NO: 114, SEQ ID NO: 115, SEQ ID NO: 116, SEQ ID NO: 118, SEQ ID NO: 119, SEQ ID NO: 120
(14) SEQ ID NO: 122, SEQ ID NO: 123, SEQ ID NO: 124, SEQ ID NO: 126, SEQ ID NO: 127, SEQ ID NO: 128
(15) SEQ ID NO: 130, SEQ ID NO: 131, SEQ ID NO: 132, SEQ ID NO: 134, SEQ ID NO: 135, SEQ ID NO: 136
[0612](Combination of heavy chain variable region and light chain variable region)
(16) SEQ ID NO: 97, SEQ ID NO: 101
(17) SEQ ID NO: 105, SEQ ID NO: 109
(18) SEQ ID NO: 113, SEQ ID NO: 117
(19) SEQ ID NO: 121, SEQ ID NO: 125
(20) SEQ ID NO: 129, SEQ ID NO: 133
[0613]Note here that (1), (6), (11) and (16) correspond to 052-033 antibody; (2), (7), (12), and (17) correspond to 053-042 antibody; (3), (8), (13), and (18) correspond to 053-051 antibody; (4), (9), (14), and (19) correspond to 053-059 antibody; as well as (5), (10), (15), and (20) correspond to 053-085 antibody. Therefore, the antibody of the present invention is expected to have high specificity to ICAM1.
[0614]The sixth embodiment of this aspect provides an isolated antibody having a specific binding property to ALCAM. The antibody of this form includes the heavy chain variable region CDR3 and the light chain variable region CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the group consisting of the following (1) to (5). Preferably, it includes the heavy chain variable regions CDR2 and CDR3 and the light chain variable regions CDR2 and CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the group consisting of the following (6) to (10). Furthermore, preferably, it includes the heavy chain variable regions CDR1 to CDR3 and the light chain variable regions CDR1 to CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the group consisting of the following (11) to (15) and (21) to (28). The most preferably, it includes the heavy chain variable region and the light chain variable region specified by the combination of SEQ ID NOs (SEQ ID NO showing the heavy chain variable region and SEQ ID NO showing the light chain variable region) selected from the group consisting of the following (16) to (20) and (29) to (36).
(Combination of CDR3)
(1) SEQ ID NO: 140, SEQ ID NO: 144
(2) SEQ ID NO: 148, SEQ ID NO: 152
(3) SEQ ID NO: 156, SEQ ID NO: 160
(4) SEQ ID NO: 164, SEQ ID NO: 168
(5) SEQ ID NO: 172, SEQ ID NO: 176
(Combination of CDR2 and CDR3)
(6) SEQ ID NO: 139, SEQ ID NO: 140, SEQ ID NO: 143, SEQ ID NO: 144
(7) SEQ ID NO: 147, SEQ ID NO: 148, SEQ ID NO: 151, SEQ ID NO: 152
(8) SEQ ID NO: 155, SEQ ID NO: 156, SEQ ID NO: 159, SEQ ID NO: 160
(9) SEQ ID NO: 163, SEQ ID NO: 164, SEQ ID NO: 167, SEQ ID NO: 168
(10) SEQ ID NO: 171, SEQ ID NO: 172, SEQ ID NO: 175, SEQ ID NO: 176
(Combination of CDR1 to CDR3)
(11) SEQ ID NO: 138, SEQ ID NO: 139, SEQ ID NO: 140, SEQ ID NO: 142, SEQ ID NO: 143, SEQ ID NO: 144
(12) SEQ ID NO: 146, SEQ ID NO: 147, SEQ ID NO: 148, SEQ ID NO: 150, SEQ ID NO: 151, SEQ ID NO: 152
(13) SEQ ID NO: 154, SEQ ID NO: 155, SEQ ID NO: 156, SEQ ID NO: 158, SEQ ID NO: 159, SEQ ID NO: 160
(14) SEQ ID NO: 162, SEQ ID NO: 163, SEQ ID NO: 164, SEQ ID NO: 166, SEQ ID NO: 167, SEQ ID NO: 168
(15) SEQ ID NO: 170, SEQ ID NO: 171, SEQ ID NO: 172, SEQ ID NO: 174, SEQ ID NO: 175, SEQ ID NO: 176
(21) SEQ ID NO: 780 (VH CDR1), SEQ ID NO: 781 (VH CDR2), SEQ ID NO 782 (VH CDR3), SEQ ID NO: 784 (VL CDR1), SEQ ID NO: 785 (VL CDR2), SEQ ID NO: 786 (VL CDR3)
(22) SEQ ID NO: 788 (VH CDR1), SEQ ID NO: 789 (VH CDR2), SEQ ID NO: 790 (VH CDR3), SEQ ID NO: 792 (VL CDR1), SEQ ID NO: 793 (yL CDR2), SEQ ID NO: 794 (VL CDR3)
(23) SEQ ID NO: 796 (VH CDR1), SEQ ID NO: 797 (VH CDR2), SEQ ID NO: 798 (VH CDR3), SEQ ID NO: 800 (VL CDR1), SEQ ID NO: 801 (VL CDR2), SEQ ID NO: 802 (VL CDR3)
(24) SEQ ID NO: 804 (VH CDR1), SEQ ID NO: 805 (VH CDR2), SEQ ID NO: 806 (VH CDR3), SEQ ID NO: 808 (VL CDR1), SEQ ID NO: 809 (VL CDR2), SEQ ID NO: 810 (VL CDR3)
(25) SEQ ID NO: 812 (VH CDR1), SEQ ID NO: 813 (VH CDR2), SEQ ID NO: 814 (VH CDR3), SEQ ID NO: 816 (VL CDR1), SEQ ID NO: 817 (VL CDR2), SEQ ID NO: 818 (VL CDR3)
(26) SEQ ID NO: 820 (VH CDR1), SEQ ID NO: 821 (VH CDR2), SEQ ID NO: 822 (VH CDR3), SEQ ID NO: 824 (VL CDR1), SEQ ID NO: 825 (VL CDR2), SEQ ID NO: 826 (VL CDR3)
(27) SEQ ID NO: 828 (VH CDR1), SEQ ID NO: 829 (VH CDR2), SEQ ID NO: 830 (VH CDR3), SEQ ID NO: 832 (VL CDR1), SEQ ID NO: 833 (VL CDR2), SEQ ID NO: 834 (VL CDR3)
(28) SEQ ID NO: 836 (VH CDR1), SEQ ID NO: 837 (VH CDR2), SEQ ID NO: 838 (VH CDR3), SEQ ID NO: 840 (VL CDR1), SEQ ID NO: 841 (VL CDR2), SEQ ID NO: 842 (VL CDR3)
(Combination of Heavy Chain Variable Region and Light Chain Variable Region)
(16) SEQ ID NO: 137, SEQ ID NO: 141
(17) SEQ ID NO: 145, SEQ ID NO: 149
(18) SEQ ID NO: 153, SEQ ID NO: 157
(19) SEQ ID NO: 161, SEQ ID NO: 165
(20) SEQ ID NO: 169, SEQ ID NO: 173
(29) SEQ ID NO: 779 (VH), SEQ ID NO: 783 (VL)
(30) SEQ ID NO: 787 (VH), SEQ ID NO: 791 (VL)
(31) SEQ ID NO: 795 (VH), SEQ ID NO: 799 (VL)
(32) SEQ ID NO: 803 (VH), SEQ ID NO: 807 (VL)
(33) SEQ ID NO: 811 (VH), SEQ ID NO: 815 (VL)
(34) SEQ ID NO: 819 (VH), SEQ ID NO: 823 (VL)
(35) SEQ ID NO: 827 (VH), SEQ ID NO: 831 (VL)
(36) SEQ ID NO: 835 (VH), SEQ ID NO: 839 (VL)
[0615]Note here that (1), (6), (11), and (16) correspond to 035-234 antibody; (2), (7), (12), and (17) correspond to 040-107 antibody; (3), (8), (13), and (18) correspond to 041-118 antibody; (4), (9), (14), and (19) correspond to 066-174 antibody; (5), (10), (15), and (20) correspond to 083-040 antibody; (21) and (29) correspond to 029-143 antibody; (22) and (30) correspond to 045-134 antibody; (23) and (31) correspond to 062-101 antibody; (24) and (32) correspond to 062-109 antibody; (25) and (33) correspond to 084-103 antibody; (26) and (34) correspond to 052-274 antibody; (27) and (35) correspond to 029-067 antibody; as well as (28) and (36) correspond to 083-131 antibody. Therefore, the antibody of the present invention is expected to have high specificity to ALCAM.
[0616]The seventh embodiment of this aspect provides an isolated antibody having a specific binding property to a CD147 antigen. The antibody of this form includes the heavy chain variable region CDR3 and the light chain variable region CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) shown in the following (1). Preferably, it includes the heavy chain variable regions CDR2 and CDR3 and the light chain variable regions CDR2 and CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the following (2). Furthermore, preferably, it includes the heavy chain variable regions CDR1 to CDR3 and the light chain variable regions CDR1 to CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2, and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) shown in the following (3). The most preferably, it includes the heavy chain variable region and the light chain variable region specified by the combination of SEQ ID NOs (SEQ ID NO showing the heavy chain variable region and SEQ ID NO showing the light chain variable region) shown in the following (4).
(Combination of CDR3)
(1) SEQ ID NO: 180, SEQ ID NO: 184
(Combination of CDR2 and CDR3)
(2) SEQ ID NO: 179, SEQ ID NO: 180, SEQ ID NO: 183, SEQ ID NO: 184
(Combination of CDR1 to CDR3)
(3) SEQ ID NO: 178, SEQ ID NO: 179, SEQ ID NO: 180, SEQ ID NO: 182, SEQ ID NO: 183, SEQ ID NO: 184
[0617](Combination of heavy chain variable region and light chain variable region)
(4) SEQ ID NO: 177, SEQ ID NO: 181
[0618]Note here that (1) to (4) correspond to 059-053 antibody. Therefore, the antibody of the present invention is expected to have high specificity to a CD147 antigen.
[0619]The eighth embodiment of this aspect provides an isolated antibody having a specific binding property to C1qR. The antibody of this form includes the heavy chain variable regions CDR1 to CDR3 and the light chain variable regions CDR1 to CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2 and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) shown in the following (1). Preferably, it includes the heavy chain variable region and the light chain variable region specified by the combination of SEQ ID NOs (SEQ ID NO showing the heavy chain variable region and SEQ ID NO showing the light chain variable region) shown in the following (2).
(Combination of CDR3)
(1) SEQ ID NO: (VH CDR1) 452, SEQ ID NO: 453 (VH CDR2), SEQ ID NO: 454 (VH CDR3), SEQ ID NO: (VL CDR1) 456, SEQ ID NO: 457 (VL CDR2), SEQ ID NO: 458 (VL CDR3)
(Combination of Heavy Chain Variable Region and Light Chain Variable Region)
(2) SEQ ID NO: 451 (VH), SEQ ID NO: 455 (VL)
[0620]Note here that (1) and (2) correspond to 070-016 antibody. Therefore, the antibody of the present invention is expected to have high specificity to C1qR.
[0621]The ninth embodiment of this aspect provides an isolated antibody having a specific binding property to CD44. The antibody of this form includes the heavy chain variable regions CDR1 to CDR3 and the light chain variable regions CDR1 to CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2 and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) shown in the following (1). Preferably, it includes the heavy chain variable region and the light chain variable region specified by the combination of SEQ ID NOs (SEQ ID NO showing the heavy chain variable region and SEQ ID NO showing the light chain variable region) shown in the following (2).
(Combination of CDR1 to CDR3)
(1) SEQ ID NO: 460 (VH CDR1), SEQ ID NO: 461 (VH CDR2), SEQ ID NO: 462 (VH CDR3), SEQ ID NO: 464 (VL CDR1), SEQ ID NO: 465 (VL CDR2), SEQ ID NO: 466 (VL CDR3)
(Combination of Heavy Chain Variable Region and Light Chain Variable Region)
(2) SEQ ID NO: 459 (VH), SEQ ID NO: 463 (VL)
[0622]Note here that (1) and (2) correspond to 064-003 antibody. Therefore, the antibody of the present invention is expected to have high specificity to CD44.
[0623]The tenth embodiment of this aspect provides an isolated antibody having a specific binding property to CD73. The antibody of this form includes the heavy chain variable regions CDR1 to CDR3 and the light chain variable regions CDR1 to CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2 and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) shown in the following (1). Preferably, it includes the heavy chain variable region and the light chain variable region specified by the combination of SEQ ID NOs (SEQ ID NO showing the heavy chain variable region and SEQ ID NO showing the light chain variable region) shown in the following (2).
(Combination of CDR1 to CDR3)
(1) SEQ ID NO: 468 (VH CDR1), SEQ ID NO: 469 (VH CDR2), SEQ ID NO: 470 (VH CDR3), SEQ ID NO: 472 (VL CDR1), SEQ ID NO: 473 (VL CDR2), SEQ ID NO: 474 (VL CDR3)
(Combination of Heavy Chain Variable Region and Light Chain Variable Region)
(2) SEQ ID NO: 467 (VH), SEQ ID NO: 471 (VL)
[0624]Note here that (1) and (2) correspond to 067-213 antibody. Therefore, the antibody of the present invention is expected to have high specificity to CD73.
[0625]The eleventh embodiment of this aspect provides an isolated antibody having a specific binding property to EpCAM. The antibody of this form includes the heavy chain variable regions CDR1 to CDR3 and the light chain variable regions CDR1 to CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2 and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) shown in the following (1). Preferably, it includes the heavy chain variable region and the light chain variable region specified by the combination of SEQ ID NOs (SEQ ID NO showing the heavy chain variable region and SEQ ID NO showing the light chain variable region) shown in the following (2).
(Combination of CDR1 to CDR3)
(1) SEQ ID NO: 476 (VH CDR1), SEQ ID NO: 477 (VH CDR2), SEQ ID NO: 478 (VH CDR3), SEQ ID NO: 480 (VL CDR1), SEQ ID NO: 481 (VL CDR2), SEQ ID NO: 482 (VL CDR3)
(Combination of Heavy Chain Variable Region and Light Chain Variable Region)
(2) SEQ ID NO: 475 (VH), SEQ ID NO: 479 (VL)
[0626]Note here that (1) and (2) correspond to 067-153 antibody. Therefore, the antibody of the present invention is expected to have high specificity to EpCAM.
[0627]The twelfth embodiment of this aspect provides an isolated antibody having a specific binding property to HGFR. The antibody of this form includes the heavy chain variable regions CDR1 to CDR3 and the light chain variable regions CDR1 to CDR3 specified by the combination of SEQ ID NOs (SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR2, SEQ ID NO showing the amino acid sequence of the heavy chain variable region CDR3, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR1, SEQ ID NO showing the amino acid sequence of the light chain variable region CDR2 and SEQ ID NO showing the amino acid sequence of the light chain variable region CDR3) selected from the group consisting of the following (1) to (3). Preferably, it includes the heavy chain variable region and the light chain variable region specified by the combination of SEQ ID NOs (SEQ ID NO showing the heavy chain variable region and SEQ ID NO showing the light chain variable region) selected from the group consisting of the following (4) to (6).
(Combination of CDR1 to CDR3)
(1) SEQ ID NO: 652 (VH CDR1), SEQ ID NO: 653 (VH CDR2), SEQ ID NO: 654 (VH CDR3), SEQ ID NO: 656 (VL CDR1), SEQ ID NO: 657 (VL CDR2), SEQ ID NO: 658 (VL CDR3)
(2) SEQ ID NO: 660 (VH CDR1), SEQ ID NO: 661 (VH CDR2), SEQ ID NO: 662 (VH CDR3), SEQ ID NO: 664 (VL CDR1), SEQ ID NO: 665 (VL CDR2), SEQ ID NO: 666 (VL CDR3)
(3) SEQ ID NO: 668 (VH CDR1), SEQ ID NO: 669 (VH CDR2), SEQ ID NO: 670 (VH CDR3), SEQ ID NO: 672 (VL CDR1), SEQ ID NO: 673 (VL CDR2), SEQ ID NO: 674 (VL CDR3)
[0628](Combination of heavy chain variable region and light chain variable region)
(4) SEQ ID NO: 651 (VH), SEQ ID NO: 655 (VL)
(5) SEQ ID NO: 659 (VH), SEQ ID NO: 663 (VL)
(6) SEQ ID NO: 667 (VH), SEQ ID NO: 671 (VL)
[0629]Note here that (1) and (4) correspond to 067-126 antibody; (2) and (5) correspond to 067-133 antibody; and (3) and (6) correspond to 067-287 antibody. Therefore, the antibody of the present invention is expected to have high specificity to HGFR.
[0630]The 13rd embodiment of this aspect provides an isolated antibody having a specific binding property to LAR. The antibody of this form includes the heavy chain variable region and the light chain variable region specified by the combination of SEQ ID NOs (SEQ ID NO showing the heavy chain variable region and SEQ ID NO showing the light chain variable region) selected from the group consisting of the following (1) to (5).
(Combination of heavy chain variable region and light chain variable region)
(1) SEQ ID NO: 944 (VH), SEQ ID NO: 945 (VL)
(2) SEQ ID NO: 946 (VH), SEQ ID NO: 947 (VL)
(3) SEQ ID NO: 948 (VH), SEQ ID NO: 949 (VL)
(4) SEQ ID NO: 950 (VH), SEQ ID NO: 951 (VL)
(5) SEQ ID NO: 952 (VH), SEQ ID NO: 953 (VL)
[0631]Note here that (1) corresponds to 064-044 antibody; (2) corresponds to 065-030 antibody; (3) corresponds to 065-358 antibody; (4) corresponds to 066-019 antibody; and (5) corresponds to 079-085 antibody. Therefore, the antibody of the present invention is expected to have high specificity to LAR.
[0632]The 14th embodiment of this aspect provides an isolated antibody having a specific binding property to BCAM. The antibody of this form includes the heavy chain variable region and the light chain variable region specified by the combination of SEQ ID NOs (SEQ ID NO showing the heavy chain variable region and SEQ ID NO showing the light chain variable region) shown in the following (1).
(Combination of heavy chain variable region and light chain variable region)
(1) SEQ ID NO: 954 (VH), SEQ ID NO: 955 (VL)
[0633]Note here that (1) corresponds to 067-024 antibody. Therefore, the antibody of the present invention is expected to have high specificity to BCAM.
[0634]In the variable region of the antibody of the present invention, the sequence of the framework region (FR region) is not particularly limited as long as it does not substantially affect the specific binding property with respect to corresponding antigen. For example, when the antibody of the present invention is constructed as a humanized antibody, the FR region of a known human antibody can be used. Furthermore, when the antibody of the present invention is constructed as an antibody used as a reagent for detection or used for application to non-human animal species, in some cases, an effect can be expected even if the human antibody FR region is not used, or the use of the human antibody FR region may not appropriate. In such cases, the FR region from non-human animal species (for example, mouse or rat) can be used.
[0635]In one embodiment of the antibody of the present invention, a constant region (for example, in the case of an IgG type antibody) is included in addition to the variable region. The sequence of the constant region in this embodiment is not particularly limited. For example, as mentioned below, when the antibody of the present invention is constructed as a humanized antibody, the constant region of a known human antibody can be used. Furthermore, similar to the above-mentioned FR region, a constant region from non-human animal species (for example, mouse or rat) can be used.
[0636]One embodiment of the antibody of the present invention relates to a humanized antibody. The "humanized antibody" herein denotes an antibody that is allowed to resemble the structure of the human antibody. It includes a humanized chimeric antibody in which only a constant region is replaced by that of human antibody, and a humanized CDR-grafted antibody in which a part other than the CDR (complementarity determining region) existing in the constant region and the variable region is replaced by that of human antibody (P. T. Johons et al., Nature 321, 522 (1986)). In order to improve the antigen binding activity of the humanized CDR-grafted antibody, improved techniques of a method of selecting a human antibody FR that is highly homologous to a mouse antibody, a method of producing a humanized antibody having high homology, and a method of transplanting a human antibody to a mouse CDR and then replacing amino acid in the FR region have been already developed (see, for example, U.S. Pat. Nos. 5,585,089, 5,693,761, 5,693,762, and 6180370, European Patent Nos. 451216 and 682040, and U.S. Pat. No. 2,828,340) and such techniques can be used for producing the humanized antibody of the present invention.
[0637]The humanized chimeric antibody can be produced by, for example, replacing the constant region of an antibody having the above-mentioned structure of H chain variable region and/or structure of L chain variable region by the constant region of a human antibody. As the constant region of the human antibody, known region can be employed. Hereinafter, one example of the method of producing the humanized chimeric antibody is described.
[0638]Firstly, mRNA is extracted from the hybridoma producing a mouse antibody to certain antigens (for example, antigens expressing certain cancers, which have been determined this time, HER1, HER2, CD46, ITGA3, ICAM1, ALCAM, CD147, or the like), and cDNA is synthesized according to the usual procedure. The synthesized cDNA is inserted into a vector so as to construct a cDNA library. From this cDNA library, an H chain gene fragment and an L chain gene fragment are used as a probe, a vector containing an H chain gene and an L chain gene is selected. By sequencing the insertion sequence of the selected vector, the sequences of the gene in the H chain variable region and the L chain variable region can be determined. Based on the thus obtained sequence data, DNA encoding H chain variable region is produced by a chemical synthesis, biochemical cleavage/recombination and the like. DNA encoding the obtained H chain variable region is ligated with DNA encoding a human H chain constant region so as to incorporate it into an expression vector. Thereby, H chain expression vector is produced. As the expression vector, for example, an SV40 virus based vector, an EB virus based vector, and a BPV (papilloma virus) based vector can be used but not limited to these vectors alone. On the other hand, by the similar method, an L chain expression vector is produced. With such H chain expression vector and L chain expression vector, host cells are co-transformed. As the host cell, CHO cell (Chinese hamster ovary cell) (A. Wright & S. L. Morrison, J. Immunol. 160, 3393-3402 (1998)), SP2/0 cell (myeloma) (K. Motmans et al., Eur. J. Cancer Prev. 5, 512-519 (1996), R. P. Junghans et al., Cancer Res. 50, 1495-1502 (1990)), and the like can be suitably used. Furthermore, for transformation, a Lipofectin method (R. W. Malone et al., Proc. Natl. Acad. Sci. USA 86, 6077 (1989), P. L. Felgner et al., Proc. Natl. Acad. Sci. USA 84, 7413 (1987), an electroporation method, a calcium phosphate method (F. L. Graham & A. J. van der Eb, Virology 52, 456-467 (1973)), a DEAE-Dextran method, and the like, are suitably used.
[0639]After the transformant is cultured, a humanized chimeric antibody is separated from the cells of transformant or the culture solution. For separation and purification, methods such as centrifugation, ammonium sulfate fractionation, salting out, ultrafiltration, affinity chromatography, ion-exchange chromatography, and gel filtration chromatography can be appropriately combined and used.
[0640]On the other hand, the humanized CDR-grafted antibody can be produced by, for example, the following method. Firstly, by the method described in the production method of chimeric antibody, the amino acid sequences of the H chain variable region and L chain variable region of the antibody to the certain antigen and the base sequences encoding the amino acid sequences are determined. In addition, the amino acid sequence and the base sequence of each CDR region are determined.
[0641]As the base sequence of the specific CDRs, any of the following combinations are used. Note here that they are shown by SEQ ID NO showing the base sequence of the heavy chain variable region CDR1, SEQ ID NO showing the base sequence of the heavy chain variable region CDR2, SEQ ID NO showing the base sequence of the heavy chain variable region CDR3, SEQ ID NO showing the base sequence of the light chain variable region CDR1, SEQ ID NO showing the base sequence of the light chain variable region CDR2, and SEQ ID NO showing the base sequence of the light chain variable region CDR3, in this order.
(1) SEQ ID NO 186, SEQ ID NO 187, SEQ ID NO 188, SEQ ID NO 190, SEQ ID NO 191, SEQ ID NO 192
(2) SEQ ID NO 194, SEQ ID NO 195, SEQ ID NO 196, SEQ ID NO 198, SEQ ID NO 199, SEQ ID NO 200
(3) SEQ ID NO: 202, SEQ ID NO: 203, SEQ ID NO: 204, SEQ ID NO: 206, SEQ ID NO: 207, SEQ ID NO: 208
(4) SEQ ID NO: 210, SEQ ID NO: 211, SEQ ID NO: 212, SEQ ID NO: 214, SEQ ID NO: 215, SEQ ID NO: 126
(5) SEQ ID NO: 218, SEQ ID NO: 219, SEQ ID NO: 220, SEQ ID NO: 222, SEQ ID NO: 223, SEQ ID NO: 224
(6) SEQ ID NO: 226, SEQ ID NO: 227, SEQ ID NO: 228, SEQ ID NO: 230, SEQ ID NO: 231, SEQ ID NO: 232
(7) SEQ ID NO: 234, SEQ ID NO: 235, SEQ ID NO: 236, SEQ ID NO: 238, SEQ ID NO: 239, SEQ ID NO: 240
(8) SEQ ID NO: 242, SEQ ID NO: 243, SEQ ID NO: 244, SEQ ID NO: 246, SEQ ID NO: 247, SEQ ID NO: 248
(9) SEQ ID NO: 250, SEQ ID NO: 251, SEQ ID NO: 252, SEQ ID NO: 254, SEQ ID NO: 255, SEQ ID NO: 256
(10) SEQ ID NO: 258, SEQ ID NO: 259, SEQ ID NO: 260, SEQ ID NO: 262, SEQ ID NO: 263, SEQ ID NO: 264
(11) SEQ ID NO: 266, SEQ ID NO: 267, SEQ ID NO: 268, SEQ ID NO: 270, SEQ ID NO: 271, SEQ ID NO: 272
(12) SEQ ID NO: 274, SEQ ID NO: 275, SEQ ID NO: 276, SEQ ID NO: 278, SEQ ID NO: 279, SEQ ID NO: 280
(13) SEQ ID NO: 282, SEQ ID NO: 283, SEQ ID NO: 284, SEQ ID NO: 286, SEQ ID NO: 287, SEQ ID NO: 288
(14) SEQ ID NO: 290, SEQ ID NO: 291, SEQ ID NO: 292, SEQ ID NO: 294, SEQ ID NO: 295, SEQ ID NO: 296
(15) SEQ ID NO: 298, SEQ ID NO: 299, SEQ ID NO: 300, SEQ ID NO: 302, SEQ ID NO: 303, SEQ ID NO: 304
(16) SEQ ID NO: 306, SEQ ID NO: 307, SEQ ID NO: 308, SEQ ID NO: 310, SEQ ID NO: 311, SEQ ID NO: 312
(17) SEQ ID NO: 314, SEQ ID NO: 315, SEQ ID NO: 316, SEQ ID NO: 318, SEQ ID NO: 319, SEQ ID NO: 320
(18) SEQ ID NO: 322, SEQ ID NO: 323, SEQ ID NO: 324, SEQ ID NO: 326, SEQ ID NO: 327, SEQ ID NO: 328
(19) SEQ ID NO: 330, SEQ ID NO: 331, SEQ ID NO: 332, SEQ ID NO: 334, SEQ ID NO: 335, SEQ ID NO: 336
(20) SEQ ID NO: 338, SEQ ID NO: 339, SEQ ID NO: 340, SEQ ID NO: 342, SEQ ID NO: 343, SEQ ID NO: 344
(21) SEQ ID NO: 346, SEQ ID NO: 347, SEQ ID NO: 348, SEQ ID NO: 350, SEQ ID NO: 351, SEQ ID NO: 352
(22) SEQ ID NO: 354, SEQ ID NO: 355, SEQ ID NO: 356, SEQ ID NO: 358, SEQ ID NO: 359, SEQ ID NO: 360
(23) SEQ ID NO: 362, SEQ ID NO: 363, SEQ ID NO: 364, SEQ ID NO: 366, SEQ ID NO: 367, SEQ ID NO: 368
[0642]Note here that these combinations correspond to the combination in CDR1 to CDR3 in 048-006 antibody, 057-091 antibody, and 059-152 antibody (which are antibodies to HER1), 015-126 antibody (which is antibody to HER2), 035-224 antibody, 045-011 antibody, 051-144 antibody, 052-053 antibody, 052-073 antibody, 053-049 antibody, and 3172-120 antibody (which are antibodies to CD46), 015-003 antibody (which is antibody to ITGA3), 052-033 antibody, 053-042 antibody, 053-051 antibody, 053-059 antibody, and 053-085 antibody (which are antibodies to ICAM1), 035-234 antibody, 040-107 antibody, 041-118 antibody, 066-174 antibody, and 083-040 antibody (which are antibodies to ALCAM), 059-053 antibody (which is antibody to CD147).
[0643]Next, FRs (framework regions) sandwiching the CDR region are selected. For selecting the FR, approximately three methods can be employed. The first method is a method using a human antibody frame whose three dimensional structure has been already identified, for example, NEWM, REI, and the like (Riechmann L. et al., Nature 332, 323-3Z7 (1988); Tempst, P R. et al., Protein Engineering 7, 1501-1507 (1994); Ellis J H. et al., J. Immunol. 155, 925-937 (1995)). The second method includes selecting a variable region of a human antibody having the highest homology to a variable region of the intended mouse antibody from database, and using the FR thereof (Queen C. et al., Proc Natl Acad Sci USA 86, 10029-10033 (1989); Rozak M J. et al., J Biol Chem 271, 22611-22618 (1996); Shearman C W. et al., J. Immunol. 147, 4366-4373 (1991)). The third method is a method of selecting amino acid most commonly used in the FR of the human antibody (Sato K. et al., Mol Immunol 31, 371-381 (1994); Kobinger F. et al., Protein Engineering 6, 971-980 (1993); Kettleborough C A. et al., Protein Engineering 4, 773-783 (1991)). The present invention can use any of these methods.
[0644]Even if the amino acid sequence is an amino acid sequence obtained by modifying the amino acid sequence of the selected human FR, it can be used as an amino acid sequence of the FR as long as a finally obtained humanized CDR-grafted antibody has a specific binding property to the corresponding antigens (HER1, HER2, CD46, ITGA3, ICAM1, ALCAM, CD147, and the like). In particular, a part of the amino acid of the selected human FR is changed to the amino acid of the FR of the antibody of the origin of CDR, the property of the antibody may be maintained. The number of the amino acid to be modified is preferably 30% or less relative to the entire FR, further preferably 20% or less relative to the entire FR, and yet further preferably 10% or less relative to the entire FR.
[0645]Next, by combining the FR selected by any of these methods and the above-mentioned CDR, DAN encoding the H chain variable region and L chain variable region is designed. Based on this design, DNA encoding H chain variable region and DNA encoding L chain variable region are produced by the chemical synthesis, biochemical cleavage/recombination, and the like, respectively. Then, DAN encoding the H chain variable region together with the DNA encoding H chain constant region of a human immunoglobulin is incorporated into an expression vector so as to construct an H chain expression vector. Similarly, DAN encoding the L chain variable region together with the DNA encoding L chain constant region of a human immunoglobulin is incorporated into an expression vector so as to construct an L chain expression vector. As the expression vector, for example, an SV40 virus based vector, an EB virus based vector, a BPV (papilloma virus) based vector, and the like can be used but not necessarily limited to these vectors.
[0646]With the H chain expression vector and L chain expression vector that are produced by the above-mentioned method, host cells are co-transformed. As the host cell, CHO cell (Chinese hamster ovary cell) (A. Wright & S. L. Morrison, J. Immunol. 160, 3393-3402 (1998)), SP2/0 cell (myeloma) (K. Motmans et al., Eur. J. Cancer Prev. 5, 512-519 (1996), R. P. Junghans et al., Cancer Res. 50, 1495-1502 (1990)), and the like can be suitably used. Furthermore, for transformation, a Lipofectin method (R. W. Malone et al., Proc. Natl. Acad. Sci. USA 86, 6077 (1989), P. L. Feigner et al., Proc. Natl. Acad. Sci. USA 84, 7413 (1987), an electroporation method, a calcium phosphate method (F. L. Graham & A. J. van der Eb, Virology 52, 456-467 (1973)), a DEAE-Dextran method, and the like, are suitably used.
[0647]After the transformant is cultured, a humanized CDR-grafted antibody is separated from the cells of transformant or the culture solution. For separation and purification, methods such as centrifugation, ammonium sulfate fractionation, salting out, ultrafiltration, affinity chromatography, ion-exchange chromatography, and gel filtration chromatography can be appropriately combined and used.
[0648]Based on the antibody of the present invention or based on the sequence information on the genes encoding the antibody of the present invention, an antibody fragment can be produced. The antibody fragment can include Fab, Fab', F(ab')2, scFv, and dsFv antibodies.
[0649]Fab is a fragment that is obtained by digesting IgG with papain in the presence of cysteine; includes L chain and H chain variable regions as well as an H chain fragment consisting of a CH1 domain and a part of hinge portion; and has a molecular weight of about 50000. In the present invention, it can be obtained by digesting the antibody with papain. Furthermore, DNA encoding a part of the H chain of the above-mentioned antibody and L chain is incorporated into an appropriate vector, and the vector is used for transforming so as to obtain a transformant. From this transformant, Fab can be prepared.
[0650]Fab' is a fragment having a molecular weight of about 50000, which can be obtained by cleaving the disulfide bond between H chains of F(ab')2 mentioned below. In the present invention, it can be obtained by digesting the above-mentioned antibody with pepsin and cleaving the disulfide bond by the use of a reducing agent. Furthermore, similar to Fab, it can also be prepared by gene engineering with the use of DNA encoding Fab'.
[0651]F(ab')2 is a fragment that is obtained by digesting IgG with pepsin; a fragment (Fab') is linked by disulfide bond including L chain and H chain variable regions as well as an H chain fragment consisting of a CH1 domain and a part of hinge portion; and has a molecular weight of about 100000. In the present invention, it can be obtained by digesting the antibody with pepsin. Furthermore, similar to Fab, it can also be prepared by gene engineering with the use of DNA encoding F(ab')2.
[0652]scFv is an antibody fragment obtained by linking Fv including an H chain variable region and an L chain variable region to C terminal of one of the chains and N terminal of the other of the chains by using an appropriate peptide linker so as to produce a single chain antibody fragment. As the peptide linker, for example, highly flexible (GGGGS)3 can be used. For example, DNA encoding an scFv antibody is constructed by using DNA encoding H chain variable region and L chain variable region of the above-mentioned antibody and DNA encoding the peptide linker is constructed. This is incorporated into an appropriate vector and this vector is used to obtain a transformant. From this transformant, scFv can be prepared.
[0653]dsFv is an Fv fragment obtained by introducing a Cys residue into an appropriate positions of the H chain variable region and L chain variable region and stabilizing the H chain variable region and chain variable region by disulfide bond. The position in which the Cys residue is introduced in each chain can be determined based on the three dimensional structure anticipated by molecule modeling. In the present invention, for example, the three dimensional structure is anticipated from the amino acid sequence of the H chain variable region and the L chain variable region of the above-mentioned antibody. DNA encoding the H chain variable region and L chain variable region into which difference based on such anticipation is constructed and the constructed DNA is incorporated into the appropriate vector. The vector is used to obtain a transformant. From this transformant, dsFv can be prepared.
[0654]Note here that an antibody fragment can be multimerized by linking an scFv antibody and a dcFv antibody and the like with the use of an appropriate linker, or by allowing streptavidin to be fused.
[0655]By fusing or linking a low molecule compound, protein, a label material, and the like to the antibody of the present invention (including an antibody fragment), a fused antibody or labeled antibody can be formed. An example of the label material may include radioactive material such as 125I, peroxidase, β-D-galactosidase, micro peroxidase, horseradish peroxidase (HRP), fluorescein isothiocyanate (FITC), rhodamine isothiocyanate (RITC), alkaline phosphatase, biotin, and the like.
[0656]The antibody of the present invention (including an antibody fragment) specifically binds to a cancer cell that specifically expresses the antigen by the specific binding property to the corresponding antigen. The use of this property makes it possible to label and detect a cancer cell (or cancer tissue). By gene recombination technology, VH and VL having such a specific binding capacity can be fused to a constant region (Fc region) of IgG so as to transform into an IgG type antibody. The thus obtained IgG type antibody is expected to exhibit a cytotoxic effect via Fc receptor on NK cells. The IgG constant region has subclass. As to the binding of Fc receptor of each IgG subclass of human, IgG1 and IgG3 have the strongest binding, IgG4 has moderate binding and IgG2 has weak binding. In transforming into IgG type antibodies, it is preferable to select a constant region in consideration of this point. Note here that the present inventors have proposed an assay of cytotoxic effect via the secondary antibody instead of IgG type antibody in the previous applications (Japanese Patent Unexamined Publication No. 2005-185281 and PCT/JP2006/303195).
[0657]Actually, as shown in the below-mentioned Examples, since 015-003 antibody as anti-ITGA3 antibody, 048-006 antibody as anti-HER1 antibody, and 015-126 antibody as anti-HER2 antibody are recognized to have an ADCC activity, they themselves can be used for damaging (killing) cancer cells. Herein, when the antibody of the present invention that has transformed into human or human IgG antibody is used, it is less attacked and excluded by the immune system, thus enabling the expected effect to be well exhibited and serious side effects to be avoided.
[0658]Furthermore, the antibody of the present invention can be used as a medium (carrier) for delivering a drug, and the like, to a specific cancer. That is to say, an anticipated application of use of the antibody of the present invention includes DDS (Drug delivery system) targeting a specific cancer cell.
[0659]Note here that each application of the antibody of the present invention is described in detail below.
(Diagnosis Application)
[0660]Another aspect of the present invention relates to a use as a diagnosis marker of based on the findings of the expression (distribution) of CD46 antigen, ITGA3, ALCAM and CD147 antigen. Specifically, one embodiment of this aspect provides a testing method of gallbladder and liver cancer or pancreas cancer based on the findings that a CD46 antigen is expressed in the gallbladder and liver cancer and the pancreas cancer. The method includes the following steps.
[0661]Step (1): preparing subject cells or tissues separated from a living body.
[0662]Step (2): detecting a CD46 antigen in the subject cells or tissues.
[0663]Information obtained by the testing method of the present invention is useful for diagnosis of gallbladder and liver cancer or pancreas cancer. For example, information obtained by subjecting the above-mentioned method to patients with gallbladder and liver cancer can be used for evaluating or grasping the pathologic condition of patients and for evaluating the therapeutic effect. For example, when the present invention is carried out concurrently with the treatment of gallbladder and liver cancer, based on the resultant information, the therapeutic effect can be evaluated. Specifically, when the method of the present invention is carried out after administering drugs, the change in the expression amount of CD46 antigen in the liver cells is examined and the therapeutic effect can be determined from the increase and decrease of the expression amount. Thus, the method of the present invention may be used for monitoring the therapeutic effect.
[0664]On the other hand, information obtained when the subjects are persons other than the patient, that is, persons that have not recognized to have gallbladder and liver cancer can be used for determination of the presence or absence of contraction of gallbladder and liver cancer, evaluation of contraction risk, and the like. Since the method of the present invention permits diagnosis of liver cancer based on the amount of expression amount of genes, i.e., an objective indicator, its value is extremely high.
[0665]Hereinafter, the steps constituting the present invention are respectively described in detail.
1. Step (1)
[0666]In the step (1), cells or tissues separated from a subject (a subject person, a living body) are prepared. The subjects herein may include not only patients (gallbladder and liver cancer patients or pancreas cancer patients) but also healthy persons (including persons having a risk of contracting gallbladder and liver cancer or pancreas cancer). For example, a part of tissues collected from a subject by biopsy can be used as subject cells or tissues in the method of the present invention.
[0667]The "subject cells or tissues" in the present invention are cells or tissues that are samples (subjects) in the detection in the method of the present invention. The subject cells or tissues are separated from a living body. That is to say, the present invention is applied to the subject cells or tissues in the state in which it is separated from the living body. The term "separated from a living body" means a state in which a part of the living tissue in which subject cells or tissues exist is extracted, thereby the subject cells or tissues are completed separated from the origin living body. In the step (2), when an immunological detection method is employed, the subject cells are generally prepared in a state in which they are present in a living body, that is, in a state in which they are linked to the surrounding cells (as tissue), and used for the method of the present invention. Note here that the subject cells may be used for the method of the present invention after they are separated (isolated) from the surrounding cells.
2. Step (2)
[0668]In the step (2), a CD46 antigen is detected in the prepared subject cells or tissues as subjects. The term "CD46 antigen is detected" means examining whether or not the CD46 antigen is expressed (presence or absence of expression), or figuring out the expression amount of the CD46 antigen as an absolute value or a relative value. The reference of the relative amount herein can be, for example, an amount of CD46 antigen of the reference samples prepared according to the grade of malignancy. In general, the presence of expression of CD46 antigen and the amount if expressed are examined. In detecting the CD46 antigen, it is not essential to determine the amount of CD46 antigens strictly.
[0669]In one embodiment of the present invention, a detection method targeting mRNA that is a transcriptional product of the CD46 antigen is carried out. For the detection (measurement) of mRNA, routine procedures such as an RT-PCR method and various hybridization methods using specific probes (for example, northern hybridization, in situ hybridization) can be employed. In another embodiment of the present invention, a detection method targeting the expression product of the CD46 antigen (protein) is carried out.
[0670]It is preferable that CD46 antigen is detected by immunologic procedures (for example, immunohistochemical staining technique). In the immunologic procedure, anti-CD46 antigen antibody is used, CD46 antigen protein is detected by using the bonding property (binding amount) of the antibodies as an indicator. The immunological detection method permits rapid and sensitive detection. Also, the operation is simple. An example of the detection methods may include ELISA method, radioimmunoassay, FCM, an immunoprecipitation method, immunoblotting, and the like.
[0671]The immunohistochemical staining technique permits rapid and sensitive detection of CD46 antigens. Also, the operation is simple. Therefore, burdens to a subject person (patient) accompanying the detection of CD46 antigen is reduced. In the immunohistochemical staining technique, in general, firstly, a step of bringing the subject cells into contact with the anti-CD46 antibody is carried out. Then, the binding amount of the anti-CD46 antibody is examined. Specifically, according to the above-mentioned immunohistochemical staining technique, the method of the present invention can be carried out.
[0672]The kind or origin of the anti-CD46 antibody to be used in immunostaining procedure is not particularly limited as long as it has a specific binding property to the CD46 antigen. The anti-CD46 antibody may be any of a polyclonal antibody, an oligoclonal antibody (a mixture of several kinds to several tens of antibodies) and a monoclonal antibody. As the polyclonal antibody or the oligoclonal antibody, affinity purification antibody by antigen can be used besides an IgG fraction derived from anti-serum obtained by immunizing an animal so as to obtain. The anti-CD46 antibody may be antibody fragments such as Fab, Fab', F(ab')2, scFv, and dsFv antibodies.
[0673]The anti-CD46 antibody can be prepared by using an immunologic procedure, phage display technique, ribosome display method, and the like.
[0674]The preparation of a polyclonal antibody by the immunologic procedure can be prepared by the following procedures. An antigen (CD46 or a part thereof) is prepared. An animal such as a rabbit is immunized with this antigen. As this antigen, not only human CD46 but also non-human CD46 such as mouse CD46 can be used. Such CD46 can be obtained by purifying a living body sample. Furthermore, recombinant CD46 may be used. The recombinant human CD46 can be prepared by, for example, introducing a gene encoding CD46 (which may include a part of gene) in an appropriate host by using a vector and expressing the gene within the obtained recombinant cells.
[0675]In order to strengthen the immunity inducing effect, an antigen to which a carrier protein is attached may be used. As the carrier protein, KLH (Keyhole Limpet Hemocyanin), BSA (Bovine Serum Albumin), OVA (Ovalbumin), and the like are used. For binding of the carrier protein, a carbodiimide method, a glutaraldehyde method, a diazo condensation method, an MBS (maleimidobenzoyl oxy succinimide) method, and the like, can be used. On the other hand, an antigen expressing CD46 (or a part thereof) as fusion protein with GST, β galactosidase, maltose bonded protein, or histidine (His) tag, and the like, can be used. Such a fusion protein can be purified by a general method in a simple manner.
[0676]If necessary, immunization is repeated. When the antibody titer is sufficiently increased, blood is collected and subjected to centrifugation so as to obtain serum. The obtained anti-serum is subjected to affinity purification. Thus, a polyclonal antibody is obtained.
[0677]On the other hand, a monoclonal antibody can be prepared by the following procedures. Firstly, an immunization operation is carried out by the similar method to the above-mentioned procedures. If necessary, immunization is repeated. When the antibody titer is sufficiently increased, antibody-producing cells are extracted from an immunized animal. Next, the obtained antibody-producing cells and myeloma cells are fused to each other so as to obtain a hybridoma. Subsequently, this hybridoma is made to be monoclonal. Then, a clone producing antibody showing high specificity to the target protein is selected. A culture solution of the selected clone is purified, thereby the target antibody can be obtained. On the other hand, hybridoma is proliferated into a predetermined number of more, then, transplanted in the abdominal cavity of an animal (for example, mouse), proliferated in the abdominal dropsy. By purifying the abdominal dropsy, the target antibody can be obtained. For purification of the above-mentioned culture solution or purification of the abdominal dropsy, affinity chromatography using protein G, protein A, and the like, is preferably used. Furthermore, affinity chromatography in which an antigen is made into a solid phase can be used. Furthermore, methods such as ion-exchange chromatography, gel filtration chromatography, ammonium sulfate fractionation, and centrifugation can be used. These methods are used singly or in arbitrary combination thereof.
[0678]On the conditions that the specific binding property to CD46 antigen is maintained, the obtained antibody may be subjected to various modifications. In the present invention, such a modified antibody may be used.
[0679]When a labeled antibody is used as an anti-CD46 antibody, the amount of bound antibody can be directly detected by using the labeled amount as an indicator. Therefore, the method is more simplified. On the contrary, it is necessary to prepare an anti-CD46 antibody to which a label material is bound and furthermore, and furthermore, the detection sensitivity is generally reduced. Therefore, it is preferable that indirect methods such as a method using a secondary antibody to which a label material is linked, a method using a polymer to which a secondary antibody and a label material are linked are used. The secondary antibody herein is an antibody having a specific binding property to the anti-CD46 antibody. For example, when an anti-CD46 antibody is prepared as a rabbit antibody, an anti-rabbit IgG antibody can be used. Label secondary antibodies that can be used for various species such as rabbit, goat, and mouse are commercially available (for example, Funakoshi Corporation, COSMO BIO Co., Ltd., etc.). Proper antibodies can be appropriately selected depending upon the anti-CD46 antibody used in the present invention.
[0680]For the label material, the label material arbitrarily selected from the group consisting of peroxidase, β-D-galactosidase, micro peroxidase, horseradish peroxidase (HRP), fluorescein isothiocyanate (FITC), rhodamine isothiocyanate (RITC), alkaline phosphatase, biotin, and radioactive material is preferably used. In particular, a method of using biotin as the label material and reacting avidin peroxidase permits highly sensitive detection.
[0681]The above-mentioned antibody of the present invention may be used as the anti-CD46 antibody. Specifically, for example, antibodies (035-224 antibody, 045-011 antibody, 051-144 antibody, 052-053 antibody, 052-073 antibody, 053-049 antibody, or 3172-120 antibody), which the present inventors have succeeded in obtaining, can be used.
[0682]Another embodiment of this aspect provides a testing method of gallbladder and liver cancer or pancreas cancer based on the findings that ITGA3 is expressed in gallbladder and liver cancer and pancreas cancer. The method includes the following steps.
[0683]Step (1): preparing subject cells or tissues separated from a living body
[0684]Step (2): detecting ITGA3 in the subject cells or tissues
[0685]Information obtained by the testing method of the present invention is useful for diagnosis of gallbladder and liver cancer or diagnosis of pancreas cancer. Since the using method and details of each step are the same as in the case of the CD46 antigen, the description thereof is not mentioned here.
[0686]A further embodiment of this aspect provides an obtaining method of information for diagnosis of kidney cancer, hepatic cell carcinoma or gallbladder and liver cancer based on the findings that ALCAM is expressed in kidney cancer, hepatic cell carcinoma and gallbladder and liver cancer. The method includes the following steps.
[0687]Step (1): preparing subject cells or tissues separated from a living body
[0688]Step (2): detecting ALCAM in the subject cells or tissues
[0689]Information obtained by the testing method of the present invention is useful for diagnosis of kidney cancer, diagnosis of hepatic cell carcinoma, or diagnosis of gallbladder and liver cancer. Since the using method and details of each step are the same as in the case of the CD46 antigen, the description thereof is not mentioned here.
[0690]A yet further embodiment of this aspect provides a testing method of kidney cancer based on the findings that CD147 antigen is expressed in kidney cancer. The method includes the following steps.
[0691]Step (1): preparing subject cells or tissues separated from a living body
[0692]Step (2): detecting a CD147 antigen in the subject cells or tissues
[0693]Information obtained by the testing method of the present invention is useful for diagnosis of kidney cancer. Since the using method and details of each step are the same as in the case of the CD46 antigen, the description thereof is not mentioned here.
(Treatment Application)
[0694]As mentioned in the below-mentioned Examples, the present inventor have succeeded in obtaining antibodies exhibiting Antibody-Dependent Cell-mediated Cytotoxicity (hereinafter, abbreviated as "ADCC") activity to certain antibodies. Furthermore, the present inventors have transformed these antibodies into IgG type and investigated the probability of application to an antibody therapeutic agent. Any antibodies show excellent anti-tumor effect. Based on these findings, the further aspect of the present invention relates to an application of the antibodies successfully obtained by the present inventors in treatment of cancer.
[0695]This aspect firstly provides a drug (cancer therapeutic agent) capable of affecting and damaging in a cancer cell-specific manner using by using ITGA3, HER1, HER2, ALCAM, EpCAM or HGFR as a target, and the treatment method using the same. One embodiment of the drug of the present invention contains anti-ITGA3 antibody as an active ingredient. One preferable embodiment of the drug of the present invention contains an anti-ITGA3 antibody having an ADCC activity as an active ingredient. The drugs of this embodiment can obtain the therapeutic effect by the cytotoxicity using the ADCC activity. As anti-ITGA3 antibody having the ADCC activity, 015-003 antibody (the specific binding property to ITGA3 and it may be partially modified as long as the ADCC activity is maintained) shown in the below-mentioned Examples or different types of antibodies constructed based on the 015-003 antibody (for example, IgG type antibody) can be used. This antibody has both the specific binding property to ITGA3 and the ADCC activity. Therefore, it specifically binds to the cancer cells expressing ITGA3 and then expresses the ADCC activity. Thus, it can damage a cancer cell. The target cancer cell of the drug of this embodiment is not particularly limited, but can target, for example, gallbladder and liver cancer cells and pancreas cancer cells.
[0696]In another embodiment of the present invention, an anti-HER1 antibody is contained as an active ingredient. In the drug of one preferable embodiment of the present invention, anti-HER1 antibody having an ADCC activity is contained as an active ingredient. In the drug of this embodiment, the therapeutic effect can be obtained by the cytotoxicity using the ADCC activity. In the drug of the further preferable embodiment, in addition to the cytotoxicity using the ADCC activity, since inhibition of binding of EGF as a ligand to HER1 and/or inhibition of phosphorylation signal by HER1 are provided, higher therapeutic effect can be obtained. As anti-HER1 antibody having such an ADCC activity, 048-006 antibody, 059-152 antibody, 055-147 antibody or 059-173 antibody shown in the below-mentioned Example (which may be partially modified as long as the specific binding property to HER1 and the ADCC activity are maintained) or different types of antibodies constructed based on them (for example, IgG type) can be used. These antibodies have the specific binding property to HER1, inhibition of binding of EGF to HER1, inhibition of phosphorylation signal of HER1 and ADCC activity. Therefore, they can specifically bind to a cancer cell expressing HER1 and inhibit HER1 activity by inhibition of binding of EGF to HER1 and/or inhibition of phosphorylation signal of HER1, thereafter, exhibit the ADCC activity so as to damage a cancer cell. Furthermore, it is confirmed that the antibody exhibits suppression effect to cancer cells and an anti-tumor effect in animal model, so that the antibody is greatly expected to be used in antibody medicine. The target cancer cell by the drug of this embodiment is not particularly limited, but it can target, for example, cells of kidney cancer, hepatic cell carcinoma, gallbladder and liver cancer, lung squamous cell carcinoma, pulmonary adenocarcinoma, and pancreas cancer.
[0697]In a further embodiment of the present invention, an anti-HER2 antibody is contained as an active ingredient. In the drug of one preferable embodiment of the present invention, anti-HER2 antibody having an ADCC activity is contained as an active ingredient. In the drug of this embodiment, the therapeutic effect can be obtained by the cytotoxicity using the ADCC activity. As anti-HER2 antibody having such an ADCC activity, 015-126 antibody shown in the below-mentioned Example (which may be partially modified as long as the specific binding property to HER2 and the ADCC activity are maintained) or different types of antibodies constructed based on them (for example, IgG type) can be used. This antibody has the specific binding property to HER2 and ADCC activity. Therefore, they can specifically bind to a cancer cell expressing HER2 then exhibits the ADCC activity so as to damage a cancer cell. Furthermore, it is confirmed that the antibody exhibits suppression effect to cancer cells, so that the antibody is greatly expected to be used in antibody medicine. The target cancer cell by the drug of this embodiment is not particularly limited, but it can target, for example, cells of kidney cancer, liver cancer, and pulmonary adenocarcinoma.
[0698]In a further embodiment of the present invention, an anti-ALCAM antibody is contained as an active ingredient. In the drug of one preferable embodiment of the present invention, anti-ALCAM antibody having an ADCC activity is contained as an active ingredient. As anti-ALCAM antibody having such an ADCC activity, 041-118 antibody or 066-174 antibody shown in the below-mentioned Example (which may be partially modified as long as the specific binding property to ALCAM and the ADCC activity are maintained) or different types of antibodies constructed based on them (for example, IgG type) can be used. This antibody has the specific binding property to ALCAM and ADCC activity. Therefore, they can specifically bind to a cancer cell expressing ALCAM then exhibits the ADCC activity so as to damage a cancer cell. The target cancer cell by the drug of this embodiment is not particularly limited, but it can target, for example, cells of pulmonary adenocarcinoma, ovarian cancer, and large bowel cancer.
[0699]In a yet further embodiment of the present invention, an anti-EpCAM antibody is contained as an active ingredient. In the drug of one preferable embodiment of the present invention, anti-EpCAM antibody having an ADCC activity is contained as an active ingredient. As anti-EpCAM antibody having such an ADCC activity, 067-153 antibody shown in the below-mentioned Example (which may be partially modified as long as the specific binding property to EpCAM and the ADCC activity are maintained) or different types of antibodies constructed based on them (for example, IgG type) can be used. This antibody has the specific binding property to EpCAM and ADCC activity. Therefore, they can specifically bind to a cancer cell expressing EpCAM then exhibits the ADCC activity so as to damage a cancer cell. The target cancer cell by the drug of this embodiment is not particularly limited, but it can target, for example, cells of gastric solid-type adenocarcinoma, colon adenocarcinoma, and pulmonary adenocarcinoma cell.
[0700]In a yet further embodiment of the present invention, an anti-CD147 antibody is contained as an active ingredient. In the drug of one preferable embodiment of the present invention, anti-CD147 antibody having an ADCC activity is contained as an active ingredient. As anti-CD147 antibody having such an ADCC activity, 059-053 antibody shown in the below-mentioned Example (which may be partially modified as long as the specific binding property to CD147 and the ADCC activity are maintained) or different types of antibodies constructed based on them (for example, IgG type) can be used. This antibody has the specific binding property to CD147 and ADCC activity. Therefore, they can specifically bind to a cancer cell expressing CD147 then exhibits the ADCC activity so as to damage a cancer cell. The target cancer cell by the drug of this embodiment is not particularly limited, but it can target, for example, kidney cancer cells.
[0701]In a yet further embodiment of the present invention, an anti-CD44 antibody is contained as an active ingredient. In the drug of one preferable embodiment of the present invention, anti-CD44 antibody having an ADCC activity is contained as an active ingredient. As anti-CD44 antibody having such an ADCC activity, 064-003 antibody shown in the below-mentioned Example (which may be partially modified as long as the specific binding property to CD44 and the ADCC activity are maintained) or different types of antibodies constructed based on them (for example, IgG type) can be used. This antibody has the specific binding property to CD44 and ADCC activity. Therefore, they can specifically bind to a cancer cell expressing CD44 then exhibits the ADCC activity so as to damage a cancer cell. The target cancer cell by the drug of this embodiment is not particularly limited, but it can target, for example, pulmonary adenocarcinoma cells.
[0702]In a yet further embodiment of the present invention, an anti-HGFR antibody is contained as an active ingredient. In the drug of one preferable embodiment of the present invention, anti-HGFR antibody having an ADCC activity is contained as an active ingredient. As anti-HGFR antibody having such an ADCC activity, 067-133 antibody shown in the below-mentioned Example (which may be partially modified as long as the specific binding property to HGFR and the ADCC activity are maintained) or different types of antibodies constructed based on them (for example, IgG type) can be used. This antibody has the specific binding property to HGFR and ADCC activity. Therefore, they can specifically bind to a cancer cell expressing HGFR then exhibits the ADCC activity so as to damage a cancer cell. The target cancer cell by the drug of this embodiment is not particularly limited, but it can target, for example, pulmonary adenocarcinoma cells.
[0703]The present invention furthermore provides a method of reducing the grade of malignancy of a target cell or promoting the normalization by damaging or suppressing the expression of HER1, HER2, CD46, ITGA3, ICAM1, ALCAM, or CD147 in the target cell.
[0704]Herein, the present inventors have investigated and recognized specific expression of CD46 in gallbladder and liver cancer and pancreas cancer, which had not been particularly reported about the relationship with respect to CD46 (see the below-mentioned Example). Similarly, the relationship between gallbladder and liver cancer and pancreas cancer and the expression of ITGA3; the relationship between kidney cancer, hepatic cell carcinoma and gallbladder and liver cancer and ALCAM; as well as the relationship between kidney cancer and CD147 have been clarified (see the below-mentioned Example). Based on the findings, a novel and effective target cell of CD46 is a gallbladder and liver cancer cell and a pancreas cancer cell; a novel and effective target cell of ITGA3 is a gallbladder and liver cancer cell and a pancreas cancer cell; and a novel and effective target cell of CD147 is a kidney cancer cell.
[0705]Note here that the inhibition or suppression of each antigen can be carried out by using an antisense method or RNA interference, or by using ribozyme.
[0706]In the case where expression inhibition by the antisense method is carried out, for example, when transcription is carried out in the target cell, an antisense-construct for generating RNA that is complementary to a portion specific to mRNA encoding this protein is used. Such an antisense-construct is introduced into the target cells, for example, in a form of an expression plasmid. On the other hand, when it is introduced in to the target cells as the antisense-construct, it is possible to employ an oligonucleotide-probe that is hybridized with mRNA or genome DNA sequence encoding this protein and inhibits the expression thereof. As such an oligonucleotide-probe, one having a low resistance to endogenous nuclease such as exonuclease and/or endonuclease is preferably used.
[0707]When DNA molecule is used as an antisense nucleic acid, it is preferable that oligodeoxyribonucleotide derived from a region (for example, a region from -10 to +10) including a translation initiation site of mRNA encoding this protein is used.
[0708]It is preferable that the complementation between the antisense nucleic acid and the target nucleic acid is strict. However, some mismatch may be accepted. The hybridization performance of the antisense nucleic acid with respect to the target nucleic acid is generally dependent upon both the degree of complementation of both nucleic acids and the length thereof. In general, as the antisense nucleic acid to be used is longer, even if the number of mismatch is increased, stable two heavy chains (or three heavy chains) can be formed between the antisense nucleic acid and the target nucleic acid. Persons skilled in the art can confirm the degree of permissible degree of the mismatch by using a standard technique.
[0709]The antisense nucleic acid may be DNA, RNA or a chimera mixture thereof, or derivative or modified type thereof. Furthermore, it may be single stranded or double stranded. By modifying a base portion, a sugar portion or a skeleton portion of phosphoric acid, the stability and hybridization performance and the like of the antisense nucleic acid can be improved. Furthermore, to the antisense nucleic acid, materials for urging the cell membrane transportation (for example, see Letsinger et al., 1989, Proc. Natl. Acad. Sci. U.S.A. 86:6553-6556; Lemaitre et al., 1987, Proc. Natl. Acad. Sci. 84:648-652; PCT Publication No. WO88/09810, published Dec. 15, 1988) or materials capable of enhancing the affinity with respect to certain cells may be added.
[0710]The antisense nucleic acid can be synthesized by a conventional method, for example, by using commercially available automated DNA synthesizer (for example, Applied Biosystems, and the like). For producing the modulated product or derivative of nucleic acid, you can see, for example, Stein et al. (1988), Nucl. Acids Res. 16:3209, or Sarin et al., (1988), Proc. Natl. Acad. Sci. U.S.A. 85:7448-7451.
[0711]In order to enhance the effect of antisense nucleic acid in the target cells, a strong promoter such as pol II and pol III can be used. That is to say, if a construct including antisense nucleic acid disposed under control of such promoters is introduced into the target cells, it is possible to secure the transcription of sufficient amount of antisense nucleic acid by the effect of the promoter.
[0712]The antisense nucleic acid can be expressed by using any promoters (derivative promoters or constitutive promoters) known to function in the mammalian cells (preferably, human cells). For example, promoters such as a SV40 initial promoter region (Bernoist and Chambon, 1981, Nature 290:304-310), a promoter derived from the 3'-terminal region of Rous sarcoma virus (Yamamoto et al., 1980, Cell 22:787-797), a Herpetic Thymidine Kinase promoter (Wagner et al., 1981, Proc. Natl. Acad. Sci. U.S.A. 78: 1441-1445), and the like, can be used.
[0713]In one embodiment of the present invention, the expression of the protein is inhibited by RNA interference (RNAi). RNAi is a process of a sequence specific post-transcriptional gene suppression that can be caused in the eukaryote. In the RNA interference, double stranded RNA (dsRNA) having a sequence corresponding to the sequence of the target mRNA is used. It is known that mammalian cells have two routes (a sequence specific route and a sequence nonspecific route) affected by dsRNA. In the sequence specific route, relatively long dsRNA is divided into short interference RNAs (siRNAs). Each of the siRNAs has sense and antisense chains of about 21 nucleotides that form siRNA of about 19 nucleotides having protruding portions at the 3' terminal portion. On the other hand, it is thought that a sequence nonspecific route can be caused by arbitrary dsRNA regardless of the sequence as long as it has a predetermined length or longer. In this route, dsRNA, two enzymes, that is, PKR, which becomes an active from and stops whole synthesis of proteins by phosphorylating the translation initiation factor eIF2, and 2', 5' oligoadenylate synthetase, which is involved in the synthesis of an RNAase L activated molecule are activated. In the method of the present invention, in order to minimize the progress of this nonspecific route, it is preferable to use dsRNA including about 30 base pairs or less (see, for example, Hunter et al. (1975) J Biol Chem 250: 409-17; Manche et al. (1992) Mol Cell Biol 12: 5239-48; Minks et al. (1979) J Biol Chem 254: 10180-3; and Elbashir et al. (2001) Nature 411: 494-8).
[0714]Note here that it is confirmed that RNAi is an effective means for reducing the gene expression in various cells (for example, a HeLa cell, a NIH/3T3 cell, a COS cell, a 293 cell, and the like). Furthermore, in general, it can inhibit expression more effectively than by the antisense method.
[0715]The dsRNA used in RNAi can be prepared in vitro or in vivo by chemical synthesis or by using an appropriate expression vector. In the latter method, it is particularly effective to prepare a relatively long dsRNA. For designing dsRNA, in general, sequence peculiar to the target nucleic acid (continuous sequence) is used. Note here that a program and algorithm for selecting an appropriate target sequence have been developed.
[0716]In another embodiment of the present invention, the expression of ITGA3 is carried out by using ribozyme. By using ribozyme for cleave mRNA at the site specific recognition sequence, it is possible to destroy mRNA encoding the protein. However, preferably, a hammerhead ribozyme is used. A method for constructing the hammerhead ribozyme can be seen in, for example, Haseloff and Gerlach, 1988, Nature, 334: 585-591.
[0717]Similar to the antisense method, for example, for the purpose of the stability and target performance, by using a modified oligonucleotide, ribozyme may be constructed. In order to produce an effective amount of ribozyme in the target cells, for example, under the control of a strong promoter (for example, pol II and pol III), it is preferable that the nucleic acid construct in which DNA encoding ribozyme is disposed is used.
[0718]Drugs used for the treatment method (including a method of urging to reducing or normalizing the grade of malignancy of cancer cells, and the like) of the present invention can be formulated according to the conventional method. In formulation, other ingredients acceptable for formulation (for example, carrier, vehicle, disintegrating agents, buffer agent, emulsifying agent, suspending agent, soothing agent, stabilizer, preservative, preservative, physiological saline, and the like) can be contained. An example of the vehicle may include lactose, starch, sorbitol, D-mannitol, and sucrose. An example of the disintegrating agents may include starch, carboxymethyl cellulose, calcium carbonate, and the like. An example of the buffer agent may include phosphate, citrate, acetate, and the like. An example of the emulsifying agent may include gum Arabic, alginate sodium, tragacanth, and the like. An example of the suspending agent may include glyceryl monostearate, aluminum monostearate, methylcellulose, carboxymethyl cellulose, hydroxymethyl cellulose, sodium lauryl sulfate, and the like. An example of the soothing agent may include benzyl alcohol, chlorobutanol, sorbitol, and the like. An example of the stabilizer may include propylene glycol, diethylene sulfite, ascorbic acid, and the like. An example of the preservative may include phenol, benzalkonium chloride, benzyl alcohol, chlorobutanol, methylparaben, and the like. An example of the preservative may include benzalkonium chloride, parahydroxybenzoate, chlorobutanol, and the like.
[0719]The dosage form in the formulation is not particularly limited. An example of the dosage form may include tablet, powdered drug, fine subtilae, granule, capsules, syrup, injectable drug, external preparation, and suppository.
[0720]In the treatment using the drug of the present invention, the drug of the present invention is administered to a subject (patient) with a cancer cell or adult T cell leukemia. The drug of the present invention can be administered to a subject (patient) by oral administration or parenteral administration (intravenous, intra-arterial, subcutaneous, intramuscular, intraperitoneal injection, direct introduction to the target cell, and the like) depending upon the dosage form.
[0721]The dosage amount of the drug of the present invention will vary depending on the symptoms, age, sex, body weight, and the like, of the patient, but the person skilled in the art can set an appropriate dosage amount. For example, the dosage amount can be set so that the dosage amount of effective ingredient for adult (body weight: about 60 kg) per day is about 0.001 mg to about 100 mg. The administration schedule can include, for example, once to several times a day, once per two days, or once per three days. For setting the administration schedule, conditions of a patient, efficacy duration time of the drug, and the like, can be considered.
[0722]In another embodiment, the drug of the present invention uses anti-HER1 antibody, anti-HER2 antibody, anti-CD46 antibody, anti-ITGA3 antibody, anti-ICAM1 antibody, anti-ALCAM antibody, anti-CD147 antibody as a carrier for DDS. That is to say, this embodiment provides an immunocomplex obtained by combining a drug (cytotoxin and the like), radioactive isotope, or the like (these are also referred to as "active ingredient" together) to anti-HER1 antibody, and others. The immunocomplex containing a drug (cytotoxin) having a cell-killing activity or a cytotoxic activity is generally referred to as immunotoxin. An example of the cytotoxin may include Taxol, cytochalasin. B, gramicidin D, ethidium bromide, emetine, mitomycin, etoposide, tenoposide, vincristine, vinblastine, colchicines, doxorubicin, daunorubicin, dihydroxy-anthracene-dione, mitoxantrone, methramycin, actinomycin D, 1-dehydrotestosterone, glucocorticoid, procaine, tetracaine, lidocaine, propranolol, and puromycin as well as analogue or homologue thereof.
[0723]As the active ingredient contained in the immunocomplex of the present invention, protein or peptide having a desirable biological activity may be used. An example of the candidate for protein and the like that can be used for such a purpose may include abrin, ricin A, Pseudomonas-exotoxin, diphteria toxin, tumor necrosis factor, interferon-γ, interleukin I (IL-1), interleukin 2 (IL-2), interleukin 6 (IL-6), a granulocyte macrophage colony stimulating factor (GM-CSF), a granulocyte colony stimulating factor (G-CSF) lymphokine.
[0724]A technology for combining an active component to an antibody is well known and you can see in, for example, Monoclonal Antibodies And Cancer Therapy, Reisfeld et al. (eds.), pp. 243-56 (Alan R. Liss, Inc. 1985), Controlled Drug Delivery (2nd edition.), Robinson et al. (eds.), pp. 623-53 (Marcel Dekker, Inc. 1987), Monoclonal Antibodies For Cancer Detection And Therapy, Baldwin et al. (eds.), pp. 303-16 (Academic Press 1985), Thorpe et al., "The Preparation And Cytotoxic Properties Of antibody-Toxin Conjugates", Immunol. Rev., 62:119-58 (1982).
(Kit Used in the Present Invention)
[0725]Each method of the present invention (a method for obtaining information for diagnosis, and the like) may be carried out by using a kit of reagent and the like. Another aspect of the present invention provides a kit used for such a purpose. For example, nucleic acid (probe and primer), reaction reagent, dilution, a reactor vessel, and the like, that are used for the method of the present invention can be contained in the kit. Note here that the kit of the present invention is generally includes instruction.
[0726]The user of a kit makes it possible to allow the method of the present invention to be carried out in a simple way and for a short time.
Example
1. Production of Vector for Producing ScFv Antibody Gene Library
[0727]1-1 Production of Vector for Producing scFV Antibody Gene Library
[0728]As conceptually shown in FIG. 5, pelB (signal sequence) of M13 phage, His6 tag sequence, cp3 protein of M13 phage (Δcp3 (198aa-406aa) N-terminal deleted capsid protein 3) sequence, protein A protein sequence were incorporated in an appropriate restriction enzyme site of a pTZ19R phagemid vector (Pharmacia) so as to from a vector pAALFab (see Iba Y. et al., Gene 194: 35-46, 1997.). A vector pFCAH9-E8d for incorporation was produced from this pAALFab.
[0729]Genes of a heavy chain and a light chain are inserted into the predetermined position of this vector, thereby completing an actual antibody protein expression vector. The shape of the antibody expressed by the completed vector is a scFv and a light chain constant region CL gene is bonded to the aforementioned cp3 gene. As a result, expression protein has a shape of scFv-CL-cp3. Specifically, the below-mentioned operation is carried out.
Used primer:
TABLE-US-00001 527 Reverse (SEQ ID NO: 377): 5'-CAGGAAACAGCTATGAC-3' 599 E8VHf-PstR: (SEQ ID NO: 378) 3'-CGGCTCCAAGTCGACGTCGTCA-5' 544 E8VHf-PstF: (SEQ ID NO: 379) 5'-CAGCTGCAGCAGTCTGGGGCAGAGCTTGTGAAGCCAGGGGCCTCAGT CAAGTTGTCCTGCACAGCTTCTGGCTTCAACATTAA-3' 545 E8VHf-XbaR: (SEQ ID NO: 380) 3'-AGACCGAAGTTGTAATTTCTGTGGATATACGTGACCCACTTCGTCTC CGGACTTTTCCCAGATCTCACCTAACCTTCCTAA-5' 546 E8VHf-XbaF: (SEQ ID NO: 381) 5'-AAGGGTCTAGAGTGGATTGGAAGGATTGATCCTGCGAGTGGTAATAC TAAATATGACCCGAAGGACAAGGCCACTATAACAGCA-3' 547 E8VHf-EcoR (SEQ ID NO: 382) 3'-TTCCTGTTCCGGTGATATTGTCGTCTGTGTAGGAGGTTGTGTCGGAT GGATGTCGACTTAAGGGAC-5' 548 E8VHf-EcoF (SEQ ID NO: 383) 5'-CAGCTGAATTCCCTGACATCTGAGGACACTGCCGTCTATTACTGTGC TGGT-3' 549 E8VHf-BstR (SEQ ID NO: 384): 3'-CAGATAATGACACGACCAATACTAATGCCGTTGAAACTGATGACCCC GGTTCCGTGGTGCCAGTGGCACAAGG-5' 590 His6-SmaR (SEQ ID NO: 385): 3'-GGTTCTCTAACAGTAGTGGTAGTAGTGGTAATTATTCTCGATAGGGC CCTCGAA-5' 542 E8VLf-SacF (SEQ ID NO: 386): 5'-GACATCGAGCTCACCCAGTCTCCAGCCTCCCTTTCTGCGTCTGTGGG AGAAACTGTCACCATCACATGT-3' 539 E8VLf-KpnR (SEQ ID NO: 387): 3'-TGACAGTGGTAGTGTACAGCTCGTTCACCCTTATAAGTGTTAATAAA TCGTACCATGGTCGTC-5' 542 E8VLf-KpnF (SEQ ID NO: 388): 5'-GCATGGTACCAGCAGAAACCAGGGAAATCTCCTCAGCTCCTGGTCTA T-3' 543 E8VLf-BamR (SEQ ID NO: 389): 3'-GGAGTCGAGGACCAGATATTACGTTTTTGGAATCGTCTACCACACGG TAGTTCCAAGTCACCGTCACCTAGGCCTTGTGTT-5' 562 E8VLf-XhoR (SEQ ID NO: 390): 3'-TCATGAGGCACCTGCAAGCCACCTCCGTGGTTCGAGCTCTAGTT T-5' 563 E8VLf-XhoF (SEQ ID NO: 391): 5'-AGTACTCCGTGGACGTTCGGTGGAGGCACCAAGCTCGAGATCAA A-3' 613 NheR (SEQ ID NO: 392): 3'-ATCGACAGCT-5' 600 E8VLKpnXhoR (SEQ ID NO: 393): 3'-AAGCCACCTCCATGGTTCGAGCTCTAGTTT-5' LCP3ASC (SEQ ID NO: 394): 3'-TCGAAGTTGTCCTTACTCACAAGCCGCGCGGTCAGCTGAGGTAA-5' hCH1Bst (SEQ ID NO: 395): 5'-ACCCTGGTCACCGTCTCCTCAGCCTCCACCAAGGGCCCATCGGTCTT CCCCCTGG-3' hCH1midAS (SEQ ID NO: 396): 3'-GGGAGTCGTCGCAGCACTGGCACGGGAGGTCGTCGAA-5' hCH1midS (SEQ ID NO: 397): 5'-GGACTCTACTCCCTCAGCAGCGTCGTGACCGTGCCC-3' hCH1H6 (SEQ ID NO: 398): 3'-GGGTCGTTGTGGTTCCACCTGTTCTTTCAACTCGGGTTTAGAACAGT AGTGGTAGTAGTGGTA-5' hCH1H6Sma (SEQ ID NO: 399): 3'-GGGTTTAGAACAGTAGTGGTAGTAGTGGTAATTATTCTCGATAGGGC CCTCGAACG-5' 702 BstXhoF (SEQ ID NO: 400): 5'-GGCACCACGGTCACCGTCTCGAGCGCCTCCACC-3'
<Production of pFCAH3-E8T H Chain Part>1) By using pAALFab as a template, PCR using 527-599 and PCR using 547-590 were carried out so as to produce a DNA fragment.2) PCR using 544-545, 546-547, and 548-549 was carried out so as to produce a DNA fragment.3) 1) and 2) were mixed and PCR by 527,590 was carried out, which was cloned to a HindIII-SmaI site of pAALFab.<pFCAH3-E8T L Chain Part>4) PCR using 542-562 and 561-613 was carried out so as to produce a DNA fragment.5) PCR using 538-539 and 542-543 was carried out so as to produce a DNA fragment.6) 4) and 5) were mixed and PCR by 538, 562 was carried out, which was cloned to a SacI-NheI site of pAALFab.<pFCAH9-E8d>
7) Production of VH Stuffer Part
[0730]pFCAH3-E8T was digested with XbaI and EcoRI and a klenow fragment was acted thereon so as to be blunted. Thereafter, the self ligation was carried out so as to produce a stuffer of the VH part.
8) Production of VL Stuffer Part
[0731]By using pFCAH3-E8T as a template, PCR with 527-600 was carried out, which was cloned to the HindIII-XhoI site in 7).
9) This was digested with KpnI and subjected to self ligation so as to produce a stuffer of a VL part.
10) Introduction of SfiI, NcoI, SpeI Sites
[0732]By using pFCAH3-E8T as a template, PCR with 527-663 was carried out, which was cloned to the HindIII-SacI site in 1).
11) Introduction of AscI Site
[0733]By using pFCAH3-E8T as a template, PCR with 527-LCP3ASC was carried out, which was cloned to 2) which was completely digested with SacI and partially digested with SalI.
12) Transform of GammaCH1 Part into Human Gene
[0734]Since human gamma CH1 part has BstPI site, cloning was carried out so as to design this site. By using tonsil cDNA as a template, PCR with hCH1Bst-hCH1midS, hCH1midAS-hCH1H6 was carried out and then mixed. PCR with hCH1Bst-hCH16Sma was carried out and the DNA fragment was cloned to the BstPI-Sma site in 3).
13) Introduction of Xho Site
[0735]By using 12) as a template, PCR with 702-663 was carried out and this was cloned to the BstPI-SacI site in 12).
<Production of pscFvCA9-E8VHdVLd>
[0736]pFCAH9-E8d 3 μg (3 μL) (see FIG. 5D) was mixed with BstPI (3 U/μL) (3 μL), 10×H buffer (5 μL), DW (39 μL) and subjected to restriction enzyme treatment at 37° C. for two hours. After treatment, precipitates obtained by ethanol precipitation were dissolved in 10 μL of TE buffer. To this solution, SacI (10 U/μL) (1 μL), 10×L buffer (5 μL) and DW (34 μL) were mixed. Then, this mixture was subjected to restriction enzyme treatment at 37° C. for two hours and to agarose gel electrophoresis. Thus, 4.7 kb fragment was recovered. The recovered products were subjected to ethanol precipitation to give 10 μL (pFCAH9-E8d BstPI-SacI fragment).
[0737]On the other hand, a primer linF (100 pmol/μL) (5 μL) and a primer linR (100 pmol/μL) (5 μL) were mixed and heated at 94° C. for 5 minutes, and then annealed at 80° C. for 5 minutes, at 70° C. for 5 minutes, and at room temperature for 30 minutes. Two μL of which was mixed with the above-obtained pFCAH9-E8d BstPI-SacI fragment (1 μL), 10× ligation buffer (1.5 μL), DW (9.5 μL), and T4DNA ligase (I μL) and reacted at 16° C. for 16 hours. After reaction, the reacted product was subjected to ethanol precipitation to concentrate to 3 μL. 1.5 μL of them was used to transform E. coli DH12S competent cells (20 μL) by electroporation. The obtained plasmid clone was extracted and the base sequence thereof was confirmed. This was named pscFvCA9-E8VHdVLd. FIG. 6 schematically shows a structure of pscFvCA9-E8VHdVLd. Furthermore, FIGS. 7-1 to 7-2 show the base sequence (SEQ ID NO: 401) of the insert part of pscFvCA9-E8VHdVLd and the amino acid sequence (SEQ ID NO: 402) encoded thereby, respectively.
TABLE-US-00002 primer linF (SEQ ID NO: 403) GTCACCGTCTCGAGAGGCGGTGGCGGATCAGGTGGCGGTGGAAGTGGCGG TGGTGGGTCCATGGCCGACATCGAGCT primer linR (SEQ ID NO: 404) CGATGTCGGCCATGGACCCACCACCGCCACTTCCACCGCCACCTGATCCG CCACCGCCTCTCGAGACG
1-2 Production of Vector for Temporarily Cloning Heavy Chain Variable Region (VH)
[0738]According to the well-known technique (see Iba Y. et al., Gene 194:35-46, 1997.), firstly, a pAALFab vector (FIG. 5A) was produced. A portion between XbaI and EcoRI was deleted from the pAALFab vector, and the restriction enzyme cut sites Kpn I, Sfi I, Nco I, and Spe I were newly added. Through pFCAH3-E8T (FIG. 5B), a vector pscFvCA-E8VHd (FIG. 5C) capable of cloning VH (heavy chain variable region) was produced. Thus, a vector for temporarily cloning the heavy chain variable region (VH) was obtained. FIGS. 8-1 to 8-2 show the base sequence (SEQ ID NO: 405) of the insert of pscFvCA-E8VHd, the restriction enzyme site and the amino acid sequence (SEQ ID NO: 406) encoded by the base sequence.
[0739]Specifically, the primer 610 and the primer 611 were annealed and annealed produced was cloned to a BstPI-SacI site of pFCAH3-E8T. Thus, a single chain was produced. Furthermore, PCR with the primer 527 and the primer 619 was carried out and this was further cloned to a HindIII-PstI site. Thus, introduction of SfiI, NcoI site was carried out. Hereinafter, primer sequences used for producing the vector are shown.
TABLE-US-00003 610 scBstSpeSacF (SEQ ID NO: 407): 5'-CACCACGGTCACCGTCTCCTCAGGCGGTGGCGGATCAGGTGGCGGTG GAAGTGGCGGTGGTGGGTCTACTAGTGACATCGAGCTCACCCAG-3' 611 scBstSpeSacR (SEQ ID NO: 408): 3'-GTGGTGCCAGTGGCAGAGGAGTCCGCCACCGCCTAGTCCACCGCCAC CTTCACCGCCACCACCCAGATGATCACTGTAGCTCGAGTGGGTC-5' 527 Reverse (SEQ ID NO: 409): 5'-CAGGAAACAGCTATGAC-3' 619 E8VHf-SftNcoPstR (SEQ ID NO: 410): 3'-GACGCCGGGTCGGCCGGTACCGGCTCCAAGTCGACGTCGTCA-5'
2. Production of Immunoglobulin Light Chain Library
2-1 Isolation of Immunoglobulin Light Chain Gene by Using PCR
[0740]From bone marrow cells (sample No. 59) 4×107 cells, and lymphocytes of cord blood and peripheral blood, by using a commercially available kit (Pharmacia Biotech, QuickPrep Micro mRNA Purification Kit), 2.6 μg of mRNA was obtained. From this mRNA, cDNA was produced. The cDNA was produced by using SuperScriptPreamplification System (GibcoBRL). As a primer, oligo dT was used. PCR using the obtained cDNA as a template was carried out by using 5' primer (κ1-κ6, λ1-λ6) and 3' primer (hCKASC primer or hCLASC primer) for obtaining light chain genes. The PCR product was treated with phenol, subjected to ethanol precipitation and suspended in 10 μL of TE buffer. The base sequence of primer and conditions of PCR are shown below. In the base sequence of a primer for obtaining light chain genes, underline part represents NcoI site and AscI site.
TABLE-US-00004 5' primer κ1-κ6 hVK1a (SEQ ID NO: 411): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCCGACATCCAGATGACCCA GTCTCC hVK2a (SEQ ID NO: 412): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC GATGTTGTGATGACTC AGTCTCC hVK3a (SEQ ID NO: 413): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC GAAATTGTGTTGACGC AGTCTCC hVK4a (SEQ ID NO: 414): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC GACATCGTGATGACCC AGTCTCC hVK5a (SEQ ID NO: 415): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC GAAACGACACTCACGC AGTCTCC hVK6a (SEQ ID NO: 416): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC GAAATTGTGCTGACTC AGTCTCC 5' primer λ1-λ6 hVL1 (SEQ ID NO: 417): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC CAGTCTGTGTTGACGC AGCCGCC hVL2 (SEQ ID NO: 418): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC CAGTCTGCCCTGACTC AGCCTGC hVK3a (SEQ ID NO: 419): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC TCCTATGTGCTGACTC AGCCACC hVL3b (SEQ ID NO: 420): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC TCTTCTGAGCTGACTC AGGACCC hVL4 (SEQ ID NO: 421): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC CACGTTATACTGACTC AACCGCC hVL5 (SEQ ID NO: 422): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC CAGGCTGTGCTCACTC AGCCGCC hVL6 (SEQ ID NO: 423): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC AATTTTATGCTGACTC AGCCCCA 3'-primer hCKASC (SEQ ID NO: 424): TCGACTGGCGCGCCGAACACTCTCCCCTGTTGAAGCTCTTTGTG 3'-primer HCLASC (SEQ ID NO: 425): TCGACTGGCGCGCCGAACATTCTGTAGGGGCCACTGTCTTCTC
Conditions of PCR
TABLE-US-00005 [0741] cDNA 2 μL 10 × buffer # 1 (attached to KOD) 10 μL dNTP mix (2.0 mM) 10 μL 25 mM MgCl2 4 μL 5' side primer (100 pmol/μL) 1 μL 3' side primer (100 pmol/μL) 1 μL sterilized MilliQ 71 μL
[0742]KOD DNA polymerase (TYOBO CO LTD., 2.5 U/mL) 1 μL
[0743]35 cycles, each cycle includes 94° C. for one minute, 55° C. for two minutes and 74° C. for one minute
2-2-1 Incorporation of Light Chain Gene into Phagemid
[0744]The PCR product obtained in 1 was treated with a restriction enzyme in the following conditions.
TABLE-US-00006 PCR product 10 μL 10 × NEB4 (attached to AscI) 5 μL Sterilized MilliQ 33 μL AscI (NEB, 10 U/μL) 1 μL NcoI (TAKARA SHUZO, 10 U/μL) 1 μL
[0745]After the reaction at 37° C. for one hour and at 50° C. for one hour, 10 μL of the reacted product was subjected to agarose gel electrophoresis and 600 bp band was cut out to be purified by using geneclean II kit (Funakoshi Corporation). Similar to the PCR product, restriction enzyme-treated pscFvCA9-E8VHdVLd was purified by using geneclean II kit and reacted with the restriction enzyme-treated PCR product at 16° C. for four hours to overnight in the following conditions, thereby carrying out ligation.
TABLE-US-00007 restriction enzyme-treated pscFvCA9-E8VHdVLd 2 μL restriction enzyme-treated PCR product 1 μL 10 × ligation buffer 1.5 μL (attached to T4 DNA ligase) 10 mM ATP 1.5 μL sterilized MilliQ 8 μL T4 DNA Iigase (TAKARA SHUZO 10 U/μL) 1 μL
2-2-2 Introduction of Phagemid into E. coli
[0746]The obtained ligated DNA was used so as to transform E. coli DH12S as follows. That is to say, ligated DNA was subjected to ethanol precipitation once, and dissolved in 3 μL of 1/5 TE (TE that was 5-fold diluted with sterilized MilliQ). 1.5 μL of them was suspended in 20 μL of competent cell DH12S (GIBCO BRL), which was subjected to electroporation in the following conditions.
TABLE-US-00008 Electroporator Cell-Porator (Cat. series 1600), product of BRL Setting conditions; voltage booster 4 kΩ capacitance 330 μF DC volts LowΩ charge rate Fast
[0747]The above-mentioned transformed E. coli was planted on a transformation medium (SOB) (2 mL) and shaking cultured at 37° C. for one hour. Then, a part of the cultured product was planted on agar medium (Amp plate) and a remaining part was cultured in a 2×TY medium containing 0.1% glucose and 100 μg/mL ampicillin to form glycerine stock. The agar medium was incubated at 30° C. and growing colony was separated by picking by a picker. A plasmid was prepared, respectively. Then, the light chain gene and the base sequence were examined.
[0748]SOB medium: to 950 mL of purified water, the following components were added and shaken so as to be dissolved completely. Thereafter, 250 mM KCl solution (10 mL) was added so as to adjust to pH 7.0 with 5N NaOH. Purified water was added to adjust to 1000 mL, then sterilized for 20 minutes in the autoclave. Immediately before the use, 5 mL of 2M sterilized MgCl2 was added.
TABLE-US-00009 bacto-tryptone 20 g bacto-yeast extract 5 g NaCl 0.5 g
[0749]2×YT medium: to 900 mL of purified water, the following components were added and shaken so as to be dissolved completely. Thereafter, 5 N NaOH was added so as to adjust to pH 7.0 with 5N NaOH. Purified water was added to adjust to 1000 mL, then sterilized for 20 minutes in the autoclave and used.
TABLE-US-00010 bacto-tryptone 16 g bacto-yeast extract 10 g NaCl 5 g
[0750]The other reagents were purchased form the following suppliers.
(Manufacture/Product name are described in this order)SIGMA/ampicillin sodium
Wako Pure Chemical/phenol
SIGMA/BSA
[0751]DIFCO/2×YT mediumWako Pure Chemical/kanamycin sulfatenacalai tesque/polyethylene glycol 6000nacalai tesque/Tween 20
KATAYAMA CHEMICAL/NaCl
Wako Pure Chemical/IPTG
[0752]Wako Pure Chemical/skim milkWako Pure Chemical/sodium azide
Wako Pure Chemical/triethylamine
[0753]Wako Pure Chemical/hydrogen peroxideWako Pure Chemical/OPD tablet
Wako Pure Chemical/ethanol
[0754]The above-mentioned operation is carried out with respect to all of κ1, κ2, κ3, κ4, κ5, and κ6, as well as λ1, λ2, λ3a, λ3b, λ4, λ5, λ6, λ7, λ8, λ9, and λ10 are operated so as to confirm whether or not the intended clones are obtained. Then, for example, κ1 and κ2, clones in each group, were mixed so that the ratio becomes near the frequency of use. The rate of expression of each group of these light chains in an actual living body is already known. These gene clones amplified by PCR method and incorporated into a vector are mixed so that the ratio becomes near the frequency of use. Thus, VL library was obtained. Constituent ratio in each family in VL library is shown below.
[Table I]
TABLE-US-00011 [0755]TABLE 1 vκ Usage Constitutive Constitutive frequency in ratio in VL ratio in family vivo (%)* library (%) KL200 (%) Vκ1 39 37 30.7 Vκ2 12 12 19.8 Vκ3 36 35 33.7 Vκ4 12 12 10.9 Vκ5 1 2 5.0 Vκ6 --** 2*** 0.0 *Griffith AD et al. EMBO J. (1994) 13, 3245-60. **Published data is not shown ***equal amount of cDNA produced with primer VK6-2 and cDNA produced with primer VK6-3 were mixed.
[Table 2]
TABLE-US-00012 [0756]TABLE 2 Vλ Usage Constitutive Constitutive frequency in ratio in VL ratio in family vivo (%)* library (%) KL200 (%) Vλ1 43 41 34.1 Vλ2 15 .sup. 15*3 15.2 Vλ3 34 .sup. 32*4 25.3 Vλ4 0 .sup. 1.6*5 0.0 Vλ5 0 .sup. 1.0*6 11.1 Vλ6 0 1.0 14.1 Vλ7 6 6 0.0 Vλ8 1 1 0.0 Vλ9 1 1 0.0 Vλ10 --*2 1 0.0 *Griffith AD et al. EMBO J. (1994) 13, 3245-60. *2Published data is not shown *3cDNA produced with primer VL2 (5%) and cDNA produced with primer VL2-2 (10%) were mixed. *4cDNA produced with primer VL3a-2 (17%) and cDNA produced with primer VL3b (15%) *5cDNA produced with primer VL4a (0.5%), cDNA produced with primer VL4b (0.5%) and cDNA produced with primer VL4c (0.5%) were mixed. *6cDNA produced with primer VLSabde (0.5%) and cDNA produced with cDNA (0.5%) were mixed.
3. Production of Combinatorial Library of Light Chain Gene Library and Heavy Chain Gene Library (scFv Antibody Gene Library)
3-1-1 Isolation of Immunoglobulin Heavy Chain Gene Using PCR
[0757]By the procedure similar to 2-1, cDNA was prepared by using cord blood, bone marrow fluid, and lymphocyte of peripheral blood as well as a human μ primer (below-mentioned primer, 634) from the tonsil or random hexamer. By using this cDNA as a template, a mixture of equal amount of 5' primer (VH1 to VH7) and 3' primer (four kinds of human JH primers are mixed in equal amount, below-mentioned primers 697 to 700) for obtaining a human antibody heavy chain gene, or human u primer (below-mentioned primer 634) were subjected to PCR. In Table, underlined parts show the SfiI site. Since hVH2a did not correspond to a germ line VH2 family, VH2a-2 was newly designed. Furthermore, since hhVH4a did not correspond to the entire VH4 family, hVH4a-2 was newly designed. Also, VH5a did not correspond to a germ line VH5 subfamily, VH5a-2 was newly designed. Furthermore, as a primer corresponding to VH7, hVH7 was designed. These were also subjected to gene amplification and incorporated into pscFvCA-E8VHd. Then, as to the obtained genes, the base sequence was determined. Since the sequence of hVH5a-2 is extremely similar to that of hVH1a and it is expected that the gene product similar to that amplified with hVH1a, this was not used. The PCR products were subjected to phenol treatment and then ethanol precipitation, and thereafter suspended in 10 μL of TE buffer.
TABLE-US-00013 634 hum μ CH1R (SEQ ID NO: 426): ATGGAGTCGGGAAGGAAGTC Primers used for amplification of each VH family Human VH primer, SfiI site is underlined. 628 hVH1a (SEQ ID NO: 427): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC CAGGTGCAGCTGGTGC AGTCTGG 629 hVH2a (SEQ ID NO: 428): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC CAGGTCAACTTAAGGG AGTCTGG 630 hVH3a (SEQ ID NO: 429): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC GAGGTGCAGCTGGTGG AGTCTGG 631 hVH4a (SEQ ID NO: 430): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC CAGGTGCAGCTGCAGG AGTCGGG 632 hVH5a (SEQ ID NO: 431): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC CAGGTGCAGCTGTTGC AGTCTGC 633 hVH6a (SEQ ID NO: 432): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC CAGGTACAGCTGCAGC AGTCAGG 629-2 hVH2a-2 (SEQ ID NO: 433): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC CAGRTCACCTTGAAGG AGTCTGGTCC 631-2 hVH4a-2 (SEQ ID NO: 434): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC CAGGTGCAGCTACAGC AGTGGGG 632-2 hVH5a-2 (SEQ ID NO: 435): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC GAGGTGCAGCTGGTGC AGTCTGG 712 hVH7 (SEQ ID NO: 436): GTCCTCGCAACTGCGGCCCAGCCGGCCATGGCC CAGGTGCAGCTGGTGC AATCTGGGTCTGAGT Human JH primer, BstPI and XhoI sites underlined. 697 hJH1-2 (SEQ ID NO: 437): GGTGGAGGCACTCGAGACGGTGACCAGGGTGC 698 hJH3 (SEQ ID NO: 438): GGTGGAGGCACTCGAGACGGTGACCATTGTCC 699 hJH4-5 (SEQ ID NO: 439): GGTGGAGGCACTCGAGACGGTGACCAGGGTTC 700 hJH6 (SEQ ID NO: 440): GGTGGAGGCACTCGAGACGGTGACCGTGGTCC
TABLE-US-00014 cDNA 2 μL 10 × buffer # 1 (attached to KOD) 10 μL dNTP mix (2.0 mM) 10 μL 25 mM MgCl2 4 μL 5' primer (100 pmol/(μL) 1 μL 3' primer (100 pmol/μL) 1 μL sterilized MilliQ 71 μL KOD DNA polymerase (TYOBO CO LTD., 2.5 U/μL) 1 μL
[0758]PCR conditions: 35 cycles, each cycle includes 94° C. for one minute, 55° C. for two minutes and 74° C. for one minute
3-1-2 Production of Heavy Chain Gene Library
[0759]The PCR product obtained in 3-1-1 was treated with a restriction enzyme in the following conditions.
TABLE-US-00015 PCR product 10 μL 10 × K buffer NEB4 (TAKARA SHUZO) 5 μL Sterilized MilliQ 33 μL SfiI (NEB, 10 U/μL) 1 μL XhoI (TAKARA SHUZO, 12 U/μL) 1 μL
[0760]After the reaction at 37° C. for two hours, 10 μL of the reacted product was subjected to agarose electrophoresis and 400 bp band was cut out to be purified by using geneclean II kit (Funakoshi Corporation). Similar to the PCR product, restriction enzyme-treated pscFvCA-E8VHd was purified by using geneclean II kit and reacted with the restriction enzyme-treated PCR product at 16° C. for four hours to overnight in the following conditions, thereby carrying out ligation.
TABLE-US-00016 restriction enzyme-treated pscFvCA-E8VHd 2 μL restriction enzyme-treated PCR product 1 μL 10 × ligation buffer 1.5 μL (attached to T4 DNA ligase) 10 mM ATP 1.5 μL sterilized MilliQ 8 μL T4 DNA ligase (TAKARA SHUZO 10 U/μL) 1 μL
3-1-3 Introduction of Phagemid into E. coli
[0761]The obtained DNA was transformed into E. coli DH12S. Specifically, DNA was subjected to ethanol precipitation once, and dissolved in 3 μL of 1/5 TE (TE that was 5-fold diluted with sterilized MilliQ). 1.5 μL of them was suspended in 20 μL of competent cell DH12S (GIBCO BRL), which was subjected to electroporation.
TABLE-US-00017 Electroporator Cell-Porator (Cat. series 1600), product of BRL Setting conditions; voltage booster 4 kΩ capacitance 330 μF DC volts LowΩ charge rate Fast
[0762]The above-mentioned transformed E. coli was planted on a transformation medium (SOB) (2 mL) and shaking cultured at 37° C. for one hour. Then, a part of the cultured product was planted on agar medium (Amp plate) and a remaining part was cultured in a 2×YT medium containing 0.1% glucose and 100 μg/mL ampicillin to form glycerine stock. The agar medium was incubated at 30° C. and growing colony was separated by picking by a picker. A plasmid was prepared, respectively. Then, the heavy chain gene and the base sequence were examined. All of the VH1 to VH7 were treated in the same way to confirm whether or not the target clone was obtained. These clones of each group (family) were mixed so that the ratio was near the use frequency in vivo. Thus, VH library was produced. The constitution ratio of each family in the VH library is shown below.
[Table 3]
TABLE-US-00018 [0763] TABLE 3 Usage Constitutive frequency in ratio in VH family vivo (%)* library (%) VH1 25 29** VH2 6.6 7 VH3 40 40 VH4 19 19*** VH5 5 --** VH6 3.8 4 VH7 1.2 2 *Griffith AD et al. EMBO J. (1994) 13, 3245-60. **Actually, since VH1 and VH5 are amplified with the same primer, they cannot be counted separately. ***cDNA produced with VH4 primer and cDNA produced with VH4-2 primer were mixed in this ratio.
3-2 Production of Combinatorial Gene Library
[0764]VH library (200 μg) was digested with HindIII and XhoI under the following conditions and heavy chain gene is cut out and purified by using geneclean II kit.
TABLE-US-00019 VH library 200 μg 100 μL 10 × K buffer (TAKARA SHUZO) 40 μL sterilized MilliQ 205 μL HindIII (TAKARA SHUZO40 U/μL) 30 μL XhoI (TAKARA SHUZO50 U/μL) 25 μL
[0765]A vector pscFvCA9-E8VHdVLd in which a VL library had been inserted was was digested with HindIII and XhoI under the following conditions, and a fragment containing a light chain gene was purified by using geneclean II kit.
TABLE-US-00020 pscFvCA9-E8VHdVLd in which a VL 100 μg, 100 μL library had been inserted 10 × K buffer (TAKARA SHUZO) 40 μL sterilized Milli-Q 230 μL HindIII (TAKARA SHUZO 40 U/μL) 15 μL XhoI (TAKARA SHUZO 50 U/μL) 15 μL
[0766]Next, a VH gene library fragment and a pscFvCA9-E8VHdVLd vector into which a light chain gene has been inserted were reacted at 16° C. overnight in the following conditions so as to be ligated.
TABLE-US-00021 restriction enzyme-treated VH library fragment 10 μg 50 μL pscFvCA9-E8VHdVLd containing restriction 40 μg 50 μL enzyme-treated VL library fragment 10 × ligation buffer (attached to T4 DNA ligase) 100 μL 10 mM ATP 100 μL Sterilized MilliQ 670 μL T4 DNA ligase (TAKARA SHUZO 10 U/μL) 30 μL
[0767]The DNA in which the reaction had been completed was used to transform E. coli DH12S. Specifically, DNA was subjected to ethanol precipitation once, and dissolved in 30 μL of 1/5 TE (TE 5-fold diluted with sterilized MilliQ). This was suspended in 500 μL of competent cell DH12S (GIBCO BRL), and electroporation was carried out.
TABLE-US-00022 Electroporator Cell-Porator (Cat. series 1600), product of BRL Setting conditions; voltage booster 4 kΩ capacitance 330 μF DC volts LowΩ charge rate Fast
[0768]The above-mentioned transformed E. coli was planted on a transformation medium (SOB) (12 mL) and shaking cultured at 37° C. for one hour. Then, a part of the cultured product was planted on agar medium (Amp plate) and a remaining part was cultured in a 2×YT medium (500 mL) containing 0.1% glucose and 100 μg/mL ampicillin to form glycerine stock. The agar medium was incubated at 30° C. and the number of clones were estimated from the number of growing colonies. 8.5×1010 clones were obtained.
4. Production of scFv-CL Antibody Phage Library from scFv-CL Antibody Gene Library
[0769]To 16 of 5-liter flasks containing 300 mL of 2×YT medium to which 1% glucose and 100 μg/mL ampicillin had been added, 2.5 mL of AIMS-5 suspension was added and shaking cultured at 37° C. Every one hour, the absorbance at the wavelength of 600 nm was measured and the culture solution was proliferated until the absorbance became 1.0. To the culture solution, 12 mL each of helper phage solution (M13KO7) was added for each flask so as to infect the helper phage, culture at 37° C. for two hours. Thus, phage infected DH12S was obtained.
[0770]To 24 of 5-L flasks, 2×YT medium (600 mL), 100 μg/mL ampicillin (0.6 mL), 50 μg/m 38 L kanamycin (0.8 mL), and helper phage infected DH12S (200 mL) were added and shaking cultured at 37° C. for 20 hours.
[0771]The bacterial cells were centrifuged at 8000 rpm at 4° C. for 10 minutes, and supernatant was recovered. 4 L of 20% polyethylene glycol/2.5M NaCl was added to the supernatant, after it was quietly stirred for about 20 minutes, centrifuged at 8000 rpm at 4° C. for 20 minutes. The precipitate was dissolved in 1 L of PBS, 200 mL of 20% polyethylene glycol/2.5M NaCl was added thereto, after it was quietly stirred for about 20 minutes, and centrifuged at 8000 rpm at 4° C. for 20 minutes. The supernatant was discarded and further, centrifuged at 8000 rpm at 4° C. for 3 minutes, and the precipitate was recovered. The precipitate was dissolved in PBS to which 0.05% NaN3 was added, after it was centrifuged at 1000 rpm at 4° C. for 15 minutes and the supernatant was recovered, further, centrifuged at 8000 rpm at 4° C. for 3 minutes and the supernatant was recovered.
[0772]The titer of the recovered phage solution was checked as followings: the phage solution was diluted with PBS in 106, 107 and 108-fold, out of these, 10 μL was infected with 990 μL of DH12S, cultured at 37° C. for one hour. 100 μL of them was plated on LBGA plate and cultured at 30° C. for 18 hours. The titer of the stock solution before dilution was calculated by counting the number of colonies. The stock solution of the phage solution was suspended in PBS containing 0.05% NaN3 so as to be 2×1014/mL.
5. Obtaining of Antibody Clone Specific to Cancer Cell
5-1 Phage Antibody Screening Using Cancer Cell Line
[0773]Phage antibodies of various cancer cell lines or clinical specimens were isolated by the following procedure. Kinds of used cell lines are described below. The culture conditions of the cell line are show in Table of FIG. 38.
[0774]pancreatic cancer cell lines PANC-1, MIA-Paca2
[0775]kidney cancer cell lines CCFRC1, Caki-1, CCFRC1, Caki-1, ACHN
[0776]ovarian cancer cell lines KF28, RMG-1, RMG-2, SKOv3
[0777]stomach cancer cell lines SNU-5, MKN45, NCI-N87
[0778]lung squamous cell carcinoma lines RERF-LC-AI, EBC1
[0779]pulmonary adenocarcinoma cell lines Calu-3, NCI-H441, A549, PC14
[0780]hepatic cell carcinoma cell lines HepG2, OCTH, Hep3B
[0781]hepatic cell carcinoma clinical specimen (HCV positive),
[0782]intrahepatic bile duct cell carcinoma cell line RBE
[0783]stomach cancer cell lines SNU5, MKN45, NCI-N87
[0784]large bowel cancer cell lines CW2, CaCo2
[0785]acute myelocytic leukemia, AML clinical specimen
[0786]An adherent cell line group in 6 well plate (Falcon 3516) and a suspended cell line such as ATL-derived cell line in suspended culture flask (70 ml (slant neck)), which had been cultured in a medium (RPMI-1640: Sigma-Aldrich, 10% fetal calf serum, 1% penicillin-streptomycin solution) in a CO2 incubator at 37° C., were used.
[0787]The adherent cell line was dissociated from culture dish with 2 mg/ml collagenase I (Gibco BRL)/cell dissociation buffer (Gibco BRL), and then recovered with 10% FBS/DMEM. On the other hand, the suspended cells were, as they were, centrifuged (400×g, 4° C., two minutes) to remove the medium once.
[0788]After such operation, each cell was washed with 1% BSA, 0.05% NaN3/PBS (BSA solution) and centrifuged (400×g, 4° C., two minutes) to remove the supernatant.
[0789]Cells from the clinical specimen derived from clinical tissue material prepared in 6 well plate (Falcon 3516), which had been cultured in a medium (RPMI-1640: Sigma-Aldrich, 10% fetal calf serum, 1% penicillin-streptomycin solution) in a CO2 incubator at 37° C., were used.
[0790]Cells were washed with cooled PBS and 4×107 of cells were used for screening. This was mixed with 1×1013 cfu of human antibody phage library, so that the final concentration of the reaction solution was made to be 1% BSA-0.1% NaN3/MEM and the volume was made to be 1.6 ml. The reaction was carried out while rotating slowly at 4° C. for four hours. After the reaction was completed, the reaction solution was divided into two parts and each part was stratified on 0.6 ml of organic solution (dibutyl phtalate cycloheximide 9:1) and subjected to centrifugation at centrifugal force of 3000 rpm by using a micro-centrifugal machine for two minutes, so that cells were allowed to precipitate at the bottom of the tube. From each tube, the solution was discarded and cells were suspended in 0.7 ml of 1% BSA/MEM, stratified on 0.7 ml of organic solvent and subjected to centrifugation. This operation was repeated again. Then the solution was discarded and cells were suspended in 0.3 ml PBS, frozen with liquid nitrogen and melted at 37° C.
[0791]This was infected with 20 ml of E. coli DH12S (OD 0.5) for one hour, the part of it was plated on an Ampicillin plate and the titer of the collected phage was calculated. Phage infected E. coli was cultured over night in 600 ml of 2×YTGA culture medium (2×YT, 200 μg/ml ampicillin sulfate, 1% glucose) at 30° C. overnight. The cultured product (10 ml) that had been cultured over night was mixed with 200 ml of 2×YTA culture medium (2×YT, 200 μg/ml ampicillin sulfate) and cultured at 37° C. for 1 hour. Thereafter, helper phage κ07 (1×1011) was placed and cultured at 37° C. for 1.5 hour. Then, 800 ml of 2×YTGAK (2×YT, 200 μg/ml ampicillin sulfate, 0.05% glucose, 50 μg/ml kanamycin) was placed and cultured over night at 30° C. This was centrifuged at 8000 rpm for ten minutes so as to prepare 1 l of supernatant. To this, 200 ml of PEG solution (20% polyetyleneglycol 6000, 2.5M NaCl) was mixed and agitated sufficiently. Thereafter, the mixture was centrifuged at 8000 rpm for 10 minutes so as to precipitate phage. This was suspended in 10 ml of PBS/0.05% NaN3 and the part of it was used so as to examine the number of infected E. coli. This is the phase of the I st screening.
[0792]For the 2nd screening, 2×107 of cells and 1×1010 cfu of the 1st screening phages were used, so that the volume of the reaction solution was made to be 0.8 ml. The reaction solution was 1% BSA-0.1% NaN3/MEM and the entire scale was carried out equal to that of the 1st screening.
[0793]The 3rd screening was carried out in the same conditions as those of the 2nd screening except that 1×109 cfu of 2nd phages were used.
[0794]When the recovering rate of the phages is increased, the screening round is stopped at the time. When the recovering rate is not increased, the 4th screening or later are carried out in the same manner by using the phage recovered immediately before round and by using 1×109 cfu of phages.
[0795]The screening of various cell lines was carried out by the same method as that of the screening mentioned above.
5-2 Selection of Antibody Clone
[0796]In the screening of HepG2 as an example, because the recovering rate of HepG2 was increased in the 3rd screening (FIG. 9), it was judged that HepG2 cell specific antibody clone was concentrated in this stage, and several hundreds clones were picked up. Next, when the base sequence of H-chain portions of these positive clones was analyzed, antibodies obtained by removing the overlap from the kinds of base sequences were classified. These were examined for expression. Furthermore, expression positive clones were selected by the following procedures.
6. Base Sequence Determination of Antibody Clone
[0797]E. coli, infected with antibody phage, obtained by screening was diluted and plated on a nutrient agar medium containing 100 Hg/ml of ampicillin. The obtained colonies were picked up and cultured in 2×YTGA culture medium at 30° C. overnight. DNA was extracted by using KURABO PI-50 and the base sequence was determined by a dideoxy method. The overlapped clones having the same base sequence were removed. Furthermore, this culture medium cultured overnight (0.05 ml) was plated on 1.2 ml of 2×YTAI (2×YT, 200 μg/ml ampicillin sulfate, 0.5 mM IPTG) and cultured overnight at 30° C., centrifuged by using a micro-centrifugal machine at 15000 rpm for 5 minutes, and supernatant was obtained.
7. Confirmation of Expression of Antibody Clone
7-1 Selection of Antibody Clone
[0798]Since the antibody was expressed as cp3 fused protein, the expression using the protein was examined. That is to say, firstly, the supernatant obtained in the previous paragraph was reacted in Maxisorp (NUNC) at 37° C. for two hours, liquid was discarded, and blocking was carried out by reacting 5% BSA/PBS/0.05% NaN3 at 37° C. for two hours. The liquid was discarded and a rabbit anti-cp3 antibody (Medical & Biological Laboratories Co., Ltd.) that had been diluted 5000-fold with 0.05% Tween/PBS was reacted at room temperature for one hour, followed by washing with PBS. Then, a HRP labeled goat anti-rabbit IgG antibody (Medical & Biological Laboratories Co., Ltd.) that had been diluted 2000-fold with 0.05% Tween/PBS was reacted at room temperature for one hour, followed by washing with PBS. Then, 100 μl of OPD solution was reacted at room temperature for 2 to 10 minutes, and the reaction was terminated by using 2N sulfuric acid, and by using SPECTRAmax 340PC (Molecular Devices), the absorbance at 492 nm of wavelength was measured.
[0799]In negative well in which the supernatant was not reacted was made to be a control. It was judged that a control whose absorbance did not become two times or more did not express. Such a control was removed from the later analysis.
7-2 Preparation of Antibody Sample
[0800]7-2-1 Production of cp Type Antibody Expression E. coli
[0801]E. coli (10 ml) infected with phage corresponding to expressing antibody clones was introduced was plated on YTGA and shaking cultured at 30° C. one day and one night (pre-culture solution). This was added to 4 l of YT 0.05GA and cultured at 30° C. When O.D. of the bacterial cells became 0.5, 4 ml of 1M IPTG was added and shaking cultured at 30° C. one day and one night. After the culture was terminated, the bacterial cells were centrifuged by using a cooling centrifugal machine at 10000 g, 4° C. for 10 minutes. To the obtained culture supernatant, an equal amount of saturated ammonium sulfate aqueous solution was added and stirred at room temperature for one hour. This solution was centrifuged by using a cooling centrifugal machine at 10000 g, 4° C. for 15 minutes, then supernatant was discarded, the obtained precipitate was suspended in 20 ml of PBS-NaN3 solution, centrifuged by using a cooling centrifugal machine at 10000 g, 4° C. for 5 minutes, and supernatant was recovered. This was dialyzed with PBS one day and one night. To this, a supernatant antibody cp3 mouse monoclonal antibody (Medical & Biological Laboratories Co., Ltd.) that had been balanced with 0.05% NaN3/PBS was chemically immobilized. Antibody affinity column was produced by using sepharose beads. The supernatant was naturally dropped, and the components that had not reacted with beads were allowed to pass through the column. This column was washed with 100 ml of PBS twice, washed with 0.1% Tween 20/PBS (30 ml) four times, and washed with 100 ml of PBS twice. To this, 0.2M Glycine-HCl (pH 3, 4 ml) was slowly added three times and the eluted component was recovered. Then, 3M Tris (80 μl) was added and neutralize (antibody solution). This was filtrated through a MILLEX-GP 0.22 μm filter, O.D. was measured, and the yield of antibodies was calculated.
7-2-2 Production of pp Type Antibody Expressing E. coli
[0802]The obtained antibody clone is originally cp3 type clone. This DNA was extracted by using KURABO PI-50, digested with a restriction enzyme SalI, self reconnected, then, introduced into E. coli DH 12S for transformation. Then, it was plated on a LBGA plate and cultured at 30° C. overnight at. The obtained E. coli colonies were cultured in 2×YTGA overnight and a pp type antibody expressing E. coli solution was obtained.
[0803]E. coli (10 ml) into which a plasmid expressing pp type antibody clones was introduced was plated on YTGA and shaking cultured at 30° C. one day and one night (pre-culture solution). This was added to 4 l of YT 0.05GA and cultured at 30° C. When O.D. of the bacterial cells became 0.5, 4 ml of 1 M IPTG was added and shaking cultured at 30° C. one day and one night. After the culture was terminated, the bacterial cells were centrifuged by using a cooling centrifugal machine at 10000 g, 4° C. for 10 minutes. To the obtained culture supernatant, an equal amount of saturated ammonium sulfate aqueous solution was added and stirred at room temperature for one hour. This solution was centrifuged by using a cooling centrifugal machine at 10000 g, 4° C. for 15 minutes, then supernatant was discarded, the obtained precipitate was suspended in 20 ml of PBS-NaN3 solution, centrifuged by using a cooling centrifugal machine at 10000 g, 4° C. for 5 minutes, and supernatant was recovered. This was dialyzed with PBS one day and one night. To this, 2 ml of IgG sepharose 6 Fast Flow (Amersham Biosciences) balanced with 0.05% NaN3/PBS was added and reacted while shaking at 4° C. one day and one night. This mixture solution was transferred to a column and naturally dropped. The components that were not reacted with beads were allowed to pass through the column. This column was washed with 100 ml of PBS twice, washed with 0.1% Tween 20/PBS (30 ml) four times, and washed with 100 ml of PBS twice. To this, 0.2M Glycine-HCl (pH 3, 4 ml) was slowly added three times and the eluted component was recovered. Then, 3M Tris (80 μl) was added and neutralize (antibody solution). This was filtrated through a MILLEX-GP 0.22 μm filter, O.D. was measured, and the yield of antibodies was calculated.
8. Reactivity to Various Cell Lines of Antibody Clone
8-1 FCM (Flow Cytometry) Analysis
[0804]The reactivity of various isolated antibody clones to various cell lines was confirmed by FCM. Experiment operation was as follows. Firstly, an adherent cell line in 6 well plate (Falcon 3516) and a suspended cell line such as ATL-derived cell line in suspended culture flask (70 ml (slant neck)), which had been cultured in a medium (RPMI-1640: Sigma-Aldrich, 10% fetal calf serum, 1% penicillin-streptomycin solution) in a CO2 incubator at 37° C., were used.
[0805]i) Adherent cell line was dissociated from a culture plate with 2 mg/ml collagenase I (Gibco BRL)/cell dissociation buffer (Gibco BRL), and then recovered with 10% FBS/DMEM. On the other hand, the suspended cells were, as they were, centrifuged (400×g, 4° C., two minutes) to remove the medium once. After such operation, each cell was washed with 2.5% BSA, 0.05% NaN3/PBS (BSA solution), suspended in 100 μl of 2.5% normal goat serum/BSA solution and stood still on ice for 30 minutes, dispensed to 106 cells/well, and then centrifuged (400×g, 4° C., two minutes) to remove the supernatant.
[0806]ii-1) In the case of cp3 antibodies, they were added so that the concentration became 5 μg/ml and left on ice for one hour. This was washed with a BSA solution once, then suspended in 100 μl of 5 μg/ml BSA solution of anti-cp3 mouse monoclonal antibody (Medical & Biological Laboratories Co., Ltd.) and left on ice for one hour. This was washed with a BSA solution once, then suspended in 100 μl of 5 μg/ml BSA solution of Alexa 488 binding anti-mouse IgG goat antibody (Molecularprobe) and left on ice for one hour. This was washed with BSA solution twice, and then suspended in 500 μl of BSA solution. To this solution, 50 μl of fixation solution (formaldehyde) was added and it was left for 10 minutes. Thereafter, 150 μl of PBS was added, treated by using Cell Strainer (Becton Dickinson), and then the fluorescence intensity of the group of cells was analyzed by using FACScaliver (FCM) (Becton Dickinson) ((1) to (3)).
[0807]ii-2) In the case of the pp type (protein A type) antibodies, they were added so that the concentration became 5 μg/ml and left on ice for one hour. This was washed with a BSA solution once, then suspended in 100 μl of 5 μg/ml BSA solution of Alexa 488 binding anti-mouse IgG goat antibody (Molecularprobe) and left on ice for one hour. This was washed with BSA solution twice, and then suspended in 500 μl of BSA solution. To this solution, 50 μl of fixation solution (formaldehyde) was added and it was left for 10 minutes. Thereafter, 150 μl of PBS was added, treated by using Cell Strainer (Becton Dickinson), and then the fluorescence intensity of the group of cells was analyzed by using FACScaliver (FCM) (Becton Dickinson).
[0808]In the analysis, detection antibody was labeled with fluorescent dye (Alexa 488, etc.) in advance. After sample antibodies and cells were reacted, they were reacted with detection antibodies. The difference in the antibody amount occurs depending upon the amount of antigen existing on the surface of the cell, and as a result, the fluorescence intensity became different. Thus, the affinity with respect to the antigen existing on the surface of the cells and the amount of antigen can be estimated. Furthermore, in order to remove dead cells and debris, and the like, Forward Scatter: FSC is expressed in X-axis and Side Scatter: SSC is expressed in Y-axis, and a group of living cells (substantially the same group because cultured cells were used) in data obtained by dot plot expansion were gated, the fluorescence intensity only in this gate was measured.
8-2 Production of Panel
[0809]From the results of FCM, a histogram showing the relationship between the antibody binding amount and the number of cells was formed. One-parameter histogram using the antibody binding amount a parameter was drawn. The one-parameter histogram is one of the display methods in the flow cytometry. The one-parameter histogram is generally shown in a graph in which X-axis represents one indicator (parameter) and Y-axis represents the number of cells.
[0810]Typical examples of the results of FCM are shown in FIGS. 10 to 12. As shown in these figures, basically, the behavior of the FCM becomes unique according to the combination of cells and antibodies. FIGS. 10 and 11 show histogram (right) and cell fluorescence cytology image (left), respectively, which show the reactivity between the scFv antibody and the undifferentiated malignant liver cancer cell line HLF obtained in the above-mentioned method. In all the antibodies (five antibodies), positive patterns are obtained but each has very unique shape of peak. Such shapes of peaks are thought to reflect the uniqueness of epitope of antigen. FIG. 12 shows a plurality of histograms (antibodies to be used was different in each case) which are overwritten. It is shown that the peak of each histogram has its own unique shape. However, during the comprehensive FCM analysis, an antibody group providing histogram having an extremely high similarity as shown in FIGS. 13 to 15 are observed. Furthermore, as shown in FIG. 16, an antibody group consistently providing histogram having a high similarity regardless of cell lines to be used in the FCM analysis was observed. FIG. 16 show comparison of histograms obtained in three kinds of antibodies (035-234 antibody, 040-107 antibody, and 041-118 antibody). According to the later investigation, it is determined that these three kinds of antibodies recognize ALCAM.
[0811]FIG. 17 shows a method for classifying the antibody group based on the results of the FCM analysis. That is to say regardless of kinds of cells to be used, a plurality of antibodies having similar behavior (shape of histogram) in the FCM analysis are shown as the same group in a panel. Basically, a plurality of antibodies having the same shape of histogram (peaks are overlapped when the shapes are overwritten) are defined as one group. However, a plurality of antibodies may be classified into groups on the basis of the factors such as the median value, mode (peak value), and kurtosis of the histogram.
[0812]A plurality of antibodies are classified based on the above-mentioned technique. Firstly, the histograms obtained in the antibody clones are overwritten for each cell line to be used, and thereby the histograms are compared with each other. Thus, similarly between the antibody clones and the reactivity between antibody clones are obtained. Then, based on the similarity and the reactivity, antibody clones are classified and summarized in table (FIG. 18). Thus, eight antibody groups (in the description hereinafter, groups are named 1, 2, 3, 4, 5, 6, 7, and 8 sequentially in this order) are obtained. In FIG. 18, information on antigen identified later is also displayed. Each mark in Table shows a shift amount relative from the histogram (reference histogram) of the negative control antibody. Double circle mark represents that the shift amount is 20 times or more (the peak value of the is 20 times or more of the reference histogram); "o" (circle mark) represents that the shift amount is 10 times or more; "Δ" (triangle mark) represents that the shift amount is 3 times or more; and "x" represents that the shift amount is less than 3, respectively (an oblique line means no data is obtained). The larger the shift amount is, the higher the reactivity is.
[0813]Next, by the following procedure, it is verified that antigens of each antibody group in the produced panel are common.
9. Identification of Protein (Antigen) Recognized by Antibody Clone
9-1 Preparation of Solid Phase Antibody for Immunoprecipitation
[0814]Firstly, a pp type antibody solution was dialyzed with a coupling buffer solution (0.1M NaHCO3--NaOH, pH 9). That is to say, an antibody solution was enclosed with a dialysis membrane (Snake Skin Pleated Dialysis Tubing 10,000 MWCO) and this was allowed to be sunk in 1.5 L of the coupling buffer solution (0.1 M NaHCO3--NaOH, pH 9) and stirred by using a stirrer at 4° C. for two to three hours. Then, the buffer solution was replaced with new one and dialyzed for two to three hours. Thereafter, the buffer solution was replaced with new one again and dialyzed one day and one night.
[0815]Next, activated CNBr-activated Sepharose 4B used for making solid phase was adjusted. That is to say, CNBr-activated Sepharose 4B (Amersham Biosciences) was swollen with 1 mM HCl, then sucked by using an aspirator. To this, 50 ml of coupling buffer solution was added, stirred, and then sucked by using an aspirator. In this sucked state, a coupling buffer solution was added.
[0816]An antibody was made to be solid phased as follows. That is to say, to 5 mg antibody solution (10 ml), activated gel (1 ml) was added to cause a reaction at room temperature for two hours. After the reaction was terminated, the gel was transferred to a column and washed with a coupling buffer solution (1 ml) ten times. The presence of non-reacted antibodies was confirmed by measuring the O.D. The solid phased gel was substituted by 0.2M Glycine-NaOH pH8 solution (5 ml) twice, the same solution (5 ml) was further added and left at room temperature for two hours, this solution was naturally dropped, to this, 0.2M Glycine-HCl (pH 3, 5 ml) was added and substituted, the same solution (5 ml) was further added and left for 5 minutes, and then naturally dropped. Finally, the column was substituted by 20 ml of PBS, then naturally dropped, and 1% NP40, protease inhibitor, and 0.05% NaN3/PBS were added, and gel was recovered.
9-2 Biotin Label of Protein on Cell Membrane and Production of Cell Lysate
[0817]Biotin labeling of the cultured liver cancer cell line was carried out as follows. That is to say, cultured cells HLF that had been cultured in five 15 cm-dishes were washed with PBS twice, and collagenase I (GIBCO) whose concentration had been adjusted to 5 mg/ml by using a cell dissociation buffer (GIBCO) was added and reacted in a CO2 incubator at 37° C., so that cells were liberated. Thereafter, cells were recovered in a culture medium and washed with PBS(-) twice. Then, the number of cells was counted by using a hemocytometer. The cells were suspended in PBS(-) so that the counted number became about 5×107/ml. To this, an equal amount of EZ-Link Sulfo-NHS-LC-Biotinylation Kit (PIERCE) was added so that the concentration had been adjusted to 1 mg/ml with PBS, left at room temperature for 30 minutes and then washed with PBS twice.
[0818]The cell lysate of biotin labeled cells was adjusted as follows. That is to say, to the above-mentioned biotin labeled cells, 4 ml of lysis buffer (1% NP40/detergent base solution, the composition of the detergent base solution: 20 mM HEPES, pH 8.0, 140 mM NaCl, protease inhibitor) was added and cells were suspended. This suspension was placed and homogenized in a cooled Dounce homogenizer. To the solution, 1/2 amount (2 ml) of a detergent mix solution (1% NP40, tritonX-100, b-D-Maltoside, n-Octyl b-D-Glucoside, n-Octyl b-D-Maltoside, n-Decyl b-D-Maltoside, deoxycholic acid, each 0.5%/detergent base solution) was added and mixed at 4° C. for four hours. This solution was centrifuged at 100,000 rpm for 30 minutes and filtrated through MILLEX-GP 0.22 μm filter.
9-3 Immunoprecipitation Reaction
[0819]Firstly, about 60 μl parts (about 150 μl solution parts) of the solid-phased antibodies (hereinafter, referred to as "antibody beads") were placed in a 2 ml-tube and 1/10 volume (about 15 μl) of 4 mM biotin was added to the tube. A mixture of 0.5 culture dishs of lysate (600 μl) and 60 μl of biotin solution was added to the tube and reacted while stirring at 4° C. for several hours. Then, the tube was centrifuged (5500 g, one minute, 4° C.) and supernatant was removed. To this, 800 μl of washing biotin/lysis-T buffer (0.5 mM biotin, 0.1% Tween 20/PBS) was added and mixed while falling two or three times, then the tube was centrifuged (5500 g, one minute, 4° C.), and supernatant was removed. This washing operation was carried out again, then 30 μl of citric acid solution (50 mM citric acid, pH 2.5) for elution was added to the antibody beads and stirred. Then, the tube was centrifuged (5500 g, 1 min, 4° C.) and supernatant was recovered. To the remaining antibody beads, 30 μl of citric acid solution for elution was added and stirred. The tube was centrifuged (5500 g, 1 min, 4° C.) and supernatant was recovered. This elution operation was repeated further three times, and a sample solution was recovered and 3M Tris was added to the solution for neutralization. This sample was migrated by SDS-PAGE so as to confirm the band by silver staining. At the same time, this sample was subjected to western blotting by using streptavidin-HRP (Anti-Streptavidin, IgG Fraction, Conjugated to Peroxidase CORTEX biochem) so as to detect a band of the biotin membrane protein.
9-4 Mass Spectrometry of Cut-Out Band
9-4-1 Trypsin Digestion in Gel
[0820]A portion corresponding to detected membrane protein was digested with trypsin in a gel and peptide was recovered. SDS polyacrylamide gel electrophoresis was carried out in accordance with a usual method and a band that had been obtained by staining with Coomassie Brilliant Blue was cut out. This was soaked in 200 mM ammonium bicarbonate 50% acetonitrile solution, shaken at 37° C. for 45 minutes. Then, the solution was discarded and the operation was repeated twice, thereby removing the Coomassie Brilliant Blue. This gel was dried under reduced pressure, and 4 μl of trypsin (20 μg/ml) dissolved in 40 mM ammonium bicarbonate (pH 8.1)-10% acetonitrile was added per unit area (mm2) of gel slice, and left at room temperature for one hour and sufficiently infiltrated. To this, a trypsin solution was added in an amount that was 25 times as much as the previously added amount, and left at 37° C. for 18 hours. This was filtrated by a tube having a filter whose power size was 0.22 μm, and peptide in which an antigen had been cut with trypsin was recovered.
9-4-2 Identification of Antigen by Mass Spectrometry
[0821]A specimen obtained by in-gel trypsin digestion was subjected to HPLC linked with an electrospray ionization type ion trap quadrupole mass spectrometer. From the reversed phase chromatography column of HPLC, according to the change of linear concentration gradient of 0% to 80% acetonitrile containing 0.1% TFA, each peptide that had been eluted sequentially depending upon the hydrophobic property was ionized by an electrospray method. The mass of each peptide was analyzed.
[0822]At the same time, the mass of limited digested product of each peptide generated by collision with helium atoms placed in the middle of the fight route of ions was analyzed. When one amino acid is removed by limited digestion, since ion that is smaller by a part of the mass of the removed amino acid is observed, the kind of the removed amino acid can be identified according to the difference in mass. Furthermore, another amino acid is removed, since ion that is smaller by a part of the mass of the removed amino acid is observed, the kind of the removed amino acid can be identified according to the difference in mass. By proceeding the same analysis of the experimental data, an inner amino acid sequence can be determined. The set of the inner sequence of the obtained amino acid was retrieved by using a published amino acid sequence database and antigen was identified. As a result, as shown below, antigen of each antibody clone was identified and it is confirmed that the antibodies in the same group have the common antigen. The identification results was confirmed because the total amount of the identified protein that had been analogized from the amino acid sequence was not contradictory to the experimental data of the molecular weight of the SDS polyacrylamide electrophoresis of antigen before carrying out the trypsin digestion.
[0823]Antigen of antibodies belonging to group 1: HER1 (also known as: ErbB1, c-erbB-1, EGFR (Epidermal Growth Factor Receptor), v-erbB)
[0824]Antigen of antibodies belonging to group 2: HER-2 (also known as: ErbB2, c-erbB-2, neu)
[0825]Antigen of antibodies belonging to group 3: CD46 antigen (also known as: MCP (membrane cofactor protein), gp45-70, HuLY-m5, measles virus receptor, MIC10, TLX-B antigen, TRA2, trophoblast leucocyte common antigen, trophoblast-lymphocyte cross-reactive antigen)
[0826]Antigen of antibodies belonging to group 4: ITGA3 (integrin alpha3) (also known as: alpha3beta1 Epiligrin Receptor, alpha3beta1 Integrin, Epiligrin Receptor, CD49c, VLA-3, Gap b3, Galactoprotein b3, Laminin-5 Receptor)
[0827]Antigen of antibodies belonging to group 5: ICAM1 (Intercellular adhesion molecule-1) (also known as: Intercellular Adhesion Molecule 1, CD54 Antigen)
[0828]Antigen of antibodies belonging to group 6: ALCAM (Activated leukocyte cell adhesion molecule) (also known as: KG-CAM, CD166 Antigen, CD6 Ligand,
Activated Leukocyte Cell Adhesion Molecule, Neurolin)
[0829]Antigen of antibodies belonging to group 7: CD147 antigen (also known as: BSG, TCSF (Tumor cell-derived collagenase stimulatory factor), 5F7 protein, OK blood group protein, basigin protein, collagenase stimulatory factor protein, EMMPRIN (Extracellular matrix metalloproteinase Inducer), M6 activation antigen, human leukocyte activation antigen M6)
[0830]Antigen of antibodies belonging to group 8: IgSF4 (also known as: BL2, ST17, NECL2, TSLC1, IGSF4A, SYNCAM, sTSLC-1)
[0831]From the above-mentioned identification results, it has been clarified that it was possible obtain three antibody clones to HER1 (048-006 antibody, 057-091 antibody, and 059-152 antibody), one antibody clone to HER-2 (015-126 antibody), seven antibody clones to CD46 antigen (035-224 antibody, 045-011 antibody, 051-144 antibody, 052-053 antibody, 052-073 antibody, 053-049 antibody, and 3172-120 antibody), one antibody clone to ITGA3 (015-003 antibody), five antibody clones to ICAM1 (052-033 antibody, 053-042 antibody, 053-051 antibody, 053-059 antibody, and 053-085 antibody), five antibody clones to ALCAM (035-234 antibody, 040-107 antibody, 041-118 antibody, 066-174 antibody, and 083-040 antibody), one antibody clone to CD147 antigen (059-053 antibody), and ten antibody clones to IgSF4. The Note here that the amino acid sequences of the antibody clones have been identified as mentioned below (antibody clones to IgSF4 are omitted).
TABLE-US-00023 <Antibodies belonging to group 1> (1) 048-006 antibody SEQ ID NO: 1 (VH), SEQ ID NO: 2 (VH CDR1), SEQ ID NO: 3 (VH CDR2), SEQ ID NO: 4 (VH CDR3), SEQ ID NO: 5 (VL), SEQ ID NO: 6 (VL CDR1), SEQ ID NO: 7 (VL CDR2), SEQ ID NO: 8 (VL CDR3) (2) 057-091 antibody SEQ ID NO: 9 (VH), SEQ ID NO: 10 (VH CDR1), SEQ ID NO: 11 (VH CDR2), SEQ ID NO: 12 (VH CDR3), SEQ ID NO: 13 (VL), SEQ ID NO: 14 (VL CDR1), SEQ ID NO: 15 (VL CDR2), SEQ ID NO: 16 (VL CDR3) (3) 059-152 antibody SEQ ID NO: 17 (VH), SEQ ID NO: 18 (VH CDR1), SEQ ID NO: 19 (VH CDR2), SEQ ID NO: 20 (VH CDR3), SEQ ID NO: 21 (VL), SEQ ID NO: 22 (VL CDR1), SEQ ID NO: 23 (VL CDR2), SEQ ID NO: 24 (VL CDR3) <Antibody belonging to group 2> (1) 015-126 antibody SEQ ID NO: 25 (VH), SEQ ID NO: 26 (VH CDR1), SEQ ID NO: 27 (VH CDR2), SEQ ID NO: 28 (VH CDR3), SEQ ID NO: 29 (VL), SEQ ID NO: 30 (VL CDR1), SEQ ID NO: 31 (VL CDR2), SEQ ID NO: 32 (VL CDR3) <Antibodies belonging to group 3> (1) 035-224 antibody SEQ ID NO: 33 (VH), SEQ ID NO: 34 (VH CDR1), SEQ ID NO: 35 (VH CDR2), SEQ ID NO: 36 (VH CDR3), SEQ ID NO: 37 (VL), SEQ ID NO: 38 (VL CDR1), SEQ ID NO: 39 (VL CDR2), SEQ ID NO: 40 (VL CDR3) (2) 045-011 antibody SEQ ID NO: 41 (VH), SEQ ID NO: 42 (VH CDR1), SEQ ID NO: 43 (VH CDR2), SEQ ID NO: 44 (VH CDR3), SEQ ID NO: 45 (VL), SEQ ID NO: 46 (VL CDR1), SEQ ID NO: 47 (VL CDR2), SEQ ID NO: 48 (VL CDR3) (3) 051-144 antibody SEQ ID NO: 49 (VH), SEQ ID NO: 50 (VH CDR1), SEQ ID NO: 51 (VH CDR2), SEQ ID NO: 52 (VH CDR3), SEQ ID NO: 53 (VL), SEQ ID NO: 54 (VL CDR1), SEQ ID NO: 55 (VL CDR2), SEQ ID NO: 56 (VL CDR3) (4) 052-053 antibody SEQ ID NO: 57 (VH), SEQ ID NO: 58 (VH CDR1), SEQ ID NO: 59 (VH CDR2), SEQ ID NO: 60 (VH CDR3), SEQ ID NO: 61 (VL), SEQ ID NO: 62 (VL CDR1), SEQ ID NO: 63 (VL CDR2), SEQ ID NO: 64 (VL CDR3) (5) 052-073 antibody SEQ ID NO: 65 (VH), SEQ ID NO: 66 (VH CDR1), SEQ ID NO: 67 (VH CDR2), SEQ ID NO: 68 (VH CDR3), SEQ ID NO: 69 (VL), SEQ ID NO: 70 (VL CDR1), SEQ ID NO: 71 (VL CDR2), SEQ ID NO: 72 (VL CDR3) (6) 053-049 antibody SEQ ID NO: 73 (VH), SEQ ID NO: 74 (VH CDR1), SEQ ID NO: 75 (VH CDR2), SEQ ID NO: 76 (VH CDR3), SEQ ID NO: 77 (VL), SEQ ID NO: 78 (VL CDR1), SEQ ID NO: 79 (VL CDR2), SEQ ID NO: 80 (VL CDR3) (7) 3172-120 antibody SEQ ID NO: 81 (VH), SEQ ID NO: 82 (VH CDR1), SEQ ID NO: 83 (VH CDR2), SEQ ID NO: 84 (VH CDR3), SEQ ID NO: 85 (VL), SEQ ID NO: 86 (VL CDR1), SEQ ID NO: 87 (VL CDR2), SEQ ID NO: 88 (VL CDR3) <Antibody belonging to group 4> (1) 015-003 antibody SEQ ID NO: 89 (VH), SEQ ID NO: 90 (VH CDR1), SEQ ID NO: 91 (VH CDR2), SEQ ID NO: 92 (VH CDR3), SEQ ID NO: 93 (VL), SEQ ID NO: 94 (VL CDR1), SEQ ID NO: 95 (VL CDR2), SEQ ID NO: 96 (VL CDR3) <<Antibodies belonging to group 5> (1) 052-033 antibody SEQ ID NO: 97 (VH), SEQ ID NO: 98 (VH CDR1), SEQ ID NO: 99 (VH CDR2), SEQ ID NO: 100 (VH CDR3), SEQ ID NO: 101 (VL), SEQ ID NO: 102 (VL CDR1), SEQ ID NO: 103 (VL CDR2), SEQ ID NO: 104 (VL CDR3) (2) 053-042 antibody SEQ ID NO: 105 (VH), SEQ ID NO: 106 (VH CDR1), SEQ ID NO: 107 (VH CDR2), SEQ ID NO: 108 (VH CDR3), SEQ ID NO: 109 (VL), SEQ ID NO: 110 (VL CDR1), SEQ ID NO: 111 (VL CDR2), SEQ ID NO: 112 (VL CDR3) (3) 053-051 antibody SEQ ID NO: 113 (VH), SEQ ID NO: 114 (VH CDR1), SEQ ID NO: 115 (VH CDR2),
SEQ ID NO: 116 (VH CDR3), SEQ ID NO: 117 (VL), SEQ ID NO: 118 (VL CDR1), SEQ ID NO: 119 (VL CDR2), SEQ ID NO: 120 (VL CDR3) (4) 053-059 antibody SEQ ID NO: 121 (VH), SEQ ID NO: 122 (VH CDR1), SEQ ID NO: 123 (VH CDR2), SEQ ID NO: 124 (VH CDR3), SEQ ID NO: 125 (VL), SEQ ID NO: 126 (VL CDR1), SEQ ID NO: 127 (VL CDR2), SEQ ID NO: 128 (VL CDR3) (5) 053-085 antibody SEQ ID NO: 129 (VH), SEQ ID NO: 130 (VH CDR1), SEQ ID NO: 131 (VH CDR2), SEQ ID NO: 132 (VH CDR3), SEQ ID NO: 133 (VL), SEQ ID NO: 134 (VL CDR1), SEQ ID NO: 135 (VL CDR2), SEQ ID NO: 136 (VL CDR3) <Antibodies belonging to group 6> (1) 035-234 antibody SEQ ID NO: 137 (VH), SEQ ID NO: 138 (VH CDR1), SEQ ID NO: 139 (VH CDR2), SEQ ID NO: 140 (VH CDR3), SEQ ID NO: 141 (VL), SEQ ID NO: 142 (VL CDR1), SEQ ID NO: 143 (VL CDR2), SEQ ID NO: 144 (VL CDR3) (2) 040-107 antibody SEQ ID NO: 145 (VH), SEQ ID NO: 146 (VH CDR1), SEQ ID NO: 147 (VH CDR2), SEQ ID NO: 148 (VH CDR3), SEQ ID NO: 149 (VL), SEQ ID NO: 150 (VL CDR1), SEQ ID NO: 151 (VL CDR2), SEQ ID NO: 152 (VL CDR3) (3) 041-118 antibody SEQ ID NO: 153 (VH), SEQ ID NO: 154 (VH CDR1), SEQ ID NO: 155 (VH CDR2), SEQ ID NO: 156 (VH CDR3), SEQ ID NO: 157 (VL), SEQ ID NO: 158 (VL CDR1), SEQ ID NO: 159 (VL CDR2), SEQ ID NO: 160 (VLCDR3) (4) 066-174 antibody SEQ ID NO: 161 (VH), SEQ ID NO: 162 (VH CDR1), SEQ ID NO: 163 (VH CDR2), SEQ ID NO: 164 (VH CDR3), SEQ ID NO: 165 (VL), SEQ ID NO: 166 (VL CDR1), SEQ ID NO: 167 (VL CDR2), SEQ ID NO: 168 (VLCDR3) (5) 083-040 antibody SEQ ID NO: 169 (VH), SEQ ID NO: 170 (VH CDR1), SEQ ID NO: 171 (VH CDR2), SEQ ID NO: 172 (VH CDR3), SEQ ID NO: 173 (VL), SEQ ID NO: 174 (VL CDR1), SEQ ID NO: 175 (VL CDR2), SEQ ID NO: 176 (VL CDR3) <<Antibody belonging to group 7> (1) 059-053 antibody SEQ ID NO: 177 (VH), SEQ ID NO: 178 (VH CDR1), SEQ ID NO: 179 (VH CDR2), SEQ ID NO: 180 (VH CDR3), SEQ ID NO: 181 (VL), SEQ ID NO: 182 (VL CDR1), SEQ ID NO: 183 (VL CDR2), SEQ ID NO: 184 (VL CDR3)
10. Confirmation of Antigen by RNAi and Immunostaining
[0832]In order to reconfirm that the isolated antibodies recognize the identified antigen, double stranded oligo RNA was allowed to act on cells so as to carry out antigen gene knockdown. Thus, the immunostaining property of the antibody identified by the isolated antigen with respect to the cell was examined.
[0833]Firstly, cells were cultured in a 6-well culture dish to about 30% confluent. To this, a mixture including Lipofectamin 2000 (5 μl) (Invitrogen) and the following oligo RNA (100 pmol) was acted. At day 2, cells were peeled off by using collagenase and recovered. To this, cp3 type purified antibody for verification was acted at the concentration of 5 μg/ml. After washing, a rabbit anti-cp3 antibody was acted at the concentration of 2 μg/ml. After washing, Alexa488 labeled anti-rabbit IgG was acted at 2 μg/ml. This was washed and then immobilized in OptiLyse (NOTECH) (50 μl) for ten minutes. This was diluted by adding 1 ml of PBS and this was measured by using FACS Caliver (Beckmann). As the antibody reaction solution and washing solution, 2.5% BSA/PBS solution was used.
[0834]Subject antigen: CD147
[0835]Sequence of the used oligo RNA:
TABLE-US-00024 CAGAGCUACACAUUGAGAACCUGAA (SEQ ID NO: 441)
[0836]Subject cell: clear cell renal cell carcinoma CCFRC1 cell
[0837]Verified antibody: 059-053 cp3 antibody
[0838]Subject antigen: CD166
[0839]Sequence of the used oligo RNA:
TABLE-US-00025 UACCUAUGUGCAGAGGAAUUAUGAU (SEQ ID NO: 442)
[0840]Subject cell: clear cell renal cell carcinoma CCFRC1 cell
[0841]Verified antibody: 035-234 cp3 antibody
[0842]Subject antigen: CD166
[0843]Sequence of the used oligo RNA:
TABLE-US-00026 GCAACCAUCUAAACCUGAAAUUGUA (SEQ ID NO: 443)
[0844]Subject cell: hepatic cell carcinoma HLF cell
[0845]Verified antibody: 048-006 cp3 antibody
[0846]Subject antigen: HER2
[0847]Sequence of the used oligo RNA:
TABLE-US-00027 UAAUAGAGGUUGUCGAAGGCUGGGC (SEQ ID NO: 444)
[0848]Subject cell: ovarian cancer SKOv-3 cell
[0849]Verified antibody: 015-126 cp3 antibody
[0850]Subject antigen: IgSF4
[0851]Sequence of the used oligo RNA:
TABLE-US-00028 CCCAACAGGCAGACCAUUUAUUUCA (SEQ ID NO: 445)
[0852]Subject cell: hepatic cell carcinoma HLF cell
[0853]Verified antibody: 035-273 cp3 antibody Results are shown in FIGS. 19 to 23. FIG. 19 shows the results of RNAi in which CD147 is a subject antigen. FIG. 20 shows the results of RNAi in which CD166 is a subject antigen. FIG. 21 shows the results of RNAi in which HER1 is a subject antigen. FIG. 22 shows the results of RNAi in which HER2 is a subject antigen. FIG. 23 shows the results of RNAi in which IgSF4 is a subject antigen. As is apparent from these results, in any of the verified antibodies, in the cell population that had been subjected to RNAi, as compared with the cell population that had not been subjected to RNAi, the staining property by antibodies (i.e., reactivity) was significantly reduced. In this way, by RNAi experiment using oligo RNA for knocking down the corresponding antigen it is reconfirmed again that each of the isolated antibodies recognizes the identified antigen.
11. Investigation of Reactivity of Each Antibody by Cell Staining and Tissue Staining
11-1 Experiment Method
(1) Cell Staining
[0854]Cells were dissociated from a culture dish by using 2 mg/ml collagenase I (Gibco BRL)/cell dissociation buffer (Gibco BRL), then collected by using 10% FBS/DMEM, and 1×105 of the cells were used. These were washed with 2.5% BSA, 0.05% NaN3/PBS (BSA solution), then suspended in 100 μl of 2.5% normal goat serum/BSA solution and left on ice for 30 minutes. Thereafter, cp3 type antibodies were added so that the concentration was 5 Hg/ml and left on ice for one hour. This was washed with a BSA solution once, then suspended in 100 μl of 5 μg/ml BSA solution of anti-cp3 mouse monoclonal antibody (Medical & Biological Laboratories Co., Ltd.) and left on ice for one hour. This was washed with a BSA solution once, then suspended in 1001i of 5 μg/ml BSA solution of ALEXA488 binding anti-mouse IgG goat antibody (Molecularprobe), and left on ice for one hour. This was washed with BSA solution twice, and then supernatant was discarded. To this, 501l of OptiLyse B (BECKMAN COULTER) was added and left at room temperature for ten minutes so as to fix the cells. To this, 950 μl of 1 ng DAPI/BSA solution was added, left at room temperature for 10 minutes, and subjected to centrifugation for collecting cells. The cells were mounted on MULTITEST SLIDE (ICN) and observed under microscopy.
(2) Tissue Staining
(2-1) Preparation of Antibody Sample
[0855]E. coli solution cultured overnight (0.5 ml) was planted in 6 ml of 2×YTAI (2×YT, 200 μg/ml ampicillin sulfate, 0.5 mM IPTG), cultured overnight at 30° C. and centrifuged at 10000 rpm for 5 minutes by using a micro-centrifugal machine, and supernatant was recovered. To this, an equal amount of saturated ammonium sulfate was added and left at room temperature for 30 minutes. Then, it was centrifuged at room temperature at 10000 rpm for 5 minutes and supernatant was discarded. The obtained precipitates were suspended in 0.6 ml of PBS-0.05% NaN3, complete solution and centrifuged at 4° C. at 15000 rpm for 5 minutes, and supernatant was recovered.
(2) Production of Section
[0856]The extracted tissue was cut into about 5 mm×5 mm×10 mm, placed in 4% PFA/0.01% glutaraldehyde/0.1 Mcacodylic acid buffer (4° C.) (PFA is a product by Wako Pure Chemical Institute, glutaraldehyde is a product by KANTO CHEMICAL CO., INC., sodium cacodylate is a product by SIGMA). By using a microwave oven (SHARP), it was microwave-fixed. Then, it was fixed again in this fixation solution at 4° C. for one hour. Then, it was transferred into 10% sucrose/PBS and immersed therein at 4° C. for four hours, then substituted by 15% sucrose/PBS and immersed therein at 4° C. for four hours, and then substituted by 20% sucrose/PBS and immersed at one night. It was embedded in an OTC compound and rapidly frozen in dry ice/hexane. This was thinly cut into 4 μm thickness by using cryostat (Reichert-Jung 2800 FRIGCUT E), attached to silane coated slide glass (MATSUNAMI) and dried by using a cold wind drier for 30 minutes.
(2-3) Staining
[0857]The slide glass to which a section was attached was immersed in PBS three times for five minutes each so as to make hydrophilic. Next, 50 μl of 0.3% H2O2/0.1% NaN3 was dropped so as to cause a reaction at room temperature for ten minutes and blocking of endogenous peroxidase was carried out. Then, it was washed with PBS three times for five minutes each. Then, it was reacted in 2% BSA/PBS at room temperature for 10 minutes, and blocking of a non-specific reaction was carried out. Then, excess liquid was dropped off and 50 μl of antibody sample was dropped thereto so as to cause a reaction at room temperature for one hour, followed by washing with PBS three times for 5 minutes each. Next, 501l of anti-CP3 rabbit antibody (5 μg/ml) was dropped to cause a secondary antibody reaction at room temperature for 45 minutes, followed by washing with PBS three times for 5 minutes each. Then, 50 μl of peroxidase labeled dextran binding anti-rabbit immunoglobulin-goat polyclonal antibody (DAKO) was dropped so as to cause a tertiary antibody reaction. This was washed with PBS three times for 5 minutes each, and the 50 μl of DAB-H2O coloring solution was dropped. After the color became brown, this was transferred to a vat filled with distilled water so as to terminate the reaction. Thereafter, obtained product was washed with water for 10 minutes, followed by staining nuclear with hematoxylin. Thereafter, dehydration and penetration were carried out, encapsulation with marinol and observation under microscopy were carried out.
11-2 Experiment Results
(1) Anti-HER1 Antibody Group (Group 1)
[0858]Cancers showing positive in the cell line staining (containing FACS): [0859]pancreatic cancer cell line PANC-1, kidney cancer cell line CCFRC1, kidney cancer cell line Caki-1, ovarian cancer cell line KF28, stomach cancer cell line SNU-5, lung squamous cell carcinoma line RERF-LC-AI, ovarian cancer cell line RMG-1, undifferentiated hepatic cell carcinoma cell line HLF, ovarian cancer cell line SKOv3, pulmonary adenocarcinoma cell line PCI4, kidney cancer cell line ACHN, lung squamous cell carcinoma line EBC1, vulva mucosal epithelial cell line A431, pulmonary adenocarcinoma cell line H1373, hepatic cell carcinoma cell line HepG2, cell line established from kidney clinical specimen
[0860]Cancers showing negative in the cell line staining (containing FACS): [0861]breast cancer cell line BT474, hamster ovarian cancer cell line CHO
[0862]Cancers showing positive in the tissue staining: [0863]kidney cancer, hepatic cell carcinoma, intrahepatic bile duct cancer, lung squamous cell cancer, pulmonary adenocarcinoma, pancreas cancer
(2) Anti-HER2 Antibody Group (Group 2)
[0864]Cancers showing positive in the cell line staining (containing FACS): [0865]pulmonary adenocarcinoma cell line Calu-3, ovarian cancer cell line SKOv3, breast cancer cell line BT474
[0866]Cancers showing negative in the cell line staining (containing FACS): [0867]hepatic cell carcinoma cell line. HLF, pulmonary adenocarcinoma cell line PC14, kidney cancer cell line ACHN, kidney cancer cell line 293T, hamster ovarian cancer cell line CHO, kidney cancer cell line Caki-1, kidney and stomach cancer cell line CCFRC1, cell line established from kidney clinical specimen
(3) Anti-CD46 Antibody Group
[0868]Cancers showing positive in the cell line staining (containing FACS): [0869]large bowel cancer cell line CaCo2, stomach cancer cell line MKN45, undifferentiated hepatic cell carcinoma cell line HLF, liver cancer cell line HepG2, intrahepatic bile duct cell cancer cell line RBE, pancreas cancer cell line PANC1, kidney cancer cell line CCFRC1, kidney cancer cell line Caki-1, pulmonary adenocarcinoma cell line NCI-H441, lung squamous cell cancer EBC1, stomach cancer cell line NCI-N87, stomach cancer cell line SNU-5, lung squamous cell carcinoma line RERF-LC-AI, hepatic cell carcinoma clinical specimen, breast cancer cell line BT474, kidney cancer cell line 293T, pulmonary adenocarcinoma cell line PC14, kidney cancer cell line ACHN, pulmonary adenocarcinoma cell line H1373
[0870]Cancers showing negative in the cell line staining (containing FACS): [0871]hamster ovarian cancer cell line CHO, vulva mucosal epithelial cell line A431
[0872]Cancers showing positive in the tissue staining: [0873]kidney cancer, hepatic cell carcinoma, intrahepatic bile duct cancer, pulmonary adenocarcinoma, pancreas cancer
[0874]Specific expression of CD46 in intrahepatic bile duct cancer and pancreas cancer, which had not been particularly reported about the relationship with respect to CD46 was recognized.
(4) Anti-ITGA3 Antibody Group (Group 4)
[0875]Cancers showing positive in the cell line staining (containing FACS): [0876]undifferentiated hepatic cell carcinoma cell line HLF, ovarian cancer cell line SKOv3, kidney cancer cell line ACHN, kidney cancer cell line Caki-1, pulmonary adenocarcinoma cell line H1373, lung squamous cell cancer EBC1, vulva mucosal epithelial cell line A431, breast cancer cell line BT474, pulmonary adenocarcinoma cell line PCI4, kidney cancer cell line CCFRC1, hepatic cell carcinoma cell line OCTH, intrahepatic bile duct cell cancer cell line RBE, pancreas cancer cell line PANC-1, pancreas cancer cell line MIA-Paca2, pulmonary adenocarcinoma cell line A549, pulmonary adenocarcinoma cell line NCI-N441, pulmonary adenocarcinoma cell line Calu-3, lung squamous cell carcinoma line RERF-LC-AI, stomach cancer cell line SNU5, stomach cancer cell line MKN45, stomach cancer cell line NCI-N87, large bowel cancer cell line CW2, ovarian cancer cell line SKOv3, ovarian cancer cell line KF-28, ovarian cancer cell line RMG-1, ovarian cancer cell line RMG-2
[0877]Cancers showing negative in the cell line staining (containing FACS): [0878]kidney cancer cell line 293T, hepatic cell carcinoma cell line HepG2, hamster ovarian cancer cell line CHO
[0879]Cancers showing positive in the tissue staining: [0880]intrahepatic bile duct cancer, pancreas cancer
[0881]Specific expression of ITGA3 in gallbladder and liver cancer and pancreas cancer, which had not been particularly reported about the relationship with respect to ITGA3 was recognized.
(5) Anti-ICAM I Antibody Group (Group 5)
[0882]Cancers showing positive in the cell line staining (containing FACS): [0883]Liver cancer cell line HepG2, pulmonary adenocarcinoma cell line PC14, cell line established from kidney clinical specimen
[0884]Cancers showing negative in the cell line staining (containing FACS): [0885]undifferentiated hepatic cell carcinoma cell line HLF, ovarian cancer cell line SKOv3, breast cancer cell line BT474, kidney cancer cell line 293T, kidney cancer cell line ACHN, kidney cancer cell line Caki-1, pulmonary adenocarcinoma cell line PC 14, kidney cancer cell line CCFRC1, hamster ovarian cancer cell line CHO
[0886]Cancers showing positive in the tissue staining: [0887]hepatic cell carcinoma
(6) Anti-ALCAM Antibody Group (Group 6)
[0888]Cancers showing positive in the cell line staining (containing FACS): [0889]Liver cancer cell line HepG2, OCTH, Hep3B, and HLF, kidney cancer cell line Caki-1, CCFRC1, ACHN, 293T, and cell line established from clinical specimen, lung cancer cell line PC14, NCI-H441, EBC-1, RERF-LC-AI, A549, and H1373, ovarian cancer cell line SKOv3, KF-28, RMG1, and RMG2, stomach cancer cell line NCI-N87, large bowel cancer cell line CW2, breast cancer cell line BT474, acute myelocytic leukemia AMLclinical specimen, hamster ovarian cancer cell line CHO
[0890]Cancers showing negative in the cell line staining (containing FACS): [0891]vulva mucosal epithelial cell line A431
[0892]Cancers showing positive in the tissue staining: [0893]kidney cancer, hepatic cell carcinoma, intrahepatic bile duct cancer, lung squamous cell cancer, alveolar cell carcinoma, adenocarcinoma [0894]Specific expression of ALCAM in kidney cancer, hepatic cell carcinoma, and gallbladder and liver cancer, which had not been particularly reported about the relationship with respect to ALCAM was recognized.
(7) Anti-CD147 Antibody Group (Group 7)
[0895]Cancers showing positive in the cell line staining (containing FACS): [0896]liver cancer cell line HepG2, kidney cancer cell line CCFRC1, kidney cancer cell line ACHN, kidney cancer cell line Caki-1, pulmonary adenocarcinoma PCI4, cell line established from kidney cancer clinical specimen
[0897]Cancers showing negative in the cell line staining (containing FACS): [0898]hamster ovarian cancer cell line CHO
[0899]Cancers showing positive in the tissue staining: [0900]kidney cancer [0901]Specific expression of CD147 in kidney cancer, which had not been particularly reported about the relationship with respect to CD147 was recognized.
12. Conversion into IgG Type Antibody
12-1 Construction of IgG Type Antibody Gene
[0902]In order to investigate the efficacy as an antibody medicine, a part of antibodies is converted into IgG type
[0903]Firstly, by using VH and VL genes of scFVcp3 type antibody, it is confirmed that there was not a restriction enzyme site necessary for cloning them to Fc region of IgG1 and the base sequence of the gene. PCR was carried out by using an antibody gene as a template and using a primer for amplifying the H chain and L chain were used. The amplified product was ligated to the downstream of CMV promoter of the IgG1 construction vector and a plasmid DNA containing an IgG type antibody gene was obtained.
12-2 Expression of IgG Type Antibody
[0904]For transfection of plasmid DNA into CHO-K1 cell, GenePORTER Reagent (Gene Therapy Systems: T201007) was used. Firstly, CHO-K1 cells were prepared in a 60 mm-culture dish the day before the transfection so that they became 2×104 cells/ml (a medium, in which α-MEM (Invitrogen: 12561-056) to which 10% FCS (Equitech: 268-1) had been added, was used).
[0905]The plasmid DNA (8 μg) was dissolved in 250 μL of serum free medium (hereinafter, abbreviate as "SFM") (Invtrogen: 12052-098 CHO--S--SFMII)) and subjected to 0.22 μm filter. GenePORTER Reagent (40 μL) was added to SFM (250
[0906]The plasmid DNA and GenePORTER Reagent dissolved in SFM were rapidly stirred and stood still at room temperature for 30 minutes.
[0907]The cells were washed with SFM (2 ml) twice, and the plasmid DNA-GenePORTER mixture (Transfection Medium) was slowly poured in a plate containing cells and cultured in an incubator at 37° C. for five hours.
[0908]The medium for transfection was sucked and washed with αMEM 10% FCS twice, then 5 ml of αMEM 10% FCS was added, which was cultured in an incubator at 37° C. for 48 hours.
[0909]The medium was replaced by a medium (10 mL) of αMEM 10% FCS+700 μg/ml G418 (Sigma: G7034) and selection was started (hereinafter, as a medium, αMEM 110% FCS+700 μg/mL G418 was used). After cultured at 37° C. for 48 hours, cells were washed with PBS (10 mL), treated with 0.25% Trypsin-EDTA (Sigma T4049) (750 μL), αMEM (5 mL) was added. Then, cultured product was peeled off and recovered from the plate. The number of cells was measured. Based on the results, limiting dilution was carried out under the conditions of 10 cells/200 μL/well (two sheets of 96 well plates). After culturing for 14 days, ELISA was carried out by using a culture supernatant of each well and the expression of an IgG type antibody was confirmed.
12-3 Purification of Expression Protein (IgG) from Culture Supernatant
[0910]Protein G Sepharose 4 Fast Flow (amersham pharmacia biotech: 17-0618-01) (1 mL) was packed in a column and balanced in PBS (5 mL). The culture supernatant was applied, sent at the flow rate of 1 drop/2 seconds, and allowed the expressed protein (IgG) to be bonded to a column. PBS (10 mL) was sent at the flow rate of 1 drop/2 seconds, non-adsorbed components were washed, then 6 mL of elute buffer (0.2M glycine-HCl, pH 3) was sent at the flow rate of 1 drop/second, and 1 mL each of eluate was collected in a 1.5 ml tube. To the collection tube, neutralizing buffer (3M Tris-HCl) (400 μL) was added in advance. Neutralization was carried out at the same time of collection. The eluate was collected and concentrated to 750 μL, and solution substitution (PBS, complete, 0.01% NaN3) was carried out. Then, the concentration of the antibody protein was calculated by SDS-PAGE.
13. Experiment of Inhibition of Binding of EGF by Successfully Obtained Anti-HER1 Antibody (048-006 Antibody)
13-1 Experimental Procedure
[0911]A431 cells were cultured in 15φ culture dish (medium: DMEM containing 10% FBS and 1% PS), and the cells were peeled off with the use of cell dissociation buffer (GIBCO: 13151-014) and recovered at 90% confluence. Two ml of PBS containing 1.0% BSA and 0.05% NaN3 was added and the recovered cells were suspended. The suspension was stood still at 4° C. for 30 minutes and then 100 μl each (about 2.5×105 cells) was dispensed into each well of a 96-well V-bottom plate. It was centrifuged (650 G) for 2 minutes, and the cells were precipitated to remove the supernatant. Each antibody solution (HR1-007 [10 μg/ml], 48-006 [10 μg/ml, 5 μg/ml, 1 μg/ml], and 59-152 [10 μg/ml, 5 μg/ml, 1 μg/ml]) (200 μl), which had been prepared by using PBS containing 1.0% BSA, was added and the cells were suspended. The suspension was stood still at 4° C. for one hour, and then, biotin labeled EGF (biotinated EGF: 50 μg/ml) was added to each well so that the final concentration became 1 μg/ml, so that the cells were suspended. Note here that the biotinated EGF was produced by the following method. Firstly, to EGF (prepared to 1 mg/ml with PBS(-); AUSTRAL Biologicals: GF-010-5) (50 μl), EZ-Link Sulfo-NHS-LC-Biotin (prepared to 2 mg/ml with PBS(-); PIERCE: 21335) (25 μl) was added. After it was stood still at room temperature for 30 minutes, 1M glycine (pH=7.0 to 8.0) (10 μl) was added. After it was stood still at room temperature for 30 minutes, PBS(-) (15 μl) was added and stored at 4° C. (final concentration: 500 μg/ml). This was 10-fold diluted with PBS containing 1.0% BSA and used for experiment.
[0912]This was stood still at 4° C. for one hour, and centrifuged (650 G) for 2 minutes so as to remove the supernatant. PBS containing 1.0% BSA (180 μl) was added and centrifuged (650 G) for 2 minutes so as to remove the supernatant. HRP-labeled streptavidin (0.2 μg/ml (PBS containing 1.0% BSA); PIERCE: 21126) (100 μl) was added and cells were suspended at 4° C. for one hour, and centrifuged (650 G) for 2 minutes so as to remove the supernatant. PBS containing 1.0% BSA (180 μl) was added and centrifuged (650 G) for 2 minutes so as to remove the supernatant. This operation was carried out again. OPD (Wako: 154-01673) coloring solution (100 μl) was added and cells were suspended. After four minutes at room temperature, coloring stop solution (2NH2SO4) (100 μl) was added and centrifuged (650 G) for 2 minutes. Then, the supernatant was transferred to a flat-bottom plate. By using a plate reader, the absorbance at 192 nm (A492) was measured and represented by a numeric value.
13-2 Results
[0913]The results are shown in FIG. 24. HR1-007 as a control does not affect the binding of EGF. 048-006 antibody and 059-152 antibody inhibit the binding of EGF. 048-006 antibody can inhibit the binding of EGF substantially completely while 059-152 antibody cannot completely inhibit the binding even if the temperature is increased. Note here that 048-006 antibody shows an inhibition effect even at the low level of about 0.02 μg/ml (FIG. 24C).
[0914]The results suggest that the antagonism activity between each antibody (048-006 antibody and 059-152 antibody) and EGF provides a part of the pharmacological effect such as anti-tumor property.
14. Experiment of Phosphorylation Signal Inhibition of HER1 by Successfully Obtained Anti-HER1 Antibody (048-006 Antibody)
[0915]By using a phosphorylation antibody, it was determined whether or not the successfully obtained anti-HER1 antibody (048-006 antibody) inhibited the phosphorylation signal of HER1. Specifically, by using three kinds of cells (renal cell carcinoma (CCF-RC1, Caki-1) and epidermoid cancer (A431)), the inhibition effect of 048-006 antibody and 059-152 antibody and the inhibition effect of ERBITUX were compared with each other.
14-1 Experimental Procedure
[0916]Each of cells was cultured in 6-well culture dish, and at 60% confluence, a medium (DMEM containing 10% FBS and 1% PS) was substituted to DMEM. After 16 hours, each antibody (HR1-007, 048-006, 059-152 (prepared to 2 mg/ml with PBS(-))) and ERBITUX were added to each well so that the final concentration became 10 μg/ml or 1 μg/ml. After 30 minutes, EGF (prepared to 20 μg/ml with PBS(-)) was added to each well so that the final concentration became 1 μg/ml. After 30 minutes, each well was washed with PBS(-) and quickly frozen together with the culture dish by using liquid nitrogen. To each well, lysis buffer (50 mM Tris (pH 7.4), 150 mM NaCl, 1 mM Na3VO4, 10 mM NaF, 1% Triton×100, complete (Roche: 11836145001)) were added, and the cells were suspended and transferred to centrifugation tube. Centrifugation (10000G) was carried out for 10 minutes so as to precipitate cell debris. A part of the supernatant was subjected to SDS-PAGE, which was transferred to a membrane. Western blotting using an anti-phosphorylation tyrosine antibody (11 g/ml; upstate: 05-321) or an anti-β-actin antibody (1 μg/ml; abcam: ab25139) as a primary antibody, and a secondary antibody reaction: HRP labeled anti-mouse IgG as a secondary antibody was carried out. A431 cells were required to be exposed to light for 1 to 2 seconds; CCF-RC1 for 10 seconds; and Caki-1 for one minute (there was originally large difference in cell sensitivity to external stimulation).
14-2 Results
[0917]The results are shown in FIG. 25 (A and B: the results of Western blotting using A431 cells; C to E: comparison effect of inhibiting HER1 phosphorylation signal between the successfully obtained antibody and ERBITUX). In CCF-RC1 and A-431 cell lines, HR1-007 as a control does not affect the phosphorylation signal of HER1. However, 048-006 antibody and 059-152 antibody inhibit signal in a concentration-dependent manner. 048-006 antibody can inhibits the binding of EGF substantially completely and also inhibit self phosphorylation of HER1 substantially completely. 059-152 antibody inhibits the binding of EGF about 50%. Furthermore, 059-152 antibody inhibits self phosphorylation of HER1 although it is weaker than 048-006 antibody. 048-006 antibody and 059-152 antibody have inhibition capabilities superior to that by ERBITUX. In particular, the inhibition capability of 048-006 antibody is remarkable.
[0918]The sensitivity to external stimulation by EGF differs depending upon the kinds of cells. Therefore, when a cell like Caki-1 that does not show sensitivity to external stimulation by EGF is used, the difference in signal inhibition effect by the antibody is not observed.
[0919]The results suggest that each antibody (048-006 antibody and 059-152 antibody) has an activity of suppress the tyrosine kinase circuit of HER1 with respect to sensitive cells of HER1 by EGF, and exhibits pharmacological effects such as proliferation suppression and anti-tumor property.
15. Measurement of Binding Constant by BIAcore
[0920]As to the successfully obtained antibodies 048-006 and 059-152, the dissociation constant with respect to the expression Her1 was measured.
15-1 Experimental Procedure
(1) Forced Expression of Partial Sequence of Her1
[0921]A sequence from a region after the signal of HER1 to immediately before the transmembrane region (621 amino acid of the expression sites from positions 26 to 645 (SEQ ID NO: 943) was cloned. For cloning and expression, a pSecTagII vector (Invitrogen) was used. When this vector is inserted, myc and his tags are added.
(2) Recovering of Expressed Cells
[0922]One 15φ-culture dish (80 confluent) in which 293T cells were cultured was prepared. The medium was replaced with new one so that cells were not peeled off, and then cultured. Thus, a state in which cells were aggregated at 90-100% confluence was formed. The day before recovering cells, final medium replacement was carried out. DNA (75 μl) was added to D-MEM (serum free) (1.9 ml) and subjected to tapping adjustment so as to make the solution A. Furthermore, Lipo (75 μl) was added to D-MEM (serum free) (1.9 ml) and subjected to tapping adjustment (50 ml, Falcon) so as to make the solution B.
[0923]One minute after the formation of the solution B, the solution B was added to the solution A by using a 5 ml-pipette, subjected to pipetting, and incubated at room temperature for 20 minutes. 22.5 ml of D-MEM (serum free) was measured and taken out into a 50 ml culture container (Falcon) and 2.5 ml of serum was added thereto, which was incubated at 37° C. for 15 minutes so as to obtain D-MEM (containing serum).
[0924]The medium was removed from a 15φ-culture dish in which 293T cells were aggregated, and D-MEM (serum free) (25 ml) was added along the wall of the dish carefully so that cells are not peeled off. The added D-MEM (serum free) was sucked by using an aspirator and D-MEM (containing serum) (25 ml) was added. Twenty minutes after D-MEM (containing serum) was formed, the mixture solution (3.8 ml) of solution A and solution B was added to the cells by using 25 ml-pipette and the cells were peeled off. The cells were separated from each other by pipetting, the cells were stood still in a CO2 incubator for 2 days. Two days after, the supernatant was recovered and subjected to protein purification.
(3) Secretory Protein Purification (Ni-NTA)
[0925]Ni-NTA agarose gel (QIAGEN) (2 ml) (bed volume of 1 ml) was packed in a column and balanced in PBS. Then, the culture supernatant recovered in (2) was applied thereto. A flow-through solution was again applied to a column. The column was washed with 5 ml of PBS, and eluted in stages with 20, 50, 100, 250, and 500 mM imidazole/PBS (5 ml each) so that the absorbance (280 nm)<0.005 was satisfied. Furthermore, it was eluted with 0.5M EDTA/PBS (10 ml). The solution was replaced by new one by dialysis so as to obtain BIAcore immobilized sample.
(4) BIAcore Measurement
[0926]The interaction between the antibody clone and the expressed Her1 was examined so as to determine KD (dissociation constant; kd/ka). For analysis, BIAcore 1000 biosensor device was used.
[0927]A carboxymethyldextran (Sensor Chip CM5, Research grade, BIACORE) sensor chip was used. With the electrostatic adsorption to a CM5 matrix and a covalent linkage between a lysyl group on CM5 and an activated carboxyl group, antigen (Her1) was immobilized on the chip. By EDC/NHS coupling chemical reaction, a carboxyl group was activated.
[0928]In the condition of HBS-EP (BIACORE) at a flow rate of 5 μL/minute by using EDC/NHS (amine coupling kit, BIACORE was mixed with equal amount of EDC and NHS), after the lysyl group on CM5 was activated (contact time: 2.4 minutes), chip was washed with HBS-EP (BIACORE). Subsequently, Her1 (20 μg/mL: Sigma, 0.6 mg protein/ml was diluted with 10 mM acetic acid (pH 4.0)) was added to the chip. The chip was washed with HBS-EP, then, 1M ethanolamine (pH 8.5) was added so as to deactivate the remaining activated carboxyl group. Thereafter, the chip was washed with 50 mM NaOH so as to remove all Her1 that were not linked covalently. Note here that all the analysis experiments were carried out under the conditions of HBS-EP (BIACORE) at a flow rate of 35 μl/minute at 25° C. Reproduction was carried out by using 50 mM NaOH (one minute).
[0929]059-152 antibody or 048-006 antibody were reacted at each concentration shown in the figure and HBS-EP at flow rate of 35 μl/minute, so that the binding constant was analyzed.
15-2 Results
[0930]The results are shown in FIG. 26. 048-006 antibody shows extremely strong binding force of more than KD=10-11 (M) at every measurement point. The actual value of Global fitting based on each detection value was 4.8×10-13 (M). This is beyond the reliable measurement limit of BIAcore. As to 059-152 antibody, a bond dissociation curve cannot be detected. This is thought to be because this antibody cannot recognize the higher-order structure of artificially produced forced expression product. In other words, it is suggested that this antibody recognize a higher-order structure of a complex or a higher-order structure that can be observed only on an intact membrane.
16. Cytotoxicity test of anti-HER1 antibody, anti-HER2 antibody, anti-ITGA3 antibody, Anti-ALCAM Antibody, and Anti-ICAM Antibody (ADCC Activity Measurement)
[0931]Antibody-Dependent Cell-mediated Cytotoxicity (ADCC) is an immune reaction of killing and attacking cells harmful to a human body, for example, virus infected cells, in which "effector cells" mainly consisting of natural-killer cell or monocyte attacks cells to which antibodies are bonded widely on the membrane surface as a target. The cytotoxicity by ADCC occurs depending upon the combination of an antibody specifically bonded to a surface of the cell membrane antigen and an effector cell.
[0932]Some of antibodies specifically bonded to a tumor surface antigen have an anti-tumor effect and a therapeutic effect to cancer and sold as antibody medicine. It has been reported that the main mechanism of action of these antibodies are ADCC. Then, in order to evaluate whether or not the cancer antigen specific antibodies successfully isolated by the present inventor have an anti-tumor effect, that is, they have promising as a cancer treatment antibody, the detection of ADCC was carried out. In the below-mentioned experiments, human IgG type antibody clone recognizing a subject antigen is reacted to a target cell to present it to an effector cell. In the detection of ADCC, the degree of cytotoxicity is calculated by using a cytotoxicity detection kit which, in principle, detects the enzymatic activity of lactate dehydrogenase leaking into the medium from the target cancer cell attached by the effector cell by using the coloring of the reagent.
16-1 Induction of ADCC I (Case of 015-003 Antibody as Anti-ITGA3 Antibody: scFVcp3 Type Antibody is Used)
[0933]Regarding 015-003 antibody as anti-ITGA3 antibody, a scFVcp3 type antibody was used and the ADCC activity was investigated by an assay combining an anti-M13 pIII rabbit antibody. Furthermore, liver cancer HLF cell is used as the target subject cultured cell. The operation procedure is described below.
[0934](1) By the following procedures, peripheral blood is collected from a volunteer and mononucleosis is separated. Firstly, heparin-added peripheral blood (30 ml) collected from a volunteer is diluted with PBS to 80 ml and superposed quietly on 10 ml each of lymphocyte isolation reagent Ficoll Paque Plus (Amersham Bioscience), which have been dispensed in four centrifugation tubes, and centrifuged (400×g, 20° C. for 40 minutes). The mononucleosis fractions (including lymphocyte and monocyte) are recovered, diluted with cooled PBS to 80 ml and centrifuged (200×g, 4° C. for 15 minutes).
[0935](2) (1) is suspended in a cooled cytotoxicity test medium (Cytotoxic Medium, hereinafter, abbreviated as "CTM", RPMI-1640 medium, 1% (v/v) fetal calf serum, 1% (v/v) Penicillin-Streptomycin Solution, 1% (v/v) 1M HEPES buffer (pH 7.0): Invitrogen) so that the final density becomes 5.0×106 cells/ml to obtain an effector cell.
[0936](3) In a culture dish having a diameter of 150 mm, a target subject cultured cell is grown in a culture medium I (Minimum Essential Medium Alpha Medium:Invitrogen, 10% (v/v) fetal calf serum: Equitic-Bio, 1% (v/v) Penicillin-Streptomycin Solution Sigma-Aldrich). A liquid medium is removed and cells are washed with PBS (10 ml) twice so as to remove the solution. Thereafter, 4% (w/v) collagenase Type IV (Invitrogen) (5 ml) is added and stored keeping warm at 37° C. for 10 minutes, so that cells are peeled off from the culture dish. Furthermore, 5 ml of liquid medium 2 (RPMI-1640, 10% (v/v) fetal calf serum, 1% (v/v) Penicillin--Streptomycin Solution:Sigma-Aldrich) (RPMI-1640: Sigma-Aldrich, 10% fetal calf serum, 1% penicillin-streptomycin solution) is added to stop a collagenase reaction. Then, suspended cells are recovered to obtain cell suspension.
(4) The cell density of the cell suspension of (3) is measured. The supernatant is removed by centrifugation and the cells are suspended in a cooled CTM medium so that the final density becomes 1.5×105 cells/ml.(5) 100 μl each of target cells is dispensed in a 96-well V-bottom multi plate on ice.(6) 2 μg/ml scFv-pIII phage antibody-CTM solution (100 μl each) is dispensed and reacted on ice for 60 minutes.(7) Centrifugation (Swing rotor: 500×g, 4° C. for 10 min) is carried out to remove the supernatant.(8) Cell pellet is suspended in 5 μg/ml anti-M13 pill rabbit polyclonal antibody-CTM solution (150 μl each), a part of 100 μl is transferred to a 96-well U-bottom multi plate.(9) The effector cell of (2) (or 2% Triton X-100-CTM solution) is added and then centrifuged (Swing rotor: 50×g, 4° C. for 5 min).(10) Reaction is carried out in 5% CO2 at 37° C. for 4 hours.(11) After the reaction, centrifugation (Swing rotor: 500×g, 4° C. for 10 min) is carried out and the supernatant (100 μl) is transferred to a flat-bottom 96 well multi plate.(12) LDH activity measurement reagent (Roche) (100 μl) is added and reaction is carried out at room temperature for 30 min.(13) OD490 and OD690 are measured by using a micro plate absorptiometer.16-2 Induction of ADCC 2 (case of 048-006 antibody as anti-HER1 antibody, 015-126 antibody as anti-HER2 antibody, 066-174 antibody, 035-234 antibody and 041-118 antibody as anti-ALCAM antibody, 053-051 antibody, 053-059 antibody and 053-085 antibody as anti-ICAM1 antibody, 067-153 antibody as anti-EpCAM antibody, 067-133 antibody as anti-HGFR antibody: IgG type antibody is used)
[0937]Regarding 048-006 antibody as anti-HER1 antibody, an IgG type antibody was used and the ADCC activity was investigated. A-431 and A549 (epidermoid tumor), ACHN and CCF-RC-1 (kidney cancer), NCI-H1373 (lung cancer), as well as SK-OV-3 (ovarian cancer) were used as the target subject cultured cell.
[0938]Also regarding 015-126 antibody as anti-HER2 antibody, an IgG type antibody was used, and the ADCC activity was investigated. Breast cancer BT-474 was used as the target subject cultured cell.
[0939]Regarding 066-174 antibody and 035-234 antibody as anti-ALCAM antibody, an IgG type antibody was used, and the ADCC activity was investigated. NCI-H1373 (pulmonary adenocarcinoma), CW2 (large bowel cancer), or NCI-H441 (lung cancer) was used as the target subject cultured cell.
[0940]Regarding 053-051 antibody, 053-059 antibody and 053-085 antibody as anti-ICAM1 antibody, an IgG type antibody was used, and the ADCC activity was investigated. HepG2 (hepatic cell carcinoma) and NCI-H441 (lung cancer) were used as the target subject cultured cell.
[0941]Furthermore, regarding the effect of 048-006 antibody or 059-152 antibody as anti-HER1 antibody on CCF-RC-1 (kidney cancer), NCI-H1373 (lung cancer) and A-431 (epidermoid cancer), the antibody dosage dependence of the ADCC activity was investigated so that the final concentration of the IgG type antibody was in the range from 0.01 to 10 μg/ml.
[0942]Regarding 041-118 antibody as anti-ALCAM antibody, an IgG type antibody was used and the antibody dosage dependence of the ADCC activity was investigated. NCI-H1373 (pulmonary adenocarcinoma) was used as the target subject cultured cell.
[0943]Regarding 067-153 antibody as anti-EpCAM antibody, an IgG type antibody was used and the antibody dosage dependence of the ADCC activity was investigated. MKN-45 (solid-type gastric adenocarcinoma), HT-29 (colon adenocarcinoma) and NCI-H1373 (lung cancer) were used as the target subject cultured cell.
[0944]Regarding 067-133 antibody as anti-HGFR antibody, an IgG type antibody was used and the antibody dosage dependence of the ADCC activity was investigated. NCI-H1373 (lung cancer) was used as antibody dosage dependence of the ADCC activity.
[0945]The antibody dosage dependence of the ADCC activity was basically measured at the E/T Ratio (ratio of effector cell:target cell) of 80:1 at final antibody concentration in the solution of 0.01 μg/ml to 10 μg/ml or 10-6 μg/ml to 10 μg/ml.
[0946]At each measurement point, the antibody and the effector cell were added to the target cell, and four hours later, the ADCC activity was measured. Regarding NCI-H1373, the ADCC activity was measured at the E/T Ratio of 100:1.
[0947]The operation procedure was carried out in accordance with the procedures described in 16-1. The detail of the reaction was made to be as follows. 66 μl/well of the target cells (2×104 cells) were placed in 96-well U-bottom plate (Becton Dickinson) and 66 μl of IgG type antibody (3 μg/ml) was added and then 66 μl of peripheral blood mononucleosis suspension (7.5×105 cells) was added. The E/T
[0948]Ratio (ratio of effector cell:target cell) was made to be 20. In order to promote the association of cells, centrifugation (60×g, 4° C., 5 minutes) so as to allow the cells to sink, which was stored keeping warm 240 minutes in a culture container that had been set to the conditions of 37° C. and 5% CO2. Thus ADCC reaction was induced. Each antibody sample was prepared as a CTM solution. Furthermore, in each sample, CTM was used as a negative control and target cell to which 100 μl of 2% Triton X-100-CTM solution was added was used as a control of maximum liberation of lactate dehydrogenase (cells had been destroyed by Triton X-100 in advance). Furthermore, three wells were used for each experiment groups.
16-3 Measurement of ADCC Activity
[0949]In both the assay using a scFVcp3 type antibody and assay using an IgG type antibody, the ADCC activity was an indicator of the damage to the target cell, which is in proportion to the degree of coloring, that is, the concentration of lactate dehydrogenase liberating to the culture supernatant. Thirty minutes after the coloring starts, absorbance (OD490-OD620 (background absorbance)) was measured by using a spectrophotometer. In each experiment group, absorbance values in the three wells were averaged to calculate the cytotoxic Index. In advance, the absorbance of only a medium was subtracted and the calculation was carried out by the following calculation equation.
Relative LDH activity=OD490-OD690
LDH activity derived from cell=experimental value-(control containing only solution)
Cytotoxicity (%)=(experimental value-effector cell control-target cell control)/(cell+Triton X-100 control-target cell control)×100 [Equation 1]
[0950]Note here that when the antibody does not have any cytotoxic activity, the cytotoxicity calculated by this method may be minus value due to a measurement error because the measurement is carried in experiments using the living body components.
16-4 Measurement Result
[0951]The measurement results of the ADCC activity are shown in FIG. 27 (anti-ITGA3 antibody was used; the target culture cell was HLF), FIG. 28 (anti-HER1 antibody was used; the target culture cell was A-431), FIG. 29 (anti-HER1 antibody was used; the target culture cell was A549), FIG. 30 (anti-HER1 antibody was used; the target culture cell was ACHN), FIG. 31 (anti-HER1 antibody was used; the target culture cell was CCF-RC-1), FIG. 32 (anti-HER1 antibody was used; the target culture cell was NCI-H1373), FIG. 33 (anti-HER1 antibody was used; the target culture cell was SK-OV-3), FIG. 34 (anti-HER2 antibody was used; the target culture cell was BT-474), FIG. 35 (066-174 as anti-ALCAM antibody was used; the target culture cell was NCI-H1373, CW2, or NCI-H441), FIG. 36 (035-234 as anti-ALCAM antibody was used; the target culture cell was CW2 or NCI-H441), FIG. 37 (053-051 as anti-ICAM1 antibody was used; the target culture cell was NCI-H441 and HepG2), FIG. 38 (053-059 as anti-ICAM1 antibody was used; the target culture cell was NCI-H441 and HepG2), and FIG. 39 (053-085 as anti-ICAM1 antibody was used; the target culture cell was NCI-H441 and HepG2). I
[0952]The measurement results of the antibody dosage dependence of the ADCC activity are shown in FIG. 40 (048-006 or 059-152 antibody as anti-HER I antibody was used; the target culture cell was CCF-RC-1), FIG. 41 (048-006 or 059-152 antibody as anti-HER1 antibody was used; the target culture cell was NCI-H1373), FIG. 42 (048-006 or 059-152 antibody as anti-HER1 antibody was used; the target culture cell was A-431), FIG. 43 (041-118 antibody as anti-ALCAM antibody was used; the target culture cell was NCI-H1373), FIG. 44 (067-153 antibody as anti-EpCAM antibody was used; the target culture cell was MKN-45), FIG. 45 (067-153 antibody as anti-EpCAM antibody was used; the target culture cell was HT-29), FIG. 46 (067-153 antibody as anti-EpCAM antibody was used; the target culture cell was NCI-H1373), and FIG. 47 (067-133 antibody as anti-HGFR antibody was used; the target culture cell was NCI-H1373).
[0953]Similarly, the measurement results of the antibody dosage dependence of the ADCC activity are shown in FIG. 48 (055-147 antibody or 059-173 antibody as anti-HER1 antibody was used; the target culture cell was CCF-RC1), FIG. 49 (048-006 antibody, 059-152 antibody, 055-147 antibody or 059-173 antibody as anti-HER1 antibody was used; the target culture cell was HT-29), FIG. 50 (048-006 antibody, 055-147 antibody or 059-173 antibody as anti-HER1 antibody was used; the target culture cell was A431), FIG. 51 (048-006 antibody or 059-152 antibody as anti-HER1 antibody was used; the target culture cell was ACHN), FIG. 52 (035-234 antibody or 066-174 antibody as anti-ALCAM antibody was used; the target culture cell was NCI-H1373), FIG. 53 (035-234 antibody or 066-174 antibody as anti-ALCAM antibody was used; the target culture cell was target cell SKOv3), FIG. 54 (035-234 antibody or 066-174 antibody as anti-ALCAM antibody was used; the target culture cell was CW-2), FIG. 55 (041-118 antibody as anti-ALCAM antibody was used; the target culture cell was EBC-1), FIG. 56 (080-040 antibody as anti-ALCAM antibody was used; the target culture cell was NCI-H1373), FIG. 57 (053-042 antibody as anti-ICAM1 antibody was used; the target culture cell was NCI-H1373), FIG. 58 (053-051 antibody, 053-059 antibody or 053-085 antibody as anti-ICAM I antibody was used; the target culture cell was NCI-H1373), FIG. 59 (067-153 antibody as anti-EpCAM antibody was used; the target culture cell was EBC-1), FIG. 60 (067-133 antibody as anti-HGFR antibody was used; the target culture cell was MKN-45), FIG. 61 (067-133 antibody as anti-HGFR antibody was used; the target culture cell was EBC-1), FIG. 62 (015-003 antibody as anti-ITGA3 antibody was used; the target culture cell was ACHN), FIG. 63 (059-053 antibody as anti-CD147 antibody was used; the target culture cell was CCF-RC1), FIG. 64 (059-053 antibody as anti-CD147 antibody was used; the target culture cell was ACHN), FIG. 65 (064-044 antibody or 079-085 antibody as anti-PTP-LAR antibody was used; the target culture cell was PC-14), and FIG. 66 (064-003 antibody as anti-CD44 antibody was used; the target culture cell was PC-14).
[0954]In any of anti-ITGA3 antibody (015-003), anti-HER1 antibody (048-006) and anti-HER2 antibody (015-126), anti-CD44 antibody (064-003), the cytotoxicity was increased in the experiment groups in which the effector cell was added. That is to say, in any of the antibodies, the cytotoxic activity caused by the effector cell that recognizes an antibody to which a target cell has been specifically-bound and attacks the target cell was observed.
[0955]Note here that an anti-habu venom antibody (control antibody) HR1-007 that is not related to the surface antigen or the experiment group in which the antibody clone is not added, the increase in the cytotoxicity is not observed. In any of anti-ALCAM antibodies (066-174, 035-234, 041-118, and 083-040), anti-ICAM1 antibody (053-051, 053-059, 053-085, and 053-042), and anti-CD147 antibody (059-053), the cytotoxicity is increased more significantly than in the anti-habu venom antibody (control antibody) HR1-007 experiment group. As mentioned above, it is clearly shown that the antibody dependent cytotoxicity is higher than that of the control antibody (HR1-007) with a significant difference.
[0956]From the above-mentioned results, it has been confirmed that an antibody capable of specifically recognizing a cancer cell and exhibiting a damaging effect by the ADCC activity has been obtained for each of HER1, HER2, ITGA3, ALCAM, ICAM, CD44, CD147, EPCAM and HGFR. In other words, an antibody that is a promising as the antibody medicine targeting each of cancer cells has been obtained.
[0957]In the results of the antibody dosage dependence test, anti-HER1 antibody (048-006) shows a significant effect even if the dosage is 0.01 μg/ml. It is determined that the effect is expected with low dosage.
[0958]It is observed that the 048-006 antibody and 059-152 antibody tend to have a strong ADCC activity in the cell line in which HER1 is expressed. However, the activity differs depending upon the concentration range of the antibody to be used or the kinds of antibodies. To A431 cell, with 0.001 Hg/ml, the difference in the activity was observed. Generally, in the low concentration range, the activity of 059-152 antibody was more significant than that of 048-006 antibody. Furthermore, 055-147 antibody and 059-173 antibody shows higher ADCC activity than ERBITUX® that is commercially available drug and is more useful.
[0959]Furthermore, 067-153 antibody as anti-EpCAM antibody shows an excellent ADCC activity to MKN-45 (solid-type gastric adenocarcinoma) cell line at the concentration of 0.01 μg/ml or more, and it shows an excellent ADCC activity to HT-29 (colon adenocarcinoma) cell line at the concentration of 10 μg/ml or more with an amazing score of 80% or more in the ADCC activity in the concentration range of about 1 μg/ml. It shows an amazing score of 50% or more in the ADCC activity in NCI-H1373 (pulmonary adenocarcinoma) cell line at the concentration of 0.01 μg/ml or more.
[0960]Furthermore, 041-118 antibody as anti-ALCAM antibody shows a remarkable effect to NCI-H1373 (pulmonary adenocarcinoma) cell line at the concentration of 0.01 μg/ml or more. It is determined that the effect can be expected with low dosage.
[0961]Furthermore, 066-174 antibody as anti-ALCAM antibody shows high ADCC activity to various cells such as NCI-H1373 (pulmonary adenocarcinoma) cell line, SKOv3 (ovarian cancer) cell line, and CW-2 (large bowel cancer) cell line. Wide application is expected.
[0962]Furthermore, 067-133 antibody as anti-HGFR antibody shows a remarkable effect to NCI-H1373 (pulmonary adenocarcinoma) cell line at the concentration of 0.01 μg/ml or more with strong activity of 40% or more at the concentration of 10 μg/ml or more.
[0963]Furthermore, 059-053 antibody as anti-CD147 antibody shows an excellent ADCC effect to CCF-RC1 (kidney cancer) cell line and ACHN (kidney cancer) cell line, which shows near the upper limit value at the low concentration. Therefore, it can be expected to show the maximum activity at a low concentration.
[0964]From the above-mentioned results, it is confirmed that a promising antibody group showing a sufficient ADCC activity even with low dosage (at low concentration) can be obtained successfully. Also in the similar experiments using a plurality of lymphocyte fractions derived from human, the same results as mentioned above can be obtained. The high reproducibility is confirmed.
17. Cancer Cell Proliferation Inhibition Test
[0965]Some antibody medicines exhibit the efficacy by an effect of inhibiting the proliferation of cancer instead of the ADCC effect (or in addition to the ADCC effect). Thus, in order to further investigate the efficacy of antibody medicine, the activity of inhibiting the proliferation of cancer by antibodies that have been successfully isolated have been investigated according to the following procedure.
17-1 Testing Method
[0966](1) Target culture cells that have grown in a culture dish are peeled off with 4% Collagenase and suspended in the used medium.(2) The cell density is measured and then the supernatant is removed by centrifugation and suspended in a RPMI-1640 (10% FBS, 1% Penicillin--Streptomycin) medium so that the final density is 1.0×104 cells/ml.(3) 100 μl each of target cells is dispensed in a flat-bottom 96 well multi plate.(4) 100 μl each of 20 μg/ml human IgG monoclonal antibody solution is dispensed.(5) Reaction is carried out in 5% CO2 at 37° C. for 5 days.(6) Medium is removed, and living cell measurement reagent (XTT: Roche) is dispensed in each well (150 μl each).(7) Reaction is carried out in 5% CO2 at 37° C. for 4 hours.(8) After reaction, OD490 and OD690 are measured by using a micro plate absorptiometer. Then, the number of living cells is calculated according to the following equation.
XTT reduction amount (degree of coloring)=OD490-OD690
XTT reducing activity derived from cells=(experimental value)-(control value using only a solution) [Equation 2]
[0967]Note here that the XTT reducing activity derived from cells is in proportion to the number of living cells.
17-2 Results
[0968]The results are shown in FIG. 67 (anti-HER1 antibody (048-006) was used; the target subject cultured cell was A-431), FIG. 68 (anti-HER1 antibody (048-006) was used; the target subject cultured cell was ACHN), FIG. 69 (anti-HER1 antibody (048-006) was used; the target subject cultured cell was NCI-H1373), FIG. 70 (anti-HER1 antibody (048-006) was used; the target subject cultured cell wasSK-OV-3), and FIG. 71 (anti-HER2 antibody (015-126) was used; the target subject cultured cell was BT-474).
[0969]As is apparent from these drawings, it is confirmed that the antibodies inhibiting the proliferation of cancer cell can be successfully obtained. In other words, it is shown that these antibodies may be effective as antibody medicine of suppressing the proliferation of cancer cells.
18. Antitumor Experiment Using Mouse
[0970]Next, whether or not the antibodies that have been successfully isolated show an anti-tumor activity in vivo is confirmed by using a cancer cell-transplanted mouse.
18-1 Animals and Cell Line to be Used
[0971]Four-week old female BALB/c nude mouse (Charles River Japan) was acclimated and bred for one week and then used for experiment. The animals were bred under the SPF environment and fed with sterilized water and feed.
[0972]Human lung cancer cell H1373 or epidermoid tumor A-431, which had been subcultured in a RPMI medium containing 10% FBS at 37° C. in the presence of 5% CO2, were used.
18-2 Method of Antitumor Experiment
[0973]Human lung cancer cells, H1373 cells (1×107 cells) were transplanted in the dorsolateral subcutaneous portion of a nude mouse so as to produce a tumor. At the time the tumor volume was 1 cm3, the tumor was cut into a size of 3 mm×3 mm, and is was successive-transplanted to the dorsal subcutaneous portion of the prepared nude mouse. After transplantation, when a volume of the tumor was estimated to be 200 mm3, administration of the antibody was started. The diameter of the tumor and body weight were measure twice a week, estimated tumor volume was calculated from the equation: W=a×b2/2 (W: estimated tumor volume (mm3), a: major axis (mm), b: minor axis (mm)). The experiment group was divided into a control group (PBS was administered) and 048-006 IgG administered group (0.5 mg/individual). The administration pathway was made to be an intraperitoneal administration. Administration was carried out twice a week eight times in total. Then, the anti-tumor effect was examined.
[0974]Furthermore, ERBITUX (Cetuximab, Bristol-Myers Squibb Company) was used as a comparative group or an additivity examining group. When ERBITUX is used singly, the dosage amount was made to be 0.25 mg/individual. ERBITUX was used together with 048-006 IgG, the dosage amount of ERBITUX was made to be 0.25 mg/individual and the dosage amount of 048-006 IgG was made to be 0.25 mg/individual. After administration, the follow-up was carried out.
[0975]When epidermoid tumor A-431 is used, epidermoid tumors A-431 (5×106) were similarly transplanted in the dorsolateral subcutaneous portion of a five-week old female BALB/c nude mouse nude mouse so as to produce a tumor. At the time the tumor volume was estimated to be 200 mm3, administration of the antibody was started. The administration pathway was made to be an intraperitoneal administration. 048-006 IgG type antibody was administered twice a week six times in total. Then, the anti-tumor effect was examined. 059-152 IgG type antibody administered group (0.25 mg or 1.00 mg of antibody was diluted in 0.5 ml PBS/individual) twice a week six times in total. Then, the anti-tumor effect was examined. And the follow up was also carried out.
18-3 Results
[0976]In the antibody (048-006 IgG type antibody) administered group, estimated tumor volume was significantly reduced as compared with the control group (PBS was administered), showing a clear anti-tumor effect. It was confirmed that the effect was comparative to ERBITUX (see FIGS. 72 to 75). On the other hand, in the antibody (059-152 IgG type antibody) administered group, estimated tumor volume was significantly reduced as compared with the control group (PBS was administered), showing a clear anti-tumor effect. The effect was more excellent than that of ERBITUX (see FIG. 75). 059-152 antibody shows stronger tumor suppression effect than 048-006 antibody and commercially available ERBITUX. Thus, it was confirmed that the successfully obtained antibodies exhibited the anti-tumor effect in also in vivo model. In other words, they are shown to be an extremely promising as the antibody medicine.
[0977]19. Analysis by Three Dimensional ELISA
(1) Expression of Antibody by Culturing Screened Clone Group and Preparation of Antibody Mixture
[0978]Clones (about 4000 clones) of phage-infected E. coli, which were screened by the methods described in 1 to 5, were transferred to 41 sheets of 96-well plates at 1 clone/well, and they were shaking cultured in 100 μl/well YTGA medium (YT medium+1% Glucose+200 μg/ml Ampicillin) at 30° C. overnight. Next, 10 μl each of culture solution was mixed in all wells of the first to sixth columns for each plate to make one group (however, as to the 28th plate, the first to seventh columns are made to be one group). Forty-one plates of the mixed antibodies were obtained in total. As to 7th to 12th columns were also made into one group (excluding the 28th plate and 35th plate). Thirty-nine plates of the mixed antibodies in total were obtained. Furthermore, after the plates were divided into 7 groups (3, 6 or 7 sheets per group), for each group, 10 μl each of culture solution was mixed in all wells in each row and they were made to one group. Thus, 56 rows of the mixed antibodies in total were obtained. Finally, after the plates were divided into 5 groups (3, 9 or 10 sheets per group), for each group, 10 μl each of culture solution was mixed in all wells in each column and they were made to one group. Thus, 54 columns of the mixed antibodies in total (in a part, two columns were made to one group) were obtained.
[0979]A YT0.05GA medium (YT medium+0.05% Glucose+200 μg/ml Ampicillin) (100 ml) was added to each mixed antibody, and shaking cultured at 30° C. until OD600 nm was about 0.3 to 0.5. Thereafter, IPTG was added so that the final concentration was 0.5 mM and further shaking cultured at 30° C. The mixture was centrifuged at 10000 rpm at 4° C. for 15 minutes, and the culture supernatant was recovered. Then, ammonium sulfate (29.1 g) was slowly added and mixed, mixture was centrifuged at 10000 rpm at 4° C. for 20 minutes, and sediment was recovered. The sediment was suspended in 5 ml of PBS/NaN3/complete. The suspension was centrifuged at 10000 rpm at 4° C. for 20 minutes, and the culture supernatant was recovered. Thus, 20-fold concentrated mixed antibodies (190 types) were obtained.
(2) Measurement by Three-Dimensional ELISA
[0980]Three dimensional ELISA was carried out by using the obtained 20-fold concentrated mixed antibodies (190 types). Firstly, 50 μl/well of antigen whose concentration was adjusted to be 20 μg/ml with PBS was added to Maxisorp (Nunc) and reacted at 37° C. for two hours to be sensitized. After the liquid in each well was removed, 5% skim milk/PBS (200 μg/well) was added and reacted at 37° C. for two hours for blocking. The liquid in each well was removed and washed with PBS, and 20-fold concentrated mixed antibody (100 μl/well) was added and reacted at 37° C. for one hour. The reacted product was washed with PBS, and a rabbit anti-cp3 antibody (MBL) that had been 5000-fold diluted with 0.05% Tween/PBS was added (100 μl/well) and reacted at 37° C. for one hour. The mixture was washed with PBS, and an HRP labeled goat anti-rabbit IgG antibody (MBL) that had been 2000-fold diluted with 0.05% Tween/PBS was added (100 μl/well) and reacted at 37° C. for one hour. The reacted product was washed with PBS and a substrate solution (100 μl/well) was added. The substrate solution was produced as followed. That is to say, to 12 ml of 0.1 M citric acid-disodium hydrogen-phosphate (pH 5.1), H2O2 was added so that the final concentration became 0.01% and furthermore, OPD tablet (Wako Pure Chemical) was added.
[0981]2N sulfuric acid (100 μl/well) was added to stop the reaction and the absorbance at 492 nm was measured by using a plate reader (Wako Pure Chemical, SUNRISE Remote).
[0982]The measurement results are shown in FIGS. 79 to 81 (ELISA using CK147 as an antigen) and FIGS. 82 to 84 (ELISA using HER1 as an antigen).
[0983]Based on the results of the above-mentioned three dimensional ELISA, positive clones were selected. That is to say, from information of plate, row and column providing positive results, intersection point was searched and antibody clones existing in the intersection point were selected. The selected antibody clones were shaking cultured in 75 μl/well YTGA medium at 30° C. overnight. In 200 μl/well YT0.05GA medium, the culture solution was plated and standing cultured at 37° C. for four hours. Thereafter, IPTG was added so that the final concentration became 1 mM and shaking cultured at 30° C. overnight. The culture was centrifuged at 3000 rpm at 4° C. for 10 minutes and the culture supernatant was recovered.
(3) Reactivity of Selected Antibody Clones
[0984]50 μl/well of antigen (CD147 or HER1) whose concentration was adjusted to be 10 μg/ml with PBS was added to Maxisorp (Nunc) and reacted at 37° C. for two hours to be sensitized. After the liquid in each well was removed, 5% skim milk/PBS (200 μl/well) was added and reacted at 37° C. for two hours for blocking. The liquid in each well was removed and washed with PBS. The culture supernatant of the selected clones (100 μl/well) was added and reacted at 37° C. for one hour. The reacted product was washed with PBS, and a rabbit anti-cp3 antibody (MBL) that had been 5000-fold diluted with 0.05% Tween/PBS was added (1001l/well) and reacted at 37° C. for one hour. The mixture was washed with PBS, and an HRP labeled goat anti-rabbit IgG antibody (MBL) that had been 2000-fold diluted with 0.05% Tween/PBS was added (100 μl/well) and reacted at 37° C. for one hour. The reacted product was washed with PBS and a substrate solution (100 μl/well) was added. 2N sulfuric acid (100 μl/well) was added to stop the reaction and the absorbance at 492 nm was measured by using a plate reader (Wako Pure Chemical, SUNRISE Remote). The results of ELISA using HER1 as an antigen is show in FIG. 85. As is apparent from the graph of FIG. 85, a large number of monoclonal antibodies to HER1 were obtained.
20. Newly Obtained Antibodies
[0985]By using the classifying method and identification method of the present invention, it was possible to obtain the following antibodies successfully.
(1) Antibody to C1qR
[0986]070-016 antibody
(a) Amino Acid Sequence
[0987]SEQ ID NO: 451 (VH); SEQ ID NO: (VH CDR1) 452; SEQ ID NO: 453 (VH CDR2); SEQ ID NO: 454 (VH CDR3), SEQ ID NO: 455 (VL); SEQ ID NO: (VL CDR1) 456; SEQ ID NO: 457 (VL CDR2); and SEQ ID NO: 458 (VL CDR3)
(b) Base Sequence
[0988]SEQ ID NO: 843 (VH); and SEQ ID NO: 844 (VL)
(2) Antibody to CD44
[0989]064-003 antibody
(a) Amino Acid Sequence
[0990]SEQ ID NO: 459 (VH); SEQ ID NO: 460 (VH CDR1); SEQ ID NO: 461 (VH CDR2); SEQ ID NO: 462 (VH CDR3); SEQ ID NO: 463 (VL); SEQ ID NO: 464 (VL CDR1); SEQ ID NO: 465 (VL CDR2); and SEQ ID NO: 466 (VL CDR3)
(b) Base Sequence
[0991]SEQ ID NO: 845 (VH); and SEQ ID NO: 846 (VL)
(3) Antibody to CD73
[0992]067-213 antibody
(a) Amino Acid Sequence
[0993]SEQ ID NO: 467 (VH); SEQ ID NO: 468 (VH CDR1); SEQ ID NO: 469 (VH CDR2); SEQ ID NO: 470 (VH CDR3); SEQ ID NO: 471 (VL); SEQ ID NO: 472 (VL CDR1); SEQ ID NO: 473 (VL CDR2); and SEQ ID NO: 474 (VL CDR3)
(b) Base Sequence
[0994]SEQ ID NO: 847 (VH); and SEQ ID NO: 848 (VL)
(4) Antibody to EpCAM
[0995]067-153 Antibody
(a) Amino Acid Sequence
[0996]SEQ ID NO: 475 (VH); SEQ ID NO: 476 (VH CDR1); SEQ ID NO: 477 (VH CDR2); SEQ ID NO: 478 (VH CDR3); SEQ ID NO: 479 (VL); SEQ ID NO: 480 (VL CDR1); SEQ ID NO: 481 (VL CDR2); and SEQ ID NO: 482 (VL CDR3)
(b) Base Sequence
[0997]SEQ ID NO: 849 (VH); and SEQ ID NO: 850 (VL)
(5) Antibody to HER1
[0998]048-040 antibody
(a) Amino Acid Sequence
[0999]SEQ ID NO: 483 (VH); SEQ ID NO: 484 (VH CDR1); SEQ ID NO: 485 (VH CDR2); SEQ ID NO: 486 (VH CDR3); SEQ ID NO: 487 (VL); SEQ ID NO: 488 (VL CDR1); SEQ ID NO: 489 (VL CDR2); and SEQ ID NO: 490 (VL CDR3)
(b) Base Sequence
[1000]SEQ ID NO: 851 (VH); and SEQ ID NO: 852 (VL)
[1001]054-101 antibody
(a) Amino Acid Sequence
[1002]SEQ ID NO: 491 (VH); SEQ ID NO: 492 (VH CDR1); SEQ ID NO: 493 (VH CDR2); SEQ ID NO: 494 (VH CDR3); SEQ ID NO: 495 (VL); SEQ ID NO: 496 (VL CDR1); SEQ ID NO: 497 (VL CDR2); and SEQ ID NO: 498 (VL CDR3)
(b) Base Sequence
[1003]SEQ ID NO: 853 (VH); and SEQ ID NO: 854 (VL)
[1004]055-147 antibody
(a) Amino Acid Sequence
[1005]SEQ ID NO: 499 (VH); SEQ ID NO: 500 (VH CDR1); SEQ ID NO: 501 (VH CDR2); SEQ ID NO: 502 (VH CDR3); SEQ ID NO: 503 (VL); SEQ ID NO: 504 (VL CDR1); SEQ ID NO: 505 (VL CDR2); and SEQ ID NO: 506 (VL CDR3)
(b) Base Sequence
[1006]SEQ ID NO: 855 (VH); and SEQ ID NO: 856 (VL)
[1007]059-173 antibody
(a) Amino Acid Sequence
[1008]SEQ ID NO: 507 (VH); SEQ ID NO: 508 (VH CDR1); SEQ ID NO: 509 (VH CDR2); SEQ ID NO: 510 (VH CDR3); SEQ ID NO: 511 (VL); SEQ ID NO: 512 (VL CDR1); SEQ ID NO: 513 (VL CDR2); and SEQ ID NO: 514 (VL CDR3)
(b) Base Sequence
[1009]SEQ ID NO: 857 (VH); and SEQ ID NO: 858 (VL)
[1010]067-149 antibody
(a) Amino Acid Sequence
[1011]SEQ ID NO: 515 (VH); SEQ ID NO: 516 (VH CDR1); SEQ ID NO: 517 (VH CDR2); SEQ ID NO: 518 (VH CDR3); SEQ ID NO: 519 (VL); SEQ ID NO: 520 (VL CDR1); SEQ ID NO: 521 (VL CDR2); and SEQ ID NO: 522 (VL CDR3)
(b) Base Sequence
[1012]SEQ ID NO: 859 (VH); and SEQ ID NO: 860 (VL)
[1013]067-176 antibody
(a) Amino Acid Sequence
[1014]SEQ ID NO: 523 (VH); SEQ ID NO: 524 (VH CDR1); SEQ ID NO: 525 (VH CDR2); SEQ ID NO: 526 (VH CDR3); SEQ ID NO: 527 (VL); SEQ ID NO: 528 (VL CDR1); SEQ ID NO: 529 (VL CDR2); and SEQ ID NO: 530 (VL CDR3)
(b) Base Sequence
[1015]SEQ ID NO: 861 (VH); and SEQ ID NO: 862 (VL)
(6) Antibody to HER2
[1016]015-044 antibody
(a) Amino Acid Sequence
[1017]SEQ ID NO: 531 (VH); SEQ ID NO: 532 (VH CDR1); SEQ ID NO: 533 (VH CDR2); SEQ ID NO: 534 (VH CDR3); SEQ ID NO: 535 (VL); SEQ ID NO: 536 (VL CDR1); SEQ ID NO: 537 (VL CDR2); and SEQ ID NO: 538 (VL CDR3)
(b) Base Sequence
[1018]SEQ ID NO: 863 (VH); and SEQ ID NO: 864 (VL)
[1019]015-102 antibody
(a) Amino Acid Sequence
[1020]SEQ ID NO: 539 (VH); SEQ ID NO: 540 (VH CDR1); SEQ ID NO: 541 (VH CDR2); SEQ ID NO: 542 (VH CDR3); SEQ ID NO: 543 (VL); SEQ ID NO: 544 (VL CDR1); SEQ ID NO: 545 (VL CDR2); and SEQ ID NO: 546 (VL CDR3)
(b) Base Sequence
[1021]SEQ ID NO: 865 (VH); and SEQ ID NO: 866 (VL)
[1022]015-136 antibody
(a) Amino Acid Sequence
[1023]SEQ ID NO: 547 (VH); SEQ ID NO: 548 (VH CDR1); SEQ ID NO: 549 (VH CDR2); SEQ ID NO: 550 (VH CDR3); SEQ ID NO: 551 (VL); SEQ ID NO: 552 (VL CDR1); SEQ ID NO: 553 (VL CDR2); and SEQ ID NO: 554 (VL CDR3)
(b) Base Sequence
[1024]SEQ ID NO: 867 (VH); SEQ ID NO: 868 (VL)
[1025]015-143 antibody
(a) Amino Acid Sequence
[1026]SEQ ID NO: 555 (VH); SEQ ID NO: 556 (VH CDR1); SEQ ID NO: 557 (VH CDR2); SEQ ID NO: 558 (VH CDR3); SEQ ID NO: 559 (VL); SEQ ID NO: 560 (VL CDR1); SEQ ID NO: 561 (VL CDR2); SEQ ID NO: 562 (VL CDR3)
(b) Base Sequence
[1027]SEQ ID NO: 869 (VH); SEQ ID NO: 870 (VL)
[1028]015-209 antibody
(a) Amino Acid Sequence
[1029]SEQ ID NO: 563 (VH); SEQ ID NO: 564 (VH CDR1); SEQ ID NO: 565 (VH CDR2); SEQ ID NO: 566 (VH CDR3); SEQ ID NO: 567 (VL); SEQ ID NO: 568 (VL CDR1); SEQ ID NO: 569 (VL CDR2); SEQ ID NO: 570 (VL CDR3)
(b) Base Sequence
[1030]SEQ ID NO: 871 (VH); SEQ ID NO: 872 (VL)
[1031]039-016 antibody
(a) Amino Acid Sequence
[1032]SEQ ID NO: 571 (VH); SEQ ID NO: 572 (VH CDR1); SEQ ID NO: 573 (VH CDR2); SEQ ID NO: 574 (VH CDR3); SEQ ID NO: 575 (VL); SEQ ID NO: 576 (VL CDR1); SEQ ID NO: 577 (VL CDR2); SEQ ID NO: 578 (VL CDR3)
(b) Base Sequence
[1033]SEQ ID NO: 873 (VH); SEQ ID NO: 874 (VL)
[1034]053-216 antibody
(a) Amino Acid Sequence
[1035]SEQ ID NO: 579 (VH); SEQ ID NO: 580 (VH CDR1); SEQ ID NO: 581 (VH CDR2); SEQ ID NO: 582 (VH CDR3); SEQ ID NO: 583 (VL); SEQ ID NO: 584 (VL CDR1); SEQ ID NO: 585 (VL CDR2); SEQ ID NO: 586 (VL CDR3)
(b) Base Sequence
[1036]SEQ ID NO: 875 (VH); SEQ ID NO: 876 (VL)
[1037]075-024 antibody
(a) Amino Acid Sequence
[1038]SEQ ID NO: 587 (VH); SEQ ID NO: 588 (VH CDR1); SEQ ID NO: 589 (VH CDR2); SEQ ID NO: 590 (VH CDR3); SEQ ID NO: 591 (VL); SEQ ID NO: 592 (VL CDR1); SEQ ID NO: 593 (VL CDR2); SEQ ID NO: 594 (VL CDR3)
(b) Base Sequence
[1039]SEQ ID NO: 877 (VH); SEQ ID NO: 878 (VL)
[1040]075-110 antibody
(a) Amino Acid Sequence
[1041]SEQ ID NO: 595 (VH); SEQ ID NO: 596 (VH CDR1); SEQ ID NO: 597 (VH CDR2); SEQ ID NO: 598 (VH CDR3); SEQ ID NO: 599 (VL); SEQ ID NO: 600 (VL CDR1); SEQ ID NO: 601 (VL CDR2); SEQ ID NO: 602 (VL CDR3)
(b) Base Sequence
[1042]SEQ ID NO: 879 (VH); SEQ ID NO: 880 (VL)
[1043]086-032 antibody
(a) Amino Acid Sequence
[1044]SEQ ID NO: 603 (VH); SEQ ID NO: 604 (VH CDR1); SEQ ID NO: 605 (VH CDR2); SEQ ID NO: 606 (VH CDR3); SEQ ID NO: 607 (VL); SEQ ID NO: 608 (VL CDR1); SEQ ID NO: 609 (VL CDR2); SEQ ID NO: 610 (VL CDR3)
(b) Base Sequence
[1045]SEQ ID NO: 881 (VH); SEQ ID NO: 882 (VL)
[1046]086-035 antibody
(a) Amino Acid Sequence
[1047]SEQ ID NO: 611 (VH); SEQ ID NO: 612 (VH CDR1); SEQ ID NO: 613 (VH CDR2); SEQ ID NO: 614 (VH CDR3); SEQ ID NO: 615 (VL); SEQ ID NO: 616 (VL CDR1); SEQ ID NO: 617 (VL CDR2); SEQ ID NO: 618 (VL CDR3)
(b) Base Sequence
[1048]SEQ ID NO: 883 (VH); SEQ ID NO: 884 (VL)
[1049]086-036 antibody
(a) Amino Acid Sequence
[1050]SEQ ID NO: 619 (VH); SEQ ID NO: 620 (VH CDR1); SEQ ID NO: 621 (VH CDR2); SEQ ID NO: 622 (VH CDR3); SEQ ID NO: 623 (VL); SEQ ID NO: 624 (VL CDR1); SEQ ID NO: 625 (VL CDR2); SEQ ID NO: 626 (VL CDR3)
(b) Base Sequence
[1051]SEQ ID NO: 885 (VH); SEQ ID NO: 886 (VL)
[1052]086-061 antibody
(a) Amino Acid Sequence
[1053]SEQ ID NO: 627 (VH); SEQ ID NO: 628 (VH CDR1); SEQ ID NO: 629 (VH CDR2); SEQ ID NO: 630 (VH CDR3); SEQ ID NO: 631 (VL); SEQ ID NO: 632 (VL CDR1); SEQ ID NO: 633 (VL CDR2); SEQ ID NO: 634 (VL CDR3)
(b) Base Sequence
[1054]SEQ ID NO: 887 (VH); SEQ ID NO: 888 (VL)
[1055]086-138 antibody
(a) Amino Acid Sequence
[1056]SEQ ID NO: 635 (VH); SEQ ID NO: 636 (VH CDR1); SEQ ID NO: 637 (VH CDR2); SEQ ID NO: 638 (VH CDR3); SEQ ID NO: 639 (VL); SEQ ID NO: 640 (VL CDR1); SEQ ID NO: 641 (VL CDR2); SEQ ID NO: 642 (VL CDR3)
(b) Base Sequence
[1057]SEQ ID NO: 889 (VH); SEQ ID NO: 890 (VL)
[1058]086-182 antibody
(a) Amino Acid Sequence
[1059]SEQ ID NO: 643 (VH); SEQ ID NO: 644 (VH CDR1); SEQ ID NO: 645 (VH CDR2); SEQ ID NO: 646 (VH CDR3); SEQ ID NO: 647 (VL); SEQ ID NO: 648 (VL CDR1); SEQ ID NO: 649 (VL CDR2); SEQ ID NO: 650 (VL CDR3,
(b) Base Sequence
[1060]SEQ ID NO: 891 (VH); SEQ ID NO: 892 (VL)
(7) Antibody to HGFR 067-126 antibody
(a) Amino Acid Sequence
[1061]SEQ ID NO: 651 (VH); SEQ ID NO: 652 (VH CDR1); SEQ ID NO: 653 (VH CDR2); SEQ ID NO: 654 (VH CDR3); SEQ ID NO: 655 (VL); SEQ ID NO: 656 (VL CDR1); SEQ ID NO: 657 (VL CDR2); SEQ ID NO: 658 (VL CDR3)
(b) Base Sequence
[1062]SEQ ID NO: 893 (VH); SEQ ID NO: 894 (VL)
[1063]067-133 antibody
(a) Amino Acid Sequence
[1064]SEQ ID NO: 659 (VH); SEQ ID NO: 660 (VH CDR1); SEQ ID NO: 661 (VH CDR2); SEQ ID NO: 662 (VH CDR3); SEQ ID NO: 663 (VL); SEQ ID NO: 664 (VL CDR1); SEQ ID NO: 665 (VL CDR2); SEQ ID NO: 666 (VL CDR3)
(b) Base Sequence
[1065]SEQ ID NO: 895 (VH); SEQ ID NO: 896 (VL)
[1066]067-287 antibody
(a) Amino Acid Sequence
[1067]SEQ ID NO: 667 (VH); SEQ ID NO: 668 (VH CDR1); SEQ ID NO: 669 (VH CDR2); SEQ ID NO: 670 (VH CDR3); SEQ ID NO: 671 (VL); SEQ ID NO: 672 (VL CDR1); SEQ ID NO: 673 (VL CDR2); SEQ ID NO: 674 (VL CDR3)
(b) Base Sequence
[1068]SEQ ID NO: 897 (VH); SEQ ID NO: 898 (VL)
(8) Antibody to ITGA3
[1069]064-002 antibody
(a) Amino Acid Sequence
[1070]SEQ ID NO: 675 (VH); SEQ ID NO: 676 (VH CDR1); SEQ ID NO: 677 (VH CDR2); SEQ ID NO: 678 (VH CDR3); SEQ ID NO: 679 (VL); SEQ ID NO: 680 (VL CDR1); SEQ ID NO: 681 (VL CDR2); SEQ ID NO: 682 (VL CDR3)
(b) Base Sequence
[1071]SEQ ID NO: 899 (VH); SEQ ID NO: 900 (VL)
[1072]064-006 antibody
(a) Amino Acid Sequence
[1073]SEQ ID NO: 683 (VH); SEQ ID NO: 684 (VH CDR1); SEQ ID NO: 685 (VH CDR2); SEQ ID NO: 686 (VH CDR3); SEQ ID NO: 687 (VL); SEQ ID NO: 688 (VL CDR1); SEQ ID NO: 689 (VL CDR2); SEQ ID NO: 690 (VL CDR3)
(b) Base Sequence
[1074]SEQ ID NO: 901 (VH); SEQ ID NO: 902 (VL)
[1075]064-012a antibody
(a) Amino Acid Sequence
[1076]SEQ ID NO: 691 (VH); SEQ ID NO: 692 (VH CDR1); SEQ ID NO: 693 (VH CDR2); SEQ ID NO: 694 (VH CDR3); SEQ ID NO: 695 (VL); SEQ ID NO: 696 (VL CDR1); SEQ ID NO: 697 (VL CDR2); SEQ ID NO: 698 (VL CDR3)
(b) Base Sequence
[1077]SEQ ID NO: 903 (VH); SEQ ID NO: 904 (VL)
[1078]064-012b antibody
(a) Amino Acid Sequence
[1079]SEQ ID NO: 699 (VH); SEQ ID NO: 700 (VH CDR1); SEQ ID NO: 701 (VH CDR2); SEQ ID NO: 702 (VH CDR3); SEQ ID NO: 703 (VL); SEQ ID NO: 704 (VL CDR1); SEQ ID NO: 705 (VL CDR2); SEQ ID NO: 706 (VL CDR3)
(b) Base Sequence
[1080]SEQ ID NO: 905 (VH); SEQ ID NO: 906 (VL)
[1081]064-014 antibody
(a) Amino Acid Sequence
[1082]SEQ ID NO: 707 (VH); SEQ ID NO: 708 (VH CDR1); SEQ ID NO: 709 (VH CDR2); SEQ ID NO: 710 (VH CDR3); SEQ ID NO: 711 (VL); SEQ ID NO: 712 (VL CDR1); SEQ ID NO: 713 (VL CDR2); SEQ ID NO: 714 (VL CDR3)
(b) Base Sequence
[1083]SEQ ID NO: 907 (VH); SEQ ID NO: 908 (VL)
[1084]064-054 antibody
(a) Amino Acid Sequence
[1085]SEQ ID NO: 715 (VH); SEQ ID NO: 716 (VH CDR1); SEQ ID NO: 717 (VH CDR2); SEQ ID NO: 718 (VH CDR3); SEQ ID NO: 719 (VL); SEQ ID NO: 720 (VL CDR1); SEQ ID NO: 721 (VL CDR2); SEQ ID NO: 722 (VL CDR3)
(b) Base Sequence
[1086]SEQ ID NO: 909 (VH); SEQ ID NO: 910 (VL)
[1087]064-085 antibody
(a) Amino Acid Sequence
[1088]SEQ ID NO: 723 (VH); SEQ ID NO: 724 (VH CDR1); SEQ ID NO: 725 (VH CDR2); SEQ ID NO: 726 (VH CDR3); SEQ ID NO: 727 (VL); SEQ ID NO: 728 (VL CDR1); SEQ ID NO: 729 (VL CDR2); SEQ ID NO: 730 (VL CDR3)
(b) Base Sequence
[1089]SEQ ID NO: 911 (VH); SEQ ID NO: 912 (VL)
[1090]064-093 antibody
(a) Amino Acid Sequence
[1091]SEQ ID NO: 731 (VH); SEQ ID NO: 732 (VH CDR1); SEQ ID NO: 733 (VH CDR2); SEQ ID NO: 734 (VH CDR3); SEQ ID NO: 735 (VL); SEQ ID NO: 736 (VL CDR1); SEQ ID NO: 737 (VL CDR2); SEQ ID NO: 738 (VL CDR3)
(b) Base Sequence
[1092]SEQ ID NO: 913 (VH); SEQ ID NO: 914 (VL)
[1093]064-116 antibody
(a) Amino Acid Sequence
[1094]SEQ ID NO: 739 (VH); SEQ ID NO: 740 (VH CDR1); SEQ ID NO: 741 (VH CDR2); SEQ ID NO: 742 (VH CDR3); SEQ ID NO: 743 (VL); SEQ ID NO: 744 (VL CDR1); SEQ ID NO: 745 (VL CDR2); SEQ ID NO: 746 (VL CDR3)
(b) Base Sequence
[1095]SEQ ID NO: 915 (VH); SEQ ID NO: 916 (VL)
[1096]065-183 antibody
(a) Amino Acid Sequence
[1097]SEQ ID NO: 747 (VH); SEQ ID NO: 748 (VH CDR1); SEQ ID NO: 749 (VH CDR2); SEQ ID NO: 750 (VH CDR3); SEQ ID NO: 751 (VL); SEQ ID NO: 752 (VL CDR1); SEQ ID NO: 753 (VL CDR2); SEQ ID NO: 754 (VL CDR3)
(b) Base Sequence
[1098]SEQ ID NO: 917 (VH); SEQ ID NO: 918 (VL)
[1099]067-142 antibody
(a) Amino Acid Sequence
[1100]SEQ ID NO: 763 (VH); SEQ ID NO: 764 (VH CDR1); SEQ ID NO: 765 (VH CDR2); SEQ ID NO: 766 (VH CDR3); SEQ ID NO: 767 (VL); SEQ ID NO: 768 (VL CDR1); SEQ ID NO: 769 (VL CDR2); SEQ ID NO: 770 (VL CDR3)
(b) Base Sequence
[1101]SEQ ID NO: 921 (VH); SEQ ID NO: 922 (VL)
[1102]068-007 antibody
(a) Amino Acid Sequence
[1103]SEQ ID NO: 771 (VH); SEQ ID NO: 772 (VH CDR1); SEQ ID NO: 773 (VH CDR2); SEQ ID NO: 774 (VH CDR3); SEQ ID NO: 775 (VL); SEQ ID NO: 776 (VL CDR1); SEQ ID NO: 777 (VL CDR2); SEQ ID NO: 778 (VL CDR3)
(b) Base Sequence
[1104]SEQ ID NO: 923 (VH); SEQ ID NO: 924 (VL)
(9) Antibody to ALCAM 029-143 antibody
(a) Amino Acid Sequence
[1105]SEQ ID NO: 779 (VH); SEQ ID NO: 780 (VH CDR1); SEQ ID NO: 781 (VH CDR2); SEQ ID NO 782 (VH CDR3); SEQ ID NO: 783 (VL); SEQ ID NO: 784 (VL CDR1); SEQ ID NO: 785 (VL CDR2); SEQ ID NO: 786 (VL CDR3)
(b) Base Sequence
[1106]SEQ ID NO: 925 (VH); SEQ ID NO: 926 (VL)
[1107]045-1.34 antibody
(a) Amino Acid Sequence
[1108]SEQ ID NO: 787 (VH); SEQ ID NO: 788 (VH CDR1); SEQ ID NO: 789 (VH CDR2); SEQ ID NO: 790 (VH CDR3); SEQ ID NO: 791 (VL); SEQ ID NO: 792 (VL CDR1); SEQ ID NO: 793 (VL CDR2); SEQ ID NO: 794 (VL CDR3)
(b) Base Sequence
[1109]SEQ ID NO: 927 (VH); SEQ ID NO: 928 (VL)
[1110]062-101 antibody
(a) Amino Acid Sequence
[1111]SEQ ID NO: 795 (VH); SEQ ID NO: 796 (VH CDR1); SEQ ID NO: 797 (VH CDR2); SEQ ID NO: 798 (VH CDR3); SEQ ID NO: 799 (VL); SEQ ID NO: 800 (VL CDR1); SEQ ID NO: 801 (VL CDR2); SEQ ID NO: 802 (VL CDR3)
(b) Base Sequence
[1112]SEQ ID NO: 929 (VH); SEQ ID NO: 930 (VL)
[1113]062-109 antibody
(a) Amino Acid Sequence
[1114]SEQ ID NO: 803 (VH); SEQ ID NO: 804 (VH CDR1); SEQ ID NO: 805 (VH CDR2); SEQ ID NO: 806 (VH CDR3); SEQ ID NO: 807 (VL); SEQ ID NO: 808 (VL CDR1); SEQ ID NO: 809 (VL CDR2); SEQ ID NO: 810 (VL CDR3)
(b) Base Sequence
[1115]SEQ ID NO: 931 (VH); SEQ ID NO: 932 (VL)
[1116]084-103 antibody
(a) Amino Acid Sequence
[1117]SEQ ID NO: 811 (VH); SEQ ID NO: 812 (VH CDR1); SEQ ID NO: 813 (VH CDR2); SEQ ID NO: 814 (VH CDR3); SEQ ID NO: 815 (VL); SEQ ID NO: 816 (VL CDR1); SEQ ID NO: 817 (VL CDR2); SEQ ID NO: 818 (VL CDR3)
(b) Base Sequence
[1118]SEQ ID NO: 933 (VH); SEQ ID NO: 934 (VL)
[1119]052-274 antibody
(a) Amino Acid Sequence
[1120]SEQ ID NO: 819 (VH); SEQ ID NO: 820 (VH CDR1); SEQ ID NO: 821 (VH CDR2); SEQ ID NO: 822 (VH CDR3); SEQ ID NO: 823 (VL); SEQ ID NO: 824 (VL CDR1); SEQ ID NO: 825 (VL CDR2); SEQ ID NO: 826 (VL CDR3)
(b) Base Sequence
[1121]SEQ ID NO: 935 (VH); SEQ ID NO: 936 (VL)
[1122]029-067 antibody
(a) Amino Acid Sequence
[1123]SEQ ID NO: 827 (VH); SEQ ID NO: 828 (VH CDR1); SEQ ID NO: 829 (VH CDR2); SEQ ID NO: 830 (VH CDR3); SEQ ID NO: 831 (VL); SEQ ID NO: 832 (VL CDR1); SEQ ID NO: 833 (VL CDR2); SEQ ID NO: 834 (VL CDR3)
(b) Base Sequence
[1124]SEQ ID NO: 937 (VH); SEQ ID NO: 938 (VL)
[1125]083-131 antibody
(a) Amino Acid Sequence
[1126]SEQ ID NO: 835 (VH); SEQ ID NO: 836 (VH CDR1); SEQ ID NO: 837 (VH CDR2); SEQ ID NO: 838 (VH CDR3); SEQ ID NO: 839 (VL); SEQ ID NO: 840 (VL CDR1); SEQ ID NO: 841 (VL CDR2); SEQ ID NO: 842 (VL CDR3)
(b) Base Sequence
[1127]SEQ ID NO: 939 (VH); SEQ ID NO: 940 (VL)
(10) Antibody to CD46
[1128]066-069 antibody
(a) Amino Acid Sequence
[1129]SEQ ID NO: 755 (VH); SEQ ID NO: 756 (VH CDR1); SEQ ID NO: 757 (VH CDR2); SEQ ID NO: 758 (VH CDR3); SEQ ID NO: 759 (VL); SEQ ID NO: 760 (VL CDR1); SEQ ID NO: 761 (VL CDR2); SEQ ID NO: 762 (VL CDR3)
(b) Base Sequence
[1130]SEQ ID NO: 919 (VH); SEQ ID NO: 920 (VL)
(11) Antibody to LAR 064-044 antibody
(a) Amino Acid Sequence
[1131]SEQ ID NO: 944 (VH); SEQ ID NO: 945 (VL)
(b) Base Sequence
[1132]SEQ ID NO: 956 (VH); SEQ ID NO: 957 (VL)
[1133]065-030 antibody
(a) Amino Acid Sequence
[1134]SEQ ID NO: 946 (VH); SEQ ID NO: 947 (VL)
(b) Base Sequence
[1135]SEQ ID NO: 958 (VH); SEQ ID NO: 959 (VL)
[1136]065-358 antibody
(a) Amino Acid Sequence
[1137]SEQ ID NO: 948 (VH); SEQ ID NO: 949 (VL)
(b) Base Sequence
[1138]SEQ ID NO: 960 (VH); SEQ ID NO: 961 (VL)
[1139]066-019 antibody
(a) Amino Acid Sequence
[1140]SEQ ID NO: 950 (VH); SEQ ID NO: 951 (VL)
(b) Base Sequence
[1141]SEQ ID NO: 962 (VH); SEQ ID NO: 963 (VL)
[1142]079-085 antibody
(a) Amino Acid Sequence
[1143]SEQ ID NO: 952 (VH); SEQ ID NO: 953 (VL)
(b) Base Sequence
[1144]SEQ ID NO: 964 (VH); SEQ ID NO: 965 (VL)
(12) Antibody to BCAM 067-024 antibody
(a) Amino Acid Sequence
[1145]SEQ ID NO: 954 (VH); SEQ ID NO: 955 (VL)
(b) Base Sequence
[1146]SEQ ID NO: 966 (VH); SEQ ID NO: 967 (VL)
(13) Antibody to IgSF4
[1147]076-048 antibody
(a) Amino Acid Sequence
[1148]SEQ ID NO: 968 (VH); SEQ ID NO: 969 (VL)
(b) Base Sequence
[1149]SEQ ID NO: 970 (VH); SEQ ID NO: 971 (VL)
21. Experiment to Confirm ITGA3 Antibody
[1150]From the results of the immunoprecipitation-mass spectrometry, a part of the antibody group, it was shown that the antibody included therein recognized a VLA complex. However, in a strict sense, it was not possible to determine what the antibody was, that is, whether the antigen was ITGA3 or ITGB1 or other molecules forming a complex such as CD151. Then, the antibody clones (015-003, 064-002, 064-006, 064-012, 064-014, 064-054, 064-085, 064-091, 064-093, 064-116, 065-183, 067-142, and 068-007) were subjected to RNAI in order to confirm antigens.
21-1 Experiment Procedure
[1151]ITGA3 stealth oligo RNA (400 pmol) purchased from Invitrogen and lipofect RNAi MAX (100 μl) (product of Invitrogen) were mixed with Opti-MEMI (8 ml) (product of GIBCO-BRL) and the mixture was stood still at a room temperature for 10 minutes. To this mixture, 4 ml of SKOv-3 cell solution (2×106 cells) and 28 ml of RPMI1640-10% FBS were added. This mixture was planted on four 10-cm culture dishes and cultured in a CO2 incubator for two days. 1% trypsin solution was allowed to act on the cultured cells so as to liberate cells. The cells were recovered in 5% BSA/PBS solution so as to produce 1 ml of cell suspension. The same experiment was carried out with respect to ITGB1. As to a group without RNAi (control group), the same experiment was carried out except that stealth oligo is not allowed to act.
[1152]To the recovered cells (50 μl), 2.5 μl of normal goat serum was added, and then primary antibody solution was added, so that the final amount was made to be 100 μl by using 5% BSA/PBS. The using amount of the primary antibody (anti-ITGA3 antibody or anti-ITGB1 antibody (mouse monoclonal antibody, product of CHEMICON)) was made to be 1 μl. As to the subjected sample (for example, 015-003 cp3 type antibody), 7 μl of 10-fold concentrated supernatant was used.
[1153]Next, the mixture was stood still at a room temperature for 10 minutes and then subjected to centrifugation. The supernatant was discarded, followed by washing with 5% BSA/PBS (200 μl). Next, as to the sample 015-003 cp3 type antibody, 100 μl of anti-cp3 mouse monoclonal antibody (MBL), which had been diluted with 5% BSA/PBS so that the concentration became 5 μg/ml, was added. The mixture was stood still at a room temperature for 10 minutes. After centrifugation, the supernatant was discarded, followed by washing with 5% BSA/PBS (200 μl). Then, ALEXA488 labeled anti-mouse IgG goat antibody (100 μl), which had been 1000-folded diluted with 5% BSA/PBS, was reacted. The reacted product was stood still at a room temperature for 10 minutes and then subjected to centrifugation. The supernatant was discarded, followed by washing with 5% BSA/PBS (200 μl). The thus obtained cells were suspended in 50 μl of OptilyseB (BECKMAN COULTER). This was stood still for 10 minutes, and then 600 μl of PBS was added to be diluted. Subsequently, the diluted product was treated with Cell-Strainer (BD Falcon) and subjected to measurement using FACS Caliber (BECKMAN COULTER).
21-2 Results
[1154]The results of the above-mentioned RNAi experiment are shown in FIG. 86. It is shown that A (results of FCM using anti-ITGA3 antibody) and B (results of FCM using anti-ITGB1 antibody) have different peak patterns. The samples (015-003, 064-002, 064-006, 064-012, 064-014, 064-054, 064-085, 064-091, 064-093, 064-116, 065-183, 067-142, and 068-007) show the peak patterns (C) similar to A. From this result, it is confirmed that antigen recognized by these antibody clones is ITGA3.
[1155]When the same RNAi experiment is carried out in each antibody obtained as an anti-HER1 antibody, an anti-HER2 antibody, an anti-HGFR antibody, an anti-IgSF4 antibody, an anti-EpCAM antibody, an anti-CD147 antibody, an anti-CD166 antibody, or anti-MCP antibody, antigen is not wrong, and it is confirmed that the method (method using a panel, three-dimensional ELISA)) of the present invention is useful.
22. Cancer Tissue Specificity of Each Antibody Clone
[1156]When the immunostaining property of the obtained antibody clones with respect to clinical cancer specimens were examined by the same method as described in the above column 11, results shown in FIG. 87 were obtained. These antibody clones are useful for studying and diagnosing the corresponding cancers. Furthermore, clinical specimens in different stages in some cancers were prepared and the immunostaining property of the antibody clones with respect to the specimens was obtained. As a result, some antibody clones showed the staining property specific to stages in addition to the staining property specific to cancer (see FIG. 88). Thus, in the actual clinical tissues, there are differences in the reactivity to each antibody clone even if the tissue is from the same cancer or in the same grade of malignancy. This results show that the use of the antibody set provided by the present invention enables new tailor-maid diagnosis in cancers to be carried out and diagnosis that is more detail than conventional criterion to be carried out. In other words, it is shown that staging of cancer and re-classification of pathologic conditions can be realized. On the other hand, the staging of cancer and the re-classification of pathologic conditions by using the antibody set are useful for determining a treatment plan. Furthermore, antibodies recognized to have specific reactivity can be useful as antibodies for treatment and useful as a tool for drug screening. Thus, the antibody set provided by the present invention can realize not only tailor-made diagnosis of cancers but also tailor-made treatment of cancers. Thus, the antibody set provides extremely great values and significance.
INDUSTRIAL APPLICABILITY
[1157]The present invention provides a method of classifying a plurality of antibodies to cell surface antigens rapidly. Also, the present invention provides a method of rapidly identifying an antigen to an antibody. The use of these methods makes it possible to obtain an antibody useful for treatment and diagnosis of cancers, or study of the onset mechanism of cancers, and the like. Furthermore, when the classifying method and the identification method of an antigen of the present invention are used, a panel on which a useful antibody set and its characteristics are displayed can be provided, which is expected to greatly contribute to tailor-made medicine. On the other hand, the present invention provides antibodies recognizing antigens expressing in a cancer-specific manner. Such antibodies are expected to be used as antibody for treatment, antibody for diagnosis, antibody for study, and the like, which target cancer cells specifically expressing cancer surface membrane protein recognized by the antibodies.
[1158]The present invention is not limited only to the description of the above embodiments. A variety of modifications which are within the scopes of the following claims and which are achieved easily by a person skilled in the art are included in the present invention.
[1159]Contents of the theses, Publication of Patent Applications, Patent Publications, and other published documents referred to in this specification are herein incorporated by reference in its entity.
Sequence CWU
1
9711120PRTHomo sapiens 1Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val
Lys Pro Ser Gln1 5 10
15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Gly Ser Ile Ser Ser Gly
20 25 30Gly Tyr Tyr Trp Ser Trp Ile
Arg Gln His Pro Gly Lys Gly Leu Glu 35 40
45Trp Ile Gly Tyr Ile Tyr Tyr Ser Gly Ser Thr Tyr Tyr Asn Pro
Ser 50 55 60Leu Lys Ser Arg Val Thr
Ile Ser Val Asp Thr Ser Lys Asn Gln Phe65 70
75 80Ser Leu Lys Leu Ser Ser Val Thr Ala Ala Asp
Thr Ala Val Tyr Tyr 85 90
95Cys Ala Arg Thr Pro Trp Glu Leu Leu Ala Phe Asp Ile Trp Gly Gln
100 105 110Gly Thr Met Val Thr Val
Ser Arg 115 12027PRTHomo sapiens 2Ser Gly Gly Tyr
Tyr Trp Ser1 5316PRTHomo sapiens 3Tyr Ile Tyr Tyr Ser Gly
Ser Thr Tyr Tyr Asn Pro Ser Leu Lys Ser1 5
10 15410PRTHomo sapiens 4Thr Pro Trp Glu Leu Leu Ala
Phe Asp Ile1 5 105109PRTHomo sapiens 5Ser
Tyr Glu Leu Thr Gln Pro Pro Ser Val Ser Val Ala Pro Gly Lys1
5 10 15Thr Thr Arg Ile Thr Cys Gly
Gly Asn Asn Ile Gly Ser Lys Ser Ala 20 25
30His Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Val Leu Val
Ile Tyr 35 40 45Tyr Asp Ser Asp
Arg Pro Ser Gly Ile Pro Glu Arg Phe Ser Gly Ser 50 55
60Asn Ser Gly Asn Thr Ala Thr Leu Thr Ile Ser Arg Val
Glu Ala Gly65 70 75
80Asp Glu Ala Asp Tyr Tyr Cys Gln Val Trp Asp Ser Ser Ser Asp His
85 90 95Trp Val Phe Gly Gly Gly
Thr Lys Leu Thr Val Leu Gly 100 105611PRTHomo
sapiens 6Gly Gly Asn Asn Ile Gly Ser Lys Ser Ala His1 5
1077PRTHomo sapiens 7Tyr Asp Ser Asp Arg Pro Ser1
589PRTHomo sapiens 8Gln Val Trp Asp Ser Ser Ser Asp His1
59126PRTHomo sapiens 9Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys
Lys Thr Gly Ser1 5 10
15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Ser Phe Ser Ser Ser
20 25 30Ala Ile Ser Trp Val Arg Gln
Ala Pro Gly His Gly Leu Glu Trp Leu 35 40
45Gly Gly Ile Ile Pro Thr Phe Gly Thr Pro Asn His Ala Gln Lys
Phe 50 55 60Gln Gly Arg Val Thr Ile
Thr Ala Asp Glu Ser Thr Gly Thr Ala Tyr65 70
75 80Met Glu Leu Ser Gly Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90
95Ala Arg Ala His Cys Gly Gly Gly Arg Cys Tyr Asp Tyr Thr Asp Ala
100 105 110Phe His Phe Trp Gly Gln
Gly Thr Met Val Thr Val Ser Arg 115 120
125105PRTHomo sapiens 10Ser Ser Ala Ile Ser1
51117PRTHomo sapiens 11Gly Ile Ile Pro Thr Phe Gly Thr Pro Asn His Ala
Gln Lys Phe Gln1 5 10
15Gly1217PRTHomo sapiens 12Ala His Cys Gly Gly Gly Arg Cys Tyr Asp Tyr
Thr Asp Ala Phe His1 5 10
15Phe13109PRTHomo sapiens 13Ser Tyr Glu Leu Thr Gln Pro Pro Ser Val Ser
Val Ala Pro Gly Lys1 5 10
15Thr Ala Arg Ile Thr Cys Gly Gly Asp Asn Ile Gly Asn Arg Ser Val
20 25 30His Trp Tyr Gln Gln Lys Pro
Gly Gln Ala Pro Val Leu Leu Ile Tyr 35 40
45Tyr Asp Ser Asp Arg Pro Ser Gly Ile Pro Lys Arg Phe Ser Gly
Ser 50 55 60Asn Ser Gly Asn Thr Ala
Thr Leu Thr Ile Ser Arg Val Glu Ala Gly65 70
75 80Asp Glu Ala Asp Tyr Tyr Cys Gln Val Trp Asp
Ser Thr Ser Asp His 85 90
95Val Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Arg 100
1051411PRTHomo sapiens 14Gly Gly Asp Asn Ile Gly Asn Arg Ser
Val His1 5 10157PRTHomo sapiens 15Tyr Asp
Ser Asp Arg Pro Ser1 5169PRTHomo sapiens 16Gln Val Trp Asp
Ser Thr Ser Asp His1 517120PRTHomo sapiens 17Gln Val Gln
Leu Gln Glu Ser Gly Pro Gly Leu Val Lys Pro Ser Glu1 5
10 15Thr Leu Ser Leu Thr Cys Thr Val Ser
Gly Gly Ser Ile Ser Ser Ser 20 25
30Ser Tyr Tyr Trp Gly Trp Ile Arg Gln Pro Pro Gly Lys Gly Leu Glu
35 40 45Trp Ile Gly Ser Ile Tyr Tyr
Ser Gly Ser Thr Tyr Tyr Asn Pro Ser 50 55
60Leu Lys Ser Arg Val Thr Ile Ser Val Asp Thr Ser Lys Asn Gln Phe65
70 75 80Ser Leu Lys Leu
Ser Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr 85
90 95Cys Ala Arg Leu Pro Met Val Thr Met Ser
Phe Asp Tyr Trp Gly Gln 100 105
110Gly Thr Leu Val Thr Val Ser Arg 115
120187PRTHomo sapiens 18Ser Ser Ser Tyr Tyr Trp Gly1
51916PRTHomo sapiens 19Ser Ile Tyr Tyr Ser Gly Ser Thr Tyr Tyr Asn Pro
Ser Leu Lys Ser1 5 10
152010PRTHomo sapiens 20Leu Pro Met Val Thr Met Ser Phe Asp Tyr1
5 1021109PRTHomo sapiens 21Ser Tyr Val Leu Thr Gln
Pro Pro Ser Val Ser Val Ala Pro Gly Lys1 5
10 15Thr Ala Arg Ile Thr Cys Gly Gly Asn Asn Ile Gly
Ser Lys Ser Val 20 25 30His
Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Val Leu Val Ile Tyr 35
40 45Tyr Asp Ser Asp Arg Pro Ser Gly Ile
Pro Glu Arg Phe Ser Gly Ser 50 55
60Asn Ser Gly Asn Thr Ala Thr Leu Thr Ile Ser Arg Val Glu Ala Gly65
70 75 80Asp Glu Ala Asp Tyr
Tyr Cys Gln Val Trp Asp Ser Ser Ser Asp His 85
90 95Val Val Phe Gly Gly Gly Thr Lys Leu Thr Val
Leu Gly 100 1052211PRTHomo sapiens 22Gly Gly
Asn Asn Ile Gly Ser Lys Ser Val His1 5
10237PRTHomo sapiens 23Tyr Asp Ser Asp Arg Pro Ser1
5249PRTHomo sapiens 24Gln Val Trp Asp Ser Ser Ser Asp His1
525121PRTHomo sapiens 25Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys
Lys Pro Gly Glu1 5 10
15Ser Leu Lys Ile Ser Cys Lys Gly Ser Gly Tyr Ser Phe Thr Ser Tyr
20 25 30Trp Ile Gly Trp Val Arg Gln
Met Pro Gly Lys Gly Leu Glu Trp Met 35 40
45Gly Ile Ile Tyr Pro Gly Asp Ser Asp Thr Arg Tyr Ser Pro Ser
Phe 50 55 60Gln Gly Gln Val Thr Ile
Ser Ala Asp Lys Ser Ile Ser Thr Ala Tyr65 70
75 80Leu Gln Trp Ser Ser Leu Lys Ala Ser Asp Thr
Ala Met Tyr Tyr Cys 85 90
95Ala Arg Leu Thr Leu Ser Tyr Ser Ser Ser Trp Phe Asp Tyr Trp Gly
100 105 110Gln Gly Thr Leu Val Thr
Val Ser Arg 115 120265PRTHomo sapiens 26Ser Tyr
Trp Ile Gly1 52717PRTHomo sapiens 27Ile Ile Tyr Pro Gly Asp
Ser Asp Thr Arg Tyr Ser Pro Ser Phe Gln1 5
10 15Gly2812PRTHomo sapiens 28Leu Thr Leu Ser Tyr Ser
Ser Ser Trp Phe Asp Tyr1 5 1029111PRTHomo
sapiens 29Gln Thr Val Val Thr Gln Glu Pro Ser Phe Ser Val Ser Pro Gly
Gly1 5 10 15Thr Val Thr
Leu Thr Cys Gly Leu Ser Ser Gly Ser Val Ser Thr Ser 20
25 30Tyr Tyr Pro Ser Trp Tyr Gln Gln Thr Pro
Gly Gln Ala Pro Arg Thr 35 40
45Leu Ile Tyr Ser Thr Asn Thr Arg Ser Ser Gly Val Pro Asp Arg Phe 50
55 60Ser Gly Ser Ile Leu Gly Asn Lys Ala
Ala Leu Thr Ile Thr Gly Ala65 70 75
80Gln Ala Asp Asp Glu Ser Asp Tyr Tyr Cys Val Leu Tyr Met
Gly Ser 85 90 95Gly Ile
Ser Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly 100
105 1103014PRTHomo sapiens 30Gly Leu Ser Ser Gly
Ser Val Ser Thr Ser Tyr Tyr Pro Ser1 5
10317PRTHomo sapiens 31Ser Thr Asn Thr Arg Ser Ser1
5328PRTHomo sapiens 32Val Leu Tyr Met Gly Ser Gly Ile1
533118PRTHomo sapiens 33Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys
Lys Pro Gly Glu1 5 10
15Ser Leu Lys Ile Ser Cys Lys Gly Ser Gly Tyr Ser Phe Thr Ser Tyr
20 25 30Trp Ile Gly Trp Val Arg Gln
Met Pro Gly Lys Gly Leu Glu Trp Met 35 40
45Gly Ile Ile Tyr Pro Gly Asp Ser Asp Thr Arg Tyr Ser Pro Ser
Phe 50 55 60Gln Gly Gln Val Thr Ile
Ser Ala Asp Lys Ser Ile Ser Thr Ala Tyr65 70
75 80Leu Gln Trp Ser Ser Leu Lys Ala Ser Asp Thr
Ala Met Tyr Tyr Cys 85 90
95Ala Arg Leu Leu Gly Ile Gly Ala Phe Asp Ile Trp Gly Gln Gly Thr
100 105 110Thr Val Thr Val Ser Arg
115345PRTHomo sapiens 34Ser Tyr Trp Ile Gly1
53517PRTHomo sapiens 35Ile Ile Tyr Pro Gly Asp Ser Asp Thr Arg Tyr Ser
Pro Ser Phe Gln1 5 10
15Gly369PRTHomo sapiens 36Leu Leu Gly Ile Gly Ala Phe Asp Ile1
537110PRTHomo sapiens 37Ser Ser Glu Leu Thr Gln Asp Pro Ala Val Ser
Val Ala Leu Gly Gln1 5 10
15Thr Val Arg Ile Thr Cys Gln Gly Asp Ser Leu Arg Ser Tyr Tyr Ala
20 25 30Ser Trp Tyr Gln Gln Lys Pro
Gly Gln Ala Pro Val Leu Val Ile Tyr 35 40
45Gly Lys Asn Asn Arg Pro Ser Gly Ile Pro Asp Arg Phe Ser Gly
Ser 50 55 60Ser Ser Gly Asn Thr Ala
Ser Leu Thr Ile Thr Gly Ala Gln Ala Glu65 70
75 80Asp Glu Ala Asp Tyr Tyr Cys Asn Ser Arg Asp
Ser Ser Gly Asn His 85 90
95His Tyr Val Phe Gly Thr Gly Thr Lys Val Thr Val Leu Gly 100
105 1103811PRTHomo sapiens 38Gln Gly Asp
Ser Leu Arg Ser Tyr Tyr Ala Ser1 5
10397PRTHomo sapiens 39Gly Lys Asn Asn Arg Pro Ser1
54010PRTHomo sapiens 40Asn Ser Arg Asp Ser Ser Gly Asn His His1
5 1041116PRTHomo sapiens 41Gln Val Gln Leu Val Gln
Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5
10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr
Phe Asn Ser Tyr 20 25 30Gly
Ile Thr Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45Gly Trp Ile Ser Val Tyr Thr Gly Lys
Thr Asn Tyr Ala Gln Lys Phe 50 55
60Gln Gly Arg Val Thr Met Thr Thr Asp Thr Ser Thr Ser Thr Ala Tyr65
70 75 80Leu Asp Leu Arg Ser
Leu Thr Ser Asp Asp Thr Ala Val Tyr Tyr Cys 85
90 95Ala Arg Gly Gly Asp His His Glu Tyr Trp Gly
Gln Gly Thr Leu Val 100 105
110Thr Val Ser Arg 115425PRTHomo sapiens 42Ser Tyr Gly Ile Thr1
54317PRTHomo sapiens 43Trp Ile Ser Val Tyr Thr Gly Lys Thr Asn
Tyr Ala Gln Lys Phe Gln1 5 10
15Gly447PRTHomo sapiens 44Gly Gly Asp His His Glu Tyr1
545107PRTHomo sapiens 45Asn Phe Met Leu Thr Gln Pro Leu Ser Val Ser Val
Ala Leu Gly Gln1 5 10
15Thr Ala Arg Ile Thr Cys Gly Gly Asn Asn Ile Gly Ser Lys Asn Val
20 25 30His Trp Tyr Gln Gln Lys Pro
Gly Gln Ala Pro Val Leu Val Ile Tyr 35 40
45Arg Asp Ser Asn Arg Pro Ser Gly Ile Pro Glu Arg Phe Ser Gly
Ser 50 55 60Asn Ser Gly Asn Thr Ala
Thr Leu Thr Ile Ser Arg Ala Gln Ala Gly65 70
75 80Asp Glu Ala Asp Tyr Tyr Cys Gln Val Trp Asp
Ser Ser Thr Val Val 85 90
95Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly 100
1054611PRTHomo sapiens 46Gly Gly Asn Asn Ile Gly Ser Lys Asn Val His1
5 10477PRTHomo sapiens 47Arg Asp Ser Asn
Arg Pro Ser1 5487PRTHomo sapiens 48Gln Val Trp Asp Ser Ser
Thr1 549128PRTHomo sapiens 49Glu Val Gln Leu Val Glu Ser
Gly Gly Gly Leu Val Gln Pro Gly Gly1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe
Ser Ser Tyr 20 25 30Ala Met
Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35
40 45Ser Gly Ile Ser Gly Ser Gly Gly Arg Thr
Tyr Tyr Ala Asp Ser Val 50 55 60Lys
Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr65
70 75 80Leu Gln Met Asn Ser Leu
Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys 85
90 95Ala Lys Asp Pro Leu Ala Leu Arg Asp Phe Asp Trp
Leu Ser Pro Gly 100 105 110Arg
Asp Phe Asp Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser Arg 115
120 125505PRTHomo sapiens 50Ser Tyr Ala Met
Ser1 55117PRTHomo sapiens 51Gly Ile Ser Gly Ser Gly Gly Arg
Thr Tyr Tyr Ala Asp Ser Val Lys1 5 10
15Gly5219PRTHomo sapiens 52Asp Pro Leu Ala Leu Arg Asp Phe
Asp Trp Leu Ser Pro Gly Arg Asp1 5 10
15Phe Asp Tyr53111PRTHomo sapiens 53Gln Ser Val Leu Thr Gln
Pro Pro Ser Val Ser Ala Ala Pro Gly Gln1 5
10 15Lys Val Thr Ile Ser Cys Ser Gly Ser His Ser Asn
Ile Gly Asn Asn 20 25 30Tyr
Val Ser Trp Ser Gln Gln Leu Pro Gly Thr Ala Pro Lys Leu Leu 35
40 45Ile Tyr Asp Asn Asn Lys Arg Pro Ser
Gly Ile Pro Asp Arg Phe Ser 50 55
60Gly Ser Lys Ser Gly Thr Ser Ala Thr Leu Asp Ile Ala Gly Leu Gln65
70 75 80Thr Gly Asp Glu Ala
Asp Tyr Tyr Cys Gly Ala Trp Asp Thr Ser Leu 85
90 95Ser Ser Tyr Val Phe Gly Ala Gly Thr Lys Val
Thr Val Leu Gly 100 105
1105413PRTHomo sapiens 54Ser Gly Ser His Ser Asn Ile Gly Asn Asn Tyr Val
Ser1 5 10557PRTHomo sapiens 55Asp Asn Asn
Lys Arg Pro Ser1 5569PRTHomo sapiens 56Gly Ala Trp Asp Thr
Ser Leu Ser Ser1 557120PRTHomo sapiens 57Glu Val Gln Leu
Val Glu Ser Gly Ala Glu Val Lys Lys Pro Gly Ser1 5
10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly
Gly Thr Phe Ser Ser Tyr 20 25
30Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met
35 40 45Gly Gly Ile Ile Pro Ile Phe Gly
Thr Ala Asn Tyr Ala Gln Lys Phe 50 55
60Gln Gly Arg Val Thr Ile Thr Ala Asp Glu Ser Thr Ser Thr Ala Tyr65
70 75 80Met Glu Leu Ser Ser
Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85
90 95Ala Arg Glu Gly Leu Ser Gly Gly Tyr Gly Met
Asp Val Trp Gly Gln 100 105
110Gly Thr Thr Val Thr Val Ser Ser 115
120585PRTHomo sapiens 58Ser Tyr Ala Ile Ser1 55917PRTHomo
sapiens 59Gly Ile Ile Pro Ile Phe Gly Thr Ala Asn Tyr Ala Gln Lys Phe
Gln1 5 10
15Gly6011PRTHomo sapiens 60Glu Gly Leu Ser Gly Gly Tyr Gly Met Asp Val1
5 1061112PRTHomo sapiens 61Gln Ser Val Leu
Thr Gln Pro Pro Ser Val Ser Gly Ala Pro Gly Gln1 5
10 15Arg Val Thr Ile Ser Cys Thr Gly Ser Ser
Ser Asn Ile Gly Ala Gly 20 25
30Tyr Asp Val His Trp Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys Leu
35 40 45Leu Ile Tyr Arg Asn Asn Asn Arg
Pro Ser Gly Val Pro Asp Arg Phe 50 55
60Ser Gly Ser Asn Ser Gly Thr Ser Ala Ser Leu Ala Ile Thr Gly Leu65
70 75 80Arg Ala Glu Asp Glu
Ala Asp Tyr Tyr Cys Gln Ser Tyr Asp Ser Ser 85
90 95Leu Ser Ser Tyr Val Phe Gly Thr Gly Thr Lys
Val Thr Val Leu Gly 100 105
1106214PRTHomo sapiens 62Thr Gly Ser Ser Ser Asn Ile Gly Ala Gly Tyr Asp
Val His1 5 10637PRTHomo sapiens 63Arg Asn
Asn Asn Arg Pro Ser1 5649PRTHomo sapiens 64Gln Ser Tyr Asp
Ser Ser Leu Ser Ser1 565125PRTHomo sapiens 65Glu Val Gln
Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1 5
10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser
Gly Phe Thr Phe Ser Ser Tyr 20 25
30Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45Ser Ala Ile Ser Gly Ser Gly
Gly Ser Thr Tyr Tyr Ala Asp Ser Val 50 55
60Lys Gly Arg Phe Thr Asn Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr65
70 75 80Leu Gln Met Asn
Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys 85
90 95Ala Lys Ala Tyr Tyr Asp Ile Leu Thr Gly
Tyr Phe Tyr Asn Gly Met 100 105
110Asp Val Trp Gly Gln Gly Thr Met Val Thr Val Ser Ser 115
120 125665PRTHomo sapiens 66Ser Tyr Ala Met Ser1
56717PRTHomo sapiens 67Ala Ile Ser Gly Ser Gly Gly Ser Thr
Tyr Tyr Ala Asp Ser Val Lys1 5 10
15Gly6816PRTHomo sapiens 68Ala Tyr Tyr Asp Ile Leu Thr Gly Tyr
Phe Tyr Asn Gly Met Asp Val1 5 10
1569112PRTHomo sapiens 69Gln Ala Val Leu Thr Gln Pro Ser Ser Val
Ser Gly Ser Pro Gly Gln1 5 10
15Ser Ile Thr Ile Ser Cys Thr Gly Thr Ser Ser Asp Val Gly Gly Tyr
20 25 30Asp Tyr Val Ser Trp Tyr
Gln Gln His Pro Gly Lys Ala Pro Lys Leu 35 40
45Met Ile His Asp Val Arg Asn Arg Pro Ser Gly Val Ser Asn
Arg Phe 50 55 60Ser Gly Ser Lys Phe
Gly Asn Thr Ala Ser Leu Thr Ile Ser Gly Leu65 70
75 80Gln Thr Glu Asp Glu Ala Asp Tyr Tyr Cys
Ser Ser Tyr Thr Ser Ser 85 90
95Ser Thr His Val Leu Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly
100 105 1107014PRTHomo sapiens
70Thr Gly Thr Ser Ser Asp Val Gly Gly Tyr Asp Tyr Val Ser1
5 10717PRTHomo sapiens 71Asp Val Arg Asn Arg Pro Ser1
5729PRTHomo sapiens 72Ser Ser Tyr Thr Ser Ser Ser Thr His1
573123PRTHomo sapiens 73Gln Val Gln Leu Val Gln Ser Gly Gly
Gly Leu Val Gln Pro Gly Arg1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Asp Asp
Tyr 20 25 30Ala Met His Trp
Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35
40 45Ser Gly Ile Ser Trp Asn Ser Gly Ser Ile Gly Tyr
Ala Asp Ser Val 50 55 60Lys Gly Arg
Phe Thr Ile Ser Arg Asp Asn Ala Lys Ser Ser Leu Tyr65 70
75 80Leu Gln Met Asn Ser Leu Arg Ala
Glu Asp Thr Ala Leu Tyr Tyr Cys 85 90
95Ala Lys Gly His Ser Pro Tyr Ser Ser Gly Trp Ser Asp Phe
Asp Tyr 100 105 110Trp Gly Gln
Gly Thr Leu Val Thr Val Ser Ser 115 120745PRTHomo
sapiens 74Asp Tyr Ala Met His1 57517PRTHomo sapiens 75Gly
Ile Ser Trp Asn Ser Gly Ser Ile Gly Tyr Ala Asp Ser Val Lys1
5 10 15Gly7614PRTHomo sapiens 76Gly
His Ser Pro Tyr Ser Ser Gly Trp Ser Asp Phe Asp Tyr1 5
1077112PRTHomo sapiens 77Gln Ser Val Leu Thr Gln Pro Pro Ser
Val Ser Gly Ala Pro Gly Gln1 5 10
15Arg Val Thr Ile Ser Cys Thr Gly Ser Ser Ser Asn Ile Gly Ala
Gly 20 25 30Tyr Asp Val Gln
Trp Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys Leu 35
40 45Leu Ile His Ala Asn Lys Asn Arg Pro Ser Gly Val
Pro Asp Arg Ile 50 55 60Ser Gly Ser
Lys Ser Gly Thr Thr Ala Ser Leu Ala Ile Thr Gly Phe65 70
75 80Gln Ala Glu Asp Glu Ala Asp Tyr
Tyr Cys Gln Ser Tyr Asp Ser Ser 85 90
95Leu Thr Gly Tyr Val Phe Gly Thr Gly Thr Lys Val Thr Val
Leu Gly 100 105 1107814PRTHomo
sapiens 78Thr Gly Ser Ser Ser Asn Ile Gly Ala Gly Tyr Asp Val Gln1
5 10797PRTHomo sapiens 79Ala Asn Lys Asn Arg Pro
Ser1 5809PRTHomo sapiens 80Gln Ser Tyr Asp Ser Ser Leu Thr
Gly1 581125PRTHomo sapiens 81Glu Val Gln Leu Val Glu Ser
Gly Gly Gly Leu Val Gln Pro Gly Arg1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe
Ala Asp Tyr 20 25 30Ala Met
His Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35
40 45Ser Ser Ile Ser Trp Asn Ser Gly Ser Ile
Ala Tyr Ala Asp Ser Val 50 55 60Lys
Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65
70 75 80Leu Gln Met Asn Ser Leu
Arg Ala Glu Asp Thr Ala Leu Tyr Tyr Cys 85
90 95Ala Lys Ala Ser Ala Ala Gly Thr Glu Tyr Tyr His
Tyr Tyr Gly Met 100 105 110Asp
Val Trp Gly Gln Gly Thr Thr Val Thr Val Ser Arg 115
120 125825PRTHomo sapiens 82Asp Tyr Ala Met His1
58317PRTHomo sapiens 83Ser Ile Ser Trp Asn Ser Gly Ser Ile Ala Tyr
Ala Asp Ser Val Lys1 5 10
15Gly8416PRTHomo sapiens 84Ala Ser Ala Ala Gly Thr Glu Tyr Tyr His Tyr
Tyr Gly Met Asp Val1 5 10
1585111PRTHomo sapiens 85Ser Tyr Val Leu Thr Gln Pro Pro Ser Ala Ser Gly
Thr Pro Gly Gln1 5 10
15Arg Val Thr Ile Ser Cys Ser Gly Ser Ser Ser Asn Ile Gly Ser Asn
20 25 30Thr Ile Asn Trp Tyr Gln Gln
Leu Pro Gly Thr Ala Pro Lys Leu Leu 35 40
45Ile Tyr Asn Asn His Gln Arg Pro Ser Gly Val Pro Asp Arg Phe
Ser 50 55 60Gly Ser Lys Ser Gly Thr
Ser Ala Ser Leu Ala Ile Ser Gly Leu Gln65 70
75 80Ser Ala Asp Glu Ala Asp Tyr Tyr Cys Gly Ala
Trp Asn Asp Ser Leu 85 90
95Asn Val Tyr Val Phe Gly Thr Gly Thr Lys Val Thr Val Leu Gly
100 105 1108613PRTHomo sapiens 86Ser Gly
Ser Ser Ser Asn Ile Gly Ser Asn Thr Ile Asn1 5
10877PRTHomo sapiens 87Asn Asn His Gln Arg Pro Ser1
5889PRTHomo sapiens 88Gly Ala Trp Asn Asp Ser Leu Asn Val1
589124PRTHomo sapiens 89Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys
Lys Pro Gly Ala1 5 10
15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser Tyr
20 25 30Gly Ile Ser Trp Val Arg Gln
Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40
45Gly Trp Ile Ser Ala Tyr Asn Gly Asn Thr Asn Tyr Ala Gln Lys
Leu 50 55 60Gln Gly Arg Val Thr Met
Thr Thr Asp Thr Ser Thr Ser Thr Ala Tyr65 70
75 80Met Glu Leu Arg Ser Leu Arg Ser Asp Asp Thr
Ala Val Tyr Tyr Cys 85 90
95Ala Thr Ala Ala Ser Leu Lys Tyr Tyr Tyr Asp Ser Ser Gly Tyr Tyr
100 105 110Tyr Trp Gly Gln Gly Thr
Leu Val Thr Val Ser Arg 115 120905PRTHomo sapiens
90Ser Tyr Gly Ile Ser1 59117PRTHomo sapiens 91Trp Ile Ser
Ala Tyr Asn Gly Asn Thr Asn Tyr Ala Gln Lys Leu Gln1 5
10 15Gly9215PRTHomo sapiens 92Ala Ala Ser
Leu Lys Tyr Tyr Tyr Asp Ser Ser Gly Tyr Tyr Tyr1 5
10 1593110PRTHomo sapiens 93Ser Ser Glu Leu Thr
Gln Asp Pro Ala Val Ser Val Ala Leu Gly Gln1 5
10 15Thr Val Arg Ile Thr Cys Gln Gly Asp Ser Leu
Arg Ser Tyr Tyr Ala 20 25
30Ser Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Val Leu Val Ile Ser
35 40 45Gly Lys Asn Asn Arg Pro Ser Gly
Ile Pro Asp Arg Phe Ser Gly Ser 50 55
60Ser Ser Gly Asp Thr Ala Ser Leu Thr Ile Thr Gly Ala Gln Ala Glu65
70 75 80Asp Glu Ala Asn Tyr
Tyr Cys Asn Ser Arg Asp Ser Ser Gly Tyr Pro 85
90 95Ser Trp Val Phe Gly Gly Gly Thr Lys Leu Thr
Val Leu Gly 100 105
1109411PRTHomo sapiens 94Gln Gly Asp Ser Leu Arg Ser Tyr Tyr Ala Ser1
5 10957PRTHomo sapiens 95Gly Lys Asn Asn Arg
Pro Ser1 59610PRTHomo sapiens 96Asn Ser Arg Asp Ser Ser Gly
Tyr Pro Ser1 5 1097130PRTHomo sapiens
97Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys
Lys Gly Ser Gly Tyr Val Phe Asn Ser Tyr 20 25
30Gly Ile Thr Trp Val Arg Gln Ala Pro Gly Gln Gly Leu
Glu Trp Met 35 40 45Gly Trp Ile
Ser Ala Tyr Asn Gly Tyr Thr Asp Tyr Ala Gln Lys Val 50
55 60Gln Gly Arg Val Thr Met Thr Thr Glu Thr Ser Thr
Ser Thr Ala Tyr65 70 75
80Met Glu Leu Arg Ser Leu Arg Ser Asp Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Asp Tyr Tyr Asp
Ser Ser Thr Tyr Tyr Ser Ser Asp Tyr Phe 100
105 110Gln Tyr Tyr Gly Met Asp Val Trp Gly Gln Gly Thr
Thr Val Thr Val 115 120 125Ser Ser
130985PRTHomo sapiens 98Ser Tyr Gly Ile Thr1
59917PRTHomo sapiens 99Trp Ile Ser Ala Tyr Asn Gly Tyr Thr Asp Tyr Ala
Gln Lys Val Gln1 5 10
15Gly10021PRTHomo sapiens 100Asp Tyr Tyr Asp Ser Ser Thr Tyr Tyr Ser Ser
Asp Tyr Phe Gln Tyr1 5 10
15Tyr Gly Met Asp Val 20101111PRTHomo sapiens 101Gln Ala Val
Leu Thr Gln Pro Ala Ser Val Ser Gly Ser Pro Gly Gln1 5
10 15Ser Ile Thr Ile Ser Cys Thr Gly Thr
Ser Ser Asp Val Gly Ala Tyr 20 25
30Asn Tyr Val Ser Trp Tyr Gln Gln His Pro Gly Lys Ala Pro Lys Leu
35 40 45Met Ile Ser Asp Val Ser Arg
Arg Pro Ser Gly Val Ser Asn Arg Phe 50 55
60Ser Gly Ser Lys Ser Gly Asn Thr Ala Ser Leu Thr Ile Ser Gly Leu65
70 75 80Gln Thr Glu Asp
Glu Ala Asp Tyr Tyr Cys Ser Ser Tyr Thr Ser Ser 85
90 95Asn Thr Val Leu Phe Gly Gly Gly Thr Lys
Leu Thr Val Leu Gly 100 105
11010214PRTHomo sapiens 102Thr Gly Thr Ser Ser Asp Val Gly Ala Tyr Asn
Tyr Val Ser1 5 101037PRTHomo sapiens
103Asp Val Ser Arg Arg Pro Ser1 51048PRTHomo sapiens 104Ser
Ser Tyr Thr Ser Ser Asn Thr1 5105122PRTHomo sapiens 105Gln
Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Glu1
5 10 15Ser Leu Arg Ile Ser Cys Lys
Gly Ser Gly Tyr Ser Phe Thr Asn Tyr 20 25
30Trp Ile Gly Trp Val Arg Gln Met Pro Gly Lys Gly Leu Glu
Trp Met 35 40 45Gly Ile Ile Tyr
Pro Gly Asp Ser Asp Thr Arg Tyr Ser Pro Ser Phe 50 55
60Gln Gly Gln Val Thr Ile Ala Ala Asp Lys Ser Ile Ser
Thr Ala Tyr65 70 75
80Leu Gln Trp Ser Ser Leu Met Ala Ser Asp Thr Ala Met Tyr Tyr Cys
85 90 95Ala Arg Arg Gly Ser Arg
Ser Ser Gly Glu Asp Ala Phe Glu Val Trp 100
105 110Gly Gln Gly Thr Thr Val Thr Val Ser Ser 115
1201065PRTHomo sapiens 106Asn Tyr Trp Ile Gly1
510717PRTHomo sapiens 107Ile Ile Tyr Pro Gly Asp Ser Asp Thr Arg Tyr
Ser Pro Ser Phe Gln1 5 10
15Gly10813PRTHomo sapiens 108Arg Gly Ser Arg Ser Ser Gly Glu Asp Ala Phe
Glu Val1 5 10109110PRTHomo sapiens 109Ser
Tyr Glu Leu Thr Gln Pro Pro Ser Val Ser Val Ala Pro Gly Lys1
5 10 15Thr Ala Thr Ile Thr Cys Gly
Gly Asp Asn Ile Gly Ser Lys Ser Val 20 25
30His Trp Tyr Gln Gln Arg Pro Gly Gln Ala Pro Val Leu Val
Ile Asn 35 40 45Tyr Asp Ser Asp
Arg Pro Ser Gly Ile Pro Glu Arg Phe Ser Gly Ser 50 55
60Asn Ser Gly Asn Thr Ala Thr Leu Thr Ile Ser Arg Val
Glu Ala Gly65 70 75
80Asp Glu Ala Asp Tyr Tyr Cys Gln Val Glu Asp Arg Arg Gly Gly Tyr
85 90 95His Val Val Phe Gly Gly
Gly Thr Lys Leu Thr Val Leu Gly 100 105
11011011PRTHomo sapiens 110Gly Gly Asp Asn Ile Gly Ser Lys Ser
Val His1 5 101117PRTHomo sapiens 111Tyr
Asp Ser Asp Arg Pro Ser1 511210PRTHomo sapiens 112Gln Val
Glu Asp Arg Arg Gly Gly Tyr His1 5
10113125PRTHomo sapiens 113Gln Val Gln Leu Val Gln Ser Gly Gly Gly Leu
Val Gln Pro Gly Gly1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Val Ser Ser Asn
20 25 30Tyr Met Ser Trp Val Arg Gln
Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40
45Ser Val Ile Tyr Ser Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val
Lys 50 55 60Gly Arg Phe Thr Ile Ser
Arg Asp Asn Ser Lys Asn Thr Leu Tyr Leu65 70
75 80Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala
Val Tyr Tyr Cys Ala 85 90
95Arg Glu Gly Tyr Cys Ser Gly Gly Ser Cys Tyr Ser Tyr Gly Ala Phe
100 105 110Asp Ile Trp Gly Gln Gly
Thr Thr Val Thr Val Ser Ser 115 120
1251145PRTHomo sapiens 114Ser Asn Tyr Met Ser1
511516PRTHomo sapiens 115Val Ile Tyr Ser Gly Gly Ser Thr Tyr Tyr Ala Asp
Ser Val Lys Gly1 5 10
1511617PRTHomo sapiens 116Glu Gly Tyr Cys Ser Gly Gly Ser Cys Tyr Ser Tyr
Gly Ala Phe Asp1 5 10
15Ile117108PRTHomo sapiens 117Asp Ile Val Met Thr Gln Ser Pro Ser Ser Val
Ser Ala Ser Val Gly1 5 10
15Asp Arg Val Thr Ile Ser Cys Arg Ala Ser Gln Asn Ile Ala Asn Trp
20 25 30Leu Ala Trp Tyr Gln Gln Lys
Pro Gly Lys Ala Pro Lys Leu Leu Ile 35 40
45Tyr Ala Ala Ser Asn Leu Gln Ser Gly Val Pro Ser Arg Phe Ser
Gly 50 55 60Ser Gly Ser Gly Thr Asp
Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70
75 80Glu Asp Phe Ala Ser Tyr Tyr Cys Gln Gln Gly
Asn Ser Phe Pro Arg 85 90
95Val Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100
10511811PRTHomo sapiens 118Arg Ala Ser Gln Asn Ile Ala Asn Trp
Leu Ala1 5 101197PRTHomo sapiens 119Ala
Ala Ser Asn Leu Gln Ser1 51208PRTHomo sapiens 120Gln Gln
Gly Asn Ser Phe Pro Arg1 5121130PRTHomo sapiens 121Gln Val
Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5
10 15Ser Val Lys Val Ser Cys Lys Ala
Ser Gly Tyr Val Phe Asn Ser Tyr 20 25
30Gly Ile Thr Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp
Met 35 40 45Gly Trp Ile Ser Ala
Tyr Asn Gly Tyr Thr Asp Tyr Ala Gln Lys Val 50 55
60Gln Gly Arg Val Thr Met Thr Thr Glu Thr Ser Thr Ser Thr
Ala Tyr65 70 75 80Met
Glu Leu Arg Ser Leu Arg Ser Asp Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Asp Tyr Tyr Asp Ser
Ser Thr Tyr Tyr Ser Ser Asp Tyr Phe 100 105
110Lys Tyr Tyr Gly Met Asp Val Trp Gly Gln Gly Thr Thr Val
Thr Val 115 120 125Ser Ser
1301225PRTHomo sapiens 122Ser Tyr Gly Ile Thr1
512317PRTHomo sapiens 123Trp Ile Ser Ala Tyr Asn Gly Tyr Thr Asp Tyr Ala
Gln Lys Val Gln1 5 10
15Gly12421PRTHomo sapiens 124Asp Tyr Tyr Asp Ser Ser Thr Tyr Tyr Ser Ser
Asp Tyr Phe Lys Tyr1 5 10
15Tyr Gly Met Asp Val20125111PRTHomo sapiens 125Gln Ala Val Leu Thr Gln
Pro Ser Ser Val Ser Gly Ser Pro Gly Gln1 5
10 15Ser Ile Thr Ile Ser Cys Ser Gly Thr Ser Ser Asp
Val Gly Ala Tyr 20 25 30Asn
Tyr Val Ser Trp Tyr Gln Gln His Pro Gly Lys Ala Pro Arg Leu 35
40 45Leu Thr Phe Asp Val Asn Arg Arg Pro
Ser Gly Ser Ser Ser Arg Phe 50 55
60Ser Gly Ser Lys Ser Gly Asn Thr Ala Ser Leu Thr Ile Ser Gly Leu65
70 75 80Gln Ala Glu Asp Glu
Ala Asp Tyr Tyr Cys Ser Ser Tyr Thr Asn Ser 85
90 95Asn Thr Val Val Phe Gly Gly Gly Thr Arg Leu
Thr Val Leu Ser 100 105
11012614PRTHomo sapiens 126Ser Gly Thr Ser Ser Asp Val Gly Ala Tyr Asn
Tyr Val Ser1 5 101277PRTHomo sapiens
127Asp Val Asn Arg Arg Pro Ser1 51288PRTHomo sapiens 128Ser
Ser Tyr Thr Asn Ser Asn Thr1 5129127PRTHomo sapiens 129Gln
Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Lys Pro Ser Glu1
5 10 15Thr Leu Ala Leu Thr Cys Thr
Val Ser Gly Gly Ser Ile Ser Asn Tyr 20 25
30Tyr Trp Ser Trp Ile Arg Gln Pro Pro Gly Lys Gly Leu Glu
Trp Ile 35 40 45Gly Tyr Ile Tyr
Asp Ile Glu Asn Thr Asn Tyr Asn Pro Ser Leu Lys 50 55
60Ser Arg Val Thr Ile Ser Val Asp Thr Ser Lys Asn Gln
Phe Ser Leu65 70 75
80Lys Leu Ser Ser Val Thr Ala Asp Asp Thr Ala Val Tyr Tyr Cys Ala
85 90 95Arg Asp Ser Arg Val Ile
Arg Phe Leu Glu Gly Tyr Ser Tyr Tyr Tyr 100
105 110Gly Val Asp Val Trp Gly Gln Gly Thr Met Val Thr
Val Ser Ser 115 120 1251305PRTHomo
sapiens 130Asn Tyr Tyr Trp Ser1 513116PRTHomo sapiens
131Tyr Ile Tyr Asp Ile Glu Asn Thr Asn Tyr Asn Pro Ser Leu Lys Ser1
5 10 1513219PRTHomo sapiens
132Asp Ser Arg Val Ile Arg Phe Leu Glu Gly Tyr Ser Tyr Tyr Tyr Gly1
5 10 15Val Asp
Val133111PRTHomo sapiens 133Ser Tyr Glu Leu Thr Gln Pro Pro Ser Ala Ser
Gly Thr Pro Gly Gln1 5 10
15Thr Val Ile Ile Ser Cys Ser Gly Ser Arg Ser Asn Ile Gly Gly His
20 25 30Gly Val Asn Trp His Gln Gln
Val Pro Gly Thr Ala Pro Lys Leu Leu 35 40
45Ile Tyr Arg Asn Asp Arg Arg Pro Ser Gly Val Pro Asp Arg Phe
Ser 50 55 60Gly Ser Lys Ser Gly Thr
Ser Ala Ser Leu Val Ile Ser Gly Leu Gln65 70
75 80Phe Glu Asp Glu Ala Asp Tyr Tyr Cys Val Ala
Trp Glu Asp Ser Leu 85 90
95Asp Gly Pro Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly
100 105 11013413PRTHomo sapiens 134Ser
Gly Ser Arg Ser Asn Ile Gly Gly His Gly Val Asn1 5
101357PRTHomo sapiens 135Arg Asn Asp Arg Arg Pro Ser1
51369PRTHomo sapiens 136Val Ala Trp Glu Asp Ser Leu Asp Gly1
5137115PRTHomo sapiens 137Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu
Val Gln Pro Gly Gly1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr
20 25 30Ala Met Ser Trp Val Arg Gln
Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40
45Ser Thr Ile Ser Gly Ser Gly Gly Ser Thr Tyr Tyr Ala Asp Ser
Val 50 55 60Lys Gly Arg Phe Thr Ile
Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr65 70
75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90
95Ala Arg Gly Ile Val Ala Thr Ser Trp Gly Gln Gly Thr Leu Val Thr
100 105 110Val Ser Arg
1151385PRTHomo sapiens 138Ser Tyr Ala Met Ser1
513917PRTHomo sapiens 139Thr Ile Ser Gly Ser Gly Gly Ser Thr Tyr Tyr Ala
Asp Ser Val Lys1 5 10
15Gly1406PRTHomo sapiens 140Gly Ile Val Ala Thr Ser1
5141108PRTHomo sapiens 141Ser Tyr Glu Leu Thr Gln Pro Pro Ser Val Ser Val
Ala Pro Gly Gln1 5 10
15Thr Ala Arg Ile Thr Cys Gly Gly Asn Lys Ile Gly Ser Lys Ser Val
20 25 30His Trp Tyr Gln Gln Lys Gln
Gly Gln Ala Pro Val Leu Val Ile Tyr 35 40
45Leu Asp Arg Asp Arg Pro Ser Gly Ile Pro Glu Arg Phe Ser Gly
Ser 50 55 60Asn Ser Gly Asn Thr Ala
Thr Leu Thr Ile Thr Arg Val Glu Ala Glu65 70
75 80Asp Glu Ala Asp Tyr Tyr Cys His Leu Trp Asp
Ser Gly Ser Asp Gln 85 90
95Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly 100
10514211PRTHomo sapiens 142Gly Gly Asn Lys Ile Gly Ser Lys Ser
Val His1 5 101437PRTHomo sapiens 143Leu
Asp Arg Asp Arg Pro Ser1 51448PRTHomo sapiens 144His Leu
Trp Asp Ser Gly Ser Asp1 5145114PRTHomo sapiens 145Glu Val
Gln Leu Val Glu Ser Gly Gly Gly Val Val Gln Pro Gly Gly1 5
10 15Ser Leu Arg Leu Ser Cys Ala Ala
Ser Gly Phe Asn Phe Asp Val Tyr 20 25
30Gly Met Asn Trp Val Arg Gln Val Pro Gly Lys Gly Leu Glu Trp
Val 35 40 45Ser Leu Ile Asn Gly
Asp Gly Gly Leu Arg Tyr Tyr Ala Asp Ser Val 50 55
60Lys Gly Arg Phe Thr Val Ser Arg Asp Asn Ser Arg Asn Ser
Leu Tyr65 70 75 80Leu
Gln Met Asn Ser Leu Arg Ser Glu Asp Thr Ala Leu Tyr Tyr Cys
85 90 95Val Lys Gly Asn Phe Gln Gln
Trp Gly Gln Gly Thr Leu Val Thr Val 100 105
110Ser Arg1465PRTHomo sapiens 146Val Tyr Gly Met Asn1
514717PRTHomo sapiens 147Leu Ile Asn Gly Asp Gly Gly Leu Arg Tyr
Tyr Ala Asp Ser Val Lys1 5 10
15Gly1485PRTHomo sapiens 148Gly Asn Phe Gln Gln1
5149109PRTHomo sapiens 149Ser Tyr Val Leu Thr Gln Pro Pro Ser Val Ser Val
Ala Pro Gly Lys1 5 10
15Thr Ala Arg Ile Thr Cys Gly Gly Asn Asn Ile Gly Ser Lys Ser Val
20 25 30His Trp Tyr Gln Gln Lys Pro
Gly Gln Ala Pro Val Leu Val Ile Tyr 35 40
45Tyr Asp Ser Asp Arg Pro Ser Gly Ile Pro Glu Arg Phe Ser Gly
Ser 50 55 60Asn Ser Gly Asn Thr Ala
Thr Leu Thr Ile Ser Arg Val Glu Ala Gly65 70
75 80Asp Glu Ala Asp Tyr Tyr Cys Gln Val Trp Asp
Ser Ser Ser Asp His 85 90
95Val Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly 100
10515011PRTHomo sapiens 150Gly Gly Asn Asn Ile Gly Ser Lys
Ser Val His1 5 101517PRTHomo sapiens
151Tyr Asp Ser Asp Arg Pro Ser1 51529PRTHomo sapiens 152Gln
Val Trp Asp Ser Ser Ser Asp His1 5153115PRTHomo sapiens
153Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1
5 10 15Ser Leu Arg Leu Ser Cys
Ala Ala Ser Gly Phe Thr Phe Asp Asp Tyr 20 25
30Ala Met His Trp Val Arg Gln Ala Pro Gly Lys Gly Leu
Glu Trp Val 35 40 45Ser Leu Ile
Ser Gly Asp Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val 50
55 60Lys Asp Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys
Asn Ser Leu Tyr65 70 75
80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Gly Asn Tyr Phe
Asp Tyr Trp Gly Gln Gly Thr Leu Val Thr 100
105 110Val Ser Arg 1151545PRTHomo sapiens 154Asp
Tyr Ala Met His1 515517PRTHomo sapiens 155Leu Ile Ser Gly
Asp Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val Lys1 5
10 15Asp1566PRTHomo sapiens 156Gly Asn Tyr Phe
Asp Tyr1 5157106PRTHomo sapiens 157Ser Tyr Glu Leu Thr Gln
Pro Leu Ser Val Ser Val Ala Leu Gly Gln1 5
10 15Thr Ala Arg Ile Thr Cys Gly Gly Asn Asn Ile Gly
Ser Lys Asn Val 20 25 30His
Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Val Leu Val Ile Tyr 35
40 45Arg Asp Ser Asn Arg Pro Ser Gly Ile
Pro Glu Arg Phe Ser Gly Ser 50 55
60Asn Ser Gly Asn Thr Ala Thr Leu Thr Ile Ser Arg Ala Gln Ala Gly65
70 75 80Asp Glu Ala Asp Tyr
Tyr Cys Gln Val Trp Asp Ser Ser Val Val Phe 85
90 95Gly Gly Gly Thr Lys Leu Thr Val Leu Gly
100 10515811PRTHomo sapiens 158Gly Gly Asn Asn Ile
Gly Ser Lys Asn Val His1 5 101597PRTHomo
sapiens 159Arg Asp Ser Asn Arg Pro Ser1 51606PRTHomo
sapiens 160Gln Val Trp Asp Ser Ser1 5161122PRTHomo sapiens
161Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1
5 10 15Ser Leu Arg Leu Ser Cys
Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20 25
30Ser Met Asn Trp Val Arg Gln Ala Pro Gly Lys Gly Leu
Glu Trp Val 35 40 45Ser Tyr Ile
Ser Ser Ser Ser Ser Thr Ile Tyr Tyr Ala Asp Ser Val 50
55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys
Asn Ser Leu Tyr65 70 75
80Leu Gln Met Asn Ser Leu Arg Asp Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Val Met Pro Ser
Tyr Tyr Tyr Tyr Tyr Gly Met Asp Val Trp 100
105 110Gly Gln Gly Thr Thr Val Thr Val Ser Arg 115
1201625PRTHomo sapiens 162Ser Tyr Ser Met Asn1
516317PRTHomo sapiens 163Tyr Ile Ser Ser Ser Ser Ser Thr Ile Tyr Tyr
Ala Asp Ser Val Lys1 5 10
15Gly16413PRTHomo sapiens 164Val Met Pro Ser Tyr Tyr Tyr Tyr Tyr Gly Met
Asp Val1 5 10165111PRTHomo sapiens 165Asn
Phe Met Leu Thr Gln Pro His Ser Val Ser Glu Ser Pro Gly Lys1
5 10 15Thr Val Thr Ile Ser Cys Thr
Gly Ser Ser Gly Ser Ile Ala Ser Asn 20 25
30Tyr Val Gln Trp Tyr Gln Gln Arg Pro Gly Ser Ala Pro Thr
Thr Val 35 40 45Ile Tyr Glu Asp
Ser Glu Arg Pro Ser Gly Val Pro Asp Arg Phe Ser 50 55
60Gly Ser Ile Asp Ser Ser Ser Asn Ser Ala Ser Leu Thr
Ile Ser Gly65 70 75
80Leu Lys Thr Gln Asp Glu Ala Asp Tyr Tyr Cys Gln Ser Tyr Asp Gly
85 90 95Val Asn Trp Val Phe Gly
Gly Gly Thr Lys Leu Thr Val Leu Gly 100 105
11016613PRTHomo sapiens 166Thr Gly Ser Ser Gly Ser Ile Ala
Ser Asn Tyr Val Gln1 5 101677PRTHomo
sapiens 167Glu Asp Ser Glu Arg Pro Ser1 51687PRTHomo
sapiens 168Gln Ser Tyr Asp Gly Val Asn1 5169114PRTHomo
sapiens 169Glu Val Gln Leu Val Glu Ser Gly Gly Gly Val Val Gln Pro Gly
Arg1 5 10 15Ser Leu Arg
Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20
25 30Gly Met His Trp Val Arg Gln Ala Pro Gly
Lys Gly Leu Glu Trp Val 35 40
45Ala Val Ile Ser Tyr Asp Gly Ser Asn Lys Tyr Tyr Ala Asp Ser Val 50
55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp
Asn Ser Lys Asn Thr Leu Tyr65 70 75
80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr
Tyr Cys 85 90 95Ala Asp
Tyr Gly Met Asp Val Trp Gly Gln Gly Thr Thr Val Thr Val 100
105 110Ser Arg 1705PRTHomo sapiens 170Ser
Tyr Gly Met His1 517117PRTHomo sapiens 171Val Ile Ser Tyr
Asp Gly Ser Asn Lys Tyr Tyr Ala Asp Ser Val Lys1 5
10 15Gly1725PRTHomo sapiens 172Tyr Gly Met Asp
Val1 5173107PRTHomo sapiens 173Asp Ile Gln Met Thr Gln Ser
Pro Ser Ser Leu Ser Ala Ser Val Gly1 5 10
15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Gln Ser Ile
Ser Ser Tyr 20 25 30Leu Asn
Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu Leu Ile 35
40 45Tyr Ala Ala Ser Ser Leu Gln Ser Gly Val
Pro Ser Arg Phe Ser Gly 50 55 60Ser
Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65
70 75 80Glu Asp Phe Ala Thr Tyr
Tyr Cys Gln Gln Ser Tyr Ser Thr Pro Val 85
90 95Thr Phe Gly Gln Gly Thr Lys Val Glu Ile Lys
100 10517411PRTHomo sapiens 174Arg Ala Ser Gln Ser
Ile Ser Ser Tyr Leu Asn1 5 101757PRTHomo
sapiens 175Ala Ala Ser Ser Leu Gln Ser1 51767PRTHomo
sapiens 176Gln Gln Ser Tyr Ser Thr Pro1 5177120PRTHomo
sapiens 177Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Lys Pro Ser
Gln1 5 10 15Thr Leu Ser
Leu Thr Cys Thr Val Ser Gly Gly Ser Ile Ser Ser Gly 20
25 30Gly Tyr Tyr Trp Ser Trp Ile Arg Gln His
Pro Gly Lys Gly Leu Glu 35 40
45Trp Ile Gly Tyr Ile Tyr Tyr Ser Gly Ser Thr Tyr Tyr Asn Pro Ser 50
55 60Leu Lys Ser Arg Val Thr Ile Ser Val
Asp Thr Ser Lys Asn Gln Phe65 70 75
80Ser Leu Lys Leu Ser Ser Val Thr Ala Ala Asp Thr Ala Val
Tyr Tyr 85 90 95Cys Ala
Arg Asp Arg Gly Thr Gly Asp Ala Phe Asp Ile Trp Gly Gln 100
105 110Gly Thr Met Val Thr Val Ser Arg
115 1201787PRTHomo sapiens 178Ser Gly Gly Tyr Tyr Trp
Ser1 517916PRTHomo sapiens 179Tyr Ile Tyr Tyr Ser Gly Ser
Thr Tyr Tyr Asn Pro Ser Leu Lys Ser1 5 10
1518010PRTHomo sapiens 180Asp Arg Gly Thr Gly Asp Ala
Phe Asp Ile1 5 10181111PRTHomo sapiens
181Gln Ser Val Leu Thr Gln Pro Pro Ser Val Ser Gly Ala Pro Arg Gln1
5 10 15Thr Val Thr Ile Ser Cys
Ser Gly Ser Ser Ser Asn Ile Gly Gln Asn 20 25
30Ser Val Thr Trp Tyr Gln Arg Leu Pro Gly Glu Ala Pro
Lys Leu Leu 35 40 45Ile Tyr Tyr
Asp Asp Leu Leu His Ser Gly Val Ser Asp Arg Phe Ser 50
55 60Gly Ser Lys Ser Gly Thr Ser Ala Ser Leu Ala Ile
Ser Gly Leu Gln65 70 75
80Ser Glu Asp Glu Ala Glu Tyr Tyr Cys Ala Ser Trp Asp Asp Ser Leu
85 90 95Lys Gly Pro Val Phe Gly
Gly Gly Thr Lys Leu Thr Val Leu Gly 100 105
11018213PRTHomo sapiens 182Ser Gly Ser Ser Ser Asn Ile Gly
Gln Asn Ser Val Thr1 5 101837PRTHomo
sapiens 183Tyr Asp Asp Leu Leu His Ser1 51849PRTHomo
sapiens 184Ala Ser Trp Asp Asp Ser Leu Lys Gly1
5185360DNAHomo sapiens 185caggtgcagc tgcaggagtc gggcccagga ctggtgaagc
cttcacagac cctgtccctc 60acctgcactg tctctggtgg ctccatcagc agtggtggtt
actactggag ctggatccgc 120cagcacccag ggaagggcct ggagtggatt gggtacatct
attacagtgg gagcacctac 180tacaacccgt ccctcaagag tcgagtcacc atatcagtag
acacgtctaa gaaccagttc 240tccctgaagc tgagctctgt gactgccgcg gacacggccg
tgtattactg tgcgaggaca 300ccgtgggagc tactagcttt tgatatctgg ggccaaggga
caatggtcac cgtctcgaga 36018621DNAHomo sapiens 186agtggtggtt actactggag
c 2118748DNAHomo sapiens
187tacatctatt acagtgggag cacctactac aacccgtccc tcaagagt
4818830DNAHomo sapiens 188acaccgtggg agctactagc ttttgatatc
30189327DNAHomo sapiens 189tcctatgagc tgactcagcc
accctcagtg tcagtggccc caggaaagac gaccaggatt 60acctgtgggg gaaacaacat
tggaagtaaa agtgcgcact ggtaccagca gaagccaggc 120caggcccctg tgctggtcat
ctattatgat agcgaccggc cctcagggat ccctgagcga 180ttctctggct ccaactctgg
gaacacggcc accctgacca tcagcagggt cgaagccggg 240gatgaggccg actattactg
tcaggtgtgg gatagtagta gtgatcattg ggtgttcggc 300ggagggacca agctgaccgt
cctaggt 32719033DNAHomo sapiens
190gggggaaaca acattggaag taaaagtgcg cac
3319121DNAHomo sapiens 191tatgatagcg accggccctc a
2119227DNAHomo sapiens 192caggtgtggg atagtagtag
tgatcat 27193378DNAHomo sapiens
193caggtgcagc tggtgcagtc tggggctgag gtgaagaaga ctgggtcctc ggtgaaggtc
60tcctgcaagg cctctggagg ctccttcagc agctctgcaa tcagctgggt gcgacaggcc
120cctggacacg ggcttgaatg gctgggaggg atcatcccta cctttggtac accaaaccac
180gcacagaagt tccagggcag agtcacaatt accgcggacg aatcaacggg cacagcctac
240atggagctga gtggcctgag atctgaggac acggccgtgt attactgtgc gagagcccat
300tgtggtggtg gtaggtgtta cgactacact gatgcttttc atttctgggg ccaagggaca
360atggtcaccg tctcgaga
37819415DNAHomo sapiens 194agctctgcaa tcagc
1519551DNAHomo sapiens 195gggatcatcc ctacctttgg
tacaccaaac cacgcacaga agttccaggg c 5119651DNAHomo sapiens
196gcccattgtg gtggtggtag gtgttacgac tacactgatg cttttcattt c
51197327DNAHomo sapiens 197tcctatgagc tgactcagcc accctcagtg tcagtggccc
caggaaagac ggccaggatt 60acctgtgggg gagacaacat tggaaataga agtgtgcact
ggtaccagca gaagccaggc 120caggcccctg tgctgcttat ctattatgat agcgaccggc
cctcagggat ccctaagcga 180ttctctggct ccaactctgg gaacacggcc accctgacca
tcagcagggt cgaagccggg 240gatgaggccg actattactg tcaggtgtgg gatagtacta
gtgatcatgt ggtattcggc 300ggagggacca agctgaccgt cctacgt
32719833DNAHomo sapiens 198gggggagaca acattggaaa
tagaagtgtg cac 3319921DNAHomo sapiens
199tatgatagcg accggccctc a
2120027DNAHomo sapiens 200caggtgtggg atagtactag tgatcat
27201360DNAHomo sapiens 201caggtgcagc tgcaggagtc
gggcccagga ctggtgaagc cttcggagac cctgtccctc 60acctgcactg tctctggtgg
ctccatcagc agtagtagtt actactgggg ctggatccgc 120cagcccccag ggaaggggct
ggagtggatt gggagtatct attatagtgg gagcacctac 180tacaacccgt ccctcaagag
tcgagtcacc atatccgtag acacgtccaa gaaccagttc 240tccctgaagc tgagctctgt
gaccgccgca gacacggctg tgtattactg tgcgagactt 300cctatggtta cgatgtcctt
tgactactgg ggccagggaa ccctggtcac cgtctcgaga 36020221DNAHomo sapiens
202agtagtagtt actactgggg c
2120348DNAHomo sapiens 203agtatctatt atagtgggag cacctactac aacccgtccc
tcaagagt 4820430DNAHomo sapiens 204cttcctatgg ttacgatgtc
ctttgactac 30205327DNAHomo sapiens
205tcctatgtgc tgactcagcc accctcagtg tcagtggccc caggaaagac ggccaggatt
60acctgtgggg gaaacaacat tggaagtaaa agtgtgcact ggtaccagca gaagccaggc
120caggcccctg tgctggtcat ctattatgat agcgaccggc cctcagggat ccctgagcga
180ttctctggct ccaactctgg gaacacggcc accctgacca tcagcagggt cgaagccggg
240gatgaggccg actattactg tcaggtgtgg gatagtagta gtgatcatgt ggtattcggc
300ggagggacca agctgaccgt cctaggt
32720633DNAHomo sapiens 206gggggaaaca acattggaag taaaagtgtg cac
3320721DNAHomo sapiens 207tatgatagcg accggccctc a
2120827DNAHomo sapiens
208caggtgtggg atagtagtag tgatcat
27209363DNAHomo sapiens 209caggtgcagc tggtgcagtc tggagcagag gtgaaaaagc
ccggggagtc tctgaagatc 60tcctgtaagg gttctggata cagctttacc agctactgga
tcggctgggt gcgccagatg 120cccgggaaag gcctggagtg gatggggatc atctatcctg
gtgactctga taccagatac 180agcccgtcct tccaaggcca ggtcaccatc tcagccgaca
agtccatcag caccgcctac 240ctgcagtgga gcagcctgaa ggcctcggac accgccatgt
attactgtgc gagacttact 300ttgtcttata gcagcagctg gtttgactac tggggccagg
gaaccctggt caccgtctcg 360aga
36321015DNAHomo sapiens 210agctactgga tcggc
1521151DNAHomo sapiens
211atcatctatc ctggtgactc tgataccaga tacagcccgt ccttccaagg c
5121236DNAHomo sapiens 212cttactttgt cttatagcag cagctggttt gactac
36213333DNAHomo sapiens 213cagactgtgg tgacccagga
gccatcgttc tcagtgtccc ctggagggac agtcacactc 60acttgtggct tgagctctgg
ctcagtctct actagttact accccagctg gtaccagcag 120accccaggcc aggctccacg
cacgctcatc tacagcacaa acactcgctc ttctggggtc 180cctgatcgct tctctggctc
catccttggg aacaaagctg ccctcaccat cacgggggcc 240caggcagatg atgaatctga
ttattactgt gtgctgtata tgggtagtgg catttcggtg 300ttcggcggag ggaccaagct
gaccgtccta ggt 33321442DNAHomo sapiens
214ggcttgagct ctggctcagt ctctactagt tactacccca gc
4221521DNAHomo sapiens 215agcacaaaca ctcgctcttc t
2121624DNAHomo sapiens 216gtgctgtata tgggtagtgg
catt 24217354DNAHomo sapiens
217caggtgcagc tggtgcagtc tggagcagag gtgaaaaagc ccggggagtc tctgaagatc
60tcctgtaagg gttctggata cagctttacc agctactgga tcggctgggt gcgccagatg
120cccgggaaag gcctggagtg gatggggatc atctatcctg gtgactctga taccagatac
180agcccgtcct tccaaggcca ggtcaccatc tcagccgaca agtccatcag caccgcctac
240ctgcagtgga gcagcctgaa ggcctcggac accgccatgt attactgtgc gagacttctg
300gggataggcg cttttgatat ctggggccaa gggaccacgg tcaccgtctc gaga
35421815DNAHomo sapiens 218agctactgga tcggc
1521951DNAHomo sapiens 219atcatctatc ctggtgactc
tgataccaga tacagcccgt ccttccaagg c 5122027DNAHomo sapiens
220cttctgggga taggcgcttt tgatatc
27221330DNAHomo sapiens 221tcttctgagc tgactcagga ccctgctgtg tctgtggcct
tgggacagac agtcaggatc 60acatgccaag gagacagcct cagaagctat tatgcaagct
ggtaccagca gaagccagga 120caggcccctg tacttgtcat ctatggtaaa aacaaccggc
cctcagggat cccagaccga 180ttctctggct ccagctcagg aaacacagct tccttgacca
tcactggggc tcaggcggaa 240gatgaggctg actattactg taactcccgg gacagcagtg
gtaaccatca ttatgtcttc 300ggaactggga ccaaggtcac cgtcctaggt
33022233DNAHomo sapiens 222caaggagaca gcctcagaag
ctattatgca agc 3322321DNAHomo sapiens
223ggtaaaaaca accggccctc a
2122430DNAHomo sapiens 224aactcccggg acagcagtgg taaccatcat
30225678DNAHomo sapiens 225caggtgcagc tggtgcagtc
tggagctgag gtgaagaagc ctggggcctc agtgaaggtc 60tcctgtaagg cttctggtta
cacctttaat agctatggta ttacttgggt gcgacaggcc 120cctggacaag ggcttgagtg
gatgggatgg atcagcgttt acactggtaa gacaaactat 180gcacagaagt tccagggcag
agtcaccatg accacagaca catccacgag tacagcctac 240ctggacctga ggagcctgac
atctgacgac acggccgttt attactgtgc gagaggaggg 300gatcaccatg aatattgggg
ccagggaacc ctggtcaccg tctcgagatc ttctgagctg 360actcaggacc ctgctgtgtc
tgtggccttg ggacagacag tcaggatcac atgccaagga 420gacagcctca gaagctatta
tgcaagctgg taccagcaga agccaggaca ggcccctgta 480cttgtcatct atggtaaaaa
caaccggccc tcagggatcc cagaccgatt ctctggctcc 540agctcaggaa acacagcttc
cttgaccatc actggggctc aggcggaaga tgaggctgac 600tattactgta actcccggga
cagcagtggt aaccatcatt atgtcttcgg aactgggacc 660aaggtcaccg tcctaggt
67822615DNAHomo sapiens
226agctatggta ttact
1522751DNAHomo sapiens 227tggatcagcg tttacactgg taagacaaac tatgcacaga
agttccaggg c 5122821DNAHomo sapiens 228ggaggggatc accatgaata
t 21229321DNAHomo sapiens
229aattttatgc tgactcagcc actctcagtg tcagtggccc tgggacagac ggccaggatt
60acctgtgggg gaaacaacat tggaagtaaa aatgtgcact ggtaccagca gaagccaggc
120caggcccctg tgctggtcat ctatagggat agcaaccggc cctctgggat ccctgagcga
180ttctctggct ccaactcggg gaacacggcc accctgacca tcagcagagc ccaagccggg
240gatgaggctg actattactg tcaggtgtgg gacagcagca ctgtggtatt cggcggaggg
300accaagctga ccgtcctagg t
32123033DNAHomo sapiens 230gggggaaaca acattggaag taaaaatgtg cac
3323121DNAHomo sapiens 231agggatagca accggccctc t
2123221DNAHomo sapiens
232caggtgtggg acagcagcac t
21233384DNAHomo sapiens 233gaggtgcagc tggtggagtc tgggggaggc ttggtacagc
cgggggggtc cctgagactc 60tcctgtgcag cctctggatt cacctttagc agctatgcca
tgagctgggt ccgccaggct 120ccagggaagg ggctggagtg ggtctcaggt attagtggta
gtggtggtag aacatactac 180gcagactccg tgaagggccg gttcaccatc tccagagaca
attctaagaa cacgctgtat 240ctgcaaatga acagcctgag agccgaggac acggccgtat
attactgtgc gaaagatccc 300ctcgcattac gagattttga ctggttatcc cccgggcggg
actttgatta ctggggccag 360ggaaccctgg tcaccgtctc gaga
38423415DNAHomo sapiens 234agctatgcca tgagc
1523551DNAHomo sapiens
235ggtattagtg gtagtggtgg tagaacatac tacgcagact ccgtgaaggg c
5123657DNAHomo sapiens 236gatcccctcg cattacgaga ttttgactgg ttatcccccg
ggcgggactt tgattac 57237333DNAHomo sapiens 237cagtctgtgt
tgacgcagcc gccctcagtg tctgcggccc cgggacagaa ggtcaccatc 60tcctgctctg
gaagccactc caacattgga aataattatg tatcgtggtc ccagcaactc 120ccaggaacag
cccccaaact cctcatttat gacaataata agcgaccctc agggattcct 180gaccgattct
ctggctccaa gtctggcacg tcagccaccc tggacatcgc cgggctccag 240actggggacg
aggccgatta ttactgcgga gcatgggata ccagcctgag ttcttatgtc 300ttcggagctg
ggaccaaggt caccgtccta ggt 33323839DNAHomo
sapiens 238tctggaagcc actccaacat tggaaataat tatgtatcg
3923921DNAHomo sapiens 239gacaataata agcgaccctc a
2124027DNAHomo sapiens 240ggagcatggg
ataccagcct gagttct
27241360DNAHomo sapiens 241gaggtgcagc tggtggagtc tggggctgag gtgaagaagc
ctgggtcctc ggtgaaggtc 60tcctgcaagg cttctggagg caccttcagc agctatgcta
tcagctgggt gcgacaggcc 120cctggacaag ggcttgagtg gatgggaggg atcatcccta
tctttggtac agcaaactac 180gcacagaagt tccagggcag agtcacgatt accgcggacg
aatccacgag cacagcctac 240atggagctga gcagcctgag atctgaggac acggccgtgt
attactgtgc gagagaaggt 300ttatcgggtg ggtacggtat ggacgtctgg ggccaaggga
ccacggtcac cgtctcgagc 36024215DNAHomo sapiens 242agctatgcta tcagc
1524351DNAHomo sapiens
243gggatcatcc ctatctttgg tacagcaaac tacgcacaga agttccaggg c
5124433DNAHomo sapiens 244gaaggtttat cgggtgggta cggtatggac gtc
33245336DNAHomo sapiens 245cagtctgtgt tgacgcagcc
gccctcagtg tctggggccc cagggcagag ggtcaccatc 60tcctgcactg ggagcagctc
caacatcggg gcaggttatg atgtacactg gtaccagcag 120cttccaggaa cagcccccaa
actcctcatc tatcgtaaca acaatcggcc ctcaggggtc 180cctgaccgat tctctggctc
caactctggc acctcagcct ccctggccat cactgggctc 240cgggctgaag atgaggctga
ttattactgc cagtcctatg acagcagcct gagtagttat 300gtcttcggaa ctgggaccaa
ggtcaccgtc ctaggt 33624642DNAHomo sapiens
246actgggagca gctccaacat cggggcaggt tatgatgtac ac
4224721DNAHomo sapiens 247cgtaacaaca atcggccctc a
2124827DNAHomo sapiens 248cagtcctatg acagcagcct
gagtagt 27249375DNAHomo sapiens
249gaggtgcagc tggtggagtc tgggggaggc ttggtacagc ctggggggtc cctgagactc
60tcctgtgcag cctctggatt cacctttagc agctatgcca tgagctgggt ccgccaggct
120ccagggaagg ggctggagtg ggtctcagct attagtggta gtggtggtag cacatactac
180gcagactccg tgaagggccg gttcaccaac tccagagaca attccaagaa cacgctgtat
240ctgcaaatga acagcctgag agccgaggac acggccgtat attactgtgc gaaagcgtat
300tacgatattt tgactggtta tttttacaac ggtatggacg tctggggcca agggacaatg
360gtcaccgtct cgagc
37525015DNAHomo sapiens 250agctatgcca tgagc
1525151DNAHomo sapiens 251gctattagtg gtagtggtgg
tagcacatac tacgcagact ccgtgaaggg c 5125248DNAHomo sapiens
252gcgtattacg atattttgac tggttatttt tacaacggta tggacgtc
48253336DNAHomo sapiens 253caggctgtgc tcactcagcc gtcttccgtg tctgggtctc
ctggacagtc gatcaccatc 60tcctgcactg gaaccagcag tgacgttggt ggttatgact
atgtctcctg gtaccaacaa 120cacccaggca aagcccccaa actcatgatt catgatgtca
ggaatcggcc ctcaggggtt 180tctaatcgct tctctggctc caagtttggc aacacggcct
ccctgaccat ctctgggctc 240cagactgagg acgaggctga ttattactgc agttcatata
caagcagcag cactcatgtg 300ctattcggcg gagggaccaa gctgaccgtc ctaggt
33625442DNAHomo sapiens 254actggaacca gcagtgacgt
tggtggttat gactatgtct cc 4225521DNAHomo sapiens
255gatgtcagga atcggccctc a
2125627DNAHomo sapiens 256agttcatata caagcagcag cactcat
27257369DNAHomo sapiens 257caggtgcagc tggtgcagtc
tgggggaggc ttggtacagc ctggcaggtc cctgagactc 60tcctgtgcag cctctggatt
cacctttgat gattatgcca tgcactgggt ccggcaagct 120ccagggaagg gcctggagtg
ggtctcaggt attagttgga atagtggtag cataggctat 180gcggactctg tgaagggccg
attcaccatc tccagagaca acgccaagag ctccctgtat 240ctgcaaatga acagtctgag
agctgaggac acggccttgt attactgtgc aaaaggccac 300tctccgtata gcagtggctg
gtctgacttt gactactggg gccagggaac cctggtcacc 360gtctcgagc
36925815DNAHomo sapiens
258gattatgcca tgcac
1525951DNAHomo sapiens 259ggtattagtt ggaatagtgg tagcataggc tatgcggact
ctgtgaaggg c 5126042DNAHomo sapiens 260ggccactctc cgtatagcag
tggctggtct gactttgact ac 42261336DNAHomo sapiens
261cagtctgtgt tgacgcagcc gccctcagtg tctggggccc cagggcagag ggtcaccatc
60tcctgcaccg ggagcagctc caacatcggg gcaggttatg atgttcagtg gtaccagcag
120ctcccaggaa cagcccccaa actcctcatc catgctaaca agaatcggcc ctcaggggtc
180cctgaccgaa tctctggctc caagtctggc accacagcct ccctggccat cactgggttc
240caggctgagg atgaggctga ttattactgc cagtcctatg acagcagcct gactggttat
300gtcttcggaa ctgggaccaa ggtcaccgtc ctaggt
33626242DNAHomo sapiens 262accgggagca gctccaacat cggggcaggt tatgatgttc ag
4226321DNAHomo sapiens 263gctaacaaga atcggccctc a
2126427DNAHomo sapiens
264cagtcctatg acagcagcct gactggt
27265375DNAHomo sapiens 265gaggtgcagc tggtggagtc tgggggaggc ttggtacagc
ctggcaggtc cctgagactc 60tcctgtgcag cctctggatt cacctttgct gattatgcca
tgcactgggt ccggcaagct 120ccagggaagg gcctggagtg ggtctcaagt attagttgga
atagtggtag catagcctat 180gcggactctg tgaagggccg attcaccatc tccagagaca
acgccaagaa ctccctgtat 240ctgcaaatga acagtctgag agctgaggac acggccttgt
attactgtgc aaaagcctca 300gcagctggta ctgaatacta ccactactac ggtatggacg
tctggggcca agggaccacg 360gtcaccgtct cgaga
37526615DNAHomo sapiens 266gattatgcca tgcac
1526751DNAHomo sapiens
267agtattagtt ggaatagtgg tagcatagcc tatgcggact ctgtgaaggg c
5126848DNAHomo sapiens 268gcctcagcag ctggtactga atactaccac tactacggta
tggacgtc 48269333DNAHomo sapiens 269tcctatgtgc
tgactcagcc accctcagcg tctgggaccc ccgggcagag ggtcaccatc 60tcttgttctg
gaagcagctc caacatcgga agtaatacta taaactggta ccagcagctc 120ccaggaacgg
cccccaaact cctcatctat aataatcatc agcggccctc aggggtccct 180gaccgattct
ctggctcaaa gtctggcacc tcagcctccc tggccatcag tgggctccag 240tctgcggatg
aggctgatta ttactgtgga gcgtggaatg acagcctgaa tgtctatgtc 300ttcggaactg
ggaccaaggt caccgtccta ggt 33327039DNAHomo
sapiens 270tctggaagca gctccaacat cggaagtaat actataaac
3927121DNAHomo sapiens 271aataatcatc agcggccctc a
2127227DNAHomo sapiens 272ggagcgtgga
atgacagcct gaatgtc
27273372DNAHomo sapiens 273caggtgcagc tggtgcagtc tggagctgag gtgaagaagc
ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta cacctttacc agctatggta
tcagctgggt gcgacaggcc 120cctggacaag ggcttgagtg gatgggatgg atcagcgctt
acaatggtaa cacaaactat 180gcacagaagc tccagggcag agtcaccatg accacagaca
catccacgag cacagcctac 240atggagctga ggagcctgag atctgacgac acggccgtgt
attactgtgc gacagcagca 300agcctaaagt attactatga tagtagtggt tattactact
ggggccaggg aaccctggtc 360accgtctcga ga
37227415DNAHomo sapiens 274agctatggta tcagc
1527551DNAHomo sapiens
275tggatcagcg cttacaatgg taacacaaac tatgcacaga agctccaggg c
5127645DNAHomo sapiens 276gcagcaagcc taaagtatta ctatgatagt agtggttatt
actac 45277330DNAHomo sapiens 277tcttctgagc
tgactcagga ccctgctgtg tctgtggcct tgggacagac agtcaggatc 60acatgccaag
gagacagcct cagaagctat tatgcaagct ggtaccagca gaagccagga 120caggcccctg
tacttgtcat ctctggtaaa aacaaccggc cctcagggat cccagaccga 180ttctctggct
ccagctcagg agacacagct tccttgacca tcactggggc tcaggcggaa 240gatgaggcta
actattactg taactctcgg gacagcagtg gttacccctc ttgggtgttc 300ggcggaggga
ccaagctgac cgtcctaggc 33027833DNAHomo
sapiens 278caaggagaca gcctcagaag ctattatgca agc
3327921DNAHomo sapiens 279ggtaaaaaca accggccctc a
2128030DNAHomo sapiens 280aactctcggg
acagcagtgg ttacccctct
30281390DNAHomo sapiens 281caggtgcagc tggtgcagtc tggggctgag gtgaagaagc
ctggggcctc agtgaaggtc 60tcctgcaagg gttctggtta cgtttttaac agttatggta
ttacctgggt gcgacaggcc 120ccaggacaag ggcttgagtg gatgggatgg atcagcgctt
acaatggtta cacagactat 180gcacagaagg tccagggcag agtcaccatg accacagaga
catccacgag cacagcctac 240atggagctga ggagcctgag atctgacgac acggccgttt
attattgtgc gagggattac 300tatgatagta gtacttatta ctccagtgat tacttccagt
actacggtat ggacgtctgg 360ggccaaggga ccacggtcac cgtctcgagc
39028215DNAHomo sapiens 282agttatggta ttacc
1528351DNAHomo sapiens
283tggatcagcg cttacaatgg ttacacagac tatgcacaga aggtccaggg c
5128463DNAHomo sapiens 284gattactatg atagtagtac ttattactcc agtgattact
tccagtacta cggtatggac 60gtc
63285333DNAHomo sapiens 285caggctgtgc tgactcagcc
ggcttccgtg tctgggtctc ctggacagtc gatcaccatc 60tcctgcactg gaaccagcag
tgacgttggt gcttataact atgtctcctg gtaccaacaa 120cacccaggca aagcccccaa
actcatgatt tctgatgtca gtaggcggcc ctcaggggtt 180tctaatcgct tctctggctc
caagtctggc aacacggcct ccctgaccat ctctgggctc 240cagactgagg acgaggctga
ttattactgc agctcatata caagcagcaa cactgtctta 300ttcggcggag ggaccaagct
gaccgtccta ggt 33328642DNAHomo sapiens
286actggaacca gcagtgacgt tggtgcttat aactatgtct cc
4228721DNAHomo sapiens 287gatgtcagta ggcggccctc a
2128824DNAHomo sapiens 288agctcatata caagcagcaa
cact 24289333DNAHomo sapiens
289caggctgtgc tgactcagcc ggcttccgtg tctgggtctc ctggacagtc gatcaccatc
60tcctgcactg gaaccagcag tgacgttggt gcttataact atgtctcctg gtaccaacaa
120cacccaggca aagcccccaa actcatgatt tctgatgtca gtaggcggcc ctcaggggtt
180tctaatcgct tctctggctc caagtctggc aacacggcct ccctgaccat ctctgggctc
240cagactgagg acgaggctga ttattactgc agctcatata caagcagcaa cactgtctta
300ttcggcggag ggaccaagct gaccgtccta ggt
33329015DNAHomo sapiens 290aactactgga tcggc
1529151DNAHomo sapiens 291atcatctatc ctggtgactc
tgataccagg tacagtccgt cattccaagg c 5129239DNAHomo sapiens
292cggggctccc gtagtagtgg tgaagatgct tttgaagtc
39293330DNAHomo sapiens 293tcctatgagc tgactcagcc accctcagtg tcagtggccc
caggaaagac ggccacaatt 60acgtgtgggg gcgacaacat tggaagtaag agtgtgcact
ggtaccagca gaggccaggc 120caggcccctg tgttggtcat caattatgat agtgaccggc
cctcagggat ccctgagcga 180ttctctggct ccaactctgg gaacacggcc accctgacca
tcagcagggt cgaagccggg 240gatgaggccg actattactg tcaggtggaa gatcgccgtg
gtggttatca tgtggtattc 300ggcggaggga ccaagctgac cgtcctaggt
33029433DNAHomo sapiens 294gggggcgaca acattggaag
taagagtgtg cac 3329521DNAHomo sapiens
295tatgatagtg accggccctc a
2129630DNAHomo sapiens 296caggtggaag atcgccgtgg tggttatcat
30297375DNAHomo sapiens 297caggtgcagc tggtgcagtc
tgggggaggc ttggtccagc ctggggggtc cctgagactc 60tcctgtgcag cctctggatt
caccgtcagt agcaactaca tgagctgggt ccgccaggct 120ccagggaagg ggctggagtg
ggtctcagtt atttatagcg gtggtagcac atactacgca 180gactccgtga agggcagatt
caccatctcc agagacaatt ccaagaacac gctgtatctt 240caaatgaaca gcctgagagc
cgaggacacg gctgtgtatt actgtgcgag ggaaggatat 300tgtagtggtg gtagctgcta
ctcctacggc gcttttgata tctggggcca agggaccacg 360gtcaccgtct cgagc
37529815DNAHomo sapiens
298agcaactaca tgagc
1529948DNAHomo sapiens 299gttatttata gcggtggtag cacatactac gcagactccg
tgaagggc 4830051DNAHomo sapiens 300gaaggatatt gtagtggtgg
tagctgctac tcctacggcg cttttgatat c 51301375DNAHomo sapiens
301caggtgcagc tggtgcagtc tgggggaggc ttggtccagc ctggggggtc cctgagactc
60tcctgtgcag cctctggatt caccgtcagt agcaactaca tgagctgggt ccgccaggct
120ccagggaagg ggctggagtg ggtctcagtt atttatagcg gtggtagcac atactacgca
180gactccgtga agggcagatt caccatctcc agagacaatt ccaagaacac gctgtatctt
240caaatgaaca gcctgagagc cgaggacacg gctgtgtatt actgtgcgag ggaaggatat
300tgtagtggtg gtagctgcta ctcctacggc gcttttgata tctggggcca agggaccacg
360gtcaccgtct cgagc
37530233DNAHomo sapiens 302cgggcgagtc agaatattgc caactggtta gcc
3330321DNAHomo sapiens 303gctgcatcca atttgcaaag t
2130424DNAHomo sapiens
304caacagggta acagtttccc tcgg
24305390DNAHomo sapiens 305caggtgcagc tggtgcagtc tggagctgag gtgaagaagc
ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta cgtttttaac agttatggta
ttacctgggt gcgacaggcc 120ccaggacaag ggcttgagtg gatgggatgg atcagcgctt
acaatggtta cacagactat 180gcacagaagg tccagggcag agtcaccatg accacagaga
catccacgag cacagcctac 240atggagctga ggagcctgag atctgacgac acggccgttt
attattgtgc gagggactac 300tatgatagta gtacttatta ctccagtgat tacttcaagt
actacggtat ggacgtctgg 360ggccaaggga ccacggtcac cgtctcgagc
39030615DNAHomo sapiens 306agttatggta ttacc
1530751DNAHomo sapiens
307tggatcagcg cttacaatgg ttacacagac tatgcacaga aggtccaggg c
5130863DNAHomo sapiens 308gactactatg atagtagtac ttattactcc agtgattact
tcaagtacta cggtatggac 60gtc
63309333DNAHomo sapiens 309caggctgtgc tcactcagcc
gtcttccgtg tctgggtctc ctggacagtc gatcaccatc 60tcctgctctg gaaccagcag
tgacgttggt gcttataact atgtctcctg gtaccaacaa 120cacccaggca aagcccccag
actcctgact tttgatgtca ataggcgtcc ctcagggtct 180tctagtcgct tctctggctc
caagtctggc aacacggcct ccctgactat ctctgggctc 240caggctgagg acgaggctga
ctattactgc agttcatata caaacagcaa cactgtcgtg 300ttcggcggag ggaccaggct
gaccgtccta agt 33331042DNAHomo sapiens
310tctggaacca gcagtgacgt tggtgcttat aactatgtct cc
4231121DNAHomo sapiens 311gatgtcaata ggcgtccctc a
2131224DNAHomo sapiens 312agttcatata caaacagcaa
cact 24313381DNAHomo sapiens
313caggtgcagc tgcaggagtc ggggccagga ctggtgaagc cgtcggagac cctggccctc
60acctgcactg tctctggtgg ctccatcagt aactactact ggagttggat ccggcagccc
120ccagggaagg gactggagtg gattgggtat atctatgaca ttgagaatac caactacaac
180ccctccctca agagtcgagt caccatatca gtggacacgt ccaagaacca gttctccctg
240aagttgagct ctgtgaccgc tgatgacacg gccgtatatt actgtgcgag agattcaagg
300gtcattcgat ttttggaggg gtactcctac tactacggtg tggacgtctg gggccaaggg
360acaatggtca ccgtctcgag c
38131415DNAHomo sapiens 314aactactact ggagt
1531548DNAHomo sapiens 315tatatctatg acattgagaa
taccaactac aacccctccc tcaagagt 4831657DNAHomo sapiens
316gattcaaggg tcattcgatt tttggagggg tactcctact actacggtgt ggacgtc
57317333DNAHomo sapiens 317tcctatgagc tgactcagcc accctcagcg tctgggaccc
ccgggcagac ggtcatcatc 60tcttgttctg gaagcaggtc caacatcgga ggtcatggtg
taaattggca ccagcaggtt 120ccaggaacgg cccccaaact cctcatctac cgtaatgatc
gccggccctc aggggtcccg 180gaccgattct ctggctccaa gtctggcact tcagcctccc
tggtcatcag tggactgcag 240tttgaggatg aggctgatta ttactgtgta gcatgggaag
acagcctgga tggtccggtg 300ttcggcggag ggaccaagct gactgtccta ggt
33331839DNAHomo sapiens 318tctggaagca ggtccaacat
cggaggtcat ggtgtaaat 3931921DNAHomo sapiens
319cgtaatgatc gccggccctc a
2132027DNAHomo sapiens 320gtagcatggg aagacagcct ggatggt
27321345DNAHomo sapiens 321gaggtgcagc tggtggagtc
tgggggaggc ttggtacagc ctggggggtc cctgagactc 60tcctgtgcag cctctggatt
cacctttagc agctatgcca tgagctgggt ccgccaggct 120ccagggaagg ggctggagtg
ggtctcaact attagtggta gtggtggtag tacatactac 180gcagactccg tgaagggccg
attcaccatc tccagagaca attccaagaa cacgctatat 240cttcaaatga acagcctgag
agccgaggac acggccgtct attattgtgc gagaggtata 300gtggctacta gctggggcca
gggaaccctg gtcaccgtct cgaga 34532215DNAHomo sapiens
322agctatgcca tgagc
1532351DNAHomo sapiens 323actattagtg gtagtggtgg tagtacatac tacgcagact
ccgtgaaggg c 5132418DNAHomo sapiens 324ggtatagtgg ctactagc
18325324DNAHomo sapiens
325tcctatgagc tgactcagcc accctcagtg tcagtggccc caggacagac ggccagaatt
60acctgtgggg gaaacaagat tggaagcaaa agtgtgcact ggtaccagca gaagcaaggc
120caggcccctg tattggtcat ctatttggat cgcgaccggc cctcagggat ccctgaacga
180ttctctggct ccaactctgg gaacacggcc accctgacca tcaccagggt cgaagccgag
240gatgaggccg actattattg tcacctgtgg gatagtggta gtgatcaggt gttcggcgga
300gggaccaaac tgaccgtcct gggt
32432633DNAHomo sapiens 326gggggaaaca agattggaag caaaagtgtg cac
3332721DNAHomo sapiens 327ttggatcgcg accggccctc a
2132824DNAHomo sapiens
328cacctgtggg atagtggtag tgat
24329342DNAHomo sapiens 329gaggtgcagc tggtggagtc tgggggaggc gtagtacagc
ctggggggtc cctgagactc 60tcctgtgcag cctcaggatt caactttgat gtttatggca
tgaactgggt ccgtcaagtt 120ccagggaagg gtctggagtg ggtctctctt atcaacgggg
atggcggttt aagatattac 180gcagactctg tgaagggccg attcaccgtc tccagagaca
acagcaggaa ttccctatat 240ctgcaaatga acagtctcag aagtgaggac accgccctgt
attattgtgt aaagggaaac 300ttccagcagt ggggccaggg aaccctggtc accgtctcga
ga 34233015DNAHomo sapiens 330gtttatggca tgaac
1533151DNAHomo sapiens
331cttatcaacg gggatggcgg tttaagatat tacgcagact ctgtgaaggg c
5133215DNAHomo sapiens 332ggaaacttcc agcag
15333327DNAHomo sapiens 333tcctatgtgc tgactcagcc
accctcagtg tcagtggccc caggaaagac ggccaggatt 60acctgtgggg gaaacaacat
tggaagtaaa agtgtgcact ggtaccagca gaagccaggc 120caggcccctg tgctggtcat
ctattatgat agcgaccggc cctcagggat ccctgagcga 180ttctctggct ccaactctgg
gaacacggcc accctgacca tcagcagggt cgaagccggg 240gatgaggccg actattactg
tcaggtgtgg gatagtagta gtgatcatgt ggtattcggc 300ggagggacca agctgaccgt
cctaggt 32733433DNAHomo sapiens
334gggggaaaca acattggaag taaaagtgtg cac
3333521DNAHomo sapiens 335tatgatagcg accggccctc a
2133627DNAHomo sapiens 336caggtgtggg atagtagtag
tgatcat 27337345DNAHomo sapiens
337gaggtgcagc tggtggagtc tgggggaggc ttggtacagc ctggggggtc cctgagactc
60tcctgtgcag cctctggatt cacctttgat gattatgcca tgcactgggt ccgtcaagct
120ccagggaagg gtctggagtg ggtctctctt attagtgggg atggtggtag cacatactat
180gcagactctg tgaaggaccg attcaccatc tccagagaca acagcaaaaa ctccctgtat
240ctgcaaatga acagcctgag agccgaggac acggccgtgt attactgtgc gaggggaaac
300tactttgact actggggcca gggaaccctg gtcaccgtct cgaga
34533815DNAHomo sapiens 338gattatgcca tgcac
1533951DNAHomo sapiens 339cttattagtg gggatggtgg
tagcacatac tatgcagact ctgtgaagga c 5134018DNAHomo sapiens
340ggaaactact ttgactac
18341318DNAHomo sapiens 341tcctatgagc tgactcagcc actctcagtg tcagtggccc
tgggacagac ggccaggatt 60acctgtgggg gaaacaacat tggaagtaaa aatgtgcact
ggtaccagca gaagccaggc 120caggcccctg tgctggtcat ctatagggat agcaaccggc
cctctgggat ccctgagcga 180ttctctggct ccaactcggg gaacacggcc accctgacca
tcagcagagc ccaagccggg 240gatgaggctg actattactg tcaggtgtgg gacagcagcg
tggtattcgg cggagggacc 300aagctgaccg tcctaggt
31834233DNAHomo sapiens 342gggggaaaca acattggaag
taaaaatgtg cac 3334321DNAHomo sapiens
343agggatagca accggccctc t
2134418DNAHomo sapiens 344caggtgtggg acagcagc
18345366DNAHomo sapiens 345gaggtgcagc tggtggagtc
tgggggaggc ttggtacagc ctggggggtc cctgagactc 60tcctgtgcag cctctggatt
caccttcagt agctatagca tgaactgggt ccgccaggct 120ccagggaagg ggctggagtg
ggtttcatac attagtagta gtagtagtac catatactac 180gcagactctg tgaagggccg
attcaccatc tccagagaca atgccaagaa ctcactgtat 240ctgcagatga acagcctgag
agacgaggac acggctgtgt attactgtgc gagagtgatg 300ccgagttact actactacta
cggtatggac gtctggggcc aagggaccac ggtcaccgtc 360tcgaga
36634615DNAHomo sapiens
346agctatagca tgaac
1534751DNAHomo sapiens 347tacattagta gtagtagtag taccatatac tacgcagact
ctgtgaaggg c 5134839DNAHomo sapiens 348gtgatgccga gttactacta
ctactacggt atggacgtc 39349333DNAHomo sapiens
349aattttatgc tgactcagcc ccactctgtg tcggagtctc cggggaagac ggtaaccatc
60tcctgcaccg gcagcagtgg cagcattgcc agcaactatg tgcagtggta ccagcagcgc
120ccgggcagtg cccccaccac tgtgatctat gaggatagtg aaagaccctc tggggtccct
180gatcggttct ctggctccat cgacagctcc tccaactctg cctccctcac catctctgga
240ctgaagactc aggacgaggc tgactactac tgtcagtctt atgatggcgt caattgggtg
300ttcggcggag ggaccaagct gaccgtccta ggt
33335039DNAHomo sapiens 350accggcagca gtggcagcat tgccagcaac tatgtgcag
3935121DNAHomo sapiens 351gaggatagtg aaagaccctc t
2135221DNAHomo sapiens
352cagtcttatg atggcgtcaa t
21353342DNAHomo sapiens 353gaggtgcagc tggtggagtc tgggggaggc gtggtccagc
ctgggaggtc cctgagactc 60tcctgtgcag cctctggatt caccttcagt agctatggca
tgcactgggt ccgccaggct 120ccaggcaagg ggctggagtg ggtggcagtt atatcatatg
atggaagtaa taaatactat 180gcagactccg tgaagggccg attcaccatc tccagagaca
attccaagaa cacgctgtat 240ctgcaaatga acagcctgag agctgaggac acggctgtgt
attactgtgc ggactacggt 300atggacgtct ggggccaagg gaccacggtc accgtctcga
ga 34235415DNAHomo sapiens 354agctatggca tgcac
1535551DNAHomo sapiens
355gttatatcat atgatggaag taataaatac tatgcagact ccgtgaaggg c
5135615DNAHomo sapiens 356tacggtatgg acgtc
15357321DNAHomo sapiens 357gacatccaga tgacccagtc
tccatcctcc ctgtctgcat ctgtaggaga cagagtcacc 60atcacttgcc gggcaagtca
gagcattagc agctatttaa attggtatca gcagaaacca 120gggaaagccc ctaagctcct
gatctatgct gcatccagtt tgcaaagtgg ggtcccatca 180aggttcagtg gcagtggatc
tgggactgat ttcactctca ccatcagcag tctgcaacct 240gaagattttg caacttacta
ctgtcaacag agttacagta cccccgtgac gttcggccaa 300gggaccaagg tggaaatcaa a
32135833DNAHomo sapiens
358cgggcaagtc agagcattag cagctattta aat
3335921DNAHomo sapiens 359gctgcatcca gtttgcaaag t
2136021DNAHomo sapiens 360caacagagtt acagtacccc c
21361360DNAHomo sapiens
361caggtgcagc tgcaggagtc gggcccggga ctggtgaagc cttcacagac cctgtccctc
60acctgcactg tctctggtgg ctccatcagc agtggtggtt actactggag ctggatccgc
120cagcacccag ggaagggcct ggagtggatt gggtacatct attacagtgg gagcacctac
180tacaacccgt ccctcaagag tcgagttacc atatcagtag acacgtctaa gaaccagttc
240tccctgaagc tgagctctgt gactgccgcg gacacggccg tgtattactg tgcgagagat
300cgggggactg gggatgcttt tgatatctgg ggccaaggga caatggtcac cgtctcgaga
36036221DNAHomo sapiens 362agtggtggtt actactggag c
2136348DNAHomo sapiens 363tacatctatt acagtgggag
cacctactac aacccgtccc tcaagagt 4836430DNAHomo sapiens
364gatcggggga ctggggatgc ttttgatatc
30365333DNAHomo sapiens 365cagtctgtgt tgacgcagcc gccctcggtg tctggggccc
cccggcagac ggtcaccatc 60tcctgctctg ggagcagctc caacatcgga caaaattctg
ttacctggta ccagcgcctc 120ccgggtgagg ctcccaaact cctcatctac tatgatgatc
tcttgcactc aggagtctct 180gaccgattct ctggctccaa gtctggcacc tcagcctcac
tggccatcag tggactccag 240tctgaggatg aggctgagta ctactgtgcg tcatgggatg
acagcctgaa aggtccggta 300ttcggcggag ggaccaaact gaccgtccta ggt
33336639DNAHomo sapiens 366tctgggagca gctccaacat
cggacaaaat tctgttacc 3936721DNAHomo sapiens
367tatgatgatc tcttgcactc a
2136827DNAHomo sapiens 368gcgtcatggg atgacagcct gaaaggt
273691200PRTHomo sapiens 369Met Arg Pro Ser Gly Thr
Ala Gly Ala Ala Leu Leu Ala Leu Leu Ala1 5
10 15Ala Leu Cys Pro Ala Ser Arg Ala Leu Glu Glu Lys
Lys Val Cys Gln 20 25 30Gly
Thr Ser Asn Lys Leu Thr Gln Leu Gly Thr Phe Glu Asp His Phe 35
40 45Leu Ser Leu Gln Arg Met Phe Asn Asn
Cys Glu Val Val Leu Gly Asn 50 55
60Leu Glu Ile Thr Tyr Val Gln Arg Asn Tyr Asp Leu Ser Phe Leu Lys65
70 75 80Thr Ile Gln Glu Val
Ala Gly Tyr Val Leu Ile Ala Leu Asn Thr Val 85
90 95Glu Arg Ile Pro Leu Glu Asn Leu Gln Ile Ile
Arg Gly Asn Met Tyr 100 105
110Tyr Glu Asn Ser Tyr Ala Leu Ala Gly Leu Lys Glu Leu Pro Met Arg
115 120 125Asn Leu Gln Glu Ile Leu His
Gly Ala Val Arg Phe Ser Asn Asn Pro 130 135
140Ala Leu Cys Asn Val Glu Ser Ile Gln Trp Arg Asp Ile Val Ser
Ser145 150 155 160Asp Phe
Leu Ser Asn Met Ser Met Asp Phe Gln Asn His Leu Gly Ser
165 170 175Cys Gln Lys Cys Asp Pro Ser
Cys Pro Asn Gly Ser Cys Trp Gly Ala 180 185
190Gly Glu Glu Asn Cys Gln Lys Leu Thr Lys Ile Ile Cys Ala
Gln Gln 195 200 205Cys Ser Gly Arg
Cys Arg Gly Lys Ser Pro Ser Asp Cys Cys His Asn 210
215 220Gln Cys Ala Ala Gly Cys Thr Gly Pro Arg Glu Ser
Asp Cys Leu Val225 230 235
240Cys Arg Lys Phe Arg Asp Glu Ala Thr Cys Lys Asp Thr Cys Pro Pro
245 250 255Leu Met Leu Tyr Asn
Pro Thr Thr Tyr Gln Met Asp Val Asn Pro Glu 260
265 270Gly Lys Tyr Ser Phe Gly Ala Thr Cys Val Lys Lys
Cys Pro Arg Asn 275 280 285Tyr Val
Val Thr Asp His Gly Ser Cys Val Arg Ala Cys Gly Ala Asp 290
295 300Ser Tyr Glu Met Glu Glu Asp Gly Val Arg Lys
Cys Lys Lys Cys Glu305 310 315
320Gly Pro Cys Arg Lys Val Cys Asn Gly Ile Gly Ile Gly Glu Phe Lys
325 330 335Asp Ser Leu Ser
Ile Asn Ala Thr Asn Ile Lys His Phe Lys Asn Cys 340
345 350Thr Ser Ile Ser Gly Asp Leu His Ile Leu Pro
Val Ala Phe Arg Gly 355 360 365Asp
Ser Phe Thr His Thr Pro Pro Leu Asp Pro Gln Glu Leu Asp Ile 370
375 380Leu Lys Thr Val Lys Glu Ile Thr Gly Phe
Leu Leu Ile Gln Ala Trp385 390 395
400Pro Glu Asn Arg Thr Asp Leu His Ala Phe Glu Asn Leu Glu Ile
Ile 405 410 415Arg Gly Arg
Thr Lys Gln His Gly Gln Phe Ser Leu Ala Val Val Ser 420
425 430Leu Asn Ile Thr Ser Leu Gly Leu Arg Ser
Leu Lys Glu Ile Ser Asp 435 440
445Gly Asp Val Ile Ile Ser Gly Asn Lys Asn Leu Cys Tyr Ala Asn Thr 450
455 460Ile Asn Trp Lys Lys Leu Phe Gly
Thr Ser Gly Gln Lys Thr Lys Ile465 470
475 480Ile Ser Asn Arg Gly Glu Asn Ser Cys Lys Ala Thr
Gly Gln Val Cys 485 490
495His Ala Leu Cys Ser Pro Glu Gly Cys Trp Gly Pro Glu Pro Arg Asp
500 505 510Cys Val Ser Cys Arg Asn
Val Ser Arg Gly Arg Glu Cys Val Asp Lys 515 520
525Cys Asn Leu Leu Glu Gly Glu Pro Arg Glu Phe Val Glu Asn
Ser Glu 530 535 540Cys Ile Gln Cys His
Pro Glu Cys Leu Pro Gln Ala Met Asn Ile Thr545 550
555 560Cys Thr Gly Arg Gly Pro Asp Asn Cys Ile
Gln Cys Ala His Tyr Ile 565 570
575Asp Gly Pro His Cys Val Lys Thr Cys Pro Ala Gly Val Met Gly Glu
580 585 590Asn Asn Thr Leu Val
Trp Lys Tyr Ala Asp Ala Gly His Val Cys His 595
600 605Leu Cys His Pro Asn Cys Thr Tyr Gly Cys Thr Gly
Pro Gly Leu Glu 610 615 620Gly Cys Pro
Thr Asn Gly Pro Lys Ile Pro Ser Ile Ala Thr Gly Met625
630 635 640Val Gly Ala Leu Leu Leu Leu
Leu Val Val Ala Leu Gly Ile Gly Leu 645
650 655Phe Met Arg Arg Arg His Ile Val Arg Lys Arg Thr
Leu Arg Arg Leu 660 665 670Leu
Gln Glu Arg Glu Leu Val Glu Pro Leu Thr Pro Ser Gly Glu Ala 675
680 685Pro Asn Gln Ala Leu Leu Arg Ile Leu
Lys Glu Thr Glu Phe Lys Lys 690 695
700Ile Lys Val Leu Gly Ser Gly Ala Phe Gly Thr Val Tyr Lys Gly Leu705
710 715 720Trp Ile Pro Glu
Gly Glu Lys Val Lys Ile Pro Val Ala Ile Lys Glu 725
730 735Leu Arg Glu Ala Thr Ser Pro Lys Ala Asn
Lys Glu Ile Leu Asp Glu 740 745
750Ala Tyr Val Met Ala Ser Val Asp Asn Pro His Val Cys Arg Leu Leu
755 760 765Gly Ile Cys Leu Thr Ser Thr
Val Gln Leu Ile Thr Gln Leu Met Pro 770 775
780Phe Gly Cys Leu Leu Asp Tyr Val Arg Glu His Lys Asp Asn Ile
Gly785 790 795 800Ser Gln
Tyr Leu Leu Asn Trp Cys Val Gln Ile Ala Lys Gly Met Asn
805 810 815Tyr Leu Glu Asp Arg Arg Leu
Val His Arg Asp Leu Ala Ala Arg Asn 820 825
830Val Leu Val Lys Thr Pro Gln His Val Lys Ile Thr Asp Phe
Gly Leu 835 840 845Ala Lys Leu Leu
Gly Ala Glu Glu Lys Glu Tyr His Ala Glu Gly Gly 850
855 860Lys Val Pro Ile Lys Trp Met Ala Leu Glu Ser Ile
Leu His Arg Ile865 870 875
880Tyr Thr His Gln Ser Asp Val Trp Ser Tyr Gly Val Thr Val Trp Glu
885 890 895Leu Met Thr Phe Gly
Ser Lys Pro Tyr Asp Gly Ile Pro Ala Ser Glu 900
905 910Ile Ser Ser Ile Leu Glu Lys Gly Glu Arg Leu Pro
Gln Pro Pro Ile 915 920 925Cys Thr
Ile Asp Val Tyr Met Ile Met Val Lys Cys Trp Met Ile Asp 930
935 940Ala Asp Ser Arg Pro Lys Phe Arg Glu Leu Ile
Ile Glu Phe Ser Lys945 950 955
960Met Ala Arg Asp Pro Gln Arg Tyr Leu Val Ile Gln Gly Asp Glu Arg
965 970 975Met His Leu Pro
Ser Pro Thr Asp Ser Asn Phe Tyr Arg Ala Leu Met 980
985 990Asp Glu Glu Asp Met Asp Asp Val Val Asp Ala
Asp Glu Tyr Leu Ile 995 1000
1005Pro Gln Gln Gly Phe Phe Ser Ser Pro Ser Thr Ser Arg Thr Pro
1010 1015 1020Leu Leu Ser Ser Leu Ser
Ala Thr Ser Asn Asn Ser Thr Val Ala 1025 1030
1035Cys Ile Asp Arg Asn Gly Leu Gln Ser Cys Pro Ile Lys Glu
Asp 1040 1045 1050Ser Phe Leu Gln Arg
Tyr Ser Ser Asp Pro Thr Gly Ala Leu Thr 1055 1060
1065Glu Asp Ser Ile Asp Asp Thr Phe Leu Pro Val Pro Glu
Tyr Ile 1070 1075 1080Asn Gln Ser Val
Pro Lys Arg Pro Ala Gly Ser Val Gln Asn Pro 1085
1090 1095Val Tyr His Asn Gln Pro Leu Asn Pro Ala Pro
Ser Arg Asp Pro 1100 1105 1110His Tyr
Gln Asp Pro His Ser Thr Ala Val Gly Asn Pro Glu Tyr 1115
1120 1125Leu Asn Thr Val Gln Pro Thr Cys Val Asn
Ser Thr Phe Asp Ser 1130 1135 1140Pro
Ala His Trp Ala Gln Lys Gly Ser His Gln Ile Ser Leu Asp 1145
1150 1155Asn Pro Asp Tyr Gln Gln Asp Phe Phe
Pro Lys Glu Ala Lys Pro 1160 1165
1170Asn Gly Ile Phe Lys Gly Ser Thr Ala Glu Asn Ala Glu Tyr Leu
1175 1180 1185Arg Val Ala Pro Gln Ser
Ser Glu Phe Ile Gly Ala 1190 1195
12003701255PRTHomo sapiens 370Met Glu Leu Ala Ala Leu Cys Arg Trp Gly Leu
Leu Leu Ala Leu Leu1 5 10
15Pro Pro Gly Ala Ala Ser Thr Gln Val Cys Thr Gly Thr Asp Met Lys
20 25 30Leu Arg Leu Pro Ala Ser Pro
Glu Thr His Leu Asp Met Leu Arg His 35 40
45Leu Tyr Gln Gly Cys Gln Val Val Gln Gly Asn Leu Glu Leu Thr
Tyr 50 55 60Leu Pro Thr Asn Ala Ser
Leu Ser Phe Leu Gln Asp Ile Gln Glu Val65 70
75 80Gln Gly Tyr Val Leu Ile Ala His Asn Gln Val
Arg Gln Val Pro Leu 85 90
95Gln Arg Leu Arg Ile Val Arg Gly Thr Gln Leu Phe Glu Asp Asn Tyr
100 105 110Ala Leu Ala Val Leu Asp
Asn Gly Asp Pro Leu Asn Asn Thr Thr Pro 115 120
125Val Thr Gly Ala Ser Pro Gly Gly Leu Arg Glu Leu Gln Leu
Arg Ser 130 135 140Leu Thr Glu Ile Leu
Lys Gly Gly Val Leu Ile Gln Arg Asn Pro Gln145 150
155 160Leu Cys Tyr Gln Asp Thr Ile Leu Trp Lys
Asp Ile Phe His Lys Asn 165 170
175Asn Gln Leu Ala Leu Thr Leu Ile Asp Thr Asn Arg Ser Arg Ala Cys
180 185 190His Pro Cys Ser Pro
Met Cys Lys Gly Ser Arg Cys Trp Gly Glu Ser 195
200 205Ser Glu Asp Cys Gln Ser Leu Thr Arg Thr Val Cys
Ala Gly Gly Cys 210 215 220Ala Arg Cys
Lys Gly Pro Leu Pro Thr Asp Cys Cys His Glu Gln Cys225
230 235 240Ala Ala Gly Cys Thr Gly Pro
Lys His Ser Asp Cys Leu Ala Cys Leu 245
250 255His Phe Asn His Ser Gly Ile Cys Glu Leu His Cys
Pro Ala Leu Val 260 265 270Thr
Tyr Asn Thr Asp Thr Phe Glu Ser Met Pro Asn Pro Glu Gly Arg 275
280 285Tyr Thr Phe Gly Ala Ser Cys Val Thr
Ala Cys Pro Tyr Asn Tyr Leu 290 295
300Ser Thr Asp Val Gly Ser Cys Thr Leu Val Cys Pro Leu His Asn Gln305
310 315 320Glu Val Thr Ala
Glu Asp Gly Thr Gln Arg Cys Glu Lys Cys Ser Lys 325
330 335Pro Cys Ala Arg Val Cys Tyr Gly Leu Gly
Met Glu His Leu Arg Glu 340 345
350Val Arg Ala Val Thr Ser Ala Asn Ile Gln Glu Phe Ala Gly Cys Lys
355 360 365Lys Ile Phe Gly Ser Leu Ala
Phe Leu Pro Glu Ser Phe Asp Gly Asp 370 375
380Pro Ala Ser Asn Thr Ala Pro Leu Gln Pro Glu Gln Leu Gln Val
Phe385 390 395 400Glu Thr
Leu Glu Glu Ile Thr Gly Tyr Leu Tyr Ile Ser Ala Trp Pro
405 410 415Asp Ser Leu Pro Asp Leu Ser
Val Phe Gln Asn Leu Gln Val Ile Arg 420 425
430Gly Arg Ile Leu His Asn Gly Ala Tyr Ser Leu Thr Leu Gln
Gly Leu 435 440 445Gly Ile Ser Trp
Leu Gly Leu Arg Ser Leu Arg Glu Leu Gly Ser Gly 450
455 460Leu Ala Leu Ile His His Asn Thr His Leu Cys Phe
Val His Thr Val465 470 475
480Pro Trp Asp Gln Leu Phe Arg Asn Pro His Gln Ala Leu Leu His Thr
485 490 495Ala Asn Arg Pro Glu
Asp Glu Cys Val Gly Glu Gly Leu Ala Cys His 500
505 510Gln Leu Cys Ala Arg Gly His Cys Trp Gly Pro Gly
Pro Thr Gln Cys 515 520 525Val Asn
Cys Ser Gln Phe Leu Arg Gly Gln Glu Cys Val Glu Glu Cys 530
535 540Arg Val Leu Gln Gly Leu Pro Arg Glu Tyr Val
Asn Ala Arg His Cys545 550 555
560Leu Pro Cys His Pro Glu Cys Gln Pro Gln Asn Gly Ser Val Thr Cys
565 570 575Phe Gly Pro Glu
Ala Asp Gln Cys Val Ala Cys Ala His Tyr Lys Asp 580
585 590Pro Pro Phe Cys Val Ala Arg Cys Pro Ser Gly
Val Lys Pro Asp Leu 595 600 605Ser
Tyr Met Pro Ile Trp Lys Phe Pro Asp Glu Glu Gly Ala Cys Gln 610
615 620Pro Cys Pro Ile Asn Cys Thr His Ser Cys
Val Asp Leu Asp Asp Lys625 630 635
640Gly Cys Pro Ala Glu Gln Arg Ala Ser Pro Leu Thr Ser Ile Ile
Ser 645 650 655Ala Val Val
Gly Ile Leu Leu Val Val Val Leu Gly Val Val Phe Gly 660
665 670Ile Leu Ile Lys Arg Arg Gln Gln Lys Ile
Arg Lys Tyr Thr Met Arg 675 680
685Arg Leu Leu Gln Glu Thr Glu Leu Val Glu Pro Leu Thr Pro Ser Gly 690
695 700Ala Met Pro Asn Gln Ala Gln Met
Arg Ile Leu Lys Glu Thr Glu Leu705 710
715 720Arg Lys Val Lys Val Leu Gly Ser Gly Ala Phe Gly
Thr Val Tyr Lys 725 730
735Gly Ile Trp Ile Pro Asp Gly Glu Asn Val Lys Ile Pro Val Ala Ile
740 745 750Lys Val Leu Arg Glu Asn
Thr Ser Pro Lys Ala Asn Lys Glu Ile Leu 755 760
765Asp Glu Ala Tyr Val Met Ala Gly Val Gly Ser Pro Tyr Val
Ser Arg 770 775 780Leu Leu Gly Ile Cys
Leu Thr Ser Thr Val Gln Leu Val Thr Gln Leu785 790
795 800Met Pro Tyr Gly Cys Leu Leu Asp His Val
Arg Glu Asn Arg Gly Arg 805 810
815Leu Gly Ser Gln Asp Leu Leu Asn Trp Cys Met Gln Ile Ala Lys Gly
820 825 830Met Ser Tyr Leu Glu
Asp Val Arg Leu Val His Arg Asp Leu Ala Ala 835
840 845Arg Asn Val Leu Val Lys Ser Pro Asn His Val Lys
Ile Thr Asp Phe 850 855 860Gly Leu Ala
Arg Leu Leu Asp Ile Asp Glu Thr Glu Tyr His Ala Asp865
870 875 880Gly Gly Lys Val Pro Ile Lys
Trp Met Ala Leu Glu Ser Ile Leu Arg 885
890 895Arg Arg Phe Thr His Gln Ser Asp Val Trp Ser Tyr
Gly Val Thr Val 900 905 910Trp
Glu Leu Met Thr Phe Gly Ala Lys Pro Tyr Asp Gly Ile Pro Ala 915
920 925Arg Glu Ile Pro Asp Leu Leu Glu Lys
Gly Glu Arg Leu Pro Gln Pro 930 935
940Pro Ile Cys Thr Ile Asp Val Tyr Met Ile Met Val Lys Cys Trp Met945
950 955 960Ile Asp Ser Glu
Cys Arg Pro Arg Phe Arg Glu Leu Val Ser Glu Phe 965
970 975Ser Arg Met Ala Arg Asp Pro Gln Arg Phe
Val Val Ile Gln Asn Glu 980 985
990Asp Leu Gly Pro Ala Ser Pro Leu Asp Ser Thr Phe Tyr Arg Ser Leu
995 1000 1005Leu Glu Asp Asp Asp Met
Gly Asp Leu Val Asp Ala Glu Glu Tyr 1010 1015
1020Leu Val Pro Gln Gln Gly Phe Phe Cys Pro Asp Pro Ala Pro
Gly 1025 1030 1035Ala Gly Gly Met Val
His His Arg His Arg Ser Ser Ser Thr Arg 1040 1045
1050Ser Gly Gly Gly Asp Leu Thr Leu Gly Leu Glu Pro Ser
Glu Glu 1055 1060 1065Glu Ala Pro Arg
Ser Pro Leu Ala Pro Ser Glu Gly Ala Gly Ser 1070
1075 1080Asp Val Phe Asp Gly Asp Leu Gly Met Gly Ala
Ala Lys Gly Leu 1085 1090 1095Gln Ser
Leu Pro Thr His Asp Pro Ser Pro Leu Gln Arg Tyr Ser 1100
1105 1110Glu Asp Pro Thr Val Pro Leu Pro Ser Glu
Thr Asp Gly Tyr Val 1115 1120 1125Ala
Pro Leu Thr Cys Ser Pro Gln Pro Glu Tyr Val Asn Gln Pro 1130
1135 1140Asp Val Arg Pro Gln Pro Pro Ser Pro
Arg Glu Gly Pro Leu Pro 1145 1150
1155Ala Ala Arg Pro Ala Gly Ala Thr Leu Glu Arg Pro Lys Thr Leu
1160 1165 1170Ser Pro Gly Lys Asn Gly
Val Val Lys Asp Val Phe Ala Phe Gly 1175 1180
1185Gly Ala Val Glu Asn Pro Glu Tyr Leu Thr Pro Gln Gly Gly
Ala 1190 1195 1200Ala Pro Gln Pro His
Pro Pro Pro Ala Phe Ser Pro Ala Phe Asp 1205 1210
1215Asn Leu Tyr Tyr Trp Asp Gln Asp Pro Pro Glu Arg Gly
Ala Pro 1220 1225 1230Pro Ser Thr Phe
Lys Gly Thr Pro Thr Ala Glu Asn Pro Glu Tyr 1235
1240 1245Leu Gly Leu Asp Val Pro Val 1250
1255371392PRTHomo sapiens 371Met Glu Pro Pro Gly Arg Arg Glu Cys Pro
Phe Pro Ser Trp Arg Phe1 5 10
15Pro Gly Leu Leu Leu Ala Ala Met Val Leu Leu Leu Tyr Ser Phe Ser
20 25 30Asp Ala Cys Glu Glu Pro
Pro Thr Phe Glu Ala Met Glu Leu Ile Gly 35 40
45Lys Pro Lys Pro Tyr Tyr Glu Ile Gly Glu Arg Val Asp Tyr
Lys Cys 50 55 60Lys Lys Gly Tyr Phe
Tyr Ile Pro Pro Leu Ala Thr His Thr Ile Cys65 70
75 80Asp Arg Asn His Thr Trp Leu Pro Val Ser
Asp Asp Ala Cys Tyr Arg 85 90
95Glu Thr Cys Pro Tyr Ile Arg Asp Pro Leu Asn Gly Gln Ala Val Pro
100 105 110Ala Asn Gly Thr Tyr
Glu Phe Gly Tyr Gln Met His Phe Ile Cys Asn 115
120 125Glu Gly Tyr Tyr Leu Ile Gly Glu Glu Ile Leu Tyr
Cys Glu Leu Lys 130 135 140Gly Ser Val
Ala Ile Trp Ser Gly Lys Pro Pro Ile Cys Glu Lys Val145
150 155 160Leu Cys Thr Pro Pro Pro Lys
Ile Lys Asn Gly Lys His Thr Phe Ser 165
170 175Glu Val Glu Val Phe Glu Tyr Leu Asp Ala Val Thr
Tyr Ser Cys Asp 180 185 190Pro
Ala Pro Gly Pro Asp Pro Phe Ser Leu Ile Gly Glu Ser Thr Ile 195
200 205Tyr Cys Gly Asp Asn Ser Val Trp Ser
Arg Ala Ala Pro Glu Cys Lys 210 215
220Val Val Lys Cys Arg Phe Pro Val Val Glu Asn Gly Lys Gln Ile Ser225
230 235 240Gly Phe Gly Lys
Lys Phe Tyr Tyr Lys Ala Thr Val Met Phe Glu Cys 245
250 255Asp Lys Gly Phe Tyr Leu Asp Gly Ser Asp
Thr Ile Val Cys Asp Ser 260 265
270Asn Ser Thr Trp Asp Pro Pro Val Pro Lys Cys Leu Lys Val Leu Pro
275 280 285Pro Ser Ser Thr Lys Pro Pro
Ala Leu Ser His Ser Val Ser Thr Ser 290 295
300Ser Thr Thr Lys Ser Pro Ala Ser Ser Ala Ser Gly Pro Arg Pro
Thr305 310 315 320Tyr Lys
Pro Pro Val Ser Asn Tyr Pro Gly Tyr Pro Lys Pro Glu Glu
325 330 335Gly Ile Leu Asp Ser Leu Asp
Val Trp Val Ile Ala Val Ile Val Ile 340 345
350Ala Ile Val Val Gly Val Ala Val Ile Cys Val Val Pro Tyr
Arg Tyr 355 360 365Leu Gln Arg Arg
Lys Lys Lys Gly Thr Tyr Leu Thr Asp Glu Thr His 370
375 380Arg Glu Val Lys Phe Thr Ser Leu385
3903721066PRTHomo sapiens 372Met Gly Pro Gly Pro Ser Arg Ala Pro Arg Ala
Pro Arg Leu Met Leu1 5 10
15Cys Ala Leu Ala Leu Met Val Ala Ala Gly Gly Cys Val Val Ser Ala
20 25 30Phe Asn Leu Asp Thr Arg Phe
Leu Val Val Lys Glu Ala Gly Asn Pro 35 40
45Gly Ser Leu Phe Gly Tyr Ser Val Ala Leu His Arg Gln Thr Glu
Arg 50 55 60Gln Gln Arg Tyr Leu Leu
Leu Ala Gly Ala Pro Arg Glu Leu Ala Val65 70
75 80Pro Asp Gly Tyr Thr Asn Arg Thr Gly Ala Val
Tyr Leu Cys Pro Leu 85 90
95Thr Ala His Lys Asp Asp Cys Glu Arg Met Asn Ile Thr Val Lys Asn
100 105 110Asp Pro Gly His His Ile
Ile Glu Asp Met Trp Leu Gly Val Thr Val 115 120
125Ala Ser Gln Gly Pro Ala Gly Arg Val Leu Val Cys Ala His
Arg Tyr 130 135 140Thr Gln Val Leu Trp
Ser Gly Ser Glu Asp Gln Arg Arg Met Val Gly145 150
155 160Lys Cys Tyr Val Arg Gly Asn Asp Leu Glu
Leu Asp Ser Ser Asp Asp 165 170
175Trp Gln Thr Tyr His Asn Glu Met Cys Asn Ser Asn Thr Asp Tyr Leu
180 185 190Glu Thr Gly Met Cys
Gln Leu Gly Thr Ser Gly Gly Phe Thr Gln Asn 195
200 205Thr Val Tyr Phe Gly Ala Pro Gly Ala Tyr Asn Trp
Lys Gly Asn Ser 210 215 220Tyr Met Ile
Gln Arg Lys Glu Trp Asp Leu Ser Glu Tyr Ser Tyr Lys225
230 235 240Asp Pro Glu Asp Gln Gly Asn
Leu Tyr Ile Gly Tyr Thr Met Gln Val 245
250 255Gly Ser Phe Ile Leu His Pro Lys Asn Ile Thr Ile
Val Thr Gly Ala 260 265 270Pro
Arg His Arg His Met Gly Ala Val Phe Leu Leu Ser Gln Glu Ala 275
280 285Gly Gly Asp Leu Arg Arg Arg Gln Val
Leu Glu Gly Ser Gln Val Gly 290 295
300Ala Tyr Phe Gly Ser Ala Ile Ala Leu Ala Asp Leu Asn Asn Asp Gly305
310 315 320Trp Gln Asp Leu
Leu Val Gly Ala Pro Tyr Tyr Phe Glu Arg Lys Glu 325
330 335Glu Val Gly Gly Ala Ile Tyr Val Phe Met
Asn Gln Ala Gly Thr Ser 340 345
350Phe Pro Ala His Pro Ser Leu Leu Leu His Gly Pro Ser Gly Ser Ala
355 360 365Phe Gly Leu Ser Val Ala Ser
Ile Gly Asp Ile Asn Gln Asp Gly Phe 370 375
380Gln Asp Ile Ala Val Gly Ala Pro Phe Glu Gly Leu Gly Lys Val
Tyr385 390 395 400Ile Tyr
His Ser Ser Ser Lys Gly Leu Leu Arg Gln Pro Gln Gln Val
405 410 415Ile His Gly Glu Lys Leu Gly
Leu Pro Gly Leu Ala Thr Phe Gly Tyr 420 425
430Ser Leu Ser Gly Gln Met Asp Val Asp Glu Asn Phe Tyr Pro
Asp Leu 435 440 445Leu Val Gly Ser
Leu Ser Asp His Ile Val Leu Leu Arg Ala Arg Pro 450
455 460Val Ile Asn Ile Val His Lys Thr Leu Val Pro Arg
Pro Ala Val Leu465 470 475
480Asp Pro Ala Leu Cys Thr Ala Thr Ser Cys Val Gln Val Glu Leu Cys
485 490 495Phe Ala Tyr Asn Gln
Ser Ala Gly Asn Pro Asn Tyr Arg Arg Asn Ile 500
505 510Thr Leu Ala Tyr Thr Leu Glu Ala Asp Arg Asp Arg
Arg Pro Pro Arg 515 520 525Leu Arg
Phe Ala Gly Ser Glu Ser Ala Val Phe His Gly Phe Phe Ser 530
535 540Met Pro Glu Met Arg Cys Gln Lys Leu Glu Leu
Leu Leu Met Asp Asn545 550 555
560Leu Arg Asp Lys Leu Arg Pro Ile Ile Ile Ser Met Asn Tyr Ser Leu
565 570 575Pro Leu Arg Met
Pro Asp Arg Pro Arg Leu Gly Leu Arg Ser Leu Asp 580
585 590Ala Tyr Pro Ile Leu Asn Gln Ala Gln Ala Leu
Glu Asn His Thr Glu 595 600 605Val
Gln Phe Gln Lys Glu Cys Gly Pro Asp Asn Lys Cys Glu Ser Asn 610
615 620Leu Gln Met Arg Ala Ala Phe Val Ser Glu
Gln Gln Gln Lys Leu Ser625 630 635
640Arg Leu Gln Tyr Ser Arg Asp Val Arg Lys Leu Leu Leu Ser Ile
Asn 645 650 655Val Thr Asn
Thr Arg Thr Ser Glu Arg Ser Gly Glu Asp Ala His Glu 660
665 670Ala Leu Leu Thr Leu Val Val Pro Pro Ala
Leu Leu Leu Ser Ser Val 675 680
685Arg Pro Pro Gly Ala Cys Gln Ala Asn Glu Thr Ile Phe Cys Glu Leu 690
695 700Gly Asn Pro Phe Lys Arg Asn Gln
Arg Met Glu Leu Leu Ile Ala Phe705 710
715 720Glu Val Ile Gly Val Thr Leu His Thr Arg Asp Leu
Gln Val Gln Leu 725 730
735Gln Leu Ser Thr Ser Ser His Gln Asp Asn Leu Trp Pro Met Ile Leu
740 745 750Thr Leu Leu Val Asp Tyr
Thr Leu Gln Thr Ser Leu Ser Met Val Asn 755 760
765His Arg Leu Gln Ser Phe Phe Gly Gly Thr Val Met Gly Glu
Ser Gly 770 775 780Met Lys Thr Val Glu
Asp Val Gly Ser Pro Leu Lys Tyr Glu Phe Gln785 790
795 800Val Gly Pro Met Gly Glu Gly Leu Val Gly
Leu Gly Thr Leu Val Leu 805 810
815Gly Leu Glu Trp Pro Tyr Glu Val Ser Asn Gly Lys Trp Leu Leu Tyr
820 825 830Pro Thr Glu Ile Thr
Val His Gly Asn Gly Ser Trp Pro Cys Arg Pro 835
840 845Pro Gly Asp Leu Ile Asn Pro Leu Asn Leu Thr Leu
Ser Asp Pro Gly 850 855 860Asp Arg Pro
Ser Ser Pro Gln Arg Arg Arg Arg Gln Leu Asp Pro Gly865
870 875 880Gly Gly Gln Gly Pro Pro Pro
Val Thr Leu Ala Ala Ala Lys Lys Ala 885
890 895Lys Ser Glu Thr Val Leu Thr Cys Ala Thr Gly Arg
Ala His Cys Val 900 905 910Trp
Leu Glu Cys Pro Ile Pro Asp Ala Pro Val Val Thr Asn Val Thr 915
920 925Val Lys Ala Arg Val Trp Asn Ser Thr
Phe Ile Glu Asp Tyr Arg Asp 930 935
940Phe Asp Arg Val Arg Val Asn Gly Trp Ala Thr Leu Phe Leu Arg Thr945
950 955 960Ser Ile Pro Thr
Ile Asn Met Glu Asn Lys Thr Thr Trp Phe Ser Val 965
970 975Asp Ile Asp Ser Glu Leu Val Glu Glu Leu
Pro Ala Glu Ile Glu Leu 980 985
990Trp Leu Val Leu Val Ala Val Gly Ala Gly Leu Leu Leu Leu Gly Leu
995 1000 1005Ile Ile Leu Leu Leu Trp
Lys Cys Gly Phe Phe Lys Arg Thr Arg 1010 1015
1020Tyr Tyr Gln Ile Met Pro Lys Tyr His Ala Val Arg Ile Arg
Glu 1025 1030 1035Glu Glu Arg Tyr Pro
Pro Pro Gly Ser Thr Leu Pro Thr Lys Lys 1040 1045
1050His Trp Val Thr Ser Trp Gln Thr Arg Asp Gln Tyr Tyr
1055 1060 1065373532PRTHomo sapiens
373Met Ala Pro Ser Ser Pro Arg Pro Ala Leu Pro Ala Leu Leu Val Leu1
5 10 15Leu Gly Ala Leu Phe Pro
Gly Pro Gly Asn Ala Gln Thr Ser Val Ser 20 25
30Pro Ser Lys Val Ile Leu Pro Arg Gly Gly Ser Val Leu
Val Thr Cys 35 40 45Ser Thr Ser
Cys Asp Gln Pro Lys Leu Leu Gly Ile Glu Thr Pro Leu 50
55 60Pro Lys Lys Glu Leu Leu Leu Pro Gly Asn Asn Arg
Lys Val Tyr Glu65 70 75
80Leu Ser Asn Val Gln Glu Asp Ser Gln Pro Met Cys Tyr Ser Asn Cys
85 90 95Pro Asp Gly Gln Ser Thr
Ala Lys Thr Phe Leu Thr Val Tyr Trp Thr 100
105 110Pro Glu Arg Val Glu Leu Ala Pro Leu Pro Ser Trp
Gln Pro Val Gly 115 120 125Lys Asn
Leu Thr Leu Arg Cys Gln Val Glu Gly Gly Ala Pro Arg Ala 130
135 140Asn Leu Thr Val Val Leu Leu Arg Gly Glu Lys
Glu Leu Lys Arg Glu145 150 155
160Pro Ala Val Gly Glu Pro Ala Glu Val Thr Thr Thr Val Leu Val Arg
165 170 175Arg Asp His His
Gly Ala Asn Phe Ser Cys Arg Thr Glu Leu Asp Leu 180
185 190Arg Pro Gln Gly Leu Glu Leu Phe Glu Asn Thr
Ser Ala Pro Tyr Gln 195 200 205Leu
Gln Thr Phe Val Leu Pro Ala Thr Pro Pro Gln Leu Val Ser Pro 210
215 220Arg Val Leu Glu Val Asp Thr Gln Gly Thr
Val Val Cys Ser Leu Asp225 230 235
240Gly Leu Phe Pro Val Ser Glu Ala Gln Val His Leu Ala Leu Gly
Asp 245 250 255Gln Arg Leu
Asn Pro Thr Val Thr Tyr Gly Asn Asp Ser Phe Ser Ala 260
265 270Lys Ala Ser Val Ser Val Thr Ala Glu Asp
Glu Gly Thr Gln Arg Leu 275 280
285Thr Cys Ala Val Ile Leu Gly Asn Gln Ser Gln Glu Thr Leu Gln Thr 290
295 300Val Thr Ile Tyr Ser Phe Pro Ala
Pro Asn Val Ile Leu Thr Lys Pro305 310
315 320Glu Val Ser Glu Gly Thr Glu Val Thr Val Lys Cys
Glu Ala His Pro 325 330
335Arg Ala Lys Val Thr Leu Asn Gly Val Pro Ala Gln Pro Leu Gly Pro
340 345 350Arg Ala Gln Leu Leu Leu
Lys Ala Thr Pro Glu Asp Asn Gly Arg Ser 355 360
365Phe Ser Cys Ser Ala Thr Leu Glu Val Ala Gly Gln Leu Ile
His Lys 370 375 380Asn Gln Thr Arg Glu
Leu Arg Val Leu Tyr Gly Pro Arg Leu Asp Glu385 390
395 400Arg Asp Cys Pro Gly Asn Trp Thr Trp Pro
Glu Asn Ser Gln Gln Thr 405 410
415Pro Met Cys Gln Ala Trp Gly Asn Pro Leu Pro Glu Leu Lys Cys Leu
420 425 430Lys Asp Gly Thr Phe
Pro Leu Pro Ile Gly Glu Ser Val Thr Val Thr 435
440 445Arg Asp Leu Glu Gly Thr Tyr Leu Cys Arg Ala Arg
Ser Thr Gln Gly 450 455 460Glu Val Thr
Arg Lys Val Thr Val Asn Val Leu Ser Pro Arg Tyr Glu465
470 475 480Ile Val Ile Ile Thr Val Val
Ala Ala Ala Val Ile Met Gly Thr Ala 485
490 495Gly Leu Ser Thr Tyr Leu Tyr Asn Arg Gln Arg Lys
Ile Lys Lys Tyr 500 505 510Arg
Leu Gln Gln Ala Gln Lys Gly Thr Pro Met Lys Pro Asn Thr Gln 515
520 525Ala Thr Pro Pro 530374583PRTHomo
sapiens 374Met Glu Ser Lys Gly Ala Ser Ser Cys Arg Leu Leu Phe Cys Leu
Leu1 5 10 15Ile Ser Ala
Thr Val Phe Arg Pro Gly Leu Gly Trp Tyr Thr Val Asn 20
25 30Ser Ala Tyr Gly Asp Thr Ile Ile Ile Pro
Cys Arg Leu Asp Val Pro 35 40
45Gln Asn Leu Met Phe Gly Lys Trp Lys Tyr Glu Lys Pro Asp Gly Ser 50
55 60Pro Val Phe Ile Ala Phe Arg Ser Ser
Thr Lys Lys Ser Val Gln Tyr65 70 75
80Asp Asp Val Pro Glu Tyr Lys Asp Arg Leu Asn Leu Ser Glu
Asn Tyr 85 90 95Thr Leu
Ser Ile Ser Asn Ala Arg Ile Ser Asp Glu Lys Arg Phe Val 100
105 110Cys Met Leu Val Thr Glu Asp Asn Val
Phe Glu Ala Pro Thr Ile Val 115 120
125Lys Val Phe Lys Gln Pro Ser Lys Pro Glu Ile Val Ser Lys Ala Leu
130 135 140Phe Leu Glu Thr Glu Gln Leu
Lys Lys Leu Gly Asp Cys Ile Ser Glu145 150
155 160Asp Ser Tyr Pro Asp Gly Asn Ile Thr Trp Tyr Arg
Asn Gly Lys Val 165 170
175Leu His Pro Leu Glu Gly Ala Val Val Ile Ile Phe Lys Lys Glu Met
180 185 190Asp Pro Val Thr Gln Leu
Tyr Thr Met Thr Ser Thr Leu Glu Tyr Lys 195 200
205Thr Thr Lys Ala Asp Ile Gln Met Pro Phe Thr Cys Ser Val
Thr Tyr 210 215 220Tyr Gly Pro Ser Gly
Gln Lys Thr Ile His Ser Glu Gln Ala Val Phe225 230
235 240Asp Ile Tyr Tyr Pro Thr Glu Gln Val Thr
Ile Gln Val Leu Pro Pro 245 250
255Lys Asn Ala Ile Lys Glu Gly Asp Asn Ile Thr Leu Lys Cys Leu Gly
260 265 270Asn Gly Asn Pro Pro
Pro Glu Glu Phe Leu Phe Tyr Leu Pro Gly Gln 275
280 285Pro Glu Gly Ile Arg Ser Ser Asn Thr Tyr Thr Leu
Met Asp Val Arg 290 295 300Arg Asn Ala
Thr Gly Asp Tyr Lys Cys Ser Leu Ile Asp Lys Lys Ser305
310 315 320Met Ile Ala Ser Thr Ala Ile
Thr Val His Tyr Leu Asp Leu Ser Leu 325
330 335Asn Pro Ser Gly Glu Val Thr Arg Gln Ile Gly Asp
Ala Leu Pro Val 340 345 350Ser
Cys Thr Ile Ser Ala Ser Arg Asn Ala Thr Val Val Trp Met Lys 355
360 365Asp Asn Ile Arg Leu Arg Ser Ser Pro
Ser Phe Ser Ser Leu His Tyr 370 375
380Gln Asp Ala Gly Asn Tyr Val Cys Glu Thr Ala Leu Gln Glu Val Glu385
390 395 400Gly Leu Lys Lys
Arg Glu Ser Leu Thr Leu Ile Val Glu Gly Lys Pro 405
410 415Gln Ile Lys Met Thr Lys Lys Thr Asp Pro
Ser Gly Leu Ser Lys Thr 420 425
430Ile Ile Cys His Val Glu Gly Phe Pro Lys Pro Ala Ile Gln Trp Thr
435 440 445Ile Thr Gly Ser Gly Ser Val
Ile Asn Gln Thr Glu Glu Ser Pro Tyr 450 455
460Ile Asn Gly Arg Tyr Tyr Ser Lys Ile Ile Ile Ser Pro Glu Glu
Asn465 470 475 480Val Thr
Leu Thr Cys Thr Ala Glu Asn Gln Leu Glu Arg Thr Val Asn
485 490 495Ser Leu Asn Val Ser Ala Ile
Ser Ile Pro Glu His Asp Glu Ala Asp 500 505
510Glu Ile Ser Asp Glu Asn Arg Glu Lys Val Asn Asp Gln Ala
Lys Leu 515 520 525Ile Val Gly Ile
Val Val Gly Leu Leu Leu Ala Ala Leu Val Ala Gly 530
535 540Val Val Tyr Trp Leu Tyr Met Lys Lys Ser Lys Thr
Ala Ser Lys His545 550 555
560Val Asn Lys Asp Leu Gly Asn Met Glu Glu Asn Lys Lys Leu Glu Glu
565 570 575Asn Asn His Lys Thr
Glu Ala 580375385PRTHomo sapiens 375Met Ala Ala Ala Leu Phe
Val Leu Leu Gly Phe Ala Leu Leu Gly Thr1 5
10 15His Gly Ala Ser Gly Ala Ala Gly Phe Val Gln Ala
Pro Leu Ser Gln 20 25 30Gln
Arg Trp Val Gly Gly Ser Val Glu Leu His Cys Glu Ala Val Gly 35
40 45Ser Pro Val Pro Glu Ile Gln Trp Trp
Phe Glu Gly Gln Gly Pro Asn 50 55
60Asp Thr Cys Ser Gln Leu Trp Asp Gly Ala Arg Leu Asp Arg Val His65
70 75 80Ile His Ala Thr Tyr
His Gln His Ala Ala Ser Thr Ile Ser Ile Asp 85
90 95Thr Leu Val Glu Glu Asp Thr Gly Thr Tyr Glu
Cys Arg Ala Ser Asn 100 105
110Asp Pro Asp Arg Asn His Leu Thr Arg Ala Pro Arg Val Lys Trp Val
115 120 125Arg Ala Gln Ala Val Val Leu
Val Leu Glu Pro Gly Thr Val Phe Thr 130 135
140Thr Val Glu Asp Leu Gly Ser Lys Ile Leu Leu Thr Cys Ser Leu
Asn145 150 155 160Asp Ser
Ala Thr Glu Val Thr Gly His Arg Trp Leu Lys Gly Gly Val
165 170 175Val Leu Lys Glu Asp Ala Leu
Pro Gly Gln Lys Thr Glu Phe Lys Val 180 185
190Asp Ser Asp Asp Gln Trp Gly Glu Tyr Ser Cys Val Phe Leu
Pro Glu 195 200 205Pro Met Gly Thr
Ala Asn Ile Gln Leu His Gly Pro Pro Arg Val Lys 210
215 220Ala Val Lys Ser Ser Glu His Ile Asn Glu Gly Glu
Thr Ala Met Leu225 230 235
240Val Cys Lys Ser Glu Ser Val Pro Pro Val Thr Asp Trp Ala Trp Tyr
245 250 255Lys Ile Thr Asp Ser
Glu Asp Lys Ala Leu Met Asn Gly Ser Glu Ser 260
265 270Arg Phe Phe Val Ser Ser Ser Gln Gly Arg Ser Glu
Leu His Ile Glu 275 280 285Asn Leu
Asn Met Glu Ala Asp Pro Gly Gln Tyr Arg Cys Asn Gly Thr 290
295 300Ser Ser Lys Gly Ser Asp Gln Ala Ile Ile Thr
Leu Arg Val Arg Ser305 310 315
320His Leu Ala Ala Leu Trp Pro Phe Leu Gly Ile Val Ala Glu Val Leu
325 330 335Val Leu Val Thr
Ile Ile Phe Ile Tyr Glu Lys Arg Arg Lys Pro Glu 340
345 350Asp Val Leu Asp Asp Asp Asp Ala Gly Ser Ala
Pro Leu Lys Ser Ser 355 360 365Gly
Gln His Gln Asn Asp Lys Gly Lys Asn Val Arg Gln Arg Asn Ser 370
375 380Ser385376442PRTHomo sapiens 376Met Ala
Ser Val Val Leu Pro Ser Gly Ser Gln Cys Ala Ala Ala Ala1 5
10 15Ala Ala Ala Ala Pro Pro Gly Leu
Arg Leu Arg Leu Leu Leu Leu Leu 20 25
30Phe Ser Ala Ala Ala Leu Ile Pro Thr Gly Asp Gly Gln Asn Leu
Phe 35 40 45Thr Lys Asp Val Thr
Val Ile Glu Gly Glu Val Ala Thr Ile Ser Cys 50 55
60Gln Val Asn Lys Ser Asp Asp Ser Val Ile Gln Leu Leu Asn
Pro Asn65 70 75 80Arg
Gln Thr Ile Tyr Phe Arg Asp Phe Arg Pro Leu Lys Asp Ser Arg
85 90 95Phe Gln Leu Leu Asn Phe Ser
Ser Ser Glu Leu Lys Val Ser Leu Thr 100 105
110Asn Val Ser Ile Ser Asp Glu Gly Arg Tyr Phe Cys Gln Leu
Tyr Thr 115 120 125Asp Pro Pro Gln
Glu Ser Tyr Thr Thr Ile Thr Val Leu Val Pro Pro 130
135 140Arg Asn Leu Met Ile Asp Ile Gln Arg Asp Thr Ala
Val Glu Gly Glu145 150 155
160Glu Ile Glu Val Asn Cys Thr Ala Met Ala Ser Lys Pro Ala Thr Thr
165 170 175Ile Arg Trp Phe Lys
Gly Asn Thr Glu Leu Lys Gly Lys Ser Glu Val 180
185 190Glu Glu Trp Ser Asp Met Tyr Thr Val Thr Ser Gln
Leu Met Leu Lys 195 200 205Val His
Lys Glu Asp Asp Gly Val Pro Val Ile Cys Gln Val Glu His 210
215 220Pro Ala Val Thr Gly Asn Leu Gln Thr Gln Arg
Tyr Leu Glu Val Gln225 230 235
240Tyr Lys Pro Gln Val His Ile Gln Met Thr Tyr Pro Leu Gln Gly Leu
245 250 255Thr Arg Glu Gly
Asp Ala Leu Glu Leu Thr Cys Glu Ala Ile Gly Lys 260
265 270Pro Gln Pro Val Met Val Thr Trp Val Arg Val
Asp Asp Glu Met Pro 275 280 285Gln
His Ala Val Leu Ser Gly Pro Asn Leu Phe Ile Asn Asn Leu Asn 290
295 300Lys Thr Asp Asn Gly Thr Tyr Arg Cys Glu
Ala Ser Asn Ile Val Gly305 310 315
320Lys Ala His Ser Asp Tyr Met Leu Tyr Val Tyr Asp Pro Pro Thr
Thr 325 330 335Ile Pro Pro
Pro Thr Thr Thr Thr Thr Thr Thr Thr Thr Thr Thr Thr 340
345 350Thr Ile Leu Thr Ile Ile Thr Asp Ser Arg
Ala Gly Glu Glu Gly Ser 355 360
365Ile Arg Ala Val Asp His Ala Val Ile Gly Gly Val Val Ala Val Val 370
375 380Val Phe Ala Met Leu Cys Leu Leu
Ile Ile Leu Gly Arg Tyr Phe Ala385 390
395 400Arg His Lys Gly Thr Tyr Phe Thr His Glu Ala Lys
Gly Ala Asp Asp 405 410
415Ala Ala Asp Ala Asp Thr Ala Ile Ile Asn Ala Glu Gly Gly Gln Asn
420 425 430Asn Ser Glu Glu Lys Lys
Glu Tyr Phe Ile 435 44037717DNAArtificialAn
artificially synthesized primer sequence 377caggaaacag ctatgac
1737822DNAArtificialAn
artificially synthesized primer sequence 378cggctccaag tcgacgtcgt ca
2237983DNAArtificialAn
artificially synthesized primer sequence 379cagctgcagc agtctggggc
agagcttgtg aagccagggg cctcagtcaa gttgtcctgc 60acagcttctg gcttcaacat
taa 8338081DNAArtificialAn
artificially synthesized primer sequence 380agaccgaagt tgtaatttct
gtggatatac gtgacccact tcgtctccgg acttttccca 60gatctcacct aaccttccta a
8138184DNAArtificialAn
artificially synthesized primer sequence 381aagggtctag agtggattgg
aaggattgat cctgcgagtg gtaatactaa atatgacccg 60aaggacaagg ccactataac
agca 8438266DNAArtificialAn
artificially synthesized primer sequence 382ttcctgttcc ggtgatattg
tcgtctgtgt aggaggttgt gtcggatgga tgtcgactta 60agggac
6638351DNAArtificialAn
artificially synthesized primer sequence 383cagctgaatt ccctgacatc
tgaggacact gccgtctatt actgtgctgg t 5138473DNAArtificialAn
artificially synthesized primer sequence 384cagataatga cacgaccaat
actaatgccg ttgaaactga tgaccccggt tccgtggtgc 60cagtggcaca agg
7338554DNAArtificialAn
artificially synthesized primer sequence 385ggttctctaa cagtagtggt
agtagtggta attattctcg atagggccct cgaa 5438669DNAArtificialAn
artificially synthesized primer sequence 386gacatcgagc tcacccagtc
tccagcctcc ctttctgcgt ctgtgggaga aactgtcacc 60atcacatgt
6938763DNAArtificialAn
artificially synthesized primer sequence 387tgacagtggt agtgtacagc
tcgttcaccc ttataagtgt taataaatcg taccatggtc 60gtc
6338848DNAArtificialAn
artificially synthesized primer sequence 388gcatggtacc agcagaaacc
agggaaatct cctcagctcc tggtctat 4838981DNAArtificialAn
artificially synthesized primer sequence 389ggagtcgagg accagatatt
acgtttttgg aatcgtctac cacacggtag ttccaagtca 60ccgtcaccta ggccttgtgt t
8139045DNAArtificialAn
artificially synthesized primer sequence 390tcatgaggca cctgcaagcc
acctccgtgg ttcgagctct agttt 4539145DNAArtificialAn
artificially synthesized primer sequence 391agtactccgt ggacgttcgg
tggaggcacc aagctcgaga tcaaa 4539210DNAArtificialAn
artificially synthesized primer sequence 392atcgacagct
1039330DNAArtificialAn
artificially synthesized primer sequence 393aagccacctc catggttcga
gctctagttt 3039444DNAArtificialAn
artificially synthesized primer sequence 394tcgaagttgt ccttactcac
aagccgcgcg gtcagctgag gtaa 4439555DNAArtificialAn
artificially synthesized primer sequence 395accctggtca ccgtctcctc
agcctccacc aagggcccat cggtcttccc cctgg 5539637DNAArtificialAn
artificially synthesized primer sequence 396gggagtcgtc gcagcactgg
cacgggaggt cgtcgaa 3739736DNAArtificialAn
artificially synthesized primer sequence 397ggactctact ccctcagcag
cgtcgtgacc gtgccc 3639863DNAArtificialAn
artificially synthesized primer sequence 398gggtcgttgt ggttccacct
gttctttcaa ctcgggttta gaacagtagt ggtagtagtg 60gta
6339956DNAArtificialAn
artificially synthesized primer sequence 399gggtttagaa cagtagtggt
agtagtggta attattctcg atagggccct cgaacg 5640033DNAArtificialAn
artificially synthesized primer sequence 400ggcaccacgg tcaccgtctc
gagcgcctcc acc
334011916DNAArtificialInsert sequence of pscFvCA9-E8VHdVLd 401aagcttgcat
gcaaattcta tttcaaggag acagtcataa tgaaatacct attgcctacg 60gcagccgctg
gattgttatt actcgctgcc caaccagcga tggcccaggt gcagctgcag 120cagtctgggg
cagagcttgt gaagccaggg gcctcagtca agttgtcctg cacagcttct 180ggcttcaaca
ttaaagacac ctatatgcac tgggtgaagc agaggcctga aaagggtcta 240gaattccctg
acatctgagg acactgccgt ctattactgt gctggttatg attacggcaa 300ctttgactac
tggggccaag gcaccacggt caccgtctcg agaggcggtg gcggatcagg 360tggcggtgga
agtggcggtg gtgggtccat ggccgacatc gagctcaccc agtctccagc 420ctccctttct
gcgtctgtgg gagaaactgt caccatcaca tgtcgagcaa gtgggaatat 480tcacaattat
ttagcatggt accaagctcg agatcaaacg ggctgatgct gcaccaactg 540tatccatctt
cccaccatcc agtgagcagt taacatctgg aggtgcctca gtcgtgtgct 600tcttgaacag
cttctacccc aaagacatca atgtcaagtg gaagattgat ggcagtgaac 660gacaaaatgg
cgtcctgaac agttggactg atcaggacag caaagacagc acctacagca 720tgagcagcac
cctcacgttg accaaggacg agtatgaacg acataacagc tatacctgtg 780aggccactca
caagacatca acttcaccca ttgtcaagag cttcaacagg aatgagtgtt 840cggcgcgcca
gtcgactcca ttcgtttgtg aatatcaagg ccaatcgtct gacctgcctc 900aacctcctgt
caatgctggc ggcggctctg gtggtggttc tggtggcggc tctgagggtg 960gtggctctga
gggtggcggt tctgagggtg gcggctctga gggaggcggt tccggtggtg 1020gctctggttc
cggtgatttt gattatgaaa agatggcaaa cgctaataag ggggctatga 1080ccgaaaatgc
cgatgaaaac gcgctacagt cagacgctaa aggcaaactt gattctgtcg 1140ctactgatta
cggtgctgct atcgatggtt tcattggtga cgtttccggc cttgctaatg 1200gtaatggtgc
tactggtgat tttgctggct ctaattccca aatggctcaa gtcggtgacg 1260gtgataattc
acctttaatg aataatttcc gtcaatattt accttccctc cctcaatcgg 1320ttgaatgtcg
cccttttgtc tttggcgctg gtaaaccata tgaattttct attgattgtg 1380acaaaataaa
cttattccgt ggtgtctttg cgtttctttt atatgttgcc acctttatgt 1440atgtattttc
tacgtttgct aacatactgc gtaataagga gtcttaatca tgccagttct 1500tttgggtgct
agctgtcgac tgcgcaacac gatgaagccg tagacaacaa attcaacaaa 1560gaacaacaaa
acgcgttcta tgagatctta catttaccta acttaaacga agaacaacga 1620aacgccttca
tccaaagttt aaaagatgac ccaagccaaa gcgctaacct tttagcagaa 1680gctaaaaagc
taaatgatgc tcaggcgccg aaagtagaca acaaattcaa caaagaacaa 1740caaaacgcgt
tctatgagat cttacattta cctaacttaa acgaagaaca acgaaacgcc 1800ttcatccaaa
gtttaaaaga tgacccaagc caaagcgcta accttttagc agaagctaaa 1860aagctaaatg
atgctcaggc gccgaaagta gacgcgaatt agctgggaat taattc
191640266PRTArtificialAmino acid sequence encoded by insert sequence
of pscFvCA9-E8VHdVLd 402Met Lys Tyr Leu Leu Pro Thr Ala Ala Ala Gly Leu
Leu Leu Leu Ala1 5 10
15Ala Gln Pro Ala Met Ala Gln Val Gln Leu Gln Gln Ser Gly Ala Glu
20 25 30Leu Val Lys Pro Gly Ala Ser
Val Lys Leu Ser Cys Thr Ala Ser Gly 35 40
45Phe Asn Ile Lys Asp Thr Tyr Met His Trp Val Lys Gln Arg Pro
Glu 50 55 60Lys
Gly6540377DNAArtificialAn artificially synthesized primer sequence
403gtcaccgtct cgagaggcgg tggcggatca ggtggcggtg gaagtggcgg tggtgggtcc
60atggccgaca tcgagct
7740468DNAArtificialAn artificially synthesized primer sequence
404cgatgtcggc catggaccca ccaccgccac ttccaccgcc acctgatccg ccaccgcctc
60tcgagacg
684051774DNAArtificialInsert sequence of pscFvCA-E8VHd 405aagcttgcat
gcaaattcta tttcaaggag acagtcataa tgaaatacct attgcctacg 60gcagccgctg
gattgttatt actcgcggcc cagccggcca tggcccaggt gcagctgcag 120cagtctgggg
cagagcttgt gaagccaggg gcctcagtca agttgtcctg cacagcttct 180ggcttcaaca
ttaaagacac ctatatgcac tgggtgaagc agaggcctga aaagggtcta 240gaattccctg
acatctgagg acactgccgt ctattactgt gctggttatg attacggcaa 300ctttgactac
tggggccaag gcaccacggt caccgtctcc tcaggcggtg gcggatcagg 360tggcggtgga
agtggcggtg gtgggtctac tagtgacatc gagctcaccc agtctccagc 420ctccctttct
gcgtctgtgg gagaaactgt caccatcaca tgtcgagcaa gtgggaatat 480tcacaattat
ttagcatggt accagcagaa accagggaaa tctcctcagc tcctggtcta 540taatgcaaaa
accttagcag atggtgtgcc atcaaggttc agtggcagtg gatccggaac 600acaatattct
ctcaagatca acagcctgca gcctgaagat tttgggagtt attactgtca 660acatttttgg
agtactccgt ggacgttcgg tggaggtacc aagctcgagt cgactccatt 720cgtttgtgaa
tatcaaggcc aatcgtctga cctgcctcaa cctcctgtca atgctggcgg 780cggctctggt
ggtggttctg gtggcggctc tgagggtggt ggctctgagg gtggcggttc 840tgagggtggc
ggctctgagg gaggcggttc cggtggtggc tctggttccg gtgattttga 900ttatgaaaag
atggcaaacg ctaataaggg ggctatgacc gaaaatgccg atgaaaacgc 960gctacagtca
gacgctaaag gcaaacttga ttctgtcgct actgattacg gtgctgctat 1020cgatggtttc
attggtgacg tttccggcct tgctaatggt aatggtgcta ctggtgattt 1080tgctggctct
aattcccaaa tggctcaagt cggtgacggt gataattcac ctttaatgaa 1140taatttccgt
caatatttac cttccctccc tcaatcggtt gaatgtcgcc cttttgtctt 1200tggcgctggt
aaaccatatg aattttctat tgattgtgac aaaataaact tattccgtgg 1260tgtctttgcg
tttcttttat atgttgccac ctttatgtat gtattttcta cgtttgctaa 1320catactgcgt
aataaggagt cttaatcatg ccagttcttt tgggtgctag ctgtcgactg 1380cgcaacacga
tgaagccgta gacaacaaat tcaacaaaga acaacaaaac gcgttctatg 1440agatcttaca
tttacctaac ttaaacgaag aacaacgaaa cgccttcatc caaagtttaa 1500aagatgaccc
aagccaaagc gctaaccttt tagcagaagc taaaaagcta aatgatgctc 1560aggcgccgaa
agtagacaac aaattcaaca aagaacaaca aaacgcgttc tatgagatct 1620tacatttacc
taacttaaac gaagaacaac gaaacgcctt catccaaagt ttaaaagatg 1680acccaagcca
aagcgctaac cttttagcag aagctaaaaa gctaaatgat gctcaggcgc 1740cgaaagtaga
cgcgaattag ctgggaatta attc
177440666PRTArtificialAmino acid sequence encoded by insert sequence
of pscFvCA-E8VHd 406Met Lys Tyr Leu Leu Pro Thr Ala Ala Ala Gly Leu Leu
Leu Leu Ala1 5 10 15Ala
Gln Pro Ala Met Ala Gln Val Gln Leu Gln Gln Ser Gly Ala Glu 20
25 30Leu Val Lys Pro Gly Ala Ser Val
Lys Leu Ser Cys Thr Ala Ser Gly 35 40
45Phe Asn Ile Lys Asp Thr Tyr Met His Trp Val Lys Gln Arg Pro Glu
50 55 60Lys Gly6540791DNAArtificialAn
artificially synthesized primer sequence 407caccacggtc accgtctcct
caggcggtgg cggatcaggt ggcggtggaa gtggcggtgg 60tgggtctact agtgacatcg
agctcaccca g 9140891DNAArtificialAn
artificially synthesized primer sequence 408gtggtgccag tggcagagga
gtccgccacc gcctagtcca ccgccacctt caccgccacc 60acccagatga tcactgtagc
tcgagtgggt c 9140917DNAArtificialAn
artificially synthesized primer sequence 409caggaaacag ctatgac
1741042DNAArtificialAn
artificially synthesized primer sequence 410gacgccgggt cggccggtac
cggctccaag tcgacgtcgt ca 4241156DNAArtificialAn
artificially synthesized primer sequence 411gtcctcgcaa ctgcggccca
gccggccatg gccgacatcc agatgaccca gtctcc 5641256DNAArtificialAn
artificially synthesized primer sequence 412gtcctcgcaa ctgcggccca
gccggccatg gccgatgttg tgatgactca gtctcc 5641356DNAArtificialAn
artificially synthesized primer sequence 413gtcctcgcaa ctgcggccca
gccggccatg gccgaaattg tgttgacgca gtctcc 5641456DNAArtificialAn
artificially synthesized primer sequence 414gtcctcgcaa ctgcggccca
gccggccatg gccgacatcg tgatgaccca gtctcc 5641556DNAArtificialAn
artificially synthesized primer sequence 415gtcctcgcaa ctgcggccca
gccggccatg gccgaaacga cactcacgca gtctcc 5641656DNAArtificialAn
artificially synthesized primer sequence 416gtcctcgcaa ctgcggccca
gccggccatg gccgaaattg tgctgactca gtctcc 5641756DNAArtificialAn
artificially synthesized primer sequence 417gtcctcgcaa ctgcggccca
gccggccatg gcccagtctg tgttgacgca gccgcc 5641856DNAArtificialAn
artificially synthesized primer sequence 418gtcctcgcaa ctgcggccca
gccggccatg gcccagtctg ccctgactca gcctgc 5641956DNAArtificialAn
artificially synthesized primer sequence 419gtcctcgcaa ctgcggccca
gccggccatg gcctcctatg tgctgactca gccacc 5642056DNAArtificialAn
artificially synthesized primer sequence 420gtcctcgcaa ctgcggccca
gccggccatg gcctcttctg agctgactca ggaccc 5642156DNAArtificialAn
artificially synthesized primer sequence 421gtcctcgcaa ctgcggccca
gccggccatg gcccacgtta tactgactca accgcc 5642256DNAArtificialAn
artificially synthesized primer sequence 422gtcctcgcaa ctgcggccca
gccggccatg gcccaggctg tgctcactca gccgcc 5642356DNAArtificialAn
artificially synthesized primer sequence 423gtcctcgcaa ctgcggccca
gccggccatg gccaatttta tgctgactca gcccca 5642444DNAArtificialAn
artificially synthesized primer sequence 424tcgactggcg cgccgaacac
tctcccctgt tgaagctctt tgtg 4442543DNAArtificialAn
artificially synthesized primer sequence 425tcgactggcg cgccgaacat
tctgtagggg ccactgtctt ctc 4342620DNAArtificialAn
artificially synthesized primer sequence 426atggagtcgg gaaggaagtc
2042756DNAArtificialAn
artificially synthesized primer sequence 427gtcctcgcaa ctgcggccca
gccggccatg gcccaggtgc agctggtgca gtctgg 5642856DNAArtificialAn
artificially synthesized primer sequence 428gtcctcgcaa ctgcggccca
gccggccatg gcccaggtca acttaaggga gtctgg 5642956DNAArtificialAn
artificially synthesized primer sequence 429gtcctcgcaa ctgcggccca
gccggccatg gccgaggtgc agctggtgga gtctgg 5643056DNAArtificialAn
artificially synthesized primer sequence 430gtcctcgcaa ctgcggccca
gccggccatg gcccaggtgc agctgcagga gtcggg 5643156DNAArtificialAn
artificially synthesized primer sequence 431gtcctcgcaa ctgcggccca
gccggccatg gcccaggtgc agctgttgca gtctgc 5643256DNAArtificialAn
artificially synthesized primer sequence 432gtcctcgcaa ctgcggccca
gccggccatg gcccaggtac agctgcagca gtcagg 5643359DNAArtificialAn
artificially synthesized primer sequence 433gtcctcgcaa ctgcggccca
gccggccatg gcccagrtca ccttgaagga gtctggtcc 5943456DNAArtificialAn
artificially synthesized primer sequence 434gtcctcgcaa ctgcggccca
gccggccatg gcccaggtgc agctacagca gtgggg 5643556DNAArtificialAn
artificially synthesized primer sequence 435gtcctcgcaa ctgcggccca
gccggccatg gccgaggtgc agctggtgca gtctgg 5643664DNAArtificialAn
artificially synthesized primer sequence 436gtcctcgcaa ctgcggccca
gccggccatg gcccaggtgc agctggtgca atctgggtct 60gagt
6443732DNAArtificialAn
artificially synthesized primer sequence 437ggtggaggca ctcgagacgg
tgaccagggt gc 3243832DNAArtificialAn
artificially synthesized primer sequence 438ggtggaggca ctcgagacgg
tgaccattgt cc 3243932DNAArtificialAn
artificially synthesized primer sequence 439ggtggaggca ctcgagacgg
tgaccagggt tc 3244032DNAArtificialAn
artificially synthesized primer sequence 440ggtggaggca ctcgagacgg
tgaccgtggt cc 3244125RNAArtificialoligo
RNA 441cagagcuaca cauugagaac cugaa
2544225RNAArtificialoligo RNA 442uaccuaugug cagaggaauu augau
2544325RNAArtificialoligo RNA
443gcaaccaucu aaaccugaaa uugua
2544425RNAArtificialoligo RNA 444uaauagaggu ugucgaaggc ugggc
2544525RNAArtificialoligo RNA 445cccaacaggc
agaccauuua uuuca
25446652PRTHomo sapiens 446Met Ala Thr Ser Met Gly Leu Leu Leu Leu Leu
Leu Leu Leu Leu Thr1 5 10
15Gln Pro Gly Ala Gly Thr Gly Ala Asp Thr Glu Ala Val Val Cys Val
20 25 30Gly Thr Ala Cys Tyr Thr Ala
His Ser Gly Lys Leu Ser Ala Ala Glu 35 40
45Ala Gln Asn His Cys Asn Gln Asn Gly Gly Asn Leu Ala Thr Val
Lys 50 55 60Ser Lys Glu Glu Ala Gln
His Val Gln Arg Val Leu Ala Gln Leu Leu65 70
75 80Arg Arg Glu Ala Ala Leu Thr Ala Arg Met Ser
Lys Phe Trp Ile Gly 85 90
95Leu Gln Arg Glu Lys Gly Lys Cys Leu Asp Pro Ser Leu Pro Leu Lys
100 105 110Gly Phe Ser Trp Val Gly
Gly Gly Glu Asp Thr Pro Tyr Ser Asn Trp 115 120
125His Lys Glu Leu Arg Asn Ser Cys Ile Ser Lys Arg Cys Val
Ser Leu 130 135 140Leu Leu Asp Leu Ser
Gln Pro Leu Leu Pro Ser Arg Leu Pro Lys Trp145 150
155 160Ser Glu Gly Pro Cys Gly Ser Pro Gly Ser
Pro Gly Ser Asn Ile Glu 165 170
175Gly Phe Val Cys Lys Phe Ser Phe Lys Gly Met Cys Arg Pro Leu Ala
180 185 190Leu Gly Gly Pro Gly
Gln Val Thr Tyr Thr Thr Pro Phe Gln Thr Thr 195
200 205Ser Ser Ser Leu Glu Ala Val Pro Phe Ala Ser Ala
Ala Asn Val Ala 210 215 220Cys Gly Glu
Gly Asp Lys Asp Glu Thr Gln Ser His Tyr Phe Leu Cys225
230 235 240Lys Glu Lys Ala Pro Asp Val
Phe Asp Trp Gly Ser Ser Gly Pro Leu 245
250 255Cys Val Ser Pro Lys Tyr Gly Cys Asn Phe Asn Asn
Gly Gly Cys His 260 265 270Gln
Asp Cys Phe Glu Gly Gly Asp Gly Ser Phe Leu Cys Gly Cys Arg 275
280 285Pro Gly Phe Arg Leu Leu Asp Asp Leu
Val Thr Cys Ala Ser Arg Asn 290 295
300Pro Cys Ser Ser Ser Pro Cys Arg Gly Gly Ala Thr Cys Ala Leu Gly305
310 315 320Pro His Gly Lys
Asn Tyr Thr Cys Arg Cys Pro Gln Gly Tyr Gln Leu 325
330 335Asp Ser Ser Gln Leu Asp Cys Val Asp Val
Asp Glu Cys Gln Asp Ser 340 345
350Pro Cys Ala Gln Glu Cys Val Asn Thr Pro Gly Gly Phe Arg Cys Glu
355 360 365Cys Trp Val Gly Tyr Glu Pro
Gly Gly Pro Gly Glu Gly Ala Cys Gln 370 375
380Asp Val Asp Glu Cys Ala Leu Gly Arg Ser Pro Cys Ala Gln Gly
Cys385 390 395 400Thr Asn
Thr Asp Gly Ser Phe His Cys Ser Cys Glu Glu Gly Tyr Val
405 410 415Leu Ala Gly Glu Asp Gly Thr
Gln Cys Gln Asp Val Asp Glu Cys Val 420 425
430Gly Pro Gly Gly Pro Leu Cys Asp Ser Leu Cys Phe Asn Thr
Gln Gly 435 440 445Ser Phe His Cys
Gly Cys Leu Pro Gly Trp Val Leu Ala Pro Asn Gly 450
455 460Val Ser Cys Thr Met Gly Pro Val Ser Leu Gly Pro
Pro Ser Gly Pro465 470 475
480Pro Asp Glu Glu Asp Lys Gly Glu Lys Glu Gly Ser Thr Val Pro Arg
485 490 495Ala Ala Thr Ala Ser
Pro Thr Arg Gly Pro Glu Gly Thr Pro Lys Ala 500
505 510Thr Pro Thr Thr Ser Arg Pro Ser Leu Ser Ser Asp
Ala Pro Ile Thr 515 520 525Ser Ala
Pro Leu Lys Met Leu Ala Pro Ser Gly Ser Ser Gly Val Trp 530
535 540Arg Glu Pro Ser Ile His His Ala Thr Ala Ala
Ser Gly Pro Gln Glu545 550 555
560Pro Ala Gly Gly Asp Ser Ser Val Ala Thr Gln Asn Asn Asp Gly Thr
565 570 575Asp Gly Gln Lys
Leu Leu Leu Phe Tyr Ile Leu Gly Thr Val Val Ala 580
585 590Ile Leu Leu Leu Leu Ala Leu Ala Leu Gly Leu
Leu Val Tyr Arg Lys 595 600 605Arg
Arg Ala Lys Arg Glu Glu Lys Lys Glu Lys Lys Pro Gln Asn Ala 610
615 620Ala Asp Ser Tyr Ser Trp Val Pro Glu Arg
Ala Glu Ser Arg Ala Met625 630 635
640Glu Asn Gln Tyr Ser Pro Thr Pro Gly Thr Asp Cys
645 650447719PRTHomo sapiens 447Leu Asn Ile Thr Cys Arg
Phe Ala Gly Val Phe His Val Glu Lys Asn1 5
10 15Gly Arg Tyr Ser Ile Ser Arg Thr Glu Ala Ala Asp
Leu Cys Lys Ala 20 25 30Phe
Asn Ser Thr Leu Pro Thr Met Ala Gln Met Glu Lys Ala Leu Ser 35
40 45Ile Gly Phe Glu Thr Cys Arg Tyr Gly
Phe Ile Glu Gly His Val Val 50 55
60Ile Pro Arg Ile His Pro Asn Ser Ile Cys Ala Ala Asn Asn Thr Gly65
70 75 80Val Tyr Ile Leu Thr
Ser Asn Thr Ser Gln Tyr Asp Thr Tyr Cys Phe 85
90 95Asn Ala Ser Ala Pro Pro Glu Glu Asp Cys Thr
Ser Val Thr Asp Leu 100 105
110Pro Asn Ala Phe Asp Gly Pro Ile Thr Ile Thr Ile Val Asn Arg Asp
115 120 125Gly Thr Arg Tyr Val Gln Lys
Gly Glu Tyr Arg Thr Asn Pro Glu Asp 130 135
140Ile Tyr Pro Ser Asn Pro Thr Asp Asp Asp Val Ser Ser Gly Ser
Ser145 150 155 160Ser Glu
Arg Ser Ser Thr Ser Gly Gly Tyr Ile Phe Tyr Thr Phe Ser
165 170 175Thr Val His Pro Ile Pro Asp
Glu Asp Ser Pro Trp Ile Thr Asp Ser 180 185
190Thr Asp Arg Ile Pro Ala Thr Thr Leu Met Ser Thr Ser Ala
Thr Ala 195 200 205Thr Glu Thr Ala
Thr Lys Arg Gln Glu Thr Trp Asp Trp Phe Ser Trp 210
215 220Leu Phe Leu Pro Ser Glu Ser Lys Asn His Leu His
Thr Thr Thr Gln225 230 235
240Met Ala Gly Thr Ser Ser Asn Thr Ile Ser Ala Gly Trp Glu Pro Asn
245 250 255Glu Glu Asn Glu Asp
Glu Arg Asp Arg His Leu Ser Phe Ser Gly Ser 260
265 270Gly Ile Asp Asp Asp Glu Asp Phe Ile Ser Ser Thr
Ile Ser Thr Thr 275 280 285Pro Arg
Ala Phe Asp His Thr Lys Gln Asn Gln Asp Trp Thr Gln Trp 290
295 300Asn Pro Ser His Ser Asn Pro Glu Val Leu Leu
Gln Thr Thr Thr Arg305 310 315
320Met Thr Asp Val Asp Arg Asn Gly Thr Thr Ala Tyr Glu Gly Asn Trp
325 330 335Asn Pro Glu Ala
His Pro Pro Leu Ile His His Glu His His Glu Glu 340
345 350Glu Glu Thr Pro His Ser Thr Ser Thr Ile Gln
Ala Thr Pro Ser Ser 355 360 365Thr
Thr Glu Glu Thr Ala Thr Gln Lys Glu Gln Trp Phe Gly Asn Arg 370
375 380Trp His Glu Gly Tyr Arg Gln Thr Pro Lys
Glu Asp Ser His Ser Thr385 390 395
400Thr Gly Thr Ala Ala Ala Ser Ala His Thr Ser His Pro Met Gln
Gly 405 410 415Arg Thr Thr
Pro Ser Pro Glu Asp Ser Ser Trp Thr Asp Phe Phe Asn 420
425 430Pro Ile Ser His Pro Met Gly Arg Gly His
Gln Ala Gly Arg Arg Met 435 440
445Asp Met Asp Ser Ser His Ser Ile Thr Leu Gln Pro Thr Ala Asn Pro 450
455 460Asn Thr Gly Leu Val Glu Asp Leu
Asp Arg Thr Gly Pro Leu Ser Met465 470
475 480Thr Thr Gln Gln Ser Asn Ser Gln Ser Phe Ser Thr
Ser His Glu Gly 485 490
495Leu Glu Glu Asp Lys Asp His Pro Thr Thr Ser Thr Leu Thr Ser Ser
500 505 510Asn Arg Asn Asp Val Thr
Gly Gly Arg Arg Asp Pro Asn His Ser Glu 515 520
525Gly Ser Thr Thr Leu Leu Glu Gly Tyr Thr Ser His Tyr Pro
His Thr 530 535 540Lys Glu Ser Arg Thr
Phe Ile Pro Val Thr Ser Ala Lys Thr Gly Ser545 550
555 560Phe Gly Val Thr Ala Val Thr Val Gly Asp
Ser Asn Ser Asn Val Asn 565 570
575Arg Ser Leu Ser Gly Asp Gln Asp Thr Phe His Pro Ser Gly Gly Ser
580 585 590His Thr Thr His Gly
Ser Glu Ser Asp Gly His Ser His Gly Ser Gln 595
600 605Glu Gly Gly Ala Asn Thr Thr Ser Gly Pro Ile Arg
Thr Pro Gln Ile 610 615 620Pro Glu Trp
Leu Ile Ile Leu Ala Ser Leu Leu Ala Leu Ala Leu Ile625
630 635 640Leu Ala Val Cys Ile Ala Val
Asn Ser Arg Arg Arg Cys Gly Gln Lys 645
650 655Lys Lys Leu Val Ile Asn Ser Gly Asn Gly Ala Val
Glu Asp Arg Lys 660 665 670Pro
Ser Gly Leu Asn Gly Glu Ala Ser Lys Ser Gln Glu Met Val His 675
680 685Leu Val Asn Lys Glu Ser Ser Glu Thr
Pro Asp Gln Phe Met Thr Ala 690 695
700Asp Glu Thr Arg Asn Leu Gln Asn Val Asp Met Lys Ile Gly Val705
710 715448574PRTHomo sapiens 448Met Cys Pro Arg
Ala Ala Arg Ala Pro Ala Thr Leu Leu Leu Ala Leu1 5
10 15Gly Ala Val Leu Trp Pro Ala Ala Gly Ala
Trp Glu Leu Thr Ile Leu 20 25
30His Thr Asn Asp Val His Ser Arg Leu Glu Gln Thr Ser Glu Asp Ser
35 40 45Ser Lys Cys Val Asn Ala Ser Arg
Cys Met Gly Gly Val Ala Arg Leu 50 55
60Phe Thr Lys Val Gln Gln Ile Arg Arg Ala Glu Pro Asn Val Leu Leu65
70 75 80Leu Asp Ala Gly Asp
Gln Tyr Gln Gly Thr Ile Trp Phe Thr Val Tyr 85
90 95Lys Gly Ala Glu Val Ala His Phe Met Asn Ala
Leu Arg Tyr Asp Ala 100 105
110Met Ala Leu Gly Asn His Glu Phe Asp Asn Gly Val Glu Gly Leu Ile
115 120 125Glu Pro Leu Leu Lys Glu Ala
Lys Phe Pro Ile Leu Ser Ala Asn Ile 130 135
140Lys Ala Lys Gly Pro Leu Ala Ser Gln Ile Ser Gly Leu Tyr Leu
Pro145 150 155 160Tyr Lys
Val Leu Pro Val Gly Asp Glu Val Val Gly Ile Val Gly Tyr
165 170 175Thr Ser Lys Glu Thr Pro Phe
Leu Ser Asn Pro Gly Thr Asn Leu Val 180 185
190Phe Glu Asp Glu Ile Thr Ala Leu Gln Pro Glu Val Asp Lys
Leu Lys 195 200 205Thr Leu Asn Val
Asn Lys Ile Ile Ala Leu Gly His Ser Gly Phe Glu 210
215 220Met Asp Lys Leu Ile Ala Gln Lys Val Arg Gly Val
Asp Val Val Val225 230 235
240Gly Gly His Ser Asn Thr Phe Leu Tyr Thr Gly Asn Pro Pro Ser Lys
245 250 255Glu Val Pro Ala Gly
Lys Tyr Pro Phe Ile Val Thr Ser Asp Asp Gly 260
265 270Arg Lys Val Pro Val Val Gln Ala Tyr Ala Phe Gly
Lys Tyr Leu Gly 275 280 285Tyr Leu
Lys Ile Glu Phe Asp Glu Arg Gly Asn Val Ile Ser Ser His 290
295 300Gly Asn Pro Ile Leu Leu Asn Ser Ser Ile Pro
Glu Asp Pro Ser Ile305 310 315
320Lys Ala Asp Ile Asn Lys Trp Arg Ile Lys Leu Asp Asn Tyr Ser Thr
325 330 335Gln Glu Leu Gly
Lys Thr Ile Val Tyr Leu Asp Gly Ser Ser Gln Ser 340
345 350Cys Arg Phe Arg Glu Cys Asn Met Gly Asn Leu
Ile Cys Asp Ala Met 355 360 365Ile
Asn Asn Asn Leu Arg His Thr Asp Glu Met Phe Trp Asn His Val 370
375 380Ser Met Cys Ile Leu Asn Gly Gly Gly Ile
Arg Ser Pro Ile Asp Glu385 390 395
400Arg Asn Asn Gly Thr Ile Thr Trp Glu Asn Leu Ala Ala Val Leu
Pro 405 410 415Phe Gly Gly
Thr Phe Asp Leu Val Gln Leu Lys Gly Ser Thr Leu Lys 420
425 430Lys Ala Phe Glu His Ser Val His Arg Tyr
Gly Gln Ser Thr Gly Glu 435 440
445Phe Leu Gln Val Gly Gly Ile His Val Val Tyr Asp Leu Ser Arg Lys 450
455 460Pro Gly Asp Arg Val Val Lys Leu
Asp Val Leu Cys Thr Lys Cys Arg465 470
475 480Val Pro Ser Tyr Asp Pro Leu Lys Met Asp Glu Val
Tyr Lys Val Ile 485 490
495Leu Pro Asn Phe Leu Ala Asn Gly Gly Asp Gly Phe Gln Met Ile Lys
500 505 510Asp Glu Leu Leu Arg His
Asp Ser Gly Asp Gln Asp Ile Asn Val Val 515 520
525Ser Thr Tyr Ile Ser Lys Met Lys Val Ile Tyr Pro Ala Val
Glu Gly 530 535 540Arg Ile Lys Phe Ser
Thr Gly Ser His Cys His Gly Ser Phe Ser Leu545 550
555 560Ile Phe Leu Ser Leu Trp Ala Val Ile Phe
Val Leu Tyr Gln 565 570449314PRTHomo
sapiens 449Met Ala Pro Pro Gln Val Leu Ala Phe Gly Leu Leu Leu Ala Ala
Ala1 5 10 15Thr Ala Thr
Phe Ala Ala Ala Gln Glu Glu Cys Val Cys Glu Asn Tyr 20
25 30Lys Leu Ala Val Asn Cys Phe Val Asn Asn
Asn Arg Gln Cys Gln Cys 35 40
45Thr Ser Val Gly Ala Gln Asn Thr Val Ile Cys Ser Lys Leu Ala Ala 50
55 60Lys Cys Leu Val Met Lys Ala Glu Met
Asn Gly Ser Lys Leu Gly Arg65 70 75
80Arg Ala Lys Pro Glu Gly Ala Leu Gln Asn Asn Asp Gly Leu
Tyr Asp 85 90 95Pro Asp
Cys Asp Glu Ser Gly Leu Phe Lys Ala Lys Gln Cys Asn Gly 100
105 110Thr Ser Thr Cys Trp Cys Val Asn Thr
Ala Gly Val Arg Arg Thr Asp 115 120
125Lys Asp Thr Glu Ile Thr Cys Ser Glu Arg Val Arg Thr Tyr Trp Ile
130 135 140Ile Ile Glu Leu Lys His Lys
Ala Arg Glu Lys Pro Tyr Asp Ser Lys145 150
155 160Ser Leu Arg Thr Ala Leu Gln Lys Glu Ile Thr Thr
Arg Tyr Gln Leu 165 170
175Asp Pro Lys Phe Ile Thr Ser Ile Leu Tyr Glu Asn Asn Val Ile Thr
180 185 190Ile Asp Leu Val Gln Asn
Ser Ser Gln Lys Thr Gln Asn Asp Val Asp 195 200
205Ile Ala Asp Val Ala Tyr Tyr Phe Glu Lys Asp Val Lys Gly
Glu Ser 210 215 220Leu Phe His Ser Lys
Lys Met Asp Leu Thr Val Asn Gly Glu Gln Leu225 230
235 240Asp Leu Asp Pro Gly Gln Thr Leu Ile Tyr
Tyr Val Asp Glu Lys Ala 245 250
255Pro Glu Phe Ser Met Gln Gly Leu Lys Ala Gly Val Ile Ala Val Ile
260 265 270Val Val Val Val Met
Ala Val Val Ala Gly Ile Val Val Leu Val Ile 275
280 285Ser Arg Lys Lys Arg Met Ala Lys Tyr Glu Lys Ala
Glu Ile Lys Glu 290 295 300Met Gly Glu
Met His Arg Glu Leu Asn Ala305 3104501390PRTHomo sapiens
450Met Lys Ala Pro Ala Val Leu Ala Pro Gly Ile Leu Val Leu Leu Phe1
5 10 15Thr Leu Val Gln Arg Ser
Asn Gly Glu Cys Lys Glu Ala Leu Ala Lys 20 25
30Ser Glu Met Asn Val Asn Met Lys Tyr Gln Leu Pro Asn
Phe Thr Ala 35 40 45Glu Thr Pro
Ile Gln Asn Val Ile Leu His Glu His His Ile Phe Leu 50
55 60Gly Ala Thr Asn Tyr Ile Tyr Val Leu Asn Glu Glu
Asp Leu Gln Lys65 70 75
80Val Ala Glu Tyr Lys Thr Gly Pro Val Leu Glu His Pro Asp Cys Phe
85 90 95Pro Cys Gln Asp Cys Ser
Ser Lys Ala Asn Leu Ser Gly Gly Val Trp 100
105 110Lys Asp Asn Ile Asn Met Ala Leu Val Val Asp Thr
Tyr Tyr Asp Asp 115 120 125Gln Leu
Ile Ser Cys Gly Ser Val Asn Arg Gly Thr Cys Gln Arg His 130
135 140Val Phe Pro His Asn His Thr Ala Asp Ile Gln
Ser Glu Val His Cys145 150 155
160Ile Phe Ser Pro Gln Ile Glu Glu Pro Ser Gln Cys Pro Asp Cys Val
165 170 175Val Ser Ala Leu
Gly Ala Lys Val Leu Ser Ser Val Lys Asp Arg Phe 180
185 190Ile Asn Phe Phe Val Gly Asn Thr Ile Asn Ser
Ser Tyr Phe Pro Asp 195 200 205His
Pro Leu His Ser Ile Ser Val Arg Arg Leu Lys Glu Thr Lys Asp 210
215 220Gly Phe Met Phe Leu Thr Asp Gln Ser Tyr
Ile Asp Val Leu Pro Glu225 230 235
240Phe Arg Asp Ser Tyr Pro Ile Lys Tyr Val His Ala Phe Glu Ser
Asn 245 250 255Asn Phe Ile
Tyr Phe Leu Thr Val Gln Arg Glu Thr Leu Asp Ala Gln 260
265 270Thr Phe His Thr Arg Ile Ile Arg Phe Cys
Ser Ile Asn Ser Gly Leu 275 280
285His Ser Tyr Met Glu Met Pro Leu Glu Cys Ile Leu Thr Glu Lys Arg 290
295 300Lys Lys Arg Ser Thr Lys Lys Glu
Val Phe Asn Ile Leu Gln Ala Ala305 310
315 320Tyr Val Ser Lys Pro Gly Ala Gln Leu Ala Arg Gln
Ile Gly Ala Ser 325 330
335Leu Asn Asp Asp Ile Leu Phe Gly Val Phe Ala Gln Ser Lys Pro Asp
340 345 350Ser Ala Glu Pro Met Asp
Arg Ser Ala Met Cys Ala Phe Pro Ile Lys 355 360
365Tyr Val Asn Asp Phe Phe Asn Lys Ile Val Asn Lys Asn Asn
Val Arg 370 375 380Cys Leu Gln His Phe
Tyr Gly Pro Asn His Glu His Cys Phe Asn Arg385 390
395 400Thr Leu Leu Arg Asn Ser Ser Gly Cys Glu
Ala Arg Arg Asp Glu Tyr 405 410
415Arg Thr Glu Phe Thr Thr Ala Leu Gln Arg Val Asp Leu Phe Met Gly
420 425 430Gln Phe Ser Glu Val
Leu Leu Thr Ser Ile Ser Thr Phe Ile Lys Gly 435
440 445Asp Leu Thr Ile Ala Asn Leu Gly Thr Ser Glu Gly
Arg Phe Met Gln 450 455 460Val Val Val
Ser Arg Ser Gly Pro Ser Thr Pro His Val Asn Phe Leu465
470 475 480Leu Asp Ser His Pro Val Ser
Pro Glu Val Ile Val Glu His Thr Leu 485
490 495Asn Gln Asn Gly Tyr Thr Leu Val Ile Thr Gly Lys
Lys Ile Thr Lys 500 505 510Ile
Pro Leu Asn Gly Leu Gly Cys Arg His Phe Gln Ser Cys Ser Gln 515
520 525Cys Leu Ser Ala Pro Pro Phe Val Gln
Cys Gly Trp Cys His Asp Lys 530 535
540Cys Val Arg Ser Glu Glu Cys Leu Ser Gly Thr Trp Thr Gln Gln Ile545
550 555 560Cys Leu Pro Ala
Ile Tyr Lys Val Phe Pro Asn Ser Ala Pro Leu Glu 565
570 575Gly Gly Thr Arg Leu Thr Ile Cys Gly Trp
Asp Phe Gly Phe Arg Arg 580 585
590Asn Asn Lys Phe Asp Leu Lys Lys Thr Arg Val Leu Leu Gly Asn Glu
595 600 605Ser Cys Thr Leu Thr Leu Ser
Glu Ser Thr Met Asn Thr Leu Lys Cys 610 615
620Thr Val Gly Pro Ala Met Asn Lys His Phe Asn Met Ser Ile Ile
Ile625 630 635 640Ser Asn
Gly His Gly Thr Thr Gln Tyr Ser Thr Phe Ser Tyr Val Asp
645 650 655Pro Val Ile Thr Ser Ile Ser
Pro Lys Tyr Gly Pro Met Ala Gly Gly 660 665
670Thr Leu Leu Thr Leu Thr Gly Asn Tyr Leu Asn Ser Gly Asn
Ser Arg 675 680 685His Ile Ser Ile
Gly Gly Lys Thr Cys Thr Leu Lys Ser Val Ser Asn 690
695 700Ser Ile Leu Glu Cys Tyr Thr Pro Ala Gln Thr Ile
Ser Thr Glu Phe705 710 715
720Ala Val Lys Leu Lys Ile Asp Leu Ala Asn Arg Glu Thr Ser Ile Phe
725 730 735Ser Tyr Arg Glu Asp
Pro Ile Val Tyr Glu Ile His Pro Thr Lys Ser 740
745 750Phe Ile Ser Gly Gly Ser Thr Ile Thr Gly Val Gly
Lys Asn Leu Asn 755 760 765Ser Val
Ser Val Pro Arg Met Val Ile Asn Val His Glu Ala Gly Arg 770
775 780Asn Phe Thr Val Ala Cys Gln His Arg Ser Asn
Ser Glu Ile Ile Cys785 790 795
800Cys Thr Thr Pro Ser Leu Gln Gln Leu Asn Leu Gln Leu Pro Leu Lys
805 810 815Thr Lys Ala Phe
Phe Met Leu Asp Gly Ile Leu Ser Lys Tyr Phe Asp 820
825 830Leu Ile Tyr Val His Asn Pro Val Phe Lys Pro
Phe Glu Lys Pro Val 835 840 845Met
Ile Ser Met Gly Asn Glu Asn Val Leu Glu Ile Lys Gly Asn Asp 850
855 860Ile Asp Pro Glu Ala Val Lys Gly Glu Val
Leu Lys Val Gly Asn Lys865 870 875
880Ser Cys Glu Asn Ile His Leu His Ser Glu Ala Val Leu Cys Thr
Val 885 890 895Pro Asn Asp
Leu Leu Lys Leu Asn Ser Glu Leu Asn Ile Glu Trp Lys 900
905 910Gln Ala Ile Ser Ser Thr Val Leu Gly Lys
Val Ile Val Gln Pro Asp 915 920
925Gln Asn Phe Thr Gly Leu Ile Ala Gly Val Val Ser Ile Ser Thr Ala 930
935 940Leu Leu Leu Leu Leu Gly Phe Phe
Leu Trp Leu Lys Lys Arg Lys Gln945 950
955 960Ile Lys Asp Leu Gly Ser Glu Leu Val Arg Tyr Asp
Ala Arg Val His 965 970
975Thr Pro His Leu Asp Arg Leu Val Ser Ala Arg Ser Val Ser Pro Thr
980 985 990Thr Glu Met Val Ser Asn
Glu Ser Val Asp Tyr Arg Ala Thr Phe Pro 995 1000
1005Glu Asp Gln Phe Pro Asn Ser Ser Gln Asn Gly Ser
Cys Arg Gln 1010 1015 1020Val Gln Tyr
Pro Leu Thr Asp Met Ser Pro Ile Leu Thr Ser Gly 1025
1030 1035Asp Ser Asp Ile Ser Ser Pro Leu Leu Gln Asn
Thr Val His Ile 1040 1045 1050Asp Leu
Ser Ala Leu Asn Pro Glu Leu Val Gln Ala Val Gln His 1055
1060 1065Val Val Ile Gly Pro Ser Ser Leu Ile Val
His Phe Asn Glu Val 1070 1075 1080Ile
Gly Arg Gly His Phe Gly Cys Val Tyr His Gly Thr Leu Leu 1085
1090 1095Asp Asn Asp Gly Lys Lys Ile His Cys
Ala Val Lys Ser Leu Asn 1100 1105
1110Arg Ile Thr Asp Ile Gly Glu Val Ser Gln Phe Leu Thr Glu Gly
1115 1120 1125Ile Ile Met Lys Asp Phe
Ser His Pro Asn Val Leu Ser Leu Leu 1130 1135
1140Gly Ile Cys Leu Arg Ser Glu Gly Ser Pro Leu Val Val Leu
Pro 1145 1150 1155Tyr Met Lys His Gly
Asp Leu Arg Asn Phe Ile Arg Asn Glu Thr 1160 1165
1170His Asn Pro Thr Val Lys Asp Leu Ile Gly Phe Gly Leu
Gln Val 1175 1180 1185Ala Lys Gly Met
Lys Tyr Leu Ala Ser Lys Lys Phe Val His Arg 1190
1195 1200Asp Leu Ala Ala Arg Asn Cys Met Leu Asp Glu
Lys Phe Thr Val 1205 1210 1215Lys Val
Ala Asp Phe Gly Leu Ala Arg Asp Met Tyr Asp Lys Glu 1220
1225 1230Tyr Tyr Ser Val His Asn Lys Thr Gly Ala
Lys Leu Pro Val Lys 1235 1240 1245Trp
Met Ala Leu Glu Ser Leu Gln Thr Gln Lys Phe Thr Thr Lys 1250
1255 1260Ser Asp Val Trp Ser Phe Gly Val Val
Leu Trp Glu Leu Met Thr 1265 1270
1275Arg Gly Ala Pro Pro Tyr Pro Asp Val Asn Thr Phe Asp Ile Thr
1280 1285 1290Val Tyr Leu Leu Gln Gly
Arg Arg Leu Leu Gln Pro Glu Tyr Cys 1295 1300
1305Pro Asp Pro Leu Tyr Glu Val Met Leu Lys Cys Trp His Pro
Lys 1310 1315 1320Ala Glu Met Arg Pro
Ser Phe Ser Glu Leu Val Ser Arg Ile Ser 1325 1330
1335Ala Ile Phe Ser Thr Phe Ile Gly Glu His Tyr Val His
Val Asn 1340 1345 1350Ala Thr Tyr Val
Asn Val Lys Cys Val Ala Pro Tyr Pro Ser Leu 1355
1360 1365Leu Ser Ser Glu Asp Asn Ala Asp Asp Glu Val
Asp Thr Arg Pro 1370 1375 1380Ala Ser
Phe Trp Glu Thr Ser 1385 1390451121PRTHomo sapiens
451Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys
Lys Ala Ser Gly Tyr Thr Phe Thr Gly Tyr 20 25
30Tyr Met His Trp Val Arg Gln Ala Pro Gly Gln Gly Leu
Glu Trp Met 35 40 45Gly Trp Ile
Asn Pro Asn Ser Gly Gly Thr Asn Tyr Ala Gln Lys Phe 50
55 60Gln Gly Arg Val Thr Met Thr Arg Asp Thr Ser Ile
Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Arg Leu Arg Ser Asp Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Ser Asp Ile Leu
Thr Gly Tyr Tyr Ala Phe Asp Ile Trp Gly 100
105 110Gln Gly Thr Met Val Thr Val Ser Arg 115
1204525PRTHomo sapiens 452Gly Tyr Tyr Met His1
545317PRTHomo sapiens 453Trp Ile Asn Pro Asn Ser Gly Gly Thr Asn Tyr Ala
Gln Lys Phe Gln1 5 10
15Gly45412PRTHomo sapiens 454Ser Asp Ile Leu Thr Gly Tyr Tyr Ala Phe Asp
Ile1 5 10455107PRTHomo sapiens 455Asp Ile
Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1 5
10 15Asp Arg Val Thr Ile Thr Cys Arg
Ala Ser Gln Ser Ile Ser Ser Tyr 20 25
30Leu Asn Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu Leu
Ile 35 40 45Tyr Ala Ala Ser Ser
Leu Gln Ser Gly Val Pro Ser Arg Phe Ser Gly 50 55
60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu
Gln Pro65 70 75 80Glu
Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Ser Tyr Ser Thr Thr Trp
85 90 95Thr Phe Gly Gln Gly Thr Lys
Val Glu Ile Lys 100 10545611PRTHomo sapiens
456Arg Ala Ser Gln Ser Ile Ser Ser Tyr Leu Asn1 5
104577PRTHomo sapiens 457Ala Ala Ser Ser Leu Gln Ser1
54587PRTHomo sapiens 458Gln Gln Ser Tyr Ser Thr Thr1
5459129PRTHomo sapiens 459Glu Val Gln Leu Val Glu Ser Gly Ala Glu Val Lys
Lys Pro Gly Ala1 5 10
15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Gly Tyr
20 25 30Tyr Met His Trp Val Arg Gln
Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40
45Gly Trp Ile Asn Pro Asn Ser Gly Gly Thr Asn Tyr Ala Gln Lys
Phe 50 55 60Gln Gly Arg Val Thr Met
Thr Arg Asp Thr Ser Ile Ser Thr Ala Tyr65 70
75 80Met Glu Leu Ser Arg Leu Arg Ser Asp Asp Thr
Ala Val Tyr Tyr Cys 85 90
95Ala Arg Asp Leu Trp Tyr Tyr Tyr Gly Ser Gly Ser Ser Leu Tyr Tyr
100 105 110Tyr Tyr Gly Met Asp Val
Trp Gly Gln Gly Thr Thr Val Thr Val Ser 115 120
125Arg4605PRTHomo sapiens 460Gly Tyr Tyr Met His1
546117PRTHomo sapiens 461Trp Ile Asn Pro Asn Ser Gly Gly Thr Asn Tyr
Ala Gln Lys Phe Gln1 5 10
15Gly46220PRTHomo sapiens 462Asp Leu Trp Tyr Tyr Tyr Gly Ser Gly Ser Ser
Leu Tyr Tyr Tyr Tyr1 5 10
15Gly Met Asp Val 20463112PRTHomo sapiens 463Gln Ser Val Leu
Thr Gln Pro Pro Ser Val Ser Gly Ala Pro Gly Gln1 5
10 15Arg Val Thr Ile Ser Cys Thr Gly Ser Ser
Ser Asn Ile Gly Ala Gly 20 25
30Tyr Asp Val His Trp Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys Leu
35 40 45Leu Ile Tyr Gly Asn Ser Asn Arg
Pro Ser Gly Val Pro Asp Arg Phe 50 55
60Ser Gly Ser Lys Ser Gly Thr Ser Ala Ser Leu Ala Ile Thr Gly Leu65
70 75 80Gln Ala Glu Asp Glu
Ala Asp Tyr Tyr Cys Gln Ser Tyr Asp Ser Ser 85
90 95Leu Ser Gly Val Val Phe Gly Gly Gly Thr Lys
Leu Thr Val Leu Gly 100 105
11046414PRTHomo sapiens 464Thr Gly Ser Ser Ser Asn Ile Gly Ala Gly Tyr
Asp Val His1 5 104657PRTHomo sapiens
465Gly Asn Ser Asn Arg Pro Ser1 54669PRTHomo sapiens 466Gln
Ser Tyr Asp Ser Ser Leu Ser Gly1 5467122PRTHomo sapiens
467Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Arg1
5 10 15Ser Leu Arg Leu Ser Cys
Ala Ala Ser Gly Phe Thr Phe Asp Asp Tyr 20 25
30Ala Met His Trp Val Arg Gln Ala Pro Gly Lys Gly Leu
Glu Trp Val 35 40 45Ser Gly Ile
Ser Trp Asn Ser Gly Ser Ile Gly Tyr Ala Asp Ser Val 50
55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys
Asn Ser Leu Tyr65 70 75
80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Leu Tyr Tyr Cys
85 90 95Val Arg Ser Gly Ser Tyr
Asn Tyr Tyr Tyr Tyr Gly Met Asp Val Trp 100
105 110Gly Gln Gly Thr Thr Val Thr Val Ser Arg 115
1204685PRTHomo sapiens 468Asp Tyr Ala Met His1
546917PRTHomo sapiens 469Gly Ile Ser Trp Asn Ser Gly Ser Ile Gly Tyr
Ala Asp Ser Val Lys1 5 10
15Gly47013PRTHomo sapiens 470Ser Gly Ser Tyr Asn Tyr Tyr Tyr Tyr Gly Met
Asp Val1 5 10471111PRTHomo sapiens 471Gln
Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro Gly Gln1
5 10 15Arg Val Thr Ile Ser Cys Ser
Gly Ser Ser Ser Asn Ile Gly Ser Asn 20 25
30Thr Val Asn Trp Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys
Leu Leu 35 40 45Ile Tyr Ser Asn
Asn Gln Arg Pro Ser Gly Val Pro Asp Arg Phe Ser 50 55
60Gly Ser Lys Ser Gly Thr Ser Ala Ser Leu Ala Ile Ser
Gly Leu Gln65 70 75
80Ser Glu Asp Glu Ala Asp Tyr Tyr Cys Ala Ala Trp Asp Asp Ser Leu
85 90 95Asn Gly Trp Val Phe Gly
Gly Gly Thr Lys Leu Thr Val Leu Gly 100 105
11047213PRTHomo sapiens 472Ser Gly Ser Ser Ser Asn Ile Gly
Ser Asn Thr Val Asn1 5 104737PRTHomo
sapiens 473Ser Asn Asn Gln Arg Pro Ser1 54749PRTHomo
sapiens 474Ala Ala Trp Asp Asp Ser Leu Asn Gly1
5475123PRTHomo sapiens 475Gln Val Gln Leu Gln Glu Ser Gly Gly Gly Leu Val
Gln Pro Gly Gly1 5 10
15Ser Leu Arg Leu Ser Cys Thr Thr Ser Gly Phe Thr Phe Thr Ser Tyr
20 25 30Ala Met Ser Trp Val Arg Gln
Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40
45Ser Ser Ile Ser Gly Ser Gly Gly Ile Thr Tyr Tyr Ala Asp Ser
Val 50 55 60Lys Gly Arg Phe Thr Ile
Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr65 70
75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90
95Ala Lys Asp Arg Val Leu Val Pro Ala Ser Ser Ser Tyr Phe Asp Tyr
100 105 110Trp Gly Gln Gly Thr Leu
Val Thr Val Ser Arg 115 1204765PRTHomo sapiens
476Ser Tyr Ala Met Ser1 547717PRTHomo sapiens 477Ser Ile
Ser Gly Ser Gly Gly Ile Thr Tyr Tyr Ala Asp Ser Val Lys1 5
10 15Gly47814PRTHomo sapiens 478Asp Arg
Val Leu Val Pro Ala Ser Ser Ser Tyr Phe Asp Tyr1 5
10479109PRTHomo sapiens 479Ser Ser Glu Leu Thr Gln Asp Pro Ala
Val Ser Val Ala Leu Gly Gln1 5 10
15Thr Val Arg Ile Thr Cys Gln Gly Asp Ser Leu Arg Ser Tyr Tyr
Ala 20 25 30Ser Trp Tyr Gln
Gln Lys Pro Gly Gln Ala Pro Val Leu Val Ile Tyr 35
40 45Gly Lys Asn Asn Arg Pro Ser Gly Ile Pro Asp Arg
Phe Ser Gly Ser 50 55 60Ser Ser Gly
Asn Thr Ala Ser Leu Thr Ile Thr Gly Ala Gln Ala Glu65 70
75 80Asp Glu Ala Asp Tyr Tyr Cys Asn
Ser Arg Asp Ser Ser Gly Asn His 85 90
95Val Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly
100 10548011PRTHomo sapiens 480Gln Gly Asp Ser Leu
Arg Ser Tyr Tyr Ala Ser1 5 104817PRTHomo
sapiens 481Gly Lys Asn Asn Arg Pro Ser1 54829PRTHomo
sapiens 482Asn Ser Arg Asp Ser Ser Gly Asn His1
5483120PRTHomo sapiens 483Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val
Lys Pro Ser Glu1 5 10
15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Gly Ser Ile Ser Ser Ser
20 25 30Ser Tyr Tyr Trp Gly Trp Ile
Arg Gln Pro Pro Gly Lys Gly Leu Glu 35 40
45Trp Ile Gly Ser Ile Tyr Tyr Ser Gly Ser Thr Tyr Tyr Asn Pro
Ser 50 55 60Leu Lys Ser Arg Val Thr
Ile Ser Val Asp Thr Ser Lys Asn Gln Phe65 70
75 80Ser Leu Lys Leu Ser Ser Val Thr Ala Ala Asp
Thr Ala Val Tyr Tyr 85 90
95Cys Ala Arg Arg Pro Leu Thr Phe Asn Ala Phe Asp Ile Trp Gly Gln
100 105 110Gly Thr Met Val Thr Val
Ser Arg 115 1204847PRTHomo sapiens 484Ser Ser Ser
Tyr Tyr Trp Gly1 548516PRTHomo sapiens 485Ser Ile Tyr Tyr
Ser Gly Ser Thr Tyr Tyr Asn Pro Ser Leu Lys Ser1 5
10 1548610PRTHomo sapiens 486Arg Pro Leu Thr
Phe Asn Ala Phe Asp Ile1 5
10487109PRTHomo sapiens 487Ser Tyr Val Leu Thr Gln Pro Pro Ser Val Ser
Val Ala Pro Gly Lys1 5 10
15Thr Ala Arg Ile Thr Cys Gly Gly Asn Asn Ile Gly Ser Lys Ser Val
20 25 30His Trp Tyr Gln Gln Lys Pro
Gly Gln Ala Pro Val Leu Val Ile Tyr 35 40
45Tyr Asp Ser Asp Arg Pro Ser Gly Ile Pro Glu Arg Phe Ser Gly
Ser 50 55 60Asn Ser Gly Asn Thr Ala
Thr Leu Thr Ile Ser Arg Val Glu Ala Gly65 70
75 80Asp Glu Ala Asp Tyr Tyr Cys Gln Val Trp Asp
Ser Ser Ser Asp His 85 90
95Val Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly 100
10548811PRTHomo sapiens 488Gly Gly Asn Asn Ile Gly Ser Lys
Ser Val His1 5 104897PRTHomo sapiens
489Tyr Asp Ser Asp Arg Pro Ser1 54909PRTHomo sapiens 490Gln
Val Trp Asp Ser Ser Ser Asp His1 5491120PRTHomo sapiens
491Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Lys Pro Ser Glu1
5 10 15Thr Leu Ser Leu Thr Cys
Thr Val Ser Gly Gly Ser Ile Ser Ser Ser 20 25
30Ser Tyr Tyr Trp Gly Trp Ile Arg Gln Pro Pro Gly Lys
Gly Leu Glu 35 40 45Trp Ile Gly
Ser Ile Tyr Tyr Ser Gly Ser Thr Tyr Tyr Asn Pro Ser 50
55 60Leu Lys Ser Arg Val Thr Ile Ser Val Asp Thr Ser
Lys Asn Gln Phe65 70 75
80Ser Leu Lys Leu Ser Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr
85 90 95Cys Ala Arg Ile Pro Met
Tyr Ser Ser Ser Val Asp Tyr Trp Gly Gln 100
105 110Gly Thr Leu Val Thr Val Ser Arg 115
1204927PRTHomo sapiens 492Ser Ser Ser Tyr Tyr Trp Gly1
549316PRTHomo sapiens 493Ser Ile Tyr Tyr Ser Gly Ser Thr Tyr Tyr Asn
Pro Ser Leu Lys Ser1 5 10
1549410PRTHomo sapiens 494Ile Pro Met Tyr Ser Ser Ser Val Asp Tyr1
5 10495109PRTHomo sapiens 495Ser Tyr Val Leu
Thr Gln Pro Pro Ser Val Ser Val Ala Pro Gly Lys1 5
10 15Thr Ala Arg Ile Thr Cys Gly Gly Asn Asn
Ile Gly Ser Lys Ser Val 20 25
30His Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Val Leu Val Ile Tyr
35 40 45Tyr Asp Ser Asp Arg Pro Ser Gly
Ile Pro Glu Arg Phe Ser Gly Ser 50 55
60Asn Ser Gly Asn Thr Ala Thr Leu Thr Ile Ser Arg Val Glu Ala Gly65
70 75 80Asp Glu Ala Asp Tyr
Tyr Cys Gln Val Trp Asp Ser Ser Ser Asp His 85
90 95Val Val Phe Gly Gly Gly Thr Lys Leu Thr Val
Leu Gly 100 10549611PRTHomo sapiens 496Gly Gly
Asn Asn Ile Gly Ser Lys Ser Val His1 5
104977PRTHomo sapiens 497Tyr Asp Ser Asp Arg Pro Ser1
54989PRTHomo sapiens 498Gln Val Trp Asp Ser Ser Ser Asp His1
5499120PRTHomo sapiens 499Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu
Val Lys Pro Ser Glu1 5 10
15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Gly Ser Ile Ser Ser Ser
20 25 30Ser Tyr Tyr Trp Gly Trp Ile
Arg Gln Pro Pro Gly Lys Gly Leu Glu 35 40
45Trp Ile Gly Ser Ile Tyr Tyr Ser Gly Ser Thr Tyr Tyr Asn Pro
Ser 50 55 60Leu Lys Ser Arg Val Thr
Ile Ser Val Asp Thr Ser Lys Asn Gln Phe65 70
75 80Ser Leu Lys Leu Ser Ser Val Thr Ala Ala Asp
Thr Ala Val Tyr Tyr 85 90
95Cys Ala Arg Arg Pro Leu Thr Phe Asn Ala Phe Asp Ile Trp Gly Gln
100 105 110Gly Thr Thr Val Thr Val
Ser Arg 115 1205007PRTHomo sapiens 500Ser Ser Ser
Tyr Tyr Trp Gly1 550116PRTHomo sapiens 501Ser Ile Tyr Tyr
Ser Gly Ser Thr Tyr Tyr Asn Pro Ser Leu Lys Ser1 5
10 1550210PRTHomo sapiens 502Arg Pro Leu Thr
Phe Asn Ala Phe Asp Ile1 5
10503109PRTHomo sapiens 503Ser Tyr Val Leu Thr Gln Pro Pro Ser Val Ser
Val Ala Pro Gly Lys1 5 10
15Thr Ala Arg Ile Thr Cys Gly Gly Asn Asn Ile Gly Ser Lys Ser Val
20 25 30His Trp Tyr Gln Gln Lys Pro
Gly Gln Ala Pro Val Leu Val Ile Tyr 35 40
45Tyr Asp Ser Asp Arg Pro Ser Gly Ile Pro Glu Arg Phe Ser Gly
Ser 50 55 60Asn Ser Gly Asn Thr Ala
Thr Leu Thr Ile Ser Arg Val Glu Ala Gly65 70
75 80Asp Glu Ala Asp Tyr Tyr Cys Gln Val Trp Asp
Ser Ser Ser Asp His 85 90
95Val Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly 100
10550411PRTHomo sapiens 504Gly Gly Asn Asn Ile Gly Ser Lys
Ser Val His1 5 105057PRTHomo sapiens
505Tyr Asp Ser Asp Arg Pro Ser1 55069PRTHomo sapiens 506Gln
Val Trp Asp Ser Ser Ser Asp His1 5507122PRTHomo sapiens
507Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1
5 10 15Ser Leu Arg Leu Ser Cys
Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20 25
30Ala Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu
Glu Trp Val 35 40 45Ser Ala Ile
Ser Gly Ser Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val 50
55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys
Asn Thr Leu Tyr65 70 75
80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Lys Asp Gly Gly Trp
Phe Gly Glu Leu Asp Tyr Phe Gln His Trp 100
105 110Gly Gln Gly Thr Leu Val Thr Val Ser Arg 115
1205085PRTHomo sapiens 508Ser Tyr Ala Met Ser1
550917PRTHomo sapiens 509Ala Ile Ser Gly Ser Gly Gly Ser Thr Tyr Tyr
Ala Asp Ser Val Lys1 5 10
15Gly51013PRTHomo sapiens 510Asp Gly Gly Trp Phe Gly Glu Leu Asp Tyr Phe
Gln His1 5 10511111PRTHomo sapiens 511Gln
Ser Val Leu Thr Gln Pro Pro Ser Val Ser Ala Ala Pro Gly Gln1
5 10 15Lys Val Thr Val Ser Cys Thr
Gly Ser Asn Ser Asn Ile Glu Lys Asn 20 25
30Asp Val Ser Trp Tyr Gln Gln Gly Pro Gly Ala Ala Pro Lys
Leu Leu 35 40 45Ile Ser Asp Thr
Asp Arg Arg Pro Ser Gly Ile Pro Asp Arg Phe Ser 50 55
60Gly Ser Lys Ser Gly Thr Ser Ala Thr Leu Ala Ile Ala
Gly Leu Gln65 70 75
80Ala Glu Asp Glu Ala Asp Tyr Tyr Cys Gln Ser His Asp Thr Thr Leu
85 90 95Ser Gly Pro Ile Phe Gly
Gly Gly Thr Gln Leu Thr Val Leu Gly 100 105
11051213PRTHomo sapiens 512Thr Gly Ser Asn Ser Asn Ile Glu
Lys Asn Asp Val Ser1 5 105137PRTHomo
sapiens 513Asp Thr Asp Arg Arg Pro Ser1 55149PRTHomo
sapiens 514Gln Ser His Asp Thr Thr Leu Ser Gly1
5515124PRTHomo sapiens 515Gln Val Gln Leu Gln Glu Ser Gly Gly Gly Leu Val
Lys Pro Gly Gly1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Ser Phe Ser Asp Tyr
20 25 30Tyr Met Ser Trp Val Arg Gln
Ala Pro Gly Lys Gly Leu Glu Trp Ile 35 40
45Ser Tyr Ile Thr Ser Ser Ser Ser Asp Thr Asp Tyr Ala Asp Ser
Val 50 55 60Lys Gly Arg Phe Thr Ile
Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70
75 80Leu Gln Met Asn Ser Leu Arg Ala Asp Asp Thr
Ala Val Tyr Tyr Cys 85 90
95Ala Arg Val Gly Tyr Tyr Tyr Asp Tyr Tyr Tyr Tyr Tyr Tyr Met Asp
100 105 110Val Trp Gly Lys Gly Thr
Thr Val Thr Val Ser Arg 115 1205165PRTHomo sapiens
516Asp Tyr Tyr Met Ser1 551717PRTHomo sapiens 517Tyr Ile
Thr Ser Ser Ser Ser Asp Thr Asp Tyr Ala Asp Ser Val Lys1 5
10 15Gly51815PRTHomo sapiens 518Val Gly
Tyr Tyr Tyr Asp Tyr Tyr Tyr Tyr Tyr Tyr Met Asp Val1 5
10 15519108PRTHomo sapiens 519Ser Ser Glu
Leu Thr Gln Asp Pro Ala Val Ser Val Ala Leu Gly Gln1 5
10 15Thr Val Arg Ile Thr Cys Gln Gly Asp
Ser Leu Arg Gly Tyr Tyr Ala 20 25
30Ser Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Val Leu Val Val Tyr
35 40 45Asp Glu Asn Asn Arg Pro Ser
Gly Ile Pro Asp Arg Phe Ser Gly Ser 50 55
60Ser Ser Gly Asn Thr Ala Ser Leu Thr Ile Thr Gly Ala Gln Ala Glu65
70 75 80Asp Glu Ala Asp
Tyr Tyr Cys Asn Ser Arg Asp Ile Asn Leu Asp Trp 85
90 95Val Phe Gly Gly Gly Thr Lys Leu Thr Val
Leu Gly 100 10552011PRTHomo sapiens 520Gln Gly
Asp Ser Leu Arg Gly Tyr Tyr Ala Ser1 5
105217PRTHomo sapiens 521Asp Glu Asn Asn Arg Pro Ser1
55228PRTHomo sapiens 522Asn Ser Arg Asp Ile Asn Leu Asp1
5523120PRTHomo sapiens 523Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val
Lys Pro Ser Glu1 5 10
15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Gly Ser Ile Ser Ser Ser
20 25 30Ser Tyr Tyr Trp Gly Trp Ile
Arg Gln Pro Pro Gly Lys Gly Leu Glu 35 40
45Trp Ile Gly Ser Ile Tyr Tyr Ser Gly Ser Thr Tyr Tyr Asn Pro
Ser 50 55 60Leu Lys Ser Arg Val Thr
Ile Ser Val Asp Thr Ser Lys Asn Gln Phe65 70
75 80Ser Leu Lys Leu Ser Ser Val Thr Ala Ala Asp
Thr Ala Val Tyr Tyr 85 90
95Cys Ala Arg Arg Val Ile Val Trp Gly Ser Tyr Asp Tyr Trp Gly Gln
100 105 110Gly Thr Leu Val Thr Val
Ser Arg 115 1205247PRTHomo sapiens 524Ser Ser Ser
Tyr Tyr Trp Gly1 552516PRTHomo sapiens 525Ser Ile Tyr Tyr
Ser Gly Ser Thr Tyr Tyr Asn Pro Ser Leu Lys Ser1 5
10 1552610PRTHomo sapiens 526Arg Val Ile Val
Trp Gly Ser Tyr Asp Tyr1 5
10527109PRTHomo sapiens 527Ser Tyr Val Leu Thr Gln Pro Pro Ser Val Ser
Val Ala Pro Gly Lys1 5 10
15Thr Ala Arg Ile Thr Cys Gly Gly Asn Asn Ile Gly Ser Lys Ser Val
20 25 30His Trp Tyr Gln Gln Lys Pro
Gly Gln Ala Pro Val Leu Val Ile Tyr 35 40
45Tyr Asp Ser Asp Arg Pro Ser Gly Ile Pro Glu Arg Phe Ser Gly
Ser 50 55 60Asn Ser Gly Asn Thr Ala
Thr Leu Thr Ile Ser Arg Val Glu Ala Gly65 70
75 80Asp Glu Ala Asp Tyr Tyr Cys Gln Val Trp Asp
Ser Ser Ser Asp His 85 90
95Val Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly 100
10552811PRTHomo sapiens 528Gly Gly Asn Asn Ile Gly Ser Lys
Ser Val His1 5 105297PRTHomo sapiens
529Tyr Asp Ser Asp Arg Pro Ser1 55309PRTHomo sapiens 530Gln
Val Trp Asp Ser Ser Ser Asp His1 5531120PRTHomo sapiens
531Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Lys Pro Ser Glu1
5 10 15Thr Leu Ser Leu Thr Cys
Thr Val Ser Gly Gly Ser Ile Ser Ser Ser 20 25
30Ser Tyr Tyr Trp Gly Trp Ile Arg Gln Pro Pro Gly Lys
Gly Leu Glu 35 40 45Trp Ile Gly
Ser Ile Tyr Tyr Ser Gly Ser Thr Tyr Tyr Asn Pro Ser 50
55 60Leu Lys Ser Arg Val Thr Ile Ser Val Asp Thr Ser
Lys Asn Gln Phe65 70 75
80Ser Leu Lys Leu Ser Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr
85 90 95Cys Ala Met Arg Ala Tyr
Gly Ser Gly Ser Tyr Asp Tyr Trp Gly Gln 100
105 110Gly Thr Leu Val Thr Val Ser Arg 115
1205327PRTHomo sapiens 532Ser Ser Ser Tyr Tyr Trp Gly1
553316PRTHomo sapiens 533Ser Ile Tyr Tyr Ser Gly Ser Thr Tyr Tyr Asn
Pro Ser Leu Lys Ser1 5 10
1553410PRTHomo sapiens 534Arg Ala Tyr Gly Ser Gly Ser Tyr Asp Tyr1
5 10535112PRTHomo sapiens 535Asp Val Val Met
Thr Gln Ser Pro Leu Ser Leu Pro Val Thr Pro Gly1 5
10 15Glu Pro Ala Ser Ile Ser Cys Arg Ser Ser
Gln Ser Leu Leu His Ser 20 25
30Asn Gly Tyr Asn Tyr Leu Asp Trp Tyr Leu Gln Lys Pro Gly Gln Ser
35 40 45Pro Gln Leu Leu Ile Tyr Leu Gly
Ser Asn Arg Ala Ser Gly Val Pro 50 55
60Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Lys Ile65
70 75 80Ser Arg Val Glu Ala
Glu Asp Val Gly Val Tyr Tyr Cys Met Gln Ala 85
90 95Leu Gln Thr Pro Phe Thr Phe Gly Pro Gly Thr
Lys Val Asp Ile Lys 100 105
11053616PRTHomo sapiens 536Arg Ser Ser Gln Ser Leu Leu His Ser Asn Gly
Tyr Asn Tyr Leu Asp1 5 10
155377PRTHomo sapiens 537Leu Gly Ser Asn Arg Ala Ser1
55387PRTHomo sapiens 538Met Gln Ala Leu Gln Thr Pro1
5539121PRTHomo sapiens 539Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val
Gln Pro Gly Gly1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr
20 25 30Ala Met Ser Trp Val Arg Gln
Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40
45Ser Ala Ile Ser Gly Ser Gly Gly Ser Thr Tyr Tyr Ala Asp Ser
Val 50 55 60Lys Gly Arg Phe Thr Ile
Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr65 70
75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp Thr
Ala Ala Tyr Tyr Cys 85 90
95Ala Lys Gly Leu Arg Tyr Ser Ser Ala Trp Thr Phe Asp Tyr Trp Gly
100 105 110Gln Gly Thr Leu Val Thr
Val Ser Arg 115 1205405PRTHomo sapiens 540Ser Tyr
Ala Met Ser1 554117PRTHomo sapiens 541Ala Ile Ser Gly Ser
Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val Lys1 5
10 15Gly54212PRTHomo sapiens 542Gly Leu Arg Tyr Ser
Ser Ala Trp Thr Phe Asp Tyr1 5
10543112PRTHomo sapiens 543Gln Ser Ala Leu Thr Gln Pro Ala Ser Val Ser
Gly Ser Pro Gly Gln1 5 10
15Ser Ile Thr Ile Ser Cys Thr Gly Thr Ser Ser Asp Val Gly Gly Tyr
20 25 30Asn Tyr Val Ser Trp Tyr Gln
Gln His Pro Gly Lys Ala Pro Lys Leu 35 40
45Met Ile Tyr Asp Val Ser Asn Arg Pro Ser Gly Val Ser Asn Arg
Phe 50 55 60Ser Gly Ser Lys Ser Gly
Asn Thr Ala Ser Leu Thr Ile Ser Gly Leu65 70
75 80Gln Ala Glu Asp Glu Ala Asp Tyr Tyr Cys Ser
Ser Tyr Thr Ser Ser 85 90
95Ser Thr Pro Val Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly
100 105 11054414PRTHomo sapiens
544Thr Gly Thr Ser Ser Asp Val Gly Gly Tyr Asn Tyr Val Ser1
5 105457PRTHomo sapiens 545Asp Val Ser Asn Arg Pro Ser1
55469PRTHomo sapiens 546Ser Ser Tyr Thr Ser Ser Ser Thr
Pro1 5547119PRTHomo sapiens 547Gln Val Gln Leu Val Gln Ser
Gly Ser Glu Leu Lys Lys Pro Gly Ala1 5 10
15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Ile Phe
Thr Arg Tyr 20 25 30Gly Ile
Asn Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45Gly Trp Ile Asn Thr Asn Thr Gly Asn Pro
Thr Tyr Ala Gln Gly Phe 50 55 60Thr
Gly Arg Val Val Phe Ser Leu Asp Thr Ser Val Ser Thr Ala Tyr65
70 75 80Leu Gln Ile Ser Ser Leu
Lys Ala Glu Asp Thr Ala Met Tyr Tyr Cys 85
90 95Ala Ile Ser Ser Gly Phe Gly Tyr Tyr Phe Asp Tyr
Trp Gly Gln Gly 100 105 110Thr
Leu Val Thr Val Ser Arg 1155485PRTHomo sapiens 548Arg Tyr Gly Ile
Asn1 554917PRTHomo sapiens 549Trp Ile Asn Thr Asn Thr Gly
Asn Pro Thr Tyr Ala Gln Gly Phe Thr1 5 10
15Gly55010PRTHomo sapiens 550Ser Ser Gly Phe Gly Tyr Tyr
Phe Asp Tyr1 5 10551107PRTHomo sapiens
551Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Val Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr
Cys Arg Ala Ser Gln Ser Ile Ser Arg Trp 20 25
30Leu Ala Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys
Phe Leu Ile 35 40 45Tyr Ala Gly
Ser Ser Leu Gln Ser Gly Val Pro Ser Arg Phe Ser Gly 50
55 60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser
Ser Leu Gln Pro65 70 75
80Glu Asp Phe Ala Thr Tyr Tyr Cys Gln Gln Thr Asn Ser Phe Pro Leu
85 90 95Thr Phe Gly Gly Gly Thr
Lys Val Glu Ile Lys 100 10555211PRTHomo
sapiens 552Arg Ala Ser Gln Ser Ile Ser Arg Trp Leu Ala1 5
105537PRTHomo sapiens 553Ala Gly Ser Ser Leu Gln Ser1
55547PRTHomo sapiens 554Gln Gln Thr Asn Ser Phe Pro1
5555120PRTHomo sapiens 555Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val
Lys Lys Pro Gly Ala1 5 10
15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser Tyr
20 25 30Gly Ile Ser Trp Val Arg Gln
Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40
45Gly Trp Ile Ser Ala Tyr Asn Gly Asn Thr Asn Tyr Ala Gln Lys
Leu 50 55 60Gln Gly Arg Val Thr Met
Thr Thr Asp Thr Ser Thr Ser Thr Ala Tyr65 70
75 80Met Glu Leu Arg Ser Leu Arg Ser Asp Asp Thr
Ala Val Tyr Tyr Cys 85 90
95Ala Arg Ala Phe Arg Asp Trp Gly Ser Leu Arg Asp Tyr Trp Gly Gln
100 105 110Gly Thr Leu Val Thr Val
Ser Arg 115 1205565PRTHomo sapiens 556Ser Tyr Gly
Ile Ser1 555717PRTHomo sapiens 557Trp Ile Ser Ala Tyr Asn
Gly Asn Thr Asn Tyr Ala Gln Lys Leu Gln1 5
10 15Gly55811PRTHomo sapiens 558Ala Phe Arg Asp Trp Gly
Ser Leu Arg Asp Tyr1 5 10559111PRTHomo
sapiens 559Gln Ser Val Leu Thr Gln Pro Pro Ser Val Ser Ala Ala Pro Gly
Gln1 5 10 15Lys Val Thr
Ile Ser Cys Ser Gly Ser Ser Ser Asn Ile Gly Asn Asn 20
25 30Tyr Val Ser Trp Tyr Gln Gln Leu Pro Gly
Thr Ala Pro Lys Leu Leu 35 40
45Ile Tyr Asp Asn Asn Lys Arg Pro Ser Gly Ile Pro Asp Arg Phe Ser 50
55 60Gly Ser Lys Ser Gly Thr Ser Ala Thr
Leu Gly Ile Thr Gly Leu Gln65 70 75
80Thr Gly Asp Glu Ala Asp Tyr Tyr Cys Gly Thr Trp Asp Ser
Ser Leu 85 90 95Ser Ala
Val Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly 100
105 11056013PRTHomo sapiens 560Ser Gly Ser Ser
Ser Asn Ile Gly Asn Asn Tyr Val Ser1 5
105617PRTHomo sapiens 561Asp Asn Asn Lys Arg Pro Ser1
55629PRTHomo sapiens 562Gly Thr Trp Asp Ser Ser Leu Ser Ala1
5563121PRTHomo sapiens 563Glu Val Gln Leu Val Glu Ser Gly Ala Glu Val
Lys Lys Pro Gly Glu1 5 10
15Ser Leu Lys Ile Ser Cys Lys Gly Ser Gly Tyr Ser Phe Thr Ser Tyr
20 25 30Trp Ile Gly Trp Val Arg Gln
Met Pro Gly Lys Gly Leu Glu Trp Met 35 40
45Gly Ile Ile Tyr Pro Gly Asp Ser Asp Thr Arg Tyr Ser Pro Ser
Phe 50 55 60Gln Gly Gln Val Thr Ile
Ser Ala Asp Lys Ser Ile Ser Thr Ala Tyr65 70
75 80Leu Gln Trp Ser Ser Leu Lys Ala Ser Asp Thr
Ala Met Tyr Tyr Cys 85 90
95Ala Arg Leu Ala Val Gly Ala Tyr Gln Tyr Tyr Phe Asp Tyr Trp Gly
100 105 110Gln Gly Thr Leu Val Thr
Val Ser Arg 115 1205645PRTHomo sapiens 564Ser Tyr
Trp Ile Gly1 556517PRTHomo sapiens 565Ile Ile Tyr Pro Gly
Asp Ser Asp Thr Arg Tyr Ser Pro Ser Phe Gln1 5
10 15Gly56612PRTHomo sapiens 566Leu Ala Val Gly Ala
Tyr Gln Tyr Tyr Phe Asp Tyr1 5
10567111PRTHomo sapiens 567Gln Thr Val Val Thr Gln Glu Pro Ser Phe Ser
Val Ser Pro Gly Gly1 5 10
15Thr Val Thr Leu Thr Cys Gly Leu Ser Ser Gly Ser Val Ser Thr Ser
20 25 30Tyr Tyr Pro Ser Trp Tyr Gln
Gln Thr Pro Gly Gln Ala Pro Arg Thr 35 40
45Leu Ile Tyr Ser Thr Asn Thr Arg Ser Ser Gly Val Pro Asp Arg
Phe 50 55 60Ser Gly Ser Ile Leu Gly
Asn Lys Ala Ala Leu Thr Ile Thr Gly Ala65 70
75 80Gln Ala Asp Asp Glu Ser Asp Tyr Tyr Cys Val
Leu Tyr Met Gly Ser 85 90
95Gly Ile Ser Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly
100 105 11056814PRTHomo sapiens 568Gly
Leu Ser Ser Gly Ser Val Ser Thr Ser Tyr Tyr Pro Ser1 5
105697PRTHomo sapiens 569Ser Thr Asn Thr Arg Ser Ser1
55708PRTHomo sapiens 570Val Leu Tyr Met Gly Ser Gly Ile1
5571119PRTHomo sapiens 571Glu Val Gln Leu Val Glu Ser Gly Gly Asn Leu
Val Gln Pro Gly Gly1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Arg Tyr
20 25 30Trp Met Thr Trp Val Arg Gln
Ala Pro Gly Lys Gly Leu Glu Trp Ile 35 40
45Ser Ser Val Ser Ser Ser Gly Ser Thr Ile Tyr Tyr Ala Asp Ser
Val 50 55 60Lys Gly Arg Phe Thr Ile
Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70
75 80Leu Gln Met Asn Ser Leu Arg Val Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90
95Ala Arg Asp Tyr Trp Pro Gly Trp Tyr Phe Asp Leu Trp Gly Arg Gly
100 105 110Thr Leu Val Thr Val Ser
Arg 1155725PRTHomo sapiens 572Arg Tyr Trp Met Thr1
557317PRTHomo sapiens 573Ser Val Ser Ser Ser Gly Ser Thr Ile Tyr Tyr Ala
Asp Ser Val Lys1 5 10
15Gly57410PRTHomo sapiens 574Asp Tyr Trp Pro Gly Trp Tyr Phe Asp Leu1
5 10575116PRTHomo sapiens 575Gln Ala Val Leu
Thr Gln Pro Ser Ser Leu Ser Ala Ser Pro Gly Ala1 5
10 15Ser Ala Ser Leu Thr Cys Thr Phe Arg Ser
Asp Ile Ser Val Gly Ser 20 25
30Tyr Arg Ile Tyr Trp Tyr Gln Gln Lys Pro Gly Ser Pro Pro Gln Phe
35 40 45Leu Leu Lys Tyr Thr Ser Asp Ser
Asp Lys Gln Gln Gly Ser Gly Val 50 55
60Pro Ser Arg Phe Ser Gly Ser Lys Asp Val Ser Ala Asn Ala Gly Ile65
70 75 80Leu Leu Ile Ser Gly
Leu Gln Ser Glu Asp Glu Ala Asp Tyr Tyr Cys 85
90 95Met Thr Trp His Asn Thr Ala Ser Val Phe Gly
Gly Gly Thr Lys Leu 100 105
110Ala Val Leu Gly 11557614PRTHomo sapiens 576Thr Phe Arg Ser Asp
Ile Ser Val Gly Ser Tyr Arg Ile Tyr1 5
1057711PRTHomo sapiens 577Tyr Thr Ser Asp Ser Asp Lys Gln Gln Gly Ser1
5 105787PRTHomo sapiens 578Met Thr Trp His
Asn Thr Ala1 5579119PRTHomo sapiens 579Glu Val Gln Leu Val
Glu Ser Gly Gly Gly Leu Ile Arg Pro Gly Gly1 5
10 15Ser Leu Arg Leu Ser Cys Thr Ala Ser Gly Phe
Thr Phe Ser Ser Tyr 20 25
30Thr Met Asn Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Leu
35 40 45Ser Tyr Ile Ser Gly Gly Ser Gly
Thr Lys Phe Tyr Ala Asp Ser Val 50 55
60Lys Gly Arg Phe Thr Val Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65
70 75 80Leu Glu Met Asn Ser
Leu Arg Pro Glu Asp Thr Ala Val Tyr Tyr Cys 85
90 95Ala Leu Val Ser Tyr Ser Ser Pro Gly Phe Asp
Tyr Trp Gly Gln Gly 100 105
110Thr Leu Val Thr Val Ser Ser 1155805PRTHomo sapiens 580Ser Tyr
Thr Met Asn1 558117PRTHomo sapiens 581Tyr Ile Ser Gly Gly
Ser Gly Thr Lys Phe Tyr Ala Asp Ser Val Lys1 5
10 15Gly58210PRTHomo sapiens 582Val Ser Tyr Ser Ser
Pro Gly Phe Asp Tyr1 5 10583107PRTHomo
sapiens 583Ser Tyr Glu Leu Thr Gln Pro Pro Ser Val Ser Val Ser Pro Gly
Gln1 5 10 15Thr Ala Ser
Ile Thr Cys Ser Gly Asp Lys Leu Gly Asp Lys Tyr Val 20
25 30Tyr Trp Tyr Gln Gln Lys Pro Gly Gln Ser
Pro Ile Leu Leu Ile Phe 35 40
45Gln Asp Ser Glu Arg Pro Ser Gly Ile Pro Glu Arg Phe Ser Gly Ser 50
55 60Asn Ser Gly Asn Thr Ala Thr Leu Thr
Ile Ser Gly Thr Gln Ala Met65 70 75
80Asp Glu Ala Asp Tyr Tyr Cys Gln Ala Trp Asp Ser Ser Ala
Ala Val 85 90 95Phe Gly
Gly Gly Thr Lys Leu Thr Val Leu Gly 100
10558411PRTHomo sapiens 584Ser Gly Asp Lys Leu Gly Asp Lys Tyr Val Tyr1
5 105857PRTHomo sapiens 585Gln Asp Ser Glu
Arg Pro Ser1 55867PRTHomo sapiens 586Gln Ala Trp Asp Ser
Ser Ala1 5587115PRTHomo sapiens 587Gln Val Gln Leu Val Gln
Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5
10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr
Phe Thr Ser Tyr 20 25 30Gly
Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45Gly Trp Ile Ser Ala Tyr Asn Gly Asn
Thr Asn Tyr Ala Gln Lys Leu 50 55
60Gln Gly Arg Val Thr Met Thr Thr Asp Thr Ser Thr Ser Thr Ala Tyr65
70 75 80Met Glu Leu Arg Ser
Leu Arg Ser Asp Asp Thr Ala Val Tyr Tyr Cys 85
90 95Ala Arg Gly Gly Ala Phe Asp Ile Trp Gly Gln
Gly Thr Thr Val Thr 100 105
110Val Ser Arg 1155885PRTHomo sapiens 588Ser Tyr Gly Ile Ser1
558917PRTHomo sapiens 589Trp Ile Ser Ala Tyr Asn Gly Asn Thr Asn
Tyr Ala Gln Lys Leu Gln1 5 10
15Gly5906PRTHomo sapiens 590Gly Gly Ala Phe Asp Ile1
5591112PRTHomo sapiens 591Gln Ser Val Leu Thr Gln Pro Pro Ser Val Ser Gly
Ala Pro Gly Gln1 5 10
15Arg Val Thr Ile Ser Cys Thr Gly Ser Ser Ser Asn Ile Gly Ala Gly
20 25 30Tyr His Val Tyr Trp Tyr Gln
Gln Leu Pro Gly Lys Ala Pro Lys Leu 35 40
45Leu Ile Tyr Val Asn Ser Asn Arg Pro Ser Gly Val Pro Asp Arg
Phe 50 55 60Ser Gly Ser Lys Ser Gly
Thr Ser Ala Ser Leu Ala Ile Thr Gly Leu65 70
75 80Gln Ala Asp Asp Glu Ala Asp Tyr Tyr Cys Gln
Ser Tyr Asp Ser Ser 85 90
95Leu Ser Gly Arg Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly
100 105 11059214PRTHomo sapiens
592Thr Gly Ser Ser Ser Asn Ile Gly Ala Gly Tyr His Val Tyr1
5 105937PRTHomo sapiens 593Val Asn Ser Asn Arg Pro Ser1
55949PRTHomo sapiens 594Gln Ser Tyr Asp Ser Ser Leu Ser
Gly1 5595119PRTHomo sapiens 595Glu Val Gln Leu Val Glu Ser
Gly Gly Gly Leu Val Gln Pro Gly Gly1 5 10
15Ser Leu Arg Leu Ser Cys Ser Ala Ser Gly Phe Thr Phe
Ser Asn Tyr 20 25 30Trp Met
Thr Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Ile 35
40 45Ser Ser Ile Ser Ser Ser Gly Ser Thr Ile
Tyr Tyr Ala Asp Ser Val 50 55 60Lys
Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65
70 75 80Leu Gln Met Asn Ser Leu
Arg Val Glu Asp Thr Ala Val Tyr Tyr Cys 85
90 95Ala Arg Asp Tyr Trp Pro Gly Trp Tyr Phe Asp Leu
Trp Gly Arg Gly 100 105 110Thr
Leu Val Thr Val Ser Arg 1155965PRTHomo sapiens 596Asn Tyr Trp Met
Thr1 559717PRTHomo sapiens 597Ser Ile Ser Ser Ser Gly Ser
Thr Ile Tyr Tyr Ala Asp Ser Val Lys1 5 10
15Gly59810PRTHomo sapiens 598Asp Tyr Trp Pro Gly Trp Tyr
Phe Asp Leu1 5 10599116PRTHomo sapiens
599Gln Ala Val Leu Thr Gln Pro Ser Ser Leu Ser Ala Ser Pro Gly Ala1
5 10 15Ser Ala Ser Leu Thr Cys
Thr Phe Arg Ser Asp Ile Ser Val Gly Ser 20 25
30Tyr Arg Ile Tyr Trp Tyr Gln Gln Lys Pro Gly Ser Pro
Pro Gln Phe 35 40 45Leu Leu Lys
Tyr Thr Ser Asp Ser Asp Lys Gln Gln Gly Ser Gly Val 50
55 60Pro Ser Arg Phe Ser Gly Ser Lys Asp Val Ser Ala
Asn Ala Gly Ile65 70 75
80Leu Leu Ile Ser Gly Leu Gln Ser Glu Asp Glu Ala Asp Tyr Tyr Cys
85 90 95Met Thr Trp His Asn Thr
Ala Ser Val Phe Gly Gly Gly Thr Lys Leu 100
105 110Ala Val Leu Gly 11560014PRTHomo sapiens
600Thr Phe Arg Ser Asp Ile Ser Val Gly Ser Tyr Arg Ile Tyr1
5 1060111PRTHomo sapiens 601Tyr Thr Ser Asp Ser Asp Lys
Gln Gln Gly Ser1 5 106027PRTHomo sapiens
602Met Thr Trp His Asn Thr Ala1 5603127PRTHomo sapiens
603Gln Val Gln Leu Val Gln Ser Gly Ser Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys
Lys Thr Ser Gly Tyr Ile Phe Thr Asp Tyr 20 25
30Tyr Met His Trp Val Arg Gln Ala Pro Gly Lys Gly Leu
Glu Trp Met 35 40 45Gly Trp Ile
Asn Pro Asn Ser Gly Gly Thr Tyr Tyr Ala Gln Lys Phe 50
55 60His Gly Arg Val Thr Met Thr Ser Asp Thr Ser Ile
Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Asp Asp Thr Ala Ile Tyr Tyr Cys
85 90 95Ala Arg Glu Asp Tyr Asp
Ile Leu Thr Gly Tyr Tyr Pro Ala Ser Gly 100
105 110His Gly Asp Tyr Trp Gly Gln Gly Thr Leu Val Thr
Val Ser Arg 115 120 1256045PRTHomo
sapiens 604Asp Tyr Tyr Met His1 560517PRTHomo sapiens
605Trp Ile Asn Pro Asn Ser Gly Gly Thr Tyr Tyr Ala Gln Lys Phe His1
5 10 15Gly60618PRTHomo sapiens
606Glu Asp Tyr Asp Ile Leu Thr Gly Tyr Tyr Pro Ala Ser Gly His Gly1
5 10 15Asp Tyr607110PRTHomo
sapiens 607Gln Ala Val Val Thr Gln Glu Pro Ser Leu Thr Val Ser Pro Gly
Gly1 5 10 15Thr Val Thr
Leu Thr Cys Ala Ser Ser Thr Gly Ala Val Thr Ser Gly 20
25 30Phe Leu Ala Asn Trp Phe Gln Gln Lys Pro
Gly Gln Thr Pro Arg Ser 35 40
45Leu Ile Tyr Lys Thr Ser Asn Lys His Pro Trp Thr Pro Ala Arg Phe 50
55 60Ser Gly Ser Leu Leu Gly Gly Lys Ala
Ala Leu Thr Leu Ser Gly Val65 70 75
80Gln Pro Glu Asp Glu Ala Asp Tyr Tyr Cys Leu Leu Leu Ser
Gly Gly 85 90 95Ala Trp
Val Phe Gly Gly Gly Thr Lys Leu Ser Val Leu Gly 100
105 11060814PRTHomo sapiens 608Ala Ser Ser Thr Gly
Ala Val Thr Ser Gly Phe Leu Ala Asn1 5
106097PRTHomo sapiens 609Lys Thr Ser Asn Lys His Pro1
56107PRTHomo sapiens 610Leu Leu Leu Ser Gly Gly Ala1
5611122PRTHomo sapiens 611Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys
Lys Pro Gly Glu1 5 10
15Ser Leu Lys Ile Ser Cys Glu Gly Ser Gly Tyr Thr Phe Thr Ser Tyr
20 25 30Trp Ile Gly Trp Val Arg Gln
Met Pro Gly Lys Asp Leu Glu Trp Met 35 40
45Gly Ile Ile Tyr Pro Gly Asp Ser Asp Thr Arg Tyr Ser Pro Ser
Phe 50 55 60Gln Gly Gln Val Thr Ile
Ser Val Asp Lys Ser Ile Ser Thr Ala Tyr65 70
75 80Leu Gln Trp Ser Ser Leu Lys Ala Ser Asp Thr
Ala Met Tyr Tyr Cys 85 90
95Ala Arg His Asp Val Val Asp Gly Tyr Asn Thr Gly Met Asp Val Trp
100 105 110Gly Gln Gly Thr Thr Val
Thr Val Ser Arg 115 1206125PRTHomo sapiens 612Ser
Tyr Trp Ile Gly1 561317PRTHomo sapiens 613Ile Ile Tyr Pro
Gly Asp Ser Asp Thr Arg Tyr Ser Pro Ser Phe Gln1 5
10 15Gly61413PRTHomo sapiens 614His Asp Val Val
Asp Gly Tyr Asn Thr Gly Met Asp Val1 5
10615111PRTHomo sapiens 615Gln Thr Val Val Thr Gln Glu Pro Ser Phe Thr
Val Ser Pro Gly Gly1 5 10
15Thr Val Thr Leu Thr Cys Gly Leu Ser Ser Gly Ser Val Ser Thr Ser
20 25 30Tyr Tyr Pro Ser Trp Tyr Gln
Gln Thr Pro Gly Gln Ala Pro Arg Thr 35 40
45Leu Ile Ser Ser Thr Asn Thr Arg Ser Ser Gly Val Pro Asp Arg
Phe 50 55 60Ser Gly Ser Ile Leu Gly
Asn Arg Ala Ala Leu Thr Ile Thr Gly Ala65 70
75 80Gln Ala Asp Asp Glu Ser Asp Tyr Tyr Cys Val
Leu Tyr Met Gly Ser 85 90
95Gly Ile Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly
100 105 11061614PRTHomo sapiens 616Gly
Leu Ser Ser Gly Ser Val Ser Thr Ser Tyr Tyr Pro Ser1 5
106177PRTHomo sapiens 617Ser Thr Asn Thr Arg Ser Ser1
56188PRTHomo sapiens 618Val Leu Tyr Met Gly Ser Gly Ile1
5619128PRTHomo sapiens 619Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val
Lys Lys Pro Gly Glu1 5 10
15Ser Leu Lys Ile Ser Cys Lys Gly Ser Gly Tyr Ser Phe Thr Ser Tyr
20 25 30Trp Ile Gly Trp Val Arg Gln
Met Pro Gly Lys Gly Leu Glu Trp Met 35 40
45Gly Ile Ile Tyr Pro Gly Asp Ser Asp Ala Arg Tyr Ser Pro Ser
Phe 50 55 60Gln Gly Gln Val Thr Ile
Ser Ala Asp Lys Ser Ile Ser Thr Ala Tyr65 70
75 80Leu Gln Trp Ser Ser Leu Lys Ala Ser Asp Thr
Ala Met Tyr Tyr Cys 85 90
95Ala Arg Leu Thr Gly Ser Ser Tyr Tyr Asp Ser Ser Gly Tyr Ser Ser
100 105 110Tyr Gly Met Asp Val Trp
Gly Gln Gly Thr Thr Val Thr Val Ser Arg 115 120
1256205PRTHomo sapiens 620Ser Tyr Trp Ile Gly1
562117PRTHomo sapiens 621Ile Ile Tyr Pro Gly Asp Ser Asp Ala Arg Tyr Ser
Pro Ser Phe Gln1 5 10
15Gly62219PRTHomo sapiens 622Leu Thr Gly Ser Ser Tyr Tyr Asp Ser Ser Gly
Tyr Ser Ser Tyr Gly1 5 10
15Met Asp Val623111PRTHomo sapiens 623Gln Thr Val Val Thr Gln Glu Pro
Ser Phe Ser Val Ser Pro Gly Gly1 5 10
15Thr Val Thr Leu Thr Cys Gly Leu Ser Ser Gly Ser Val Ser
Thr Ser 20 25 30Tyr Tyr Pro
Ser Trp Phe Gln Gln Thr Pro Gly Gln Ala Pro Arg Thr 35
40 45Leu Ile Tyr Ser Thr Asn Thr Arg Ser Ser Gly
Val Pro Asp Arg Phe 50 55 60Ser Gly
Ser Ile Leu Gly Asn Lys Ala Ala Leu Thr Ile Thr Gly Ala65
70 75 80Gln Ala Asp Asp Glu Ser Asp
Tyr Tyr Cys Val Leu Tyr Met Gly Ser 85 90
95Gly Ile Ser Val Phe Gly Gly Gly Thr Gln Leu Thr Val
Leu Gly 100 105
11062414PRTHomo sapiens 624Gly Leu Ser Ser Gly Ser Val Ser Thr Ser Tyr
Tyr Pro Ser1 5 106257PRTHomo sapiens
625Ser Thr Asn Thr Arg Ser Ser1 56268PRTHomo sapiens 626Val
Leu Tyr Met Gly Ser Gly Ile1 5627121PRTHomo sapiens 627Gln
Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Glu1
5 10 15Ser Leu Lys Ile Ser Cys Lys
Gly Ser Gly Tyr Ser Phe Thr Ser Tyr 20 25
30Trp Ile Gly Trp Val Arg Gln Met Pro Gly Lys Gly Leu Glu
Trp Met 35 40 45Gly Ile Ile Tyr
Pro Gly Asp Ser Asp Thr Arg Tyr Ser Pro Ser Phe 50 55
60Gln Gly Gln Val Thr Ile Ser Ala Asp Lys Ser Ile Ser
Thr Ala Tyr65 70 75
80Leu Gln Trp Ser Ser Leu Lys Ala Ser Asp Thr Ala Met Tyr Tyr Cys
85 90 95Ala Arg His Asn Ser Asn
Tyr Tyr Tyr Tyr Tyr Met Asp Val Trp Gly 100
105 110Lys Gly Thr Leu Val Thr Val Ser Arg 115
1206285PRTHomo sapiens 628Ser Tyr Trp Ile Gly1
562917PRTHomo sapiens 629Ile Ile Tyr Pro Gly Asp Ser Asp Thr Arg Tyr Ser
Pro Ser Phe Gln1 5 10
15Gly63012PRTHomo sapiens 630His Asn Ser Asn Tyr Tyr Tyr Tyr Tyr Met Asp
Val1 5 10631111PRTHomo sapiens 631Gln Thr
Val Val Thr Gln Glu Pro Ser Phe Ser Val Ser Pro Gly Gly1 5
10 15Thr Val Thr Leu Thr Cys Gly Leu
Ser Ser Gly Ser Val Ser Thr Ser 20 25
30Tyr Tyr Pro Ser Trp Tyr Gln Gln Thr Pro Gly Gln Ala Pro Arg
Thr 35 40 45Leu Ile Tyr Ser Thr
Asn Thr Arg Ser Ser Gly Val Pro Asp Arg Phe 50 55
60Ser Gly Ser Ile Leu Gly Asn Lys Ala Ala Leu Thr Ile Thr
Gly Ala65 70 75 80Gln
Ala Asp Asp Glu Ser Asp Tyr Tyr Cys Val Leu Tyr Met Gly Ser
85 90 95Gly Ile Ser Val Phe Gly Gly
Gly Thr Lys Leu Thr Val Leu Gly 100 105
11063214PRTHomo sapiens 632Gly Leu Ser Ser Gly Ser Val Ser Thr
Ser Tyr Tyr Pro Ser1 5 106337PRTHomo
sapiens 633Ser Thr Asn Thr Arg Ser Ser1 56348PRTHomo
sapiens 634Val Leu Tyr Met Gly Ser Gly Ile1 5635127PRTHomo
sapiens 635Gln Val Gln Leu Val Gln Ser Gly Ser Glu Val Lys Lys Pro Gly
Ala1 5 10 15Ser Val Lys
Val Ser Cys Lys Thr Ser Gly Tyr Ile Phe Thr Asp Tyr 20
25 30Tyr Met His Trp Val Arg Gln Ala Pro Gly
Lys Gly Leu Glu Trp Met 35 40
45Gly Trp Ile Asn Pro Asn Ser Gly Gly Thr Tyr Tyr Ala Gln Lys Phe 50
55 60His Gly Arg Val Thr Met Thr Ser Asp
Thr Ser Ile Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Asp Asp Thr Ala Ile Tyr
Tyr Cys 85 90 95Ala Arg
Glu Asp Tyr Asp Ile Leu Thr Gly Phe Tyr Pro Ala Ser Gly 100
105 110His Gly Asp Tyr Trp Gly Gln Gly Thr
Leu Val Thr Val Ser Arg 115 120
1256365PRTHomo sapiens 636Asp Tyr Tyr Met His1
563717PRTHomo sapiens 637Trp Ile Asn Pro Asn Ser Gly Gly Thr Tyr Tyr Ala
Gln Lys Phe His1 5 10
15Gly63818PRTHomo sapiens 638Glu Asp Tyr Asp Ile Leu Thr Gly Phe Tyr Pro
Ala Ser Gly His Gly1 5 10
15Asp Tyr639107PRTHomo sapiens 639Glu Ile Val Leu Thr Gln Ser Pro Gly
Thr Leu Ser Leu Ser Pro Gly1 5 10
15Glu Arg Ala Thr Leu Ser Cys Arg Ala Ser Gln Ser Val Ser Ser
Ser 20 25 30Tyr Leu Ala Trp
Tyr Gln Gln Lys Pro Gly Gln Ala Pro Arg Leu Leu 35
40 45Ile Tyr Gly Ala Ser Ser Arg Ala Thr Gly Ile Pro
Asp Arg Phe Ser 50 55 60Gly Ser Gly
Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Arg Leu Glu65 70
75 80Pro Glu Asp Phe Ala Val Tyr Tyr
Cys Gln Gln Tyr Gly Ser Ser Leu 85 90
95Thr Phe Gly Gly Gly Thr Lys Val Glu Ile Lys 100
10564012PRTHomo sapiens 640Arg Ala Ser Gln Ser Val Ser
Ser Ser Tyr Leu Ala1 5 106417PRTHomo
sapiens 641Gly Ala Ser Ser Arg Ala Thr1 56426PRTHomo
sapiens 642Gln Gln Tyr Gly Ser Ser1 5643122PRTHomo sapiens
643Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Glu1
5 10 15Ser Leu Lys Ile Ser Cys
Lys Gly Ser Gly Tyr Ser Phe Thr Ser Tyr 20 25
30Trp Ile Gly Trp Val Arg Gln Met Pro Gly Lys Gly Leu
Glu Trp Met 35 40 45Gly Ile Ile
Tyr Pro Gly Asp Ser Asp Thr Arg Tyr Ser Pro Ser Phe 50
55 60Gln Gly Gln Val Thr Ile Ser Ala Asp Lys Ser Ile
Ser Thr Ala Tyr65 70 75
80Leu Gln Trp Ser Ser Leu Lys Ala Ser Asp Thr Ala Met Tyr Tyr Cys
85 90 95Ala Arg His Gly Met Thr
Ser Gly Tyr Val Ala His Asn Asp Tyr Trp 100
105 110Gly Gln Gly Thr Leu Val Thr Val Ser Arg 115
1206445PRTHomo sapiens 644Ser Tyr Trp Ile Gly1
564517PRTHomo sapiens 645Ile Ile Tyr Pro Gly Asp Ser Asp Thr Arg Tyr
Ser Pro Ser Phe Gln1 5 10
15Gly64613PRTHomo sapiens 646His Gly Met Thr Ser Gly Tyr Val Ala His Asn
Asp Tyr1 5 10647111PRTHomo sapiens 647Gln
Thr Val Val Thr Gln Glu Pro Ser Phe Ser Val Ser Pro Gly Gly1
5 10 15Thr Val Thr Leu Thr Cys Gly
Leu Ser Ser Gly Ser Val Ser Thr Ser 20 25
30Tyr Tyr Pro Ser Trp Tyr Gln Gln Thr Pro Gly Gln Ala Pro
Arg Thr 35 40 45Leu Ile Tyr Ser
Thr Asn Thr Arg Ser Ser Gly Val Pro Asp Arg Phe 50 55
60Ser Gly Ser Ile Leu Gly Asn Lys Ala Ala Leu Thr Ile
Thr Gly Ala65 70 75
80Gln Ala Asp Asp Glu Ser Asp Tyr Tyr Cys Val Leu Tyr Met Gly Ser
85 90 95Gly Ile Trp Val Phe Gly
Gly Gly Thr Lys Leu Thr Val Leu Gly 100 105
11064814PRTHomo sapiens 648Gly Leu Ser Ser Gly Ser Val Ser
Thr Ser Tyr Tyr Pro Ser1 5 106497PRTHomo
sapiens 649Ser Thr Asn Thr Arg Ser Ser1 56508PRTHomo
sapiens 650Val Leu Tyr Met Gly Ser Gly Ile1 5651117PRTHomo
sapiens 651Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Lys Pro Ser
Gly1 5 10 15Thr Leu Ser
Leu Thr Cys Ala Val Ser Gly Gly Ser Ile Ser Ser Ser 20
25 30Asn Trp Trp Ser Trp Val Arg Gln Pro Pro
Gly Lys Gly Leu Glu Trp 35 40
45Ile Gly Glu Ile Tyr His Ser Gly Ser Thr Asn Tyr Asn Pro Ser Leu 50
55 60Lys Ser Arg Val Thr Ile Ser Val Asp
Lys Ser Lys Asn Gln Phe Ser65 70 75
80Leu Lys Leu Ser Ser Val Thr Ala Ala Asp Thr Ala Val Tyr
Tyr Cys 85 90 95Ala Arg
Gly Gly Ser Tyr Tyr Phe Asp Tyr Trp Gly Gln Gly Thr Leu 100
105 110Val Thr Val Ser Arg
1156526PRTHomo sapiens 652Ser Ser Asn Trp Trp Ser1
565316PRTHomo sapiens 653Glu Ile Tyr His Ser Gly Ser Thr Asn Tyr Asn Pro
Ser Leu Lys Ser1 5 10
156548PRTHomo sapiens 654Gly Gly Ser Tyr Tyr Phe Asp Tyr1
5655112PRTHomo sapiens 655Gln Ser Ala Leu Thr Gln Pro Ala Ser Val Ser Gly
Ser Pro Gly Gln1 5 10
15Ser Ile Thr Ile Ser Cys Thr Gly Thr Ile Ser Asp Val Gly Gly Tyr
20 25 30Asp Phe Val Ser Trp Tyr Gln
His His Pro Gly Lys Ala Pro Lys Leu 35 40
45Leu Ile Tyr Asp Val Asn Asn Arg Pro Ser Gly Val Ser His Arg
Phe 50 55 60Ser Gly Ser Lys Ser Gly
Asn Thr Ala Ser Leu Thr Ile Ser Gly Leu65 70
75 80Gln Ala Glu Asp Glu Ala Thr Tyr Tyr Cys Ser
Ser Tyr Ser Asn Arg 85 90
95His Ser Leu Ile Val Phe Gly Ser Gly Thr Gln Val Val Gly Leu Gly
100 105 11065614PRTHomo sapiens
656Thr Gly Thr Ile Ser Asp Val Gly Gly Tyr Asp Phe Val Ser1
5 106577PRTHomo sapiens 657Asp Val Asn Asn Arg Pro Ser1
56589PRTHomo sapiens 658Ser Ser Tyr Ser Asn Arg His Ser
Leu1 5659121PRTHomo sapiens 659Gln Val Gln Leu Gln Gln Trp
Gly Ala Gly Leu Leu Lys Pro Ser Glu1 5 10
15Thr Leu Ser Leu Thr Cys Ala Val Tyr Gly Gly Ser Phe
Ser Gly Tyr 20 25 30Tyr Trp
Ser Trp Ile Arg Gln Pro Pro Gly Lys Gly Leu Glu Trp Ile 35
40 45Gly Glu Ile Asn His Ser Gly Ser Thr Asn
Tyr Asn Pro Ser Leu Lys 50 55 60Ser
Arg Val Thr Ile Ser Val Asp Thr Ser Lys Asn Gln Phe Ser Leu65
70 75 80Lys Leu Ser Ser Val Thr
Ala Ala Asp Thr Ala Val Tyr Tyr Cys Ala 85
90 95Arg Gly Gly Gly Val Glu Ala Leu Asn Tyr Gly Met
Asp Val Trp Gly 100 105 110Gln
Gly Thr Thr Val Thr Val Ser Arg 115 1206605PRTHomo
sapiens 660Gly Tyr Tyr Trp Ser1 566116PRTHomo sapiens
661Glu Ile Asn His Ser Gly Ser Thr Asn Tyr Asn Pro Ser Leu Lys Ser1
5 10 1566213PRTHomo sapiens
662Gly Gly Gly Val Glu Ala Leu Asn Tyr Gly Met Asp Val1 5
10663112PRTHomo sapiens 663Gln Ser Ala Leu Thr Gln Pro
Ala Ser Val Ser Gly Ser Pro Gly Gln1 5 10
15Thr Ile Thr Ile Ser Cys Thr Gly Thr Ile Ser Asp Val
Gly Gly Tyr 20 25 30Asp Phe
Val Ser Trp Tyr Gln His His Pro Gly Lys Ala Pro Lys Leu 35
40 45Leu Ile Tyr Asp Val Asn Asn Arg Pro Ser
Gly Val Ser His Arg Phe 50 55 60Ser
Gly Ser Lys Ser Gly Asn Thr Ala Ser Leu Thr Ile Ser Gly Leu65
70 75 80Gln Ala Glu Asp Glu Ala
Thr Tyr Tyr Cys Ser Ser Tyr Ser Asn Arg 85
90 95His Ser Leu Ile Val Phe Gly Ser Gly Thr Gln Val
Val Gly Leu Gly 100 105
11066414PRTHomo sapiens 664Thr Gly Thr Ile Ser Asp Val Gly Gly Tyr Asp
Phe Val Ser1 5 106657PRTHomo sapiens
665Asp Val Asn Asn Arg Pro Ser1 56669PRTHomo sapiens 666Ser
Ser Tyr Ser Asn Arg His Ser Leu1 5667119PRTHomo sapiens
667Gln Val Gln Leu Gln Gln Trp Gly Ala Gly Leu Leu Lys Pro Ser Glu1
5 10 15Thr Leu Ser Leu Thr Cys
Thr Val His Gly Gly Ser Phe Asp Asp Tyr 20 25
30Tyr Trp Thr Trp Ile Arg Gln Pro Pro Gly Gly Gly Leu
Glu Trp Ile 35 40 45Gly Glu Met
Asn Ser Gly Arg Thr Tyr Asn Tyr Asn Pro Phe Leu Glu 50
55 60Ser Arg Ala Ser Ile Asp Val Asp Thr Phe Lys Lys
Gln Phe Ser Leu65 70 75
80Ala Leu Arg Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr Cys Ala
85 90 95Arg Gly Ala Tyr Val Asn
Tyr Tyr Tyr Ile Asp Val Trp Gly Asp Gly 100
105 110Thr Thr Val Thr Val Ser Arg 1156685PRTHomo
sapiens 668Asp Tyr Tyr Trp Thr1 566916PRTHomo sapiens
669Glu Met Asn Ser Gly Arg Thr Tyr Asn Tyr Asn Pro Phe Leu Glu Ser1
5 10 1567011PRTHomo sapiens
670Gly Ala Tyr Val Asn Tyr Tyr Tyr Ile Asp Val1 5
10671112PRTHomo sapiens 671Gln Ser Ala Leu Thr Gln Pro Ala Ser
Val Ser Gly Ser Pro Gly Gln1 5 10
15Thr Ile Thr Ile Ser Cys Thr Gly Thr Ile Ser Asp Val Gly Gly
Tyr 20 25 30Asp Phe Val Ser
Trp Tyr Gln His His Pro Gly Lys Ala Pro Lys Leu 35
40 45Leu Ile Tyr Asp Val Asn Asn Arg Pro Ser Gly Val
Ser His Arg Phe 50 55 60Ser Gly Ser
Lys Ser Gly Asn Thr Ala Ser Leu Thr Ile Ser Gly Leu65 70
75 80Gln Ala Glu Asp Glu Ala Thr Tyr
Tyr Cys Ser Ser Tyr Ser Asn Arg 85 90
95His Ser Leu Ile Val Phe Gly Ser Gly Thr Gln Val Val Gly
Leu Gly 100 105
11067214PRTHomo sapiens 672Thr Gly Thr Ile Ser Asp Val Gly Gly Tyr Asp
Phe Val Ser1 5 106737PRTHomo sapiens
673Asp Val Asn Asn Arg Pro Ser1 56749PRTHomo sapiens 674Ser
Ser Tyr Ser Asn Arg His Ser Leu1 5675125PRTHomo sapiens
675Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Lys Pro Ser Glu1
5 10 15Thr Leu Ser Leu Thr Cys
Thr Val Ser Gly Gly Ser Ile Ser Ser Tyr 20 25
30Tyr Trp Ser Trp Ile Arg Gln Pro Ala Gly Lys Gly Leu
Glu Trp Ile 35 40 45Gly Arg Ile
Tyr Thr Ser Gly Ser Thr Asn Tyr Asn Pro Ser Leu Lys 50
55 60Ser Arg Val Thr Met Ser Val Asp Thr Ser Lys Asn
Gln Phe Ser Leu65 70 75
80Lys Leu Ser Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr Cys Ala
85 90 95Arg Glu Arg Ala Tyr Cys
Ser Ser Thr Ser Cys Tyr Arg Asn Ala Phe 100
105 110Asp Ile Trp Gly Gln Gly Thr Thr Val Thr Val Ser
Arg 115 120 1256765PRTHomo sapiens
676Ser Tyr Tyr Trp Ser1 567716PRTHomo sapiens 677Arg Ile
Tyr Thr Ser Gly Ser Thr Asn Tyr Asn Pro Ser Leu Lys Ser1 5
10 1567817PRTHomo sapiens 678Glu Arg
Ala Tyr Cys Ser Ser Thr Ser Cys Tyr Arg Asn Ala Phe Asp1 5
10 15Ile679113PRTHomo sapiens 679Gln
Ser Val Leu Thr Gln Pro Pro Ser Val Ser Gly Ala Pro Gly Gln1
5 10 15Arg Val Asn Ile Ser Cys Ala
Gly Ser Ser Ser Asn Ile Gly Ala Gly 20 25
30Tyr Asp Val His Trp Tyr Gln Gln Ile Pro Gly Thr Ala Pro
Lys Leu 35 40 45Leu Met Tyr Gly
Asn Ser Asn Arg Pro Ser Gly Val Pro Asp Arg Phe 50 55
60Ser Gly Ser Lys Ser Gly Ala Ser Ala Ser Leu Ala Ile
Thr Arg Leu65 70 75
80Gln Ala Glu Asp Glu Ala Asp Tyr Tyr Cys Gln Ser Tyr Asp Ser Ser
85 90 95Leu Ser Gly Ser Arg Val
Phe Gly Thr Gly Thr Lys Val Thr Val Leu 100
105 110Gly68014PRTHomo sapiens 680Ala Gly Ser Ser Ser Asn
Ile Gly Ala Gly Tyr Asp Val His1 5
106817PRTHomo sapiens 681Gly Asn Ser Asn Arg Pro Ser1
568210PRTHomo sapiens 682Gln Ser Tyr Asp Ser Ser Leu Ser Gly Ser1
5 10683122PRTHomo sapiens 683Glu Val Gln Leu Val
Glu Ser Gly Gly Gly Val Val Gln Pro Gly Arg1 5
10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe
Thr Phe Ser Ser Tyr 20 25
30Gly Met His Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45Thr Val Ile Ser Phe Asp Gly Ser
Asn Lys Tyr Tyr Ala Asp Ser Val 50 55
60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr65
70 75 80Leu Gln Met Asn Ser
Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys 85
90 95Ala Lys Ala Tyr Thr Asn Thr Trp Trp Pro Asp
Ala Phe Asp Ile Trp 100 105
110Gly Gln Gly Thr Thr Val Thr Val Ser Arg 115
1206845PRTHomo sapiens 684Ser Tyr Gly Met His1
568517PRTHomo sapiens 685Val Ile Ser Phe Asp Gly Ser Asn Lys Tyr Tyr Ala
Asp Ser Val Lys1 5 10
15Gly68613PRTHomo sapiens 686Ala Tyr Thr Asn Thr Trp Trp Pro Asp Ala Phe
Asp Ile1 5 10687107PRTHomo sapiens 687Asp
Ile Gln Met Thr Gln Ser Pro Ser Ser Leu Ser Ala Ser Val Gly1
5 10 15Asp Arg Val Thr Ile Thr Cys
Gln Ala Ser Gln Asp Ile Ser Asn Tyr 20 25
30Leu Asn Trp Tyr Gln Gln Lys Pro Gly Lys Ala Pro Lys Leu
Leu Ile 35 40 45Tyr Asp Ala Ser
Asn Leu Glu Thr Gly Val Pro Ser Arg Phe Ser Gly 50 55
60Ser Gly Ser Gly Thr Asp Phe Thr Phe Thr Ile Ser Ser
Leu Gln Pro65 70 75
80Glu Asp Ile Ala Thr Tyr Tyr Cys Gln Gln Tyr Asp Asn Leu Pro Pro
85 90 95Thr Phe Gly Pro Gly Thr
Lys Val Asp Ile Lys 100 10568811PRTHomo
sapiens 688Gln Ala Ser Gln Asp Ile Ser Asn Tyr Leu Asn1 5
106897PRTHomo sapiens 689Asp Ala Ser Asn Leu Glu Thr1
56907PRTHomo sapiens 690Gln Gln Tyr Asp Asn Leu Pro1
5691118PRTHomo sapiens 691Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu
Val Lys Pro Ser Glu1 5 10
15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Ser Ser Ile Ser Ser Tyr
20 25 30Tyr Trp Ser Trp Ile Arg Gln
Pro Pro Gly Lys Gly Leu Glu Trp Ile 35 40
45Gly Tyr Ile Tyr Tyr Ser Gly Ser Thr Asn Tyr Asn Pro Ser Leu
Lys 50 55 60Ser Arg Val Thr Ile Ser
Val Asp Thr Ser Lys Asn Gln Phe Ser Leu65 70
75 80Lys Leu Ser Ser Val Thr Ala Ala Asp Thr Ala
Val Tyr Tyr Cys Ala 85 90
95Arg Asp Leu Tyr Trp Asn Asp Ala Phe Asp Ile Trp Gly Gln Gly Thr
100 105 110Thr Val Thr Val Ser Arg
1156925PRTHomo sapiens 692Ser Tyr Tyr Trp Ser1
569316PRTHomo sapiens 693Tyr Ile Tyr Tyr Ser Gly Ser Thr Asn Tyr Asn Pro
Ser Leu Lys Ser1 5 10
1569410PRTHomo sapiens 694Asp Leu Tyr Trp Asn Asp Ala Phe Asp Ile1
5 10695111PRTHomo sapiens 695Gln Ser Ala Leu Thr
Gln Pro Leu Ser Ala Ser Gly Thr Pro Gly Gln1 5
10 15Arg Val Thr Ile Ser Cys Ser Gly Ser Ser Ser
Asn Ile Gly Ser Asn 20 25
30Thr Val Asn Trp Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys Leu Leu
35 40 45Ile Tyr Ser Asn Asn Gln Arg Pro
Ser Gly Val Pro Asp Arg Phe Ser 50 55
60Gly Ser Lys Ser Gly Thr Ser Ala Ser Leu Ala Ile Ser Gly Leu Gln65
70 75 80Ser Glu Asp Glu Ala
Asp Tyr Tyr Cys Ala Ala Trp Asp Asp Ser Leu 85
90 95Asn Gly Pro Val Phe Gly Gly Gly Thr Lys Leu
Thr Val Leu Gly 100 105
11069613PRTHomo sapiens 696Ser Gly Ser Ser Ser Asn Ile Gly Ser Asn Thr
Val Asn1 5 106977PRTHomo sapiens 697Ser
Asn Asn Gln Arg Pro Ser1 56989PRTHomo sapiens 698Ala Ala
Trp Asp Asp Ser Leu Asn Gly1 5699118PRTHomo sapiens 699Gln
Val Gln Leu Gln Glu Ser Gly Pro Gly Leu Val Lys Pro Ser Glu1
5 10 15Thr Leu Ser Leu Thr Cys Thr
Val Ser Gly Ser Ser Ile Ser Ser Tyr 20 25
30Tyr Trp Ser Trp Ile Arg Gln Pro Pro Gly Lys Gly Leu Glu
Trp Ile 35 40 45Gly Tyr Ile Tyr
Tyr Ser Gly Ser Thr Asn Tyr Asn Pro Ser Leu Lys 50 55
60Ser Arg Val Thr Ile Ser Val Asp Thr Ser Lys Asn Gln
Phe Ser Leu65 70 75
80Lys Leu Ser Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr Cys Ala
85 90 95Arg Asp Leu Tyr Trp Asn
Asp Ala Phe Asp Ile Trp Gly Gln Gly Thr 100
105 110Thr Val Thr Val Ser Arg 1157005PRTHomo
sapiens 700Ser Tyr Tyr Trp Ser1 570116PRTHomo sapiens
701Tyr Ile Tyr Tyr Ser Gly Ser Thr Asn Tyr Asn Pro Ser Leu Lys Ser1
5 10 1570210PRTHomo sapiens
702Asp Leu Tyr Trp Asn Asp Ala Phe Asp Ile1 5
10703111PRTHomo sapiens 703Gln Ser Val Leu Thr Gln Pro Pro Ser Ala
Ser Gly Thr Pro Gly Gln1 5 10
15Ser Val Thr Ile Ser Cys Ser Gly Ser Ser Ser Asn Ile Gly Ser Asn
20 25 30Thr Val Asn Trp Tyr Gln
Gln Leu Pro Gly Thr Ala Pro Lys Leu Leu 35 40
45Ile Tyr Ser Asn Asn Gln Arg Pro Ser Gly Val Pro Asp Arg
Phe Ser 50 55 60Gly Ser Lys Ser Gly
Thr Ser Ala Ser Leu Ala Ile Ser Gly Leu Gln65 70
75 80Ser Glu Asp Glu Ala Asp Tyr Tyr Cys Ala
Ala Trp Asp Asp Ser Leu 85 90
95Asn Gly Pro Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly
100 105 11070413PRTHomo sapiens
704Ser Gly Ser Ser Ser Asn Ile Gly Ser Asn Thr Val Asn1 5
107057PRTHomo sapiens 705Ser Asn Asn Gln Arg Pro Ser1
57069PRTHomo sapiens 706Ala Ala Trp Asp Asp Ser Leu Asn Gly1
5707126PRTHomo sapiens 707Gln Val Gln Leu Gln Gln Ser Gly Pro
Gly Leu Val Lys Pro Ser Gln1 5 10
15Thr Leu Ser Leu Thr Cys Ala Ile Ser Gly Asp Ser Val Ser Ser
Asn 20 25 30Ser Ala Ala Trp
Asn Trp Ile Arg Gln Ser Pro Ser Arg Gly Leu Glu 35
40 45Trp Leu Gly Arg Thr Tyr Tyr Arg Ser Lys Trp Tyr
Asn Asp Tyr Ala 50 55 60Val Ser Val
Lys Ser Arg Ile Thr Ile Asn Pro Asp Thr Ser Lys Asn65 70
75 80Gln Phe Ser Leu Gln Leu Asn Ser
Val Thr Pro Glu Asp Thr Ala Val 85 90
95Tyr Tyr Cys Ala Arg Glu Ser Trp Leu Trp Gly Ile Gly Gly
Asp Ala 100 105 110Phe Asp Ile
Trp Gly Gln Gly Thr Thr Val Thr Val Ser Arg 115
120 1257087PRTHomo sapiens 708Ser Asn Ser Ala Ala Trp
Asn1 570918PRTHomo sapiens 709Arg Thr Tyr Tyr Arg Ser Lys
Trp Tyr Asn Asp Tyr Ala Val Ser Val1 5 10
15Lys Ser71014PRTHomo sapiens 710Glu Ser Trp Leu Trp Gly
Ile Gly Gly Asp Ala Phe Asp Ile1 5
10711111PRTHomo sapiens 711Gln Ser Val Leu Thr Gln Pro Pro Ser Val Ser
Gly Ala Pro Arg Gln1 5 10
15Thr Val Thr Ile Ser Cys Ser Gly Ser Ser Ser Asn Ile Gly Gln Asn
20 25 30Ser Val Thr Trp Tyr Gln Arg
Leu Pro Gly Glu Ala Pro Lys Leu Leu 35 40
45Ile Tyr Tyr Asp Asp Leu Leu His Ser Gly Val Ser Asp Arg Phe
Ser 50 55 60Gly Ser Lys Ser Gly Thr
Ser Ala Ser Leu Ala Ile Ser Gly Leu Gln65 70
75 80Ser Glu Asp Glu Ala Glu Tyr Tyr Cys Ala Ser
Trp Asp Asp Ser Leu 85 90
95Lys Gly Pro Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly
100 105 11071213PRTHomo sapiens 712Ser
Gly Ser Ser Ser Asn Ile Gly Gln Asn Ser Val Thr1 5
107137PRTHomo sapiens 713Tyr Asp Asp Leu Leu His Ser1
57149PRTHomo sapiens 714Ala Ser Trp Asp Asp Ser Leu Lys Gly1
5715112PRTHomo sapiens 715Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu
Val Gln Pro Gly Gly1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Val Ser Ser Asn
20 25 30Tyr Met Ser Trp Val Arg Gln
Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40
45Ser Val Ile Tyr Ser Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val
Lys 50 55 60Gly Arg Phe Thr Ile Ser
Arg Asp Asn Ser Lys Asn Thr Leu Tyr Leu65 70
75 80Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala
Val Tyr Tyr Cys Ala 85 90
95Arg Glu Phe Asp Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser Arg
100 105 1107165PRTHomo sapiens 716Ser
Asn Tyr Met Ser1 571716PRTHomo sapiens 717Val Ile Tyr Ser
Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val Lys Gly1 5
10 157184PRTHomo sapiens 718Glu Phe Asp
Tyr1719105PRTHomo sapiens 719Asp Ile Gln Met Thr Gln Ser Pro Ser Ser Leu
Ser Ala Ser Val Gly1 5 10
15Asp Arg Val Thr Ile Thr Cys Gln Ala Ser Gln Asp Ile Ser Asn Tyr
20 25 30Leu Asn Trp Tyr Gln Gln Lys
Pro Gly Lys Ala Pro Lys Leu Leu Ile 35 40
45Tyr Asp Ala Ser Asn Leu Glu Thr Gly Val Pro Ser Arg Phe Ser
Gly 50 55 60Ser Gly Ser Gly Thr Asp
Phe Thr Phe Thr Ile Ser Ser Leu Gln Pro65 70
75 80Glu Asp Ile Ala Thr Tyr Tyr Cys Gln Gln Tyr
Asp Asn Pro Ala Phe 85 90
95Gly Gly Gly Thr Lys Val Glu Ile Lys 100
10572011PRTHomo sapiens 720Gln Ala Ser Gln Asp Ile Ser Asn Tyr Leu Asn1
5 107217PRTHomo sapiens 721Asp Ala Ser Asn
Leu Glu Thr1 57225PRTHomo sapiens 722Gln Gln Tyr Asp Asn1
5723121PRTHomo sapiens 723Gln Val Gln Leu Val Gln Ser Gly Ala
Glu Val Lys Lys Pro Gly Ala1 5 10
15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser
Tyr 20 25 30Gly Ile Ser Trp
Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45Gly Trp Ile Ser Ala Tyr Asn Gly Asn Thr Asn Tyr
Ala Gln Lys Leu 50 55 60Gln Gly Arg
Val Thr Met Thr Thr Asp Thr Ser Thr Ser Thr Ala Tyr65 70
75 80Met Glu Leu Arg Ser Leu Arg Ser
Asp Asp Thr Ala Val Tyr Tyr Cys 85 90
95Ala Arg Asp Arg Tyr Tyr Gly Ser Gly Phe Gly Met Asp Val
Trp Gly 100 105 110Gln Gly Thr
Met Val Thr Val Ser Arg 115 1207245PRTHomo sapiens
724Ser Tyr Gly Ile Ser1 572517PRTHomo sapiens 725Trp Ile
Ser Ala Tyr Asn Gly Asn Thr Asn Tyr Ala Gln Lys Leu Gln1 5
10 15Gly72612PRTHomo sapiens 726Asp Arg
Tyr Tyr Gly Ser Gly Phe Gly Met Asp Val1 5
10727109PRTHomo sapiens 727Ser Ser Glu Leu Thr Gln Asp Pro Ala Met Ser
Val Ala Leu Gly Gln1 5 10
15Thr Val Lys Ile Thr Cys Gln Gly Asp Ser Leu Thr Asn Tyr Tyr Pro
20 25 30Ser Trp Tyr Gln Gln Lys Pro
Gly Gln Ala Pro Val Leu Val Met Tyr 35 40
45Gly Lys Asp Ser Arg Pro Ser Gly Ile Ser Asp Arg Phe Ser Gly
Ser 50 55 60Ser Ser Gly Ile Ser Ala
Ser Leu Thr Ile Thr Gly Ala Gln Ala Glu65 70
75 80Asp Glu Ala Asp Tyr Tyr Cys Asn Ser Arg Asp
Gly Ser Ala His Arg 85 90
95Leu Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly 100
10572811PRTHomo sapiens 728Gln Gly Asp Ser Leu Thr Asn Tyr
Tyr Pro Ser1 5 107297PRTHomo sapiens
729Gly Lys Asp Ser Arg Pro Ser1 57309PRTHomo sapiens 730Asn
Ser Arg Asp Gly Ser Ala His Arg1 5731124PRTHomo sapiens
731Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys
Lys Ala Ser Gly Tyr Thr Phe Thr Asn Tyr 20 25
30Gly Leu Thr Trp Val Arg Gln Ala Pro Gly Gln Gly Leu
Glu Trp Met 35 40 45Gly Trp Ile
Ser Thr Tyr Asn Ser Asn Thr Asn Tyr Ala Glu Lys Leu 50
55 60Gln Gly Arg Val Thr Met Thr Thr Asp Thr Ser Thr
Ser Thr Ala Tyr65 70 75
80Met Glu Leu Arg Ser Leu Thr Ser Asp Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Gly Pro Thr Tyr
Ser Phe Asp Ser Ser Gly Tyr Phe Phe Asp 100
105 110Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser Arg
115 1207325PRTHomo sapiens 732Asn Tyr Gly Leu Thr1
573317PRTHomo sapiens 733Trp Ile Ser Thr Tyr Asn Ser Asn Thr
Asn Tyr Ala Glu Lys Leu Gln1 5 10
15Gly73415PRTHomo sapiens 734Gly Pro Thr Tyr Ser Phe Asp Ser Ser
Gly Tyr Phe Phe Asp Tyr1 5 10
15735109PRTHomo sapiens 735Ser Ser Glu Leu Thr Gln Asp Pro Ala Val
Ser Val Ala Leu Gly Gln1 5 10
15Thr Val Arg Ile Thr Cys Gln Gly Asp Ser Leu Arg Ser Tyr Tyr Ala
20 25 30Ser Trp Tyr Gln Gln Lys
Pro Gly Gln Ala Pro Val Leu Val Ile Tyr 35 40
45Gly Lys Asn Asn Arg Pro Ser Gly Ile Pro Asp Arg Phe Ser
Gly Ser 50 55 60Ser Ser Gly Asn Thr
Ala Ser Leu Thr Ile Thr Gly Ala Gln Ala Glu65 70
75 80Asp Glu Ala Asp Tyr Tyr Cys Asn Ser Arg
Asp Ser Ser Gly Asn His 85 90
95Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly 100
10573611PRTHomo sapiens 736Gln Gly Asp Ser Leu Arg Ser
Tyr Tyr Ala Ser1 5 107377PRTHomo sapiens
737Gly Lys Asn Asn Arg Pro Ser1 57389PRTHomo sapiens 738Asn
Ser Arg Asp Ser Ser Gly Asn His1 5739120PRTHomo sapiens
739Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys
Lys Ala Ser Gly Tyr Thr Phe Thr Ser Tyr 20 25
30Gly Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu
Glu Trp Met 35 40 45Gly Trp Ile
Ser Ala Tyr Asn Gly Asn Thr Asn Tyr Ala Gln Lys Leu 50
55 60Gln Gly Arg Val Thr Met Thr Thr Asp Thr Ser Thr
Ser Thr Ala Tyr65 70 75
80Met Glu Leu Arg Ser Leu Arg Ser Asp Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Asp Thr Tyr Ser
Ser Gly Trp Tyr Phe Asp Tyr Trp Gly Gln 100
105 110Gly Thr Leu Val Thr Val Ser Arg 115
1207405PRTHomo sapiens 740Ser Tyr Gly Ile Ser1
574117PRTHomo sapiens 741Trp Ile Ser Ala Tyr Asn Gly Asn Thr Asn Tyr Ala
Gln Lys Leu Gln1 5 10
15Gly74211PRTHomo sapiens 742Asp Thr Tyr Ser Ser Gly Trp Tyr Phe Asp Tyr1
5 10743110PRTHomo sapiens 743Ser Ser Glu
Leu Thr Gln Asp Pro Ala Val Ser Val Ala Leu Gly Gln1 5
10 15Thr Val Arg Ile Thr Cys Gln Gly Asp
Ser Leu Arg Asn Tyr Tyr Ala 20 25
30Ser Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Val Leu Val Ile Tyr
35 40 45Gly Lys Asn Asn Arg Pro Ser
Gly Ile Pro Asp Arg Phe Ser Gly Ser 50 55
60Ser Ser Gly Asn Thr Ala Ser Leu Thr Ile Thr Gly Ala Gln Ala Glu65
70 75 80Asp Glu Ala Asp
Tyr Tyr Cys Asn Ser Arg Asp Ser Ser Gly Asn His 85
90 95Leu Tyr Val Phe Gly Thr Gly Thr Lys Val
Thr Val Leu Gly 100 105
11074411PRTHomo sapiens 744Gln Gly Asp Ser Leu Arg Asn Tyr Tyr Ala Ser1
5 107457PRTHomo sapiens 745Gly Lys Asn Asn
Arg Pro Ser1 574610PRTHomo sapiens 746Asn Ser Arg Asp Ser
Ser Gly Asn His Leu1 5 10747120PRTHomo
sapiens 747Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys Lys Pro Gly
Ala1 5 10 15Ser Val Lys
Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser Tyr 20
25 30Gly Ile Ser Trp Val Arg Gln Ala Pro Gly
Gln Gly Leu Glu Trp Met 35 40
45Gly Trp Ile Ser Ala Tyr Asn Gly Asn Thr Asn Tyr Ala Gln Lys Leu 50
55 60Gln Gly Arg Val Thr Met Thr Thr Asp
Thr Ser Thr Ser Thr Ala Tyr65 70 75
80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr Ala Val Tyr
Tyr Cys 85 90 95Ala Thr
Leu Asn Ile Ser Gly Ser Tyr Tyr Phe Asp Tyr Trp Gly Gln 100
105 110Gly Thr Leu Val Thr Val Ser Arg
115 1207485PRTHomo sapiens 748Ser Tyr Gly Ile Ser1
574917PRTHomo sapiens 749Trp Ile Ser Ala Tyr Asn Gly Asn Thr Asn
Tyr Ala Gln Lys Leu Gln1 5 10
15Gly75011PRTHomo sapiens 750Leu Asn Ile Ser Gly Ser Tyr Tyr Phe Asp
Tyr1 5 10751110PRTHomo sapiens 751Ser Ser
Glu Leu Thr Gln Asp Pro Ala Val Ser Val Ala Leu Gly Gln1 5
10 15Thr Val Arg Ile Thr Cys Gln Gly
Asp Ser Leu Arg Ser Tyr Tyr Ala 20 25
30Thr Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Val Leu Val Ile
Ser 35 40 45Gly Lys Asn Asn Arg
Pro Ser Gly Ile Pro Asp Arg Phe Ser Gly Ser 50 55
60Ser Ser Gly Asn Thr Ala Ser Leu Thr Ile Thr Gly Ala Gln
Ala Glu65 70 75 80Asp
Glu Ala Asp Tyr Tyr Cys Asn Ser Arg Asp Ser Ser Gly Tyr Pro
85 90 95Ser Trp Val Phe Gly Gly Gly
Thr Lys Leu Thr Val Leu Gly 100 105
11075211PRTHomo sapiens 752Gln Gly Asp Ser Leu Arg Ser Tyr Tyr Ala
Thr1 5 107537PRTHomo sapiens 753Gly Lys
Asn Asn Arg Pro Ser1 575410PRTHomo sapiens 754Asn Ser Arg
Asp Ser Ser Gly Tyr Pro Ser1 5
10755118PRTHomo sapiens 755Gln Val Gln Leu Val Gln Ser Glu Ala Glu Val
Lys Lys Pro Gly Glu1 5 10
15Ser Leu Lys Ile Ser Cys Lys Gly Ser Gly Tyr Ser Phe Thr Ser Tyr
20 25 30Trp Ile Gly Trp Val Arg Gln
Met Pro Gly Lys Gly Leu Glu Trp Met 35 40
45Gly Ile Ile Tyr Pro Gly Asp Ser Asp Thr Arg Tyr Ser Pro Ser
Phe 50 55 60Gln Gly Gln Val Thr Ile
Ser Ala Asp Lys Ser Ile Ser Thr Ala Tyr65 70
75 80Leu Gln Trp Ser Ser Leu Lys Ala Ser Asp Thr
Ala Met Tyr Tyr Cys 85 90
95Ala Arg Thr Ile Gly Leu Gly Ala Phe Asp Ile Trp Gly Gln Gly Thr
100 105 110Thr Val Thr Val Ser Arg
1157565PRTHomo sapiens 756Ser Tyr Trp Ile Gly1
575717PRTHomo sapiens 757Ile Ile Tyr Pro Gly Asp Ser Asp Thr Arg Tyr Ser
Pro Ser Phe Gln1 5 10
15Gly7589PRTHomo sapiens 758Thr Ile Gly Leu Gly Ala Phe Asp Ile1
5759110PRTHomo sapiens 759Ser Ser Glu Leu Thr Gln Asp Pro Ala Val
Ser Val Ala Leu Gly Gln1 5 10
15Thr Val Arg Ile Thr Cys Gln Gly Asp Ser Leu Arg Ser Tyr Tyr Ala
20 25 30Ser Trp Tyr Gln Gln Lys
Pro Gly Gln Ala Pro Val Leu Val Ile Tyr 35 40
45Gly Lys Asn Asn Arg Pro Ser Gly Ile Pro Asp Arg Phe Ser
Gly Ser 50 55 60Ser Ser Gly Asn Thr
Ala Ser Leu Thr Ile Thr Gly Ala Gln Ala Glu65 70
75 80Asp Glu Ala Asp Tyr Tyr Cys Asn Ser Arg
Asp Ser Ser Gly Asn His 85 90
95His Tyr Val Phe Gly Thr Gly Thr Lys Val Thr Val Leu Gly
100 105 11076011PRTHomo sapiens 760Gln
Gly Asp Ser Leu Arg Ser Tyr Tyr Ala Ser1 5
107617PRTHomo sapiens 761Gly Lys Asn Asn Arg Pro Ser1
576210PRTHomo sapiens 762Asn Ser Arg Asp Ser Ser Gly Asn His His1
5 10763119PRTHomo sapiens 763Gln Val Gln Leu Gln
Glu Ser Gly Pro Gly Leu Val Lys Pro Ser Glu1 5
10 15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Gly
Ser Ile Ser Ser Ser 20 25
30Ser Tyr Tyr Trp Gly Trp Ile Arg Gln Pro Pro Gly Lys Gly Leu Glu
35 40 45Trp Ile Gly Ser Ile Tyr Tyr Ser
Gly Ser Thr Tyr Tyr Asn Pro Ser 50 55
60Leu Lys Ser Arg Val Thr Ile Ser Val Asp Thr Ser Lys Asn Gln Phe65
70 75 80Ser Leu Lys Leu Ser
Ser Val Thr Ala Ala Asp Thr Ala Val Tyr Tyr 85
90 95Cys Ala Arg His Trp Gly Asn Tyr Ala Phe Asp
Ile Trp Gly Gln Gly 100 105
110Thr Thr Val Thr Val Ser Arg 1157647PRTHomo sapiens 764Ser Ser
Ser Tyr Tyr Trp Gly1 576516PRTHomo sapiens 765Ser Ile Tyr
Tyr Ser Gly Ser Thr Tyr Tyr Asn Pro Ser Leu Lys Ser1 5
10 157669PRTHomo sapiens 766His Trp Gly Asn
Tyr Ala Phe Asp Ile1 5767111PRTHomo sapiens 767Gln Ser Val
Leu Thr Gln Pro Pro Ser Ala Ser Gly Thr Pro Gly Gln1 5
10 15Arg Val Thr Ile Ser Cys Ser Gly Ser
Ser Ser Asn Ile Gly Ser Asn 20 25
30Tyr Val Tyr Trp Tyr Gln Gln Leu Pro Gly Thr Ala Pro Lys Leu Leu
35 40 45Ile Tyr Ser Asn Asn Gln Arg
Pro Ser Gly Val Pro Asp Arg Phe Ser 50 55
60Gly Ser Lys Ser Gly Thr Ser Ala Ser Leu Ala Ile Ser Gly Leu Arg65
70 75 80Ser Glu Asp Glu
Ala Asp Tyr Tyr Cys Ala Ala Trp Asp Asp Ser Leu 85
90 95Ser Gly Gln Val Phe Gly Gly Gly Thr Gln
Leu Thr Val Leu Gly 100 105
11076813PRTHomo sapiens 768Ser Gly Ser Ser Ser Asn Ile Gly Ser Asn Tyr
Val Tyr1 5 107697PRTHomo sapiens 769Ser
Asn Asn Gln Arg Pro Ser1 57709PRTHomo sapiens 770Ala Ala
Trp Asp Asp Ser Leu Ser Gly1 5771120PRTHomo sapiens 771Glu
Val Gln Leu Val Glu Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1
5 10 15Ser Val Lys Val Ser Cys Lys
Ala Ser Gly Tyr Thr Phe Thr Ser Tyr 20 25
30Gly Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu
Trp Met 35 40 45Gly Trp Ile Ser
Ala Tyr Asn Gly Asn Thr Asn Tyr Ala Gln Lys Leu 50 55
60Gln Gly Arg Val Thr Met Thr Thr Asp Thr Ser Thr Ser
Thr Ala Tyr65 70 75
80Met Glu Leu Arg Ser Leu Arg Ser Asp Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Glu Lys Tyr Ser
Ser Gly Trp Tyr Phe Asp Tyr Trp Gly Gln 100
105 110Gly Thr Leu Val Thr Val Ser Arg 115
1207725PRTHomo sapiens 772Ser Tyr Gly Ile Ser1
577317PRTHomo sapiens 773Trp Ile Ser Ala Tyr Asn Gly Asn Thr Asn Tyr Ala
Gln Lys Leu Gln1 5 10
15Gly77411PRTHomo sapiens 774Glu Lys Tyr Ser Ser Gly Trp Tyr Phe Asp Tyr1
5 10775110PRTHomo sapiens 775Ser Ser Glu
Leu Thr Gln Asp Pro Ala Val Ser Val Ala Leu Gly Gln1 5
10 15Thr Val Arg Ile Thr Cys Gln Gly Asp
Ser Leu Arg Ser Tyr Tyr Ala 20 25
30Ser Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Val Leu Val Ile Tyr
35 40 45Gly Lys Asn Asn Arg Pro Ser
Gly Ile Pro Asp Arg Phe Ser Gly Ser 50 55
60Ser Ser Gly Asn Thr Ala Ser Leu Thr Ile Thr Gly Ala Gln Ala Glu65
70 75 80Asp Glu Ala Asp
Tyr Tyr Cys Asn Ser Arg Asp Ser Ser Gly Asn His 85
90 95His Tyr Val Phe Gly Thr Gly Thr Lys Val
Thr Val Leu Gly 100 105
11077611PRTHomo sapiens 776Gln Gly Asp Ser Leu Arg Ser Tyr Tyr Ala Ser1
5 107777PRTHomo sapiens 777Gly Lys Asn Asn
Arg Pro Ser1 577810PRTHomo sapiens 778Asn Ser Arg Asp Ser
Ser Gly Asn His His1 5 10779115PRTHomo
sapiens 779Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly
Gly1 5 10 15Ser Leu Arg
Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20
25 30Ala Met Ser Trp Val Arg Gln Ala Pro Gly
Lys Gly Leu Glu Trp Val 35 40
45Ser Ala Ile Ser Gly Ser Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val 50
55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp
Asn Ser Lys Asn Thr Leu Tyr65 70 75
80Leu Gln Ile Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr
Tyr Cys 85 90 95Ala Lys
Gly Ser Ile Ala Ala Asp Trp Gly Gln Gly Thr Leu Val Thr 100
105 110Val Ser Arg 1157805PRTHomo
sapiens 780Ser Tyr Ala Met Ser1 578117PRTHomo sapiens
781Ala Ile Ser Gly Ser Gly Gly Ser Thr Tyr Tyr Ala Asp Ser Val Lys1
5 10 15Gly7826PRTHomo sapiens
782Gly Ser Ile Ala Ala Asp1 5783108PRTHomo sapiens 783Ser
Ser Glu Leu Thr Gln Asp Pro Ala Val Ser Val Ala Leu Gly Gln1
5 10 15Thr Val Arg Ile Thr Cys Gln
Gly Asp Ser Leu Arg Ser Tyr Tyr Ala 20 25
30Ser Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Val Leu Val
Ile Tyr 35 40 45Gly Lys Asn Asn
Arg Pro Ser Gly Ile Pro Asp Arg Phe Ser Gly Ser 50 55
60Ser Ser Gly Asn Thr Ala Ser Leu Thr Ile Thr Gly Ala
Gln Ala Glu65 70 75
80Asp Glu Ala Asp Tyr Tyr Cys Asn Ser Arg Asp Ser Ser Gly Asn Val
85 90 95Val Phe Gly Gly Gly Thr
Lys Leu Thr Val Leu Gly 100 10578411PRTHomo
sapiens 784Gln Gly Asp Ser Leu Arg Ser Tyr Tyr Ala Ser1 5
107857PRTHomo sapiens 785Gly Lys Asn Asn Arg Pro Ser1
57868PRTHomo sapiens 786Asn Ser Arg Asp Ser Ser Gly Asn1
5787115PRTHomo sapiens 787Glu Val Gln Leu Val Glu Ser Gly Gly Gly
Leu Val Gln Pro Gly Gly1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr
20 25 30Ala Met Ser Trp Val Arg
Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40
45Ser Ala Ile Ser Gly Ser Gly Gly Ser Thr Tyr Tyr Ala Asp
Ser Val 50 55 60Lys Gly Arg Phe Thr
Ile Ser Arg Asp Asn Ser Lys Asn Ser Leu Tyr65 70
75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90
95Ala Ser Glu Gln Ala Gly Asp Tyr Trp Gly Gln Gly Thr Leu Val Thr
100 105 110Val Ser Arg
1157885PRTHomo sapiens 788Ser Tyr Ala Met Ser1
578917PRTHomo sapiens 789Ala Ile Ser Gly Ser Gly Gly Ser Thr Tyr Tyr Ala
Asp Ser Val Lys1 5 10
15Gly7906PRTHomo sapiens 790Glu Gln Ala Gly Asp Tyr1
5791111PRTHomo sapiens 791Gln Ser Val Leu Thr Gln Pro Pro Ser Ala Ser Gly
Thr Pro Gly Gln1 5 10
15Arg Val Thr Ile Ser Cys Ser Gly Ser Ser Ser Asn Ile Gly Ser Asn
20 25 30Tyr Val Tyr Trp Tyr Gln Gln
Leu Pro Gly Thr Ala Pro Lys Leu Leu 35 40
45Ile Tyr Arg Asn Asn Gln Arg Pro Ser Gly Val Pro Asp Arg Phe
Ala 50 55 60Gly Ser Lys Ser Gly Thr
Ser Ala Thr Leu Gly Ile Thr Gly Leu Gln65 70
75 80Thr Gly Asp Glu Ala Asp Tyr Tyr Cys Gly Thr
Trp Asp Ser Ser Leu 85 90
95Ser Ala Gly Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly
100 105 11079213PRTHomo sapiens 792Ser
Gly Ser Ser Ser Asn Ile Gly Ser Asn Tyr Val Tyr1 5
107937PRTHomo sapiens 793Arg Asn Asn Gln Arg Pro Ser1
57949PRTHomo sapiens 794Gly Thr Trp Asp Ser Ser Leu Ser Ala1
5795119PRTHomo sapiens 795Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val
Lys Lys Pro Gly Ala1 5 10
15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser Tyr
20 25 30Tyr Met His Trp Val Arg Gln
Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40
45Gly Ile Ile Asn Pro Ser Gly Gly Ser Thr Ser Tyr Ala Gln Lys
Phe 50 55 60Gln Gly Arg Val Thr Met
Thr Arg Asp Thr Ser Thr Ser Thr Val Tyr65 70
75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90
95Ala Arg Glu Leu Ser Ala Ala Thr Ala Phe Asp Ile Trp Gly Gln Gly
100 105 110Thr Met Val Thr Val Ser
Arg 1157965PRTHomo sapiens 796Ser Tyr Tyr Met His1
579717PRTHomo sapiens 797Ile Ile Asn Pro Ser Gly Gly Ser Thr Ser Tyr Ala
Gln Lys Phe Gln1 5 10
15Gly79810PRTHomo sapiens 798Glu Leu Ser Ala Ala Thr Ala Phe Asp Ile1
5 10799107PRTHomo sapiens 799Glu Thr Thr Leu
Thr Gln Ser Pro Phe Ser Val Ser Ala Ser Val Gly1 5
10 15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser
Gln Tyr Ile Ser Arg Trp 20 25
30Leu Ala Trp Tyr Gln Gln Arg Pro Gly Lys Ala Pro Lys Leu Leu Ile
35 40 45Tyr Ala Ala Ala Ser Leu Gln Ser
Gly Val Pro Ser Arg Phe Ser Gly 50 55
60Ser Gly Ser Gly Thr Asp Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65
70 75 80Glu Asp Phe Ala Thr
Tyr Tyr Cys Gln Gln Ala Asn Ser Phe Pro Val 85
90 95Thr Phe Gly Gln Gly Thr Arg Leu Glu Ile Lys
100 10580011PRTHomo sapiens 800Arg Ala Ser Gln
Tyr Ile Ser Arg Trp Leu Ala1 5
108017PRTHomo sapiens 801Ala Ala Ala Ser Leu Gln Ser1
58027PRTHomo sapiens 802Gln Gln Ala Asn Ser Phe Pro1
5803120PRTHomo sapiens 803Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys
Lys Pro Gly Ala1 5 10
15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr Ser Tyr
20 25 30Gly Ile Ser Trp Val Arg Gln
Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40
45Gly Trp Ile Ser Ala Tyr Asn Gly Asn Thr Asn Tyr Ala Gln Lys
Leu 50 55 60Gln Gly Arg Val Thr Met
Thr Thr Asp Thr Ser Thr Ser Thr Ala Tyr65 70
75 80Met Glu Leu Arg Ser Leu Arg Ser Asp Asp Thr
Ala Val Tyr Tyr Cys 85 90
95Ala Arg Val Gly Tyr Tyr Tyr Tyr Tyr Gly Met Asp Val Trp Gly Gln
100 105 110Gly Thr Thr Val Thr Val
Ser Arg 115 1208045PRTHomo sapiens 804Ser Tyr Gly
Ile Ser1 580517PRTHomo sapiens 805Trp Ile Ser Ala Tyr Asn
Gly Asn Thr Asn Tyr Ala Gln Lys Leu Gln1 5
10 15Gly80611PRTHomo sapiens 806Val Gly Tyr Tyr Tyr Tyr
Tyr Gly Met Asp Val1 5 10807111PRTHomo
sapiens 807Gln Ser Ala Leu Thr Gln Pro Ala Ser Val Ser Gly Ser Pro Gly
Gln1 5 10 15Ser Ile Thr
Ile Ser Cys Thr Gly Thr Ser Ser Asp Val Gly Ser Tyr 20
25 30Asn Leu Val Ser Trp Tyr Gln Gln His Pro
Gly Lys Ala Pro Lys Leu 35 40
45Met Ile Tyr Glu Gly Ser Lys Arg Pro Ser Gly Val Ser Asn Arg Phe 50
55 60Ser Gly Ser Lys Ser Gly Asn Thr Ala
Ser Leu Thr Ile Ser Gly Leu65 70 75
80Gln Ala Glu Asp Glu Ala Asp Tyr Tyr Cys Ser Ser Tyr Thr
Ser Ser 85 90 95Ser Thr
Trp Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly 100
105 11080814PRTHomo sapiens 808Thr Gly Thr Ser
Ser Asp Val Gly Ser Tyr Asn Leu Val Ser1 5
108097PRTHomo sapiens 809Glu Gly Ser Lys Arg Pro Ser1
58108PRTHomo sapiens 810Ser Ser Tyr Thr Ser Ser Ser Thr1
5811118PRTHomo sapiens 811Gln Val Gln Leu Val Gln Ser Gly Ala Glu Val Lys
Lys Pro Gly Ser1 5 10
15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Gly Thr Phe Ser Ser Tyr
20 25 30Ala Ile Ser Trp Val Arg Gln
Ala Pro Gly Gln Gly Leu Glu Trp Met 35 40
45Gly Trp Met Asn Pro Asn Ser Gly Asn Thr Gly Tyr Ala Gln Lys
Phe 50 55 60Gln Gly Arg Val Thr Met
Thr Arg Asn Thr Ser Ile Ser Thr Ala Tyr65 70
75 80Met Glu Leu Ser Ser Leu Arg Ser Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90
95Ala Arg Asp Val Gly Ile Gly Val Phe Asp Tyr Trp Gly Gln Gly Thr
100 105 110Leu Val Thr Val Ser Arg
1158125PRTHomo sapiens 812Ser Tyr Ala Ile Ser1
581317PRTHomo sapiens 813Trp Met Asn Pro Asn Ser Gly Asn Thr Gly Tyr Ala
Gln Lys Phe Gln1 5 10
15Gly8149PRTHomo sapiens 814Asp Val Gly Ile Gly Val Phe Asp Tyr1
5815109PRTHomo sapiens 815Ser Tyr Val Leu Thr Gln Pro Pro Ser Val
Ser Val Ala Pro Gly Lys1 5 10
15Thr Ala Arg Ile Thr Cys Gly Gly Asn Asn Ile Gly Ser Lys Ser Val
20 25 30His Trp Tyr Gln Gln Lys
Pro Gly Gln Ala Pro Val Leu Val Ile Tyr 35 40
45Tyr Asp Ser Asp Arg Pro Ser Gly Ile Pro Glu Arg Phe Ser
Gly Ser 50 55 60Asn Ser Gly Asn Thr
Ala Thr Leu Thr Ile Ser Arg Val Glu Ala Gly65 70
75 80Asp Glu Ala Asp Tyr Tyr Cys Gln Val Trp
Asp Ser Ser Ser Asp His 85 90
95Val Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly 100
10581611PRTHomo sapiens 816Gly Gly Asn Asn Ile Gly Ser
Lys Ser Val His1 5 108177PRTHomo sapiens
817Tyr Asp Ser Asp Arg Pro Ser1 58189PRTHomo sapiens 818Gln
Val Trp Asp Ser Ser Ser Asp His1 5819121PRTHomo sapiens
819Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1
5 10 15Ser Leu Arg Leu Ser Cys
Ala Ala Ser Gly Phe Thr Phe Ser Asp Tyr 20 25
30Ala Met Asn Trp Val Arg Gln Ala Pro Gly Lys Gly Leu
Glu Trp Val 35 40 45Ser Ala Thr
Ser Gly Ser Gly Gly Ser Thr Phe Tyr Ala Asp Ser Val 50
55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys
Ser Thr Leu Tyr65 70 75
80Leu Gln Met Asn Ser Leu Arg Asp Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Lys Gly Thr Leu Pro
His Tyr Tyr Asp Ser Ser Gly Ile Gly Gly 100
105 110Gln Gly Thr Leu Val Thr Val Ser Ser 115
1208205PRTHomo sapiens 820Asp Tyr Ala Met Asn1
582117PRTHomo sapiens 821Ala Thr Ser Gly Ser Gly Gly Ser Thr Phe Tyr Ala
Asp Ser Val Lys1 5 10
15Gly82212PRTHomo sapiens 822Gly Thr Leu Pro His Tyr Tyr Asp Ser Ser Gly
Ile1 5 10823113PRTHomo sapiens 823Gln Ala
Val Leu Thr Gln Pro Ser Ser Val Ser Gly Ser Pro Gly Gln1 5
10 15Ser Ile Thr Ile Ser Cys Thr Gly
Thr Ser Ser Asp Val Gly Gly Tyr 20 25
30Asn Tyr Val Ser Trp Tyr Gln Gln His Pro Gly Lys Ala Pro Lys
Leu 35 40 45Met Ile Tyr Asp Val
Gly Lys Arg Pro Ser Gly Val Ser Asn Arg Phe 50 55
60Ser Gly Ser Lys Ser Gly Asn Thr Ala Ser Leu Thr Ile Ser
Gly Leu65 70 75 80Arg
Ala Glu Asp Glu Ala Asn Tyr Tyr Cys Ser Ser Tyr Thr Ser Ser
85 90 95Ser Thr Trp Phe Val Val Phe
Gly Gly Gly Thr Lys Leu Thr Val Leu 100 105
110Gly82414PRTHomo sapiens 824Thr Gly Thr Ser Ser Asp Val
Gly Gly Tyr Asn Tyr Val Ser1 5
108257PRTHomo sapiens 825Asp Val Gly Lys Arg Pro Ser1
582610PRTHomo sapiens 826Ser Ser Tyr Thr Ser Ser Ser Thr Trp Phe1
5 10827121PRTHomo sapiens 827Gln Val Gln Leu Gln
Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ala1 5
10 15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr
Thr Phe Thr Thr Tyr 20 25
30Tyr Met His Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met
35 40 45Gly Ile Ile Asn Pro Ser Gly Glu
Asn Thr Asn Tyr Ala Gln Lys Phe 50 55
60Gln Gly Arg Val Thr Met Thr Arg Asp Thr Ser Thr Thr Thr Val Tyr65
70 75 80Met Glu Leu Ser Ser
Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85
90 95Ala Arg Ser Phe Ile Gly Thr Arg Gly Gly Gly
Leu Asp Val Trp Gly 100 105
110Gln Gly Thr Thr Val Thr Val Ser Arg 115
1208285PRTHomo sapiens 828Thr Tyr Tyr Met His1
582917PRTHomo sapiens 829Ile Ile Asn Pro Ser Gly Glu Asn Thr Asn Tyr Ala
Gln Lys Phe Gln1 5 10
15Gly83012PRTHomo sapiens 830Ser Phe Ile Gly Thr Arg Gly Gly Gly Leu Asp
Val1 5 10831113PRTHomo sapiens 831Gln Ser
Val Leu Thr Gln Pro Pro Ser Val Ser Gly Ala Pro Gly Gln1 5
10 15Arg Val Asn Ile Ser Cys Ala Gly
Ser Ser Ser Asn Ile Gly Ala Gly 20 25
30Tyr Asp Val His Trp Tyr Gln Gln Ile Pro Gly Thr Ala Pro Lys
Leu 35 40 45Leu Met Tyr Gly Asn
Ser Asn Arg Pro Ser Gly Val Pro Asp Arg Phe 50 55
60Ser Gly Ser Lys Ser Gly Ala Ser Ala Ser Leu Ala Ile Thr
Arg Leu65 70 75 80Gln
Ala Glu Asp Glu Ala Asp Tyr Tyr Cys Gln Ser Tyr Asp Ser Ser
85 90 95Leu Ser Gly Ser Arg Val Phe
Gly Thr Gly Thr Lys Val Thr Val Leu 100 105
110Gly83214PRTHomo sapiens 832Ala Gly Ser Ser Ser Asn Ile
Gly Ala Gly Tyr Asp Val His1 5
108337PRTHomo sapiens 833Gly Asn Ser Asn Arg Pro Ser1
583410PRTHomo sapiens 834Gln Ser Tyr Asp Ser Ser Leu Ser Gly Ser1
5 10835117PRTHomo sapiens 835Glu Val Gln Leu Val
Glu Ser Gly Gly Gly Val Val Gln Pro Gly Arg1 5
10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe
Thr Phe Asp Asp Tyr 20 25
30Ala Met His Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45Ser Gly Ile Ser Trp Asn Ser Gly
Ser Ile Gly Tyr Ala Asp Ser Val 50 55
60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65
70 75 80Leu Gln Met Asn Ser
Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys 85
90 95Ala Thr Glu Glu Trp Trp Arg Phe Asp Leu Trp
Gly Arg Gly Thr Leu 100 105
110Val Thr Val Ser Arg 1158365PRTHomo sapiens 836Asp Tyr Ala Met
His1 583717PRTHomo sapiens 837Gly Ile Ser Trp Asn Ser Gly
Ser Ile Gly Tyr Ala Asp Ser Val Lys1 5 10
15Gly8388PRTHomo sapiens 838Glu Glu Trp Trp Arg Phe Asp
Leu1 5839111PRTHomo sapiens 839Gln Ser Ala Leu Thr Gln Pro
Pro Ser Ala Ser Gly Ser Pro Gly Gln1 5 10
15Ser Val Thr Ile Ser Cys Thr Gly Thr Ser Ser Asp Val
Asp Asp Tyr 20 25 30Asn Tyr
Val Ser Trp Tyr Gln Gln His Pro Gly Lys Ala Pro Lys Leu 35
40 45Met Ile Tyr Glu Val Thr Lys Arg Pro Ser
Gly Val Pro Asp Arg Phe 50 55 60Ser
Gly Ser Lys Ser Gly Asn Thr Ala Ser Leu Thr Ile Ser Gly Leu65
70 75 80Gln Ala Glu Asp Glu Ala
Asp Tyr Tyr Cys Ser Ser Tyr Val Gly Ser 85
90 95Asp Asn Arg Val Phe Gly Thr Gly Thr Lys Val Thr
Val Pro Gly 100 105
11084014PRTHomo sapiens 840Thr Gly Thr Ser Ser Asp Val Asp Asp Tyr Asn
Tyr Val Ser1 5 108417PRTHomo sapiens
841Glu Val Thr Lys Arg Pro Ser1 58428PRTHomo sapiens 842Ser
Ser Tyr Val Gly Ser Asp Asn1 5843363DNAHomo sapiens
843caggtgcagc tggtgcagtc tggggctgag gtgaagaagc ctggggcctc agtgaaggtc
60tcctgcaagg cttctggata caccttcacc ggctactata tgcactgggt gcgacaggcc
120cctggacaag ggcttgagtg gatgggatgg atcaacccta acagtggtgg cacaaactat
180gcacagaagt ttcagggcag ggtcaccatg accagggaca cgtccatcag cacagcctac
240atggagctga gcaggctgag atctgacgac acggccgtgt attactgtgc gagatccgat
300attttgactg gttattatgc ttttgatatc tggggccaag ggacaatggt caccgtctcg
360aga
363844321DNAHomo sapiens 844gacatccaga tgacccagtc tccatcctcc ctgtctgcat
ctgtaggaga cagagtcacc 60atcacttgcc gggcaagtca gagcattagc agctatttaa
attggtatca gcagaaacca 120gggaaagccc ctaagctcct gatctatgct gcatccagtt
tgcaaagtgg ggtcccatca 180aggttcagtg gcagtggatc tgggacagat ttcactctca
ccatcagcag tctgcaacct 240gaagattttg caacttacta ctgtcaacag agttacagta
ccacgtggac gttcggccaa 300gggaccaagg tggaaatcaa a
321845387DNAHomo sapiens 845gaggtgcagc tggtggagtc
tggggctgag gtgaagaagc ctggggcctc agtgaaggtc 60tcctgcaagg cttctggata
caccttcacc ggctactata tgcactgggt gcgacaggcc 120cctggacaag ggcttgagtg
gatgggatgg atcaacccta acagtggtgg cacaaactat 180gcacagaagt ttcagggcag
ggtcaccatg accagggaca cgtccatcag cacagcctac 240atggagctga gcaggctgag
atctgacgac acggccgtgt attactgtgc gagagattta 300tggtattact atggttcggg
gagttcactg tactactact acggtatgga cgtctggggc 360caagggacca cggtcaccgt
ctcgaga 387846336DNAHomo sapiens
846cagtctgtgt tgacgcagcc gccctcagtg tctggggccc cagggcagag ggtcaccatc
60tcctgcactg ggagcagctc caacatcggg gcaggttatg atgtacactg gtaccagcag
120cttccaggaa cagcccccaa actcctcatc tatggtaaca gcaatcggcc ctcaggggtc
180cctgaccgat tctctggctc caagtctggc acctcagcct ccttggccat cactgggctc
240caggctgagg atgaggctga ttattactgc cagtcctatg acagcagcct gagtggtgtg
300gtattcggcg gagggaccaa gctgaccgtc ctaggt
336847366DNAHomo sapiens 847gaggtgcagc tggtggagtc tgggggaggc ttggtacagc
ctggcaggtc cctgagactc 60tcctgtgcag cctctggatt cacctttgat gattatgcca
tgcactgggt ccggcaagct 120ccagggaagg gcctggagtg ggtctcaggt attagttgga
atagtggtag cataggctat 180gcggactctg tgaagggccg attcaccatc tccagagaca
acgccaagaa ctccctgtat 240ctgcaaatga acagtctgag agctgaggac acggccttgt
attactgtgt acggagtggg 300agctacaact actactacta cggtatggac gtctggggcc
aagggaccac ggtcaccgtc 360tcgaga
366848333DNAHomo sapiens 848cagtctgtgt tgacgcagcc
gccctcagcg tctgggaccc ccgggcagag ggtcaccatc 60tcttgttctg gaagcagctc
caacatcgga agtaatactg taaactggta ccagcagctc 120ccaggaacgg cccccaaact
cctcatctat agtaataatc agcggccctc aggggtccct 180gaccgattct ctggctccaa
gtctggcacc tcagcctccc tggccatcag tgggctccag 240tctgaggatg aggctgatta
ttactgtgca gcatgggatg acagcctgaa tggttgggtg 300ttcggcggag ggaccaagct
gaccgtccta ggt 333849369DNAHomo sapiens
849caggtgcagc tgcaggagtc ggggggaggc ttggtacagc ctggggggtc cctgagactc
60tcctgtacaa cctctggatt cacctttacc agctatgcca tgagctgggt ccgccaggct
120ccagggaagg ggctggagtg ggtctcatct attagtggta gtggtggtat cacatactac
180gcagactccg tgaagggccg gttcaccatc tccagagaca attccaagaa cacactgtat
240ctgcaaatga acagcctaag agccgaggac acggccgtat attactgtgc gaaagatagg
300gttctagtcc cagcttcctc ttcgtacttt gactactggg gccagggaac cctggtcacc
360gtctcgaga
369850327DNAHomo sapiens 850tcttctgagc tgactcagga ccctgctgtg tctgtggcct
tgggacagac agtcaggatc 60acatgccaag gagacagcct cagaagctat tatgcaagct
ggtaccagca gaagccagga 120caggcccctg tacttgtcat ctatggtaaa aacaaccggc
cctcagggat cccagaccga 180ttctctggct ccagctcagg aaacacagct tccttgacca
tcactggggc tcaggcggaa 240gatgaggctg actattactg taactcccgg gacagcagtg
gtaaccatgt ggtattcggc 300ggagggacca agctgaccgt cctaggt
327851360DNAHomo sapiens 851caggtgcagc tgcaggagtc
gggcccagga ctggtgaagc cttcggagac cctgtccctc 60acctgcactg tctctggtgg
ctccatcagc agtagtagtt actactgggg ctggatccgc 120cagcccccag ggaaggggct
ggagtggatt gggagtatct attatagtgg gagcacctac 180tacaacccgt ccctcaagag
tcgagtcacc atatccgtag acacgtccaa gaaccagttc 240tccctgaagc tgagctctgt
gaccgccgca gacacggctg tgtattactg tgcgagacgt 300cccttaacct ttaatgcttt
tgatatctgg ggccaaggga caatggtcac cgtctcgaga 360852327DNAHomo sapiens
852tcctatgtgc tgactcagcc accctcagtg tcagtggccc caggaaagac ggccaggatt
60acctgtgggg gaaacaacat tggaagtaaa agtgtgcact ggtaccagca gaagccaggc
120caggcccctg tgctggtcat ctattatgat agcgaccggc cctcagggat ccctgagcga
180ttctctggct ccaactctgg gaacacggcc accctgacca tcagcagggt cgaagccggg
240gatgaggccg actattactg tcaggtgtgg gatagtagta gtgatcatgt ggtattcggc
300ggagggacca agctgaccgt cctaggt
327853360DNAHomo sapiens 853caggtgcagc tgcaggagtc gggcccagga ctggtgaagc
cttcggagac cctgtccctc 60acctgcactg tctctggtgg ctccatcagc agtagtagtt
actactgggg ctggatccgc 120cagcccccag ggaaggggct ggagtggatt gggagtatct
attatagtgg gagcacctac 180tacaacccgt ccctcaagag tcgagtcacc atatccgtag
acacgtccaa gaaccagttc 240tccctgaagc tgagctctgt gaccgccgca gacacggctg
tgtattactg tgcgaggatt 300cccatgtata gcagctcggt tgactactgg ggccagggaa
ccctggtcac cgtctcgaga 360854327DNAHomo sapiens 854tcctatgtgc
tgactcagcc accctcagtg tcagtggccc caggaaagac ggccaggatt 60acctgtgggg
gaaacaacat tggaagtaaa agtgtgcact ggtaccagca gaagccaggc 120caggcccctg
tgctggtcat ctattatgat agcgaccggc cctcagggat ccctgagcga 180ttctctggct
ccaactctgg gaacacggcc accctgacca tcagcagggt cgaagccggg 240gatgaggccg
actattactg tcaggtgtgg gatagtagta gtgatcatgt ggtattcggc 300ggagggacca
agctgaccgt cctaggt
327855360DNAHomo sapiens 855caggtgcagc tgcaggagtc gggcccagga ctggtgaagc
cttcggagac cctgtccctc 60acctgcactg tctctggtgg ctccatcagc agtagtagtt
actactgggg ctggatccgc 120cagcccccag ggaaggggct ggagtggatt gggagtatct
attatagtgg gagcacctac 180tacaacccgt ccctcaagag tcgagtcacc atatccgtag
acacgtccaa gaaccagttc 240tccctgaagc tgagctctgt gaccgccgca gacacggctg
tgtattactg tgcgagacgt 300cccttaacct ttaatgcttt tgatatctgg ggccaaggga
ccacggtcac cgtctcgaga 360856327DNAHomo sapiens 856tcctatgtgc
tgactcagcc accctcagtg tcagtggccc caggaaagac ggccaggatt 60acctgtgggg
gaaacaacat tggaagtaaa agtgtgcact ggtaccagca gaagccaggc 120caggcccctg
tgctggtcat ctattatgat agcgaccggc cctcagggat ccctgagcga 180ttctctggct
ccaactctgg gaacacggcc accctgacca tcagcagggt cgaagccggg 240gatgaggccg
actattactg tcaggtgtgg gatagtagta gtgatcatgt ggtattcggc 300ggagggacca
agctgaccgt cctaggt
327857366DNAHomo sapiens 857gaggtgcagc tggtggagtc tgggggaggc ttggtacagc
ctggggggtc cctgagactc 60tcctgtgcag cctctggatt cacctttagc agctatgcca
tgagctgggt ccgccaggct 120ccagggaagg ggctggagtg ggtctcagct attagtggta
gtggtggtag cacatactac 180gcagactccg tgaagggccg gttcaccatc tccagagaca
attccaagaa cacgctgtat 240ctgcaaatga acagcctgag agccgaggac acggccgtat
attactgtgc gaaagatggg 300ggatggttcg gggagttaga ttacttccag cactggggcc
agggcaccct ggtcaccgtc 360tcgaga
366858333DNAHomo sapiens 858cagtctgtgt tgacgcagcc
gccctcagtg tctgcggccc caggacagaa ggtcaccgtc 60tcctgcactg gaagcaactc
caacattgag aagaatgatg tttcctggta ccagcaggga 120ccaggagcag cccccaaact
cctcatttct gacactgata ggcgaccctc agggattcct 180gaccgattct ctggctccaa
gtctggcacg tcagccaccc tggccatcgc tgggctccag 240gctgaggatg aggctgatta
ttactgccag tcccatgaca ccactctgag tggtccgatc 300ttcggcgggg ggacccagct
gaccgtccta ggt 333859372DNAHomo sapiens
859caggtgcagc tgcaggagtc ggggggaggc ttggtcaagc ctggagggtc cctgagactc
60tcctgtgcag cctctggatt cagtttcagt gactactaca tgagctgggt ccgccaggct
120ccagggaagg ggctggagtg gatttcatat attactagta gtagtagtga cacagactac
180gcagactctg tgaagggccg attcaccatc tccagagaca acgccaagaa ctcactatat
240ctgcaaatga acagcctgag agccgacgac acggccgtgt attactgtgc gagagtgggc
300tattattatg attactacta ctactactac atggacgtct ggggcaaagg gaccacggtc
360accgtctcga ga
372860324DNAHomo sapiens 860tcttctgagc tgactcagga ccctgctgtg tctgtggcct
tgggacagac agtcaggatc 60acatgccaag gagacagcct cagaggctat tatgcaagct
ggtaccagca gaagccagga 120caggcccctg tccttgtcgt ctatgatgaa aacaaccggc
cctcagggat cccagaccga 180ttctctggct ccagctcagg aaacacagct tccttgacca
tcactggggc tcaggcggaa 240gatgaggctg actattactg taactcccgg gacattaacc
ttgattgggt gttcggcgga 300gggaccaagt tgaccgtcct aggt
324861360DNAHomo sapiens 861caggtgcagc tgcaggattc
gggcccagga ctggtgaagc cttcggatac cctgtccctc 60acctgcactg tctctggtgg
ctccatcagc agtattagtt actactgggg ctggatccgc 120cagcccccag ggaaggggct
ggagtgtatt gggagtatct attatagtgg gagcacctac 180tacaacccgt ccctcaagag
tcgagtcacc atatccgtag acacgtccaa gaaccagttc 240tccctgaagc tgagctctgt
gaccgccgca gacacggctg tgtattactg tgcgagacgg 300gtcatagtgt gggggagtga
tgactactgg ggccagggaa ccctggtcac cgtctcgaga 360862327DNAHomo sapiens
862tcctatgtgc tgactcagcc accctcagtg tcagtggccc caggaaagac ggccaggatt
60acctgtgggg gaaacaacat tggaagtaaa agtgtgcact ggtaccagca gaagccaggc
120caggcccctg tgctggtcat ctattatgat agcgaccggc cctcagggat ccctgagcga
180ttctctggct ccaactctgg gaacacggcc accctgacca tcagcagggt cgaagccggg
240gatgaggccg actattactg tcaggtgtgg gatagtagta gtgatcatgt ggtattcggc
300ggagggacca agctgaccgt cctaggt
327863360DNAHomo sapiens 863caggtgcagc tgcaggagtc gggcccagga ctggtgaagc
cttcggagac cctgtccctc 60acctgcactg tctctggtgg ctccatcagc agtagtagtt
actactgggg ctggatccgc 120cagcccccag ggaaggggct ggagtggatt gggagtatct
attatagtgg gagcacctac 180tacaacccgt ccctcaagag tcgagtcacc atatccgtag
acacgtccaa gaaccagttc 240tccctgaagc tgagctctgt gaccgccgca gacacggctg
tgtattactg tgcgatgcgg 300gcctatggtt cagggagtta tgactactgg ggccagggaa
ccctggtcac cgtctcgaga 360864336DNAHomo sapiens 864gatgttgtga
tgactcagtc tccactctcc ctgcccgtca cccctggaga gccggcctcc 60atctcctgca
ggtctagtca gagcctcctc catagtaatg gatacaacta tttggattgg 120tacctgcaga
agccagggca gtctccgcag ctcctgatct atttgggttc taatcgggcc 180tccggggtcc
ctgacaggtt cagtggcagt ggatcaggca cagattttac actgaaaatc 240agcagagtgg
aggctgagga tgttggggtt tattactgca tgcaagctct acaaacccca 300ttcactttcg
gccctgggac caaagtggat atcaaa
336865363DNAHomo sapiens 865gaggtgcagc tggtggagtc tgggggaggc ttggtacagc
ctggggggtc cctgagactc 60tcctgtgcag cctctggatt cacctttagc agctatgcca
tgagctgggt ccgccaggct 120ccagggaagg ggctggagtg ggtctcagct attagtggta
gtggtggtag cacatattac 180gcagactccg tgaagggccg gttcaccatc tccagagaca
attccaagaa cacgctgtat 240ctgcaaatga acagcctgag agccgaggac acggccgcat
attactgtgc gaaaggtctt 300aggtatagca gtgcctggac gtttgactac tggggccagg
gaaccctggt caccgtctcg 360aga
363866336DNAHomo sapiens 866cagtctgccc tgactcagcc
tgcctccgtg tctgggtctc ctggacagtc gatcaccatc 60tcctgcactg gaaccagcag
tgacgttggt ggttataact atgtctcctg gtaccaacaa 120cacccaggca aagcccccaa
actcatgatt tatgatgtca gtaatcggcc ctcaggggtt 180tctaatcgct tctctggctc
caagtctggc aacacggcct ccctgaccat ctctggactc 240caggctgagg acgaggctga
ttattactgc agctcatata caagcagtag cactcctgtg 300gtattcggcg gagggaccaa
gctgaccgtc ctaggt 336867357DNAHomo sapiens
867caggtgcagc tggtgcaatc tgggtctgag ttgaagaagc ctggggcctc agtgaaggtt
60tcctgcaagg cttctggata catcttcact agatatggca taaattgggt gcgacaggcc
120cctggacaag ggcttgagtg gatgggatgg atcaacacca acactgggaa cccaacgtat
180gcccagggct tcacaggccg ggttgtcttc tccttggaca cctctgtcag cacggcatat
240ctgcagatca gcagcctaaa ggctgaggac actgccatgt attactgtgc gatcagcagt
300ggctttgggt actactttga ctactggggc cagggaaccc tggtcaccgt ctcgaga
357868321DNAHomo sapiens 868gacatccaga tgacccagtc tccatcgtcc gtgtctgcat
ctgtgggaga cagagtcacc 60atcacttgtc gggcgagtca aagtattagt aggtggttag
cctggtatca gcagaaacca 120gggaaagccc ctaagttcct gatctatgct ggatccagtt
tgcaaagtgg ggtcccatcg 180aggttcagcg gcagtgggtc tgggacagat ttcactctca
ccatcagcag cctgcagcct 240gaagactttg caacttatta ttgtcaacag actaacagtt
tccctctcac cttcggcgga 300gggaccaagg tggagatcaa a
321869360DNAHomo sapiens 869caggtgcagc tggtgcagtc
tggggctgag gtgaagaagc ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta
cacctttacc agctacggta tcagctgggt gcgacaggcc 120cctggacaag ggcttgagtg
gatgggatgg atcagcgctt acaatggtaa cacaaactat 180gcacagaagc tccagggcag
agtcaccatg accacagaca catccacgag cacagcctac 240atggagctga ggagcctgag
atctgacgac acggccgtgt attactgtgc gagagctttt 300agggactggg gatctcttag
ggactactgg ggccagggca ccctggtcac cgtctcgaga 360870333DNAHomo sapiens
870cagtctgtgt tgacgcagcc gccctcagtg tctgcggccc caggacagaa ggtcaccatc
60tcctgctctg gaagcagctc caacattggg aataattatg tatcctggta ccagcagctc
120ccaggaacag cccccaaact cctcatttat gacaataata agcgaccctc agggattcct
180gaccgattct ctggctccaa gtctggcacg tcagccaccc tgggcatcac cggactccag
240actggggacg aggccgatta ttactgcgga acatgggata gcagcctgag tgccgtggta
300ttcggcggag ggaccaagct gaccgtccta ggt
333871363DNAHomo sapiens 871gaggtgcagc tggtggagtc tggagcagag gtgaaaaagc
ccggggagtc tctgaagatc 60tcctgtaagg gttctggata cagctttacc agctactgga
tcggctgggt gcgccagatg 120cccgggaaag gcctggagtg gatggggatc atctatcctg
gtgactctga taccagatac 180agcccgtcct tccaaggcca ggtcaccatc tcagccgaca
agtccatcag caccgcctac 240ctgcagtgga gcagcctgaa ggcctcggac accgccatgt
attactgtgc gagactggca 300gtgggagcct accagtacta ctttgactac tggggccagg
gaaccctggt caccgtctcg 360aga
363872333DNAHomo sapiens 872cagactgtgg tgacccagga
gccatcgttc tcagtgtccc ctggagggac agtcacactc 60acttgtggct tgagctctgg
ctcagtctct actagttact accccagctg gtaccagcag 120accccaggcc aggctccacg
cacgctcatc tacagcacaa acactcgctc ttctggggtc 180cctgatcgct tctctggctc
catccttggg aacaaagctg ccctcaccat cacgggggcc 240caggcagatg atgaatctga
ttattactgt gtgctgtata tgggtagtgg catttcggtg 300ttcggcggag ggaccaagct
gaccgtccta ggt 333873357DNAHomo sapiens
873gaggtgcagc tggtggagtc tgggggaaac ttggtccagc cgggggggtc cctgagactc
60tcctgtgcag cctctggatt caccttcagt aggtattgga tgacctgggt ccgccaggct
120ccagggaagg ggctggagtg gatttcatcc gttagtagta gcggcagtac catatactac
180gcagactctg tgaagggccg attcaccatc tccagagaca acgccaagaa ctcactgtat
240ctgcaaatga acagcctgag agtcgaggac acggctgttt attactgtgc gcgagactat
300tggcctggct ggtacttcga tctctggggc cgtggaaccc tggtcaccgt ctcgaga
357874348DNAHomo sapiens 874caggctgtgc tcactcagcc gtcttccctc tctgcatctc
ctggagcatc agccagtctc 60acctgcacct tccgcagtga catcagtgtt ggttcctata
ggatatactg gtaccagcag 120aagccaggga gtcctcccca gtttctcctg aaatatacgt
cagactcaga taagcagcag 180ggctctggag tccccagccg cttctctgga tccaaagatg
tttcggccaa tgctggcatt 240ttactcatct ctgggctcca gtctgaggat gaggctgact
attactgtat gacttggcac 300aacaccgctt cggtattcgg cggagggacc aagctggccg
tcctaggt 348875357DNAHomo sapiens 875gaggtacagc
tggtggagtc tgggggaggt ttgataaggc cgggggggtc cctgagactc 60tcctgtacag
cctctggatt caccttcagt agttatacta tgaattgggt ccgccaggct 120ccagggaagg
ggctggagtg gctttcatat ataagtggtg gcagtggtac caaattctac 180gcagactctg
tgaagggccg gttcaccgtc tccagagaca atgccaagaa ttcattgtat 240ctggaaatga
acagcctgag acccgaggac acggctgtct attactgtgc gctagtgtca 300tatagttcgc
cgggctttga ctactggggc cagggcaccc tggtcaccgt ctcgagc
357876321DNAHomo sapiens 876tcctatgagc tgactcagcc accctcagtg tccgtgtccc
caggacagac agccagcatc 60acctgctctg gagataaatt gggcgataaa tatgtttact
ggtatcaaca gaagccaggc 120cagtccccta tattgctcat ctttcaagat agcgagcggc
cctcagggat ccctgagcga 180ttctctggct ccaactctgg gaacacagcc actctgacca
tcagcgggac ccaggctatg 240gatgaggctg actattactg tcaggcgtgg gacagcagcg
ctgcggtgtt cggcggaggg 300accaagctga ccgtcctagg t
321877345DNAHomo sapiens 877caggtgcagc tggtgcagtc
tggagcagag gtgaaaaagc ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta
cacctttacc agctatggta tcagctgggt gcgacaggcc 120cctggacaag ggcttgagtg
gatgggatgg atcagcgctt acaatggtaa cacaaactat 180gcacagaagc tccagggcag
agtcaccatg accacagaca catccacgag cacagcctac 240atggagctga ggagcctgag
atctgacgac acggccgtgt attactgtgc gagagggggt 300gcttttgata tctggggcca
agggaccacg gtcaccgtct cgaga 345878336DNAHomo sapiens
878cagtctgtgt tgacgcagcc gccctcagtg tctggggccc cagggcagag ggtcaccatc
60tcctgcactg ggagcagctc caacatcggg gcaggttatc atgtatactg gtaccagcag
120cttccaggaa aagcccccaa actcctcatc tacgttaaca gcaatcggcc ctcaggggtc
180cctgaccgat tctctggctc caagtctggc acctcagcct ccctggccat cactgggctc
240caggctgacg atgaggctga ttactactgc cagtcctatg acagcagcct gagtggtagg
300gttttcggcg gagggaccaa gctgaccgtc ttaggt
336879357DNAHomo sapiens 879gaggtgcagc tggtggagtc tgggggcggc ttggtccagc
cgggggggtc cctgagactc 60tcctgttcag cctctggatt cacctttagt aactattgga
tgacctgggt ccgtcaggct 120ccagggaagg ggctggagtg gatttcatcc attagtagta
gcggcagtac catatactac 180gcagactctg tgaagggccg attcaccatc tccagagaca
acgccaagaa ctcactgtat 240ctgcaaatga acagcctgag agtcgaggac acggctgttt
attactgtgc gcgagactat 300tggcctggct ggtacttcga tctctggggc cgtggcaccc
tggtcaccgt ctcgaga 357880348DNAHomo sapiens 880caggctgtgc
tcactcagcc gtcttccctc tctgcatctc ctggagcatc agccagtctc 60acctgcacct
tccgcagtga catcagtgtt ggttcctata ggatatactg gtaccagcag 120aagccaggga
gtcctcccca gtttctcctg aaatatacgt cagactcaga taagcagcag 180ggctctggag
tccccagccg cttctctgga tccaaagatg tttcggccaa tgctggcatt 240ttactcatct
ctgggctcca gtctgaggat gaggctgact attactgtat gacttggcac 300aacaccgctt
cggtattcgg cggagggacc aagctggccg tcctaggt
348881381DNAHomo sapiens 881caggtgcagc tggtgcagtc tgggtctgag gtgaagaagc
ctggggcctc agtgaaggtc 60tcctgcaaga cttctggata catcttcacc gactactata
tgcactgggt gcgacaggcc 120cctggaaaag ggcttgagtg gatgggatgg attaacccta
acagtggtgg cacatactat 180gcacagaagt ttcacggcag ggtcaccatg accagtgaca
cgtccatcag cacagcctac 240atggagctga gcagtctgag atctgacgac acggccatat
attactgtgc gagagaggat 300tacgatattt tgactggtta ttatcccgcg tccggccacg
gggactactg gggccaggga 360accctggtca ccgtctcgag a
381882330DNAHomo sapiens 882caggctgtgg tgactcagga
gccctcactg actgtgtccc caggagggac agtcactctc 60acctgtgcat ccagcactgg
agcagtcacc agtggtttcc ttgcaaactg gttccagcag 120aaacctggac aaacacccag
gtcactgatt tataaaacaa gcaacaaaca tccctggacc 180cctgcccggt tctcaggctc
cctccttggg ggcaaagctg ccctgacact gtcaggtgtg 240cagcctgagg acgaggctga
ctattactgc ctgctcttat ctggtggtgc atgggtgttt 300ggcggaggga ccaagctgag
tgtcctaggt 330883366DNAHomo sapiens
883caggtgcagc tggtgcagtc tggagcagag gtgaaaaagc ccggagagtc tctgaagatc
60tcgtgtgagg gttctggata cacctttacc agctactgga tcggctgggt gcgccagatg
120cccgggaaag acctggagtg gatggggatc atctatcctg gtgactctga taccagatac
180agcccgtcct tccaaggcca ggtcaccatc tcagtcgaca agtctatcag caccgcctac
240ctgcagtgga gcagcctgaa ggcctcggac accgccatgt attactgtgc gagacacgac
300gtagttgatg gctacaatac cggtatggac gtctggggcc aagggaccac ggtcaccgtc
360tcgaga
366884333DNAHomo sapiens 884cagactgtgg tgacccagga gccatcgttc acagtgtccc
ctggagggac agtcacactc 60acttgtggct tgagctctgg ctcagtctct actagttact
accccagctg gtaccagcag 120accccaggcc aggctccacg cacgctcatc tccagcacaa
acactcgctc ttctggggtc 180cctgatcgct tctctggctc catccttggg aacagagctg
ccctcaccat cacgggggcc 240caggcagatg atgagtctga ttattactgt gtgctgtata
tgggtagtgg catttgggtg 300ttcggcggag ggaccaagct gaccgtccta ggt
333885384DNAHomo sapiens 885caggtgcagc tggtgcagtc
tggagcagag gtgaaaaagc ccggggagtc tctgaagatc 60tcctgtaagg gttctggata
cagctttacc agctactgga tcggctgggt gcgccagatg 120cccgggaaag gcctggagtg
gatggggatc atctatcctg gtgactctga tgccagatac 180agcccgtcct tccaaggcca
ggtcaccatc tcagccgaca agtccatcag caccgcctac 240ctgcagtgga gcagcctgaa
ggcctcggac accgccatgt attactgtgc gagactcacg 300ggcagttctt actatgatag
tagtggttat tcctcctacg gtatggacgt ctggggccaa 360gggaccacgg tcaccgtctc
gaga 384886333DNAHomo sapiens
886cagactgtgg tgacccagga gccatcgttc tcagtgtccc ctggagggac agtcacactc
60acttgtggct tgagctctgg ctcagtctct actagttact accccagctg gttccagcag
120accccaggcc aggctccacg cacgctcatc tacagcacaa acactcgctc ttctggggtc
180cctgatcgct tctctggctc catccttggg aacaaagctg ccctcaccat cacgggggcc
240caggcagatg atgaatctga ttattactgt gtgctgtata tgggtagtgg catttctgtg
300ttcggaggag gcacccagct gaccgtcctc ggt
333887363DNAHomo sapiens 887caggtgcagc tggtgcagtc tggagcagag gtgaaaaagc
ccggggagtc tctgaagatc 60tcctgtaagg gttctggata cagctttacc agctactgga
tcggctgggt gcgccagatg 120cccgggaaag gcctggagtg gatggggatc atctatcctg
gtgactctga taccagatac 180agcccgtcct tccaaggcca ggtcaccatc tcagccgaca
agtccatcag caccgcctac 240ctgcagtgga gcagcctgaa ggcctcggac accgccatgt
attactgtgc gagacataac 300agtaactact actactacta catggacgtc tggggcaaag
gaaccctggt caccgtctcg 360aga
363888333DNAHomo sapiens 888cagactgtgg tgacccagga
gccatcgttc tcagtgtccc ctggagggac agtcacactc 60acttgtggct tgagctctgg
ctcagtctct actagttact accccagctg gtaccagcag 120accccaggcc aggctccacg
cacgctcatc tacagcacaa acactcgctc ttctggggtc 180cctgatcgct tctctggctc
catccttggg aacaaagctg ccctcaccat cacgggggcc 240caggcagatg atgaatctga
ttattactgt gtgctgtata tgggtagtgg catttcggtg 300ttcggcggag ggaccaagct
gaccgtccta ggt 333889381DNAHomo sapiens
889caggtgcagc tggtgcagtc tgggtctgag gtgaagaagc ctggggcctc agtgaaggtc
60tcctgcaaga cttctggata catcttcacc gactactata tgcactgggt gcgacaggcc
120cctggaaaag ggcttgagtg gatgggatgg attaacccta acagtggtgg cacatactat
180gcacagaagt ttcacggcag ggtcaccatg accagtgaca cgtccatcag cacagcctac
240atggagctga gcagtctgag atctgacgac acggccatat attactgtgc gagagaggat
300tacgatattt tgactggttt ttatcccgcg tccggccacg gggactactg gggccaggga
360accctggtca ccgtctcgag a
381890321DNAHomo sapiens 890gaaattgtgt tgacgcagtc tccaggcacc ctgtctttgt
ctccagggga aagagccacc 60ctctcctgca gggccagtca gagtgttagc agcagctact
tagcctggta ccagcagaaa 120cctggccagg ctcccaggct cctcatctat ggtgcatcca
gcagggccac tggcatccca 180gacaggttca gtggcagtgg gtctgggaca gacttcactc
tcaccatcag cagactggag 240cctgaagatt ttgcagtgta ttactgtcag cagtatggta
gctcactcac tttcggcgga 300gggaccaagg tggagatcaa a
321891366DNAHomo sapiens 891caggtgcagc tggtgcagtc
tggagcagag gtgaaaaagc ccggggagtc tctgaagatc 60tcctgtaagg gttctggata
cagctttacc agctactgga tcggctgggt gcgccagatg 120cccgggaaag gcctggagtg
gatggggatc atctatcctg gtgactctga taccagatac 180agcccgtcct tccaaggcca
ggtcaccatc tcagccgaca agtccatcag caccgcctac 240ctgcagtgga gcaccctgaa
ggcctcggac accgccatgt attactgtgc gagacatggg 300atgactagtg gctacgtcgc
gcacaatgac tactggggcc agggaaccct ggtcaccgtc 360tcgaga
366892333DNAHomo sapiens
892cagactgtgg tgacccagga gccatcgttc tcagtgtccc ctggagggac agtcacactc
60acttgtggct tgagctctgg ctcagtctct actagttact accccagctg gtaccagcag
120accccaggcc aggctccacg cacgctcatc tacagcacaa acactcgctc ttctggggtc
180cctgatcgct tctctggctc catccttggg aacaaagctg ccctcaccat cacgggggcc
240caggcagatg atgaatctga ttattactgt gtgctgtata tgggtagtgg catttgggtg
300ttcggcggag ggaccaagct gaccgtccta ggt
333893351DNAHomo sapiens 893caggtgcagc tgcaggagtc gggcccagga ctggtgaagc
cttcggggac cctgtccctc 60acctgcgctg tctctggtgg ctccatcagc agtagtaact
ggtggagttg ggtccgccag 120cccccaggga aggggctgga gtggattggg gaaatctatc
atagtgggag caccaactac 180aacccgtccc tcaagagtcg agtcaccata tcagtagaca
agtccaagaa ccagttctcc 240ctgaagctga gctctgtgac cgccgcggac acggccgtgt
attactgtgc gagaggtggg 300agctactact ttgactactg gggccaggga accctggtca
ccgtctcgag a 351894336DNAHomo sapiens 894cagtctgccc
tgactcagcc tgcctccgtg tctgggtctc ctggacagtc gatcaccatc 60tcctgcactg
gaaccatcag tgacgttggt ggttatgact ttgtctcctg gtaccaacac 120caccccggca
aagcccccaa actcctgatt tatgatgtca ataatcggcc ctctggggtt 180tctcatcgct
tctctggctc caagtctggc aacacggcct ccctgaccat ctctgggctc 240caggctgagg
acgaggctac ttattactgc agttcatatt caaacagaca ttctctcatc 300gtcttcggat
ctgggaccca ggtcgtcggc ctaggt
336895363DNAHomo sapiens 895caggtgcagc tacagcagtg gggcgcagga ctgttgaagc
cttcggagac cctgtccctc 60acctgcgctg tctatggtgg gtccttcagt ggttactact
ggagctggat ccgccagccc 120ccagggaagg ggctggagtg gattggggaa atcaatcata
gtggaagcac caactacaac 180ccgtccctca agagtcgagt caccatatca gtagacacgt
ccaagaacca gttctccctg 240aagctgagct ctgtgaccgc cgcggacacg gctgtgtatt
actgtgcgag aggcgggggg 300gttgaggcgt tgaactacgg tatggacgtc tggggccaag
ggaccacggt caccgtctcg 360aga
363896336DNAHomo sapiens 896cagtctgccc tgactcagcc
tgcctccgtg tctgggtctc ctggacagac gatcaccatc 60tcctgcactg gaaccatcag
tgacgttggt ggttatgact ttgtctcctg gtaccaacac 120caccccggca aagcccccaa
actcctgatt tatgatgtca ataatcggcc ctctggggtt 180tctcatcgct tctctggctc
caagtctggc aacacggcct ccctgaccat ctctgggctc 240caggctgagg acgaggctac
ttattactgc agttcatatt caaacagaca ttctctcatc 300gtcttcggat ctgggaccca
ggtcgtcggc ctaggt 336897357DNAHomo sapiens
897caggtgcagc tacagcagtg gggcgcaggg ctgttgaagc cttcggaaac cctgtccctc
60acctgcactg tccatggtgg gtccttcgat gattactact ggacctggat ccgccagccc
120ccaggggggg ggctggaatg gattggggaa atgaattccg gtagaactta caactacaac
180ccgttcctgg agagtcgagc ctccatagat gttgacacgt tcaagaagca gttctccctg
240gcattgcgtt ctgtgaccgc cgcggacaca gctgtctatt actgtgcgcg gggcgcctat
300gtcaactact actacataga cgtctggggc gacgggacca cggtcaccgt ctcgaga
357898336DNAHomo sapiens 898cagtctgccc tgactcagcc tgcctccgtg tctgggtctc
ctggacagac gatcaccatc 60tcctgcactg gaaccatcag tgacgttggt ggttatgact
ttgtctcctg gtaccaacac 120caccccggca aagcccccaa actcctgatt tatgatgtca
ataatcggcc ctctggggtt 180tctcatcgct tctctggctc caagtctggc aacacggcct
ccctgaccat ctctgggctc 240caggctgagg acgaggctac ttattactgc agttcatatt
caaacagaca ttctctcatc 300gtcttcggat ctgggaccca ggtcgtcggc ctaggt
336899375DNAHomo sapiens 899caggtgcagc tgcaggagtc
gggcccagga ctggtgaagc cttcggagac cctgtccctc 60acctgcactg tctctggtgg
ctccatcagt agttactact ggagctggat ccggcagccc 120gccgggaagg gactggagtg
gattgggcgt atctatacca gtgggagcac caactacaac 180ccctccctca agagtcgagt
caccatgtca gtagacacgt ccaagaacca gttctccctg 240aagctgagct ctgtgaccgc
cgcggacacg gccgtgtatt actgtgcgag agagagggca 300tattgtagta gtaccagctg
ctatcgaaat gcttttgata tctggggcca agggaccacg 360gtcaccgtct cgaga
375900357DNAHomo sapiens
900cagtctgtgt tgacgcagcc gccctcagtg tctggggccc cagggcagag ggtcaacatc
60tcctgcgctg ggagcagctc caacatcggg gcgggttatg atgttcactg gtaccagcag
120attccaggaa cagcccccaa actcctcatg tatggtaata gtaatcggcc ctcaggggtc
180cctgaccgat tctctggctc caagtctggc gcctcagcct ccctggccat cactaggctc
240caggctgagg atgaggctga ttattactgc cagtcctatg acagcagcct gagtggttcg
300agggtcttcg gaactgggac caaggtcacc gtcctaggtc agtccaacgt cctaggt
357901366DNAHomo sapiens 901gaggtgcagc tggtggagtc tgggggaggc gtggtccagc
ctgggaggtc cctgagactc 60tcctgtgcag cctctggatt caccttcagt agctatggca
tgcactgggt ccgccaggct 120ccaggcaagg ggctggagtg ggtgacagtt atatcatttg
atggaagtaa taaatactat 180gcagactccg tgaagggccg attcaccatc tccagagaca
attccaagaa cacgctgtat 240ctgcaaatga acagcctgag agctgaggac acggctgtgt
attactgtgc gaaagcgtat 300accaacacct ggtggcctga tgcttttgat atctggggcc
aagggaccac ggtcaccgtc 360tcgaga
366902321DNAHomo sapiens 902gacatccaga tgacccagtc
tccatcctcc ctgtctgcat ctgtaggaga cagagtcacc 60atcacttgcc aggcgagtca
ggacattagc aactatttaa attggtatca gcagaaacca 120gggaaagccc ctaagctcct
gatctacgat gcatccaatt tggaaacagg ggtcccatca 180aggttcagtg gaagtggatc
tgggacagat tttactttca ccatcagcag cctgcagcct 240gaagatattg caacatatta
ctgtcaacag tatgataatc tccctcccac tttcggccct 300gggaccaaag tggatatcaa a
321903354DNAHomo sapiens
903caggtgcagc tgcaggagtc gggcccagga ctggtgaagc cttcggagac cctgtccctc
60acctgcactg tctctggtag ctccatcagt agttactact ggagctggat ccggcagccc
120ccagggaagg gactggagtg gattgggtat atctattaca gtgggagcac caactacaac
180ccctccctca agagtcgagt caccatatca gtagacacgt ccaagaacca gttctccctg
240aagctgagct ctgtgaccgc tgcggacacg gccgtgtatt actgtgcgag agatctgtac
300tggaacgacg cttttgatat ctggggccaa gggaccacgg tcaccgtctc gaga
354904333DNAHomo sapiens 904cagtctgccc tgactcagcc tctctcagcg tctgggaccc
ccgggcagag ggtcaccatc 60tcttgttctg gaagcagctc caacatcgga agtaatactg
taaactggta ccagcagctc 120ccaggaacgg cccccaaact cctcatctat agtaataatc
agcggccctc aggggtccct 180gaccgattct ctggctccaa gtctggcacc tcagcctccc
tggccatcag tgggctccag 240tctgaggatg aggctgatta ttactgcgca gcatgggatg
acagcctgaa tggtccggta 300ttcggcggag ggaccaagct gaccgtccta ggt
333905354DNAHomo sapiens 905caggtgcagc tgcaggagtc
gggcccagga ctggtgaagc cttcggagac cctgtccctc 60acctgcactg tctctggtag
ctccatcagt agttactact ggagctggat ccggcagccc 120ccagggaagg gactggagtg
gattgggtat atctattaca gtgggagcac caactacaac 180ccctccctca agagtcgagt
caccatatca gtagacacgt ccaagaacca gttctccctg 240aagctgagct ctgtgaccgc
tgcggacacg gccgtgtatt actgtgcgag agatctgtac 300tggaacgacg cttttgatat
ctggggccaa gggaccacgg tcaccgtctc gaga 354906333DNAHomo sapiens
906cagtctgtgt tgacgcagcc gccctcagcg tctgggaccc ccgggcagag tgtcaccatc
60tcttgttctg gaagcagctc caacatcgga agtaatactg taaactggta ccagcagctc
120ccaggaacgg cccccaaact cctcatctat agtaataatc agcggccctc aggggtccct
180gaccgattct ctggctccaa gtctggcacc tcagcctccc tggccatcag tgggctccag
240tctgaggatg aggctgatta ttactgtgca gcatgggatg acagcctgaa tggtccggtg
300ttcggcggag ggaccaagct gaccgtccta ggt
333907378DNAHomo sapiens 907caggtacagc tgcagcagtc aggtccagga ctggtgaagc
cctcgcagac cctctcactc 60acctgtgcca tctccgggga cagtgtctct agcaacagtg
ctgcttggaa ctggatcagg 120cagtccccat cgagaggcct tgagtggctg ggaaggacat
actacaggtc caagtggtat 180aatgattatg cagtatctgt gaaaagtcga ataaccatca
acccagacac atccaagaac 240cagttctccc tgcagctgaa ctctgtgact cccgaggaca
cggctgtgta ttactgtgca 300agagaatcat ggctttgggg gattgggggg gatgcttttg
atatctgggg ccaagggacc 360acggtcaccg tctcgaga
378908333DNAHomo sapiens 908cagtctgtgt tgacgcagcc
gccctcggtg tctggggccc cccggcagac ggtcaccatc 60tcctgctctg ggagcagctc
caacatcgga caaaattctg ttacctggta ccagcgcctc 120ccgggtgagg ctcccaaact
cctcatctac tatgatgatc tcttgcactc aggagtctct 180gaccgattct ctggctccaa
gtctggcacc tcagcctcac tggccatcag tggactccag 240tctgaggatg aggctgagta
ctactgtgcg tcatgggatg acagcctgaa aggtccggta 300ttcggcggag ggaccaaact
gaccgtccta ggt 333909336DNAHomo sapiens
909gaggtgcagc tggtggagtc tgggggaggc ttggtccagc ctggggggtc cctgagactc
60tcctgtgcag cctctggatt caccgtcagt agcaactaca tgagctgggt ccgccaggct
120ccagggaagg ggctggagtg ggtctcagtt atttatagcg gtggtagcac atactacgca
180gactccgtga agggcagatt caccatctcc agagacaatt ccaagaacac gctgtatctt
240caaatgaaca gcctgagagc cgaggacacg gctgtgtatt actgtgcgag ggagtttgac
300tactggggcc agggaaccct ggtcaccgtc tcgaga
336910315DNAHomo sapiens 910gacatccaga tgacccagtc tccatcctcc ctgtctgcat
ctgtaggaga cagagtcacc 60atcacttgcc aggcgagtca ggacattagc aactatttaa
attggtatca gcagaaacca 120gggaaagccc ctaagctcct gatctacgat gcatccaatt
tggaaacagg ggtcccatca 180aggttcagtg gaagtggatc tgggacagat tttactttca
ccatcagcag cctgcagcct 240gaagatatcg caacatatta ctgtcaacag tatgataatc
ccgctttcgg cggagggacc 300aaggtggaga tcaaa
315911363DNAHomo sapiens 911caggtgcagc tggtgcagtc
tggagctgag gtgaagaagc ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta
cacctttacc agctatggta tcagctgggt gcgacaggcc 120cctggacaag ggcttgagtg
gatgggatgg atcagcgctt acaatggtaa cacaaactat 180gcacagaagc tccagggcag
agtcaccatg accacagaca catccacgag cacagcctac 240atggagctga ggagcctgag
atctgacgac acggccgtgt attactgtgc gagagatcgt 300tactatggtt cggggttcgg
tatggacgtc tggggccaag ggacaatggt caccgtctcg 360aga
363912327DNAHomo sapiens
912tcttctgagc tgactcagga ccctgctatg tctgtggcct tgggacagac agtcaaaatc
60acttgccaag gagacagcct cacaaactat tatccaagtt ggtatcagca gaagccagga
120caggcccctg tccttgtcat gtatggaaaa gacagccggc cctcagggat ctcagaccga
180ttctctggct ccagctcagg aatctcagct tccttgacca tcactggggc tcaggcggaa
240gatgaggctg actactactg taactcccga gacggcagtg ctcaccgtct ggttttcggc
300ggagggacca agttgaccgt cctgggt
327913372DNAHomo sapiens 913caggtgcagc tggtgcagtc tggagctgag gtgaagaagc
ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta cacctttacc aactatggtc
tcacctgggt gcgacaggcc 120cctggacaag ggcttgagtg gatgggatgg atcagcactt
acaatagtaa cacaaactat 180gcagagaagc tccagggcag agtcaccatg accacagaca
catccacgag cacagcctac 240atggagttga ggagcctgac atctgacgac acggccgtgt
attactgtgc gagaggcccc 300acatattcct ttgatagtag tggttatttt tttgactact
ggggccaggg aaccctggtc 360accgtctcga ga
372914327DNAHomo sapiens 914tcttctgagc tgactcagga
ccctgctgtg tctgtggcct tgggacagac agtcaggatc 60acatgccaag gagacagcct
cagaagctat tatgcaagct ggtaccagca gaagccagga 120caggcccctg tacttgtcat
ctatggtaaa aacaaccggc cctcagggat cccagaccga 180ttctctggct ccagctcagg
aaacacagct tccttgacca tcactggggc tcaggcggaa 240gatgaggctg actattactg
taactcccgg gacagcagtg gtaaccattg ggtgttcggc 300ggagggacca agctgaccgt
cctaggt 327915360DNAHomo sapiens
915caggtgcagc tggtgcagtc tggagctgag gtgaagaagc ctggggcctc agtgaaggtc
60tcctgcaagg cttctggtta cacctttacc agctatggta tcagctgggt gcgacaggcc
120cctggacaag ggcttgagtg gatgggatgg atcagcgctt acaatggtaa cacaaactat
180gcacagaagc tccagggcag agtcaccatg accacagaca catccacgag cacagcctac
240atggagctga ggagcctgag atctgacgac acggccgtgt attactgtgc gagagatacg
300tatagcagtg gctggtactt tgactactgg ggccagggca ccctggtcac cgtctcgaga
360916330DNAHomo sapiens 916tcttctgagc tgactcagga ccctgctgtg tctgtggcct
tgggacagac agtcaggatc 60acatgccaag gagacagcct cagaaactat tatgcaagct
ggtaccagca gaagccagga 120caggcccctg tacttgtcat ctatggtaaa aacaaccggc
cctcagggat cccagaccga 180ttctctggct ccagctcagg aaacacagct tccttgacca
tcactggggc tcaggcggaa 240gatgaggctg actattactg taactcccga gacagcagtg
gtaaccatct ttatgtcttc 300ggaactggga ccaaggtcac cgtcctaggt
330917360DNAHomo sapiens 917caggtgcagc tggtgcagtc
tggagctgag gtgaagaagc ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta
cacctttacc agctatggta tcagctgggt gcgacaggcc 120cctggacaag ggcttgagtg
gatgggatgg atcagcgctt acaatggtaa cacaaactat 180gcacagaagc tccagggcag
agtcaccatg accacagaca catccacgag cacagcctac 240atggagctga gcagcctgag
atctgaggac acggccgtgt attactgtgc aacccttaac 300attagtggga gctactactt
tgactactgg ggccagggaa ccctggtcac cgtctcgaga 360918344DNAHomo sapiens
918tcttctgagc tgactcagga ccctgctgtg tctgtggcct tgggacagac agtcaggatc
60acatgccaag gagacagcct cagaagctat tatgcaacct ggtaccagca gaagccagga
120caggcccctg tacttgtcat ctctggtaaa aacaaccggc cctcagggat cccagaccga
180ttctctggct ccagctcagg aaacacagct tccttgacca tcactggggc tcaggcggaa
240gatgaggctg actattactg taactcccgg gacagcagtg gttacccctc ttgggtgttc
300ggcggaggga ccaagctgac cggaccaagc tgaccgtcct aggt
344919354DNAHomo sapiens 919caggtgcagc tggtgcagtc tgaagcagag gtgaaaaagc
ccggggagtc tctgaagatc 60tcctgtaagg gttctggata cagctttacc agctactgga
tcggctgggt gcgccagatg 120cccgggaaag gcctggagtg gatggggatc atctatcctg
gtgactctga taccagatac 180agcccgtcct tccaaggcca ggtcaccatc tcagccgaca
agtccatcag caccgcctac 240ctgcagtgga gcagcctgaa ggcctcggac accgccatgt
attactgtgc gagaacgatc 300gggcttggtg cttttgatat ctggggccaa gggaccacgg
tcaccgtctc gaga 354920330DNAHomo sapiens 920tcttctgagc
tgactcagga ccctgctgtg tctgtggcct tgggacagac agtcaggatc 60acatgccaag
gagacagcct cagaagctat tatgcaagct ggtaccagca gaagccagga 120caggcccctg
tacttgtcat ctatggtaaa aacaaccggc cctcagggat cccagaccga 180ttctctggct
ccagctcagg aaacacagct tccttgacca tcactggggc tcaggcggaa 240gatgaggctg
actattactg taactcccgg gacagcagtg gtaaccatca ttatgtcttc 300ggaactggga
ccaaggtcac cgtcctaggt
330921357DNAHomo sapiens 921caggtgcagc tgcaggagtc gggcccagga ctggtgaagc
cttcggagac cctgtccctc 60acctgcactg tctctggtgg ctccatcagc agtagtagtt
actactgggg ctggatccgc 120cagcccccag ggaaggggct ggagtggatt gggagtatct
attatagtgg gagcacctac 180tacaacccgt ccctcaagag tcgagtcacc atatccgtag
acacgtccaa gaaccagttc 240tccctgaagc tgagctctgt gaccgccgca gacacggctg
tgtattactg tgcgagacac 300tggggaaact atgcttttga tatctggggc caagggacca
cggtcaccgt ctcgaga 357922333DNAHomo sapiens 922cagtctgtgt
tgacgcagcc gccctcagcg tctgggaccc ccgggcagag ggtcaccatc 60tcttgttctg
gaagcagctc caacatcgga agtaattatg tatactggta ccaacagctc 120ccaggaacgg
cccccaaact cctcatctat agtaataatc agcggccctc aggggtccct 180gaccgattct
ctggctccaa gtctggcacc tcagcctccc tggccatcag tgggctccgg 240tccgaggatg
aggctgatta ttactgtgca gcatgggatg acagcctgag tggtcaagtg 300ttcggaggag
gcacccagct gaccgtcctc ggt
333923360DNAHomo sapiens 923gaggtgcagc tggtggagtc tggagctgag gtgaagaagc
ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta cacctttacc agctatggta
tcagctgggt gcgacaggcc 120cctggacaag ggcttgagtg gatgggatgg atcagcgctt
acaatggtaa cacaaactat 180gcacagaagc tccagggcag agtcaccatg accacagaca
catccacgag cacagcctac 240atggagctga ggagcctgag atctgacgac acggccgtgt
attactgtgc gagagagaag 300tatagcagtg gctggtactt tgactactgg ggccagggca
ccctggtcac cgtctcgaga 360924330DNAHomo sapiens 924tcttctgagc
tgactcagga ccctgctgtg tctgtggcct tgggacagac agtcaggatc 60acatgccaag
gagacagcct cagaagctat tatgcaagct ggtaccagca gaagccagga 120caggcccctg
tacttgtcat ctatggtaaa aacaaccggc cctcagggat cccagaccga 180ttctctggct
ccagctcagg aaacacagct tccttgacca tcactggggc tcaggcggaa 240gatgaggctg
actattactg taactcccgg gacagcagtg gtaaccatca ttatgtcttc 300ggaactggga
ccaaggtcac cgtcctaggt
330925345DNAHomo sapiens 925gaggtgcagc tggtggagtc tgggggaggc ttggtacagc
ctggggggtc cctgagactc 60tcctgtgcag cctctggatt cacctttagc agctatgcca
tgagctgggt ccgccaggct 120ccagggaagg ggctggagtg ggtctcagct attagtggta
gtggtggtag cacatactac 180gcagactccg tgaagggccg gttcaccatc tccagagaca
attccaagaa cacgctgtat 240ctgcaaataa acagcctgag agccgaggac acggccgtat
attactgtgc gaaaggttct 300atagcagcgg actggggcca gggaaccctg gtcaccgtct
cgaga 345926324DNAHomo sapiens 926tcttctgagc
tgactcagga ccctgctgtg tctgtggcct tgggacagac agtcaggatc 60acatgccaag
gagacagcct cagaagctat tatgcaagct ggtaccagca gaagccagga 120caggcccctg
tacttgtcat ctatggtaaa aacaaccggc cctcagggat cccagaccga 180ttctctggct
ccagctcagg aaacacagct tccttgacca tcactggggc tcaggcggaa 240gatgaggctg
actattactg taactcccgg gacagcagtg gtaacgtggt attcggcgga 300gggaccaagc
tgaccgtcct aggt
324927345DNAHomo sapiens 927gaggtgcagc tggtggagtc tgggggaggc ttggtacagc
ctggggggtc cctgagactc 60tcctgtgcag cctctggatt cacctttagc agctatgcca
tgagctgggt ccgccaggct 120ccagggaagg ggctggagtg ggtctcagct attagtggta
gtggtggtag cacatactac 180gcagactccg tgaagggccg gttcaccatc tccagagaca
attccaagaa ctcactgtat 240ctgcaaatga acagcctgag agccgaggac acggctgtgt
attactgtgc gagtgagcag 300gctggggact actggggcca gggaaccctg gtcaccgtct
cgaga 345928333DNAHomo sapiens 928cagtctgtgt
tgacgcagcc gccctcagcg tctgggaccc ccgggcagag ggtcaccatc 60tcttgttctg
gaagcagctc caacatcgga agtaattatg tatactggta ccagcagctc 120ccaggaacgg
cccccaaact cctcatctat aggaataatc agcggccctc aggggtccct 180gaccgattcg
ctggctccaa gtctggcacg tcagccaccc tgggcatcac cggactccag 240actggggacg
aggccgatta ttactgcgga acatgggata gcagcctgag tgctggggta 300ttcggcggag
ggaccaagct gaccgtccta ggt
333929357DNAHomo sapiens 929caggtgcagc tggtgcagtc tggggctgag gtgaagaagc
ctggggcctc agtgaaggtt 60tcctgcaagg catctggata caccttcacc agctactata
tgcactgggt gcgacaggcc 120cctggacaag ggcttgagtg gatgggaata atcaacccta
gtggtggtag cacaagctac 180gcacagaagt tccagggcag agtcaccatg accagggaca
cgtccacgag cacagtctac 240atggagctga gcagcctgag atctgaggac acggccgtgt
attactgtgc gagagaattg 300tcggctgcta ctgcttttga tatctggggc caagggacaa
tggtcaccgt ctcgaga 357930321DNAHomo sapiens 930gaaacgacac
tcacgcagtc tccattttct gtgtctgcat ctgtaggaga cagagtcacc 60atcacttgtc
gggcgagtca gtatattagc agatggctag cctggtatca gcagagacca 120gggaaagccc
ctaagctcct gatctatgct gcagccagtt tgcaaagtgg ggtcccatca 180aggttcagcg
gcagtggatc tgggacagat ttcactctca ctatcagcag cctgcaacct 240gaagattttg
caacttacta ttgccaacag gctaacagtt tccccgtcac cttcggccaa 300gggacacgac
tggagattaa a
321931360DNAHomo sapiens 931caggtgcagc tggtgcagtc tggagctgag gtgaagaagc
ctggggcctc agtgaaggtc 60tcctgcaagg cttctggtta cacctttacc agctatggta
tcagctgggt gcgacaggcc 120cctggacaag ggcttgagtg gatgggatgg atcagcgctt
acaatggtaa cacaaactat 180gcacagaagc tccagggcag agtcaccatg accacagaca
catccacgag cacagcctac 240atggagctga ggagcctgag atctgacgac acggccgtgt
attactgtgc gagagtaggt 300tattactact actacggtat ggacgtctgg ggccaaggga
ccacggtcac cgtctcgaga 360932333DNAHomo sapiens 932cagtctgccc
tgactcagcc tgcctccgtg tctgggtctc ctggacagtc gatcaccatc 60tcctgcactg
gaaccagcag tgatgttggg agttataacc ttgtctcctg gtaccaacag 120cacccaggca
aagcccccaa actcatgatt tatgagggca gtaagcggcc ctcaggggtt 180tctaatcgct
tctctggctc caagtctggc aacacggcct ccctgaccat ctctgggctc 240caggctgagg
acgaggctga ttattactgc agctcatata caagcagcag cacttgggtg 300ttcggcggag
ggaccaagct gaccgtccta ggt
333933354DNAHomo sapiens 933caggtgcagc tggtgcagtc tggggctgag gtgaagaagc
ctgggtcctc ggtgaaggtc 60tcctgcaagg cttctggagg caccttcagc agctatgcta
tcagctgggt gcgacaggcc 120cctggacaag ggcttgagtg gatgggatgg atgaacccta
acagtggtaa cacaggctat 180gcacagaagt tccagggcag agtcaccatg accaggaaca
cctccataag cacagcctac 240atggagctga gcagcctgag atctgaggac acggccgtgt
attactgtgc gagagacgtt 300gggattgggg tctttgacta ctggggccag ggaaccctgg
tcaccgtctc gaga 354934327DNAHomo sapiens 934tcctatgtgc
tgactcagcc accctcagtg tcagtggccc caggaaagac ggccaggatt 60acctgtgggg
gaaacaacat tggaagtaaa agtgtgcact ggtaccagca gaagccaggc 120caggcccctg
tgctggtcat ctattatgat agcgaccggc cctcagggat ccctgagcga 180ttctctggct
ccaactctgg gaacacggcc accctgacca tcagcagggt cgaagccggg 240gatgaggccg
actattactg tcaggtgtgg gatagtagta gtgatcatgt ggtattcggc 300ggagggacca
agctgaccgt cctaggt
327935363DNAHomo sapiens 935gaggtgcagc tggtggagtc tgggggaggc ttggtacagc
ctggggggtc cctgagactc 60tcctgtgcag cctctggatt cacctttagc gactatgcca
tgaactgggt ccgccaggct 120ccagggaagg ggctggagtg ggtctcagct actagtggta
gtggaggcag cacattctac 180gcagactccg tgaagggccg gttcaccatc tccagagaca
attccaagag tacgttgtat 240ctgcaaatga acagcctgag agacgaggac acggccgtgt
attactgtgc gaaagggacc 300ttaccgcatt actatgatag tagtggtata gggggccagg
gcaccctggt caccgtctcg 360agc
363936339DNAHomo sapiens 936caggctgtgc tcactcagcc
gtcttccgtg tctgggtctc ctggacagtc gatcaccatc 60tcctgcactg gaaccagcag
tgacgttggt ggttataact atgtctcctg gtaccaacag 120cacccaggca aagcccccaa
actcatgatt tatgatgtcg gtaagcggcc ctcaggggtt 180tctaatcgct tctctggctc
caagtctggc aacacggcct ccctgaccat ctctgggctc 240cgggctgagg acgaggctaa
ttattactgc agctcatata caagcagcag cacttggttt 300gtggtattcg gcggagggac
caagctgacc gtcctaggt 339937363DNAHomo sapiens
937caggtacagc tgcagcagtc aggggctgag gtgaagaagc ctggggcctc agtgaaggtt
60tcctgcaagg catctggata caccttcacc acctactata tgcactgggt gcgacaggcc
120cctggacaag ggcttgagtg gatgggaata atcaatccta gtggtgaaaa tacaaactac
180gcacagaagt tccagggcag agtcaccatg accagggaca catccacgac cacagtctac
240atggagctga gcagcctgag atctgaggac acggccgtgt attactgtgc gagatctttc
300ataggaactc gtgggggcgg tttggacgtc tggggccaag ggaccacggt caccgtctcg
360aga
363938339DNAHomo sapiens 938cagtctgtgt tgacgcagcc gccctcagtg tctggggccc
cagggcagag ggtcaacatc 60tcctgcgctg ggagcagctc caacatcggg gcgggttatg
atgttcactg gtaccagcag 120attccaggaa cagcccccaa actcctcatg tatggtaata
gtaatcggcc ctcaggggtc 180cctgaccgat tctctggctc caagtctggc gcctcagcct
ccctggccat cactaggctc 240caggctgagg atgaggctga ttattactgc cagtcctatg
acagcagcct gagtggttcg 300agggtcttcg gaactgggac caaggtcacc gtcctaggt
339939351DNAHomo sapiens 939gaggtgcagc tggtggagtc
tgggggaggc gtggtccagc ctgggaggtc cctgagactc 60tcctgtgcag cctctggatt
cacctttgat gattatgcca tgcactgggt ccggcaagct 120ccggggaagg gcctggagtg
ggtctcaggt attagttgga atagtggtag cataggctat 180gcggactctg tgaagggccg
attcaccatc tccagagaca acgccaagaa ctccctgtat 240ctgcaaatga acagtctgag
agctgaggac acggctgtgt attactgtgc gacggaagaa 300tggtggcgct tcgatctctg
gggccgtggc accctggtca ccgtctcgag a 351940333DNAHomo sapiens
940cagtctgccc tgactcagcc tccctccgcg tccgggtctc ctggacagtc agtcaccatc
60tcctgcactg gaaccagcag tgacgttgat gattacaact atgtctcctg gtaccaacag
120cacccaggca aagcccccaa actcatgatt tatgaggtca ctaagcggcc ctcaggggtc
180cctgatcgct tctctggctc caagtctggc aacacggcct ccctgaccat ctctgggctc
240caggctgagg atgaggctga ttattactgc agctcatatg tgggcagcga caatagagtc
300ttcggaactg ggaccaaggt caccgtccca ggt
3339411897PRTHomo sapiens 941Met Val Pro Leu Val Pro Ala Leu Val Met Leu
Gly Leu Val Ala Gly1 5 10
15Ala His Gly Asp Ser Lys Pro Val Phe Ile Lys Val Pro Glu Asp Gln
20 25 30Thr Gly Leu Ser Gly Gly Val
Ala Ser Phe Val Cys Gln Ala Thr Gly 35 40
45Glu Pro Lys Pro Arg Ile Thr Trp Met Lys Lys Gly Lys Lys Val
Ser 50 55 60Ser Gln Arg Phe Glu Val
Ile Glu Phe Asp Asp Gly Ala Gly Ser Val65 70
75 80Leu Arg Ile Gln Pro Leu Arg Val Gln Arg Asp
Glu Ala Ile Tyr Glu 85 90
95Cys Thr Ala Thr Asn Ser Leu Gly Glu Ile Asn Thr Ser Ala Lys Leu
100 105 110Ser Val Leu Glu Glu Glu
Gln Leu Pro Pro Gly Phe Pro Ser Ile Asp 115 120
125Met Gly Pro Gln Leu Lys Val Val Glu Lys Ala Arg Thr Ala
Thr Met 130 135 140Leu Cys Ala Ala Gly
Gly Asn Pro Asp Pro Glu Ile Ser Trp Phe Lys145 150
155 160Asp Phe Leu Pro Val Asp Pro Ala Thr Ser
Asn Gly Arg Ile Lys Gln 165 170
175Leu Arg Ser Gly Ala Leu Gln Ile Glu Ser Ser Glu Glu Ser Asp Gln
180 185 190Gly Lys Tyr Glu Cys
Val Ala Thr Asn Ser Ala Gly Thr Arg Tyr Ser 195
200 205Ala Pro Ala Asn Leu Tyr Val Arg Val Arg Arg Val
Ala Pro Arg Phe 210 215 220Ser Ile Pro
Pro Ser Ser Gln Glu Val Met Pro Gly Gly Ser Val Asn225
230 235 240Leu Thr Cys Val Ala Val Gly
Ala Pro Met Pro Tyr Val Lys Trp Met 245
250 255Met Gly Ala Glu Glu Leu Thr Lys Glu Asp Glu Met
Pro Val Gly Arg 260 265 270Asn
Val Leu Glu Leu Ser Asn Val Val Arg Ser Ala Asn Tyr Thr Cys 275
280 285Val Ala Ile Ser Ser Leu Gly Met Ile
Glu Ala Thr Ala Gln Val Thr 290 295
300Val Lys Ala Leu Pro Lys Pro Pro Ile Asp Leu Val Val Thr Glu Thr305
310 315 320Thr Ala Thr Ser
Val Thr Leu Thr Trp Asp Ser Gly Asn Ser Glu Pro 325
330 335Val Thr Tyr Tyr Gly Ile Gln Tyr Arg Ala
Ala Gly Thr Glu Gly Pro 340 345
350Phe Gln Glu Val Asp Gly Val Ala Thr Thr Arg Tyr Ser Ile Gly Gly
355 360 365Leu Ser Pro Phe Ser Glu Tyr
Ala Phe Arg Val Leu Ala Val Asn Ser 370 375
380Ile Gly Arg Gly Pro Pro Ser Glu Ala Val Arg Ala Arg Thr Gly
Glu385 390 395 400Gln Ala
Pro Ser Ser Pro Pro Arg Arg Val Gln Ala Arg Met Leu Ser
405 410 415Ala Ser Thr Met Leu Val Gln
Trp Glu Pro Pro Glu Glu Pro Asn Gly 420 425
430Leu Val Arg Gly Tyr Arg Val Tyr Tyr Thr Pro Asp Ser Arg
Arg Pro 435 440 445Pro Asn Ala Trp
His Lys His Asn Thr Asp Ala Gly Leu Leu Thr Thr 450
455 460Val Gly Ser Leu Leu Pro Gly Ile Thr Tyr Ser Leu
Arg Val Leu Ala465 470 475
480Phe Thr Ala Val Gly Asp Gly Pro Pro Ser Pro Thr Ile Gln Val Lys
485 490 495Thr Gln Gln Gly Val
Pro Ala Gln Pro Ala Asp Phe Gln Ala Glu Val 500
505 510Glu Ser Asp Thr Arg Ile Gln Leu Ser Trp Leu Leu
Pro Pro Gln Glu 515 520 525Arg Ile
Ile Met Tyr Glu Leu Val Tyr Trp Ala Ala Glu Asp Glu Asp 530
535 540Gln Gln His Lys Val Thr Phe Asp Pro Thr Ser
Ser Tyr Thr Leu Glu545 550 555
560Asp Leu Lys Pro Asp Thr Leu Tyr Arg Phe Gln Leu Ala Ala Arg Ser
565 570 575Asp Met Gly Val
Gly Val Phe Thr Pro Thr Ile Glu Ala Arg Thr Ala 580
585 590Gln Ser Thr Pro Ser Ala Pro Pro Gln Lys Val
Met Cys Val Ser Met 595 600 605Gly
Ser Thr Thr Val Arg Val Ser Trp Val Pro Pro Pro Ala Asp Ser 610
615 620Arg Asn Gly Val Ile Thr Gln Tyr Ser Val
Ala His Glu Ala Val Asp625 630 635
640Gly Glu Asp Arg Gly Arg His Val Val Asp Gly Ile Ser Arg Glu
His 645 650 655Ser Ser Trp
Asp Leu Val Gly Leu Glu Lys Trp Thr Glu Tyr Arg Val 660
665 670Trp Val Arg Ala His Thr Asp Val Gly Pro
Gly Pro Glu Ser Ser Pro 675 680
685Val Leu Val Arg Thr Asp Glu Asp Val Pro Ser Gly Pro Pro Arg Lys 690
695 700Val Glu Val Glu Pro Leu Asn Ser
Thr Ala Val His Val Tyr Trp Lys705 710
715 720Leu Pro Val Pro Ser Lys Gln His Gly Gln Ile Arg
Gly Tyr Gln Val 725 730
735Thr Tyr Val Arg Leu Glu Asn Gly Glu Pro Arg Gly Leu Pro Ile Ile
740 745 750Gln Asp Val Met Leu Ala
Glu Ala Gln Trp Arg Pro Glu Glu Ser Glu 755 760
765Asp Tyr Glu Thr Thr Ile Ser Gly Leu Thr Pro Glu Thr Thr
Tyr Ser 770 775 780Val Thr Val Ala Ala
Tyr Thr Thr Lys Gly Asp Gly Ala Arg Ser Lys785 790
795 800Pro Lys Ile Val Thr Thr Thr Gly Ala Val
Pro Gly Arg Pro Thr Met 805 810
815Met Ile Ser Thr Thr Ala Met Asn Thr Ala Leu Leu Gln Trp His Pro
820 825 830Pro Lys Glu Leu Pro
Gly Glu Leu Leu Gly Tyr Arg Leu Gln Tyr Cys 835
840 845Arg Ala Asp Glu Ala Arg Pro Asn Thr Ile Asp Phe
Gly Lys Asp Asp 850 855 860Gln His Phe
Thr Val Thr Gly Leu His Lys Gly Thr Thr Tyr Ile Phe865
870 875 880Arg Leu Ala Ala Lys Asn Arg
Ala Gly Leu Gly Glu Glu Phe Glu Lys 885
890 895Glu Ile Arg Thr Pro Glu Asp Leu Pro Ser Gly Phe
Pro Gln Asn Leu 900 905 910His
Val Thr Gly Leu Thr Thr Ser Thr Thr Glu Leu Ala Trp Asp Pro 915
920 925Pro Val Leu Ala Glu Arg Asn Gly Arg
Ile Ile Ser Tyr Thr Val Val 930 935
940Phe Arg Asp Ile Asn Ser Gln Gln Glu Leu Gln Asn Ile Thr Thr Asp945
950 955 960Thr Arg Phe Thr
Leu Thr Gly Leu Lys Pro Asp Thr Thr Tyr Asp Ile 965
970 975Lys Val Arg Ala Trp Thr Ser Lys Gly Ser
Gly Pro Leu Ser Pro Ser 980 985
990Ile Gln Ser Arg Thr Met Pro Val Glu Gln Val Phe Ala Lys Asn Phe
995 1000 1005Arg Val Ala Ala Ala Met
Lys Thr Ser Val Leu Leu Ser Trp Glu 1010 1015
1020Val Pro Asp Ser Tyr Lys Ser Ala Val Pro Phe Lys Ile Leu
Tyr 1025 1030 1035Asn Gly Gln Ser Val
Glu Val Asp Gly His Ser Met Arg Lys Leu 1040 1045
1050Ile Ala Asp Leu Gln Pro Asn Thr Glu Tyr Ser Phe Val
Leu Met 1055 1060 1065Asn Arg Gly Ser
Ser Ala Gly Gly Leu Gln His Leu Val Ser Ile 1070
1075 1080Arg Thr Ala Pro Asp Leu Leu Pro His Lys Pro
Leu Pro Ala Ser 1085 1090 1095Ala Tyr
Ile Glu Asp Gly Arg Phe Asp Leu Ser Met Pro His Val 1100
1105 1110Gln Asp Pro Ser Leu Val Arg Trp Phe Tyr
Ile Val Val Val Pro 1115 1120 1125Ile
Asp Arg Val Gly Gly Ser Met Leu Thr Pro Arg Trp Ser Thr 1130
1135 1140Pro Glu Glu Leu Glu Leu Asp Glu Leu
Leu Glu Ala Ile Glu Gln 1145 1150
1155Gly Gly Glu Glu Gln Arg Arg Arg Arg Arg Gln Ala Glu Arg Leu
1160 1165 1170Lys Pro Tyr Val Ala Ala
Gln Leu Asp Val Leu Pro Glu Thr Phe 1175 1180
1185Thr Leu Gly Asp Lys Lys Asn Tyr Arg Gly Phe Tyr Asn Arg
Pro 1190 1195 1200Leu Ser Pro Asp Leu
Ser Tyr Gln Cys Phe Val Leu Ala Ser Leu 1205 1210
1215Lys Glu Pro Met Asp Gln Lys Arg Tyr Ala Ser Ser Pro
Tyr Ser 1220 1225 1230Asp Glu Ile Val
Val Gln Val Thr Pro Ala Gln Gln Gln Glu Glu 1235
1240 1245Pro Glu Met Leu Trp Val Thr Gly Pro Val Leu
Ala Val Ile Leu 1250 1255 1260Ile Ile
Leu Ile Val Ile Ala Ile Leu Leu Phe Lys Arg Lys Arg 1265
1270 1275Thr His Ser Pro Ser Ser Lys Asp Glu Gln
Ser Ile Gly Leu Lys 1280 1285 1290Asp
Ser Leu Leu Ala His Ser Ser Asp Pro Val Glu Met Arg Arg 1295
1300 1305Leu Asn Tyr Gln Thr Pro Gly Met Arg
Asp His Pro Pro Ile Pro 1310 1315
1320Ile Thr Asp Leu Ala Asp Asn Ile Glu Arg Leu Lys Ala Asn Asp
1325 1330 1335Gly Leu Lys Phe Ser Gln
Glu Tyr Glu Ser Ile Asp Pro Gly Gln 1340 1345
1350Gln Phe Thr Trp Glu Asn Ser Asn Leu Glu Val Asn Lys Pro
Lys 1355 1360 1365Asn Arg Tyr Ala Asn
Val Ile Ala Tyr Asp His Ser Arg Val Ile 1370 1375
1380Leu Thr Ser Ile Asp Gly Val Pro Gly Ser Asp Tyr Ile
Asn Ala 1385 1390 1395Asn Tyr Ile Asp
Gly Tyr Arg Lys Gln Asn Ala Tyr Ile Ala Thr 1400
1405 1410Gln Gly Pro Leu Pro Glu Thr Met Gly Asp Phe
Trp Arg Met Val 1415 1420 1425Trp Glu
Gln Arg Thr Ala Thr Val Val Met Met Thr Arg Leu Glu 1430
1435 1440Glu Lys Ser Arg Val Lys Cys Asp Gln Tyr
Trp Pro Ala Arg Gly 1445 1450 1455Thr
Glu Thr Cys Gly Leu Ile Gln Val Thr Leu Leu Asp Thr Val 1460
1465 1470Glu Leu Ala Thr Tyr Thr Val Arg Thr
Phe Ala Leu His Lys Ser 1475 1480
1485Gly Ser Ser Glu Lys Arg Glu Leu Arg Gln Phe Gln Phe Met Ala
1490 1495 1500Trp Pro Asp His Gly Val
Pro Glu Tyr Pro Thr Pro Ile Leu Ala 1505 1510
1515Phe Leu Arg Arg Val Lys Ala Cys Asn Pro Leu Asp Ala Gly
Pro 1520 1525 1530Met Val Val His Cys
Ser Ala Gly Val Gly Arg Thr Gly Cys Phe 1535 1540
1545Ile Val Ile Asp Ala Met Leu Glu Arg Met Lys His Glu
Lys Thr 1550 1555 1560Val Asp Ile Tyr
Gly His Val Thr Cys Met Arg Ser Gln Arg Asn 1565
1570 1575Tyr Met Val Gln Thr Glu Asp Gln Tyr Val Phe
Ile His Glu Ala 1580 1585 1590Leu Leu
Glu Ala Ala Thr Cys Gly His Thr Glu Val Pro Ala Arg 1595
1600 1605Asn Leu Tyr Ala His Ile Gln Lys Leu Gly
Gln Val Pro Pro Gly 1610 1615 1620Glu
Ser Val Thr Ala Met Glu Leu Glu Phe Lys Leu Leu Ala Ser 1625
1630 1635Ser Lys Ala His Thr Ser Arg Phe Ile
Ser Ala Asn Leu Pro Cys 1640 1645
1650Asn Lys Phe Lys Asn Arg Leu Val Asn Ile Met Pro Tyr Glu Leu
1655 1660 1665Thr Arg Val Cys Leu Gln
Pro Ile Arg Gly Val Glu Gly Ser Asp 1670 1675
1680Tyr Ile Asn Ala Ser Phe Leu Asp Gly Tyr Arg Gln Gln Lys
Ala 1685 1690 1695Tyr Ile Ala Thr Gln
Gly Pro Leu Ala Glu Ser Thr Glu Asp Phe 1700 1705
1710Trp Arg Met Leu Trp Glu His Asn Ser Thr Ile Ile Val
Met Leu 1715 1720 1725Thr Lys Leu Arg
Glu Met Gly Arg Glu Lys Cys His Gln Tyr Trp 1730
1735 1740Pro Ala Glu Arg Ser Ala Arg Tyr Gln Tyr Phe
Val Val Asp Pro 1745 1750 1755Met Ala
Glu Tyr Asn Met Pro Gln Tyr Ile Leu Arg Glu Phe Lys 1760
1765 1770Val Thr Asp Ala Arg Asp Gly Gln Ser Arg
Thr Ile Arg Gln Phe 1775 1780 1785Gln
Phe Thr Asp Trp Pro Glu Gln Gly Val Pro Lys Thr Gly Glu 1790
1795 1800Gly Phe Ile Asp Phe Ile Gly Gln Val
His Lys Thr Lys Glu Gln 1805 1810
1815Phe Gly Gln Asp Gly Pro Ile Thr Val His Cys Ser Ala Gly Val
1820 1825 1830Gly Arg Thr Gly Val Phe
Ile Thr Leu Ser Ile Val Leu Glu Arg 1835 1840
1845Met Arg Tyr Glu Gly Val Val Asp Met Phe Gln Thr Val Lys
Thr 1850 1855 1860Leu Arg Thr Gln Arg
Pro Ala Met Val Gln Thr Glu Asp Gln Tyr 1865 1870
1875Gln Leu Cys Tyr Arg Ala Ala Leu Glu Tyr Leu Gly Ser
Phe Asp 1880 1885 1890His Tyr Ala Thr
1895942628PRTHomo sapiens 942Met Glu Pro Pro Asp Ala Pro Ala Gln Ala
Arg Gly Ala Pro Arg Leu1 5 10
15Leu Leu Leu Ala Val Leu Leu Ala Ala His Pro Asp Ala Gln Ala Glu
20 25 30Val Arg Leu Ser Val Pro
Pro Leu Val Glu Val Met Arg Gly Lys Ser 35 40
45Val Ile Leu Asp Cys Thr Pro Thr Gly Thr His Asp His Tyr
Met Leu 50 55 60Glu Trp Phe Leu Thr
Asp Arg Ser Gly Ala Arg Pro Arg Leu Ala Ser65 70
75 80Ala Glu Met Gln Gly Ser Glu Leu Gln Val
Thr Met His Asp Thr Arg 85 90
95Gly Arg Ser Pro Pro Tyr Gln Leu Asp Ser Gln Gly Arg Leu Val Leu
100 105 110Ala Glu Ala Gln Val
Gly Asp Glu Arg Asp Tyr Val Cys Val Val Arg 115
120 125Ala Gly Ala Ala Gly Thr Ala Glu Ala Thr Ala Arg
Leu Asn Val Phe 130 135 140Ala Lys Pro
Glu Ala Thr Glu Val Ser Pro Asn Lys Gly Thr Leu Ser145
150 155 160Val Met Glu Asp Ser Ala Gln
Glu Ile Ala Thr Cys Asn Ser Arg Asn 165
170 175Gly Asn Pro Ala Pro Lys Ile Thr Trp Tyr Arg Asn
Gly Gln Arg Leu 180 185 190Glu
Val Pro Val Glu Met Asn Pro Glu Gly Tyr Met Thr Ser Arg Thr 195
200 205Val Arg Glu Ala Ser Gly Leu Leu Ser
Leu Thr Ser Thr Leu Tyr Leu 210 215
220Arg Leu Arg Lys Asp Asp Arg Asp Ala Ser Phe His Cys Ala Ala His225
230 235 240Tyr Ser Leu Pro
Glu Gly Arg His Gly Arg Leu Asp Ser Pro Thr Phe 245
250 255His Leu Thr Leu His Tyr Pro Thr Glu His
Val Gln Phe Trp Val Gly 260 265
270Ser Pro Ser Thr Pro Ala Gly Trp Val Arg Glu Gly Asp Thr Val Gln
275 280 285Leu Leu Cys Arg Gly Asp Gly
Ser Pro Ser Pro Glu Tyr Thr Leu Phe 290 295
300Arg Leu Gln Asp Glu Gln Glu Glu Val Leu Asn Val Asn Leu Glu
Gly305 310 315 320Asn Leu
Thr Leu Glu Gly Val Thr Arg Gly Gln Ser Gly Thr Tyr Gly
325 330 335Cys Arg Val Glu Asp Tyr Asp
Ala Ala Asp Asp Val Gln Leu Ser Lys 340 345
350Thr Leu Glu Leu Arg Val Ala Tyr Leu Asp Pro Leu Glu Leu
Ser Glu 355 360 365Gly Lys Val Leu
Ser Leu Pro Leu Asn Ser Ser Ala Val Val Asn Cys 370
375 380Ser Val His Gly Leu Pro Thr Pro Ala Leu Arg Trp
Thr Lys Asp Ser385 390 395
400Thr Pro Leu Gly Asp Gly Pro Met Leu Ser Leu Ser Ser Ile Thr Phe
405 410 415Asp Ser Asn Gly Thr
Tyr Val Cys Glu Ala Ser Leu Pro Thr Val Pro 420
425 430Val Leu Ser Arg Thr Gln Asn Phe Thr Leu Leu Val
Gln Gly Ser Pro 435 440 445Glu Leu
Lys Thr Ala Glu Ile Glu Pro Lys Ala Asp Gly Ser Trp Arg 450
455 460Glu Gly Asp Glu Val Thr Leu Ile Cys Ser Ala
Arg Gly His Pro Asp465 470 475
480Pro Lys Leu Ser Trp Ser Gln Leu Gly Gly Ser Pro Ala Glu Pro Ile
485 490 495Pro Gly Arg Gln
Gly Trp Val Ser Ser Ser Leu Thr Leu Lys Val Thr 500
505 510Ser Ala Leu Ser Arg Asp Gly Ile Ser Cys Glu
Ala Ser Asn Pro His 515 520 525Gly
Asn Lys Arg His Val Phe His Phe Gly Thr Val Ser Pro Gln Thr 530
535 540Ser Gln Ala Gly Val Ala Val Met Ala Val
Ala Val Ser Val Gly Leu545 550 555
560Leu Leu Leu Val Val Ala Val Phe Tyr Cys Val Arg Arg Lys Gly
Gly 565 570 575Pro Cys Cys
Arg Gln Arg Arg Glu Lys Gly Ala Pro Pro Pro Gly Glu 580
585 590Pro Gly Leu Ser His Ser Gly Ser Glu Gln
Pro Glu Gln Thr Gly Leu 595 600
605Leu Met Gly Gly Ala Ser Gly Gly Ala Arg Gly Gly Ser Gly Gly Phe 610
615 620Gly Asp Glu Cys625943621PRTHomo
sapiens 943Leu Glu Glu Lys Lys Val Cys Gln Gly Thr Ser Asn Lys Leu Thr
Gln1 5 10 15Leu Gly Thr
Phe Glu Asp His Phe Leu Ser Leu Gln Arg Met Phe Asn 20
25 30Asn Cys Glu Val Val Leu Gly Asn Leu Glu
Ile Thr Tyr Val Gln Arg 35 40
45Asn Tyr Asp Leu Ser Phe Leu Lys Thr Ile Gln Glu Val Ala Gly Tyr 50
55 60Val Leu Ile Ala Leu Asn Thr Val Glu
Arg Ile Pro Leu Glu Asn Leu65 70 75
80Gln Ile Ile Arg Gly Asn Met Tyr Tyr Glu Asn Ser Tyr Ala
Leu Ala 85 90 95Val Leu
Ser Asn Tyr Asp Ala Asn Lys Thr Gly Leu Lys Glu Leu Pro 100
105 110Met Arg Asn Leu Gln Glu Ile Leu His
Gly Ala Val Arg Phe Ser Asn 115 120
125Asn Pro Ala Leu Cys Asn Val Glu Ser Ile Gln Trp Arg Asp Ile Val
130 135 140Ser Ser Asp Phe Leu Ser Asn
Met Ser Met Asp Phe Gln Asn His Leu145 150
155 160Gly Ser Cys Gln Lys Cys Asp Pro Ser Cys Pro Asn
Gly Ser Cys Trp 165 170
175Gly Ala Gly Glu Glu Asn Cys Gln Lys Leu Thr Lys Ile Ile Cys Ala
180 185 190Gln Gln Cys Ser Gly Arg
Cys Arg Gly Lys Ser Pro Ser Asp Cys Cys 195 200
205His Asn Gln Cys Ala Ala Gly Cys Thr Gly Pro Arg Glu Ser
Asp Cys 210 215 220Leu Val Cys Arg Lys
Phe Arg Asp Glu Ala Thr Cys Lys Asp Thr Cys225 230
235 240Pro Pro Leu Met Leu Tyr Asn Pro Thr Thr
Tyr Gln Met Asp Val Asn 245 250
255Pro Glu Gly Lys Tyr Ser Phe Gly Ala Thr Cys Val Lys Lys Cys Pro
260 265 270Arg Asn Tyr Val Val
Thr Asp His Gly Ser Cys Val Arg Ala Cys Gly 275
280 285Ala Asp Ser Tyr Glu Met Glu Glu Asp Gly Val Arg
Lys Cys Lys Lys 290 295 300Cys Glu Gly
Pro Cys Arg Lys Val Cys Asn Gly Ile Gly Ile Gly Glu305
310 315 320Phe Lys Asp Ser Leu Ser Ile
Asn Ala Thr Asn Ile Lys His Phe Lys 325
330 335Asn Cys Thr Ser Ile Ser Gly Asp Leu His Ile Leu
Pro Val Ala Phe 340 345 350Arg
Gly Asp Ser Phe Thr His Thr Pro Pro Leu Asp Pro Gln Glu Leu 355
360 365Asp Ile Leu Lys Thr Val Lys Glu Ile
Thr Gly Phe Leu Leu Ile Gln 370 375
380Ala Trp Pro Glu Asn Arg Thr Asp Leu His Ala Phe Glu Asn Leu Glu385
390 395 400Ile Ile Arg Gly
Arg Thr Lys Gln His Gly Gln Phe Ser Leu Ala Val 405
410 415Val Ser Leu Asn Ile Thr Ser Leu Gly Leu
Arg Ser Leu Lys Glu Ile 420 425
430Ser Asp Gly Asp Val Ile Ile Ser Gly Asn Lys Asn Leu Cys Tyr Ala
435 440 445Asn Thr Ile Asn Trp Lys Lys
Leu Phe Gly Thr Ser Gly Gln Lys Thr 450 455
460Lys Ile Ile Ser Asn Arg Gly Glu Asn Ser Cys Lys Ala Thr Gly
Gln465 470 475 480Val Cys
His Ala Leu Cys Ser Pro Glu Gly Cys Trp Gly Pro Glu Pro
485 490 495Arg Asp Cys Val Ser Cys Arg
Asn Val Ser Arg Gly Arg Glu Cys Val 500 505
510Asp Lys Cys Asn Leu Leu Glu Gly Glu Pro Arg Glu Phe Val
Glu Asn 515 520 525Ser Glu Cys Ile
Gln Cys His Pro Glu Cys Leu Pro Gln Ala Met Asn 530
535 540Ile Thr Cys Thr Gly Arg Gly Pro Asp Asn Cys Ile
Gln Cys Ala His545 550 555
560Tyr Ile Asp Gly Pro His Cys Val Lys Thr Cys Pro Ala Gly Val Met
565 570 575Gly Glu Asn Asn Thr
Leu Val Trp Lys Tyr Ala Asp Ala Gly His Val 580
585 590Cys His Leu Cys His Pro Asn Cys Thr Tyr Gly Cys
Thr Gly Pro Gly 595 600 605Leu Glu
Gly Cys Pro Thr Asn Gly Pro Lys Ile Pro Ser 610 615
620944123PRTHomo sapiens 944Gln Val Gln Leu Val Gln Ser Gly
Ala Glu Val Lys Lys Pro Gly Ala1 5 10
15Ser Val Lys Val Ser Cys Lys Val Ser Gly Tyr Thr Leu Thr
Glu Leu 20 25 30Ser Met His
Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Met 35
40 45Gly Gly Phe Asp Pro Glu Asp Gly Glu Thr Ile
Tyr Ala Gln Lys Phe 50 55 60Gln Gly
Arg Val Thr Met Thr Glu Asp Thr Ser Thr Asp Thr Ala Tyr65
70 75 80Met Glu Leu Ser Ser Leu Arg
Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90
95Ala Thr Asp Leu Asp Tyr Tyr Gly Ser Gly Ser Tyr Ala
Phe Asp Ile 100 105 110Trp Gly
Gln Gly Thr Thr Val Thr Val Ser Arg 115
120945111PRTHomo sapiens 945Gln Ser Ala Leu Thr Gln Pro Ala Ser Val Ser
Gly Ser Pro Gly Gln1 5 10
15Ser Ile Thr Ile Ser Cys Thr Gly Thr Ser Ser Asp Val Gly Ser Tyr20
25 30Asn Leu Val Ser Trp Tyr Gln Gln His Pro
Gly Lys Ala Pro Lys Leu35 40 45Met Ile
Tyr Glu Gly Ser Lys Arg Pro Ser Gly Val Ser Asn Arg Phe50
55 60Ser Gly Ser Lys Ser Gly Asn Thr Ala Ser Leu Thr
Ile Ser Gly Leu65 70 75
80Gln Ala Glu Asp Glu Ala Asp Tyr Tyr Cys Ser Ser Tyr Thr Ser Ser
85 90 95Ser Thr Trp Val Phe Gly
Gly Gly Thr Lys Leu Thr Val Leu Gly 100 105
110946123PRTHomo sapiens 946Gln Val Gln Leu Gln Gln Trp Gly
Ala Gly Leu Leu Lys Pro Ser Glu1 5 10
15Thr Leu Ser Leu Thr Cys Ala Ala Tyr Gly Gly Ser Phe Ser
Gly Tyr 20 25 30Tyr Trp Ser
Trp Ile Arg Gln Pro Pro Gly Lys Gly Leu Glu Trp Ile 35
40 45Gly Glu Ile Asn His Ser Gly Ser Thr Asn Tyr
Asn Pro Ser Leu Lys 50 55 60Ser Arg
Val Thr Ile Ser Val Asp Thr Ser Lys Asn Gln Phe Ser Leu65
70 75 80Lys Leu Ser Ser Val Thr Ala
Ala Asp Thr Ala Val Tyr Tyr Cys Ala 85 90
95Arg Thr Tyr Tyr Gly Ser Gly Ser Tyr Gln Tyr Asn Trp
Phe Asp Pro 100 105 110Trp Gly
Gln Gly Thr Leu Val Thr Val Ser Arg 115
120947112PRTHomo sapiens 947Gln Ser Val Leu Thr Gln Pro Pro Ser Val Ser
Gly Ala Pro Gly Gln1 5 10
15Arg Val Thr Ile Ser Cys Thr Gly Ser Ser Ser Asn Ile Gly Ala Gly
20 25 30Tyr Asp Val His Trp Tyr Gln
Gln Leu Pro Gly Thr Ala Pro Lys Leu 35 40
45Leu Ile Tyr Gly Asn Ser Asn Arg Pro Ser Gly Val Pro Asp Arg
Phe 50 55 60Ser Gly Ser Lys Ser Gly
Thr Ser Ala Ser Leu Ala Ile Thr Gly Leu65 70
75 80Gln Ala Glu Asp Glu Ala Asp Tyr Tyr Cys Gln
Ser Tyr Asp Ser Ser 85 90
95Leu Ser Gly Val Val Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly
100 105 110948117PRTHomo sapiens
948Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly1
5 10 15Ser Leu Arg Leu Ser Cys
Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr 20 25
30Ser Met Asn Trp Val Arg Gln Ala Pro Gly Lys Gly Leu
Glu Trp Val 35 40 45Ser Tyr Ile
Ser Ser Ser Ser Ser Thr Ile Tyr Tyr Ala Asp Ser Val 50
55 60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys
Asn Ser Leu Tyr65 70 75
80Leu Gln Met Asn Ser Leu Arg Asp Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Ala Arg Asp Asn Leu Glu
Gly Leu Asp Tyr Trp Gly Gln Gly Thr Leu 100
105 110Val Thr Val Ser Arg 115949109PRTHomo
sapiens 949Ser Tyr Val Leu Thr Gln Pro Pro Ser Val Ser Val Ala Pro Gly
Lys1 5 10 15Thr Ala Arg
Ile Thr Cys Gly Gly Asn Asn Ile Gly Ser Lys Ser Val 20
25 30His Trp Tyr Gln Gln Lys Pro Gly Gln Ala
Pro Val Leu Val Ile Tyr 35 40
45Tyr Asp Ser Asp Arg Pro Ser Gly Ile Pro Glu Arg Phe Ser Gly Ser 50
55 60Asn Ser Gly Asn Thr Ala Thr Leu Thr
Ile Ser Arg Val Glu Ala Gly65 70 75
80Asp Glu Ala Asp Tyr Tyr Cys Gln Val Trp Asp Ser Ser Ser
Asp His 85 90 95Val Val
Phe Gly Gly Gly Thr Lys Leu Thr Val Leu Gly 100
105950120PRTHomo sapiens 950Gln Val Gln Leu Gln Glu Ser Gly Pro Gly Leu
Val Lys Pro Ser Gln1 5 10
15Thr Leu Ser Leu Thr Cys Thr Val Ser Gly Gly Ser Ile Ser Ser Gly
20 25 30Asp Tyr Tyr Trp Ser Trp Ile
Arg Gln Pro Pro Gly Lys Gly Leu Glu 35 40
45Trp Ile Gly Tyr Ile Tyr Tyr Ser Gly Ser Thr Tyr Tyr Asn Pro
Ser 50 55 60Leu Lys Ser Arg Val Thr
Ile Ser Val Asp Thr Ser Lys Asn Gln Phe65 70
75 80Ser Leu Lys Leu Ser Ser Val Thr Ala Ala Asp
Thr Ala Val Tyr Tyr 85 90
95Cys Ala Arg Gly Thr Gly Asp Leu Glu Trp Phe Asp Pro Trp Gly Gln
100 105 110Gly Thr Leu Val Thr Val Ser
Arg 115 120951111PRTHomo sapiens 951Gln Ser Val Leu
Thr Gln Pro Pro Ser Val Ser Gly Ala Pro Arg Gln1 5
10 15Thr Val Thr Ile Ser Cys Ser Gly Ser Ser
Ser Asn Ile Gly Gln Asn 20 25
30Ser Val Thr Trp Tyr Gln Arg Leu Pro Gly Glu Ala Pro Lys Leu Leu
35 40 45Ile Tyr Tyr Asp Asp Leu Leu His
Ser Gly Val Ser Asp Arg Phe Ser 50 55
60Gly Ser Lys Ser Gly Thr Ser Ala Ser Leu Ala Ile Ser Gly Leu Gln65
70 75 80Ser Glu Asp Glu Ala
Glu Tyr Tyr Cys Ala Ser Trp Asp Asp Ser Leu 85
90 95Lys Gly Pro Val Phe Gly Gly Gly Thr Lys Leu
Thr Val Leu Gly 100 105
110952121PRTHomo sapiens 952Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu
Val Gln Pro Gly Gly1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Ser Tyr
20 25 30Ser Met Asn Trp Val Arg Gln
Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40
45Ser Tyr Ile Ser Ser Ser Ser Ser Thr Ile Tyr Tyr Ala Asp Ser
Val 50 55 60Lys Gly Arg Phe Thr Ile
Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr65 70
75 80Leu Gln Met Asn Ser Leu Arg Asp Glu Asp Thr
Ala Val Tyr Tyr Cys 85 90
95Ala Arg Glu Gly Pro Arg Gly Ser Tyr Tyr Tyr Phe Asp Tyr Trp Gly
100 105 110Gln Gly Thr Leu Val Thr
Val Ser Arg 115 120953111PRTHomo sapiens 953Asn
Phe Met Leu Thr Gln Pro His Ser Val Ser Glu Ser Pro Gly Lys1
5 10 15Thr Val Thr Ile Ser Cys Thr
Arg Ser Ser Gly Ser Ile Ala Ser Asn 20 25
30Tyr Val Gln Trp Tyr Gln Gln Arg Pro Gly Ser Ala Pro Thr
Thr Val 35 40 45Ile Tyr Glu Asp
Asn Gln Arg Pro Ser Gly Val Pro Asp Arg Phe Ser 50 55
60Gly Ser Ile Asp Ser Ser Ser Asn Ser Ala Ser Leu Thr
Ile Ser Gly65 70 75
80Leu Lys Thr Glu Asp Glu Ala Asp Tyr Tyr Cys Gln Ser Tyr Asp Ser
85 90 95Ser Asn Trp Val Phe Gly
Gly Gly Thr Lys Leu Thr Val Leu Gly 100 105
110954119PRTHomo sapiens 954Gln Val Gln Leu Val Gln Ser Gly
Ala Glu Val Lys Lys Pro Gly Ala1 5 10
15Ser Val Lys Val Ser Cys Lys Ala Ser Gly Tyr Thr Phe Thr
Ser Tyr 20 25 30Tyr Met His
Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met 35
40 45Gly Ile Ile Asn Pro Ser Gly Gly Ser Thr Ser
Tyr Ala Gln Lys Phe 50 55 60Gln Gly
Arg Val Thr Met Thr Arg Asp Thr Ser Thr Ser Thr Val Tyr65
70 75 80Met Glu Leu Ser Ser Leu Arg
Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85 90
95Ala Arg Ala Lys Arg Arg Gly Ser Ala Phe Asp Ile Trp
Gly Gln Gly 100 105 110Thr Thr
Val Thr Val Ser Arg 115955109PRTHomo sapiens 955Ser Ser Glu Leu
Thr Gln Asp Pro Ala Val Ser Val Ala Leu Gly Gln1 5
10 15Thr Val Arg Ile Thr Cys Gln Gly Asp Ser
Leu Arg Ser Tyr Tyr Ala 20 25
30Ser Trp Tyr Gln Gln Lys Pro Gly Gln Ala Pro Val Leu Val Ile Tyr
35 40 45Gly Lys Asn Asn Arg Pro Ser Gly
Ile Pro Asp Arg Phe Ser Gly Ser 50 55
60Ser Ser Gly Asn Thr Ala Ser Leu Thr Ile Thr Gly Ala Gln Ala Glu65
70 75 80Asp Glu Ala Asp Tyr
Tyr Cys Asn Ser Arg Asp Ser Ser Gly Asn His 85
90 95Val Val Phe Gly Gly Gly Thr Lys Leu Thr Val
Leu Gly 100 105956369DNAHomo sapiens
956caggtgcagc tggtgcagtc tggagctgag gtgaagaagc ctggggcctc agtgaaggtc
60tcctgcaagg tttccggata caccctcact gaattatcca tgcactgggt gcgacaggct
120cctggaaaag ggcttgagtg gatgggaggt tttgatcctg aagatggtga aacaatctac
180gcacagaagt tccagggcag agtcaccatg accgaggaca catctacaga cacagcctac
240atggagctga gcagcctgag atctgaggac acggccgtgt attactgtgc aacagatctc
300gattactatg gttcggggag ttatgctttt gatatctggg gccaagggac cacggtcacc
360gtctcgaga
369957333DNAHomo sapiens 957cagtctgccc tgactcagcc tgcctccgtg tctgggtctc
ctggacagtc gatcaccatc 60tcctgcactg gaaccagcag tgatgttggg agttataacc
ttgtctcctg gtaccaacag 120cacccaggca aagcccccaa actcatgatt tatgagggca
gtaagcggcc ctcaggggtt 180tctaatcgct tctctggctc caagtctggc aacacggcct
ccctgaccat ctctgggctc 240caggctgagg acgaggctga ttattactgc agctcatata
caagcagcag cacttgggtg 300ttcggcggag ggaccaagct gaccgtccta ggt
333958369DNAHomo sapiens 958caggtgcagc tacagcagtg
gggcgcagga ctgttgaagc cttcggagac cctgtccctc 60acctgcgctg cctatggtgg
gtccttcagt ggttactact ggagctggat ccgccagccc 120ccagggaagg ggctggagtg
gattggggaa atcaatcata gtggaagcac caactacaac 180ccgtccctca agagtcgagt
caccatatca gtagacacgt ccaagaacca gttctccctg 240aagctgagct ctgtgaccgc
cgcggacacg gctgtgtatt actgtgcgag gacttactat 300ggttcgggga gttatcagta
caactggttc gacccctggg gccagggaac cctggtcacc 360gtctcgaga
369959336DNAHomo sapiens
959cagtctgtgt tgacgcagcc gccctcagtg tctggggccc cagggcagag ggtcaccatc
60tcctgcactg ggagcagctc caacatcggg gcaggttatg atgtacactg gtaccagcag
120cttccaggaa cagcccccaa actcctcatc tatggtaaca gcaatcggcc ctcaggggtc
180cctgaccgat tctctggctc caagtctggc acctcagcct ccttggccat cactgggctc
240caggctgagg atgaggctga ttattactgc cagtcctatg acagcagcct gagtggtgtg
300gtattcggcg gagggaccaa gctgaccgtc ctaggt
336960351DNAHomo sapiens 960gaggtgcagc tggtggagtc tgggggaggc ttggtacagc
ctggggggtc cctgagactc 60tcctgtgcag cctctggatt caccttcagt agctatagca
tgaactgggt ccgccaggct 120ccagggaagg ggctggagtg ggtttcatat attagtagta
gtagtagtac catatactac 180gcagactctg tgaagggccg attcaccatc tccagagaca
atgccaagaa ctcactgtat 240ctgcaaatga acagcctgag agacgaggac acggctgtgt
attactgtgc gagagataat 300cttgaaggcc tggactactg gggccaggga accctggtca
ccgtctcgag a 351961327DNAHomo sapiens 961tcctatgtgc
tgactcagcc accctcagtg tcagtggccc caggaaagac ggccaggatt 60acctgtgggg
gaaacaacat tggaagtaaa agtgtgcact ggtaccagca gaagccaggc 120caggcccctg
tgctggtcat ctattatgat agcgaccggc cctcagggat ccctgagcga 180ttctctggct
ccaactctgg gaacacggcc accctgacca tcagcagggt cgaagccggg 240gatgaggccg
actattactg tcaggtgtgg gatagtagta gtgatcatgt ggtattcggc 300ggagggacca
agctgaccgt cctaggt
327962360DNAHomo sapiens 962caggtgcagc tgcaggagtc gggcccagga ctggtgaagc
cttcacagac cctgtccctc 60acctgcactg tctctggtgg ctccatcagc agtggtgatt
actactggag ttggatccgc 120cagcccccag ggaagggcct ggagtggatt gggtacatct
attacagtgg gagcacctac 180tacaacccgt ccctcaagag tcgagttacc atatcagtag
acacgtccaa gaaccagttc 240tccctgaagc tgagctctgt gactgccgca gacacggccg
tgtattactg tgccagaggg 300actggggatc ttgagtggtt cgacccctgg ggccagggca
ccctggtcac cgtctcgaga 360963333DNAHomo sapiens 963cagtctgtgt
tgacgcagcc gccctcggtg tctggggccc cccggcagac ggtcaccatc 60tcctgctctg
ggagcagctc caacatcgga caaaattctg ttacctggta ccagcgcctc 120ccgggtgagg
ctcccaaact cctcatctac tatgatgatc tcttgcactc aggagtctct 180gaccgattct
ctggctccaa gtctggcacc tcagcctcac tggccatcag tggactccag 240tctgaggatg
aggctgagta ctactgtgcg tcatgggatg acagcctgaa aggtccggta 300ttcggcggag
ggaccaaact gaccgtccta ggt
333964363DNAHomo sapiens 964gaggtgcagc tggtggagtc tgggggaggc ttggtacagc
ctggggggtc cctgagactc 60tcctgtgcag cctctggatt caccttcagt agctatagca
tgaactgggt ccgccaggct 120ccagggaagg ggctggagtg ggtttcatac attagtagta
gtagtagtac catatactac 180gcagactctg tgaagggccg attcaccatc tccagagaca
atgccaagaa ctcactgtat 240ctgcaaatga acagcctgag agacgaggac acggctgtgt
attactgtgc gagagagggc 300ccaaggggga gctactacta ctttgactac tggggccagg
gaaccctggt caccgtctcg 360aga
363965333DNAHomo sapiens 965aattttatgc tgactcagcc
gcactctgtg tcggagtctc cggggaagac ggtaaccatc 60tcctgcaccc gcagcagtgg
cagcattgcc agcaactatg tgcagtggta ccagcagcgc 120ccgggcagtg cccccaccac
tgtgatctat gaggataacc aaagaccctc tggggtccct 180gatcggttct ctggctccat
cgacagctcc tccaactctg cctccctcac catctctgga 240ctgaagactg aggacgaggc
tgactactac tgtcagtctt atgatagcag caattgggtg 300ttcggcggag ggaccaagct
gaccgtccta ggt 333966357DNAHomo sapiens
966caggtgcagc tggtgcagtc tggggctgag gtgaagaagc ctggggcctc agtgaaggtt
60tcctgcaagg catctggata caccttcacc agctactata tgcactgggt gcgacaggcc
120cctggacaag ggcttgagtg gatgggaata atcaacccta gtggtggtag cacaagctac
180gcacagaagt tccagggcag agtcaccatg accagggaca cgtccacgag cacagtctac
240atggagctga gcagcctgag atctgaggac acggccgtgt attactgtgc gagagcgaag
300agaaggggat ctgcttttga tatctggggc caagggacca cggtcaccgt ctcgaga
357967327DNAHomo sapiens 967tcttctgagc tgactcagga ccctgctgtg tctgtggcct
tgggacagac agtcaggatc 60acatgccaag gagacagcct cagaagctat tatgcaagct
ggtaccagca gaagccagga 120caggcccctg tacttgtcat ctatggtaaa aacaaccggc
cctcagggat cccagaccga 180ttctctggct ccagctcagg aaacacagct tccttgacca
tcactggggc tcaggcggaa 240gatgaggctg actattactg taactcccgg gacagcagtg
gtaaccatgt ggtattcggc 300ggagggacca agctgaccgt cctaggt
327968124PRTHomo sapiens 968Gln Val Gln Leu Val
Gln Ser Gly Ala Glu Val Lys Lys Pro Gly Ser1 5
10 15Ser Val Lys Val Ser Cys Arg Ala Ser Gly Gly
Thr Phe Ser Ser Tyr 20 25
30Ala Ile Ser Trp Val Arg Gln Ala Pro Gly Gln Gly Leu Glu Trp Met
35 40 45Gly Gly Ile Ile Pro Ile Phe Gly
Thr Ala Asn Tyr Ala Gln Lys Phe 50 55
60Gln Gly Arg Val Thr Ile Thr Ala Asp Lys Ser Thr Ser Thr Ala Tyr65
70 75 80Met Glu Leu Ser Ser
Leu Arg Ser Glu Asp Thr Ala Val Tyr Tyr Cys 85
90 95Ala Gly Asp Ile Ser Arg Gly Ser Ser Trp Tyr
Gly Tyr Tyr Phe Asp 100 105
110Tyr Trp Gly Gln Gly Thr Leu Val Thr Val Ser Arg 115
120969106PRTHomo sapiens 969Asp Ile Gln Met Thr Gln Ser Pro Ser Thr
Leu Ala Ala Ser Val Gly1 5 10
15Asp Arg Val Thr Ile Thr Cys Arg Ala Ser Gln Ser Ile Ser Ser Trp
20 25 30Leu Ala Trp Tyr Gln Gln
Lys Pro Gly Lys Ala Pro Lys Leu Leu Ile 35 40
45Tyr Lys Ala Ser Ser Leu Glu Ser Gly Val Pro Ser Arg Phe
Ser Gly 50 55 60Ser Gly Ser Gly Thr
Glu Phe Thr Leu Thr Ile Ser Ser Leu Gln Pro65 70
75 80Asp Asp Phe Ala Thr Tyr Tyr Cys Gln Gln
Tyr Asn Ser Tyr Ser Thr 85 90
95Phe Gly Gln Gly Thr Lys Leu Glu Ile Lys 100
105970372DNAHomo sapiens 970caggtgcagc tggtgcagtc tggggctgag
gtgaagaagc ctgggtcctc ggtgaaggtc 60tcctgcaggg cttctggagg caccttcagc
agctatgcta tcagctgggt gcgacaggcc 120cctggacaag ggcttgagtg gatgggaggg
atcatcccta tctttggtac agcaaactac 180gcacagaagt tccagggcag agtcacgatt
accgcggaca aatccacgag cacagcctac 240atggagctga gcagcctgag atctgaggac
acggccgtgt attactgtgc gggagatatc 300agccgaggca gcagctggta cgggtactac
tttgactact ggggccaggg aaccctggtc 360accgtctcga ga
372971318DNAHomo sapiens 971gacatccaga
tgacccagtc tccttccacc ctggctgcat ctgtaggaga cagagtcacc 60atcacttgcc
gggccagtca gagtattagt agctggttgg cctggtatca gcagaaacca 120gggaaagccc
ctaagctcct gatctataag gcgtctagtt tagaaagtgg ggtcccatca 180aggttcagcg
gcagtggatc tgggacagaa ttcactctca ccatcagcag cctgcagcct 240gatgattttg
caacttatta ctgccaacag tataatagtt attccacttt tggccagggg 300accaagctgg
agatcaaa 318
User Contributions:
comments("1"); ?> comment_form("1"); ?>Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
User Contributions:
Comment about this patent or add new information about this topic: