Patent application title: Human complement C3 derivates with cobra venom factor-like function
Inventors:
Carl-Wilhelm Vogel (Honolulu, HI, US)
David C. Fritzinger (Honolulu, HI, US)
IPC8 Class: AA61K3800FI
USPC Class:
514 12
Class name: Designated organic active ingredient containing (doai) peptide containing (e.g., protein, peptones, fibrinogen, etc.) doai 25 or more peptide repeating units in known peptide chain structure
Publication date: 2008-09-25
Patent application number: 20080234191
Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
Patent application title: Human complement C3 derivates with cobra venom factor-like function
Inventors:
Carl-Wilhelm Vogel
David C. Fritzinger
Agents:
MORRISON & FOERSTER LLP
Assignees:
Origin: PALO ALTO, CA US
IPC8 Class: AA61K3800FI
USPC Class:
514 12
Abstract:
A modified human complement C3 protein (C3) is disclosed comprising a
substitution of a portion of a human C3 protein, with a corresponding
portion of a Cobra Venom Factor protein (CVF) which results in a human C3
protein with CVF functions, but with substantially reduced
immunogenicity. Advantageously, the C3 protein can be manipulated to
contain at least one of the following CVF functions: increased stability
of the C3 convertase and increased resistance to the actions of factors H
and/or I. A large number of hybrid C3 proteins containing substitutions
in the C-terminal portion of the alpha chain of C3 are presented and
tested for the above functions. Methods of treatment of diseases such as
reperfusion injury, autoimmune diseases, and other diseases of increased
complement activation are presented as well as methods of increasing the
effectiveness of gene therapeutics and other therapeutics.Claims:
1. A modified human complement C3 protein, comprising a substitution of a
portion of a human C3 protein, with a corresponding portion of a Cobra
Venom Factor protein of a sequence substantially related thereto.
2. The modified C3 protein of claim 1, wherein the substituted portion of the CVF is within the alpha chain of C3.
3. The modified C3 protein of claim 2, wherein the substituted portion of the CVF is a C-terminal portion of the alpha chain of C3.
4. The modified C3 protein of claim 3, wherein the substituted C-terminal portion includes amino acid 1663 of the human C3 protein.
5. The modified C3 protein of claim 3, wherein the substituted C-terminal portion is an internal portion that does not extend through the entire C-terminus of the human C3 protein.
6. The modified C3 protein of claim 1, wherein the modified protein has substantially the same number of amino acid residues as an unmodified human C3 protein.
7. The modified C3 protein of claim 1, wherein the substitution comprises any positions within amino acid positions 700-1663 of the human C3 protein.
8. The modified C3 protein of claim 1, wherein the substitution comprises a first position and a last position, wherein the first position is selected from the group consisting of 749, 874, 936, ', 1264, 1348, 1496, 1504, and 1550, and wherein the last position is selected from the group consisting of 784, 921, 970 1324, 1550, 1617, and 1663.
9. The modified C3 protein of claim 8, wherein the substitution is selected from the group consisting of amino acids: 1550-1663, 1504-1663, 1348-1663, 1550-1617, 1504-1617, 1496-1663, 1348-1617, 1496-1617, 1264-1324, 749-784, 874-921, 994-1663, 994-1550 and 936-970.
10. The modified C3 protein of claim 1, wherein the modified C3 protein has an affinity for factor B and supports formation of an active convertase.
11. The modified C3 protein of claim 10, wherein the convertase has an intrinsic half-life of at least about 15 minutes at 37.degree. C.
12. The modified C3 protein of claim 1, wherein the modified protein has an additional 1 to 19 amino acids at the N-terminus that are not encoded by C3 or CVF.
13. The modified C3 protein of claim 1, wherein the modified protein is substantially non-immunogenic.
14. A method for depleting complement, comprising administering the modified C3 protein of claim 1 to a patient in an amount effective for the depletion of complement.
15. A method for increasing the efficiency and/or effectiveness of gene therapy, comprising: delivering the modified C3 protein of claim 1 in an amount sufficient to deplete complement; providing the gene therapy; and observing an enhanced result therefrom.
16. A method of increasing delivery of a therapeutic or diagnostic agent, comprising: delivering the modified C3 protein of claim 1 in an amount sufficient to increase blood flow; and providing the therapeutic or diagnostic agent.
17. The method of any of claims 14-16, further comprising chemically linking the modified C3 protein to an antibody with an affinity for a specific tissue prior to the delivering step.
18. A method of treating a condition or disease associated with undesirable complement activation, comprising administering the modified C3 protein in an amount sufficient to deplete complement.
19. The method of claim 18, wherein the condition or disease is selected from the group consisting of asthma, systemic lupus erythematosus, glomerulonephritis, rheumatoid arthritis, Alzheimer's disease, multiple sclerosis, myocardial ischemia, reperfusion, sepsis, hyperacute rejection, transplant rejection, cardiopulmonary bypass, myocardial infarction, angioplasty, nephritis, dermatomyositis, pemphigoid, spinal cord injury and Parkinson's disease
20. A method of selecting a modified C3 protein, comprising: characterizing at least one property of the modified C3 protein to form a function profile of the modified protein; and matching the function profile with a disease or condition to be treated.
21. A nucleic acid sequence encoding the modified C3 protein of claim 1.
22. A composition comprising the modified human complement C3 protein of claim 1 and a pharmaceutically acceptable carrier.
23. A expression system expressing the modified C3 protein of claim 1.
Description:
RELATED APPLICATIONS
[0001]This International application claims priority to U.S. Provisional Applications 60/567,069, filed Apr. 30, 2004, 60/653,247, filed Feb. 14, 2005, and 60/667,352, filed Mar. 30, 2005, each of which is incorporated by reference in its entirety.
FIELD OF THE INVENTION
[0002]The invention relates generally to chimeric derivatives of Human Complement C3 having a substitution of a portion of a human C3 protein with a corresponding portion of a Cobra Venom Factor (CVF) protein. Preferably, a portion of the alpha chain of C3 is substituted with the corresponding portion of CVF.
BACKGROUND OF THE INVENTION
[0003]The third component of complement, C3, plays a pivotal role in both the classical and alternative pathways of complement activation, and many of the physiologic C3 activation products have important functions in the immune response and host defense. In the alternative pathway, the activated form of C3, C3b, is a structural subunit of the C3 convertase. This bimolecular enzyme consists of C3b and Bb, the activated form of factor B. This enzyme is formed by the binding of C3b to factor B that is subsequently cleaved by factor D, resulting in the formation of the C3 convertase, C3b,Bb, and the release of the activation peptide Ba. The C3 convertase activates C3 by cleaving the molecule into C3b and the anaphylatoxin, C3a. The C3b molecule will bind to a cell or particle in close proximity to the C3 convertase. Eventually, the bound C3b will allow for the activation of C5 into C5b and the anaphylatoxin, C5a. C5 activation occurs by the same C3b,Bb enzyme that can cleave C5 when it is bound to an additional C3b molecule to produce a trimolecular complex composed of (C3b)2,Bb. This C5-cleaving trimolecular enzyme is called C5 convertase. Inasmuch as the activation of both C3 and C5 occurs at the identical active site in the Bb subunit, the enzyme is also called C3/C5 convertase; and only one EC number has been assigned (EC 3.4.21.47).
[0004]Cobra venom contains a structural and functional analog of C3 called cobra venom factor (CVF). This molecule can bind factor B in human and mammalian serum to form the complex, CVF,B, which is also cleaved by factor D into the bimolecular enzyme CVF,Bb and Ba. The bimolecular complex CVF,Bb is a C3/C5 convertase that activates C3 and C5 analogously to the C3/C5 convertase formed with C3b. Although the two C3/C5 convertases, C3b,Bb and CVF,Bb, share the same molecular architecture, the active site-bearing Bb subunit, and the substrate specificity, the two enzymes exhibit significant functional differences. The CVF,Bb enzyme is physiochemically far more stable than C3b,Bb, it is resistant to inactivation by the regulatory proteins factors H and I, it exhibits different kinetic properties, and it does not require additional C3b for C5 cleavage.
[0005]CVF and mammalian C3 have been shown to exhibit several structural similarities including immunologic cross-reactivity, amino acid composition, circular dichroism spectra, secondary structure, electron microscopic ultrastructure, and amino acid sequence. Nevertheless, significant structural differences exist between the two molecules. Whereas C3 is a two-chain molecule with an apparent molecular mass, dependent on the species, of 170 to 190 kDa, CVF is a three-chain molecule with an apparent molecular mass of 149 kDa that resembles C3c, one of the physiologic activation products of C3. Another significant structural difference between C3 and CVF lies in their glycosylation: CVF has a 7.4% (w/w) carbohydrate content consisting mainly of N-linked complex-type chains with unusual α-galactosyl residues at the non-reducing termini. In contrast, human and rat C3 exhibit a lower extent of glycosylation with different structures of their oligosaccharide chains.
[0006]Whereas CVF,Bb and C3b,Bb are both C3/C5 convertases, they exhibit important differences. The CVF-containing enzyme is far more stable than the C3-containing enzyme. Both convertases will spontaneously decay into their two respective subunits. However, the intrinsic half-life (stability) of the CVF-containing convertase is approximately 7 hours at 37° C., several hundred times longer than the C3-containing enzyme with an intrinsic half-life of approximately 1.5 minutes. Furthermore, the CVF-containing enzyme as well as free CVF are not subject to regulation by the complement regulatory proteins factors H and I. The combination of the long intrinsic half-life and the resistance to regulation of the CVF-containing enzymes allows CVF to continuously activate C3 and C5 (and subsequently other complement components), ultimately resulting in depletion of the serum complement activity.
[0007]Based on the involvement of the complement system in multiple diseases, including diseases of major prevalence, the last decade has seen the development of multiple anti-complementary agents to interfere with the unwanted complement activation process in these disease states. All complement-oriented drug development attempts are based on inhibiting the activation of complement, while CVF acts by depleting complement in serum. Of interest for the treatment of diseases of complement activation is a C3-type molecule which combines the non- or low immunogenicity of C3, with the complement-depleting function of CVF.
SUMMARY OF THE INVENTION
[0008]The following listing of embodiments is a nonlimiting statement of various aspects of the invention. Other aspects and variations will be evident in light of the entire disclosure.
[0009]Some embodiments include one or more modified human complement C3 proteins, which can have a substitution of a portion of a human C3 protein, with a corresponding portion of a Cobra Venom Factor protein of a sequence substantially related thereto. In some embodiments the substituted portion of the CVF can be within the alpha chain of C3. In other embodiments, the substituted portion of the CVF can be a C-terminal portion of the alpha chain of C3. In some embodiments, the substituted C-terminal portion can include amino acid 1663 of the human C3 protein. In some embodiments, the substituted C-terminal portion can be an internal portion that does not extend through the entire C-terminus of the human C3 protein. In further embodiments, the modified protein can have substantially the same number of amino acid residues as an unmodified human C3 protein. In some embodiments, the substitution can include any positions within amino acid positions 700-1663 of the human C3 protein. Other embodiments are human complement C3 proteins, which can have a substitution of a portion of a human C3 protein, with a corresponding portion of a Cobra Venom Factor protein of a sequence substantially related thereto and which can have at least two substitutions. In some embodiments, the substitution has a selected beginning position and a selected last position; in some such embodiments, the beginning position can be, for example, 749, 874, 936, 994, 1264, 1348, 1496, 1504, 1550, and the like; the last position can be, for example, 784, 921, 970 1324, 1550, 1617, 1663, and the like. In preferred embodiments, the one or more substitutions can include any of amino acids: 1550-1663, 1504-1663, 1348-1663, 1550-1617, 1504-1617, 1496-1663, 1348-1617, 1496-1617, 1264-1324, 749-784, 874-921, 994-1663, 994-1550 and 936-970. In some embodiments, the substituted portion of CVF can be within the beta chain of C3.
[0010]In some embodiments, the modified C3 protein can have an affinity for factor B and can support formation of a convertase. In some embodiments, the resulting convertase can cleave C3 and not C5. In further embodiments, the convertase can have an intrinsic half-life between about 1.5 minutes and about 7 hours at 37° C. In some embodiments, the resulting convertase can have an intrinsic half-life of at least about 7 hours at 37° C.
[0011]In some embodiments, the modified C3 protein can be expressed as a single chain protein. In some embodiments, the modified C3 protein can be cleaved into at least two chains in a form that resembles C3. In further embodiments, the modified C3 protein can be cleaved to release a C3a portion therefrom. In some embodiments, the modified protein can have an additional 1 to about 19 amino acids at the N-terminus that are not encoded by C3 or CVF. In some embodiments, the modified protein can include a non-C3 signal peptide, such as a Drosophila Bip signal sequence. In some embodiments, the modified C3 protein can have modified affinity for factor B and/or factor D. In some embodiments, the modified protein can show partial or complete resistance to Factor H and/or Factor I. In some embodiments, the modified C3 protein can be substantially non-immunogenic.
[0012]Other embodiments can include a method for depleting complement by administering a modified C3 protein to a patient in an amount effective for the depletion of complement. In some embodiments, the administration can be local. In further embodiments, the local administration can be into an organ, subcutaneously, into a cavity, or into a tissue. In other embodiments, the local administration can employ a targeting function capable of concentrating the modified C3 protein in a desired location. In further embodiments, the targeting function can include using an antibody conjugated to the modified C3 protein. In some embodiments, the administration can be a systemic administration, such as intravenous or intraperitoneal.
[0013]Further embodiments can be methods for avoiding or ameliorating reperfusion injury in a patient by delivering a modified C3 protein to the patient, sufficient to deplete complement; and permitting reperfusion in the patient. In some embodiments, the delivering step can include injecting the modified C3 protein into an artery. In other embodiments, the delivering step can include a local delivery of the modified C3 protein. In other embodiments, the delivering step can include a systemic delivery of the modified C3 protein. In some embodiments, reperfusion can include opening a blocked artery. In some embodiments, the reperfusion can occur in connection with transplantation of an organ.
[0014]Some embodiments can include methods for increasing the efficiency and/or effectiveness of gene therapy by delivering a modified C3 protein in an amount sufficient to deplete complement, providing the gene therapy; and observing an enhanced result therefrom.
[0015]Some embodiments can include methods of increasing delivery of a therapeutic or diagnostic agent by delivering a modified C3 protein sufficient to increase blood flow; and providing the therapeutic or diagnostic agent. In some embodiments, the method can include chemically linking the modified C3 protein to an antibody with an affinity for a specific tissue prior to the delivering step. In some embodiments, the antibody can be attached to the modified C3 by recombinant DNA technology. In some embodiments the antibody can be a monoclonal antibody.
[0016]Some embodiments include methods of treating an autoimmune disease, comprising administering the modified C3 protein sufficient to deplete complement. In some embodiments, the administration can be episodic and corresponds to periods of at least one elevated disease symptom. In some embodiments, the autoimmune disease can be any of asthma, systemic lupus erythematosus, glomerulonephritis, rheumatoid arthritis, Alzheimer's disease, multiple sclerosis, myocardial ischemia, reperfusion, sepsis, hyperacute rejection, transplant rejection, cardiopulmonary bypass, myocardial infarction, angioplasty, nephritis, dermatomyositis, pemphigoid, spinal cord injury and Parkinson's disease.
[0017]Some embodiments include methods of mimicking the properties of CVF in a human protein, by, for example, substituting a portion of a human complement C3 protein with the corresponding portion of a Cobra Venom Factor (CVF) protein. In some embodiments the portion can be within the alpha chain of C3. In other embodiments, the portion can be a C-terminal portion of the alpha chain of C3. In some embodiments, the C-terminal portion can include amino acid 1663 of the human C3 protein. In some embodiments, the substituted C-terminal portion can be an internal portion that does not extend through the entire C-terminus of the human C3 protein.
[0018]Other embodiments include methods of selecting a modified C3 protein, by characterizing at least one property of the modified C3 protein to form a function profile of the modified protein; and matching the function profile with a disease or condition to be treated. In some embodiments, at least one property can be selected from the group consisting of: convertase activity, convertase formation, convertase stability, susceptibility to Factor H, susceptibility to Factor 1, ability to cleave C3, and ability to cleave C5. In some embodiments, the selected C3 protein participates in formation of a convertase adapted for treatment of a chronic condition. In some embodiments, the adaptation can include any of, for example, long plasma half-life, high stability, resistance to Factor H, resistance to Factor I, and the like. In some embodiments, the convertase can be adapted for treatment of a reperfusion injury. In other embodiments, the adaptation can be any of, for example, high convertase activity, resistance to Factor H, resistance to Factor I and the like.
[0019]Some embodiments include a nucleic acid sequence encoding a modified C3 protein, and/or a vector including the nucleic acid and/or a host cell containing the vector. In some embodiments, the host cell can be any of: a Drosophila S2 cell, an Sf9 cell, a CHO cell, a COS-7 cell, a HiFive cell, a yeast cell, a BHK cell, an HEK293 cell, and an E. coli cell.
[0020]Some embodiments include a composition that can include the modified human complement C3 protein and a pharmaceutically acceptable carrier and/or the nucleic acid.
[0021]Some embodiments include an expression system expressing the modified C3 protein. In some embodiment, the expression system include a cell selected from the group consisting of: a Drosophila S2 cell, an Sf9 cell, a CHO cell, a COS-7 cell, a HiFive cell, a yeast cell, a BHK cell, an HEK293 cell, and an E. coli cell.
BRIEF DESCRIPTION OF THE DRAWINGS
[0022]FIG. 1 depicts the chain structures of C3 and CVF with shaded portions present in the mature proteins.
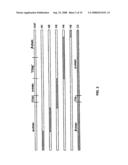
[0023]FIG. 2 shows a map of the original CVF/cobra C3 hybrid proteins, showing the region of CVF that was substituted with cobra C3 sequences in each of the five hybrid proteins.
[0024]FIGS. 3A-3G show the cDNA and derived amino acid sequence of CVF1 (SEQ ID NOs:3 and 4). The NH2-- and C-termini of the α-, γ-, and β-chains, functionally important regions, and known ligand binding sites are indicated. Amino acid residue numbering starts at the NH2-terminus of the pro-CVF1 molecule;
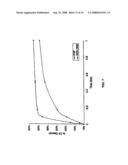
[0025]FIG. 4 shows results of a complement depletion assay in human serum of the three modified human C3 proteins (HC3-1348, HC3-1504, and HC3-1550) as compared to CVF.
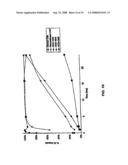
[0026]FIG. 5 shows results of an assay to measure the ability of the modified human C3 proteins (HC3-1348, HC3-1504, and HC3-1550) to activate factor B and to form a C3/C5 convertase as compared to CVF and C3b.
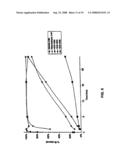
[0027]FIG. 6 shows results of an assay to measure the activity of C3/C5 convertase to activate C3 for the three modified human C3 proteins (HC3-1348, HC3-1504, and HC3-1550) as compared to CVF.
[0028]FIG. 7 shows results of an assay to measure C3 cleavage (using 20% of the amount of convertase used in the experiment in FIG. 6) of the modified human C3 protein HC3-1348 as compared to CVF.
[0029]FIG. 8 is a summary of the results for the activity measurements of the three modified human C3 proteins: HC3-1348, HC3-1504, and HC3-1550 as compared to CVF and Human C3b.
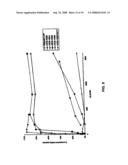
[0030]FIG. 9 is a graph showing the results of complement depletion of Natural CVF and a variety of human C3 proteins.
[0031]FIG. 10 is a graph showing the results of Cleavage of factor B by C3b, Recombinant CVF and a variety of modified human C3 proteins.
[0032]FIG. 11 is a graph showing the results of C3 cleavage of Natural CVF and a variety of modified human C3 proteins. The inset graph is of the same reaction performed using 20% of the convertase in the main graph.
[0033]FIG. 12 is a graph showing the results of cleavage of selected hybrid proteins by factors H and I.
[0034]FIG. 13 is a picture of a gel showing the results of C5 conversion by Human C3/CVF hybrid proteins.
DETAILED DESCRIPTION OF THE PREFERRED EMBODIMENTS
[0035]The replacement of human Complement C3 sequences with CVF sequences representing key structural requirements for CVF specific functions allows the creation of C3 derivatives with CVF-like functions. Preferred embodiments of the invention provide human C3 derivatives that exhibit the CVF-specific function of depleting complement by forming a stable convertase for use as a novel therapeutic agent to deplete complement in clinical situations where complement activation is part of the pathogenesis. Because the structural changes in human C3 caused by CVF-specific sequences are minimal, the modified C3 molecule will exhibit significantly reduced or even absent immunogenicity.
[0036]A number of modified human complement C3 proteins (C3) are disclosed that have a substitution of a portion of a human C3 protein, with a corresponding portion of a Cobra Venom Factor protein (CVF) which results in a human C3 protein with CVF functions, but with substantially reduced immunogenicity. The C3 protein may be the pro-protein (a single chain protein) or the cleaved protein by removal of four arginine residues. An additional two chain form may be formed by the removal of the C3a portion. Advantageously, the C3 protein can be manipulated to contain at least one of the following CVF functions: increased ability to form the C3 convertase, increased stability of the C3 convertase, increased resistance to the actions of factors H and/or I, increased activity to cleave C3 and C5, and increased plasma half-life. In one embodiment, the C-terminal portion of the alpha chain of C3 is substituted with a corresponding portion of a CVF protein. A wide variety of specific regions and specific chimerae are disclosed herein. Further, a method of identifying and/or selecting a modified C3 protein is disclosed which involves characterizing the properties of the modified C3 protein to form a function profile of the modified protein, and matching the function profile with a disease or condition to be treated. The function profile may include one or more of the following functions: ability to form a convertase, susceptibility to H and/or I, the ability to cleave C3, the ability to cleave C5, the relative activity of the convertase, the stability of the convertase, and the plasma half-life. The modified C3 protein, also referred to as a chimeric C3 protein, a C3/CVF chimera, or a C3/CVF hybrid protein, can be used to treat a variety of diseases and/or conditions that result from local or systemic complement activation. These diseases include, but are not limited to: reperfusion injury, autoimmune diseases such as rheumatoid arthritis and lupus, and myasthenia gravis and other diseases that result from antibodies that recognize and direct an immune reaction against proteins or structures of the human body.
The Complement System and Disease
[0037]The complement system is a component of the vertebrate immune system which is involved in maintaining health through its roles in host defense and immune response. However, complement activation is also involved in the pathogenesis of a multitude of diseases. Examples of these diseases include autoimmune hemolytic anemias, rheumatoid arthritis and other immune complex diseases, and reperfusion injuries in which tissue damage occurs after blood flow has been temporarily suspended. Examples of reperfusion injuries are tissue damage after the reopening of blocked vessels (e.g. heart attack, stroke) and reperfusion with a recipient's blood after organ transplantation. Based on the involvement of the complement system in multiple diseases, including diseases of major prevalence, the last decade has seen the development of multiple anti-complementary agents to interfere with the unwanted complement activation process in these disease states. All of these drug development attempts are based on inhibiting the activation of complement.
Complement Protein C3, Cobra Venom Factor CVF, Similarities and Differences
[0038]The third component of complement, C3, plays a pivotal role in both the classical and alternative pathways of complement activation, and many of the physiological C3 activation products have important functions in the immune response and host defense (for review see Muller-Eberhard, H. J. (1988) "Molecular Organization and Function of the Complement System," Ann. Rev. Biochem., 57:321-347, herein incorporated by reference in its entirety). Human C3 is a two-chain glycoprotein with a molecular weight of approximately 185,000. It is synthesized as single-chain pre-pro-C3 which undergoes subsequent processing by removal of four arginine residues between the β- and α-chains. The primary sequence of human C3 is known from molecular cloning. Full or partial sequence information of C3 from other mammalian species as well as non-mammalian species is available including mouse, rat, guinea pig, chicken, cobra, Xenopus, and lamprey. The human C3 gene is 42 kb in length and includes 41 exons, ranging in size from 52 to 213 bp.
[0039]The major activation product of C3 is C3b. C3b plays a central role in the formation of the alternative pathway C3/C5 convertase. The formation of this enzyme requires initial binding of C3b to Factor B. The weak complex C3b,B is subsequently cleaved by Factor D in the presence of Mg2+, resulting in the enzymatically active C3/C5 convertase C3b,Bb and in release of the activation peptide Ba. The C3b,Bb enzyme is very labile, exhibiting spontaneous decay-dissociation into the two subunits C3b and Bb with an intrinsic half life of 1.5 minutes at 37° C. The C3b,Bb enzyme is stabilized by properdin. The C3/C5 convertase cleaves C3 and C5 by hydrolyzing a single peptide bond in the α-chains of the two substrates. In order for the enzyme to cleave C5, C5 has to be bound to another C3b molecule bound to the convertase. In addition to the fast spontaneous decay-dissociation, the C3b,Bb enzyme is subject to stringent control. The enzyme is disassembled by Factor H, and C3b is inactivated by the combined action of factors H and I. In the presence of Factor H, Factor I cleaves the α'-chain of C3b at two cleavage sites. The resulting C3b derivative, called iC3b, can no longer form a convertase with Factor B. Factor I can cleave the α'-chain at a third site, which causes the generation of the two C3 fragments C3c and C3dg. For the third cleavage by Factor I, the C3b receptor, CR1 serves as co-factor.
[0040]An unusual structural property of C3 is the presence of an intramolecular thioester in the α-chain. Upon activation of C3 to C3b, the thioester becomes highly reactive and is responsible for the covalent attachment of C3b to cellular and other particular targets. The structural change which accompanies cleavage of the thioester allows the subsequent binding of Factor B and its activation.
[0041]The thioester in C3 undergoes slow spontaneous hydrolysis, resulting in the formation of a form of C3 called iC3 or C3(H2O). iC3 assumes C3b-like functions and can form a fluid-phase convertase with Factors B and D in serum. The resulting convertase iC3,Bb is similarly labile as the C3b,Bb convertase and subject to control by factors H and I. However, spontaneous hydrolysis of the thioester and the ensuing low grade activation of C3 by the iC3,Bb convertase is believed to be responsible for the initial deposition of C3b on target cells or particles, leading to activation of the alternative pathway on so-called activator surfaces.
[0042]C3 is a highly unusual multi-functional protein. The protein including its various activation products specifically interacts with approximately twenty different plasma proteins or cell surface receptors. This multifunctionality has spurred significant interest in a detailed structure/function analysis of the molecule. For some ligands of C3, including Factor H, properdin, Factor B, and the complement receptors CR1, CR2, CR3, and C3a receptor binding sites have been proposed or assigned to more or less defined regions of the C3 polypeptide.
[0043]Cobra venom contains a structural and functional analog of C3 called cobra venom factor (CVF). Functionally, CVF resembles C3b in that it can bind Factor B in human serum and virtually all vertebrate sera to form a weak complex CVF,B, which is subsequently cleaved by Factor D in the presence of Mg2+ into the bimolecular enzyme CVF,Bb and Ba. The bimolecular complex CVF,Bb is a C3/C5 convertase that activates C3 and C5 analogously to the C3/C5 convertase formed with C3b.
[0044]CVF is a three-chain glycoprotein with a molecular mass of approximately 150,000 Da. CVF and mammalian C3 have been shown to exhibit several structural similarities including immunological cross-reactivity, amino acid composition, circular dichroism spectra and secondary structure, and electron microscopic ultrastructure. Initial N-terminal amino acid sequence comparisons have demonstrated sequence homology with C3 and have led to the suggestion that CVF structurally resembles C3c. The structural homology between CVF and C3 and the chain relationships were confirmed by the molecular cloning of CVF, which revealed an overall similarity at the protein level of approximately 70 percent to mammalian C3s and over 90 percent when compared to cobra C3.
[0045]Despite these functional and structural similarities between CVF and C3, the two molecules and the resulting convertases exhibit important functional differences: [0046]1. Both enzymes exhibit spontaneous decay-dissociation into the respective subunits which abolishes the enzymatic activity. Whereas the C3b,Bb enzyme is very short-lived and decays with an intrinsic half-life of 1.5 minutes at 37° C., the CVF,Bb enzyme is orders of magnitude more stable, decaying with an intrinsic half-life of approximately seven hours. [0047]2. The C3b,Bb enzyme is subject to regulation by factors H and I. In contrast, CVF,Bb and CVF are completely resistant to the regulatory actions of these two proteins. [0048]3. The C3b,Bb enzyme generated during complement activation is surface bound. In contrast, the CVF,Bb enzyme is a fluid-phase enzyme (like iC3,Bb). [0049]4. Another functional difference between C3b,Bb and CVF,Bb lies in the C5 convertase activities. In order for C5 to be cleaved by a C5 convertase, it needs to be bound to either C3b or CVF. However, for C5 cleavage to occur by the C3b,Bb enzyme, C5 has to be bound to a different C3b molecule than the one that is part of the C3b,Bb enzyme. In contrast, C5 is bound by the same CVF molecule that carries the Bb catalytic subunit. This property of the CVF,Bb enzyme to bind C5 is probably the reason for its ability to exhibit fluid phase C5 convertase activity, whereas the C5 convertase activity of the C3b,Bb enzyme is confined to the surface of a particle. [0050]5. Both enzymes have been shown to differ somewhat in their kinetics of C3 hydrolysis. Based on the kcat/Km, the catalytic efficiency is approximately eight-fold greater for C3b,Bb compared to CVF,Bb.
[0051]In terms of functional consequences, the two most significant differences between CVF,Bb and C3b,Bb are the intrinsic stability of the CVF,Bb enzyme and its resistance to the regulatory proteins factors H and I. Once the CVF,Bb enzyme has formed, it will continue to activate C3 and C5, leading to complement consumption. Ever since it was demonstrated over 30 years ago that CVF can be administered safely to laboratory animals in order to deplete their plasma complement, CVF has become an important investigational tool to study the various biological functions of complement in immune response, host defense, and pathogenesis of disease by comparing normal (complement-sufficient) animals with CVF-treated (complement-depleted) animals.
C3/CVF Derivatives.
[0052]CVF is a complement inhibitor that acts through a mechanism of exhaustive activation which subsequently leads to depletion. As a matter of fact, CVF is frequently used as the standard to evaluate the anti-complement activity of other drugs. Whereas CVF exhibits this powerful anti-complement activity, it is not suitable for human application because of its immunogenicity. For this reason it is desirable to prepare a substantially non-immunogenic CVF by taking advantage of the extensive structural similarity between CVF and C3, and to generate human C3 derivatives with the desired complement-depleting function of CVF by substituting the functionally important regions of the CVF molecule into human C3 by recombinant means. This has been accomplished as described herein.
[0053]A number of human C3 derivatives have been produced and/or designed, in which portions of the C3 sequence were replaced with homologous CVF sequences, mainly sequences in the CVF beta chain. CVF and C3 have been successfully expressed by recombinant means in eukaryotic expression systems. The alpha chain region of C3, and preferably the C-terminal portion of this chain, was chosen based on previous work in which five hybrid proteins were constructed between CVF and cobra C3 (because of its greater similarity to CVF than human C3), collectively spanning the entire CVF sequence (Mol. Immunol. 40:199 (2003), incorporated herein by reference in its entirety). The other line of work involved the limited proteolysis of the CVF protein (Mol. Immunol. 30, Suppl. 1, 113 (1993) U.S. Pat. No. 5,174,344, incorporated herein by reference in its entirety). Using the results of this previous work, three human C3 preferred derivatives were created by replacing amino acid residues 1550-1663, 1504-1663, and 1348-1663, respectively, with the homologous sequences from CVF. The C3 derivatives are designated by the first amino acid residue and if the last amino acid residue is not indicated, it is understood to be 1663. If the endpoint of the CVF insertion is before the C-terminus of the protein, it is designated with the location (for example: HC3-1550/1617). The C3 derivative which was created by replacing amino acids residues 1348-1663 was formerly referred to as 1325-1663, but substitution mapping has subsequently shown that the substitution was at 1348 rather than 1325. The three human C3 derivatives are referred to as HC3-1550, HC3-1504, and HC3-1348. The three human C3 derivatives were shown to exhibit the desired CVF activity: all three proteins are able to form an active C3/C5 convertase. This is demonstrated by the ability of the three proteins to support the activation of factor B, and by the ability of the three resulting convertases to cleave C3. All three proteins form stable convertases, although the intrinsic stability and at least one (HC3-1550) exhibits a lower intrinsic stability compared to CVF. Unexpected was the observation that two of the three proteins first tested (HC3-1550 and HC3-1348) were actually able to decomplement guinea pig serum, although this activity was clearly less than that of CVF, and four of five proteins were able to deplete complement in human serum, with HC3-1348 and HC3-1496 able to decomplement human sera almost as well as CVF. HC3-1550/1617 was not able to deplete complement in human serum. The ability to at least partially deplete complement indicates that the C3 derivative does not only form a stable convertase but is at least partially resistant to the regulatory proteins factors H and/or 1, thereby exhibiting CVF-like activity. The human C3 derivative HC3-1550 differs from human C3 in less than 4% of the amino acid residues, which greatly reduces or eliminates its predicted immunogenicity compared to CVF. Various C3 hybrids can be preferred, based upon their various characteristics, depending upon the disease to be treated. For example, for the treatment of chronic diseases, the objective is to provide a C3 hybrid having extremely low or no immunogenicity and high convertase stability, facilitating its persistence in the body of the patient suffering from the chronic disease. In such a situation, the other characteristics, such as the activity of the convertase, are of comparatively less importance. In contrast, for treatment to avoid complement-associated reperfusion injury, a high convertase activity is of greater importance than convertase stability. Thus, a multiplicity of different hybrids, each having a particular array of properties, permits selection of a preferred hybrid for treatment of a given condition that can be treated by complement activation and/or complement depletion.
[0054]In one embodiment, the present invention relates to modified complement C3 proteins that exhibit at least one of the following CVF or CVF,Bb qualities: decaying with an intrinsic half-life of longer than 1.5 minutes, increased resistance to the regulatory actions of factors H and/or I, fluid-phase C3 convertase and fluid-phase C5 convertase activity. In addition to these factors, in some embodiments the catalytic efficiency may be reduced relative to C3b,Bb, since the catalytic efficiency is approximately eight-fold greater for C3b,Bb compared to CVF,Bb. In other embodiments the catalytic efficiency is not reduced or is elevated in comparison to C3b,Bb. Although many preferred C3 hybrids have little or no immunogenicity, other embodiments, which are nevertheless suitable for many applications, may display detectable to moderate immunogenicity in some cases.
[0055]In some embodiments, the intrinsic half-life of the convertase formed with the modified C3 protein is greater than 1.5 minutes, preferably greater than 10 minutes. In further embodiments, the intrinsic half-life can fall generally between that of the CVF-containing convertase (7 hours or longer) and that of C3 (1.5 minutes), including but not limited to about: 2 minutes, 10 minutes, 20 minutes, 30 minutes, 40 minutes, 50 minutes, 60 minutes, 90 minutes, 2 hours, 2.5 hours, 3 hours, 3.5 hours, 4 hours, 4.5 hours, 5 hours, 5.5 hours, 6 hours, 6.5 hours and 7 hours, or more. Modified C3 proteins with short convertase intrinsic half-lives and/or short plasma half-lives will be useful for some applications, while C3 proteins with long convertase intrinsic half-lives and/or long plasma half-lives will be useful for different applications.
[0056]In further embodiments, the resistance of the C3 hybrid to factors H and/or I is greater than that for the unmodified C3 and is in some embodiments as good as the resistance of CVF. However, in some embodiments, further modification in other parts of the molecule can be necessary to achieve resistance to factors H and/or 1.
[0057]While many embodiments described herein are directed to specific substitutions of one or more discrete regions of C3 with corresponding regions of CVF, other embodiments include substitutions employing a sequence that is substantially related, but not identical, to a CVF sequence. That is, there are positions within a CVF region selected for substitution, in which changes to one or more amino acids can be made without a loss of any desirable feature or function of the selected CVF region, and in some cases such changes can confer an enhanced feature or function. All such changes are considered to be embodiments of the invention.
[0058]In a further embodiment, the catalytic activity of convertase containing the modified C3 protein is in some embodiments at least 50% that of the convertase containing CVF, and may be greater than that of the convertase containing unmodified C3. In a further embodiment, the catalytic activity is 60%, 70%, 80% 90% or 100% that of the CVF convertase. Both enzymes have been shown to differ somewhat in their kinetics of C3 hydrolysis. Thus, in many embodiments, convertases containing the modified C3 can have a catalytic activity that falls between the two, or that exceeds the activity of the convertase containing unmodified C3. Thus, in some embodiments, such activity of the convertase containing the modified C3 can be from 10% to 1000%, or more, that of the convertase containing CVF, including but not limited to 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, and 110%, 135%, 150%, 200%, 300%, 400%, 500%, 750%, 1000% and more.
[0059]Based on the kcat/Km, the catalytic efficiency is approximately eight-fold greater for C3b,Bb compared to CVF,Bb when cleaving C3. Thus, in some embodiments, the catalytic efficiency of convertase containing the modified C3 protein is in some embodiments at least 50% that of the convertase containing CVF, and may be greater than that of the convertase containing unmodified C3b. Both enzymes have been shown to differ somewhat in their kinetics of C3 hydrolysis. Thus, in many embodiments, convertases containing the modified C3 can have a catalytic efficiency that falls between the two, or that exceeds the efficiency of the convertase containing unmodified C3. Thus, in some embodiments, such efficiency of the convertase containing the modified C3 can be from 10% to 1000%, or more, that of the convertase containing CVF, including but not limited to 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, and 110%, 135%, 150%, 200%, 300%, 400%, 500%, 750%, 1000% and more.
[0060]In further embodiments, the C5 cleaving activity of the modified C3 proteins is increased. The C5 cleaving activity can be from about 10% to 400% of the activity of CVF or C3, including but not limited to: 15%, 20%, 25%, 30%, 35%, 40%, 45%, 50%, 55%, 60%, 65%, 70%, 75%, 80%, 85%, 90%, 95%, 99%, 100%, 150%, 200%, 250%, 300%, 350%, and 375%.
[0061]In further embodiments, the binding of the modified C3 proteins to Factor B and/or its subsequent cleavage by factor D may be reduced. However, as long as at least a functional amount of the catalytic activity and the complement depleting activity remain, the modified C3 protein is useful.
[0062]In further embodiments, convertases having the modified C3 proteins exhibit substantially the same complement-activating activity of those containing natural CVF. By the term "exhibit substantially the same complement-activating activity of natural CVF it is meant that the C3 derivatives of the present invention have from 0.1 to 97%, preferably from 50 to 97%, preferable from 80 to 97% of the level of the complement activating activity of natural CVF as measured by the method of Cochran et al. ((1970) J. Immunol. 105(1)), 55-69, herein incorporated by reference in its entirety).
[0063]In some embodiments, the modified complement C3 proteins are C3 molecules in which some or all of the C-terminal region has been replaced with the corresponding region of CVF. In some embodiments, only the C-terminal portion of the alpha chain of C3 is replaced with the corresponding region of CVF. In some embodiments some or all of amino acids 700-1663 are replaced, including but not limited to regions of from 20 to about 1000 amino acids, including but not limited to: 30, 40, 50, 60, 70, 80, 90, 100, 110, 120, 130, 150, 175, 180, 190, 200, 250, 275, 300, 350, 375, 400, 450, 475, 500, 550, 575, 600, 650, 675, 700, 750, 775, 800, 850, 875, 900, 950 975, 1000. In some embodiments, the beta chain of C3 is intact, that is, is the same as in natural C3. However, in some embodiments, in addition to the substitutions in the alpha chain of C3, other substitutions can be made to provide such functions as increased stability (for example, the sites which relate to Factor H binding and Factor I cleavage can be mutated or substituted). Specific substitutions include but are not limited to amino acids 1550-1663, 1504-1663, 1348-1663, 1550-1617, 1504-1617, 1470-1663, 1348-1617, 1470-1617, 1264-1324, 1348-1386, 749-784, 874-921, 1496-1663, 1496-1617 and 936-970. In some embodiments, still smaller substitutions in these areas can correspond to smaller regions that result in a C3 with the desired CVF functions or qualities.
[0064]The immunogenicity of preferred embodiments remains low or absent--comparable to that of C3. In some embodiments, the modified C3 protein is substantially non-immunogenic. Substantially non-immunogenic means that the protein can still exhibit complement activating function when injected into a human patient. Further, the modified C3 protein may be as non-immunogenic as C3 or may be from about 75% non-immunogenic to about 100% non-immunogenic, including but not limited to 80%, 85%, 90%, 95%, and 99%.
Methods of Treatment Systemically and Locally
[0065]The modified human complement C3 proteins produced by the methods disclosed herein can be used to deplete complement locally or systemically as follows:
[0066]Local treatment may be effected in a number of ways to produce a result of depletion of complement or activation of complement, depending on the desired effect. In one embodiment, local depletion is effected when the modified C3 proteins are administered locally to an organ, tissue, cavity, or intradermally. This results in a temporary and complete depletion of complement in the area. Local depletion or activation may also be effected using an insulin-type pump that produces an intermittent or constant flow of the modified C3 protein to a selected site. Alternatively, local activation of complement may employ a specific monoclonal antibody which, when chemically attached to the modified C3, can localize it to a specific tissue, a disease, or an infected cell to cause continuous activation of complement in that area. In other embodiments, the antibody can be attached to the modified C3 protein via recombinant DNA technology.
[0067]Systemic depletion is effected when the modified C3 proteins are administered systemically, for example, intravenously or intraperitoneally. This results in a temporary and complete depletion of complement systemically. This method can be used for reperfusion injury, coronary heart surgery, transplantation and or systemic disease, particularly during a flare-up or episodical activity. This is further discussed in the Examples provided herein.
[0068]Many of the most advantageous qualities of the modified human C3 proteins used in each of these cases and in specific disease states can vary considerably. For example, in those cases when an immediate, but temporary removal of complement is desired, a modified protein which has a shorter plasma half-life and/or stability, but high complement activation activity (high C3/C5 convertase activity) would be most suited. In the case where a chronic disease is being treated, a long plasma half-life and/or stability and a low or sluggish activity would be most suitable.
Convertase Activities/Functions
[0069]When producing and analyzing the best use for the modified C3 proteins herein, a number of functions and activities can be used for such an analysis, including but not limited to the following:
[0070]Affinity for Factor B--Differences in how efficiently the modified C3 proteins support the activation of factor B (cleavage into Ba and Bb) can be detected and taken advantage of. There are multiple factors beyond just affinity for factor B that go into supporting factor B activation, but once the convertase is formed, the subsequent key properties that are generally most important are stability, regulation by H and I, and activity of cleaving C3 and C5.
[0071]Stability--Both the C3b,Bb and CVF,Bb enzymes exhibit spontaneous decay-dissociation into the respective subunits which abolishes the enzymatic activity. Whereas the C3b,Bb enzyme is very short-lived and decays with an intrinsic half-life of 1.5 minutes at 37° C., the CVF,Bb enzyme is orders of magnitude more stable, decaying with an intrinsic half-life of approximately seven hours.
[0072]Regulation by Factors H and/or I. The C3b,Bb enzyme is subject to regulation by factors H and I. In contrast, CVF,Bb and CVF are completely resistant to the regulatory actions of these two proteins.
[0073]Activity of Cleaving C3--Both enzymes have been shown to differ somewhat in their kinetics of C3 hydrolysis. Based on the kcat/Km, the catalytic efficiency is approximately eight-fold greater for C3b,Bb compared to CVF,Bb.
[0074]Immunogenicity--Because the structural changes in human C3 caused by CVF-specific sequences are minimal, modified C3 can exhibit significantly reduced or even absent immunogenicity as compared with CVF. In addition, as is typical in species far removed from human, a transferase enzyme is produced in cobras that transfers an alpha-bonded galactose to the end of CVF oligosaccharide chains. Because humans do not produce this transferase, the alpha bonded galactose residues are seen as non-self and are highly immunogenic. Thus, anti-alpha-gal antibodies are produced that recognize the specific CHO moieties produced on CVF. Human C3 and CVF lack identity at 52% of their positions, and carbohydrate differences are even more pronounced. Thus, because preferred modified C3 proteins have less than 10%, and in some embodiments about 4%, of the C3 amino acids replaced with those of CVF, and also have no cobra-derived carbohydrate structures (specifically alpha-bonded galactoses on the glycans), the C3 hybrids of the invention are significantly less immunogenic. In some embodiments, the C3 hybrids of the preferred embodiments are significantly less immunogenic than CVF. In other embodiments, the C3 hybrids of the preferred embodiments are at least 50% less immunogenic than CVF. In some embodiments, the recombinant proteins are produced in either insect or mammalian cell lines, which will result in carbohydrate moieties that are either simple or very similar to what would be found in humans.
EXAMPLES
[0075]Five CVF/cobra C3 loss-of-function hybrid proteins were produced in which large portions of the CVF sequence were replaced by homologous portions of cobra C3. Preliminary characterization of these hybrid proteins showed that substitutions in the α-chain of CVF (β-chain of C3) with cobra C3 sequences did not appear to change the functional property of depleting serum complement activity, whereas replacing portions of the β- and γ-chains had a major effect. Hybrid proteins H4 and H5, in which CVF residues 978 through 1642 were replaced by cobra C3 sequence, exhibited a significant reduction in bystander lysis (a measure for C5 cleavage) activity in addition to the impaired ability to deplete serum complement. This suggested that the alpha chain of C3 (corresponding to the β- and β-chains of CVF) may be a major site for the differences between the activity of CVF and C3. Another line of experimental support strongly suggests the C-terminal portion of the CVF β-chain is important for CVF function comes from the experiments of limited proteolysis of CVF with chymotrypsin (Grunwald, et al. (1993) Mol. Immunol. 30, Supp. 1, 30, herein incorporated by reference in its entirety). Thus, a large number of modified human complement C3 proteins (C3) have been produce or identified that have a substitution of a portion of the human C3 protein, with a corresponding portion of the Cobra Venom Factor protein (CVF). These substitutions result in a human C3 protein with CVF-type functions, but with substantially less immunogenicity than CVF. The production and testing of these proteins is set forth below in Examples 1-3. Various uses and assays for these proteins are provided in Examples 4-10.
Example 1
Production of Human Complement C3/CVF Hybrid Proteins
[0076]The replacement of human C3 sequences with CVF sequences representing important structural features for CVF specific functions allows the creation of C3 derivatives with CVF-like functions. Thus, embodiments of the invention are directed to generation of human C3 derivatives that exhibit the CVF-specific function of depleting complement by forming a stable convertase for use as a novel therapeutic agent to deplete complement in clinical situations where complement activation is part of the pathogenesis. Because the structural changes in human C3 caused by CVF-specific sequences are minimal, it is clear that the modified C3 molecules can exhibit significantly reduced or event absent immunogenicity.
[0077]The human C3 molecules in Table 1 are engineered to contain specific CVF sequences so as to create human C3 derivatives with CVF functions. Some embodiments of the invention provide smaller substitutions in these areas, to define small regions that result in modified C3 proteins that can form a relatively stable convertase and that exhibit little or no immunogenicity.
TABLE-US-00001 TABLE 1 Exemplary Human C3/CVF clones HC3-1550 HC3-1504 HC3-1348 HC3-1550/1617 HC3-1504/1617 HC3-1470 HC3-1348/1617 HC3-1470/1617 HC3-1264/1324 HC3-1348/1386 HC3-749/784 HC3-874/921 HC3-936/970 HC3-1496 HC3-1496/1617 HC3-994 HC3-994/1617 HC3-1617
[0078]Certain modified C3 proteins (also called hybrid proteins or chimerae) were produced by site-directed mutagenesis as described below. Briefly, for site-directed mutagenesis replacing small portions of human C3 sequence with CVF, the procedure of Ho et al. was used (Ho, S. N., Hunt, H. D., Horton, R. M., Pullen, J. K. and Pease, L. R. (1989) "Site-Directed Mutagenesis by Overlap Extension Using the Polymerase Chain Reaction" Gene, 77:51-59, herein incorporated by reference in its entirety). In this method, two PCR reactions were performed, one with a forward primer somewhere upstream from the site of the desired mutation. The second, reverse primer contained the mutation. The second PCR of this round had a forward primer containing the desired mutation, with the reverse primer being downstream from the mutation site. At least one unique restriction site is preferably present in each of the PCR products from this step so that it is possible to transfer the modified DNA back into the original clone. Amplification was done using a large amount of template DNA, and a low number of cycles to minimize mutations introduced by the PCR process. More specifically, the first round of PCR, the reaction for the 5' product used a human C3 plasmid as the template, while the other PCR used a CVF plasmid. In this case the middle, "mutagenesis" primers partially consisted of C3 sequences and partially of CVF sequences, providing a "bridge" between the two sequences.
[0079]After amplification, the two products were purified by gel electrophoresis and isolated from the gel using a Qiaquick Gel extraction kit from Qiagen. Then, the fragments were combined, and another PCR reaction done, using the two fragments as the template, and the outside primers as the amplification primers. Again, the PCR reaction was performed using a high concentration of template DNA and only cycled for a few cycles to minimize PCR-caused mutations. The resulting PCR product was cut with the two unique restriction enzymes, size purified on an agarose gel, and the fragment of interest isolated using the Qiaquick column. The fragments were then cut with the appropriate enzymes, and cloned into either pBS-HuC3 or pHC3-1550(-sig) that had been cut with the same enzymes.
[0080]The first hybrid plasmid, pHC3-1550 contained CVF sequences replacing the homologous C3 sequences from position 1550 to the C-terminus of the protein. The second hybrid plasmid, pHC3-1504, codes for a hybrid protein containing CVF sequences replacing human C3 sequences from position 1504 to the C-terminus of the protein. The third hybrid plasmid, pHC3-1348, contained CVF sequences replacing the homologous C3 sequences from position 1348 to the C terminus of the protein. To prepare the first plasmid, two initial PCR reactions were performed. Both hybrids had about 19 amino acid vector-coded sequence having some or all of the following RSPWPGVPTSPVWWNSADA (SEQ ID NO:5). Because of the method of cloning of the C3 gene, these amino acids coded for by the cloning vector, were N-terminal to the N-terminus of human C3. These extra amino acids did not affect the activity of the protein, so they could be removed with no ill effect. The first plasmid was prepared using pBS-HC3-2 as a template, and using the following oligonucleotides as primers: HC3H5-1 (GGATGCCACTATGTCTATATTGGACATATCC--SEQ ID NO:6), and HC3H5-2 (TCTTCTATTCGAACCAGTCGGGTCTTGTAC--SEQ ID NO:7). The second PCR used pCVF-FL3Δ as a template, and the following oligonucleotides as primers: HuC3H5-3 (GTACAAGACCCGACTGGTTCGAATAGAAGAACAAG--SEQ ID NO:8) and HuC3H5-4 (TATCATGTAAGCGGCCGCGTATAAACAATTTAAGGG--SEQ ID NO:9). Both reactions were performed in an Eppendorf thermocycler, using the following program: 95° C. for 5 min., followed by five cycles of 95° C. for 30 sec., 58° C. for 30 sec. (gradient from 50 to 65° C.), and 72° C. for 1 min., followed by 20 cycles of 95° C. for 30 sec., 57.5° C. for 30 sec. (gradient from 55 to 60° C.), 72° C. for 1 min, followed by 72° C. for 10 min. The two fragments were purified using a Qiagen PCR cleanup kit, and joined in a second PCR reaction, using HC3H5-1 and HC3H5-2 as primers, and the two primary PCR products as template. The cycling conditions used for this reaction were: 95° C. for 5 min, followed by 5 cycles of; 95° C. for 30 sec., 51° C. for 30 sec (gradient from 46 to 56° C.), and 72° C. for 1.5 min, followed by 20 cycles of; 95° C. for 30 sec., 57° C. for 30 sec (Gradient from 52 to 62° C.), and 72° C. for 1.5 min. This was followed by a 10 min incubation at 72° C. The PCR fragment was purified as described above, cut with BsrGI and NotI, gel purified, and isolated using a Qiagen Gel Isolation kit. This fragment was cloned into pBS-HuC3-2 that had been cut with the same enzymes, and the resulting clones were screened for insertion of the correct fragment by digestion with EcoRI. All clones with the correct EcoRI digestion pattern were sequenced to ascertain that no PCR-induced mutations were inserted. A large-scale preparation of one clone (pHC3-1550) with the expected sequence was performed, and the insert excised by digestion with HindIII (and end repaired with T4 DNA polymerase) and NotI. The fragment was purified by gel electrophoresis, isolated from the gel as described above, and cloned into the Drosophila expression vector, pMT/V5-HisA that had been cut with EcoRV and NotI. Attempts to obtain expression of hybrid proteins from this construct resulted in very low yields of protein. For this reason, a construct was made having the human C3 signal sequence removed from pHC3-1550, while inserting a new, unique AfeI site. To do this, pHC3-1550 was amplified with the following two primers: HC3SigRemF; AGATCTCCATGGAAGCTTAGCGCTGGGAGTCCCATGTACTCTATCATC (SEQ ID NO:10, and HC3SigRemR: GCGTCCCGCCTfCAACAGCC (SEQ ID NO:11). After amplification, the fragment was purified as described above, cut with HindIII and SpeI. The 150 bp band was gel isolated, and cloned into pHC3-1550 that had been cut with the same enzymes. DNA from transformants was screened by cutting with AfeI, and all positive clones were confirmed by DNA sequencing. This plasmid was called pHC3-1550(-sig).
[0081]The insert was excised from the plasmid by digestion with AfeI, DraI (to fragment the plasmid), and NotI. The digest was run on a gel, and the 5 kb fragment isolated as described above. It was then ligated into pMT-Bip/V5-HisA that had been digested with EcoRV and NotI. The resulting plasmid was called pMB/HC3-1550.
[0082]The plasmid for the production of the second hybrid protein, HC3-1504, was produced in a similar manner, as follows. Two PCR reactions were performed to obtain the human C3 and CVF portions of the coding sequence. In the first, pBS-HuC3-2 was used as a template, with the following oligonucleotides used as primers: HC3H5-3-F1(TCTGTGTGGCAGACCCCTTCGAGG--SEQ ID NO:12) and HC3H5-3--R1 (CGTrACCAATACATATCTTGTTCAGCTTTCCATCC--SEQ ID NO:13). The second PCR used pCVF-FL3Δ as a template, and the following oligonucleotides as primers: HuCC3H5-3-F2 (GGATGGAAAGCTGAACAAGATATGTATTGGTAACG--SEQ ID NO:14), and HuC3H5-3-R2 (CATCCATGACATAGATATCATTACCATCTTG--SEQ ID NO:15). The resulting two PCR products were joined in a PCR reaction, using HuC3H5-3-F1 and HuC3H5-3-R2 as primers, and the two PCR fragments as the template. After the second PCR reaction, the product was purified using a Qiagen PCR cleanup kit, it was then cut with NspV, and cloned into pHC3-1550(-sig) that had been cut with the same enzyme and that had also been treated with Calf intestine alkaline phosphate. Resulting clones were cut with EcoRI to determine the orientation of the insert and sequenced to ascertain that no PCR induced modifications were present. The resulting plasmid was called pHC3-1504. The insert from this plasmid was then isolated as described above, and cloned into pMT-Bip/V5-HisA as described above. This plasmid was called pMB/HC3-1504.
[0083]The plasmid for the production of the third construct, HC3-1348, was constructed in a similar manner to that used for HC3-1504. The only difference is that the two mutation primers were HuC3H5-5-1R (GCAACTGTGCGTTATACATTGTCACCACCGAC--SEQ ID NO:16) and HuC3H5-5-2F (GTCGGTGGTGACAATGTATAACGCACAGTTGC--SEQ ID NO:17). For the primary PCR reactions, the primers used were HuC3H5-3-1F and HuC3H5-5-1R, and the template was pBS-HuC3-2, while the primers used for the second primary PCR reaction were HuC3H5-5-2F and HuC3H5-3-2R, using pCVF-FL3Δ as the template. After the primary PCR, the two fragments were purified and used as templates for the secondary PCR reaction as described for the construction of pHC3-1504. The secondary PCR reaction product was purified, cut with NspV, and cloned into pHC3-1550, the sequence confirmed and the insert cloned into pMT-Bip/V5-HisA) as described above.
[0084]The plasmid for the production of the fourth hybrid protein, HC3-1496, was produced in a similar manner, as follows. Two PCR reactions were performed to obtain the human C3 and CVF portions of the coding sequence. In the first, pBS-HuC3-2 was used as a template, with the following oligonucleotides used as primers: HC3H5-3-F1(TCTGTGTGGCAGACCCCTTCGAGG--SEQ ID NO:12) and HC3H5-4-R1 GAGAAGGCCTGTTCCTTTATCCGGATGGTAGAACCGGGTAC (SEQ ID NO:18) and. The second PCR used pCVF-FL3Δ as a template, and the following oligonucleotides as primers: HuCC3H5-4-F2 CCGGTTCTACCATCCGGATAAAGGAACAGGCCTTC (SEQ ID NO:19), and HuC3H5-3-R2 (CATCCATGACATAGATATCATTACCATCTTG--SEQ ID NO:20). The resulting two PCR products were joined in a PCR reaction, using HuC3H5-3-F1 and HuC3H5-3-R2 as primers, and the two PCR fragments as the template. After the second PCR reaction, the product was purified using a Qiagen PCR cleanup kit. It was then cut with NspV, and cloned into pHC3-1550(-sig) that had been cut with the same enzyme and been Calf intestine alkaline phosphate treated. The resulting plasmid was called pHC3-1496. The insert from this plasmid was then isolated as described above, and cloned into pMT-Bip/V5-HisA as described above. This plasmid was called pMB/HC3-1496.
[0085]The plasmid for the production of the fifth hybrid protein, HC3-1550/1617, in which the C-terminal 46 amino acid residues of HC3-1550 are replaced with human C3 sequences, is described below. Again, two PCR reactions were done to obtain the CVF and human C3 portions of the coding sequence. In the first, pHC3-1550 was amplified, using the following two primers; HuC3H5-F1 GGATGCCACTATGTCTATATTGGACATATCC (SEQ ID NO:21), and HuC3H5-2R1, CCCGATGATGTAGCTGAGTTTATCTTTCGTGGG (SEQ ID NO:22). The second PCR was performed using pCVF-FL3Δ as the template, and HuC3H5-2F2 (CCCACGAAAGATAAACTCAGCTACATCATCGGG--SEQ ID NO:23) and HuC3H5-2-R2 (AATTGGAGCTCCACCGCGGTGG--SEQ ID NO:24) as the primers. After the first PCR, the fragments were joined in a second PCR reaction, using HuC3H5-F1 and HuC3H5-2-R2 as the primers, and the two PCR fragments as the template. Following this PCR, the amplified fragment was purified using Qiagen PCR purification columns, cut with BsrGI and NotI, and cloned into pHC3-1550(-sig) that had been cut with the same enzymes. The resulting plasmid was sequenced to ascertain the correct sequence. It was called pHC3-1550/1617. The insert was isolated as described above and cloned into pMT-Bip/V5-HisA as described above. This plasmid was called pMB/HC3-1550/1617.
[0086]In some cases, portions of the human C3 are additionally exchanged with CVF sequence at more than one site to generate a modified C3 that is useful for one or more of the purposes presented herein. Thus, modified C3 proteins are generated wherein CVF-specific sequence is inserted in more than one region, or known regions are mutagenized by various means. For example, in addition to a site or sites required for the formation of a physically stable convertase with factor B, it can also be desirable to change a Factor I cleavage site in human C3. Factor I-resistant mutants of human C3 have been successfully described previously by Fecke, et al., 1998 (Fecke, W., Farries, T. C., D'Cruz, L. G., Napper, C. M., and Harrison, R. A. (1998) Xenotransplantation 5:29-34, herein incorporated by reference in its entirety). For example, replacement of select sequences required for specific CVF functions allows one to engineer novel C3 derivatives wherein specific subsets of CVF functions are present or eliminated, respectively (e.g. a C3 derivative or CVF derivative that forms a stable C3 convertase but does not cleave C5). A modified C3 molecule that does not activate C5 may have particular advantage for therapy as it prevents the generation of the pro-inflammatory C5a anaphylatoxin.
Example 2
Expression of Modified human C3 Proteins
[0087]The proteins were produced in the Drosophila S2 cell system, using the Drosophila Bip signal sequence for secretion of the proteins. Briefly, the plasmids pMB/HC3-1550, pMB/HC3-1504, pMB/HC3-1496, pMB/HC3-1550/1617, and pMB/HC3-1348 were transfected into Drosophila S2 cells using the calcium phosphate method of Chen and Okayama (Chen, C., and Okayama, H. (1987) Mol. Cell. Biol. 7(8), 2745-2752, herein incorporated by reference in its entirety). S2 cells were transfected with a mixture of expression plasmid and pCoBlast, using a ratio of 19:1 (w:w). Following transfection, cells containing both plasmids were selected using blasticidin (25 μg/ml). For expression, 1-liter cultures of transfected cells were grown in serum-free medium (Hi-Five plus L-glutamine), in the absence of blasticidin. When the cells reached a density of 5×106 cells/ml., production of the recombinant proteins was induced by the addition of CuSO4 to a final concentration of 25 μM. Cultures were allowed to express recombinant proteins for 4-5 days. Hybrid proteins were then purified from the media by a combination of ANX, Sephacryl H-300, and CM-FF chromatography.
[0088]Because of the method of cloning of the C3 gene, there are several amino acids (approximately 19) coded for by the cloning vector, that are N-terminal to the N-terminus of human C3. These extra amino acids do not affect the activity of the protein, so they can be removed with no ill effect. In some cases, it can be preferred for there to be at least two amino acids that are an artifact of the restriction sites needed to clone the final construct into the expression vector. In various embodiments, a variety of signal sequences can be used, including the native signal sequence of human C3, as well as any other signal sequence effective in directing entry of the nascent polypeptide into the endoplasmic reticulum.
[0089]Other expression systems that can be used, include but are not limited to: Baculovirus infection of Sf9 or HiFive cells (other insect expression systems), CHO cells, COS-7 cells (mammalian expression systems), E. coli, BHK, HEK293 cells, and various yeast expression systems, including the Hanselula yeast expression system.
Example 3
Results of the Activity Measurements of the Modified Human Complement C3 Proteins
[0090]The purified modified human C3 protein hybrids were subjected to a number of functional analyses as follows.
[0091]Complement Depletion:
[0092]This assay measures the ability of a protein to deplete complement in human (or other) serum. The assay was done in two steps. In the first step, the protein of interest was diluted to the desired concentrations in buffer, usually by serial dilution (typically from less than a nanogram/microliter up to approximately 320 ng/microliter or 3.2 μg in the 10 microliters used in the assay). Then, a 10 μl aliquot of the diluted protein was mixed with undiluted serum. The mixture was incubated at 37° C. for 3 hours, which allows the protein to activate complement by forming a C3 convertase. The convertases formed were then able to activate C3 in the serum. Then, to measure the amount of complement activity left, the serum was diluted and mixed with antibody-sensitized sheep erythrocytes, which are easily lysed by complement when it is present in serum. This reaction was allowed to proceed for 30 minutes, and was stopped by diluting the mixture in cold buffer. The cells were centrifuged, and the lysed cells quantified by measuring the hemoglobin released. The results are shown in FIG. 4 and FIG. 9.
[0093]As expected, very small amounts of CVF were able to completely deplete the complement in human serum. 800 ng of either of the proteins HC3-1348 and HC3-1496 was able to completely deplete 10 μl of human serum. Other hybrid proteins were less active, requiring about 34 μg of protein to partially deplete 10 μl of human serum of complement. One hybrid protein, HC3-1550/1617 was apparently unable to deplete complement at the concentrations examined.
[0094]Unexpected was the observation that two of the proteins (HC3-1550 and HC3-1348) were actually able to deplete complement in guinea pig serum, although this activity was clearly less than that of CVF. Notably, preferred embodiments HC3-1550, HC3-1504, HC3-1496 and HC3-1348 were all able to deplete complement in human serum.
Factor B Activation Assay.
[0095]This was an assay to measure the ability of a hybrid protein to activate factor B, and form a C3/C5 convertase. The convertase formation was measured as a function of the cleavage of factor B into Bb and Ba. In the assay, purified hybrid proteins were incubated with a three-fold excess of factor B and factor D (all highly purified) in the presence of magnesium at 37° C. At various times, aliquots of the reaction were withdrawn, and the reaction stopped by adding EDTA, which chelates the magnesium. The reaction products were run on a non-reducing SDS-polyacrylamide gel, which was stained for proteins with Coomassie Blue. The amount of Factor B converted was quantified by scanning the gel into a specialized computer program and measuring the amount of protein in the factor B and Bb bands.
[0096]The results in FIGS. 5 and 10 show that Factor B was activated very rapidly in the presence of Human C3, for two reasons. Human C3 was able to bind factor B very rapidly, which makes it available for cleavage by factor D. However, the resulting convertase was very unstable, and falls apart rapidly, making the C3b available to bind more factor B. The reaction in the presence of CVF was much slower, which was a result of the lower affinity of CVF for factor B, and the greater stability of the CVF containing convertase. The conversion of factor B in the presence of HC3-1504 or HC3-1496 was quite similar to that of CVF. HC3-1550 and HC3-1550/1617 were able to convert factor B much more rapidly than CVF, but slower than C3b. This was probably a result of the convertase being less stable than the CVF containing enzyme, but more stable than the C3b containing convertase. In addition, it is likely that the initial binding of factor B by HC3-1550 is much more rapid than by CVF. Finally, HC3-1348 supports the cleavage of factor B less well than the other proteins discussed. This is probably a combination of the resulting convertase being more stable than the C3b containing enzyme, and the initial binding of factor B being less rapid.
C3 Convertase Activity Assay
[0097]This assay measures the activity of C3/C5 convertases containing hybrid proteins to activate human C3, by cleaving off the C3a peptide. To perform this assay, convertases were formed as described above, and the reaction stopped by the addition of EDTA. The convertase was then mixed with human C3, and the reaction incubated at 37° C. At the indicated times, aliquots were removed, and the reaction stopped by mixing with gel loading buffer containing SDS and β-mercaptoethanol. The SDS denatures the proteins, and the β-mercaptoethanol reduces the disulfide bonds between cysteines in the proteins. After electrophoresis under reducing conditions, the gel was stained with Coomassie Blue dye, and the relative amounts of the C3 α-chain and C3 α'-chain quantified as described above. Care was taken to use the same amount of convertase in each reaction.
[0098]FIG. 6 and FIG. 11 show that in this assay, CVF and HC3-1550 both were able to convert human C3 at approximately equal rates. HC3-1504 was markedly slower than CVF, but still was able to completely convert the amount of C3 present within one hour. HC3-1348 and HC3-1496 appeared to convert C3 at a rate faster than CVF, while HC3-1550/1617 appeared to convert C3 at an initially high rate which became slower after about 10 minutes. To further investigate this phenomenon, the C3 conversion assay was repeated, using all proteins except HC3-1504. In an effort to reduce the speed of the reaction, the amount of convertase present was reduced by a factor of five. These results are shown in FIG. 7 and in the insert of FIG. 11. In this assay, it was clear that the HC3-1348 and HC3-1496 formed convertases that were markedly efficient than CVF. HC3-1550/1617 formed a convertase that was initially active, but appears to become mostly inactive after about 10 minutes.
C5 Conversion Assay
[0099]The assay for C5 conversion activity was done essentially as described by Petrella et al., (1987) J. Immunol. Methods 104(1-2), 159-172, herein incorporated by reference in its entirety. In this assay, C5 convertase was formed as described above, using a total of 3 μg protein. After convertase formation, the reaction was stopped by the addition of EDTA to a final concentration of 5 mM. Then, 5 μl of this reaction was added to a 25 μl reaction containing 7 μg C5 in PBS. The reaction was incubated at 37° C. for 24 hours, and the reaction stopped by the addition of 7 μl Laemmli gel loading buffer, followed by boiling for 5 minutes. The reaction products were separated by reducing SDS-PAGE, and the gel was stained with Coomassie Blue dye, and the relative amounts of the C5 α-chain and C5 α'-chain quantified as described above. None of the current proteins are able to form an active C5 convertase.
[0100]The assay for degradation of the proteins by factors H and I was performed essentially according to the method of Oran and Isenman ((1999) J. Biol. Chem. 274 (8), 5120-5130, herein incorporated by reference in its entirety). In this method, 12 μg of each protein was incubated with 4.3 μg factor H and 0.3 μg factor I at 37° C. in a total volume of 60 μl. At the indicated times, 10 μl aliquots were withdrawn, and reactions stopped by the addition of 5 μl 5× Laemli gel loading buffer. Reaction products were separated by SDS-PAGE on a 4-20% gradient gel under reducing conditions. These data show that all proteins are partially resistant to digestion by factor I in the presence of factor H. In a similar assay, C3b was nearly digested to completion at the 0 timepoint.
Example 4
A Method for Depleting Complement Locally or Systemically
[0101]The modified human Complement C3 proteins produced by the methods disclosed herein are used to deplete complement locally or systemically as follows:
[0102]Local depletion is effected when the modified C3 proteins are administered locally to an organ, tissue, cavity, or intradermally. This results in a temporary and complete depletion of complement in the area. Alternatively, local depletion may use a specific monoclonal antibody which, when chemically attached to the modified C3, would localize it to a specific tissue, a disease site, or an infected cell to cause continuous depletion of complement in that area.
[0103]Systemic depletion is effected when the modified C3 proteins are administered systemically, for example, intravenously or intraperitoneally. This results in a temporary and complete depletion of complement systemically. This method can be used for reperfusion injury, coronary heart surgery, transplantation and/or systemic disease, particularly during a flare-up of symptoms or during episodical activity.
[0104]Some of the most advantageous qualities of the modified human C3 proteins used in each of these cases and in specific disease states can vary considerably. For example, in those cases for which an immediate, but temporary depletion of complement is desired, a modified protein having a shorter plasma half-life and/or lower stability, but high complement activation activity is preferred. In treating a chronic disease, a long plasma half-life and/or high stability, even if accompanied by a low or sluggish activity, would be preferred. Further, a modified C3 molecule that does not activate C5 can be particularly advantageous for certain therapies as it prevents the generation of the pro-inflammatory C5a anaphylatoxin.
Example 5
A Method for the Treatment of Reperfusion Injury
[0105]Examples of reperfusion injuries are tissue damage after the reopening of blocked vessels (e.g. after a heart attack or ischemic stroke), and reperfusion of a transplanted organ with a recipients' blood. For a clogged coronary artery or for reperfusion after organ transplantation, it can be desirable in many cases to deplete the complement before the transplant is reperfused or before the opening of the blocked vessel. The ability to avoid complement activation can avoid the tissue damage it causes, so the only remaining primary source of tissue damage is oxygen starvation. Typically, tissue damage due to complement activation during reperfusion is twice as great as tissue damage due to oxygen starvation--that is, roughly 2/3 of tissue damage is attributable to complement activation while 1/3 is attributable to oxygen starvation. Thus, reperfusion injury can be greatly diminished by depleting complement prior to reperfusion, as is possible with embodiments of the present invention. In this case it is preferable to use the highest activity convertase and to use a high dosage. The stability of the convertase is less important than it would be for a chronic disease.
[0106]In general terms, the methods of these embodiments of the invention involve administering an effective amount of a modified C3 protein systemically allowing enough time for depletion of complement, and then performing the surgery.
Example 6
A Method for Increasing the Effectiveness and/or Efficiency of Gene Therapy
[0107]This method relies on the depletion of complement in order to help prolong survival of a useful virus in the body. Because complement has been found to help the removal from the body of certain viral vectors used in gene therapy, it is desirable to reduce the amount of circulating complement prior to administration of a gene therapy vector. This may be done locally or systemically depending upon the type of gene therapy that is being used.
[0108]The method involves administering an effective amount of a modified C3 protein systemically or to the local area where the gene therapy is being administered, allowing time for depletion of complement, and then administering the gene therapy.
Example 7
A Method for Increasing Delivery of a Therapeutic (e.g. Chemotherapeutic) or Diagnostic Agent
[0109]To increase the blood flow to an area where a therapeutic is being administered, a modified C3 protein is chemically linked to a monoclonal antibody with an affinity for the tissue of choice. In this example, the modified C3 is one that forms a highly active C3/C5 convertase. The modified C3/antibody is targeted to the tissue, resulting in local complement activation in the region, causing vessel permeability due to complement activation. The vessel permeability continues as long as the active convertase/antibody complexes are bound to the target, because new, non-activated complement is continually supplied by the blood arriving at the target.
[0110]For use of this method during the treatment of lung cancer, a modified C3 protein-monoclonal antibody hybrid is administered to a lung cancer patient, using a monoclonal antibody that recognizes a lung-specific antigen. The antibody binds to lung tissue and activates complement locally. This increases the vessel permeability in the lung, permitting the chemotherapeutic agent to more efficiently and effectively act on the lung cancer. As described above, this method permits continuous local activation of complement, which permits a persistent local elevation in blood-vessel permeability.
Example 8
A Method of Treating Rheumatoid Arthritis, Lupus and Other Autoimmune or Immune Complex Diseases
[0111]This method uses the example of rheumatoid arthritis as one of several conditions in which pain and inflammation arise from complement activation in a local area. For treatment, a modified C3 protein is administered systemically or locally, diminishing the complement response/activation by depleting the complement. This can reduce the symptoms of the disease and the progression of the disease. It can also be beneficial to have episodical depletion in combination with a longer term lowering of the activity of the complement system, especially when there are episodes of exacerbation of the symptoms of the disease. Further, this method can be used with other diseases with circulating immune complexes, such as lupus and other autoimmune diseases.
[0112]For example, when a complement-activating autoantibody is produced and is directed against the body's own proteins, the disease effect is primarily due to the binding of the antibody to the target causing complement activation and tissue damage as well as other interference with normal function. Myasthenia Gravis is an example of this type of disease. Autoantibody binds to the neuromuscular endplate where the nerve comes into contact with the muscle. Complement is activated and blocks the neurotransmitter, resulting in paralysis. Employing the method of this embodiment of the invention, the systemic depletion of complement, either continuously or during exacerbations in the disease, can markedly reduce symptoms and progression thereof.
Example 9
Method of Selecting a Modified C3 Protein
[0113]Various modified C3 proteins are useful for different diseases, and methods of treatment. Thus, it is useful to analyze the functional qualities of the modified C3 proteins of embodiments of the invention and to use them accordingly. The following methods are employed to analyze the function of purified modified C3 proteins produced as in Example 2. The methods described herein, as well as others that are known to those of skill in the art, may be used.
[0114]Assays to determine convertase activity. In addition to the specific assays as mentioned below, two hemolytic assays for depletion of serum complement activity and induction of bystander lysis can be employed for screening.
[0115]Complement depletion assay. To measure the anticomplementary (complement consumption) activity of modified C3 proteins, a small volume of human serum is incubated with CVF or hybrid proteins for three hours or shorter periods of time at 37° C. at a protein concentration of 5 μg/ml, to allow the proteins to deplete complement. The remaining complement hemolytic activity is subsequently measured using sensitized sheep erythrocytes using methods known to one of skill in the art including that of Cochrane et al., 1970 (Cochrane et al., 1970, J. Immunol. 117:630-4, herein incorporated by reference in its entirety).
[0116]Bystander lysis assay. The bystander lysis assay is performed by incubating 20 μl of normal guinea pig serum at 37° C. with 20 μl of CVF or hybrid proteins at a concentration of 5 μg/ml and 20 μl guinea pig erythrocytes (5×108/ml). The CVF or hybrid proteins participate in fluid-phase activation of C5, which leads to lysis of the erythrocytes. Thus, presence of hemoglobin in the supernatant is indicative of C5 activation. The reaction is incubated at 37° C. for 30 minutes, and is stopped by the addition of 1 ml of cold buffer. After centrifugation, the released hemoglobin is measured spectrophotometrically. (Vogel, C. W., and Muller-Eberhard, H. J. (1984) J. Immunol. Methods 73(1), 203-220, herein incorporated by reference in its entirety).
[0117]C3 convertase formation/Factor B activation. To detect cleavage of Factor B into Ba and Bb, a hybrid protein (at 1 μM) is incubated for up to twenty four hours in the presence of a three-fold molar excess of Factor B and 0.5 μM of Factor D in the presence of MgCl2 at 37° C. The reaction mixtures are analyzed by electrophoresis on 7.5% (w/v) SDS polyacrylamide gels under non-reducing conditions to monitor the disappearance of Factor B and the appearance of the cleavage products Ba and Bb. If necessary, a subsequent western blot can be performed to detect the Ba and Bb cleavage fragments. Controls can include native CVF, pro-CVF, cobra C3, iC3, human C3, iC3, C3b, and EDTA (Vogel and Muller-Eberhard, 1982, J. Biol. Chem. 257:8292-9, herein incorporated by reference in its entirety).
[0118]C3 cleaving activity. To examine C3 cleaving activity, a C3 convertase is pre-formed as described herein, in reference to "C3 convertase formation/Factor B activation," using the hybrid proteins and human Factor B and Factor D. The convertase formation is stopped by the addition of EDTA, and purified human C3 is added The reaction mixture is incubated at 37° C. for one hour or for any other appropriate period of time. Aliquots are taken and immediately transferred into an ice water bath to stop further C3 activation. C3 cleavage is monitored by the disappearance of the C3 α-chain and appearance of the C3 α'-chain by running the reaction products on a 7.5% (w/v) SDS polyacrylamide gel under reducing conditions. If necessary, a subsequent western blot using anti-C3 antiserum is performed. Controls include native CVF, pro-CVF, and human and cobra iC3 or C3b (Vogel and MUller-Eberhard, 1982, J. Biol. Chem. 257:8292-9, herein incorporated by reference in its entirety).
[0119]C5 cleaving assay. The C5 cleaving assay is performed as described above for the C3 cleaving assay using purified human C5 as substrate (Petrella et al., 1987, J. Immunol. 164:4742-4751, herein incorporated by reference in its entirety). See, for example, FIG. 12.
[0120]Assay for convertase stability. Bimolecular convertases are pre-formed using the hybrid proteins and purified human Factor B and Factor D as described above. After addition of EDTA, the mixture is incubated at 37° C. and aliquots are removed over a period of 24 hours or shorter periods of time if appropriate, and each aliquot is immediately placed in an ice water bath. Subsequently, the C3 convertase activity is determined by the C3 cleaving assay described above. From the reduction of the C3 cleaving activity over time, the half-life of the spontaneous decay-dissociation of the various convertases is calculated. If insufficient quantities of hybrid proteins for this assay are available, the enzymatic activity is determined using the fluorogenic tripeptide t-butyloxy-carbonyl-leucyl-glycyl-arginyl-aminomethylcoumarin. (Caporale, L. H., Gabaer, S. S., Kell, W., and Gotze, O. 1981 J. Immunol. 126(5), 1963-1965, herein incorporated by reference in its entirety).
[0121]Assay for Factor H binding. Factor H binding to hybrid proteins is determined using an ELISA assay. Hybrid proteins are adsorbed onto microtiter plates. After blocking with ovalbumin and BSA, purified human Factor H at 10 μg/ml is added and incubated for 30 minutes at room temperature. After washing, bound Factor H is detected with anti-Factor-H antibody followed with an appropriate phosphatase-linked secondary antibody. If factor H can bind to the protein, an appropriate color change is observed. Controls can include native CVF, pro-CVF, cobra and human C3, as well as cobra and human iC3 (Alsenz et al., 1992, Dev. Comp. Immunol. 16:63-76, herein incorporated by reference in its entirety).
[0122]Assay for Factor I cleavage. Hybrid proteins are incubated with purified human Factor H and Factor I at 37° C. for several hours. The reactions are analyzed by subsequent 4-20% (w/v) SDS polyacrylamide gel electrophoresis under reducing conditions. Factor I activity is determined by the reduction in the strength of the 105 kDa α'-chain band, and appearance of bands with a molecular weight of 37 and 40 kDa. If necessary, a subsequent western blot is performed using anti-CVF and/or anti-C3 antibodies. Alternatively, hybrid proteins are labeled with 125I using the iodogen method (Fraker and Speck, 1978). Cleavage products are detected after SDS polyacrylamide gel electrophoresis by autoradiography (Lambris et al., 1996, J. Immunol. 156:4821-32, herein incorporated by reference in its entirety). See, for example, FIG. 12.
[0123]Assays for Immunogenicity. Various methods can be used to analyze immunogenicity, including but not limited to, skin tests, testing the modified C3 protein in transgenic animals which have been genetically engineered to have human immune systems, in vitro methods, including RIA tests using serum generated in such transgenic animals, Radioimmunoprecipitation assays, ELISA assays, Electrochemiluminescence, and Surface Plasmon Resonance. In addition, mouse, rate or guinea pig analogs of some proteins are constructed, using either mouse, rate or guinea pig C3 and CVF sequences. These are injected into the appropriate animal, and serum is collected and analyzed for the production of antibodies against the hybrid proteins.
Example 10
Method of Measuring Plasma Half-Life
[0124]There are many factors that can affect the plasma half-life of the modified C3 protein, including but not limited to specific antibodies produced by the immune system, proteases circulating within the serum, non-specific immune responses, and specific regulatory factors such as Factors H and I. In order for the modified C3 proteins to be able to activate and subsequently deplete complement, preferred C3 proteins will persist within human plasma for at least a minimum amount of time. Thus, it is of interest to identify the plasma half-life of the modified C3 proteins to determine how useful they will be for treatment of certain diseases.
[0125]This method measures the stability of the modified C3 protein in plasma in three ways. However it is to be understood that one or all of the methods can be used as well as any other methods known to one of skill in the art.
[0126]The first method measures the stability in serum in vitro. Human serum is isolated and separated from the whole blood of a patient. Aliquots of different concentrations of the modified C3 proteins are added to the serum and allowed to incubate. Aliquots of the serum are removed at various time intervals and the amount of modified C3 that persists is identified in an ELISA assay using a monoclonal antibody which is specific to C3.
[0127]A second method allows for the identification of stability in serum in a humanized animal. The modified C3 protein is administered to the animal and blood samples are taken over time. The amount of modified C3 protein is identified in an ELISA assay using specific antibodies to the protein.
[0128]A third method allows for the identification of stability in a human patient. The modified C3 is administered to the patient and blood samples are removed over time. The amount of modified C3 protein is identified using an ELISA assay. This will give a clear indication of how long the modified C3 protein circulates within the plasma of a patient.
[0129]In some embodiments, the antibody that is used need not be specific for the modified C3. For example, antibodies that recognize normal C3 can be tested and used to identify the modified C3 in an ELISA procedure.
Example 11
C3 Convertase Formation of Modified Human Complement C3 Proteins as Measured Using Surface Plasmon Resonance
[0130]Table 2 shows relative binding of complement proteins to C3, CVF, and recombinant human C3/CVF proteins. In all cases, a higher number indicates that the protein interaction is tighter. The proteins (C3b, etc.) were bound to a BIACORE CHIP®, and then contacted with complement factors. The amount of each of the complement factors bound to the chip (and thus bound to the modified C3 proteins) was determined by surface plasmon resonance. The results showed that: 1) Neither of the recombinant proteins bound C5, consistent with the inability to form a convertase capable of cleaving C5. 2) Both proteins bound factor H, though CVF does not. Factor H is one of the regulatory proteins that is capable of dissociating the C3b,Bb convertase complex and directing a second complement protein, factor I, to inactivate C3b through cleavage. 3) The affinity of the proteins for factor B in the presence of factor D and magnesium was approximately proportional to the rate the proteins were able to form a C3 convertase (as measured by their ability to mediate factor B cleavage--see FIG. 10). Both HC3-1550 and HC3-1348 were cleaved but slower than C3b. CVF is not cleaved at all by H or I, because natural CVF has no H or I sites. It is interesting that recombinant CVF, a 2 chain molecule, has 2 or 3 I sites, but is still not cleaved by factor I.
TABLE-US-00002 TABLE 2 Relative binding of Complement proteins to C3, CVF, and recombinant human C3/CVF proteins. Amount of protein bound (RU)-2nd column corrected for MW Protein on Factor B Factor B Factor B Chip C5 Factor H (in EDTA) (with Mg) (with fD and Mg) C3b 607 646 243 285 625 HC3-1550 66 550 9 264 340 HC3-1348 18 219 5 276 162 CVF 617 -14 99 298 370
[0131]Table 3 shows the stability of the C3 convertases formed by the modified human Complement C3 proteins as measured by surface plasmon resonance on a BIACORE machine at 25° C. The results showed that both proteins were able to form C3 convertases that were substantially more stable than the C3b-containing convertase. The table also shows that the HC3-1348-containing convertase was actually more stable than the CVF-containing enzyme. These results are consistent with other measurements of half life at higher temperatures which showed that CVF had a half life of 7 hours at 37° C. and C3b had a half life of 1.5 minutes at 37° C.
TABLE-US-00003 TABLE 3 C3 convertase formation Protein on Chip T1/2 of C3 convertase (min) C3b 4.3 HC3-1550 119.3 HC3-1348 1720.0 CVF 1100.0
Example 12
Factor B Cleavage of Modified Human Complement C3 Proteins
[0132]The factor B cleavage graph in FIG. 10 shows a time course of the ability of the modified human complement C3b proteins to mediate the cleavage of factor B in the presence of magnesium ion and factor D. This can be used to measure the ability of the protein to form a C3/C5 convertase. Of note in this assay was that C3b formed a convertasc very efficiently, both because it bound factor B very efficiently in the presence of factor D and magnesium (see the data in Example 11), and because the resulting complex was very unstable. CVF, HC3-1496 and HC3-1504 all formed a convertase at approximately the same rate, while HC3-1550 and HC3-1550/1617 were intermediate between C3b and CVF in their ability to form a convertase. HC3-1348 was very slow to bind factor B. This is most probably explained by a combination of a longer half-life of the convertase once formed and a lower affinity of the hybrid protein for factor B.
Example 13
C3 Cleavage of Modified Human Complement C3 Proteins
[0133]FIG. 11 shows the ability of the convertases to cleave C3, and shows a time course of the cleavage reaction. The inset is a time course performed with 20% of the convertase used in the main graph. The results showed that CVF and HC3-1550 formed convertases that were approximately equally efficient at C3 cleavage, while HC3-1348 and HC3-1496 both formed convertases that were approximately 5-fold more efficient than CVF at cleaving C3. It is interesting that the HC3-1550/1617-containing convertase appeared to be quite unstable, and did not support C3 cleavage after about 10 minutes.
Example 14
Complement Depletion of Modified Human Complement C3 Proteins
[0134]The data in the complement depletion chart in FIG. 9 show that all proteins except HC3-1550/1617 were able to deplete complement, though with very different efficiencies. Both HC3-1550 and HC3-1504 were quite inefficient at the depletion of complement, while HC3-1496 and HC3-1348 were able to deplete complement quite efficiently, though not quite as well as natural or recombinant CVF. HC3-1348 was formerly referred to as HC3-1325 in a previous application, but substitution mapping has subsequently shown that the substitution was at 1348 rather than 1325.
[0135]In conclusion, HC3-1496 is an interesting protein because, although it only has an insert 8 amino acids longer than HC3-1504, it acts much more like HC3-1348 with respect to complement depletion and C3 cleavage and more like HC3-1504 with respect to factor B cleavage. It forms a convertase as well as HC3-1504 or CVF, but the resulting convertase is more active at cleaving C3.
[0136]HC31550/1617 was made by replacing the region from 1550 to the end of C3 with the CVF region, then taking away the last 46 amino acids and replacing them with C3. This chimeric molecule showed no complement depletion, formed a C3 convertase as well as HC3-1550, and the C3 convertase formed was nearly as active as HC3-550, but had an apparently shorter half-life.
Example 15
Methods of Producing Variants in the Modified C3 Proteins
[0137]Variants of the modified C3 proteins are produced that have advantageous qualities. By advantageous it is meant that the variants enhance one or more activities of the protein, including but not limited to: C3 convertase activity, serum decomplementation, factor B binding, cleavage of C3, Binding of Bb and Binding of C3. The variants can include one or more mutations in each region and can include one or more amino acids. Some specific variants are set out below:
[0138]Because the C-terminus is involved in stabilization of the C3 convertase, mutations are produced in the C-terminus (after aa 1617) of any modified C3 proteins. The mutations can be insertions, deletions, and substitutions. However, the mutations are preferably substitutions. The mutations stabilize the C3 convertase. Examples of mutations at the C terminus include but are not limited to: mutations at positions 1633, 1654, and 1658. Non-conservative as well as conservative mutations, in addition to amino acid changes that affect the conformation, are included among the embodiments. However, preferably the mutations result in a non-conservative amino acid change.
[0139]There are three changes between CVF and human C3 in the span of 1496 and 1504 (8 amino acid residues) that appear to be responsible for a large change in the activity of the convertase. Thus, mutations between positions 1496 and 1504 of any of the modified C3 proteins are included that result in increased activity of the convertase in cleaving C3 and/or in decomplementing serum. The mutations can be changes to one or more amino acids within that region. Preferably, the mutations are substitutions of one or more amino acids in that region, in particular, mutations that result in non-conservative amino acid changes.
[0140]Mutations between positions 1348 and 1496 in any modified C3 protein are produced that modify the ability of the protein to bind factor B (and specifically, the ability for C3b and factor B to bind), preferably the modification results in an increased ability to bind to factor B. Whole regions from CVF are switched out to C3 and vice versa. Further mutations are included that result in amino acid substitutions. More specifically, the regions from 1367-1379 are switched for CVF and for C3 and specific amino acids within this region are substituted.
[0141]Sequence changes in the region of around 1550 in CVF and 1570-1584 in C3 that may be responsible for the activity of the C3 convertase in cleaving C3, either from binding Bb or binding the target C3 molecule. Using CVF/cobra C3 substitutions, a series of 4 amino acid residues were identified (Q1550G, E1554R, P1556A and R1557Q-positions numbered according to CVF sequence numbering) that resulted in a protein with a much lower activity in cleaving C3. Therefore, variants are produced in the amino acids from position 1570-1584 that result in an increased activity of the C3 convertase in cleaving C3 for the modified C3 proteins. The variants are preferably amino acid substitutions of one or more amino acids in that region.
[0142]The various methods and techniques described above provide a number of ways to carry out the invention. Of course, it is to be understood that not necessarily all objectives or advantages described may be achieved in accordance with any particular embodiment described herein. Thus, for example, those skilled in the art will recognize that the methods may be performed in a manner that achieves or optimizes one advantage or group of advantages as taught herein without necessarily achieving other objectives or advantages as may be taught or suggested herein.
[0143]Furthermore, the skilled artisan will recognize the interchangeability of various features from different embodiments. Similarly, the various features and steps discussed above, as well as other known equivalents for each such feature or step, can be combined and/or exchanged by one of ordinary skill in this art to perform methods in accordance with principles described herein. Each patent, journal reference, and the like, cited herein is hereby incorporated by reference in its entirety.
[0144]Although the invention has been disclosed in the context of certain embodiments and examples, it is understood by those skilled in the art that the invention extends beyond the specifically disclosed embodiments to other alternative embodiments and/or uses and obvious modifications and equivalents thereof. Accordingly, the invention is not intended to be limited by the specific disclosures of preferred embodiments herein.
Sequence CWU
1
2415067DNAHomo sapiensCDS(61)...(5052) 1ctcctcccca tcctctccct ctgtccctct
gtccctctga ccctgcactg tcccagcacc 60atg gga ccc acc tca ggt ccc agc
ctg ctg ctc ctg cta cta acc cac 108Met Gly Pro Thr Ser Gly Pro Ser
Leu Leu Leu Leu Leu Leu Thr His1 5 10
15ctc ccc ctg gct ctg ggg agt ccc atg tac tct atc atc acc
ccc aac 156Leu Pro Leu Ala Leu Gly Ser Pro Met Tyr Ser Ile Ile Thr
Pro Asn 20 25 30atc ttg cgg
ctg gag agc gag gag acc atg gtg ctg gag gcc cac gac 204Ile Leu Arg
Leu Glu Ser Glu Glu Thr Met Val Leu Glu Ala His Asp 35
40 45gcg caa ggg gat gtt cca gtc act gtt act gtc
cac gac ttc cca ggc 252Ala Gln Gly Asp Val Pro Val Thr Val Thr Val
His Asp Phe Pro Gly 50 55 60aaa aaa
cta gtg ctg tcc agt gag aag act gtg ctg acc cct gcc acc 300Lys Lys
Leu Val Leu Ser Ser Glu Lys Thr Val Leu Thr Pro Ala Thr65
70 75 80aac cac atg ggc aac gtc acc
ttc acg atc cca gcc aac agg gag ttc 348Asn His Met Gly Asn Val Thr
Phe Thr Ile Pro Ala Asn Arg Glu Phe 85 90
95aag tca gaa aag ggg cgc aac aag ttc gtg acc gtg cag
gcc acc ttc 396Lys Ser Glu Lys Gly Arg Asn Lys Phe Val Thr Val Gln
Ala Thr Phe 100 105 110ggg acc
caa gtg gtg gag aag gtg gtg ctg gtc agc ctg cag agc ggg 444Gly Thr
Gln Val Val Glu Lys Val Val Leu Val Ser Leu Gln Ser Gly 115
120 125tac ctc ttc atc cag aca gac aag acc atc
tac acc cct ggc tcc aca 492Tyr Leu Phe Ile Gln Thr Asp Lys Thr Ile
Tyr Thr Pro Gly Ser Thr 130 135 140gtt
ctc tat cgg atc ttc acc gtc aac cac aag ctg cta ccc gtg ggc 540Val
Leu Tyr Arg Ile Phe Thr Val Asn His Lys Leu Leu Pro Val Gly145
150 155 160cgg acg gtc atg gtc aac
att gag aac ccg gaa ggc atc ccg gtc aag 588Arg Thr Val Met Val Asn
Ile Glu Asn Pro Glu Gly Ile Pro Val Lys 165
170 175cag gac tcc ttg tct tct cag aac cag ctt ggc gtc
ttg ccc ttg tct 636Gln Asp Ser Leu Ser Ser Gln Asn Gln Leu Gly Val
Leu Pro Leu Ser 180 185 190tgg
gac att ccg gaa ctc gtc aac atg ggc cag tgg aag atc cga gcc 684Trp
Asp Ile Pro Glu Leu Val Asn Met Gly Gln Trp Lys Ile Arg Ala 195
200 205tac tat gaa aac tca cca cag cag gtc
ttc tcc act gag ttt gag gtg 732Tyr Tyr Glu Asn Ser Pro Gln Gln Val
Phe Ser Thr Glu Phe Glu Val 210 215
220aag gag tac gtg ctg ccc agt ttc gag gtc ata gtg gag cct aca gag
780Lys Glu Tyr Val Leu Pro Ser Phe Glu Val Ile Val Glu Pro Thr Glu225
230 235 240aaa ttc tac tac
atc tat aac gag aag ggc ctg gag gtc acc atc acc 828Lys Phe Tyr Tyr
Ile Tyr Asn Glu Lys Gly Leu Glu Val Thr Ile Thr 245
250 255gcc agg ttc ctc tac ggg aag aaa gtg gag
gga act gcc ttt gtc atc 876Ala Arg Phe Leu Tyr Gly Lys Lys Val Glu
Gly Thr Ala Phe Val Ile 260 265
270ttc ggg atc cag gat ggc gaa cag agg att tcc ctg cct gaa tcc ctc
924Phe Gly Ile Gln Asp Gly Glu Gln Arg Ile Ser Leu Pro Glu Ser Leu
275 280 285aag cgc att ccg att gag gat
ggc tcg ggg gag gtt gtg ctg agc cgg 972Lys Arg Ile Pro Ile Glu Asp
Gly Ser Gly Glu Val Val Leu Ser Arg 290 295
300aag gta ctg ctg gac ggg gtg cag aac ctc cga gca gaa gac ctg gtg
1020Lys Val Leu Leu Asp Gly Val Gln Asn Leu Arg Ala Glu Asp Leu Val305
310 315 320ggg aag tct ttg
tac gtg tct gcc acc gtc atc ttg cac tca ggc agt 1068Gly Lys Ser Leu
Tyr Val Ser Ala Thr Val Ile Leu His Ser Gly Ser 325
330 335gac atg gtg cag gca gag cgc agc ggg atc
ccc atc gtg acc tct ccc 1116Asp Met Val Gln Ala Glu Arg Ser Gly Ile
Pro Ile Val Thr Ser Pro 340 345
350tac cag atc cac ttc acc aag aca ccc aag tac ttc aaa cca gga atg
1164Tyr Gln Ile His Phe Thr Lys Thr Pro Lys Tyr Phe Lys Pro Gly Met
355 360 365ccc ttt gac ctc atg gtg ttc
gtg acg aac cct gat ggc tct cca gcc 1212Pro Phe Asp Leu Met Val Phe
Val Thr Asn Pro Asp Gly Ser Pro Ala 370 375
380tac cga gtc ccc gtg gca gtc cag ggc gag gac act gtg cag tct cta
1260Tyr Arg Val Pro Val Ala Val Gln Gly Glu Asp Thr Val Gln Ser Leu385
390 395 400acc cag gga gat
ggc gtg gcc aaa ctc agc atc aac aca cac ccc agc 1308Thr Gln Gly Asp
Gly Val Ala Lys Leu Ser Ile Asn Thr His Pro Ser 405
410 415cag aag ccc ttg agc atc acg gtg cgc acg
aag aag cag gag ctc tcg 1356Gln Lys Pro Leu Ser Ile Thr Val Arg Thr
Lys Lys Gln Glu Leu Ser 420 425
430gag gca gag cag gct acc agg acc atg cag gct ctg ccc tac agc acc
1404Glu Ala Glu Gln Ala Thr Arg Thr Met Gln Ala Leu Pro Tyr Ser Thr
435 440 445gtg ggc aac tcc aac aat tac
ctg cat ctc tca gtg cta cgt aca gag 1452Val Gly Asn Ser Asn Asn Tyr
Leu His Leu Ser Val Leu Arg Thr Glu 450 455
460ctc aga ccc ggg gag acc ctc aac gtc aac ttc ctc ctg cga atg gac
1500Leu Arg Pro Gly Glu Thr Leu Asn Val Asn Phe Leu Leu Arg Met Asp465
470 475 480cgc gcc cac gag
gcc aag atc cgc tac tac acc tac ctg atc atg aac 1548Arg Ala His Glu
Ala Lys Ile Arg Tyr Tyr Thr Tyr Leu Ile Met Asn 485
490 495aag ggc agg ctg ttg aag gcg gga cgc cag
gtg cga gag ccc ggc cag 1596Lys Gly Arg Leu Leu Lys Ala Gly Arg Gln
Val Arg Glu Pro Gly Gln 500 505
510gac ctg gtg gtg ctg ccc ctg tcc atc acc acc gac ttc atc cct tcc
1644Asp Leu Val Val Leu Pro Leu Ser Ile Thr Thr Asp Phe Ile Pro Ser
515 520 525ttc cgc ctg gtg gcg tac tac
acg ctg atc ggt gcc agc ggc cag agg 1692Phe Arg Leu Val Ala Tyr Tyr
Thr Leu Ile Gly Ala Ser Gly Gln Arg 530 535
540gag gtg gtg gcc gac tcc gtg tgg gtg gac gtc aag gac tcc tgc gtg
1740Glu Val Val Ala Asp Ser Val Trp Val Asp Val Lys Asp Ser Cys Val545
550 555 560ggc tcg ctg gtg
gta aaa agc ggc cag tca gaa gac cgg cag cct gta 1788Gly Ser Leu Val
Val Lys Ser Gly Gln Ser Glu Asp Arg Gln Pro Val 565
570 575cct ggg cag cag atg acc ctg aag ata gag
ggt gac cac ggg gcc cgg 1836Pro Gly Gln Gln Met Thr Leu Lys Ile Glu
Gly Asp His Gly Ala Arg 580 585
590gtg gta ctg gtg gcc gtg gac aag ggc gtg ttc gtg ctg aat aag aag
1884Val Val Leu Val Ala Val Asp Lys Gly Val Phe Val Leu Asn Lys Lys
595 600 605aac aaa ctg acg cag agt aag
atc tgg gac gtg gtg gag aag gca gac 1932Asn Lys Leu Thr Gln Ser Lys
Ile Trp Asp Val Val Glu Lys Ala Asp 610 615
620atc ggc tgc acc ccg ggc agt ggg aag gat tac gcc ggt gtc ttc tcc
1980Ile Gly Cys Thr Pro Gly Ser Gly Lys Asp Tyr Ala Gly Val Phe Ser625
630 635 640gac gca ggg ctg
acc ttc acg agc agc agt ggc cag cag acc gcc cag 2028Asp Ala Gly Leu
Thr Phe Thr Ser Ser Ser Gly Gln Gln Thr Ala Gln 645
650 655agg gca gaa ctt cag tgc ccg cag cca gcc
gcc cgc cga cgc cgt tcc 2076Arg Ala Glu Leu Gln Cys Pro Gln Pro Ala
Ala Arg Arg Arg Arg Ser 660 665
670gtg cag ctc acg gag aag cga atg gac aaa gtc ggc aag tac ccc aag
2124Val Gln Leu Thr Glu Lys Arg Met Asp Lys Val Gly Lys Tyr Pro Lys
675 680 685gag ctg cgc aag tgc tgc gag
gac ggc atg cgg gag aac ccc atg agg 2172Glu Leu Arg Lys Cys Cys Glu
Asp Gly Met Arg Glu Asn Pro Met Arg 690 695
700ttc tcg tgc cag cgc cgg acc cgt ttc atc tcc ctg ggc gag gcg tgc
2220Phe Ser Cys Gln Arg Arg Thr Arg Phe Ile Ser Leu Gly Glu Ala Cys705
710 715 720aag aag gtc ttc
ctg gac tgc tgc aac tac atc aca gag ctg cgg cgg 2268Lys Lys Val Phe
Leu Asp Cys Cys Asn Tyr Ile Thr Glu Leu Arg Arg 725
730 735cag cac gcg cgg gcc agc cac ctg ggc ctg
gcc agg agt aac ctg gat 2316Gln His Ala Arg Ala Ser His Leu Gly Leu
Ala Arg Ser Asn Leu Asp 740 745
750gag gac atc att gca gaa gag aac atc gtt tcc cga agt gag ttc cca
2364Glu Asp Ile Ile Ala Glu Glu Asn Ile Val Ser Arg Ser Glu Phe Pro
755 760 765gag agc tgg ctg tgg aac gtt
gag gac ttg aaa gag cca ccg aaa aat 2412Glu Ser Trp Leu Trp Asn Val
Glu Asp Leu Lys Glu Pro Pro Lys Asn 770 775
780gga atc tct acg aag ctc atg aat ata ttt ttg aaa gac tcc atc acc
2460Gly Ile Ser Thr Lys Leu Met Asn Ile Phe Leu Lys Asp Ser Ile Thr785
790 795 800acg tgg gag att
ctg gct gtc agc atg tcg gac aag aaa ggg atc tgt 2508Thr Trp Glu Ile
Leu Ala Val Ser Met Ser Asp Lys Lys Gly Ile Cys 805
810 815gtg gca gac ccc ttc gag gtc aca gta atg
cag gac ttc ttc atc gac 2556Val Ala Asp Pro Phe Glu Val Thr Val Met
Gln Asp Phe Phe Ile Asp 820 825
830ctg cgg cta ccc tac tct gtt gtt cga aac gag cag gtg gaa atc cga
2604Leu Arg Leu Pro Tyr Ser Val Val Arg Asn Glu Gln Val Glu Ile Arg
835 840 845gcc gtt ctc tac aat tac cgg
cag aac caa gag ctc aag gtg agg gtg 2652Ala Val Leu Tyr Asn Tyr Arg
Gln Asn Gln Glu Leu Lys Val Arg Val 850 855
860gaa cta ctc cac aat cca gcc ttc tgc agc ctg gcc acc acc aag agg
2700Glu Leu Leu His Asn Pro Ala Phe Cys Ser Leu Ala Thr Thr Lys Arg865
870 875 880cgt cac cag cag
acc gta acc atc ccc ccc aag tcc tcg ttg tcc gtt 2748Arg His Gln Gln
Thr Val Thr Ile Pro Pro Lys Ser Ser Leu Ser Val 885
890 895cca tat gtc atc gtg ccg cta aag acc ggc
ctg cag gaa gtg gaa gtc 2796Pro Tyr Val Ile Val Pro Leu Lys Thr Gly
Leu Gln Glu Val Glu Val 900 905
910aag gct gcc gtc tac cat cat ttc atc agt gac ggt gtc agg aag tcc
2844Lys Ala Ala Val Tyr His His Phe Ile Ser Asp Gly Val Arg Lys Ser
915 920 925ctg aag gtc gtg ccg gaa gga
atc aga atg aac aaa act gtg gct gtt 2892Leu Lys Val Val Pro Glu Gly
Ile Arg Met Asn Lys Thr Val Ala Val 930 935
940cgc acc ctg gat cca gaa cgc ctg ggc cgt gaa gga gtg cag aaa gag
2940Arg Thr Leu Asp Pro Glu Arg Leu Gly Arg Glu Gly Val Gln Lys Glu945
950 955 960gac atc cca cct
gca gac ctc agt gac caa gtc ccg gac acc gag tct 2988Asp Ile Pro Pro
Ala Asp Leu Ser Asp Gln Val Pro Asp Thr Glu Ser 965
970 975gag acc aga att ctc ctg caa ggg acc cca
gtg gcc cag atg aca gag 3036Glu Thr Arg Ile Leu Leu Gln Gly Thr Pro
Val Ala Gln Met Thr Glu 980 985
990gat gcc gtc gac gcg gaa cgg ctg aag cac ctc att gtg acc ccc tcg
3084Asp Ala Val Asp Ala Glu Arg Leu Lys His Leu Ile Val Thr Pro Ser
995 1000 1005ggc tgc ggg gaa cag aac atg
atc ggc atg acg ccc acg gtc atc gct 3132Gly Cys Gly Glu Gln Asn Met
Ile Gly Met Thr Pro Thr Val Ile Ala 1010 1015
1020gtg cat tac ctg gat gaa acg gag cag tgg gag aag ttc ggc cta gag
3180Val His Tyr Leu Asp Glu Thr Glu Gln Trp Glu Lys Phe Gly Leu Glu1025
1030 1035 1040aag cgg cag
ggg gcc ttg gag ctc atc aag aag ggg tac acc cag cag 3228Lys Arg Gln
Gly Ala Leu Glu Leu Ile Lys Lys Gly Tyr Thr Gln Gln 1045
1050 1055ctg gcc ttc aga caa ccc agc tct gcc
ttt gcg gcc ttc gtg aaa cgg 3276Leu Ala Phe Arg Gln Pro Ser Ser Ala
Phe Ala Ala Phe Val Lys Arg 1060 1065
1070gca ccc agc acc tgg ctg acc gcc tac gtg gtc aag gtc ttc tct ctg
3324Ala Pro Ser Thr Trp Leu Thr Ala Tyr Val Val Lys Val Phe Ser Leu
1075 1080 1085gct gtc aac ctc atc gcc atc
gac tcc caa gtc ctc tgc ggg gct gtt 3372Ala Val Asn Leu Ile Ala Ile
Asp Ser Gln Val Leu Cys Gly Ala Val 1090 1095
1100aaa tgg ctg atc ctg gag aag cag aag ccc gac ggg gtc ttc cag gag
3420Lys Trp Leu Ile Leu Glu Lys Gln Lys Pro Asp Gly Val Phe Gln Glu1105
1110 1115 1120gat gcg ccc
gtg ata cac caa gaa atg att ggt gga tta cgg aac aac 3468Asp Ala Pro
Val Ile His Gln Glu Met Ile Gly Gly Leu Arg Asn Asn 1125
1130 1135aac gag aaa gac atg gcc ctc acg gcc
ttt gtt ctc atc tcg ctg cag 3516Asn Glu Lys Asp Met Ala Leu Thr Ala
Phe Val Leu Ile Ser Leu Gln 1140 1145
1150gag gct aaa gat att tgc gag gag cag gtc aac agc ctg cca ggc agc
3564Glu Ala Lys Asp Ile Cys Glu Glu Gln Val Asn Ser Leu Pro Gly Ser
1155 1160 1165atc act aaa gca gga gac ttc
ctt gaa gcc aac tac atg aac cta cag 3612Ile Thr Lys Ala Gly Asp Phe
Leu Glu Ala Asn Tyr Met Asn Leu Gln 1170 1175
1180aga tcc tac act gtg gcc att gct ggc tat gct ctg gcc cag atg ggc
3660Arg Ser Tyr Thr Val Ala Ile Ala Gly Tyr Ala Leu Ala Gln Met Gly1185
1190 1195 1200agg ctg aag
ggg cct ctt ctt aac aaa ttt ctg acc aca gcc aaa gat 3708Arg Leu Lys
Gly Pro Leu Leu Asn Lys Phe Leu Thr Thr Ala Lys Asp 1205
1210 1215aag aac cgc tgg gag gac cct ggt aag
cag ctc tac aac gtg gag gcc 3756Lys Asn Arg Trp Glu Asp Pro Gly Lys
Gln Leu Tyr Asn Val Glu Ala 1220 1225
1230aca tcc tat gcc ctc ttg gcc cta ctg cag cta aaa gac ttt gac ttt
3804Thr Ser Tyr Ala Leu Leu Ala Leu Leu Gln Leu Lys Asp Phe Asp Phe
1235 1240 1245gtg cct ccc gtc gtg cgt tgg
ctc aat gaa cag aga tac tac ggt ggt 3852Val Pro Pro Val Val Arg Trp
Leu Asn Glu Gln Arg Tyr Tyr Gly Gly 1250 1255
1260ggc tat ggc tct acc cag gcc acc ttc atg gtg ttc caa gcc ttg gct
3900Gly Tyr Gly Ser Thr Gln Ala Thr Phe Met Val Phe Gln Ala Leu Ala1265
1270 1275 1280caa tac caa
aag gac gcc cct gac cac cag gaa ctg aac ctt gat gtg 3948Gln Tyr Gln
Lys Asp Ala Pro Asp His Gln Glu Leu Asn Leu Asp Val 1285
1290 1295tcc ctc caa ctg ccc agc cgc agc tcc
aag atc acc cac cgt atc cac 3996Ser Leu Gln Leu Pro Ser Arg Ser Ser
Lys Ile Thr His Arg Ile His 1300 1305
1310tgg gaa tct gcc agc ctc ctg cga tca gaa gag acc aag gaa aat gag
4044Trp Glu Ser Ala Ser Leu Leu Arg Ser Glu Glu Thr Lys Glu Asn Glu
1315 1320 1325ggt ttc aca gtc aca gct gaa
gga aaa ggc caa ggc acc ttg tcg gtg 4092Gly Phe Thr Val Thr Ala Glu
Gly Lys Gly Gln Gly Thr Leu Ser Val 1330 1335
1340gtg aca atg tac cat gct aag gcc aaa gat caa ctc acc tgt aat aaa
4140Val Thr Met Tyr His Ala Lys Ala Lys Asp Gln Leu Thr Cys Asn Lys1345
1350 1355 1360ttc gac ctc
aag gtc acc ata aaa cca gca ccg gaa aca gaa aag agg 4188Phe Asp Leu
Lys Val Thr Ile Lys Pro Ala Pro Glu Thr Glu Lys Arg 1365
1370 1375cct cag gat gcc aag aac act atg atc
ctt gag atc tgt acc agg tac 4236Pro Gln Asp Ala Lys Asn Thr Met Ile
Leu Glu Ile Cys Thr Arg Tyr 1380 1385
1390cgg gga gac cag gat gcc act atg tct ata ttg gac ata tcc atg atg
4284Arg Gly Asp Gln Asp Ala Thr Met Ser Ile Leu Asp Ile Ser Met Met
1395 1400 1405act ggc ttt gct cca gac aca
gat gac ctg aag cag ctg gcc aat ggt 4332Thr Gly Phe Ala Pro Asp Thr
Asp Asp Leu Lys Gln Leu Ala Asn Gly 1410 1415
1420gtt gac aga tac atc tcc aag tat gag ctg gac aaa gcc ttc tcc gat
4380Val Asp Arg Tyr Ile Ser Lys Tyr Glu Leu Asp Lys Ala Phe Ser Asp1425
1430 1435 1440agg aac acc
ctc atc atc tac ctg gac aag gtc tca cac tct gag gat 4428Arg Asn Thr
Leu Ile Ile Tyr Leu Asp Lys Val Ser His Ser Glu Asp 1445
1450 1455gac tgt cta gct ttc aaa gtt cac caa
tac ttt aat gta gag ctt atc 4476Asp Cys Leu Ala Phe Lys Val His Gln
Tyr Phe Asn Val Glu Leu Ile 1460 1465
1470cag cct gga gca gtc aag gtc tac gcc tat tac aac ctg gag gaa agc
4524Gln Pro Gly Ala Val Lys Val Tyr Ala Tyr Tyr Asn Leu Glu Glu Ser
1475 1480 1485tgt acc cgg ttc tac cat ccg
gaa aag gag gat gga aag ctg aac aag 4572Cys Thr Arg Phe Tyr His Pro
Glu Lys Glu Asp Gly Lys Leu Asn Lys 1490 1495
1500ctc tgc cgt gat gaa ctg tgc cgc tgt gct gag gag aat tgc ttc ata
4620Leu Cys Arg Asp Glu Leu Cys Arg Cys Ala Glu Glu Asn Cys Phe Ile1505
1510 1515 1520caa aag tcg
gat gac aag gtc acc ctg gaa gaa cgg ctg gac aag gcc 4668Gln Lys Ser
Asp Asp Lys Val Thr Leu Glu Glu Arg Leu Asp Lys Ala 1525
1530 1535tgt gag cca gga gtg gac tat gtg tac
aag acc cga ctg gtc aag gtt 4716Cys Glu Pro Gly Val Asp Tyr Val Tyr
Lys Thr Arg Leu Val Lys Val 1540 1545
1550cag ctg tcc aat gac ttt gac gag tac atc atg gcc att gag cag acc
4764Gln Leu Ser Asn Asp Phe Asp Glu Tyr Ile Met Ala Ile Glu Gln Thr
1555 1560 1565atc aag tca ggc tcg gat gag
gtg cag gtt gga cag cag cgc acg ttc 4812Ile Lys Ser Gly Ser Asp Glu
Val Gln Val Gly Gln Gln Arg Thr Phe 1570 1575
1580atc agc ccc atc aag tgc aga gaa gcc ctg aag ctg gag gag aag aaa
4860Ile Ser Pro Ile Lys Cys Arg Glu Ala Leu Lys Leu Glu Glu Lys Lys1585
1590 1595 1600cac tac ctc
atg tgg ggt ctc tcc tcc gat ttc tgg gga gag aag ccc 4908His Tyr Leu
Met Trp Gly Leu Ser Ser Asp Phe Trp Gly Glu Lys Pro 1605
1610 1615aac ctc agc tac atc atc ggg aag gac
act tgg gtg gag cac tgg cct 4956Asn Leu Ser Tyr Ile Ile Gly Lys Asp
Thr Trp Val Glu His Trp Pro 1620 1625
1630gag gag gac gaa tgc caa gac gaa gag aac cag aaa caa tgc cag gac
5004Glu Glu Asp Glu Cys Gln Asp Glu Glu Asn Gln Lys Gln Cys Gln Asp
1635 1640 1645ctc ggc gcc ttc acc gag agc
atg gtt gtc ttt ggg tgc ccc aac tga 5052Leu Gly Ala Phe Thr Glu Ser
Met Val Val Phe Gly Cys Pro Asn * 1650 1655
1660ccacaccccc attcc
506721663PRTHomo sapiens 2Met Gly Pro Thr Ser Gly Pro Ser Leu Leu Leu
Leu Leu Leu Thr His1 5 10
15Leu Pro Leu Ala Leu Gly Ser Pro Met Tyr Ser Ile Ile Thr Pro Asn
20 25 30Ile Leu Arg Leu Glu Ser Glu
Glu Thr Met Val Leu Glu Ala His Asp 35 40
45Ala Gln Gly Asp Val Pro Val Thr Val Thr Val His Asp Phe Pro Gly
50 55 60Lys Lys Leu Val Leu Ser Ser
Glu Lys Thr Val Leu Thr Pro Ala Thr65 70
75 80Asn His Met Gly Asn Val Thr Phe Thr Ile Pro Ala
Asn Arg Glu Phe 85 90
95Lys Ser Glu Lys Gly Arg Asn Lys Phe Val Thr Val Gln Ala Thr Phe
100 105 110Gly Thr Gln Val Val Glu Lys
Val Val Leu Val Ser Leu Gln Ser Gly 115 120
125Tyr Leu Phe Ile Gln Thr Asp Lys Thr Ile Tyr Thr Pro Gly Ser
Thr 130 135 140Val Leu Tyr Arg Ile Phe
Thr Val Asn His Lys Leu Leu Pro Val Gly145 150
155 160Arg Thr Val Met Val Asn Ile Glu Asn Pro Glu
Gly Ile Pro Val Lys 165 170
175Gln Asp Ser Leu Ser Ser Gln Asn Gln Leu Gly Val Leu Pro Leu Ser
180 185 190Trp Asp Ile Pro Glu Leu Val
Asn Met Gly Gln Trp Lys Ile Arg Ala 195 200
205Tyr Tyr Glu Asn Ser Pro Gln Gln Val Phe Ser Thr Glu Phe Glu
Val 210 215 220Lys Glu Tyr Val Leu Pro
Ser Phe Glu Val Ile Val Glu Pro Thr Glu225 230
235 240Lys Phe Tyr Tyr Ile Tyr Asn Glu Lys Gly Leu
Glu Val Thr Ile Thr 245 250
255Ala Arg Phe Leu Tyr Gly Lys Lys Val Glu Gly Thr Ala Phe Val Ile
260 265 270Phe Gly Ile Gln Asp Gly Glu
Gln Arg Ile Ser Leu Pro Glu Ser Leu 275 280
285Lys Arg Ile Pro Ile Glu Asp Gly Ser Gly Glu Val Val Leu Ser
Arg 290 295 300Lys Val Leu Leu Asp Gly
Val Gln Asn Leu Arg Ala Glu Asp Leu Val305 310
315 320Gly Lys Ser Leu Tyr Val Ser Ala Thr Val Ile
Leu His Ser Gly Ser 325 330
335Asp Met Val Gln Ala Glu Arg Ser Gly Ile Pro Ile Val Thr Ser Pro
340 345 350Tyr Gln Ile His Phe Thr Lys
Thr Pro Lys Tyr Phe Lys Pro Gly Met 355 360
365Pro Phe Asp Leu Met Val Phe Val Thr Asn Pro Asp Gly Ser Pro
Ala 370 375 380Tyr Arg Val Pro Val Ala
Val Gln Gly Glu Asp Thr Val Gln Ser Leu385 390
395 400Thr Gln Gly Asp Gly Val Ala Lys Leu Ser Ile
Asn Thr His Pro Ser 405 410
415Gln Lys Pro Leu Ser Ile Thr Val Arg Thr Lys Lys Gln Glu Leu Ser
420 425 430Glu Ala Glu Gln Ala Thr Arg
Thr Met Gln Ala Leu Pro Tyr Ser Thr 435 440
445Val Gly Asn Ser Asn Asn Tyr Leu His Leu Ser Val Leu Arg Thr
Glu 450 455 460Leu Arg Pro Gly Glu Thr
Leu Asn Val Asn Phe Leu Leu Arg Met Asp465 470
475 480Arg Ala His Glu Ala Lys Ile Arg Tyr Tyr Thr
Tyr Leu Ile Met Asn 485 490
495Lys Gly Arg Leu Leu Lys Ala Gly Arg Gln Val Arg Glu Pro Gly Gln
500 505 510Asp Leu Val Val Leu Pro Leu
Ser Ile Thr Thr Asp Phe Ile Pro Ser 515 520
525Phe Arg Leu Val Ala Tyr Tyr Thr Leu Ile Gly Ala Ser Gly Gln
Arg 530 535 540Glu Val Val Ala Asp Ser
Val Trp Val Asp Val Lys Asp Ser Cys Val545 550
555 560Gly Ser Leu Val Val Lys Ser Gly Gln Ser Glu
Asp Arg Gln Pro Val 565 570
575Pro Gly Gln Gln Met Thr Leu Lys Ile Glu Gly Asp His Gly Ala Arg
580 585 590Val Val Leu Val Ala Val Asp
Lys Gly Val Phe Val Leu Asn Lys Lys 595 600
605Asn Lys Leu Thr Gln Ser Lys Ile Trp Asp Val Val Glu Lys Ala
Asp 610 615 620Ile Gly Cys Thr Pro Gly
Ser Gly Lys Asp Tyr Ala Gly Val Phe Ser625 630
635 640Asp Ala Gly Leu Thr Phe Thr Ser Ser Ser Gly
Gln Gln Thr Ala Gln 645 650
655Arg Ala Glu Leu Gln Cys Pro Gln Pro Ala Ala Arg Arg Arg Arg Ser
660 665 670Val Gln Leu Thr Glu Lys Arg
Met Asp Lys Val Gly Lys Tyr Pro Lys 675 680
685Glu Leu Arg Lys Cys Cys Glu Asp Gly Met Arg Glu Asn Pro Met
Arg 690 695 700Phe Ser Cys Gln Arg Arg
Thr Arg Phe Ile Ser Leu Gly Glu Ala Cys705 710
715 720Lys Lys Val Phe Leu Asp Cys Cys Asn Tyr Ile
Thr Glu Leu Arg Arg 725 730
735Gln His Ala Arg Ala Ser His Leu Gly Leu Ala Arg Ser Asn Leu Asp
740 745 750Glu Asp Ile Ile Ala Glu Glu
Asn Ile Val Ser Arg Ser Glu Phe Pro 755 760
765Glu Ser Trp Leu Trp Asn Val Glu Asp Leu Lys Glu Pro Pro Lys
Asn 770 775 780Gly Ile Ser Thr Lys Leu
Met Asn Ile Phe Leu Lys Asp Ser Ile Thr785 790
795 800Thr Trp Glu Ile Leu Ala Val Ser Met Ser Asp
Lys Lys Gly Ile Cys 805 810
815Val Ala Asp Pro Phe Glu Val Thr Val Met Gln Asp Phe Phe Ile Asp
820 825 830Leu Arg Leu Pro Tyr Ser Val
Val Arg Asn Glu Gln Val Glu Ile Arg 835 840
845Ala Val Leu Tyr Asn Tyr Arg Gln Asn Gln Glu Leu Lys Val Arg
Val 850 855 860Glu Leu Leu His Asn Pro
Ala Phe Cys Ser Leu Ala Thr Thr Lys Arg865 870
875 880Arg His Gln Gln Thr Val Thr Ile Pro Pro Lys
Ser Ser Leu Ser Val 885 890
895Pro Tyr Val Ile Val Pro Leu Lys Thr Gly Leu Gln Glu Val Glu Val
900 905 910Lys Ala Ala Val Tyr His His
Phe Ile Ser Asp Gly Val Arg Lys Ser 915 920
925Leu Lys Val Val Pro Glu Gly Ile Arg Met Asn Lys Thr Val Ala
Val 930 935 940Arg Thr Leu Asp Pro Glu
Arg Leu Gly Arg Glu Gly Val Gln Lys Glu945 950
955 960Asp Ile Pro Pro Ala Asp Leu Ser Asp Gln Val
Pro Asp Thr Glu Ser 965 970
975Glu Thr Arg Ile Leu Leu Gln Gly Thr Pro Val Ala Gln Met Thr Glu
980 985 990Asp Ala Val Asp Ala Glu Arg
Leu Lys His Leu Ile Val Thr Pro Ser 995 1000
1005Gly Cys Gly Glu Gln Asn Met Ile Gly Met Thr Pro Thr Val Ile
Ala 1010 1015 1020Val His Tyr Leu Asp Glu
Thr Glu Gln Trp Glu Lys Phe Gly Leu Glu1025 1030
1035 1040Lys Arg Gln Gly Ala Leu Glu Leu Ile Lys Lys
Gly Tyr Thr Gln Gln 1045 1050
1055Leu Ala Phe Arg Gln Pro Ser Ser Ala Phe Ala Ala Phe Val Lys Arg
1060 1065 1070Ala Pro Ser Thr Trp Leu
Thr Ala Tyr Val Val Lys Val Phe Ser Leu 1075 1080
1085Ala Val Asn Leu Ile Ala Ile Asp Ser Gln Val Leu Cys Gly
Ala Val 1090 1095 1100Lys Trp Leu Ile Leu
Glu Lys Gln Lys Pro Asp Gly Val Phe Gln Glu1105 1110
1115 1120Asp Ala Pro Val Ile His Gln Glu Met Ile
Gly Gly Leu Arg Asn Asn 1125 1130
1135Asn Glu Lys Asp Met Ala Leu Thr Ala Phe Val Leu Ile Ser Leu Gln
1140 1145 1150Glu Ala Lys Asp Ile
Cys Glu Glu Gln Val Asn Ser Leu Pro Gly Ser 1155
1160 1165Ile Thr Lys Ala Gly Asp Phe Leu Glu Ala Asn Tyr
Met Asn Leu Gln 1170 1175 1180Arg Ser Tyr
Thr Val Ala Ile Ala Gly Tyr Ala Leu Ala Gln Met Gly1185
1190 1195 1200Arg Leu Lys Gly Pro Leu Leu
Asn Lys Phe Leu Thr Thr Ala Lys Asp 1205
1210 1215Lys Asn Arg Trp Glu Asp Pro Gly Lys Gln Leu Tyr
Asn Val Glu Ala 1220 1225 1230Thr
Ser Tyr Ala Leu Leu Ala Leu Leu Gln Leu Lys Asp Phe Asp Phe 1235
1240 1245Val Pro Pro Val Val Arg Trp Leu Asn
Glu Gln Arg Tyr Tyr Gly Gly 1250 1255
1260Gly Tyr Gly Ser Thr Gln Ala Thr Phe Met Val Phe Gln Ala Leu Ala1265
1270 1275 1280Gln Tyr Gln Lys
Asp Ala Pro Asp His Gln Glu Leu Asn Leu Asp Val 1285
1290 1295Ser Leu Gln Leu Pro Ser Arg Ser Ser Lys
Ile Thr His Arg Ile His 1300 1305
1310Trp Glu Ser Ala Ser Leu Leu Arg Ser Glu Glu Thr Lys Glu Asn Glu
1315 1320 1325Gly Phe Thr Val Thr Ala Glu
Gly Lys Gly Gln Gly Thr Leu Ser Val 1330 1335
1340Val Thr Met Tyr His Ala Lys Ala Lys Asp Gln Leu Thr Cys Asn
Lys1345 1350 1355 1360Phe
Asp Leu Lys Val Thr Ile Lys Pro Ala Pro Glu Thr Glu Lys Arg
1365 1370 1375Pro Gln Asp Ala Lys Asn Thr
Met Ile Leu Glu Ile Cys Thr Arg Tyr 1380 1385
1390Arg Gly Asp Gln Asp Ala Thr Met Ser Ile Leu Asp Ile Ser
Met Met 1395 1400 1405Thr Gly Phe Ala
Pro Asp Thr Asp Asp Leu Lys Gln Leu Ala Asn Gly 1410
1415 1420Val Asp Arg Tyr Ile Ser Lys Tyr Glu Leu Asp Lys
Ala Phe Ser Asp1425 1430 1435
1440Arg Asn Thr Leu Ile Ile Tyr Leu Asp Lys Val Ser His Ser Glu Asp
1445 1450 1455Asp Cys Leu Ala Phe
Lys Val His Gln Tyr Phe Asn Val Glu Leu Ile 1460
1465 1470Gln Pro Gly Ala Val Lys Val Tyr Ala Tyr Tyr Asn
Leu Glu Glu Ser 1475 1480 1485Cys Thr
Arg Phe Tyr His Pro Glu Lys Glu Asp Gly Lys Leu Asn Lys 1490
1495 1500Leu Cys Arg Asp Glu Leu Cys Arg Cys Ala Glu
Glu Asn Cys Phe Ile1505 1510 1515
1520Gln Lys Ser Asp Asp Lys Val Thr Leu Glu Glu Arg Leu Asp Lys Ala
1525 1530 1535Cys Glu Pro Gly
Val Asp Tyr Val Tyr Lys Thr Arg Leu Val Lys Val 1540
1545 1550Gln Leu Ser Asn Asp Phe Asp Glu Tyr Ile Met
Ala Ile Glu Gln Thr 1555 1560 1565Ile
Lys Ser Gly Ser Asp Glu Val Gln Val Gly Gln Gln Arg Thr Phe 1570
1575 1580Ile Ser Pro Ile Lys Cys Arg Glu Ala Leu
Lys Leu Glu Glu Lys Lys1585 1590 1595
1600His Tyr Leu Met Trp Gly Leu Ser Ser Asp Phe Trp Gly Glu Lys
Pro 1605 1610 1615Asn Leu Ser
Tyr Ile Ile Gly Lys Asp Thr Trp Val Glu His Trp Pro 1620
1625 1630Glu Glu Asp Glu Cys Gln Asp Glu Glu Asn
Gln Lys Gln Cys Gln Asp 1635 1640
1645Leu Gly Ala Phe Thr Glu Ser Met Val Val Phe Gly Cys Pro Asn 1650
1655 166035948DNANaja kaouthiaCDS(4)...(4932)
3ccc atg gag agg atg gct ctc tat ctg gtg gct gct cta ttg att ggt
48Met Glu Arg Met Ala Leu Tyr Leu Val Ala Ala Leu Leu Ile Gly 1
5 10 15ttt cca ggg tct tct cat ggg
gct ctc tac acc ctc atc acc cct gct 96Phe Pro Gly Ser Ser His Gly
Ala Leu Tyr Thr Leu Ile Thr Pro Ala 20 25
30gtt ttg cga aca gac aca gaa gag caa att ttg gtg gag gcc
cat gga 144Val Leu Arg Thr Asp Thr Glu Glu Gln Ile Leu Val Glu Ala
His Gly 35 40 45gac agt act cca
aaa cag ctt gac atc ttt gtt cat gat ttt cca cgg 192Asp Ser Thr Pro
Lys Gln Leu Asp Ile Phe Val His Asp Phe Pro Arg 50
55 60aag cag aaa acc ttg ttc caa acc aga gta gat atg aat
cca gca gga 240Lys Gln Lys Thr Leu Phe Gln Thr Arg Val Asp Met Asn
Pro Ala Gly 65 70 75ggc atg ctt gtc act
cca act ata gag att cca gca aaa gaa gtg agt 288Gly Met Leu Val Thr
Pro Thr Ile Glu Ile Pro Ala Lys Glu Val Ser80 85
90 95acg gac tcc agg caa aat caa tat gtg gtt
gtg caa gta act ggt cct 336Thr Asp Ser Arg Gln Asn Gln Tyr Val Val
Val Gln Val Thr Gly Pro 100 105
110caa gtg aga ttg gaa aag gtg gtt ctc ctt tct tac cag agt agc ttt
384Gln Val Arg Leu Glu Lys Val Val Leu Leu Ser Tyr Gln Ser Ser Phe
115 120 125ctg ttt atc cag aca gat aaa
ggc atc tat aca cca ggg tct cca gta 432Leu Phe Ile Gln Thr Asp Lys
Gly Ile Tyr Thr Pro Gly Ser Pro Val 130 135
140ctc tat cgt gtt ttt tct atg gat cac aac aca agc aag atg aac aaa
480Leu Tyr Arg Val Phe Ser Met Asp His Asn Thr Ser Lys Met Asn Lys145
150 155act gtg att gtt gag ttt cag act cca
gaa ggc att ctt gtc agt tct 528Thr Val Ile Val Glu Phe Gln Thr Pro
Glu Gly Ile Leu Val Ser Ser160 165 170
175aat tca gtt gac cta aac ttc ttc tgg cct tac aat tta cca
gac ctt 576Asn Ser Val Asp Leu Asn Phe Phe Trp Pro Tyr Asn Leu Pro
Asp Leu 180 185 190gtc agt ttg
ggg act tgg agg att gtg gcc aaa tat gaa cat tcc cca 624Val Ser Leu
Gly Thr Trp Arg Ile Val Ala Lys Tyr Glu His Ser Pro 195
200 205gag aat tat act gca tat ttt gat gtc agg aaa
tat gtg ttg cca agc 672Glu Asn Tyr Thr Ala Tyr Phe Asp Val Arg Lys
Tyr Val Leu Pro Ser 210 215 220ttt gaa
gtc cgt ctg caa cca tca gag aag ttt ttt tac att gac ggc 720Phe Glu
Val Arg Leu Gln Pro Ser Glu Lys Phe Phe Tyr Ile Asp Gly225
230 235aat gaa aat ttc cac gtg tct atc act gca agg tac
ttg tat gga gag 768Asn Glu Asn Phe His Val Ser Ile Thr Ala Arg Tyr
Leu Tyr Gly Glu240 245 250
255gaa gtg gaa ggt gtg gcc ttt gtc ctc ttt gga gtg aaa ata gat gat
816Glu Val Glu Gly Val Ala Phe Val Leu Phe Gly Val Lys Ile Asp Asp
260 265 270gct aaa aag agt att cca
gac tca ctc acg aga att ccg att att gat 864Ala Lys Lys Ser Ile Pro
Asp Ser Leu Thr Arg Ile Pro Ile Ile Asp 275 280
285gga gat ggg aaa gca aca cta aaa aga gat aca ttc cgt tct
cga ttt 912Gly Asp Gly Lys Ala Thr Leu Lys Arg Asp Thr Phe Arg Ser
Arg Phe 290 295 300cca aat ctc aat gag
ctt gtt ggg cat act ctg tat gca tct gta aca 960Pro Asn Leu Asn Glu
Leu Val Gly His Thr Leu Tyr Ala Ser Val Thr305 310
315gtc atg aca gaa tca ggc agt gat atg gta gtg act gag caa agc
ggc 1008Val Met Thr Glu Ser Gly Ser Asp Met Val Val Thr Glu Gln Ser
Gly320 325 330 335att cat
att gtg gca tct ccc tat cag atc cac ttc aca aaa acc ccc 1056Ile His
Ile Val Ala Ser Pro Tyr Gln Ile His Phe Thr Lys Thr Pro 340
345 350aaa tat ttc aag cca gga atg cca tat
gaa ctg acg gtg tat gtt acc 1104Lys Tyr Phe Lys Pro Gly Met Pro Tyr
Glu Leu Thr Val Tyr Val Thr 355 360
365aac cct gat ggc tca cca gct gcc cat gtg cca gtg gta tca gag gcc
1152Asn Pro Asp Gly Ser Pro Ala Ala His Val Pro Val Val Ser Glu Ala
370 375 380ttt cat tct atg gga acc act
ttg agt gat ggg act gct aag ctc atc 1200Phe His Ser Met Gly Thr Thr
Leu Ser Asp Gly Thr Ala Lys Leu Ile385 390
395ctg aac ata cca ttg aat gct caa agc cta cca atc act gtt aga act
1248Leu Asn Ile Pro Leu Asn Ala Gln Ser Leu Pro Ile Thr Val Arg Thr400
405 410 415aac cat gga gac
ctc cca aga gaa cgc cag gca aca aag tcc atg aca 1296Asn His Gly Asp
Leu Pro Arg Glu Arg Gln Ala Thr Lys Ser Met Thr 420
425 430gcc ata gcc tac caa acc cag gga gga tct gga
aac tat ctt cat gta 1344Ala Ile Ala Tyr Gln Thr Gln Gly Gly Ser Gly
Asn Tyr Leu His Val 435 440 445gcc
att aca tct aca gag att aag ccc gga gat aac tta cct gtc aat 1392Ala
Ile Thr Ser Thr Glu Ile Lys Pro Gly Asp Asn Leu Pro Val Asn 450
455 460ttc aat gtg aag ggc aat gca aat tca ctg
aag cag atc aaa tat ttc 1440Phe Asn Val Lys Gly Asn Ala Asn Ser Leu
Lys Gln Ile Lys Tyr Phe465 470 475aca tac
ctc ata ttg aat aaa ggg aag att ttc aag gtt ggc agg caa 1488Thr Tyr
Leu Ile Leu Asn Lys Gly Lys Ile Phe Lys Val Gly Arg Gln480
485 490 495ccc agg aga gat ggg cag aat
ctg gtg acc atg aat ctg cat atc act 1536Pro Arg Arg Asp Gly Gln Asn
Leu Val Thr Met Asn Leu His Ile Thr 500 505
510cca gat ctc atc cct tcc ttc cgg ttt gtg gct tac tac caa
gtg gga 1584Pro Asp Leu Ile Pro Ser Phe Arg Phe Val Ala Tyr Tyr Gln
Val Gly 515 520 525aac aac gaa att
gtg gct gat tct gtc tgg gtg gat gtg aag gat acc 1632Asn Asn Glu Ile
Val Ala Asp Ser Val Trp Val Asp Val Lys Asp Thr 530
535 540tgc atg gga acg ttg gtt gtg aaa gga gac aat cta
ata caa atg cca 1680Cys Met Gly Thr Leu Val Val Lys Gly Asp Asn Leu
Ile Gln Met Pro545 550 555gga gct gca atg
aaa atc aaa ttg gaa ggg gat cca ggt gct cgg gtt 1728Gly Ala Ala Met
Lys Ile Lys Leu Glu Gly Asp Pro Gly Ala Arg Val560 565
570 575ggt ctt gtg gct gtg gac aaa gca gta
tat gtt ctc aat gat aaa tat 1776Gly Leu Val Ala Val Asp Lys Ala Val
Tyr Val Leu Asn Asp Lys Tyr 580 585
590aag att agc caa gct aag ata tgg gac aca ata gaa aag agt gac ttt
1824Lys Ile Ser Gln Ala Lys Ile Trp Asp Thr Ile Glu Lys Ser Asp Phe
595 600 605ggc tgt aca gct ggc agt ggc
cag aat aat ctg ggt gtg ttt gaa gat 1872Gly Cys Thr Ala Gly Ser Gly
Gln Asn Asn Leu Gly Val Phe Glu Asp 610 615
620gct gga ctg gct ctg aca acc agc act aat ctc aac acc aaa cag aga
1920Ala Gly Leu Ala Leu Thr Thr Ser Thr Asn Leu Asn Thr Lys Gln Arg625
630 635tca gct gca aag tgt cct cag cct gca
aat cgg agg cgt cgc agt tct 1968Ser Ala Ala Lys Cys Pro Gln Pro Ala
Asn Arg Arg Arg Arg Ser Ser640 645 650
655gtt ttg ctg ctt gac agc aac gca agc aaa gcg gca gaa ttt
cag gat 2016Val Leu Leu Leu Asp Ser Asn Ala Ser Lys Ala Ala Glu Phe
Gln Asp 660 665 670caa gac ctg
cgt aaa tgc tgt gaa gat gtc atg cat gag aac ccc atg 2064Gln Asp Leu
Arg Lys Cys Cys Glu Asp Val Met His Glu Asn Pro Met 675
680 685ggg tac act tgt gaa aag cgt gca aaa tac atc
cag gag gga gat gct 2112Gly Tyr Thr Cys Glu Lys Arg Ala Lys Tyr Ile
Gln Glu Gly Asp Ala 690 695 700tgt aag
gct gcc ttc ctt gaa tgc tgt cgc tac atc aag ggg gtc cga 2160Cys Lys
Ala Ala Phe Leu Glu Cys Cys Arg Tyr Ile Lys Gly Val Arg705
710 715gat gaa aac caa cgg gag agc gag ttg ttt ctg gca
aga gat gat aat 2208Asp Glu Asn Gln Arg Glu Ser Glu Leu Phe Leu Ala
Arg Asp Asp Asn720 725 730
735gaa gat ggt ttc ata gca gat agt gat atc atc tca agg tct gat ttc
2256Glu Asp Gly Phe Ile Ala Asp Ser Asp Ile Ile Ser Arg Ser Asp Phe
740 745 750ccc aag agt tgg ttg tgg
cta aca aag gac ttg acc gag gag cct aac 2304Pro Lys Ser Trp Leu Trp
Leu Thr Lys Asp Leu Thr Glu Glu Pro Asn 755 760
765agt caa ggg att tca agc aag aca atg tct ttt tat ctg agg
gat tcc 2352Ser Gln Gly Ile Ser Ser Lys Thr Met Ser Phe Tyr Leu Arg
Asp Ser 770 775 780atc aca acc tgg gtg
gtg ctg gct gta agc ttt aca ccc acc aaa ggg 2400Ile Thr Thr Trp Val
Val Leu Ala Val Ser Phe Thr Pro Thr Lys Gly785 790
795atc tgt gtg gct gaa cct tat gaa ata aga gtc atg aaa gtc ttc
ttc 2448Ile Cys Val Ala Glu Pro Tyr Glu Ile Arg Val Met Lys Val Phe
Phe800 805 810 815att gat
ctt caa atg cca tat tca gta gtg aag aat gag cag gtg gag 2496Ile Asp
Leu Gln Met Pro Tyr Ser Val Val Lys Asn Glu Gln Val Glu 820
825 830att cga gct att ctg cac aac tac gtt
aac gag gat att tat gtg cga 2544Ile Arg Ala Ile Leu His Asn Tyr Val
Asn Glu Asp Ile Tyr Val Arg 835 840
845gtg gaa ctg tta tac aac cca gcc ttc tgc agt gct tcc aca aaa gga
2592Val Glu Leu Leu Tyr Asn Pro Ala Phe Cys Ser Ala Ser Thr Lys Gly
850 855 860caa aga tac cga cag cag ttc
cca att aaa gcc ctg tcc tcc aga gca 2640Gln Arg Tyr Arg Gln Gln Phe
Pro Ile Lys Ala Leu Ser Ser Arg Ala865 870
875gta ccg ttt gtg ata gtc cca tta gag caa gga ttg cat gat gtt gag
2688Val Pro Phe Val Ile Val Pro Leu Glu Gln Gly Leu His Asp Val Glu880
885 890 895att aaa gca agt
gtc cag gaa gcg ttg tgg tca gac ggt gtg agg aag 2736Ile Lys Ala Ser
Val Gln Glu Ala Leu Trp Ser Asp Gly Val Arg Lys 900
905 910aaa ctg aaa gtt gta cct gaa ggg gta cag aaa
tcc att gtg act att 2784Lys Leu Lys Val Val Pro Glu Gly Val Gln Lys
Ser Ile Val Thr Ile 915 920 925gtt
aaa ctg gac cca agg gca aaa gga gtt ggt gga aca cag cta gaa 2832Val
Lys Leu Asp Pro Arg Ala Lys Gly Val Gly Gly Thr Gln Leu Glu 930
935 940gtg atc aaa gcc cgc aaa tta gat gac aga
gtg cct gac aca gaa att 2880Val Ile Lys Ala Arg Lys Leu Asp Asp Arg
Val Pro Asp Thr Glu Ile945 950 955gaa acc
aag att atc atc caa ggt gac cct gtg gct cag att att gaa 2928Glu Thr
Lys Ile Ile Ile Gln Gly Asp Pro Val Ala Gln Ile Ile Glu960
965 970 975aac tca att gat gga agt aaa
ctc aac cat ctc att atc act cct tct 2976Asn Ser Ile Asp Gly Ser Lys
Leu Asn His Leu Ile Ile Thr Pro Ser 980 985
990ggc tgt ggg gag caa aat atg atc cgc atg gcc gca cca gtt
att gcc 3024Gly Cys Gly Glu Gln Asn Met Ile Arg Met Ala Ala Pro Val
Ile Ala 995 1000 1005acc tac tac
ctg gac acc aca gag cag tgg gag act ctc ggc ata aat 3072Thr Tyr Tyr
Leu Asp Thr Thr Glu Gln Trp Glu Thr Leu Gly Ile Asn 1010
1015 1020cgc agg act gaa gct gtc aat cag atc gtg act ggt
tat gcc cag cag 3120Arg Arg Thr Glu Ala Val Asn Gln Ile Val Thr Gly
Tyr Ala Gln Gln1025 1030 1035atg gtg tac
aag aaa gca gat cat tcc tat gca gca ttt aca aac cgt 3168Met Val Tyr
Lys Lys Ala Asp His Ser Tyr Ala Ala Phe Thr Asn Arg1040
1045 1050 1055gca tct agt tct tgg cta aca
gca tat gtc gta aaa gtc ttt gcc atg 3216Ala Ser Ser Ser Trp Leu Thr
Ala Tyr Val Val Lys Val Phe Ala Met 1060 1065
1070gct gcc aaa atg gta gca ggc att agt cat gaa atc att tgt
gga ggt 3264Ala Ala Lys Met Val Ala Gly Ile Ser His Glu Ile Ile Cys
Gly Gly 1075 1080 1085gtg agg tgg
ctg att ctg aac agg caa caa cca gat gga gcg ttc aaa 3312Val Arg Trp
Leu Ile Leu Asn Arg Gln Gln Pro Asp Gly Ala Phe Lys 1090
1095 1100gaa aat gcc cct gta ctt tct gga aca atg cag gga
gga att caa ggt 3360Glu Asn Ala Pro Val Leu Ser Gly Thr Met Gln Gly
Gly Ile Gln Gly1105 1110 1115gct gaa gaa
gaa gta tat tta aca gct ttc att ctg gtt gcg ttg ttg 3408Ala Glu Glu
Glu Val Tyr Leu Thr Ala Phe Ile Leu Val Ala Leu Leu1120
1125 1130 1135gaa tcc aaa aca atc tgc aat
gac tat gtc aat agt cta gac agc agc 3456Glu Ser Lys Thr Ile Cys Asn
Asp Tyr Val Asn Ser Leu Asp Ser Ser 1140 1145
1150atc aag aag gcc aca aat tat tta ctc aaa aag tat gag aaa
ctg caa 3504Ile Lys Lys Ala Thr Asn Tyr Leu Leu Lys Lys Tyr Glu Lys
Leu Gln 1155 1160 1165agg cct tac
act aca gcc ctc aca gcc tat gct ttg gct gct gca gac 3552Arg Pro Tyr
Thr Thr Ala Leu Thr Ala Tyr Ala Leu Ala Ala Ala Asp 1170
1175 1180caa ctc aat gat gac agg gta ctc atg gca gca tca
aca gga agg gat 3600Gln Leu Asn Asp Asp Arg Val Leu Met Ala Ala Ser
Thr Gly Arg Asp1185 1190 1195cat tgg gaa
gaa tac aat gct cac acc cac aac att gaa ggc act tcc 3648His Trp Glu
Glu Tyr Asn Ala His Thr His Asn Ile Glu Gly Thr Ser1200
1205 1210 1215tat gcc ttg ttg gcc ctg ctg
aaa atg aag aaa ttt gat caa act ggt 3696Tyr Ala Leu Leu Ala Leu Leu
Lys Met Lys Lys Phe Asp Gln Thr Gly 1220 1225
1230ccc ata gtc aga tgg ctg aca gat cag aat ttt tat ggg gaa
aca tat 3744Pro Ile Val Arg Trp Leu Thr Asp Gln Asn Phe Tyr Gly Glu
Thr Tyr 1235 1240 1245gga caa acc
caa gca aca gtt atg gca ttt caa gct ctt gct gaa tat 3792Gly Gln Thr
Gln Ala Thr Val Met Ala Phe Gln Ala Leu Ala Glu Tyr 1250
1255 1260gag att cag atg cct acc cat aag gac tta aac tta
gat att act att 3840Glu Ile Gln Met Pro Thr His Lys Asp Leu Asn Leu
Asp Ile Thr Ile1265 1270 1275gaa ctg cca
gat cga gaa gta cct ata agg tac aga att aat tat gaa 3888Glu Leu Pro
Asp Arg Glu Val Pro Ile Arg Tyr Arg Ile Asn Tyr Glu1280
1285 1290 1295aat gct ctc ctg gct cgg aca
gta gag acc aaa ctc aac caa gac atc 3936Asn Ala Leu Leu Ala Arg Thr
Val Glu Thr Lys Leu Asn Gln Asp Ile 1300 1305
1310act gtg aca gca tca ggt gat gga aaa gca aca atg acc att
ttg aca 3984Thr Val Thr Ala Ser Gly Asp Gly Lys Ala Thr Met Thr Ile
Leu Thr 1315 1320 1325ttc tat aac
gca cag ttg cag gag aag gca aat gtt tgc aat aaa ttt 4032Phe Tyr Asn
Ala Gln Leu Gln Glu Lys Ala Asn Val Cys Asn Lys Phe 1330
1335 1340cat ctt aat gtt tct gtt gaa aac atc cac ttg aat
gca atg gga gcc 4080His Leu Asn Val Ser Val Glu Asn Ile His Leu Asn
Ala Met Gly Ala1345 1350 1355aag gga gcc
ctc atg ctc aag atc tgc aca agg tat ctg gga gaa gtt 4128Lys Gly Ala
Leu Met Leu Lys Ile Cys Thr Arg Tyr Leu Gly Glu Val1360
1365 1370 1375gat tct aca atg aca ata att
gat att tct atg ctg act ggt ttt ctc 4176Asp Ser Thr Met Thr Ile Ile
Asp Ile Ser Met Leu Thr Gly Phe Leu 1380 1385
1390cct gat gct gaa gac ctt aca agg ctt tct aaa gga gtg gac
aga tac 4224Pro Asp Ala Glu Asp Leu Thr Arg Leu Ser Lys Gly Val Asp
Arg Tyr 1395 1400 1405atc tcc aga
tat gaa gtt gac aat aat atg gct cag aaa gta gct gtt 4272Ile Ser Arg
Tyr Glu Val Asp Asn Asn Met Ala Gln Lys Val Ala Val 1410
1415 1420atc att tac tta aac aag gtc tcc cac tct gaa gat
gaa tgc ctg cac 4320Ile Ile Tyr Leu Asn Lys Val Ser His Ser Glu Asp
Glu Cys Leu His1425 1430 1435ttt aag att
ctc aag cat ttt gaa gtt ggc ttc att cag cca gga tca 4368Phe Lys Ile
Leu Lys His Phe Glu Val Gly Phe Ile Gln Pro Gly Ser1440
1445 1450 1455gtc aag gtg tac agc tac tac
aat cta gat gaa aaa tgt acc aag ttc 4416Val Lys Val Tyr Ser Tyr Tyr
Asn Leu Asp Glu Lys Cys Thr Lys Phe 1460 1465
1470tac cat cca gat aaa gga aca ggc ctt ctc aat aag ata tgt
att ggt 4464Tyr His Pro Asp Lys Gly Thr Gly Leu Leu Asn Lys Ile Cys
Ile Gly 1475 1480 1485aac gtt tgc
cga tgt gca gga gaa acc tgt tcc tcg ctc aac cat cag 4512Asn Val Cys
Arg Cys Ala Gly Glu Thr Cys Ser Ser Leu Asn His Gln 1490
1495 1500gaa agg att gat gtt cca tta caa att gaa aaa gcc
tgc gag acg aat 4560Glu Arg Ile Asp Val Pro Leu Gln Ile Glu Lys Ala
Cys Glu Thr Asn1505 1510 1515gtg gat tat
gtc tac aaa acc aag ctg ctt cga ata gaa gaa caa gat 4608Val Asp Tyr
Val Tyr Lys Thr Lys Leu Leu Arg Ile Glu Glu Gln Asp1520
1525 1530 1535ggt aat gat atc tat gtc atg
gat gtt tta gaa gtt att aaa caa ggt 4656Gly Asn Asp Ile Tyr Val Met
Asp Val Leu Glu Val Ile Lys Gln Gly 1540 1545
1550act gac gaa aat cca cga gca aag acc cac cag tac ata agt
caa agg 4704Thr Asp Glu Asn Pro Arg Ala Lys Thr His Gln Tyr Ile Ser
Gln Arg 1555 1560 1565aaa tgc cag
gag gct ctg aat ctg aag gtg aat gat gat tat ctg atc 4752Lys Cys Gln
Glu Ala Leu Asn Leu Lys Val Asn Asp Asp Tyr Leu Ile 1570
1575 1580tgg ggt tcc agg agt gac ctg ttg ccc acg aaa gat
aaa att tcc tac 4800Trp Gly Ser Arg Ser Asp Leu Leu Pro Thr Lys Asp
Lys Ile Ser Tyr1585 1590 1595atc att aca
aag aac aca tgg att gag aga tgg cca cat gaa gac gaa 4848Ile Ile Thr
Lys Asn Thr Trp Ile Glu Arg Trp Pro His Glu Asp Glu1600
1605 1610 1615tgt cag gaa gaa gaa ttc caa
aag ttg tgt gat gac ttt gct cag ttt 4896Cys Gln Glu Glu Glu Phe Gln
Lys Leu Cys Asp Asp Phe Ala Gln Phe 1620 1625
1630agc tac aca ttg act gag ttt ggc tgc cct act taa
aagttcagaa 4942Ser Tyr Thr Leu Thr Glu Phe Gly Cys Pro Thr *
1635 1640gaatcaatga taggaaggaa attctcagaa gacagatttt
tgagccaatg catatatgtt 5002actttgcctc ttgatctttt agttttatgt caatttgctc
tgttattttc ccttaaattg 5062tttatacata aaataaataa tcgatttctt actttgatat
gttcttgatt tttaataaac 5122aatggtgatt catgattatt tttttcttct tctgatccgt
ccaatatttg aagtgctctg 5182aacagagcac ttatggagta atgttttagt gatggatgaa
taagttggtg agtcaatatt 5242atcaggccct atatactctt atggaagatc gatttgtacc
caaagaaaca tagattgaaa 5302tgtgttactt tgaaaacaga ggtttcagtt gtatatgttt
acacttggat acaatcttaa 5362ctcttaataa acactgatct cagaacattt aacagctgct
atttaataat gacaaaatat 5422tctttgactg cacccacaga aaacattgca ttacattaga
atgggtttta tcagatgact 5482aagtctgcta gacttgccat ctgtcaaaat gtgcctcttc
cccagctcca actttaagga 5542tagtaactaa tagatgttct ctcattggct cctgacagag
gtgtggtagc cactgagttt 5602ccctggatga cactagaagc tggcagcaca ctgcagcctg
gtggaggggc ctcttttgct 5662atcccatgag cttctattca tcctcttatc tgttgggatg
gggatgggac gtctctgatt 5722ttccaggtat acaggtgatc tcatttacta acatcaccac
taacttcaag gattggttga 5782ggggttatgc caatgtgatt gaaggtttca cccatgtgaa
tctattctcc aatcccaatg 5842ctgtatctat gctgctcatt tctgcttgta aaaatggtat
aaaaagaata aacactgccc 5902aggcagtcag acatctttgg acactgaaaa aaaaaaaaaa
aaaaaa 594841642PRTNaja kaouthia 4Met Glu Arg Met Ala
Leu Tyr Leu Val Ala Ala Leu Leu Ile Gly Phe1 5
10 15Pro Gly Ser Ser His Gly Ala Leu Tyr Thr Leu Ile
Thr Pro Ala Val 20 25 30Leu
Arg Thr Asp Thr Glu Glu Gln Ile Leu Val Glu Ala His Gly Asp 35
40 45Ser Thr Pro Lys Gln Leu Asp Ile Phe Val
His Asp Phe Pro Arg Lys 50 55 60Gln
Lys Thr Leu Phe Gln Thr Arg Val Asp Met Asn Pro Ala Gly Gly65
70 75 80Met Leu Val Thr Pro Thr
Ile Glu Ile Pro Ala Lys Glu Val Ser Thr 85
90 95Asp Ser Arg Gln Asn Gln Tyr Val Val Val Gln Val Thr
Gly Pro Gln 100 105 110Val Arg
Leu Glu Lys Val Val Leu Leu Ser Tyr Gln Ser Ser Phe Leu 115
120 125Phe Ile Gln Thr Asp Lys Gly Ile Tyr Thr
Pro Gly Ser Pro Val Leu 130 135 140Tyr
Arg Val Phe Ser Met Asp His Asn Thr Ser Lys Met Asn Lys Thr145
150 155 160Val Ile Val Glu Phe Gln
Thr Pro Glu Gly Ile Leu Val Ser Ser Asn 165
170 175Ser Val Asp Leu Asn Phe Phe Trp Pro Tyr Asn Leu
Pro Asp Leu Val 180 185 190Ser
Leu Gly Thr Trp Arg Ile Val Ala Lys Tyr Glu His Ser Pro Glu 195
200 205Asn Tyr Thr Ala Tyr Phe Asp Val Arg
Lys Tyr Val Leu Pro Ser Phe 210 215
220Glu Val Arg Leu Gln Pro Ser Glu Lys Phe Phe Tyr Ile Asp Gly Asn225
230 235 240Glu Asn Phe His
Val Ser Ile Thr Ala Arg Tyr Leu Tyr Gly Glu Glu 245
250 255Val Glu Gly Val Ala Phe Val Leu Phe Gly
Val Lys Ile Asp Asp Ala 260 265
270Lys Lys Ser Ile Pro Asp Ser Leu Thr Arg Ile Pro Ile Ile Asp Gly
275 280 285Asp Gly Lys Ala Thr Leu Lys
Arg Asp Thr Phe Arg Ser Arg Phe Pro 290 295
300Asn Leu Asn Glu Leu Val Gly His Thr Leu Tyr Ala Ser Val Thr Val305
310 315 320Met Thr Glu
Ser Gly Ser Asp Met Val Val Thr Glu Gln Ser Gly Ile 325
330 335His Ile Val Ala Ser Pro Tyr Gln Ile
His Phe Thr Lys Thr Pro Lys 340 345
350Tyr Phe Lys Pro Gly Met Pro Tyr Glu Leu Thr Val Tyr Val Thr Asn
355 360 365Pro Asp Gly Ser Pro Ala Ala
His Val Pro Val Val Ser Glu Ala Phe 370 375
380His Ser Met Gly Thr Thr Leu Ser Asp Gly Thr Ala Lys Leu Ile Leu385
390 395 400Asn Ile Pro
Leu Asn Ala Gln Ser Leu Pro Ile Thr Val Arg Thr Asn 405
410 415His Gly Asp Leu Pro Arg Glu Arg Gln
Ala Thr Lys Ser Met Thr Ala 420 425
430Ile Ala Tyr Gln Thr Gln Gly Gly Ser Gly Asn Tyr Leu His Val Ala
435 440 445Ile Thr Ser Thr Glu Ile Lys
Pro Gly Asp Asn Leu Pro Val Asn Phe 450 455
460Asn Val Lys Gly Asn Ala Asn Ser Leu Lys Gln Ile Lys Tyr Phe Thr465
470 475 480Tyr Leu Ile
Leu Asn Lys Gly Lys Ile Phe Lys Val Gly Arg Gln Pro 485
490 495Arg Arg Asp Gly Gln Asn Leu Val Thr
Met Asn Leu His Ile Thr Pro 500 505
510Asp Leu Ile Pro Ser Phe Arg Phe Val Ala Tyr Tyr Gln Val Gly Asn
515 520 525Asn Glu Ile Val Ala Asp Ser
Val Trp Val Asp Val Lys Asp Thr Cys 530 535
540Met Gly Thr Leu Val Val Lys Gly Asp Asn Leu Ile Gln Met Pro Gly545
550 555 560Ala Ala Met
Lys Ile Lys Leu Glu Gly Asp Pro Gly Ala Arg Val Gly 565
570 575Leu Val Ala Val Asp Lys Ala Val Tyr
Val Leu Asn Asp Lys Tyr Lys 580 585
590Ile Ser Gln Ala Lys Ile Trp Asp Thr Ile Glu Lys Ser Asp Phe Gly
595 600 605Cys Thr Ala Gly Ser Gly Gln
Asn Asn Leu Gly Val Phe Glu Asp Ala 610 615
620Gly Leu Ala Leu Thr Thr Ser Thr Asn Leu Asn Thr Lys Gln Arg Ser625
630 635 640Ala Ala Lys
Cys Pro Gln Pro Ala Asn Arg Arg Arg Arg Ser Ser Val 645
650 655Leu Leu Leu Asp Ser Asn Ala Ser Lys
Ala Ala Glu Phe Gln Asp Gln 660 665
670Asp Leu Arg Lys Cys Cys Glu Asp Val Met His Glu Asn Pro Met Gly
675 680 685Tyr Thr Cys Glu Lys Arg Ala
Lys Tyr Ile Gln Glu Gly Asp Ala Cys 690 695
700Lys Ala Ala Phe Leu Glu Cys Cys Arg Tyr Ile Lys Gly Val Arg Asp705
710 715 720Glu Asn Gln
Arg Glu Ser Glu Leu Phe Leu Ala Arg Asp Asp Asn Glu 725
730 735Asp Gly Phe Ile Ala Asp Ser Asp Ile
Ile Ser Arg Ser Asp Phe Pro 740 745
750Lys Ser Trp Leu Trp Leu Thr Lys Asp Leu Thr Glu Glu Pro Asn Ser
755 760 765Gln Gly Ile Ser Ser Lys Thr
Met Ser Phe Tyr Leu Arg Asp Ser Ile 770 775
780Thr Thr Trp Val Val Leu Ala Val Ser Phe Thr Pro Thr Lys Gly Ile785
790 795 800Cys Val Ala
Glu Pro Tyr Glu Ile Arg Val Met Lys Val Phe Phe Ile 805
810 815Asp Leu Gln Met Pro Tyr Ser Val Val
Lys Asn Glu Gln Val Glu Ile 820 825
830Arg Ala Ile Leu His Asn Tyr Val Asn Glu Asp Ile Tyr Val Arg Val
835 840 845Glu Leu Leu Tyr Asn Pro Ala
Phe Cys Ser Ala Ser Thr Lys Gly Gln 850 855
860Arg Tyr Arg Gln Gln Phe Pro Ile Lys Ala Leu Ser Ser Arg Ala Val865
870 875 880Pro Phe Val
Ile Val Pro Leu Glu Gln Gly Leu His Asp Val Glu Ile 885
890 895Lys Ala Ser Val Gln Glu Ala Leu Trp
Ser Asp Gly Val Arg Lys Lys 900 905
910Leu Lys Val Val Pro Glu Gly Val Gln Lys Ser Ile Val Thr Ile Val
915 920 925Lys Leu Asp Pro Arg Ala Lys
Gly Val Gly Gly Thr Gln Leu Glu Val 930 935
940Ile Lys Ala Arg Lys Leu Asp Asp Arg Val Pro Asp Thr Glu Ile Glu945
950 955 960Thr Lys Ile
Ile Ile Gln Gly Asp Pro Val Ala Gln Ile Ile Glu Asn 965
970 975Ser Ile Asp Gly Ser Lys Leu Asn His
Leu Ile Ile Thr Pro Ser Gly 980 985
990Cys Gly Glu Gln Asn Met Ile Arg Met Ala Ala Pro Val Ile Ala Thr
995 1000 1005Tyr Tyr Leu Asp Thr Thr
Glu Gln Trp Glu Thr Leu Gly Ile Asn Arg 1010 1015
1020Arg Thr Glu Ala Val Asn Gln Ile Val Thr Gly Tyr Ala Gln Gln
Met1025 1030 1035 1040Val
Tyr Lys Lys Ala Asp His Ser Tyr Ala Ala Phe Thr Asn Arg Ala
1045 1050 1055Ser Ser Ser Trp Leu Thr Ala
Tyr Val Val Lys Val Phe Ala Met Ala 1060 1065
1070Ala Lys Met Val Ala Gly Ile Ser His Glu Ile Ile Cys Gly
Gly Val 1075 1080 1085Arg Trp Leu Ile
Leu Asn Arg Gln Gln Pro Asp Gly Ala Phe Lys Glu 1090
1095 1100Asn Ala Pro Val Leu Ser Gly Thr Met Gln Gly Gly
Ile Gln Gly Ala1105 1110 1115
1120Glu Glu Glu Val Tyr Leu Thr Ala Phe Ile Leu Val Ala Leu Leu Glu
1125 1130 1135Ser Lys Thr Ile Cys
Asn Asp Tyr Val Asn Ser Leu Asp Ser Ser Ile 1140
1145 1150Lys Lys Ala Thr Asn Tyr Leu Leu Lys Lys Tyr Glu
Lys Leu Gln Arg 1155 1160 1165Pro Tyr
Thr Thr Ala Leu Thr Ala Tyr Ala Leu Ala Ala Ala Asp Gln 1170
1175 1180Leu Asn Asp Asp Arg Val Leu Met Ala Ala Ser
Thr Gly Arg Asp His1185 1190 1195
1200Trp Glu Glu Tyr Asn Ala His Thr His Asn Ile Glu Gly Thr Ser Tyr
1205 1210 1215Ala Leu Leu Ala
Leu Leu Lys Met Lys Lys Phe Asp Gln Thr Gly Pro 1220
1225 1230Ile Val Arg Trp Leu Thr Asp Gln Asn Phe Tyr
Gly Glu Thr Tyr Gly 1235 1240 1245Gln
Thr Gln Ala Thr Val Met Ala Phe Gln Ala Leu Ala Glu Tyr Glu 1250
1255 1260Ile Gln Met Pro Thr His Lys Asp Leu Asn
Leu Asp Ile Thr Ile Glu1265 1270 1275
1280Leu Pro Asp Arg Glu Val Pro Ile Arg Tyr Arg Ile Asn Tyr Glu
Asn 1285 1290 1295Ala Leu Leu
Ala Arg Thr Val Glu Thr Lys Leu Asn Gln Asp Ile Thr 1300
1305 1310Val Thr Ala Ser Gly Asp Gly Lys Ala Thr
Met Thr Ile Leu Thr Phe 1315 1320
1325Tyr Asn Ala Gln Leu Gln Glu Lys Ala Asn Val Cys Asn Lys Phe His
1330 1335 1340Leu Asn Val Ser Val Glu Asn
Ile His Leu Asn Ala Met Gly Ala Lys1345 1350
1355 1360Gly Ala Leu Met Leu Lys Ile Cys Thr Arg Tyr Leu
Gly Glu Val Asp 1365 1370
1375Ser Thr Met Thr Ile Ile Asp Ile Ser Met Leu Thr Gly Phe Leu Pro
1380 1385 1390Asp Ala Glu Asp Leu Thr
Arg Leu Ser Lys Gly Val Asp Arg Tyr Ile 1395 1400
1405Ser Arg Tyr Glu Val Asp Asn Asn Met Ala Gln Lys Val Ala
Val Ile 1410 1415 1420Ile Tyr Leu Asn Lys
Val Ser His Ser Glu Asp Glu Cys Leu His Phe1425 1430
1435 1440Lys Ile Leu Lys His Phe Glu Val Gly Phe
Ile Gln Pro Gly Ser Val 1445 1450
1455Lys Val Tyr Ser Tyr Tyr Asn Leu Asp Glu Lys Cys Thr Lys Phe Tyr
1460 1465 1470His Pro Asp Lys Gly
Thr Gly Leu Leu Asn Lys Ile Cys Ile Gly Asn 1475
1480 1485Val Cys Arg Cys Ala Gly Glu Thr Cys Ser Ser Leu
Asn His Gln Glu 1490 1495 1500Arg Ile Asp
Val Pro Leu Gln Ile Glu Lys Ala Cys Glu Thr Asn Val1505
1510 1515 1520Asp Tyr Val Tyr Lys Thr Lys
Leu Leu Arg Ile Glu Glu Gln Asp Gly 1525
1530 1535Asn Asp Ile Tyr Val Met Asp Val Leu Glu Val Ile
Lys Gln Gly Thr 1540 1545 1550Asp
Glu Asn Pro Arg Ala Lys Thr His Gln Tyr Ile Ser Gln Arg Lys 1555
1560 1565Cys Gln Glu Ala Leu Asn Leu Lys Val
Asn Asp Asp Tyr Leu Ile Trp 1570 1575
1580Gly Ser Arg Ser Asp Leu Leu Pro Thr Lys Asp Lys Ile Ser Tyr Ile1585
1590 1595 1600Ile Thr Lys Asn
Thr Trp Ile Glu Arg Trp Pro His Glu Asp Glu Cys 1605
1610 1615Gln Glu Glu Glu Phe Gln Lys Leu Cys Asp
Asp Phe Ala Gln Phe Ser 1620 1625
1630Tyr Thr Leu Thr Glu Phe Gly Cys Pro Thr 1635
1640519PRTArtificial SequenceVector coded sequence. 5Arg Ser Pro Trp Pro
Gly Val Pro Thr Ser Pro Val Trp Trp Asn Ser1 5
10 15Ala Asp Ala631DNAArtificial SequencePrimer
HC3H5-1. 6ggatgccact atgtctatat tggacatatc c
31730DNAArtificial SequencePrimer HC3H5-2. 7tcttctattc gaaccagtcg
ggtcttgtac 30835DNAArtificial
SequencePrimer HuC3H5-3. 8gtacaagacc cgactggttc gaatagaaga acaag
35936DNAArtificial SequencePrimer HuC3H5-4.
9tatcatgtaa gcggccgcgt ataaacaatt taaggg
361048DNAArtificial SequencePrimer HC3SigRemF. 10agatctccat ggaagcttag
cgctgggagt cccatgtact ctatcatc 481120DNAArtificial
SequencePrimer HC3SigRemR. 11gcgtcccgcc ttcaacagcc
201224DNAArtificial SequencePrimer HC3H5-3-F1.
12tctgtgtggc agaccccttc gagg
241335DNAArtificial SequencePrimer HC3H5-5-R1. 13cgttaccaat acatatcttg
ttcagctttc catcc 351435DNAArtificial
SequencePrimer HuCC3H5-3-F2. 14ggatggaaag ctgaacaaga tatgtattgg taacg
351531DNAArtificial SequencePrimer
HuC3H5-3-R2. 15catccatgac atagatatca ttaccatctt g
311632DNAArtificial SequenceHuC3H5-5-1R. 16gcaactgtgc
gttatacatt gtcaccaccg ac
321732DNAArtificial SequenceHuC3H5-5-2F. 17gtcggtggtg acaatgtata
acgcacagtt gc 321841DNAArtificial
SequenceHC3H5-4-R1. 18gagaaggcct gttcctttat ccggatggta gaaccgggta c
411935DNAArtificial SequenceHCC3H5-4-F2. 19ccggttctac
catccggata aaggaacagg ccttc
352031DNAArtificial SequenceHC3H5-3-R2. 20catccatgac atagatatca
ttaccatctt g 312131DNAArtificial
SequenceHuC3H5-F1. 21ggatgccact atgtctatat tggacatatc c
312233DNAArtificial SequenceHuC3H5-2R1. 22cccgatgatg
tagctgagtt tatctttcgt ggg
332333DNAArtificial SequenceHuC3H5-2F2. 23cccacgaaag ataaactcag
ctacatcatc ggg 332422DNAArtificial
SequenceHuC3H5-2-R2. 24aattggagct ccaccgcggt gg
22
User Contributions:
comments("1"); ?> comment_form("1"); ?>Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
User Contributions:
Comment about this patent or add new information about this topic: