Patent application title: Method for Treating CCR4-Related Diseases
Inventors:
Shigeru Iida (Tokyo, JP)
Mitsuo Satoh (Tokyo, JP)
Miho Urakubo (Tokyo, JP)
Masako Wakitani (Tokyo, JP)
Kazuhisa Uchida (Tokyo, JP)
Rinpei Niwa (Tokyo, JP)
Rinpei Niwa (Tokyo, JP)
Kenya Shitara (Tokyo, JP)
IPC8 Class: AA61K39395FI
USPC Class:
4241331
Class name: Drug, bio-affecting and body treating compositions immunoglobulin, antiserum, antibody, or antibody fragment, except conjugate or complex of the same with nonimmunoglobulin material structurally-modified antibody, immunoglobulin, or fragment thereof (e.g., chimeric, humanized, cdr-grafted, mutated, etc.)
Publication date: 2008-09-04
Patent application number: 20080213266
Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
Patent application title: Method for Treating CCR4-Related Diseases
Inventors:
Shigeru Iida
Mitsuo Satoh
Miho Urakubo
Masako Wakitani
Kazuhisa Uchida
Rinpei Niwa
Kenya Shitara
Agents:
NIXON & VANDERHYE, PC
Assignees:
Origin: ARLINGTON, VA US
IPC8 Class: AA61K39395FI
USPC Class:
4241331
Abstract:
The present invention provides an antibody composition comprising an
antibody molecule which specifically binds to human CC chemokine receptor
4 (CCR4) and has complex type N-glycoside-linked sugar chains in the Fc
region, wherein the complex type N-glycoside-linked sugar chains have a
structure in which fucose is not bound to N-acetylglucosamine in the
reducing end in the sugar chains; a transformant which produces the
antibody composition; a process for producing the antibody composition;
and a pharmaceutical composition comprising the antibody composition.Claims:
1-53. (canceled)
54. A method for treating diseases related to a human CC chemokine receptor 4 (CCR4), which comprises administering a composition comprising an antibody to a subject in need of such treatment, the antibody being a recombinant antibody molecule which specifically binds to an extracellular region of the CCR4 and has complex type N-glycoside-linked sugar chains in the Fc region, wherein the complex type N-glycoside-linked sugar chains have a structure in which fucose is not bound to 6-position of N-acetylglucosamine in the reducing end through an α-bond in the sugar chains, wherein 100% of the recombinant antibody molecules in the composition do not possess N-glycoside-linked sugar chains in which fucose is bound to 6-position of N-acetylglucosamine in the reducing end through an α-bond, and the disease related to a human CCR4 being selected from the group consisting of blood cancers and Th2 cells-associated inflammatory diseases.
55-56. (canceled)
57. The method according to claim 54, wherein the extracellular region is an extracellular region selected from the group consisting of the sequences at positions 1 to 39, positions 98 to 112, positions 176 to 206 and positions 271 to 284 of the amino acid sequence represented by SEQ ID NO:36.
58. The method according to claim 54, wherein the extracellular region is an epitope existing at positions 2 to 29 of the amino acid sequence represented by SEQ ID NO:36.
59. The method according to claim 54, wherein the extracellular region is an epitope existing at positions 13 to 29 of the amino acid sequence represented by SEQ ID NO:36.
60. The method according to claim 54, wherein the extracellular region is an epitope existing at positions 13 to 25 of the amino acid sequence represented by SEQ ID NO:36.
61. The method according to claim 60, wherein the antibody has lower binding activity to a peptide comprising the sequence at positions 13 to 25 of the amino acid sequence represented by SEQ ID NO:36 wherein at least one of tyrosine residues at positions 16, 19, 20 and 22 is sulfated, in comparison with its binding activity to a peptide comprising the sequence at positions 13 to 25 of the amino acid sequence represented by SEQ ID NO:36.
62. The method according to claim 54, wherein the antibody has no reactivity to a human blood platelet.
63. The method according to claim 54, wherein the antibody has no activity of inhibiting a binding of a CCR4 ligand to the CCR4, the CCR4 ligand being TARC (thymus and activation-regulated chemokine) or MDC (macrophage-derived chemokine).
64. The method according to claim 54, wherein the antibody has cytotoxic activity against a CCR4-expressing cell.
65. The method according to claim 64, wherein the antibody has higher cytotoxic activity against a CCR4-expressing cell than a monoclonal antibody produced by a non-human animal-derived hybridoma.
66. The method according to claim 64, wherein the CCR4-expressing cell is a helper T cell.
67. The method according to claim 64, wherein the cytotoxic activity is antibody-dependent cell-mediated cytotoxic (ADCC) activity.
68. The method according to claim 54, wherein the antibody comprises complementarity determining region (CDR) 1, CDR 2 and CDR 3 of an antibody molecule heavy chain (H chain) variable region (V region) consisting of the amino acid sequences represented by SEQ ID NOs:14, 15 and 16, respectively; and CDR 1, CDR 2 and CDR 3 of an antibody molecule light chain (L chain) V region consisting of the amino acid sequences represented by SEQ ID NOs:17, 18 and 19, respectively.
69. The method according to claim 54, wherein the antibody is a human chimeric antibody, a human CDR-grafted antibody or a human antibody.
70. The method according to claim 69, wherein the antibody comprises CDRs of heavy chain (H chain) variable region (V region) and light chain (L chain) V region of a monoclonal antibody which specifically binds to the CCR4.
71. The method according to claim 70, wherein the heavy chain (H chain) variable region (V region) of the human chimeric antibody comprises the amino acid sequence represented by SEQ ID NO:21.
72. The method according to claim 70, wherein the light chain (L chain) variable region (V region) of the human chimeric antibody comprises the amino acid sequence represented by SEQ ID NO:23.
73. The method according to claim 70, wherein the heavy chain (H chain) variable region (V region) of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:21 and the light chain (L chain) V region of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:23.
74. The method according to claim 69, wherein the human CDR-grafted antibody comprises CDRs of heavy chain (H chain) variable region (V region) and light chain (L chain) V region of a monoclonal antibody which specifically binds to the CCR4, and framework regions (FRs) of H chain V region and L chain V region of a human antibody.
75. The method according to claim 69, wherein the human CDR-grafted antibody comprises CDRs of heavy chain (H chain) variable region (V region) and light chain (L chain) V region of a monoclonal antibody which specifically binds to the CCR4, FRs of H chain V region and L chain V region of a human antibody, and H chain constant region (C region) and L chain C region of a human antibody.
76. The method according to claim 70, wherein the heavy chain (H chain) variable region (V region) of the human CDR-grafted antibody comprises the amino acid sequence represented by SEQ ID NO:24 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ala at position 40, Gly at position 42, Lys at position 43, Gly at position 44, Lys at position 76 and Ala at position 97 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:24.
77. The method according to claim 70, wherein the heavy chain (H chain) variable region (V region) of the human CDR-grafted antibody comprises the amino acid sequence represented by SEQ ID NO:25 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Thr at position 28 and Ala at position 97 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:25.
78. The method according to claim 70, wherein the light chain (L chain) variable region (V region) of the human CDR-grafted antibody comprises the amino acid sequence represented by SEQ ID NO:26 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ile at position 2, Val at position 3, GIn at position 50 and Val at position 88 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:26.
79. The method according to claim 70, wherein the heavy chain (H chain) variable region (V region) of the human CDR-grafted antibody comprises the amino acid sequence represented by SEQ ID NO:24 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ala at position 40, Gly at position 42, Lys at position 43, Gly at position 44, Lys at position 76 and Ala at position 97 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:24; and the light chain (L chain) V region of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:26 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ile at position 2, Val at position 3, GIn at position 50 and Val at position 88 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:26.
80. The method according to claim 70, wherein the heavy chain (H chain) variable region (V region) of the human CDR-grafted antibody comprises the amino acid sequence represented by SEQ ID NO:25 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Thr at position 28 and Ala at position 97 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:25; and the light chain (L chain) V region of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:26 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ile at position 2, Val at position 3, GIn at position 50 and Val at position 88 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:26.
81. The method according to claim 70, wherein the heavy chain (H chain) variable region (V region) of the human CDR-grafted antibody comprises an amino acid sequence selected from the group consisting of the amino acid sequences represented by SEQ ID NOs:24, 25, 27, 28, 29, 30, 31 and 32.
82. The method according to claim 70, wherein the light (L chain) variable region (V region) of the human CDR-grafted antibody comprises an amino acid sequence selected from the group consisting of the amino acid sequences represented by SEQ ID NOs:26, 33, 34 and 35.
83. The method according to claim 70, wherein the heavy chain (H chain) variable region (V region) of the human CDR-grafted antibody comprises an amino acid sequence selected from the group consisting of the amino acid sequences represented by SEQ ID NOs:24, 25, 27, 28, 29, 30, 31 and 32; and the light chain (L chain) V region of the antibody molecule comprises an amino acid sequence selected from the group consisting of the amino acid sequences represented by SEQ ID NOs:33, 34 and 35.
84. The method according to claim 70, wherein the heavy chain (H chain) variable region (V region) of the human CDR-grafted antibody comprises the amino acid sequence represented by SEQ ID NO:27 or 28, and the light chain (L chain) V region of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:35.
Description:
[0001]This is a Divisional of application Ser. No. 10/959,310 filed Oct.
7, 2004, which claims priority of provisional application No. 60/572,784
filed May 2, 2004 and claims priority to Japanese patent application No.
2003-350,162 filed with the Japanese Patent Office on Oct. 8, 2003.
BACKGROUND OF THE INVENTION
[0002]1. Field of the Invention
[0003]The present invention relates to an antibody composition comprising a recombinant antibody molecule which specifically binds to human CC chemokine receptor 4 (hereinafter referred to as CCR4) and has complex type N-glycoside-linked sugar chains in the Fc region, wherein the complex type N-glycoside-linked sugar chains have a structure in which fucose is not bound to N-acetylglucosamine in the reducing end in the sugar chains; a transformant which produces the antibody composition; a process for producing the antibody composition; and a pharmaceutical composition comprising the antibody composition.
[0004]2. Brief Description of the Background Art
[0005]Eosinophils, mast cells, and various factors such as IgE, cytokines and chemokines are involved in the pathology of allergic diseases such as bronchial asthma (Am. J. Respir. Crit. Care Med., 152, 2059 (1995), Immunol. Today, 15, 19 (1994)).
[0006]Major cytokine-producing cells are CD4-positive helper T cells (hereinafter referred to as Th cells). The activated Th cells are increased in the airway cells and peripheral blood of patients with bronchial asthma (Am. Rev. Respir. Dis., 145, S22 (1992)).
[0007]In patients with atopic dermatitis or asthma or animal models of allergic inflammation, IL-4 and IL-5 are increased (Clin. Immunol. Immunopathol, 75, 75 (1995)). These cytokines are produced by Th2 cells among helper T cells. Accordingly, Th2 cells are involved in allergic diseases.
[0008]Methods so far developed for the treatment of immunodiseases mediated by Th2 cells include 1) antagonists of cytokines and chemokines, 2) cytokine/chemokine production inhibitors, 3) regulators of inflammatory cell function, and 4) chemical mediator antagonists. However, they inhibit only a part of the complicated network among cytokines, chemokines and inflammatory cells and are not radical treatments.
[0009]Thus, in order to inhibit the whole of the complicated network among cytokines, chemokines and inflammatory cells, it is necessary to control the upstream part of allergic reactions, that is, Th2 cells.
[0010]Anti-CD4 antibodies are known to control T cells. However, since CD4 is widely expressed in immunocytes, they lack in specificity and have the defect of accompanying potent immunosuppressive effects (Int. Arch. Aller. Immunol., 118, 133 (1999)). Currently, steroid administration is mainly employed for the treatment of Th2-mediated immunodiseases, but steroids have strong side effects.
[0011]Human CC chemokine receptor 4 (hereinafter referred to as CCR4) is a seven membrane-spanning G protein-coupled chemokine receptor cloned as K5-5 from human immature basophilic cell line KU-812 (WO96/23068, J. Biol. Chem., 270, 19495 (1995)). Expression of CCR4 in Th2 cells has been confirmed (J. Exp. Med., 187, 129 (1998), J. Immunol, 161, 5111 (1998)) and it is reported that CCR4 is expressed in effector/memory T cells (Int. Immunol., 11, 81 (1999)) and in memory T cells involved in the systemic immunity (Nature, 400, 776 (1999)). These facts strongly suggest that memory T cells activated upon induction of inflammation migrate to the inflammatory site via CCR4 by the action of ligands such as MDC (macrophage-derived chemokine) and TARC (thymus and activation-regulated chemokine) to accelerate activation of other inflammatory cells. It is also reported that, besides immunodiseases, CCR4 is expressed on the surface of tumor cells in leukemia and lymphoma such as adult T cell leukemia and mycosis fungoides (Blood, 96, 685 (2000)).
[0012]From the foregoing, therapeutic agents for removing the cells expressing CCR4 from the body of a patient are expected to be effective as therapeutic agents not only for immunodiseases mediated by Th2 cells but also for inflammatory diseases and leukemia mediated by CCR4. As examples of such therapeutic agents targeting at specific molecules, the following antibodies against CCR4 are known.
[0013]As antibodies against CCR4, 1G1 antibody and 2B10 antibody having the activity to neutralize receptors (WO00/42074) and an antibody having antagonist activity (WO02/15666) have been reported, which are antibodies produced by hybridomas. An anti-CCR4 human CDR-grafted antibody belonging to a human IgG1 type which specifically reacts to the extracellular region of human CCR4 but does not react with a blood platelet and which shows cytotoxic activity against human CCR4-expressing cells is known (WO03/18635).
[0014]It is generally known that when an antibody derived from a non-human animal is administered to human, it is recognized as a foreign substance, whereby side effects are induced (J. Clin. Oncol., 2, 881 (1984), Blood, 65, 1349 (1985), J. Natl. Cancer Inst., 80, 932 (1988), Proc. Natl. Acad. Sci. U.S.A., 82, 1242 (1985)), disappearance of the antibody from the body is accelerated (Blood, 65, 1349 (1985), J. Nucl Med., 26, 1011 (1985), J. Natl. Cancer Inst., 80, 937 (1988)) and thereby the therapeutic effects of the antibody is reduced (J. Immunol., 135, 1530 (1985), Cancer Res., 46, 6489 (1986)).
[0015]In order to solve these problems, attempts have been made to convert an antibody derived from a non-human animal into a humanized antibody, such as a human complementarity determining region (hereinafter referred to as CDR)-grafted antibody utilizing recombinant DNA techniques (Nature, 321, 522 (1986)). Humanized antibodies are reported to have reduced immunogenicity (Proc. Natl. Acad. Sci. U.S.A., 86, 4220 (1989)) and have prolonged therapeutic effects (Cancer Res., 56, 1118 (1996), Immunol., 85, 668 (1995)), as compared with antibodies derived from non-human animals.
[0016]Since humanized antibodies are prepared utilizing recombinant DNA techniques, they can be prepared as molecules in various forms. For example, a humanized antibody having high effector function can be prepared (Cancer Res., 56, 1118 (1996)).
[0017]In recent years, in the treatment of non Hodgkin's leukemia patients by Rituxan and the treatment of mammary cancer patients by Herceptin, when a therapeutic antibody induces high ADCC activity in effector cells of the patients, higher therapeutic effects can be obtained (Blood, 99, 754 (2002); J. Clin. Oncol., 21, 3940 (2003); Clin. Cancer Res., 10, 5650 (2004)).
[0018]Antibodies of the human IgG1 subclass express ADCC activity and CDC activity via interactions between the Fc region thereof and antibody receptors (hereinafter referred to as FcγR) or various complement components. It is suggested that in the binding of an antibody to FcγR, the hinge region of the antibody and a sugar chain bound to the second domain of the C region (hereinafter referred to as Cβ2 domain) are important (Chemical Immunology, 65, 88 (1997)).
[0019]It is known that there is diversity regarding the addition of galactose to the non-reducing end in a complex type N-glycoside-linked sugar chain bound to the Fc region of an IgG antibody molecule and the addition of fucose to N-acetylglucosamine at the reducing end (Biochemistry, 36, 130 (1997)). In particular, it is reported that the addition of fucose to the N-acetylglucosamine at the reducing end in the sugar chain causes significant decrease of the ADCC activity of the antibody (WO00/61739, J. Biol. Chem., 278, 3466 (2003)).
[0020]In general, most of the antibody compositions utilized as pharmaceutical compositions are prepared by recombinant DNA techniques using animal cells such as Chinese hamster ovary tissue-derived CHO cells as host cells, and the sugar chain structure of the expressed antibody compositions differs depending host cells.
[0021]It is possible to increase the ratio of sugar chains having a structure in which fucose is not bound to N-acetylglucosamine in the reducing end in the complex type N-glycoside-linked sugar chains bound to the Fc region of antibody molecules in an antibody composition by decreasing or deleting the activity of α1,6-fucosyltransferase (hereinafter referred to as FUT8), GDP-mannose 4,6-dehydratase (hereinafter referred to as GMD) or GDP-4-keto-6-deoxy-D-mannose-3,5-epimerase (hereinafter referred to as Fx) of antibody-producing cells (WO02/31140).
SUMMARY OF THE INVENTION
[0022]An object of the present invention is to provide an antibody composition comprising a recombinant antibody molecule which specifically binds to human CC chemokine receptor 4 (CCR4) and has complex type N-glycoside-linked sugar chains in the Fc region, wherein the complex type N-glycoside-linked sugar chains have a structure in which fucose is not bound to N-acetylglucosamine in the reducing end in the sugar chains; a transformant which produces the antibody composition; a process for producing the antibody composition; and a pharmaceutical composition comprising the antibody composition. Since the anti-CCR4 antibody composition of the present invention includes no antibody molecule having sugar chains to which fucose is bound, it has enhanced effector function. Therefore, it is effective for treatment to reduce CCR4-expressing Th2 cells from patients. The therapeutic use of the antibody in which the effector function is enhanced is also expected to be effective for weakening side effects of patients because it does not require combination use of chemotherapeutic agents or radioisotope-labeled antibodies. Furthermore, reduction of burden on patients and the like are expected by decreasing the dose of the therapeutic agent to patients.
[0023]The present invention relates to the following (1) to (55).
(1) An antibody composition comprising a recombinant antibody molecule which specifically binds to human CC chemokine receptor 4 (CCR4) and has complex type N-glycoside-linked sugar chains in the Fc region, wherein the complex type N-glycoside-linked sugar chains have a structure in which fucose is not bound to N-acetylglucosamine in the reducing end in the sugar chains.(2) The antibody composition according to (1), wherein the complex type N-glycoside-linked sugar chains are sugar chains in which 1-position of fucose is not bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in the sugar chains.(3) The antibody composition according to (1), which specifically binds to an extracellular region of human CC chemokine receptor 4 (CCR4).(4) The antibody composition according to (3), wherein the extracellular region is an extracellular region selected from the group consisting of the sequences at positions 1 to 39, positions 98 to 112, positions 176 to 206 and positions 271 to 284 of the amino acid sequence represented by SEQ ID NO:36.(5) The antibody composition according to (3) or (4), wherein the extracellular region is an epitope existing at positions 2 to 29 of the amino acid sequence represented by SEQ ID NO:36.(6) The antibody composition according to any one of (3) to (5), wherein the extracellular region is an epitope existing at positions 13 to 29 of the amino acid sequence represented by SEQ ID NO:36.(7) The antibody composition according to any one of (3) to (6), wherein the extracellular region is an epitope existing at positions 13 to 25 of the amino acid sequence represented by SEQ ID NO:36.(8) The antibody composition according to (7), which has lower binding activity to a peptide comprising the sequence at positions 13 to 25 of the amino acid sequence represented by SEQ ID NO:36 wherein at least one of tyrosine residues at positions 16, 19, 20 and 22 is sulfated, in comparison with its binding activity to a peptide comprising the sequence at positions 13 to 25 of the amino acid sequence represented by SEQ ID NO:36.(9) The antibody composition according to any one of (1) to (8), which has no reactivity to a human blood platelet.(10) The antibody composition according to any one of (1) to (9), which specifically binds to an extracellular region of human CC chemokine receptor 4 (CCR4) and has no inhibition activity on binding of a CCR4 ligand which is TARC (thymus and activation-regulated chemokine) or MDC (macrophage-derived chemokine) to CCR4.(11) The antibody composition according to any one of (1) to (10), which specifically binds to a human CC chemokine receptor 4 (CCR4)-expressing cell.(12) The antibody composition according to any one of (1) to (11), which has cytotoxic activity against a human CC chemokine receptor 4 (CCR4)-expressing cell.(13) The antibody composition according to any one of (1) to (12), which has higher cytotoxic activity against a human CC chemokine receptor 4 (CCR4)-expressing cell than a monoclonal antibody produced by a non-human animal-derived hybridoma.(14) The antibody composition according to any one of (11) to (13), wherein the human CC chemokine receptor 4 (CCR4)-expressing cell is a helper T cell.(15) The antibody composition according to (12) or (13), wherein the cytotoxic activity is antibody-dependent cell-mediated cytotoxic (ADCC) activity.(16) The antibody composition according to any one of (1) to (15), which comprises complementarity determining region (CDR) 1, CDR 2 and CDR 3 of an antibody molecule heavy chain (H chain) variable region (V region) consisting of the amino acid sequences represented by SEQ ID NOs:14, 15 and 16, respectively.(17) The antibody composition according to any one of (1) to (16), which comprises complementarity determining region (CDR) 1, CDR 2 and CDR 3 of an antibody molecule light chain (L chain) variable region (V region) consisting of the amino acid sequences represented by SEQ ID NOs:17, 18 and 19, respectively.(18) The antibody composition according to any one of (1) to (17), which comprises complementarity determining region (CDR) 1, CDR 2 and CDR 3 of an antibody molecule heavy chain (H chain) variable region (V region) consisting of the amino acid sequences represented by SEQ ID NOs:14, 15 and 16, respectively; and CDR 1, CDR 2 and CDR 3 of an antibody molecule light chain (L chain) V region consisting of the amino acid sequences represented by SEQ ID NOs:17, 18 and 19, respectively.(19) The antibody composition according to any one of (1) to (18), wherein the recombinant antibody is a human chimeric antibody or a human CDR-grafted antibody.(20) The antibody composition according to (19), wherein the human chimeric antibody comprises CDRs of heavy chain (H chain) variable region (V region) and light chain (L chain) V region of a monoclonal antibody which specifically binds to human CC chemokine receptor 4 (CCR4).(21) The antibody composition according to (20), wherein the heavy chain (H chain) variable region (V region) of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:21.(22) The antibody composition according to (20), wherein the light chain (L chain) variable region (V region) of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:23.(23) The antibody composition according to any one of (20) to (22), wherein the heavy chain (H chain) variable region (V region) of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:21 and the light chain (L chain) V region of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:23.(24) The antibody composition according to (19), wherein the human CDR-grafted antibody comprises CDRs of H chain V region and L chain V region of a monoclonal antibody which specifically binds to human CC chemokine receptor 4 (CCR4).(25) The antibody composition according to (24), wherein the human CDR-grafted antibody comprises CDRs of heavy chain (H chain) variable region (V region) and light chain (L chain) V region of a monoclonal antibody which specifically binds to human CC chemokine receptor 4 (CCR4), and framework regions (FRs) of H chain V region and L chain V region of a human antibody.(26) The antibody composition according to (24) or (25), which comprises CDRs of heavy chain (H chain) variable region (V region) and light chain (L chain) V region of a monoclonal antibody which specifically binds to human CC chemokine receptor 4 (CCR4), FRs of H chain V region and L chain V region of a human antibody, and H chain constant region (C region) and L chain C region of a human antibody.(27) The antibody composition according to any one of (24) to (26), wherein the heavy chain (H chain) variable region (V region) of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:24 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ala at position 40, Gly at position 42, Lys at position 43, Gly at position 44, Lys at position 76 and Ala at position 97 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:24.(28) The antibody composition according to any one of (24) to (26), wherein the heavy chain (H chain) variable region (V region) of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:25 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Thr at position 28 and Ala at position 97 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:25.(29) The antibody composition according to any one of (24) to (26), wherein the light chain (L chain) variable region (V region) of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:26 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ile at position 2, Val at position 3, Gln at position 50 and Val at position 88 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:26.(30) The antibody composition according to any one of (24) to (27) and (29), wherein the heavy chain (H chain) variable region (V region) of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:24 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ala at position 40, Gly at position 42, Lys at position 43, Gly at position 44, Lys at position 76 and Ala at position 97 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:24; and the light chain (L chain) V region of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:26 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ile at position 2, Val at position 3, Gln at position 50 and Val at position 88 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:26.(31) The antibody composition according to any one of (24) to (26), (28) and (29), wherein the heavy chain (H chain) variable region (V region) of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:25 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Thr at position 28 and Ala at position 97 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:25; and the light chain (L chain) V region of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:26 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ile at position 2, Val at position 3, Gln at position 50 and Val at position 88 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:26.(32) The antibody composition according to any one of (24) to (28) and (29) to (31), wherein the heavy chain (H chain) variable region (V region) of the antibody molecule comprises an amino acid sequence selected from the group consisting of the amino acid sequences represented by SEQ ID NOs:24, 25, 27, 28, 29, 30, 31 and 32.(33) The antibody composition according to any one of (24) to (26) and (29) to (31), wherein the light (L chain) variable region (V region) of the antibody molecule comprises an amino acid sequence selected from the group consisting of the amino acid sequences represented by SEQ ID NOs:26, 33, 34 and 35.(34) The antibody composition according to any one of (24) to (26), (32) and (33), wherein the heavy chain (H chain) variable region (V region) of the antibody molecule comprises an amino acid sequence selected from the group consisting of the amino acid sequences represented by SEQ ID NOs:24, 25, 27, 28, 29, 30, 31 and 32; and the light chain (L chain) V region of the antibody molecule comprises an amino acid sequence selected from the group consisting of the amino acid sequences represented by SEQ ID NOs:33, 34 and 35.(35) The antibody composition according to any one of (24) to (26), wherein the heavy chain (H chain) variable region (V region) of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:27 or 28, and the light chain (L chain) V region of the antibody molecule comprises the amino acid sequence represented by SEQ ID NO:35.(36) A transformant producing the antibody composition according to any one of (1) to (35), which is obtained by introducing a DNA encoding an antibody molecule which specifically binds to human CC chemokine receptor 4 (CCR4) into a host cell.(37) The transformant according to (36), wherein the host cell is a cell in which genome is modified so as to have deleted activity of an enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose, or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain.(38) The transformant according to (36), wherein the host cell is a cell in which all of alleles on a genome encoding an enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose, or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain are knocked out.(39) The transformant according to (37) or (38), wherein the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose, is an enzyme selected from the group consisting of GDP-mannose 4,6-dehydratase (GMD) and GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase (Fx).(40) The transformant according to (39), wherein the GDP-mannose 4,6-dehydratase is a protein encoded by a DNA selected from the group consisting of the following (a) and (b):(a) a DNA comprising the nucleotide sequence represented by SEQ ID NO:1;(b) a DNA which hybridizes with the DNA consisting of the nucleotide sequence represented by SEQ ID NO:1 under stringent conditions and which encodes a protein having GDP-mannose 4,6-dehydratase activity.(41) The transformant according to (39), wherein the GDP-mannose 4,6-dehydratase is a protein selected from the group consisting of the following (a) to (c):(a) a protein comprising the amino acid sequence represented by SEQ ID NO:2;(b) a protein consisting of an amino acid sequence wherein one or more amino acid residues are deleted, substituted, inserted and/or added in the amino acid sequence represented by SEQ ID NO:2 and having GDP-mannose 4,6-dehydratase activity;(c) a protein consisting of an amino acid sequence which has 80% or more homology to the amino acid sequence represented by SEQ ID NO:2 and having GDP-mannose 4,6-dehydratase activity.(42) The transformant according to (39), wherein the GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase is a protein encoded by a DNA selected from the group consisting of the following (a) and (b):(a) a DNA comprising the nucleotide sequence represented by SEQ ID NO:3;(b) a DNA which hybridizes with the DNA consisting of the nucleotide sequence represented by SEQ ID NO:3 under stringent conditions and which encodes a protein having GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase activity.(43) The transformant according to (39), wherein the GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase is a protein selected from the group consisting of the following (a) to (c):(a) a protein comprising the amino acid sequence represented by SEQ ID NO:4;(b) a protein consisting of an amino acid sequence wherein one or more amino acid residues are deleted, substituted, inserted and/or added in the amino acid sequence represented by SEQ ID NO:4 and having GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase activity;(c) a protein consisting of an amino acid sequence which has 80% or more homology to the amino acid sequence represented by SEQ ID NO:4 and having GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase activity.(44) The transformant according to (37) or (38), wherein the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain is α1,6-fucosyltransferase.(45) The transformant according to (44), wherein the α1,6-fucosyltransferase is a protein encoded by a DNA selected from the group consisting of the following (a) to (d):(a) a DNA comprising the nucleotide sequence represented by SEQ ID NO:5;(b) a DNA comprising the nucleotide sequence represented by SEQ ID NO:6;(c) a DNA which hybridizes with the DNA consisting of the nucleotide sequence represented by SEQ ID NO:5 under stringent conditions and which encodes a protein having α1,6-fucosyltransferase activity;(d) a DNA which hybridizes with the DNA consisting of the nucleotide sequence represented by SEQ ID NO:6 under stringent conditions and which encodes a protein having α1,6-fucosyltransferase activity.(46) The transformant according to (44), wherein the α1,6-fucosyltransferase is a protein selected from the group consisting of the following (a) to (f):(a) a protein comprising the amino acid sequence represented by SEQ ID NO:7;(b) a protein comprising the amino acid sequence represented by SEQ ID NO:8;(c) a protein consisting of an amino acid sequence wherein one or more amino acid residues are deleted, substituted, inserted and/or added in the amino acid sequence represented by SEQ ID NO:7 and having α1,6-fucosyltransferase activity;(d) a protein consisting of an amino acid sequence wherein one or more amino acid residues are deleted, substituted, inserted and/or added in the amino acid sequence represented by SEQ ID NO:8 and having α1,6-fucosyltransferase activity;(e) a protein consisting of an amino acid sequence which has 80% or more homology to the amino acid sequence represented by SEQ ID NO:7 and having α1,6-fucosyltransferase activity;(f) a protein consisting of an amino acid sequence which has 80% or more homology to the amino acid sequence represented by SEQ ID NO:8 and having α1,6-fucosyltransferase activity.(47) The transformant according to (46), wherein the transformant is FERM BP-8467.(48) The transformant according to any one of (36) to (47), wherein the host cell is a cell
selected from the group consisting of the following (a) to (i):(a) a CHO cell derived from Chinese hamster ovary tissue;(b) a rat myeloma cell line YB2/3HL.P2.G11.16Ag.20 cell;(c) a mouse myeloma cell line NS0 cell;(d) a mouse myeloma cell line SP2/0-Ag14 cell;(e) a BHK cell derived from Syrian hamster kidney tissue;(f) an antibody-producing hybridoma cell;(g) a human leukemia cell line Namalwa cell;(h) an embryonic stem cell;(i) a fertilized egg cell.(49) A process for producing the antibody composition according to any one of (1) to (35), which comprises culturing the transformant according to any one of (36) to (48) in a medium to form and accumulate the antibody composition in the culture, and recovering and purifying the antibody composition from the culture.(50) The antibody composition according to any one of (1) to (35), which is obtained by the process according to (49).(51) A pharmaceutical composition comprising the antibody composition according to any one of (1) to (35) and (50) as an active ingredient.(52) A therapeutic agent for diseases relating to a human CC chemokine receptor 4 (CCR4), comprising the antibody composition according to any one of (1) to (35) and (50) as an active ingredient.(53) The therapeutic agent according to (52), wherein the diseases relating to a human CC chemokine receptor 4 (CCR4) are cancer or inflammatory diseases.(54) A method for treating diseases related to a human CC chemokine receptor 4 (CCR4), which comprises administering to a patient the antibody composition according to any one of (1) to (35) and (50).(55) Use of the antibody composition according to any one of (1) to (35) and (50) to produce a therapeutic agent for diseases related to a human CC chemokine receptor 4 (CCR4).
BRIEF DESCRIPTION OF THE DRAWINGS
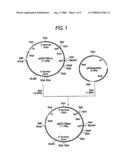
[0024]FIG. 1 shows the steps for constructing plasmid pKOFUT8Neo.
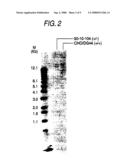
[0025]FIG. 2 shows the result of genomic Southern analysis of a hemi-knockout clone wherein one copy of the FUT8 allele was disrupted in CHO/DG44 cell. The lanes respectively show the following, from left to right: molecular weight marker, hemi-knockout clone 50-10-104, and parent cell CHO/DG44.
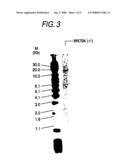
[0026]FIG. 3 shows the result of genomic Southern analysis of double-knockout clone WK704 wherein both FUT8 alleles were disrupted in CHO/DG44 cell. The arrow indicates the detection spot of a positive fragment resulting from homologous recombination.
[0027]FIG. 4 shows the result of genomic Southern analysis of a clone obtained by removing a drug-resistance gene from a double-knockout clone wherein both FUT8 alleles were disrupted in CHO/DG44 cell. The lanes respectively show the following, from left to right: molecular weight marker, drug resistance gene-removed double-knockout clone 4-5-C3, double-knockout clone WK704, hemi-knockout clone 50-10-104, and parent cell CHO/DG44.
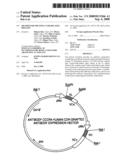
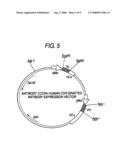
[0028]FIG. 5 shows the schematic view of the produced expression vector for the anti-CCR4 human CDR-grafted antibody.
[0029]FIG. 6 shows the reactivity of purified Ms705/CCR4 antibody and DG44/CCR4 antibody at varied concentrations to a CCR4-partial peptide measured by ELISA. The numbers on the abscissa indicate the antibody concentration and those on the ordinate indicate the absorbance at each antibody concentration. quadrature corresponds to the DG44/CCR4 antibody, and .box-solid. corresponds to the Ms705/CCR4 antibody.
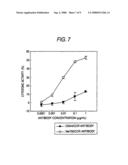
[0030]FIG. 7 shows the ADCC activity of purified Ms705/CCR4 antibody and DG44/CCR4 antibody at varied concentrations to human CCR4-highly expressing cells (CCR4/EL-4). The numbers on the abscissa indicate the antibody concentration and those on the ordinate indicate the cytotoxic activity at each antibody concentration. corresponds to the DG44/CCR4 antibody, and ∘ corresponds to the Ms705/CCR4 antibody.
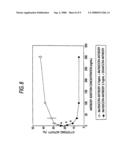
[0031]FIG. 8 shows in vitro ADCC activities of anti-CCR4 human CDR-grafted antibody compositions prepared by adding 0 to 300 ng/ml of the DG44/CCR4 antibody or the Ms705/CCR4 antibody to 3.7 ng/ml of the Ms705/CCR4 antibody, to the CCR4/EL4 cells. The ordinate shows the cytotoxic activity, and the abscissa shows the added antibody concentration. corresponds to the activity of the antibody composition prepared by adding the DG44/CCR4 antibody to 3.7 ng/ml of the Ms705/CCR4 antibody, and ∘ corresponds to the activity of the antibody composition prepared by adding the Ms705/CCR4 antibody to 3.7 ng/ml of the Ms705/CCR4 antibody. In the drawing, * corresponds to an antibody composition in which the ratio of an antibody having a sugar chain in which fucose is not bound is 20% or more, among the antibody compositions prepared by adding the DG44/CCR4 antibody to 3.7 ng/ml of the Ms705/CCR4 antibody.
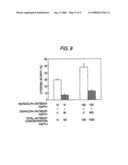
[0032]FIG. 9 shows in vitro ADCC activities of an antibody composition comprising the Ms705/CCR4 antibody alone, or an antibody composition prepared by mixing the Ms705/CCR4 antibody with a 9-fold amount of the DG44/CCR4 antibody, to CCR4/EL4 cells. The ordinate shows the cytotoxic activity. The numerical values plotted as the abscissa show the concentration of the Ms705/CCR4 antibody (ng/ml), the concentration of the added DG44/CCR4 antibody (ng/ml) and the total antibody concentration (ng/ml), respectively, from the upper row. quadrature corresponds to the activity of the antibody composition comprising the Ms705/CCR4 antibody alone, and .box-solid. corresponds to the activity of the antibody composition prepared by mixing the Ms705/CCR4 antibody with a 9-fold amount of the DG44/CCR4 antibody.
DETAILED DESCRIPTION OF THE INVENTION
[0033]An example of the antibody composition of the present invention comprising a recombinant antibody molecule which specifically binds to human CCR4 and has complex type N-glycoside-linked sugar chains in the Fc region, wherein the complex type N-glycoside-linked sugar chains have a structure in which fucose is not bound to N-acetylglucosamine in the reducing end in the sugar chains, is an antibody composition wherein the complex type N-glycoside linked sugar chains have a structure in which 1-position of fucose is not bound to 6-position of N-acetylglucosamine in the reducing end through α-bond.
[0034]An antibody molecule has the Fc region, to which N-glycoside-linked sugar chains are bound. Therefore, two sugar chains are bound to one antibody molecule.
[0035]The N-glycoside-linked sugar chains include complex type sugar chains having one or multiple number of parallel galactose-N-acetylglucosamine (hereinafter referred to as Gal-GlcNAc) side chains in the non-reducing end of the core structure and having sialic acid, bisecting N-acetylglucosamine or the like in the non-reducing end of Gal-GlcNAc.
[0036]In the present invention, the complex type N-glycoside-linked sugar chain is represented by the following chemical formula 1.
[0037]In the present invention, the sugar chain to which fucose is not bound includes a sugar chain represented by the above chemical formula in which fucose is not bound to N-acetylglucosamine in the reducing end. The sugar chain in the non-reducing end may have any structure.
[0038]Accordingly, the antibody composition of the present invention comprises an antibody molecule having the same sugar chain structure or antibody molecules having different sugar chain structures, so long as the antibody composition has the above sugar chain structure.
[0039]The expression "fucose is not bound to the N-acetylglucosamine in the reducing end in the sugar chains" as used herein means that fucose is not substantially bound thereto. The "antibody composition in which fucose is not substantially bound" specifically refers to an antibody composition in which fucose is not substantially detected, i.e., the content of fucose is below the detection limit, when subjected to the sugar chain analysis described in 4 below. The antibody composition of the present invention in which fucose is not bound to the N-acetylglucosamine in the reducing end in the sugar chains has high ADCC activity.
[0040]The ratio of an antibody molecule having sugar chains in which fucose is not bound to the N-acetylglucosamine in the reducing end in an antibody composition comprising an antibody molecule having complex type N-glycoside-linked sugar chains in the Fc region can be determined by releasing the sugar chains from the antibody molecule by known methods such as hydrazinolysis and enzyme digestion [Seibutsukagaku Jikkenho (Biochemical Experimentation Methods) 23--Totanpakushitsu Tosa Kenkyuho (Methods of Studies on Glycoprotein Sugar Chains), Gakkai Shuppan Center, edited by Reiko Takahashi (1989)], labeling the released sugar chains with a fluorescent substance or radioisotope, and separating the labeled sugar chains by chromatography. Alternatively, the released sugar chains may be analyzed by the HPAED-PAD method [J. Liq. Chromatogr., 6, 1577 (1983)] to determine the ratio.
[0041]The antibody compositions of the present invention include recombinant antibody compositions comprising a recombinant antibody molecule which specifically binds to an extracellular region of human CCR4, wherein the complex type N-glycoside-linked sugar chains have a structure in which fucose is not bound to N-acetylglucosamine in the reducing end in the sugar chains. Among these, the antibody composition which has no reactivity to a human blood platelet is preferred.
[0042]Examples of the extracellular region of human CCR4 include a polypeptide or peptide comprising the amino acid sequence at positions 1 to 39, 98 to 112, 176 to 206 or 271 to 284 of the amino acid sequence represented by SEQ ID NO:36, preferably, comprising the sequence at positions 2 to 29 of the amino acid sequence represented by SEQ ID NO:36 (SEQ ID NO:37), more preferably, comprising the sequence at positions 12 to 29 of the amino acid sequence represented by SEQ ID NO:36 (SEQ ID NO:38), and most preferably, represented by the sequence at positions 12 to 25 of the amino acid sequence represented by SEQ ID NO:36. Accordingly, the antibody composition of the present invention is preferably an antibody composition which specifically binds to a polypeptide or peptide comprising the amino acid sequence at positions 1 to 39, 98 to 112, 176 to 206 or 271 to 284 of the amino acid sequence represented by SEQ ID NO:36, preferably, comprising the sequence at positions 2 to 29 of the amino acid sequence represented by SEQ ID NO:36 (SEQ ID NO:37), more preferably, comprising the sequence at positions 12 to 29 of the amino acid sequence represented by SEQ ID NO:36 (SEQ ID NO:38), and most preferably, comprising the sequence at positions 12 to 25 of the amino acid sequence represented by SEQ ID NO:36, wherein the complex type N-glycoside-linked sugar chains have a structure in which fucose is not bound to N-acetylglucosamine in the reducing end in the sugar chains.
[0043]The expression "having no reactivity to a human blood platelet" means that an antibody does not substantially react with a human blood platelet, and specifically, the reactivity is not observed in an analysis using an apparatus capable of detecting the binding of an antibody to a cell such as a flow cytometer.
[0044]Also, preferably, the antibody compositions of the present invention include an antibody composition which has lower binding activity to a peptide comprising the sequence at positions 13 to 25 of the amino acid sequence represented by SEQ ID NO:36 wherein at least one of tyrosine residues at positions 16, 19, 20 and 22 is sulfated, in comparison with its binding activity to a peptide comprising the sequence at positions 13 to 25 of the amino acid sequence represented by SEQ ID NO:36.
[0045]The antibody compositions of the present invention also include a recombinant antibody composition which have no activity of inhibiting binding of TARC or MDC, which are CCR4 ligands, to CCR4. The expression "not having the activity to inhibit the binding of CCR4 ligands such as TARC and MDC to CCR4" means that the antibody composition is not substantially capable of inhibiting the binding of the ligand to the receptor, and specifically, the antibody composition even at a high concentration does not affect the binding of the ligand to the receptor. Herein, the high concentration means an antibody composition at 50 μg/ml or more.
[0046]Furthermore, the antibody compositions of the present invention include antibody compositions comprising a recombinant antibody molecule which specifically binds to a cell in which human CCR4 is expressed (hereinafter referred to as human CCR4-expressing cell) and has complex type N-glycoside-linked sugar chains in the Fc region, wherein the complex type N-glycoside-linked sugar chains have a structure in which fucose is not bound to N-acetylglucosamine in the reducing end in the sugar chains, and the like. Among such antibody compositions, they are preferably antibody compositions having cytotoxic activity against a human CCR4-expressing cell.
[0047]The human CCR4-expressing cells include helper T2 cells (hereinafter referred to as Th2 cells) and the like. Th2 cells are cells producing interleukin-4 (hereinafter referred to as IL-4), interleukin-5 (hereinafter referred to as IL-5) and interleukin-13 (hereinafter referred to as IL-13).
[0048]The cytotoxic activity includes complement-dependent cytotoxic activity (hereinafter referred to as CDC activity), antibody-dependent cell-mediated cytotoxic activity (hereinafter referred to as ADCC activity), and the like.
[0049]The antibody compositions having cytotoxic activity against a human CCR4-expressing cell, wherein the complex type N-glycoside-linked sugar chains have a structure in which fucose is not bound to N-acetylglucosamine in the reducing end in the sugar chains are capable of injuring Th2 cells which are CCR4-expressing cells by the cytotoxic activity possessed by the antibody composition to thereby eliminate the Th2 cells. Accordingly, the antibody composition can suppress production of interleukins such as IL-4, IL-5 and IL-13 from the Th-2 cells.
[0050]The antibody compositions of the present invention include compositions of human chimeric antibodies, compositions of human CDR-grafted antibodies, compositions of human antibodies and compositions of fragments of such antibodies.
[0051]The "human chimeric antibody" refers to an antibody comprising VH and VL of an antibody derived from a non-human animal, and CH and CL of a human antibody. As the non-human animal, any animal can be used so long as hybridomas can be prepared from the animal. Suitable animals include mouse, rat, hamster, rabbit and the like.
[0052]The human chimeric antibody composition of the present invention can be produced by obtaining cDNAs encoding VH and VL of a non-human animal-derived antibody which specifically binds to human CCR4, inserting the cDNAs into an expression vector for animal cells which carries genes encoding CH and CL of a human antibody to construct a human chimeric antibody expression vector, and introducing the vector into an animal cell to induce expression.
[0053]As the CH for the human chimeric antibody, any CH of antibodies belonging to human immunoglobulin (hereinafter referred to as hIg) may be used. Preferred are those of antibodies belonging to the hIgG class, which may be of any subclass, e.g., hIgG1, hIgG2, hIgG3 and hIgG4. As the CL for the human chimeric antibody, any CL of antibodies belonging to hIg, such as class κ or class λ, may be used.
[0054]Examples of the human chimeric antibody compositions of the present invention which specifically bind to human CCR4 include: an anti-human CCR4 chimeric antibody comprising CDR1, CDR2 and CDR3 of VH consisting of the amino acid sequences represented by SEQ ID NOs:14, 15 and 16, respectively, and/or CDR1, CDR2 and CDR3 of VL consisting of the amino acid sequences represented by SEQ ID NOs:17, 18 and 19, respectively; an anti-human CCR4 chimeric antibody wherein the VH of the antibody comprises the amino acid sequence represented by SEQ ID NO:21 and/or the VL of the antibody comprises the amino acid sequence represented by SEQ ID NO:23; and an anti-human CCR4 chimeric antibody composition wherein the VH of the antibody consists of the amino acid sequence represented by SEQ ID NO:21, the CH of the human antibody consists of an amino acid sequence of the hIgG1 subclass, the VL of the antibody consists of the amino acid sequence represented by SEQ ID NO:23, and the CL of the human antibody consists of an amino acid sequence of the K class.
[0055]The "human CDR-grafted antibody" refers to an antibody in which CDRs of VH and VL of an antibody derived from a non-human animal are grafted into appropriate sites in VH and VL of a human antibody.
[0056]The human CDR-grafted antibody composition of the present invention can be produced by constructing cDNAs encoding V regions in which CDRs of VH and VL of a non-human animal-derived antibody which specifically binds to CCR4 are grafted into frameworks (hereinafter referred to as FR) of VH and VL of an arbitrary human antibody, inserting the resulting cDNAs into an expression vector for animal cells which has DNAs encoding the heavy chain constant region (hereinafter referred to as CH) and the light chain constant region (hereinafter referred to as CL) of a human antibody to construct a human CDR-grafted antibody expression vector, and introducing the expression vector into an animal cell to induce expression.
[0057]As the FR amino acid sequences of VH and VL of a human antibody, any of those derived from human antibodies can be used. Suitable sequences include the FR amino acid sequences of VH and VL of human antibodies registered in databases such as Protein Data Bank, and the amino acid sequences common to all FR subgroups of VH and VL of human antibodies (Sequences of Proteins of Immunological Interest, US Dept. Health and Human Services, 1991).
[0058]As the CH for the antibody of the present invention, any CH of antibodies belonging to hIg may be used. Preferred are those of antibodies belonging to the hIgG class, which may be of any subclass, e.g., hIgG1, hIgG2, hIgG3 and hIgG4. As the CL for the human CDR-grafted antibody, any CL of antibodies belonging to hIg, e.g., class K or class A, may be used.
[0059]An example of the human CDR-grafted antibody composition of the present invention is a human CDR-grafted antibody or antibody fragment composition comprising CDR1, CDR2 and CDR3 of VH consisting of the amino acid sequences represented by SEQ ID NOs:14, 15 and 16, respectively; and/or CDR1, CDR2 and CDR3 of VL consisting of the amino acid sequences represented by SEQ ID NOs:17, 18 and 19, respectively.
[0060]Preferred human CDR-grafted antibody compositions include: a human CDR-grafted antibody composition, wherein the VH of the antibody comprises the amino acid sequence represented by SEQ ID NO:24 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ala at position 40, Gly at position 42, Lys at position 43, Gly at position 44, Lys at position 76 and Ala at position 97 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:24, a human CDR-grafted antibody composition, wherein the VH of the antibody comprises the amino acid sequence represented by SEQ ID NO:25 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Thr at position 28 and Ala at position 97 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:25, and a human CDR-grafted antibody composition, wherein the VL of the antibody comprises the amino acid sequence represented by SEQ ID NO:26 or an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ile at position 2, Val at position 3, Gln at position 50 and Val at position 88 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:26. More preferred are the following antibody compositions: a human CDR-grafted antibody composition, wherein the VH of the antibody comprises an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ala at position 40, Gly at position 42, Lys at position 43, Gly at position 44, Lys at position 76 and Ala at position 97 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:24; and the VL of the antibody comprises an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ile at position 2, Val at position 3, Gln at position 50 and Val at position 88 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:26, and a human CDR-grafted antibody composition, wherein the VH of the antibody comprises an amino acid sequence in which at least one amino acid residue selected from the group consisting of Thr at position 28 and Ala at position 97 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:25; and the VL of the antibody comprises an amino acid sequence in which at least one amino acid residue selected from the group consisting of Ile at position 2, Val at position 3, Gln at position 50 and Val at position 88 is substituted with another amino acid residue in the amino acid sequence represented by SEQ ID NO:26.
[0061]A specific example of the human CDR-grafted antibody composition is a human CDR-grafted antibody composition wherein the VH of the antibody comprises an amino acid sequence selected from the group consisting of the amino acid sequences represented by SEQ ID NOs:24, 25, 27, 28, 29, 30, 31 and 32; a human CDR-grafted antibody composition wherein the VL of the antibody comprises an amino acid sequence selected from the group consisting of the amino acid sequences represented by SEQ ID NOs:26, 33, 34 and 35; and a human CDR-grafted antibody composition wherein the VH of the antibody comprises an amino acid sequence selected from the group consisting of the amino acid sequences represented by SEQ ID NOs: 24, 25, 27, 28, 29, 30, 31 and 32, and the VL of the antibody comprises an amino acid sequence selected from the group consisting of the amino acid sequences represented by SEQ ID NOs: 26, 33, 34 and 35.
[0062]The human CDR-grated antibody composition of the present invention is most preferably a CDR-grafted antibody composition wherein the VH of the antibody comprises the amino acid sequence represented by SEQ ID NO:27 or 28, and the VL of the antibody comprises the amino acid sequence represented by SEQ ID NO:35.
[0063]Also included within the scope of the present invention are antibodies and antibody fragments which specifically bind to an extracellular region of human CCR4 and have no reactivity to a human blood platelet, and consist of amino acid sequences wherein one or more amino acid residues are deleted, added, substituted and/or inserted in the above amino acid sequences.
[0064]The number of amino acid residues which are deleted, substituted, inserted and/or added is one or more and is not specifically limited, but it is within the range where deletion, substitution or addition is possible by known methods such as site-directed mutagenesis described in Molecular Cloning, A Laboratory Manual, Second Edition; Current Protocols in Molecular Biology; Nucleic Acids Research, 10, 6487 (1982); Proc. Natl. Acad. Sci. USA, 79, 6409 (1982); Gene, 34, 315 (1985); Nucleic Acids Research, 13, 4431 (1985); Proc. Natl. Acad. Sci. USA, 82, 488 (1985), etc. The suitable number is 1 to dozens, preferably 1 to 20, more preferably 1 to 10, further preferably 1 to 5.
[0065]The expression "one or more amino acid residues are deleted, substituted, inserted or added in the amino acid sequence of the antibody composition of the present invention" means that the amino acid sequence of the antibody composition contains deletion, substitution, insertion or addition of a single or plural amino acid residues at a single or plural residues at arbitrary positions therein. Deletion, substitution, insertion and addition may be simultaneously contained in one sequence, and amino acid residues to be substituted, inserted or added may be either natural or not. Examples of the natural amino acid residues are L-alanine, L-asparagine, L-aspartic acid, L-glutamine, L-glutamic acid, glycine, L-histidine, L-isoleucine, L-leucine, L-lysine, L-methionine, L-phenylalanine, L-proline, L-serine, L-threonine, L-tryptophan, L-tyrosine, L-valine and L-cysteine.
[0066]The followings are preferred examples of the amino acid residues capable of mutual substitution. The amino acid residues in the same group shown below can be mutually substituted. [0067]Group A: leucine, isoleucine, norleucine, valine, norvaline, alanine, 2-aminobutanoic acid, methionine, O-methylserine, t-butylglycine, t-butylalanine, cyclohexylalanine [0068]Group B: aspartic acid, glutamic acid, isoaspartic acid, isoglutamic acid, 2-aminoadipic acid, 2-aminosuberic acid [0069]Group C: asparagine, glutamine [0070]Group D: lysine, arginine, ornithine, 2,4-diaminobutanoic acid, 2,3-diaminopropionic acid [0071]Group E: proline, 3-hydroxyproline, 4-hydroxyproline [0072]Group F: serine, threonine, homoserine [0073]Group G: phenylalanine, tyrosine
[0074]The recombinant antibody fragment compositions of the present invention include compositions of antibody fragments which specifically bind to human CCR4 and which contain a part or the whole of the antibody Fc region in which fucose is not bound to the N-acetylglucosamine in the reducing end in complex type N-glycoside-linked sugar chains.
[0075]The antibody fragment compositions of the present invention include compositions of antibody fragments, e.g., Fab, Fab', F(ab')2, scFv, diabody, dsFv and a peptide comprising CDR, containing a part or the whole of the antibody Fc region in which fucose is not bound to the N-acetylglucosamine in the reducing end in complex type N-glycoside-linked sugar chains. When the antibody fragment composition does not contain a part or the whole of the antibody Fc region, the antibody fragment may be fused with a part or the whole of the Fc region of the antibody having sugar chains in which fucose is not bound to N-acetylglucosamine in the reducing end in the complex type N-glycoside-linked sugar chains as a fusion protein, or the antibody fragment may be used as a fusion protein composition with a protein comprising a part or the whole of the Fc region.
[0076]An Fab fragment is one of the fragments obtained by treatment of IgG with the proteolytic enzyme, papain (cleavage at amino acid residue 224 of H chain). It is an antibody fragment with a molecular weight of approximately 50,000 having antigen-binding activity and composed of the N-terminal half of H chain and the entire L chain linked by a disulfide bond.
[0077]The Fab fragment of the present invention can be obtained by treating the antibody composition of the present invention which specifically binds to human CCR4 with the proteolytic enzyme, papain. Alternatively, the Fab fragment may be produced by inserting DNA encoding the Fab fragment of the antibody into an expression vector for prokaryote or eukaryote, and introducing the vector into a prokaryote or eukaryote to induce expression.
[0078]An F(ab')2 fragment is one of the fragments obtained by treatment of IgG with the proteolytic enzyme, pepsin (cleavage at amino acid residue 234 of H chain). It is an antibody fragment with a molecular weight of approximately 100,000 having antigen-binding activity, which is slightly larger than the Fab fragments linked together by a disulfide bond at the hinge region.
[0079]The F(ab')2 fragment of the present invention can be obtained by treating the antibody composition of the present invention which specifically binds to human CCR4 with the proteolytic enzyme, pepsin. Alternatively, the F(ab')2 fragment may be prepared by binding Fab' fragments described below by a thioether bond or a disulfide bond.
[0080]An Fab' fragment is an antibody fragment with a molecular weight of approximately 50,000 having antigen-binding activity, which is obtained by cleaving the disulfide bond at the hinge region of the above F(ab')2 fragment.
[0081]The Fab' fragment of the present invention can be obtained by treating the F(ab')2 fragment composition of the present invention which specifically binds to human CCR4 with a reducing agent, dithiothreitol. Alternatively, the Fab' fragment may be produced by inserting DNA encoding the Fab' fragment of the antibody into an expression vector for prokaryote or eukaryote, and introducing the vector into a prokaryote or eukaryote to induce expression.
[0082]An scFv fragment is a VH-P-VL or VL-P-VH polypeptide in which one VH and one VL are linked via an appropriate peptide linker (hereinafter referred to as P) and which has antigen-binding activity.
[0083]The scFv fragment of the present invention can be produced by obtaining cDNAs encoding the VH and VL of the antibody composition of the present invention which specifically binds to human CCR4, constructing DNA encoding the scFv fragment, inserting the DNA into an expression vector for prokaryote or eukaryote, and introducing the expression vector into a prokaryote or eukaryote to induce expression.
[0084]A diabody is an antibody fragment which is an scFv dimer showing bivalent antigen binding activity, which may be either monospecific or bispecific.
[0085]The diabody of the present invention can be produced by obtaining cDNAs encoding the VH and VL of the antibody composition of the present invention which specifically binds to human CCR4, constructing DNA encoding scFv fragments with P having an amino acid sequence of 8 or less amino acid residues, inserting the DNA into an expression vector for prokaryote or eukaryote, and introducing the expression vector into a prokaryote or eukaryote to induce expression.
[0086]A dsFv fragment is an antibody fragment wherein polypeptides in which one amino acid residue of each of VH and VL is substituted with a cysteine residue are linked by a disulfide bond between the cysteine residues. The amino acid residue to be substituted with a cysteine residue can be selected based on antibody tertiary structure prediction according to the method proposed by Reiter, et al. (Protein Engineering, 7, 697-704 (1994)).
[0087]The dsFv fragment of the present invention can be produced by obtaining cDNAs encoding the VH and VL of the antibody composition of the present invention which specifically binds to human CCR4, constructing DNA encoding the dsFv fragment, inserting the DNA into an expression vector for prokaryote or eukaryote, and introducing the vector into a prokaryote or eukaryote to induce expression.
[0088]A peptide comprising CDR comprises one or more region CDR of VH or VL. A peptide comprising plural CDRs can be prepared by binding CDRs directly or via an appropriate peptide linker.
[0089]The peptide comprising CDR of the present invention can be produced by constructing DNA encoding CDR of VH and VL of the antibody composition of the present invention which specifically binds to human CCR4, inserting the DNA into an expression vector for prokaryote or eukaryote, and introducing the expression vector into a prokaryote or eukaryote to induce expression.
[0090]The peptide comprising CDR can also be produced by chemical synthesis methods such as the Fmoc method (fluorenylmethyloxycarbonyl method) and the tBoc method (t-butyloxycarbonyl method).
[0091]The transformant of the present invention includes any transformant that is obtained by introducing DNA encoding an antibody molecule which specifically binds to human CCR4 into a host cell and that produces the antibody composition of the present invention. Examples of such transformants include those obtained by introducing DNA encoding an antibody molecule which specifically binds to human CCR4 into host cells such as the following (a) or (b):
(a) a cell in which genome is modified so as to have deleted activity of an enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose;(b) a cell in which genome is modified so as to have deleted activity of an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain.
[0092]Specifically, the "modification of genome so as to have deleted activity of an enzyme" refers to introduction of mutation into an expression regulation region of a gene encoding the enzyme so as to delete the expression of the enzyme or introduction of mutation in the amino acid sequence of a gene encoding the enzyme so as to inactivate the enzyme. The "introduction of mutation" refers to carrying out modification of the nucleotide sequence on the genome such as deletion, substitution, insertion and/or addition in the nucleotide sequence. Complete inhibition of the expression or activity of the thus modified genomic gene refers to "knock out of the genomic gene".
[0093]Examples of the enzymes relating to the synthesis of the intracellular sugar nucleotide GDP-fucose include GDP-mannose 4,6-dehydratase (GMD), GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase (Fx) and the like.
[0094]Examples of the GDP-mannose 4,6-dehydratase include proteins encoded by the DNAs of the following (a) and (b):
(a) a DNA comprising the nucleotide sequence represented by SEQ ID NO:1;(b) a DNA which hybridizes with DNA consisting of the nucleotide sequence represented by SEQ ID NO:1 under stringent conditions and which encodes a protein having GDP-mannose 4,6-dehydratase activity.
[0095]Examples of the GDP-mannose 4,6-dehydratase also include proteins of the following (a) to (c):
(a) a protein comprising the amino acid sequence represented by SEQ ID NO:2;(b) a protein consisting of an amino acid sequence wherein one or more amino acid residues are deleted, substituted, inserted and/or added in the amino acid sequence represented by SEQ ID NO:2 and having GDP-mannose 4,6-dehydratase activity;(c) a protein consisting of an amino acid sequence which has 80% or more homology to the amino acid sequence represented by SEQ ID NO:2 and having GDP-mannose 4,6-dehydratase activity.
[0096]Examples of the GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase include proteins encoded by the DNAs of the following (a) and (b):
(a) a DNA comprising the nucleotide sequence represented by SEQ ID NO:3;(b) a DNA which hybridizes with DNA consisting of the nucleotide sequence represented by SEQ ID NO:3 under stringent conditions and which encodes a protein having GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase activity.
[0097]Examples of the GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase also include proteins of the following (a) to (c):
(a) a protein comprising the amino acid sequence represented by SEQ ID NO:4;(b) a protein consisting of an amino acid sequence wherein one or more amino acid residues are deleted, substituted, inserted and/or added in the amino acid sequence represented by SEQ ID NO:4 and having GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase activity;(c) a protein consisting of an amino acid sequence which has 80% or more homology to the amino acid sequence represented by SEQ ID NO:4 and having GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase activity.
[0098]An example of the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain is α1,6-fucosyltransferase.
[0099]In the present invention, examples of the α1,6-fucosyltransferase include proteins encoded by the DNAs of the following (a) to (d):
(a) a DNA comprising the nucleotide sequence represented by SEQ ID NO:5;(b) a DNA comprising the nucleotide sequence represented by SEQ ID NO:6;(c) a DNA which hybridizes with DNA consisting of the nucleotide sequence represented by SEQ ID NO:5 under stringent conditions and which encodes a protein having α1,6-fucosyltransferase activity;(d) a DNA which hybridizes with DNA consisting of the nucleotide sequence represented by SEQ ID NO:6 under stringent conditions and which encodes a protein having α1,6-fucosyltransferase activity, or proteins of the following (e) to (j):(e) a protein comprising the amino acid sequence represented by SEQ ID NO:7;(f) a protein comprising the amino acid sequence represented by SEQ ID NO:8;(g) a protein consisting of an amino acid sequence wherein one or more amino acid residues are deleted, substituted, inserted and/or added in the amino acid sequence represented by SEQ ID NO:7 and having α1,6-fucosyltransferase activity;(h) a protein consisting of an amino acid sequence wherein one or more amino acid residues are deleted, substituted, inserted and/or added in the amino acid sequence represented by SEQ ID NO:8 and having α1,6-fucosyltransferase activity;(i) a protein consisting of an amino acid sequence which has 80% or more homology to the amino acid sequence represented by SEQ ID NO:7 and having α1,6-fucosyltransferase activity;(j) a protein consisting of an amino acid sequence which has 80% or more homology to the amino acid sequence represented by SEQ ID NO:8 and having α1,6-fucosyltransferase activity.
[0100]The DNAs encoding the amino acid sequences of the enzymes relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose include a DNA comprising the nucleotide sequence represented by SEQ ID NO:1 or 3, and DNA which hybridizes with a DNA comprising the nucleotide sequence represented by SEQ ID NO:1 or 3 under stringent conditions and which encodes a protein having the enzyme activity relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose.
[0101]The DNAs encoding the amino acid sequences of α1,6-fucosyltransferase include a DNA comprising the nucleotide sequence represented by SEQ ID NO:5 or 6, and a DNA which hybridizes with DNA comprising the nucleotide sequence represented by SEQ ID NO:5 or 6 under stringent conditions and which encodes a protein having α1,6-fucosyltransferase activity.
[0102]In the present invention, the DNA which hybridizes under stringent conditions refers to a DNA which is obtained by colony hybridization, plaque hybridization, Southern hybridization or the like using, for example, a DNA consisting of the nucleotide sequence represented by SEQ ID NO:1, 3, 5 or 6 or a fragment thereof as a probe. A specific example of such DNA is a DNA which can be identified by performing hybridization at 65° C. in the presence of 0.7 to 1.0 M sodium chloride using a filter with colony- or plaque-derived DNA immobilized thereon, and then washing the filter at 65° C. with a 0.1 to 2-fold concentration SSC solution (1-fold concentration SSC solution: 150 mM sodium chloride and 15 mM sodium citrate). Hybridization can be carried out according to the methods described in Molecular Cloning, A Laboratory Manual, Second Edition, Cold Spring Harbor Laboratory Press (1989) (hereinafter referred to as Molecular Cloning, Second Edition); Current Protocols in Molecular Biology, John Wiley & Sons (1987-1997) (hereinafter referred to as Current Protocols in Molecular Biology); DNA Cloning 1: Core Techniques, A Practical Approach, Second Edition, Oxford University (1995), etc. Specifically, the DNA capable of hybridization under stringent conditions includes DNA having at least 60% or more homology, preferably 70% or more homology, more preferably 80% or more homology, further preferably 90% or more homology, particularly preferably 95% or more homology, most preferably 98% or more homology to the nucleotide sequence represented by SEQ ID NO:1, 3, 5 or 6.
[0103]In the present invention, the protein consisting of an amino acid sequence wherein one or more amino acid residues are deleted, substituted, inserted and/or added in the amino acid sequence represented by SEQ ID NO:2 or 4 and having the activity of an enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose, or the protein consisting of an amino acid sequence wherein one or more amino acid residues are deleted, substituted, inserted and/or added in the amino acid sequence represented by SEQ ID NO:7 or 8 and having α1,6-fucosyltransferase activity can be obtained, for example, by introducing a site-directed mutation into DNA having the nucleotide sequence represented by SEQ ID NO:1, 3, 5 or 6 by site-directed mutagenesis described in Molecular Cloning, Second Edition; Current Protocols in Molecular Biology; Nucleic Acids Research, 10, 6487 (1982); Proc. Natl. Acad. Sci. USA, 79, 6409 (1982); Gene, 34, 315 (1985); Nucleic Acids Research, 13, 4431 (1985); Proc. Natl. Acad. Sci. USA, 82, 488 (1985), etc. The number of amino acid residues which are deleted, substituted, inserted and/or added is one or more, and is not specifically limited, but it is within the range where deletion, substitution or addition is possible by known methods such as the above site-directed mutagenesis. The suitable number is 1 to dozens, preferably 1 to 20, more preferably 1 to 10, further preferably 1 to 5.
[0104]The protein consisting of an amino acid sequence which has 80% or more homology to the amino acid sequence represented by SEQ ID NO:2, 4, 7 or 8 and having GDP-mannose 4,6-dehydratase activity, GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase activity or α1,6-fucosyltransferase activity includes a protein having at least 80% or more homology, preferably 85% or more homology, more preferably 90% or more homology, further preferably 95% or more homology, particularly preferably 97% or more homology, most preferably 99% or more homology to the amino acid sequence represented by SEQ ID NO:2, 4, 7 or 8, respectively, as calculated by use of analysis software such as BLAST [J. Mol. Biol., 215, 403 (1990)] or FASTA [Methods in Enzymology, 183, 63 (1990)].
[0105]The host cell used in the present invention, that is, the host cell in which the activity of an enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain is deleted may be obtained by any technique capable of deleting the above enzyme activity. For example, the following techniques can be employed for deleting the above enzyme activity:
(a) gene disruption targeting at a gene encoding the enzyme;(b) introduction of a dominant-negative mutant of a gene encoding the enzyme;(c) introduction of a mutation into the enzyme;(d) inhibition of transcription or translation of a gene encoding the enzyme;(e) selection of a cell line resistant to a lectin which recognizes a sugar chain structure in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain.
[0106]As the lectin which recognizes a sugar chain structure in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain, any lectin capable of recognizing the sugar chain structure can be used. Specific examples include lentil lectin LCA (lentil agglutinin derived from Lens culinaris), pea lectin PSA (pea lectin derived from Pisum sativum), broad bean lectin VFA (agglutinin derived from Vicia faba), Aleuria aurantia lectin AAL (lectin derived from Aleuria aurantia) and the like.
[0107]The "cell resistant to a lectin" refers to a cell in which growth is not inhibited by the presence of a lectin at an effective concentration. The "effective concentration" is a concentration higher than the lowest concentration that does not allow the normal growth of a cell prior to the genome modification (hereinafter referred to also as parent cell line), preferably equal to the lowest concentration that does not allow the normal growth of a cell prior to the genome modification, more preferably 2 to 5 times, further preferably 10 times, most preferably 20 or more times the lowest concentration that does not allow the normal growth of a cell prior to the modification of the genomic gene.
[0108]The effective concentration of lectin that does not inhibit growth may be appropriately determined according to each cell line. It is usually 10 μg/ml to 10 mg/ml, preferably 0.5 mg/ml to 2.0 mg/ml.
[0109]The host cell for producing the antibody composition of the present invention may be any of the above host cells capable of expressing the antibody composition of the present invention. For example, yeast cells, animal cells, insect cells and plant cells can be used. Examples of the cells include those described in 1 below. Specifically, preferred among animal cells are CHO cell derived from Chinese hamster ovary tissue, rat myeloma cell line YB2/3HL.P2.G11.16Ag.20, mouse myeloma cell line NS0, mouse myeloma cell line SP2/0-Ag14, BHK cell derived from Syrian hamster kidney tissue, an antibody-producing hybridoma cell, human leukemia cell line Namalwa, an embryonic stem cell, and a fertilized egg cell.
[0110]A specific example of the transformant of the present invention is Ms705/CCR4, which is a transformant derived from Chinese hamster ovary tissue-derived CHO cell line CHO/DG44 and carrying an introduced gene of the anti-CCR4 antibody of the present invention. The transformant Ms705/CCR4 derived from CHO cell line CHO/DG44 was deposited with International Patent Organism Depositary, National Institute of Advanced Industrial Science and Technology, Central 6, 1, Higashi 1-chome, Tsukuba-shi, Ibaraki, Japan, on Sep. 9, 2003 with accession No. FERM BP-8467.
[0111]Described below are the method for preparing a cell producing the antibody composition of the present invention, the method for producing the antibody composition of the present invention, the method for analyzing the antibody composition of the present invention and the method for utilizing the antibody composition of the present invention.
1. Preparation of a Cell Producing the Antibody Composition of the Present Invention
[0112]The cell producing the antibody composition of the present invention (hereinafter referred to as the cell of the present invention) can be prepared by preparing a host cell used for the production of the antibody composition of the present invention by the following techniques and then introducing a gene encoding the anti-human CCR4 antibody into the host cell by the method described in 2 below.
(1) Gene Disruption Technique Targeting at a Gene Encoding an Enzyme
[0113]The host cell used for the production of the antibody composition of the present invention can be prepared by a gene disruption technique targeting a gene encoding an enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain. Examples of the enzymes relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose include GDP-mannose 4,6-dehydratase (hereinafter referred to as GMD) and GDP-4-keto-6-deoxy-D-mannose 3,5-epimerase (hereinafter referred to as Fx). Examples of the enzymes relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain include α1,6-fucosyltransferase and α-L-fucosidase.
[0114]The gene as used herein includes DNA and RNA.
[0115]The method of gene disruption may be any method capable of disrupting the gene encoding the target enzyme. Useful methods include the antisense method, the ribozyme method, the homologous recombination method, the RNA-DNA oligonucleotide method (hereinafter referred to as the RDO method), the RNA interference method (hereinafter referred to as the RNAi method), the method using a retrovirus and the method using a transposon. These methods are specifically described below.
(a) Preparation of the Host Cell for the Production of the Antibody Composition of the Present Invention by the Antisense Method or the Ribozyme Method
[0116]The host cell used for the production of the antibody composition of the present invention can be prepared by the antisense method or the ribozyme method described in Cell Technology, 12, 239 (1993); BIO/TECHNOLOGY, 17, 1097 (1999); Hum. Mol. Genet., 5, 1083 (1995); Cell Technology, 13, 255 (1994); Proc. Natl. Acad. Sci. U.S.A., 96, 1886 (1999); etc. targeting at a gene encoding an enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain, for example, in the following manner.
[0117]A cDNA or a genomic DNA encoding an enzyme relating to the synthesis of the intracellular sugar nucleotide, GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain is prepared.
[0118]The nucleotide sequence of the prepared cDNA or genomic DNA is determined.
[0119]Based on the determined DNA sequence, an antisense gene or a ribozyme of appropriate length is designed which comprises a DNA moiety encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain, non-translated regions and introns.
[0120]In order to express the antisense gene or ribozyme in a cell, a recombinant vector is prepared by inserting a fragment or full-length of the prepared DNA into a site downstream of a promoter in an appropriate expression vector.
[0121]The recombinant vector is introduced into a host cell suited for the expression vector to obtain a transformant.
[0122]The host cell used for the production of the antibody composition of the present invention can be obtained by selecting a transformant using, as a marker, the activity of the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain. The host cell used for the production of the antibody composition of the present invention can also be obtained by selecting a transformant using, as a marker, the sugar chain structure of a glycoprotein on the cell membrane or the sugar chain structure of the produced antibody molecule.
[0123]As the host cell used for the production of the antibody composition of the present invention, any yeast cell, animal cell, insect cell, plant cell, or the like can be used so long as it has a gene encoding the target enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain. Examples of the host cells include those described in 2 below.
[0124]The expression vectors that can be employed are those capable of autonomous replication or integration into the chromosome in the above host cells and comprising a promoter at a position appropriate for the transcription of the designed antisense gene or ribozyme. Examples of the expression vectors include those described in 2 below.
[0125]Introduction of a gene into various host cells can be carried out by the methods suitable for introducing a recombinant vector into various host cells described in 2 below.
[0126]Selection of a transformant using, as a marker, the activity of an enzyme relating to the synthesis of an intracellular sugar nucleotide GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be carried out, for example, by the following methods.
Methods for Selecting a Transformant
[0127]A cell in which the activity of an enzyme relating to the synthesis of the intracellular sugar nucleotide GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain is deleted can be selected by measuring the activity of the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain using biochemical methods or genetic engineering techniques described in Shin Seikagaku Jikken Koza (New Lectures on Experiments in Biochemistry) 3--Saccharides I, Glycoprotein (Tokyo Kagaku Dojin), edited by The Japanese Biochemical Society (1988); Cell Technology, Extra Edition, Experimental Protocol Series, Glycobiology Experimental Protocol, Glycoprotein, Glycolipid and Proteoglycan (Shujunsha), edited by Naoyuki Taniguchi, Akemi Suzuki, Kiyoshi Furukawa and Kazuyuki Sugawara (1996); Molecular Cloning, Second Edition; Current Protocols in Molecular Biology; and the like. An example of the biochemical methods is a method in which the enzyme activity is evaluated using an enzyme-specific substrate. Examples of the genetic engineering techniques include Northern analysis and RT-PCR in which the amount of mRNA for a gene encoding the enzyme is measured.
[0128]Selection of a transformant using, as a marker, the sugar chain structure of a glycoprotein on the cell membrane can be carried out, for example, by the method described in 1(5) below. Selection of a transformant using, as a marker, the sugar chain structure of a produced antibody molecule can be carried out, for example, by the methods described in 4 or 5 below.
[0129]Preparation of a cDNA encoding an enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be carried out, for example, by the following method.
Preparation of cDNA
[0130]Total RNA or mRNA is prepared from a various host cell tissue or cell.
[0131]A cDNA library is prepared from the total RNA or mRNA.
[0132]Degenerative primers are prepared based on the amino acid sequence of an enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain, and a gene fragment encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain is obtained by PCR using the prepared cDNA library as a template.
[0133]A DNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be obtained by screening the cDNA library using the obtained gene fragment as a probe.
[0134]As the mRNA of a human or non-human animal tissue or cell, commercially available one (for example, manufactured by Clontech) may be use, or it may be prepared from a human or non-human animal tissue or cell in the following manner.
[0135]The methods for preparing total RNA from a human or non-human animal tissue or cell include the guanidine thiocyanate-cesium trifluoroacetate method [Methods in Enzymology, 154, 3 (1987)], the acidic guanidine thiocyanate-phenol-chloroform (AGPC) method [Analytical Biochemistry, 162, 156 (1987); Experimental Medicine, 9, 1937 (1991)] and the like.
[0136]The methods for preparing mRNA as poly(A)+RNA from the total RNA include the oligo (dT) immobilized cellulose column method (Molecular Cloning, Second Edition).
[0137]It is also possible to prepare mRNA by using a commercially available kit such as Fast Track mRNA Isolation Kit® (manufactured by Invitrogen) or Quick Prep mRNA Purification Kit® (manufactured by Pharmacia).
[0138]A cDNA library is prepared from the obtained mRNA of a human or non-human animal tissue or cell. The methods for preparing the cDNA library include the methods described in Molecular Cloning, Second Edition; Current Protocols in Molecular Biology; A Laboratory Manual, 2nd Ed. (1989); etc., and methods using commercially available kits such as SuperScript Plasmid System for cDNA Synthesis and Plasmid Cloning® (manufactured by Life Technologies) and ZAP-cDNA Synthesis Kit® (manufactured by STRATAGENE).
[0139]As the cloning vector for preparing the cDNA library, any vectors, e.g. phage vectors and plasmid vectors, can be used so long as they are autonomously replicable in Escherichia coli K12. Examples of suitable vectors include ZAP Express® [manufactured by STRATAGENE; Strategies, 5, 58 (1992)], pBluescript II SK(+) [Nucleic Acids Research, 17, 9494 (1989)], λZAP II® (manufactured by STRATAGENE), λgt10, λgt11 [DNA Cloning, A Practical Approach, 1, 49 (1985)], λTriplEx® (manufactured by Clontech), λExCell® (manufactured by Pharmacia), pT7T318U® (manufactured by Pharmacia), pcD2 [Mol Cell Biol., 3, 280 (1983)] and pUC18 [Gene, 33, 103 (1985)].
[0140]Any microorganism can be used as the host microorganism for preparing the cDNA library, but Escherichia coli is preferably used. Examples of suitable host microorganisms are Escherichia coli XL1-Blue MRF'® [manufactured by STRATAGENE; Strategies, 5, 81 (1992)], Escherichia coli C600 [Genetics, 39, 440 (1954)], Escherichia coli Y1088 [Science, 222, 778 (1983)], Escherichia coli Y1090 [Science, 222, 778 (1983)], Escherichia coli NM522 [J. Mol. Biol., 166, 1 (1983)], Escherichia coli K802 [J. Mol. Biol., 16, 118 (1966)] and Escherichia coli JM105 [Gene, 38, 275 (1985)].
[0141]The cDNA library may be used as such in the following analysis. Alternatively, in order to efficiently obtain full-length cDNAs by decreasing the ratio of partial cDNAs, a cDNA library prepared using the oligo-cap method developed by Sugano, et al. [Gene, 138, 171 (1994); Gene, 200, 149 (1997); Protein, Nucleic Acid and Enzyme, 41, 603 (1996); Experimental Medicine, 11, 2491 (1993); cDNA Cloning (Yodosha) (1996); Methods for Preparing Gene Libraries (Yodosha) (1994)] may be used in the following analysis.
[0142]Degenerative primers specific for the 5'-terminal and 3'-terminal nucleotide sequences of a nucleotide sequence presumed to encode the amino acid sequence of an enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain are prepared based on the amino acid sequence of the enzyme. A gene fragment encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be obtained by DNA amplification by PCR [PCR Protocols, Academic Press (1990)] using the prepared cDNA library as a template.
[0143]It can be confirmed that the obtained gene fragment is a cDNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain by analyzing the nucleotide sequence by generally employed methods such as the dideoxy method of Sanger, et al. [Proc. Natl. Acad. Sci. U.S.A., 74, 5463 (1977)] or by use of nucleotide sequencers such as ABI PRISM 377 DNA Sequencer (manufactured by Applied Biosystems).
[0144]A DNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be obtained from the cDNA or cDNA library synthesized from the mRNA contained in a human or non-human animal tissue or cell by colony hybridization or plaque hybridization (Molecular Cloning, Second Edition) using the above gene fragment as a probe.
[0145]A cDNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can also be obtained by amplification by PCR using the cDNA or cDNA library synthesized from the mRNA contained in a human or non-human animal tissue or cell as a template and using the primers used for obtaining the gene fragment encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain.
[0146]The nucleotide sequence of the obtained cDNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be determined by generally employed sequencing methods such as the dideoxy method of Sanger, et al. [Proc. Natl. Acad. Sci. U.S.A., 74, 5463 (1977)] or by use of nucleotide sequencers such as ABI PRISM 377 DNA Sequencer (manufactured by Applied Biosystems).
[0147]By carrying out a search of nucleotide sequence databases such as GenBank, EMBL or DDBJ using a homology search program such as BLAST based on the determined nucleotide sequence of the cDNA, it can be confirmed that the obtained DNA is a gene encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain among the genes in the nucleotide sequence database.
[0148]Examples of the nucleotide sequences of the genes encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose obtained by the above methods include the nucleotide sequences represented by SEQ ID NO:1 or 3.
[0149]Examples of the nucleotide sequences of the genes encoding the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain obtained by the above methods include the nucleotide sequences represented by SEQ ID NO:5 or 6.
[0150]The cDNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can also be obtained by chemical synthesis with a DNA synthesizer such as DNA Synthesizer Model 392 (manufactured by Perkin Elmer) utilizing the phosphoramidite method based on the determined nucleotide sequence of the DNA.
[0151]Preparation of a genomic DNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be carried out, for example, by the following method.
Method for Preparing Genomic DNA
[0152]The genomic DNA can be prepared by known methods described in Molecular Cloning, Second Edition; Current Protocols in Molecular Biology; etc. In addition, the genomic DNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be obtained by using a kit such as Genomic DNA Library Screening System® (manufactured by Genome Systems) or Universal GenomeWalker® Kits (manufactured by CLONTECH).
[0153]The nucleotide sequence of the obtained DNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be determined by generally employed sequencing methods such as the dideoxy method of Sanger, et al. [Proc. Natl. Acad. Sci. U.S.A., 74, 5463 (1977)] or by use of nucleotide sequencers such as ABI PRISM 377 DNA Sequencer (manufactured by Applied Biosystems).
[0154]By carrying out a search of nucleotide sequence databases such as GenBank, EMBL or DDBJ using a homology search program such as BLAST based on the determined nucleotide sequence of the genomic DNA, it can be confirmed that the obtained DNA is a gene encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain among the genes in the nucleotide sequence database.
[0155]The genomic DNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can also be obtained by chemical synthesis with a DNA synthesizer such as DNA Synthesizer Model 392 (manufactured by Perkin Elmer) utilizing the phosphoamidite method based on the determined nucleotide sequence of the DNA.
[0156]Examples of the nucleotide sequences of the genomic DNAs encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose obtained by the above methods include the nucleotide sequences represented by SEQ ID NOs:9, 10, 11 and 12.
[0157]An example of the nucleotide sequence of the genomic DNA encoding the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain obtained by the above methods is the nucleotide sequence represented by SEQ ID NO:13.
[0158]The host cell used for the production of the antibody composition of the present invention can also be obtained without using an expression vector by directly introducing into a host cell an antisense oligonucleotide or ribozyme designed based on the nucleotide sequence encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain.
[0159]The antisense oligonucleotide or ribozyme can be prepared by known methods or by using a DNA synthesizer. Specifically, based on the sequence information on an oligonucleotide having a sequence corresponding to 5 to 150, preferably 5 to 60, more preferably 10 to 40 contiguous nucleotides in the nucleotide sequence of the cDNA or genomic DNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain, an oligonucleotide corresponding to the sequence complementary to the above oligonucleotide (antisense oligonucleotide) or a ribozyme comprising the oligonucleotide sequence can be synthesized.
[0160]The oligonucleotide includes oligo RNA and derivatives of the oligonucleotide (hereinafter referred to as oligonucleotide derivatives).
[0161]The oligonucleotide derivatives include an oligonucleotide derivative wherein the phosphodiester bond in the oligonucleotide is converted to a phosophorothioate bond, an oligonucleotide derivative wherein the phosphodiester bond in the oligonucleotide is converted to an N3'-P5' phosphoamidate bond, an oligonucleotide derivative wherein the ribose-phosphodiester bond in the oligonucleotide is converted to a peptide-nucleic acid bond, an oligonucleotide derivative wherein the uracil in the oligonucleotide is substituted with C-5 propynyluracil, an oligonucleotide derivative wherein the uracil in the oligonucleotide is substituted with C-5 thiazolyluracil, an oligonucleotide derivative wherein the cytosine in the oligonucleotide is substituted with C-5 propynylcytosine, an oligonucleotide derivative wherein the cytosine in the oligonucleotide is substituted with phenoxazine-modified cytosine, an oligonucleotide derivative wherein the ribose in the oligonucleotide is substituted with 2'-O-propylribose, and an oligonucleotide derivative wherein the ribose in the oligonucleotide is substituted with 2'-methoxyethoxyribose [Cell Technology, 16, 1463 (1997)].
(b) Preparation of the Host Cell for the Production of the Antibody Composition of the Present Invention by the Homologous Recombination Method
[0162]The host cell used for the production of the antibody composition of the present invention can be prepared by modifying a target gene on the chromosome by the homologous recombination method targeting a gene encoding an enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain.
[0163]Modification of the target gene on the chromosome can be carried out by using the methods described in Manipulating the Mouse Embryo, A Laboratory Manual, Second Edition, Cold Spring Harbor Laboratory Press (1994) (hereinafter referred to as Manipulating the Mouse Embryo, A Laboratory Manual); Gene Targeting, A Practical Approach, IRL Press at Oxford University Press (1993); Biomanual Series 8, Gene Targeting, Preparation of Mutant Mice Using ES Cells, Yodosha (1995) (hereinafter referred to as Preparation of Mutant Mice Using ES Cells); etc., for example, in the following manner.
[0164]A genomic DNA encoding an enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain is prepared.
[0165]Based on the nucleotide sequence of the genomic DNA, a target vector is prepared for homologous recombination of a target gene to be modified (e.g., the structural gene or promoter gene for the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain).
[0166]The host cell used for the preparation of the cell of the present invention can be prepared by introducing the prepared target vector into a host cell and selecting a cell in which homologous recombination occurred between the target gene on the chromosome and the target vector.
[0167]As the host cell, any yeast cell, animal cell, insect cell, plant cell, or the like can be used so long as it has a gene encoding the target enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain. Examples of the host cells include those described in 2 below.
[0168]The genomic DNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be prepared by the methods for preparing a genomic DNA described in the above 1 (1) (a), etc.
[0169]Examples of the nucleotide sequences of the genomic DNAs encoding the enzyme relating to the synthesis of the intracellular sugar nucleotide GDP-fucose obtained by the above methods include the nucleotide sequences represented by SEQ ID NOs:9, 10, 11 and 12.
[0170]An example of the nucleotide sequence of the genomic DNA encoding the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain obtained by the above methods is the nucleotide sequence represented by SEQ ID NO:13.
[0171]The target vector for use in the homologous recombination of the target gene on the chromosome can be prepared according to the methods described in Gene Targeting, A Practical Approach, IRL Press at Oxford University Press (1993); Preparation of Mutant Mice Using ES Cells; etc. The target vector may be either a replacement-type one or an insertion-type one.
[0172]Introduction of the target vector into various host cells can be carried out by the methods suitable for introducing a recombinant vector into various host cells described in 3 below.
[0173]The methods for efficiently selecting a homologous recombinant include positive selection, promoter selection, negative selection and polyA selection described in Gene Targeting, A Practical Approach, IRL Press at Oxford University Press (1993); Preparation of Mutant Mice Using ES Cells; etc. The methods for selecting the desired homologous recombinant from the selected cell lines include Southern hybridization (Molecular Cloning, Second Edition) and PCR [PCR Protocols, Academic Press (1990)] with the genomic DNA.
(c) Preparation of the Host Cell for the Production of the Antibody Composition of the Present Invention by the RDO Method
[0174]The host cell used for the production of the antibody composition of the present invention can be prepared by the RDO method targeting a gene encoding an enzyme relating to the synthesis of the intracellular sugar nucleotide GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain, for example, in the following manner.
[0175]A cDNA or a genomic DNA encoding an enzyme relating to the synthesis of the intracellular sugar nucleotide GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain is prepared by the methods described in the above 1 (1) (a).
[0176]The nucleotide sequence of the prepared cDNA or genomic DNA is determined.
[0177]Based on the determined DNA sequence, an RDO construct of appropriate length which comprises a DNA moiety encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain, non-translated regions and introns is designed and synthesized.
[0178]The host cell of the present invention can be obtained by introducing the synthesized RDO into a host cell and then selecting a transformant in which a mutation occurred in the target enzyme, that is, the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain.
[0179]As the host cell, any yeast cell, animal cell, insect cell, plant cell, or the like can be used so long as it has a gene encoding the target enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain. Examples of the host cells include those described in 2 below.
[0180]Introduction of the RDO into various host cells can be carried out by the methods suitable for introducing a recombinant vector into various host cells described in 2 below.
[0181]The cDNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be prepared by the methods for preparing a cDNA described in the above 1 (1) (a) or the like.
[0182]The genomic DNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be prepared by the methods for preparing a genomic DNA described in the above 1 (1) (b) or the like.
[0183]After DNA is cleaved with appropriate restriction enzymes, the nucleotide sequence of the DNA can be determined by cloning the DNA fragments into a plasmid such as pBluescript SK(-)® (manufactured by Stratagene), subjecting the clones to the reaction generally used as a method for analyzing a nucleotide sequence such as the dideoxy method of Sanger et al. [Proc. Natl. Acad. Sci. USA, 74, 5463 (1977)] or the like, and then analyzing the clones by using an automatic nucleotide sequence analyzer such as ABI PRISM 377 DNA Sequencer (manufactured by Applied Biosystems) or the like.
[0184]The RDO can be prepared by conventional methods or by using a DNA synthesizer.
[0185]The methods for selecting a cell in which a mutation occurred by introducing the RDO into the host cell, in the gene encoding the target enzyme, that is, the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain include the methods for directly detecting mutations in chromosomal genes described in Molecular Cloning, Second Edition; Current Protocols in Molecular Biology; etc.
[0186]For the selection of the transformant, the following methods can also be employed: the method using, as a marker, the activity of the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain described in the above 1 (1) (a); the method using, as a marker, the sugar chain structure of a glycoprotein on the cell membrane described in 1 (5) below; and the method using, as a marker, the sugar chain structure of a produced antibody molecule described in 4 and 5 below.
[0187]The RDO can be designed according to the descriptions in Science, 273, 1386 (1996); Nature Medicine, 4, 285 (1998); Hepatology, 25, 1462 (1997); Gene Therapy, 5, 1960 (1999); Gene Therapy, 5, 1960 (1999); J. Mol. Med., 75, 829 (1997); Proc. Natl. Acad. Sci. USA, 96, 8774 (1999); Proc. Natl. Acad. Sci. USA, 96, 8768 (1999); Nuc. Acids Res., 27, 1323 (1999); Invest. Dermatol., 111, 1172 (1998); Nature Biotech., 16, 1343 (1998); Nature Biotech., 18, 43 (2000); Nature Biotech., 18, 555 (2000); etc.
(d) Preparation of the Host Cell for the Production of the Antibody Composition of the Present Invention by the RNAi Method
[0188]The host cell used for the production of the antibody composition of the present invention can be prepared by the RNAi method targeting a gene encoding an enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain, for example, in the following manner.
[0189]A cDNA encoding an enzyme relating to the synthesis of the intracellular sugar nucleotide GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain is prepared by the methods described in the above 1 (1) (a).
[0190]The nucleotide sequence of the prepared cDNA is determined.
[0191]Based on the determined cDNA sequence, an RNAi gene of appropriate length is designed which comprises a moiety encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain, or non-translated regions.
[0192]In order to express the RNAi gene in a cell, a recombinant vector is prepared by inserting a fragment or full-length of the prepared cDNA into a site downstream of a promoter in an appropriate expression vector.
[0193]The recombinant vector is introduced into a host cell suited for the expression vector to obtain a transformant.
[0194]The host cell used for the preparation of the cell of the present invention can be obtained by selecting a transformant using, as a marker, the activity of the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain, or the sugar chain structure of a produced antibody molecule or a glycoprotein on the cell membrane.
[0195]As the host cell, any yeast cell, animal cell, insect cell, plant cell, or the like can be used so long as it has a gene encoding the target enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain. Examples of the host cells include those described in 2 below.
[0196]The expression vectors that can be employed are those capable of autonomous replication or integration into the chromosome in the above host cells and comprising a promoter at a position appropriate for the transcription of the designed RNAi gene. Examples of the expression vectors include those described in 2 below.
[0197]Introduction of a gene into various host cells can be carried out by the methods suitable for introducing a recombinant vector into various host cells described in 2 below.
[0198]The methods for selecting the transformant using, as a marker, the activity of the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain include the methods described in the above 1 (1) (a).
[0199]The methods for selecting the transformant using, as a marker, the sugar chain structure of a glycoprotein on the cell membrane include the method described in 1 (5). The methods for selecting the transformant using, as a marker, the sugar chain structure of a produced antibody molecule include the methods described in 4 or 5 below.
[0200]The cDNA encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be prepared by the methods for preparing a cDNA described in the above 1 (1) (a), etc.
[0201]The host cell used for the preparation of the cell of the present invention can also be obtained without using an expression vector by directly introducing into a host cell the RNAi gene designed based on the nucleotide sequence encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain.
[0202]The RNAi gene can be prepared by known methods or by using a DNA synthesizer.
[0203]The RNAi gene construct can be designed according to the descriptions in Nature, 391, 806 (1998); Proc. Natl. Acad. Sci. USA, 95, 15502 (1998); Nature, 395, 854 (1998); Proc. Natl. Acad. Sci. USA, 96, 5049 (1999); Cell, 95, 1017 (1998); Proc. Natl. Acad. Sci. USA, 96, 1451 (1999); Proc. Natl. Acad. Sci. USA, 95, 13959 (1998); Nature Cell Biol., 2, 70 (2000); etc.
(e) Preparation of the Host Cell for the Production of the Antibody Composition of the Present Invention by the Method Using a Transposon
[0204]The host cell used for the production of the antibody composition of the present invention can be prepared by using the transposon system described in Nature Genet., 25, 35 (2000), etc., and then selecting a mutant using, as a marker, the activity of the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain, or the sugar chain structure of a produced antibody molecule or a glycoprotein on the cell membrane.
[0205]The transposon system is a system for inducing a mutation by random insertion of an exogenous gene into the chromosome, wherein usually an exogenous gene inserted into a transposon is used as a vector for inducing a mutation and a transposase expression vector for randomly inserting the gene into the chromosome is introduced into the cell at the same time.
[0206]Any transposase can be used so long as it is suitable for the sequence of the transposon to be used.
[0207]As the exogenous gene, any gene can be used so long as it can induce a mutation in the DNA of a host cell.
[0208]As the host cell, any yeast cell, animal cell, insect cell, plant cell, or the like can be used so long as it has a gene encoding the target enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain. Examples of the host cells include those described in 2 below. Introduction of the gene into various host cells can be carried out by the methods suitable for introducing a recombinant vector into various host cells described in 2 below.
[0209]The methods for selecting the mutant using, as a marker, the activity of the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain include the methods described in the above 1 (1) (a).
[0210]The methods for selecting the mutant using, as a marker, the sugar chain structure of a glycoprotein on the cell membrane include the method described in 1 (5). The methods for selecting the mutant using, as a marker, the sugar chain structure of a produced antibody molecule include the methods described in 4 or 5 below.
(2) Technique of Introducing a Dominant-Negative Mutant of a Gene Encoding an Enzyme
[0211]The host cell used for the production of the antibody composition of the present invention can be prepared by using the method of introducing a dominant-negative mutant of a target gene, i.e., a gene encoding an enzyme relating to the synthesis of the intracellular sugar nucleotide GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain. Examples of the enzymes relating to the synthesis of the intracellular sugar nucleotide GDP-fucose include GMD and Fx. Examples of the enzymes relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain include α1,6-fucosyltransferase and α-L-fucosidase.
[0212]These enzymes have substrate specificity and catalyze specific reactions. By disrupting the active center of such enzymes having substrate specificity and catalytic action, their dominant-negative mutants can be prepared. Preparation of a dominant-negative mutant is described in detail below, using for an example GMD among the target enzymes.
[0213]As a result of the analysis of the tertiary structure of GMD derived from Escherichia coli, it has been revealed that four amino acids (threonine at position 133, glutamic acid at position 135, tyrosine at position 157 and lysine at position 161) have an important function for the enzyme activity (Structure, 8, 2, 2000). That is, the mutants prepared by substituting the above four amino acids by other amino acids based on the tertiary structure information all showed significantly decreased enzyme activity. On the other hand, little change was observed in the ability of the mutants to bind to the GMD coenzyme NADP or the substrate GDP-mannose. Accordingly, a dominant-negative mutant can be prepared by substituting the four amino acids which are responsible for the enzyme activity of GMD. On the basis of the result of preparation of a dominant-negative mutant of GMD derived from Escherichia coli, dominant-negative mutants of other GMDs can be prepared by performing homology comparison and tertiary structure prediction using the amino acid sequence information. For example, in the case of GMD derived from CHO cell (SEQ ID NO:2), a dominant-negative mutant can be prepared by substituting threonine at position 155, glutamic acid at position 157, tyrosine at position 179 and lysine at position 183 by other amino acids. Preparation of such a gene carrying introduced amino acid substitutions can be carried out by site-directed mutagenesis described in Molecular Cloning, Second Edition; Current Protocols in Molecular Biology; etc.
[0214]The host cell used for the production of the antibody composition of the present invention can be prepared according to the method of gene introduction described in Molecular Cloning, Second Edition; Current Protocols in Molecular Biology; Manipulating the Mouse Embryo, Second Edition; etc. using a gene encoding a dominant-negative mutant of a target enzyme (hereinafter abbreviated as dominant-negative mutant gene) prepared as above, for example, in the following manner.
[0215]A dominant-negative mutant gene encoding the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain is prepared.
[0216]Based on the full-length DNA of the prepared dominant-negative mutant gene, a DNA fragment of appropriate length containing a region encoding the protein is prepared according to need.
[0217]A recombinant vector is prepared by inserting the DNA fragment or full-length DNA into a site downstream of a promoter in an appropriate expression vector.
[0218]The recombinant vector is introduced into a host cell suited for the expression vector to obtain a transformant.
[0219]The host cell used for the preparation of the cell of the present invention can be obtained by selecting a transformant using, as a marker, the activity of the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain, or the sugar chain structure of a produced antibody molecule or a glycoprotein on the cell membrane.
[0220]As the host cell, any yeast cell, animal cell, insect cell, plant cell, or the like can be used so long as it has a gene encoding the target enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain. Examples of the host cells include those described in 2 below.
[0221]The expression vectors that can be employed are those capable of autonomous replication or integration into the chromosome in the above host cells and comprising a promoter at a position appropriate for the transcription of the DNA encoding the desired dominant-negative mutant. Examples of the expression vectors include those described in 2 below.
[0222]Introduction of a gene into various host cells can be carried out by the methods suitable for introducing a recombinant vector into various host cells described in 2 below.
[0223]The methods for selecting the transformant using, as a marker, the activity of the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain include the methods described in the above 1 (1) (a).
[0224]The methods for selecting the transformant using, as a marker, the sugar chain structure of a glycoprotein on the cell membrane include the method described in 1 (5) below. The methods for selecting the transformant using, as a marker, the sugar chain structure of a produced antibody molecule include the methods described in 4 or 5 below.
(3) Technique of Introducing a Mutation into an Enzyme
[0225]The host cell used for the production of the antibody composition of the present invention can be prepared by introducing a mutation into a gene encoding an enzyme relating to the synthesis of the intracellular sugar nucleotide GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain, and then selecting a desired cell line in which the mutation occurred in the enzyme.
[0226]Examples of the enzymes relating to the synthesis of the intracellular sugar nucleotide GDP-fucose include GMD and Fx. Examples of the enzymes relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain include α1,6-fucosyltransferase and α-L-fucosidase.
[0227]The methods for introducing a mutation into the enzyme include: 1) a method in which a desired cell line is selected from mutants obtained by subjecting a parent cell line to mutagenesis or by spontaneous mutation using, as a marker, the activity of the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain; 2) a method in which a desired cell line is selected from mutants obtained by subjecting a parent cell line to mutagenesis or by spontaneous mutation using, as a marker, the sugar chain structure of a produced antibody molecule; and 3) a method in which a desired cell line is selected from mutants obtained by subjecting a parent cell line to mutagenesis or by spontaneous mutation using, as a marker, the sugar chain structure of a glycoprotein on the cell membrane.
[0228]Mutagenesis may be carried out by any method capable of inducing a point mutation, a deletion mutation or a frameshift mutation in DNA of a cell of a parent cell line.
[0229]Suitable methods include treatment with ethyl nitrosourea, nitrosoguanidine, benzopyrene or an acridine dye and radiation treatment. Various alkylating agents and carcinogens are also useful as mutagens. A mutagen is allowed to act on a cell by the methods described in Soshiki Baiyo no Gijutsu (Tissue Culture Techniques), Third Edition (Asakura Shoten), edited by The Japanese Tissue Culture Association (1996); Nature Genet., 24, 314 (2000); etc.
[0230]Examples of the mutants generated by spontaneous mutation include spontaneous mutants obtained by continuing subculture under usual cell culture conditions without any particular treatment for mutagenesis.
[0231]The methods for measuring the activity of the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain include the methods described in the above 1 (1) (a). The methods for determining the sugar chain structure of a produced antibody molecule include the methods described in 4 or 5 below. The methods for determining the sugar chain structure of a glycoprotein on the cell membrane include the method described in 1 (5).
(4) Technique of Suppressing Transcription or Translation of a Gene Encoding an Enzyme
[0232]The host cell used for the production of the antibody composition of the present invention can be prepared by inhibiting transcription or translation of a target gene, i.e., a gene encoding an enzyme relating to the synthesis of the intracellular sugar nucleotide GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain using the antisense RNA/DNA technique [Bioscience and Industry, 50, 322 (1992); Chemistry, 46, 681 (1991); Biotechnology, 9, 358 (1992); Trends in Biotechnology, 10, 87 (1992); Trends in Biotechnology, 10, 152 (1992); Cell Technology, 16, 1463 (1997)], the triple helix technique [Trends in Biotechnology, 10, 132 (1992)], etc.
[0233]Examples of the enzymes relating to the synthesis of the intracellular sugar nucleotide GDP-fucose include GMD and Fx. Examples of the enzymes relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain include α1,6-fucosyltransferase and α-L-fucosidase.
[0234]The methods for measuring the activity of the enzyme relating to the synthesis of an intracellular sugar nucleotide, GDP-fucose or the enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain include the methods described in the above 1 (1) (a).
[0235]The methods for determining the sugar chain structure of a glycoprotein on the cell membrane include the method described in 1 (5). The methods for determining the sugar chain structure of a produced antibody molecule include the methods described in 4 or 5 below.
(5) Technique of Selecting a Cell Line Resistant to a Lectin which Recognizes a Sugar Chain Structure in which 1-Position of Fucose is Bound to 6-Position of N-Acetylglucosamine in The Reducing End Through α-Bond in a Complex Type N-Glycoside-Linked Sugar Chain
[0236]The host cell used for the production of the antibody composition of the present invention can be prepared by selecting a cell line resistant to a lectin which recognizes a sugar chain structure in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain.
[0237]Selection of a cell line resistant to a lectin which recognizes a sugar chain structure in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be carried out, for example, by the method using a lectin described in Somatic Cell Mol. Genet., 12, 51 (1986), etc.
[0238]As the lectin, any lectin can be used so long as it recognizes a sugar chain structure in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain. Specific examples include lentil lectin LCA (lentil agglutinin derived from Lens culinaris), pea lectin PSA (pea lectin derived from Pisum sativum), broad bean lectin VFA (agglutinin derived from Vicia faba) and Aleuria aurantia lectin AAL (lectin derived from Aleuria aurantia).
[0239]Specifically, the cell line of the present invention resistant to a lectin which recognizes a sugar chain structure in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain can be selected by culturing cells in a medium containing the above lectin at a concentration of 1 μg/ml to 1 mg/ml for one day to 2 weeks, preferably one day to one week, subculturing surviving cells or picking up a colony and transferring it into a culture vessel, and subsequently continuing the culturing using the medium containing the lectin.
2. Process for Producing the Antibody Composition
[0240]The antibody composition of the present invention can be obtained by expressing it in a host cell using the methods described in Molecular Cloning, Second Edition; Current Protocols in Molecular Biology; Antibodies, A Laboratory Manual, Cold Spring Harbor Laboratory, 1988 (hereinafter referred to as Antibodies); Monoclonal Antibodies: Principles and Practice, Third Edition, Acad. Press, 1993 (hereinafter referred to as Monoclonal Antibodies); Antibody Engineering, A Practical Approach, IRL Press at Oxford University Press, 1996 (hereinafter referred to as Antibody Engineering); etc., for example, in the following manner.
[0241]A full-length cDNA encoding an anti-human CCR4 antibody molecule is prepared, and a DNA fragment of appropriate length comprising a region encoding the antibody molecule is prepared.
[0242]A recombinant vector is prepared by inserting the DNA fragment or full-length DNA into a site downstream of a promoter in an appropriate expression vector.
[0243]The recombinant vector is introduced into a host cell suited for the expression vector to obtain a transformant producing the antibody molecule.
[0244]As the host cell, any yeast cells, animal cells, insect cells, plant cells, etc. that are capable of expressing the desired gene can be used.
[0245]Also useful are cells obtained by selecting cells in which the activity of an enzyme relating to the modification of an N-glycoside-linked sugar chain bound to the Fc region of an antibody molecule, i.e., an enzyme relating to the synthesis of an intracellular sugar nucleotide GDP-fucose or an enzyme relating to the modification of a sugar chain in which 1-position of fucose is bound to 6-position of N-acetylglucosamine in the reducing end through α-bond in a complex type N-glycoside-linked sugar chain is deleted, and cells obtained by various artificial techniques described in the above 1.
[0246]The expression vectors that can be employed are those capable of autonomous replication or integration into the chromosome in the above host cells and comprising a promoter at a position appropriate for the transcription of the DNA encoding the desired antibody molecule.
[0247]The cDNA can be prepared from a human or non-human animal tissue or cell according to the methods for preparing a cDNA described in the above 1 (1) (a) using, e.g., a probe or primers specific for the desired antibody molecule.
[0248]When yeast is used as the host cell, YEP13 (ATCC 37115), YEp24 (ATCC 37051), YCp50 (ATCC 37419), etc. can be used as the expression vector.
[0249]As the promoter, any promoters capable of expressing in yeast strains can be used. Suitable promoters include promoters of genes of the glycolytic pathway such as hexokinase, PHO5 promoter, PGK promoter, GAP promoter, ADH promoter, gal 1 promoter, gal 10 promoter, heat shock protein promoter, MFα1 promoter and CUP 1 promoter.
[0250]Examples of suitable host cells are microorganisms belonging to the genera Saccharomyces, Schizosaccharomyces, Kluyveromyces, Trichosporon and Schwanniomyces, and specifically, Saccharomyces cerevisiae, Schizosaccharomyces pombe, Kluyveromyces lactis, Trichosporon pullulans and Schwanniomyces alluvius.
[0251]Introduction of the recombinant vector can be carried out by any of the methods for introducing DNA into yeast, for example, electroporation [Methods Enzymol., 194, 182 (1990)], the spheroplast method [Proc. Natl. Acad. Sci. USA, 84, 1929 (1978)], the lithium acetate method [J. Bacteriology, 153, 163 (1983)] and the method described in Proc. Natl. Acad. Sci. USA, 75, 1929 (1978).
[0252]When an animal cell is used as the host cell, pcDNAI, pcDM8 (commercially available from Funakoshi Co., Ltd.), pAGE107 [Japanese Published Unexamined Patent Application No. 22979/91; Cytotechnology, 3, 133 (1990)], pAS3-3 (Japanese Published Unexamined Patent Application No. 227075/90), pCDM8 [Nature, 329, 840 (1987)], pcDNAI/Amp (manufactured by Invitrogen Corp.), pREP4 (manufactured by Invitrogen Corp.), pAGE103 [J. Biochemistry, 101, 1307 (1987)], pAGE210, etc. can be used as the expression vector.
[0253]As the promoter, any promoters capable of expressing in animal cells can be used. Suitable promoters include the promoter of IE (immediate early) gene of cytomegalovirus (CMV), SV40 early promoter, the promoter of a retrovirus, metallothionein promoter, heat shock promoter, SRα promoter, etc. The enhancer of IE gene of human CMV may be used in combination with the promoter.
[0254]Examples of suitable host cells are human-derived Namalwa cells, monkey-derived COS cells, Chinese hamster-derived CHO cells, HBT5637 (Japanese Published Unexamined Patent Application No. 299/88), rat myeloma cells, mouse myeloma cells, cells derived from Syrian hamster kidney, embryonic stem cells and fertilized egg cells.
[0255]Introduction of the recombinant vector can be carried out by any of the methods for introducing DNA into animal cells, for example, electroporation [Cytotechnology, 3, 133 (1990)], the calcium phosphate method (Japanese Published Unexamined Patent Application No. 227075/90), lipofection [Proc. Natl. Acad. Sci. USA, 84, 7413 (1987)], the injection method (Manipulating the Mouse Embryo, A Laboratory Manual), the method using particle gun (gene gun) (Japanese Patent Nos. 2606856 and 2517813), the DEAE-dextran method [Biomanual Series 4--Methods of Gene Transfer, Expression and Analysis (Yodosha), edited by Takashi Yokota and Kenichi Arai (1994)] and the virus vector method (Manipulating the Mouse Embryo, Second Edition).
[0256]When an insect cell is used as the host cell, the protein can be expressed by the methods described in Current Protocols in Molecular Biology; Baculovirus Expression Vectors, A Laboratory Manual, W.H. Freeman and Company, New York (1992); Bio/Technology, 6, 47 (1988), etc.
[0257]That is, the recombinant vector and a baculovirus are cotransfected into insect cells to obtain a recombinant virus in the culture supernatant of the insect cells, and then insect cells are infected with the recombinant virus, whereby the protein can be expressed.
[0258]The gene transfer vectors useful in this method include pVL1392, pVL1393 and pBlueBacIII® (products of Invitrogen Corp.).
[0259]An example of the baculovirus is Autographa californica nuclear polyhedrosis virus, which is a virus infecting insects belonging to the family Barathra.
[0260]Examples of the insect cells are Spodoptera frugiperda ovarian cells Sf9 and Sf21 [Current Protocols in Molecular Biology; Baculovirus Expression Vectors, A Laboratory Manual, W.H. Freeman and Company, New York (1992)] and Trichoplusia ni ovarian cell High 5 (manufactured by Invitrogen Corp.).
[0261]Cotransfection of the above recombinant vector and the above baculovirus into insect cells for the preparation of the recombinant virus can be carried out by the calcium phosphate method (Japanese Published Unexamined Patent Application No. 227075/90), lipofection [Proc. Natl. Acad. Sci. USA, 84, 7413 (1987)], etc.
[0262]When a plant cell is used as the host cell, Ti plasmid, tobacco mosaic virus vector, etc. can be used as the expression vector.
[0263]As the promoter, any promoters capable of expressing in plant cells can be used. Suitable promoters include 35S promoter of cauliflower mosaic virus (CaMV), rice actin 1 promoter, etc.
[0264]Examples of suitable host cells are cells of plants such as tobacco, potato, tomato, carrot, soybean, rape, alfalfa, rice, wheat and barley.
[0265]Introduction of the recombinant vector can be carried out by any of the methods for introducing DNA into plant cells, for example, the method using Agrobacterium (Japanese Published Unexamined Patent Application Nos. 140885/84 and 70080/85, WO94/00977), electroporation (Japanese Published Unexamined Patent Application No. 251887/85) and the method using particle gun (gene gun) (Japanese Patent Nos. 2606856 and 2517813).
[0266]Expression of the antibody gene can be carried out not only by direct expression but also by secretory production, expression of a fusion protein of the Fc region and another protein, etc. according to the methods described in Molecular Cloning, Second Edition, etc.
[0267]When the gene is expressed in yeast, an animal cell, an insect cell or a plant cell carrying an introduced gene relating to the synthesis of a sugar chain, an antibody molecule to which a sugar or a sugar chain is added by the introduced gene can be obtained.
[0268]The antibody composition can be produced by culturing the transformant obtained as above in a medium, allowing the antibody molecules to form and accumulate in the culture, and recovering them from the culture. Culturing of the transformant in a medium can be carried out by conventional methods for culturing the host cell.
[0269]For the culturing of the transformant obtained by using a eucaryote such as yeast as the host, any of natural media and synthetic media can be used insofar as it is a medium suitable for efficient culturing of the transformant which contains carbon sources, nitrogen sources, inorganic salts, etc. which can be assimilated by the host used.
[0270]As the carbon sources, any carbon sources that can be assimilated by the host can be used. Examples of suitable carbon sources include carbohydrates such as glucose, fructose, sucrose, molasses containing them, starch and starch hydrolyzate; organic acids such as acetic acid and propionic acid; and alcohols such as ethanol and propanol.
[0271]As the nitrogen sources, ammonia, ammonium salts of organic or inorganic acids such as ammonium chloride, ammonium sulfate, ammonium acetate and ammonium phosphate, and other nitrogen-containing compounds can be used as well as peptone, meat extract, yeast extract, corn steep liquor, casein hydrolyzate, soybean cake, soybean cake hydrolyzate, and various fermented microbial cells and digested products thereof.
[0272]Examples of the inorganic salts include potassium dihydrogenphosphate, dipotassium hydrogenphosphate, magnesium phosphate, magnesium sulfate, sodium chloride, ferrous sulfate, manganese sulfate, copper sulfate and calcium carbonate.
[0273]Culturing is usually carried out under aerobic conditions, for example, by shaking culture or submerged spinner culture under aeration. The culturing temperature is preferably 15 to 40° C., and the culturing period is usually 16 hours to 7 days. The pH is maintained at 3.0 to 9.0 during the culturing. The pH adjustment is carried out by using an organic or inorganic acid, an alkali solution, urea, calcium carbonate, ammonia, etc.
[0274]If necessary, antibiotics such as ampicillin and tetracycline may be added to the medium during the culturing.
[0275]When a microorganism transformed with a recombinant vector comprising an inducible promoter is cultured, an inducer may be added to the medium, if necessary. For example, in the case of a microorganism transformed with a recombinant vector comprising lac promoter, isopropyl-β-D-thiogalactopyranoside or the like may be added to the medium; and in the case of a microorganism transformed with a recombinant vector comprising trp promoter, indoleacrylic acid or the like may be added.
[0276]For the culturing of the transformant obtained by using an animal cell as the host cell, generally employed media such as RPMI1640 medium [The Journal of the American Medical Association, 199, 519 (1967)], Eagle's MEM [Science, 122, 501 (1952)], Dulbecco's modified MEM [Virology, 8, 396 (1959)], 199 medium [Proceeding of the Society for the Biological Medicine, 73, 1 (1950)] and Whitten's medium [Developmental Engineering Experimentation Manual--Preparation of Transgenic Mice (Kodansha), edited by Motoya Katsuki (1987)], media prepared by adding fetal calf serum or the like to these media, etc. can be used as the medium.
[0277]Culturing is usually carried out under conditions of pH 6.0 to 8.0 at 30 to 40° C. for 1 to 7 days in the presence of 5% CO2.
[0278]If necessary, antibiotics such as kanamycin and penicillin may be added to the medium during the culturing.
[0279]For the culturing of the transformant obtained by using an insect cell as the host cell, generally employed media such as TNM-FH Medium® (manufactured by Pharmingen, Inc.), Sf-900 II SFM Medium® (manufactured by Life Technologies, Inc.), ExCell 400® and ExCell 405® (manufactured by JRH Biosciences, Inc.) and Grace's Insect Medium [Nature, 195, 788 (1962)] can be used as the medium.
[0280]Culturing is usually carried out under conditions of pH 6.0 to 7.0 at 25 to 30° C. for 1 to 5 days.
[0281]If necessary, antibiotics such as gentamicin may be added to the medium during the culturing.
[0282]The transformant obtained by using a plant cell as the host cell may be cultured in the form of cells as such or after differentiation into plant cells or plant organs. For the culturing of such transformant, generally employed media such as Murashige-Skoog (MS) medium and White medium, media prepared by adding phytohormones such as auxin and cytokinin to these media, etc. can be used as the medium.
[0283]Culturing is usually carried out under conditions of pH 5.0 to 9.0 at 20 to 40° C. for 3 to 60 days.
[0284]If necessary, antibiotics such as kanamycin and hygromycin may be added to the medium during the culturing.
[0285]As described above, the antibody composition can be produced by culturing, according to a conventional culturing method, the transformant derived from an animal cell or a plant cell and carrying an expression vector into which DNA encoding the antibody molecule has been inserted, allowing the antibody composition to form and accumulate, and recovering the antibody composition from the culture.
[0286]Expression of the antibody gene can be carried out not only by direct expression but also by secretory production, fusion protein expression, etc. according to the methods described in Molecular Cloning, Second Edition.
[0287]The antibody composition may be produced by intracellular production by host cells, extracellular secretion by host cells or production on outer membranes by host cells. A desirable production method can be adopted by changing the kind of the host cells used or the structure of the antibody molecule to be produced.
[0288]When the antibody composition is produced in host cells or on outer membranes of host cells, it is possible to force the antibody composition to be secreted outside the host cells by applying the method of Paulson, et al. [J. Biol. Chem., 264, 17619 (1989)], the method of Lowe, et al. [Proc. Natl. Acad. Sci. USA, 86, 8227 (1989); Genes Develop., 4, 1288 (1990)], or the methods described in Japanese Published Unexamined Patent Application No. 336963/93, WO94/23021, etc.
[0289]That is, it is possible to force the desired antibody molecule to be secreted outside the host cells by inserting DNA encoding the antibody molecule and DNA encoding a signal peptide suitable for the expression of the antibody molecule into an expression vector, introducing the expression vector into the host cells, and then expressing the antibody molecule by use of recombinant DNA techniques.
[0290]It is also possible to increase the production of the antibody composition by utilizing a gene amplification system using a dihydrofolate reductase gene or the like according to the method described in Japanese Published Unexamined Patent Application No. 227075/90.
[0291]Further, the antibody composition can be produced using an animal having an introduced gene (non-human transgenic animal) or a plant having an introduced gene (transgenic plant) constructed by redifferentiation of animal or plant cells carrying the introduced gene.
[0292]When the transformant is an animal or plant, the antibody composition can be produced by raising or culturing the animal or plant in a usual manner, allowing the antibody composition to form and accumulate therein, and recovering the antibody composition from the animal or plant.
[0293]Production of the antibody composition using an animal can be carried out, for example, by producing the desired antibody composition in an animal constructed by introducing the gene according to known methods [American Journal of Clinical Nutrition, 63, 639S (1996); American Journal of Clinical Nutrition, 63, 627S (1996); Bio/Technology, 9, 830 (1991)].
[0294]In the case of an animal, the antibody composition can be produced, for example, by raising a non-human transgenic animal carrying the introduced DNA encoding the antibody molecule, allowing the antibody composition to form and accumulate in the animal, and recovering the antibody composition from the animal. The places where the antibody composition is formed and accumulated include milk (Japanese Published Unexamined Patent Application No. 309192/88), egg, etc. of the animal. As the promoter in this process, any promoters capable of expressing in an animal can be used. Preferred promoters include mammary gland cell-specific promoters such as α casein promoter, β casein promoter, β lactoglobulin promoter and whey acidic protein promoter.
[0295]Production of the antibody composition using a plant can be carried out, for example, by culturing a transgenic plant carrying the introduced DNA encoding the antibody molecule according to known methods [Soshiki Baiyo (Tissue Culture), 20 (1994); Soshiki Baiyo (Tissue Culture), 21 (1995); Trends in Biotechnology, 15, 45 (1997)], allowing the antibody composition to form and accumulate in the plant, and recovering the antibody composition from the plant.
[0296]When the antibody composition produced by the transformant carrying the introduced gene encoding the antibody molecule is expressed in a soluble form in cells, the cells are recovered by centrifugation after the completion of culturing and suspended in an aqueous buffer, followed by disruption using a sonicator, French press, Manton Gaulin homogenizer, Dynomill or the like to obtain a cell-free extract. A purified preparation of the antibody composition can be obtained by centrifuging the cell-free extract to obtain the supernatant and then subjecting the supernatant to ordinary means for isolating and purifying enzymes, e.g., extraction with a solvent, salting-out with ammonium sulfate, etc., desalting, precipitation with an organic solvent, anion exchange chromatography using resins such as diethylaminoethyl (DEAE)-Sepharose and DIAION HPA-75® (manufactured by Mitsubishi Chemical Corporation), cation exchange chromatography using resins such as S-Sepharose FF® (manufactured by Pharmacia), hydrophobic chromatography using resins such as butyl Sepharose and phenyl Sepharose, gel filtration using a molecular sieve, affinity chromatography, chromatofocusing, and electrophoresis such as isoelectric focusing, alone or in combination.
[0297]When the antibody composition is expressed as an inclusion body in cells, the cells are similarly recovered and disrupted, followed by centrifugation to recover the inclusion body of the antibody composition as a precipitate fraction. The recovered inclusion body of the antibody composition is solubilized with a protein-denaturing agent. The solubilized antibody solution is diluted or dialyzed, whereby the antibody composition is renatured to have normal conformation. Then, a purified preparation of the antibody composition can be obtained by the same isolation and purification steps as described above.
[0298]When the antibody composition is extracellularly secreted, the antibody composition or its derivative can be recovered in the culture supernatant. That is, the culture is treated by the same means as above, e.g., centrifugation, to obtain the culture supernatant. A purified preparation of the antibody composition can be obtained from the culture supernatant by using the same isolation and purification methods as described above.
[0299]As an example of the methods for obtaining the antibody composition of the present invention, the method for producing a humanized antibody composition is specifically described below. Other antibody compositions can also be obtained in a similar manner.
(1) Construction of a Vector for Expression of Humanized Antibody
[0300]A vector for expression of humanized antibody is an expression vector for animal cells carrying inserted genes encoding CH and CL of a human antibody, which can be constructed by cloning each of the genes encoding CH and CL of a human antibody into an expression vector for animal cells.
[0301]The C regions of a human antibody may be CH and CL of any human antibody. Examples of the C regions include the C region of IgG1 subclass human antibody H chain (hereinafter referred to as hCγ1) and the C region of κ class human antibody L chain (hereinafter referred to as hCκ).
[0302]As the genes encoding CH and CL of a human antibody, a genomic DNA comprising exons and introns can be used. Also useful is a cDNA prepared by reverse transcription of an mRNA.
[0303]As the expression vector for animal cells, any vector for animal cells can be used so long as it is capable of inserting and expressing the gene encoding the C region of a human antibody. Suitable vectors include pAGE107 [Cytotechnology, 3, 133 (1990)], pAGE103 [J. Biochem., 101, 1307 (1987)], pHSG274 [Gene, 27, 223 (1984)], pKCR [Proc. Natl. Acad. Sci. USA, 78, 1527 (1981)] and pSG1βd2-4 [Cytotechnology, 4, 173 (1990)]. Examples of the promoter and enhancer for use in the expression vector for animal cells include SV40 early promoter and enhancer [J. Biochem., 101, 1307 (1987)], LTR of Moloney mouse leukemia virus [Biochem. Biophys. Res. Commun., 149, 960 (1987)] and immunoglobulin H chain promoter [Cell, 41, 479 (1985)] and enhancer [Cell, 33, 717 (1983)].
[0304]The vector for expression of humanized antibody may be either of the type in which the genes encoding antibody H chain and L chain exist on separate vectors or of the type in which both genes exist on the same vector (hereinafter referred to as tandem-type). The tandem-type ones are preferred in view of the easiness of construction of the vector for expression of humanized antibody, the easiness of introduction into animal cells, the balance between the expression of antibody H chain and that of antibody L chain in animal cells, etc. [J. Immunol. Methods, 167, 271 (1994)]. Examples of the tandem-type humanized antibody expression vectors include pKANTEX93 [Mol. Immunol., 37, 1035 (2000)] and pEE18 [Hybridoma, 17, 559 (1998)].
[0305]The constructed vector for expression of humanized antibody can be used for the expression of a human chimeric antibody and a human CDR-grafted antibody in animal cells.
(2) Obtaining of cDNA Encoding V Region of an Antibody Derived from a Non-Human Animal
[0306]cDNAs encoding VH and VL of an antibody derived from a non-human animal, e.g., a mouse antibody can be obtained in the following manner.
[0307]A cDNA is synthesized using, as a template, an mRNA extracted from a hybridoma cell producing a non-human animal-derived antibody which specifically binds to human CCR4. The synthesized cDNA is inserted into a vector such as a phage or a plasmid to prepare a cDNA library. A recombinant phage or recombinant plasmid carrying a cDNA encoding VH and a recombinant phage or recombinant plasmid carrying a cDNA encoding VL are isolated from the cDNA library using DNA encoding the C region or V region of a known mouse antibody as a probe. The entire nucleotide sequences of VH and VL of the desired mouse antibody on the recombinant phages or recombinant plasmids are determined, and the whole amino acid sequences of VH and VL are deduced from the nucleotide sequences.
[0308]Hybridoma cells producing a non-human animal-derived antibody which specifically binds to human CCR4 can be obtained by immunizing a non-human animal with human CCR4 represented by SEQ ID NO:43, preparing hybridomas from antibody-producing cells of the immunized animal and myeloma cells according to a known method (Antibodies: A Laboratory Manual, Cold Spring Harbor Laboratory, Chapter 14, 1998), selecting cloned hybridomas, culturing the selected hybridomas and purifying cells from the culture supernatant.
[0309]As the non-human animal, any animal can be used so long as hybridoma cells can be prepared from the animal. Suitable animals include mouse, rat, hamster and rabbit.
[0310]The methods for preparing total RNA from a hybridoma cell include the guanidine thiocyanate-cesium trifluoroacetate method [Methods in Enzymol., 154, 3 (1987)], and the methods for preparing mRNA from the total RNA include the oligo (dT) immobilized cellulose column method (Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Lab. Press New York, 1989). Examples of the kits for preparing mRNA from a hybridoma cell include Fast Track mRNA Isolation Kit® (manufactured by Invitrogen) and Quick Prep mRNA Purification Kit® (manufactured by Pharmacia).
[0311]The methods for synthesizing the cDNA and preparing the cDNA library include conventional methods (Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Lab. Press New York, 1989; Current Protocols in Molecular Biology, Supplement 1-34), or methods using commercially available kits such as SuperScript® Plasmid System for cDNA Synthesis and Plasmid Cloning® (manufactured by GIBCO BRL) and ZAP-cDNA Synthesis Kit® (manufactured by STRATAGENE).
[0312]In preparing the cDNA library, the vector for inserting the cDNA synthesized using the mRNA extracted from a hybridoma cell as a template may be any vector so long as the cDNA can be inserted. Examples of suitable vectors include ZAP Express® [Strategies, 5, 58 (1992)], pBluescript II SK(+)® [Nucleic Acids Research, 17, 9494 (1989)], λZAP II® (manufactured by STRATAGENE), λgt10, λgt11 [DNA Cloning: A Practical Approach, I, 49 (1985)], Lambda BlueMid® (manufactured by Clontech), λExCell, pT7T3 18U (manufactured by Pharmacia), pcD2 [Mol. Cell. Biol., 3, 280 (1983)] and pUC18 [Gene, 33, 103 (1985)].
[0313]As Escherichia coli for introducing the cDNA library constructed with a phage or plasmid vector, any Escherichia coli can be used so long as the cDNA library can be introduced, expressed and maintained. Examples of suitable Escherichia coli include XL1-Blue MRF' [Strategies, 5, 81 (1992)], C600 [Genetics, 39, 440 (1954)], Y1088, Y1090 [Science, 222, 778 (1983)], NM522 [J. Mol. Biol., 166, 1 (1983)], K802 [J. Mol. Biol., 16, 118 (1966)] and JM105 [Gene, 38, 275 (1985)].
[0314]The methods for selecting the cDNA clones encoding VH and VL of a non-human animal-derived antibody from the cDNA library include colony hybridization or plaque hybridization (Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Lab. Press New York, 1989) using an isotope- or fluorescence-labeled probe. It is also possible to prepare the cDNAs encoding VH and VL by preparing primers and performing PCR (Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Lab. Press New York, 1989; Current Protocols in Molecular Biology, Supplement 1-34) using the cDNA or cDNA library as a template.
[0315]The nucleotide sequences of the cDNAs selected by the above methods can be determined by cleaving the cDNAs with appropriate restriction enzymes, cloning the fragments into a plasmid such as pBluescript SK(-)® (manufactured by STRATAGENE), and then analyzing the sequences by generally employed sequencing methods such as the dideoxy method of Sanger, et al. [Proc. Natl. Acad. Sci. USA, 74, 5463 (1977)] or by use of nucleotide sequencers such as ABI PRISM 377 DNA Sequencer (manufactured by Applied Biosystems).
[0316]The whole amino acid sequences of VH and VL are deduced from the determined nucleotide sequences and compared with the entire amino acid sequences of VH and VL of a known antibody (Sequences of Proteins of Immunological Interest, US Dept. Health and Human Services, 1991), whereby it can be confirmed that the obtained cDNAs encode amino acid sequences which completely comprise VH and VL of the antibody including secretory signal sequences.
[0317]Further, when the amino acid sequence of an antibody variable region or the nucleotide sequence of DNA encoding the variable region is already known, the DNA can be obtained by the following methods.
[0318]When the amino acid sequence is known, the desired DNA can be obtained by designing a DNA sequence encoding the variable region taking into consideration the frequency of occurrence of codons (Sequences of Proteins of Immunological Interest, US Dept. Health and Human Services, 1991), synthesizing several synthetic DNAs constituting approximately 100-nucleotides based on the designed DNA sequence, and carrying out PCR using the synthetic DNAs. When the nucleotide sequence is known, the desired DNA can be obtained by synthesizing several synthetic DNAs constituting approximately 100-nucleotides based on the nucleotide sequence information and carrying out PCR using the synthetic DNAs.
(3) Analysis of the Amino Acid Sequence of the V Region of an Antibody Derived from a Non-Human Animal
[0319]By comparing the whole amino acid sequences of VH and VL of the antibody including secretory signal sequences with the amino acid sequences of VH and VL of a known antibody (Sequences of Proteins of Immunological Interest, US Dept. Health and Human Services, 1991), it is possible to deduce the length of the secretory signal sequences and the N-terminal amino acid sequences and further to know the subgroup to which the antibody belongs. In addition, the amino acid sequences of CDRs of VH and VL can be deduced in a similar manner.
(4) Construction of a Human Chimeric Antibody Expression Vector
[0320]A human chimeric antibody expression vector can be constructed by inserting the cDNAs encoding VH and VL of an antibody derived from a non-human animal into sites upstream of the genes encoding CH and CL of a human antibody in the vector for expression of humanized antibody described in the above 2 (1). For example, a human chimeric antibody expression vector can be constructed by ligating the cDNAs encoding VH and VL of an antibody derived from a non-human animal respectively to synthetic DNAs comprising the 3'-terminal nucleotide sequences of VH and VL of an antibody derived from a non-human animal and the 5'-terminal nucleotide sequences of CH and CL of a human antibody and also having recognition sequences for appropriate restriction enzymes at both ends, and inserting them into sites upstream of the genes encoding CH and CL of a human antibody in the vector for humanized antibody expression described in the above 2 (1) so as to express them in an appropriate form.
(5) Construction of cDNA Encoding V Region of a Human CDR-Grafted Antibody
[0321]cDNAs encoding VH and VL of a human CDR-grafted antibody can be constructed in the following manner. First, amino acid sequences of FRs of VH and VL of a human antibody for grafting CDRs of VH and VL of a non-human animal-derived antibody are selected. The amino acid sequences of FRs of VH and VL of a human antibody may be any of those derived from human antibodies. Suitable sequences include the amino acid sequences of FRs of VHs and VLs of human antibodies registered at databases such as Protein Data Bank, and the amino acid sequences common to subgroups of FRs of VHs and VLs of human antibodies (Sequences of Proteins of Immunological Interest, US Dept. Health and Human Services, 1991). In order to prepare a human CDR-grafted antibody having a sufficient activity, it is preferred to select amino acid sequences having as high a homology as possible (at least 60% or more) with the amino acid sequences of FRs of VH and VL of the non-human animal-derived antibody of interest.
[0322]Next, the amino acid sequences of CDRs of VH and VL of the non-human animal-derived antibody of interest are grafted to the selected amino acid sequences of FRs of VH and VL of a human antibody to design amino acid sequences of VH and VL of a human CDR-grafted antibody. The designed amino acid sequences are converted into DNA sequences taking into consideration the frequency of occurrence of codons in the nucleotide sequences of antibody genes (Sequences of Proteins of Immunological Interest, US Dept. Health and Human Services, 1991), and DNA sequences encoding the amino acid sequences of VH and VL of the human CDR-grafted antibody are designed. Several synthetic DNAs constituting approximately 100-nucleotides are synthesized based on the designed DNA sequences, and PCR is carried out using the synthetic DNAs. It is preferred to design 4 to 6 synthetic DNAs for each of the H chain and the L chain in view of the reaction efficiency of PCR and the lengths of DNAs that can be synthesized.
[0323]Cloning into the vector for humanized antibody expression constructed in the above 2 (1) can be easily carried out by introducing recognition sequences for appropriate restriction enzymes to the 5' ends of synthetic DNAs present on both ends. After the PCR, the amplification products are cloned into a plasmid such as pBluescript SK(-)® (manufactured by STRATAGENE) and the nucleotide sequences are determined by the method described in the above 2 (2) to obtain a plasmid carrying DNA sequences encoding the amino acid sequences of VH and VL of the desired human CDR-grafted antibody.
(6) Modification of the Amino Acid Sequence of V Region of a Human CDR-Grafted Antibody
[0324]It is known that a human CDR-grafted antibody prepared merely by grafting CDRs of VH and VL of a non-human animal-derived antibody to FRs of VH and VL of a human antibody has a lower antigen-binding activity compared with the original non-human animal-derived antibody [BIO/TECHNOLOGY, 9, 266 (1991)]. This is probably because in VH and VL of the original non-human animal-derived antibody, not only CDRs but also some of the amino acid residues in FRs are involved directly or indirectly in the antigen-binding activity, and such amino acid residues are replaced by amino acid residues derived from FRs of VH and VL of the human antibody by CDR grafting. In order to solve this problem, attempts have been made in the preparation of a human CDR-grafted antibody to raise the lowered antigen-binding activity by identifying the amino acid residues in the amino acid sequences of FRs of VH and VL of a human antibody which are directly relating to the binding to an antigen or which are indirectly relating to it through interaction with amino acid residues in CDRs or maintenance of the tertiary structure of antibody, and modifying such amino acid residues to those derived from the original non-human animal-derived antibody [BIO/TECHNOLOGY, 9, 266 (1991)].
[0325]In the preparation of a human CDR-grafted antibody, it is most important to efficiently identify the amino acid residues in FR which are relating to the antigen-binding activity. For the efficient identification, construction and analyses of the tertiary structures of antibodies have been carried out by X ray crystallography [J. Mol. Biol., 112, 535 (1977)], computer modeling [Protein Engineering, 7, 1501 (1994)], etc. Although these studies on the tertiary structures of antibodies have provided much information useful for the preparation of human CDR-grafted antibodies, there is no established method for preparing a human CDR-grafted antibody that is adaptable to any type of antibody. That is, at present, it is still necessary to make trial-and-error approaches, e.g., preparation of several modifications for each antibody and examination of each modification for the relationship with the antigen-binding activity.
[0326]Modification of the amino acid residues in FRs of VH and VL of a human antibody can be achieved by PCR as described in the above 2 (5) using synthetic DNAs for modification. The nucleotide sequence of the PCR amplification product is determined by the method described in the above 2 (2) to confirm that the desired modification has been achieved.
(7) Construction of a Human CDR-Grafted Antibody Expression Vector
[0327]A human CDR-grafted antibody expression vector can be constructed by inserting the cDNAs encoding VH and VL of the human CDR-grafted antibody constructed in the above 2 (5) and (6) into sites upstream of the genes encoding CH and CL of a human antibody in the vector for humanized antibody expression described in the above 2 (1). For example, a human CDR-grafted antibody expression vector can be constructed by introducing recognition sequences for appropriate restriction enzymes to the 5' ends of synthetic DNAs present on both ends among the synthetic DNAs used for constructing VH and VL of the human CDR-grafted antibody in the above 2 (5) and (6), and inserting them into sites upstream of the genes encoding CH and CL of a human antibody in the vector for humanized antibody expression described in the above 2 (1) so as to express them in an appropriate form.
(8) Stable Production of a Humanized Antibody
[0328]Transformants capable of stably producing a human chimeric antibody and a human CDR-grafted antibody (hereinafter collectively referred to as humanized antibody) can be obtained by introducing the humanized antibody expression vectors described in the above 2 (4) and (7) into appropriate animal cells.
[0329]Introduction of the humanized antibody expression vector into an animal cell can be carried out by electroporation [Japanese Published Unexamined Patent Application No. 257891/90; Cytotechnology, 3, 133 (1990)], etc.
[0330]As the animal cell for introducing the humanized antibody expression vector, any animal cell capable of producing a humanized antibody can be used.
[0331]Examples of the animal cells include mouse myeloma cell lines NS0 and SP2/0, Chinese hamster ovary cells CHO/dhfr- and CHO/DG44, rat myeloma cell lines YB2/0 and IR983F, Syrian hamster kidney-derived BHK cell, and human myeloma cell line Namalwa. Preferred are Chinese hamster ovary cell CHO/DG44 and rat myeloma cell line YB2/0.
[0332]After the introduction of the humanized antibody expression vector, the transformant capable of stably producing the humanized antibody can be selected using a medium for animal cell culture containing a compound such as G418 sulfate (hereinafter referred to as G418; manufactured by SIGMA) according to the method described in Japanese Published Unexamined Patent Application No. 257891/90. Examples of the media for animal cell culture include RPMI1640 medium (manufactured by Nissui Pharmaceutical Co., Ltd.), GIT Medium® (manufactured by Nihon Pharmaceutical Co., Ltd.), EX-CELL 302 Medium® (manufactured by JRH), IMDM medium (manufactured by GIBCO BRL), Hybridoma-SFM Medium® (manufactured by GIBCO BRL), and media prepared by adding various additives such as fetal calf serum (hereinafter referred to as FCS) to these media. By culturing the obtained transformant in the medium, the humanized antibody can be formed and accumulated in the culture supernatant. The amount and the antigen-binding activity of the humanized antibody produced in the culture supernatant can be measured by enzyme-linked immunosorbent assay (hereinafter referred to as ELISA; Antibodies: A Laboratory Manual, Cold Spring Harbor Laboratory, Chapter 14, 1998; Monoclonal Antibodies: Principles and Practice, Academic Press Limited, 1996) or the like. The production of the humanized antibody by the transformant can be increased by utilizing a DHFR gene amplification system or the like according to the method described in Japanese Published Unexamined Patent Application No. 257891/90.
[0333]The humanized antibody can be purified from the culture supernatant of the transformant using a protein A column (Antibodies: A Laboratory Manual, Cold Spring Harbor Laboratory, Chapter 8, 1988; Monoclonal Antibodies: Principles and Practice, Academic Press Limited, 1996). In addition, purification methods generally employed for the purification of proteins can also be used. For example, the purification can be carried out by combinations of gel filtration, ion exchange chromatography, ultrafiltration and the like. The molecular weight of the H chain, L chain or whole antibody molecule of the purified humanized antibody can be measured by SDS-denatured polyacrylamide gel electrophoresis [hereinafter referred to as SDS-PAGE; Nature, 227, 680 (1970)], Western blotting (Antibodies: A Laboratory Manual, Cold Spring Harbor Laboratory, Chapter 12, 1988; Monoclonal Antibodies: Principles and Practice, Academic Press Limited, 1996), etc.
[0334]Shown above is the method for producing the antibody composition using an animal cell as the host. As described above, the antibody composition can also be produced using yeast, an insect cell, a plant cell, an animal or a plant by similar methods.
[0335]When a host cell inherently has the ability to express the antibody molecule, the antibody composition of the present invention can be produced by preparing a cell expressing the antibody molecule using the method described in the above 1, culturing the cell, and then purifying the desired antibody composition from the culture.
3. Evaluation of the Activity of the Antibody Composition
[0336]The protein amount, antigen-binding activity and effector function of the purified antibody composition can be measured using the known methods described in Monoclonal Antibodies, Antibody Engineering, etc.
[0337]Specifically, when the antibody composition is a humanized antibody, the activity to bind to an antigen or an antigenically positive cultured cell line can be measured by ELISA, the fluorescent antibody technique [Cancer Immunol. Immunother., 36, 373 (1993)], etc. The cytotoxic activity against an antigenically positive cultured cell line can be evaluated by measuring CDC activity, ADCC activity, etc. [Cancer Immunol. Immunother., 36, 373 (1993)].
[0338]The safety and therapeutic effect of the antibody composition in human can be evaluated using an appropriate animal model of a species relatively close to human, e.g., cynomolgus monkey.
4. Analysis of Sugar Chains in the Antibody Composition
[0339]The sugar chain structure of antibody molecules expressed in various cells can be analyzed according to general methods of analysis of the sugar chain structure of glycoproteins. For example, a sugar chain bound to an IgG molecule consists of neutral sugars such as galactose, mannose and fucose, amino sugars such as N-acetylglucosamine, and acidic sugars such as sialic acid, and can be analyzed by techniques such as sugar composition analysis and sugar chain structure analysis using two-dimensional sugar chain mapping.
(1) Analysis of Neutral Sugar and Amino Sugar Compositions
[0340]The sugar chain composition of an antibody molecule can be analyzed by carrying out acid hydrolysis of sugar chains with trifluoroacetic acid or the like to release neutral sugars or amino sugars and analyzing the composition ratio.
[0341]Specifically, the analysis can be carried out by a method using a carbohydrate analysis system (BioLC®; product of Dionex). BioLC® is a system for analyzing the sugar composition by HPAEC-PAD (high performance anion-exchange chromatography-pulsed amperometric detection) [J. Liq. Chromatogr., 6, 1577 (1983)].
[0342]The composition ratio can also be analyzed by the fluorescence labeling method using 2-aminopyridine. Specifically, the composition ratio can be calculated by fluorescence labeling an acid-hydrolyzed sample by 2-aminopyridylation according to a known method [Agric. Biol. Chem., 55(1), 283-284 (1991)] and then analyzing the composition by HPLC.
(2) Analysis of Sugar Chain Structure
[0343]The sugar chain structure of an antibody molecule can be analyzed by two-dimensional sugar chain mapping [Anal. Biochem., 171, 73 (1988); Seibutsukagaku Jikkenho (Biochemical Experimentation Methods) 23--Totanpakushitsu Tosa Kenkyuho (Methods of Studies on Glycoprotein Sugar Chains), Gakkai Shuppan Center, edited by Reiko Takahashi (1989)]. The two-dimensional sugar chain mapping is a method of deducing a sugar chain structure, for example, by plotting the retention time or elution position of a sugar chain by reversed phase chromatography as the X axis and the retention time or elution position of the sugar chain by normal phase chromatography as the Y axis, and comparing them with the results on known sugar chains.
[0344]Specifically, a sugar chain is released from an antibody by hydrazinolysis of the antibody and subjected to fluorescence labeling with 2-aminopyridine (hereinafter referred to as PA) [J. Biochem., 95, 197 (1984)]. After being separated from an excess PA-treating reagent by gel filtration, the sugar chain is subjected to reversed phase chromatography. Then, each peak of the sugar chain is subjected to normal phase chromatography. The sugar chain structure can be deduced by plotting the obtained results on a two-dimensional sugar chain map and comparing them with the spots of a sugar chain standard (manufactured by Takara Shuzo Co., Ltd.) or those in the literature [Anal. Biochem., 171, 73 (1988)].
[0345]The structure deduced by the two-dimensional sugar chain mapping can be confirmed by carrying out mass spectrometry, e.g., MALDI-TOF-MS, of each sugar chain.
5. Immunoassay for Determining the Sugar Chain Structure of an Antibody Molecule
[0346]An antibody composition comprises an antibody molecule having different sugar chain structures binding to the Fc region of antibody. The antibody composition of the present invention, in which the ratio of a sugar chain in which fucose is not bound to the N-acetylglucosamine in the reducing end to the total complex type N-glycoside-linked sugar chains bound to the Fc region is 100%, has high ADCC activity. Such an antibody composition can be identified using the method for analyzing the sugar chain structure of an antibody molecule described in the above 4. Further, it can also be identified by immunoassays using lectins.
[0347]Discrimination of the sugar chain structure of an antibody molecule by immunoassays using lectins can be made according to the immunoassays such as Western staining, RIA (radioimmunoassay), VIA (viroimmunoassay), EIA (enzymoimmunoassay), FIA (fluoroimmunoassay) and MIA (metalloimmunoassay) described in the literature [Monoclonal Antibodies: Principles and Applications, Wiley-Liss, Inc. (1995); Enzyme Immunoassay, 3rd Ed., Igaku Shoin (1987); Enzyme Antibody Technique, Revised Edition, Gakusai Kikaku (1985); etc.], for example, in the following manner.
[0348]A lectin recognizing the sugar chain structure of an antibody molecule is labeled, and the labeled lectin is subjected to reaction with a sample antibody composition, followed by measurement of the amount of a complex of the labeled lectin with the antibody molecule.
[0349]Examples of lectins useful for determining the sugar chain structure of an antibody molecule include WGA (wheat-germ agglutinin derived from T. vulgaris), ConA (concanavalin A derived from C. ensiformis), RIC (toxin derived from R. communis), L-PHA (leukoagglutinin derived from P. vulgaris), LCA (lentil agglutinin derived from L. culinaris), PSA (pea lectin derived from P. sativum), AAL (Aleuria aurantia lectin), ACL (Amaranthus caudatus lectin), BPL (Bauhinia purpurea lectin), DSL (Datura stramonium lectin), DBA (Dolichos biflorus agglutinin), EBL (Elderberry balk lectin), ECL (Erythrina cristagalli lectin), EEL (Euonymus europaeus lectin), GNL (Galanthus nivalis lectin), GSL (Griffonia simplicifolia lectin), HPA (Helix pomatia agglutinin), HHL (Hippeastrum hybrid lectin), Jacalin, LTL (Lotus tetragonolobus lectin), LEL (Lycopersicon esculentum lectin), MAL (Maackia amurensis lectin), MPL (Maclura pomifera lectin), NPL (Narcissus pseudonarcissus lectin), PNA (peanut agglutinin), E-PHA (Phaseolus vulgaris erythroagglutinin), PTL (Psophocarpus tetragonolobus lectin), RCA (Ricinus communis agglutinin), STL (Solanum tuberosum lectin), SJA (Sophora japonica agglutinin), SBA (soybean agglutinin), UEA (Ulex europaeus agglutinin), VVL (Vicia villosa lectin) and WFA (Wisteria floribunda agglutinin).
[0350]It is preferred to use lectins specifically recognizing a sugar chain structure wherein fucose is bound to the N-acetylglucosamine in the reducing end in complex type N-glycoside-linked sugar chains. Examples of such lectins include lentil lectin LCA (lentil agglutinin derived from Lens culinaris), pea lectin PSA (pea lectin derived from Pisum sativum), broad bean lectin VFA (agglutinin derived from Vicia faba) and Aleuria aurantia lectin AAL (lectin derived from Aleuria aurantia).
6. Utilization of the Antibody Composition of the Present Invention
[0351]Since the antibody composition of the present invention specifically binds to human CCR4 and has high antibody-dependent cell-mediated cytotoxic activity, it is useful for the prevention and treatment of various diseases in which CCR4-expressing cells are concerned, including cancer and inflammatory diseases.
[0352]Examples of the cancer for which treatment by the antibody composition of the present invention is effective include blood cancers, particularly leukemia or lymphoma. Specific examples include adult T cell leukemia, ATL, mycosis fungoides, Sezary's syndrome, undifferentiated large cell lymphoma, non-specific T cell lymphoma and the like.
[0353]Examples of the inflammatory diseases for which treatment by the antibody composition of the present invention is effective include immunodiseases mediated by Th2 cells, such as acute or chronic bronchial hyperresponsiveness, bronchial asthma, atopic skin diseases including atopic dermatitis, chronic sinusitis, Churg-Strauss syndrome, nettle rash, pemphigus, eosinophilic myocarditis, allergic enterogastritis, allergic granulomatous angitis, allergic rhinitis and pollenosis.
[0354]In cancer such as malignant tumor, cancer cells proliferate abnormally. For example, in B cell lymphoma, specific B cells proliferate abnormally. Ordinary anti-tumor agents are characterized by the suppression of the growth of cancer cells. In contrast, antibodies having high antibody-dependent cell-mediated cytotoxic activity can treat cancer by injuring cancer cells which express antigens due to cell-killing effect, and therefore, they are more effective as therapeutic agents for cancer than ordinary anti-tumor agents. At present, therapeutic antibodies used as therapeutic agents for cancer have only insufficient anti-tumor effect when used alone and thus are used in combination with chemotherapy [Science, 280, 1197 (1998)]. If more potent anti-tumor effect is provided by the antibody composition of the present invention alone, the dependency on chemotherapy will be decreased and side effects will be reduced as well.
[0355]In allergic inflammatory diseases such as chronic airway hypersensitivity asthma, bronchial asthma, atopic skin diseases including atopic dermatitis, allergic rhinitis and pollinosis, inflammatory cells such as eosinophils and mast cells are induced to proliferate or differentiate by cytokines and chemokines such as IL-4, IL-5 and IL-13 produced by Th2 cells, and tissue injury or allergic reaction is induced via biofunctional molecules produced by these inflammatory cells. As therapeutic agents for the treatment of such immunodiseases mediated by Th2 cells, antagonists to cytokine such as IL-4, IL-5 or IL-13 produced by Th2 cells, cytokine production inhibitors thereof, and inhibitors against inflammatory cells and inflammatory biofunctional molecules have been developed. However, these therapeutic agents inhibit only a part of the complicated network among cytokines, chemokines and inflammatory cells and are not complete treatments. Since the antibody composition of the present invention specifically binds to the extracellular region of CCR4 and shows potent cytotoxic activity against CCR4-expressing cells, it can selectively eliminate Th2 cells which are CCR4-expressing cells present in the upstream part of allergic reactions and it further suppresses production of IL-4, IL-5, and IL-13 from Th2 cells. Accordingly, it is effective as a therapeutic agent for the above inflammatory diseases.
[0356]Since the antibody composition of the present invention includes no antibody molecule comprising sugar chains to which fucose is bound, its cytotoxic activity is enhanced. Therefore, patients of the above cancers, inflammatory diseases and the like who cannot be treated by an antibody composition which comprises an antibody molecule having sugar chains to which fucose is bound can be treated by the antibody composition of the present invention.
[0357]Especially, among the above diseases, since it is difficult to deliver a drug to the infiltration region of CCR4-expressing cells in the diseases such as cancers, bronchial asthma and chronic sinusitis, it is preferable that therapeutic effects can be obtained by a small amount of drug. Since the antibody composition of the present invention has high ADCC activity at a small amount, it is effective for treatment of these diseases.
[0358]A pharmaceutical composition comprising the antibody composition of the present invention may be administered alone as a therapeutic agent. However, it is preferably mixed with one or more pharmaceutically acceptable carriers and provided as a pharmaceutical preparation produced by an arbitrary method well known in the technical field of pharmaceutics.
[0359]It is desirable to administer the pharmaceutical composition by the route that is most effective for the treatment. Suitable administration routes include oral administration and parenteral administration such as intraoral administration, intratracheal administration, intrarectal administration, subcutaneous administration, intramuscular administration and intravenous administration. In the case of an antibody preparation, intravenous administration is preferable.
[0360]The pharmaceutical preparation may be in the form of spray, capsules, tablets, granules, syrup, emulsion, suppository, injection, ointment, tape, and the like.
[0361]The pharmaceutical preparations suitable for oral administration include emulsions, syrups, capsules, tablets, powders and granules.
[0362]Liquid preparations such as emulsions and syrups can be prepared using, as additives, water, sugars (e.g., sucrose, sorbitol and fructose), glycols (e.g., polyethylene glycol and propylene glycol), oils (e.g., sesame oil, olive oil and soybean oil), antiseptics (e.g., p-hydroxybenzoates), flavors (e.g., strawberry flavor and peppermint), and the like.
[0363]Capsules, tablets, powders, granules, etc. can be prepared using, as additives, excipients (e.g., lactose, glucose, sucrose and mannitol), disintegrators (e.g., starch and sodium alginate), lubricants (e.g., magnesium stearate and talc), binders (e.g., polyvinyl alcohol, hydroxypropyl cellulose and gelatin), surfactants (e.g., fatty acid esters), plasticizers (e.g., glycerin), and the like.
[0364]The pharmaceutical preparations suitable for parenteral administration include injections, suppositories and sprays.
[0365]Injections can be prepared using carriers comprising a salt solution, a glucose solution, or a mixture thereof, etc. It is also possible to prepare powder injections by freeze-drying the antibody composition according to a conventional method and adding sodium chloride thereto.
[0366]Suppositories can be prepared using carriers such as cacao butter, hydrogenated fat and carboxylic acid.
[0367]The antibody composition may be administered as such in the form of spray, but sprays may be prepared using carriers which do not stimulate the oral or airway mucous membrane of a recipient and which can disperse the antibody composition as fine particles to facilitate absorption thereof.
[0368]Suitable carriers include lactose and glycerin. It is also possible to prepare aerosols, dry powders, etc. according to the properties of the antibody composition and the carriers used. In preparing these parenteral preparations, the above-mentioned additives for the oral preparations may also be added.
[0369]The dose and administration frequency will vary depending on the desired therapeutic effect, the administration route, the period of treatment, the patient's age and body weight, etc. However, an appropriate dose of the active ingredient for an adult person is generally 10 μg/kg to 20 mg/kg per day.
[0370]The anti-tumor effect of the antibody composition against various tumor cells can be examined by in vitro tests such as CDC activity measurement and ADCC activity measurement and in vivo tests such as anti-tumor experiments using tumor systems in experimental animals (e.g., mice).
[0371]The CDC activity and ADCC activity measurements and anti-tumor experiments can be carried out according to the methods described in the literature [Cancer Immunology Immunotherapy, 36, 373 (1993); Cancer Research, 54, 1511 (1994); etc.].
[0372]Certain embodiments of the present invention are illustrated in the following examples. These examples are not to be construed as limiting the scope of the present invention.
EXAMPLE 1
Construction of CHO/DG44 Cell Line in which Both Alleles of α1,6-Fucosyltransferase (Hereinafter Referred to as FUT8) on the Genome have been Disrupted
[0373]The CHO/DG44 cell line comprising the deletion of a genome region for both alleles of FUT8 including the translation initiation codons was constructed according to the following steps.
1. Construction of Targeting Vector pKOFUT8Neo Comprising Exon 2 of Chinese Hamster FUT8 Gene
[0374]pKOFUT8Neo was constructed in the following manner using targeting vector pKOFUT8Puro comprising exon 2 of Chinese hamster FUT8 gene constructed by the method described in Example 13-1 of WO02/31140, and pKOSelectNeo® (manufactured by Lexicon).
[0375]pKOSelectNeo® (manufactured by Lexicon) was digested with the restriction enzyme AscI (manufactured by New England Biolabs) and subjected to agarose gel electrophoresis, and approximately 1.6 Kb AscI fragment comprising the neomycin resistance gene expression unit was recovered using GENECLEAN Spin Kit® (manufactured by BIO 101).
[0376]After pKOFUT8Puro was digested with the restriction enzyme AscI (manufactured by New England Biolabs), the end of the DNA fragment with alkaline phosphatase derived from Escherichia coli C15 (manufactured by Takara Shuzo Co., Ltd.) was dephosphorylated. After the reaction, the DNA fragment was purified by phenol/chloroform extraction and ethanol precipitation.
[0377]Sterilized water was added to 0.1 μg of the pKOSelectNeo-derived AscI fragment (approximately 1.6 Kb) and 0.1 μg of the pKOFUT8Puro-derived AscI fragment (approximately 10.1 Kb) obtained above to make up to 5 μl, and 5 μl of Ligation High (manufactured by Toyobo Co., Ltd.) was added thereto. The ligation reaction was carried out at 16° C. for 30 minutes. Escherichia coli DH5α was transformed using the resulting reaction mixture, and a plasmid DNA was prepared from each of the obtained ampicillin-resistant clones. The plasmid DNA was subjected to reaction using BigDye Terminator Cycle Sequencing Ready Reaction Kit v2.0® (manufactured by Applied Biosystems) according to the attached instructions, and the nucleotide sequence was analyzed using DNA Sequencer ABI PRISM 377 (manufactured by Applied Biosystems). The thus obtained plasmid pKOFUT8Neo shown in FIG. 1 was used as a targeting vector for the subsequent preparation of FUT8 gene-hemi-knockout CHO cell line.
2. Preparation of Hemi-Knockout Cell Line in which One Copy of the FUT8 Gene on the Genome has been Disrupted(1) Obtaining of a Cell Line in which the Targeting Vector pKOFUT8Neo has been Introduced
[0378]The Chinese hamster FUT8 genome region targeting vector pKOFUT8Neo constructed in Example 1-1 was introduced into Chinese hamster ovary-derived CHO/DG44 cells deficient in the dihydrofolate reductase gene (dhfr) [Somataic Cell and Molecular Genetics, 12, 555 (1986)] in the following manner.
[0379]pKOFUT8Neo was digested with the restriction enzyme SalI (manufactured by New England Biolabs) for linearization, and 4 μg of the linearized pKOFUT8Neo was introduced into 1.6×106 CHO/DG44 cells by electroporation [Cytotechnology, 3, 133 (1990)]. The resulting cells were suspended in IMDM-dFBS (10)-HT(1) [IMDM medium (manufactured by Invitrogen) containing 10% dialysis FBS (manufactured by Invitrogen) and 1-fold concentration HT supplement (manufactured by Invitrogen)] and then seeded on a 10-cm dish for adherent cell culture (manufactured by Falcon). After culturing in a 5% CO2 incubator at 37° C. for 24 hours, the medium was replaced with 10 ml of IMDM-dFBS(10) (IMDM medium containing 10% dialysis FBS) containing 600 μg/ml G418 (manufactured by Nacalai Tesque, Inc.). Culturing was carried out in a 5% CO2 incubator at 37° C. for 15 days during which the above medium replacement was repeated every 3 to 4 days to obtain G418-resistant clones.
(2) Confirmation of Homologous Recombination by Genomic PCR
[0380]Confirmation of the homologous recombination in the G418-resistant clones obtained in the above (1) was carried out by PCR using genomic DNA in the following manner.
[0381]The G418-resistant clones on a 96-well plate were subjected to trypsinization, and a 2-fold volume of a frozen medium (20% DMSO, 40% fetal calf serum and 40% IMDM) was added to each well to suspend the cells. One half of the cell suspension in each well was seeded on a flat-bottomed 96-well plate for adherent cells (manufactured by Asahi Techno Glass) to prepare a replica plate, while the other half was stored by cryopreservation as a master plate.
[0382]The neomycin-resistant clones on the replica plate were cultured using IMDM-dFBS(10) containing 600 μg/ml G418 in a 5% CO2 incubator at 37° C. for one week, followed by recovery of cells. The genomic DNA of each clone was prepared from the recovered cells according to a known method [Analytical Biochemistry, 201, 331 (1992)] and then dissolved overnight in 30 μl of TE-RNase buffer (pH 8.0) (10 mmol/l Tris-HCL, 1 mmol/l EDTA, 200 μg/ml RNase A).
[0383]Primers used in the genomic PCR were designed as follows. Primers respectively having the sequences represented by SEQ ID NOs:39 and 40, which are contained in the sequence of the FUT8 genome region obtained by the method described in Example 12 of WO03/31140 (SEQ ID NO:13), were employed as forward primers. Primers respectively having the sequences represented by SEQ ID NOs:41 and 42 which specifically bind to the loxP sequence of the targeting vector were employed as reverse primers in the following polymerase chain reaction (PCR). A reaction mixture [25 μl; DNA polymerase ExTaq® (manufactured by Takara Shuzo Co., Ltd.), ExTaq Buffer® (manufactured by Takara Shuzo Co., Ltd.), 0.2 mmol/l dNTPs, 0.5 μmol/l each of the above primers (a combination of a forward primer and a reverse primer)] containing 10 μl of each genomic DNA solution prepared above was prepared, and PCR was carried out, after heating at 94° C. for 3 minutes, by cycles, one cycle consisting of reaction at 94° C. for one minute, reaction at 60° C. for one minute and reaction at 72° C. for 2 minutes.
[0384]After the PCR, the reaction mixture was subjected to 0.8% (w/v) agarose gel electrophoresis, and cell lines with which a specific amplification product (approximately 1.7 Kb) resulting from the homologous recombination was observed were judged to be positive clones.
(3) Confirmation of Homologous Recombination by Genomic Southern Blotting
[0385]Confirmation of the homologous recombination in the positive clones obtained in the above (2) was carried out by Southern blotting using genomic DNA in the following manner.
[0386]From the master plates stored by cryopreservation in the above (2), a 96-well plate containing the positive clones found in (2) was selected. After the plate was allowed to stand in a 5% CO2 incubator at 37° C. for 10 minutes, the cells in the wells corresponding to the positive clones were seeded on a flat-bottomed 24-well plate for adherent cells (manufactured by Greiner). After culturing using IMDM-dFBS(10) containing 600 μg/ml G418 in a 5% CO2 incubator at 37° C. for one week, the cells were seeded on a flat-bottomed 6-well plate for adherent cells (manufactured by Greiner). The plate was subjected to culturing in a 5% CO2 incubator at 37° C. and the cells were recovered. The genomic DNA of each clone was prepared from the recovered cells according to a known method [Nucleic Acids Research, 3, 2303 (1976)] and then dissolved overnight in 150 μl of TE-RNase buffer (pH 8.0).
[0387]The genomic DNA prepared above (12 μg) was digested with the restriction enzyme BamHI (manufactured by New England Biolabs), and a DNA fragment recovered by ethanol precipitation was dissolved in 20 μl of TE buffer (pH 8.0) (10 mmol/l Tris-HCL, 1 mmol/l EDTA) and then subjected to 0.6% (w/v) agarose gel electrophoresis. After the electrophoresis, the genomic DNA was transferred to a nylon membrane according to a known method [Proc. Natl. Acad. Sci. USA, 76, 3683 (1979)], followed by heat treatment of the nylon membrane at 80° C. for 2 hours for immobilization.
[0388]Separately, a probe used in the Southern blotting was prepared in the following manner. Primers respectively having the sequences represented by SEQ ID NOs:43 and 44, which are contained in the sequence of the FUT8 genome region obtained by the method described in Example 12 of WO03/31140 (SEQ ID NO:13), were prepared and used in the following PCR. A reaction mixture [20 μl; DNA polymerase ExTaq® (manufactured by Takara Shuzo Co., Ltd.), ExTaq Buffer® (manufactured by Takara Shuzo Co., Ltd.), 0.2 mmol/l dNTPs, 0.5 μmol/l each of the above primers] containing 4.0 ng of pFUT8fgE2-2 described in Example 12 of WO02/31140 as a template was prepared, and PCR was carried out, after heating at 94° C. for one minute, by 25 cycles, one cycle consisting of reaction at 94° C. for 30 seconds, reaction at 55° C. for 30 seconds and reaction at 74° C. for one minute.
[0389]After the PCR, the reaction mixture was subjected to 1.75% (w/v) agarose gel electrophoresis, and approximately 230 bp probe DNA fragment was recovered using GENECLEAN Spin Kit® (manufactured by BIO101). A 5-μl portion of the obtained probe DNA solution was subjected to radiolabeling using [α-32P] dCTP 1.75 MBq and Megaprime DNA Labelling system, dCTP® (manufactured by Amersham Pharmacia Biotech).
[0390]Hybridization was carried out in the following manner. The above nylon membrane to which the genomic DNA digestion product had been transferred was put into a roller bottle and 15 ml of a hybridization solution [5×SSPE, 50×Denhardt's solution, 0.5% (w/v) SDS, 100 μg/ml salmon sperm DNA] was added thereto. Prehybridization was carried out at 65° C. for 3 hours. Then, the 32P-labeled probe DNA was heat-denatured and put into the bottle, and hybridization was carried out at 65° C. overnight.
[0391]After the hybridization, the nylon membrane was immersed in 50 ml of a primary washing solution [2×SSC-0.1% (w/v) SDS] and washed by heating at 65° C. for 15 minutes. After this washing step was repeated twice, the nylon membrane was immersed in 50 ml of a secondary washing solution [0.2×SSC-0.1% (w/v) SDS] and washed by heating at 65° C. for 15 minutes. Then, the nylon membrane was exposed to an X-ray film at -80° C. for development.
[0392]FIG. 2 shows the results of the analysis of the genomic DNAs of the parent cell line CHO/DG44 and the 50-10-104 cell line, which is the positive clone obtained in the above (2), according to the present method. In the CHO/DG44 cell line, only approximately 25.5 Kb fragment derived from the wild-type FUT8 allele was detected. On the other hand, in the positive clone, i.e. 50-10-104 cell line, approximately 20.0 Kb fragment peculiar to the allele which underwent homologous recombination was detected in addition to approximately 25.5 Kb fragment derived from the wild-type FUT8 allele. The quantitative ratio of these two kinds of fragments was 1:1, whereby it was confirmed that the 50-10-104 cell line was a hemi-knockout clone wherein one copy of the FUT8 allele was disrupted.
3. Preparation of CHO/DG44 Cell Line in which the FUT8 Gene on the Genome has been Double-Knocked Out(1) Preparation of a Cell Line in which Targeting Vector pKOFUT8Puro has been Introduced
[0393]In order to disrupt the other FUT8 allele in the FUT8 gene-hemi-knockout clone obtained in the above 2, the Chinese hamster FUT8 gene exon 2 targeting vector pKOFUT8Puro described in Example 13-1 of WO02/31140 was introduced into the clone in the following manner.
[0394]pKOFUT8Puro was digested with the restriction enzyme SalI (manufactured by New England Biolabs) for linearization, and 4 μg of the linearized pKOFUT8Puro was introduced into 1.6×106 cells of the FUT8 gene-hemi-knockout clone by electroporation [Cytotechnology, 3, 133 (1990)]. The resulting cells were suspended in IMDM-dFBS(10)-HT(1) and then seeded on a 10-cm dish for adherent cell culture (manufactured by Falcon). After culturing in a 5% CO2 incubator at 37° C. for 24 hours, the medium was replaced with 10 ml of IMDM-dFBS(10)-HT(1) containing 15 μg/ml puromycin (manufactured by SIGMA). Culturing was carried out in a 5% CO2 incubator at 37° C. for 15 days during which the above medium replacement was repeated every 7 days to obtain puromycin-resistant clones.
(2) Confirmation of Homologous Recombination by Genomic Southern Blotting
[0395]Confirmation of the homologous recombination in the drug-resistant clones obtained in the above (1) was carried out by Southern blotting using genomic DNA in the following manner.
[0396]The puromycin-resistant clones were recovered into a flat-bottomed plate for adherent cells (manufactured by Asahi Techno Glass) according to a known method [Gene Targeting, Oxford University Press (1993)], followed by culturing using IMDM-dFBS(10)-HT(1) containing 15 μg/ml puromycin (manufactured by SIGMA) in a 5% CO2 incubator at 37° C. for one week.
[0397]After the culturing, each clone on the above plate was subjected to trypsinization and the resulting cells were seeded on a flat-bottomed 24-well plate for adherent cells (manufactured by Greiner). After culturing using IMDM-dFBS(10)-HT(1) containing 15 μg/ml puromycin (manufactured by SIGMA) in a 5% CO2 incubator at 37° C. for one week, the cells were subjected to trypsinization again and then seeded on a flat-bottomed 6-well plate for adherent cells (manufactured by Greiner). The plate was subjected to culturing in a 5% CO2 incubator at 37° C. and the cells were recovered. The genomic DNA of each clone was prepared from the recovered cells according to a known method [Nucleic Acids Research, 3, 2303 (1976)] and then dissolved overnight in 150 μl of TE-RNase buffer (pH 8.0).
[0398]The genomic DNA prepared above (12 μg) was digested with the restriction enzyme BamHI (manufactured by New England Biolabs), and a DNA fragment recovered by ethanol precipitation was dissolved in 20 μl of TE buffer (pH 8.0) and then subjected to 0.6% (w/v) agarose gel electrophoresis. After the electrophoresis, the genomic DNA was transferred to a nylon membrane according to a known method [Proc. Natl. Acad. Sci. USA, 76, 3683 (1979)], followed by heat treatment of the nylon membrane at 80° C. for 2 hours for immobilization.
[0399]Separately, a probe used in the Southern blotting was prepared in the following manner. Primers respectively having the sequences represented by SEQ ID NOs:45 and 46, which specifically bind to the sequences closer to the 5' end than the FUT8 genome region contained in the targeting vector, were prepared and used in the following PCR. A reaction mixture [20 μl; DNA polymerase ExTaq (manufactured by Takara Shuzo Co., Ltd.), ExTaq Buffer® (manufactured by Takara Shuzo Co., Ltd.), 0.2 mmol/l dNTPs, 0.5 μmol/l each of the above primers] containing 4.0 ng of the plasmid pFUT8fgE2-2 described in Example 12 of WO02/31140 as a template was prepared, and PCR was carried out, after heating at 94° C. for one minute, by 25 cycles, one cycle consisting of reaction at 94° C. for 30 seconds, reaction at 55° C. for 30 seconds and reaction at 74° C. for one minute.
[0400]After the PCR, the reaction mixture was subjected to 1.75% (w/v) agarose gel electrophoresis, and approximately 230 bp probe DNA fragment was purified using GENECLEAN Spin Kit® (manufactured by BIO101). A 5-μl portion of the obtained probe DNA solution was subjected to radiolabeling using [α-32P] dCTP 1.75 MBq and Megaprime DNA Labelling System®, dCTP (manufactured by Amersham Pharmacia Biotech).
[0401]Hybridization was carried out in the following manner. The above nylon membrane to which the genomic DNA digestion product had been transferred was put into a roller bottle and 15 ml of a hybridization solution [5×SSPE, 50×Denhardt's solution, 0.5% (w/v) SDS, 100 μg/ml salmon sperm DNA] was added thereto. Prehybridization was carried out at 65° C. for 3 hours. Then, the 32P-labeled probe DNA was heat-denatured and put into the bottle, and hybridization was carried out at 65° C. overnight.
[0402]After the hybridization, the nylon membrane was immersed in 50 ml of a primary washing solution [2×SSC-0.1% (w/v) SDS] and washed by heating at 65° C. for 15 minutes. After this washing step was repeated twice, the nylon membrane was immersed in 50 ml of a secondary washing solution [0.2×SSC-0.1% (w/v) SDS] and washed by heating at 65° C. for 15 minutes. Then, the nylon membrane was exposed to an X-ray film at -80° C. for development.
[0403]FIG. 3 shows the result of the analysis of the genomic DNA of the WK704 cell line, which is one of the puromycin-resistant clones obtained from the 50-10-104 cell line by the method described in the above (1), according to the present method. In the WK704 cell line, approximately 25.5 Kb fragment derived from the wild-type FUT8 allele was not detected and only approximately 20.0 Kb fragment specific to the allele which underwent homologous recombination (indicated by arrow in the figure) was detected. From this result, it was confirmed that the WK704 cell line was a clone wherein both FUT8 alleles were disrupted.
4. Removal of the Drug Resistance Genes from FUT8 Gene-Double-Knockout Cells
(1) Introduction of Cre Recombinase Expression Vector
[0404]For the purpose of removing the drug resistance genes from the FUT8 gene-double-knockout clone obtained in the above item 3, the Cre recombinase expression vector pBS185® (manufactured by Life Technologies) was introduced into the clone in the following manner.
[0405]pBS185 (4 μg) was introduced into 1.6×106 cells of the FUT8 gene-double-knockout clone by electroporation [Cytotechnology, 3, 133 (1990)]. The resulting cells were suspended in 10 ml of IMDM-dFBS(10)-HT(1) and the suspension was diluted 20000-fold with the same medium. The diluted suspension was seeded on seven 10-cm dishes for adherent cell culture (manufactured by Falcon), followed by culturing in a 5% CO2 incubator at 37° C. for 10 days to form colonies.
(2) Obtaining of a Cell Line in which the Cre Recombinase Expression Vector has been Introduced
[0406]Clones arbitrarily selected from the colonies obtained in the above (1) were recovered into a flat-bottomed plate for adherent cells (manufactured by Asahi Techno Glass) according to a known method [Gene Targeting, Oxford University Press (1993)], followed by culturing using IMDM-dFBS(10)-HT(1) in a 5% CO2 incubator at 37° C. for one week.
[0407]After the culturing, each clone on the above plate was subjected to trypsinization, and a 2-fold volume of a frozen medium (20% DMSO, 40% fetal calf serum and 40% IMDM) was added to each well to suspend the cells. One half of the cell suspension in each well was seeded on a flat-bottomed 96-well plate for adherent cells (manufactured by Asahi Techno Glass) to prepare a replica plate, while the other half was stored by cryopreservation as a master plate.
[0408]The cells on the replica plate were cultured using IMDM-dFBS(10)-HT(1) containing 600 μg/ml G418 and 15 μg/ml puromycin in a 5% CO2 incubator at 37° C. for one week. Positive clones in which the drug resistance genes inserted between loxP sequences has been removed by the expression of Cre recombinase have died in the presence of G418 and puromycin. The positive clones were selected in this manner.
(3) Confirmation of Removal of the Drug Resistance Genes by Genomic Southern Blotting
[0409]Confirmation of the removal of the drug resistance genes in the positive clones selected in the above (2) was carried out by genomic Southern blotting in the following manner.
[0410]From the master plates stored by cryopreservation in the above (2), a 96-well plate containing the above positive clones was selected. After the plate was allowed to stand in a 5% CO2 incubator at 37° C. for 10 minutes, the cells in the wells corresponding to the above clones were seeded on a flat-bottomed 24-well plate for adherent cells (manufactured by Greiner). After culturing using IMDM-dFBS(10)-HT(1) for one week, the cells were subjected to trypsinization and then seeded on a flat-bottomed 6-well plate for adherent cells (manufactured by Greiner). The plate was subjected to culturing in a 5% CO2 incubator at 37° C. and the proliferated cells were recovered. The genomic DNA of each clone was prepared from the recovered cells according to a known method [Nucleic Acids Research, 3, 2303 (1976)] and then dissolved overnight in 150 μl of TE-RNase buffer (pH 8.0).
[0411]The genomic DNA prepared above (12 μg) was digested with the restriction enzyme NheI (manufactured by New England Biolabs), and a DNA fragment recovered by ethanol precipitation was dissolved in 20 μl of TE buffer (pH 8.0) and then subjected to 0.6% (w/v) agarose gel electrophoresis. After the electrophoresis, the genomic DNA was transferred to a nylon membrane according to a known method [Proc. Natl. Acad. Sci. USA, 76, 3683 (1979)], followed by heat treatment of the nylon membrane at 80° C. for 2 hours for immobilization.
[0412]Separately, a probe used in the Southern blotting was prepared in the following manner. PCR was carried out using primers respectively having the sequences represented by SEQ ID NOs:45 and 46, which specifically bind to the sequences closer to the 5' end than the FUT8 genome region contained in the targeting vector. That is, a reaction mixture [20 μl; DNA polymerase ExTaq (manufactured by Takara Shuzo Co., Ltd.), ExTaq Buffer® (manufactured by Takara Shuzo Co., Ltd.), 0.2 mmol/l dNTPs, 0.5 μmol/l each of the above primers] containing 4.0 ng of the plasmid pFUT8fgE2-2 described in Example 12 of WO02/31140 as a template was prepared, and PCR was carried out, after heating at 94° C. for one minute, by 25 cycles, one cycle consisting of reaction at 94° C. for 30 seconds, reaction at 55° C. for 30 seconds and reaction at 74° C. for one minute.
[0413]After the PCR, the reaction mixture was subjected to 1.75% (w/v) agarose gel electrophoresis, and approximately 230 bp probe DNA fragment was purified using GENECLEAN Spin Kit® (manufactured by BIO101). A 5-μl portion of the obtained probe DNA solution was subjected to radiolabeling using [α-32P] dCTP 1.75 MBq and Megaprime DNA Labelling System®, dCTP (manufactured by Amersham Pharmacia Biotech).
[0414]Hybridization was carried out in the following manner. The above nylon membrane to which the genomic DNA digestion product had been transferred was put into a roller bottle and 15 ml of a hybridization solution [5×SSPE, 50×Denhardt's solution, 0.5% (w/v) SDS, 100 μg/ml salmon sperm DNA] was added thereto. Prehybridization was carried out at 65° C. for 3 hours. Then, the 32P-labeled probe DNA was heat-denatured and put into the bottle, and hybridization was carried out at 65° C. overnight.
[0415]After the hybridization, the nylon membrane was immersed in 50 ml of a primary washing solution [2×SSC-0.1% (w/v) SDS] and washed by heating at 65° C. for 15 minutes. After this washing step was repeated twice, the nylon membrane was immersed in 50 ml of a secondary washing solution [0.2×SSC-0.1% (w/v) SDS] and washed by heating at 65° C. for 15 minutes. Then, the nylon membrane was exposed to an X-ray film at -80° C. for development.
[0416]FIG. 4 shows the results of the analysis of the genomic DNAs of the parent cell line CHO/DG44, the 50-10-104 cell line described in the above item 2, the WK704 cell line described in the above item 3, and the 4-5-C3 cell line, which is one of the drug-sensitive clones obtained from the WK704 cell line by the method described in the above (2), according to the present method. In the CHO/DG44 cell line, only approximately 8.0 Kb DNA fragment derived from the wild-type FUT8 allele was detected. In the 50-10-104 cell line and the WK704 cell line, approximately 9.5 Kb DNA fragment derived from the allele which underwent homologous recombination was observed. On the other hand, in the 4-5-C3 cell line, only approximately 8.0 Kb DNA fragment resulting from the removal of the neomycin resistance gene (approximately 1.6 Kb) and the puromycin resistance gene (approximately 1.5 Kb) from the allele which underwent homologous recombination was detected. From the above results, it was confirmed that the drug resistance genes had been removed by Cre recombinase in the 4-5-C3 cell line.
[0417]Besides the 4-5-C3 cell line, plural FUT8 gene-double-knockout clones in which the drug-resistance gene had been removed (hereinafter referred to as FUT8 gene-double-knockout cells) were obtained.
EXAMPLE 2
Expression of an Anti-CCR4 Human CDR-Grafted Antibody Composition in FUT8 Gene-Double-Knockout Cell
1. Stable Expression in FUT8 Gene-Double-Knockout Cell
[0418]Cells stably producing an anti-CCR4 human CDR-grafted antibody were prepared by the method described in Example 1-2 (2) of WO02/31140 using the FUT8 gene-double-knockout cell described in Example 1-4 above and the parent cell CHO/DG44 as host cells. The anti-CCR4 human CDR-grafted antibody expression vector was prepared by the method described in Example 1 of WO03/18635. FIG. 5 shows the schematic view of the produced expression vector for the anti-CCR4 human CDR-grafted antibody.
[0419]As a result, transformants which are capable of growing in IMDM-dFBS(10) containing 500 μg/ml G418 and which produce the anti-CCR4 human CDR-grafted antibody were obtained. The transformants obtained from the parent cell CHO/DG44 and the FUT8 gene-double-knockout cell were designated DG44/CCR4 cell line and Ms705/CCR4 cell line, respectively.
2. Measurement of the Human IgG Antibody Concentration in Culture Supernatant (ELISA)
[0420]Goat anti-human IgG (manufactured by H & L) antibody (manufactured by American Qualex) was diluted with Phosphate Buffered Saline (hereinafter referred to as PBS) (manufactured by Invitrogen) to a concentration of 1 μg/ml and put into wells of a 96-well plate for ELISA (manufactured by Greiner) in an amount of 50 μl/well, followed by standing at 4° C. overnight for adsorption. After washing with PBS, PBS containing 1% BSA (hereinafter referred to as 1% BSA-PBS) (manufactured by Wako Pure Chemical Industries, Ltd.) was added to the wells in an amount of 100 μL/well, followed by reaction at room temperature for one hour to block the remaining active groups. Then, the 1% BSA-PBS was discarded, and 50 μl each of the culture supernatant of transformant or variously diluted solutions of an antibody purified from the culture supernatant were respectively added to the wells, followed by reaction at room temperature for one hour. After the reaction, the wells were washed with PBS containing 0.05% Tween 20 (hereinafter referred to as Tween-PBS) (manufactured by Wako Pure Chemical Industries, Ltd.). To each well was added 50 μl of peroxidase-labeled goat anti-human IgG (manufactured by H & L) antibody solution (manufactured by American Qualex) diluted 2000-fold with 1% BSA-PBS as a secondary antibody solution, followed by reaction at room temperature for one hour. After the reaction, the wells were washed with Tween-PBS, and 50 μl of ABTS substrate solution [a solution prepared by dissolving 0.55 g of 2,2'-azino-bis(3-ethylbenzothiazoline-6-sulfonic acid) ammonium (manufactured by Wako Pure Chemical Industries, Ltd.) in 1 liter of 0.1 M citrate buffer (pH 4.2) and adding thereto, just before use, 1 μl ml hydrogen peroxide (manufactured by Wako Pure Chemical Industries, Ltd.)] was added to each well to develop color. Then, the absorbance at 415 nm (hereinafter referred to as OD 415) was measured.
3. Purification of Anti-CCR4 Human CDR-Grafted Antibody Compositions
[0421]Anti-CCR4 human CDR-grafted antibody compositions produced by the transformants DG44/CHO and Ms705/CCR4 obtained in Example 2-1 were purified in the following manner.
[0422]Each transformant was suspended in IMDM-dFBS(10) containing 500 μg/ml G418 and 30 ml of the suspension was put into a 182-cm2 flask (manufactured by Greiner), followed by culturing in a 5% CO2 incubator at 37° C. for several days. When the cells became confluent, the culture supernatant was removed and the cells were washed with 25 ml of PBS, followed by addition of 30 ml of EXCELL301 Medium® (manufactured by JRH Biosciences). After culturing in a 5% CO2 incubator at 37° C. for 7 days, the cell suspension was recovered and subjected to centrifugation at 3000 rpm at 4° C. for 5 minutes to recover the supernatant. The supernatant was filtered through Millex GV Filter® (pore size: 0.22 μm, manufactured by Millipore) for sterilization. The anti-CCR4 human CDR-grafted antibody composition was purified from the culture supernatant thus obtained using Mab Select® (manufactured by Amersham Biosciences) column according to the attached instructions. The purified anti-CCR4 human CDR-grafted antibody compositions obtained from the DG44/CCR4 cell line and the Ms705/CCR4 cell line were designated DG44/CCR4 antibody and Ms705/CCR4 antibody, respectively. Also, the thus obtained Ms705/CCR4 cell line was deposited with International Patent Organism Depositary, National Institute of Advanced Industrial Science and Technology (Central 6, 1, Higashi 1-chome, Tsukuba-shi, Ibaraki, Japan) on Sep. 9, 2003 with accession No. FERM BP-8467.
EXAMPLE 3
Biological Activities of Anti-CCR4 Human CDR-Grafted Antibody Produced by FUT8 Gene-Double-Knockout Cell
1. Binding Activity of Anti-CCR4 Human CDR-Grafted Antibody to Human CCR4 Antigen (ELISA)
[0423]The binding activity of the DG44/CCR4 antibody and the Ms705/CCR4 antibody purified in Example 2-3 to human CCR4 antigen was measured in the following manner.
[0424]Compound 1 (SEQ ID NO:38) was selected as the peptide of the extracellular region of human CCR4 which can react with the anti-CCR4 human CDR-grafted antibody. In order to use Compound 1 in the activity measurement by ELISA, its conjugate with BSA (bovine serum albumin) (manufactured by Nacalai Tesque, Inc.) was prepared in the following manner and was used as the antigen. That is, 100 μl of 25 mg/ml SMCC [4-(N-maleimidomethyl)cyclohexane-1-carboxylic acid N-hydroxysuccinimide ester](manufactured by Sigma)-DMSO solution was added dropwise to 900 μl of PBS solution containing 10 mg of BSA under vortex, followed by gentle stirring for 30 minutes. After 1 ml of the reaction mixture was applied to a gel filtration column such as NAP-10 column equilibrated with 25 ml of PBS (manufactured by Invitrogen), the eluate eluted with 1.5 ml of PBS was used as a BSA-SMCC solution (BSA concentration was calculated by A280 measurement). To 0.5 mg of Compound 1 were added 250 μl of PBS and then 250 μl of DMF to completely dissolve Compound 1, and then the above BSA-SMCC solution (BSA amount: 1.25 mg) was added under vortex, followed by gentle stirring for 3 hours. The reaction mixture was dialyzed against PBS at 4° C. overnight and sodium azide was added thereto to give a final concentration of 0.05%. The resulting mixture was filtered through a 0.22 μm filter to obtain a BSA-Compound 1 solution.
[0425]The BSA-Compound 1 solution thus prepared in the above was diluted with PBS to 1 μg/ml and put into wells of a 96-well plate for ELISA (manufactured by Greiner) in an amount of 50 μl/well, followed by standing at 4° C. overnight for adsorption. After washing with PBS, 1% BSA-PBS was added to the wells in an amount of 100 μl/well, followed by reaction at room temperature for one hour to block the remaining active groups. After the 1% BSA-PBS was discarded and each well was washed with Tween-PBS, 50 μl each of variously diluted solutions of the DG44/CCR4 antibody or the Ms705/CCR4 antibody prepared in Example 2-3 were respectively added to the wells, followed by reaction at room temperature for 2 hours. After the reaction, the wells were washed with Tween-PBS. To each well was added 50 μl of peroxidase-labeled goat anti-human IgG (H & L) antibody solution diluted 2000-fold with 1% BSA-PBS as a secondary antibody solution, followed by reaction at room temperature for one hour. After the reaction, the wells were washed with Tween-PBS, and 50 μl of ABTS substrate solution was added to each well to develop color, followed by measurement of OD415.
[0426]FIG. 6 shows the binding activity of the DG44/CCR4 antibody and the Ms705/CCR4 antibody to the CCR4 extracellular region peptide. Each antibody had an equal binding activity to the CCR4 extracellular region peptide.
2. In Vitro Cytotoxic Activity (ADCC Activity) of Anti-CCR4 Human CDR-Grafted Antibody Composition
[0427]The in vitro cytotoxic activity of the DG44/CCR4 antibody and the Ms705/CCR4 antibody obtained in Example 2-3 was measured in the following manner.
(1) Preparation of a Target Cell Suspension
[0428]CCR4/EL4 cells which are mouse thymoma cell E4 cell line in which human CCR4 is highly expressed as described in WO01/64754 were washed with RPMI 1640-FCS(5) medium (RPMI 1640 medium (manufactured by GIBCO BRL) containing 5% FCS) by centrifugation and suspension and then adjusted to a density of 2×105 cells/ml by using RPMI 1640-FCS(5) medium and used as the target cell suspension.
(2) Preparation of an Effector Cell Suspension
[0429]Venous blood (50 ml) was collected from a healthy person and gently mixed with 0.5 ml of heparin sodium (manufactured by Shimizu Pharmaceutical Co., Ltd.). The monocyte layer was separated from this mixture using Lymphoprep® (manufactured by AXIS SHIELD) according to the attached instructions. After being washed three times with RPMI1640-FCS(5) medium through centrifugation, the cells were suspended in the same medium at a density of 5×106 cells/ml to give an effector cell suspension.
(3) Measurement of ADCC Activity
[0430]The target cell suspension prepared in the above (1) (50 μl) was put into each well of a 96-well U-shaped bottom plate (manufactured by Falcon) (1×104 cells/well). Then, 50 μl of the effector cell suspension prepared in (2) was added to each well (2.5×105 cells/well; the ratio of effector cells to target cells becomes 25:1). Subsequently, each of the various anti-CCR4 human CDR-grafted antibodies was added to give a final concentration of 0.1 to 1000 ng/ml and to make a total volume of 150 μl, followed by reaction at 37° C. for 4 hours. After the reaction, the plate was subjected to centrifugation, and the lactate dehydrogenase (LDH) activity of the supernatant was measured by obtaining absorbance data using CytoTox96 Non-Radioactive Cytotoxicity Assay® (manufactured by Promega) according to the attached instructions. The absorbance data for target cell spontaneous release were obtained by the same procedure as above using only the medium instead of the effector cell suspension and the antibody solution, and those for effector cell spontaneous release were obtained by the same procedure using only the medium instead of the target cell suspension and the antibody solution. The absorbance data for target cell total release were obtained by the same procedure as above using the medium instead of the antibody solution and the effector cell suspension, adding 15 μl of 9% Triton X-100 solution 45 minutes before the completion of the reaction, and measuring the LDH activity of the supernatant. The ADCC activity was calculated according to the following equation.
Cytotoxic activity ( % ) = ( Absorbance of sample ) - ( Absorbance for effector cell spontaneous release ) - ( Absorbance for target cell spontaneous release ) ( Absorbance for target cell total release ) - ( Absorbance for target cell spontaneous release ) × 100
[0431]FIG. 7 shows the cytotoxic activities of the DG44/CCR4 antibody and the Ms705/CCR4 antibody against the CCR4/EL4 cells. The Ms705/CCR4 antibody showed higher ADCC activity than the DG44/CCR4 antibody at any antibody concentration.
EXAMPLE 4
Analysis of Monosaccharide Composition of Anti-CCR4 Human CDR-Grafted Antibody Composition Produced by FUT8 Gene-Double-Knockout Cell
[0432]Analysis of the neutral sugar and amino sugar composition of the DG44/CCR4 antibody and the Ms705/CCR4 antibody purified in Example 2-3 was carried out in the following manner.
[0433]After the antibody was dried under reduced pressure using a centrifugal concentrator, a 2.0 to 4.0 M trifluoroacetic acid solution was added thereto and acid hydrolysis was carried out at 100° C. for 2 to 4 hours to release neutral sugars and amino sugars from the protein. The trifluoroacetic acid solution was removed with a centrifugal concentrator, and the sugars were redissolved in deionized water and subjected to analysis using a carbohydrate analysis system (DX-500® manufactured by Dionex). The analysis was carried out according to the elution program shown in Table 1 using CarboPac PA-1 Column® and CarboPac PA-1 guard Column® (manufactured by Dionex), a 10 to 20 mM solution of sodium hydroxide in deionized water as an eluting solution and a 500 mM solution of sodium hydroxide in deionized water as a washing solution.
TABLE-US-00001 TABLE 1 Elution program for neutral sugar and amino sugar composition analysis Time (min.) 0 35 35.1 45 45.1 58 Eluting solution (%) 100 100 0 0 100 100 Washing solution (%) 0 0 100 100 0 0
[0434]From the peak areas of neutral and amino sugar components in the obtained elution profile, the composition ratio of components (fucose, galactose and mannose) was calculated, regarding the value of N-acetylglucosamine as 4.
[0435]Table 2 shows the ratio of sugar chains having a structure in which fucose is not bound to the N-acetylglucosamine in the reducing end among the total complex type N-glycoside-linked sugar chains as calculated from the monosaccharide composition ratio of each antibody. In the DG44/CCR4 antibody, the ratio of sugar chains having a structure in which fucose is not bound was 8%. On the other hand, in the Ms705/CCR4 antibody, the peak of fucose was below the detection limit, whereby the ratio of sugar chains having a structure in which fucose is not bound was estimated to be close to 100%.
[0436]The above result indicates that fucose is not bound to the N-acetylglucosamine in the reducing end in complex type N-glycoside-linked sugar chains in the Ms705/CCR4 antibody.
TABLE-US-00002 TABLE 2 Ratio of sugar chains to which fucose is not bound in anti-CCR4 human CDR-grafted antibody compositions Ratio of sugar chains to which Antibody fucose is not bound DG44/CCR4 antibody 2% Ms705/CCR4 antibody 100%
EXAMPLE 5
Analysis of Biological Activity of Anti-CCR4 Human CDR-Grafted Antibody Composition Having Sugar Chains to which Fucose is not Bound
[0437]In order to further reveal superiority of the anti-CCR4 human CDR-grafted antibody composition of the present invention, biological activity of an antibody composition having sugar chains to which fucose is not bound was compared with that of an antibody composition in which an antibody molecule having sugar chains to which fucose is not bound was mixed with an antibody molecule having sugar chains to which fucose is bound. Specifically, changes in the cytotoxic activity were examined in antibody compositions in which the Ms705/CCR4 antibody which is an anti-CCR4 human CDR-grafted antibody having sugar chains to which fucose is not bound was mixed with an anti-CCR4 human CDR-grafted antibody having sugar chains to which fucose is bound.
(1) Preparation of Target Cell Suspension
[0438]The preparation was carried out according to the method described in Example 3-2(1).
(2) Preparation of Effector Cell Suspension
[0439]A monocyte layer was separated according to the method described in Example 3-2(2) and the monocytes were suspended by using RPMI 1640-FCS(5) medium to a density of 4×106 cells/ml to give the effector cell suspension.
(3) Measurement of ADCC Activity
[0440]The target cell suspension prepared in the above (1) was dispensed at 50 μl into each well of a 96-well U bottom plate (manufactured by Falcon) (1×104 cells/well). Next, the effector cell suspension prepared in the above (2) was added at 50 μl (2×105 cells/well, the ratio of effector cells to target cells becomes 20:1). Subsequently, the Ms705/CCR4 antibody and the DG44/CCR4 antibody were added independently or as a mixture of both of them, adjusted to a total volume of 150 μl and then allowed to react at 37° C. for 4 hours. After the reaction, the plate was centrifuged, and lactate dehydrogenase (LDH) activity in the supernatant was measured using LDH-Cytotoxic Test Wako® (manufactured by Wako Pure Chemical Industries) in accordance with the instructions attached thereto. The ADCC activity was calculated in accordance with the method described in Example 3-2.
[0441]An anti-CCR4 human CDR-grafted antibody composition in which the ratio of an antibody having sugar chains to which fucose is not bound was changed was prepared by adding DG44/CCR4 antibody to a predetermined amount of the Ms705/CCR4 antibody, and its ADCC activity was measured. Specifically, an anti-CCR4 human CDR-grafted antibody composition in which 0 to 300 ng/ml of the DG44/CCR4 antibody was added to 3.7 ng/ml of Ms705/CCR4 antibody was prepared. The ADCC activity of the thus prepared antibody composition is shown in FIG. 8.
[0442]When the Ms705/CCR4 antibody was further added to 3.7 ng/ml of the Ms705/CCR4 antibody, increase of the ADCC activity was observed with increase of the total antibody concentration, but when the DG44/CCR4 antibody was further added to 3.7 ng/ml of the Ms705/CCR4 antibody, the ADCC activity of the thus prepared antibody composition was reduced on the contrary regardless of the increased total antibody concentration. This result showed that an antibody molecule having a sugar chain to which fucose is bound inhibits ADCC activity of an antibody molecule having a sugar chain to which fucose is not bound. Also, in the case of antibody compositions in which an antibody molecule having sugar chains to which fucose is bound was mixed with an antibody molecule having sugar chains to which fucose is not bound, an antibody composition in which the ratio of the antibody molecule having sugar chains to which fucose is not bound was 20% or more showed markedly higher ADCC activity than an antibody composition in which said ratio was less than 20%. ADCC activities of an antibody sample of 10 ng/ml of the Ms705/CCR4 antibody and an antibody sample prepared by mixing 10 ng/ml of the Ms705/CCR4 antibody with a 9-fold amount, namely 90 ng/ml, of the DG44/CCR4 antibody are shown as a graph in FIG. 9. The ADCC activity of the Ms705/CCR4 antibody was sharply reduced by the addition of DG44/CCR4 antibody. Even when antibody concentration of the antibody composition was increased to 100 times or more while keeping the existing ratio of the Ms705/CCR4 antibody and the DG44/CCR4 antibody at 1/9, the ADCC activity was still fell short of that of the 10 ng/ml Ms705/CCR4 antibody sample. Based on the above, it was found that an antibody molecule having sugar chains to which fucose is bound inhibits the ADCC activity of an antibody molecule having sugar chains to which fucose is not bound, and that the conventional antibody compositions cannot exert its ADCC activity similar to that of the antibody composition of the present invention.
[0443]Accordingly, patients who were unable to be treated by the conventional antibody compositions can be treated by the antibody composition of the present invention. Since the antibody composition of the present invention has high activity of injuring CCR4-expressing cells, it is clinically useful as a pharmaceutical composition for the treatment of CCR4-related diseases such as cancer and inflammatory diseases.
[0444]While the invention has been described in detail and with reference to specific embodiments thereof, it will be apparent to one skill in the art that various changes and modifications can be made therein without departing from the spirit and scope thereof. All references cited herein are incorporated in their entirety.
[0445]This application is based on Japanese application No. 2003-350162 filed on Oct. 8, 2003 and U.S. provisional patent application No. 60/572,784 filed on May 21, 2004, the entire contents of which are incorporated hereinto by reference.
Sequence CWU
1
4611504DNACricetulus griseusCDS(1)..(1119) 1atg gct cac gct ccc gct agc
tgc ccg agc tcc agg aac tct ggg gac 48Met Ala His Ala Pro Ala Ser
Cys Pro Ser Ser Arg Asn Ser Gly Asp 1 5
10 15ggc gat aag ggc aag ccc agg aag gtg gcg ctc atc acg
ggc atc acc 96Gly Asp Lys Gly Lys Pro Arg Lys Val Ala Leu Ile Thr
Gly Ile Thr 20 25 30ggc cag
gat ggc tca tac ttg gca gaa ttc ctg ctg gag aaa gga tac 144Gly Gln
Asp Gly Ser Tyr Leu Ala Glu Phe Leu Leu Glu Lys Gly Tyr 35
40 45gag gtt cat gga att gta cgg cga tcc agt
tca ttt aat aca ggt cga 192Glu Val His Gly Ile Val Arg Arg Ser Ser
Ser Phe Asn Thr Gly Arg 50 55 60att
gaa cat tta tat aag aat cca cag gct cat att gaa gga aac atg 240Ile
Glu His Leu Tyr Lys Asn Pro Gln Ala His Ile Glu Gly Asn Met 65
70 75 80aag ttg cac tat ggt gac
ctc acc gac agc acc tgc cta gta aaa atc 288Lys Leu His Tyr Gly Asp
Leu Thr Asp Ser Thr Cys Leu Val Lys Ile 85
90 95atc aat gaa gtc aaa cct aca gag atc tac aat ctt
ggt gcc cag agc 336Ile Asn Glu Val Lys Pro Thr Glu Ile Tyr Asn Leu
Gly Ala Gln Ser 100 105 110cat
gtc aag att tcc ttt gac tta gca gag tac act gca gat gtt gat 384His
Val Lys Ile Ser Phe Asp Leu Ala Glu Tyr Thr Ala Asp Val Asp 115
120 125gga gtt ggc acc ttg cgg ctt ctg gat
gca att aag act tgt ggc ctt 432Gly Val Gly Thr Leu Arg Leu Leu Asp
Ala Ile Lys Thr Cys Gly Leu 130 135
140ata aat tct gtg aag ttc tac cag gcc tca act agt gaa ctg tat gga
480Ile Asn Ser Val Lys Phe Tyr Gln Ala Ser Thr Ser Glu Leu Tyr Gly145
150 155 160aaa gtg caa gaa
ata ccc cag aaa gag acc acc cct ttc tat cca agg 528Lys Val Gln Glu
Ile Pro Gln Lys Glu Thr Thr Pro Phe Tyr Pro Arg 165
170 175tcg ccc tat gga gca gcc aaa ctt tat gcc
tat tgg att gta gtg aac 576Ser Pro Tyr Gly Ala Ala Lys Leu Tyr Ala
Tyr Trp Ile Val Val Asn 180 185
190ttt cga gag gct tat aat ctc ttt gcg gtg aac ggc att ctc ttc aat
624Phe Arg Glu Ala Tyr Asn Leu Phe Ala Val Asn Gly Ile Leu Phe Asn
195 200 205cat gag agt cct aga aga gga
gct aat ttt gtt act cga aaa att agc 672His Glu Ser Pro Arg Arg Gly
Ala Asn Phe Val Thr Arg Lys Ile Ser 210 215
220cgg tca gta gct aag att tac ctt gga caa ctg gaa tgt ttc agt ttg
720Arg Ser Val Ala Lys Ile Tyr Leu Gly Gln Leu Glu Cys Phe Ser Leu225
230 235 240gga aat ctg gac
gcc aaa cga gac tgg ggc cat gcc aag gac tat gtc 768Gly Asn Leu Asp
Ala Lys Arg Asp Trp Gly His Ala Lys Asp Tyr Val 245
250 255gag gct atg tgg ctg atg tta caa aat gat
gaa cca gag gac ttt gtc 816Glu Ala Met Trp Leu Met Leu Gln Asn Asp
Glu Pro Glu Asp Phe Val 260 265
270ata gct act ggg gaa gtt cat agt gtc cgt gaa ttt gtt gag aaa tca
864Ile Ala Thr Gly Glu Val His Ser Val Arg Glu Phe Val Glu Lys Ser
275 280 285ttc atg cac att gga aag acc
att gtg tgg gaa gga aag aat gaa aat 912Phe Met His Ile Gly Lys Thr
Ile Val Trp Glu Gly Lys Asn Glu Asn 290 295
300gaa gtg ggc aga tgt aaa gag acc ggc aaa att cat gtg act gtg gat
960Glu Val Gly Arg Cys Lys Glu Thr Gly Lys Ile His Val Thr Val Asp305
310 315 320ctg aaa tac tac
cga cca act gaa gtg gac ttc ctg cag gga gac tgc 1008Leu Lys Tyr Tyr
Arg Pro Thr Glu Val Asp Phe Leu Gln Gly Asp Cys 325
330 335tcc aag gcg cag cag aaa ctg aac tgg aag
ccc cgc gtt gcc ttt gac 1056Ser Lys Ala Gln Gln Lys Leu Asn Trp Lys
Pro Arg Val Ala Phe Asp 340 345
350gag ctg gtg agg gag atg gtg caa gcc gat gtg gag ctc atg aga acc
1104Glu Leu Val Arg Glu Met Val Gln Ala Asp Val Glu Leu Met Arg Thr
355 360 365aac ccc aac gcc tga
gcacctctac aaaaaaattc gcgagacatg gactatggtg 1159Asn Pro Asn Ala
370cagagccagc caaccagagt ccagccactc ctgagaccat cgaccataaa ccctcgactg
1219cctgtgtcgt ccccacagct aagagctggg ccacaggttt gtgggcacca ggacggggac
1279actccagagc taaggccact tcgcttttgt caaaggctcc tctcaatgat tttgggaaat
1339caagaagttt aaaatcacat actcatttta cttgaaatta tgtcactaga caacttaaat
1399ttttgagtct tgagattgtt tttctctttt cttattaaat gatctttcta tgacccagca
1459aaaaaaaaaa aaaaaaggga tataaaaaaa aaaaaaaaaa aaaaa
15042372PRTCricetulus griseus 2Met Ala His Ala Pro Ala Ser Cys Pro Ser
Ser Arg Asn Ser Gly Asp 1 5 10
15Gly Asp Lys Gly Lys Pro Arg Lys Val Ala Leu Ile Thr Gly Ile Thr
20 25 30Gly Gln Asp Gly Ser
Tyr Leu Ala Glu Phe Leu Leu Glu Lys Gly Tyr 35
40 45Glu Val His Gly Ile Val Arg Arg Ser Ser Ser Phe Asn
Thr Gly Arg 50 55 60Ile Glu His Leu
Tyr Lys Asn Pro Gln Ala His Ile Glu Gly Asn Met 65 70
75 80Lys Leu His Tyr Gly Asp Leu Thr Asp
Ser Thr Cys Leu Val Lys Ile 85 90
95Ile Asn Glu Val Lys Pro Thr Glu Ile Tyr Asn Leu Gly Ala Gln
Ser 100 105 110His Val Lys Ile
Ser Phe Asp Leu Ala Glu Tyr Thr Ala Asp Val Asp 115
120 125Gly Val Gly Thr Leu Arg Leu Leu Asp Ala Ile Lys
Thr Cys Gly Leu 130 135 140Ile Asn Ser
Val Lys Phe Tyr Gln Ala Ser Thr Ser Glu Leu Tyr Gly145
150 155 160Lys Val Gln Glu Ile Pro Gln
Lys Glu Thr Thr Pro Phe Tyr Pro Arg 165
170 175Ser Pro Tyr Gly Ala Ala Lys Leu Tyr Ala Tyr Trp
Ile Val Val Asn 180 185 190Phe
Arg Glu Ala Tyr Asn Leu Phe Ala Val Asn Gly Ile Leu Phe Asn 195
200 205His Glu Ser Pro Arg Arg Gly Ala Asn
Phe Val Thr Arg Lys Ile Ser 210 215
220Arg Ser Val Ala Lys Ile Tyr Leu Gly Gln Leu Glu Cys Phe Ser Leu225
230 235 240Gly Asn Leu Asp
Ala Lys Arg Asp Trp Gly His Ala Lys Asp Tyr Val 245
250 255Glu Ala Met Trp Leu Met Leu Gln Asn Asp
Glu Pro Glu Asp Phe Val 260 265
270Ile Ala Thr Gly Glu Val His Ser Val Arg Glu Phe Val Glu Lys Ser
275 280 285Phe Met His Ile Gly Lys Thr
Ile Val Trp Glu Gly Lys Asn Glu Asn 290 295
300Glu Val Gly Arg Cys Lys Glu Thr Gly Lys Ile His Val Thr Val
Asp305 310 315 320Leu Lys
Tyr Tyr Arg Pro Thr Glu Val Asp Phe Leu Gln Gly Asp Cys
325 330 335Ser Lys Ala Gln Gln Lys Leu
Asn Trp Lys Pro Arg Val Ala Phe Asp 340 345
350Glu Leu Val Arg Glu Met Val Gln Ala Asp Val Glu Leu Met
Arg Thr 355 360 365Asn Pro Asn Ala
37031316DNACricetulus griseus 3gccccgcccc ctccacctgg accgagagta
gctggagaat tgtgcaccgg aagtagctct 60tggactggtg gaaccctgcg caggtgcagc
aacaatgggt gagccccagg gatccaggag 120gatcctagtg acagggggct ctggactggt
gggcagagct atccagaagg tggtcgcaga 180tggcgctggc ttacccggag aggaatgggt
gtttgtctcc tccaaagatg cagatctgac 240ggatgcagca caaacccaag ccctgttcca
gaaggtacag cccacccatg tcatccatct 300tgctgcaatg gtaggaggcc ttttccggaa
tatcaaatac aacttggatt tctggaggaa 360gaatgtgcac atcaatgaca acgtcctgca
ctcagctttc gaggtgggca ctcgcaaggt 420ggtctcctgc ctgtccacct gtatcttccc
tgacaagacc acctatccta ttgatgaaac 480aatgatccac aatggtccac cccacagcag
caattttggg tactcgtatg ccaagaggat 540gattgacgtg cagaacaggg cctacttcca
gcagcatggc tgcaccttca ctgctgtcat 600ccctaccaat gtctttggac ctcatgacaa
cttcaacatt gaagatggcc atgtgctgcc 660tggcctcatc cataaggtgc atctggccaa
gagtaatggt tcagccttga ctgtttgggg 720tacagggaaa ccacggaggc agttcatcta
ctcactggac ctagcccggc tcttcatctg 780ggtcctgcgg gagtacaatg aagttgagcc
catcatcctc tcagtgggcg aggaagatga 840agtctccatt aaggaggcag ctgaggctgt
agtggaggcc atggacttct gtggggaagt 900cacttttgat tcaacaaagt cagatgggca
gtataagaag acagccagca atggcaagct 960tcgggcctac ttgcctgatt tccgtttcac
acccttcaag caggctgtga aggagacctg 1020tgcctggttc accgacaact atgagcaggc
ccggaagtga agcatgggac aagcgggtgc 1080tcagctggca atgcccagtc agtaggctgc
agtctcatca tttgcttgtc aagaactgag 1140gacagtatcc agcaacctga gccacatgct
ggtctctctg ccagggggct tcatgcagcc 1200atccagtagg gcccatgttt gtccatcctc
gggggaaggc cagaccaaca ccttgtttgt 1260ctgcttctgc cccaacctca gtgcatccat
gctggtcctg ctgtcccttg tctaga 13164321PRTCricetulus griseus 4Met Gly
Glu Pro Gln Gly Ser Arg Arg Ile Leu Val Thr Gly Gly Ser 1 5
10 15Gly Leu Val Gly Arg Ala Ile Gln
Lys Val Val Ala Asp Gly Ala Gly 20 25
30Leu Pro Gly Glu Glu Trp Val Phe Val Ser Ser Lys Asp Ala Asp
Leu 35 40 45Thr Asp Ala Ala Gln
Thr Gln Ala Leu Phe Gln Lys Val Gln Pro Thr 50 55
60His Val Ile His Leu Ala Ala Met Val Gly Gly Leu Phe Arg
Asn Ile 65 70 75 80Lys
Tyr Asn Leu Asp Phe Trp Arg Lys Asn Val His Ile Asn Asp Asn
85 90 95Val Leu His Ser Ala Phe Glu
Val Gly Thr Arg Lys Val Val Ser Cys 100 105
110Leu Ser Thr Cys Ile Phe Pro Asp Lys Thr Thr Tyr Pro Ile
Asp Glu 115 120 125Thr Met Ile His
Asn Gly Pro Pro His Ser Ser Asn Phe Gly Tyr Ser 130
135 140Tyr Ala Lys Arg Met Ile Asp Val Gln Asn Arg Ala
Tyr Phe Gln Gln145 150 155
160His Gly Cys Thr Phe Thr Ala Val Ile Pro Thr Asn Val Phe Gly Pro
165 170 175His Asp Asn Phe Asn
Ile Glu Asp Gly His Val Leu Pro Gly Leu Ile 180
185 190His Lys Val His Leu Ala Lys Ser Asn Gly Ser Ala
Leu Thr Val Trp 195 200 205Gly Thr
Gly Lys Pro Arg Arg Gln Phe Ile Tyr Ser Leu Asp Leu Ala 210
215 220Arg Leu Phe Ile Trp Val Leu Arg Glu Tyr Asn
Glu Val Glu Pro Ile225 230 235
240Ile Leu Ser Val Gly Glu Glu Asp Glu Val Ser Ile Lys Glu Ala Ala
245 250 255Glu Ala Val Val
Glu Ala Met Asp Phe Cys Gly Glu Val Thr Phe Asp 260
265 270Ser Thr Lys Ser Asp Gly Gln Tyr Lys Lys Thr
Ala Ser Asn Gly Lys 275 280 285Leu
Arg Ala Tyr Leu Pro Asp Phe Arg Phe Thr Pro Phe Lys Gln Ala 290
295 300Val Lys Glu Thr Cys Ala Trp Phe Thr Asp
Asn Tyr Glu Gln Ala Arg305 310 315
320Lys52008DNACricetulus griseus 5aacagaaact tattttcctg
tgtggctaac tagaaccaga gtacaatgtt tccaattctt 60tgagctccga gaagacagaa
gggagttgaa actctgaaaa tgcgggcatg gactggttcc 120tggcgttgga ttatgctcat
tctttttgcc tgggggacct tattgtttta tataggtggt 180catttggttc gagataatga
ccaccctgac cattctagca gagaactctc caagattctt 240gcaaagctgg agcgcttaaa
acaacaaaat gaagacttga ggagaatggc tgagtctctc 300cgaataccag aaggccctat
tgatcagggg acagctacag gaagagtccg tgttttagaa 360gaacagcttg ttaaggccaa
agaacagatt gaaaattaca agaaacaagc taggaatgat 420ctgggaaagg atcatgaaat
cttaaggagg aggattgaaa atggagctaa agagctctgg 480ttttttctac aaagtgaatt
gaagaaatta aagaaattag aaggaaacga actccaaaga 540catgcagatg aaattctttt
ggatttagga catcatgaaa ggtctatcat gacagatcta 600tactacctca gtcaaacaga
tggagcaggt gagtggcggg aaaaagaagc caaagatctg 660acagagctgg tccagcggag
aataacatat ctgcagaatc ccaaggactg cagcaaagcc 720agaaagctgg tatgtaatat
caacaaaggc tgtggctatg gatgtcaact ccatcatgtg 780gtttactgct tcatgattgc
ttatggcacc cagcgaacac tcatcttgga atctcagaat 840tggcgctatg ctactggagg
atgggagact gtgtttagac ctgtaagtga gacatgcaca 900gacaggtctg gcctctccac
tggacactgg tcaggtgaag tgaaggacaa aaatgttcaa 960gtggtcgagc tccccattgt
agacagcctc catcctcgtc ctccttactt acccttggct 1020gtaccagaag accttgcaga
tcgactcctg agagtccatg gtgatcctgc agtgtggtgg 1080gtatcccagt ttgtcaaata
cttgatccgt ccacaacctt ggctggaaag ggaaatagaa 1140gaaaccacca agaagcttgg
cttcaaacat ccagttattg gagtccatgt cagacgcact 1200gacaaagtgg gaacagaagc
agccttccat cccattgagg aatacatggt acacgttgaa 1260gaacattttc agcttctcga
acgcagaatg aaagtggata aaaaaagagt gtatctggcc 1320actgatgacc cttctttgtt
aaaggaggca aagacaaagt actccaatta tgaatttatt 1380agtgataact ctatttcttg
gtcagctgga ctacacaacc gatacacaga aaattcactt 1440cggggcgtga tcctggatat
acactttctc tcccaggctg acttccttgt gtgtactttt 1500tcatcccagg tctgtagggt
tgcttatgaa atcatgcaaa cactgcatcc tgatgcctct 1560gcaaacttcc attctttaga
tgacatctac tattttggag gccaaaatgc ccacaaccag 1620attgcagttt atcctcacca
acctcgaact aaagaggaaa tccccatgga acctggagat 1680atcattggtg tggctggaaa
ccattggaat ggttactcta aaggtgtcaa cagaaaacta 1740ggaaaaacag gcctgtaccc
ttcctacaaa gtccgagaga agatagaaac agtcaaatac 1800cctacatatc ctgaagctga
aaaatagaga tggagtgtaa gagattaaca acagaattta 1860gttcagacca tctcagccaa
gcagaagacc cagactaaca tatggttcat tgacagacat 1920gctccgcacc aagagcaagt
gggaaccctc agatgctgca ctggtggaac gcctctttgt 1980gaagggctgc tgtgccctca
agcccatg 200861728DNAMus musculus
6atgcgggcat ggactggttc ctggcgttgg attatgctca ttctttttgc ctgggggacc
60ttgttatttt atataggtgg tcatttggtt cgagataatg accaccctga tcactccagc
120agagaactct ccaagattct tgcaaagctt gaacgcttaa aacagcaaaa tgaagacttg
180aggcgaatgg ctgagtctct ccgaatacca gaaggcccca ttgaccaggg gacagctaca
240ggaagagtcc gtgttttaga agaacagctt gttaaggcca aagaacagat tgaaaattac
300aagaaacaag ctagaaatgg tctggggaag gatcatgaaa tcttaagaag gaggattgaa
360aatggagcta aagagctctg gttttttcta caaagcgaac tgaagaaatt aaagcattta
420gaaggaaatg aactccaaag acatgcagat gaaattcttt tggatttagg acaccatgaa
480aggtctatca tgacagatct atactacctc agtcaaacag atggagcagg ggattggcgt
540gaaaaagagg ccaaagatct gacagagctg gtccagcgga gaataacata tctccagaat
600cctaaggact gcagcaaagc caggaagctg gtgtgtaaca tcaataaagg ctgtggctat
660ggttgtcaac tccatcacgt ggtctactgt ttcatgattg cttatggcac ccagcgaaca
720ctcatcttgg aatctcagaa ttggcgctat gctactggtg gatgggagac tgtgtttaga
780cctgtaagtg agacatgtac agacagatct ggcctctcca ctggacactg gtcaggtgaa
840gtaaatgaca aaaacattca agtggtcgag ctccccattg tagacagcct ccatcctcgg
900cctccttact taccactggc tgttccagaa gaccttgcag accgactcct aagagtccat
960ggtgaccctg cagtgtggtg ggtgtcccag tttgtcaaat acttgattcg tccacaacct
1020tggctggaaa aggaaataga agaagccacc aagaagcttg gcttcaaaca tccagttatt
1080ggagtccatg tcagacgcac agacaaagtg ggaacagaag cagccttcca ccccatcgag
1140gagtacatgg tacacgttga agaacatttt cagcttctcg cacgcagaat gcaagtggat
1200aaaaaaagag tatatctggc tactgatgat cctactttgt taaaggaggc aaagacaaag
1260tactccaatt atgaatttat tagtgataac tctatttctt ggtcagctgg actacacaat
1320cggtacacag aaaattcact tcggggtgtg atcctggata tacactttct ctcacaggct
1380gactttctag tgtgtacttt ttcatcccag gtctgtcggg ttgcttatga aatcatgcaa
1440accctgcatc ctgatgcctc tgcgaacttc cattctttgg atgacatcta ctattttgga
1500ggccaaaatg cccacaatca gattgctgtt tatcctcaca aacctcgaac tgaagaggaa
1560attccaatgg aacctggaga tatcattggt gtggctggaa accattggga tggttattct
1620aaaggtatca acagaaaact tggaaaaaca ggcttatatc cctcctacaa agtccgagag
1680aagatagaaa cagtcaagta tcccacatat cctgaagctg aaaaatag
17287575PRTCricetulus griseus 7Met Arg Ala Trp Thr Gly Ser Trp Arg Trp
Ile Met Leu Ile Leu Phe 1 5 10
15Ala Trp Gly Thr Leu Leu Phe Tyr Ile Gly Gly His Leu Val Arg Asp
20 25 30Asn Asp His Pro Asp
His Ser Ser Arg Glu Leu Ser Lys Ile Leu Ala 35
40 45Lys Leu Glu Arg Leu Lys Gln Gln Asn Glu Asp Leu Arg
Arg Met Ala 50 55 60Glu Ser Leu Arg
Ile Pro Glu Gly Pro Ile Asp Gln Gly Thr Ala Thr 65 70
75 80Gly Arg Val Arg Val Leu Glu Glu Gln
Leu Val Lys Ala Lys Glu Gln 85 90
95Ile Glu Asn Tyr Lys Lys Gln Ala Arg Asn Asp Leu Gly Lys Asp
His 100 105 110Glu Ile Leu Arg
Arg Arg Ile Glu Asn Gly Ala Lys Glu Leu Trp Phe 115
120 125Phe Leu Gln Ser Glu Leu Lys Lys Leu Lys Lys Leu
Glu Gly Asn Glu 130 135 140Leu Gln Arg
His Ala Asp Glu Ile Leu Leu Asp Leu Gly His His Glu145
150 155 160Arg Ser Ile Met Thr Asp Leu
Tyr Tyr Leu Ser Gln Thr Asp Gly Ala 165
170 175Gly Glu Trp Arg Glu Lys Glu Ala Lys Asp Leu Thr
Glu Leu Val Gln 180 185 190Arg
Arg Ile Thr Tyr Leu Gln Asn Pro Lys Asp Cys Ser Lys Ala Arg 195
200 205Lys Leu Val Cys Asn Ile Asn Lys Gly
Cys Gly Tyr Gly Cys Gln Leu 210 215
220His His Val Val Tyr Cys Phe Met Ile Ala Tyr Gly Thr Gln Arg Thr225
230 235 240Leu Ile Leu Glu
Ser Gln Asn Trp Arg Tyr Ala Thr Gly Gly Trp Glu 245
250 255Thr Val Phe Arg Pro Val Ser Glu Thr Cys
Thr Asp Arg Ser Gly Leu 260 265
270Ser Thr Gly His Trp Ser Gly Glu Val Lys Asp Lys Asn Val Gln Val
275 280 285Val Glu Leu Pro Ile Val Asp
Ser Leu His Pro Arg Pro Pro Tyr Leu 290 295
300Pro Leu Ala Val Pro Glu Asp Leu Ala Asp Arg Leu Leu Arg Val
His305 310 315 320Gly Asp
Pro Ala Val Trp Trp Val Ser Gln Phe Val Lys Tyr Leu Ile
325 330 335Arg Pro Gln Pro Trp Leu Glu
Arg Glu Ile Glu Glu Thr Thr Lys Lys 340 345
350Leu Gly Phe Lys His Pro Val Ile Gly Val His Val Arg Arg
Thr Asp 355 360 365Lys Val Gly Thr
Glu Ala Ala Phe His Pro Ile Glu Glu Tyr Met Val 370
375 380His Val Glu Glu His Phe Gln Leu Leu Glu Arg Arg
Met Lys Val Asp385 390 395
400Lys Lys Arg Val Tyr Leu Ala Thr Asp Asp Pro Ser Leu Leu Lys Glu
405 410 415Ala Lys Thr Lys Tyr
Ser Asn Tyr Glu Phe Ile Ser Asp Asn Ser Ile 420
425 430Ser Trp Ser Ala Gly Leu His Asn Arg Tyr Thr Glu
Asn Ser Leu Arg 435 440 445Gly Val
Ile Leu Asp Ile His Phe Leu Ser Gln Ala Asp Phe Leu Val 450
455 460Cys Thr Phe Ser Ser Gln Val Cys Arg Val Ala
Tyr Glu Ile Met Gln465 470 475
480Thr Leu His Pro Asp Ala Ser Ala Asn Phe His Ser Leu Asp Asp Ile
485 490 495Tyr Tyr Phe Gly
Gly Gln Asn Ala His Asn Gln Ile Ala Val Tyr Pro 500
505 510His Gln Pro Arg Thr Lys Glu Glu Ile Pro Met
Glu Pro Gly Asp Ile 515 520 525Ile
Gly Val Ala Gly Asn His Trp Asn Gly Tyr Ser Lys Gly Val Asn 530
535 540Arg Lys Leu Gly Lys Thr Gly Leu Tyr Pro
Ser Tyr Lys Val Arg Glu545 550 555
560Lys Ile Glu Thr Val Lys Tyr Pro Thr Tyr Pro Glu Ala Glu Lys
565 570 5758575PRTMus
musculus 8Met Arg Ala Trp Thr Gly Ser Trp Arg Trp Ile Met Leu Ile Leu Phe
1 5 10 15Ala Trp Gly Thr
Leu Leu Phe Tyr Ile Gly Gly His Leu Val Arg Asp 20
25 30Asn Asp His Pro Asp His Ser Ser Arg Glu Leu
Ser Lys Ile Leu Ala 35 40 45Lys
Leu Glu Arg Leu Lys Gln Gln Asn Glu Asp Leu Arg Arg Met Ala 50
55 60Glu Ser Leu Arg Ile Pro Glu Gly Pro Ile
Asp Gln Gly Thr Ala Thr 65 70 75
80Gly Arg Val Arg Val Leu Glu Glu Gln Leu Val Lys Ala Lys Glu
Gln 85 90 95Ile Glu Asn
Tyr Lys Lys Gln Ala Arg Asn Gly Leu Gly Lys Asp His 100
105 110Glu Ile Leu Arg Arg Arg Ile Glu Asn Gly
Ala Lys Glu Leu Trp Phe 115 120
125Phe Leu Gln Ser Glu Leu Lys Lys Leu Lys His Leu Glu Gly Asn Glu 130
135 140Leu Gln Arg His Ala Asp Glu Ile
Leu Leu Asp Leu Gly His His Glu145 150
155 160Arg Ser Ile Met Thr Asp Leu Tyr Tyr Leu Ser Gln
Thr Asp Gly Ala 165 170
175Gly Asp Trp Arg Glu Lys Glu Ala Lys Asp Leu Thr Glu Leu Val Gln
180 185 190Arg Arg Ile Thr Tyr Leu
Gln Asn Pro Lys Asp Cys Ser Lys Ala Arg 195 200
205Lys Leu Val Cys Asn Ile Asn Lys Gly Cys Gly Tyr Gly Cys
Gln Leu 210 215 220His His Val Val Tyr
Cys Phe Met Ile Ala Tyr Gly Thr Gln Arg Thr225 230
235 240Leu Ile Leu Glu Ser Gln Asn Trp Arg Tyr
Ala Thr Gly Gly Trp Glu 245 250
255Thr Val Phe Arg Pro Val Ser Glu Thr Cys Thr Asp Arg Ser Gly Leu
260 265 270Ser Thr Gly His Trp
Ser Gly Glu Val Asn Asp Lys Asn Ile Gln Val 275
280 285Val Glu Leu Pro Ile Val Asp Ser Leu His Pro Arg
Pro Pro Tyr Leu 290 295 300Pro Leu Ala
Val Pro Glu Asp Leu Ala Asp Arg Leu Leu Arg Val His305
310 315 320Gly Asp Pro Ala Val Trp Trp
Val Ser Gln Phe Val Lys Tyr Leu Ile 325
330 335Arg Pro Gln Pro Trp Leu Glu Lys Glu Ile Glu Glu
Ala Thr Lys Lys 340 345 350Leu
Gly Phe Lys His Pro Val Ile Gly Val His Val Arg Arg Thr Asp 355
360 365Lys Val Gly Thr Glu Ala Ala Phe His
Pro Ile Glu Glu Tyr Met Val 370 375
380His Val Glu Glu His Phe Gln Leu Leu Ala Arg Arg Met Gln Val Asp385
390 395 400Lys Lys Arg Val
Tyr Leu Ala Thr Asp Asp Pro Thr Leu Leu Lys Glu 405
410 415Ala Lys Thr Lys Tyr Ser Asn Tyr Glu Phe
Ile Ser Asp Asn Ser Ile 420 425
430Ser Trp Ser Ala Gly Leu His Asn Arg Tyr Thr Glu Asn Ser Leu Arg
435 440 445Gly Val Ile Leu Asp Ile His
Phe Leu Ser Gln Ala Asp Phe Leu Val 450 455
460Cys Thr Phe Ser Ser Gln Val Cys Arg Val Ala Tyr Glu Ile Met
Gln465 470 475 480Thr Leu
His Pro Asp Ala Ser Ala Asn Phe His Ser Leu Asp Asp Ile
485 490 495Tyr Tyr Phe Gly Gly Gln Asn
Ala His Asn Gln Ile Ala Val Tyr Pro 500 505
510His Lys Pro Arg Thr Glu Glu Glu Ile Pro Met Glu Pro Gly
Asp Ile 515 520 525Ile Gly Val Ala
Gly Asn His Trp Asp Gly Tyr Ser Lys Gly Ile Asn 530
535 540Arg Lys Leu Gly Lys Thr Gly Leu Tyr Pro Ser Tyr
Lys Val Arg Glu545 550 555
560Lys Ile Glu Thr Val Lys Tyr Pro Thr Tyr Pro Glu Ala Glu Lys
565 570 5759383DNACricetulus griseus
9gttaactggg gctcttttaa accctgaatt tttctaaatc cccacctcca agagtttggt
60ttaaactgat ttttttaatg aatacctttt gaagaataga gcattgtctc atcatgcaaa
120gcttctcagg gattcagcta gcatgttgaa gaaacataag ggtgttaaat tgtttgtcac
180aagtgctgaa taaatattga cgtagtcttc agctattcta tactggaagt agatgatatt
240ctcattggaa attctgttag gaagtaaccc ttcttgtctt cttacctgca tagaatccca
300ggatataaaa cttgtgcttg tcgcccttgc cattgtctct cactggtggc ctttattgca
360tctcatatct gccttctctt tcc
38310564DNACricetulus griseus 10taagaattcc tgtgcccagc tgtatgtgag
gctctctgca ggtgtaggga tgtttctgct 60ttctttctgc acatgcttca cagctgaagt
cctttgggtg tgagattgac attcagatag 120actaaagtga ctggacttgt tgggaaacat
actgtatgca ttattgccgt tgcctccagg 180tgaaattaac acctcattca ccaatccctg
ttcatccaaa ctttctaccc acatcacttt 240aaatagaaat tagacccaat atgactcctt
ttttcctaag ctgtttatag agattgtgct 300ggagcagtga gcttttgtgt ttgtttgttt
gttttgtaat tttccccatg aaaatttctc 360taaactcaaa cctaagaggg aaaaaaaaaa
aacagactta tatgtgccac acttgtaaaa 420aaaaatcatg aaagatgtat atgatatttt
taaacagttt gaatattaag atcacaattt 480ctattttaaa aacaatcttg ttttacatat
caatcaccca attcccttgc cttcccatcc 540tcccattccc cccactgatc cccc
56411120DNACricetulus griseus
11atgaatgttc attctttggg tatatgccca agagtagaat tgctaaatat tgaggtagac
60tgattcccat tttcttgagg agtcgccata ttgatttcca aagtgactgt acaagttaac
12012274DNACricetulus griseus 12aggcactagg taaatatttt tgaagaaaga
atgagtatct cctatttcag aaaaactttt 60attgacttaa atttaggata tcagaattag
aaaacagtaa aaatttatag gagagttttt 120aatgaatgtt attttaaggt tccatacaaa
tagtaattaa aacttacaca aactatttgt 180agtaatgatt cagtctggta taccctgatg
agcattatac acttttaaat tctttttgta 240aattttttta ttagttcaaa ttaggaacaa
gctt 274139196DNACricetulus griseus
13tctagaccag gctggtctcg aactcacaga gaaccacctg cctctgccac ctgagtgctg
60ggattaaagg tgtgcaccac caccgcccgg cgtaaaatca tatttttgaa tattgtgata
120atttacatta taattgtaag taaaaatttt cagcctattt tgttatacat ttttgcgtaa
180attattcttt tttgaaagtt ttgttgtcca taatagtcta gggaaacata aagttataat
240ttttgtctat gtatttgcat atatatctat ttaatctcct aatgtccagg aaataaatag
300ggtatgtaat agcttcaaca tgtggtatga tagaattttt cagtgctata taagttgtta
360cagcaaagtg ttattaattc atatgtccat atttcaattt tttatgaatt attaaattga
420atccttaagc tgccagaact agaattttat tttaatcagg aagccccaaa tctgttcatt
480ctttctatat atgtggaaag gtaggcctca ctaactgatt cttcacctgt tttagaacat
540ggtccaagaa tggagttatg taaggggaat tacaagtgtg agaaaactcc tagaaaacaa
600gatgagtctt gtgaccttag tttctttaaa aacacaaaat tcttggaatg tgttttcatg
660ttcctcccag gtggatagga gtgagtttat ttcagattat ttattacaac tggctgttgt
720tacttgtttc tatgtcttta tagaaaaaca tatttttttt gccacatgca gcttgtcctt
780atgattttat acttgtgtga ctcttaactc tcagagtata aattgtctga tgctatgaat
840aaagttggct attgtatgag acttcagccc acttcaatta ttggcttcat tctctcagat
900cccaccacct ccagagtggt aaacaacttg aaccattaaa cagactttag tctttatttg
960aatgatagat ggggatatca gatttatagg cacagggttt tgagaaaggg agaaggtaaa
1020cagtagagtt taacaacaac aaaaagtata ctttgtaaac gtaaaactat ttattaaagt
1080agtagacaag acattaaata ttccttggga ttagtgcttt ttgaattttg ctttcaaata
1140atagtcagtg agtatacccc tcccccattc tatattttag cagaaatcag aataaatggt
1200gtttctggta cattcttttg tagagaattt attttctttg ggtttttgtg catttaaagt
1260caataaaaat taaggttcag taatagaaaa aaaactctga tttttggaat cccctttctt
1320cagcttttct atttaatctc ttaatgataa tttaatttgt ggccatgtgg tcaaagtata
1380tagccttgta tatgtaaatg ttttaaccaa cctgccttta cagtaactat ataattttat
1440tctataatat atgacttttc ttccatagct ttagagttgc ccagtcactt taagttacat
1500tttcatatat gttctttgtg ggaggagata attttatttc taagagaatc ctaagcatac
1560tgattgagaa atggcaaaca aaacacataa ttaaagctga taaagaacga acatttggag
1620tttaaaatac atagccaccc taagggttta actgttgtta gccttctttt ggaattttta
1680ttagttcata tagaaaaatg gattttatcg tgacatttcc atatatgtat ataatatatt
1740tacatcatat ccacctgtaa ttattagtgt ttttaaatat atttgaaaaa ataatggtct
1800ggtttgatcc atttgaacct tttgatgttt ggtgtggttg ccaattggtt gatggttatg
1860ataacctttg cttctctaag gttcaagtca gtttgagaat atgtcctcta aaaatgacag
1920gttgcaagtt aagtagtgag atgacagcga gatggagtga tgagaatttg tagaaatgaa
1980ttcacttata ctgagaactt gttttgcttt tagataatga acatattagc ctgaagtaca
2040tagccgaatt gattaattat tcaaagatat aatcttttaa tccctataaa agaggtatta
2100cacaacaatt caagaaagat agaattagac ttccagtatt ggagtgaacc atttgttatc
2160aggtagaacc ctaacgtgtg tggttgactt aaagtgttta ctttttacct gatactgggt
2220agctaattgt ctttcagcct cctggccaaa gataccatga aagtcaactt acgttgtatt
2280ctatatctca aacaactcag ggtgtttctt actctttcca cagcatgtag agcccaggaa
2340gcacaggaca agaaagctgc ctccttgtat caccaggaag atctttttgt aagagtcatc
2400acagtatacc agagagacta attttgtctg aagcatcatg tgttgaaaca acagaaactt
2460attttcctgt gtggctaact agaaccagag tacaatgttt ccaattcttt gagctccgag
2520aagacagaag ggagttgaaa ctctgaaaat gcgggcatgg actggttcct ggcgttggat
2580tatgctcatt ctttttgcct gggggacctt attgttttat ataggtggtc atttggttcg
2640agataatgac caccctgacc attctagcag agaactctcc aagattcttg caaagctgga
2700gcgcttaaaa caacaaaatg aagacttgag gagaatggct gagtctctcc ggtaggtttg
2760aaatactcaa ggatttgatg aaatactgtg cttgaccttt aggtataggg tctcagtctg
2820ctgttgaaaa atataatttc tacaaaccgt ctttgtaaaa ttttaagtat tgtagcagac
2880tttttaaaag tcagtgatac atctatatag tcaatatagg tttacatagt tgcaatctta
2940ttttgcatat gaatcagtat atagaagcag tggcatttat atgcttatgt tgcatttaca
3000attatgttta gacgaacaca aactttatgt gatttggatt agtgctcatt aaattttttt
3060attctatgga ctacaacaga gacataaatt ttgaaaggct tagttactct taaattctta
3120tgatgaaaag caaaaattca ttgttaaata gaacagtgca tccggaatgt gggtaattat
3180tgccatattt ctagtctact aaaaattgtg gcataactgt tcaaagtcat cagttgtttg
3240gaaagccaaa gtctgattta aatggaaaac ataaacaatg atatctattt ctagatacct
3300ttaacttgca gttactgagt ttacaagttg tctgacaact ttggattctc ttacttcata
3360tctaagaatg atcatgtgta cagtgcttac tgtcacttta aaaaactgca gggctagaca
3420tgcagatatg aagactttga cattagatgt ggtaattggc actaccagca agtggtatta
3480agatacagct gaatatatta ctttttgagg aacataattc atgaatggaa agtggagcat
3540tagagaggat gccttctggc tctcccacac cactgtttgc atccattgca tttcacactg
3600cttttagaac tcagatgttt catatggtat attgtgtaac tcaccatcag ttttatcttt
3660aaatgtctat ggatgataat gttgtatgtt aacactttta caaaaacaaa tgaagccata
3720tcctcggtgt gagttgtgat ggtggtaatt gtcacaatag gattattcag caaggaacta
3780agtcagggac aagaagtggg cgatactttg ttggattaaa tcattttact ggaagttcat
3840cagggagggt tatgaaagtt gtggtctttg aactgaaatt atatgtgatt cattattctt
3900gatttaggcc ttgctaatag taactatcat ttattgggaa tttgtcatat gtgccaattt
3960gtcatgggcc agacagcgtg ttttactgaa tttctagata tctttatgag attctagtac
4020tgttttcagc cattttacag atgaagaatc ttaaaaaatg ttaaataatt tagtttgccc
4080aagattatac gttaacaaat ggtagaacct tctttgaatt ctggcagtat ggctacacag
4140tccgaactct tatcttccta agctgaaaac agaaaaagca atgacccaga aaattttatt
4200taaaagtctc aggagagact tcccatcctg agaagatctc ttttcccttt tataatttag
4260gctcctgaat aatcactgaa ttttctccat gttccatcta tagtactgtt atttctgttt
4320tccttttttc ttaccacaaa gtatcttgtt tttgctgtat gaaagaaaat gtgttattgt
4380aatgtgaaat tctctgtccc tgcagggtcc cacatccgcc tcaatcccaa ataaacacac
4440agaggctgta ttaattatga aactgttggt cagttggcta gggcttctta ttggctagct
4500ctgtcttaat tattaaacca taactactat tgtaagtatt tccatgtggt cttatcttac
4560caaggaaagg gtccagggac ctcttactcc tctggcgtgt tggcagtgaa gaggagagag
4620cgatttccta tttgtctctg cttattttct gattctgctc agctatgtca cttcctgcct
4680ggccaatcag ccaatcagtg ttttattcat tagccaataa aagaaacatt tacacagaag
4740gacttccccc atcatgttat ttgtatgagt tcttcagaaa atcatagtat cttttaatac
4800taatttttat aaaaaattaa ttgtattgaa aattatgtgt atatgtgtct gtgtgtcgat
4860ttgtgctcat aagtagcatg gagtgcagaa gagggaatca gatctttttt taagggacaa
4920agagtttatt cagattacat tttaaggtga taatgtatga ttgcaaggtt atcaacatgg
4980cagaaatgtg aagaagctgg tcacattaca tccagagtca agagtagaga gcaatgaatt
5040gatgcatgca ttcctgtgct cagctcactt ttcctggagc tgagctgatt gtaagccatc
5100tgatgtcttt gctgggaact aactcaaagg caagttcaaa acctgttctt aagtataagc
5160catctctcca gtccctcata tggtctctta agacactttc tttatattct tgtacataga
5220aattgaattc ctaacaactg cattcaaatt acaaaatagt ttttaaaagc tgatataata
5280aatgtaaata caatctagaa catttttata aataagcata ttaactcagt aaaaataaat
5340gcatggttat tttccttcat tagggaagta tgtctcccca ggctgttctc tagattctac
5400tagtaatgct gtttgtacac catccacagg ggttttattt taaagctaag acatgaatga
5460tggacatgct tgttagcatt tagacttttt tccttactat aattgagcta gtatttttgt
5520gctcagtttg atatctgtta attcagataa atgtaatagt aggtaatttc tttgtgataa
5580aggcatataa attgaagttg gaaaacaaaa gcctgaaatg acagttttta agattcagaa
5640caataatttt caaaagcagt tacccaactt tccaaataca atctgcagtt ttcttgatat
5700gtgataaatt tagacaaaga aatagcacat tttaaaatag ctatttactc ttgatttttt
5760tttcaaattt aggctagttc actagttgtg tgtaaggtta tggctgcaaa catctttgac
5820tcttggttag ggaatccagg atgatttacg tgtttggcca aaatcttgtt ccattctggg
5880tttcttctct atctaggtag ctagcacaag ttaaaggtgt ggtagtattg gaaggctctc
5940aggtatatat ttctatattc tgtatttttt tcctctgtca tatatttgct ttctgtttta
6000ttgatttcta ctgttagttt gatacttact ttcttacact ttctttggga tttattttgc
6060tgttctaaga tttcttagca agttcatatc actgatttta acagttgctt cttttgtaat
6120atagactgaa tgccccttat ttgaaatgct tgggatcaga aactcagatt tgaacttttc
6180ttttttaata tttccatcaa gtttaccagc tgaatgtcct gatccaagaa tatgaaatct
6240gaaatgcttt gaaatctgaa acttttagag tgataaagct tccctttaaa ttaatttgtg
6300ttctatattt tttgacaatg tcaacctttc attgttatcc aatgagtgaa catattttca
6360atttttttgt ttgatctgtt atattttgat ctgaccatat ttataaaatt ttatttaatt
6420tgaatgttgt gctgttactt atctttatta ttatttttgc ttattttcta gccaaatgaa
6480attatattct gtattatttt agtttgaatt ttactttgtg gcttagtaac tgccttttgt
6540tggtgaatgc ttaagaaaaa cgtgtggtct actgatattg gttctaatct tatatagcat
6600gttgtttgtt aggtagttga ttatgctggt cagattgtct tgagtttatg caaatgtaaa
6660atatttagat gcttgttttg ttgtctaaga acaaagtatg cttgctgtct cctatcggtt
6720ctggtttttc cattcatctc ttcaagctgt tttgtgtgtt gaatactaac tccgtactat
6780cttgttttct gtgaattaac cccttttcaa aggtttcttt tctttttttt tttaagggac
6840aacaagttta ttcagattac attttaagct gataatgtat gattgcaagg ttatcaacat
6900ggcagaaatg tgaagaagct aggcacatta catccacatg gagtcaagag cagagagcag
6960tgaattaatg catgcattcc tgtggtcagc tcacttttcc tattcttaga tagtctagga
7020tcataaacct ggggaatagt gctaccacaa tgggcatatc cacttacttc agttcatgca
7080atcaaccaag gcacatccac aggaaaaact gatttagaca acctctcatt gagactcttc
7140ccagatgatt agactgtgtc aagttgacaa ttaaaactat cacacctgaa gccatcacta
7200gtaaatataa tgaaaatgtt gattatcacc ataattcatc tgtatccctt tgttattgta
7260gattttgtga agttcctatt caagtccctg ttccttcctt aaaaacctgt tttttagtta
7320aataggtttt ttagtgttcc tgtctgtaaa tactttttta aagttagata ttattttcaa
7380gtatgttctc ccagtctttg gcttgtattt tcatcccttc aatacatata tttttgtaat
7440ttattttttt tatttaaatt agaaacaaag ctgcttttac atgtcagtct cagttccctc
7500tccctcccct cctcccctgc tccccaccta agccccaatt ccaactcctt tcttctcccc
7560aggaagggtg aggccctcca tgggggaaat cttcaatgtc tgtcatatca tttggagcag
7620ggcctagacc ctccccagtg tgtctaggct gagagagtat ccctctatgt ggagagggct
7680cccaaagttc atttgtgtac taggggtaaa tactgatcca ctatcagtgg ccccatagat
7740tgtccggacc tccaaactga cttcctcctt cagggagtct ggaacagttc tatgctggtt
7800tcccagatat cagtctgggg tccatgagca accccttgtt caggtcagtt gtttctgtag
7860gtttccccag cccggtcttg acccctttgc tcatcacttc tccctctctg caactggatt
7920ccagagttca gctcagtgtt tagctgtggg tgtctgcatc tgcttccatc agctactgga
7980tgagggctct aggatggcat ataaggtagt catcagtctc attatcagag aagggctttt
8040aaggtagcct cttgattatt gcttagattg ttagttgggg tcaaccttgt aggtctctgg
8100acagtgacag aattctcttt aaacctataa tggctccctc tgtggtggta tcccttttct
8160tgctctcatc cgttcctccc ctgactagat cttcctgctc cctcatgtcc tcctctcccc
8220tccccttctc cccttctctt tcttctaact ccctctcccc tccacccacg atccccatta
8280gcttatgaga tcttgtcctt attttagcaa aacctttttg gctataaaat taattaattt
8340aatatgctta tatcaggttt attttggcta gtatttgtat gtgtttggtt agtgttttta
8400accttaattg acatgtatcc ttatatttag acacagattt aaatatttga agtttttttt
8460tttttttttt ttaaagattt atttattttt tatgtcttct gcctgcatgc cagaagaggg
8520caccagatct cattcaaggt ggttgtgagc caccatgtgg ttgctgggaa ttgaactcag
8580gacctctgga agaacagtca gtgctcttaa ccgctgagcc atctctccag cccctgaagt
8640gtttctttta aagaggatag cagtgcatca tttttccctt tgaccaatga ctcctacctt
8700actgaattgt tttagccatt tatatgtaat gctgttacca ggtttacatt ttcttttatc
8760ttgctaaatt tcttccctgt ttgtctcatc tcttattttt gtctgttgga ttatataggc
8820ttttattttt ctgtttttac agtaagttat atcaaattaa aattatttta tggaatgggt
8880gtgttgacta catgtatgtc tgtgcaccat gtgctgacct ggtcttggcc agaagaaggt
8940gtcatattct ctgaaactgg tattgtggat gttacgaact gccatagggt gctaggaatc
9000aaaccccagc tcctctggaa aagcagccac tgctctgagc cactgagtcc tctcttcaag
9060caggtgatgc caacttttaa tggttaccag tggataagag tgcttgtatc tctagcaccc
9120atgaaaattt atgcattgct atatgggctt gtcacttcag cattgtgtga cagagacagg
9180aggatcccaa gagctc
9196145PRTMus musculus 14Asn Tyr Gly Met Ser 1 51517PRTMus
musculus 15Thr Ile Ser Ser Ala Ser Thr Tyr Ser Tyr Tyr Pro Asp Ser Val
Lys 1 5 10
15Gly1610PRTMus musculus 16His Ser Asp Gly Asn Phe Ala Phe Gly Tyr 1
5 101716PRTMus musculus 17Arg Ser Ser Arg Asn
Ile Val His Ile Asn Gly Asp Thr Tyr Leu Glu 1 5
10 15187PRTMus musculus 18Lys Val Ser Asn Arg Phe
Ser 1 5199PRTMus musculus 19Phe Gln Gly Ser Leu Leu Phe
Trp Thr 1 520414DNAMus musculusCDS(1)..(414) 20atg aac ctc
ggg ctc agt ttg att ttc ctt gcc ctc att tta aaa ggt 48Met Asn Leu
Gly Leu Ser Leu Ile Phe Leu Ala Leu Ile Leu Lys Gly 1 5
10 15gtc cag tgt gag gtg cag ctg gtg gag
tct ggg gga gac tta atg aag 96Val Gln Cys Glu Val Gln Leu Val Glu
Ser Gly Gly Asp Leu Met Lys 20 25
30cct gga ggg tcc ctg aaa atc tcc tgt gca gcc tct gga ttc att ttc
144Pro Gly Gly Ser Leu Lys Ile Ser Cys Ala Ala Ser Gly Phe Ile Phe
35 40 45agt aat tat ggc atg tct tgg
gtt cgc cag act cca gac atg agg ctg 192Ser Asn Tyr Gly Met Ser Trp
Val Arg Gln Thr Pro Asp Met Arg Leu 50 55
60gaa tgg gtc gca acc att agt agt gct agt act tat tcc tat tat cca
240Glu Trp Val Ala Thr Ile Ser Ser Ala Ser Thr Tyr Ser Tyr Tyr Pro 65
70 75 80gac agt gtg aag
gga cga ttc acc ata tcc agg gac aac gcc gag aac 288Asp Ser Val Lys
Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Glu Asn 85
90 95tcc cta tat ctg caa atg aat agt ctg agg
tct gag gac aca ggc ata 336Ser Leu Tyr Leu Gln Met Asn Ser Leu Arg
Ser Glu Asp Thr Gly Ile 100 105
110tat tac tgt gga aga cat agc gat gga aac ttc gcg ttt ggt tat tgg
384Tyr Tyr Cys Gly Arg His Ser Asp Gly Asn Phe Ala Phe Gly Tyr Trp
115 120 125ggc cga ggg act ctg gtc act
gtc tct gca 414Gly Arg Gly Thr Leu Val Thr
Val Ser Ala 130 13521119PRTMus musculus 21Glu Val Gln
Leu Val Glu Ser Gly Gly Asp Leu Met Lys Pro Gly Gly 1 5
10 15Ser Leu Lys Ile Ser Cys Ala Ala Ser
Gly Phe Ile Phe Ser Asn Tyr 20 25
30Gly Met Ser Trp Val Arg Gln Thr Pro Asp Met Arg Leu Glu Trp Val
35 40 45Ala Thr Ile Ser Ser Ala
Ser Thr Tyr Ser Tyr Tyr Pro Asp Ser Val 50 55
60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Glu Asn Ser Leu
Tyr 65 70 75 80Leu Gln
Met Asn Ser Leu Arg Ser Glu Asp Thr Gly Ile Tyr Tyr Cys
85 90 95Gly Arg His Ser Asp Gly Asn Phe
Ala Phe Gly Tyr Trp Gly Arg Gly 100 105
110Thr Leu Val Thr Val Ser Ala 11522396DNAMus
musculusCDS(1)..(396) 22atg aag ttg cct gtt agg ctg ttg gtg ctg atg ttc
tgg att cct gct 48Met Lys Leu Pro Val Arg Leu Leu Val Leu Met Phe
Trp Ile Pro Ala 1 5 10
15tcc agc agt gat gtt ttg atg acc caa act cca ctc tcc ctg cct gtc
96Ser Ser Ser Asp Val Leu Met Thr Gln Thr Pro Leu Ser Leu Pro Val
20 25 30agt ctt gga gat caa gcc tcc
atc tct tgc aga tct agt cgg aac att 144Ser Leu Gly Asp Gln Ala Ser
Ile Ser Cys Arg Ser Ser Arg Asn Ile 35 40
45gtt cat att aat ggt gac aca tat tta gaa tgg tac ctg cag aga
ccg 192Val His Ile Asn Gly Asp Thr Tyr Leu Glu Trp Tyr Leu Gln Arg
Pro 50 55 60ggc cag tct cca aag ctc
cta atc tac aaa gtt tcc aac cga ttt tct 240Gly Gln Ser Pro Lys Leu
Leu Ile Tyr Lys Val Ser Asn Arg Phe Ser 65 70
75 80ggg gtc cca gac agg ttc agt ggc agt gga tca
ggg aca gat ttc aca 288Gly Val Pro Asp Arg Phe Ser Gly Ser Gly Ser
Gly Thr Asp Phe Thr 85 90
95ctc aag atc agc aga gtg gag gct gag gat ctg gga gtt tat tac tgc
336Leu Lys Ile Ser Arg Val Glu Ala Glu Asp Leu Gly Val Tyr Tyr Cys
100 105 110ttt caa ggt tca ctt ctt
ccg tgg acg ttc ggt gga ggc acc agg ctg 384Phe Gln Gly Ser Leu Leu
Pro Trp Thr Phe Gly Gly Gly Thr Arg Leu 115 120
125gaa atc aga cgg
396Glu Ile Arg Arg 13023113PRTMus musculus 23Asp Val Leu
Met Thr Gln Thr Pro Leu Ser Leu Pro Val Ser Leu Gly 1 5
10 15Asp Gln Ala Ser Ile Ser Cys Arg Ser
Ser Arg Asn Ile Val His Ile 20 25
30Asn Gly Asp Thr Tyr Leu Glu Trp Tyr Leu Gln Arg Pro Gly Gln Ser
35 40 45Pro Lys Leu Leu Ile Tyr
Lys Val Ser Asn Arg Phe Ser Gly Val Pro 50 55
60Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu Lys
Ile 65 70 75 80Ser Arg
Val Glu Ala Glu Asp Leu Gly Val Tyr Tyr Cys Phe Gln Gly
85 90 95Ser Leu Leu Pro Trp Thr Phe Gly
Gly Gly Thr Arg Leu Glu Ile Arg 100 105
110Arg24119PRTArtificial SequenceSynthetic peptide 24Glu Val Gln
Leu Val Glu Ser Gly Gly Asp Leu Val Gln Pro Gly Arg 1 5
10 15Ser Leu Arg Leu Ser Cys Ala Ala Ser
Gly Phe Ile Phe Ser Asn Tyr 20 25
30Gly Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp Val
35 40 45Ala Thr Ile Ser Ser Ala
Ser Thr Tyr Ser Tyr Tyr Pro Asp Ser Val 50 55
60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu
Tyr 65 70 75 80Leu Gln
Met Asn Ser Leu Arg Val Glu Asp Thr Ala Leu Tyr Tyr Cys
85 90 95Ala Arg His Ser Asp Gly Asn Phe
Ala Phe Gly Tyr Trp Gly Gln Gly 100 105
110Thr Leu Val Thr Val Ser Ser 11525119PRTArtificial
SequenceSynthetic peptide 25Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu
Val Gln Pro Gly Gly 1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Thr Phe Ser Asn Tyr
20 25 30Gly Met Ser Trp Val Arg
Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40
45Ser Thr Ile Ser Ser Ala Ser Thr Tyr Ser Tyr Tyr Pro Asp
Ser Val 50 55 60Lys Gly Arg Phe Thr
Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr 65 70
75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90
95Ala Arg His Ser Asp Gly Asn Phe Ala Phe Gly Tyr Trp Gly Gln Gly
100 105 110Thr Leu Val Thr Val
Ser Ser 11526112PRTArtificial SequenceSynthetic peptide 26Asp Ile
Val Met Thr Gln Ser Pro Leu Ser Leu Pro Val Thr Pro Gly 1
5 10 15Glu Pro Ala Ser Ile Ser Cys Arg
Ser Ser Arg Asn Ile Val His Ile 20 25
30Asn Gly Asp Thr Tyr Leu Glu Trp Tyr Leu Gln Lys Pro Gly Gln
Ser 35 40 45Pro Gln Leu Leu Ile
Tyr Lys Val Ser Asn Arg Phe Ser Gly Val Pro 50 55
60Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu
Lys Ile 65 70 75 80Ser
Arg Val Glu Ala Glu Asp Val Gly Val Tyr Tyr Cys Phe Gln Gly
85 90 95Ser Leu Leu Pro Trp Thr Phe
Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105
11027119PRTArtificial SequenceSynthetic peptide 27Glu Val
Gln Leu Val Glu Ser Gly Gly Asp Leu Val Gln Pro Gly Arg 1
5 10 15Ser Leu Arg Leu Ser Cys Ala Ala
Ser Gly Phe Ile Phe Ser Asn Tyr 20 25
30Gly Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp
Val 35 40 45Ala Thr Ile Ser Ser
Ala Ser Thr Tyr Ser Tyr Tyr Pro Asp Ser Val 50 55
60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser
Leu Tyr 65 70 75 80Leu
Gln Met Asn Ser Leu Arg Val Glu Asp Thr Ala Leu Tyr Tyr Cys
85 90 95Gly Arg His Ser Asp Gly Asn
Phe Ala Phe Gly Tyr Trp Gly Gln Gly 100 105
110Thr Leu Val Thr Val Ser Ser 11528119PRTArtificial
SequenceSynthetic peptide 28Glu Val Gln Leu Val Glu Ser Gly Gly Asp Leu
Val Gln Pro Gly Arg 1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Ile Phe Ser Asn Tyr
20 25 30Gly Met Ser Trp Val Arg
Gln Ala Pro Asp Lys Arg Leu Glu Trp Val 35 40
45Ala Thr Ile Ser Ser Ala Ser Thr Tyr Ser Tyr Tyr Pro Asp
Ser Val 50 55 60Lys Gly Arg Phe Thr
Ile Ser Arg Asp Asn Ala Lys Asn Ser Leu Tyr 65 70
75 80Leu Gln Met Asn Ser Leu Arg Val Glu Asp
Thr Ala Leu Tyr Tyr Cys 85 90
95Ala Arg His Ser Asp Gly Asn Phe Ala Phe Gly Tyr Trp Gly Gln Gly
100 105 110Thr Leu Val Thr Val
Ser Ser 11529119PRTArtificial SequenceSynthetic peptide 29Glu Val
Gln Leu Val Glu Ser Gly Gly Asp Leu Val Gln Pro Gly Arg 1
5 10 15Ser Leu Arg Leu Ser Cys Ala Ala
Ser Gly Phe Ile Phe Ser Asn Tyr 20 25
30Gly Met Ser Trp Val Arg Gln Ala Pro Asp Lys Arg Leu Glu Trp
Val 35 40 45Ala Thr Ile Ser Ser
Ala Ser Thr Tyr Ser Tyr Tyr Pro Asp Ser Val 50 55
60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ala Lys Asn Ser
Leu Tyr 65 70 75 80Leu
Gln Met Asn Ser Leu Arg Val Glu Asp Thr Ala Leu Tyr Tyr Cys
85 90 95Gly Arg His Ser Asp Gly Asn
Phe Ala Phe Gly Tyr Trp Gly Gln Gly 100 105
110Thr Leu Val Thr Val Ser Ser 11530119PRTArtificial
SequenceSynthetic peptide 30Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu
Val Gln Pro Gly Gly 1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Ile Phe Ser Asn Tyr
20 25 30Gly Met Ser Trp Val Arg
Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40
45Ser Thr Ile Ser Ser Ala Ser Thr Tyr Ser Tyr Tyr Pro Asp
Ser Val 50 55 60Lys Gly Arg Phe Thr
Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr 65 70
75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90
95Ala Arg His Ser Asp Gly Asn Phe Ala Phe Gly Tyr Trp Gly Gln Gly
100 105 110Thr Leu Val Thr Val
Ser Ser 11531119PRTArtificial SequenceSynthetic peptide 31Glu Val
Gln Leu Val Glu Ser Gly Gly Gly Leu Val Gln Pro Gly Gly 1
5 10 15Ser Leu Arg Leu Ser Cys Ala Ala
Ser Gly Phe Thr Phe Ser Asn Tyr 20 25
30Gly Met Ser Trp Val Arg Gln Ala Pro Gly Lys Gly Leu Glu Trp
Val 35 40 45Ser Thr Ile Ser Ser
Ala Ser Thr Tyr Ser Tyr Tyr Pro Asp Ser Val 50 55
60Lys Gly Arg Phe Thr Ile Ser Arg Asp Asn Ser Lys Asn Thr
Leu Tyr 65 70 75 80Leu
Gln Met Asn Ser Leu Arg Ala Glu Asp Thr Ala Val Tyr Tyr Cys
85 90 95Gly Arg His Ser Asp Gly Asn
Phe Ala Phe Gly Tyr Trp Gly Gln Gly 100 105
110Thr Leu Val Thr Val Ser Ser 11532119PRTArtificial
SequenceSynthetic peptide 32Glu Val Gln Leu Val Glu Ser Gly Gly Gly Leu
Val Gln Pro Gly Gly 1 5 10
15Ser Leu Arg Leu Ser Cys Ala Ala Ser Gly Phe Ile Phe Ser Asn Tyr
20 25 30Gly Met Ser Trp Val Arg
Gln Ala Pro Gly Lys Gly Leu Glu Trp Val 35 40
45Ser Thr Ile Ser Ser Ala Ser Thr Tyr Ser Tyr Tyr Pro Asp
Ser Val 50 55 60Lys Gly Arg Phe Thr
Ile Ser Arg Asp Asn Ser Lys Asn Thr Leu Tyr 65 70
75 80Leu Gln Met Asn Ser Leu Arg Ala Glu Asp
Thr Ala Val Tyr Tyr Cys 85 90
95Gly Arg His Ser Asp Gly Asn Phe Ala Phe Gly Tyr Trp Gly Gln Gly
100 105 110Thr Leu Val Thr Val
Ser Ser 11533112PRTArtificial SequenceSynthetic peptide 33Asp Val
Val Met Thr Gln Ser Pro Leu Ser Leu Pro Val Thr Pro Gly 1
5 10 15Glu Pro Ala Ser Ile Ser Cys Arg
Ser Ser Arg Asn Ile Val His Ile 20 25
30Asn Gly Asp Thr Tyr Leu Glu Trp Tyr Leu Gln Lys Pro Gly Gln
Ser 35 40 45Pro Gln Leu Leu Ile
Tyr Lys Val Ser Asn Arg Phe Ser Gly Val Pro 50 55
60Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu
Lys Ile 65 70 75 80Ser
Arg Val Glu Ala Glu Asp Val Gly Val Tyr Tyr Cys Phe Gln Gly
85 90 95Ser Leu Leu Pro Trp Thr Phe
Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105
11034112PRTArtificial SequenceSynthetic peptide 34Asp Ile
Leu Met Thr Gln Ser Pro Leu Ser Leu Pro Val Thr Pro Gly 1
5 10 15Glu Pro Ala Ser Ile Ser Cys Arg
Ser Ser Arg Asn Ile Val His Ile 20 25
30Asn Gly Asp Thr Tyr Leu Glu Trp Tyr Leu Gln Lys Pro Gly Gln
Ser 35 40 45Pro Gln Leu Leu Ile
Tyr Lys Val Ser Asn Arg Phe Ser Gly Val Pro 50 55
60Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu
Lys Ile 65 70 75 80Ser
Arg Val Glu Ala Glu Asp Val Gly Val Tyr Tyr Cys Phe Gln Gly
85 90 95Ser Leu Leu Pro Trp Thr Phe
Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105
11035112PRTArtificial SequenceSynthetic peptide 35Asp Val
Leu Met Thr Gln Ser Pro Leu Ser Leu Pro Val Thr Pro Gly 1
5 10 15Glu Pro Ala Ser Ile Ser Cys Arg
Ser Ser Arg Asn Ile Val His Ile 20 25
30Asn Gly Asp Thr Tyr Leu Glu Trp Tyr Leu Gln Lys Pro Gly Gln
Ser 35 40 45Pro Gln Leu Leu Ile
Tyr Lys Val Ser Asn Arg Phe Ser Gly Val Pro 50 55
60Asp Arg Phe Ser Gly Ser Gly Ser Gly Thr Asp Phe Thr Leu
Lys Ile 65 70 75 80Ser
Arg Val Glu Ala Glu Asp Val Gly Val Tyr Tyr Cys Phe Gln Gly
85 90 95Ser Leu Leu Pro Trp Thr Phe
Gly Gln Gly Thr Lys Val Glu Ile Lys 100 105
11036360PRTHomo sapiens 36Met Asn Pro Thr Asp Ile Ala Asp
Thr Thr Leu Asp Glu Ser Ile Tyr 1 5 10
15Ser Asn Tyr Tyr Leu Tyr Glu Ser Ile Pro Lys Pro Cys Thr
Lys Glu 20 25 30Gly Ile Lys
Ala Phe Gly Glu Leu Phe Leu Pro Pro Leu Tyr Ser Leu 35
40 45Val Phe Val Phe Gly Leu Leu Gly Asn Ser Val
Val Val Leu Val Leu 50 55 60Phe Lys
Tyr Lys Arg Leu Arg Ser Met Thr Asp Val Tyr Leu Leu Asn 65
70 75 80Leu Ala Ile Ser Asp Leu Leu
Phe Val Phe Ser Leu Pro Phe Trp Gly 85
90 95Tyr Tyr Ala Ala Asp Gln Trp Val Phe Gly Leu Gly Leu
Cys Lys Met 100 105 110Ile Ser
Trp Met Tyr Leu Val Gly Phe Tyr Ser Gly Ile Phe Phe Val 115
120 125Met Leu Met Ser Ile Asp Arg Tyr Leu Ala
Ile Val His Ala Val Phe 130 135 140Ser
Leu Arg Ala Arg Thr Leu Thr Tyr Gly Val Ile Thr Ser Leu Ala145
150 155 160Thr Trp Ser Val Ala Val
Phe Ala Ser Leu Pro Gly Phe Leu Phe Ser 165
170 175Thr Cys Tyr Thr Glu Arg Asn His Thr Tyr Cys Lys
Thr Lys Tyr Ser 180 185 190Leu
Asn Ser Thr Thr Trp Lys Val Leu Ser Ser Leu Glu Ile Asn Ile 195
200 205Leu Gly Leu Val Ile Pro Leu Gly Ile
Met Leu Phe Cys Tyr Ser Met 210 215
220Ile Ile Arg Thr Leu Gln His Cys Lys Asn Glu Lys Lys Asn Lys Ala225
230 235 240Val Lys Met Ile
Phe Ala Val Val Val Leu Phe Leu Gly Phe Trp Thr 245
250 255Pro Tyr Asn Ile Val Leu Phe Leu Glu Thr
Leu Val Glu Leu Glu Val 260 265
270Leu Gln Asp Cys Thr Phe Glu Arg Tyr Leu Asp Tyr Ala Ile Gln Ala
275 280 285Thr Glu Thr Leu Ala Phe Val
His Cys Cys Leu Asn Pro Ile Ile Tyr 290 295
300Phe Phe Leu Gly Glu Lys Phe Arg Lys Tyr Ile Leu Gln Leu Phe
Lys305 310 315 320Thr Cys
Arg Gly Leu Phe Val Leu Cys Gln Tyr Cys Gly Leu Leu Gln
325 330 335Ile Tyr Ser Ala Asp Thr Pro
Ser Ser Ser Tyr Thr Gln Ser Thr Met 340 345
350Asp His Asp Leu His Asp Ala Leu 355
3603728PRTHomo sapiens 37Asn Pro Thr Asp Ile Ala Asp Thr Thr Leu Asp Glu
Ser Ile Tyr Ser 1 5 10
15Asn Tyr Tyr Leu Tyr Glu Ser Ile Pro Lys Pro Cys 20
253818PRTHomo sapiens 38Asp Glu Ser Ile Tyr Ser Asn Tyr Tyr Leu
Tyr Glu Ser Ile Pro Lys 1 5 10
15Pro Cys3928DNAArtificial SequenceDescription of Artificial
Sequence Synthetic DNA 39gagacttcag cccacttcaa ttattggc
284025DNAArtificial SequenceDescription of
Artificial Sequence Synthetic DNA 40cttgtgtgac tcttaactct cagag
254125DNAArtificial
SequenceDescription of Artificial Sequence Synthetic DNA
41gaggccactt gtgtagcgcc aagtg
254223DNAArtificial SequenceDescription of Artificial Sequence
Synthetic DNA 42ccctcgagat aacttcgtat agc
234318DNAArtificial SequenceDescription of Artificial
Sequence Synthetic DNA 43ggtaggcctc actaactg
184425DNAArtificial SequenceDescription of
Artificial Sequence Synthetic DNA 44catagaaaca agtaacaaca gccag
254521DNAArtificial
SequenceDescription of Artificial Sequence Synthetic DNA
45gtgagtccat ggctgtcact g
214620DNAArtificial SequenceDescription of Artificial Sequence
Synthetic DNA 46cctgacttgg ctattctcag
20
User Contributions:
comments("1"); ?> comment_form("1"); ?>Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
User Contributions:
Comment about this patent or add new information about this topic:
| People who visited this patent also read: | |
| Patent application number | Title |
|---|---|
| 20130332813 | Generic Workspace Awareness Support for Collaborative Web Applications |
| 20130332812 | METHOD AND SYSTEM TO GENERATE A PROCESS FLOW DIAGRAM |
| 20130332811 | PROGRESSIVE LOADING FOR WEB-BASED SPREADSHEET APPLICATIONS |
| 20130332810 | MANAGING OBJECTS IN PANORAMA DISPLAY TO NAVIGATE SPREADSHEET |
| 20130332809 | WINE CLASSIFICATION SYSTEMS AND METHODS OF DISPLAYING WINES |