Patent application title: NUCLEIC ACID MOLECULES AND METHODS FOR EXCHANGING EXON(S) BY TRANSSPLICING
Inventors:
Luis Garcia (Bailly, FR)
Luis Garcia (Bailly, FR)
Stephanie Lorain (Vincennes, FR)
Assignees:
ASSOCIATION INSTITUT DE MYOLOGIE
Centre National De La Recherche Scientifique (CNRS)
INTITUT NATIONAL DE LA SANTE ET DE LA RECHERCHE MEDICALE (INSERM)
Universite Pierre Et Marie Curie (Paris 6)
IPC8 Class: AC07H2100FI
USPC Class:
514 44 R
Class name:
Publication date: 2013-03-14
Patent application number: 20130065948
Abstract:
The present invention provides methods and compositions for generating
novel nucleic acid molecules through targeted spliceosome mediated simple
or double trans-splicing. The compositions of the invention include
pre-trans-splicing molecules (PTMs) designed to interact with a target
precursor messenger RNA molecule (target pre-mRNA) and to mediate a
simple or double trans-splicing reaction resulting in the generation of a
novel chimeric RNA molecule (chimeric RNA).Claims:
1. A nucleic acid molecule comprising: a) two target binding domains AS
and AS' that target the binding of the nucleic acid molecule to a target
pre-mRNA, wherein the two target binding domains AS and AS' are located
respectively at the 5'-end and at the 3'-end of the nucleic acid
molecule, b) a 3' splice region comprising a branch point, a
polypyrimidine tract and a 3' splice acceptor site, c) a 5' splice region
comprising a 5' splice donor site, d) a spacer sequence that separates
the 3' splice region from the 5'-end target binding domain AS, e) a
spacer sequence that separates the 5' splice region from the 3'-end
target binding domain AS', and f) a nucleotide sequence to be
trans-spliced to the target pre-mRNA, wherein said nucleotide sequence
encodes at least a part of a normal polypeptide, and is located between
the 3' splice region and the 5' splice region of said nucleic acid.
2. The nucleic acid molecule according to claim 1, wherein the two target binding domains AS and AS' target binding of the nucleic acid molecule to the pre-mRNA of the dystrophin gene (DMD), and wherein the nucleotide sequence to be trans-spliced encodes at least a part of the normal dystrophin polypeptide.
3. The nucleic acid molecule according to claims 1 and 2, wherein each of the target binding domains AS and AS' comprises between about 100 and about 200 nucleotides, preferably about 150 nucleotides.
4. The nucleic acid molecule according to claims 2 and 3, wherein the nucleotide sequence to be trans-spliced comprises at least one exon of the normal DMD gene.
5. The nucleic acid molecule according to claims 2 to 4, wherein the nucleotide sequence to be trans-spliced comprises at least the sequence of exon 23 of the normal DMD gene, or the sequence of exon 70 of the normal DMD gene.
6. The nucleic acid molecule according to claim 5, wherein the nucleotide sequence to be trans-spliced comprises the exon 23 of the human gene (SEQ ID NO 60).
7. The nucleic acid molecule according to claims 1 to 6, wherein the 5'-end target binding domain AS targets the binding of the nucleic acid to the intron 22 of the pre-mRNA of the DMD gene.
8. The nucleic acid molecule according to claims 1 to 7, wherein the 3'-end target binding domain AS' targets the binding of the nucleic acid to the intron 23 of the pre-RNA of the DMD gene.
9. The nucleic acid molecule according to claims 1 to 8, wherein the 5'-end target binding domain AS comprises at least 20 successive nucleotides of one of the nucleotide sequence chosen among: SEQ ID NO 13 and SEQ ID NO 14.
10. The nucleic acid molecule according to claims 1 to 9, wherein the 3'-end target binding domain AS' targets the binding of the nucleic acid to a nucleotide sequence located in the 5'-half of the nucleotide sequence of intron 23.
11. The nucleic acid molecule according to claim 10, wherein the 3'-end target binding domain AS' comprises at least 20 successive nucleotides of one of the nucleotide sequences chosen among: SEQ ID NO 16, SEQ ID NO 19, and SEQ ID NO 20.
12. The nucleic acid molecule according to claims 1 to 11, wherein the 3'-end target binding domain AS' comprises at least 20 successive nucleotide of the SEQ ID NO 21.
13. The nucleic acid molecule according to claims 1 to 12, wherein the two spacers comprise between 10 and 100 nucleotides, preferably between 30 and 50 nucleotides.
14. The nucleic acid molecule according to claims 1 to 13, wherein the branch point is a conserved yeast branch point sequence, and is preferably SEQ ID NO 25.
15. The nucleic acid molecule according to claims 1 to 14, wherein the spacer separating the 5' splice donor site and the 3'-end target binding domain AS' comprises a downstream intronic splice enhancer (DISE), preferably the DISE sequence from the rat FGFR2 gene (SEQ ID NO 27).
16. A recombinant vector comprising the nucleic acid of anyone of claims 1 to 15.
17. The vector according to claim 16, wherein it is an eukaryotic expression vector.
18. A cell comprising the nucleic acid molecule according to anyone of claims 1 to 15, or the recombinant vector of claims 16 and 17.
19. The cell according to claim 18, wherein it is an eukaryotic cell.
20. An in vitro method of replacing a mutated endogenous exon of the DMD gene within a cell, comprising contacting the cellular pre-mRNA of the DMD gene with the nucleic acid molecule of anyone of claims 1 to 15, under conditions in which the nucleotide sequence to be trans-spliced is trans-spliced to the target pre-mRNA of the DMD gene to form a chimeric mRNA within the cell.
21. A method for correcting a DMD genetic defect in a subject, comprising administering to said subject the nucleic acid molecule of anyone of claims 1 to 15, or the vector of claim 16 or 17, or the cell of claim 18 or 19.
22. A method for correcting at least one genetic mutation present in at least one of the DMD gene in a subject in need thereof, comprising administering to said subject the nucleic acid of anyone of claims 1 to 15.
23. A method for treating the Duchenne muscular dystrophy in a subject in need thereof, comprising administering to said subject the nucleic acid of anyone of claims 1 to 15.
24. The method according to claim 23, wherein the nucleotide sequence to be trans-spliced comprises at least the exon 23 of the DMD gene (SEQ ID NO:8).
25. A nucleic acid molecule comprising: a) one target binding domain (AS) that target binding of the nucleic acid molecule to the pre-mRNA of the dystrophin gene (DMD), b) a 3' splice region comprising a branch point, a polypyrimidine tract and a 3' splice acceptor site, c) a spacer sequence that separates the 3' splice region from the target binding domain AS, d) a nucleotide sequence to be trans-spliced to the target pre-mRNA wherein said nucleotide sequence encodes at least a part of the DMD polypeptide.
26. The nucleic acid according to claim 25, wherein the sequence to be trans-spliced to the pre-mRNA of the dystrophin gene (DMD) comprises at least one exon of the normal DMD gene.
27. The nucleic acid according to claims 25 and 26, wherein the sequence to be trans-spliced to the pre-mRNA of the dystrophin gene (DMD) comprises at least one exon chosen among exons 59 to 79 of the DMD gene, preferably exon 70 of the DMD gene.
28. A recombinant vector comprising the nucleic acid of anyone of claims 25 to 27.
29. A cell comprising the nucleic acid molecule according to anyone of claims 25 to 27, or the recombinant vector of claim 28.
30. An in vitro method of replacing a mutated endogenous exon of the DMD gene within a cell, comprising contacting the cellular pre-mRNA of the DMD gene with the nucleic acid molecule of anyone of claims 25 to 27, under conditions in which the nucleotide sequence to be trans-spliced is trans-spliced to the target pre-mRNA of the DMD gene to form a chimeric mRNA within the cell.
31. A method for treating a patient suffering from the Duchenne muscular dystrophy comprising administering a pharmaceutical composition comprising the nucleic acid of anyone of claims 25 to 27 or the vector of claim 28, or the cell of claim 29.
32. A method for correcting a DMD genetic defect in a subject, comprising administering to said subject the nucleic acid molecule of anyone of claims 25 to 27, or the vector of claim 28, or the cell of claim 29.
33. The method for treating the Duchenne muscular dystrophy according to claim 31, or for correcting a DMD genetic defect in a subject according to claim 32, wherein the nucleotide sequence to be trans-spliced comprises at least one exon of the DMD gene.
34. A method for correcting at least one genetic mutation present in at least one exon of the DMD gene in a subject in need thereof, comprising administering to said subject the nucleic acid of anyone of claims 25 to 27.
35. The method according to claim 34, wherein the at least one mutation is in the exon 70 of the DMD gene, and the nucleotide sequence to be trans-spliced comprises the exon 70 of the normal human DMD gene (SEQ ID NO 72).
36. The method according to claim 35, wherein the nucleotide sequence to be trans-spliced comprises the cDNA from exon 59 to exon 79 of the DMD human gene (SEQ ID NO 70).
Description:
[0001] The present invention provides methods and compositions for
generating novel nucleic acid molecules through targeted spliceosome
mediated simple or double trans-splicing. The compositions of the
invention include pre-trans-splicing molecules (PTMs, herein also called
"TS molecule" for "Trans-Splicing molecules") designed to interact with a
target precursor messenger RNA molecule (target pre-mRNA) and to mediate
a simple or double trans-splicing reaction resulting in the generation of
a novel chimeric RNA molecule (chimeric RNA). This approach enables to
replace whole nucleotide sequences such as exonic sequences in a targeted
mRNA and is therefore very interesting to address disorders caused by
dominant mutations while preserving levels and tissue specificity. This
RNA repair strategy is thus useful to replace mutated nucleic acid
sequences into the normal ones and thereby treat many genetic disorders.
[0002] In particular, the PTMs of the present invention include those genetically engineered to interact with DMD target pre-mRNA so as to result in correction of DMD genetic defects responsible for the Duchenne muscular dystrophy (DMD).
[0003] The compositions of the invention further include recombinant vector systems capable of expressing the PTMs of the invention and cells expressing said PTMs. The methods of the invention encompass contacting the PTMs of the invention with a DMD target pre-mRNA under conditions in which a portion of the PTM is trans-spliced to a portion of the target pre-mRNA to form a mRNA molecule wherein the genetic defect in the DMD gene has been corrected. The methods and compositions of the present invention can be used in gene therapy for correction of neuromuscular disorders such as the Duchenne muscular dystrophy. The principle of this treatment can also be applied to any genetic disease where the pathogenic mutation involves an alteration of the transcript that can be corrected by simple or double trans-splicing.
BACKGROUND OF THE INVENTION
[0004] DNA sequences in the chromosome are transcribed into pre-mRNAs which contain coding regions (exons) and generally also contain intervening non-coding regions (introns). Introns are removed from pre-mRNAs in a precise process called cis-splicing. Splicing takes place as a coordinated interaction of several small nuclear ribonucleoprotein particles (snRNPs) and many protein factors that assemble to form an enzymatic complex known as the spliceosome (Staley and Guthrie, 1998).
[0005] In most cases, the splicing reaction occurs within the same pre-mRNA molecule, which is termed cis-splicing. Splicing between two independently transcribed pre-mRNAs is termed trans-splicing. Trans-splicing was first discovered in trypanosomes and subsequently in nematodes, flatworms and in plant mitochondria, drosophila, mice an humans (Takayuki Horiuchi and Toshiro Aigaki, 2006).
[0006] The mechanism of splice leader trans-splicing, which is nearly identical to that of conventional cis-splicing, proceeds via two phosphoryl transfer reactions. The first causes the formation of a 2'-5'phosphodiester bond producing a `Y` shaped branched intermediate, equivalent to the lariat intermediate in cis-splicing. The second reaction, exon ligation, proceeds as in conventional cis-splicing. In addition, sequences at the 3' splice site and some of the snRNPs which catalyze the trans-splicing reaction, closely resemble their counterparts involved in cis-splicing.
[0007] Trans-splicing may also refer to a different process, where an intron of one pre-mRNA interacts with an intron of a second pre-mRNA, enhancing the recombination of splice sites between two conventional pre-mRNAs. This type of trans-splicing was postulated to account for transcripts encoding a human immunoglobulin variable region sequence linked to the endogenous constant region in a transgenic mouse (Shimizu et al., 1989). In addition, trans-splicing of c-myb pre-RNA has been demonstrated (Vellard, M. et al. 1992) and more recently, RNA transcripts from cloned SV40 trans-spliced to each other were detected in cultured cells and nuclear extracts (Eul et al., 1995). However, naturally occurring trans-splicing of mammalian pre-mRNAs is thought to be a rare event (Finta, C. et al., 2002).
[0008] In vitro trans-splicing has been used as a model system to examine the mechanism of splicing by several. Reasonably efficient trans-splicing (30% of cis-spliced analog) was achieved between RNAs capable of base pairing to each other, whereas splicing of RNAs not tethered by base pairing was further diminished by a factor of 10. Other in vitro trans-splicing reactions not requiring obvious RNA-RNA interactions among the substrates were observed for example by Chiara & Reed (1995, Nature). These reactions occur at relatively low frequencies and require specialized elements, such as a downstream 5' splice site or exonic splicing enhancers.
[0009] The present invention relates to the use of targeted trans-splicing mediated by native mammalian splicing machinery, i. e., spliceosomes, to reprogram or alter the coding sequence of a targeted m-RNA.
[0010] A lot of studies have already described PTMs that can mediate "simple" trans-splicing, that is, a technology that enables to replace either the 3' part of a transcript, or, more rarely, the 5' part (Mansfield et al, 2003; Kierlin-Duncan & Sullenger, 2007). For example, U.S. Pat. Nos. 6,083,702, 6,013,487 and 6,280,978 describe the use of PTMs to mediate a "simple" trans-splicing reaction by contacting a target precursor mRNA to generate novel chimeric RNAs. Importantly, the "simple" trans-splicing technologies enable to correct a number of mutations using minigenes or endogenous transcripts in genetic disease context like hemophilia A (Chao et al., 2003), spinal muscular atrophy (Coady et al., 2008), X-linked immunodeficiency (Tahara et al., 2004) and cystic fibrosis where the widespread mutation CFTRΔF508 was replaced efficiently in vivo by the normal sequence via a trans-splicing reaction (Liu et al., 2005).
[0011] As opposed to "simple" trans-splicing, "double" trans-splicing enables to replace or introduce a sequence, such as exonic sequences, in a targeted mRNA (herein called ExChange: concomitant 3' and 5' trans-splicing reactions). More precisely, double trans-splicing can modify a given or replace a missing exonic sequence within a given gene transcript while at the same time preserving the regulatory intronic sequences which are present 5' and 3' of the exonic sequence targeted by double trans-splicing, thereby allowing for alternative transcripts to occur. Conversely to conventional gene therapy, "double" trans-splicing approaches would be very interesting to address disorders caused by dominant mutations, while preserving levels and tissue specificity. Exon exchange (ExChange) using double trans-splicing, at both sides of a targeted exon, would have the advantage of minimizing exogenous material as well as preserving full regulatory elements potentially present in 5' and/or 3' untranslated domains of the rescued mRNA. As a RNA repair strategy, the ExChange approach will produce the corrected protein where it is naturally expressed. It has the supplementary advantage upon other RNA surgery strategies of correcting precisely the sequence defect without changing anything to the whole messenger sequence (i.e. the open reading frame and untranslated regions). Hence, the regulatory sequences present in 5' and 3' UTRs are preserved, something which never happens in classical gene therapy where cDNAs are amputated of their non coding sequences. These regions are now known to be essential for mRNA stability and translation regulation; in particular, they are targets for miRNAs which play important role in a variety of disease (Zhang & Farwell, 2008).
[0012] However no study has ever described so far an efficient PTM enabling to perform a "double" trans-splicing, that is a PTM containing both a 3' splice region and a 5' splice region and a nucleotide sequence to be inserted into a target mRNA. Furthermore, no study has ever demonstrated that such PTM might be able to replace with high efficiency a nucleotide sequence inside a target mRNA.
[0013] The herein presented results demonstrate for the first time an efficient PTM enabling high level of ExChange (i.e. concomitant 3' and 5' trans-splicing reactions), and therefore the possibility of rescuing mutated transcripts by specifically replacing a mutated exon by its normal version during a double trans-splicing reaction. By using the PTM presented in the present invention, one can obtain an ExChange efficiency that reaches a level of at least about 50%, for example 53%, of repaired transcripts with DMD minigene as target. Importantly, no non-specific end products were ever detected, suggesting that the chosen annealing sequences of the PTM did not decipher cryptic splicing site nor obstruct splicing events.
[0014] The trans-splicing technology uses a trans-splicing molecule that "tricks" the spliceosome into using it as a substrate for splicing. In the ExChange approach, the game is more "tricky" since the spliceosome must realize a double trans-splicing between the pre-messenger transcript and the PTM. After having tested various combinations of antisenses, it was found by the present inventors that the dogmas: i) blocking endogenous splicing signals on the nascent pre-mRNA transcript via base-pairing or ii) at the opposite, bringing the replacing exon closer to endogenous splice site to be joined, did not produce the best results, as shown in the present examples. Indeed, the best antisense for the first trans-splicing (3' replacement) matched with the middle of the first intron while the second one (5' replacement) was better when located close to the 3' end of the exon to be replaced.
FIGURE LEGENDS
[0015] The FIG. 1 exposes the trans-splicing strategy for the 3' replacement:
[0016] (A) Exons 22 to 24 (boxes E22 to E24) with natural introns (lines with black balls illustrating the splice sites) on the DMD minigene. The cross represents the nonsense mdx mutation in E23. The trans-splicing (TS) molecule AS-E24 comprises a 150 nt antisense sequence (AS) complementary to intron 22 as well as a spacer, a strong conserved yeast branch point sequence, a polypyrimidine tract, a 3' acceptor site (the three last elements are represented as blacks balls) and E24. TS constructs were made with three different antisense sequences, AS1 to AS3. Arrows indicate the positions of the forward A and reverse B PCR primers in the minigene and the TS molecule. (B) Expected transcripts generated by cis-splicing (E23 inclusion and skipping) and trans-splicing, and the predicted sizes of the corresponding PCR amplification products detected using the RT-PCR strategy illustrated in (A). (C) RT-PCR analysis using PCR primers A and B of NIH3T3 cells cotransfected with DMD minigene and constructions pSMD2-GFP (Ctrl), pSMD2-U7-SD23-BP22 (U7), pSMD2-E24 (AS-), pSMD2-AS1-E24 (AS1), pSMD2-AS2-E24 (AS2) and pSMD2-AS3-E24 (AS3). RT-AS2: samples containing DMD minigene and pSMD2-AS2-E24 without reverse transcription; H2O: PCR negative control. (D) An exact E22-E24 junction was confirmed by sequencing of the 310 bp product.
[0017] The FIG. 2 shows the exon replacement approach on DMD reporter transcripts.
[0018] (A) The exon exchange molecule (EX) AS-E24-AS' comprises the same elements as the TS molecule (see FIG. 1A) followed by a 5' donor site (black ball) and a second antisens sequence (AS') of 150 nt complementary to intron 23. EX constructs were made with five different AS' antisense sequences, AS4 to AS8. Arrows indicate the positions of the forward A and reverse C PCR primers in the minigene. (B) Expected transcripts generated by cis-splicing (E23 inclusion and skipping) and exon exchange, and predicted sizes of the corresponding PCR amplification products detected using the RT-PCR strategy illustrated in (A). (C) RT-PCR analysis using primers A and C of NIH3T3 cells cotransfected with DMD minigene and constructions pSMD2-GFP (Ctrl), pSMD2-U7-SD23-BP22 (U7), the TS constructions pSMD2-AS1-E24 (AS1), pSMD2-AS2-E24 (AS2) and EX molecules pSMD2-AS-E24-AS' containing AS1 or AS2 and AS4 to AS8. AS2-2XAS4, EX plasmid pSMD2-AS2-E24-2XAS4 containing two AS4 copies; H20: PCR negative control. (D) Accurate E22-E24 and E24-E24 junctions were confirmed by sequencing of the 408 bp product.
[0019] The FIG. 3 shows the effect of intronic splice enhancer sequences on exon replacement efficiency.
[0020] (A) Exon exchange molecules AS-E24-AS' with intronic splice enhancers ISE or DISE sequences. (B) RT-PCR analysis using primers A and C of NIH3T3 cells cotransfected with DMD minigene and constructs pSMD2-GFP (Ctrl), pSMD2-U7-SD23-BP22 (U7) and the following EX plasmids with AS4 or AS8: pSMD2-AS2-E24-AS' (-), pSMD2-AS2-ISE-E24-AS' (ISE), pSMD2-AS2-E24-DISE-AS' (DISE) and pSMD2-AS2-E24-2XAS' (2XAS'). H20: PCR negative control. (C) Efficiency of DMD exon exchange induced by AS4 containing EX molecules analyzed by absolute quantitative real-time RT-PCR.
[0021] The FIG. 4 shows the trans-splicing strategy for dystrophin transcript repair with the simple transsplicing molecule of the invention SEQ ID NO 71 (example 2)
[0022] The endogenous pre-messenger dystrophin transcript is illustrated on the top with boxes representing exons, and black lines representing introns. The trans-splicing mRNA molecule (second line) comprise a 150 nt antisense sequence complementary to intron 58 of the DMD gene as well as a spacer, a strong conserved yeast branch point sequence (BP), a polypyrimidine tract (PPT), a 3' splice acceptor (SA) and the normal human dystrophin cDNA from exon 59 to the exon 79 STOP codon.
[0023] The FIG. 5 shows the detection of repaired dystrophin transcripts with the simple transsplicing molecule of the invention SEQ ID NO 71 (example 2)
[0024] (A) Mutated dystrophin mRNA is represented on the top, as well as the trans-splicing mRNA molecule SEQ ID NO: (second line), and expected dystrophin transcripts generated by cis-splicing and trans-splicing (third line). Arrows indicate the positions of the forward Fo/Fi and reverse Ro/Ri PCR primers designed to detect only the repaired dystrophin cDNA by generating a 2443 bp PCR product. (B) RT-PCR analysis using PCR primers Fo/Fi and Ro/Ri of total RNAs extracted from patient myotubes (DMD) transduced by lentivirus expressing the simple trans-splicing mRNA molecules (TsM). Lane "-": DMD non transduced myotubes.
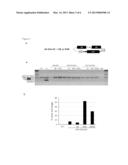
[0025] The FIG. 6 shows the Dystrophin rescue in DMD cells using the simple transsplicing molecule of the invention SEQ ID NO 71 (example 2)
[0026] Western blot of total protein extracted from DMD patient myotubes transduced by lentivirus expressing the TSM molecules, stained with the NCL-DYS1 monoclonal antibody. The full-length 427 kD dystrophin is indicated as detected in normal CHQ myotubes sample (WT). Lane "-": DMD non transduced myotubes. Each lane was loaded with 50 μg of total protein except Ctrl, 5 μg. Red panel: visualization of total proteins present on the same membrane by Ponceau red staining.
SUMMARY OF THE INVENTION
[0027] In a first aspect, the present invention is drawn to a nucleic acid molecule comprising:
[0028] a) two target binding domains AS and AS' that target the binding of the nucleic acid molecule to a target pre-mRNA, wherein the two target binding domains AS and AS' are located respectively at the 5'-end and at the 3'-end of the nucleic acid molecule,
[0029] b) a 3' splice region comprising a branch point, a polypyrimidine tract and a 3' splice acceptor site,
[0030] c) a 5' splice region comprising a 5' splice donor site,
[0031] d) a spacer sequence that separates the 3' splice region from the 5'-end target binding domain AS,
[0032] e) a spacer sequence that separates the 5' splice region from the 3'-end target binding domain AS', and
[0033] f) a nucleotide sequence to be trans-spliced to the target pre-mRNA, wherein said nucleotide sequence encodes at least a part of a normal polypeptide, and is located between the 3' splice region and the 5' splice region of said nucleic acid.
This nucleic acid molecule is hereafter referred to as "double trans-splicing molecule", or "double PTM" of the invention. Preferably, said nucleic acid molecule comprises:
[0034] a) two target binding domains AS and AS' that target binding of the nucleic acid molecule to the pre-mRNA of the dystrophin gene (DMD), wherein the two target binding domains AS and AS' are located respectively at the 5'-end and at the 3'-end of the nucleic acid molecule,
[0035] b) a 3' splice region comprising a branch point, a polypyrimidine tract and a 3' splice acceptor site,
[0036] c) a 5' splice region comprising a 5' splice donor site,
[0037] d) a spacer sequence that separates the 3' splice region from the 5'-end target binding domain AS,
[0038] e) a spacer sequence that separates the 5' splice region from the 3'-end target binding domain AS', and
[0039] f) a nucleotide sequence to be trans-spliced to the target pre-mRNA of the dystrophin gene (DMD), wherein said nucleotide sequence encodes at least a part of the normal dystrophin polypeptide, and is located between the 3' splice region and the 5' splice region of said nucleic acid.
[0040] In a second aspect, the present invention is drawn to a nucleic acid molecule comprising:
[0041] a) one target binding domain (AS) that target binding of the nucleic acid molecule to the pre-mRNA of the dystrophin gene (DMD),
[0042] b) a 3' splice region comprising a branch point, a polypyrimidine tract and a 3' splice acceptor site,
[0043] c) a spacer sequence that separates the 3' splice region from the target binding domain AS,
[0044] d) a nucleotide sequence to be trans-spliced to the target pre-mRNA wherein said nucleotide sequence encodes at least a part of the DMD polypeptide.
[0045] This nucleic acid molecule is hereafter referred to as "simple trans-splicing molecule", or "simple PTM" of the invention.
[0046] The present invention is also drawn to a recombinant vector comprising the PTMs of the invention and to a cell comprising the PTMs of the invention, or the recombinant vector of comprising the PTMs of the invention.
[0047] The compositions and methods can be used to provide a gene encoding a functional biologically active molecule to cells of an individual with an inherited genetic disorder where expression of the missing or mutant gene product produces a normal phenotype.
[0048] Specifically, the compositions and methods can be used to replace in vitro a mutated endogenous exon 23 or exon 70 of the DMD gene within a cell, comprising contacting the cellular pre-mRNA of the DMD gene with the PTMs of the present invention, under conditions in which the nucleotide sequence to be trans-spliced is trans-spliced to the target pre-mRNA of the DMD gene to form a chimeric mRNA within the cell.
[0049] To go further, the present invention also discloses a method for in vivo correcting a DMD genetic defect in a subject, comprising administering to said subject the PTMs of the invention, or the vector comprising the PTMs of the invention, or the cell comprising the PTMs of the invention.
[0050] More specifically, the present invention discloses a method for correcting at least one genetic mutation present in exon 23 or 70 of the DMD gene in a subject in need thereof, comprising administering to said subject the PTMs of the invention, wherein the nucleotide sequence to be trans-spliced is at least the exon 23 or at least the exon 70 of the DMD gene.
[0051] The present invention also provides pharmaceutical compositions comprising an effective amount of the PTMs of the invention and a pharmaceutically acceptable carrier.
DETAILED DESCRIPTION OF THE PREFERRED EMBODIMENTS
[0052] In a first aspect, the present invention relies on the designing and the optimization of a double PTM dedicated to concomitant 3' and 5' trans-splicing reactions in order to replace specific nucleotide sequence, and for example a mutated exon, or to replace missing exons in the case of deletion mutations. Such a double PTM therefore necessarily contains a 3' splice region and a 5' splice region. Also, the double PTM must contain at least two distinct target binding domains that enable the PTM to recognize and get very close to the target mRNA. In the context of the invention, these two target binding domains are called "AS" (for Anti Sens).
[0053] In this first aspect, the present invention is thus drawn to a nucleic acid molecule comprising:
[0054] a) two target binding domains AS and AS' that target the binding of the nucleic acid molecule to a target pre-mRNA, wherein the two target binding domains AS and AS' are located respectively at the 5'-end and at the 3'-end of the nucleic acid molecule,
[0055] b) a 3' splice region comprising a branch point, a polypyrimidine tract and a 3' splice acceptor site,
[0056] c) a 5' splice region comprising a 5' splice donor site,
[0057] d) a spacer sequence that separates the 3' splice region from the 5'-end target binding domain AS,
[0058] e) a spacer sequence that separates the 5' splice region from the 3'-end target binding domain AS', and
[0059] f) a nucleotide sequence to be trans-spliced to the target pre-mRNA, wherein said nucleotide sequence encodes at least a part of a normal polypeptide, and is located between the 3' splice region and the 5' splice region of said nucleic acid.
[0060] This nucleic acid molecule is hereafter designated by the "double trans-splicing molecule of the invention" or "the double PTM of the invention".
[0061] The target binding domain of a PTM endows the PTM with a binding affinity for the target pre-mRNA. As used herein, a target binding domain is defined as any molecule, i. e., nucleotide, protein, chemical compound, etc., that confers specificity of binding and anchors the pre-mRNA closely in space to the PTM so that the spliceosome processing machinery of the nucleus can trans-splice a portion of the PTM to a portion of the pre-mRNA. The target binding domains of the PTM are preferably nucleotide sequences which are complementary to and in anti-sense orientation to the targeted region of the selected target pre-mRNA. The target binding domains may comprise up to several thousand nucleotides. In preferred embodiments of the invention the target binding domains may comprise between about 100 and 200 nucleotides, and preferably about 150 nucleotides.
[0062] A sequence "complementary" to a portion of an RNA, as referred to herein, means a sequence having sufficient complementarity to be able to hybridize with the target pre-mRNA, forming a stable duplex. The ability to hybridize will depend on both the degree of complementarity and the length of the nucleic acid (See, for example, Sambrook et al., 1989, Molecular Cloning, A Laboratory Manual, 2d Ed., Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.). Generally, the longer the hybridizing nucleic acid, the more base mismatches with an RNA it may contain and still form a stable duplex. One skilled in the art can ascertain a tolerable degree of mismatch or length of duplex by use of standard procedures to determine the stability of the hybridized complex. Binding domains may encompass any or all sequences located within the target intron and flanking exons and may consist of contiguous sequence or contain sequence gaps ranging in size from a few to several hundred nucleotides in length. In such cases, the binding domain may be considered to be comprised of multiple, smaller binding domains that are positioned within the PTM in either orientation (sense or antisense) relative to the target sequence or to each other. Any or all sequence elements within the binding domain may contain significant complementarity to the target region.
[0063] After having tested various combinations of antisens nucleotide sequences, the present inventors have found that, contrary to what was commonly taught in the art, it is no use to block the endogenous splicing signals on the nascent pre-mRNA transcript via base-pairing. Therefore, in a preferred embodiment, the target binding domains of the double PTM of the present invention are not blocking the endogenous splicing signals on the nascent pre-RNA transcript.
[0064] In a first embodiment, the target pre-RNA is a mutated exon of a gene, and the nucleotide sequence to be trans-spliced to the target pre-mRNA, is the corresponding normal exon, or exons, of said gene.
[0065] The gene targeted by double-trans-splicing must be a gene that is actively transcribed in the cell targeted by the procedure. It must be composed of several exons that are transcribed into a pre-mRNA molecule.
[0066] In this case, it has been shown here for the first time that bringing the replacing exon closer to endogenous splice site to be joined does not produce the best results. Therefore, in the context of the invention, the target binding domains AS and AS' are preferably not complementary to sequences that are close to the endogenous splice sites of the mutated exon to be replaced, that is, below 200 nucleotides from the endogenous splice sites.
[0067] The double PTM molecule also contains a 3' splice region that includes a branch point sequence, a polypyrimidine tract (such as SEQ ID NO 28) and a 3' splice acceptor site. The double PTM molecule also contains a 5' splice region.
[0068] Consensus sequences for the 5' splice donor site and the 3' splice region used in RNA splicing are well known in the art (See, Moore, et al., 1993, The RNA World, Cold Spring Harbor Laboratory Press, p. 303-358). In addition, modified consensus sequences that maintain the ability to function as 5' donor splice sites and 3' splice regions may be used in the practice of the invention. Briefly, the mammalian consensus sequences for the 5' donor splice site and the 3' acceptor splice site are respectively: GTAAGT and TCCCTCCAG. For example, the 5' donor splice site of the double PTM of the invention can be GTAAGA (SEQ ID NO: 30) and the 3' acceptor splice site of the double PTM of the invention can be GGAAAACAG (SEQ ID NO: 29).
[0069] The branch point consensus sequence in mammals is YNYURAC (Y=pyrimidine; N=any nucleotide; R=purine). For example, the branch point can be TACTAAC (SEQ ID NO:25) corresponding to the well conserved yeast branch point (Mansfield et al 2000). The A is the site of branch formation. A polypyrimidine tract is located between the branch point and the 3' splice site acceptor and is important for different branch point utilization and 3' splice site recognition. Recently, pre-mRNA introns beginning with the dinucleotide AU and ending with the dinucleotide AC have been identified and referred to as U12 introns. U12 intron sequences as well as any sequences that function as splice acceptor/donor sequences may also be used to generate the double PTMs of the invention.
[0070] A spacer region to separate the RNA splice site from the target binding domain is also included in the double PTM. The double PTM of the invention contains at least two different spacers: a spacer sequence that separates the 3' splice region from the 5'-end target binding domain AS, and a spacer sequence that separates the 5' splice region from the 3'-end target binding domain AS'. They are preferably non coding sequences and comprise between 10 and 100 nucleotides, preferably between 20 and 70 nucleotides, more preferably between 30 and 50 nucleotides. The spacer regions may be designed to include features such as stop codons which would block any translation of a spliced PTM. In an embodiment of the invention, splicing enhancers such as, for example, sequences referred to as exonic splicing enhancers may also be included in the double PTM design. Transacting splicing factors, namely the serine/arginine-rich (SR) proteins, have been shown to interact with such exonic splicing enhancers and modulate splicing (Tacke et al., 1999). Also, the G-rich intronic splice enhancer from the human GH-1 gene (SEQ ID NO: 26) and/or the DISE sequence from the rat FGFR2 gene (SEQ ID NO: 27) can be used as splicing enhancers.
[0071] In a more preferred embodiment, the spacer sequence that separates the 3' splice region from the 5'-end target binding domain AS contains a ISE, for example the G-rich intronic splice enhancer from the human GH-1 gene (SEQ ID NO: 26), and the spacer sequence that separates the 5' splice region from the 3'-end target binding domain AS contains a DISE sequence, for example the DISE sequence from the rat FGFR2 gene (SEQ ID NO: 27), preferably in close vicinity of the nucleotide sequence to be trans-spliced, that is the DISE sequence should be located not farer than 60 nucleotides from the nucleotide sequence to be trans-spliced.
[0072] Additional features can be added to the double PTM molecule either after, or before, the nucleotide sequence encoding a translatable protein, such as polyadenylation signals to modify RNA expression/stability, or 5'splice sequences to enhance splicing, additional binding regions, "safety"-self complementary regions, additional splice sites, or protective groups to modulate the stability of the molecule and prevent degradation. In addition, stop codons may be included in the PTM structure to prevent translation of unspliced PTMs. Further elements such as a 3' hairpin structure, circularized RNA, nucleotide base modification, or synthetic analogs can be incorporated into double PTMs to promote or facilitate nuclear localization and spliceosomal incorporation, and intra-cellular stability.
[0073] As an example, the present invention relates to the design and the optimization of ExChange constructs (PTMs) designed for rescuing mutated mRNAs from very large genes such as the dystrophin gene DMD (for example the human DMD gene identified as NC--000023.10 or the mouse DMD gene identified as NC--000086.6). Mutations in the dystrophin gene DMD cause the Duchenne muscular dystrophy (DMD), the most common severe childhood muscular pathology. Recently, exon skipping strategies have proven to be efficacious in restoring functional dystrophin expression in models of muscular dystrophy including the mdx mouse, the GRMD dog and muscle stem cells from DMD patients and in four DMD patients by local intramuscular injection (Goyenvalle et al., 2004; Denti et al., 2006; Yokota et al., 2009). Indeed, the modular structure of the dystrophin, with its central rod-domain made of 24 spectrin-like repeats, tolerates large truncations. However, exon skipping strategies only concern patients for whom forced splicing would generate a shorter but still functional protein. Many pathological situations escape this prerequisite. In this context, ExChange strategies could be of great interest for replacing precisely a mutated exon of DMD by a normal corresponding exon, and for example the mutated exon 23 which carries a stop mutation in the mdx mouse model of DMD (cf. SEQ ID NO 9), or a genetic anomaly present between exons 59 and 79, which represent 8% of Duchenne patients. Furthermore, ExChange strategies could be useful for replacing missing exons, thereby producing a full length gene product instead of a truncated gene product as results from exon skipping approaches.
[0074] In such a view, the present invention shows here for the first time a PTM enabling an efficient exon exchange (for example one exon of the DMD gene, and more particularly the exon 23 of the DMD gene) importantly demonstrating that the ExChange of a specific exon is possible and efficient with the "double" trans-splicing technology.
[0075] In a preferred embodiment, the present invention is thus drawn to a nucleic acid molecule comprising:
[0076] a) two target binding domains AS and AS' that target binding of the nucleic acid molecule to the pre-mRNA of the dystrophin gene (DMD), wherein the two target binding domains AS and AS' are located respectively at the 5'-end and at the 3'-end of the nucleic acid molecule,
[0077] b) a 3' splice region comprising a branch point, a polypyrimidine tract and a 3' splice acceptor site,
[0078] c) a 5' splice region comprising a 5' splice donor site,
[0079] d) a spacer sequence that separates the 3' splice region from the 5'-end target binding domain AS,
[0080] e) a spacer sequence that separates the 5' splice region from the 3'-end target binding domain AS', and
[0081] f) a nucleotide sequence to be trans-spliced to the target pre-mRNA of the dystrophin gene (DMD), wherein said nucleotide sequence encodes at least a part of the normal dystrophin polypeptide, and is located between the 3' splice region and the 5' splice region of said nucleic acid.
[0082] The general design, construction and genetic engineering of PTMs and demonstration of their ability to mediate successful trans-splicing reactions within the cell are described in detail in U.S. Pat. Nos. 6,083,702, 6,013,487 and 6,280,978 as well as patent Ser. Nos. 09/941,492, 09/756,095, 09/756,096 and 09/756,097 the disclosures of which are incorporated by reference in their entirety herein.
[0083] In a particular embodiment, in the double PTM of the present invention, the nucleotide sequence to be trans-spliced comprises at least one exon of the normal DMD gene, preferably the sequence of exon 23 of the normal DMD gene, that is SEQ ID NO 8 (from the mouse gene) or SEQ ID NO 60 (from the human gene), or the exon 70 of the normal DMD gene (SEQ ID NO 72 for the human gene, SEQ ID NO 73 for the mouse gene).
[0084] In a preferred embodiment of the invention, the 5'-end target binding domain AS targets the binding of the nucleic acid to the intron 22 of the pre-mRNA of the DMD gene (SEQ ID NO 11 for the mouse gene, or SEQ ID NO 61 for the human gene) and the 3'-end target binding domain AS' targets the binding of the nucleic acid to the intron 23 of the pre-RNA of the DMD gene (SEQ ID NO 12 for the mouse gene, SEQ ID NO 62 for the human gene).
[0085] In an embodiment of the invention, the target binding domains AS and AS' comprises between about 100 and about 200 nucleotides, preferably about 150 nucleotides.
[0086] In a preferred embodiment, the 5'-end target binding domain AS comprises at least 20 successive nucleotides of one of the nucleotide sequence chosen among: SEQ ID NO 13 (hereafter called "AS1") and SEQ ID NO 14 (hereafter called "AS2"). Preferably, the 5'-end target binding domain AS comprises at least 20 successive nucleotides of SEQ ID NO 15.
[0087] On the other hand, the 3'-end target binding domain AS' targets preferably the binding of the nucleic acid to a nucleotide sequence located in the 5'-half of the nucleotide sequence of intron 23, and, more preferably, to a nucleotide sequence located in SEQ ID NO 22 (for the mouse gene) and comprises at least 20 successive nucleotides of one of the nucleotide sequences chosen among: SEQ ID NO 16 (hereafter called "AS4"), SEQ ID NO 19 (herafter called "AS 7"), SEQ ID NO 20 (hereafter called "AS8") and SEQ ID NO 21 (hereafter called "2XAS4"). In a preferred embodiment, the 3'-end target binding domain AS' comprises at least 20 successive nucleotides of SEQ ID NO 21.
[0088] In another embodiment, the double PTM of the invention comprises a conserved yeast branch point sequence, for example the yeast branch point of SEQ ID NO 25.
[0089] In another embodiment, the spacer separating the 5' splice donor site and the 3'-end target binding domain AS' comprises between 10 and 100 nucleotides, preferably between 20 and 70 nucleotides, more preferably between 30 and 50 nucleotides. In a preferred embodiment, this spacer comprises a downstream intronic splice enhancer (DISE), which is preferably the DISE sequence from the rat FGFR2 gene, i.e. the SEQ ID NO 27.
[0090] The spacer sequence that separates the 3' splice region from the 5'-end target binding domain AS comprises between 10 and 100 nucleotides, preferably between 20 and 70 nucleotides, more preferably between 30 and 50 nucleotides.
[0091] Such spacers are preferably non coding sequences but may be designed to include features such as stop codons which would block any translation of a spliced PTM. Examples of useful spacers are given in the experimental part of this application. They are for example SEQ ID NO 23 (3' end), and SEQ ID NO 24 (5' end).
[0092] In a specific embodiment, the present invention is drawn to a recombinant vector comprising the nucleic acid previously described. More particularly, the double PTM of interest may be recombinantly engineered into a variety of host vector systems that also provide for replication of the DNA in large scale and contain the necessary elements for directing the transcription of the double PTM. The use of such a construct to transfect target cells in the patient will result in the transcription of sufficient amounts of double PTMs that will form complementary base pairs with the endogenously expressed pre-mRNA targets, such as for example, DMD pre-mRNA target, and thereby facilitate a trans-splicing reaction between the complexed nucleic acid molecules. Such a vector can remain episomal or become chromosomally integrated, as long as it can be transcribed to produce the desired RNA, i. e., PTM. Such vectors can be constructed by recombinant DNA technology methods standard in the art. Vectors comprising the double PTM of interest can be plasmid, viral, or others known in the art, used for replication and expression in mammalian cells. Expression of the double PTM can be regulated by any promoter/enhancer sequences known in the art to act in mammalian, preferably human cells. Such promoters/enhancers can be inducible or constitutive. Such promoters include but are not limited to: the SV40 early promoter region, the promoter contained in the 3' long terminal repeat of Rous sarcoma virus, the herpes thymidine kinase promoter, the regulatory sequences of the metallothionein gene, the viral CMV promoter, the human chorionic gonadotropin-P promoter, etc. Any type of plasmid, cosmid, YAC or viral vector can be used to prepare the recombinant DNA construct which can be introduced directly into the tissue site. Alternatively, viral vectors can be used which selectively infect the desired target cell. Vectors for use in the practice of the invention include any eukaryotic expression vectors, including but not limited to viral expression vectors such as those derived from the class of retroviruses, adenoviruses or adeno-associated viruses.
[0093] In a preferred embodiment, the recombinant vector of the invention is an eukaryotic expression vector.
[0094] In another specific embodiment, the present invention comprises delivering the double PTM of the invention to a target cell. Various delivery systems are known and can be used to transfer the compositions of the invention into cells, e. g. encapsulation in liposomes, microparticles, microcapsules, recombinant cells capable of expressing the composition, receptor-mediated endocytosis, construction of a nucleic acid as part of a retroviral, adenoviral, adeno-associated viral or other vector, injection of DNA, electroporation, calcium phosphate mediated transfection, etc. In this case, the PTM which may be in any form used by one skilled in the art, for example, an RNA molecule, or a DNA vector which is transcribed into a RNA molecule, wherein said PTM binds to a pre-mRNA and mediates a double trans-splicing reaction resulting in the formation of a chimeric RNA comprising a portion of the PTM molecule spliced to a portion of the pre-mRNA. The present invention also concerns a cell comprising the double PTM of the invention, or the recombinant vector of comprising the double PTM of the invention.
[0095] In a preferred embodiment, the cell comprising the double PTM or the recombinant vector comprising the double PTM is an eukaryotic cell.
[0096] The compositions and methods can be used to provide a gene encoding a functional biologically active molecule to cells of an individual with an inherited genetic disorder where expression of the missing or mutant gene product produces a normal phenotype.
[0097] Specifically, the compositions and methods can be used to replace in vitro a mutated endogenous exon of the DMD gene within a cell comprising contacting the cellular pre-mRNA of the DMD gene with the double PTM of the present invention, under conditions in which the nucleotide sequence to be trans-spliced is trans-spliced to the target pre-mRNA of the DMD gene to form a chimeric mRNA within the cell. Said mutated exon is preferably exon 23 of the DMD gene.
[0098] To go further, the present invention also discloses a method for in vivo correcting a DMD genetic defect in a subject, comprising administering to said subject the double PTM of the invention, or the vector comprising the double PTM of the invention, or the cell comprising the double PTM of the invention.
[0099] More specifically, the present invention discloses a method for correcting at least one genetic mutation present in at least one endogenous mutated exon of the DMD gene in a subject in need thereof, comprising administering to said subject the double PTM of the invention.
[0100] Preferably, said endogenous mutated exon is exon 23 of the DMD gene, or exon 70 of the DMD gene, and the nucleotide sequence to be trans-spliced comprises at least the exon 23 of the DMD gene (SEQ ID NO 8 for the mouse gene, SEQ ID NO 60 for the human gene), or at least the exon 70 of the DMD gene (SEQ ID NO 72 for the mouse gene, SEQ ID NO 73 for the human gene).
[0101] In other words, the present invention discloses a double PTM, a vector comprising it, or a cell comprising it, for their use for correcting a DMD genetic defect in a subject in need thereof, or, more precisely, for their use for correcting at least one genetic mutation present in at least one of the DMD gene in a subject in need thereof, wherein, preferably said endogenous mutated exon is exon 23 of the DMD gene or exon 70 of the DMD gene, and the nucleotide sequence to be trans-spliced comprises at least the exon 23 of the DMD gene (SEQ ID NO 8 for the mouse gene, SEQ ID NO 60 for the human gene), or at least the exon 70 of the DMD gene (SEQ ID NO 72 for the mouse gene, SEQ ID NO 73 for the human gene).
[0102] The present invention also provides pharmaceutical compositions comprising an effective amount of the double PTM of the invention, and a pharmaceutically acceptable carrier. In a specific embodiment, the term "pharmaceutically acceptable" means approved by a regulatory agency of the Federal or a state government or listed in the U.S. Pharmacopeia or other generally recognized pharmacopeia for use in animals, and more particularly in humans. The term "carrier" refers to a diluent, adjuvant, excipient, or vehicle with which the therapeutic is administered. Examples of suitable pharmaceutical carriers are described in "Remington's Pharmaceutical sciences" by E. W. Martin.
[0103] Eventually, the present invention is drawn to a method for treating the Duchenne muscular dystrophy in a subject in need thereof, comprising administering to said subject the pharmaceutical composition comprising the double PTM of the invention. Preferably, the nucleotide sequence to be trans-spliced comprises at least an exon of the DMD gene, for example the exon 23 of the DMD gene (SEQ ID NO 8 or SEQ ID NO 60), or the exon 70 of the DMD gene (SEQ ID NO 72 or SEQ ID NO 73). In other words, the present invention covers the double PTM of the invention for its use for treating the Duchenne muscular dystrophy in a subject in need thereof. Preferably, the nucleotide sequence to be trans-spliced is at least an exon of the DMD gene, for example the exon 23 of the DMD gene (SEQ ID NO 8 or SEQ ID NO 60) or the exon 70 of the DMD gene (SEQ ID NO 72 or SEQ ID NO 73).
[0104] In a specific embodiment, it may be desirable to administer the pharmaceutical compositions of the invention locally to the area in need of treatment, i. e. in the muscles. This may be achieved by, for example, and not by way of limitation, local infusion during surgery, loco-regional infusion under high pressure in a limb where the arterial and venous blood flux is intermittently interrupted by a tourniquet, topical application, e. g., in conjunction with a wound dressing after surgery, by injection, by means of a catheter, by means of a suppository, or by means of an implant, said implant being of a porous, non-porous, or gelatinous material, including membranes, such as sialastic membranes, or fibers. Other controled release drug delivery systems exist, such as nanoparticles, matrices such as controlled-release polymers, hydrogels. The double PTM will be administered in amounts which are effective to produce the desired effect in the targeted cell. Effective dosages of the double PTMs can be determined through procedures well known to those in the art which address such parameters as biological half-life, bioavailability and toxicity. The amount of the composition of the invention which will be effective will depend on the severity of the DMD being treated, and can be determined by standard clinical techniques.
[0105] The double PTM can also be delivered to cells or stem cells ex vivo which can, in a second step after ex vivo correction, be transferred as a cell transplant to an individual with the goal of correcting an organ or an individual affected by a genetic disease through cell therapy.
[0106] In a second aspect, the present invention is drawn to a nucleic acid molecule dedicated to simple trans-splicing as it is described in the following examples.
[0107] In this particular embodiment, the nucleic acid molecule of the invention comprises:
[0108] a) one target binding domain (AS) that target binding of the nucleic acid molecule to the pre-mRNA of the dystrophin gene (DMD),
[0109] b) a 3' splice region comprising a branch point, a polypyrimidine tract and a 3' splice acceptor site,
[0110] c) a spacer sequence that separates the 3' splice region from the target binding domain AS,
[0111] d) a nucleotide sequence to be trans-spliced to the target pre-mRNA wherein said nucleotide sequence encodes at least a part of the DMD polypeptide.
[0112] This nucleic acid molecule is hereafter referred to as "simple trans-splicing molecule", or "simple PTM" of the invention.
[0113] The different parts of this simple trans-splicing molecule (3' splice acceptor site, branch point, polypyrimidine tract, spacer sequence) are the same as described above for the PTM mediating double trans-splicing, that is, for example, SEQ ID NO 29 for the 3' splice acceptor site, SEQ ID NO 25 for the branch point, SEQ ID NO 28 for the polypyrimidine tract, SEQ ID NO 23 for the 3' end spacer.
[0114] As a matter of fact, the present invention shows also here for the first time a simple PTM enabling efficient exon(s) replacement (for example replacing anormal exon 70 of the human DMD gene, by the normal one), importantly demonstrating that the replacement of specific exon(s) is possible and efficient also with a "simple" trans-splicing technology (cf. example 2).
[0115] In this simple PTM, the nucleotide sequence to be trans-spliced to the pre-mRNA of the dystrophin gene (DMD) comprises preferably one or several exon(s) of the normal DMD gene, more preferably the exon 23 or any one of the exons 59 to 79 of the DMD gene. Even more preferably, the sequence to be trans-spliced is the exon 70 of the DMD gene, or the cDNA from all the exons 59 to 79 of the DMD gene, as shown in example 2 below.
[0116] More precisely, in the simple PTM of the present invention, the nucleotide sequence to be trans-spliced is either the sequence of exon 23 of the normal DMD gene (that is SEQ ID NO 8 for the mouse gene or SEQ ID NO 60 for the human gene), or the sequence of exon 70 of the DMD gene (that is, SEQ ID NO 72 for the human gene, and SEQ ID NO 73 for the mouse gene), or the cDNA corresponding to exons 59 to 79 of the DMD gene (that is SEQ ID NO 69 for the mouse gene or SEQ ID NO 70 for the human gene).
[0117] In a preferred embodiment of the invention, the 5'end target binding domain AS of the simple PTM of the invention targets the binding of the nucleic acid to intron 22 of the pre-mRNA of the DMD gene (SEQ ID NO 11 for the mouse gene, or SEQ ID NO 61 for the human gene), or to intron 58 of the pre-mRNA of the DMD gene (SEQ ID NO: 66 for the mouse gene, SEQ ID NO 67 for the human gene).
[0118] In a more preferred embodiment, the 5' end target binding domain AS of the simple PTM of the invention has the sequence SEQ ID NO 68 (for targeting the PTM to intron 58 of the DMD gene) or SEQ ID NO 13, SEQ ID NO 14, SEQ ID NO 15, or SEQ ID NO 58 (for targeting the PTM to intron 22 of the DMD gene).
[0119] In an even more preferred embodiment, the simple trans-splicing molecule of the invention has the SEQ ID NO 71.
[0120] In a specific embodiment, the present invention is drawn to a recombinant vector comprising the simple PTM previously described. More particularly, the simple PTM of interest may be recombinantly engineered into a variety of host vector systems that also provide for replication of the DNA in large scale and contain the necessary elements for directing the transcription of the simple PTM. The use of such a construct to transfect target cells in the patient will result in the transcription of sufficient amounts of simple PTMs that will form complementary base pairs with the endogenously expressed pre-mRNA targets, such as for example, DMD pre-mRNA target, and thereby facilitate a trans-splicing reaction between the complexed nucleic acid molecules. Such a vector can remain episomal or become chromosomally integrated, as long as it can be transcribed to produce the desired RNA, i. e., PTM. Such vectors can be constructed by recombinant DNA technology methods standard in the art. Vectors comprising the simple PTM of interest can be plasmid, viral, or others known in the art, used for replication and expression in mammalian cells. Expression of the simple PTM can be regulated by any promoter/enhancer sequences known in the art to act in mammalian, preferably human cells. Such promoters/enhancers can be inducible or constitutive. Such promoters include but are not limited to: the SV40 early promoter region, the promoter contained in the 3' long terminal repeat of Rous sarcoma virus, the herpes thymidine kinase promoter, the regulatory sequences of the metallothionein gene, the viral CMV promoter, the human chorionic gonadotropin-P promoter, etc. Any type of plasmid, cosmid, YAC or viral vector can be used to prepare the recombinant DNA construct which can be introduced directly into the tissue site. Alternatively, viral vectors can be used which selectively infect the desired target cell. Vectors for use in the practice of the invention include any eukaryotic expression vectors, including but not limited to viral expression vectors such as those derived from the class of retroviruses, adenoviruses or adeno-associated viruses.
[0121] In a preferred embodiment, the recombinant vector of the invention is an eukaryotic expression vector.
[0122] In another specific embodiment, the present invention comprises delivering the simple PTM of the invention to a target cell. Various delivery systems are known and can be used to transfer the compositions of the invention into cells, e. g. encapsulation in liposomes, microparticles, microcapsules, recombinant cells capable of expressing the composition, receptor-mediated endocytosis, construction of a nucleic acid as part of a retroviral, adenoviral, adeno-associated viral or other vector, injection of DNA, electroporation, calcium phosphate mediated transfection, etc. In this case, the simple PTM which may be in any form used by one skilled in the art, for example, an RNA molecule, or a DNA vector which is transcribed into a RNA molecule, wherein said simple PTM binds to a pre-mRNA and mediates a simple trans-splicing reaction resulting in the formation of a chimeric RNA comprising a portion of the simple PTM molecule spliced to a portion of the pre-mRNA. The present invention also concerns a cell comprising the simple PTM of the invention, or the recombinant vector of comprising the simple PTM of the invention.
[0123] In a preferred embodiment, the cell comprising the simple PTM or the recombinant vector comprising the simple PTM is an eukaryotic cell.
[0124] The simple PTM of the invention appears to be a very interesting tool to target DMD patients independently of their DMD mutation. By enabling to restore mutated exons of the DMD gene, this molecule appears to be also an efficient tool to treat DMD patients.
[0125] Therefore, the present invention is also drawn to a method for treating a patient suffering from the Duchenne muscular dystrophy comprising administering to said patient a pharmaceutical composition comprising the simple trans-splicing molecule of the invention or the recombinant vector comprising it, or the cell comprising them. In other words, the present invention is drawn to the simple trans-splicing molecule of the invention, or the recombinant vector comprising it, or the cell comprising them for their use in a pharmaceutical composition for treating the Duchenne muscular dystrophy in a subject in need thereof.
[0126] More precisely, the present invention also concerns a method for correcting a DMD genetic defect in a subject, comprising administering to said subject the simple trans-splicing molecule of the invention, or the recombinant vector comprising it, or the cell comprising them. In other words, the invention is drawn to the simple trans-splicing molecule of the invention, or the recombinant vector comprising it, or the cell comprising them, for their use for correcting a DMD genetic defect in a subject in need thereof.
[0127] Preferably, the nucleotide sequence to be trans-spliced comprises at least one exon of the DMD gene, and comprises more preferably at least exon 23 of the normal DMD gene (that is SEQ ID NO 8 for the mouse gene or SEQ ID NO 60 for the human gene), or exon 70 of the DMD gene (that is, SEQ ID NO 72 for the human gene, or SEQ ID NO 73 for the murine gene) or any exon chosen among exons 59 to 79 of the DMD gene. In a preferred embodiment, it comprises the cDNA from exon 59 to 79 of DMD gene (that is, SEQ ID NO 70 for the human gene, and SEQ ID NO 69 for the mouse gene).
[0128] The present invention also concerns an in vitro method of replacing at least one mutated endogenous exon of the DMD gene within a cell, comprising contacting the cellular pre-mRNA of the DMD gene with the simple trans-splicing molecule of the invention, under conditions in which the nucleotide sequence to be trans-spliced is trans-spliced to the target pre-mRNA of the DMD gene to form a chimeric mRNA within the cell. Preferably, said mutated exon is exon 23 of the DMD gene, or exon 70 of the DMD gene or any exon chosen among exons 59 to 79 of the DMD gene. More preferably, the nucleotide sequence to be trans-spliced to the target pre-mRNA of the DMD gene thus comprises at least exon 23, or exon 70 of the DMD gene, or any exon chosen among exons 59 to 79 of the DMD gene. Even more preferably the nucleotide sequence to be trans-spliced to the target pre-mRNA of the DMD gene comprises the cDNA from exon 59 to 79 of DMD human gene.
[0129] The present invention also concerns a method for correcting at least one genetic mutation present in at least one exon of the DMD gene in a subject in need thereof, comprising administering to said subject the simple trans-splicing molecule of the invention. Preferably, said at least one exon of the DMD gene is exon 23 or exon 70, and the nucleotide sequence to be trans-spliced comprises at least the exon 23 of the DMD gene (that is SEQ ID NO 8 for the mouse gene or SEQ ID NO 60 for the human gene), or the exon 70 of the DMD gene (that is, SEQ ID NO 72 for the human gene, or SEQ ID NO 73 for the murine gene), or the cDNA from exon 59 to exon 79 of the normal DMD human gene (SEQ ID NO 70).
[0130] In this case, in other words, the invention is drawn to the simple trans-splicing molecule of the invention, or the recombinant vector comprising it, or the cell comprising them, for their use for correcting at least one genetic mutation present in exon 23 or exon 70 of the normal human DMD gene in a subject in need thereof, wherein the nucleotide sequence to be trans-spliced comprises at least the exon 23 of the normal DMD gene (that is SEQ ID NO 8 for the mouse gene or SEQ ID NO 60 for the human gene), or the exon 70 of the normal DMD gene (that is, SEQ ID NO 72 for the human gene, or SEQ ID NO 73 for the mouse gene), or the cDNA from exon 59 to exon 79 of the normal DMD human gene (SEQ ID NO 70).
[0131] The present invention also provides pharmaceutical compositions comprising an effective amount of the simple PTM of the invention, and a pharmaceutically acceptable carrier. In a specific embodiment, the term "pharmaceutically acceptable" means approved by a regulatory agency of the Federal or a state government or listed in the U.S. Pharmacopeia or other generally recognized pharmacopeia for use in animals, and more particularly in humans. The term "carrier" refers to a diluent, adjuvant, excipient, or vehicle with which the therapeutic is administered. Examples of suitable pharmaceutical carriers are described in "Remington's Pharmaceutical sciences" by E. W. Martin.
[0132] In a specific embodiment, it may be desirable to administer the pharmaceutical compositions of the invention locally to the area in need of treatment, i. e. in the muscles. This may be achieved by, for example, and not by way of limitation, local infusion during surgery, loco-regional infusion under high pressure in a limb where the arterial and venous blood flux is intermittently interrupted by a tourniquet, topical application, e. g., in conjunction with a wound dressing after surgery, by injection, by means of a catheter, by means of a suppository, or by means of an implant, said implant being of a porous, non-porous, or gelatinous material, including membranes, such as sialastic membranes, or fibers. Other controled release drug delivery systems exist, such as nanoparticles, matrices such as controlled-release polymers, hydrogels. The simple PTM will be administered in amounts which are effective to produce the desired effect in the targeted cell. Effective dosages of the simple PTMs can be determined through procedures well known to those in the art which address such parameters as biological half-life, bioavailability and toxicity. The amount of the composition of the invention which will be effective will depend on the severity of the DMD being treated, and can be determined by standard clinical techniques.
[0133] The simple PTM can also be delivered to cells or stem cells ex vivo which can, in a second step after ex vivo correction, be transferred as a cell transplant to an individual with the goal of correcting an organ or an individual affected by a genetic disease through cell therapy.
EXAMPLES
1. Double Trans-Splicing Molecule of the Invention
1.1. Materials and Methods
Plasmids Constructions
[0134] The murine DMD minigene target (3993 bp (SEQ ID NO 39)) comprising exons E22, E23, and E24 and the natural E23 flanking intronic sequences was constructed by PCR amplification from mdx genomic DNA and subcloned in pSMD2 into KpnI site. An ATG in Kozack ACCACCATGG context and a STOP codon were introduced at both sides of the minigene.
[0135] The E24 for the TS and EX molecules (114 bp (SEQ ID NO 10)) was amplified from the DMD minigene. The different domains of the TS and EX molecules detailed in Results section were constructed by PCR and subcloned in pSMD2 into HindIII and EcoRI between the CMV promoter and the polyA signal. Antisens sequences AS bind to DMD intron 22: AS1 targets nucleotides -763 to -614 (SEQ ID NO 13); AS2, -463 to -314 (SEQ ID NO 14); AS3 -159 to -10 (SEQ ID NO 15); AS3bis -159 to +5 (SEQ ID NO 58) (where nucleotide +1 is the first E23 nucleotide). The second antisens domains AS' bind to DMD intron 23: AS4, +1801 to +1950 (SEQ ID NO 16); AS5, +2101 to +2250 (SEQ ID NO 17); AS6, +2401 to +2550 (SEQ ID NO 18); AS7, -5 to +145 (SEQ ID NO 19); AS8, +151 to +300 (SEQ ID NO 20) (where nucleotide +1 is the first nucleotide of intron 23).
[0136] All expression cassettes are under the control of the strong CMV promoter and a polyA signal and were verified by sequencing.
Cell Culture and Transfection
[0137] Mouse embryonic fibroblast NIH3T3 cells were maintained in DMEM (Invitrogen) supplemented with 10% heat-inactivated FBS (Invitrogen), 100 units/ml penicillin, and 100 μg/ml streptomycin. For transfections, cells were grown to 70% confluence in 12-well plates and exposed to the DNA/Lipofectamine 2000 reagent (Invitrogen) complex for 5 h in DMEM before being returned to normal culture medium. Typically, 0.5 μg of DMD minigene and 1.5 μg of TS or EX molecules DNA were used in each transfection. Cells were routinely analyzed 72 h after transfection.
RT-PCR Analysis
[0138] Total RNA was isolated from transfected cells by using RNAeasy extraction kit (Qiagen). Reverse transcription was performed on 200 ng of RNA by using the Superscript II (Invitrogen) and the reverse primer pSMD2-R1 (see below) at 10 min at 25° C., 50 min at 42° C., and a final step of 5 min at 95° C. To detect non-repaired and repaired DMD transcripts, reverse transcribed RNA was amplified by PCR under the following conditions: 95° C. for 5 min, 30 cycles of 30 s at 95° C., 1 min at 56° C., 45 s to 1 min at 72° C., and a final step of 7 min at 72° C.
[0139] The sequences of the primers were as followed: E22-F GACACTTTACCACCAATGCGC (SEQ ID NO 36) (Primer A on FIGS. 1A-B and 2A-B), pSMD2-R1 CTTTCTGATAGGCAGCCTGC (SEQ ID NO 37) (Primer B on FIG. 1A-B) and pSMD2-R5 CTCACCCTGAAGTTCTCAGG (SEQ ID NO 38) (Primer C on FIG. 2A-B). RT-PCR products were separated by electrophoresis in 2% agarose gels with ethidium bromide and sequenced.
Quantitative Real-Time RT-PCR
[0140] mRNA levels were measured by absolute quantitative real-time RT-PCR method using Absolute SYBR Green Rox Mix (Thermo scientific). Two positive control DMD cDNA fragments, E22-E23-E24 and E22-E24-E24, were cloned into the pCR®2.1-TOPO®. As a reference samples, those plasmids were 10-fold serially diluted (from 107 to 103 copies) and used to generate standard curves. Real-time PCR was performed and analyzed on a DNA Engine Opticon 2 (Bio-Rad). In each experiment, duplicates of standard dilution series of control plasmids and first strand cDNA generated by the Superscript II (Invitrogen) from 200ng of total RNA were amplified by specific primers. Primers for E23, E23-F AGATGGCCAAGAAAGCACC (SEQ ID NO: 32) and E23-R CTTTCCACCAACTGGGAGG, (SEQ ID NO: 33) were used to measure non-repaired DMD transcript; and primers for E24-E24 junction, E24-F TGAAAAAACAGCTCAAACAATGC (SEQ ID NO: 34) and E24-R AGCATCCCCCAGGGCAGGC (SEQ ID NO: 35), for the repaired transcript.
1. 2. Results
Design of Trans-Splicing Molecules (TS Molecules)
[0141] In ExChange molecules, the replacing exon is flanked by artificial intronic sequences with strong acceptor and donor splice sites, which are connected to antisense sequences designed to anneal the target mRNA. Annealing is crucial to permit the trans-splicing reaction, although it is not enough. Ideally, the site of annealing must disturb the definition of the targeted exon in the parental pre-messenger while enhancing cross-splicing in between the two independent mRNAs. In the case of ExChange, there are more constraints since two trans-splicing reactions must be synchronized at both edges of the targeted exon.
[0142] The murine model for DMD, the mdx mouse, carries a nonsense mutation in exon 23 (E23m: SEQ ID NO 9) of the dystrophin gene. In order to locate the best site of annealing in intron 22, upstream the mutated exon, three trans-splicing (TS) molecules for 3' replacement only differing in their binding domains were designed (FIG. 1A). Antisense sequences of about 150 nucleotides (AS1=SEQ ID NO 13, AS2=SEQ ID NO 14 and AS3=SEQ ID NO 15) were chosen to match either to the 5' end, the middle or the 3' end of intron 22 (SEQ ID NO 11). The idea was to test whether getting the TS molecule close to its target, 5' donor splice site of intron 22, or at the opposite, masking the 3' acceptor splice site, would facilitate trans-splicing. In the three constructions, the artificial intron included a spacer sequence (SEQ ID NO 59), a strong conserved yeast branch point sequence (SEQ ID NO 25), a polypyrimidine tract (SEQ ID NO 28), and a canonical 3' acceptor splice site (SEQ ID NO 29). To facilitate the readout, it was also decided to employ exon 24 (E24 (SEQ ID NO 10)) in the TS molecule instead of the normal version of exon 23 (E23). Indeed, E24 is smaller than E23 (114 versus 213 bp) allowing unequivocal distinction by RT-PCR of repaired mRNA (E22-E24-E24) from non-repaired parental transcripts (E22-E23m-E24). Three TS molecules for 3' replacement were thus constructed: AS1-E24 (SEQ ID NO 40), AS2-E24 (SEQ ID NO 41), AS3-E24 (SEQ ID NO 42). As control, a trans-splicing molecule with no binding domain (AS-, SEQ ID NO 43) was used.
[0143] To facilitate the analysis of DMD splicing in tissue culture, a DMD reporter gene made of a genomic fragment of 3993 bp comprising E22 to E24 with full-length natural introns was made (SEQ ID NO 39). Cis- and trans-splicing patterns are illustrated in FIG. 1B. An RT-PCR strategy was designed to detect specifically RNA resulting from cis- and trans-splicing events by using a forward primer E22-F (SEQ ID NO 36) (arrow A in FIG. 1B) specific for E22, and a reverse primer pSMD2-R1 (SEQ ID NO 37) (arrow B) specific for a sequence upstream the polyA signal in DMD minigene and TS molecules. Importantly, these primers also allowed discriminating E22-E24 amplicons resulting from either trans-splicing or exon skipping.
3' Replacement in DMD Transcripts
[0144] DMD reporter minigene and TS plasmids were cotransfected in the mouse embryonic fibroblast NIH3T3 cell-line. Cells were harvested 72 h after transfection, and total RNA was isolated. Cis- and trans-spliced RNA patterns were assessed by RT-PCR. As expected, samples that received only the DMD minigene displayed a single 638 bp amplicon corresponding to the cis-spliced DMD transcript E22-E23m-E24 (Ctrl in FIG. 1C). Also, cDNAs from cells transfected with both DMD minigene and trans-splicing constructs (AS2-E24) gave no PCR products when reverse transcription was omitted (RT-AS2), ensuring about the specificity of the present assay. In the presence of U7-SD23-BP22 (U7) plasmids described to induce E23 skipping (Goyenvalle et al., 2004), a 425 bp band corresponding to E22-E24 transcript from cis-splicing was detected.
[0145] In samples that received DMD minigene and TS plasmids, a product of 310 bp was generated, corresponding specifically to the trans-spliced E22-E24 variant, and not to an exon skipping product as it was obtained with U7. In the presence of either AS1-E24 (SEQ ID NO 40) or AS2-E24 (SEQ ID NO 41), the E22-E23m-E24 amplicon corresponding to the parental DMD minigene had almost entirely disappeared thus confirming that, here, trans-splicing efficacies were nearly complete. Importantly, trans-splicing did not occur when AS was removed (AS-) from TS constructs demonstrating that this reaction required close interaction in between the two strands of mRNA to combine. The AS3-E24 TS molecule (SEQ ID NO: 42) appeared to be less efficient. Surprisingly, extending AS3 in order to cover the 3' acceptor site of E23m did not improve the trans-splicing reaction (not shown). These experiments show that trans-splicing could not do without AS sequences, although getting closer the two mRNAs is not sufficient.
mRNA Repair By Using ExChange
[0146] To test the possibility of mRNA repair by ExChange, several ExChange (EX) molecules AS-E24-AS' based on the efficient TS molecules mentioned above were developed, and modified to bind both intron 22 and intron 23 of the DMD reporter minigene (FIG. 2A). The EX molecules contained the same elements as previously described in AS1-E24 and AS2-E24 TS molecules followed by a 5' donor splice site (SEQ ID NO 30) and a second 150 nt antisense targeting intron 23 (SEQ ID NO 12). Five antisenses, AS 4 (SEQ ID NO 16), AS5 (SEQ ID NO 17), AS6 (SEQ ID NO 18), AS7 (SEQ ID NO 19) and AS8 (SEQ ID NO 20), were selected within intron 23 (SEQ ID NO 12), which spans over 2607 bp. The following spacer sequences were used: for the spacer sequence that separates the 3' splice region from the 5'-end target binding domain AS, spacer 2 (SEQ ID NO 24, 42 nucleotides) was used, and for the spacer sequence that separates the 5' splice region from the 3'-end target binding domain AS', spacer 1 (SEQ ID NO 23, 34 nucleotides) was used.
[0147] Finally, the following PTM were constructed: AS1-E24-AS4 (SEQ ID NO 44), AS1-E24-AS5 (SEQ ID NO 45), AS1-E24-AS6 (SEQ ID NO 46), AS1-E24-AS7 (SEQ ID NO 47), AS2-E24-AS4 (SEQ ID NO 48), AS2-E24-AS5 (SEQ ID NO 49), AS2-E24-AS6 (SEQ ID NO 50), AS2-E24-AS7 (SEQ ID NO 51), AS2-E24-AS8 (SEQ ID NO 52) and AS2-E24-2XAS4 (SEQ ID NO 53).
[0148] As previously, EX constructs and DMD minigene were cotransfected in NIH3T3 cell-line. Cells were harvested 72 h after transfection, and total RNA was extracted. To detect specifically RNA resulting from cis-splicing and/or exon exchange events, a forward primer E22-F (SEQ ID NO 36) (arrow A in FIG. 2A-B) specific of E22, and a reverse primer pSMD2-R5 (SEQ ID NO 38) (arrow C) specific of a sequence only present in the DMD minigene upstream its polyA signal were used. Targeting of AS-E24-AS' in the DMD reporter pre-mRNA is illustrated in FIG. 2A and expected sizes of the various amplification products shown in FIG. 2B.
[0149] A RT-PCR product of 408 bp was detected in samples transfected with AS2-E24-AS' plasmids (FIG. 2C). Direct sequencing confirmed that this product corresponded to the exchanged mRNA variant E22-E24-E24 (FIG. 2D). This product was absent when one of the two antisens was lacking, showing that co-targeting of intron 22 and intron 23 is crucial for ExChange. Among the antisense combinations we tried, levels of the 408 bp band were stronger with AS2-E24-2XAS4 (SEQ ID NO 53), -AS7 (SEQ ID NO 51) and -AS8 (SEQ ID NO 52). Interestingly, the AS2-E24-2XAS4 molecule (SEQ ID NO 53), which carried two AS4, was more efficient than its single AS4 counterpart. In AS2-E24-AS7 sample, a supplementary band of 294 bp was detected corresponding to E22-E24 transcript generated by exon 23 skipping. This was not surprising considering that AS7 bound the 5' donor splice site of intron 23 and would mask its recognition by the spliceosome. It is likely that AS4, AS7 and AS8 brought back EX molecules closer to E23 than AS5 and AS6 suggesting that a tight framing is essential for efficient ExChange.
Optimization of ExChange Efficacy by Adding up Intronic Splice Enhancers
[0150] In order to improve the ExChange reaction, the G-rich intronic splice enhancer (ISE, SEQ ID NO 26) from the human GH-1 gene was added upstream the 3' acceptor site of AS2-E24-AS4 and AS2-E24-AS8 (McCarthy & Phillips, 1998) and the DISE sequence from the rat FGFR2 gene downstream the 5' donor site (SEQ ID NO 27) (Kierlin-Duncan & Sullenger, 2007) (FIG. 3A). As shown in FIG. 3B, RT-PCR analysis revealed that insertion of the DISE sequence in AS2-E24-AS4 (SEQ ID NO 54) and AS2-E24-AS8 (SEQ ID NO 55) molecules increased significantly the 408 bp band corresponding to the E22-E24-E24 mRNA variant, while addition of ISE sequence did not enhance the ExChange efficacy. As expected, no ExChange was observed with control vectors lacking the downstream AS': AS2-ISE-E24 and AS2-E24-DISE. FIG. 3C shows ExChange efficacy of various vectors by using quantitative RT-PCR. The AS2-DISE-E24-AS4 (SEQ ID NO 54) molecule allowed obtaining 53% of exon exchange. Its efficacy was improved by about 7.5 folds when compared to its counterpart AS2-E24-AS4 (SEQ ID NO 48) lacking the DISE motive.
[0151] Interestingly, introduction of two redundant downstream AS' (here AS4) improved ExChange efficacy, which was about 30%. However, AS2-E24-DISE-2XAS4 (SEQ ID NO 57) was not more efficient than AS2-DISE-E24-AS4 (SEQ ID NO 54) (data not shown).
2. Simple Trans-Splicing Molecule of the Invention
2.1. Materials and Methods
Plasmid Constructions
[0152] The different domains of the TSM molecules were constructed by PCR and subcloned into pSMD2. Human dystrophin exons 59 to 79 until the STOP codon (2390 bp, see SEQ ID NO 70) were amplified from human myotubes cDNA while antisense sequences from human genomic DNA. Antisense sequence binds to dystrophin intron 58: nucleotides -445 to -295 (where nucleotide +1 is the first E59 nucleotide, SEQ ID NO 68). The trans-splicing cassettes were subcloned into a plasmid pRRL-cPPT-mcs-WPRE under hPGK promoter (Zufferey R et al, 1997). All the constructs were verified by sequencing.
Sequences
[0153] Artificial intron included a spacer sequence, a strong conserved yeast branch point sequence (BP), a polypyrimidine tract (PPT) and a canonical 3' splice acceptor site (SA): the same sequences are already described for the double PTM of example 1.
Cell Cultures
[0154] Primary DMD myoblast cell cultures were established from explants of biceps as described previously (Mouly V et al, 1993), in accordance with French ethics legislation. DMD and control myoblasts CHQ (Edom et al, 1994) were grown in Skeletal Muscle Cell Growth Medium (PromoCell). To induce differentiation, cultures were switched to DMEM 2% horse serum with apotransferrin (100 μg/ml) and insulin (10 μg/ml). All cultures were grown in humidified incubators at 37° C. in 5% CO2.
Lentiviral Productions
[0155] Lentiviral vectors pseudotyped with the VSV-G protein were produced by transient quadri-transfection into 293T cells were determined by transduction of HCT116 cells and assayed by quantitative real-time PCR on genomic DNA [Charrier S et al, 2005). Titration of the lentivirus is expressed as viral genome/mL (vg/mL) (ranging from 2.5×109 to 5×109 vg/mL). 5×106 vg were used to transduce 4×105 myoblasts plated the day before in 24 wells tissue culture dishes in 500 μL of DMEM supplemented with 10% FCS. Four hours post-transduction, the medium was diluted by adding 200 μL per well of previous medium. The dishes were incubated for 24 hr at 37° C. and 5% CO2 before washing.
RT-PCR Analysis
[0156] Total RNA was extracted from transduced myotubes with TRIzol reagent (Invitrogen). Five microgram of RNA was reverse transcribed using SuperScript III First-Strand Synthesis SuperMix (Invitrogen) and the reverse primer Pst1-WPRE-Ro, AACTGCAGCAGGCGGGGAGGCGGCCCAAAG (Ro on FIG. 2A). cDNAs were subjected to nested PCR amplification with Phusion High-Fidelity PCR Master Mix with GC buffer (Finnzymes) under the following conditions: 95° C. for 5 min, 20 cycles of 30 s at 95° C., 2 min at 56° C., 45 s to 1 min at 72° C., and a final step of 7 min at 72° C., using external primers E58-Fo CATGAGTACTCTTGAGACTG (Fo on FIG. 2A) and WPRE-Ro AGCAGCGTATCCACATAGCG (Ro). Five microliters of each of these reactions was then reamplified for 30 cycles using the internal primers E58-Fi, AGGACTAGAGAAACTCTACC (Fi), and WPRE-Ri, TTGTCGACCAGCGTTTCTAG (Ri). RT-PCR products were separated by electrophoresis in 1% agarose gels with ethidium bromide and sequenced.
Protein Analysis
[0157] Forty μg of protein were loaded onto NuPAGE® Novex 4-12% Bis-Tris Gel (Invitrogen), electrophoresed, blotted onto nitrocellulose membranes and probed with 1:50 NCL-DYS1 or NCL-DYS2 (NovoCastra), followed by incubation with 1:15000 horseradish peroxidase-conjugated secondary antibody (Jackson ImmunoResearch) and SuperSignal® West Pico Analysis System (Thermo Scientific).
2.2. Results
[0158] The aim of this work was to repair by simple trans-splicing dystrophin pre-mRNAs carrying any genetic anomalies present between exons 59 to 79, which represents 8% of Duchenne patients. These patients are not treatable by exon skipping therapy as these exons are indispensable for the protein function.
[0159] The simple trans-splicing molecule contains a 150 nucleotides antisense sequence complementary to intron 58 (SEQ ID NO 68), an artificial intron included a spacer sequence (SEQ ID NO 23), a strong conserved yeast branch point sequence (BP) (SEQ ID NO 25), a polypyrimidine tract (PPT) (SEQ ID NO 28) and a canonical 3' splice acceptor site (SA) (SEQ ID NO 29), and the normal human dystrophin cDNA from exon 59 to the exon 79 STOP codon (SEQ ID NO 70) (cf. FIG. 4). A RT-PCR strategy was designed to amplify specifically a 2443 bp product from mRNA resulting from trans-splicing events by using forward primers, E58-Fo and E58-Fi (arrows Fo/Fi in FIG. 5A), specific for E58, and reverse primers, WPRE-Ro and WPRE-Ri (Ro/Ri in FIG. 5A), specific for the WPRE element present on trans-splicing molecules.
[0160] Duchenne muscular dystrophy patient myoblasts carrying a non-sense mutation in exon 70 were transduced with lentivirus expressing trans-splicing molecules. After differentiation, myotubes were harvested, total RNA isolated and trans-spliced dystrophin transcripts investigated by specific RT-PCR. As expected, samples that did not receive lentivirus gave no PCR products (lane "-" in FIG. 5B). In samples that express trans-splicing molecules (TsM), a product of 2443 bp was generated corresponding specifically to the trans-spliced DMD variant as confirmed by direct sequencing of the amplicon (data not shown).
[0161] In order to detect specifically rescued dystrophin encoded by trans-spliced transcripts, Western blotting with the NCL-DYS1 monoclonal antibody that recognizes spectrin-like repeats R8 to R10 was used. Consistent with the generation of trans-spliced transcripts, the full-length 427 kD dystrophin protein was readily detected by Western blot on transduced myotubes extracts (the two lanes TsM in FIG. 6), whereas no band is present in the non treated DMD cells ("-").
BIBLIOGRAPHIC REFERENCES
[0162] Chao et al., 2003 Nat Med 9:1015-1019
[0163] Charrier S, et al. 2005. Gene Ther 12: 597-606.
[0164] Chiara & Reed 1995, Nature 375: 510
[0165] Coady et al., 2008 PLoS ONE 3:e3468
[0166] Denti et al., 2006; Proc Natl Acad Sci USA 103:3758-3763.
[0167] Dingwell and Laskey, 1986, Ann. Rev. Cell Biol. 2: 367-390
[0168] Edom F, et al, 1994. Dev Biol 164: 219-229.
[0169] Eul et al., 1995, EMBO. R 14: 3226
[0170] Finta, C. et al., 2002 J : Biol Chem 277: 5882-5890
[0171] Goyenvalle et al., 2004 Science 306:1796-1799
[0172] Kierlin-Duncan & Sullenger, 2007 RNA 13:1317-1327
[0173] Liu et al., 2005 Hum Gene Ther 16:1116-1123.
[0174] Mansfield et al, RNA 2003 9:1290-1297
[0175] McCarthy & Phillips, 1998 Hum Mol Genet 7:1491-1496.
[0176] Mouly V, et al, 1993, Neuromuscul Disord 3: 371-377.
[0177] Shimizu et al., 1989, Proc. Nat'l. Acad. Sci. USA 86: 8020
[0178] Staley and Guthrie, 1998, Cell 92 : 315-326
[0179] Tacke et al., 1999, Curr. Opinion. Cell Biol. 11: 358-362
[0180] Tahara et al., 2004 Nat Med 10:835-841.
[0181] Takayuki Horiuchi and Toshiro Aigaki, Biol. Cell (2006) 98, 135-140
[0182] Vellard, M. et al. Proc. Nat'l. Acad. Sci., 1992 89: 2511-2515
[0183] Yokota et al., 2009 Ann Neurol June; 65(6) :667-76
[0184] Zufferey R, et al, 1997. Nat Biotechnol 15: 871-875.
Sequence CWU
1
1
731537DNAArtificial SequenceNucleotide sequence of the Pre-Trans-Splicing
molecule protein, exon 24 1atatattttc agatttaaaa agaataagta
tacaacatgg gatttttaga atcaacaaaa 60aaattagtct ttatatgtcc tcacatcaca
gaagtttctc ttactagtat ttaatttcca 120gacttaggct attaaaataa ctactcaata
ctcgagagat ctccgcggaa cattattata 180acgttgctcg aatactaact gatatctctt
cttttttttt ttccggaaaa cagaatcaca 240taaaaacctt acagaaatgg atggctgaag
ttgatgtttt cctgaaagag gaatggcctg 300ccctggggga tgctgaaatc ctgaaaaaac
agctcaaaca atgcagagta agaacagctc 360tttctttcca tgggttggcc tgaattctta
gttccatgta attcacaaaa tcaagttata 420attgtcttct tttctaaaat ttatattgaa
aatacatgca tcacagaaaa tttccctttc 480aatatgatta aaatgtggtt aataactaca
gatttaaaaa caattgacta aatataa 5372636DNAArtificial
SequenceNucleotide sequence of the Pre-Trans-Splicing molecule
protein, exon 23 2atatattttc agatttaaaa agaataagta tacaacatgg gatttttaga
atcaacaaaa 60aaattagtct ttatatgtcc tcacatcaca gaagtttctc ttactagtat
ttaatttcca 120gacttaggct attaaaataa ctactcaata ctcgagagat ctccgcggaa
cattattata 180acgttgctcg aatactaact gatatctctt cttttttttt ttccggaaaa
caggctctgc 240aaagttcttt gaaagagcaa caaaatggct tcaactatct gagtgacact
gtgaaggaga 300tggccaagaa agcaccttca gaaatatgcc agaaatatct gtcagaattt
gaagagattg 360aggggcactg gaagaaactt tcctcccagt tggtggaaag ctgccaaaag
ctagaagaac 420atatgaataa gcttcgaaaa tttcaggtaa gaacagctct ttctttccat
gggttggcct 480gaattcttag ttccatgtaa ttcacaaaat caagttataa ttgtcttctt
ttctaaaatt 540tatattgaaa atacatgcat cacagaaaat ttccctttca atatgattaa
aatgtggtta 600ataactacag atttaaaaac aattgactaa atataa
636371PRTMus musculusmisc_featureProtein sequence of the
murine dystrophin gene DMD, exon 23 3Ala Leu Gln Ser Ser Leu Lys Glu
Gln Gln Asn Gly Phe Asn Tyr Leu1 5 10
15Ser Asp Thr Val Lys Glu Met Ala Lys Lys Ala Pro Ser Glu
Ile Cys 20 25 30Gln Lys Tyr
Leu Ser Glu Phe Glu Glu Ile Glu Gly His Trp Lys Lys 35
40 45Leu Ser Ser Gln Leu Val Glu Ser Cys Gln Lys
Leu Glu Glu His Met 50 55 60Asn Lys
Leu Arg Lys Phe Gln65 70438PRTMus
musculusmisc_featureProtein sequence of the murine dystrophin gene
DMD, exon 24 4Asn His Ile Lys Thr Leu Gln Lys Trp Met Ala Glu Val Asp Val
Phe1 5 10 15Leu Lys Glu
Glu Trp Pro Ala Leu Gly Asp Ala Glu Ile Leu Lys Lys 20
25 30Gln Leu Lys Gln Cys Arg
3553678PRTMus musculusmisc_featureProtein sequence of the murine
dystrophin gene DMD 5Met Leu Trp Trp Glu Glu Val Glu Asp Cys Tyr Glu
Arg Glu Asp Val1 5 10
15Gln Lys Lys Thr Phe Thr Lys Trp Ile Asn Ala Gln Phe Ser Lys Phe
20 25 30Gly Lys Gln His Ile Asp Asn
Leu Phe Ser Asp Leu Gln Asp Gly Lys 35 40
45Arg Leu Leu Asp Leu Leu Glu Gly Leu Thr Gly Gln Lys Leu Pro
Lys 50 55 60Glu Lys Gly Ser Thr Arg
Val His Ala Leu Asn Asn Val Asn Lys Ala65 70
75 80Leu Arg Val Leu Gln Lys Asn Asn Val Asp Leu
Val Asn Ile Gly Ser 85 90
95Thr Asp Ile Val Asp Gly Asn His Lys Leu Thr Leu Gly Leu Ile Trp
100 105 110Asn Ile Ile Leu His Trp
Gln Val Lys Asn Val Met Lys Thr Ile Met 115 120
125Ala Gly Leu Gln Gln Thr Asn Ser Glu Lys Ile Leu Leu Ser
Trp Val 130 135 140Arg Gln Ser Thr Arg
Asn Tyr Pro Gln Val Asn Val Ile Asn Phe Thr145 150
155 160Ser Ser Trp Ser Asp Gly Leu Ala Leu Asn
Ala Leu Ile His Ser His 165 170
175Arg Pro Asp Leu Phe Asp Trp Asn Ser Val Val Ser Gln His Ser Ala
180 185 190Thr Gln Arg Leu Glu
His Ala Phe Asn Ile Ala Lys Cys Gln Leu Gly 195
200 205Ile Glu Lys Leu Leu Asp Pro Glu Asp Val Ala Thr
Thr Tyr Pro Asp 210 215 220Lys Lys Ser
Ile Leu Met Tyr Ile Thr Ser Leu Phe Gln Val Leu Pro225
230 235 240Gln Gln Val Ser Ile Glu Ala
Ile Gln Glu Val Glu Met Leu Pro Arg 245
250 255Thr Ser Ser Lys Val Thr Arg Glu Glu His Phe Gln
Leu His His Gln 260 265 270Met
His Tyr Ser Gln Gln Ile Thr Val Ser Leu Ala Gln Gly Tyr Glu 275
280 285Gln Thr Ser Ser Ser Pro Lys Pro Arg
Phe Lys Ser Tyr Ala Phe Thr 290 295
300Gln Ala Ala Tyr Val Ala Thr Ser Asp Ser Thr Gln Ser Pro Tyr Pro305
310 315 320Ser Gln His Leu
Glu Ala Pro Arg Asp Lys Ser Leu Asp Ser Ser Leu 325
330 335Met Glu Thr Glu Val Asn Leu Asp Ser Tyr
Gln Thr Ala Leu Glu Glu 340 345
350Val Leu Ser Trp Leu Leu Ser Ala Glu Asp Thr Leu Arg Ala Gln Gly
355 360 365Glu Ile Ser Asn Asp Val Glu
Glu Val Lys Glu Gln Phe His Ala His 370 375
380Glu Gly Phe Met Met Asp Leu Thr Ser His Gln Gly Leu Val Gly
Asn385 390 395 400Val Leu
Gln Leu Gly Ser Gln Leu Val Gly Lys Gly Lys Leu Ser Glu
405 410 415Asp Glu Glu Ala Glu Val Gln
Glu Gln Met Asn Leu Leu Asn Ser Arg 420 425
430Trp Glu Cys Leu Arg Val Ala Ser Met Glu Lys Gln Ser Lys
Leu His 435 440 445Lys Val Leu Met
Asp Leu Gln Asn Gln Lys Leu Lys Glu Leu Asp Asp 450
455 460Trp Leu Thr Lys Thr Glu Glu Arg Thr Lys Lys Met
Glu Glu Glu Pro465 470 475
480Phe Gly Pro Asp Leu Glu Asp Leu Lys Cys Gln Val Gln Gln His Lys
485 490 495Val Leu Gln Glu Asp
Leu Glu Gln Glu Gln Val Arg Val Asn Ser Leu 500
505 510Thr His Met Val Val Val Val Asp Glu Ser Ser Gly
Asp His Ala Thr 515 520 525Ala Ala
Leu Glu Glu Gln Leu Lys Val Leu Gly Asp Arg Trp Ala Asn 530
535 540Ile Cys Arg Trp Thr Glu Asp Arg Trp Ile Val
Leu Gln Asp Ile Leu545 550 555
560Leu Lys Trp Gln His Phe Thr Glu Glu Gln Cys Leu Phe Ser Thr Trp
565 570 575Leu Ser Glu Lys
Glu Asp Ala Met Lys Asn Ile Gln Thr Ser Gly Phe 580
585 590Lys Asp Gln Asn Glu Met Met Ser Ser Leu His
Lys Ile Ser Thr Leu 595 600 605Lys
Ile Asp Leu Glu Lys Lys Lys Pro Thr Met Glu Lys Leu Ser Ser 610
615 620Leu Asn Gln Asp Leu Leu Ser Ala Leu Lys
Asn Lys Ser Val Thr Gln625 630 635
640Lys Met Glu Ile Trp Met Glu Asn Phe Ala Gln Arg Trp Asp Asn
Leu 645 650 655Thr Gln Lys
Leu Glu Lys Ser Ser Ala Gln Ile Ser Gln Ala Val Thr 660
665 670Thr Thr Gln Pro Ser Leu Thr Gln Thr Thr
Val Met Glu Thr Val Thr 675 680
685Met Val Thr Thr Arg Glu Gln Ile Met Val Lys His Ala Gln Glu Glu 690
695 700Leu Pro Pro Pro Pro Pro Gln Lys
Lys Arg Gln Ile Thr Val Asp Ser705 710
715 720Glu Leu Arg Lys Arg Leu Asp Val Asp Ile Thr Glu
Leu His Ser Trp 725 730
735Ile Thr Arg Ser Glu Ala Val Leu Gln Ser Ser Glu Phe Ala Val Tyr
740 745 750Arg Lys Glu Gly Asn Ile
Ser Asp Leu Gln Glu Lys Val Asn Ala Ile 755 760
765Ala Arg Glu Lys Ala Glu Lys Phe Arg Lys Leu Gln Asp Ala
Ser Arg 770 775 780Ser Ala Gln Ala Leu
Val Glu Gln Met Ala Asn Glu Gly Val Asn Ala785 790
795 800Glu Ser Ile Arg Gln Ala Ser Glu Gln Leu
Asn Ser Arg Trp Thr Glu 805 810
815Phe Cys Gln Leu Leu Ser Glu Arg Val Asn Trp Leu Glu Tyr Gln Thr
820 825 830Asn Ile Ile Thr Phe
Tyr Asn Gln Leu Gln Gln Leu Glu Gln Met Thr 835
840 845Thr Thr Ala Glu Asn Leu Leu Lys Thr Gln Ser Thr
Thr Leu Ser Glu 850 855 860Pro Thr Ala
Ile Lys Ser Gln Leu Lys Ile Cys Lys Asp Glu Val Asn865
870 875 880Arg Leu Ser Ala Leu Gln Pro
Gln Ile Glu Gln Leu Lys Ile Gln Ser 885
890 895Leu Gln Leu Lys Glu Lys Gly Gln Gly Pro Met Phe
Leu Asp Ala Asp 900 905 910Phe
Val Ala Phe Thr Asn His Phe Asn His Ile Phe Asp Gly Val Arg 915
920 925Ala Lys Glu Lys Glu Leu Gln Thr Ile
Phe Asp Thr Leu Pro Pro Met 930 935
940Arg Tyr Gln Glu Thr Met Ser Ser Ile Arg Thr Trp Ile Gln Gln Ser945
950 955 960Glu Ser Lys Leu
Ser Val Pro Tyr Leu Ser Val Thr Glu Tyr Glu Ile 965
970 975Met Glu Glu Arg Leu Gly Lys Leu Gln Ala
Leu Gln Ser Ser Leu Lys 980 985
990Glu Gln Gln Asn Gly Phe Asn Tyr Leu Ser Asp Thr Val Lys Glu Met
995 1000 1005Ala Lys Lys Ala Pro Ser
Glu Ile Cys Gln Lys Tyr Leu Ser Glu 1010 1015
1020Phe Glu Glu Ile Glu Gly His Trp Lys Lys Leu Ser Ser Gln
Leu 1025 1030 1035Val Glu Ser Cys Gln
Lys Leu Glu Glu His Met Asn Lys Leu Arg 1040 1045
1050Lys Phe Gln Asn His Ile Lys Thr Leu Gln Lys Trp Met
Ala Glu 1055 1060 1065Val Asp Val Phe
Leu Lys Glu Glu Trp Pro Ala Leu Gly Asp Ala 1070
1075 1080Glu Ile Leu Lys Lys Gln Leu Lys Gln Cys Arg
Leu Leu Val Gly 1085 1090 1095Asp Ile
Gln Thr Ile Gln Pro Ser Leu Asn Ser Val Asn Glu Gly 1100
1105 1110Gly Gln Lys Ile Lys Ser Glu Ala Glu Leu
Glu Phe Ala Ser Arg 1115 1120 1125Leu
Glu Thr Glu Leu Arg Glu Leu Asn Thr Gln Trp Asp His Ile 1130
1135 1140Cys Arg Gln Val Tyr Thr Arg Lys Glu
Ala Leu Lys Ala Gly Leu 1145 1150
1155Asp Lys Thr Val Ser Leu Gln Lys Asp Leu Ser Glu Met His Glu
1160 1165 1170Trp Met Thr Gln Ala Glu
Glu Glu Tyr Leu Glu Arg Asp Phe Glu 1175 1180
1185Tyr Lys Thr Pro Asp Glu Leu Gln Thr Ala Val Glu Glu Met
Lys 1190 1195 1200Arg Ala Lys Glu Glu
Ala Leu Gln Lys Glu Thr Lys Val Lys Leu 1205 1210
1215Leu Thr Glu Thr Val Asn Ser Val Ile Ala His Ala Pro
Pro Ser 1220 1225 1230Ala Gln Glu Ala
Leu Lys Lys Glu Leu Glu Thr Leu Thr Thr Asn 1235
1240 1245Tyr Gln Trp Leu Cys Thr Arg Leu Asn Gly Lys
Cys Lys Thr Leu 1250 1255 1260Glu Glu
Val Trp Ala Cys Trp His Glu Leu Leu Ser Tyr Leu Glu 1265
1270 1275Lys Ala Asn Lys Trp Leu Asn Glu Val Glu
Leu Lys Leu Lys Thr 1280 1285 1290Met
Glu Asn Val Pro Ala Gly Pro Glu Glu Ile Thr Glu Val Leu 1295
1300 1305Glu Ser Leu Glu Asn Leu Met His His
Ser Glu Glu Asn Pro Asn 1310 1315
1320Gln Ile Arg Leu Leu Ala Gln Thr Leu Thr Asp Gly Gly Val Met
1325 1330 1335Asp Glu Leu Ile Asn Glu
Glu Leu Glu Thr Phe Asn Ser Arg Trp 1340 1345
1350Arg Glu Leu His Glu Glu Ala Val Arg Lys Gln Lys Leu Leu
Glu 1355 1360 1365Gln Ser Ile Gln Ser
Ala Gln Glu Ile Glu Lys Ser Leu His Leu 1370 1375
1380Ile Gln Glu Ser Leu Glu Phe Ile Asp Lys Gln Leu Ala
Ala Tyr 1385 1390 1395Ile Thr Asp Lys
Val Asp Ala Ala Gln Met Pro Gln Glu Ala Gln 1400
1405 1410Lys Ile Gln Ser Asp Leu Thr Ser His Glu Ile
Ser Leu Glu Glu 1415 1420 1425Met Lys
Lys His Asn Gln Gly Lys Asp Ala Asn Gln Arg Val Leu 1430
1435 1440Ser Gln Ile Asp Val Ala Gln Lys Lys Leu
Gln Asp Val Ser Met 1445 1450 1455Lys
Phe Arg Leu Phe Gln Lys Pro Ala Asn Phe Glu Gln Arg Leu 1460
1465 1470Glu Glu Ser Lys Met Ile Leu Asp Glu
Val Lys Met His Leu Pro 1475 1480
1485Ala Leu Glu Thr Lys Ser Val Glu Gln Glu Val Ile Gln Ser Gln
1490 1495 1500Leu Ser His Cys Val Asn
Leu Tyr Lys Ser Leu Ser Glu Val Lys 1505 1510
1515Ser Glu Val Glu Met Val Ile Lys Thr Gly Arg Gln Ile Val
Gln 1520 1525 1530Lys Lys Gln Thr Glu
Asn Pro Lys Glu Leu Asp Glu Arg Val Thr 1535 1540
1545Ala Leu Lys Leu His Tyr Asn Glu Leu Gly Ala Lys Val
Thr Glu 1550 1555 1560Arg Lys Gln Gln
Leu Glu Lys Cys Leu Lys Leu Ser Arg Lys Met 1565
1570 1575Arg Lys Glu Met Asn Val Leu Thr Glu Trp Leu
Ala Ala Thr Asp 1580 1585 1590Thr Glu
Leu Thr Lys Arg Ser Ala Val Glu Gly Met Pro Ser Asn 1595
1600 1605Leu Asp Ser Glu Val Ala Trp Gly Lys Ala
Thr Gln Lys Glu Ile 1610 1615 1620Glu
Lys Gln Lys Ala His Leu Lys Ser Val Thr Glu Leu Gly Glu 1625
1630 1635Ser Leu Lys Met Val Leu Gly Lys Lys
Glu Thr Leu Val Glu Asp 1640 1645
1650Lys Leu Ser Leu Leu Asn Ser Asn Trp Ile Ala Val Thr Ser Arg
1655 1660 1665Val Glu Glu Trp Leu Asn
Leu Leu Leu Glu Tyr Gln Lys His Met 1670 1675
1680Glu Thr Phe Asp Gln Asn Ile Glu Gln Ile Thr Lys Trp Ile
Ile 1685 1690 1695His Ala Asp Glu Leu
Leu Asp Glu Ser Glu Lys Lys Lys Pro Gln 1700 1705
1710Gln Lys Glu Asp Ile Leu Lys Arg Leu Lys Ala Glu Met
Asn Asp 1715 1720 1725Met Arg Pro Lys
Val Asp Ser Thr Arg Asp Gln Ala Ala Lys Leu 1730
1735 1740Met Ala Asn Arg Gly Asp His Cys Arg Lys Val
Val Glu Pro Gln 1745 1750 1755Ile Ser
Glu Leu Asn Arg Arg Phe Ala Ala Ile Ser His Arg Ile 1760
1765 1770Lys Thr Gly Lys Ala Ser Ile Pro Leu Lys
Glu Leu Glu Gln Phe 1775 1780 1785Asn
Ser Asp Ile Gln Lys Leu Leu Glu Pro Leu Glu Ala Glu Ile 1790
1795 1800Gln Gln Gly Val Asn Leu Lys Glu Glu
Asp Phe Asn Lys Asp Met 1805 1810
1815Ser Glu Asp Asn Glu Gly Thr Val Asn Glu Leu Leu Gln Arg Gly
1820 1825 1830Asp Asn Leu Gln Gln Arg
Ile Thr Asp Glu Arg Lys Arg Glu Glu 1835 1840
1845Ile Lys Ile Lys Gln Gln Leu Leu Gln Thr Lys His Asn Ala
Leu 1850 1855 1860Lys Asp Leu Arg Ser
Gln Arg Arg Lys Lys Ala Leu Glu Ile Ser 1865 1870
1875His Gln Trp Tyr Gln Tyr Lys Arg Gln Ala Asp Asp Leu
Leu Lys 1880 1885 1890Cys Leu Asp Glu
Ile Glu Lys Lys Leu Ala Ser Leu Pro Glu Pro 1895
1900 1905Arg Asp Glu Arg Lys Leu Lys Glu Ile Asp Arg
Glu Leu Gln Lys 1910 1915 1920Lys Lys
Glu Glu Leu Asn Ala Val Arg Arg Gln Ala Glu Gly Leu 1925
1930 1935Ser Glu Asn Gly Ala Ala Met Ala Val Glu
Pro Thr Gln Ile Gln 1940 1945 1950Leu
Ser Lys Arg Trp Arg Gln Ile Glu Ser Asn Phe Ala Gln Phe 1955
1960 1965Arg Arg Leu Asn Phe Ala Gln Ile His
Thr Leu His Glu Glu Thr 1970 1975
1980Met Val Val Thr Thr Glu Asp Met Pro Leu Asp Val Ser Tyr Val
1985 1990 1995Pro Ser Thr Tyr Leu Thr
Glu Ile Ser His Ile Leu Gln Ala Leu 2000 2005
2010Ser Glu Val Asp His Leu Leu Asn Thr Pro Glu Leu Cys Ala
Lys 2015 2020 2025Asp Phe Glu Asp Leu
Phe Lys Gln Glu Glu Ser Leu Lys Asn Ile 2030 2035
2040Lys Asp Asn Leu Gln Gln Ile Ser Gly Arg Ile Asp Ile
Ile His 2045 2050 2055Lys Lys Lys Thr
Ala Ala Leu Gln Ser Ala Thr Ser Met Glu Lys 2060
2065 2070Val Lys Val Gln Glu Ala Val Ala Gln Met Asp
Phe Gln Gly Glu 2075 2080 2085Lys Leu
His Arg Met Tyr Lys Glu Arg Gln Gly Arg Phe Asp Arg 2090
2095 2100Ser Val Glu Lys Trp Arg His Phe His Tyr
Asp Met Lys Val Phe 2105 2110 2115Asn
Gln Trp Leu Asn Glu Val Glu Gln Phe Phe Lys Lys Thr Gln 2120
2125 2130Asn Pro Glu Asn Trp Glu His Ala Lys
Tyr Lys Trp Tyr Leu Lys 2135 2140
2145Glu Leu Gln Asp Gly Ile Gly Gln Arg Gln Ala Val Val Arg Thr
2150 2155 2160Leu Asn Ala Thr Gly Glu
Glu Ile Ile Gln Gln Ser Ser Lys Thr 2165 2170
2175Asp Val Asn Ile Leu Gln Glu Lys Leu Gly Ser Leu Ser Leu
Arg 2180 2185 2190Trp His Asp Ile Cys
Lys Glu Leu Ala Glu Arg Arg Lys Arg Ile 2195 2200
2205Glu Glu Gln Lys Asn Val Leu Ser Glu Phe Gln Arg Asp
Leu Asn 2210 2215 2220Glu Phe Val Leu
Trp Leu Glu Glu Ala Asp Asn Ile Ala Ile Thr 2225
2230 2235Pro Leu Gly Asp Glu Gln Gln Leu Lys Glu Gln
Leu Glu Gln Val 2240 2245 2250Lys Leu
Leu Ala Glu Glu Leu Pro Leu Arg Gln Gly Ile Leu Lys 2255
2260 2265Gln Leu Asn Glu Thr Gly Gly Ala Val Leu
Val Ser Ala Pro Ile 2270 2275 2280Arg
Pro Glu Glu Gln Asp Lys Leu Glu Lys Lys Leu Lys Gln Thr 2285
2290 2295Asn Leu Gln Trp Ile Lys Val Ser Arg
Ala Leu Pro Glu Lys Gln 2300 2305
2310Gly Glu Leu Glu Val His Leu Lys Asp Phe Arg Gln Leu Glu Glu
2315 2320 2325Gln Leu Asp His Leu Leu
Leu Trp Leu Ser Pro Ile Arg Asn Gln 2330 2335
2340Leu Glu Ile Tyr Asn Gln Pro Ser Gln Ala Gly Pro Phe Asp
Ile 2345 2350 2355Lys Glu Ile Glu Val
Thr Val His Gly Lys Gln Ala Asp Val Glu 2360 2365
2370Arg Leu Leu Ser Lys Gly Gln His Leu Tyr Lys Glu Lys
Pro Ser 2375 2380 2385Thr Gln Pro Val
Lys Arg Lys Leu Glu Asp Leu Arg Ser Glu Trp 2390
2395 2400Glu Ala Val Asn His Leu Leu Arg Glu Leu Arg
Thr Lys Gln Pro 2405 2410 2415Asp Arg
Ala Pro Gly Leu Ser Thr Thr Gly Ala Ser Ala Ser Gln 2420
2425 2430Thr Val Thr Leu Val Thr Gln Ser Val Val
Thr Lys Glu Thr Val 2435 2440 2445Ile
Ser Lys Leu Glu Met Pro Ser Ser Leu Leu Leu Glu Val Pro 2450
2455 2460Ala Leu Ala Asp Phe Asn Arg Ala Trp
Thr Glu Leu Thr Asp Trp 2465 2470
2475Leu Ser Leu Leu Asp Arg Val Ile Lys Ser Gln Arg Val Met Val
2480 2485 2490Gly Asp Leu Glu Asp Ile
Asn Glu Met Ile Ile Lys Gln Lys Ala 2495 2500
2505Thr Leu Gln Asp Leu Glu Gln Arg Arg Pro Gln Leu Glu Glu
Leu 2510 2515 2520Ile Thr Ala Ala Gln
Asn Leu Lys Asn Lys Thr Ser Asn Gln Glu 2525 2530
2535Ala Arg Thr Ile Ile Thr Asp Arg Ile Glu Arg Ile Gln
Ile Gln 2540 2545 2550Trp Asp Glu Val
Gln Glu Gln Leu Gln Asn Arg Arg Gln Gln Leu 2555
2560 2565Asn Glu Met Leu Lys Asp Ser Thr Gln Trp Leu
Glu Ala Lys Glu 2570 2575 2580Glu Ala
Glu Gln Val Ile Gly Gln Val Arg Gly Lys Leu Asp Ser 2585
2590 2595Trp Lys Glu Gly Pro His Thr Val Asp Ala
Ile Gln Lys Lys Ile 2600 2605 2610Thr
Glu Thr Lys Gln Leu Ala Lys Asp Leu Arg Gln Arg Gln Ile 2615
2620 2625Ser Val Asp Val Ala Asn Asp Leu Ala
Leu Lys Leu Leu Arg Asp 2630 2635
2640Tyr Ser Ala Asp Asp Thr Arg Lys Val His Met Ile Thr Glu Asn
2645 2650 2655Ile Asn Thr Ser Trp Gly
Asn Ile His Lys Arg Val Ser Glu Gln 2660 2665
2670Glu Ala Ala Leu Glu Glu Thr His Arg Leu Leu Gln Gln Phe
Pro 2675 2680 2685Leu Asp Leu Glu Lys
Phe Leu Ser Trp Ile Thr Glu Ala Glu Thr 2690 2695
2700Thr Ala Asn Val Leu Gln Asp Ala Ser Arg Lys Glu Lys
Leu Leu 2705 2710 2715Glu Asp Ser Arg
Gly Val Arg Glu Leu Met Lys Pro Trp Gln Asp 2720
2725 2730Leu Gln Gly Glu Ile Glu Thr His Thr Asp Ile
Tyr His Asn Leu 2735 2740 2745Asp Glu
Asn Gly Gln Lys Ile Leu Arg Ser Leu Glu Gly Ser Asp 2750
2755 2760Glu Ala Pro Leu Leu Gln Arg Arg Leu Asp
Asn Met Asn Phe Lys 2765 2770 2775Trp
Ser Glu Leu Gln Lys Lys Ser Leu Asn Ile Arg Ser His Leu 2780
2785 2790Glu Ala Ser Ser Asp Gln Trp Lys Arg
Leu His Leu Ser Leu Gln 2795 2800
2805Glu Leu Leu Val Trp Leu Gln Leu Lys Asp Asp Glu Leu Ser Arg
2810 2815 2820Gln Ala Pro Ile Gly Gly
Asp Phe Pro Ala Val Gln Lys Gln Asn 2825 2830
2835Asp Ile His Arg Ala Phe Lys Arg Glu Leu Lys Thr Lys Glu
Pro 2840 2845 2850Val Ile Met Ser Thr
Leu Glu Thr Val Arg Ile Phe Leu Thr Glu 2855 2860
2865Gln Pro Leu Glu Gly Leu Glu Lys Leu Tyr Gln Glu Pro
Arg Glu 2870 2875 2880Leu Pro Pro Glu
Glu Arg Ala Gln Asn Val Thr Arg Leu Leu Arg 2885
2890 2895Lys Gln Ala Glu Glu Val Asn Ala Glu Trp Asp
Lys Leu Asn Leu 2900 2905 2910Arg Ser
Ala Asp Trp Gln Arg Lys Ile Asp Glu Ala Leu Glu Arg 2915
2920 2925Leu Gln Glu Leu Gln Glu Ala Ala Asp Glu
Leu Asp Leu Lys Leu 2930 2935 2940Arg
Gln Ala Glu Val Ile Lys Gly Ser Trp Gln Pro Val Gly Asp 2945
2950 2955Leu Leu Ile Asp Ser Leu Gln Asp His
Leu Glu Lys Val Lys Ala 2960 2965
2970Leu Arg Gly Glu Ile Ala Pro Leu Lys Glu Asn Val Asn Arg Val
2975 2980 2985Asn Asp Leu Ala His Gln
Leu Thr Thr Leu Gly Ile Gln Leu Ser 2990 2995
3000Pro Tyr Asn Leu Ser Thr Leu Glu Asp Leu Asn Thr Arg Trp
Arg 3005 3010 3015Leu Leu Gln Val Ala
Val Glu Asp Arg Val Arg Gln Leu His Glu 3020 3025
3030Ala His Arg Asp Phe Gly Pro Ala Ser Gln His Phe Leu
Ser Thr 3035 3040 3045Ser Val Gln Gly
Pro Trp Glu Arg Ala Ile Ser Pro Asn Lys Val 3050
3055 3060Pro Tyr Tyr Ile Asn His Glu Thr Gln Thr Thr
Cys Trp Asp His 3065 3070 3075Pro Lys
Met Thr Glu Leu Tyr Gln Ser Leu Ala Asp Leu Asn Asn 3080
3085 3090Val Arg Phe Ser Ala Tyr Arg Thr Ala Met
Lys Leu Arg Arg Leu 3095 3100 3105Gln
Lys Ala Leu Cys Leu Asp Leu Leu Ser Leu Ser Ala Ala Cys 3110
3115 3120Asp Ala Leu Asp Gln His Asn Leu Lys
Gln Asn Asp Gln Pro Met 3125 3130
3135Asp Ile Leu Gln Ile Ile Asn Cys Leu Thr Thr Ile Tyr Asp Arg
3140 3145 3150Leu Glu Gln Glu His Asn
Asn Leu Val Asn Val Pro Leu Cys Val 3155 3160
3165Asp Met Cys Leu Asn Trp Leu Leu Asn Val Tyr Asp Thr Gly
Arg 3170 3175 3180Thr Gly Arg Ile Arg
Val Leu Ser Phe Lys Thr Gly Ile Ile Ser 3185 3190
3195Leu Cys Lys Ala His Leu Glu Asp Lys Tyr Arg Tyr Leu
Phe Lys 3200 3205 3210Gln Val Ala Ser
Ser Thr Gly Phe Cys Asp Gln Arg Arg Leu Gly 3215
3220 3225Leu Leu Leu His Asp Ser Ile Gln Ile Pro Arg
Gln Leu Gly Glu 3230 3235 3240Val Ala
Ser Phe Gly Gly Ser Asn Ile Glu Pro Ser Val Arg Ser 3245
3250 3255Cys Phe Gln Phe Ala Asn Asn Lys Pro Glu
Ile Glu Ala Ala Leu 3260 3265 3270Phe
Leu Asp Trp Met Arg Leu Glu Pro Gln Ser Met Val Trp Leu 3275
3280 3285Pro Val Leu His Arg Val Ala Ala Ala
Glu Thr Ala Lys His Gln 3290 3295
3300Ala Lys Cys Asn Ile Cys Lys Glu Cys Pro Ile Ile Gly Phe Arg
3305 3310 3315Tyr Arg Ser Leu Lys His
Phe Asn Tyr Asp Ile Cys Gln Ser Cys 3320 3325
3330Phe Phe Ser Gly Arg Val Ala Lys Gly His Lys Met His Tyr
Pro 3335 3340 3345Met Val Glu Tyr Cys
Thr Pro Thr Thr Ser Gly Glu Asp Val Arg 3350 3355
3360Asp Phe Ala Lys Val Leu Lys Asn Lys Phe Arg Thr Lys
Arg Tyr 3365 3370 3375Phe Ala Lys His
Pro Arg Met Gly Tyr Leu Pro Val Gln Thr Val 3380
3385 3390Leu Glu Gly Asp Asn Met Glu Thr Pro Val Thr
Leu Ile Asn Phe 3395 3400 3405Trp Pro
Val Asp Ser Ala Pro Ala Ser Ser Pro Gln Leu Ser His 3410
3415 3420Asp Asp Thr His Ser Arg Ile Glu His Tyr
Ala Ser Arg Leu Ala 3425 3430 3435Glu
Met Glu Asn Ser Asn Gly Ser Tyr Leu Asn Asp Ser Ile Ser 3440
3445 3450Pro Asn Glu Ser Ile Asp Asp Glu His
Leu Leu Ile Gln His Tyr 3455 3460
3465Cys Gln Ser Leu Asn Gln Asp Ser Pro Leu Ser Gln Pro Arg Ser
3470 3475 3480Pro Ala Gln Ile Leu Ile
Ser Leu Glu Ser Glu Glu Arg Gly Glu 3485 3490
3495Leu Glu Arg Ile Leu Ala Asp Leu Glu Glu Glu Asn Arg Asn
Leu 3500 3505 3510Gln Ala Glu Tyr Asp
Arg Leu Lys Gln Gln His Glu His Lys Gly 3515 3520
3525Leu Ser Pro Leu Pro Ser Pro Pro Glu Met Met Pro Thr
Ser Pro 3530 3535 3540Gln Ser Pro Arg
Asp Ala Glu Leu Ile Ala Glu Ala Lys Leu Leu 3545
3550 3555Arg Gln His Lys Gly Arg Leu Glu Ala Arg Met
Gln Ile Leu Glu 3560 3565 3570Asp His
Asn Lys Gln Leu Glu Ser Gln Leu His Arg Leu Arg Gln 3575
3580 3585Leu Leu Glu Gln Pro Gln Ala Glu Ala Lys
Val Asn Gly Thr Thr 3590 3595 3600Val
Ser Ser Pro Ser Thr Ser Leu Gln Arg Ser Asp Ser Ser Gln 3605
3610 3615Pro Met Leu Leu Arg Val Val Gly Ser
Gln Thr Ser Glu Ser Met 3620 3625
3630Gly Glu Glu Asp Leu Leu Ser Pro Pro Gln Asp Thr Ser Thr Gly
3635 3640 3645Leu Glu Glu Val Met Glu
Gln Leu Asn Asn Ser Phe Pro Ser Ser 3650 3655
3660Arg Gly Arg Asn Ala Pro Gly Lys Pro Met Arg Glu Asp Thr
Met 3665 3670 367563685PRTHomo
sapiensmisc_featureProtein sequence of the human dystrophin gene DMD
6Met Leu Trp Trp Glu Glu Val Glu Asp Cys Tyr Glu Arg Glu Asp Val1
5 10 15Gln Lys Lys Thr Phe Thr
Lys Trp Val Asn Ala Gln Phe Ser Lys Phe 20 25
30Gly Lys Gln His Ile Glu Asn Leu Phe Ser Asp Leu Gln
Asp Gly Arg 35 40 45Arg Leu Leu
Asp Leu Leu Glu Gly Leu Thr Gly Gln Lys Leu Pro Lys 50
55 60Glu Lys Gly Ser Thr Arg Val His Ala Leu Asn Asn
Val Asn Lys Ala65 70 75
80Leu Arg Val Leu Gln Asn Asn Asn Val Asp Leu Val Asn Ile Gly Ser
85 90 95Thr Asp Ile Val Asp Gly
Asn His Lys Leu Thr Leu Gly Leu Ile Trp 100
105 110Asn Ile Ile Leu His Trp Gln Val Lys Asn Val Met
Lys Asn Ile Met 115 120 125Ala Gly
Leu Gln Gln Thr Asn Ser Glu Lys Ile Leu Leu Ser Trp Val 130
135 140Arg Gln Ser Thr Arg Asn Tyr Pro Gln Val Asn
Val Ile Asn Phe Thr145 150 155
160Thr Ser Trp Ser Asp Gly Leu Ala Leu Asn Ala Leu Ile His Ser His
165 170 175Arg Pro Asp Leu
Phe Asp Trp Asn Ser Val Val Cys Gln Gln Ser Ala 180
185 190Thr Gln Arg Leu Glu His Ala Phe Asn Ile Ala
Arg Tyr Gln Leu Gly 195 200 205Ile
Glu Lys Leu Leu Asp Pro Glu Asp Val Asp Thr Thr Tyr Pro Asp 210
215 220Lys Lys Ser Ile Leu Met Tyr Ile Thr Ser
Leu Phe Gln Val Leu Pro225 230 235
240Gln Gln Val Ser Ile Glu Ala Ile Gln Glu Val Glu Met Leu Pro
Arg 245 250 255Pro Pro Lys
Val Thr Lys Glu Glu His Phe Gln Leu His His Gln Met 260
265 270His Tyr Ser Gln Gln Ile Thr Val Ser Leu
Ala Gln Gly Tyr Glu Arg 275 280
285Thr Ser Ser Pro Lys Pro Arg Phe Lys Ser Tyr Ala Tyr Thr Gln Ala 290
295 300Ala Tyr Val Thr Thr Ser Asp Pro
Thr Arg Ser Pro Phe Pro Ser Gln305 310
315 320His Leu Glu Ala Pro Glu Asp Lys Ser Phe Gly Ser
Ser Leu Met Glu 325 330
335Ser Glu Val Asn Leu Asp Arg Tyr Gln Thr Ala Leu Glu Glu Val Leu
340 345 350Ser Trp Leu Leu Ser Ala
Glu Asp Thr Leu Gln Ala Gln Gly Glu Ile 355 360
365Ser Asn Asp Val Glu Val Val Lys Asp Gln Phe His Thr His
Glu Gly 370 375 380Tyr Met Met Asp Leu
Thr Ala His Gln Gly Arg Val Gly Asn Ile Leu385 390
395 400Gln Leu Gly Ser Lys Leu Ile Gly Thr Gly
Lys Leu Ser Glu Asp Glu 405 410
415Glu Thr Glu Val Gln Glu Gln Met Asn Leu Leu Asn Ser Arg Trp Glu
420 425 430Cys Leu Arg Val Ala
Ser Met Glu Lys Gln Ser Asn Leu His Arg Val 435
440 445Leu Met Asp Leu Gln Asn Gln Lys Leu Lys Glu Leu
Asn Asp Trp Leu 450 455 460Thr Lys Thr
Glu Glu Arg Thr Arg Lys Met Glu Glu Glu Pro Leu Gly465
470 475 480Pro Asp Leu Glu Asp Leu Lys
Arg Gln Val Gln Gln His Lys Val Leu 485
490 495Gln Glu Asp Leu Glu Gln Glu Gln Val Arg Val Asn
Ser Leu Thr His 500 505 510Met
Val Val Val Val Asp Glu Ser Ser Gly Asp His Ala Thr Ala Ala 515
520 525Leu Glu Glu Gln Leu Lys Val Leu Gly
Asp Arg Trp Ala Asn Ile Cys 530 535
540Arg Trp Thr Glu Asp Arg Trp Val Leu Leu Gln Asp Ile Leu Leu Lys545
550 555 560Trp Gln Arg Leu
Thr Glu Glu Gln Cys Leu Phe Ser Ala Trp Leu Ser 565
570 575Glu Lys Glu Asp Ala Val Asn Lys Ile His
Thr Thr Gly Phe Lys Asp 580 585
590Gln Asn Glu Met Leu Ser Ser Leu Gln Lys Leu Ala Val Leu Lys Ala
595 600 605Asp Leu Glu Lys Lys Lys Gln
Ser Met Gly Lys Leu Tyr Ser Leu Lys 610 615
620Gln Asp Leu Leu Ser Thr Leu Lys Asn Lys Ser Val Thr Gln Lys
Thr625 630 635 640Glu Ala
Trp Leu Asp Asn Phe Ala Arg Cys Trp Asp Asn Leu Val Gln
645 650 655Lys Leu Glu Lys Ser Thr Ala
Gln Ile Ser Gln Ala Val Thr Thr Thr 660 665
670Gln Pro Ser Leu Thr Gln Thr Thr Val Met Glu Thr Val Thr
Thr Val 675 680 685Thr Thr Arg Glu
Gln Ile Leu Val Lys His Ala Gln Glu Glu Leu Pro 690
695 700Pro Pro Pro Pro Gln Lys Lys Arg Gln Ile Thr Val
Asp Ser Glu Ile705 710 715
720Arg Lys Arg Leu Asp Val Asp Ile Thr Glu Leu His Ser Trp Ile Thr
725 730 735Arg Ser Glu Ala Val
Leu Gln Ser Pro Glu Phe Ala Ile Phe Arg Lys 740
745 750Glu Gly Asn Phe Ser Asp Leu Lys Glu Lys Val Asn
Ala Ile Glu Arg 755 760 765Glu Lys
Ala Glu Lys Phe Arg Lys Leu Gln Asp Ala Ser Arg Ser Ala 770
775 780Gln Ala Leu Val Glu Gln Met Val Asn Glu Gly
Val Asn Ala Asp Ser785 790 795
800Ile Lys Gln Ala Ser Glu Gln Leu Asn Ser Arg Trp Ile Glu Phe Cys
805 810 815Gln Leu Leu Ser
Glu Arg Leu Asn Trp Leu Glu Tyr Gln Asn Asn Ile 820
825 830Ile Ala Phe Tyr Asn Gln Leu Gln Gln Leu Glu
Gln Met Thr Thr Thr 835 840 845Ala
Glu Asn Trp Leu Lys Ile Gln Pro Thr Thr Pro Ser Glu Pro Thr 850
855 860Ala Ile Lys Ser Gln Leu Lys Ile Cys Lys
Asp Glu Val Asn Arg Leu865 870 875
880Ser Gly Leu Gln Pro Gln Ile Glu Arg Leu Lys Ile Gln Ser Ile
Ala 885 890 895Leu Lys Glu
Lys Gly Gln Gly Pro Met Phe Leu Asp Ala Asp Phe Val 900
905 910Ala Phe Thr Asn His Phe Lys Gln Val Phe
Ser Asp Val Gln Ala Arg 915 920
925Glu Lys Glu Leu Gln Thr Ile Phe Asp Thr Leu Pro Pro Met Arg Tyr 930
935 940Gln Glu Thr Met Ser Ala Ile Arg
Thr Trp Val Gln Gln Ser Glu Thr945 950
955 960Lys Leu Ser Ile Pro Gln Leu Ser Val Thr Asp Tyr
Glu Ile Met Glu 965 970
975Gln Arg Leu Gly Glu Leu Gln Ala Leu Gln Ser Ser Leu Gln Glu Gln
980 985 990Gln Ser Gly Leu Tyr Tyr
Leu Ser Thr Thr Val Lys Glu Met Ser Lys 995 1000
1005Lys Ala Pro Ser Glu Ile Ser Arg Lys Tyr Gln Ser
Glu Phe Glu 1010 1015 1020Glu Ile Glu
Gly Arg Trp Lys Lys Leu Ser Ser Gln Leu Val Glu 1025
1030 1035His Cys Gln Lys Leu Glu Glu Gln Met Asn Lys
Leu Arg Lys Ile 1040 1045 1050Gln Asn
His Ile Gln Thr Leu Lys Lys Trp Met Ala Glu Val Asp 1055
1060 1065Val Phe Leu Lys Glu Glu Trp Pro Ala Leu
Gly Asp Ser Glu Ile 1070 1075 1080Leu
Lys Lys Gln Leu Lys Gln Cys Arg Leu Leu Val Ser Asp Ile 1085
1090 1095Gln Thr Ile Gln Pro Ser Leu Asn Ser
Val Asn Glu Gly Gly Gln 1100 1105
1110Lys Ile Lys Asn Glu Ala Glu Pro Glu Phe Ala Ser Arg Leu Glu
1115 1120 1125Thr Glu Leu Lys Glu Leu
Asn Thr Gln Trp Asp His Met Cys Gln 1130 1135
1140Gln Val Tyr Ala Arg Lys Glu Ala Leu Lys Gly Gly Leu Glu
Lys 1145 1150 1155Thr Val Ser Leu Gln
Lys Asp Leu Ser Glu Met His Glu Trp Met 1160 1165
1170Thr Gln Ala Glu Glu Glu Tyr Leu Glu Arg Asp Phe Glu
Tyr Lys 1175 1180 1185Thr Pro Asp Glu
Leu Gln Lys Ala Val Glu Glu Met Lys Arg Ala 1190
1195 1200Lys Glu Glu Ala Gln Gln Lys Glu Ala Lys Val
Lys Leu Leu Thr 1205 1210 1215Glu Ser
Val Asn Ser Val Ile Ala Gln Ala Pro Pro Val Ala Gln 1220
1225 1230Glu Ala Leu Lys Lys Glu Leu Glu Thr Leu
Thr Thr Asn Tyr Gln 1235 1240 1245Trp
Leu Cys Thr Arg Leu Asn Gly Lys Cys Lys Thr Leu Glu Glu 1250
1255 1260Val Trp Ala Cys Trp His Glu Leu Leu
Ser Tyr Leu Glu Lys Ala 1265 1270
1275Asn Lys Trp Leu Asn Glu Val Glu Phe Lys Leu Lys Thr Thr Glu
1280 1285 1290Asn Ile Pro Gly Gly Ala
Glu Glu Ile Ser Glu Val Leu Asp Ser 1295 1300
1305Leu Glu Asn Leu Met Arg His Ser Glu Asp Asn Pro Asn Gln
Ile 1310 1315 1320Arg Ile Leu Ala Gln
Thr Leu Thr Asp Gly Gly Val Met Asp Glu 1325 1330
1335Leu Ile Asn Glu Glu Leu Glu Thr Phe Asn Ser Arg Trp
Arg Glu 1340 1345 1350Leu His Glu Glu
Ala Val Arg Arg Gln Lys Leu Leu Glu Gln Ser 1355
1360 1365Ile Gln Ser Ala Gln Glu Thr Glu Lys Ser Leu
His Leu Ile Gln 1370 1375 1380Glu Ser
Leu Thr Phe Ile Asp Lys Gln Leu Ala Ala Tyr Ile Ala 1385
1390 1395Asp Lys Val Asp Ala Ala Gln Met Pro Gln
Glu Ala Gln Lys Ile 1400 1405 1410Gln
Ser Asp Leu Thr Ser His Glu Ile Ser Leu Glu Glu Met Lys 1415
1420 1425Lys His Asn Gln Gly Lys Glu Ala Ala
Gln Arg Val Leu Ser Gln 1430 1435
1440Ile Asp Val Ala Gln Lys Lys Leu Gln Asp Val Ser Met Lys Phe
1445 1450 1455Arg Leu Phe Gln Lys Pro
Ala Asn Phe Glu Gln Arg Leu Gln Glu 1460 1465
1470Ser Lys Met Ile Leu Asp Glu Val Lys Met His Leu Pro Ala
Leu 1475 1480 1485Glu Thr Lys Ser Val
Glu Gln Glu Val Val Gln Ser Gln Leu Asn 1490 1495
1500His Cys Val Asn Leu Tyr Lys Ser Leu Ser Glu Val Lys
Ser Glu 1505 1510 1515Val Glu Met Val
Ile Lys Thr Gly Arg Gln Ile Val Gln Lys Lys 1520
1525 1530Gln Thr Glu Asn Pro Lys Glu Leu Asp Glu Arg
Val Thr Ala Leu 1535 1540 1545Lys Leu
His Tyr Asn Glu Leu Gly Ala Lys Val Thr Glu Arg Lys 1550
1555 1560Gln Gln Leu Glu Lys Cys Leu Lys Leu Ser
Arg Lys Met Arg Lys 1565 1570 1575Glu
Met Asn Val Leu Thr Glu Trp Leu Ala Ala Thr Asp Met Glu 1580
1585 1590Leu Thr Lys Arg Ser Ala Val Glu Gly
Met Pro Ser Asn Leu Asp 1595 1600
1605Ser Glu Val Ala Trp Gly Lys Ala Thr Gln Lys Glu Ile Glu Lys
1610 1615 1620Gln Lys Val His Leu Lys
Ser Ile Thr Glu Val Gly Glu Ala Leu 1625 1630
1635Lys Thr Val Leu Gly Lys Lys Glu Thr Leu Val Glu Asp Lys
Leu 1640 1645 1650Ser Leu Leu Asn Ser
Asn Trp Ile Ala Val Thr Ser Arg Ala Glu 1655 1660
1665Glu Trp Leu Asn Leu Leu Leu Glu Tyr Gln Lys His Met
Glu Thr 1670 1675 1680Phe Asp Gln Asn
Val Asp His Ile Thr Lys Trp Ile Ile Gln Ala 1685
1690 1695Asp Thr Leu Leu Asp Glu Ser Glu Lys Lys Lys
Pro Gln Gln Lys 1700 1705 1710Glu Asp
Val Leu Lys Arg Leu Lys Ala Glu Leu Asn Asp Ile Arg 1715
1720 1725Pro Lys Val Asp Ser Thr Arg Asp Gln Ala
Ala Asn Leu Met Ala 1730 1735 1740Asn
Arg Gly Asp His Cys Arg Lys Leu Val Glu Pro Gln Ile Ser 1745
1750 1755Glu Leu Asn His Arg Phe Ala Ala Ile
Ser His Arg Ile Lys Thr 1760 1765
1770Gly Lys Ala Ser Ile Pro Leu Lys Glu Leu Glu Gln Phe Asn Ser
1775 1780 1785Asp Ile Gln Lys Leu Leu
Glu Pro Leu Glu Ala Glu Ile Gln Gln 1790 1795
1800Gly Val Asn Leu Lys Glu Glu Asp Phe Asn Lys Asp Met Asn
Glu 1805 1810 1815Asp Asn Glu Gly Thr
Val Lys Glu Leu Leu Gln Arg Gly Asp Asn 1820 1825
1830Leu Gln Gln Arg Ile Thr Asp Glu Arg Lys Arg Glu Glu
Ile Lys 1835 1840 1845Ile Lys Gln Gln
Leu Leu Gln Thr Lys His Asn Ala Leu Lys Asp 1850
1855 1860Leu Arg Ser Gln Arg Arg Lys Lys Ala Leu Glu
Ile Ser His Gln 1865 1870 1875Trp Tyr
Gln Tyr Lys Arg Gln Ala Asp Asp Leu Leu Lys Cys Leu 1880
1885 1890Asp Asp Ile Glu Lys Lys Leu Ala Ser Leu
Pro Glu Pro Arg Asp 1895 1900 1905Glu
Arg Lys Ile Lys Glu Ile Asp Arg Glu Leu Gln Lys Lys Lys 1910
1915 1920Glu Glu Leu Asn Ala Val Arg Arg Gln
Ala Glu Gly Leu Ser Glu 1925 1930
1935Asp Gly Ala Ala Met Ala Val Glu Pro Thr Gln Ile Gln Leu Ser
1940 1945 1950Lys Arg Trp Arg Glu Ile
Glu Ser Lys Phe Ala Gln Phe Arg Arg 1955 1960
1965Leu Asn Phe Ala Gln Ile His Thr Val Arg Glu Glu Thr Met
Met 1970 1975 1980Val Met Thr Glu Asp
Met Pro Leu Glu Ile Ser Tyr Val Pro Ser 1985 1990
1995Thr Tyr Leu Thr Glu Ile Thr His Val Ser Gln Ala Leu
Leu Glu 2000 2005 2010Val Glu Gln Leu
Leu Asn Ala Pro Asp Leu Cys Ala Lys Asp Phe 2015
2020 2025Glu Asp Leu Phe Lys Gln Glu Glu Ser Leu Lys
Asn Ile Lys Asp 2030 2035 2040Ser Leu
Gln Gln Ser Ser Gly Arg Ile Asp Ile Ile His Ser Lys 2045
2050 2055Lys Thr Ala Ala Leu Gln Ser Ala Thr Pro
Val Glu Arg Val Lys 2060 2065 2070Leu
Gln Glu Ala Leu Ser Gln Leu Asp Phe Gln Trp Glu Lys Val 2075
2080 2085Asn Lys Met Tyr Lys Asp Arg Gln Gly
Arg Phe Asp Arg Ser Val 2090 2095
2100Glu Lys Trp Arg Arg Phe His Tyr Asp Ile Lys Ile Phe Asn Gln
2105 2110 2115Trp Leu Thr Glu Ala Glu
Gln Phe Leu Arg Lys Thr Gln Ile Pro 2120 2125
2130Glu Asn Trp Glu His Ala Lys Tyr Lys Trp Tyr Leu Lys Glu
Leu 2135 2140 2145Gln Asp Gly Ile Gly
Gln Arg Gln Thr Val Val Arg Thr Leu Asn 2150 2155
2160Ala Thr Gly Glu Glu Ile Ile Gln Gln Ser Ser Lys Thr
Asp Ala 2165 2170 2175Ser Ile Leu Gln
Glu Lys Leu Gly Ser Leu Asn Leu Arg Trp Gln 2180
2185 2190Glu Val Cys Lys Gln Leu Ser Asp Arg Lys Lys
Arg Leu Glu Glu 2195 2200 2205Gln Lys
Asn Ile Leu Ser Glu Phe Gln Arg Asp Leu Asn Glu Phe 2210
2215 2220Val Leu Trp Leu Glu Glu Ala Asp Asn Ile
Ala Ser Ile Pro Leu 2225 2230 2235Glu
Pro Gly Lys Glu Gln Gln Leu Lys Glu Lys Leu Glu Gln Val 2240
2245 2250Lys Leu Leu Val Glu Glu Leu Pro Leu
Arg Gln Gly Ile Leu Lys 2255 2260
2265Gln Leu Asn Glu Thr Gly Gly Pro Val Leu Val Ser Ala Pro Ile
2270 2275 2280Ser Pro Glu Glu Gln Asp
Lys Leu Glu Asn Lys Leu Lys Gln Thr 2285 2290
2295Asn Leu Gln Trp Ile Lys Val Ser Arg Ala Leu Pro Glu Lys
Gln 2300 2305 2310Gly Glu Ile Glu Ala
Gln Ile Lys Asp Leu Gly Gln Leu Glu Lys 2315 2320
2325Lys Leu Glu Asp Leu Glu Glu Gln Leu Asn His Leu Leu
Leu Trp 2330 2335 2340Leu Ser Pro Ile
Arg Asn Gln Leu Glu Ile Tyr Asn Gln Pro Asn 2345
2350 2355Gln Glu Gly Pro Phe Asp Val Gln Glu Thr Glu
Ile Ala Val Gln 2360 2365 2370Ala Lys
Gln Pro Asp Val Glu Glu Ile Leu Ser Lys Gly Gln His 2375
2380 2385Leu Tyr Lys Glu Lys Pro Ala Thr Gln Pro
Val Lys Arg Lys Leu 2390 2395 2400Glu
Asp Leu Ser Ser Glu Trp Lys Ala Val Asn Arg Leu Leu Gln 2405
2410 2415Glu Leu Arg Ala Lys Gln Pro Asp Leu
Ala Pro Gly Leu Thr Thr 2420 2425
2430Ile Gly Ala Ser Pro Thr Gln Thr Val Thr Leu Val Thr Gln Pro
2435 2440 2445Val Val Thr Lys Glu Thr
Ala Ile Ser Lys Leu Glu Met Pro Ser 2450 2455
2460Ser Leu Met Leu Glu Val Pro Ala Leu Ala Asp Phe Asn Arg
Ala 2465 2470 2475Trp Thr Glu Leu Thr
Asp Trp Leu Ser Leu Leu Asp Gln Val Ile 2480 2485
2490Lys Ser Gln Arg Val Met Val Gly Asp Leu Glu Asp Ile
Asn Glu 2495 2500 2505Met Ile Ile Lys
Gln Lys Ala Thr Met Gln Asp Leu Glu Gln Arg 2510
2515 2520Arg Pro Gln Leu Glu Glu Leu Ile Thr Ala Ala
Gln Asn Leu Lys 2525 2530 2535Asn Lys
Thr Ser Asn Gln Glu Ala Arg Thr Ile Ile Thr Asp Arg 2540
2545 2550Ile Glu Arg Ile Gln Asn Gln Trp Asp Glu
Val Gln Glu His Leu 2555 2560 2565Gln
Asn Arg Arg Gln Gln Leu Asn Glu Met Leu Lys Asp Ser Thr 2570
2575 2580Gln Trp Leu Glu Ala Lys Glu Glu Ala
Glu Gln Val Leu Gly Gln 2585 2590
2595Ala Arg Ala Lys Leu Glu Ser Trp Lys Glu Gly Pro Tyr Thr Val
2600 2605 2610Asp Ala Ile Gln Lys Lys
Ile Thr Glu Thr Lys Gln Leu Ala Lys 2615 2620
2625Asp Leu Arg Gln Trp Gln Thr Asn Val Asp Val Ala Asn Asp
Leu 2630 2635 2640Ala Leu Lys Leu Leu
Arg Asp Tyr Ser Ala Asp Asp Thr Arg Lys 2645 2650
2655Val His Met Ile Thr Glu Asn Ile Asn Ala Ser Trp Arg
Ser Ile 2660 2665 2670His Lys Arg Val
Ser Glu Arg Glu Ala Ala Leu Glu Glu Thr His 2675
2680 2685Arg Leu Leu Gln Gln Phe Pro Leu Asp Leu Glu
Lys Phe Leu Ala 2690 2695 2700Trp Leu
Thr Glu Ala Glu Thr Thr Ala Asn Val Leu Gln Asp Ala 2705
2710 2715Thr Arg Lys Glu Arg Leu Leu Glu Asp Ser
Lys Gly Val Lys Glu 2720 2725 2730Leu
Met Lys Gln Trp Gln Asp Leu Gln Gly Glu Ile Glu Ala His 2735
2740 2745Thr Asp Val Tyr His Asn Leu Asp Glu
Asn Ser Gln Lys Ile Leu 2750 2755
2760Arg Ser Leu Glu Gly Ser Asp Asp Ala Val Leu Leu Gln Arg Arg
2765 2770 2775Leu Asp Asn Met Asn Phe
Lys Trp Ser Glu Leu Arg Lys Lys Ser 2780 2785
2790Leu Asn Ile Arg Ser His Leu Glu Ala Ser Ser Asp Gln Trp
Lys 2795 2800 2805Arg Leu His Leu Ser
Leu Gln Glu Leu Leu Val Trp Leu Gln Leu 2810 2815
2820Lys Asp Asp Glu Leu Ser Arg Gln Ala Pro Ile Gly Gly
Asp Phe 2825 2830 2835Pro Ala Val Gln
Lys Gln Asn Asp Val His Arg Ala Phe Lys Arg 2840
2845 2850Glu Leu Lys Thr Lys Glu Pro Val Ile Met Ser
Thr Leu Glu Thr 2855 2860 2865Val Arg
Ile Phe Leu Thr Glu Gln Pro Leu Glu Gly Leu Glu Lys 2870
2875 2880Leu Tyr Gln Glu Pro Arg Glu Leu Pro Pro
Glu Glu Arg Ala Gln 2885 2890 2895Asn
Val Thr Arg Leu Leu Arg Lys Gln Ala Glu Glu Val Asn Thr 2900
2905 2910Glu Trp Glu Lys Leu Asn Leu His Ser
Ala Asp Trp Gln Arg Lys 2915 2920
2925Ile Asp Glu Thr Leu Glu Arg Leu Gln Glu Leu Gln Glu Ala Thr
2930 2935 2940Asp Glu Leu Asp Leu Lys
Leu Arg Gln Ala Glu Val Ile Lys Gly 2945 2950
2955Ser Trp Gln Pro Val Gly Asp Leu Leu Ile Asp Ser Leu Gln
Asp 2960 2965 2970His Leu Glu Lys Val
Lys Ala Leu Arg Gly Glu Ile Ala Pro Leu 2975 2980
2985Lys Glu Asn Val Ser His Val Asn Asp Leu Ala Arg Gln
Leu Thr 2990 2995 3000Thr Leu Gly Ile
Gln Leu Ser Pro Tyr Asn Leu Ser Thr Leu Glu 3005
3010 3015Asp Leu Asn Thr Arg Trp Lys Leu Leu Gln Val
Ala Val Glu Asp 3020 3025 3030Arg Val
Arg Gln Leu His Glu Ala His Arg Asp Phe Gly Pro Ala 3035
3040 3045Ser Gln His Phe Leu Ser Thr Ser Val Gln
Gly Pro Trp Glu Arg 3050 3055 3060Ala
Ile Ser Pro Asn Lys Val Pro Tyr Tyr Ile Asn His Glu Thr 3065
3070 3075Gln Thr Thr Cys Trp Asp His Pro Lys
Met Thr Glu Leu Tyr Gln 3080 3085
3090Ser Leu Ala Asp Leu Asn Asn Val Arg Phe Ser Ala Tyr Arg Thr
3095 3100 3105Ala Met Lys Leu Arg Arg
Leu Gln Lys Ala Leu Cys Leu Asp Leu 3110 3115
3120Leu Ser Leu Ser Ala Ala Cys Asp Ala Leu Asp Gln His Asn
Leu 3125 3130 3135Lys Gln Asn Asp Gln
Pro Met Asp Ile Leu Gln Ile Ile Asn Cys 3140 3145
3150Leu Thr Thr Ile Tyr Asp Arg Leu Glu Gln Glu His Asn
Asn Leu 3155 3160 3165Val Asn Val Pro
Leu Cys Val Asp Met Cys Leu Asn Trp Leu Leu 3170
3175 3180Asn Val Tyr Asp Thr Gly Arg Thr Gly Arg Ile
Arg Val Leu Ser 3185 3190 3195Phe Lys
Thr Gly Ile Ile Ser Leu Cys Lys Ala His Leu Glu Asp 3200
3205 3210Lys Tyr Arg Tyr Leu Phe Lys Gln Val Ala
Ser Ser Thr Gly Phe 3215 3220 3225Cys
Asp Gln Arg Arg Leu Gly Leu Leu Leu His Asp Ser Ile Gln 3230
3235 3240Ile Pro Arg Gln Leu Gly Glu Val Ala
Ser Phe Gly Gly Ser Asn 3245 3250
3255Ile Glu Pro Ser Val Arg Ser Cys Phe Gln Phe Ala Asn Asn Lys
3260 3265 3270Pro Glu Ile Glu Ala Ala
Leu Phe Leu Asp Trp Met Arg Leu Glu 3275 3280
3285Pro Gln Ser Met Val Trp Leu Pro Val Leu His Arg Val Ala
Ala 3290 3295 3300Ala Glu Thr Ala Lys
His Gln Ala Lys Cys Asn Ile Cys Lys Glu 3305 3310
3315Cys Pro Ile Ile Gly Phe Arg Tyr Arg Ser Leu Lys His
Phe Asn 3320 3325 3330Tyr Asp Ile Cys
Gln Ser Cys Phe Phe Ser Gly Arg Val Ala Lys 3335
3340 3345Gly His Lys Met His Tyr Pro Met Val Glu Tyr
Cys Thr Pro Thr 3350 3355 3360Thr Ser
Gly Glu Asp Val Arg Asp Phe Ala Lys Val Leu Lys Asn 3365
3370 3375Lys Phe Arg Thr Lys Arg Tyr Phe Ala Lys
His Pro Arg Met Gly 3380 3385 3390Tyr
Leu Pro Val Gln Thr Val Leu Glu Gly Asp Asn Met Glu Thr 3395
3400 3405Pro Val Thr Leu Ile Asn Phe Trp Pro
Val Asp Ser Ala Pro Ala 3410 3415
3420Ser Ser Pro Gln Leu Ser His Asp Asp Thr His Ser Arg Ile Glu
3425 3430 3435His Tyr Ala Ser Arg Leu
Ala Glu Met Glu Asn Ser Asn Gly Ser 3440 3445
3450Tyr Leu Asn Asp Ser Ile Ser Pro Asn Glu Ser Ile Asp Asp
Glu 3455 3460 3465His Leu Leu Ile Gln
His Tyr Cys Gln Ser Leu Asn Gln Asp Ser 3470 3475
3480Pro Leu Ser Gln Pro Arg Ser Pro Ala Gln Ile Leu Ile
Ser Leu 3485 3490 3495Glu Ser Glu Glu
Arg Gly Glu Leu Glu Arg Ile Leu Ala Asp Leu 3500
3505 3510Glu Glu Glu Asn Arg Asn Leu Gln Ala Glu Tyr
Asp Arg Leu Lys 3515 3520 3525Gln Gln
His Glu His Lys Gly Leu Ser Pro Leu Pro Ser Pro Pro 3530
3535 3540Glu Met Met Pro Thr Ser Pro Gln Ser Pro
Arg Asp Ala Glu Leu 3545 3550 3555Ile
Ala Glu Ala Lys Leu Leu Arg Gln His Lys Gly Arg Leu Glu 3560
3565 3570Ala Arg Met Gln Ile Leu Glu Asp His
Asn Lys Gln Leu Glu Ser 3575 3580
3585Gln Leu His Arg Leu Arg Gln Leu Leu Glu Gln Pro Gln Ala Glu
3590 3595 3600Ala Lys Val Asn Gly Thr
Thr Val Ser Ser Pro Ser Thr Ser Leu 3605 3610
3615Gln Arg Ser Asp Ser Ser Gln Pro Met Leu Leu Arg Val Val
Gly 3620 3625 3630Ser Gln Thr Ser Asp
Ser Met Gly Glu Glu Asp Leu Leu Ser Pro 3635 3640
3645Pro Gln Asp Thr Ser Thr Gly Leu Glu Glu Val Met Glu
Gln Leu 3650 3655 3660Asn Asn Ser Phe
Pro Ser Ser Arg Gly Arg Asn Thr Pro Gly Lys 3665
3670 3675Pro Met Arg Glu Asp Thr Met 3680
3685713857DNAMus musculusmisc_featuremRNA sequence of the murine
dystrophin gene DMD transcript 7atcagttact atgttgactc actcagtgtt
gggctcactc acttgcccct tacaggactc 60agctcttgaa ggcaatagcc tttatagaaa
aaacgaatag gaagacttga agtgctattt 120ttttttttgt caaggctgct gaagtttatt
ggcttctcat cgtacctaag cctcctggag 180caataaaact gggagaaact tttaccaaga
tttttatccc tgccttgata tatacttttt 240cttccaaatg ctttggtggg aagaagtaga
ggactgttat gaaagagaag atgttcaaaa 300gaaaacattc acaaaatgga taaatgcaca
attttctaag tttggaaagc aacacataga 360caacctcttc agtgacctgc aggatggaaa
acgcctccta gacctcttgg aaggccttac 420agggcaaaaa ctgccaaaag aaaagggatc
tacaagagtt catgccctga acaatgtcaa 480caaggcactg cgggtcttac agaaaaataa
tgttgattta gtgaatatag gaagcactga 540catagtggat ggaaatcata aactcactct
tggtttgatt tggaatataa tcctccactg 600gcaggtcaaa aatgtgatga aaactatcat
ggctggattg cagcaaacca acagtgaaaa 660gattcttctg agctgggttc gacagtcaac
acgtaattat ccacaggtta acgtcatcaa 720cttcacctct agctggtccg acgggttggc
tttgaatgct cttatccata gtcacaggcc 780cgacctgttt gattggaata gtgtggtttc
acagcactca gccacccaaa gactggaaca 840tgccttcaac attgcaaaat gccagttagg
catagaaaaa cttcttgatc ctgaagatgt 900tgctaccact tatccagaca agaagtccat
cttaatgtac atcacatcac tctttcaagt 960tttgccacaa caagtgagca ttgaagccat
tcaagaagtg gaaatgttgc ccaggacatc 1020ttcaaaagta actagagaag aacattttca
attacatcac cagatgcatt actctcaaca 1080gatcacagtc agtctagcac agggctatga
acaaacttct tcatctccta agcctcgatt 1140caagagttat gccttcacac aggctgctta
tgttgccacc tctgattcca cacagagccc 1200ctatccttca cagcatttgg aagctcccag
agacaagtca cttgacagtt cattgatgga 1260gacggaagta aatctggata gttaccaaac
tgctttagaa gaagtacttt catggcttct 1320ttctgccgag gatacattgc gagcacaagg
agagatttca aatgatgttg aagaagtgaa 1380agaacagttt catgctcatg agggattcat
gatggatctg acatctcatc aaggacttgt 1440tggtaatgtt ctacagttag gaagtcaact
agttggaaaa gggaaattat cagaagatga 1500agaagctgaa gtgcaagaac aaatgaatct
cctaaattca agatgggaat gtctcagggt 1560agctagcatg gaaaaacaaa gcaaattaca
caaagttcta atggatctcc agaatcagaa 1620attaaaagaa ctagatgact ggttaacaaa
aactgaagag agaactaaga aaatggagga 1680agagcccttt ggacctgatc ttgaagatct
aaaatgccaa gtacaacaac ataaggtgct 1740tcaagaagat ctagaacagg agcaggtcag
ggtcaactcg ctcactcaca tggtagtagt 1800ggttgatgaa tccagcggtg atcatgcaac
agctgctttg gaagaacaac ttaaggtact 1860gggagatcga tgggcaaata tctgcagatg
gactgaagac cgctggattg ttttacaaga 1920tattcttcta aaatggcagc attttactga
agaacagtgc ctttttagta catggctttc 1980agaaaaagaa gatgcaatga agaacattca
gacaagtggc tttaaagatc aaaatgaaat 2040gatgtcaagt cttcacaaaa tatctacttt
aaaaatagat ctagaaaaga aaaagccaac 2100catggaaaaa ctaagttcac tcaatcaaga
tctactttcg gcactgaaaa ataagtcagt 2160gactcaaaag atggaaatct ggatggaaaa
ctttgcacaa cgttgggaca atttaaccca 2220aaaacttgaa aagagttcag cacaaatttc
acaggctgtc accaccactc aaccatccct 2280aacacagaca actgtaatgg aaacggtaac
tatggtgacc acaagggaac aaatcatggt 2340aaaacatgcc caagaggaac ttccaccacc
acctcctcaa aagaagaggc agataactgt 2400ggattctgaa ctcaggaaaa ggttggatgt
cgatataact gaacttcaca gttggattac 2460tcgttcagaa gctgtattac agagttctga
atttgcagtc tatcgaaaag aaggcaacat 2520ctcagacttg caagaaaaag tcaatgccat
agcacgagaa aaagcagaga agttcagaaa 2580actgcaagat gccagcagat cagctcaggc
cctggtggaa cagatggcaa atgagggtgt 2640taatgctgaa agtatcagac aagcttcaga
acaactgaac agccggtgga cagaattctg 2700ccaattgctg agtgagagag ttaactggct
agagtatcaa accaacatca ttacctttta 2760taatcagcta caacaattgg aacagatgac
aactactgcc gaaaacttgt tgaaaaccca 2820gtctaccacc ctatcagagc caacagcaat
taaaagccag ttaaaaattt gtaaggatga 2880agtcaacaga ttgtcagctc ttcagcctca
aattgagcaa ttaaaaattc agagtctaca 2940actgaaagaa aagggacagg ggccaatgtt
tctggatgca gactttgtgg cctttactaa 3000tcattttaac cacatctttg atggtgtgag
ggccaaagag aaagagctac agacaatttt 3060tgacacttta ccaccaatgc gctatcagga
gacaatgagt agcatcagga cgtggatcca 3120gcagtcagaa agcaaactct ctgtacctta
tcttagtgtt actgaatatg aaataatgga 3180ggagagactc gggaaattac aggctctgca
aagttctttg aaagagcaac aaaatggctt 3240caactatctg agtgacactg tgaaggagat
ggccaagaaa gcaccttcag aaatatgcca 3300gaaatatctg tcagaatttg aagagattga
ggggcactgg aagaaacttt cctcccagtt 3360ggtggaaagc tgccaaaagc tagaagaaca
tatgaataaa cttcgaaaat ttcagaatca 3420cataaaaacc ttacagaaat ggatggctga
agttgatgtt ttcctgaaag aggaatggcc 3480tgccctgggg gatgctgaaa tcctgaaaaa
acagctcaaa caatgcagac ttttagttgg 3540tgatattcaa acaattcagc ccagtttaaa
tagtgttaat gaaggtgggc agaagataaa 3600gagtgaagct gaacttgagt ttgcatccag
actggagaca gaacttagag agcttaacac 3660tcagtgggat cacatatgcc gccaggtcta
caccagaaag gaagccttaa aggcaggttt 3720ggataaaacc gtaagcctcc aaaaagatct
atcagagatg catgagtgga tgacacaagc 3780tgaagaagaa tatctagaga gagattttga
atataaaact ccagatgaat tacagactgc 3840tgttgaagaa atgaagagag ctaaagaaga
ggcactacaa aaagaaacta aagtgaaact 3900ccttactgag actgtaaata gtgtaatagc
tcacgctcca ccctcagcac aagaggcctt 3960aaaaaaggaa cttgaaactc tgaccaccaa
ctaccaatgg ctgtgcacca ggctgaatgg 4020aaaatgcaaa actttggaag aagtttgggc
atgttggcat gagttattgt catatttaga 4080gaaagcaaac aagtggctca atgaagtaga
attgaaactt aaaaccatgg aaaatgttcc 4140tgcaggacct gaggaaatca ctgaagtgct
agaatctctt gaaaatctga tgcatcattc 4200agaggagaac ccaaatcaga ttcgtctatt
ggcacagact cttacagatg gaggagtcat 4260ggatgaactg atcaatgagg agcttgagac
gtttaattct cgttggaggg aactacatga 4320agaggctgtg aggaaacaaa agttgcttga
acagagtatc cagtctgccc aggaaattga 4380aaagtccttg cacttaattc aggagtcgct
tgaattcatt gacaagcagt tggcagctta 4440tatcactgac aaggtggatg cagctcaaat
gcctcaggaa gcccagaaaa tccaatcaga 4500tttgacaagt catgagataa gtttagaaga
aatgaagaaa cataaccagg ggaaggatgc 4560caaccaaagg gttctttcac aaattgatgt
tgcacagaaa aaattacaag atgtctccat 4620gaaatttcga ttattccaaa aaccagccaa
ttttgaacaa cgtctagagg aaagtaagat 4680gattttagat gaagtcaaga tgcatttgcc
tgcattggaa accaagagtg ttgaacagga 4740agtaattcag tcacaactaa gtcattgtgt
gaacttgtat aaaagcctga gtgaagtcaa 4800gtctgaagtg gaaatggtga ttaaaaccgg
acgtcaaatt gtacagaaaa agcagacaga 4860aaatcccaaa gagcttgatg aacgagtaac
agctttgaaa ttgcattaca atgagttggg 4920tgcgaaggta acagagagaa agcaacagtt
ggagaaatgc ttgaagttgt cccgtaagat 4980gagaaaggaa atgaatgtct taacagaatg
gctggcagca acagatacag aattgacgaa 5040gagatcagca gttgaaggaa tgccaagtaa
tttggattct gaagttgcct ggggaaaggc 5100tactcaaaaa gagattgaga aacagaaggc
tcacttgaag agtgttacag aattaggaga 5160gtctttgaaa atggtgttgg gcaagaaaga
aaccttggta gaagataaac tgagtcttct 5220gaacagtaac tggatagctg tcacctccag
agtagaagaa tggctaaatc ttttgttgga 5280ataccagaaa cacatggaaa cctttgatca
gaacatagaa caaatcacaa agtggatcat 5340tcatgcagat gaacttttag atgagtctga
aaagaagaaa ccacaacaaa aggaagacat 5400tcttaagcgt ttaaaggctg aaatgaatga
catgcgccca aaggtggact ccacacgtga 5460ccaagcagca aaattgatgg caaaccgcgg
tgaccactgc aggaaagtag tagagcccca 5520aatctctgag ctcaaccgtc gatttgcagc
tatttctcac agaattaaga ctggaaaggc 5580ctccattcct ttgaaggaat tggagcagtt
taactcagat atacaaaaat tgcttgaacc 5640actggaggct gaaattcagc agggggtgaa
tctgaaagag gaagacttca ataaagatat 5700gagtgaagac aatgagggta ctgtaaatga
attgttgcaa agaggagaca acttacaaca 5760aagaatcaca gatgagagaa agcgagagga
aataaagata aaacagcagc tgttacagac 5820aaaacataat gctctcaagg atttgaggtc
tcaaagaaga aaaaaggccc tagaaatttc 5880tcaccagtgg tatcagtaca agaggcaggc
tgatgatctc ctgaaatgct tggatgaaat 5940tgaaaaaaaa ttagccagcc tacctgaacc
cagagatgaa agaaaattaa aggaaattga 6000tcgtgaattg cagaagaaga aagaggagct
gaatgcagtg cgcaggcaag ctgagggctt 6060gtctgagaat ggggccgcaa tggcagtgga
gccaactcag atccagctca gcaagcgctg 6120gcggcaaatt gagagcaatt ttgctcagtt
tcgaagactc aactttgcac aaattcacac 6180tctccatgaa gaaactatgg tagtgacgac
tgaagatatg cctttggatg tttcttatgt 6240gccttctact tatttgaccg agatcagtca
tatcttacaa gctctttcag aagttgatca 6300tcttctaaat actcctgaac tctgtgctaa
agattttgaa gatcttttta agcaagagga 6360gtctcttaag aatataaaag acaatttgca
acaaatctca ggtcggattg atattattca 6420caagaagaag acagcagcct tgcaaagtgc
cacctccatg gaaaaggtga aagtacagga 6480agccgtggca cagatggatt tccaggggga
aaaacttcat agaatgtaca aggaacgaca 6540agggcgattc gacagatcag ttgaaaaatg
gcgacacttt cattatgata tgaaggtatt 6600taatcaatgg ctgaatgaag ttgaacagtt
tttcaaaaag acacaaaatc ctgaaaactg 6660ggaacatgct aaatacaaat ggtatcttaa
ggaactccag gatggcattg ggcagcgtca 6720agctgttgtc agaacactga atgcaactgg
ggaagaaata attcaacagt cttcaaaaac 6780agatgtcaat attctacaag aaaaattagg
aagcttgagt ctgcggtggc acgacatctg 6840caaagagctg gcagaaagga gaaagaggat
tgaagaacaa aagaatgtct tgtcagaatt 6900tcaaagagat ttaaatgaat ttgttttgtg
gctggaagaa gcagataaca ttgctattac 6960tccacttgga gatgagcagc agctaaaaga
acaacttgaa caagtcaagt tactggcaga 7020agagttgccc ctgcgccagg gaattctaaa
acaattaaat gaaacaggag gagcagtact 7080tgtaagtgct cccataaggc cagaagagca
agataaactt gaaaagaagc tcaaacagac 7140aaatctccag tggataaagg tctccagagc
tttacctgag aaacaaggag agcttgaggt 7200tcacttaaaa gattttaggc agcttgaaga
gcagctggat cacctgcttc tgtggctctc 7260tcctattaga aaccagttgg aaatttataa
ccaaccaagt caggcaggac cgtttgacat 7320aaaggagatt gaagtaacag ttcacggtaa
acaagcggat gtggaaaggc ttttgtcgaa 7380agggcagcat ttgtataagg aaaaaccaag
cactcagcca gtgaagagga agttagaaga 7440tctgaggtct gagtgggagg ctgtaaacca
tttacttcgg gagctgagga caaagcagcc 7500tgaccgtgcc cctggactga gcactactgg
agcctctgcc agtcagactg ttactctagt 7560gacacaatct gtggttacta aggaaactgt
catctccaaa ctagaaatgc catcttcttt 7620gctgttggag gtacctgcac tggcagactt
caaccgagct tggacagaac ttacagactg 7680gctgtctctg cttgatcgag ttataaaatc
acagagagtg atggtgggtg atctggaaga 7740catcaatgaa atgatcatca aacagaaggc
aacactgcaa gatttggaac agagacgccc 7800ccaattggaa gaactcatta ctgctgccca
gaatttgaaa aacaaaacca gcaatcaaga 7860agctagaaca atcattactg atcgaattga
aagaattcag attcagtggg atgaggttca 7920agaacagctg cagaacagga gacaacagtt
gaatgaaatg ttaaaggatt caacacaatg 7980gctggaagct aaggaagaag ccgaacaggt
cataggacag gtcagaggca agcttgactc 8040atggaaagaa ggtcctcaca cagtagatgc
aatccaaaag aagatcacag aaaccaagca 8100gttggccaaa gacctccgtc aacggcagat
aagtgtagac gtggcaaatg atttggcact 8160gaaacttctt cgggactatt ctgctgatga
taccagaaaa gtacacatga taacagagaa 8220tatcaatact tcttggggaa acattcataa
aagagtaagt gagcaagagg ctgctttgga 8280agaaactcat agattactgc agcagttccc
tctggacctg gagaagtttc tttcctggat 8340tacggaagca gaaacaactg ccaatgtcct
acaggacgct tcccgtaagg agaagctcct 8400agaagactcc aggggagtca gagagctgat
gaaaccatgg caagatctcc aaggagaaat 8460tgaaactcac acagatatct atcacaatct
tgatgaaaat ggccaaaaaa tcctgagatc 8520cctggaaggt tcggatgaag cacccctgtt
acaaagacgt ttggataaca tgaatttcaa 8580gtggagtgaa cttcagaaaa agtctctcaa
cattaggtcc catttggaag caagttctga 8640ccagtggaag cgtttgcatc tttctcttca
ggaacttctt gtttggctac agctgaaaga 8700tgatgaactg agccgtcagg cacccatcgg
tggtgatttc ccagcagttc agaagcagaa 8760tgatatacat agggccttca agagggaatt
gaaaactaaa gaacctgtaa tcatgagtac 8820tctggagact gtgagaatat ttctgacaga
gcagcctttg gaaggactag agaaactcta 8880ccaggagccc agagaactgc ctcctgaaga
aagagctcag aatgtcactc ggctcctacg 8940aaagcaggct gaagaggtca acgctgaatg
ggacaaattg aacctgcgct cagctgattg 9000gcagagaaaa atagatgaag ctcttgaaag
actccaggaa cttcaggaag ctgccgatga 9060actggacctc aagttgcgcc aagctgaggt
gatcaaggga tcctggcagc cagtggggga 9120tctcctcatt gactctctgc aagatcacct
tgaaaaagtc aaggcacttc ggggagaaat 9180tgcacctctt aaagagaatg tcaatcgtgt
caatgacctt gcacatcagc tgaccacact 9240gggcattcag ctctcacctt ataacctcag
cactttggaa gatctgaata ccagatggag 9300gcttctacag gtggctgtgg aggaccgtgt
cagacagctg catgaagccc acagggactt 9360tggtcctgca tcccagcact tcctttccac
ttcagttcag ggtccctggg agagagccat 9420ctcaccaaac aaagtgccct actatatcaa
ccacgagacc caaaccactt gttgggacca 9480ccccaaaatg acagagctct accagtcttt
agctgacctg aataatgtca ggttctccgc 9540gtataggact gccatgaagc tcagaaggct
ccagaaggcc ctttgcttgg atctcttgag 9600cctgtcagct gcatgtgatg ccctggacca
gcacaacctc aagcaaaatg accagcccat 9660ggatatcctg cagataatta actgtttgac
tacaatttat gatcgtctgg agcaagagca 9720caacaatctg gtcaatgtcc ctctctgtgt
ggatatgtgt ctcaactggc ttctcaatgt 9780ttatgatacg ggacgaacag ggaggatccg
tgtcctgtct tttaaaactg gcatcatttc 9840tctgtgtaaa gcacacttgg aagacaagta
cagatacctt ttcaagcaag tggcaagttc 9900aactggcttt tgtgaccagc gtaggctggg
tcttcttctg catgattcta ttcaaatccc 9960aagacagttg ggtgaagttg cttcctttgg
gggcagtaac attgagccga gtgtcaggag 10020ctgcttccaa tttgccaata ataaacctga
gattgaagct gctctcttcc ttgactggat 10080gcgcctggaa ccccagtcta tggtgtggct
gcccgtcttg cacagagtgg ctgctgctga 10140aactgccaag catcaagcca agtgtaacat
ctgtaaggag tgtccaatca ttggattcag 10200gtacagaagc ctaaagcatt ttaattatga
catctgccaa agttgctttt tttctggccg 10260agttgcaaag ggccataaaa tgcactaccc
catggtagag tattgcactc cgactacatc 10320cggagaagat gttcgcgact tcgccaaggt
actaaaaaac aaatttcgaa ccaaaaggta 10380ttttgcgaag catccccgaa tgggctacct
gccagtgcag actgtgttag agggggacaa 10440catggaaact cccgttactc tgatcaactt
ctggccagta gattctgcgc ctgcctcgtc 10500cccccagctt tcacacgatg atactcattc
acgcattgaa cattatgcta gcaggctagc 10560agaaatggaa aacagcaatg gatcttatct
aaatgatagc atctctccta atgagagcat 10620agatgatgaa catttgttaa tccagcatta
ctgccaaagt ttgaaccagg actcccccct 10680gagccagcct cgtagtcctg cccagatctt
gatttcctta gagagtgagg aaagagggga 10740gctagagaga atcctagcag atcttgagga
agaaaacagg aatctgcaag cagaatatga 10800tcgcctgaag cagcagcatg agcataaagg
cctgtctcca ctgccatctc ctcctgagat 10860gatgcccacc tctcctcaga gtcccaggga
tgctgagctc attgctgagg ctaagctact 10920gcgccaacac aaaggacgcc tggaagccag
gatgcaaatc ctggaagacc acaataaaca 10980gctggagtct cagttacata gactgagaca
gctcctggag cagccccagg ctgaagctaa 11040ggtgaatggc accacggtgt cctctccttc
cacctctctg cagaggtcag atagcagtca 11100gcctatgctg ctccgagtgg ttggcagtca
aacttcagaa tctatgggtg aggaagatct 11160tctgagtcct ccccaggaca caagcacagg
gttagaagaa gtgatggagc aactcaacaa 11220ctccttccct agttcaagag gaagaaatgc
ccccggaaag ccaatgagag aggacacaat 11280gtaggaagcc ttttccacat ggcagatgat
ttgggcagag cgatggagtc cttagtttca 11340gtcatgacag atgaagaagg agcagaataa
atgttttaca actcctgatt cccgcatggt 11400ttttataata ttcgtacaac aaagaggatt
agacagtaag agtttacaag aaataaaatc 11460tatatttttg tgaagggtag tggtactata
ctgtagattt cagtagtttc taagtctgtt 11520attgttttgt taacaatggc aggttttaca
cgtctatgca attgtacaaa aaagttaaaa 11580gaaaacatgt aaaatcttga tagctaaata
acttgccatt tctttatatg gaacgcattt 11640tgggttgttt aaaaatttat aacagttata
aagaaagatt gtaaactaaa gtgtgcttta 11700taaaaaaagt tgtttataaa aacccctaaa
caaacacaca cgcacacaca cacacacaca 11760cacacacaca cacacacgca cacatacatg
cacgaaccca ccacacacac acacacacac 11820acacacactg aggcagcaca ttgttttgca
ttactttagc gtggtattca tatggaattc 11880atgacgtttt tttattttct tgcatacgaa
ccccaccaaa tgactgcttc atattgctct 11940tttgagaatt gttgactgag tggggctggc
tatgggcttt cattttatac atctatatgt 12000ctacaagtat ataaatacta taggtatata
gataaataga tatgaagtta cttcttcaaa 12060tgttcttgcc acttcctaat ggaaattgct
tctagtcatc tgggcttatc tgcttgggca 12120agagtgaatt ttccctggag cccaaagcca
ggagactacc gccacactaa aatattgtct 12180agggctccag atgtttctag ttttaaactt
tccactgaga gctagaggat tcattttttt 12240caaggaacat gcgaatgaat acacaggact
tactatcata gtaatttgtt ggctgatata 12300ttcaacttcc tactgttggg ttatatttaa
tgatgtttct gcaatagaac atcagatgac 12360atttttaact cccagacagt aggaggaaga
tggtaggagc taaaggttgc ggctcctcag 12420tcaatttata tgaggggagc aacaactctg
taaaagaatg gatgaatatt tacaactata 12480catataaaca tctctataat tacaactaaa
ttgttctgcc ctcttcataa actcaacctg 12540aagtgggtgg ttttgttgtt gttgttgttg
ttgttgttga tgatgatgat gaattttaga 12600ttttagattt tttgggtttt tttttcttca
ttgtgatgat tttttttttt aatgctgcaa 12660gacttaggat tactgttaag aaagtaaccc
aatcacattg tgaccctggt gaatatcagt 12720ccagaagccc atgaactgca tttgtctcct
ttgcattggt ttccctgcaa gtaactccac 12780acaggattgt gggtgagaag gcacagtggt
tggaaagttt tgagagcaaa agcgtctcca 12840aactctctgg tctagttgac gggctgaaat
gtctaaacaa atgcaagtca ttgaaccagg 12900agaaaaagtg caacagaaag ctaaggactg
ctaggaagag ctttactcct ctcatgccag 12960tttcttcttc ttagcattta aagagcattc
tctcaataga aatcactgtc ctatcatttt 13020gcaaatctgt tacctctaac gtcaagtgta
attaacttct agcgagtggg ttttgtccat 13080tattaattgt aattaacatc aaacacagct
tctcatgcta tttctacctc actttggttt 13140tggggtgttt ctagtaattg tgcacaccta
atttcacaac ttcaccactt gtctgttgtg 13200tggacaccag tttccttttt tcatttataa
tttccaaaag aaaacccaaa gctctaagat 13260aacaaattga aatttggttc tggtcttgct
ttctctctct ctctctcctt tatgtggcac 13320tgggcatttt ctttatccaa ggatttgttt
tcaccaagat ttaaaacaag gggttccttt 13380cctactaaga agttttaagt ttcattctaa
aatccaaggt agatagagtg catagttttg 13440ttttaatctt ttcgttttat cttttagata
ttagttctgg agtgaatcta tcaaaatatt 13500tgaataaaaa ctgagagctt tattgctgat
tttaagcata atttggacat catttcatgt 13560tctttataac catcaagtat taaagtgtaa
atcataatca gtgtaactga agcataatca 13620tcacatggca tgtatcatca ttgtctccag
gtactggact cttacttgag tatcataata 13680gattgtgttt taacaccaac actgtaacat
ttactaatta tttttttaaa cttcagtttt 13740actgcatttt cacaacatat cagatttcac
caaatatatg ccttactatt gtattatatt 13800actgctttac tgtgtatctc aataaagcac
gcagttatgt tacaaaaaaa taaaaaa 138578213DNAMus
musculusmisc_featureNucleotide sequence of the murine dystrophin
gene DMD, exon 23 8gctctgcaaa gttctttgaa agagcaacaa aatggcttca actatctgag
tgacactgtg 60aaggagatgg ccaagaaagc accttcagaa atatgccaga aatatctgtc
agaatttgaa 120gagattgagg ggcactggaa gaaactttcc tcccagttgg tggaaagctg
ccaaaagcta 180gaagaacata tgaataaact tcgaaaattt cag
2139213DNAMus musculusmisc_featureNucleotide sequence of the
mutated dystrophin gene DMD from the mdx mouse, exon 23 9gctctgcaaa
gttctttgaa agagcaataa aatggcttca actatctgag tgacactgtg 60aaggagatgg
ccaagaaagc accttcagaa atatgccaga aatatctgtc agaatttgaa 120gagattgagg
ggcactggaa gaaactttcc tcccagttgg tggaaagctg ccaaaagcta 180gaagaacata
tgaataaact tcgaaaattt cag 21310114DNAMus
musculusmisc_featureNucleotide sequence of the murine dystrophin
gene DMD, exon 24 10aatcacataa aaaccttaca gaaatggatg gctgaagttg
atgttttcct gaaagaggaa 60tggcctgccc tgggggatgc tgaaatcctg aaaaaacagc
tcaaacaatg caga 11411913DNAMus musculusmisc_featureNucleotide
sequence of the murine dystrophin gene DMD, intron 22 11gtctgtggac
atttgaatat cataaataac aaagaacatg tcttatcagt caagagatca 60tattgatata
ttaaacttaa ggtaataatg aaaaagtaaa gataataatg aaaaatcata 120gattatgagt
tggaaaaata aacagaacaa tttgaccaaa aacatgactt tttcttattt 180ttttctatat
attattttat aaatatacag acataaatag atatatattt ttaaattaaa 240agtactgtat
taaaggaaag gtataatttc atttcatatt tagtgacata agatatgaag 300tatgattatt
aaaattaaat cacattattt tattataatt actttatttt taattcctaa 360tttctttaag
cttaggtaaa atcaatggat ttatataatt agttagaatt taaatattaa 420caaactataa
cactatgatt aaatgcttga tattgagtag ttattttaat agcctaagtc 480tggaaattaa
atactagtaa gagaaacttc tgtgatgtga ggacatataa agactaattt 540ttttgttgat
tctaaaaatc ccatgttgta tacttattct ttttaaatct gaaaatatat 600taatcatata
ttgcctaaat gtcttaataa tgtttcactg taggtaagtt aaaatgtatc 660acatatataa
taaacatagt tattaatgca tagatattca gtaaaattat gacttctaaa 720tttctgtcta
aatataatat gccctgtaat ataatagaaa ttattcataa gaatacatat 780atattgcttt
atcagatatt ctactttgtt tagatctcta aattacataa acttttattt 840accttcttct
tgatatgaat gaaactcatc aaatatgcgt gttagtgtaa atgaacttct 900atttaatttt
gag 913122607DNAMus
musculusmisc_featureNucleotide sequence of the murine dystrophin
gene DMD, intron 23 12gtaagccgag gtttggcctt taaactatat tttttcacat
agcaattaat tggaaaatgt 60gatgggaaac agatatttta cccagagtcc ttcaaagata
ttgatgatat caaaagccaa 120atctatttca aaggattgca acttgcctat ttttcctatg
aaaacagtaa tgtgtcatac 180cttcttggat tgtctgtata aatgaattga ttttttttca
ccaactccaa gtatacttaa 240cattttaaca taataattta aaatatcctt attccattat
gttcattttt taagttgtag 300atatgattta gctcacagca tacatatata cacatgtatt
acatatgcat atattatata 360tatggcagac atatgttttc actaccatat ttcacttttg
aattatgaat atatgtttaa 420tttctgccat atttccttcc ctacattgac ttctattaat
ttagtatttc agtagttcta 480acacattaat aataacctag actcaataca gtaatctaac
aattatattt gtgcctgtaa 540ttctaagtta gttaaattca taggttgtgt ttctcatagt
tggccatttg tgaaatataa 600taatatccga aaagaaagtt caaaaatgtc atgacttcat
atagagttat tgaaacagtg 660cccttacttt cattctggcc atgctagtga cttgatcatt
cttgtatttt acagctaaaa 720cactaccaaa agtgtcaaat ccatgatcta catgtttgac
tgaggctagc agcacttatt 780ccacccttat atgaagcctt taagagaaag tatatttgtt
tgctattttt aacttcttga 840aggaacatac aatctttgtt tcaagagctc atcctctttc
atgctagtaa attttggtgg 900cattgcatcc atgtctgact ctgaatctgt ttctgtctat
cctgctccct aacactgtac 960catcttcctt tttgaaaaaa aaatattgaa ttattttatt
tatttacttt ccaaagttgc 1020tcctgcctgt tcctccttct ccaagttctt cagtcccccc
tgctccccac cgatgagagg 1080gaaaggtcct gaattcactg ggctccatgg gggtcctttt
gcattttctt aaccttctta 1140ataaaatagg ccttctagaa ttatatcata tacattgtga
tatgacaaat gataaagtat 1200attgttcaga gttttacctt gttcatattt gcaatgtccc
cctgtcatgc tggatattct 1260ttgattgggt atatttgcta acagattaag tatatttatc
ttcgttaagc agtataactt 1320attaagaaag aactctatta atatgagaaa taactaatga
aacaccactc cacaggtgat 1380ttcagccact ttatgaactg ctggaagcaa aaatgagatc
tttgcaacat gaagcagttg 1440ctcagttcat taaactgtgt tcaatatttc agccataaca
tacattagag aatgatttat 1500attgttcaaa catttggtgc tctatttttg catgacgtgg
gattaaacac agcaccaaca 1560atcaaacaat tgcaaagatg tattacaagt attttttctt
tttaaaacag gaaagtatac 1620ttatatttcc attgtccaaa ccatcatgaa agggatagag
attactgaca caaatttaga 1680gaaaggattt gagtggagta agaattaaat gaaccaaaga
agaattaatg tattcatcaa 1740gaagtcatgg aggtgaaatt ggccttgaat gataccacta
aggagagaat gttgagatcc 1800ttatatttag tcaattgttt ttaaatctgt agttattaac
cacattttaa tcatattgaa 1860agggaaattt tctgtgatgc atgtattttc aatataaatt
ttagaaaaga agacaattat 1920aacttgattt tgtgaattac atggaactaa agaaatgaca
gatttacatt tgaaaattga 1980ctgaactaaa gtacataaat aaaagtcata cagaaaaatg
tgggaggtgc ttgtccattt 2040ataaaggaca aaaatgccat ttgttgccta atcattattt
cttattggtc agaccaataa 2100gaaatcaaga gctttgactt taaaggtaag aaaatcttac
cttaaaatcc ccaactgaag 2160ggactgttta aactgtcaac tgcagaaaac aagttatgga
agttcaggtt tagggaaact 2220ataaacacac cataacattg agtttatgtg catagtttgt
tttatgtaca gtgagagtaa 2280attgttagta ttatcatgag ttgttttgaa acttcaaatt
tctctagagg ggtatgattt 2340aatgttctca agaggaacat aataaaacca tatctggtat
tagtttttat ttttaacaat 2400agcagacttc atacaccaat gttcacagtg tagaccataa
aatgcagtct tagtaaaaat 2460attattctct ataaagctac aatgagacct ccctcaaaca
tacattgttt ttttttttct 2520aacttatgtt tggatatatc atcatgatga actatgttaa
aaacaatcag agcttagtaa 2580tactttcata ttgctttttt attccag
260713150DNAArtificial SequenceAntisense sequence
AS1 13cttcatatct tatgtcacta aatatgaaat gaaattatac ctttccttta atacagtact
60tttaatttaa aaatatatat ctatttatgt ctgtatattt ataaaataat atatagaaaa
120aaataagaaa aagtcatgtt tttggtcaaa
15014150DNAArtificial SequenceAntisense sequence AS2 14atatattttc
agatttaaaa agaataagta tacaacatgg gatttttaga atcaacaaaa 60aaattagtct
ttatatgtcc tcacatcaca gaagtttctc ttactagtat ttaatttcca 120gacttaggct
attaaaataa ctactcaata
15015150DNAArtificial SequenceAntisense sequence AS3 15aaatagaagt
tcatttacac taacacgcat atttgatgag tttcattcat atcaagaaga 60aggtaaataa
aagtttatgt aatttagaga tctaaacaaa gtagaatatc tgataaagca 120atatatatgt
attcttatga ataatttcta
15016150DNAArtificial SequenceAntisense sequence AS4 16ttagttccat
gtaattcaca aaatcaagtt ataattgtct tcttttctaa aatttatatt 60gaaaatacat
gcatcacaga aaatttccct ttcaatatga ttaaaatgtg gttaataact 120acagatttaa
aaacaattga ctaaatataa
15017150DNAArtificial SequenceAntisense sequence AS5 17cacataaact
caatgttatg gtgtgtttat agtttcccta aacctgaact tccataactt 60gttttctgca
gttgacagtt taaacagtcc cttcagttgg ggattttaag gtaagatttt 120cttaccttta
aagtcaaagc tcttgatttc
15018150DNAArtificial SequenceAntisense sequence AS6 18tcatcatgat
gatatatcca aacataagtt agaaaaaaaa aaacaatgta tgtttgaggg 60aggtctcatt
gtagctttat agagaataat atttttacta agactgcatt ttatggtcta 120cactgtgaac
attggtgtat gaagtctgct
15019150DNAArtificial SequenceAntisense sequence AS7 19caagttgcaa
tcctttgaaa tagatttggc ttttgatatc atcaatatct ttgaaggact 60ctgggtaaaa
tatctgtttc ccatcacatt ttccaattaa ttgctatgtg aaaaaatata 120gtttaaaggc
caaacctcgg cttacctgaa
15020150DNAArtificial SequenceAntisense sequence AS8 20ctacaactta
aaaaatgaac ataatggaat aaggatattt taaattatta tgttaaaatg 60ttaagtatac
ttggagttgg tgaaaaaaaa tcaattcatt tatacagaca atccaagaag 120gtatgacaca
ttactgtttt cataggaaaa
15021306DNAArtificial SequenceAntisense sequence 2XAS4 21ttagttccat
gtaattcaca aaatcaagtt ataattgtct tcttttctaa aatttatatt 60gaaaatacat
gcatcacaga aaatttccct ttcaatatga ttaaaatgtg gttaataact 120acagatttaa
aaacaattga ctaaatataa gaattcttag ttccatgtaa ttcacaaaat 180caagttataa
ttgtcttctt ttctaaaatt tatattgaaa atacatgcat cacagaaaat 240ttccctttca
atatgattaa aatgtggtta ataactacag atttaaaaac aattgactaa 300atataa
306222250DNAArtificial SequenceNucleotide sequence located in the 5'-half
of the nucleotide sequence of intron 23 22gtaagccgag gtttggcctt
taaactatat tttttcacat agcaattaat tggaaaatgt 60gatgggaaac agatatttta
cccagagtcc ttcaaagata ttgatgatat caaaagccaa 120atctatttca aaggattgca
acttgcctat ttttcctatg aaaacagtaa tgtgtcatac 180cttcttggat tgtctgtata
aatgaattga ttttttttca ccaactccaa gtatacttaa 240cattttaaca taataattta
aaatatcctt attccattat gttcattttt taagttgtag 300atatgattta gctcacagca
tacatatata cacatgtatt acatatgcat atattatata 360tatggcagac atatgttttc
actaccatat ttcacttttg aattatgaat atatgtttaa 420tttctgccat atttccttcc
ctacattgac ttctattaat ttagtatttc agtagttcta 480acacattaat aataacctag
actcaataca gtaatctaac aattatattt gtgcctgtaa 540ttctaagtta gttaaattca
taggttgtgt ttctcatagt tggccatttg tgaaatataa 600taatatccga aaagaaagtt
caaaaatgtc atgacttcat atagagttat tgaaacagtg 660cccttacttt cattctggcc
atgctagtga cttgatcatt cttgtatttt acagctaaaa 720cactaccaaa agtgtcaaat
ccatgatcta catgtttgac tgaggctagc agcacttatt 780ccacccttat atgaagcctt
taagagaaag tatatttgtt tgctattttt aacttcttga 840aggaacatac aatctttgtt
tcaagagctc atcctctttc atgctagtaa attttggtgg 900cattgcatcc atgtctgact
ctgaatctgt ttctgtctat cctgctccct aacactgtac 960catcttcctt tttgaaaaaa
aaatattgaa ttattttatt tatttacttt ccaaagttgc 1020tcctgcctgt tcctccttct
ccaagttctt cagtcccccc tgctccccac cgatgagagg 1080gaaaggtcct gaattcactg
ggctccatgg gggtcctttt gcattttctt aaccttctta 1140ataaaatagg ccttctagaa
ttatatcata tacattgtga tatgacaaat gataaagtat 1200attgttcaga gttttacctt
gttcatattt gcaatgtccc cctgtcatgc tggatattct 1260ttgattgggt atatttgcta
acagattaag tatatttatc ttcgttaagc agtataactt 1320attaagaaag aactctatta
atatgagaaa taactaatga aacaccactc cacaggtgat 1380ttcagccact ttatgaactg
ctggaagcaa aaatgagatc tttgcaacat gaagcagttg 1440ctcagttcat taaactgtgt
tcaatatttc agccataaca tacattagag aatgatttat 1500attgttcaaa catttggtgc
tctatttttg catgacgtgg gattaaacac agcaccaaca 1560atcaaacaat tgcaaagatg
tattacaagt attttttctt tttaaaacag gaaagtatac 1620ttatatttcc attgtccaaa
ccatcatgaa agggatagag attactgaca caaatttaga 1680gaaaggattt gagtggagta
agaattaaat gaaccaaaga agaattaatg tattcatcaa 1740gaagtcatgg aggtgaaatt
ggccttgaat gataccacta aggagagaat gttgagatcc 1800ttatatttag tcaattgttt
ttaaatctgt agttattaac cacattttaa tcatattgaa 1860agggaaattt tctgtgatgc
atgtattttc aatataaatt ttagaaaaga agacaattat 1920aacttgattt tgtgaattac
atggaactaa agaaatgaca gatttacatt tgaaaattga 1980ctgaactaaa gtacataaat
aaaagtcata cagaaaaatg tgggaggtgc ttgtccattt 2040ataaaggaca aaaatgccat
ttgttgccta atcattattt cttattggtc agaccaataa 2100gaaatcaaga gctttgactt
taaaggtaag aaaatcttac cttaaaatcc ccaactgaag 2160ggactgttta aactgtcaac
tgcagaaaac aagttatgga agttcaggtt tagggaaact 2220ataaacacac cataacattg
agtttatgtg 22502334DNAArtificial
SequenceSpacer 1 (3' end) 23acagctcttt ctttccatgg gttggcctga attc
342442DNAArtificial SequenceSpacer 2 (5'-end)
24ctcgagagat ctccgcggaa cattattata acgttgctcg aa
42257DNAArtificial SequenceConserved yeast branch point sequence
25tactaac
72630DNAArtificial SequenceIntronic Splice Enhancer (ISE) sequence
26ggctgaggga aggactgtcc tggggactgg
302724DNAArtificial SequenceDownstream Intronic Splice Enhancer (DISE)
sequence 27ctctttcttt ccatgggttg gcct
242820DNAArtificial SequencePolypyrimidine tract sequence
28tctcttcttt ttttttttcc
20299DNAArtificial SequenceCanonical 3' acceptor splice site 29ggaaaacag
9306DNAArtificial Sequence5' donor splice site 30gtaaga
63141DNAArtificial
Sequence3' splice region of the Pre-Trans-Splicing molecule gene
31tactaactga tatctcttct tttttttttt ccggaaaaca g
413219DNAArtificial SequenceForward primer for E23 32agatggccaa gaaagcacc
193319DNAArtificial
SequenceReverse primer for E23 33ctttccacca actgggagg
193423DNAArtificial SequenceForward primer
for E24 34tgaaaaaaca gctcaaacaa tgc
233519DNAArtificial SequenceReverse primer for E24 35agcatccccc
agggcaggc
193621DNAArtificial SequenceForward primer E22-F 36gacactttac caccaatgcg
c 213720DNAArtificial
SequenceReverse primer pSMD2-R1 37ctttctgata ggcagcctgc
203820DNAArtificial SequencePrimer pSMD2-R5
38ctcaccctga agttctcagg
20393996DNAArtificial SequenceThe murine DMD minigene target comprising
exons E22, E23, and E24 39tttttgacac tttaccacca atgcgctatc aggagacaat
gagtagcatc aggacgtgga 60tccagcagtc agaaagcaaa ctctctgtac cttatcttag
tgttactgaa tatgaaataa 120tggaggagag actcgggaaa ttacaggtct gtggacattt
gaatatcata aataacaaag 180aacatgtctt atcagtcaag agatcatatt gatatattaa
acttaaggta ataatgaaaa 240agtaaagata ataatgaaaa atcatagatt atgagttgga
aaaataaaca gaacaatttg 300accaaaaaca tgactttttc ttattttttt ctatatatta
ttttataaat atacagacat 360aaatagatat atatttttaa attaaaagta ctgtattaaa
ggaaaggtat aatttcattt 420catatttagt gacataagat atgaagtatg attattaaaa
ttaaatcaca ttattttatt 480ataattactt tatttttaat tcctaatttc tttaagctta
ggtaaaatca atggatttat 540ataattagtt agaatttaaa tattaacaaa ctataacact
atgattaaat gcttgatatt 600gagtagttat tttaatagcc taagtctgga aattaaatac
tagtaagaga aacttctgtg 660atgtgaggac atataaagac taattttttt gttgattcta
aaaatcccat gttgtatact 720tattcttttt aaatctgaaa atatattaat catatattgc
ctaaatgtct taataatgtt 780tcactgtagg taagttaaaa tgtatcacat atataataaa
catagttatt aatgcataga 840tattcagtaa aattatgact tctaaatttc tgtctaaata
taatatgccc tgtaatataa 900tagaaattat tcataagaat acatatatat tgctttatca
gatattctac tttgtttaga 960tctctaaatt acataaactt ttatttacct tcttcttgat
atgaatgaaa ctcatcaaat 1020atgcgtgtta gtgtaaatga acttctattt aattttgagg
ctctgcaaag ttctttgaaa 1080gagcaataaa atggcttcaa ctatctgagt gacactgtga
aggagatggc caagaaagca 1140ccttcagaaa tatgccagaa atatctgtca gaatttgaag
agattgaggg gcactggaag 1200aaactttcct cccagttggt ggaaagctgc caaaagctag
aagaacatat gaataaactt 1260cgaaaatttc aggtaagccg aggtttggcc tttaaactat
attttttcac atagcaatta 1320attggaaaat gtgatgggaa acagatattt tacccagagt
ccttcaaaga tattgatgat 1380atcaaaagcc aaatctattt caaaggattg caacttgcct
atttttccta tgaaaacagt 1440aatgtgtcat accttcttgg attgtctgta taaatgaatt
gatttttttt caccaactcc 1500aagtatactt aacattttaa cataataatt taaaatatcc
ttattccatt atgttcattt 1560tttaagttgt agatatgatt tagctcacag catacatata
tacacatgta ttacatatgc 1620atatattata tatatggcag acatatgttt tcactaccat
atttcacttt tgaattatga 1680atatatgttt aatttctgcc atatttcctt ccctacattg
acttctatta atttagtatt 1740tcagtagttc taacacatta ataataacct agactcaata
cagtaatcta acaattatat 1800ttgtgcctgt aattctaagt tagttaaatt cataggttgt
gtttctcata gttggccatt 1860tgtgaaatat aataatatcc gaaaagaaag ttcaaaaatg
tcatgacttc atatagagtt 1920attgaaacag tgcccttact ttcattctgg ccatgctagt
gacttgatca ttcttgtatt 1980ttacagctaa aacactacca aaagtgtcaa atccatgatc
tacatgtttg actgaggcta 2040gcagcactta ttccaccctt atatgaagcc tttaagagaa
agtatatttg tttgctattt 2100ttaacttctt gaaggaacat acaatctttg tttcaagagc
tcatcctctt tcatgctagt 2160aaattttggt ggcattgcat ccatgtctga ctctgaatct
gtttctgtct atcctgctcc 2220ctaacactgt accatcttcc tttttgaaaa aaaaatattg
aattatttta tttatttact 2280ttccaaagtt gctcctgcct gttcctcctt ctccaagttc
ttcagtcccc cctgctcccc 2340accgatgaga gggaaaggtc ctgaattcac tgggctccat
gggggtcctt ttgcattttc 2400ttaaccttct taataaaata ggccttctag aattatatca
tatacattgt gatatgacaa 2460atgataaagt atattgttca gagttttacc ttgttcatat
ttgcaatgtc cccctgtcat 2520gctggatatt ctttgattgg gtatatttgc taacagatta
agtatattta tcttcgttaa 2580gcagtataac ttattaagaa agaactctat taatatgaga
aataactaat gaaacaccac 2640tccacaggtg atttcagcca ctttatgaac tgctggaagc
aaaaatgaga tctttgcaac 2700atgaagcagt tgctcagttc attaaactgt gttcaatatt
tcagccataa catacattag 2760agaatgattt atattgttca aacatttggt gctctatttt
tgcatgacgt gggattaaac 2820acagcaccaa caatcaaaca attgcaaaga tgtattacaa
gtattttttc tttttaaaac 2880aggaaagtat acttatattt ccattgtcca aaccatcatg
aaagggatag agattactga 2940cacaaattta gagaaaggat ttgagtggag taagaattaa
atgaaccaaa gaagaattaa 3000tgtattcatc aagaagtcat ggaggtgaaa ttggccttga
atgataccac taaggagaga 3060atgttgagat ccttatattt agtcaattgt ttttaaatct
gtagttatta accacatttt 3120aatcatattg aaagggaaat tttctgtgat gcatgtattt
tcaatataaa ttttagaaaa 3180gaagacaatt ataacttgat tttgtgaatt acatggaact
aaagaaatga cagatttaca 3240tttgaaaatt gactgaacta aagtacataa ataaaagtca
tacagaaaaa tgtgggaggt 3300gcttgtccat ttataaagga caaaaatgcc atttgttgcc
taatcattat ttcttattgg 3360tcagaccaat aagaaatcaa gagctttgac tttaaaggta
agaaaatctt accttaaaat 3420ccccaactga agggactgtt taaactgtca actgcagaaa
acaagttatg gaagttcagg 3480tttagggaaa ctataaacac accataacat tgagtttatg
tgcatagttt gttttatgta 3540cagtgagagt aaattgttag tattatcatg agttgttttg
aaacttcaaa tttctctaga 3600ggggtatgat ttaatgttct caagaggaac ataataaaac
catatctggt attagttttt 3660atttttaaca atagcagact tcatacacca atgttcacag
tgtagaccat aaaatgcagt 3720cttagtaaaa atattattct ctataaagct acaatgagac
ctccctcaaa catacattgt 3780tttttttttt ctaacttatg tttggatata tcatcatgat
gaactatgtt aaaaacaatc 3840agagcttagt aatactttca tattgctttt ttattccaga
atcacataaa aaccttacag 3900aaatggatgg ctgaagttga tgttttcctg aaagaggaat
ggcctgccct gggggatgct 3960gaaatcctga aaaaacagct caaacaatgc agataa
399640347DNAArtificial SequenceTS molecules AS1-E24
40cttcatatct tatgtcacta aatatgaaat gaaattatac ctttccttta atacagtact
60tttaatttaa aaatatatat ctatttatgt ctgtatattt ataaaataat atatagaaaa
120aaataagaaa aagtcatgtt tttggtcaaa ctcgagagat ctccgcggaa cattattata
180acgttgctcg aatactaact gatatctctt cttttttttt ttccggaaaa cagaatcaca
240taaaaacctt acagaaatgg atggctgaag ttgatgtttt cctgaaagag gaatggcctg
300ccctggggga tgctgaaatc ctgaaaaaac agctcaaaca atgcaga
34741347DNAArtificial SequenceTS molecule AS2-E24 41atatattttc agatttaaaa
agaataagta tacaacatgg gatttttaga atcaacaaaa 60aaattagtct ttatatgtcc
tcacatcaca gaagtttctc ttactagtat ttaatttcca 120gacttaggct attaaaataa
ctactcaata ctcgagagat ctccgcggaa cattattata 180acgttgctcg aatactaact
gatatctctt cttttttttt ttccggaaaa cagaatcaca 240taaaaacctt acagaaatgg
atggctgaag ttgatgtttt cctgaaagag gaatggcctg 300ccctggggga tgctgaaatc
ctgaaaaaac agctcaaaca atgcaga 34742347DNAArtificial
SequenceTS molecule AS3-E24 42aaatagaagt tcatttacac taacacgcat atttgatgag
tttcattcat atcaagaaga 60aggtaaataa aagtttatgt aatttagaga tctaaacaaa
gtagaatatc tgataaagca 120atatatatgt attcttatga ataatttcta ctcgagagat
ctccgcggaa cattattata 180acgttgctcg aatactaact gatatctctt cttttttttt
ttccggaaaa cagaatcaca 240taaaaacctt acagaaatgg atggctgaag ttgatgtttt
cctgaaagag gaatggcctg 300ccctggggga tgctgaaatc ctgaaaaaac agctcaaaca
atgcaga 34743185DNAArtificial SequenceTrans-Splicing
molecule with no binding domain (AS-) 43ccgcggaaca ttattataac
gttgctcgaa tactaactga tatctcttct tttttttttt 60ccggaaaaca gaatcacata
aaaaccttac agaaatggat ggctgaagtt gatgttttcc 120tgaaagagga atggcctgcc
ctgggggatg ctgaaatcct gaaaaaacag ctcaaacaat 180gcaga
18544515DNAArtificial
SequenceTS molecule AS1-E24-AS4 44cttcatatct tatgtcacta aatatgaaat
gaaattatac ctttccttta atacagtact 60tttaatttaa aaatatatat ctatttatgt
ctgtatattt ataaaataat atatagaaaa 120aaataagaaa aagtcatgtt tttggtcaaa
ctcgagagat ctccgcggaa cattattata 180acgttgctcg aatactaact gatatctctt
cttttttttt ttccggaaaa cagaatcaca 240taaaaacctt acagaaatgg atggctgaag
ttgatgtttt cctgaaagag gaatggcctg 300ccctggggga tgctgaaatc ctgaaaaaac
agctcaaaca atgcagagta agaagatctg 360aattcttagt tccatgtaat tcacaaaatc
aagttataat tgtcttcttt tctaaaattt 420atattgaaaa tacatgcatc acagaaaatt
tccctttcaa tatgattaaa atgtggttaa 480taactacaga tttaaaaaca attgactaaa
tataa 51545515DNAArtificial SequenceTS
molecule AS1-E24-AS5 45cttcatatct tatgtcacta aatatgaaat gaaattatac
ctttccttta atacagtact 60tttaatttaa aaatatatat ctatttatgt ctgtatattt
ataaaataat atatagaaaa 120aaataagaaa aagtcatgtt tttggtcaaa ctcgagagat
ctccgcggaa cattattata 180acgttgctcg aatactaact gatatctctt cttttttttt
ttccggaaaa cagaatcaca 240taaaaacctt acagaaatgg atggctgaag ttgatgtttt
cctgaaagag gaatggcctg 300ccctggggga tgctgaaatc ctgaaaaaac agctcaaaca
atgcagagta agaagatctg 360aattccacat aaactcaatg ttatggtgtg tttatagttt
ccctaaacct gaacttccat 420aacttgtttt ctgcagttga cagtttaaac agtcccttca
gttggggatt ttaaggtaag 480attttcttac ctttaaagtc aaagctcttg atttc
51546515DNAArtificial SequenceTS molecule
AS1-E24-AS6 46cttcatatct tatgtcacta aatatgaaat gaaattatac ctttccttta
atacagtact 60tttaatttaa aaatatatat ctatttatgt ctgtatattt ataaaataat
atatagaaaa 120aaataagaaa aagtcatgtt tttggtcaaa ctcgagagat ctccgcggaa
cattattata 180acgttgctcg aatactaact gatatctctt cttttttttt ttccggaaaa
cagaatcaca 240taaaaacctt acagaaatgg atggctgaag ttgatgtttt cctgaaagag
gaatggcctg 300ccctggggga tgctgaaatc ctgaaaaaac agctcaaaca atgcagagta
agaagatctg 360aattctcatc atgatgatat atccaaacat aagttagaaa aaaaaaaaca
atgtatgttt 420gagggaggtc tcattgtagc tttatagaga ataatatttt tactaagact
gcattttatg 480gtctacactg tgaacattgg tgtatgaagt ctgct
51547515DNAArtificial SequenceTS molecule AS1-E24-AS7
47cttcatatct tatgtcacta aatatgaaat gaaattatac ctttccttta atacagtact
60tttaatttaa aaatatatat ctatttatgt ctgtatattt ataaaataat atatagaaaa
120aaataagaaa aagtcatgtt tttggtcaaa ctcgagagat ctccgcggaa cattattata
180acgttgctcg aatactaact gatatctctt cttttttttt ttccggaaaa cagaatcaca
240taaaaacctt acagaaatgg atggctgaag ttgatgtttt cctgaaagag gaatggcctg
300ccctggggga tgctgaaatc ctgaaaaaac agctcaaaca atgcagagta agaagatctg
360aattccaagt tgcaatcctt tgaaatagat ttggcttttg atatcatcaa tatctttgaa
420ggactctggg taaaatatct gtttcccatc acattttcca attaattgct atgtgaaaaa
480atatagttta aaggccaaac ctcggcttac ctgaa
51548515DNAArtificial SequenceTS molecule AS2-E24-AS4 48atatattttc
agatttaaaa agaataagta tacaacatgg gatttttaga atcaacaaaa 60aaattagtct
ttatatgtcc tcacatcaca gaagtttctc ttactagtat ttaatttcca 120gacttaggct
attaaaataa ctactcaata ctcgagagat ctccgcggaa cattattata 180acgttgctcg
aatactaact gatatctctt cttttttttt ttccggaaaa cagaatcaca 240taaaaacctt
acagaaatgg atggctgaag ttgatgtttt cctgaaagag gaatggcctg 300ccctggggga
tgctgaaatc ctgaaaaaac agctcaaaca atgcagagta agaagatctg 360aattcttagt
tccatgtaat tcacaaaatc aagttataat tgtcttcttt tctaaaattt 420atattgaaaa
tacatgcatc acagaaaatt tccctttcaa tatgattaaa atgtggttaa 480taactacaga
tttaaaaaca attgactaaa tataa
51549515DNAArtificial SequenceTS molecule AS2-E24-AS5 49atatattttc
agatttaaaa agaataagta tacaacatgg gatttttaga atcaacaaaa 60aaattagtct
ttatatgtcc tcacatcaca gaagtttctc ttactagtat ttaatttcca 120gacttaggct
attaaaataa ctactcaata ctcgagagat ctccgcggaa cattattata 180acgttgctcg
aatactaact gatatctctt cttttttttt ttccggaaaa cagaatcaca 240taaaaacctt
acagaaatgg atggctgaag ttgatgtttt cctgaaagag gaatggcctg 300ccctggggga
tgctgaaatc ctgaaaaaac agctcaaaca atgcagagta agaagatctg 360aattccacat
aaactcaatg ttatggtgtg tttatagttt ccctaaacct gaacttccat 420aacttgtttt
ctgcagttga cagtttaaac agtcccttca gttggggatt ttaaggtaag 480attttcttac
ctttaaagtc aaagctcttg atttc
51550515DNAArtificial SequenceTS molecule AS2-E24-AS6 50atatattttc
agatttaaaa agaataagta tacaacatgg gatttttaga atcaacaaaa 60aaattagtct
ttatatgtcc tcacatcaca gaagtttctc ttactagtat ttaatttcca 120gacttaggct
attaaaataa ctactcaata ctcgagagat ctccgcggaa cattattata 180acgttgctcg
aatactaact gatatctctt cttttttttt ttccggaaaa cagaatcaca 240taaaaacctt
acagaaatgg atggctgaag ttgatgtttt cctgaaagag gaatggcctg 300ccctggggga
tgctgaaatc ctgaaaaaac agctcaaaca atgcagagta agaagatctg 360aattctcatc
atgatgatat atccaaacat aagttagaaa aaaaaaaaca atgtatgttt 420gagggaggtc
tcattgtagc tttatagaga ataatatttt tactaagact gcattttatg 480gtctacactg
tgaacattgg tgtatgaagt ctgct
51551515DNAArtificial SequenceTS molecule AS2-E24-AS7 51atatattttc
agatttaaaa agaataagta tacaacatgg gatttttaga atcaacaaaa 60aaattagtct
ttatatgtcc tcacatcaca gaagtttctc ttactagtat ttaatttcca 120gacttaggct
attaaaataa ctactcaata ctcgagagat ctccgcggaa cattattata 180acgttgctcg
aatactaact gatatctctt cttttttttt ttccggaaaa cagaatcaca 240taaaaacctt
acagaaatgg atggctgaag ttgatgtttt cctgaaagag gaatggcctg 300ccctggggga
tgctgaaatc ctgaaaaaac agctcaaaca atgcagagta agaagatctg 360aattccaagt
tgcaatcctt tgaaatagat ttggcttttg atatcatcaa tatctttgaa 420ggactctggg
taaaatatct gtttcccatc acattttcca attaattgct atgtgaaaaa 480atatagttta
aaggccaaac ctcggcttac ctgaa
51552515DNAArtificial SequenceTS molecule AS2-E24-AS8 52atatattttc
agatttaaaa agaataagta tacaacatgg gatttttaga atcaacaaaa 60aaattagtct
ttatatgtcc tcacatcaca gaagtttctc ttactagtat ttaatttcca 120gacttaggct
attaaaataa ctactcaata ctcgagagat ctccgcggaa cattattata 180acgttgctcg
aatactaact gatatctctt cttttttttt ttccggaaaa cagaatcaca 240taaaaacctt
acagaaatgg atggctgaag ttgatgtttt cctgaaagag gaatggcctg 300ccctggggga
tgctgaaatc ctgaaaaaac agctcaaaca atgcagagta agaagatctg 360aattcctaca
acttaaaaaa tgaacataat ggaataagga tattttaaat tattatgtta 420aaatgttaag
tatacttgga gttggtgaaa aaaaatcaat tcatttatac agacaatcca 480agaaggtatg
acacattact gttttcatag gaaaa
51553671DNAArtificial SequenceTS molecule AS2-E24-2XAS4 53atatattttc
agatttaaaa agaataagta tacaacatgg gatttttaga atcaacaaaa 60aaattagtct
ttatatgtcc tcacatcaca gaagtttctc ttactagtat ttaatttcca 120gacttaggct
attaaaataa ctactcaata ctcgagagat ctccgcggaa cattattata 180acgttgctcg
aatactaact gatatctctt cttttttttt ttccggaaaa cagaatcaca 240taaaaacctt
acagaaatgg atggctgaag ttgatgtttt cctgaaagag gaatggcctg 300ccctggggga
tgctgaaatc ctgaaaaaac agctcaaaca atgcagagta agaagatctg 360aattcttagt
tccatgtaat tcacaaaatc aagttataat tgtcttcttt tctaaaattt 420atattgaaaa
tacatgcatc acagaaaatt tccctttcaa tatgattaaa atgtggttaa 480taactacaga
tttaaaaaca attgactaaa tataagaatt cttagttcca tgtaattcac 540aaaatcaagt
tataattgtc ttcttttcta aaatttatat tgaaaataca tgcatcacag 600aaaatttccc
tttcaatatg attaaaatgt ggttaataac tacagattta aaaacaattg 660actaaatata a
67154543DNAArtificial SequenceTS molecule AS2-E24-AS4 comprising DISE
54atatattttc agatttaaaa agaataagta tacaacatgg gatttttaga atcaacaaaa
60aaattagtct ttatatgtcc tcacatcaca gaagtttctc ttactagtat ttaatttcca
120gacttaggct attaaaataa ctactcaata ctcgagagat ctccgcggaa cattattata
180acgttgctcg aatactaact gatatctctt cttttttttt ttccggaaaa cagaatcaca
240taaaaacctt acagaaatgg atggctgaag ttgatgtttt cctgaaagag gaatggcctg
300ccctggggga tgctgaaatc ctgaaaaaac agctcaaaca atgcagagta agaacagctc
360tttctttcca tgggttggcc tagatctgaa ttcttagttc catgtaattc acaaaatcaa
420gttataattg tcttcttttc taaaatttat attgaaaata catgcatcac agaaaatttc
480cctttcaata tgattaaaat gtggttaata actacagatt taaaaacaat tgactaaata
540taa
54355537DNAArtificial SequenceTS molecule AS2-E24-AS8 comprising DISE
55atatattttc agatttaaaa agaataagta tacaacatgg gatttttaga atcaacaaaa
60aaattagtct ttatatgtcc tcacatcaca gaagtttctc ttactagtat ttaatttcca
120gacttaggct attaaaataa ctactcaata ctcgagagat ctccgcggaa cattattata
180acgttgctcg aatactaact gatatctctt cttttttttt ttccggaaaa cagaatcaca
240taaaaacctt acagaaatgg atggctgaag ttgatgtttt cctgaaagag gaatggcctg
300ccctggggga tgctgaaatc ctgaaaaaac agctcaaaca atgcagagta agaacagctc
360tttctttcca tgggttggcc tgaattccta caacttaaaa aatgaacata atggaataag
420gatattttaa attattatgt taaaatgtta agtatacttg gagttggtga aaaaaaatca
480attcatttat acagacaatc caagaaggta tgacacatta ctgttttcat aggaaaa
53756545DNAArtificial SequenceTS molecule AS2-ISE-E24-AS4 56atatattttc
agatttaaaa agaataagta tacaacatgg gatttttaga atcaacaaaa 60aaattagtct
ttatatgtcc tcacatcaca gaagtttctc ttactagtat ttaatttcca 120gacttaggct
attaaaataa ctactcaata ctcgagagat ctccgcggaa cattattata 180acgttgctcg
aaggctgagg gaaggactgt cctggggact ggtactaact gatatctctt 240cttttttttt
ttccggaaaa cagaatcaca taaaaacctt acagaaatgg atggctgaag 300ttgatgtttt
cctgaaagag gaatggcctg ccctggggga tgctgaaatc ctgaaaaaac 360agctcaaaca
atgcagagta agaagatctg aattcttagt tccatgtaat tcacaaaatc 420aagttataat
tgtcttcttt tctaaaattt atattgaaaa tacatgcatc acagaaaatt 480tccctttcaa
tatgattaaa atgtggttaa taactacaga tttaaaaaca attgactaaa 540tataa
54557699DNAArtificial SequenceTS molecule AS2-E24-DISE-2XAS4 57atatattttc
agatttaaaa agaataagta tacaacatgg gatttttaga atcaacaaaa 60aaattagtct
ttatatgtcc tcacatcaca gaagtttctc ttactagtat ttaatttcca 120gacttaggct
attaaaataa ctactcaata ctcgagagat ctccgcggaa cattattata 180acgttgctcg
aatactaact gatatctctt cttttttttt ttccggaaaa cagaatcaca 240taaaaacctt
acagaaatgg atggctgaag ttgatgtttt cctgaaagag gaatggcctg 300ccctggggga
tgctgaaatc ctgaaaaaac agctcaaaca atgcagagta agaacagctc 360tttctttcca
tgggttggcc tagatctgaa ttcttagttc catgtaattc acaaaatcaa 420gttataattg
tcttcttttc taaaatttat attgaaaata catgcatcac agaaaatttc 480cctttcaata
tgattaaaat gtggttaata actacagatt taaaaacaat tgactaaata 540taagaattct
tagttccatg taattcacaa aatcaagtta taattgtctt cttttctaaa 600atttatattg
aaaatacatg catcacagaa aatttccctt tcaatatgat taaaatgtgg 660ttaataacta
cagatttaaa aacaattgac taaatataa
69958164DNAArtificial SequenceAntisense sequence AS3bis 58agagcctcaa
aattaaatag aagttcattt acactaacac gcatatttga tgagtttcat 60tcatatcaag
aagaaggtaa ataaaagttt atgtaattta gagatctaaa caaagtagaa 120tatctgataa
agcaatatat atgtattctt atgaataatt tcta
1645942DNAArtificial SequenceSpacer sequence 59ctcgagagat ctccgcggaa
cattattata acgttgctcg aa 4260213DNAHomo
sapiensmisc_featureNucleotide sequence of the human dystrophin gene
DMD, exon 23 60gctttacaaa gttctctgca agagcaacaa agtggcctat actatctcag
caccactgtg 60aaagagatgt cgaagaaagc gccctctgaa attagccgga aatatcaatc
agaatttgaa 120gaaattgagg gacgctggaa gaagctctcc tcccagctgg ttgagcattg
tcaaaagcta 180gaggagcaaa tgaataaact ccgaaaaatt cag
213613453DNAHomo sapiensmisc_featureNucleotide sequence of
the human dystrophin gene DMD, intron 22 61gtctgtgaat atttgaatgt
caaaacaata aagcacgctt atcaagcatt cacattgata 60taacctttaa ataatattag
aatttaagtc attctgacaa gtatggtagt ttgcccattg 120agcaaatgaa aatgagactt
tacagtatta tttatgcata tactctatct atggatatat 180atgtatgggc atatatacat
gctgtttttg aaaacaattg tatgatctat ttatgaagta 240tatttttaat ttagcatgca
ttttcagatg tattgatttt gattattaga aaaaccattg 300ttttattccc cacctttgct
tctttatttg taactcctgt cacattctca aactggtggg 360taaaatcagt gtacagtcat
cccttcgtat ccatggggaa gtggttccag gaactcctat 420gaatatcaga attcacagat
gttcaagtcc tttatatgaa tggtgtaata tttgcatata 480acttatgaac atgctcccat
atactttaac tcatctctgg attacttata atgcctaata 540caatgtaaat attatgaaaa
tatgtgttat actgtattat ttaggaagta gtgacaatga 600aaaagtctta catgtttagt
acagatgtag ttatttttgt caaatattat ctattagtag 660ttgactgaat ccactcattc
agaatccaca aatatggagg gctggctata tttataaaat 720tggtttaaat ttaaacatta
attttctgta atgtaatgag acatttgaat tctaatagct 780acattttttt aacatccttc
tatcaggaaa gatatactaa tatgggaaat gaaagaaata 840atttgtaaga gattttataa
tacttttcct actttatttt atttttagac ggagtttcat 900tcttgttgcc caggctggag
tgcagcagtg ctatctcggc tcactgcaac ctacgcctcc 960cgggttcaag tgattctcct
gccttagcct cctgagtagc tgggattata gtgcgtgcca 1020tcacgcctgg ctaatttttg
tatttttaaa gtaaagatgg ggtttcacca tcttggccat 1080gctggtctca aactcctgac
ctcagatgat ctgcctgcct cagcctccca aagtgctggg 1140attacaggtg tgagccactg
cgcccagccc ttttcctact tattataaaa tactacatta 1200tatgttcaat aactttttat
ttctgaagac atgaaaacgt attagttata tttagaagct 1260gagataatat gtgcatttta
agaaaaatgt ttaaaaattc aaatttaaac tttatcacat 1320agaatttttt ttggataatt
aaatattttt ttgaagtaat gtattcaaag aaaaatacat 1380ttaaacaact ttcattatgt
attgtcctat ttagatattt attctactac aataccaaat 1440ggtagctgtt tcaccttttt
tgtttccttt tgttgtggtt tttaatgctt ggcataatga 1500ttccaaagtc tatctggaaa
aatgtgttgc ctaatagaac tagtaaactt ttaaacataa 1560gaataaaaat gtgcaagaat
aagagtggag aaaacacaag aaaaggcagt tagatgagca 1620gaagagacta tttccactga
aaaagatagg aaagaaaata attgttcata aatattattg 1680ggaaatgtat taactaacta
aaaaaaaaga tatgaacatt tcccccaaag ttacaccaaa 1740ataaaaacca catacaataa
aatgttcaaa aaatgtgggg aagcccaaat tttggaaaat 1800atttaaaata ataatgttat
aaaaatttag gtggagaagc ataaaaatcg atggaaacaa 1860tcacaaacat tgtaaaaaca
ccagagacaa ttttaaagaa atatgatacc aaggaatctt 1920tttttttctg tgagtaagga
agaaagtctt gagtaaggta tcctccaaaa agagacatat 1980atgaataata aacctatgaa
taattgtaac ttaaattgta aaggaagaag taaaaaaatt 2040acaagatgag aatttttttt
gcatactaaa ttgcatacac acaaaagata taagtaccaa 2100tttgcaagtg gtgaaataaa
aattgtctct tactgctgga aatgtaaatt caatatgaat 2160tttactgagg ttaactcttc
agtgcatcct aataacatac aacattaata taatttccat 2220gaaatctgta tctgatggaa
ataatcagaa tacactcaaa gatgtaaatg gaactggaga 2280tagataggat attgattgat
tgagggatgg atggatggat tagataggta ggtagataga 2340gataaacata atttacagga
gtggaaaact agaaatgaat gtgtagaact aataatggaa 2400attgttgaac tagaaaagcc
attacaattt ttgtttttga aaaaataatt atttgacatg 2460ggcaaatgct cacactaaaa
cacctagctc aaagtgtagt tatatgctag atattgtcta 2520aataatcagt cataggggaa
caatttgaaa aagtgaaata aacttaaaaa agtaaagtaa 2580aaaaacaaat taagtaccac
caaataggca taggatacta tgaaactata aaacttagat 2640atttgacaaa ttttgggttg
atataggtca cattataggg gaaagaataa ctgtttatat 2700gtgcatacat gatcatatat
ttaatattta gctatacata gaacaaattg aaaatttata 2760taaagtacta gtagttctgg
tttttttttc catttttgtc atgagccttt attattttat 2820aaagtaaaag taacattaaa
gtaaatattg aacaggggaa gtctaaagtt atgatcctac 2880tgtaagaaat tacatgtgtt
atacagaata agtataaata aattttataa attctccaca 2940aaactatcct tgaatcccac
cataatacta cataacagaa taacattttt tgaaaatata 3000ttaagtacat agattgttca
ataagttttt ctattccata tatcaatatt attagctata 3060gttttatgta gagtttactt
tcctgaaagc tcagtataat taatttcaac tccattaact 3120gtatcacaaa tgcagttatt
aatatatata cttaattcag taaaattaga tgcaccaaaa 3180ctttttggct ctgtaagata
tagttttgag atatatttga cattgttcag aaaaatacat 3240atggagtgtt aaataccact
aaataatatt caagttactc ttaggataat atgtaaaatt 3300taaattacca ctagaagttt
ataactgata gaagatcatc tactttgttt acatgtttga 3360atcatataga tttcaagtac
agttaatttc actaaaactc atcaattatt attcatcaat 3420tagggtaaat gtatttaaaa
aattgttttt tag 3453623798DNAHomo
sapiensmisc_featureNucleotide sequence of the human dystrophin gene
DMD, intron 23 62gtaattcaag attttacttt ctaccctcat ttttatttac ttgttttttc
cctaacgata 60cactgtaaac tgtaaaggta cataagcatt tgaccttcag catctttcaa
agttagtgag 120aagatgaaaa gacatgagtc ttttacaaag gatggcgatt tgcttattcc
ttctaggaaa 180agaataggag ttttctcagt gtttttagaa tgaattttct attttcacca
ttaccaatag 240gaattaaaca tgtttttagt ttaataattt tcaaaaccaa tcatccattt
agtctggttg 300tcttttaaga tacgggtaat catttagctt atgactagaa aaaagttatg
tatatataca 360ttatacagag gtatattgta atcattttat atatataatg tatagtatgt
atatgattat 420ttcctatatt acctttcatt tctgctgttt tgctacttat aaaaagtagg
agtcaactat 480gctgtctata tgtgctatta tatataaaat atggtgatat gttcataaaa
ttgcattata 540tattcatccc attatgtata actccatata tgtatatgtg cagtagaaat
gtaggtgtat 600taatgtccta tgatactcct aagaaattag cagaaattta atggcttcaa
acaacacaaa 660tttattacat tactgttctg gaggtcagaa gttcagaatt aatctctgtg
ggctaaaatt 720aaggtgttag cagaactgtg tctgttttgg agactttgaa agagaatgtg
tgttattggc 780atttttagct tccatatata tatgtatata tatacacaca catacatata
acatgtatat 840atgtgtgtat atatatatat atatacgtgt ataatggaat tatatatcat
gtacatatat 900acatatctaa tgtaatatat atcacatatg taacctgttg tatatgtata
ttaatgttac 960atatgtaata tatagtaata acttatttta tcaccatatc gaatttgtaa
ttatatgctt 1020aaattttacc acatcacctt cttcctgaaa ccaaattccc ttgacttaat
ttgtttgaat 1080agtgccacca tactaataac cactcaagct aaagaaaaac agtaaaccaa
caattgcatt 1140aaggtctaaa agcctaaaat attcaataag gtaagtaatg tgtcccttga
gtggtcaatt 1200attgtgacaa atgttaatgc tgggaaggaa atgtcgaaaa cgttatggta
tgatggaaaa 1260acattgcaac aatatgctcc tggatttaaa tcccagtcta gctgcttagt
ttttgtgctt 1320accttttaca aaccaaagca ctaagcaaac ctaaaacatt ttttaatgca
tagcatttct 1380ttacttataa acatatgtgt acatgcatac ctatgtaaat actgcatgaa
aatatctatt 1440atttgtcaag accagtggcc taagagggca caattacatg gctcttgtta
tcaaggagcc 1500cacattctag tgaaggagga aggtaggtgt attcatttcc tatggcactc
ctaacaaatt 1560gccggaaatt taataacttc aagcaacaca aatttattag agttctggaa
gtcaaaagtt 1620caggattagt ctcagtgggc taaaattaag gtgtcagcag gactttgtct
gttttggagg 1680ctttgcgaca gagtctgttt tattggcatt tttagcttct agaagcattc
cttgtctcat 1740gtccccatcc tccatcttca aagtcagcag tgtaacatca tctagtgtca
atttctctct 1800ccctgtctat ctccctctcc ctctctttct acctctcctt ctcaccttcc
tttccctatc 1860ctgttccctg tgttgacttc cttttaatta ggacctttgt gattacatta
tgccctggtg 1920gatgatccag tataatctcc ccatctcaga atccttaatg taatcacatc
tgcaaagtcc 1980tttctggtaa tatactcaca ggtttcagag attagggtga ggacatcttg
ggggtagcat 2040tattcagcct tgcatggtaa ataaattaaa cagtatggga aaggttgtca
cagtaggcag 2100gaaggaaatc caacataaac taggaaaaag gattatgaaa tattatcgga
acataggttg 2160tctctgctat agattaaata ggtagaattt agttagctaa taccgagaaa
gaaagagttc 2220ttgagaaagc atagcagcat tttcagttaa atgatataag ttcaacactg
ggtagacaat 2280atatatgggg attaataaaa tattatggtt ggaaaactgg attgagagta
gacagtgaat 2340gtccttggat gctatgtttt acaatctgag ttttaatcta gtggtgatgg
gactgataat 2400aataacaata aaaacttaat aagaaaaaac tgctattggt taaagaggga
aataatatga 2460ttccacttgt gttttcaaaa cattcccaag atgagaggaa ggaggagagg
agattactgg 2520catcactgta gatcagtaat tagaatgagg tgagaccgag gcagatatca
aaacgagcca 2580gtgcattaat ggagtcacta atgagacctg aacagcccag gcgtagtggc
tcatgcctat 2640aatcccagca ctctgggcag ctgaggcagg tagatcacga ggtcaagaga
tcaagaccat 2700gctggccaac atggcgaaac cctgtctcta ctaaaaacac aaaaattagc
tgggcatggt 2760agtgcacgcc tgtaatccca gctacttggg aggctgaggc aggagaatgg
cgtgaacctg 2820ggaggtggag gctgcagtga gccaagattg tgccacttca ctccagcctg
ggcgacagag 2880caagactcca tctcaaaaaa aaaaaaaaaa aaaaacaaaa aacaaacaaa
aaaaaaaaga 2940cctgaacaag gacagtgaag atgtgaaata agaacaaatg tagacttgaa
agataacagc 3000aaggtgatcg tcttgaagtt ctcttagata aatagtttaa aaattattat
taataatcac 3060attaaagggg aaactttctg tggagcacat attttcaatg taaatttaag
taaaacagaa 3120gaaaataaca catgtagaca gcatagttga atcatacttt ttagaagtgg
caaaacagca 3180gaaaaaaagg taatacggga aagaatcatg ggtgagagat actggctatt
tataaatgat 3240aaaagatgat gatttttacc tgatcattac ttcgtactgg tcagacaaat
aaaagcaaaa 3300gctctgtctt tgaagatgac aaaatattag tccaaaagtt cccaactgga
aggactattt 3360aaactgtcat ctgtagaaaa taatttgtga aagttcgggt ttagggaggc
tataaagaca 3420ccattacatt gagtttattg ttcatagttt gttttatgta ctgtaaggac
acatttttag 3480tattctcatg agttgttttg taacttaaaa tttctctaga gggggatatg
atttaatgtt 3540ctcgagagta acatcataaa accacatttg gtagtaattt tgtattttta
acaatagcag 3600acttcacaca ccagtgctca tacagtagac cataaaaatg cagtcttagt
aaaaatattc 3660tttgcctcaa gaactactta gagacatcct ttaaacatgg gaattgtttt
tgggcctgtg 3720tttagacata acacaatgat gaattgtgtt aaaagtaatc agcacaccag
taatgcctta 3780taacgggtct cgtttcag
37986371PRTHomo sapiensmisc_featureHuman sequence of the
dystrophin gene DMD, exon 23 63Ala Leu Gln Ser Ser Leu Gln Glu Gln
Gln Ser Gly Leu Tyr Tyr Leu1 5 10
15Ser Thr Thr Val Lys Glu Met Ser Lys Lys Ala Pro Ser Glu Ile
Ser 20 25 30Arg Lys Tyr Gln
Ser Glu Phe Glu Glu Ile Glu Gly Arg Trp Lys Lys 35
40 45Leu Ser Ser Gln Leu Val Glu His Cys Gln Lys Leu
Glu Glu Gln Met 50 55 60Asn Lys Leu
Arg Lys Ile Gln65 7064114DNAHomo
sapiensmisc_featureHuman nucleotide sequence of the dystrophin gene
DMD, exon 24 64aatcacatac aaaccctgaa gaaatggatg gctgaagttg atgtttttct
gaaggaggaa 60tggcctgccc ttggggattc agaaattcta aaaaagcagc tgaaacagtg
caga 1146538PRTHomo sapiensmisc_featureHuman sequence of the
dystrophin gene DMD, exon 24 65Asn His Ile Gln Thr Leu Lys Lys Trp
Met Ala Glu Val Asp Val Phe1 5 10
15Leu Lys Glu Glu Trp Pro Ala Leu Gly Asp Ser Glu Ile Leu Lys
Lys 20 25 30Gln Leu Lys Gln
Cys Arg 3566608DNAMus musculusmisc_featureMurine sequence of the
dystrophin gene DMD, intron 58 66gtaattgaat gtggaactgt aataacatat
tgatagaggg ctcagtgatg agagcacagc 60ccatccatgc ttgctgccag agtctgtata
gctctcacac ttcagggtta aacagaatca 120attcaaatta aatgtaggtg ctgaaaaaga
atacagaatc acaaaccacc atgcacaatt 180ctgttttcag tcataaataa agcaatcaat
gatcagttat ttgaatattt agaaaatggt 240aagcaaaaga tcttatctgt gataagaagt
tctattgtta caaagactgc aagctaattt 300tccgtttaag ctgactgaaa taaattggag
ctgttcagca catgctaaca tttttctctc 360tgaaagctta aatgaatctt gatttacctg
attgacagaa agtagaacag attcactaag 420gaagacagtt tagctgtgct tgactatatc
aaaatttatg ccaaagtgta aaagagccat 480taatcagtaa gttggcctcg tgttaatcta
tattcctttc ctttcttcta gtctgacctt 540ttcaaccatg tttttttttt ttaacaaaaa
tggatgtgac cttaaacctt gtcatattgc 600caatttag
60867608DNAHomo
sapiensmisc_featureHuman sequence of the dystrophin gene DMD, intron
58 67gtaattgaat gtggaactat aataacatat tgatagaagg atcagtggtg acggagcagc
60ccatccattc ttgctgccag ggtctggata gctctcatat tttcttggtt aaatagaatc
120aattcaaatt aaacatagat gctgaaaaaa aaataaggac tcataaacta ccatgcataa
180atcagtttgt gcattcataa ataaacatca aaagagatta gcaatcagtt attggaacat
240ttagaaaata ataaacaaag ggagttatct gtgaggagga attctattgt tgcaaagact
300acaagttaat tttccattta agagcctgcc tgaagtaaat tagagcttgt ggtatgcatc
360ctaacgtttt tctcctcaat agcttaagtg aatcttaaat tgtcggattg atataaggta
420gaaactcagg aagatatact ttagatgttc tgggctgtat caaaatttat gccaaagtat
480aaaaaagccg ttaatcagta ggttaccctc ttgttcaact gtactctttc tttcttccag
540tatgaccttt ttgacaatgt ttaaaaaaaa agaatgtggc ctaaaacctt gtcatattgc
600caatttag
60868150DNAArtificial SequenceAntisens sequence to intron 58 of DMD
68aaaattaact tgtagtcttt gcaacaatag aattcctcct cacagataac tccctttgtt
60tattattttc taaatgttcc aataactgat tgctaatctc ttttgatgtt tatttatgaa
120tgcacaaact gatttatgca tggtagttta
150692390DNAMus musculusmisc_featureMurine sequence of the dystrophin
gene DMD, exons 59-79 69aactgcctcc tgaagaaaga gctcagaatg tcactcggct
cctacgaaag caggctgaag 60aggtcaacgc tgaatgggac aaattgaacc tgcgctcagc
tgattggcag agaaaaatag 120atgaagctct tgaaagactc caggaacttc aggaagctgc
cgatgaactg gacctcaagt 180tgcgccaagc tgaggtgatc aagggatcct ggcagccagt
gggggatctc ctcattgact 240ctctgcaaga tcaccttgaa aaagtcaagg cacttcgggg
agaaattgca cctcttaaag 300agaatgtcaa tcgtgtcaat gaccttgcac atcagctgac
cacactgggc attcagctct 360caccttataa cctcagcact ttggaagatc tgaataccag
atggaggctt ctacaggtgg 420ctgtggagga ccgtgtcaga cagctgcatg aagcccacag
ggactttggt cctgcatccc 480agcacttcct ttccacttca gttcagggtc cctgggagag
agccatctca ccaaacaaag 540tgccctacta tatcaaccac gagacccaaa ccacttgttg
ggaccacccc aaaatgacag 600agctctacca gtctttagct gacctgaata atgtcaggtt
ctccgcgtat aggactgcca 660tgaagctcag aaggctccag aaggcccttt gcttggatct
cttgagcctg tcagctgcat 720gtgatgccct ggaccagcac aacctcaagc aaaatgacca
gcccatggat atcctgcaga 780taattaactg tttgactaca atttatgatc gtctggagca
agagcacaac aatctggtca 840atgtccctct ctgtgtggat atgtgtctca actggcttct
caatgtttat gatacgggac 900gaacagggag gatccgtgtc ctgtctttta aaactggcat
catttctctg tgtaaagcac 960acttggaaga caagtacaga taccttttca agcaagtggc
aagttcaact ggcttttgtg 1020accagcgtag gctgggtctt cttctgcatg attctattca
aatcccaaga cagttgggtg 1080aagttgcttc ctttgggggc agtaacattg agccgagtgt
caggagctgc ttccaatttg 1140ccaataataa acctgagatt gaagctgctc tcttccttga
ctggatgcgc ctggaacccc 1200agtctatggt gtggctgccc gtcttgcaca gagtggctgc
tgctgaaact gccaagcatc 1260aagccaagtg taacatctgt aaggagtgtc caatcattgg
attcaggtac agaagcctaa 1320agcattttaa ttatgacatc tgccaaagtt gctttttttc
tggccgagtt gcaaagggcc 1380ataaaatgca ctaccccatg gtagagtatt gcactccgac
tacatccgga gaagatgttc 1440gcgacttcgc caaggtacta aaaaacaaat ttcgaaccaa
aaggtatttt gcgaagcatc 1500cccgaatggg ctacctgcca gtgcagactg tgttagaggg
ggacaacatg gaaactcccg 1560ttactctgat caacttctgg ccagtagatt ctgcgcctgc
ctcgtccccc cagctttcac 1620acgatgatac tcattcacgc attgaacatt atgctagcag
gctagcagaa atggaaaaca 1680gcaatggatc ttatctaaat gatagcatct ctcctaatga
gagcatagat gatgaacatt 1740tgttaatcca gcattactgc caaagtttga accaggactc
ccccctgagc cagcctcgta 1800gtcctgccca gatcttgatt tccttagaga gtgaggaaag
aggggagcta gagagaatcc 1860tagcagatct tgaggaagaa aacaggaatc tgcaagcaga
atatgatcgc ctgaagcagc 1920agcatgagca taaaggcctg tctccactgc catctcctcc
tgagatgatg cccacctctc 1980ctcagagtcc cagggatgct gagctcattg ctgaggctaa
gctactgcgc caacacaaag 2040gacgcctgga agccaggatg caaatcctgg aagaccacaa
taaacagctg gagtctcagt 2100tacatagact gagacagctc ctggagcagc cccaggctga
agctaaggtg aatggcacca 2160cggtgtcctc tccttccacc tctctgcaga ggtcagatag
cagtcagcct atgctgctcc 2220gagtggttgg cagtcaaact tcagaatcta tgggtgagga
agatcttctg agtcctcccc 2280aggacacaag cacagggtta gaagaagtga tggagcaact
caacaactcc ttccctagtt 2340caagaggaag aaatgccccc ggaaagccaa tgagagagga
cacaatgtag 2390702390DNAHomo sapiensmisc_featureHuman
sequence of the dystrophin gene DMD, exons 59-79 70agctgcctcc
tgaggagaga gcccagaatg tcactcggct tctacgaaag caggctgagg 60aggtcaatac
tgagtgggaa aaattgaacc tgcactccgc tgactggcag agaaaaatag 120atgagaccct
tgaaagactc caggaacttc aagaggccac ggatgagctg gacctcaagc 180tgcgccaagc
tgaggtgatc aagggatcct ggcagcccgt gggcgatctc ctcattgact 240ctctccaaga
tcacctcgag aaagtcaagg cacttcgagg agaaattgcg cctctgaaag 300agaacgtgag
ccacgtcaat gaccttgctc gccagcttac cactttgggc attcagctct 360caccgtataa
cctcagcact ctggaagacc tgaacaccag atggaagctt ctgcaggtgg 420ccgtcgagga
ccgagtcagg cagctgcatg aagcccacag ggactttggt ccagcatctc 480agcactttct
ttccacgtct gtccagggtc cctgggagag agccatctcg ccaaacaaag 540tgccctacta
tatcaaccac gagactcaaa caacttgctg ggaccatccc aaaatgacag 600agctctacca
gtctttagct gacctgaata atgtcagatt ctcagcttat aggactgcca 660tgaaactccg
aagactgcag aaggcccttt gcttggatct cttgagcctg tcagctgcat 720gtgatgcctt
ggaccagcac aacctcaagc aaaatgacca gcccatggat atcctgcaga 780ttattaattg
tttgaccact atttatgacc gcctggagca agagcacaac aatttggtca 840acgtccctct
ctgcgtggat atgtgtctga actggctgct gaatgtttat gatacgggac 900gaacagggag
gatccgtgtc ctgtctttta aaactggcat catttccctg tgtaaagcac 960atttggaaga
caagtacaga taccttttca agcaagtggc aagttcaaca ggattttgtg 1020accagcgcag
gctgggcctc cttctgcatg attctatcca aattccaaga cagttgggtg 1080aagttgcatc
ctttgggggc agtaacattg agccaagtgt ccggagctgc ttccaatttg 1140ctaataataa
gccagagatc gaagcggccc tcttcctaga ctggatgaga ctggaacccc 1200agtccatggt
gtggctgccc gtcctgcaca gagtggctgc tgcagaaact gccaagcatc 1260aggccaaatg
taacatctgc aaagagtgtc caatcattgg attcaggtac aggagtctaa 1320agcactttaa
ttatgacatc tgccaaagct gctttttttc tggtcgagtt gcaaaaggcc 1380ataaaatgca
ctatcccatg gtggaatatt gcactccgac tacatcagga gaagatgttc 1440gagactttgc
caaggtacta aaaaacaaat ttcgaaccaa aaggtatttt gcgaagcatc 1500cccgaatggg
ctacctgcca gtgcagactg tcttagaggg ggacaacatg gaaactcccg 1560ttactctgat
caacttctgg ccagtagatt ctgcgcctgc ctcgtcccct cagctttcac 1620acgatgatac
tcattcacgc attgaacatt atgctagcag gctagcagaa atggaaaaca 1680gcaatggatc
ttatctaaat gatagcatct ctcctaatga gagcatagat gatgaacatt 1740tgttaatcca
gcattactgc caaagtttga accaggactc ccccctgagc cagcctcgta 1800gtcctgccca
gatcttgatt tccttagaga gtgaggaaag aggggagcta gagagaatcc 1860tagcagatct
tgaggaagaa aacaggaatc tgcaagcaga atatgaccgt ctaaagcagc 1920agcacgaaca
taaaggcctg tccccactgc cgtcccctcc tgaaatgatg cccacctctc 1980cccagagtcc
ccgggatgct gagctcattg ctgaggccaa gctactgcgt caacacaaag 2040gccgcctgga
agccaggatg caaatcctgg aagaccacaa taaacagctg gagtcacagt 2100tacacaggct
aaggcagctg ctggagcaac cccaggcaga ggccaaagtg aatggcacaa 2160cggtgtcctc
tccttctacc tctctacaga ggtccgacag cagtcagcct atgctgctcc 2220gagtggttgg
cagtcaaact tcggactcca tgggtgagga agatcttctc agtcctcccc 2280aggacacaag
cacagggtta gaggaggtga tggagcaact caacaactcc ttccctagtt 2340caagaggaag
aaatacccct ggaaagccaa tgagagagga cacaatgtag
2390712617DNAArtificial SequenceSimple PTM 71aaaattaact tgtagtcttt
gcaacaatag aattcctcct cacagataac tccctttgtt 60tattattttc taaatgttcc
aataactgat tgctaatctc ttttgatgtt tatttatgaa 120tgcacaaact gatttatgca
tggtagttta ggtaccccgc ggaacattat tataacgttg 180ctcgaatact aactgatatc
tcttcttttt ttttttccgg aaaacagagc tgcctcctga 240ggagagagcc cagaatgtca
ctcggcttct acgaaagcag gctgaggagg tcaatactga 300gtgggaaaaa ttgaacctgc
actccgctga ctggcagaga aaaatagatg agacccttga 360aagactccag gaacttcaag
aggccacgga tgagctggac ctcaagctgc gccaagctga 420ggtgatcaag ggatcctggc
agcccgtggg cgatctcctc attgactctc tccaagatca 480cctcgagaaa gtcaaggcac
ttcgaggaga aattgcgcct ctgaaagaga acgtgagcca 540cgtcaatgac cttgctcgcc
agcttaccac tttgggcatt cagctctcac cgtataacct 600cagcactctg gaagacctga
acaccagatg gaagcttctg caggtggccg tcgaggaccg 660agtcaggcag ctgcatgaag
cccacaggga ctttggtcca gcatctcagc actttctttc 720cacgtctgtc cagggtccct
gggagagagc catctcgcca aacaaagtgc cctactatat 780caaccacgag actcaaacaa
cttgctggga ccatcccaaa atgacagagc tctaccagtc 840tttagctgac ctgaataatg
tcagattctc agcttatagg actgccatga aactccgaag 900actgcagaag gccctttgct
tggatctctt gagcctgtca gctgcatgtg atgccttgga 960ccagcacaac ctcaagcaaa
atgaccagcc catggatatc ctgcagatta ttaattgttt 1020gaccactatt tatgaccgcc
tggagcaaga gcacaacaat ttggtcaacg tccctctctg 1080cgtggatatg tgtctgaact
ggctgctgaa tgtttatgat acgggacgaa cagggaggat 1140ccgtgtcctg tcttttaaaa
ctggcatcat ttccctgtgt aaagcacatt tggaagacaa 1200gtacagatac cttttcaagc
aagtggcaag ttcaacagga ttttgtgacc agcgcaggct 1260gggcctcctt ctgcatgatt
ctatccaaat tccaagacag ttgggtgaag ttgcatcctt 1320tgggggcagt aacattgagc
caagtgtccg gagctgcttc caatttgcta ataataagcc 1380agagatcgaa gcggccctct
tcctagactg gatgagactg gaaccccagt ccatggtgtg 1440gctgcccgtc ctgcacagag
tggctgctgc agaaactgcc aagcatcagg ccaaatgtaa 1500catctgcaaa gagtgtccaa
tcattggatt caggtacagg agtctaaagc actttaatta 1560tgacatctgc caaagctgct
ttttttctgg tcgagttgca aaaggccata aaatgcacta 1620tcccatggtg gaatattgca
ctccgactac atcaggagaa gatgttcgag actttgccaa 1680ggtactaaaa aacaaatttc
gaaccaaaag gtattttgcg aagcatcccc gaatgggcta 1740cctgccagtg cagactgtct
tagaggggga caacatggaa actcccgtta ctctgatcaa 1800cttctggcca gtagattctg
cgcctgcctc gtcccctcag ctttcacacg atgatactca 1860ttcacgcatt gaacattatg
ctagcaggct agcagaaatg gaaaacagca atggatctta 1920tctaaatgat agcatctctc
ctaatgagag catagatgat gaacatttgt taatccagca 1980ttactgccaa agtttgaacc
aggactcccc cctgagccag cctcgtagtc ctgcccagat 2040cttgatttcc ttagagagtg
aggaaagagg ggagctagag agaatcctag cagatcttga 2100ggaagaaaac aggaatctgc
aagcagaata tgaccgtcta aagcagcagc acgaacataa 2160aggcctgtcc ccactgccgt
cccctcctga aatgatgccc acctctcccc agagtccccg 2220ggatgctgag ctcattgctg
aggccaagct actgcgtcaa cacaaaggcc gcctggaagc 2280caggatgcaa atcctggaag
accacaataa acagctggag tcacagttac acaggctaag 2340gcagctgctg gagcaacccc
aggcagaggc caaagtgaat ggcacaacgg tgtcctctcc 2400ttctacctct ctacagaggt
ccgacagcag tcagcctatg ctgctccgag tggttggcag 2460tcaaacttcg gactccatgg
gtgaggaaga tcttctcagt cctccccagg acacaagcac 2520agggttagag gaggtgatgg
agcaactcaa caactccttc cctagttcaa gaggaagaaa 2580tacccctgga aagccaatga
gagaggacac aatgtag 261772137DNAHomo
sapiensmisc_featureHuman sequence of the dystrophin gene DMD, exon
70 72actacatcag gagaagatgt tcgagacttt gccaaggtac taaaaaacaa atttcgaacc
60aaaaggtatt ttgcgaagca tccccgaatg ggctacctgc cagtgcagac tgtcttagag
120ggggacaaca tggaaac
13773137DNAMus musculusmisc_featureMurine sequence of the dystrophin gene
DMD, exon 70 73actacatccg gagaagatgt tcgcgacttc gccaaggtac
taaaaaacaa atttcgaacc 60aaaaggtatt ttgcgaagca tccccgaatg ggctacctgc
cagtgcagac tgtgttagag 120ggggacaaca tggaaac
137
User Contributions:
Comment about this patent or add new information about this topic: