Patent application title: METHOD FOR DNA BREAKPOINT ANALYSIS
Inventors:
Alexander Alan Morley (Glenegl, AU)
IPC8 Class: AC12Q168FI
USPC Class:
435 6
Class name: Chemistry: molecular biology and microbiology measuring or testing process involving enzymes or micro-organisms; composition or test strip therefore; processes of forming such composition or test strip involving nucleic acid
Publication date: 2010-08-26
Patent application number: 20100216138
Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
Patent application title: METHOD FOR DNA BREAKPOINT ANALYSIS
Inventors:
Alexander Alan Morley
Agents:
KNOBBE MARTENS OLSON & BEAR LLP
Assignees:
Origin: IRVINE, CA US
IPC8 Class: AC12Q168FI
USPC Class:
Publication date: 08/26/2010
Patent application number: 20100216138
Abstract:
The present invention relates to a method for identifying a DNA breakpoint
and agents for use therein. More particularly, the present invention
provides a method for identifying a gene translocation breakpoint based
on the application of a novel multiplex DNA amplification technique. The
method of the present invention facilitates not only the identification
of the break-point position but, further, enables the isolation of the
DNA segment across which the breakpoint occurs. This provides a valuable
opportunity to conduct further analysis of the breakpoint region, such as
to sequence across this region. The method of the present invention is
useful in a range of applications including, but not limited to,
providing a routine means to characterise the gene break-point associated
with disease onset in a patient and thereby enable the design of patient
specific probes and primers for ongoing monitoring of the subject disease
condition. In addition to monitoring the progression of a condition
characterised by the existence of the breakpoint, there is also enabled
assessment of the effectiveness of existing therapeutic drugs and/or new
therapeutic drugs and, to the extent that the condition is a neoplasm,
prediction of the likelihood of a subject's relapse from a remissive
state.Claims:
1. A method of identifying a gene breakpoint, said method comprising:(i)
contacting a DNA sample with:(a) one or more forward primers directed to
a DNA region of the flanking gene or fragment thereof located 5' relative
to the gene breakpoint, which primers are optionally operably linked at
their 5' end to an oligonucleotide tag; and(b) one or more reverse
primers directed to a DNA region of the flanking gene or fragment thereof
located 3' relative to the gene breakpoint, which primers are optionally
operably linked at their 5' end to an oligonucleotide tag;wherein the
oligonucleotide tags of the forward primers are the same relative to the
forward primer tags of step (a) and the oligonucleotide tags of the
reverse primers are the same relative to the reverse primer tags of step
(a) but which forward primer oligonucleotide tags are different relative
to the reverse primer tags;(ii) amplifying the DNA sample of step
(i);(iii) optionally contacting the amplicon generated in step (ii)
with:(a) one or more forward primers directed to a DNA region of the
flanking gene or fragment thereof located 5' relative to the gene
breakpoint, which primers are directed to DNA regions which are located
3' to one or more of the forward primers of step (i) and which primers
are optionally operably linked at their 5' end to an oligonucleotide tag;
and(b) one or more reverse primers directed to a DNA region of the
flanking gene or fragment thereof located 3' relative to the gene
breakpoint, which primers are directed to DNA regions which are located
5' to one or more of the reverse primers of step (i) and which primers
are optionally operably linked at their 5' end to an oligonucleotide
tag,wherein the oligonucleotide tags of the forward primers are the same
relative to the forward primer tags of step (iii)(a) and the
oligonucleotide tags of the reverse primers are the same relative to the
other reverse primer tags of step (iii)(a) but which forward primer
oligonucleotide tags are different relative to the reverse primer tags
and which forward and reverse primer tags of step (iii) are different
relative to the forward and reverse primer tags of step (i); and(iv)
amplifying the DNA sample of step (iii);(v) analysing said amplified DNA.
2. (canceled)
3. (canceled)
4. (canceled)
5. (canceled)
6. (canceled)
7. (canceled)
8. (canceled)
9. (canceled)
10. (canceled)
11. (canceled)
12. (canceled)
13. (canceled)
14. (canceled)
15. (canceled)
16. (canceled)
17. (canceled)
18. (canceled)
19. (canceled)
20. (canceled)
21. (canceled)
22. (canceled)
23. (canceled)
24. (canceled)
25. (canceled)
26. (canceled)
27. (canceled)
28. A method of identifying a gene translocation breakpoint, said method comprising:(i) contacting a DNA sample with:(a) one or more forward primers directed to a DNA region of the antisense strand of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag;(b) one or more reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag;wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags;(c) a primer directed to the forward primer oligonucleotide tag of step (i)(a); and(d) a primer directed to the reverse primer oligonucleotide tag of step (i)(b);(ii) amplifying the DNA sample of step (i);(iii) optionally contacting the amplicon generated in step (ii) with:(a) one or more forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are directed to DNA regions which are located 3' to one or more of the forward primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag;(b) one or more reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' to the gene breakpoint, which primers are directed to DNA regions which are located 5' to one or more of the reverse primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag;(c) a primer directed to the forward primer oligonucleotide tag of step (iii)(a); and(d) a primer directed to the reverse primer oligonucleotide tag of step (iii)(b);wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (iii)(a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (iii)(a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags and which forward and reverse primer tags of step (iii) are different relative to the forward and reverse primer tags of step (i);(iv) amplifying the DNA sample of step (iii); and(v) analysing said amplified DNA.
29. The method according to claim 1, wherein at least one set of said forward and reverse primers are linked to an oligonucleotide tag.
30. The method according to claim 1, wherein one primer is used in step (i)(a) and two or more primers are used in step (i)(b).
31. The method according to claim 1, wherein one primer is used in step (i)(b) and two or more primers are used in step (i)(a).
32. The method according to claim 1, wherein step (iii) is performed and one primer is used in step (iii)(a) and two or more primers are used in step (πi)(b).
33. The method according to claim 1, wherein step (iii) is performed and one primer is used in step (iii)(b) and two or more primers are used in step (iii)(a).
34. The method according to claim 1, wherein said gene breakpoint is a gene translocation breakpoint or a homologous recombination point.
35. The method according to claim 34, wherein said gene translocation breakpoint is a chromosomal gene translocation breakpoint.
36. The method according to claim 35, wherein said gene translocation breakpoint is selected from the group consisting of:(i) BCR-ABL translocation,(ii) PML-RARα translocation,(iii) t(2;5)(p23;q35) translocation,(iv) t(8; 14) translocation,(v) t(9;22)(q34;q11) translocation,(vi) t(11; 14) translocation,(vii) t(11;22)(q24;q11.2-12) translocation,(viii) t(14;18)(q32;q21) translocation,(ix) t(17;22) translocation,(x) t(15; 17) translocation,(xi) t(1; 12)(q21;p 13) translocation,(xii) t(9; 12)(p24;p 13) translocation,(xiii) t(X;18)(p1 1.2;q1 1.2) translocation,(xiv) t(1;1 I)(q42.1;q14.3) translocation, and(xv) t(1; 19) translocation.
37. The method according to claim 1, wherein 1-30 primers are used in step (i)(a) and 24-400 primers are used in step (i)(b).
38. The method according to claim 37, wherein step (iii) is performed and 1-30 primers are used in step (iii)(a) and 24-400 primers are used in step (iii)(b).
39. The method according to claim 1, wherein only steps (i), (ii) and (v) are performed.
40. The method according to claim 39, wherein steps (iii) and (iv) are additionally performed.
41. The method according to claim 39, wherein said amplified DNA of step (ii) is subjected to a further step of amplification, selection or enrichment.
42. The method according to claim 41, wherein said amplification is bottleneck PCR.
43. The method according to claim 40, wherein said amplified DNA of step (iv) is subjected to a further step of amplification, selection or enrichment.
44. The method according to claim 43, wherein said amplification is bottleneck PCR.
45. The method according to claim 1, wherein said gene breakpoint is a chromosomal BCR-ABL translocation and:(a) the forward primers of step (i)(a) correspond to SEQ ID NOs:10-15; and(b) the reverse primers of step (i)(b) correspond to SEQ ID NOs:23-304 and are linked to the oligonucleotide tag defined by SEQ ID NO:22and wherein if step (iii) is performed then:(a) the forward primers of step (iii)(a) correspond to SEQ ID NOs: 16-21; and(b) the reverse primers of step (iii)(b) corresponds to SEQ ID NOs:306-587 and are linked to the oligonucleotide tag defined by SEQ ID NO:305.
46. The method according to claim 1, wherein said gene breakpoint is a chromosomal PML-RARalpha translocation and:(a) the forward primers of step (i)(a) correspond to SEQ ID NOs:588-593; and(b) the reverse primers of step (i)(b) correspond to SEQ ID NOs:601-634 and are linked to the oligonucleotide tag defined by SEQ ID NO: 22and wherein step (ii) is followed by bottleneck PCR which is performed using primers corresponding to SEQ ID NOs: 594-599.
47. A method of monitoring a disease condition in a mammal, which disease condition is characterised by a gene breakpoint, said method comprising screening for the presence of said breakpoint in a biological sample derived from said mammal, which breakpoint has been identified in accordance with the method of claim 1.
48. The method according to claim 47, wherein said condition is:anaplastic large cell lymphoma,Burkitt's lymphoma,CML, ALL,Mantle cell lymphoma,Ewing's sarcoma,follicular lymphoma,dermatofibrosarcoma protuberans,acute promyelocy e leukemia,acute myelogenous leukemia,Synovial sarcoma,Schizophrenia; oracute pre-B cell leukemia.
49. A DNA primer set, which primer set is designed to amplify and/or otherwise detect a gene breakpoint, which breakpoint has been identified in accordance with the method of claim 1.
Description:
FIELD OF THE INVENTION
[0001]The present invention relates to a method for identifying a DNA breakpoint and agents for use therein. More particularly, the present invention provides a method for identifying a gene translocation breakpoint based on the application of a novel multiplex DNA amplification technique. The method of the present invention facilitates not only the identification of the breakpoint position but, further, enables the isolation of the DNA segment across which the breakpoint occurs. This provides a valuable opportunity to conduct further analysis of the breakpoint region, such as to sequence across this region. The method of the present invention is useful in a range of applications including, but not limited to, providing a routine means to characterise the gene breakpoint associated with disease onset in a patient and thereby enable the design of patient specific probes and primers for ongoing monitoring of the subject disease condition. In addition to monitoring the progression of a condition characterised by the existence of the breakpoint, there is also enabled assessment of the effectiveness of existing therapeutic drugs and/or new therapeutic drugs and, to the extent that the condition is a neoplasm, prediction of the likelihood of a subject's relapse from a remissive state.
BACKGROUND OF THE INVENTION
[0002]The reference in this specification to any prior publication (or information derived from it), or to any matter which is known, is not, and should not be taken as an acknowledgment or admission or any form of suggestion that that prior publication (or information derived from it) or known matter forms part of the common general knowledge in the field of endeavour to which this specification relates.
[0003]Bibliographic details of the publications referred to by author in this specification are collected alphabetically at the end of the description.
[0004]Chromosomal translocations bring the previously unlinked segments of the genome together by virtue of the exchange of parts between non-homologous chromosomes. Although some translocations are not associated with a new phenotype, others may result in disease due to the modulation of protein expression or the synthesis of a new fusion protein.
[0005]There are two main types of chromosomal translocations which occur, these being reciprocal translocations (also known as non-Robertsonian) and Robertsonian translocations. Further, translocations can be balanced (in an even exchange of material with no genetic information extra or missing) or unbalanced (where the exchange of chromosome material is unequal resulting in extra or missing genes).
[0006]Reciprocal (non-Robertsonian) translocations usually result in an exchange of material between non-homologous chromosomes and are found in about 1 in 600 newborns. Such translocations are usually harmless and may be found through prenatal diagnosis. However, carriers of balanced reciprocal translocations exhibit an increased risk of creating gametes with unbalanced chromosome translocations thereby leading to miscarriages or children with abnormalities.
[0007]Robertsonian translocations involve two acrocentric chromosomes that fuse near the centromere region with loss of the short arms. The resulting karyotype has only 45 chromosomes since two chromosomes have fused together. Robertsonian translocations have been observed involving all combinations of acrocentric chromosomes. The most common translocation involves chromosomes 13 and 14 and is seen in about 1 in 1300 persons. Like other translocations, carriers of Robertsonian translocations are phenotypically normal, but exhibit a risk of unbalanced gametes which lead to miscarriages or abnormal offspring. For example, carriers of Robertsonian translocations involving chromosome 21 exhibit a higher probability of having a child with Down syndrome.
Diseases which may result from the occurrence of a translocation include: [0008]Cancer--several forms of cancer are caused by translocations; this mainly having been described in leukemia (eg. acute myelogenous leukemia and chronic myelogenous leukemia). [0009](ii) Infertility--this can occur where one of the would-be parents carries a balanced translocation, where the parent is asymptomatic but conceived foetuses are not viable. [0010](iii) Down syndrome--in some cases this is caused by a Robertsonian translocation of about a third of chromosome 21 onto chromosome 14.Specific examples of chromosomal translocations and the disease with which they are associated include: [0011]t(2;5)(p23;q35)--anaplastic large cell lymphoma [0012]t(8;14) --Burkitt's lymphoma (c-myc) [0013]t(9;22)(q34;q11)--Philadelphia chromosome, CML, ALL [0014]t(11;14)--Mantle cell lymphoma (Bcl-1) [0015]t(11;22)(q24;q11.2-12)--Ewing's sarcoma [0016]t(14;18)(q32;q21)--follicular lymphoma (Bcl-2) [0017]t(17;22)--dermatofibrosarcoma protuberans [0018]t(15;17)--acute promyelocytic leukemia (pml and retinoic acid receptor genes) [0019]t(1;12)(q21;p13)--acute myelogenous leukemia [0020]t(9;12)(p24;p13)--CML, ALL (TEL-JAK2) [0021]t(X;18)(p11.2;q11.2)--Synovial sarcoma [0022]t(1;11)(q42.1;q14.3)--Schizophrenia [0023]t(1;19)--acute pre-B cell leukemia (PBX-1 and E2A genes).
[0024]The shorthand t(A;B)(p1;q2) is used to denote a translocation between chromosome A and chromosome B. The information in the second set of parentheses, when given, gives a precise location within the chromosome for chromosomes A and B respectively--with p indicating the short arm of the chromosome, q indicating the long arm, and the numbers of p and q refers to regions, bands and sub-bands seen when staining the chromosomes under microscope.
[0025]As detailed above, chronic myelogenous leukemia is an example of a neoplastic condition which is caused by a chromosomal translocation. However, unlike many neoplastic conditions, its treatment prospects are quite good if it can be effectively diagnosed and monitored.
[0026]In virtually all cases of chronic myelogenous leukemia, a specific translocation is seen. This translocation involves the reciprocal fusion of small pieces from the long arms of chromosome 9 and 22. The altered chromosome 22 is known as the Philadelphia chromosome (abbreviated as Ph1). When the breakpoint of the Ph1 chromosome was sequenced, it was found that the translocation creates a fusion gene by bringing together sequences from the c-ABL proto-oncogene and another BCR (breakpoint cluster region). The BCR-ABL fusion gene encodes a phosphoprotein (p210) that functions as a dysregulated protein tyrosine kinase and predisposes the cell to become neoplastic. This hypothesis is supported by finding that expression of p210 results in transformation of a variety of hematopoietic cell lines in vitro and that mice transgenic for the human BCR-ABL gene develop a number of hematologic malignancies. Another well studied example of a translocation generating cancer is seen in Burkitt's lymphoma. In some cases of this B cell tumor, a translocation is seen involving chromosome 8 and one of three other chromosomes (2, 14 or 22). In these cases, a fusion protein is not produced. Rather, the c-myc proto-oncogene on chromosome 8 is brought under transcriptional control of an immunoglobulin gene promoter. In B cells, immunoglobulin promoters are transcriptionally quite active, resulting in over expression of c-myc, which is known from several other systems to exhibit monogenic properties. Accordingly, this translocation results in aberrant high expression of an oncogenic protein.
[0027]The classical method of diagnosing chromosomal translocations, such as those observed in chronic myelogenic leukemia, is by karyotyping. For many translocations, however, it is now possible to detect the translocation by PCR, using primers which span the breakpoint. In some cases, the PCR technique can also be used for sensitive detection and monitoring of treatment. Monitoring to determine the effect of treatment has become increasingly important for diseases such as chronic myeloid leukemia and acute promyelocytic leukemia as increasingly effective treatment has been developed. For monitoring in these 2 diseases, the starting material for the PCR is RNA. The translocation breakpoint is within the introns of the respective genes and, as a consequence, RNA splicing removes the sequence of RNA transcribed by introns and results in only one or a very limited number of final mRNA products being produced, despite the very large number of different translocations which are present in the patient population.
[0028]However, the use of RNA as the starting material to detect and quantify the translocation by PCR suffers the disadvantage that RNA is a difficult molecule to work with due to its inherent susceptibility to degradation. DNA is a more stable molecule. However, the initial identification and characterisation of the breakpoint in the context of DNA is much more difficult since cluster regions of chromosomal fusion sites often span large introns of several tens of thousands of nucleotides. These sizes are too large for direct coverage by a single PCR reaction. There therefore exists an ongoing need to develop means for routinely conducting breakpoint analyses on DNA samples.
[0029]In work leading up to the present invention, a novel multiplex amplification reaction has been developed which enables the localisation and analysis of a breakpoint in a DNA sample. Despite the precise position of the breakpoint being unknown, the method of the present invention nevertheless enables diagnosis of the existence of the breakpoint in a DNA sample and the isolation and analysis of the breakpoint region using a relatively modest and simple multiplex amplification reaction. The design of this amplification reaction results in the advantage that generation of long PCR products is not required. Still further, the optional incorporation of a primer hybridisation tag region at the 5' end of the amplification primers enables the rapid generation of large copy numbers of the amplicons generated using these primers and therefore facilitates the isolation and analysis of the amplicons.
SUMMARY OF THE INVENTION
[0030]Throughout this specification and the claims which follow, unless the context requires otherwise, the word "comprise", and variations such as "comprises" and "comprising", will be understood to imply the inclusion of a stated integer or step or group of integers or steps but not the exclusion of any other integer or step or group of integers or steps.
[0031]As used herein, the term "derived from" shall be taken to indicate that a particular integer or group of integers has originated from the species specified, but has not necessarily been obtained directly from the specified source. Further, as used herein the singular forms of "a", "and" and "the" include plural referents unless the context clearly dictates otherwise.
[0032]Unless otherwise defined, all technical and scientific terms used herein have the same meaning as commonly understood by one of ordinary skill in the art to which this invention belongs.
[0033]The subject specification contains nucleotide sequence information prepared using the programme PatentIn Version 3.1, presented herein after the bibliography. Each nucleotide sequence is identified in the sequence listing by the numeric indicator <210> followed by the sequence identifier (eg. <210>1, <210>2, etc). The length, type of sequence (DNA, etc) and source organism for each sequence is indicated by information provided in the numeric indicator fields <211>, <212> and <213>, respectively. Nucleotide sequences referred to in the specification are identified by the indicator SEQ ID NO: followed by the sequence identifier (eg. SEQ ID NO:1, SEQ ID NO:2, etc.). The sequence identifier referred to in the specification correlates to the information provided in numeric indicator field <400> in the sequence listing, which is followed by the sequence identifier (eg. <400>1, <400>2, etc). That is SEQ ID NO:1 as detailed in the specification correlates to the sequence indicated as <400>1 in the sequence listing
One aspect the present invention is directed to a method of identifying a gene breakpoint, said method comprising:contacting a DNA sample with: [0034](a) one or more forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0035](b) one or more reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0036]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags;(ii) amplifying the DNA sample of step (i);(iii) optionally contacting the amplicon generated in step (ii) with: [0037](a) one or more forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are directed to DNA regions which are located 3' to one or more of the forward primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0038](b) one or more reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' relative to the gene breakpoint, which primers are directed to DNA regions which are located 5' to one or more of the reverse primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag [0039]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (iii)(a) and the oligonucleotide tags of the reverse primers are the same relative to the other reverse primer tags of step (iii)(a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags and which forward and reverse primer tags of step (iii) are different relative to the forward and reverse primer tags of step (i);The present invention therefore preferably provides a method of identifying a chromosomal gene translocation breakpoint, said method comprising:(i) contacting a genomic DNA sample with: [0040](a) one or more forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0041](b) one or more reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0042]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags;(ii) amplifying the DNA sample of step (i);(iii) optionally contacting the amplicon generated in step (ii) with: [0043](a) one or more forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' to the gene breakpoint, which primers are directed to DNA regions which are located 3' to one or more of the forward primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0044](b) one or more reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' to the gene breakpoint, which primers are directed to DNA regions which are located 5' to one or more of the reverse primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0045]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (iii)(a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (iii)(a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags and which forward and reverse primer tags of step (iii) are different relative to the forward and reverse primer tags of step (i);(iv) amplifying the DNA sample of step (iii);(v) analysing said amplified DNA.There is therefore preferably provided a method of identifying a gene breakpoint, said method comprising:(i) contacting a DNA sample with [0046](a) one to thirty forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0047](b) twenty-four to four hundred reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0048]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags;(ii) amplifying the DNA sample of step (i);(iii) optionally contacting the amplicon generated in step (ii) with: [0049](a) one to thirty forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are directed to DNA regions which are located 3' to one or more of the forward primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0050](b) twenty-four to four hundred reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' to the gene breakpoint, which primers are directed to DNA regions which are located 5' to one or more of the reverse primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0051]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (iii)(a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (iii)(a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags and which forward and reverse primer tags of step (iii) are different relative to the forward and reverse primer tags of step (i);(iv) amplifying the DNA sample of step (iii);(v) analysing said amplified DNA.The present invention therefore provides a method of identifying a gene translocation breakpoint, said method comprising:(i) contacting a DNA sample with: [0052](a) one to thirty forward primers directed to a DNA region of the antisense strand of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0053](b) twenty-four to four hundred reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0054]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags; [0055](c) a primer directed to the forward primer oligonucleotide tag of step (i)(a); and [0056](d) a primer directed to the reverse primer oligonucleotide tag of step (i)(b);(ii) amplifying the DNA sample of step (i);(iii) optionally contacting the amplicon generated in step (ii) with: [0057](a) one to thirty forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are directed to DNA regions which are located 3' to one or more of the forward primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0058](b) twenty-four to four hundred reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' to the gene breakpoint, which primers are directed to DNA regions which are located 5' to one or more of the reverse primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0059](c) a primer directed to the forward primer oligonucleotide tag of step (iii)(a); and [0060](d) a primer directed to the reverse primer oligonucleotide tag of step (iii)(b); [0061]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (iii)(a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (iii)(a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags and which forward and reverse primer tags of step (iii) are different relative to the forward and reverse primer tags of step (i);(iv) amplifying the DNA sample of step (iii);(v) analysing said amplified DNA.According to this preferred embodiment there is provided a method of identifying a chromosomal BCR-ABL translocation breakpoint, said method comprising:(i) contacting a DNA sample with: [0062](a) one or more forward primers directed to a DNA region of BCR or fragment thereof, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0063](b) one or more reverse primers directed to a DNA region of ABL or fragment thereof, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0064]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags;(ii) amplifying the DNA sample of step (i);(iii) optionally contacting the amplicon generated in step (ii) with: [0065](a) one or more forward primers directed to a DNA region of BCR or fragment thereof, which primers are directed to DNA regions which are located 3' to one or more of the forward primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0066](b) one or more reverse primers directed to ABL or fragment thereof, which primers are directed to DNA regions which are located 5' to one or more of the reverse primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0067]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (iii)(a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (iii)(a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags and which forward and reverse primer tags of step (iii) are different relative to the forward and reverse primer tags of step (i);(iv) amplifying the DNA sample of step (iii);(v) analysing said amplified DNA.The present invention therefore preferably provides a method of identifying a chromosomal BCR-ABL translocation breakpoint, said method comprising:(i) contacting a DNA sample with: [0068](a) one to thirty forward primers directed to a DNA region of BCR or fragment thereof, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0069](b) twenty-four to four hundred reverse primers directed to a DNA region of ABL or fragment thereof, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0070]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags; [0071](c) a primer directed to the forward primer oligonucleotide tag of step (i)(a); and [0072](d) a primer directed to the reverse primer oligonucleotide tag of step (i)(b);(ii) amplifying the DNA sample of step (i);(iii) contacting the amplicon generated in step (ii) with: [0073](a) one to thirty forward primers directed to a DNA region of BCR or fragment thereof, which primers are directed to DNA regions which are located 3' to one or more of the forward primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0074](b) twenty-four to four hundred reverse primers directed to a DNA region of ABL or fragment thereof, which primers are directed to DNA regions which are located 5' to one or more of the reverse primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0075](c) a primer directed to the forward primer oligonucleotide tag of step (iii)(a); and [0076](d) a primer directed to the reverse primer oligonucleotide tag of step (iii)(b); [0077]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (iii)(a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (iii)(a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags and which forward and reverse primer tags of step (iii) are different relative to the forward and reverse primer tags of step (i);(iv) amplifying the DNA sample of step (iii);(v) isolating and sequencing said amplified DNA.
BRIEF DESCRIPTION OF THE DRAWINGS
[0078]FIG. 1 is a schematic representation of the strategy for amplification of the breakpoint region in the first round PCR. The forward primers for gene A and the reverse primers for gene B are preferably used in pools rather than individually. Only primer pairs which closely straddle the breakpoint will produce efficient amplification. The tags and tag primers are not shown. The strategy for the second round PCR is the same although the forward and reverse primers are just internal to their corresponding primers in the first round. In the case of chronic myeloid leukemia, gene A is the BCR gene and gene B is the ABL gene. Primer binding sites are staggered so that the maximum amplicon size does not exceed 1 kilobase.
[0079]FIG. 2 is a schematic representation of a protocol for isolation of the BCR-ABL translocation breakpoint in chronic myeloid leukemia.
[0080]FIG. 3 is an image of the results of electrophoresis showing amplified material from study of one patient. NFA was the pool of 6 forward BCR primers and NFA13 and NFA14 were 2 pools each containing 12 reverse ABL primers. NFA 13/14 was a pool containing the 24 ABL primers belonging to pools 13 and 14.
[0081]FIG. 4 is a representation of the sequences of the breakpoints in 4 patients with chronic myeloid leukemia. The numbers on the left are the Genbank base numbers for the BCR and ABL genes.
[0082]FIG. 5 shows the site of the DNA breakpoints in the ABL and BCR genes in the 27 patients with breakpoints isolated and identified. Blue regions in the ABL gene represent the exons 1a, 1b and E2. Red regions in the BCR gene represent exons 13, 14 and 15.
Isolation of the BCR-ABL Breakpoint in Chronic Myeloid Leukemia (CML)
[0083]Samples from 29 CML patients have been studied using the invention. In 27 of these patients the breakpoint sequences have been isolated and detailed sequencing information obtained. For one patient it has not been possible to amplify the BCR/ABL breakpoint. For the remaining patient a suspected breakpoint has been amplified. Sequence information shows the BCR gene at the 5' end and ABL sequence at the 3' end, however this breakpoint has not been confirmed with primers made specifically for the suspected regions.
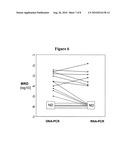
[0084]FIG. 6 is a comparison of DNA-based and RNA-based quantification of minimal residual disease (MRD) in samples of blood from 16 patients with CML. ND=not detected. Y-axis shows the number of leukemic cells as a proportion of total cells. The DNA-based PCR used patient-specific primers synthesised using knowledge of the breakpoint sequence in the patient being studied, the RNA-based PCR was the conventional approach using reverse transcription followed by PCR using generic primers. Black symbols show MRD detected by both techniques, red symbols show disease detected only by DNA-PCR and blue symbols show disease not detected. DNA-based PCR appears to be approximately 2 orders of magnitude more sensitive than RNA-based PCR.
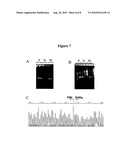
[0085]FIG. 7 is an illustration of the isolation of the PML-RARα breakpoint from a sample from the one patient with acute promyelocytic leukemia
DETAILED DESCRIPTION OF THE INVENTION
[0086]The present invention is predicated, in part, on the determination that gene translocation breakpoints can be routinely and easily identified, via DNA analysis, by sequentially performing two PCR reactions which use multiple primers directed to the genes flanking the breakpoint which are themselves tagged at their 5' end with a DNA region suitable for use as a primer hybridisation site. The simultaneous use of multiple primers facilitates the performance of a short PCR, rather than the long PCRs which have been performed to date. By sequentially performing a second PCR using primers directed to gene regions internal to those used in the first reaction, amplification of a DNA molecule spanning the breakpoint region can be achieved in a manner which enables the identification and isolation of a smaller amplification product than has been enabled to date in terms of the analysis of genomic DNA. By incorporating unique tag regions which can themselves be targeted by a primer, amplification of the initial amplicon can be rapidly achieved, thereby overcoming any disadvantage associated with the use of a low concentration of starting primer directed to the genes flanking the breakpoint. The method of the present invention therefore provides a simple yet accurate means of identifying and analysing a gene breakpoint using DNA. To this end, it would be appreciated that although the method of the present invention is exemplified by reference to chronic myelogenic leukemia, this method can be applied to any situation in which a gene breakpoint is sought to be identified via a DNA sample.
[0087]Accordingly, in one aspect the present invention is directed to a method of identifying a gene breakpoint, said method comprising:
(i) contacting a DNA sample with: [0088](a) one or more forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0089](b) one or more reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0090]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags;(ii) amplifying the DNA sample of step (i);(iii) optionally contacting the amplicon generated in step (ii) with: [0091](a) one or more forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are directed to DNA regions which are located 3' to one or more of the forward primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0092](b) one or more reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' relative to the gene breakpoint, which primers are directed to DNA regions which are located 5' to one or more of the reverse primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag [0093]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (iii)(a) and the oligonucleotide tags of the reverse primers are the same relative to the other reverse primer tags of step (iii)(a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags and which forward and reverse primer tags of step (iii) are different relative to the forward and reverse primer tags of step (i);(iv) amplifying the DNA sample of step (iii);(v) analysing said amplified DNA.
[0094]It should be understood that in a preferred embodiment of the present invention, where one primer is used in step (i)(a), it is preferable that two or more primers are used in step (i)(b). The converse applies where one primer is used in step (i)(b). Similarly, in another preferred embodiment, where one primer is used in step (iii)(a), it is preferable that two or more primers are used in step (iii)(b). The converse applies where one primer is used in step (iii)(b). Reference to the "flanking genes" 5' and 3' to the breakpoint should be understood as a reference to the genes or gene fragments on either side of the breakpoint. In terms of the 5' and 3' nomenclature which is utilised in the context of these genes/gene fragments, this should be understood as a reference to the 5'→3' orientation of the sense strand of double stranded DNA from which the DNA of interest derives. Accordingly, reference to "the flanking gene 5' to the breakpoint" should be understood as a reference to the sense strand of double stranded DNA. To this end, any reference to "gene" or "gene fragment" herein, to the extent that it is not specified, is a reference to the sense strand of double stranded DNA. Reference to the forward primer being directed to the antisense strand of the flanking gene 5' to the breakpoint therefore indicates that the forward primer bears the same DNA sequence as a region of the sense strand 5' to the breakpoint and therefore will bind to and amplify the antisense strand corresponding to that region.
[0095]Reference to "gene" should be understood as a reference to a DNA molecule which codes for a protein product, whether that be a full protein or a protein fragment. In terms of chromosomal DNA, the gene will include both intron and exon regions. However, to the extent that the DNA of interest is cDNA, such as might occur if the DNA of interest is vector DNA, there may not exist intron regions. Such DNA may nevertheless include 5' or 3' untranslated regions. Accordingly, reference to "gene" herein should be understood to encompass any form of DNA which codes for a protein or protein fragment including, for example, genomic DNA and cDNA.
[0096]Reference to a gene "breakpoint" should be understood as a reference to the point at which a fragment of one gene recombines with another gene or fragment thereof. That is, there has occurred a recombination of two genes such that either one or both genes have become linked at a point within one or both of the genes rather than the beginning or end of one gene being linked to the beginning or end of the other gene. That is, at least one of the subject genes has been cleaved and has recombined with all or part of another gene. The recombination of the two non-homologous gene regions may occur by any method including but not limited to chromosomal gene translocations or in vitro homologous recombinations (such as may occur where a DNA segment is being inserted into a vector or an artificial chromosome or where a vector portion thereof chromosomally integrates in a host cell).
[0097]Preferably, the subject gene breakpoint is a chromosomal gene translocation breakpoint. As detailed hereinbefore, chromosomal gene translocations are known to occur and, in some cases, lead to the onset of disease states. Since a gene translocation between two genes will not necessarily result in the breakpoint occurring at precisely the same nucleotide position on the two genes each time the translocation event occurs, it is not possible to assume that the breakpoint position in one patient, such as the Philadelphia chromosome breakpoint in one CML patient, will be the same in another patient. The method of the present invention enables the simple yet accurate determination of a gene breakpoint using DNA.
The present invention therefore preferably provides a method of identifying a chromosomal gene translocation breakpoint, said method comprising:(i) contacting a genomic DNA sample with: [0098](a) one or more forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0099](b) one or more reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0100]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags;(ii) amplifying the DNA sample of step (i);(iii) optionally contacting the amplicon generated in step (ii) with: [0101](a) one or more forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' to the gene breakpoint, which primers are directed to DNA regions which are located 3' to one or more of the forward primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0102](b) one or more reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' to the gene breakpoint, which primers are directed to DNA regions which are located 5' to one or more of the reverse primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0103]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (iii)(a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (iii)(a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags and which forward and reverse primer tags of step (iii) are different relative to the forward and reverse primer tags of step (i);(iv) amplifying the DNA sample of step (iii);(v) analysing said amplified DNA.
[0104]Reference to "DNA" should be understood as a reference to deoxyribonucleic acid or derivative or analogue thereof. In this regard, it should be understood to encompass all forms of DNA, including cDNA and genomic DNA. The nucleic acid molecules of the present invention may be of any origin including naturally occurring (such as would be derived from a biological sample), recombinantly produced or synthetically produced.
[0105]Reference to "derivatives" should be understood to include reference to fragments, homologs or orthologs of said DNA from natural, synthetic or recombinant sources. "Functional derivatives" should be understood as derivatives which exhibit any one or more of the functional activities of DNA. The derivatives of said DNA sequences include fragments having particular regions of the DNA molecule fused to other proteinaceous or non-proteinaceous molecules. "Analogs" contemplated herein include, but are not limited to, modifications to the nucleotide or nucleic acid molecule such as modifications to its chemical makeup or overall conformation. This includes, for example, modification to the manner in which nucleotides or nucleic acid molecules interact with other nucleotides or nucleic acid molecules such as at the level of backbone formation or complementary base pair hybridisation. The biotinylation or other form of labelling of a nucleotide or nucleic acid molecules is an example of a "functional derivative" as herein defined.
[0106]As detailed hereinbefore, the method of the present invention is predicated on the use of multiple oligonucleotide primers to facilitate the multiplexed amplification of a DNA sample of interest. In one embodiment of the present invention, the DNA sample of interest is a hybrid gene which comprises a portion of one gene (gene A) which is located 5' to the translocation breakpoint and a second gene (gene B) which is located 3' to the translocation breakpoint. In a particular embodiment, gene A is BCR and gene B is ABL. The identification of the existence and nature of a gene translocation breakpoint is achieved by using two or more forward primers directed to gene A and two or more reverse primers directed towards gene B. The primers directed to gene A are designed to hybridise at intervals along gene A and the primers directed to gene B are similarly designed to hybridise at intervals along gene B. In the first round PCR, the primers which will amplify the hybrid gene are the upstream primers which hybridise to that portion of gene A which lies 5' to the breakpoint and the downstream primers which hybridise to that portion of gene B which lies 3' to the breakpoint. Furthermore, since small amplicons are amplified more efficiently than larger amplicons, there will occur selection for amplification directed by the primer pair which hybridises closest to the breakpoint. The same principle holds for the second round primers and, since in one embodiment each second round primer corresponds to an individual first-round primer but hybridises internal to it with regard to the breakpoint, there will be further selection for amplification by the pair of the second round primers which bound the breakpoint. Without limiting the present invention in any way, the second round of PCR amplification provides additional specificity for amplification of the breakpoint region. Following the second round PCR, successful amplification of the sequence surrounding the breakpoint will be evident as a band of amplified material on electrophoresis.
[0107]Since it is not known precisely where the breakpoint lies, it is possible that one or more of the internal primers may not hybridise to their target region sequence due to this sequence having been effectively spliced out during the translocation event. However, in one embodiment, the forward and reverse primers selected for the first round amplification are directed to amplifying from the 5' and 3' end regions, respectively, of the gene fragments flanking the breakpoint. The second round primers are then directed to internal regions of the gene fragments flanking the breakpoint, that is, the regions which are closer to the breakpoint than the regions targeted by the first round primers. Again, it would be appreciated that since the precise location of the breakpoint is not known, one or more of these forward and/or reverse primers may not hybridise to the DNA sample due to their target region sequence having been spliced out. In terms of the second round "internal primers", it should be understood that this is a reference to a population of primers of which at least one primer, but preferably all the primers, are designed to amplify the subject DNA from a point which, when considered in the context of the translocated gene itself (rather than the antisense strand or the amplification product), is 3' of the most 3' of the forward primers used in the first round amplification and 5' of the most 5' of the reverse primers used in the first round amplification. By using the approach of a two step amplification using progressively more internally localised primers, amplification of DNA spanning the breakpoint region can be achieved without the requirement to perform long PCRs or to generate very long and cumbersome amplification products.
[0108]Reference to a "primer" or an "oligonucleotide primer" should be understood as a reference to any molecule comprising a sequence of nucleotides, or functional derivatives or analogues thereof, the function of which includes hybridisation to a region of a nucleic acid molecule of interest (the DNA of interest also being referred to as a "target DNA") and the amplification of the DNA sequence 5' to that region. It should be understood that the primer may comprise non-nucleic acid components. For example, the primer may also comprise a non-nucleic acid tag such as a fluorescent or enzymatic tag or some other non-nucleic acid component which facilitates the use of the molecule as a probe or which otherwise facilitates its detection or immobilisation. The primer may also comprise additional nucleic acid components, such as the oligonucleotide tag which is discussed in more detail hereinafter. In another example, the primer may be a protein nucleic acid which comprises a peptide backbone exhibiting nucleic acid side chains. preferably, said oligonucleotide primer is a DNA primer.
[0109]Reference to "forward primer" should be understood as a reference to a primer which amplifies the target DNA in the DNA sample of interest by hybridising to the antisense strand of the target DNA.
[0110]Reference to "reverse primer" should be understood as a reference to a primer which amplifies the target DNA in the DNA sample of interest and in the PCR by hybridising to the sense strand of the target DNA.
[0111]The design and synthesis of primers suitable for use in the present invention would be well known to those of skill in the art. In one embodiment, the subject primer is 4 to 60 nucleotides in length, in another embodiment 10 to 50 in length, in yet another embodiment 15 to 45 in length, in still another embodiment 20 to 40 in length, in yet another embodiment 25 to 35 in length. In yet still another embodiment, primer is about 26, 27, 28, 29, 30, 31, 32, 33 or 34 nucleotides in length. Without limiting the invention in any way, the primers are designed in one embodiment to have a TM of 65 to 70° C. This enables the PCR to use a high annealing temperature, which minimises non-specific annealing and amplification. Each forward or reverse primer for the second round PCR is designed to hybridise to a sequence which is close, either downstream for the forward primer or upstream for the reverse primer, to the hybridisation sequence for its corresponding forward or reverse first-round primer. Designing the corresponding primers to hybridise to closely adjoining sequences minimises the probability that the translocation breakpoint will involve or occur between the hybridisation sequences. even if this does occur, the sequence surrounding the translation breakpoint can still be amplified by the immediately upstream or downstream, as the case may be, primer pair.
[0112]In the exemplified embodiment described herein, primers were chosen so that their binding sites were staggered with the separation between adjacent binding sites being approximately 500 bases. This was done so that the amplified material would have range in size, up to a maximum length of approximately 1 kilobase. This strategy is in contrast to the strategy of "Long PCR" which would require fewer primers and a less complex multiplex PCR reaction. The advantages of the strategy of the present invention are that the standard shorter PCR reaction is more robust and the amplified product can be sequenced immediately rather than requiring another set of PCR reactions to break it up into smaller amplicons which are suitable for sequencing.
[0113]In terms of the number of primers which are used in the method of the invention, this can be determined by the person of skill in the art. With regard to the total number of primers, the variables which require consideration are the size of the gene region which is being targeted and the distance between the sequences to which the primers hybridise. In order to amplify PCR fragments which are no larger than about 1 kb, the primers can be designed to hybridise at intervals of approximately 500 bases. With regard to CML, nearly all BCR translocations involve one of two regions, each of approximately 3 kb in length. In this case, 12 outer forward primers and 12 corresponding inner primers may be used. The ABL gene, however, is larger, approximately 140 kb in length, and up to 280 outer reverse primers and 280 inner reverse primers may be used. In one particular embodiment, a combination of 6 forward primers and 24 reverse primers is used and in another embodiment a combination of 6 forward primers and 140 reverse primers. The primer number which is selected to be used will depend on the genes involved in the translocation and thus may vary from translocation to translocation and will involve consideration of the competing issues of the number of PCR reactions which are required to be performed versus the probability of generating non-specific products during a PCR reaction. As would be understood by the person of skill in the art, a large number of primers in each individual PCR reaction decreases the number of PCR reactions but increases the probability of non-specific amplification reactions.
[0114]In one embodiment, the method of the present invention is performed using at least three primers, in another embodiment at least four primers. In yet another embodiment said invention is performed using 6-10 primers, 6-15 primers, 6-20 primers, 6-25 primers or 6-30 primers. In still another embodiment there is used 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29 or 30 primers.
There is therefore preferably provided a method of identifying a gene breakpoint, said method comprising:(i) contacting a DNA sample with [0115](a) one to thirty forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0116](b) twenty-four to four hundred reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0117]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags;(ii) amplifying the DNA sample of step (i);(iii) optionally contacting the amplicon generated in step (ii) with: [0118](a) one to thirty forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are directed to DNA regions which are located 3' to one or more of the forward primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0119](b) twenty-four to four hundred reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' to the gene breakpoint, which primers are directed to DNA regions which are located 5' to one or more of the reverse primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0120]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (iii)(a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (iii)(a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags and which forward and reverse primer tags of step (iii) are different relative to the forward and reverse primer tags of step (i);(iv) amplifying the DNA sample of step (iii);(v) analysing said amplified DNA.preferably, said gene breakpoint is a gene translocation breakpoint and still more preferably a chromosomal gene translocation breakpoint.
[0121]The primers which are used in the method of the present invention are of a relatively low individual concentration due to the starting primer pool comprising multiple individual primers. This reduces the risk of inducing inhibition of PCR. In order to facilitate a successful amplification result, it is therefore necessary to enable the generation of sufficient amplicons for detection and isolation. In one aspect of the present invention, this can be facilitated by tagging the primers with an oligonucleotide which can be used as a primer hybridisation site. In addition to the primers directed towards genes A and B, each PCR reaction may therefore also contain concentrations of two oligonucleotides which are directed to the tag, as a primer hybridisation site. These oligonucleotide sequences act as primers and enable efficient secondary amplification of the amplicons generated by the initial hybridisation and extension of the primers directed towards genes A and B. In one embodiment, the primer which is directed to the tag exhibits a TM of 65° C.-70° C. in order to minimise non-specific amplification. Thus these primers are directed towards overcoming the potential problem posed by the low concentrations of the primers directed towards A and B. Nevertheless, in some situations it may not be necessary to use one or both tag primers. For example, when there are only six forward primers for the BCR gene each primer may be at a concentration which is sufficient for relatively efficient amplification. Still further, it should be appreciated that the oligonucleotide tags provide an additional use when they are present in the final PCR round, since the tag primers can also be used for sequencing. Accordingly, although the tag is suitable for use as a site for primer hybridisation, it should be understood that the subject tag may also be useful for other purposes, such as a probe binding site in the context of Southern gel analysis or to enable isolation of the primer or the amplicon extended therefrom. To this end, the tag may comprise a non-nucleic acid component, such as a protein molecule or biotin which would enable isolation, for example by affinity chromatography, streptavidin binding or visualisation.
[0122]In order to ensure that these tags do not interfere with the extension of the primer, the primers are linked to the oligonucleotide tag at their 5' end. Reference to "oligonucleotide tag" should therefore be understood as a reference to a nucleotide sequence of less than 50 nucleotides which is linked to the 5' end of the forward and reverse primers of the present invention. In one embodiment, the tag is 25-30 bases in length. It should also be understood that consistently with the definitions provided in relation to the forward and reverse primers, the oligonucleotide tags herein described may also comprise non-nucleic acid components such as isolation or visualisation tags eg. biotin, enzymatic labels, fluorescent labels and the like. This enables quick and simple isolation or visualisation of the tagged primers or amplicons via non-molecular methods.
[0123]That the oligonucleotide tag is "operably linked" to the primer should be understood as a reference to those regions being linked such that the functional objectives of the tagged primer, as detailed hereinbefore, can be achieved. In terms of the means by which these regions are linked and, further, the means by which the subject oligonucleotide primer binds to its target DNA region, these correspond to various types of interactions. In this regard, reference to "interaction" should be understood as a reference to any form of interaction such as hybridisation between complementary nucleotide base pairs or some other form of interaction such as the formation of bonds between any nucleic or non-nucleic acid portion of the primer molecule or tag molecule with any other nucleic acid or non-nucleic acid molecule, such as the target molecule, a visualisation means, an isolation means or the like. This type of interaction may occur via the formation of bonds such as, but not limited to, covalent bonds, hydrogen bonds, van der Wals forces or any other mechanism of interaction. preferably, to the extent that the interaction occurs between the primer and a region of the target DNA, said interaction is hybridisation between complementary nucleotide base pairs. In order to facilitate this interaction, it is preferable that the target DNA is rendered partially or fully single stranded for a time and under conditions sufficient for hybridisation with the primer to occur.
[0124]Without limiting the present invention to any one theory or mode of action, the inclusion of an oligonucleotide tag which can itself function as a primer hybridisation site can assist in facilitating the convenient and specific amplification of the amplicon generated by the forward and reverse primers of the present invention. Accordingly, this overcomes somewhat the amplification limitation which is inherent where a relatively low starting concentration of the forward and reverse primers is used. Where the starting concentration of forward and reverse primers is sufficiently high, it may not be necessary to use a tag. Accordingly, in a preferred embodiment, the DNA sample of interest is contacted with both the forward and reverse primers of the present invention and primers directed to the oligonucleotide tags of the forward and reverse primers such that the amplification reaction of step (ii) proceeds in the context of all these primers. It should be understood, however, that although it is preferred that amplification based on both the gene primers and the tag primers is performed simultaneously, the method can be adapted to perform the tag primer based amplification step after the completion of the gene primer based amplification.
[0125]The DNA sequence of the tags may be the same or different. With respect to a first round amplification, the tags may be the same if the purpose is to amplify the initial amplification product. However, if one wishes to selectively enrich for amplicons containing the sequence of one of the flanking genes, the primer directed to the tag region of the primer of the gene of interest (eg. gene A) should differ to the primer directed to the tag region of the primer of the other gene (eg. gene B). In another example, in terms of a second or subsequent round of amplification, the tags which are used for sequencing would be required to be different to prevent the simultaneous sequencing of both strands.
The present invention therefore provides a method of identifying a gene translocation breakpoint, said method comprising:(i) contacting a DNA sample with: [0126](a) one to thirty forward primers directed to a DNA region of the antisense strand of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0127](b) twenty-four to four hundred reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' relative to the gene breakpoint, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0128]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags; [0129](c) a primer directed to the forward primer oligonucleotide tag of step (i)(a); and [0130](d) a primer directed to the reverse primer oligonucleotide tag of step (i)(b);(ii) amplifying the DNA sample of step (i);(iii) optionally contacting the amplicon generated in step (ii) with: [0131](a) one to thirty forward primers directed to a DNA region of the flanking gene or fragment thereof located 5' relative to the gene breakpoint, which primers are directed to DNA regions which are located 3' to one or more of the forward primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0132](b) twenty-four to four hundred reverse primers directed to a DNA region of the flanking gene or fragment thereof located 3' to the gene breakpoint, which primers are directed to DNA regions which are located 5' to one or more of the reverse primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0133](c) a primer directed to the forward primer oligonucleotide tag of step (iii)(a); and [0134](d) a primer directed to the reverse primer oligonucleotide tag of step (iii)(b); [0135]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (iii)(a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (iii)(a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags and which forward and reverse primer tags of step (iii) are different relative to the forward and reverse primer tags of step (i);(iv) amplifying the DNA sample of step (iii);(v) analysing said amplified DNA.preferably said gene translocation breakpoint is a chromosomal gene translocation breakpoint.
[0136]It should be understood that the oligonucleotide primers and tags of the present invention should not be limited to the specific structure exemplified herein (being a linear, single-stranded molecule) but may extend to any suitable structural configuration which achieves the functional objectives detailed herein. For example, it may be desirable that all or part of the oligonucleotide is double stranded, comprises a looped region (such as a hairpin bend) or takes the form of an open circle confirmation, that is, where the nucleotide primer is substantially circular in shape but its terminal regions do not connect.
[0137]Facilitating the interaction of the nucleic acid primer with the target DNA may be performed by any suitable method. Those methods will be known to those skilled in the art.
[0138]Methods for achieving primer directed amplification are also very well known to those of skill in the art. In a preferred method, said amplification is polymerase chain reaction, NASBA or strand displacement amplification. Most preferably, said amplification is polymerase chain reaction. To this end, in one embodiment of the invention, a 20 minute hybridisation provides good amplification in the first round PCR.
[0139]Reference to a "sample" should be understood as a reference to either a biological or a non-biological sample. Examples of non-biological samples includes, for example, the nucleic acid products of synthetically produced nucleic acid populations. Reference to a "biological sample" should be understood as a reference to any sample of biological material derived from an animal, plant or microorganism (including cultures of microorganisms) such as, but not limited to, cellular material, blood, mucus, faeces, urine, tissue biopsy specimens, fluid which has been introduced into the body of an animal and subsequently removed (such as, for example, the saline solution extracted from the lung following lung lavage or the solution retrieved from an enema wash), plant material or plant propagation material such as seeds or flowers or a microorganism colony. The biological sample which is tested according to the method of the present invention may be tested directly or may require some form of treatment prior to testing. For example, a biopsy sample may require homogenisation prior to testing or it may require sectioning for in situ testing. Further, to the extent that the biological sample is not in liquid form, (if such form is required for testing) it may require the addition of a reagent, such as a buffer, to mobilise the sample.
[0140]To the extent that the target DNA is present in a biological sample, the biological sample may be directly tested or else all or some of the nucleic acid material present in the biological sample may be isolated prior to testing. It is within the scope of the present invention for the target nucleic acid molecule to be pre-treated prior to testing, for example inactivation of live virus or being run on a gel. It should also be understood that the biological sample may be freshly harvested or it may have been stored (for example by freezing) prior to testing or otherwise treated prior to testing (such as by undergoing culturing).
[0141]Reference to "contacting" the sample with the primer should be understood as a reference to facilitating the mixing of the primer with the sample such that interaction (for example, hybridisation) can occur. Means of achieving this objective would be well known to those of skill in the art.
[0142]The choice of what type of sample is most suitable for testing in accordance with the method disclosed herein will be dependent on the nature of the situation, such as the nature of the condition being monitored. For example, in a preferred embodiment a neoplastic condition is the subject of analysis. If the neoplastic condition is a lymphoid leukemia, a blood sample, lymph fluid sample or bone marrow aspirate would likely provide a suitable testing sample. Where the neoplastic condition is a lymphoma, a lymph node biopsy or a blood or marrow sample would likely provide a suitable source of tissue for testing. Consideration would also be required as to whether one is monitoring the original source of the neoplastic cells or whether the presence of metastases or other forms of spreading of the neoplasia from the point of origin is to be monitored. In this regard, it may be desirable to harvest and test a number of different samples from any one mammal. Choosing an appropriate sample for any given detection scenario would fall within the skills of the person of ordinary skill in the art.
[0143]The term "mammal" to the extent that it is used herein includes humans, primates, livestock animals (e.g. horses, cattle, sheep, pigs, donkeys), laboratory test animals (e.g. mice, rats, rabbits, guinea pigs), companion animals (eg. dogs, cats) and captive wild animals (eg. kangaroos, deer, foxes). preferably, the mammal is a human or a laboratory test animal. Even more preferably the mammal is a human.
[0144]As detailed hereinbefore, in one embodiment the method of the present invention is performed as a sequential two step amplification using multiple second round primers each of which is directed to a gene region which is either 3' (for the forward primers) or 5' (for the reverse primers) to that which is targeted by the corresponding first round primers. The person of skill in the art would appreciate that in some cases it may not be necessary to conduct a second round amplification. The necessity to perform a second round amplification may also be obviated if a selective or enrichment step as described below is performed. This situation may arise when the sequence around the breakpoint is amplified very efficiently and there is very little non-specific amplification such that a clearly defined band of amplification product is observed on electrophoresis of the product of the first round amplification or if the subsequent selection step is very efficient. In general, however, it is expected that a sequential two step amplification process would be used in order to minimise non-specific amplification and to generate a relatively short amplification product which spans the breakpoint region. In general, it is expected that the amplification product would be less than 1.5 kb, less than 1 kb, less than 0.8 kb or less than 0.5 kb. It should be understood that depending on the size of the genes which have been translocated, the method of the invention may be adapted to incorporate third or fourth round amplification steps in order to further minimise non-specific amplification. This can be an issue owing to the number of primers present in the multiplexed reaction and to the fact that one of the genes participating in the translocation often contains multiple repetitive sequences such as Alu. Nevertheless, it is expected that the need for further rounds of amplification would be unlikely.
[0145]Although the method of the present invention has been designed such that the amplification steps can be sequentially performed directly on the amplification product of a previous amplification, this should not be understood as a limitation in terms of whether any additional steps are sought to be incorporated by the skilled person, such as enrichment/selection steps. For example, one may seek to select for the desired amplicons after the first round amplification and to thereafter conduct the second round amplification on their material alone. Methods which one could utilise to select or enrich include: [0146]a selection step based on the unique oligonucleotide tags which are linked to the primers. Accordingly, since the tags themselves are also amplified and therefore form part of the amplicon, they could be used as a probe site to enable isolation of amplicons which are the result of both forward and reverse primer amplification and therefore should span the breakpoint. Alternatively, biotinylation of one of the tags provides means of identifying and isolating amplicons which have resulted from extension by either the forward or reverse primers. For example, by flooding the amplification product with biotinylated primer, the primer can act as a probe to identify the amplicons of interest and the biotinylation can provide a basis for isolating those amplicons. By ensuring that each of the primer groups of the present invention comprises a unique tag, it is possible to select out, with significant particularity, only specific amplicons of interest. In particular, the skilled person would seek to exclude amplicons which have been amplified by a forward primer but which have not then been amplified by a reverse primer, thereby indicating that the subject amplicon possibly does not extend across the breakpoint. By selecting out the amplicons which are most likely spanning the breakpoint, a subsequent round of amplification is more specifically targeted and less likely to generate unwanted amplicons as a result of either inherent cross-hybridisation of primers or the amplification of amplicons which do not flank both sides of the breakpoint. [0147](ii) One may seek to run the products on a gel and excise out only certain bands or regions which are likely to be relevant and thereafter subject these to a further amplification step. When a band is present on the gel after the second round amplification, if there are any problems in sequencing an attempt can be made to clean it up by cutting the product out of the gel and performing a series of PCR reactions using individual primers and/or smaller pools of primers. For example, one might use individual forward BCR primers and pools containing only 12 reverse ABL primers. [0148](iii) one may expose the amplified products to one or more rounds of bottleneck PCR in order to provide negative selection against non-specific amplified products.
[0149]Without limiting the application of the present invention to any one theory or mode of action, in a classical PCR, the primers and reaction conditions are designed so that primer hybridisation and extension of the forward and reverse primers occur at or close to the maximum efficiency so that the number of amplicons approximately doubles with each cycle resulting in efficient exponential amplification. Bottleneck PCR, however, is predicated on the use of forward and reverse primer sets where the primers of one set have been designed or are otherwise used under conditions wherein they do not hybridise and extend efficiently. Accordingly, although the efficient primer set will amplify normally, the inefficient set will not. As a consequence, when a sequence of interest is amplified, the number of amplicon strands is significantly less than that which would occur in a classical PCR. Efficient amplification only commences once amplicons have been generated which incorporate, at one end, the tag region of the inefficient primer. At this point, the primers directed to the tag regions effect a normal amplification rate. A "bottleneck" is therefore effectively created in terms of the generation of transcripts from the inefficient primer set.
[0150]A more severe bottleneck is usefully created where the inefficient primers are directed to commonly repeated sequences, such as an alu sequence. Amplification of unwanted product may result if such binding sites are closely apposed and if the inefficient primers can act as forward primers and reverse primers. However, owing to both primers being inefficient, amplification is initially extremely inefficient and there is a severe bottleneck. Efficient amplification only commences once amplicon strands have been generated which comprise the tag region of the inefficient primer at one end and its complement at the other. After any given number of cycles, the number of such amplicons is, however, substantially less than that which occurs during amplification of the sequence of interest. The amount of unwanted product at the end of the amplification reaction is correspondingly reduced.
[0151]Hybridisation and extension of an inefficient primer which has correctly hybridised to the sequence of interest followed in a subsequent cycle by hybridisation and extension of an efficient primer to the previously synthesised amplicon generates a template to which the tag primer can efficiently hybridise and extend. Since such molecules together with their complements provide upstream and downstream binding sites, each for an efficient primer (the tag primer and one member of the efficient set), succeeding cycles of amplification from such templates are both efficient and exponential. The result is that, after an initial lag or "bottleneck", the overall rate of amplification speeds up in later cycles so that a near doubling of amplicon number with each cycle results. However, the net result is that there is negative selection against amplification of undesired amplicons as compared to amplicons of the sequence of interest, owing to the bottleneck at each end for the former and only at one end for the latter.
[0152]Accordingly, if the same number of commencing target sequences is considered and comparison to the amplification produced by classical PCR is made, application of the bottleneck PCR will produce a lesser increase in the number of amplicons of the sequence of interest and an even lesser increase in the number of amplicons of unwanted sequences. Although amplification of both wanted and unwanted products occurs, there is relative enrichment of the sequence of interest relative to the unwanted sequences. There is an inverse relationship between absolute amplification and enrichment since decreasing the efficiency of the inefficient primer set produces increased enrichment at the expense of lesser amplification.
[0153]Once the amplification rounds have been completed, the amplicons spanning the breakpoint region can be analysed. In a preferred embodiment, the subject amplicon is isolated by excision of a gel band containing that amplicon and sequenced in order to characterise the breakpoint region. To the extent that a band excised from a gel is to be analysed, it may be necessary to further amplify the DNA contained therein in order to provide sufficient material for sequencing. The oligonucleotide tags hereinbefore described provide a suitable primer hybridisation site to facilitate further amplification of the isolated amplicons.
[0154]As detailed hereinbefore, the method of the present invention provides a simple and routine means of identifying and characterising any breakpoint region, such as the nature, accuracy and stability of a site directed insertion of a gene into a chromosome or vector (this being important in the context of gene therapy), but in particular the chromosomal gene translocation breakpoints that are characteristic of many diseases. Examples of such translocations and diseases include, but are not limited to: [0155]t(2;5)(p23;q35)--anaplastic large cell lymphoma [0156]t(8;14) --Burkitt's lymphoma (c-myc) [0157]t(9;22)(q34;q11)--Philadelphia chromosome, CML, ALL (BCR-ABL recombination) [0158]t(11;14)--Mantle cell lymphoma (Bcl-1) [0159]t(11;22)(q24;q11.2-12)--Ewing's sarcoma [0160]t(14;18)(q32;q21)--follicular lymphoma (Bcl-2) [0161]t(17;22)--dermatofibrosarcoma protuberans [0162]t(15;17)--acute promyelocytic leukemia (pml and retinoic acid receptor genes) [0163]t(1;12)(q21;p13)--acute myelogenous leukemia [0164]t(9;12)(p24;p13)--CML, ALL (TEL-JAK2) [0165]t(X;18)(p11.2;q11.2)--Synovial sarcoma [0166]t(1;11)(q42.1;q14.3)--Schizophrenia [0167]t(1;19)--acute pre-B cell leukemia (PBX-1 and E2A genes).preferably, said chromosomal gene translocation is a BCR-ABL translocation or a PML-RARalpha translocation.According to this preferred embodiment there is provided a method of identifying a chromosomal BCR-ABL translocation breakpoint, said method comprising:(i) contacting a DNA sample with: [0168](a) one or more forward primers directed to a DNA region of BCR or fragment thereof, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0169](b) one or more reverse primers directed to a DNA region of ABL or fragment thereof, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0170]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags;(ii) amplifying the DNA sample of step (i);(iii) optionally contacting the amplicon generated in step (ii) with: [0171](a) one or more forward primers directed to a DNA region of BCR or fragment thereof, which primers are directed to DNA regions which are located 3' to one or more of the forward primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; and [0172](b) one or more reverse primers directed to ABL or fragment thereof, which primers are directed to DNA regions which are located 5' to one or more of the reverse primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0173]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (iii)(a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (iii)(a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags and which forward and reverse primer tags of step (iii) are different relative to the forward and reverse primer tags of step (i);(iv) amplifying the DNA sample of step (iii);(v) analysing said amplified DNA.preferably, said amplification steps are performed using 1-30 forward primers and 24-300 reverse primers.
[0174]In terms of the embodiment of the invention exemplified herein, primers were chosen so that their binding sites were staggered with the separation between adjacent binding sites being approximately 500 bases. This was done so that the amplified material would have range in size, up to a maximum length of approximately 1 kilobase. This strategy may be contrasted to the prior art strategy of "Long PCR" which would require fewer primers and a less complex multiplex PCR reaction. One of the advantages of the strategy of the present invention is that the standard shorter PCR reaction is more robust and the amplified product can be sequenced immediately rather than requiring another set of PCR reactions to break it up into smaller amplicons which are suitable for sequencing.
The present invention therefore preferably provides a method of identifying a chromosomal BCR-ABL translocation breakpoint, said method comprising:(i) contacting a DNA sample with: [0175](a) one to thirty forward primers directed to a DNA region of BCR or fragment thereof, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0176](b) twenty-four to four hundred reverse primers directed to a DNA region of ABL or fragment thereof, which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0177]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags; [0178](c) a primer directed to the forward primer oligonucleotide tag of step (i)(a); and [0179](d) a primer directed to the reverse primer oligonucleotide tag of step (i)(b);(ii) amplifying the DNA sample of step (i);(iii) contacting the amplicon generated in step (ii) with: [0180](a) one to thirty forward primers directed to a DNA region of BCR or fragment thereof, which primers are directed to DNA regions which are located 3' to one or more of the forward primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0181](b) twenty-four to four hundred reverse primers directed to a DNA region of ABL or fragment thereof, which primers are directed to DNA regions which are located 5' to one or more of the reverse primers of step (i) and which primers are optionally operably linked at their 5' end to an oligonucleotide tag; [0182](c) a primer directed to the forward primer oligonucleotide tag of step (iii)(a); and [0183](d) a primer directed to the reverse primer oligonucleotide tag of step (iii)(b); [0184]wherein the oligonucleotide tags of the forward primers are the same relative to the forward primer tags of step (iii)(a) and the oligonucleotide tags of the reverse primers are the same relative to the reverse primer tags of step (iii)(a) but which forward primer oligonucleotide tags are different relative to the reverse primer tags and which forward and reverse primer tags of step (iii) are different relative to the forward and reverse primer tags of step (i);(iv) amplifying the DNA sample of step (iii);(v) isolating and sequencing said amplified DNA.More preferably, said DNA sequence is a blood derived sample.The method of the present invention has broad application including, but not limited to: [0185](i) enabling the design and generation of patient specific probes which can be used for the ongoing monitoring of a patient who is diagnosed with a disease condition characterised by chromosomal gene translocation. Results obtained by this means for chronic myeloid leukemia are shown in FIG. 6. [0186](ii) the analysis and monitoring of in vitro and in vivo gene transfection systems which are directed to integrating a gene or other DNA region into a chromosome, vector, plasmid, artificial chromosome or the like. Where the general site at which recombination should occur is known, the present invention can be applied to determine the specific point and nature of the integration (i.e. the breakpoint). It can also be used to monitor the ongoing stability of the genetic recombination event by virtue of enabling the generation of specific primers.
[0187]Accordingly, in yet another aspect there is provided a method of monitoring a disease condition in a mammal, which disease condition is characterised by a gene breakpoint, said method comprising screening for the presence of said breakpoint in a biological sample derived from said mammal, which breakpoint has been identified in accordance with the method hereinbefore defined.
[0188]Methods of screening for the subject breakpoint would be well known to those skilled in the art and include any suitable probe-based screening technique, such as PCR based methods. By virtue of the identification of the breakpoint region in accordance with the method of the invention, one can design an appropriate probe set to specifically amplify the subject breakpoint.
In one embodiment, said gene breakpoint is a chromosomal gene translocation breakpoint such as: [0189]t(2;5)(p23;q35) [0190]t(8;14) [0191]t(9;22)(q34;q11) [0192]t(11;14) [0193]t(11;22)(q24;q11.2-12) [0194]t(14;18)(q32;q21) [0195]t(17;22) [0196]t(15;17) [0197]t(1;12)(q21;p13) [0198]t(9;12)(p24;p13) [0199]t(X;18)(p11.2;q11.2) [0200]t(1;11)(q42.1;q14.3) [0201]t(1;19).In another embodiment, said condition is: [0202]anaplastic large cell lymphoma [0203]Burkitt's lymphoma [0204]CML, ALL [0205]Mantle cell lymphoma [0206]Ewing's sarcoma [0207]follicular lymphoma [0208]dermatofibrosarcoma protuberans [0209]acute promyelocytic leukemia [0210]acute myelogenous leukemia [0211]Synovial sarcoma [0212]Schizophrenia; or [0213]acute pre-B cell leukemia.
[0214]Still another aspect of the present invention is directed to a DNA primer set, which primer set is designed to amplify and/or otherwise detect a gene breakpoint, which breakpoint has been identified in accordance with the method hereinbefore defined.
[0215]The present invention is now described by reference to the following non-limiting examples and figures.
Example 1
Isolation of BCR/ABL Breakpoint Product from gDNA of Patient 1
[0216]Genomic DNA extracted by Qiagen Flexigene kit
[0217]1st Round PCR (50 ng genomic DNA)--all reactions performed in duplicate
Forward primer pool--FA (Contains 7 forward BCR primers BCRF1-BCRF7 each with same 5' tag sequence (A), Total 50 ng (7.14 ng each)Reverse primer pool--R3/4 (Pool of 24 oligonucleotide reverse ABL primers, each with same 5' tag sequence (C), Total 50 ng (2.08 ng each)Forward and reverse tag sequence primers (A,C) --25 ng of each
PCR Conditions
[0218]1×PCR buffer, 5 mM MgCl2, 0.75 ul dUTP (300 uM each), 0.4 ul Platinum Taq (2 U)
Cycling Conditions
[0219]95/4 min
[0220](97° C./1 min, 65° C./20 min, 72° C./1 min)×5
[0221](96° C./30 sec, 65° C./20 min, 72° C./1 min)×5
[0222](92° C./30 sec, 65° C./20 min, 72° C./1 min)×10
[0223]2nd Round PCR (1st round reaction diluted 1/200 in sterile water)
Forward primer pool--NFA (Contains 7 forward internal BCR primers BFN1-BFN7 each with same 5' tag sequence (B), Total 50 ng (7.14 ng each)Reverse primer pool--RN3/4 (Pool of 24 oligonucleotide reverse internal ABL primers, each with same 5' tag sequence (D), Total 50 ng (2.08 ng each)Forward and reverse tags (B,D)--25 ng of each
PCR Conditions
[0224]1×PCR buffer, 5 mM MgCl2, 0.75 ul dUTP (300 uM), 0.4 ul Pt Taq (2 U)
Cycling Conditions
[0225]95/4 min
[0226](94° C./30 sec, 65° C./10 min, 72° C./1 min) x10
[0227](94° C./30 sec, 65° C./5 min, 72° C./1 min) x15 [0228]PCR products (7 ul) resolved on 1.5% (v/v) agarose gel at 120 voltsIdentification of BCR/ABL Breakpoint from Patient 1 [0229]PCR products resolved on 1.5% (v/v) agarose gel at 120 volt Band excised and purified via Flexigene kit [0230]Reamplification of Bands by PCR (1/1000 Dilution of Purified Product)Forward primer--Tag B (25 ng)Reverse primer--TagD (25 ng)PCR conditions
[0231]1×PCR buffer, 5 mM MgCl2, 0.75 ul dUTP (300 uM), 0.4 ul Pt Taq (2 U)
Cycling Conditions
[0232]95/4 min
[0233](94° C./30 sec, 65° C./30 sec, 72° C./30 sec)×35 [0234]PCR Product Sequenced with TagB primer (Flinders Sequencing Facility)
Confirmation of Breakpoint by PCR
[0234] [0235]PCR performed on gDNA (50 ng) across breakpointPatient 1 gDNA vs 10× Normal gDNA (several primer combinations)Forward primer--BCR (patient specific) (25 ng)Reverse primer--ABL (patient specific) (25 ng)
PCR Conditions
[0236]1×PCR buffer, 5 mM MgCl2, 0.75 ul dUTP (300 uM), 0.4 ul Pt Taq (2 U)
Cycling Conditions
[0237]95/4 min
[0238](97° C./1 min, 65° C./30 sec, 72° C./30 sec)×5
[0239](96° C./30 sec, 65° C./30 sec, 72° C./30 sec)×5
[0240](92° C./30 sec, 65° C./30 sec, 72° C./30 sec)×25 [0241]PCR products resolved on 3% (v/v) agarose gel at 120 volt
[0242]Band excised and purified via Qiagen minElute kit
[0243]Products sequenced with 5' BCR specific primer to confirm BCR/ABL breakpoint (Flinders sequencing facility).
[0244]Nearly all translocations involve a 3 kb region of the BCR gene and 140 kb region of the ABL gene. Six forward primers used to cover the region of the BCR gene and 282 primers used to cover the region of the ABL gene. Six PCRs are set up, each containing one of the BCR primers, all of the ABL primers, and the common tag primer.
[0245]If necessary, a second round of PCR is performed with a nested internal BCR primer and 282 nested internal ABL primers Alternatively, 1-3 rounds of Bottleneck PCR are performed in order to remove non-specific amplified products and reveal the amplified translocation sequence.
[0246]The ABL gene is very rich in Alu sequences, and the BCR gene also contains one such sequence. The ABL primers have therefore undergone a selection procedure which sequentially involves, for each ABL primer: [0247]design using standard criteria [0248]pairing with each BCR primer and testing by electronic PCR for amplification off the BCR template. Primers that fail this criterion are discarded. [0249]incorporation in a pool of 12 or 24 ABL primers, pairing the pool with each BCR primer, and testing by experimental PCR using a BCR template which has been previously produced by PCR amplification. Any pool that that produces amplification and thus fails this test is further analysed by testing each of the individual ABL primers to determine which is responsible for amplification. When identified, this primer is discarded.
[0250]The BCR and ABL primers used in Example 1 are shown in Example 2.
Example 2
Primers Used for Isolation of BCR-ABL Translocation Breakpoint in Chronic Myeloid Leukemia
BCR Primers
TABLE-US-00001 [0251] 1st Rd BCRF1-FT0 cttctccctgacatccgtgg BCRF2-FT0 (-5) acacagcatacgctatgcacatgtg BCRF3-FT0 gaggttgttcagatgaccacgg BCRF4-FT1 (-10) cagctactggagctgtcagaacag BCRF5-FT0 tgggcctccctgcatcc BCRF6-FT0 tccccctgcaccccacg 2nd Rd BCRF1-FT1 tgacatccgtggagctgcagatgc BCRF2-FT1 acatgtgtccacacacaccccacc BCRF3-FT1 accacgggacacctttgaccctgg BCRF4-FT1 (-4) ctggagctgtcagaacagtgaagg BCRF5-FT1 tccctgcatccctgcatctcctcc BCRF6-FT1 cccacgacttctccagcactgagc
[0252]The second round primers were internal to the first round primers and were used either for a second round together with internal ABL primers or for performing Bottleneck PCR in order to eliminate non-specific amplified material and facilitate isolation of the translocation breakpoint.
[0253]Various combinations of the forward and reverse primers can be used. In one embodiment, the protocol that was used was to set up 6 PCRs, each containing a different BCR primer and all 282 ABL primers
282 Reverse ABL Primers Used for the First PCR Round and the Tag Sequence which was on the 5' End of Each Primer
TABLE-US-00002 Tag A gcaacactgtgacgtactggagg R1 gtctatctaaaattcacaaggaatgc R2 aggcaaagtaaaatccaagcaccc R3 cactcctgcactccagcctgg R4 caaccaccaaagtgcttttcctgg R5 atatggcatctgtaaatattaccacc R6 tgcctcggcctcccaaagtgc R7 agccaccacacccagccagg R8 aataactgttttctccccccaaaac R9 tgttttacaaaaatggggccatacc R10 acttaagcaaattctttcataaaaaggg R11 ctttcaattgttgtaccaactctcc R12 acctcctgcatctctccttttgc R13 aaataaagttttgagaaccataagtgg R14 caccatcacagctcactgcagc R15 aacctctttgagaatcggatagcc R16 aaataaagtacatacctccaattttgc R17 gacacattcctatgggtttaattcc R18 tgtaaaatatggtttcagaagggagg R19 gcaggtggataacgaggtcagg R20 ccagccaagaatttcaaagattagc R21 gaagggagatgacaaagggaacg R22 gcagaagaactgcttgaacctgg R23 gtggtcccagctactcgagagg R24 ccctcagcaaaactaactgaaaagg R25 tagaaaccaagatatctagaattccc R26 ccacgcccggcggaataaatgc R27 acaaaaaaagaggcaaaaactgagag R28 ctgggcgcagtggctcatgcc R29 tggctgtgaggctgagaactgc R30 ctgggcgacagagtgagactcc R31 aagtctggctgggcgcagtgg R32 aatggacaaaagaggtgaactggc R33 gatagagtgaaaacgcacaatggc R34 aattaaacagctaggtcaatatgagg R35 ggtctccactatcaagggacaag R36 aagcagctgttagtcatttccagg R37 aggcatcctcagattatggctcc R38 cctgagtaacactgagaccctgc R39 aacactcaagctgtcaagagacac R40 attcaggccaggcgcagtggc R41 taaatcgtaaaactgccacaaagc R42 cagaggagtaggagaaggaaaagg R43 ggtagctatctaccaagtagaatcc R44 atcagattggaaaaagtcccaaagc R45 ctcctgaaaagcacctactcagc R46 ctccttaaacctgaggtactggg R47 ttttctcctaatagaccaccattcc R48 ctgctgtattaccatcactcatgtc R49 ctggccaacatagtgaaaccacg R50 atttgaataggggttaaagtatcattg R51 cacttcagtggaagttggcatgc R52 gtttttcttcgaagtgataaacatacg R53 gctccttagtctatgtacctgtgg R54 tactctggcatggtaactggtgc R55 acaaaggactaggtctgtggagc R56 ccaagtttaccaaattaccaaagttacc R57 tgagccgatatcacgccactgc R58 tcccaataaaggttttggcccagg R59 ctgggtagcaaattagggaacagg R60 ctggccagaaaagacagttttatcc R61 ggttcccaggaagggataacacc R62 tcactccaggaggttccatttcc R63 aggcttggaaataagcagcagtgg R64 attcatacaatggaatactactcagc R65 taagtgatcctcccacctcaacc R66 tataagaggaagactggggctgg R67 tcatacttatgcaggttataggagg R68 caagatcacgccactgcactcc R69 aaaataaatagctggtgctcaagatc R70 caccagcctcattcaacagatgg R71 caatgcagcctcaacctcctgg R72 gttaggtcaggtgctcatgtctg R73 aagtttcaaaaggacatgtacaaaatg R74 tcctgaagaggctgcagcttcc R75 ctggtgcacattcccaagtgtgc R76 catgttggccatgttcttctgagg R77 ctcagcctcccgagtagctgg R78 aaagacatttaagaggagatgaggc R79 tgctgggattacaggcgtgagc R80 tgtgacttccatccgcagctcc R81 gacacttttgtggagctttcatgg R82 catgtgagggggcacgtcttgc R83 tcttctctatgagaaaagtggttgc R84 tggcaaaatgctatcgagctgcc R85 tatgaacacagccggcctcagg R86 gaggttgcagtgagctgagatcg R87 gtcaagcacccagtccgatacc R88 atctgggcttggtggcgcacg R89 gttaagcgggtcccacatcagc R90 cagccagtttcagtagaaagatgc R91 gacccaagcataaggggactagc R92 cccaaaaagtttacaagagaaattttc R93 cgcctgtagtcccagctactcg R94 cgcgtgatgcggaaaagaaatcc R95 tctactatgaaccctccttcagac R96 gtgctgggattacaggtgtgagc R97 ttatccaaatgtcccagggcagg R98 ctgccagcactgctcgccagc R99 gctactgcaggcagtgccttcc R100 catccaagcccaaggtgtcagg R101 tgtttgcatgtaatttcaggaagcc R102 gatccgtcactgttaacactcagg R103 ctcacagtcacaagctcctgagc R104 gagatgatgctggggtcacagg R105 ttagaagaatgggatcgcaaagg R106 cggtattcaaatatgaggtcaggc R107 gtaaatcctgctgccagtcttcc R108 acagggtcagacagagccttgg R109 agttattgatctaactatacaacaagc R110 aaagactaggggccggggacg R111 ctggtagaaataaagacaacaaagcc R112 gtgccaagtaattaaaagtttgaaacc R113 ggcttttgaagggagcaccacc R114 gaaggataaatacctatgatactttcc R115 ggcagggaaatactgtgcttcaag R116 gtggtgaaattccacctcagtacc R117 tcccaaagtgctgggattacagg R118 gaaattagcaaacaatgccaagacg R119 taagtattggaccgggaaggagg R120 ctatcattttgctcaaagtgtagcc R121 atttcacaaactacagaggccagg R122 tagacttctgtctctctatgctgc R123 tgagtgagctgccatgtgatacc R124 acttcacaccagcctgtccacc
R125 taactcatatcctcagagagaccc R126 agaggttcctcgattcccctgc R127 gtgtcagcgtcccaacacaaagc R128 gaaagtggatgggcaagcattgc R129 gtgatcacctcacagctgcagg R130 gtttgtttagtcaaggcatttcacc R131 cctcagcctccagagtagctgg R132 taaaagaaaactcctccttcctgg R133 aatgtgctatgtctttaaatccatgg R134 agctggcaaatctggtaatataaaag R135 gcttgaacctggaaggtggagg R136 gcaggcatgctaagaccttcagc R137 cagctccatgaataactccacagg R138 gcttgaacccaggaggcagagg R139 atcgaagatgccactgcaagagg R140 ccaaccacacttcaggggatacc R141 cacgccagtccactgatactcac R142 gggtttcaccatgttggccagg R143 cccaacaaaggctctggcctgg R144 atgacagcagaggagcttcatcc R145 gcaggctacgagtaaaaggatgg R146 cgggtaaaatcttgcctccttcc R147 aaacttaaaccaatggtggatgtgg R148 agagactgaggaactgttccagc R149 gaaacggtcttggatcactgatcc R150 tgcgcatgatatcttgtttcaggg R151 ggcctccgtttaaactgttgtgc R152 gaatgctggcccgacacagtgg R153 tcttggtatagaaaagccagctgg R154 gcaaaagcccaagagcccctgg R155 ttctcccaaaatgagccccaagg R156 gtggtgacgtaaacaaaaggtacc R157 gcaaattccatgtgaatcttattggc R158 cctgatctatggaacagtggtgg R159 gttacaaacgttgcagtttgcaacg R160 gaaccccgtcaacagtgatcacc R161 acaggacctcaaggcaaggagc R162 catacctaaaatagaaatgtctatccc R163 gagttgcatatatgttttataaatccc R164 tgagcccacatccataaagttagc R165 accgcaacctttgccgcctgg R166 taaatattttgtatggagtcaccacc R167 aaagccaggagaaaaagttatgagg R168 tcccaaagtcccaggattacagg R169 tcactatggagcatctccgatgg R170 agttccctggaagtctccgagg R171 aaaataatcacccagcccacatcc R172 acaaaactacagacacagaaagtgg R173 tttgggaggctgaggtaggtgg R174 aaagacagtgaaacatctataaggg R175 cattttgggagaccagggcagg R176 gcatgggacagacacaaagcagc R177 gaataacaaagagagccggctgg R178 taaaccttttattgaaaattgtcaaatgg R179 cgcctcagcctcccaaagtgc R180 tacattagttttataggtccagtagg R181 gaaggtttattcatattaaaatgtgcc R182 ctggcttctgtggtttgagttgg R183 acagacctacctcctaaggatgg R184 gctagcttttgtgtgtaagaatggg R185 ggcctactcacacaatagaatacc R186 gcaccattgcactccagcctgg R187 gaaattaggataaaggttgtcacagc R188 cagaagtgttcaaggtgaaactgtc R189 ctgaatcatgaaatgttctactctgc R190 tgtcaacttgactgggccatacg R191 ctcccgtatagttgggattatagg R192 gcttggagttccttgaaattcttgg R193 cctggtggctccagttttctacc R194 aactcctgacctcatgatccacc R195 gctgggattacaggcatgagcc R196 ttctcctttatccttggtgacattc R197 tcccaaagtgctgggattacagg R198 gtcataagtcagggaccatctgc R199 ctgtttcattgatttccagactggc R200 gcaatctcggctcactgcaagc R201 gaagaagtgactatatcagatctgg R202 ttcaccatgttggccaggctgg R203 catcactgaagatgacaactgagc R204 gtccagcctgggcgatagagc R205 gaggaaagtctttgaagaggaacc R206 ggtacactcaccagcagttttgc R207 gagcaactggtgtgaatacatatgg R208 caatacctggcaccacatacacc R209 gggactacaggcatgtgccacc R210 cggtggctcacgcgtgtaatcc R211 caactgttaaatctctcatggaaacc R212 gacaaaggattagaaatgcaccc R213 ggaaatgttctaaaactggattgtgg R214 aataataatagccaggtgtggtagc R215 ctggaacactcacacattgctgg R216 ctgggtgacagagcgagactcc R217 cccaaatcatccccgtgaaacatgc R218 gaccctgcaatcccaacactgg R219 ctctcaggccttcaaactacacc R220 caggaaagggctcgctcagtgg R221 atctgcaaaagcagcagagcagg R222 gtacccatgacagacaagttttagg R223 cttatcccctactgtctcctttgg R224 ggatggtctcgatctcctgacc R225 aggttagagaccttcctctaatgc R226 agctgggattacaggtgcctgc R227 gctgaggcaggttggggctgc R228 acatttaacgtctcctaacttctcc R229 gtgctgcgattacaggtgtgagc R230 tatgacagcagtattatactatcacc R231 ctggggaccaaatctgaactgcc R232 gtagctattgttatttccaaaagagg R233 gcttgggaccccaggacaagg R234 cctggccaacatggggaaatcc R235 aattgcttgaacctgggaggtgg R236 gcctaagacccaaaagctattagc R237 catattaaagggccatattcaaattgg R238 ggatgtaaccagtgtatatcacagg R239 ggaagtttagtccacatcttctagc R240 gcacccacaggacaaccacacg R241 gggacgcgcctgttaacaaagg R242 gggctgggggccacgctcc R243 cgcaaaagtgaagccctcctgg R244 gaaatcctacttgatctaaagtgagc R245 tttgagcaacttggaaaaaataagcg R246 ttcccaaaagacaaatagcacttcc R247 ccattttgaaaatcacagtgaattcc R248 gaaaagaaaaccctgaattcaaaagg R249 tgctgaaaagaagcatttaaaagtgg R250 ctcttaccagtttcagagctttcc
R251 ttttcagccaaaaatcaaggacagg R252 cttgagcccaggagtttgagacc R253 cgcctgtagtaccctctactagg R254 ggtaaagaaagaaggatttgaaaacc R255 taagagtaatgaggttaaagtttatgc R256 catttttattgtcacaggccatttgc R257 gccacgccttctcttctgccacc R258 tgcctctcctgactgcactgtg R259 ccatgctctaccacgcccttgg R260 cattcaggctggagtgcggtgg R261 cttaaaaattgtctggctaagacattg R262 ttgctcttgttgcccgggttgg R263 gagcttagaggaaaagtattatttcc R264 tggtgctgtgccagacgctgg R265 cagatctttttggctattgtcttgg R266 gaaggaaagggcctcccactgc R267 catgaaaaagcatgctggggagg R268 caaacataaaaaagctttaatagaagcc R269 tcccaactatgaaaaaatagaagacg R270 cacaaattagccgggcatggtgg R271 cttcctttactgagtctttctaaagc R272 tgtcctttgaaatgtaggtatgtgg R273 ggatcttgcaatactgacatctcc R274 atttgaaaagaactgaaggatctacc R275 gtgagctgagatctcgtctctgc R276 tttgtctgaaacagattctaaaagttgg R277 gcaggtgcctgtagtcccagc R278 gtttgagcttctaaaattcatggattc R279 gtggtaggtcaaaccgcaattcc R280 accaaatcagacatatcagctttgg R281 cacagaacggatcctcaataaagg R282 gttaactcctcccttctctttatgg
282 Reverse ABL Primers Used for the Second PCR Round and the Tag Sequence which was on the 5' End of Each Primer
TABLE-US-00003 2nd Round Tag D gtgttcagagagcttgatttccagg RN1 cccacttgatttttcccacatgg RN2 atttatttagatgaagtgaatattttcc RN3 atttagtttgtttaactgtgagtgc RN4 gtacagaagtgcttgatgcatacc RN5 aggcagataaaaattctccattagc RN6 acaagcacgagccacagcacc RN7 cgctcttgttgcccaggctgg RN8 cccaaaacagactttctagataacc RN9 ttcaaattgctttttttctactcacc RN10 gatctgaaaaaagtgacaggttgg RN11 cactgaaatttgaaaggaacatatgg RN12 tctggtgcagtggcctctagg RN13 accataagtggttttacctgatgg RN14 cccaggcgcaggtgattctcc RN15 ggtggctcacgcctgaaatcc RN16 cacagtccacgtgccacaatcc RN17 aatcatgttaacacatccctctcc RN18 gaagagagtgttgaaaggttaagc RN19 cgagaccatactggctaagatgg RN20 attagccacacaataaatgttctgg RN21 tttgaaaagcgttgcaatatgatgc RN22 ggttgcagtgagccgagatcg RN23 ggtgggaggactgcctgagc RN24 aacagagagaaaaaacacaaattacc RN25 gatatctagaattcccaaatacttgg RN26 gtgatagaattaaaggaaaaaataaacg RN27 attgttccttttctaaatattctacc RN28 cagcactttgggaggctgagg RN29 cacagaggtttcacagtgctgg RN30 aacttctgcttctgtccataatgc RN31 gcctgtaatcccagcactttgg RN32 gccagtaaacatatgaaaaggtgc RN33 aattatgtaaataaagagtgaaaagg RN34 cccctacacagaaaaaacaattcc RN35 tgagtgtcaaagaaaaatacaattgg RN36 atacacagagaaaatgagtccacc RN37 aacactccccttctctgtttagc RN38 gatattctttgcaacctaggatgc RN39 ctctaaaactaatcagcaatgtaacc RN40 cacctgtaatcccagcactttgg RN41 cgtaaaactgccacaaagcttgtagg RN42 gtggcagaggtgcaagcaagc RN43 acagaaatgacaaacgcatgtacc RN44 acactctcttagctaggctttgg RN45 gagcttggaatagggcagttcc RN46 ctgggttctttaaacatgtccagg RN47 tcaagaaaggacactgcagtggc RN48 catgcacacaaactatctcattcc RN49 tagccgggcatggtggcacg RN50 atcatgctgattgaatttcaaatagc RN51 ttggcatgcagggcagtgacc RN52 ggtggtgagataataacacctgc RN53 ttgctatataataatcatttgtgatcc RN54 cggtaactgttactctgggatgg RN55 aggctaggttcccttctcttcc RN56 gtagtgcctagcacagagaaagc RN57 ctagcctgggcaacaagagcg RN58 tctctctcctctctgggatcag RN59 gtttgaatatttgtatgcagcaagc RN60 tagaacaaattctggcttataaaagc RN61 ccactctacctttattccttgcc RN62 agaccagaatatgcaagcagagg RN63 ggacgttttgctggtgtctgcg RN64 aaggaacaaactgttgtcacatgc RN65 atgtagctgggactacaggtgc RN66 ggctcatgcctgtaatcccagc RN67 atgaggttttcacacaaaaagatgc RN68 tgggcgacagagcaagactcc RN69 aaatgtccctaaaagtgatcaacagc RN70 cagactcagttttacctcatcagc RN71 agtgatctttcctctttaacctcc RN72 ccagctattcaggaggccaagg RN73 cttaaacattatgacactgtcttgc RN74 ccaggtctatgaggccgttcc RN75 tccaaagcatccctacattatacc RN76 acatacatacatgcagtgactagc RN77 tacaggtgccagccaccatgc RN78 gcctgtaatcccagcactctgg RN79 gacagagtcccactcttgttgc RN80 gtgccttccaaagcagtgtagg RN81 tatcttactgggtatgtataatgcc RN82 caaaggaaatacgtcctaccagg RN83 ccttttctcacagacatgcttcc RN84 taaacacagtgagcagaatccc RN85 ataaagcaaacttctaaaagggtcc RN86 accactacactccagcctggg RN87 gatacctgggtcagagtaagtgc RN88 tgtaatctcagctacttgggagg RN89 gtgtcgtcttctcttcctctacg RN90 ctggctagtatgaggttggtgc RN91 ggactagccacatttcaaccagg RN92 gcagtatactgagaatttagtttcc RN93 gaggctgaggcaggagaatgg RN94 cattgtttgatgaaggtcaacagc RN95 cagacaagagtggctacggcag RN96 acgcccagccagattattcagg RN97 ggaaccagaaagaagtgcaaagg RN98 tgagccatcttggaggcaggc RN99 caggaccttcctacaaacctcc RN100 aacacaacatatctgaccttacgc RN101 gccttagaagtccagaggaaagc RN102 tgacgtacccagtagaccttcc RN103 ctctgcaagcctgggaaacagg RN104 gccttgtccccaagtcctaagg RN105 gcaaagggactcctggaattcc RN106 gctcctgcctgtaatcccagc RN107 gaaggaaacagaaaaagcagaggc RN108 cttactaccgttcttcttcactgg RN109 actattctgtttctttaggtttactgc RN110 cggtggctcacacctgtaatcc RN111 agccagagttctgtgctctagg RN112 taatttgcatttcgtgccgctcc RN113 cacttttaatacagatcccaatagg RN114 atgtattttttcttttcctgtcaagc RN115 aaatgttaacattattctccctaagg RN116 catatgcccagatcccgtctcc RN117 acaggtgtgagccgctgcacc RN118 gccaagacgtttacagttttggc RN119 aggaaacttctgaggatgatggg RN120 gctttatagggcagtctgaattcc RN121 ttagaataaaagttatctcgggagg RN122 taatttcttcagctttatccctcag RN123 cacatgactaattctctattcattcc RN124 aaagacctcaagaaaagagtcacc
RN125 gacccataaagattatatgcccag RN126 aaagtactaatgcagtgtgtcagc RN127 gaggttcctcgattcccctgc RN128 ggagagcagaggaattcacagg RN129 agtaattagaaactgattctaagacg RN130 cataccattgccaatccagttcc RN131 attacgggtgcctgccactgc RN132 cagccaggcagaggagagagg RN133 ttttcattccaagtttctgtttggg RN134 tttcaaataggaatttggataatccc RN135 taagccgagatcacaccactgc RN136 ccttcagcgcattatatcttggc RN137 ccatctaatccatcttaaattcacc RN138 gagtggagactgcgccactgc RN139 aatcatgtgccaattaaaccatggc RN140 cccagggaccagaccagacc RN141 ctcactcaccagtgaaaatcagc RN142 ggttgctctcgaactcctgacc RN143 gttcccccagctcctttctgc RN144 agaaagatgtagaagggtccagc RN145 gggaaaaggtgtattatgcaagcg RN146 ctctctcagacctaatgcaaaagc RN147 aactatacatacagtatttgtattagc RN148 aaattaatgcaatccatgatccagg RN149 ctttctccactctaagagaaccc RN150 ttttggtgtgttcatattggctgc RN151 gcttccacaaatgacagacaaagg RN152 ggctcatgcttgtaatcccagc RN153 catatgaattgttgttcctttgtagg RN154 cactggtacaagtccaagagtcc RN155 gaccctgtgtctacttcctggg RN156 tatttgaactatctcttgaaatgtcc RN157 ctgattaaaaagtattacccttggc RN158 tttgaaactgcactcaataacttgg RN159 agtaatgtgtcatgatccaatggc RN160 gaaagcatttcccaatgtctcacc RN161 caatggacaaaaggcccaactgc RN162 tccagctctggcttttttgttaag RN163 acggagtctcactccgtgacc RN164 ctatgtcatagtcaagagactttgc RN165 gttcaagcgattctcctgtctcg RN166 ccacctaatacttaaatacggaagc RN167 acaacaaacttaatagtgaagtg RN168 ttacaggcgtgagtcaccatgc RN169 aacacctccaagaggccaaacg RN170 tactattggcaaatttcaattatatgg RN171 agcccacatcctaaaattcaataag RN172 gaaagtggataagtgtttgtctgg RN173 ggccaggcattcaagaccagc RN174 agccaacaacaaaaagacacaacc RN175 ttgagcccaggagttcaagacc RN176 cagactaaagatctcagagagaaac RN177 cgcttgtaatcccagcacttgg RN178 aaaagtgaaatcagaatttgtttcc RN179 caggcgtgagcaactgtgtcc RN180 ggtccagtaggatctcgtttgc RN181 actttgaaaatgttgttatagctggg RN182 ttccctgcatctaagtcttctcc RN183 agatatctaccattgaagagtttgc RN184 agtcttcacttcactttgttgtcc RN185 ccatgcaggtatgaaatataaaagc RN186 tgggtgacagagtgagactcc RN187 acagcaataccgggttaacatgc RN188 tttatgtaaaagatgaatgcgaggc RN189 ctactctgctactgggaacagg RN190 caaacgttagtctggcaaaatgcg RN191 tgcacgctaccacacccagc RN192 aattcttggatctgtgtgtttactgc RN193 taccagttatcattctctttctgc RN194 atccacccacctcggcctcc RN195 cactctgcctggcccttaatgg RN196 atagtttgtttaatatgccactaagg RN197 gcgtgagccaccgcacctgg RN198 ctccatcacacaaattttatgtggc RN199 agacggagtctcgttctgtcgc RN200 tcccaggttcaagccattctcc RN201 tattttgagagtctcactctgtcg RN202 gtctcgaactcctgacctcagg RN203 aaggaggtgaagagtgaactacg RN204 gtctcaggttttggacttacttgg RN205 tttacagatcttaaatgcattaggac RN206 gtacactgaacaaaggagacagg RN207 ctggtagtaatgcaaaatagcacc RN208 catttaatgtgaaatgaattataagcc RN209 gagacagggtttcactatgttgg RN210 ccagcactttggaaggctgagg RN211 gaaaccaagtatcatggtaaattgc RN212 cagtgagggctgctcagttcc RN213 gccaggtgcggtggctcacg RN214 catgcctgtaatcccagctacc RN215 atgtaaatggtacagtcactttagg RN216 cccacaatacagagaactcttacc RN217 tgaaacatgcagcccagtgtcc RN218 tgttttttctcctgccttcaatcc RN219 gctttcctgggtctccatctgg RN220 gcagccgcttgaaaacaaaacagc RN221 gatcacgttacatttgggggtgg RN222 taggctgaaaaactaaaatttgttgc RN223 ctcctttgggctcctttagtcc RN224 gcctcggcctcccaaagtgc RN225 aatgcctagagagatttggcagg RN226 gagatggggtttcactatgttgg RN227 tgtgatcttgccactgcactcc RN228 acttctcctccatttgtttcttcg RN229 cgtgcccgggctcagttctac RN230 ccaaaacaataaaatcacaatttggg RN231 ctgaactgccttagagtaaatccg RN232 atttctgtatcaggtctgtgttcc RN233 ggctgaccccttcactgtttcc RN234 caaaaattagccaggcatggtgg RN235 gcagtgagcagtgatcgcacc RN236 aaagactgtgaactaacttgtttgc RN237'1 tgccaagaattacacattattaggc RN238 ggccaggatgtcattaactttcc RN239 gtaagagctgacgtgtattgtgc RN240 cccggtgaggccgcacatcc RN241 cctgcgccttaaccccctcc RN242 cggcgcctaggggccatcg RN243 acttaaggaaacgaacatgacacc RN244 gagaccgagtcttgctgtgtcg RN245 gtattaattgaagatgatttggaatgc RN246 tctttaaaagactatcgctgaggc RN247 aaaagagacatcagtagagcatcc RN248 gttcatgttttctttgacgtctcc RN249 tttcgaaagttcaggctgagtgc
RN250 gaccctcaaaacaatcctctaagg RN251 caaaacacacttagaaacaaactgc RN252 gcctgggcgacatagtgagacc RN253 ggcaggagaatggcgtgaacc RN254 tttgctcgttgcccaggctgg RN255 gcaacttaatgtgatagaataatagc RN256 cctccccttctgctgccagc RN257 ccacaacaatgtaaactcctctgg RN258 tactctccctagagttcgttccc RN259 gggtccccctttggccattcc RN260 gatcttggctcacttcaacctcc RN261 aggggaaatatttaaaccttgg RN262 aatgcaatggtgcatttacagagg RN263 tcattttatctatttctacatggtcc RN264 ggaagggaaatgcccatgaacc RN265 agtgaacattttctgcagcctcc RN266 caacaggacgtcaggcgatcc RN267 ccttcaggctgtcctgaaaagg RN268 agtctcactccatcgcccagg RN269 actgtgaacagtagttaactcagg RN270 gcatgcctgtaatccaagctgc RN271 gaaacaattctcttttcacacttgc RN272 ggctcatgcctgttatcccagc RN273 agaagaagcttagtcatatgtttgg RN274 cagatgcttgagccaaacaaatgg RN275 ctggcagacagagtgagactcc RN276 aatgtgtgaatattattcattacaggg RN277 gcaggagaattgcttgaacctgg RN278 ctttagtcaaattaaaacagtctatcc RN279 gatttctatctcctgcaaccacc RN280 ttcttgtgtaactactaaaaatctcc RN281 aaagggtcttcataaggctaatgg RN282 ctcttaaggattatttatatgaagacc
Example 3
Identification of the PML-RARalpha Breakpoint
[0254]Amplified patient DNA was electrophoresed on a 2% agarose gel. P is patient DNA, N is the normal DNA and W is the water control. The patient DNA was amplified using multiple RARα primers and a single PML primer [0255]a) Amplified patient DNA electrophoresed on a 2% agarose gel, P is patient DNA, N is the normal DNA and W is the water control. The patient DNA was amplified for one round using an RARα primer and a PML primer designed using the breakpoint sequence. [0256]b) The sequence chromatogram obtained from the patient DNA. The breakpoint between PML and RARα is shown.
Isolation of the PML-RARalpha Breakpoint in Acute Promyelocytic Leukemia
[0257]Two patients have been studied and the breakpoint has been isolated and sequenced in both. The primers used are shown in Example 4.
Example 4
Primers Used for Isolation of PML-RARalpha Translocation Breakpoint in Acute Promyelocytic Leukemia
PML Forward Primers
TABLE-US-00004 [0258] 1st Rd PML F1-FT1 caggaggagccccagagc PML F2-FT1 tcctggggatggttggatgc PML F3-FT1 tgaccccacagagtttacacagc PML F4-FT1 agtcagggcaggctctgcc PML F5-FT1 tattttggccccatccagaaagc PML F6-FT1 cacccagagtacagctttgttcc 2nd Rd PML F1-FT2 gaggagccccagagcctgc PML F2-FT2 tggggatggttggatgcttacc PML F3-FT2 cccacagagtttacacagcttgc PML F4-FT2 caggctctgcccactcacc PML F5-FT2 ccatccagaaagcccaaagcc PML F6-FT2 ccagagtacagctttgttcctcattc
[0259]The second round primers were internal to the first round primers and were used for performing Bottleneck PCR in order to eliminate non-specific amplified material and facilitate isolation of the translocation breakpoint.
[0260]Various combinations of the forward and reverse primers can be used. 2 exemplary protocols were either to set up 6 PCRs, each containing a different PML primer and all 34 RARalpha primers, or to set up 1 PCR which contained all 6 forward and all 34 reverse primers.
34 Reverse RARalpha Reverse Primers Used for the First PCR Round and the Tag Sequence which was on the 5' End of Each Primer
TABLE-US-00005 Tag R1 gcagtacaaacaacgcacagcg RAR1 ctgccaccctccacagtccc RAR2 gccaagaccatgcatgcg RAR3 cccagggacaaagagactccc RAR4 caggaagcagacagtcttctagttcc RAR5 tgcctgtaatcccaacactttgg RAR6 tccctctggccaggatggg RAR7 atggggaatgggagtaggaagc RAR8 cagatcagttctcccctccagc RAR9 acaaaaaagaaacatgctcagagagg RAR10 tggtggcatgcatctgtagtcc RAR11 aggtgctctatagatgttagcatccc RAR12 ccaggacaggatggagatctgg RAR13 agggaacctgtgcattatccttgc RAR14 cagaagtcttgctttaaggaggagg RAR15 gggtacgtgaaactcaccaagg RAR16 cagagtgtggcaagcaaggg RAR17 aacattttaaaggtacaaataacgtggg RAR18 tagggagcaacagccattaagc RAR19 ggtgcactgtccagctctgg RAR20 actctcgctgaactcgcctgg RAR21 ctcggtctctggtggtacgc RAR22 gcaagaggtccgagctggg RAR23 ggaagaagtgaaacaagagatgaagg RAR24 cccagagaacaaaccggattagg RAR25 cccttcaaccttctccaatctgc RAR26 cccatgtccagtggtttaggg RAR27 gagattggtgggagacagatgg RAR28 cttctcagctcaaagttccagcg RAR29 gaatgggagagatgaccagagg RAR30 aagggcaagggggtatgtgg RAR31 ggaaggaagcatgggaacacc RAR32 ccatcaatgctctgtctgtctgg RAR33 gtgccgtgactgtgcttgg RAR34 acatcccattgacctcatcaagc
[0261]Nearly all translocations involve a 3 kb region of the BCR gene and 140 kb region of the ABL gene. Six forward primers used to cover the region of the BCR gene and 282 primers used to cover the region of the ABL gene. Six PCRs are set up, each containing one of the BCR primers, all of the ABL primers, and the common tag primer.
[0262]If necessary, a second round of PCR is performed with a nested internal BCR primer and 282 nested internal ABL primers Alternatively, 1-3 rounds of Bottleneck PCR are performed in order to remove non-specific amplified products and reveal the amplified translocation sequence.
[0263]The ABL gene is very rich in Alu sequences, and the BCR gene also contains one such sequence. The ABL primers have therefore undergone a selection procedure which sequentially involves, for each ABL primer: [0264]design using standard criteria [0265]pairing with each BCR primer and testing by electronic PCR for amplification off the BCR template. Primers that fail this criterion are discarded. [0266]incorporation in a pool of 12 or 24 ABL primers, pairing the pool with each BCR primer, and testing by experimental PCR using a BCR template which has been previously produced by PCR amplification. Any pool that that produces amplification and thus fails this test is further analysed by testing each of the individual ABL primers to determine which is responsible for amplification. When identified, this primer is discarded.
[0267]The BCR and ABL primers used in Example 1 are shown in Example 2.
[0268]Those skilled in the art will appreciate that the invention described herein is susceptible to variations and modifications other than those specifically described. It is to be understood that the invention includes all such variations and modifications. The invention also includes all of the steps, features, compositions and compounds referred to or indicated in this specification, individually or collectively, and any and all combinations of any two or more of said steps or features.
TABLE-US-00006 TABLE 1 Sequence Identifier Sequence SEQ ID NO: 001 gtgggcccccccgtttccgtgtacagggcacctgca gggagggcaggcagctagcctgaaggctgatccccc cttcctgttagcacttttgatgggactagtggactt tggttcagaaggaagagctatgcttgttagggcctc ttgtctcctcccaggagtggacaaggtgggttagga gcagtttctccctgagtggctgc SEQ ID NO: 002 caccacgtctggctaatttttgtatttttagtagag atggggtttcaacatgttagccaggctggtctcgaa ctcctgacctcaggtgatccacccgcctgggccctc caaagtgctgggattacaggcaggagccactgtgcc cggcctgacctcatatttgaataccgagttttagtt ctggaggagctgcaggttttatgaaaagggaacaca tttgattcctcagagcagccacaggccagctctctg aagtaaagtgcacgtgtgcatgtgtgtgcacactca cacacacgtacacacacattcacaaataactgtgcc cggcctgacctcatatttgaataccgagttttagtt ctggaggagctgcagg SEQ ID NO: 003 tttgggaggctgaggcaggtggatcgcttgagctca ggagttggagaccagcctgaccaacatggtgaaacc ctgtgtctactaaaaatacaaagattagccgggcta ggcagtgggcacctgtaatcacaactgcttgggagg ctgagggaagagaatcgcttgaacccaggaggcgga ggttgcagtgagccgagcttgtgccactgcattcca gcctgggcgacagag SEQ ID NO: 004 ggtctcactctgttgaactcctggtggcctcaaggg atcctcctacctcggcctcacaaagtattggaatta caggtgtgagtcactgcagctggccttcacttatca ctgtgaggagtaaacagctgcatggtgggcttaatg ccatctaacacgagtgactccatgttcagacagtag gatcacaaatgattattatatagcaatgaatggcca caggtacatagactaaggagccacatccctgct SEQ ID NO: 005 cctccagctacctgccagccggcacttttggtcaag ctgttttgcattcactgttgcacatatgctcagtca cacacacagcatacgctatgcacatgtgtccacaca caccccacccacatcccacatcaccccgaccccctc tgctgtccttggaaccttattacacttcgagtcact ggtttgcctgtattgtgaaaccagctggatcc SEQ ID NO: 006 ttatttataacaacattttcagcgtggcaactgcag tttcagaatggtggaattataccagtcagagagaga tgcaaatgatttaaaataggaagaaagcaggtgtct ggcccagaggaccagattaagaagaccccatgagag ttacaatagttagtgaaaatggtgcttctgcaaacc tcatgtctacagaagctggt SEQ ID NO: 007 tgcaccttcataacataatctttctcctgggcccct gtctctggctgcctcataaacgctggtgtttccctc gtgggcctccctgcatccctgcatctcctcccgggt cctgtctgtgagcaatacagcgtgacaccctacgct gccccgtggtcccgggcttgtctctccttgcctccc tgttacctttctttctatctcttccttgccccg SEQ ID NO: 008 gtgagctccgcctcctgtcagatcagtggcggcatt agtttctcataggagcatgaaatctattgtgaacag tacatgcgatggatccaggttgcgtgctcctagtga gaatctaatgcctgaggatctctcattgtctcttat cactcccagataggactgtctagttgcaggaaaaca agctcagggctcccactgattctacattacagtggg ttgtataattattatatattacaatgtaataataa SEQ ID NO: 009 ggagtctgaggaggggaaggaggcaaggttggctcg gatcccagccagtaagtctgggtgtgg SEQ ID NO: 010 cttctccctgacatccgtgg SEQ ID NO: 011 acacagcatacgctatgcacatgtg SEQ ID NO: 012 gaggttgttcagatgaccacgg SEQ ID NO: 013 cagctactggagctgtcagaacag SEQ ID NO: 014 tgggcctccctgcatcc SEQ ID NO: 015 tccccctgcaccccacg SEQ ID NO: 016 tgacatccgtggagctgcagatgc SEQ ID NO: 017 acatgtgtccacacacaccccacc SEQ ID NO: 018 accacgggacacctttgaccctgg SEQ ID NO: 019 ctggagctgtcagaacagtgaagg SEQ ID NO: 020 tccctgcatccctgcatctcctcc SEQ ID NO: 021 cccacgacttctccagcactgagc SEQ ID NO: 022 gcaacactgtgacgtactggagg SEQ ID NO: 023 gtctatctaaaattcacaaggaatgc SEQ ID NO: 024 aggcaaagtaaaatccaagcaccc SEQ ID NO: 025 cactcctgcactccagcctgg SEQ ID NO: 026 caaccaccaaagtgcttttcctgg SEQ ID NO: 027 atatggcatctgtaaatattaccacc SEQ ID NO: 028 tgcctcggcctcccaaagtgc SEQ ID NO: 029 agccaccacacccagccagg SEQ ID NO: 030 aataactgttttctccccccaaaac SEQ ID NO: 031 tgttttacaaaaatggggccatacc SEQ ID NO: 032 acttaagcaaattctttcataaaaaggg SEQ ID NO: 033 ctttcaattgttgtaccaactctcc SEQ ID NO: 034 acctcctgcatctctccttttgc SEQ ID NO: 035 aaataaagttttgagaaccataagtgg SEQ ID NO: 036 caccatcacagctcactgcagc SEQ ID NO: 037 aacctctttgagaatcggatagcc SEQ ID NO: 038 aaataaagtacatacctccaattttgc SEQ ID NO: 039 gacacattcctatgggtttaattcc SEQ ID NO: 040 tgtaaaatatggtttcagaagggagg SEQ ID NO: 041 gcaggtggataacgaggtcagg SEQ ID NO: 042 ccagccaagaatttcaaagattagc SEQ ID NO: 043 gaagggagatgacaaagggaacg SEQ ID NO: 044 gcagaagaactgcttgaacctgg SEQ ID NO: 045 gtggtcccagctactcgagagg SEQ ID NO: 046 ccctcagcaaaactaactgaaaagg SEQ ID NO: 047 tagaaaccaagatatctagaattccc SEQ ID NO: 048 ccacgcccggcggaataaatgc SEQ ID NO: 049 acaaaaaaagaggcaaaaactgagag SEQ ID NO: 050 ctgggcgcagtggctcatgcc SEQ ID NO: 051 tggctgtgaggctgagaactgc SEQ ID NO: 052 ctgggcgacagagtgagactcc SEQ ID NO: 053 aagtctggctgggcgcagtgg SEQ ID NO: 054 aatggacaaaagaggtgaactggc SEQ ID NO: 055 gatagagtgaaaacgcacaatggc SEQ ID NO: 056 aattaaacagctaggtcaatatgagg SEQ ID NO: 057 ggtctccactatcaagggacaag SEQ ID NO: 058 aagcagctgttagtcatttccagg SEQ ID NO: 059 aggcatcctcagattatggctcc SEQ ID NO: 060 cctgagtaacactgagaccctgc SEQ ID NO: 061 aacactcaagctgtcaagagacac SEQ ID NO: 062 attcaggccaggcgcagtggc SEQ ID NO: 063 taaatcgtaaaactgccacaaagc SEQ ID NO: 064 cagaggagtaggagaaggaaaagg SEQ ID NO: 065 ggtagctatctaccaagtagaatcc SEQ ID NO: 066 atcagattggaaaaagtcccaaagc SEQ ID NO: 067 ctcctgaaaagcacctactcagc SEQ ID NO: 068 ctccttaaacctgaggtactggg SEQ ID NO: 069 ttttctcctaatagaccaccattcc SEQ ID NO: 070 ctgctgtattaccatcactcatgtc SEQ ID NO: 071 ctggccaacatagtgaaaccacg SEQ ID NO: 072 atttgaataggggttaaagtatcattg SEQ ID NO: 073 cacttcagtggaagttggcatgc SEQ ID NO: 074 gtttttcttcgaagtgataaacatacg SEQ ID NO: 075 gctccttagtctatgtacctgtgg SEQ ID NO: 076 tactctggcatggtaactggtgc SEQ ID NO: 077 acaaaggactaggtctgtggagc SEQ ID NO: 078 ccaagtttaccaaattaccaaagttacc SEQ ID NO: 079 tgagccgatatcacgccactgc SEQ ID NO: 080 tcccaataaaggttttggcccagg SEQ ID NO: 081 ctgggtagcaaattagggaacagg SEQ ID NO: 082 ctggccagaaaagacagttttatcc SEQ ID NO: 083 ggttcccaggaagggataacacc SEQ ID NO: 084 tcactccaggaggttccatttcc SEQ ID NO: 085 aggcttggaaataagcagcagtgg SEQ ID NO: 086 attcatacaatggaatactactcagc SEQ ID NO: 087 taagtgatcctcccacctcaacc SEQ ID NO: 088 tataagaggaagactggggctgg SEQ ID NO: 089 tcatacttatgcaggttataggagg SEQ ID NO: 090 caagatcacgccactgcactcc SEQ ID NO: 091 aaaataaatagctggtgctcaagatc SEQ ID NO: 092 caccagcctcattcaacagatgg SEQ ID NO: 093 caatgcagcctcaacctcctgg SEQ ID NO: 094 gttaggtcaggtgctcatgtctg SEQ ID NO: 095 aagtttcaaaaggacatgtacaaaatg SEQ ID NO: 096 tcctgaagaggctgcagcttcc SEQ ID NO: 097 ctggtgcacattcccaagtgtgc SEQ ID NO: 098 catgttggccatgttcttctgagg
SEQ ID NO: 099 ctcagcctcccgagtagctgg SEQ ID NO: 100 aaagacatttaagaggagatgaggc SEQ ID NO: 101 tgctgggattacaggcgtgagc SEQ ID NO: 102 tgtgacttccatccgcagctcc SEQ ID NO: 103 gacacttttgtggagctttcatgg SEQ ID NO: 104 catgtgagggggcacgtcttgc SEQ ID NO: 105 tcttctctatgagaaaagtggttgc SEQ ID NO: 106 tggcaaaatgctatcgagctgcc SEQ ID NO: 107 tatgaacacagccggcctcagg SEQ ID NO: 108 gaggttgcagtgagctgagatcg SEQ ID NO: 109 gtcaagcacccagtccgatacc SEQ ID NO: 110 atctgggcttggtggcgcacg SEQ ID NO: 111 gttaagcgggtcccacatcagc SEQ ID NO: 112 cagccagtttcagtagaaagatgc SEQ ID NO: 113 gacccaagcataaggggactagc SEQ ID NO: 114 cccaaaaagtttacaagagaaattttc SEQ ID NO: 115 cgcctgtagtcccagctactcg SEQ ID NO: 116 cgcgtgatgcggaaaagaaatcc SEQ ID NO: 117 tctactatgaaccctccttcagac SEQ ID NO: 118 gtgctgggattacaggtgtgagc SEQ ID NO: 119 ttatccaaatgtcccagggcagg SEQ ID NO: 120 ctgccagcactgctcgccagc SEQ ID NO: 121 gctactgcaggcagtgccttcc SEQ ID NO: 122 catccaagcccaaggtgtcagg SEQ ID NO: 123 tgtttgcatgtaatttcaggaagcc SEQ ID NO: 124 gatccgtcactgttaacactcagg SEQ ID NO: 125 ctcacagtcacaagctcctgagc SEQ ID NO: 126 gagatgatgctggggtcacagg SEQ ID NO: 127 ttagaagaatgggatcgcaaagg SEQ ID NO: 128 cggtattcaaatatgaggtcaggc SEQ ID NO: 129 gtaaatcctgctgccagtcttcc SEQ ID NO: 130 acagggtcagacagagccttgg SEQ ID NO: 131 agttattgatctaactatacaacaagc SEQ ID NO: 132 aaagactaggggccggggacg SEQ ID NO: 133 ctggtagaaataaagacaacaaagcc SEQ ID NO: 134 gtgccaagtaattaaaagtttgaaacc SEQ ID NO: 135 ggcttttgaagggagcaccacc SEQ ID NO: 136 gaaggataaatacctatgatactttcc SEQ ID NO: 137 ggcagggaaatactgtgcttcaag SEQ ID NO: 138 gtggtgaaattccacctcagtacc SEQ ID NO: 139 tcccaaagtgctgggattacagg SEQ ID NO: 140 gaaattagcaaacaatgccaagacg SEQ ID NO: 141 taagtattggaccgggaaggagg SEQ ID NO: 142 ctatcattttgctcaaagtgtagcc SEQ ID NO: 143 atttcacaaactacagaggccagg SEQ ID NO: 144 tagacttctgtctctctatgctgc SEQ ID NO: 145 tgagtgagctgccatgtgatacc SEQ ID NO: 146 acttcacaccagcctgtccacc SEQ ID NO: 147 taactcatatcctcagagagaccc SEQ ID NO: 148 agaggttcctcgattcccctgc SEQ ID NO: 149 gtgtcagcgtcccaacacaaagc SEQ ID NO: 150 gaaagtggatgggcaagcattgc SEQ ID NO: 151 gtgatcacctcacagctgcagg SEQ ID NO: 152 gtttgtttagtcaaggcatttcacc SEQ ID NO: 153 cctcagcctccagagtagctgg SEQ ID NO: 154 taaaagaaaactcctccttcctgg SEQ ID NO: 155 aatgtgctatgtctttaaatccatgg SEQ ID NO: 156 agctggcaaatctggtaatataaaag SEQ ID NO: 157 gcttgaacctggaaggtggagg SEQ ID NO: 158 gcaggcatgctaagaccttcagc SEQ ID NO: 159 cagctccatgaataactccacagg SEQ ID NO: 160 gcttgaacccaggaggcagagg SEQ ID NO: 161 atcgaagatgccactgcaagagg SEQ ID NO: 162 ccaaccacacttcaggggatacc SEQ ID NO: 163 cacgccagtccactgatactcac SEQ ID NO: 164 gggtttcaccatgttggccagg SEQ ID NO: 165 cccaacaaaggctctggcctgg SEQ ID NO: 166 atgacagcagaggagcttcatcc SEQ ID NO: 167 gcaggctacgagtaaaaggatgg SEQ ID NO: 168 cgggtaaaatcttgcctccttcc SEQ ID NO: 169 aaacttaaaccaatggtggatgtgg SEQ ID NO: 170 agagactgaggaactgttccagc SEQ ID NO: 171 gaaacggtcttggatcactgatcc SEQ ID NO: 172 tgcgcatgatatcttgtttcaggg SEQ ID NO: 173 ggcctccgtttaaactgttgtgc SEQ ID NO: 174 gaatgctggcccgacacagtgg SEQ ID NO: 175 tcttggtatagaaaagccagctgg SEQ ID NO: 176 gcaaaagcccaagagcccctgg SEQ ID NO: 177 ttctcccaaaatgagccccaagg SEQ ID NO: 178 gtggtgacgtaaacaaaaggtacc SEQ ID NO: 179 gcaaattccatgtgaatcttattggc SEQ ID NO: 180 cctgatctatggaacagtggtgg SEQ ID NO: 181 gttacaaacgttgcagtttgcaacg SEQ ID NO: 182 gaaccccgtcaacagtgatcacc SEQ ID NO: 183 acaggacctcaaggcaaggagc SEQ ID NO: 184 catacctaaaatagaaatgtctatccc SEQ ID NO: 185 gagttgcatatatgttttataaatccc SEQ ID NO: 186 tgagcccacatccataaagttagc SEQ ID NO: 187 accgcaacctttgccgcctgg SEQ ID NO: 188 taaatattttgtatggagtcaccacc SEQ ID NO: 189 aaagccaggagaaaaagttatgagg SEQ ID NO: 190 tcccaaagtcccaggattacagg SEQ ID NO: 191 tcactatggagcatctccgatgg SEQ ID NO: 192 agttccctggaagtctccgagg SEQ ID NO: 193 aaaataatcacccagcccacatcc SEQ ID NO: 194 acaaaactacagacacagaaagtgg SEQ ID NO: 195 tttgggaggctgaggtaggtgg SEQ ID NO: 196 aaagacagtgaaacatctataaggg SEQ ID NO: 197 cattttgggagaccagggcagg SEQ ID NO: 198 gcatgggacagacacaaagcagc SEQ ID NO: 199 gaataacaaagagagccggctgg SEQ ID NO: 200 taaaccttttattgaaaattgtcaaatgg SEQ ID NO: 201 cgcctcagcctcccaaagtgc SEQ ID NO: 202 tacattagttttataggtccagtagg SEQ ID NO: 203 gaaggtttattcatattaaaatgtgcc SEQ ID NO: 204 ctggcttctgtggtttgagttgg SEQ ID NO: 205 acagacctacctcctaaggatgg SEQ ID NO: 206 gctagcttttgtgtgtaagaatggg SEQ ID NO: 207 ggcctactcacacaatagaatacc SEQ ID NO: 208 gcaccattgcactccagcctgg SEQ ID NO: 209 gaaattaggataaaggttgtcacagc SEQ ID NO: 210 cagaagtgttcaaggtgaaactgtc SEQ ID NO: 211 ctgaatcatgaaatgttctactctgc SEQ ID NO: 212 tgtcaacttgactgggccatacg SEQ ID NO: 213 ctcccgtatagttgggattatagg SEQ ID NO: 214 gcttggagttccttgaaattcttgg SEQ ID NO: 215 cctggtggctccagttttctacc SEQ ID NO: 216 aactcctgacctcatgatccacc SEQ ID NO: 217 gctgggattacaggcatgagcc SEQ ID NO: 218 ttctcctttatccttggtgacattc SEQ ID NO: 219 tcccaaagtgctgggattacagg SEQ ID NO: 220 gtcataagtcagggaccatctgc SEQ ID NO: 221 ctgtttcattgatttccagactggc SEQ ID NO: 222 gcaatctcggctcactgcaagc SEQ ID NO: 223 gaagaagtgactatatcagatctgg SEQ ID NO: 224 ttcaccatgttggccaggctgg
SEQ ID NO: 225 catcactgaagatgacaactgagc SEQ ID NO: 226 gtccagcctgggcgatagagc SEQ ID NO: 227 gaggaaagtctttgaagaggaacc SEQ ID NO: 228 ggtacactcaccagcagttttgc SEQ ID NO: 229 gagcaactggtgtgaatacatatgg SEQ ID NO: 230 caatacctggcaccacatacacc SEQ ID NO: 231 gggactacaggcatgtgccacc SEQ ID NO: 232 cggtggctcacgcgtgtaatcc SEQ ID NO: 233 caactgttaaatctctcatggaaacc SEQ ID NO: 234 gacaaaggattagaaatgcaccc SEQ ID NO: 235 ggaaatgttctaaaactggattgtgg SEQ ID NO: 236 aataataatagccaggtgtggtagc SEQ ID NO: 237 ctggaacactcacacattgctgg SEQ ID NO: 238 ctgggtgacagagcgagactcc SEQ ID NO: 239 cccaaatcatccccgtgaaacatgc SEQ ID NO: 240 gaccctgcaatcccaacactgg SEQ ID NO: 241 ctctcaggccttcaaactacacc SEQ ID NO: 242 caggaaagggctcgctcagtgg SEQ ID NO: 243 atctgcaaaagcagcagagcagg SEQ ID NO: 244 gtacccatgacagacaagttttagg SEQ ID NO: 245 cttatcccctactgtctcctttgg SEQ ID NO: 246 ggatggtctcgatctcctgacc SEQ ID NO: 247 aggttagagaccttcctctaatgc SEQ ID NO: 248 agctgggattacaggtgcctgc SEQ ID NO: 249 gctgaggcaggttggggctgc SEQ ID NO: 250 acatttaacgtctcctaacttctcc SEQ ID NO: 251 gtgctgcgattacaggtgtgagc SEQ ID NO: 252 tatgacagcagtattatactatcacc SEQ ID NO: 253 ctggggaccaaatctgaactgcc SEQ ID NO: 254 gtagctattgttatttccaaaagagg SEQ ID NO: 255 gcttgggaccccaggacaagg SEQ ID NO: 256 cctggccaacatggggaaatcc SEQ ID NO: 257 aattgcttgaacctgggaggtgg SEQ ID NO: 258 gcctaagacccaaaagctattagc SEQ ID NO: 259 catattaaagggccatattcaaattgg SEQ ID NO: 260 ggatgtaaccagtgtatatcacagg SEQ ID NO: 261 ggaagtttagtccacatcttctagc SEQ ID NO: 262 gcacccacaggacaaccacacg SEQ ID NO: 263 gggacgcgcctgttaacaaagg SEQ ID NO: 264 gggctgggggccacgctcc SEQ ID NO: 265 cgcaaaagtgaagccctcctgg SEQ ID NO: 266 gaaatcctacttgatctaaagtgagc SEQ ID NO: 267 tttgagcaacttggaaaaaataagcg SEQ ID NO: 268 ttcccaaaagacaaatagcacttcc SEQ ID NO: 269 ccattttgaaaatcacagtgaattcc SEQ ID NO: 270 gaaaagaaaaccctgaattcaaaagg SEQ ID NO: 271 tgctgaaaagaagcatttaaaagtgg SEQ ID NO: 272 ctcttaccagtttcagagctttcc SEQ ID NO: 273 ttttcagccaaaaatcaaggacagg SEQ ID NO: 274 cttgagcccaggagtttgagacc SEQ ID NO: 275 cgcctgtagtaccctctactagg SEQ ID NO: 276 ggtaaagaaagaaggatttgaaaacc SEQ ID NO: 277 taagagtaatgaggttaaagtttatgc SEQ ID NO: 278 catttttattgtcacaggccatttgc SEQ ID NO: 279 gccacgccttctcttctgccacc SEQ ID NO: 280 tgcctctcctgactgcactgtg SEQ ID NO: 281 ccatgctctaccacgcccttgg SEQ ID NO: 282 cattcaggctggagtgcggtgg SEQ ID NO: 283 cttaaaaattgtctggctaagacattg SEQ ID NO: 284 ttgctcttgttgcccgggttgg SEQ ID NO: 285 gagcttagaggaaaagtattatttcc SEQ ID NO: 286 tggtgctgtgccagacgctgg SEQ ID NO: 287 cagatctttttggctattgtcttgg SEQ ID NO: 288 gaaggaaagggcctcccactgc SEQ ID NO: 289 catgaaaaagcatgctggggagg SEQ ID NO: 290 caaacataaaaaagctttaatagaagcc SEQ ID NO: 291 tcccaactatgaaaaaatagaagacg SEQ ID NO: 292 cacaaattagccgggcatggtgg SEQ ID NO: 293 cttcctttactgagtctttctaaagc SEQ ID NO: 294 tgtcctttgaaatgtaggtatgtgg SEQ ID NO: 295 ggatcttgcaatactgacatctcc SEQ ID NO: 296 atttgaaaagaactgaaggatctacc SEQ ID NO: 297 gtgagctgagatctcgtctctgc SEQ ID NO: 298 tttgtctgaaacagattctaaaagttgg SEQ ID NO: 299 gcaggtgcctgtagtcccagc SEQ ID NO: 300 gtttgagcttctaaaattcatggattc SEQ ID NO: 301 gtggtaggtcaaaccgcaattcc SEQ ID NO: 302 accaaatcagacatatcagctttgg SEQ ID NO: 303 cacagaacggatcctcaataaagg SEQ ID NO: 304 gttaactcctcccttctctttatgg SEQ ID NO: 305 gtgttcagagagcttgatttccagg SEQ ID NO: 306 cccacttgatttttcccacatgg SEQ ID NO: 307 atttatttagatgaagtgaatattttcc SEQ ID NO: 308 atttagtttgtttaactgtgagtgc SEQ ID NO: 309 gtacagaagtgcttgatgcatacc SEQ ID NO: 310 aggcagataaaaattctccattagc SEQ ID NO: 311 acaagcacgagccacagcacc SEQ ID NO: 312 cgctcttgttgcccaggctgg SEQ ID NO: 313 cccaaaacagactttctagataacc SEQ ID NO: 314 ttcaaattgctttttttctactcacc SEQ ID NO: 315 gatctgaaaaaagtgacaggttgg SEQ ID NO: 316 cactgaaatttgaaaggaacatatgg SEQ ID NO: 317 tctggtgcagtggcctctagg SEQ ID NO: 318 accataagtggttttacctgatgg SEQ ID NO: 319 cccaggcgcaggtgattctcc SEQ ID NO: 320 ggtggctcacgcctgaaatcc SEQ ID NO: 321 cacagtccacgtgccacaatcc SEQ ID NO: 322 aatcatgttaacacatccctctcc SEQ ID NO: 323 gaagagagtgttgaaaggttaagc SEQ ID NO: 324 cgagaccatactggctaagatgg SEQ ID NO: 325 attagccacacaataaatgttctgg SEQ ID NO: 326 tttgaaaagcgttgcaatatgatgc SEQ ID NO: 327 ggttgcagtgagccgagatcg SEQ ID NO: 328 ggtgggaggactgcctgagc SEQ ID NO: 329 aacagagagaaaaaacacaaattacc SEQ ID NO: 330 gatatctagaattcccaaatacttgg SEQ ID NO: 331 gtgatagaattaaaggaaaaaataaacg SEQ ID NO: 332 attgttccttttctaaatattctacc SEQ ID NO: 333 cagcactttgggaggctgagg SEQ ID NO: 334 cacagaggtttcacagtgctgg SEQ ID NO: 335 aacttctgcttctgtccataatgc SEQ ID NO: 336 gcctgtaatcccagcactttgg SEQ ID NO: 337 gccagtaaacatatgaaaaggtgc SEQ ID NO: 338 aattatgtaaataaagagtgaaaagg SEQ ID NO: 339 cccctacacagaaaaaacaattcc SEQ ID NO: 340 tgagtgtcaaagaaaaatacaattgg SEQ ID NO: 341 atacacagagaaaatgagtccacc SEQ ID NO: 342 aacactccccttctctgtttagc SEQ ID NO: 343 gatattctttgcaacctaggatgc SEQ ID NO: 344 ctctaaaactaatcagcaatgtaacc SEQ ID NO: 345 cacctgtaatcccagcactttgg SEQ ID NO: 346 cgtaaaactgccacaaagcttgtagg SEQ ID NO: 347 gtggcagaggtgcaagcaagc SEQ ID NO: 348 acagaaatgacaaacgcatgtacc SEQ ID NO: 349 acactctcttagctaggctttgg
SEQ ID NO: 350 gagcttggaatagggcagttcc SEQ ID NO: 351 ctgggttctttaaacatgtccagg SEQ ID NO: 352 tcaagaaaggacactgcagtggc SEQ ID NO: 353 catgcacacaaactatctcattcc SEQ ID NO: 354 tagccgggcatggtggcacg SEQ ID NO: 355 atcatgctgattgaatttcaaatagc SEQ ID NO: 356 ttggcatgcagggcagtgacc SEQ ID NO: 357 ggtggtgagataataacacctgc SEQ ID NO: 358 ttgctatataataatcatttgtgatcc SEQ ID NO: 359 cggtaactgttactctgggatgg SEQ ID NO: 360 aggctaggttcccttctcttcc SEQ ID NO: 361 gtagtgcctagcacagagaaagc SEQ ID NO: 362 ctagcctgggcaacaagagcg SEQ ID NO: 363 tctctctcctctctgggatcag SEQ ID NO: 364 gtttgaatatttgtatgcagcaagc SEQ ID NO: 365 tagaacaaattctggcttataaaagc SEQ ID NO: 366 ccactctacctttattccttgcc SEQ ID NO: 367 agaccagaatatgcaagcagagg SEQ ID NO: 368 ggacgttttgctggtgtctgcg SEQ ID NO: 369 aaggaacaaactgttgtcacatgc SEQ ID NO: 370 atgtagctgggactacaggtgc SEQ ID NO: 371 ggctcatgcctgtaatcccagc SEQ ID NO: 372 atgaggttttcacacaaaaagatgc SEQ ID NO: 373 tgggcgacagagcaagactcc SEQ ID NO: 374 aaatgtccctaaaagtgatcaacagc SEQ ID NO: 375 cagactcagttttacctcatcagc SEQ ID NO: 376 agtgatctttcctctttaacctcc SEQ ID NO: 377 ccagctattcaggaggccaagg SEQ ID NO: 378 cttaaacattatgacactgtcttgc SEQ ID NO: 379 ccaggtctatgaggccgttcc SEQ ID NO: 380 tccaaagcatccctacattatacc SEQ ID NO: 381 acatacatacatgcagtgactagc SEQ ID NO: 382 tacaggtgccagccaccatgc SEQ ID NO: 383 gcctgtaatcccagcactctgg SEQ ID NO: 384 gacagagtcccactcttgttgc SEQ ID NO: 385 gtgccttccaaagcagtgtagg SEQ ID NO: 386 tatcttactgggtatgtataatgcc SEQ ID NO: 387 caaaggaaatacgtcctaccagg SEQ ID NO: 388 ccttttctcacagacatgcttcc SEQ ID NO: 389 taaacacagtgagcagaatccc SEQ ID NO: 390 ataaagcaaacttctaaaagggtcc SEQ ID NO: 391 accactacactccagcctggg SEQ ID NO: 392 gatacctgggtcagagtaagtgc SEQ ID NO: 393 tgtaatctcagctacttgggagg SEQ ID NO: 394 gtgtcgtcttctcttcctctacg SEQ ID NO: 395 ctggctagtatgaggttggtgc SEQ ID NO: 396 ggactagccacatttcaaccagg SEQ ID NO: 397 gcagtatactgagaatttagtttcc SEQ ID NO: 398 gaggctgaggcaggagaatgg SEQ ID NO: 399 cattgtttgatgaaggtcaacagc SEQ ID NO: 400 cagacaagagtggctacggcag SEQ ID NO: 401 acgcccagccagattattcagg SEQ ID NO: 402 ggaaccagaaagaagtgcaaagg SEQ ID NO: 403 tgagccatcttggaggcaggc SEQ ID NO: 404 caggaccttcctacaaacctcc SEQ ID NO: 405 aacacaacatatctgaccttacgc SEQ ID NO: 406 gccttagaagtccagaggaaagc SEQ ID NO: 407 tgacgtacccagtagaccttcc SEQ ID NO: 408 ctctgcaagcctgggaaacagg SEQ ID NO: 409 gccttgtccccaagtcctaagg SEQ ID NO: 410 gcaaagggactcctggaattcc SEQ ID NO: 411 gctcctgcctgtaatcccagc SEQ ID NO: 412 gaaggaaacagaaaaagcagaggc SEQ ID NO: 413 cttactaccgttcttcttcactgg SEQ ID NO: 414 actattctgtttctttaggtttactgc SEQ ID NO: 415 cggtggctcacacctgtaatcc SEQ ID NO: 416 agccagagttctgtgctctagg SEQ ID NO: 417 taatttgcatttcgtgccgctcc SEQ ID NO: 418 cacttttaatacagatcccaatagg SEQ ID NO: 419 atgtattttttcttttcctgtcaagc SEQ ID NO: 420 aaatgttaacattattctccctaagg SEQ ID NO: 421 catatgcccagatcccgtctcc SEQ ID NO: 422 acaggtgtgagccgctgcacc SEQ ID NO: 423 gccaagacgtttacagttttggc SEQ ID NO: 424 aggaaacttctgaggatgatggg SEQ ID NO: 425 gctttatagggcagtctgaattcc SEQ ID NO: 426 ttagaataaaagttatctcgggagg SEQ ID NO: 427 taatttcttcagctttatccctcag SEQ ID NO: 428 cacatgactaattctctattcattcc SEQ ID NO: 429 aaagacctcaagaaaagagtcacc SEQ ID NO: 430 gacccataaagattatatgcccag SEQ ID NO: 431 aaagtactaatgcagtgtgtcagc SEQ ID NO: 432 gaggttcctcgattcccctgc SEQ ID NO: 433 ggagagcagaggaattcacagg SEQ ID NO: 434 agtaattagaaactgattctaagacg SEQ ID NO: 435 cataccattgccaatccagttcc SEQ ID NO: 436 attacgggtgcctgccactgc SEQ ID NO: 437 cagccaggcagaggagagagg SEQ ID NO: 438 ttttcattccaagtttctgtttggg SEQ ID NO: 439 tttcaaataggaatttggataatccc SEQ ID NO: 440 taagccgagatcacaccactgc SEQ ID NO: 441 ccttcagcgcattatatcttggc SEQ ID NO: 442 ccatctaatccatcttaaattcacc SEQ ID NO: 443 gagtggagactgcgccactgc SEQ ID NO: 444 aatcatgtgccaattaaaccatggc SEQ ID NO: 445 cccagggaccagaccagacc SEQ ID NO: 446 ctcactcaccagtgaaaatcagc SEQ ID NO: 447 ggttgctctcgaactcctgacc SEQ ID NO: 448 gttcccccagctcctttctgc SEQ ID NO: 449 agaaagatgtagaagggtccagc SEQ ID NO: 450 gggaaaaggtgtattatgcaagcg SEQ ID NO: 451 ctctctcagacctaatgcaaaagc SEQ ID NO: 452 aactatacatacagtatttgtattagc SEQ ID NO: 453 aaattaatgcaatccatgatccagg SEQ ID NO: 454 ctttctccactctaagagaaccc SEQ ID NO: 455 ttttggtgtgttcatattggctgc SEQ ID NO: 456 gcttccacaaatgacagacaaagg SEQ ID NO: 457 ggctcatgcttgtaatcccagc SEQ ID NO: 458 catatgaattgttgttcctttgtagg SEQ ID NO: 459 cactggtacaagtccaagagtcc SEQ ID NO: 460 gaccctgtgtctacttcctggg SEQ ID NO: 461 tatttgaactatctcttgaaatgtcc SEQ ID NO: 462 ctgattaaaaagtattacccttggc SEQ ID NO: 463 tttgaaactgcactcaataacttgg SEQ ID NO: 464 agtaatgtgtcatgatccaatggc SEQ ID NO: 465 gaaagcatttcccaatgtctcacc SEQ ID NO: 466 caatggacaaaaggcccaactgc SEQ ID NO: 467 tccagctctggcttttttgttaag SEQ ID NO: 468 acggagtctcactccgtgacc SEQ ID NO: 469 ctatgtcatagtcaagagactttgc SEQ ID NO: 470 gttcaagcgattctcctgtctcg SEQ ID NO: 471 ccacctaatacttaaatacggaagc SEQ ID NO: 472 atattcaacaaacttaatagtgaagtg SEQ ID NO: 473 ttacaggcgtgagtcaccatgc SEQ ID NO: 474 aacacctccaagaggccaaacg SEQ ID NO: 475 tactattggcaaatttcaattatatgg
SEQ ID NO: 476 agcccacatcctaaaattcaataag SEQ ID NO: 477 gaaagtggataagtgtttgtctgg SEQ ID NO: 478 ggccaggcattcaagaccagc SEQ ID NO: 479 agccaacaacaaaaagacacaacc SEQ ID NO: 480 ttgagcccaggagttcaagacc SEQ ID NO: 481 cagactaaagatctcagagagaaac SEQ ID NO: 482 cgcttgtaatcccagcacttgg SEQ ID NO: 483 aaaagtgaaatcagaatttgtttcc SEQ ID NO: 484 caggcgtgagcaactgtgtcc SEQ ID NO: 485 ggtccagtaggatctcgtttgc SEQ ID NO: 486 actttgaaaatgttgttatagctggg SEQ ID NO: 487 ttccctgcatctaagtcttctcc SEQ ID NO: 488 agatatctaccattgaagagtttgc SEQ ID NO: 489 agtcttcacttcactttgttgtcc SEQ ID NO: 490 ccatgcaggtatgaaatataaaagc SEQ ID NO: 491 tgggtgacagagtgagactcc SEQ ID NO: 492 acagcaataccgggttaacatgc SEQ ID NO: 493 tttatgtaaaagatgaatgcgaggc SEQ ID NO: 494 ctactctgctactgggaacagg SEQ ID NO: 495 caaacgttagtctggcaaaatgcg SEQ ID NO: 496 tgcacgctaccacacccagc SEQ ID NO: 497 aattcttggatctgtgtgtttactgc SEQ ID NO: 498 taccagttatcattctctttctgc SEQ ID NO: 499 atccacccacctcggcctcc SEQ ID NO: 500 cactctgcctggcccttaatgg SEQ ID NO: 501 atagtttgtttaatatgccactaagg SEQ ID NO: 502 gcgtgagccaccgcacctgg SEQ ID NO: 503 ctccatcacacaaattttatgtggc SEQ ID NO: 504 agacggagtctcgttctgtcgc SEQ ID NO: 505 tcccaggttcaagccattctcc SEQ ID NO: 506 tattttgagagtctcactctgtcg SEQ ID NO: 507 gtctcgaactcctgacctcagg SEQ ID NO: 508 aaggaggtgaagagtgaactacg SEQ ID NO: 509 gtctcaggttttggacttacttgg SEQ ID NO: 510 tttacagatcttaaatgcattaggac SEQ ID NO: 511 gtacactgaacaaaggagacagg SEQ ID NO: 512 ctggtagtaatgcaaaatagcacc SEQ ID NO: 513 catttaatgtgaaatgaattataagcc SEQ ID NO: 514 gagacagggtttcactatgttgg SEQ ID NO: 515 ccagcactttggaaggctgagg SEQ ID NO: 516 gaaaccaagtatcatggtaaattgc SEQ ID NO: 517 cagtgagggctgctcagttcc SEQ ID NO: 518 gccaggtgcggtggctcacg SEQ ID NO: 519 catgcctgtaatcccagctacc SEQ ID NO: 520 atgtaaatggtacagtcactttagg SEQ ID NO: 521 cccacaatacagagaactcttacc SEQ ID NO: 522 tgaaacatgcagcccagtgtcc SEQ ID NO: 523 tgttttttctcctgccttcaatcc SEQ ID NO: 524 gctttcctgggtctccatctgg SEQ ID NO: 525 gcagccgcttgaaaacaaaacagc SEQ ID NO: 526 gatcacgttacatttgggggtgg SEQ ID NO: 527 taggctgaaaaactaaaatttgttgc SEQ ID NO: 528 ctcctttgggctcctttagtcc SEQ ID NO: 529 gcctcggcctcccaaagtgc SEQ ID NO: 530 aatgcctagagagatttggcagg SEQ ID NO: 531 gagatggggtttcactatgttgg SEQ ID NO: 532 tgtgatcttgccactgcactcc SEQ ID NO: 533 acttctcctccatttgtttcttcg SEQ ID NO: 534 cgtgcccgggctcagttctac SEQ ID NO: 535 ccaaaacaataaaatcacaatttggg SEQ ID NO: 536 ctgaactgccttagagtaaatccg SEQ ID NO: 537 atttctgtatcaggtctgtgttcc SEQ ID NO: 538 ggctgaccccttcactgtttcc SEQ ID NO: 539 caaaaattagccaggcatggtgg SEQ ID NO: 540 gcagtgagcagtgatcgcacc SEQ ID NO: 541 aaagactgtgaactaacttgtttgc SEQ ID NO: 542 tgccaagaattacacattattaggc SEQ ID NO: 543 ggccaggatgtcattaactttcc SEQ ID NO: 544 gtaagagctgacgtgtattgtgc SEQ ID NO: 545 cccggtgaggccgcacatcc SEQ ID NO: 546 cctgcgccttaaccccctcc SEQ ID NO: 547 cggcgcctaggggccatcg SEQ ID NO: 548 acttaaggaaacgaacatgacacc SEQ ID NO: 549 gagaccgagtcttgctgtgtcg SEQ ID NO: 550 gtattaattgaagatgatttggaatgc SEQ ID NO: 551 tctttaaaagactatcgctgaggc SEQ ID NO: 552 aaaagagacatcagtagagcatcc SEQ ID NO: 553 gttcatgttttctttgacgtctcc SEQ ID NO: 554 tttcgaaagttcaggctgagtgc SEQ ID NO: 555 gaccctcaaaacaatcctctaagg SEQ ID NO: 556 caaaacacacttagaaacaaactgc SEQ ID NO: 557 gcctgggcgacatagtgagacc SEQ ID NO: 558 ggcaggagaatggcgtgaacc SEQ ID NO: 559 tttgctcgttgcccaggctgg SEQ ID NO: 560 gcaacttaatgtgatagaataatagc SEQ ID NO: 561 cctccccttctgctgccagc SEQ ID NO: 562 ccacaacaatgtaaactcctctgg SEQ ID NO: 563 tactctccctagagttcgttccc SEQ ID NO: 564 gggtccccctttggccattcc SEQ ID NO: 565 gatcttggctcacttcaacctcc SEQ ID NO: 566 aggggaaatatttaaaccttgg SEQ ID NO: 567 aatgcaatggtgcatttacagagg SEQ ID NO: 568 tcattttatctatttctacatggtcc SEQ ID NO: 569 ggaagggaaatgcccatgaacc SEQ ID NO: 570 agtgaacattttctgcagcctcc SEQ ID NO: 571 caacaggacgtcaggcgatcc SEQ ID NO: 572 ccttcaggctgtcctgaaaagg SEQ ID NO: 573 agtctcactccatcgcccagg SEQ ID NO: 574 actgtgaacagtagttaactcagg SEQ ID NO: 575 gcatgcctgtaatccaagctgc SEQ ID NO: 576 gaaacaattctcttttcacacttgc SEQ ID NO: 577 ggctcatgcctgttatcccagc SEQ ID NO: 578 agaagaagcttagtcatatgtttgg SEQ ID NO: 579 cagatgcttgagccaaacaaatgg SEQ ID NO: 580 ctggcagacagagtgagactcc SEQ ID NO: 581 aatgtgtgaatattattcattacaggg SEQ ID NO: 582 gcaggagaattgcttgaacctgg SEQ ID NO: 583 ctttagtcaaattaaaacagtctatcc SEQ ID NO: 584 gatttctatctcctgcaaccacc SEQ ID NO: 585 ttcttgtgtaactactaaaaatctcc SEQ ID NO: 586 aaagggtcttcataaggctaatgg SEQ ID NO: 587 ctcttaaggattatttatatgaagacc SEQ ID NO: 588 caggaggagccccagagc SEQ ID NO: 589 tcctggggatggttggatgc SEQ ID NO: 590 tgaccccacagagtttacacagc SEQ ID NO: 591 agtcagggcaggctctgcc SEQ ID NO: 592 tattttggccccatccagaaagc SEQ ID NO: 593 cacccagagtacagctttgttcc SEQ ID NO: 594 gaggagccccagagcctgc SEQ ID NO: 595 tggggatggttggatgcttacc SEQ ID NO: 596 cccacagagtttacacagcttgc SEQ ID NO: 597 caggctctgcccactcacc SEQ ID NO: 598 ccatccagaaagcccaaagcc SEQ ID NO: 599 ccagagtacagctttgttcctcattc SEQ ID NO: 600 gcagtacaaacaacgcacagcg
SEQ ID NO: 601 ctgccaccctccacagtccc SEQ ID NO: 602 gccaagaccatgcatgcg SEQ ID NO: 603 cccagggacaaagagactccc SEQ ID NO: 604 caggaagcagacagtcttctagttcc SEQ ID NO: 605 tgcctgtaatcccaacactttgg SEQ ID NO: 606 tccctctggccaggatggg SEQ ID NO: 607 atggggaatgggagtaggaagc SEQ ID NO: 608 cagatcagttctcccctccagc SEQ ID NO: 609 acaaaaaagaaacatgctcagagagg SEQ ID NO: 610 tggtggcatgcatctgtagtcc SEQ ID NO: 611 aggtgctctatagatgttagcatccc SEQ ID NO: 612 ccaggacaggatggagatctgg SEQ ID NO: 613 agggaacctgtgcattatccttgc SEQ ID NO: 614 cagaagtcttgctttaaggaggagg SEQ ID NO: 615 gggtacgtgaaactcaccaagg SEQ ID NO: 616 cagagtgtggcaagcaaggg SEQ ID NO: 617 aacattttaaaggtacaaataacgtggg SEQ ID NO: 618 tagggagcaacagccattaagc SEQ ID NO: 619 ggtgcactgtccagctctgg SEQ ID NO: 620 actctcgctgaactcgcctgg SEQ ID NO: 621 ctcggtctctggtggtacgc SEQ ID NO: 622 gcaagaggtccgagctggg SEQ ID NO: 623 ggaagaagtgaaacaagagatgaagg SEQ ID NO: 624 cccagagaacaaaccggattagg SEQ ID NO: 625 cccttcaaccttctccaatctgc SEQ ID NO: 626 cccatgtccagtggtttaggg SEQ ID NO: 627 gagattggtgggagacagatgg SEQ ID NO: 628 cttctcagctcaaagttccagcg SEQ ID NO: 629 gaatgggagagatgaccagagg SEQ ID NO: 630 aagggcaagggggtatgtgg SEQ ID NO: 631 ggaaggaagcatgggaacacc SEQ ID NO: 632 ccatcaatgctctgtctgtctgg SEQ ID NO: 633 gtgccgtgactgtgcttgg SEQ ID NO: 634 acatcccattgacctcatcaagc
Sequence CWU
1
6341203DNAHomo sapiensBCR region 1gtgggccccc ccgtttccgt gtacagggca
cctgcaggga gggcaggcag ctagcctgaa 60ggctgatccc cccttcctgt tagcactttt
gatgggacta gtggactttg gttcagaagg 120aagagctatg cttgttaggg cctcttgtct
cctcccagga gtggacaagg tgggttagga 180gcagtttctc cctgagtggc tgc
2032376DNAHomo sapiensABL region
2caccacgtct ggctaatttt tgtattttta gtagagatgg ggtttcaaca tgttagccag
60gctggtctcg aactcctgac ctcaggtgat ccacccgcct gggccctcca aagtgctggg
120attacaggca ggagccactg tgcccggcct gacctcatat ttgaataccg agttttagtt
180ctggaggagc tgcaggtttt atgaaaaggg aacacatttg attcctcaga gcagccacag
240gccagctctc tgaagtaaag tgcacgtgtg catgtgtgtg cacactcaca cacacgtaca
300cacacattca caaataactg tgcccggcct gacctcatat ttgaataccg agttttagtt
360ctggaggagc tgcagg
3763231DNAHomo sapiensBCR region 3tttgggaggc tgaggcaggt ggatcgcttg
agctcaggag ttggagacca gcctgaccaa 60catggtgaaa ccctgtgtct actaaaaata
caaagattag ccgggctagg cagtgggcac 120ctgtaatcac aactgcttgg gaggctgagg
gaagagaatc gcttgaaccc aggaggcgga 180ggttgcagtg agccgagctt gtgccactgc
attccagcct gggcgacaga g 2314249DNAHomo sapiensABL region
4ggtctcactc tgttgaactc ctggtggcct caagggatcc tcctacctcg gcctcacaaa
60gtattggaat tacaggtgtg agtcactgca gctggccttc acttatcact gtgaggagta
120aacagctgca tggtgggctt aatgccatct aacacgagtg actccatgtt cagacagtag
180gatcacaaat gattattata tagcaatgaa tggccacagg tacatagact aaggagccac
240atccctgct
2495212DNAHomo sapiensBCR region 5cctccagcta cctgccagcc ggcacttttg
gtcaagctgt tttgcattca ctgttgcaca 60tatgctcagt cacacacaca gcatacgcta
tgcacatgtg tccacacaca ccccacccac 120atcccacatc accccgaccc cctctgctgt
ccttggaacc ttattacact tcgagtcact 180ggtttgcctg tattgtgaaa ccagctggat
cc 2126200DNAHomo sapiensABL region
6ttatttataa caacattttc agcgtggcaa ctgcagtttc agaatggtgg aattatacca
60gtcagagaga gatgcaaatg atttaaaata ggaagaaagc aggtgtctgg cccagaggac
120cagattaaga agaccccatg agagttacaa tagttagtga aaatggtgct tctgcaaacc
180tcatgtctac agaagctggt
2007213DNAHomo sapiensBCR region 7tgcaccttca taacataatc tttctcctgg
gcccctgtct ctggctgcct cataaacgct 60ggtgtttccc tcgtgggcct ccctgcatcc
ctgcatctcc tcccgggtcc tgtctgtgag 120caatacagcg tgacacccta cgctgccccg
tggtcccggg cttgtctctc cttgcctccc 180tgttaccttt ctttctatct cttccttgcc
ccg 2138251DNAHomo sapiensABL region
8gtgagctccg cctcctgtca gatcagtggc ggcattagtt tctcatagga gcatgaaatc
60tattgtgaac agtacatgcg atggatccag gttgcgtgct cctagtgaga atctaatgcc
120tgaggatctc tcattgtctc ttatcactcc cagataggac tgtctagttg caggaaaaca
180agctcagggc tcccactgat tctacattac agtgggttgt ataattatta tatattacaa
240tgtaataata a
251963DNAHomo sapiens 9ggagtctgag gaggggaagg aggcaaggtt ggctcggatc
ccagccagta agtctgggtg 60tgg
631020DNAArtificial SequenceSynthetic
oligonucleotide 10cttctccctg acatccgtgg
201125DNAArtificial SequenceSynthetic oligonucleotide
11acacagcata cgctatgcac atgtg
251222DNAArtificial SequenceSynthetic oligonucleotide 12gaggttgttc
agatgaccac gg
221324DNAArtificial SequenceSynthetic oligonucleotide 13cagctactgg
agctgtcaga acag
241417DNAArtificial SequenceSynthetic oligonucleotide 14tgggcctccc
tgcatcc
171517DNAArtificial SequenceSynthetic oligonucleotide 15tccccctgca
ccccacg
171624DNAArtificial SequenceSynthetic oligonucleotide 16tgacatccgt
ggagctgcag atgc
241724DNAArtificial SequenceSynthetic oligonucleotide 17acatgtgtcc
acacacaccc cacc
241824DNAArtificial SequenceSynthetic oligonucleotide 18accacgggac
acctttgacc ctgg
241924DNAArtificial SequenceSynthetic oligonucleotide 19ctggagctgt
cagaacagtg aagg
242024DNAArtificial SequenceSynthetic oligonucleotide 20tccctgcatc
cctgcatctc ctcc
242124DNAArtificial SequenceSynthetic oligonucleotide 21cccacgactt
ctccagcact gagc
242223DNAArtificial SequenceSynthetic oligonucleotide 22gcaacactgt
gacgtactgg agg
232326DNAArtificial SequenceSynthetic oligonucleotide 23gtctatctaa
aattcacaag gaatgc
262424DNAArtificial SequenceSynthetic oligonucleotide 24aggcaaagta
aaatccaagc accc
242521DNAArtificial SequenceSynthetic oligonucleotide 25cactcctgca
ctccagcctg g
212624DNAArtificial SequenceSynthetic oligonucleotide 26caaccaccaa
agtgcttttc ctgg
242726DNAArtificial SequenceSynthetic oligonucleotide 27atatggcatc
tgtaaatatt accacc
262821DNAArtificial SequenceSynthetic oligonucleotide 28tgcctcggcc
tcccaaagtg c
212920DNAArtificial SequenceSynthetic oligonucleotide 29agccaccaca
cccagccagg
203025DNAArtificial SequenceSynthetic oligonucleotide 30aataactgtt
ttctcccccc aaaac
253125DNAArtificial SequenceSynthetic oligonucleotide 31tgttttacaa
aaatggggcc atacc
253228DNAArtificial SequenceSynthetic oligonucleotide 32acttaagcaa
attctttcat aaaaaggg
283325DNAArtificial SequenceSynthetic oligonucleotide 33ctttcaattg
ttgtaccaac tctcc
253423DNAArtificial SequenceSynthetic oligonucleotide 34acctcctgca
tctctccttt tgc
233527DNAArtificial SequenceSynthetic oligonucleotide 35aaataaagtt
ttgagaacca taagtgg
273622DNAArtificial SequenceSynthetic oligonucleotide 36caccatcaca
gctcactgca gc
223724DNAArtificial SequenceSynthetic oligonucleotide 37aacctctttg
agaatcggat agcc
243827DNAArtificial SequenceSynthetic oligonucleotide 38aaataaagta
catacctcca attttgc
273925DNAArtificial SequenceSynthetic oligonucleotide 39gacacattcc
tatgggttta attcc
254026DNAArtificial SequenceSynthetic oligonucleotide 40tgtaaaatat
ggtttcagaa gggagg
264122DNAArtificial SequenceSynthetic oligonucleotide 41gcaggtggat
aacgaggtca gg
224225DNAArtificial SequenceSynthetic oligonucleotide 42ccagccaaga
atttcaaaga ttagc
254323DNAArtificial SequenceSynthetic oligonucleotide 43gaagggagat
gacaaaggga acg
234423DNAArtificial SequenceSynthetic oligonucleotide 44gcagaagaac
tgcttgaacc tgg
234522DNAArtificial SequenceSynthetic oligonucleotide 45gtggtcccag
ctactcgaga gg
224625DNAArtificial SequenceSynthetic oligonucleotide 46ccctcagcaa
aactaactga aaagg
254726DNAArtificial SequenceSynthetic oligonucleotide 47tagaaaccaa
gatatctaga attccc
264822DNAArtificial SequenceSynthetic oligonucleotide 48ccacgcccgg
cggaataaat gc
224926DNAArtificial SequenceSynthetic oligonucleotide 49acaaaaaaag
aggcaaaaac tgagag
265021DNAArtificial SequenceSynthetic oligonucleotide 50ctgggcgcag
tggctcatgc c
215122DNAArtificial SequenceSynthetic oligonucleotide 51tggctgtgag
gctgagaact gc
225222DNAArtificial SequenceSynthetic oligonucleotide 52ctgggcgaca
gagtgagact cc
225321DNAArtificial SequenceSynthetic oligonucleotide 53aagtctggct
gggcgcagtg g
215424DNAArtificial SequenceSynthetic oligonucleotide 54aatggacaaa
agaggtgaac tggc
245524DNAArtificial SequenceSynthetic oligonucleotide 55gatagagtga
aaacgcacaa tggc
245626DNAArtificial SequenceSynthetic oligonucleotide 56aattaaacag
ctaggtcaat atgagg
265723DNAArtificial SequenceSynthetic oligonucleotide 57ggtctccact
atcaagggac aag
235824DNAArtificial SequenceSynthetic oligonucleotide 58aagcagctgt
tagtcatttc cagg
245923DNAArtificial SequenceSynthetic oligonucleotide 59aggcatcctc
agattatggc tcc
236023DNAArtificial SequenceSynthetic oligonucleotide 60cctgagtaac
actgagaccc tgc
236124DNAArtificial SequenceSynthetic oligonucleotide 61aacactcaag
ctgtcaagag acac
246221DNAArtificial SequenceSynthetic oligonucleotide 62attcaggcca
ggcgcagtgg c
216324DNAArtificial SequenceSynthetic oligonucleotide 63taaatcgtaa
aactgccaca aagc
246424DNAArtificial SequenceSynthetic oligonucleotide 64cagaggagta
ggagaaggaa aagg
246525DNAArtificial SequenceSynthetic oligonucleotide 65ggtagctatc
taccaagtag aatcc
256625DNAArtificial SequenceSynthetic oligonucleotide 66atcagattgg
aaaaagtccc aaagc
256723DNAArtificial SequenceSynthetic oligonucleotide 67ctcctgaaaa
gcacctactc agc
236823DNAArtificial SequenceSynthetic oligonucleotide 68ctccttaaac
ctgaggtact ggg
236925DNAArtificial SequenceSynthetic oligonucleotide 69ttttctccta
atagaccacc attcc
257025DNAArtificial SequenceSynthetic oligonucleotide 70ctgctgtatt
accatcactc atgtc
257123DNAArtificial SequenceSynthetic oligonucleotide 71ctggccaaca
tagtgaaacc acg
237227DNAArtificial SequenceSynthetic oligonucleotide 72atttgaatag
gggttaaagt atcattg
277323DNAArtificial SequenceSynthetic oligonucleotide 73cacttcagtg
gaagttggca tgc
237427DNAArtificial SequenceSynthetic oligonucleotide 74gtttttcttc
gaagtgataa acatacg
277524DNAArtificial SequenceSynthetic oligonucleotide 75gctccttagt
ctatgtacct gtgg
247623DNAArtificial SequenceSynthetic oligonucleotide 76tactctggca
tggtaactgg tgc
237723DNAArtificial SequenceSynthetic oligonucleotide 77acaaaggact
aggtctgtgg agc
237828DNAArtificial SequenceSynthetic oligonucleotide 78ccaagtttac
caaattacca aagttacc
287922DNAArtificial SequenceSynthetic oligonucleotide 79tgagccgata
tcacgccact gc
228024DNAArtificial SequenceSynthetic oligonucleotide 80tcccaataaa
ggttttggcc cagg
248124DNAArtificial SequenceSynthetic oligonucleotide 81ctgggtagca
aattagggaa cagg
248225DNAArtificial SequenceSynthetic oligonucleotide 82ctggccagaa
aagacagttt tatcc
258323DNAArtificial SequenceSynthetic oligonucleotide 83ggttcccagg
aagggataac acc
238423DNAArtificial SequenceSynthetic oligonucleotide 84tcactccagg
aggttccatt tcc
238524DNAArtificial SequenceSynthetic oligonucleotide 85aggcttggaa
ataagcagca gtgg
248626DNAArtificial SequenceSynthetic oligonucleotide 86attcatacaa
tggaatacta ctcagc
268723DNAArtificial SequenceSynthetic oligonucleotide 87taagtgatcc
tcccacctca acc
238823DNAArtificial SequenceSynthetic oligonucleotide 88tataagagga
agactggggc tgg
238925DNAArtificial SequenceSynthetic oligonucleotide 89tcatacttat
gcaggttata ggagg
259022DNAArtificial SequenceSynthetic oligonucleotide 90caagatcacg
ccactgcact cc
229126DNAArtificial SequenceSynthetic oligonucleotide 91aaaataaata
gctggtgctc aagatc
269223DNAArtificial SequenceSynthetic oligonucleotide 92caccagcctc
attcaacaga tgg
239322DNAArtificial SequenceSynthetic oligonucleotide 93caatgcagcc
tcaacctcct gg
229423DNAArtificial SequenceSynthetic oligonucleotide 94gttaggtcag
gtgctcatgt ctg
239527DNAArtificial SequenceSynthetic oligonucleotide 95aagtttcaaa
aggacatgta caaaatg
279622DNAArtificial SequenceSynthetic oligonucleotide 96tcctgaagag
gctgcagctt cc
229723DNAArtificial SequenceSynthetic oligonucleotide 97ctggtgcaca
ttcccaagtg tgc
239824DNAArtificial SequenceSynthetic oligonucleotide 98catgttggcc
atgttcttct gagg
249921DNAArtificial SequenceSynthetic oligonucleotide 99ctcagcctcc
cgagtagctg g
2110025DNAArtificial SequenceSynthetic oligonucleotide 100aaagacattt
aagaggagat gaggc
2510122DNAArtificial SequenceSynthetic oligonucleotide 101tgctgggatt
acaggcgtga gc
2210222DNAArtificial SequenceSynthetic oligonucleotide 102tgtgacttcc
atccgcagct cc
2210324DNAArtificial SequenceSynthetic oligonucleotide 103gacacttttg
tggagctttc atgg
2410422DNAArtificial SequenceSynthetic oligonucleotide 104catgtgaggg
ggcacgtctt gc
2210525DNAArtificial SequenceSynthetic oligonucleotide 105tcttctctat
gagaaaagtg gttgc
2510623DNAArtificial SequenceSynthetic oligonucleotide 106tggcaaaatg
ctatcgagct gcc
2310722DNAArtificial SequenceSynthetic oligonucleotide 107tatgaacaca
gccggcctca gg
2210823DNAArtificial SequenceSynthetic oligonucleotide 108gaggttgcag
tgagctgaga tcg
2310922DNAArtificial SequenceSynthetic oligonucleotide 109gtcaagcacc
cagtccgata cc
2211021DNAArtificial SequenceSynthetic oligonucleotide 110atctgggctt
ggtggcgcac g
2111122DNAArtificial SequenceSynthetic oligonucleotide 111gttaagcggg
tcccacatca gc
2211224DNAArtificial SequenceSynthetic oligonucleotide 112cagccagttt
cagtagaaag atgc
2411323DNAArtificial SequenceSynthetic oligonucleotide 113gacccaagca
taaggggact agc
2311427DNAArtificial SequenceSynthetic oligonucleotide 114cccaaaaagt
ttacaagaga aattttc
2711522DNAArtificial SequenceSynthetic oligonucleotide 115cgcctgtagt
cccagctact cg
2211623DNAArtificial SequenceSynthetic oligonucleotide 116cgcgtgatgc
ggaaaagaaa tcc
2311724DNAArtificial SequenceSynthetic oligonucleotide 117tctactatga
accctccttc agac
2411823DNAArtificial SequenceSynthetic oligonucleotide 118gtgctgggat
tacaggtgtg agc
2311923DNAArtificial SequenceSynthetic oligonucleotide 119ttatccaaat
gtcccagggc agg
2312021DNAArtificial SequenceSynthetic oligonucleotide 120ctgccagcac
tgctcgccag c
2112122DNAArtificial SequenceSynthetic oligonucleotide 121gctactgcag
gcagtgcctt cc
2212222DNAArtificial SequenceSynthetic oligonucleotide 122catccaagcc
caaggtgtca gg
2212325DNAArtificial SequenceSynthetic oligonucleotide 123tgtttgcatg
taatttcagg aagcc
2512424DNAArtificial SequenceSynthetic oligonucleotide 124gatccgtcac
tgttaacact cagg
2412523DNAArtificial SequenceSynthetic oligonucleotide 125ctcacagtca
caagctcctg agc
2312622DNAArtificial SequenceSynthetic oligonucleotide 126gagatgatgc
tggggtcaca gg
2212723DNAArtificial SequenceSynthetic oligonucleotide 127ttagaagaat
gggatcgcaa agg
2312824DNAArtificial SequenceSynthetic oligonucleotide 128cggtattcaa
atatgaggtc aggc
2412923DNAArtificial SequenceSynthetic oligonucleotide 129gtaaatcctg
ctgccagtct tcc
2313022DNAArtificial SequenceSynthetic oligonucleotide 130acagggtcag
acagagcctt gg
2213127DNAArtificial SequenceSynthetic oligonucleotide 131agttattgat
ctaactatac aacaagc
2713221DNAArtificial SequenceSynthetic oligonucleotide 132aaagactagg
ggccggggac g
2113326DNAArtificial SequenceSynthetic oligonucleotide 133ctggtagaaa
taaagacaac aaagcc
2613427DNAArtificial SequenceSynthetic oligonucleotide 134gtgccaagta
attaaaagtt tgaaacc
2713522DNAArtificial SequenceSynthetic oligonucleotide 135ggcttttgaa
gggagcacca cc
2213627DNAArtificial SequenceSynthetic oligonucleotide 136gaaggataaa
tacctatgat actttcc
2713724DNAArtificial SequenceSynthetic oligonucleotide 137ggcagggaaa
tactgtgctt caag
2413824DNAArtificial SequenceSynthetic oligonucleotide 138gtggtgaaat
tccacctcag tacc
2413923DNAArtificial SequenceSynthetic oligonucleotide 139tcccaaagtg
ctgggattac agg
2314025DNAArtificial SequenceSynthetic oligonucleotide 140gaaattagca
aacaatgcca agacg
2514123DNAArtificial SequenceSynthetic oligonucleotide 141taagtattgg
accgggaagg agg
2314225DNAArtificial SequenceSynthetic oligonucleotide 142ctatcatttt
gctcaaagtg tagcc
2514324DNAArtificial SequenceSynthetic oligonucleotide 143atttcacaaa
ctacagaggc cagg
2414424DNAArtificial SequenceSynthetic oligonucleotide 144tagacttctg
tctctctatg ctgc
2414523DNAArtificial SequenceSynthetic oligonucleotide 145tgagtgagct
gccatgtgat acc
2314622DNAArtificial SequenceSynthetic oligonucleotide 146acttcacacc
agcctgtcca cc
2214724DNAArtificial SequenceSynthetic oligonucleotide 147taactcatat
cctcagagag accc
2414822DNAArtificial SequenceSynthetic oligonucleotide 148agaggttcct
cgattcccct gc
2214923DNAArtificial SequenceSynthetic oligonucleotide 149gtgtcagcgt
cccaacacaa agc
2315023DNAArtificial SequenceSynthetic oligonucleotide 150gaaagtggat
gggcaagcat tgc
2315122DNAArtificial SequenceSynthetic oligonucleotide 151gtgatcacct
cacagctgca gg
2215225DNAArtificial SequenceSynthetic oligonucleotide 152gtttgtttag
tcaaggcatt tcacc
2515322DNAArtificial SequenceSynthetic oligonucleotide 153cctcagcctc
cagagtagct gg
2215424DNAArtificial SequenceSynthetic oligonucleotide 154taaaagaaaa
ctcctccttc ctgg
2415526DNAArtificial SequenceSynthetic oligonucleotide 155aatgtgctat
gtctttaaat ccatgg
2615626DNAArtificial SequenceSynthetic oligonucleotide 156agctggcaaa
tctggtaata taaaag
2615722DNAArtificial SequenceSynthetic oligonucleotide 157gcttgaacct
ggaaggtgga gg
2215823DNAArtificial SequenceSynthetic oligonucleotide 158gcaggcatgc
taagaccttc agc
2315924DNAArtificial SequenceSynthetic oligonucleotide 159cagctccatg
aataactcca cagg
2416022DNAArtificial SequenceSynthetic oligonucleotide 160gcttgaaccc
aggaggcaga gg
2216123DNAArtificial SequenceSynthetic oligonucleotide 161atcgaagatg
ccactgcaag agg
2316223DNAArtificial SequenceSynthetic oligonucleotide 162ccaaccacac
ttcaggggat acc
2316323DNAArtificial SequenceSynthetic oligonucleotide 163cacgccagtc
cactgatact cac
2316422DNAArtificial SequenceSynthetic oligonucleotide 164gggtttcacc
atgttggcca gg
2216522DNAArtificial SequenceSynthetic oligonucleotide 165cccaacaaag
gctctggcct gg
2216623DNAArtificial SequenceSynthetic oligonucleotide 166atgacagcag
aggagcttca tcc
2316723DNAArtificial SequenceSynthetic oligonucleotide 167gcaggctacg
agtaaaagga tgg
2316823DNAArtificial SequenceSynthetic oligonucleotide 168cgggtaaaat
cttgcctcct tcc
2316925DNAArtificial SequenceSynthetic oligonucleotide 169aaacttaaac
caatggtgga tgtgg
2517023DNAArtificial SequenceSynthetic oligonucleotide 170agagactgag
gaactgttcc agc
2317124DNAArtificial SequenceSynthetic oligonucleotide 171gaaacggtct
tggatcactg atcc
2417224DNAArtificial SequenceSynthetic oligonucleotide 172tgcgcatgat
atcttgtttc aggg
2417323DNAArtificial SequenceSynthetic oligonucleotide 173ggcctccgtt
taaactgttg tgc
2317422DNAArtificial SequenceSynthetic oligonucleotide 174gaatgctggc
ccgacacagt gg
2217524DNAArtificial SequenceSynthetic oligonucleotide 175tcttggtata
gaaaagccag ctgg
2417622DNAArtificial SequenceSynthetic oligonucleotide 176gcaaaagccc
aagagcccct gg
2217723DNAArtificial SequenceSynthetic oligonucleotide 177ttctcccaaa
atgagcccca agg
2317824DNAArtificial SequenceSynthetic oligonucleotide 178gtggtgacgt
aaacaaaagg tacc
2417926DNAArtificial SequenceSynthetic oligonucleotide 179gcaaattcca
tgtgaatctt attggc
2618023DNAArtificial SequenceSynthetic oligonucleotide 180cctgatctat
ggaacagtgg tgg
2318125DNAArtificial SequenceSynthetic oligonucleotide 181gttacaaacg
ttgcagtttg caacg
2518223DNAArtificial SequenceSynthetic oligonucleotide 182gaaccccgtc
aacagtgatc acc
2318322DNAArtificial SequenceSynthetic oligonucleotide 183acaggacctc
aaggcaagga gc
2218427DNAArtificial SequenceSynthetic oligonucleotide 184catacctaaa
atagaaatgt ctatccc
2718527DNAArtificial SequenceSynthetic oligonucleotide 185gagttgcata
tatgttttat aaatccc
2718624DNAArtificial SequenceSynthetic oligonucleotide 186tgagcccaca
tccataaagt tagc
2418721DNAArtificial SequenceSynthetic oligonucleotide 187accgcaacct
ttgccgcctg g
2118826DNAArtificial SequenceSynthetic oligonucleotide 188taaatatttt
gtatggagtc accacc
2618925DNAArtificial SequenceSynthetic oligonucleotide 189aaagccagga
gaaaaagtta tgagg
2519023DNAArtificial SequenceSynthetic oligonucleotide 190tcccaaagtc
ccaggattac agg
2319123DNAArtificial SequenceSynthetic oligonucleotide 191tcactatgga
gcatctccga tgg
2319222DNAArtificial SequenceSynthetic oligonucleotide 192agttccctgg
aagtctccga gg
2219324DNAArtificial SequenceSynthetic oligonucleotide 193aaaataatca
cccagcccac atcc
2419425DNAArtificial SequenceSynthetic oligonucleotide 194acaaaactac
agacacagaa agtgg
2519522DNAArtificial SequenceSynthetic oligonucleotide 195tttgggaggc
tgaggtaggt gg
2219625DNAArtificial SequenceSynthetic oligonucleotide 196aaagacagtg
aaacatctat aaggg
2519722DNAArtificial SequenceSynthetic oligonucleotide 197cattttggga
gaccagggca gg
2219823DNAArtificial SequenceSynthetic oligonucleotide 198gcatgggaca
gacacaaagc agc
2319923DNAArtificial SequenceSynthetic oligonucleotide 199gaataacaaa
gagagccggc tgg
2320029DNAArtificial SequenceSynthetic oligonucleotide 200taaacctttt
attgaaaatt gtcaaatgg
2920121DNAArtificial SequenceSynthetic oligonucleotide 201cgcctcagcc
tcccaaagtg c
2120226DNAArtificial SequenceSynthetic oligonucleotide 202tacattagtt
ttataggtcc agtagg
2620327DNAArtificial SequenceSynthetic oligonucleotide 203gaaggtttat
tcatattaaa atgtgcc
2720423DNAArtificial SequenceSynthetic oligonucleotide 204ctggcttctg
tggtttgagt tgg
2320523DNAArtificial SequenceSynthetic oligonucleotide 205acagacctac
ctcctaagga tgg
2320625DNAArtificial SequenceSynthetic oligonucleotide 206gctagctttt
gtgtgtaaga atggg
2520724DNAArtificial SequenceSynthetic oligonucleotide 207ggcctactca
cacaatagaa tacc
2420822DNAArtificial SequenceSynthetic oligonucleotide 208gcaccattgc
actccagcct gg
2220926DNAArtificial SequenceSynthetic oligonucleotide 209gaaattagga
taaaggttgt cacagc
2621025DNAArtificial SequenceSynthetic oligonucleotide 210cagaagtgtt
caaggtgaaa ctgtc
2521126DNAArtificial SequenceSynthetic oligonucleotide 211ctgaatcatg
aaatgttcta ctctgc
2621223DNAArtificial SequenceSynthetic oligonucleotide 212tgtcaacttg
actgggccat acg
2321324DNAArtificial SequenceSynthetic oligonucleotide 213ctcccgtata
gttgggatta tagg
2421425DNAArtificial SequenceSynthetic oligonucleotide 214gcttggagtt
ccttgaaatt cttgg
2521523DNAArtificial SequenceSynthetic oligonucleotide 215cctggtggct
ccagttttct acc
2321623DNAArtificial SequenceSynthetic oligonucleotide 216aactcctgac
ctcatgatcc acc
2321722DNAArtificial SequenceSynthetic oligonucleotide 217gctgggatta
caggcatgag cc
2221825DNAArtificial SequenceSynthetic oligonucleotide 218ttctccttta
tccttggtga cattc
2521923DNAArtificial SequenceSynthetic oligonucleotide 219tcccaaagtg
ctgggattac agg
2322023DNAArtificial SequenceSynthetic oligonucleotide 220gtcataagtc
agggaccatc tgc
2322125DNAArtificial SequenceSynthetic oligonucleotide 221ctgtttcatt
gatttccaga ctggc
2522222DNAArtificial SequenceSynthetic oligonucleotide 222gcaatctcgg
ctcactgcaa gc
2222325DNAArtificial SequenceSynthetic oligonucleotide 223gaagaagtga
ctatatcaga tctgg
2522422DNAArtificial SequenceSynthetic oligonucleotide 224ttcaccatgt
tggccaggct gg
2222524DNAArtificial SequenceSynthetic oligonucleotide 225catcactgaa
gatgacaact gagc
2422621DNAArtificial SequenceSynthetic oligonucleotide 226gtccagcctg
ggcgatagag c
2122724DNAArtificial SequenceSynthetic oligonucleotide 227gaggaaagtc
tttgaagagg aacc
2422823DNAArtificial SequenceSynthetic oligonucleotide 228ggtacactca
ccagcagttt tgc
2322925DNAArtificial SequenceSynthetic oligonucleotide 229gagcaactgg
tgtgaataca tatgg
2523023DNAArtificial SequenceSynthetic oligonucleotide 230caatacctgg
caccacatac acc
2323122DNAArtificial SequenceSynthetic oligonucleotide 231gggactacag
gcatgtgcca cc
2223222DNAArtificial SequenceSynthetic oligonucleotide 232cggtggctca
cgcgtgtaat cc
2223326DNAArtificial SequenceSynthetic oligonucleotide 233caactgttaa
atctctcatg gaaacc
2623423DNAArtificial SequenceSynthetic oligonucleotide 234gacaaaggat
tagaaatgca ccc
2323526DNAArtificial SequenceSynthetic oligonucleotide 235ggaaatgttc
taaaactgga ttgtgg
2623625DNAArtificial SequenceSynthetic oligonucleotide 236aataataata
gccaggtgtg gtagc
2523723DNAArtificial SequenceSynthetic oligonucleotide 237ctggaacact
cacacattgc tgg
2323822DNAArtificial SequenceSynthetic oligonucleotide 238ctgggtgaca
gagcgagact cc
2223925DNAArtificial SequenceSynthetic oligonucleotide 239cccaaatcat
ccccgtgaaa catgc
2524022DNAArtificial SequenceSynthetic oligonucleotide 240gaccctgcaa
tcccaacact gg
2224123DNAArtificial SequenceSynthetic oligonucleotide 241ctctcaggcc
ttcaaactac acc
2324222DNAArtificial SequenceSynthetic oligonucleotide 242caggaaaggg
ctcgctcagt gg
2224323DNAArtificial SequenceSynthetic oligonucleotide 243atctgcaaaa
gcagcagagc agg
2324425DNAArtificial SequenceSynthetic oligonucleotide 244gtacccatga
cagacaagtt ttagg
2524524DNAArtificial SequenceSynthetic oligonucleotide 245cttatcccct
actgtctcct ttgg
2424622DNAArtificial SequenceSynthetic oligonucleotide 246ggatggtctc
gatctcctga cc
2224724DNAArtificial SequenceSynthetic oligonucleotide 247aggttagaga
ccttcctcta atgc
2424822DNAArtificial SequenceSynthetic oligonucleotide 248agctgggatt
acaggtgcct gc
2224921DNAArtificial SequenceSynthetic oligonucleotide 249gctgaggcag
gttggggctg c
2125025DNAArtificial SequenceSynthetic oligonucleotide 250acatttaacg
tctcctaact tctcc
2525123DNAArtificial SequenceSynthetic oligonucleotide 251gtgctgcgat
tacaggtgtg agc
2325226DNAArtificial SequenceSynthetic oligonucleotide 252tatgacagca
gtattatact atcacc
2625323DNAArtificial SequenceSynthetic oligonucleotide 253ctggggacca
aatctgaact gcc
2325426DNAArtificial SequenceSynthetic oligonucleotide 254gtagctattg
ttatttccaa aagagg
2625521DNAArtificial SequenceSynthetic oligonucleotide 255gcttgggacc
ccaggacaag g
2125622DNAArtificial SequenceSynthetic oligonucleotide 256cctggccaac
atggggaaat cc
2225723DNAArtificial SequenceSynthetic oligonucleotide 257aattgcttga
acctgggagg tgg
2325824DNAArtificial SequenceSynthetic oligonucleotide 258gcctaagacc
caaaagctat tagc
2425927DNAArtificial SequenceSynthetic oligonucleotide 259catattaaag
ggccatattc aaattgg
2726025DNAArtificial SequenceSynthetic oligonucleotide 260ggatgtaacc
agtgtatatc acagg
2526125DNAArtificial SequenceSynthetic oligonucleotide 261ggaagtttag
tccacatctt ctagc
2526222DNAArtificial SequenceSynthetic oligonucleotide 262gcacccacag
gacaaccaca cg
2226322DNAArtificial SequenceSynthetic oligonucleotide 263gggacgcgcc
tgttaacaaa gg
2226419DNAArtificial SequenceSynthetic oligonucleotide 264gggctggggg
ccacgctcc
1926522DNAArtificial SequenceSynthetic oligonucleotide 265cgcaaaagtg
aagccctcct gg
2226626DNAArtificial SequenceSynthetic oligonucleotide 266gaaatcctac
ttgatctaaa gtgagc
2626726DNAArtificial SequenceSynthetic oligonucleotide 267tttgagcaac
ttggaaaaaa taagcg
2626825DNAArtificial SequenceSynthetic oligonucleotide 268ttcccaaaag
acaaatagca cttcc
2526926DNAArtificial SequenceSynthetic oligonucleotide 269ccattttgaa
aatcacagtg aattcc
2627026DNAArtificial SequenceSynthetic oligonucleotide 270gaaaagaaaa
ccctgaattc aaaagg
2627126DNAArtificial SequenceSynthetic oligonucleotide 271tgctgaaaag
aagcatttaa aagtgg
2627224DNAArtificial SequenceSynthetic oligonucleotide 272ctcttaccag
tttcagagct ttcc
2427325DNAArtificial SequenceSynthetic oligonucleotide 273ttttcagcca
aaaatcaagg acagg
2527423DNAArtificial SequenceSynthetic oligonucleotide 274cttgagccca
ggagtttgag acc
2327523DNAArtificial SequenceSynthetic oligonucleotide 275cgcctgtagt
accctctact agg
2327626DNAArtificial SequenceSynthetic oligonucleotide 276ggtaaagaaa
gaaggatttg aaaacc
2627727DNAArtificial SequenceSynthetic oligonucleotide 277taagagtaat
gaggttaaag tttatgc
2727826DNAArtificial SequenceSynthetic oligonucleotide 278catttttatt
gtcacaggcc atttgc
2627923DNAArtificial SequenceSynthetic oligonucleotide 279gccacgcctt
ctcttctgcc acc
2328022DNAArtificial SequenceSynthetic oligonucleotide 280tgcctctcct
gactgcactg tg
2228122DNAArtificial SequenceSynthetic oligonucleotide 281ccatgctcta
ccacgccctt gg
2228222DNAArtificial SequenceSynthetic oligonucleotide 282cattcaggct
ggagtgcggt gg
2228327DNAArtificial SequenceSynthetic oligonucleotide 283cttaaaaatt
gtctggctaa gacattg
2728422DNAArtificial SequenceSynthetic oligonucleotide 284ttgctcttgt
tgcccgggtt gg
2228526DNAArtificial SequenceSynthetic oligonucleotide 285gagcttagag
gaaaagtatt atttcc
2628621DNAArtificial SequenceSynthetic oligonucleotide 286tggtgctgtg
ccagacgctg g
2128725DNAArtificial SequenceSynthetic oligonucleotide 287cagatctttt
tggctattgt cttgg
2528822DNAArtificial SequenceSynthetic oligonucleotide 288gaaggaaagg
gcctcccact gc
2228923DNAArtificial SequenceSynthetic oligonucleotide 289catgaaaaag
catgctgggg agg
2329028DNAArtificial SequenceSynthetic oligonucleotide 290caaacataaa
aaagctttaa tagaagcc
2829126DNAArtificial SequenceSynthetic oligonucleotide 291tcccaactat
gaaaaaatag aagacg
2629223DNAArtificial SequenceSynthetic oligonucleotide 292cacaaattag
ccgggcatgg tgg
2329326DNAArtificial SequenceSynthetic oligonucleotide 293cttcctttac
tgagtctttc taaagc
2629425DNAArtificial SequenceSynthetic oligonucleotide 294tgtcctttga
aatgtaggta tgtgg
2529524DNAArtificial SequenceSynthetic oligonucleotide 295ggatcttgca
atactgacat ctcc
2429626DNAArtificial SequenceSynthetic oligonucleotide 296atttgaaaag
aactgaagga tctacc
2629723DNAArtificial SequenceSynthetic oligonucleotide 297gtgagctgag
atctcgtctc tgc
2329828DNAArtificial SequenceSynthetic oligonucleotide 298tttgtctgaa
acagattcta aaagttgg
2829921DNAArtificial SequenceSynthetic oligonucleotide 299gcaggtgcct
gtagtcccag c
2130027DNAArtificial SequenceSynthetic oligonucleotide 300gtttgagctt
ctaaaattca tggattc
2730123DNAArtificial SequenceSynthetic oligonucleotide 301gtggtaggtc
aaaccgcaat tcc
2330225DNAArtificial SequenceSynthetic oligonucleotide 302accaaatcag
acatatcagc tttgg
2530324DNAArtificial SequenceSynthetic oligonucleotide 303cacagaacgg
atcctcaata aagg
2430425DNAArtificial SequenceSynthetic oligonucleotide 304gttaactcct
cccttctctt tatgg
2530525DNAArtificial SequenceSynthetic oligonucleotide 305gtgttcagag
agcttgattt ccagg
2530623DNAArtificial SequenceSynthetic oligonucleotide 306cccacttgat
ttttcccaca tgg
2330728DNAArtificial SequenceSynthetic oligonucleotide 307atttatttag
atgaagtgaa tattttcc
2830825DNAArtificial SequenceSynthetic oligonucleotide 308atttagtttg
tttaactgtg agtgc
2530924DNAArtificial SequenceSynthetic oligonucleotide 309gtacagaagt
gcttgatgca tacc
2431025DNAArtificial SequenceSynthetic oligonucleotide 310aggcagataa
aaattctcca ttagc
2531121DNAArtificial SequenceSynthetic oligonucleotide 311acaagcacga
gccacagcac c
2131221DNAArtificial SequenceSynthetic oligonucleotide 312cgctcttgtt
gcccaggctg g
2131325DNAArtificial SequenceSynthetic oligonucleotide 313cccaaaacag
actttctaga taacc
2531426DNAArtificial SequenceSynthetic oligonucleotide 314ttcaaattgc
tttttttcta ctcacc
2631524DNAArtificial SequenceSynthetic oligonucleotide 315gatctgaaaa
aagtgacagg ttgg
2431626DNAArtificial SequenceSynthetic oligonucleotide 316cactgaaatt
tgaaaggaac atatgg
2631721DNAArtificial SequenceSynthetic oligonucleotide 317tctggtgcag
tggcctctag g
2131824DNAArtificial SequenceSynthetic oligonucleotide 318accataagtg
gttttacctg atgg
2431921DNAArtificial SequenceSynthetic oligonucleotide 319cccaggcgca
ggtgattctc c
2132021DNAArtificial SequenceSynthetic oligonucleotide 320ggtggctcac
gcctgaaatc c
2132122DNAArtificial SequenceSynthetic oligonucleotide 321cacagtccac
gtgccacaat cc
2232224DNAArtificial SequenceSynthetic oligonucleotide 322aatcatgtta
acacatccct ctcc
2432324DNAArtificial SequenceSynthetic oligonucleotide 323gaagagagtg
ttgaaaggtt aagc
2432423DNAArtificial SequenceSynthetic oligonucleotide 324cgagaccata
ctggctaaga tgg
2332525DNAArtificial SequenceSynthetic oligonucleotide 325attagccaca
caataaatgt tctgg
2532625DNAArtificial SequenceSynthetic oligonucleotide 326tttgaaaagc
gttgcaatat gatgc
2532721DNAArtificial SequenceSynthetic oligonucleotide 327ggttgcagtg
agccgagatc g
2132820DNAArtificial SequenceSynthetic oligonucleotide 328ggtgggagga
ctgcctgagc
2032926DNAArtificial SequenceSynthetic oligonucleotide 329aacagagaga
aaaaacacaa attacc
2633026DNAArtificial SequenceSynthetic oligonucleotide 330gatatctaga
attcccaaat acttgg
2633128DNAArtificial SequenceSynthetic oligonucleotide 331gtgatagaat
taaaggaaaa aataaacg
2833226DNAArtificial SequenceSynthetic oligonucleotide 332attgttcctt
ttctaaatat tctacc
2633321DNAArtificial SequenceSynthetic oligonucleotide 333cagcactttg
ggaggctgag g
2133422DNAArtificial SequenceSynthetic oligonucleotide 334cacagaggtt
tcacagtgct gg
2233524DNAArtificial SequenceSynthetic oligonucleotide 335aacttctgct
tctgtccata atgc
2433622DNAArtificial SequenceSynthetic oligonucleotide 336gcctgtaatc
ccagcacttt gg
2233724DNAArtificial SequenceSynthetic oligonucleotide 337gccagtaaac
atatgaaaag gtgc
2433826DNAArtificial SequenceSynthetic oligonucleotide 338aattatgtaa
ataaagagtg aaaagg
2633924DNAArtificial SequenceSynthetic oligonucleotide 339cccctacaca
gaaaaaacaa ttcc
2434026DNAArtificial SequenceSynthetic oligonucleotide 340tgagtgtcaa
agaaaaatac aattgg
2634124DNAArtificial SequenceSynthetic oligonucleotide 341atacacagag
aaaatgagtc cacc
2434223DNAArtificial SequenceSynthetic oligonucleotide 342aacactcccc
ttctctgttt agc
2334324DNAArtificial SequenceSynthetic oligonucleotide 343gatattcttt
gcaacctagg atgc
2434426DNAArtificial SequenceSynthetic oligonucleotide 344ctctaaaact
aatcagcaat gtaacc
2634523DNAArtificial SequenceSynthetic oligonucleotide 345cacctgtaat
cccagcactt tgg
2334626DNAArtificial SequenceSynthetic oligonucleotide 346cgtaaaactg
ccacaaagct tgtagg
2634721DNAArtificial SequenceSynthetic oligonucleotide 347gtggcagagg
tgcaagcaag c
2134824DNAArtificial SequenceSynthetic oligonucleotide 348acagaaatga
caaacgcatg tacc
2434923DNAArtificial SequenceSynthetic oligonucleotide 349acactctctt
agctaggctt tgg
2335022DNAArtificial SequenceSynthetic oligonucleotide 350gagcttggaa
tagggcagtt cc
2235124DNAArtificial SequenceSynthetic oligonucleotide 351ctgggttctt
taaacatgtc cagg
2435223DNAArtificial SequenceSynthetic oligonucleotide 352tcaagaaagg
acactgcagt ggc
2335324DNAArtificial SequenceSynthetic oligonucleotide 353catgcacaca
aactatctca ttcc
2435420DNAArtificial SequenceSynthetic oligonucleotide 354tagccgggca
tggtggcacg
2035526DNAArtificial SequenceSynthetic oligonucleotide 355atcatgctga
ttgaatttca aatagc
2635621DNAArtificial SequenceSynthetic oligonucleotide 356ttggcatgca
gggcagtgac c
2135723DNAArtificial SequenceSynthetic oligonucleotide 357ggtggtgaga
taataacacc tgc
2335827DNAArtificial SequenceSynthetic oligonucleotide 358ttgctatata
ataatcattt gtgatcc
2735923DNAArtificial SequenceSynthetic oligonucleotide 359cggtaactgt
tactctggga tgg
2336022DNAArtificial SequenceSynthetic oligonucleotide 360aggctaggtt
cccttctctt cc
2236123DNAArtificial SequenceSynthetic oligonucleotide 361gtagtgccta
gcacagagaa agc
2336221DNAArtificial SequenceSynthetic oligonucleotide 362ctagcctggg
caacaagagc g
2136322DNAArtificial SequenceSynthetic oligonucleotide 363tctctctcct
ctctgggatc ag
2236425DNAArtificial SequenceSynthetic oligonucleotide 364gtttgaatat
ttgtatgcag caagc
2536526DNAArtificial SequenceSynthetic oligonucleotide 365tagaacaaat
tctggcttat aaaagc
2636623DNAArtificial SequenceSynthetic oligonucleotide 366ccactctacc
tttattcctt gcc
2336723DNAArtificial SequenceSynthetic oligonucleotide 367agaccagaat
atgcaagcag agg
2336822DNAArtificial SequenceSynthetic oligonucleotide 368ggacgttttg
ctggtgtctg cg
2236924DNAArtificial SequenceSynthetic oligonucleotide 369aaggaacaaa
ctgttgtcac atgc
2437022DNAArtificial SequenceSynthetic oligonucleotide 370atgtagctgg
gactacaggt gc
2237122DNAArtificial SequenceSynthetic oligonucleotide 371ggctcatgcc
tgtaatccca gc
2237225DNAArtificial SequenceSynthetic oligonucleotide 372atgaggtttt
cacacaaaaa gatgc
2537321DNAArtificial SequenceSynthetic oligonucleotide 373tgggcgacag
agcaagactc c
2137426DNAArtificial SequenceSynthetic oligonucleotide 374aaatgtccct
aaaagtgatc aacagc
2637524DNAArtificial SequenceSynthetic oligonucleotide 375cagactcagt
tttacctcat cagc
2437624DNAArtificial SequenceSynthetic oligonucleotide 376agtgatcttt
cctctttaac ctcc
2437722DNAArtificial SequenceSynthetic oligonucleotide 377ccagctattc
aggaggccaa gg
2237825DNAArtificial SequenceSynthetic oligonucleotide 378cttaaacatt
atgacactgt cttgc
2537921DNAArtificial SequenceSynthetic oligonucleotide 379ccaggtctat
gaggccgttc c
2138024DNAArtificial SequenceSynthetic oligonucleotide 380tccaaagcat
ccctacatta tacc
2438124DNAArtificial SequenceSynthetic oligonucleotide 381acatacatac
atgcagtgac tagc
2438221DNAArtificial SequenceSynthetic oligonucleotide 382tacaggtgcc
agccaccatg c
2138322DNAArtificial SequenceSynthetic oligonucleotide 383gcctgtaatc
ccagcactct gg
2238422DNAArtificial SequenceSynthetic oligonucleotide 384gacagagtcc
cactcttgtt gc
2238522DNAArtificial SequenceSynthetic oligonucleotide 385gtgccttcca
aagcagtgta gg
2238625DNAArtificial SequenceSynthetic oligonucleotide 386tatcttactg
ggtatgtata atgcc
2538723DNAArtificial SequenceSynthetic oligonucleotide 387caaaggaaat
acgtcctacc agg
2338823DNAArtificial SequenceSynthetic oligonucleotide 388ccttttctca
cagacatgct tcc
2338922DNAArtificial SequenceSynthetic oligonucleotide 389taaacacagt
gagcagaatc cc
2239025DNAArtificial SequenceSynthetic oligonucleotide 390ataaagcaaa
cttctaaaag ggtcc
2539121DNAArtificial SequenceSynthetic oligonucleotide 391accactacac
tccagcctgg g
2139223DNAArtificial SequenceSynthetic oligonucleotide 392gatacctggg
tcagagtaag tgc
2339323DNAArtificial SequenceSynthetic oligonucleotide 393tgtaatctca
gctacttggg agg
2339423DNAArtificial SequenceSynthetic oligonucleotide 394gtgtcgtctt
ctcttcctct acg
2339522DNAArtificial SequenceSynthetic oligonucleotide 395ctggctagta
tgaggttggt gc
2239623DNAArtificial SequenceSynthetic oligonucleotide 396ggactagcca
catttcaacc agg
2339725DNAArtificial SequenceSynthetic oligonucleotide 397gcagtatact
gagaatttag tttcc
2539821DNAArtificial SequenceSynthetic oligonucleotide 398gaggctgagg
caggagaatg g
2139924DNAArtificial SequenceSynthetic oligonucleotide 399cattgtttga
tgaaggtcaa cagc
2440022DNAArtificial SequenceSynthetic oligonucleotide 400cagacaagag
tggctacggc ag
2240122DNAArtificial SequenceSynthetic oligonucleotide 401acgcccagcc
agattattca gg
2240223DNAArtificial SequenceSynthetic oligonucleotide 402ggaaccagaa
agaagtgcaa agg
2340321DNAArtificial SequenceSynthetic oligonucleotide 403tgagccatct
tggaggcagg c
2140422DNAArtificial SequenceSynthetic oligonucleotide 404caggaccttc
ctacaaacct cc
2240524DNAArtificial SequenceSynthetic oligonucleotide 405aacacaacat
atctgacctt acgc
2440623DNAArtificial SequenceSynthetic oligonucleotide 406gccttagaag
tccagaggaa agc
2340722DNAArtificial SequenceSynthetic oligonucleotide 407tgacgtaccc
agtagacctt cc
2240822DNAArtificial SequenceSynthetic oligonucleotide 408ctctgcaagc
ctgggaaaca gg
2240922DNAArtificial SequenceSynthetic oligonucleotide 409gccttgtccc
caagtcctaa gg
2241022DNAArtificial SequenceSynthetic oligonucleotide 410gcaaagggac
tcctggaatt cc
2241121DNAArtificial SequenceSynthetic oligonucleotide 411gctcctgcct
gtaatcccag c
2141224DNAArtificial SequenceSynthetic oligonucleotide 412gaaggaaaca
gaaaaagcag aggc
2441324DNAArtificial SequenceSynthetic oligonucleotide 413cttactaccg
ttcttcttca ctgg
2441427DNAArtificial SequenceSynthetic oligonucleotide 414actattctgt
ttctttaggt ttactgc
2741522DNAArtificial SequenceSynthetic oligonucleotide 415cggtggctca
cacctgtaat cc
2241622DNAArtificial SequenceSynthetic oligonucleotide 416agccagagtt
ctgtgctcta gg
2241723DNAArtificial SequenceSynthetic oligonucleotide 417taatttgcat
ttcgtgccgc tcc
2341825DNAArtificial SequenceSynthetic oligonucleotide 418cacttttaat
acagatccca atagg
2541926DNAArtificial SequenceSynthetic oligonucleotide 419atgtattttt
tcttttcctg tcaagc
2642026DNAArtificial SequenceSynthetic oligonucleotide 420aaatgttaac
attattctcc ctaagg
2642122DNAArtificial SequenceSynthetic oligonucleotide 421catatgccca
gatcccgtct cc
2242221DNAArtificial SequenceSynthetic oligonucleotide 422acaggtgtga
gccgctgcac c
2142323DNAArtificial SequenceSynthetic oligonucleotide 423gccaagacgt
ttacagtttt ggc
2342423DNAArtificial SequenceSynthetic oligonucleotide 424aggaaacttc
tgaggatgat ggg
2342524DNAArtificial SequenceSynthetic oligonucleotide 425gctttatagg
gcagtctgaa ttcc
2442625DNAArtificial SequenceSynthetic oligonucleotide 426ttagaataaa
agttatctcg ggagg
2542725DNAArtificial SequenceSynthetic oligonucleotide 427taatttcttc
agctttatcc ctcag
2542826DNAArtificial SequenceSynthetic oligonucleotide 428cacatgacta
attctctatt cattcc
2642924DNAArtificial SequenceSynthetic oligonucleotide 429aaagacctca
agaaaagagt cacc
2443024DNAArtificial SequenceSynthetic oligonucleotide 430gacccataaa
gattatatgc ccag
2443124DNAArtificial SequenceSynthetic oligonucleotide 431aaagtactaa
tgcagtgtgt cagc
2443221DNAArtificial SequenceSynthetic oligonucleotide 432gaggttcctc
gattcccctg c
2143322DNAArtificial SequenceSynthetic oligonucleotide 433ggagagcaga
ggaattcaca gg
2243426DNAArtificial SequenceSynthetic oligonucleotide 434agtaattaga
aactgattct aagacg
2643523DNAArtificial SequenceSynthetic oligonucleotide 435cataccattg
ccaatccagt tcc
2343621DNAArtificial SequenceSynthetic oligonucleotide 436attacgggtg
cctgccactg c
2143721DNAArtificial SequenceSynthetic oligonucleotide 437cagccaggca
gaggagagag g
2143825DNAArtificial SequenceSynthetic oligonucleotide 438ttttcattcc
aagtttctgt ttggg
2543926DNAArtificial SequenceSynthetic oligonucleotide 439tttcaaatag
gaatttggat aatccc
2644022DNAArtificial SequenceSynthetic oligonucleotide 440taagccgaga
tcacaccact gc
2244123DNAArtificial SequenceSynthetic oligonucleotide 441ccttcagcgc
attatatctt ggc
2344225DNAArtificial SequenceSynthetic oligonucleotide 442ccatctaatc
catcttaaat tcacc
2544321DNAArtificial SequenceSynthetic oligonucleotide 443gagtggagac
tgcgccactg c
2144425DNAArtificial SequenceSynthetic oligonucleotide 444aatcatgtgc
caattaaacc atggc
2544520DNAArtificial SequenceSynthetic oligonucleotide 445cccagggacc
agaccagacc
2044623DNAArtificial SequenceSynthetic oligonucleotide 446ctcactcacc
agtgaaaatc agc
2344722DNAArtificial SequenceSynthetic oligonucleotide 447ggttgctctc
gaactcctga cc
2244821DNAArtificial SequenceSynthetic oligonucleotide 448gttcccccag
ctcctttctg c
2144923DNAArtificial SequenceSynthetic oligonucleotide 449agaaagatgt
agaagggtcc agc
2345024DNAArtificial SequenceSynthetic oligonucleotide 450gggaaaaggt
gtattatgca agcg
2445124DNAArtificial SequenceSynthetic oligonucleotide 451ctctctcaga
cctaatgcaa aagc
2445227DNAArtificial SequenceSynthetic oligonucleotide 452aactatacat
acagtatttg tattagc
2745325DNAArtificial SequenceSynthetic oligonucleotide 453aaattaatgc
aatccatgat ccagg
2545423DNAArtificial SequenceSynthetic oligonucleotide 454ctttctccac
tctaagagaa ccc
2345524DNAArtificial SequenceSynthetic oligonucleotide 455ttttggtgtg
ttcatattgg ctgc
2445624DNAArtificial SequenceSynthetic oligonucleotide 456gcttccacaa
atgacagaca aagg
2445722DNAArtificial SequenceSynthetic oligonucleotide 457ggctcatgct
tgtaatccca gc
2245826DNAArtificial SequenceSynthetic oligonucleotide 458catatgaatt
gttgttcctt tgtagg
2645923DNAArtificial SequenceSynthetic oligonucleotide 459cactggtaca
agtccaagag tcc
2346022DNAArtificial SequenceSynthetic oligonucleotide 460gaccctgtgt
ctacttcctg gg
2246126DNAArtificial SequenceSynthetic oligonucleotide 461tatttgaact
atctcttgaa atgtcc
2646225DNAArtificial SequenceSynthetic oligonucleotide 462ctgattaaaa
agtattaccc ttggc
2546325DNAArtificial SequenceSynthetic oligonucleotide 463tttgaaactg
cactcaataa cttgg
2546424DNAArtificial SequenceSynthetic oligonucleotide 464agtaatgtgt
catgatccaa tggc
2446524DNAArtificial SequenceSynthetic oligonucleotide 465gaaagcattt
cccaatgtct cacc
2446623DNAArtificial SequenceSynthetic oligonucleotide 466caatggacaa
aaggcccaac tgc
2346724DNAArtificial SequenceSynthetic oligonucleotide 467tccagctctg
gcttttttgt taag
2446821DNAArtificial SequenceSynthetic oligonucleotide 468acggagtctc
actccgtgac c
2146925DNAArtificial SequenceSynthetic oligonucleotide 469ctatgtcata
gtcaagagac tttgc
2547023DNAArtificial SequenceSynthetic oligonucleotide 470gttcaagcga
ttctcctgtc tcg
2347125DNAArtificial SequenceSynthetic oligonucleotide 471ccacctaata
cttaaatacg gaagc
2547227DNAArtificial SequenceSynthetic oligonucleotide 472atattcaaca
aacttaatag tgaagtg
2747322DNAArtificial SequenceSynthetic oligonucleotide 473ttacaggcgt
gagtcaccat gc
2247422DNAArtificial SequenceSynthetic oligonucleotide 474aacacctcca
agaggccaaa cg
2247527DNAArtificial SequenceSynthetic oligonucleotide 475tactattggc
aaatttcaat tatatgg
2747625DNAArtificial SequenceSynthetic oligonucleotide 476agcccacatc
ctaaaattca ataag
2547724DNAArtificial SequenceSynthetic oligonucleotide 477gaaagtggat
aagtgtttgt ctgg
2447821DNAArtificial SequenceSynthetic oligonucleotide 478ggccaggcat
tcaagaccag c
2147924DNAArtificial SequenceSynthetic oligonucleotide 479agccaacaac
aaaaagacac aacc
2448022DNAArtificial SequenceSynthetic oligonucleotide 480ttgagcccag
gagttcaaga cc
2248125DNAArtificial SequenceSynthetic oligonucleotide 481cagactaaag
atctcagaga gaaac
2548222DNAArtificial SequenceSynthetic oligonucleotide 482cgcttgtaat
cccagcactt gg
2248325DNAArtificial SequenceSynthetic oligonucleotide 483aaaagtgaaa
tcagaatttg tttcc
2548421DNAArtificial SequenceSynthetic oligonucleotide 484caggcgtgag
caactgtgtc c
2148522DNAArtificial SequenceSynthetic oligonucleotide 485ggtccagtag
gatctcgttt gc
2248626DNAArtificial SequenceSynthetic oligonucleotide 486actttgaaaa
tgttgttata gctggg
2648723DNAArtificial SequenceSynthetic oligonucleotide 487ttccctgcat
ctaagtcttc tcc
2348825DNAArtificial SequenceSynthetic oligonucleotide 488agatatctac
cattgaagag tttgc
2548924DNAArtificial SequenceSynthetic oligonucleotide 489agtcttcact
tcactttgtt gtcc
2449025DNAArtificial SequenceSynthetic oligonucleotide 490ccatgcaggt
atgaaatata aaagc
2549121DNAArtificial SequenceSynthetic oligonucleotide 491tgggtgacag
agtgagactc c
2149223DNAArtificial SequenceSynthetic oligonucleotide 492acagcaatac
cgggttaaca tgc
2349325DNAArtificial SequenceSynthetic oligonucleotide 493tttatgtaaa
agatgaatgc gaggc
2549422DNAArtificial SequenceSynthetic oligonucleotide 494ctactctgct
actgggaaca gg
2249524DNAArtificial SequenceSynthetic oligonucleotide 495caaacgttag
tctggcaaaa tgcg
2449620DNAArtificial SequenceSynthetic oligonucleotide 496tgcacgctac
cacacccagc
2049726DNAArtificial SequenceSynthetic oligonucleotide 497aattcttgga
tctgtgtgtt tactgc
2649824DNAArtificial SequenceSynthetic oligonucleotide 498taccagttat
cattctcttt ctgc
2449920DNAArtificial SequenceSynthetic oligonucleotide 499atccacccac
ctcggcctcc
2050022DNAArtificial SequenceSynthetic oligonucleotide 500cactctgcct
ggcccttaat gg
2250126DNAArtificial SequenceSynthetic oligonucleotide 501atagtttgtt
taatatgcca ctaagg
2650220DNAArtificial SequenceSynthetic oligonucleotide 502gcgtgagcca
ccgcacctgg
2050325DNAArtificial SequenceSynthetic oligonucleotide 503ctccatcaca
caaattttat gtggc
2550422DNAArtificial SequenceSynthetic oligonucleotide 504agacggagtc
tcgttctgtc gc
2250522DNAArtificial SequenceSynthetic oligonucleotide 505tcccaggttc
aagccattct cc
2250624DNAArtificial SequenceSynthetic oligonucleotide 506tattttgaga
gtctcactct gtcg
2450722DNAArtificial SequenceSynthetic oligonucleotide 507gtctcgaact
cctgacctca gg
2250823DNAArtificial SequenceSynthetic oligonucleotide 508aaggaggtga
agagtgaact acg
2350924DNAArtificial SequenceSynthetic oligonucleotide 509gtctcaggtt
ttggacttac ttgg
2451026DNAArtificial SequenceSynthetic oligonucleotide 510tttacagatc
ttaaatgcat taggac
2651123DNAArtificial SequenceSynthetic oligonucleotide 511gtacactgaa
caaaggagac agg
2351224DNAArtificial SequenceSynthetic oligonucleotide 512ctggtagtaa
tgcaaaatag cacc
2451327DNAArtificial SequenceSynthetic oligonucleotide 513catttaatgt
gaaatgaatt ataagcc
2751423DNAArtificial SequenceSynthetic oligonucleotide 514gagacagggt
ttcactatgt tgg
2351522DNAArtificial SequenceSynthetic oligonucleotide 515ccagcacttt
ggaaggctga gg
2251625DNAArtificial SequenceSynthetic oligonucleotide 516gaaaccaagt
atcatggtaa attgc
2551721DNAArtificial SequenceSynthetic oligonucleotide 517cagtgagggc
tgctcagttc c
2151820DNAArtificial SequenceSynthetic oligonucleotide 518gccaggtgcg
gtggctcacg
2051922DNAArtificial SequenceSynthetic oligonucleotide 519catgcctgta
atcccagcta cc
2252025DNAArtificial SequenceSynthetic oligonucleotide 520atgtaaatgg
tacagtcact ttagg
2552124DNAArtificial SequenceSynthetic oligonucleotide 521cccacaatac
agagaactct tacc
2452222DNAArtificial SequenceSynthetic oligonucleotide 522tgaaacatgc
agcccagtgt cc
2252324DNAArtificial SequenceSynthetic oligonucleotide 523tgttttttct
cctgccttca atcc
2452422DNAArtificial SequenceSynthetic oligonucleotide 524gctttcctgg
gtctccatct gg
2252524DNAArtificial SequenceSynthetic oligonucleotide 525gcagccgctt
gaaaacaaaa cagc
2452623DNAArtificial SequenceSynthetic oligonucleotide 526gatcacgtta
catttggggg tgg
2352726DNAArtificial SequenceSynthetic oligonucleotide 527taggctgaaa
aactaaaatt tgttgc
2652822DNAArtificial SequenceSynthetic oligonucleotide 528ctcctttggg
ctcctttagt cc
2252920DNAArtificial SequenceSynthetic oligonucleotide 529gcctcggcct
cccaaagtgc
2053023DNAArtificial SequenceSynthetic oligonucleotide 530aatgcctaga
gagatttggc agg
2353123DNAArtificial SequenceSynthetic oligonucleotide 531gagatggggt
ttcactatgt tgg
2353222DNAArtificial SequenceSynthetic oligonucleotide 532tgtgatcttg
ccactgcact cc
2253324DNAArtificial SequenceSynthetic oligonucleotide 533acttctcctc
catttgtttc ttcg
2453421DNAArtificial SequenceSynthetic oligonucleotide 534cgtgcccggg
ctcagttcta c
2153526DNAArtificial SequenceSynthetic oligonucleotide 535ccaaaacaat
aaaatcacaa tttggg
2653624DNAArtificial SequenceSynthetic oligonucleotide 536ctgaactgcc
ttagagtaaa tccg
2453724DNAArtificial SequenceSynthetic oligonucleotide 537atttctgtat
caggtctgtg ttcc
2453822DNAArtificial SequenceSynthetic oligonucleotide 538ggctgacccc
ttcactgttt cc
2253923DNAArtificial SequenceSynthetic oligonucleotide 539caaaaattag
ccaggcatgg tgg
2354021DNAArtificial SequenceSynthetic oligonucleotide 540gcagtgagca
gtgatcgcac c
2154125DNAArtificial SequenceSynthetic oligonucleotide 541aaagactgtg
aactaacttg tttgc
2554225DNAArtificial SequenceSynthetic oligonucleotide 542tgccaagaat
tacacattat taggc
2554323DNAArtificial SequenceSynthetic oligonucleotide 543ggccaggatg
tcattaactt tcc
2354423DNAArtificial SequenceSynthetic oligonucleotide 544gtaagagctg
acgtgtattg tgc
2354520DNAArtificial SequenceSynthetic oligonucleotide 545cccggtgagg
ccgcacatcc
2054620DNAArtificial SequenceSynthetic oligonucleotide 546cctgcgcctt
aaccccctcc
2054719DNAArtificial SequenceSynthetic oligonucleotide 547cggcgcctag
gggccatcg
1954824DNAArtificial SequenceSynthetic oligonucleotide 548acttaaggaa
acgaacatga cacc
2454922DNAArtificial SequenceSynthetic oligonucleotide 549gagaccgagt
cttgctgtgt cg
2255027DNAArtificial SequenceSynthetic oligonucleotide 550gtattaattg
aagatgattt ggaatgc
2755124DNAArtificial SequenceSynthetic oligonucleotide 551tctttaaaag
actatcgctg aggc
2455224DNAArtificial SequenceSynthetic oligonucleotide 552aaaagagaca
tcagtagagc atcc
2455324DNAArtificial SequenceSynthetic oligonucleotide 553gttcatgttt
tctttgacgt ctcc
2455423DNAArtificial SequenceSynthetic oligonucleotide 554tttcgaaagt
tcaggctgag tgc
2355524DNAArtificial SequenceSynthetic oligonucleotide 555gaccctcaaa
acaatcctct aagg
2455625DNAArtificial SequenceSynthetic oligonucleotide 556caaaacacac
ttagaaacaa actgc
2555722DNAArtificial SequenceSynthetic oligonucleotide 557gcctgggcga
catagtgaga cc
2255821DNAArtificial SequenceSynthetic oligonucleotide 558ggcaggagaa
tggcgtgaac c
2155921DNAArtificial SequenceSynthetic oligonucleotide 559tttgctcgtt
gcccaggctg g
2156026DNAArtificial SequenceSynthetic oligonucleotide 560gcaacttaat
gtgatagaat aatagc
2656120DNAArtificial SequenceSynthetic oligonucleotide 561cctccccttc
tgctgccagc
2056224DNAArtificial SequenceSynthetic oligonucleotide 562ccacaacaat
gtaaactcct ctgg
2456323DNAArtificial SequenceSynthetic oligonucleotide 563tactctccct
agagttcgtt ccc
2356421DNAArtificial SequenceSynthetic oligonucleotide 564gggtccccct
ttggccattc c
2156523DNAArtificial SequenceSynthetic oligonucleotide 565gatcttggct
cacttcaacc tcc
2356622DNAArtificial SequenceSynthetic oligonucleotide 566aggggaaata
tttaaacctt gg
2256724DNAArtificial SequenceSynthetic oligonucleotide 567aatgcaatgg
tgcatttaca gagg
2456826DNAArtificial SequenceSynthetic oligonucleotide 568tcattttatc
tatttctaca tggtcc
2656922DNAArtificial SequenceSynthetic oligonucleotide 569ggaagggaaa
tgcccatgaa cc
2257023DNAArtificial SequenceSynthetic oligonucleotide 570agtgaacatt
ttctgcagcc tcc
2357121DNAArtificial SequenceSynthetic oligonucleotide 571caacaggacg
tcaggcgatc c
2157222DNAArtificial SequenceSynthetic oligonucleotide 572ccttcaggct
gtcctgaaaa gg
2257321DNAArtificial SequenceSynthetic oligonucleotide 573agtctcactc
catcgcccag g
2157424DNAArtificial SequenceSynthetic oligonucleotide 574actgtgaaca
gtagttaact cagg
2457522DNAArtificial SequenceSynthetic oligonucleotide 575gcatgcctgt
aatccaagct gc
2257625DNAArtificial SequenceSynthetic oligonucleotide 576gaaacaattc
tcttttcaca cttgc
2557722DNAArtificial SequenceSynthetic oligonucleotide 577ggctcatgcc
tgttatccca gc
2257825DNAArtificial SequenceSynthetic oligonucleotide 578agaagaagct
tagtcatatg tttgg
2557924DNAArtificial SequenceSynthetic oligonucleotide 579cagatgcttg
agccaaacaa atgg
2458022DNAArtificial SequenceSynthetic oligonucleotide 580ctggcagaca
gagtgagact cc
2258127DNAArtificial SequenceSynthetic oligonucleotide 581aatgtgtgaa
tattattcat tacaggg
2758223DNAArtificial SequenceSynthetic oligonucleotide 582gcaggagaat
tgcttgaacc tgg
2358327DNAArtificial SequenceSynthetic oligonucleotide 583ctttagtcaa
attaaaacag tctatcc
2758423DNAArtificial SequenceSynthetic oligonucleotide 584gatttctatc
tcctgcaacc acc
2358526DNAArtificial SequenceSynthetic oligonucleotide 585ttcttgtgta
actactaaaa atctcc
2658624DNAArtificial SequenceSynthetic oligonucleotide 586aaagggtctt
cataaggcta atgg
2458727DNAArtificial SequenceSynthetic oligonucleotide 587ctcttaagga
ttatttatat gaagacc
2758818DNAArtificial SequenceSynthetic oligonucleotide 588caggaggagc
cccagagc
1858920DNAArtificial SequenceSynthetic oligonucleotide 589tcctggggat
ggttggatgc
2059023DNAArtificial SequenceSynthetic oligonucleotide 590tgaccccaca
gagtttacac agc
2359119DNAArtificial SequenceSynthetic oligonucleotide 591agtcagggca
ggctctgcc
1959223DNAArtificial SequenceSynthetic oligonucleotide 592tattttggcc
ccatccagaa agc
2359323DNAArtificial SequenceSynthetic oligonucleotide 593cacccagagt
acagctttgt tcc
2359419DNAArtificial SequenceSynthetic oligonucleotide 594gaggagcccc
agagcctgc
1959522DNAArtificial SequenceSynthetic oligonucleotide 595tggggatggt
tggatgctta cc
2259623DNAArtificial SequenceSynthetic oligonucleotide 596cccacagagt
ttacacagct tgc
2359719DNAArtificial SequenceSynthetic oligonucleotide 597caggctctgc
ccactcacc
1959821DNAArtificial SequenceSynthetic oligonucleotide 598ccatccagaa
agcccaaagc c
2159926DNAArtificial SequenceSynthetic oligonucleotide 599ccagagtaca
gctttgttcc tcattc
2660022DNAArtificial SequenceSynthetic oligonucleotide 600gcagtacaaa
caacgcacag cg
2260120DNAArtificial SequenceSynthetic oligonucleotide 601ctgccaccct
ccacagtccc
2060218DNAArtificial SequenceSynthetic oligonucleotide 602gccaagacca
tgcatgcg
1860321DNAArtificial SequenceSynthetic oligonucleotide 603cccagggaca
aagagactcc c
2160426DNAArtificial SequenceSynthetic oligonucleotide 604caggaagcag
acagtcttct agttcc
2660523DNAArtificial SequenceSynthetic oligonucleotide 605tgcctgtaat
cccaacactt tgg
2360619DNAArtificial SequenceSynthetic oligonucleotide 606tccctctggc
caggatggg
1960722DNAArtificial SequenceSynthetic oligonucleotide 607atggggaatg
ggagtaggaa gc
2260822DNAArtificial SequenceSynthetic oligonucleotide 608cagatcagtt
ctcccctcca gc
2260926DNAArtificial SequenceSynthetic oligonucleotide 609acaaaaaaga
aacatgctca gagagg
2661022DNAArtificial SequenceSynthetic oligonucleotide 610tggtggcatg
catctgtagt cc
2261126DNAArtificial SequenceSynthetic oligonucleotide 611aggtgctcta
tagatgttag catccc
2661222DNAArtificial SequenceSynthetic oligonucleotide 612ccaggacagg
atggagatct gg
2261324DNAArtificial SequenceSynthetic oligonucleotide 613agggaacctg
tgcattatcc ttgc
2461425DNAArtificial SequenceSynthetic oligonucleotide 614cagaagtctt
gctttaagga ggagg
2561522DNAArtificial SequenceSynthetic oligonucleotide 615gggtacgtga
aactcaccaa gg
2261620DNAArtificial SequenceSynthetic oligonucleotide 616cagagtgtgg
caagcaaggg
2061728DNAArtificial SequenceSynthetic oligonucleotide 617aacattttaa
aggtacaaat aacgtggg
2861822DNAArtificial SequenceSynthetic oligonucleotide 618tagggagcaa
cagccattaa gc
2261920DNAArtificial SequenceSynthetic oligonucleotide 619ggtgcactgt
ccagctctgg
2062021DNAArtificial SequenceSynthetic oligonucleotide 620actctcgctg
aactcgcctg g
2162120DNAArtificial SequenceSynthetic oligonucleotide 621ctcggtctct
ggtggtacgc
2062219DNAArtificial SequenceSynthetic oligonucleotide 622gcaagaggtc
cgagctggg
1962326DNAArtificial SequenceSynthetic oligonucleotide 623ggaagaagtg
aaacaagaga tgaagg
2662423DNAArtificial SequenceSynthetic oligonucleotide 624cccagagaac
aaaccggatt agg
2362523DNAArtificial SequenceSynthetic oligonucleotide 625cccttcaacc
ttctccaatc tgc
2362621DNAArtificial SequenceSynthetic oligonucleotide 626cccatgtcca
gtggtttagg g
2162722DNAArtificial SequenceSynthetic oligonucleotide 627gagattggtg
ggagacagat gg
2262823DNAArtificial SequenceSynthetic oligonucleotide 628cttctcagct
caaagttcca gcg
2362922DNAArtificial SequenceSynthetic oligonucleotide 629gaatgggaga
gatgaccaga gg
2263020DNAArtificial SequenceSynthetic oligonucleotide 630aagggcaagg
gggtatgtgg
2063121DNAArtificial SequenceSynthetic oligonucleotide 631ggaaggaagc
atgggaacac c
2163223DNAArtificial SequenceSynthetic oligonucleotide 632ccatcaatgc
tctgtctgtc tgg
2363319DNAArtificial SequenceSynthetic oligonucleotide 633gtgccgtgac
tgtgcttgg
1963423DNAArtificial SequenceSynthetic oligonucleotide 634acatcccatt
gacctcatca agc 23
User Contributions:
comments("1"); ?> comment_form("1"); ?>Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
User Contributions:
Comment about this patent or add new information about this topic: