Patent application title: USE OF DECORINE FOR INCREASING MUSCLE MASS
Inventors:
Antoine Kichler (Mennecy, FR)
Daniel Scherman (Paris, FR)
Daniel Scherman (Paris, FR)
Assignees:
ASSOCIATION FRANCAISE CONTRE LES MYOPATHIES
IPC8 Class: AA61K3814FI
USPC Class:
514 209
Class name: Designated organic active ingredient containing (doai) peptide (e.g., protein, etc.) containing doai glycopeptide utilizing
Publication date: 2012-03-08
Patent application number: 20120058955
Abstract:
The invention concerns decorin for increasing muscle mass, particularly
in the treatment of muscular dystrophies.Claims:
1. Composition containing a fragment of decorin able to bind zinc to
treat diseases associated with muscle wasting.
2. Composition according to claim 1, characterised in that the diseases are selected from the group of neuromuscular diseases, to advantage muscular dystrophies such as Duchenne muscular dystrophy, and the cachexia.
3. Composition according to claim 1 or 2, characterised in that the fragment comprises the sequence SEQ ID NO: 7 or 15.
4. Composition according to claim 1 or 2, characterised in that the fragment has the sequence SEQ ID NO: 7 or 15.
5. Composition containing decorin to increase muscle mass.
6. Composition according to claim 5, characterised in that the aim of increasing muscle mass is to compensate for wasting resulting from immobilisation or old age.
7. Composition according to claim 5 or 6, characterised in that it is for use in animals.
8. Composition according to one of claim 5 or 6, characterised in that decorin is in the form of an active fragment.
9. Composition according to claim 8, characterised in that the active fragment is able to bind zinc.
10. Composition according to claim 9, characterised in that the fragment comprises the sequence SEQ ID NO: 7 or 15.
11. Composition according to claim 9, characterised by the fragment has the sequence SEQ ID NO: 7 or 15.
12. Composition according to claim 1 or 5, characterised in that the decorin is in the form of an active fragment, a recombinant protein, a fusion protein or a nucleic acid encoding such a protein or fragment.
13. Composition according to claim 1 or 5, characterised in that the decorin is for intramuscular, intraperitoneal or intravenous injection.
14. Composition according to claim 1 or 5, characterised in that the decorin is associated with other treatments, particularly gene therapy and cell grafting.
Description:
TECHNICAL DOMAIN
[0001] The aim of this invention is to increase muscle mass in humans or animals.
[0002] More specifically, it advocates the use of decorin to develop muscle mass, particularly for treating pathological conditions associated with muscular wasting, such as muscular dystrophy.
PRIOR STATE OF THE ART
[0003] Neuromuscular diseases include various conditions that are generally associated with temporary or permanent loss of muscular strength. This loss of strength is usually accompanied by muscular wasting, also known as amyotrophia.
[0004] Myopathies, which involve damage to the actual muscle fibres, are an important group of these muscular diseases, and among them, progressive muscular dystrophies are characterised by a decrease in muscular strength, generally with atrophy of the muscles, as well as abnormalities in the muscle biopsy showing modifications of the tissue. This group notably includes Duchenne muscular dystrophy (or DMD), Becker muscular dystrophy (or BMD) and the limb girdle muscular dystrophies.
[0005] Associated genetic abnormalities have been identified for some of these diseases. Duchenne or Becker muscular dystrophies are related to alterations in the gene encoding dystrophin, type 2A limb girdle muscular dystrophy (LGMD 2A or calpainopathy) to alterations in the calpain 3 gene, while the sarcoglycanopathies or the dystrophy types LGMD 2C, LGMD 2D, LGMD 2E, LGMD 2F are related to defects in the γ-, α-, β- and δ-sarcoglycan genes respectively (McNally E M, Pytel P, Muscle diseases: the muscular dystrophies. Annu Rev Pathol. 2007; Vol 2: 87-109).
[0006] In these particular cases, different gene therapy strategies are being developed but are difficult to put into practice.
[0007] Nevertheless and more generally in all cases of muscular wasting, there is a clear need to develop technical solutions to increase muscle mass and/or volume.
[0008] The document WO 2005/094446 identified antibodies against an epitope located between residues 40 and 64 of mature human myostatin which could increase muscle mass. However, this strategy based on the recognition of myostatin by an antibody is not free of problems. Alternative solutions therefore need to be found.
[0009] The present invention is based on the discovery by the inventors of this property of decorin.
[0010] Decorin belongs to the SLRP (Small Leucine-Rich Proteoglycan) family of proteins and includes an LRR (Leucine-Rich Repeat) segment. Decorin is a member of class I of the SLRPs. The members of this family are secreted with a propeptide which, in some cases, is cleaved. Decorin also has a glycosaminoglycan (GAG) chain.
[0011] Decorin is a protein of the extracellular matrix, with a similar structure to that of the protein biglycan. It plays a role in assembling the matrix and interacts with various partners, such as type I, II, III and IV collagen, or TGF-beta (Ameye L, Young M F, Mice deficient in small leucine-rich proteoglycans: novel in vivo models for osteoporosis, osteoarthritis, Ehlers-Danlos syndrome, muscular dystrophy, and corneal diseases. Glycobiology 2002; Vol 12:107 R-116R; Reed C C, Iozzo R V, The role of decorin in collagen fibrillogenesis and skin homeostasis. Glycoconj J. 2002; Vol 19(4-5): 249-55).
[0012] On the basis of its interaction with TGF-beta, WO 96/25178 proposed the use of decorin to treat diseases associated with tissue fibrosis, i.e. excessive production of extracellular matrix, without relating this to the muscle mass problem.
DETAILED DESCRIPTION OF THE INVENTION
[0013] The present invention thus concerns the use of decorin to counter muscle wasting and even to increase muscle mass.
[0014] For this invention, the term "muscle mass" could be replaced either by muscle weight or volume.
[0015] More precisely the invention also concerns: [0016] a composition containing decorin to treat diseases associated with muscle wasting; [0017] the use of decorin in the preparation of a medicinal product for treating diseases associated with muscle wasting; [0018] a composition containing decorin to increase muscle mass; [0019] the use of decorin to increase muscle mass.
[0020] There are a number of conditions in which muscle wasting occurs.
[0021] Firstly, it may result from pathological conditions, particularly in the case of neuromuscular diseases. Duchenne muscular dystrophy is a disease particularly targeted, but all forms of neuromuscular diseases, especially muscular dystrophies, can be treated.
[0022] In addition, cachexia or marasmus is also a medical condition targeted by this invention. This state is characterized by extreme thinness, especially of the muscles, caused by prolonged illness or inadequate calorie or protein intake.
[0023] This condition is particularly seen in cases of chronic diseases such as cancer or AIDS or in individuals with either heart failure, where there is atrophy of skeletal muscles in 68% of patients, or urinary incontinence.
[0024] Although not actually considered as pathological, some situations are associated with muscle wasting: ageing, prolonged immobilisation etc. Here again, therefore, there is a reason for increasing the muscle mass.
[0025] The invention also offers the possibility particularly in the area of food production of increasing animal meat production. The use of decorin is therefore of particular interest in animals.
[0026] This invention is based therefore on detection of the stimulating properties of decorin, particularly related to muscle volume.
[0027] In the invention, "decorin" is used generically to mean the protein described by Krusius et al. (Krusius T., Ruoslahti E., Primary structure of an extracellular matrix proteoglycan core protein deduced from cloned cDNA. Proc Natl Acad Sci USA 1986; Vol 83(20): 7683-87). The human protein described in this document has the sequence SEQ ID NO: 1. It is in the form of a preproprotein of 359 amino acids. Both native proteins and those deprived of their propeptide and/or their signal sequence (329 aa), are covered by this invention.
[0028] Although decorin naturally has a glycosaminoglycan (GAG) chain, a decorin without GAG (GAG-) can also be used in the context of this invention. This can, for example, be obtained by enzyme treatment.
[0029] The decorin can be obtained from any organism, but in this invention, decorin of human origin is preferred. More generally and advantageously, the protein comes from the same organism as the organism into which it will be administered. Preferably therefore, for therapeutic indications in humans, human decorin is used to advantage.
[0030] One of the primary benefits indeed of the solution proposed in this invention is that decorin is a protein naturally present in mammals, especially humans, and therefore a priori is not likely to cause side effects or immune responses.
[0031] It has also been shown for human decorin that transcriptional variants exist (variants b, c, d and e), resulting in protein isoforms, of sequence SEQ ID NO: 2, 3, 4 and 5, respectively, included in this submission.
[0032] In the context of the invention, the term "decorin" thus has a wide meaning and covers: [0033] the native protein, particularly the sequence SEQ ID NO: 1; [0034] the protein with or without the GAG chain (GAG+ or GAG-, respectively); [0035] the protein lacking the propeptide and/or the signal sequence; [0036] variants of these proteins, especially embodied in the sequences SEQ ID NOS: 2 to 5; [0037] more generally active fragments of these proteins, [0038] or active derivatives or functional equivalents.
[0039] As far as the fragments or derivatives are concerned, they are, of course, active fragments or derivatives. The activity in question which these fragments or derivatives must possess concerns the ability to increase muscle mass, which is easily assessed by using the test described in this submission.
[0040] In practice, they are to advantage 60% identical to one of sequences SEQ ID NO: 1 to 7, even more advantageously 70%, 80% or 90% identical.
[0041] Thus, by way of example for the derivatives, it could be sequence SEQ ID NO: 6 corresponding to the murine protein of 354 aa, which is 80% identical to the human sequence SEQ ID NO: 1.
[0042] According to a preferred embodiment of the invention, the decorin is in the form of an active fragment. To advantage by "fragment" we mean a peptide containing less than 100 amino acids, to even greater advantage, less than 50 amino acids.
[0043] The use of a peptide instead of the protein has certain advantages, particularly in terms of its production but also concerning the possible risk of undesirable interference in vivo.
[0044] It has been shown as part of this application that a 41 residue fragment of the N-terminal part of murine decorin (SEQ ID NO: 7) corresponding to residues 31-71 of the sequence SEQ ID NO: 6, reported to fix zinc (Yang V W, LaBrenz S R, Rosenberg L C, McQuillan D, Hook M. Decorin is a Zn2+metalloprotein. J Biol Chem. 1999, 274(18): 12454-60), had the required activity. It has also been shown that an even smaller fragment of 30 residues (residues 42 to 71 of the sequence SEQ ID NO: 6 corresponding to SEQ ID NO: 15) was also active.
[0045] The corresponding domain, present in human decorin, can be easily determined by the methodology described in this document. Such a fragment may for example have the sequence SEQ ID NO: 16.
[0046] More generally, the invention therefore concerns the use of a fragment of decorin including the zinc binding domain, in practice the residues 31 to 71, possibly 42 to 71 of the murine sequence. In a particular embodiment, the sequence of the fragment in question corresponds to sequence SEQ ID NO: 7 or SEQ ID NO: 15. In addition, fragments which are to advantage 50% identical to SEQ ID NO: 7 or 15, or even more advantageously 60%, 70%, 80% or 90% identical to them, and which retain their ability to bind zinc, are also covered by this invention.
[0047] In addition, decorin, its fragments and active derivatives may also be in the form of fusion proteins or chimeric proteins with another protein fragment at their N- or C-terminal ends, which can, for example, but without being limited to this, increase the residence time of the protein in the organism. A preferred example is the chimera consisting of the constant region of mammalian IgGs, attached via a hinge sequence to decorin or one of its fragments. Another example is human or mammalian albumin, also attached to decorin or to a protein fragment of decorin. Such combinations can be obtained both from a recombinant cDNA and by chemical bonding of the 2 proteins.
[0048] The present invention is therefore based on an exogenous supply of decorin. In fact, the composition covered by the invention consists of either the protein as such or a system producing the protein.
[0049] As far as the protein itself is concerned, it could be either native decorin, purified from an organism naturally producing this protein, or a recombinant protein produced by any of the synthesis systems available and known to those working in the field.
[0050] Alternatively, a nucleic acid sequence encoding decorin is put into an expression system, to advantage under the control of a promoter in a vector. After introduction into the body, the decorin is produced in vivo. The transfer of the nucleic acids (DNA or RNA) can be done either with viral approaches to gene transfer (e.g. adeno-associated virus or AAV) or with non-viral approaches (e.g. by simple intramuscular injection of a plasmid). Genomic DNA may be of interest since in some cases, the presence of introns stabilises the prespliced mRNA and improves its stability in the nucleus and its export, which leads to better protein expression.
[0051] Decorin, its derivatives or fragments, can thus be provided in the form of nucleic acids, particularly DNA or RNA, and may for example be in the form of transcripts occurring naturally in humans or the mouse. The following sequences are preferred: [0052] Sequence SEQ ID NO: 8, corresponding to the A1 variant (Accession Number NM--001920.3), which is the longest transcript and encodes the isoform a of the human decorin sequence SEQ ID NO: 1 (Accession Number NP--001911); [0053] Sequence SEQ ID NO: 9, corresponding to the A2 variant (Accession Number NM--133503.2), which uses an alternative exon at the 5'UTR compared with the variant A1 and encodes the same protein sequence SEQ ID NO: 1 (Accession Number NP--598010.1); [0054] Sequence SEQ ID NO: 10, corresponding to the B variant (Accession Number NM--133504.2), which lacks exons 3 and 4 in the coding region, compared with the A1 variant. This causes no change in reading frame but codes for an isoform b of the protein, which lacks an internal fragment of 109 aa, and has the sequence SEQ ID NO: 2 (Accession Number NP--598011.1); [0055] Sequence SEQ ID NO: 11, corresponding to the C variant (Accession Number NM--133505.2), which lacks exons 3, 4 and 5 in the coding region, compared with the A1 variant. This causes a change of internal reading frame and the isoform c encoded of SEQ ID NO: 3 (Accession Number NP--598012.1) is shorter than isoform a by 147 amino acids; [0056] Sequence SEQ ID NO: 12, corresponding to the D variant (Accession Number NM--133506.2), which lacks exons 4, 5, 6 and 7 in the coding region, compared with the A1 variant. This causes no change in reading frame but codes for an isoform d of the protein, which lacks an internal fragment of 187 aa, and has the sequence SEQ ID NO: 4 (Accession Number NP--598013.1); [0057] Sequence SEQ ID NO: 13, corresponding to the E variant (Accession Number NM--133507.2), which lacks exons 3, 4, 5, 6 and 7 in the coding region, compared with the A1 variant. This causes a change of internal reading frame and the isoform e encoded of SEQ ID NO: 5 (Accession Number NP--598014.1) is shorter than isoform a by 284 amino acids; [0058] Sequence SEQ ID NO: 14, encoding the murine protein sequence SEQ ID NO: 6 (Accession Number P28654).
[0059] As already mentioned, decorin is known to be a zinc metalloprotein. Owing to this and in order to potentiate its activity, one could choose to provide additional zinc to that naturally available in the organism to which the decorin is administered. Thus, according to this embodiment, the composition containing the decorin also includes zinc, e.g. as zinc chloride, preferably at a concentration between 1 and 50 μM, even equal to 15 μM.
[0060] A composition containing decorin according to the invention for the treatment of diseases associated with muscle wasting or intended to increase muscle mass may also contain any acceptable compound or excipient, particularly a pharmaceutical compound or excipient. The route of administration may be intramuscular or intravenous, or even subcutaneous, intraperitoneal or oral.
[0061] To promote the engraftment of precursor cells or stem cells, it may be advantageous to combine the administration of decorin with the cell grafts (myoblasts, stem cells etc.). This administration can be simultaneous or separated in time.
[0062] It can also be advantageous to combine gene therapy for the treatment of a neuromuscular disease with administration of decorin. In a preferred embodiment, a therapeutic gene is associated with decorin treatment. Administration of the two treatments can be simultaneous or separated in time.
[0063] The beneficial effects of decorin result in an increase in muscle volume (either mass or weight), due to an increase in the area of fibres possibly associated with an increase in the number of fibres. These positive effects can be observed in the various different skeletal muscles, both in an organism with a disease affecting its muscle mass and in a healthy individual. In principle, there are no side effects and no immune reaction.
EXAMPLES OF EMBODIMENTS
[0064] The invention and the advantages resulting from it are better illustrated by the following examples of embodiments and the attached figures. These are in no way limiting.
[0065] The invention is further illustrated by means of recombinant mouse decorin injected intramuscularly into mdx mice with a gene encoding an altered dystrophin and serving as an experimental model of Duchenne muscular dystrophy, and gamma-sarcoglycan-/-mice (mouse model of sarcoglycanopathies on a pure C57/B16 background).
LEGENDS TO THE FIGURES
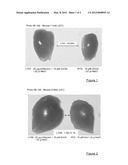
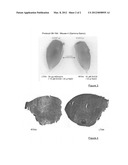
[0066] FIG. 1 is a view of the tibialis anterior muscle taken from mdx mouse 7 that had received (on the left) or not (on the right) an intramuscular injection of decorin.
[0067] FIG. 2 is a view of the tibialis anterior muscle taken from mdx mouse 8 that had received (on the left) or not (on the right) an intramuscular injection of decorin.
[0068] FIG. 3 is a view of the tibialis anterior muscle taken from gamma-sarcoglycan-/-mouse 4 at D18 that had received (on the left) or not (on the right) an intramuscular injection of decorin.
[0069] FIG. 4 is a view of a cross-section of the tibialis anterior muscle taken from gamma-sarcoglycan-/-mouse 4 at D18 that had received (LTA4 on the right of the figure) or not (RTA4 on the left of the figure) an intramuscular injection of decorin.
I) MATERIALS AND METHODS
[0070] Preparing the mDecorin Solution
[0071] The protein used was recombinant mouse decorin (mDecorin) of sequence SEQ ID NO: 6, provided by R&D Systems.
[0072] Twenty-four to forty hours before the injection, 100 μl of 150 mM sterile NaCl and 6 μl of 250 μM ZnCl2 were added to 100 μg of mDecorin powder. The final volume was 106 μl with a final concentration of approximately 1 μg/μl. For the injections into the control muscles a mixture was also prepared of 100 μl of 150 mM NaCl and 6 μl of 250 μM ZnCl2. All these solutions, after being vortexed, were stored at 4° C.
In Vivo Injection
[0073] All the mice were treated according to EU directives on human health and the use of experimental animals.
[0074] mdx dystrophic (S-linked muscular dystrophy) or gamma-sarcoglycan-/-mice were used that were at least 6 weeks old. 20 μg of mDecorin, i.e. 22 μl of the solution described above (20 μg Decorin+15 μM ZnCl2/22 μl NaCl), were injected into the left tibialis anterior (LTA), the muscle treated. 22 μl of the control solution described above (15 μM ZnCl2/22 μl NaCl) were administered into the control muscle, the right tibialis anterior (RTA). A specific number of days after injection, the mice were sacrificed and the RTA and LTA were removed, weighed then frozen for further histological study.
Preparation and Injection of the Solution Containing the Peptide mDCN 31-71:
[0075] The peptide with the sequence SEQ ID NO: 7 was synthesised by the company NeoMPS with purity >65%. It was dissolved at 2 mg/ml in 150 mM NaCl and stored at -80° C.
[0076] For injections, the preparation protocol was identical to that used for the protein, i.e. 24-40 hours before injection, the desired amount of peptide was removed from the stock solution and mixed with a solution of zinc chloride (ZnCl2) and 150 mM NaCl, to produce a final zinc concentration of 15 μM. The injection protocol was identical to that used for the protein.
Histological Analyses
[0077] Laminin Labelling:
[0078] Cryostat sections (8 μm) were made of treated and control muscles using standard techniques. The slides were fixed with Dakopen (DAKO®, ref.: S 2002) for 10 minutes open to the air and then blocked with a solution of PBS/10% goat serum for 30 min at room temperature in a humidity chamber. The rabbit anti-laminin antibody (DAKO®, ref.: Z0097) was applied to the slides at a dilution of 1:1000 for 12 hours in the humidity chamber. The slides were then rinsed in PSB (5 minutes) while being agitated and the secondary antibody (Envision HRP rabbit kit) was applied to the slides in a humidity chamber for 30 min at room temperature. After rinsing the slides in PBS (5 minutes) while being agitated, the DAB (DAKO®, ref.: K 3466) was applied to the sections for 2 to 5 minutes at room temperature in a humidity chamber. The slides were rinsed constantly and were mounted in the fume cupboard. The results were analysed using ELLIX software.
[0079] HPS Staining:
[0080] Cryostat sections (8 μm) were made of treated and control muscles using standard techniques. The slides were immersed in Harris haematoxylin for 3 minutes and then rinsed with running water. The slides were then put into acid alcohol, rinsed and soaked in Scott's tap water substitute for one minute. After rinsing, the slides were immersed in phloxine for 30 seconds, rinsed with running water and soaked in absolute alcohol for one minute. After exposure to the saffron for 3 minutes, the slides were rinsed with absolute alcohol and mounted with Eukitt resin, the solvent for which is xylene. The results were analysed using the CARTHOGRAPH program.
II) RESULTS
[0081] 1/ Weight of Muscles at Different Times after Injection into Dystrophic mdx Mice:
[0082] The RTA and LTA muscles were collected 7 (D7), 14 (D14) or 21 (D21) days after the injection and weighed. The experiment was repeated on three separate mice each time. The results are summarised in the following tables:
Day 7:
TABLE-US-00001 [0083] Growth in % Mouse Muscles Weight (g) (100*LTA/RTA) - 100 Mouse 1 RTA 1 0.0661 3.18 LTA 1 0.0682 Mouse 2 RTA 2 0.0774 0.90 LTA 2 0.0781 Mouse 3 RTA 3 0.0749 2.94 LTA 3 0.0771
Day 14:
TABLE-US-00002 [0084] Mouse Muscles Weight Growth Mouse 4 RTA 4 0.0707 58.98 LTA 4 0.1124 Mouse 5 RTA 5 0.0694 48.41 LTA 5 0.103 Mouse 6 RTA 6 0.0854 6.67 LTA 6 0.0911
Day 21:
TABLE-US-00003 [0085] Mouse Muscles Weight Growth Mouse 7 RTA 7 0.068 53.09 LTA 7 0.1041 Mouse 8 RTA 8 0.0567 66.31 LTA 8 0.0943 Mouse 9 RTA 9 0.0731 37.21 LTA 9 0.1003
[0086] The difference in muscle mass at day 21 between an mdx mouse that had received or had not received an intramuscular injection of decorin can be seen in FIGS. 1 and 2 for mice 7 and 8, respectively. There is a clear increase in muscle mass (+53.09% and +66.31%, respectively).
2/ Weight of Muscles at D18 after Injection into Dystrophic Gamma-Sarcoglycan-/-Mice:
[0087] A second series of experiments was performed on four gamma-sarcoglycan-/-mice. The protocol was identical to that described for mdx mice. The mice were sacrificed on D18. The results, shown in FIGS. 3 and 4, are presented in the following table:
TABLE-US-00004 Mouse Muscles Weight (g) Growth Mouse 1 RTA 1 0.0456 10.75 LTA 1 0.0505 Mouse 2 RTA 2 0.0413 17.43 LTA 2 0.0485 Mouse 3 RTA 3 0.0528 12.31 LTA 3 0.0593 Mouse 4 RTA 4 0.0444 24.10 LTA 4 0.0551
3/ Injection of the Peptide 31-71 Derived from the N-Terminal Part of Murine Decorin in mdx Mice:
[0088] To verify whether the N-terminal part of decorin is sufficient to produce observable increases in muscle mass, similar experiments were performed in the presence of the mDCN 31-71 peptide (SEQ ID NO: 7) corresponding to residues 31-71 of murine decorin (SEQ ID NO:6). This peptide has been described as being sufficient and necessary for binding zinc (Yang V W, LaBrenz S R, Rosenberg L C, McQuillan D, Hook M. Decorin is a Zn2+ metalloprotein. J Biol Chem. 1999, 274(18): 12454-60.).
[0089] mdx mice were injected intramuscularly into the TA with the following formulations:
LTA 1: 65 μg peptide 41 DCN+15 μM ZnCl2/33 μl NaCl;
RTA 2: 15 μM ZnCl2/33 μl NaCl.
[0090] At D18, the mice were sacrificed and the weight of the RTA and LTA muscles was measured. The results are given in the following table:
TABLE-US-00005 Muscle Weight (mg) Growth Mouse 4 RTA 4 53.6 8.77 LTA 4 58.3 Mouse 5 RTA 5 39.2 19.39 LTA 5 46.8 Mouse 6 RTA 6 40.1 24.69 LTA 6 50
[0091] These results show that an effect on muscle growth is indeed maintained in the presence of just this part of decorin.
[0092] Similar results were obtained with an even shorter peptide of 30 amino acids, with the sequence SEQ ID NO:15.
Sequence CWU
1
161359PRTartificial sequencehuman DCN (variant A) 1Met Lys Ala Thr Ile Ile
Leu Leu Leu Leu Ala Gln Val Ser Trp Ala1 5
10 15Gly Pro Phe Gln Gln Arg Gly Leu Phe Asp Phe Met
Leu Glu Asp Glu 20 25 30Ala
Ser Gly Ile Gly Pro Glu Val Pro Asp Asp Arg Asp Phe Glu Pro 35
40 45Ser Leu Gly Pro Val Cys Pro Phe Arg
Cys Gln Cys His Leu Arg Val 50 55
60Val Gln Cys Ser Asp Leu Gly Leu Asp Lys Val Pro Lys Asp Leu Pro65
70 75 80Pro Asp Thr Thr Leu
Leu Asp Leu Gln Asn Asn Lys Ile Thr Glu Ile 85
90 95Lys Asp Gly Asp Phe Lys Asn Leu Lys Asn Leu
His Ala Leu Ile Leu 100 105
110Val Asn Asn Lys Ile Ser Lys Val Ser Pro Gly Ala Phe Thr Pro Leu
115 120 125Val Lys Leu Glu Arg Leu Tyr
Leu Ser Lys Asn Gln Leu Lys Glu Leu 130 135
140Pro Glu Lys Met Pro Lys Thr Leu Gln Glu Leu Arg Ala His Glu
Asn145 150 155 160Glu Ile
Thr Lys Val Arg Lys Val Thr Phe Asn Gly Leu Asn Gln Met
165 170 175Ile Val Ile Glu Leu Gly Thr
Asn Pro Leu Lys Ser Ser Gly Ile Glu 180 185
190Asn Gly Ala Phe Gln Gly Met Lys Lys Leu Ser Tyr Ile Arg
Ile Ala 195 200 205Asp Thr Asn Ile
Thr Ser Ile Pro Gln Gly Leu Pro Pro Ser Leu Thr 210
215 220Glu Leu His Leu Asp Gly Asn Lys Ile Ser Arg Val
Asp Ala Ala Ser225 230 235
240Leu Lys Gly Leu Asn Asn Leu Ala Lys Leu Gly Leu Ser Phe Asn Ser
245 250 255Ile Ser Ala Val Asp
Asn Gly Ser Leu Ala Asn Thr Pro His Leu Arg 260
265 270Glu Leu His Leu Asp Asn Asn Lys Leu Thr Arg Val
Pro Gly Gly Leu 275 280 285Ala Glu
His Lys Tyr Ile Gln Val Val Tyr Leu His Asn Asn Asn Ile 290
295 300Ser Val Val Gly Ser Ser Asp Phe Cys Pro Pro
Gly His Asn Thr Lys305 310 315
320Lys Ala Ser Tyr Ser Gly Val Ser Leu Phe Ser Asn Pro Val Gln Tyr
325 330 335Trp Glu Ile Gln
Pro Ser Thr Phe Arg Cys Val Tyr Val Arg Ser Ala 340
345 350Ile Gln Leu Gly Asn Tyr Lys
3552250PRTartificial sequencehuman DCN (variant B) 2Met Lys Ala Thr Ile
Ile Leu Leu Leu Leu Ala Gln Val Ser Trp Ala1 5
10 15Gly Pro Phe Gln Gln Arg Gly Leu Phe Asp Phe
Met Leu Glu Asp Glu 20 25
30Ala Ser Gly Ile Gly Pro Glu Val Pro Asp Asp Arg Asp Phe Glu Pro
35 40 45Ser Leu Gly Pro Val Cys Pro Phe
Arg Cys Gln Cys His Leu Arg Val 50 55
60Val Gln Cys Ser Asp Leu Glu Leu Gly Thr Asn Pro Leu Lys Ser Ser65
70 75 80Gly Ile Glu Asn Gly
Ala Phe Gln Gly Met Lys Lys Leu Ser Tyr Ile 85
90 95Arg Ile Ala Asp Thr Asn Ile Thr Ser Ile Pro
Gln Gly Leu Pro Pro 100 105
110Ser Leu Thr Glu Leu His Leu Asp Gly Asn Lys Ile Ser Arg Val Asp
115 120 125Ala Ala Ser Leu Lys Gly Leu
Asn Asn Leu Ala Lys Leu Gly Leu Ser 130 135
140Phe Asn Ser Ile Ser Ala Val Asp Asn Gly Ser Leu Ala Asn Thr
Pro145 150 155 160His Leu
Arg Glu Leu His Leu Asp Asn Asn Lys Leu Thr Arg Val Pro
165 170 175Gly Gly Leu Ala Glu His Lys
Tyr Ile Gln Val Val Tyr Leu His Asn 180 185
190Asn Asn Ile Ser Val Val Gly Ser Ser Asp Phe Cys Pro Pro
Gly His 195 200 205Asn Thr Lys Lys
Ala Ser Tyr Ser Gly Val Ser Leu Phe Ser Asn Pro 210
215 220Val Gln Tyr Trp Glu Ile Gln Pro Ser Thr Phe Arg
Cys Val Tyr Val225 230 235
240Arg Ser Ala Ile Gln Leu Gly Asn Tyr Lys 245
2503212PRTartificial sequencehuman DCN (variant C) 3Met Lys Ala Thr
Ile Ile Leu Leu Leu Leu Ala Gln Val Ser Trp Ala1 5
10 15Gly Pro Phe Gln Gln Arg Gly Leu Phe Asp
Phe Met Leu Glu Asp Glu 20 25
30Ala Ser Gly Ile Gly Pro Glu Val Pro Asp Asp Arg Asp Phe Glu Pro
35 40 45Ser Leu Gly Pro Val Cys Pro Phe
Arg Cys Gln Cys His Leu Arg Val 50 55
60Val Gln Cys Ser Asp Leu Gly Leu Pro Pro Ser Leu Thr Glu Leu His65
70 75 80Leu Asp Gly Asn Lys
Ile Ser Arg Val Asp Ala Ala Ser Leu Lys Gly 85
90 95Leu Asn Asn Leu Ala Lys Leu Gly Leu Ser Phe
Asn Ser Ile Ser Ala 100 105
110Val Asp Asn Gly Ser Leu Ala Asn Thr Pro His Leu Arg Glu Leu His
115 120 125Leu Asp Asn Asn Lys Leu Thr
Arg Val Pro Gly Gly Leu Ala Glu His 130 135
140Lys Tyr Ile Gln Val Val Tyr Leu His Asn Asn Asn Ile Ser Val
Val145 150 155 160Gly Ser
Ser Asp Phe Cys Pro Pro Gly His Asn Thr Lys Lys Ala Ser
165 170 175Tyr Ser Gly Val Ser Leu Phe
Ser Asn Pro Val Gln Tyr Trp Glu Ile 180 185
190Gln Pro Ser Thr Phe Arg Cys Val Tyr Val Arg Ser Ala Ile
Gln Leu 195 200 205Gly Asn Tyr Lys
2104172PRTartificial sequencehuman DCN (variant D) 4Met Lys Ala Thr
Ile Ile Leu Leu Leu Leu Ala Gln Val Ser Trp Ala1 5
10 15Gly Pro Phe Gln Gln Arg Gly Leu Phe Asp
Phe Met Leu Glu Asp Glu 20 25
30Ala Ser Gly Ile Gly Pro Glu Val Pro Asp Asp Arg Asp Phe Glu Pro
35 40 45Ser Leu Gly Pro Val Cys Pro Phe
Arg Cys Gln Cys His Leu Arg Val 50 55
60Val Gln Cys Ser Asp Leu Gly Leu Asp Lys Val Pro Lys Asp Leu Pro65
70 75 80Pro Asp Thr Thr Leu
Leu Asp Leu Gln Asn Asn Lys Ile Thr Glu Ile 85
90 95Lys Asp Gly Asp Phe Lys Asn Leu Lys Asn Leu
His Val Val Tyr Leu 100 105
110His Asn Asn Asn Ile Ser Val Val Gly Ser Ser Asp Phe Cys Pro Pro
115 120 125Gly His Asn Thr Lys Lys Ala
Ser Tyr Ser Gly Val Ser Leu Phe Ser 130 135
140Asn Pro Val Gln Tyr Trp Glu Ile Gln Pro Ser Thr Phe Arg Cys
Val145 150 155 160Tyr Val
Arg Ser Ala Ile Gln Leu Gly Asn Tyr Lys 165
170575PRTartificial sequencehuman DCN (variant E) 5Met Lys Ala Thr Ile
Ile Leu Leu Leu Leu Ala Gln Val Ser Trp Ala1 5
10 15Gly Pro Phe Gln Gln Arg Gly Leu Phe Asp Phe
Met Leu Glu Asp Glu 20 25
30Ala Ser Gly Ile Gly Pro Glu Val Pro Asp Asp Arg Asp Phe Glu Pro
35 40 45Ser Leu Gly Pro Val Cys Pro Phe
Arg Cys Gln Cys His Leu Arg Val 50 55
60Val Gln Cys Ser Asp Leu Gly Cys Leu Pro Ser65 70
756354PRTartificial sequencemurine DCN 6Met Lys Ala Thr Leu
Ile Phe Phe Leu Leu Ala Gln Val Ser Trp Ala1 5
10 15Gly Pro Phe Glu Gln Arg Gly Leu Phe Asp Phe
Met Leu Glu Asp Glu 20 25
30Ala Ser Gly Ile Ile Pro Tyr Asp Pro Asp Asn Pro Leu Ile Ser Met
35 40 45Cys Pro Tyr Arg Cys Gln Cys His
Leu Arg Val Val Gln Cys Ser Asp 50 55
60Leu Gly Leu Asp Lys Val Pro Trp Asp Phe Pro Pro Asp Thr Thr Leu65
70 75 80Leu Asp Leu Gln Asn
Asn Lys Ile Thr Glu Ile Lys Glu Gly Ala Phe 85
90 95Lys Asn Leu Lys Asp Leu His Thr Leu Ile Leu
Val Asn Asn Lys Ile 100 105
110Ser Lys Ile Ser Pro Glu Ala Phe Lys Pro Leu Val Lys Leu Glu Arg
115 120 125Leu Tyr Leu Ser Lys Asn Gln
Leu Lys Glu Leu Pro Glu Lys Met Pro 130 135
140Arg Thr Leu Gln Glu Leu Arg Val His Glu Asn Glu Ile Thr Lys
Leu145 150 155 160Arg Lys
Ser Asp Phe Asn Gly Leu Asn Asn Val Leu Val Ile Glu Leu
165 170 175Gly Gly Asn Pro Leu Lys Asn
Ser Gly Ile Glu Asn Gly Ala Phe Gln 180 185
190Gly Leu Lys Ser Leu Ser Tyr Ile Arg Ile Ser Asp Thr Asn
Ile Thr 195 200 205Ala Ile Pro Gln
Gly Leu Pro Thr Ser Leu Thr Glu Val His Leu Asp 210
215 220Gly Asn Lys Ile Thr Lys Val Asp Ala Pro Ser Leu
Lys Gly Leu Ile225 230 235
240Asn Leu Ser Lys Leu Gly Leu Ser Phe Asn Ser Ile Thr Val Met Glu
245 250 255Asn Gly Ser Leu Ala
Asn Val Pro His Leu Arg Glu Leu His Leu Asp 260
265 270Asn Asn Lys Leu Leu Arg Val Pro Ala Gly Leu Ala
Gln His Lys Tyr 275 280 285Ile Gln
Val Val Tyr Leu His Asn Asn Asn Ile Ser Ala Val Gly Gln 290
295 300Asn Asp Phe Cys Arg Ala Gly His Pro Ser Arg
Lys Ala Ser Tyr Ser305 310 315
320Ala Val Ser Leu Tyr Gly Asn Pro Val Arg Tyr Trp Glu Ile Phe Pro
325 330 335Asn Thr Phe Arg
Cys Val Tyr Val Arg Ser Ala Ile Gln Leu Gly Asn 340
345 350Tyr Lys 741PRTartificial sequencemurine 31-71
DCN 7Asp Glu Ala Ser Gly Ile Ile Pro Tyr Asp Pro Asp Asn Pro Leu Ile1
5 10 15Ser Met Cys Pro Tyr
Arg Cys Gln Cys His Leu Arg Val Val Gln Cys 20
25 30Ser Asp Leu Gly Leu Asp Lys Val Pro 35
4082305DNAartificial sequencevariant A1 human DCN 8gaatctacaa
taagacaaat ttcaaatcaa gttgctccac tatactgcat aagcagttta 60gaatcttaag
cagatgcaaa aagaataaag caaatgggag gaaaaaaaag gccgataaag 120tttctggcta
caatacaaga gacatatcat taccatatga tctaatgtgg gtgtcagccg 180gattgtgttc
attgagggaa accttatttt ttaactgtgc tatggagtag aagcaggagg 240ttttcaacct
agtcacagag cagcacctac cccctcctcc tttccacacc tgcaaactct 300tttacttggg
ctgaatattt agtgtaatta catctcagct ttgagggctc ctgtggcaaa 360ttcccggatt
aaaaggttcc ctggttgtga aaatacatga gataaatcat gaaggccact 420atcatcctcc
ttctgcttgc acaagtttcc tgggctggac cgtttcaaca gagaggctta 480tttgacttta
tgctagaaga tgaggcttct gggataggcc cagaagttcc tgatgaccgc 540gacttcgagc
cctccctagg cccagtgtgc cccttccgct gtcaatgcca tcttcgagtg 600gtccagtgtt
ctgatttggg tctggacaaa gtgccaaagg atcttccccc tgacacaact 660ctgctagacc
tgcaaaacaa caaaataacc gaaatcaaag atggagactt taagaacctg 720aagaaccttc
acgcattgat tcttgtcaac aataaaatta gcaaagttag tcctggagca 780tttacacctt
tggtgaagtt ggaacgactt tatctgtcca agaatcagct gaaggaattg 840ccagaaaaaa
tgcccaaaac tcttcaggag ctgcgtgccc atgagaatga gatcaccaaa 900gtgcgaaaag
ttactttcaa tggactgaac cagatgattg tcatagaact gggcaccaat 960ccgctgaaga
gctcaggaat tgaaaatggg gctttccagg gaatgaagaa gctctcctac 1020atccgcattg
ctgataccaa tatcaccagc attcctcaag gtcttcctcc ttcccttacg 1080gaattacatc
ttgatggcaa caaaatcagc agagttgatg cagctagcct gaaaggactg 1140aataatttgg
ctaagttggg attgagtttc aacagcatct ctgctgttga caatggctct 1200ctggccaaca
cgcctcatct gagggagctt cacttggaca acaacaagct taccagagta 1260cctggtgggc
tggcagagca taagtacatc caggttgtct accttcataa caacaatatc 1320tctgtagttg
gatcaagtga cttctgccca cctggacaca acaccaaaaa ggcttcttat 1380tcgggtgtga
gtcttttcag caacccggtc cagtactggg agatacagcc atccaccttc 1440agatgtgtct
acgtgcgctc tgccattcaa ctcggaaact ataagtaatt ctcaagaaag 1500ccctcatttt
tataacctgg caaaatcttg ttaatgtcat tgctaaaaaa taaataaaag 1560ctagatactg
gaaacctaac tgcaatgtgg atgttttacc cacatgactt attatgcata 1620aagccaaatt
tccagtttaa gtaattgcct acaataaaaa gaaattttgc ctgccatttt 1680cagaatcatc
ttttgaagct ttctgttgat gttaactgag ctactagaga tattcttatt 1740tcactaaatg
taaaatttgg agtaaatata tatgtcaata tttagtaaag cttttctttt 1800ttaatttcca
ggaaaaaata aaaagagtat gagtcttctg taattcattg agcagttagc 1860tcatttgaga
taaagtcaaa tgccaaacac tagctctgta ttaatcccca tcattactgg 1920taaagcctca
tttgaatgtg tgaattcaat acaggctatg taaaattttt actaatgtca 1980ttattttgaa
aaaataaatt taaaaataca ttcaaaatta ctattgtata caagcttaat 2040tgttaatatt
ccctaaacac aattttatga agggagaaga cattggtttg ttgacaataa 2100cagtacatct
tttcaagttc tcagctattt cttctacctc tccctatctt acatttgagt 2160atggtaactt
atgtcatcta tgttgaatgt aagcttataa agcacaaagc atacatttcc 2220tgactggtct
agagaactga tgtttcaatt tacccctctg ctaaataaat attaaaacta 2280tcatgtgaaa
aaaaaaaaaa aaaaa
230592151DNAartificial sequencevariant A2 human CN 9ggaataataa gacacgccct
gaaggagtac atcgtctagt gagggacaga ccaagcacgc 60aaaacaaatt gcaatataat
gtgataagtt ctttaaaaga ggtaagagca acgtgctttg 120ggagcagaga agagggagaa
agcagcatct tgcctggatg agccagggga cacagaagag 180aagcccacta tctcatttaa
tctttacaac tctcttgcaa ggttccctgg ttgtgaaaat 240acatgagata aatcatgaag
gccactatca tcctccttct gcttgcacaa gtttcctggg 300ctggaccgtt tcaacagaga
ggcttatttg actttatgct agaagatgag gcttctggga 360taggcccaga agttcctgat
gaccgcgact tcgagccctc cctaggccca gtgtgcccct 420tccgctgtca atgccatctt
cgagtggtcc agtgttctga tttgggtctg gacaaagtgc 480caaaggatct tccccctgac
acaactctgc tagacctgca aaacaacaaa ataaccgaaa 540tcaaagatgg agactttaag
aacctgaaga accttcacgc attgattctt gtcaacaata 600aaattagcaa agttagtcct
ggagcattta cacctttggt gaagttggaa cgactttatc 660tgtccaagaa tcagctgaag
gaattgccag aaaaaatgcc caaaactctt caggagctgc 720gtgcccatga gaatgagatc
accaaagtgc gaaaagttac tttcaatgga ctgaaccaga 780tgattgtcat agaactgggc
accaatccgc tgaagagctc aggaattgaa aatggggctt 840tccagggaat gaagaagctc
tcctacatcc gcattgctga taccaatatc accagcattc 900ctcaaggtct tcctccttcc
cttacggaat tacatcttga tggcaacaaa atcagcagag 960ttgatgcagc tagcctgaaa
ggactgaata atttggctaa gttgggattg agtttcaaca 1020gcatctctgc tgttgacaat
ggctctctgg ccaacacgcc tcatctgagg gagcttcact 1080tggacaacaa caagcttacc
agagtacctg gtgggctggc agagcataag tacatccagg 1140ttgtctacct tcataacaac
aatatctctg tagttggatc aagtgacttc tgcccacctg 1200gacacaacac caaaaaggct
tcttattcgg gtgtgagtct tttcagcaac ccggtccagt 1260actgggagat acagccatcc
accttcagat gtgtctacgt gcgctctgcc attcaactcg 1320gaaactataa gtaattctca
agaaagccct catttttata acctggcaaa atcttgttaa 1380tgtcattgct aaaaaataaa
taaaagctag atactggaaa cctaactgca atgtggatgt 1440tttacccaca tgacttatta
tgcataaagc caaatttcca gtttaagtaa ttgcctacaa 1500taaaaagaaa ttttgcctgc
cattttcaga atcatctttt gaagctttct gttgatgtta 1560actgagctac tagagatatt
cttatttcac taaatgtaaa atttggagta aatatatatg 1620tcaatattta gtaaagcttt
tcttttttaa tttccaggaa aaaataaaaa gagtatgagt 1680cttctgtaat tcattgagca
gttagctcat ttgagataaa gtcaaatgcc aaacactagc 1740tctgtattaa tccccatcat
tactggtaaa gcctcatttg aatgtgtgaa ttcaatacag 1800gctatgtaaa atttttacta
atgtcattat tttgaaaaaa taaatttaaa aatacattca 1860aaattactat tgtatacaag
cttaattgtt aatattccct aaacacaatt ttatgaaggg 1920agaagacatt ggtttgttga
caataacagt acatcttttc aagttctcag ctatttcttc 1980tacctctccc tatcttacat
ttgagtatgg taacttatgt catctatgtt gaatgtaagc 2040ttataaagca caaagcatac
atttcctgac tggtctagag aactgatgtt tcaatttacc 2100cctctgctaa ataaatatta
aaactatcat gtgaaaaaaa aaaaaaaaaa a 2151101570DNAartificial
sequenceVariant B human DCN 10atgaaggcca ctatcatcct ccttctgctt gcacaagttt
cctgggctgg accgtttcaa 60cagagaggct tatttgactt tatgctagaa gatgaggctt
ctgggatagg cccagaagtt 120cctgatgacc gcgacttcga gccctcccta ggcccagtgt
gccccttccg ctgtcaatgc 180catcttcgag tggtccagtg ttctgatttg gaactgggca
ccaatccgct gaagagctca 240ggaattgaaa atggggcttt ccagggaatg aagaagctct
cctacatccg cattgctgat 300accaatatca ccagcattcc tcaaggtctt cctccttccc
ttacggaatt acatcttgat 360ggcaacaaaa tcagcagagt tgatgcagct agcctgaaag
gactgaataa tttggctaag 420ttgggattga gtttcaacag catctctgct gttgacaatg
gctctctggc caacacgcct 480catctgaggg agcttcactt ggacaacaac aagcttacca
gagtacctgg tgggctggca 540gagcataagt acatccaggt tgtctacctt cataacaaca
atatctctgt agttggatca 600agtgacttct gcccacctgg acacaacacc aaaaaggctt
cttattcggg tgtgagtctt 660ttcagcaacc cggtccagta ctgggagata cagccatcca
ccttcagatg tgtctacgtg 720cgctctgcca ttcaactcgg aaactataag taattctcaa
gaaagccctc atttttataa 780cctggcaaaa tcttgttaat gtcattgcta aaaaataaat
aaaagctaga tactggaaac 840ctaactgcaa tgtggatgtt ttacccacat gacttattat
gcataaagcc aaatttccag 900tttaagtaat tgcctacaat aaaaagaaat tttgcctgcc
attttcagaa tcatcttttg 960aagctttctg ttgatgttaa ctgagctact agagatattc
ttatttcact aaatgtaaaa 1020tttggagtaa atatatatgt caatatttag taaagctttt
cttttttaat ttccaggaaa 1080aaataaaaag agtatgagtc ttctgtaatt cattgagcag
ttagctcatt tgagataaag 1140tcaaatgcca aacactagct ctgtattaat ccccatcatt
actggtaaag cctcatttga 1200atgtgtgaat tcaatacagg ctatgtaaaa tttttactaa
tgtcattatt ttgaaaaaat 1260aaatttaaaa atacattcaa aattactatt gtatacaagc
ttaattgtta atattcccta 1320aacacaattt tatgaaggga gaagacattg gtttgttgac
aataacagta catcttttca 1380agttctcagc tatttcttct acctctccct atcttacatt
tgagtatggt aacttatgtc 1440atctatgttg aatgtaagct tataaagcac aaagcataca
tttcctgact ggtctagaga 1500actgatgttt caatttaccc ctctgctaaa taaatattaa
aactatcatg tgaaaaaaaa 1560aaaaaaaaaa
1570111456DNAartificial sequencevariant C human DCN
11atgaaggcca ctatcatcct ccttctgctt gcacaagttt cctgggctgg accgtttcaa
60cagagaggct tatttgactt tatgctagaa gatgaggctt ctgggatagg cccagaagtt
120cctgatgacc gcgacttcga gccctcccta ggcccagtgt gccccttccg ctgtcaatgc
180catcttcgag tggtccagtg ttctgatttg ggtcttcctc cttcccttac ggaattacat
240cttgatggca acaaaatcag cagagttgat gcagctagcc tgaaaggact gaataatttg
300gctaagttgg gattgagttt caacagcatc tctgctgttg acaatggctc tctggccaac
360acgcctcatc tgagggagct tcacttggac aacaacaagc ttaccagagt acctggtggg
420ctggcagagc ataagtacat ccaggttgtc taccttcata acaacaatat ctctgtagtt
480ggatcaagtg acttctgccc acctggacac aacaccaaaa aggcttctta ttcgggtgtg
540agtcttttca gcaacccggt ccagtactgg gagatacagc catccacctt cagatgtgtc
600tacgtgcgct ctgccattca actcggaaac tataagtaat tctcaagaaa gccctcattt
660ttataacctg gcaaaatctt gttaatgtca ttgctaaaaa ataaataaaa gctagatact
720ggaaacctaa ctgcaatgtg gatgttttac ccacatgact tattatgcat aaagccaaat
780ttccagttta agtaattgcc tacaataaaa agaaattttg cctgccattt tcagaatcat
840cttttgaagc tttctgttga tgttaactga gctactagag atattcttat ttcactaaat
900gtaaaatttg gagtaaatat atatgtcaat atttagtaaa gcttttcttt tttaatttcc
960aggaaaaaat aaaaagagta tgagtcttct gtaattcatt gagcagttag ctcatttgag
1020ataaagtcaa atgccaaaca ctagctctgt attaatcccc atcattactg gtaaagcctc
1080atttgaatgt gtgaattcaa tacaggctat gtaaaatttt tactaatgtc attattttga
1140aaaaataaat ttaaaaatac attcaaaatt actattgtat acaagcttaa ttgttaatat
1200tccctaaaca caattttatg aagggagaag acattggttt gttgacaata acagtacatc
1260ttttcaagtt ctcagctatt tcttctacct ctccctatct tacatttgag tatggtaact
1320tatgtcatct atgttgaatg taagcttata aagcacaaag catacatttc ctgactggtc
1380tagagaactg atgtttcaat ttacccctct gctaaataaa tattaaaact atcatgtgaa
1440aaaaaaaaaa aaaaaa
1456121336DNAArtificial sequencevariant D human DCN 12atgaaggcca
ctatcatcct ccttctgctt gcacaagttt cctgggctgg accgtttcaa 60cagagaggct
tatttgactt tatgctagaa gatgaggctt ctgggatagg cccagaagtt 120cctgatgacc
gcgacttcga gccctcccta ggcccagtgt gccccttccg ctgtcaatgc 180catcttcgag
tggtccagtg ttctgatttg ggtctggaca aagtgccaaa ggatcttccc 240cctgacacaa
ctctgctaga cctgcaaaac aacaaaataa ccgaaatcaa agatggagac 300tttaagaacc
tgaagaacct tcacgttgtc taccttcata acaacaatat ctctgtagtt 360ggatcaagtg
acttctgccc acctggacac aacaccaaaa aggcttctta ttcgggtgtg 420agtcttttca
gcaacccggt ccagtactgg gagatacagc catccacctt cagatgtgtc 480tacgtgcgct
ctgccattca actcggaaac tataagtaat tctcaagaaa gccctcattt 540ttataacctg
gcaaaatctt gttaatgtca ttgctaaaaa ataaataaaa gctagatact 600ggaaacctaa
ctgcaatgtg gatgttttac ccacatgact tattatgcat aaagccaaat 660ttccagttta
agtaattgcc tacaataaaa agaaattttg cctgccattt tcagaatcat 720cttttgaagc
tttctgttga tgttaactga gctactagag atattcttat ttcactaaat 780gtaaaatttg
gagtaaatat atatgtcaat atttagtaaa gcttttcttt tttaatttcc 840aggaaaaaat
aaaaagagta tgagtcttct gtaattcatt gagcagttag ctcatttgag 900ataaagtcaa
atgccaaaca ctagctctgt attaatcccc atcattactg gtaaagcctc 960atttgaatgt
gtgaattcaa tacaggctat gtaaaatttt tactaatgtc attattttga 1020aaaaataaat
ttaaaaatac attcaaaatt actattgtat acaagcttaa ttgttaatat 1080tccctaaaca
caattttatg aagggagaag acattggttt gttgacaata acagtacatc 1140ttttcaagtt
ctcagctatt tcttctacct ctccctatct tacatttgag tatggtaact 1200tatgtcatct
atgttgaatg taagcttata aagcacaaag catacatttc ctgactggtc 1260tagagaactg
atgtttcaat ttacccctct gctaaataaa tattaaaact atcatgtgaa 1320aaaaaaaaaa
aaaaaa
1336131223DNAArtificial sequenceVariant E human DCN 13atgaaggcca
ctatcatcct ccttctgctt gcacaagttt cctgggctgg accgtttcaa 60cagagaggct
tatttgactt tatgctagaa gatgaggctt ctgggatagg cccagaagtt 120cctgatgacc
gcgacttcga gccctcccta ggcccagtgt gccccttccg ctgtcaatgc 180catcttcgag
tggtccagtg ttctgatttg ggttgtctac cttcataaca acaatatctc 240tgtagttgga
tcaagtgact tctgcccacc tggacacaac accaaaaagg cttcttattc 300gggtgtgagt
cttttcagca acccggtcca gtactgggag atacagccat ccaccttcag 360atgtgtctac
gtgcgctctg ccattcaact cggaaactat aagtaattct caagaaagcc 420ctcattttta
taacctggca aaatcttgtt aatgtcattg ctaaaaaata aataaaagct 480agatactgga
aacctaactg caatgtggat gttttaccca catgacttat tatgcataaa 540gccaaatttc
cagtttaagt aattgcctac aataaaaaga aattttgcct gccattttca 600gaatcatctt
ttgaagcttt ctgttgatgt taactgagct actagagata ttcttatttc 660actaaatgta
aaatttggag taaatatata tgtcaatatt tagtaaagct tttctttttt 720aatttccagg
aaaaaataaa aagagtatga gtcttctgta attcattgag cagttagctc 780atttgagata
aagtcaaatg ccaaacacta gctctgtatt aatccccatc attactggta 840aagcctcatt
tgaatgtgtg aattcaatac aggctatgta aaatttttac taatgtcatt 900attttgaaaa
aataaattta aaaatacatt caaaattact attgtataca agcttaattg 960ttaatattcc
ctaaacacaa ttttatgaag ggagaagaca ttggtttgtt gacaataaca 1020gtacatcttt
tcaagttctc agctatttct tctacctctc cctatcttac atttgagtat 1080ggtaacttat
gtcatctatg ttgaatgtaa gcttataaag cacaaagcat acatttcctg 1140actggtctag
agaactgatg tttcaattta cccctctgct aaataaatat taaaactatc 1200atgtgaaaaa
aaaaaaaaaa aaa
1223141065DNAArtificial sequenceMurine DCN 14atgaaggcaa ctctcatctt
cttccttctg gcacaagtct cttgggctgg accatttgaa 60cagagaggct tatttgactt
catgctagaa gatgaggctt ctggcataat cccttatgac 120cctgacaatc ccctgatatc
tatgtgcccc taccgatgcc agtgtcatct tcgagtggtg 180cagtgttctg atctgggttt
ggacaaagtg ccctgggatt ttccacccga cacaaccttg 240ctagacctgc aaaacaacaa
aattacagag atcaaagaag gggccttcaa gaacctgaag 300gacttgcata ccttgatcct
tgtcaacaac aagatcagca aaatcagtcc agaggcattc 360aaacctctcg tgaagttgga
aaggctttac ctgtctaaga accaactaaa ggaactgcct 420gaaaaaatgc ccagaactct
ccaggaactt cgtgtccatg agaatgagat caccaagctg 480cggaaatccg acttcaatgg
actgaacaat gtgcttgtca tagaactggg cggcaaccca 540ctgaaaaact ctgggattga
aaacggagcc ttccagggac tgaagagtct ctcatacatt 600cgcatctcag acaccaacat
aactgcgatc cctcaaggtc tgcctacttc tctcactgaa 660gtgcatctag atggcaacaa
gatcaccaag gttgatgcac ccagcctgaa aggactgatt 720aatttgtcta aactgggatt
gagcttcaac agcatcaccg ttatggagaa tggcagtctg 780gccaatgttc ctcatctgag
ggaactccac ttggacaaca acaaactcct cagggtgcct 840gctgggctgg cacagcataa
gtatatccag gtcgtctacc ttcacaacaa caacatctcc 900gcagttgggc aaaatgactt
ctgccgagct ggacacccct ctcgaaaggc ttcctactcg 960gctgtgagtc tttacggcaa
ccctgtccgg tattgggaaa tctttccaaa caccttcaga 1020tgtgtctatg tgcgttctgc
cattcaactt ggaaactaca agtaa 10651530PRTartificial
sequencemurine 42-71 DCN 15Asp Asn Pro Leu Ile Ser Met Cys Pro Tyr Arg
Cys Gln Cys His Leu1 5 10
15Arg Val Val Gln Cys Ser Asp Leu Gly Leu Asp Lys Val Pro 20
25 301641PRTartificial sequencehuman DCN
fragment 16Ile Gly Pro Glu Val Pro Asp Asp Arg Asp Phe Glu Pro Ser Leu
Gly1 5 10 15Pro Val Cys
Pro Phe Arg Cys Gln Cys His Leu Arg Val Val Gln Cys 20
25 30Ser Asp Leu Gly Leu Asp Lys Val Pro
35 40
User Contributions:
Comment about this patent or add new information about this topic: