Patent application title: TARGETING PAX2 FOR THE INDUCTION OF DEFB1-MEDIATED TUMOR IMMUNITY AND CANCER THERAPY
Inventors:
Carlton D. Donald (Mount Pleasant, SC, US)
Carlton D. Donald (Mount Pleasant, SC, US)
Assignees:
Medical University of South Carolina
IPC8 Class: AA61K39395FI
USPC Class:
4241361
Class name: Immunoglobulin, antiserum, antibody, or antibody fragment, except conjugate or complex of the same with nonimmunoglobulin material structurally-modified antibody, immunoglobulin, or fragment thereof (e.g., chimeric, humanized, cdr-grafted, mutated, etc.) bispecific or bifunctional, or multispecific or multifunctional, antibody or fragment thereof
Publication date: 2011-07-21
Patent application number: 20110177075
Abstract:
Provided is a method of treating cancer in a subject by inhibiting
expression of PAX2. An example of a cancer treated by the present method
is prostate cancer. In the cancer treatment methods disclosed, the method
of inhibiting expression of PAX2 can be by administration of a nucleic
acid encoding an siRNA for PAX2. A method of treating cancer in a subject
by administering DEFB1 is also provided. Similarly, provided is a method
of treating cancer in a subject by increasing expression of DEFB1 in the
subject.Claims:
1-11. (canceled)
12. A method for treating cancer in a subject, comprising: administering an effective amount of an anti-PAX2 agent to said subject, wherein said anti-PAX2 agent comprises an anti-PAX2 antibody.
13. The method of claim 12, wherein said cancer is prostate cancer.
14. The method of claim 12, wherein said anti-PAX2 agent is administered with a pharmaceutically acceptable carrier.
15. The method of claim 12, wherein said anti-PAX2 agent is administered parenterally.
16. The method of claim 12, wherein said anti-PAX2 agent is administered subcutaneously.
17. The method of claim 12, wherein said anti-PAX2 agent is administered intramuscularly.
18. The method of claim 12, wherein said anti-PAX2 agent is administered intravenously.
19. The method of claim 12, wherein said anti-PAX2 agent is formulated for sustained release.
20. The method of claim 12, wherein said anti-PAX2 agent is conjugated to an antibody to target a particular cell type.
21. The method of claim 12, wherein said anti-PAX2 agent is conjugated to a receptor to target a particular cell type.
22. The method of claim 12, wherein said anti-PAX2 agent is conjugated to a receptor ligand to target a particular cell type.
23. A composition for treating cancer, comprising: an effective amount of an anti-PAX2 antibody; and a pharmaceutically acceptable carrier, wherein said composition is formulated for parenteral administration.
24. The composition of claim 23, wherein said composition is formulated for intramuscular administration.
25. The composition of claim 23, wherein said composition is formulated for intravenous administration.
26. The composition of claim 23, wherein said composition is formulated for subcutaneous administration.
27. The composition of claim 23, wherein said composition is formulated for sustained release.
28. The composition of claim 23, wherein said anti-PAX2 agent is conjugated to an antibody to target a particular cell type.
29. The composition of claim 23, wherein said anti-PAX2 agent is conjugated to a receptor to target a particular cell type.
30. The composition of claim 23, wherein said anti-PAX2 agent is conjugated to a receptor ligand to target a particular cell type.
31. The composition of claim 23, wherein said pharmaceutically acceptable carrier comprises semipermeable matrices of solid hydrophobic polymers and wherein said matrices are in the form of shaped articles.
Description:
[0001] This application is a continuation application of U.S. patent
application Ser. No. 12/090,191, filed on Sep. 15, 2008 as the national
entry of PCT Application No. PCT/US2006/040215, filed on Oct. 16, 2006,
which claims priority to U.S. Patent Application No. 60/726,921, filed on
Oct. 14, 2005, all of the aforementioned applications are incorporated
herein by reference in their entirety.
SUMMARY
[0003] Current anticancer chemotherapies that are based on alkylating agents, anti-metabolites and natural products are heterogeneous in their mechanism of action. Consequently, most of them also act against normal cells resulting in severe side effects and toxicity to the patient. Disclosed herein is a method for the treatment of advanced prostate cancer using human beta defensin-1 (DEFB1), which is a naturally component of the innate immune system, to induce prostate cancer tumor immunity. This is accomplished through endogenously added DEFB1, ectopically expressed DEFB1 or de novo expression of EFB1 by inhibiting the transcriptional repressor PAX2 by a variety of mechanisms or agents. Inhibiting PAX2 expression by siRNA therapy turns on DEFB1 expression and generates DEFB1-mediated cell death in prostate cancer. With this, the technology described here is used for the design of small molecules to specifically block PAX2 expression. Alternatively provided are molecules containing the CCTTG (SEQ ID NO:1) recognition sequence (in either forward of reverse orientation) that to bind to the DNA-binding domain of PAX2 preventing its binding to the DEFB1 promoter through competitive inhibition. This permits DEFB1 expression, triggering both an innate and adaptive immune response, and resulting in the killing of prostate cancer cells and the suppression of prostate tumor formation. In conclusion, these modulators of innate tumor immunity, PAX2 and DEFB1, and the molecular therapies based on them provide for the treatment of prostate cancer with little toxicity to the patient.
BRIEF DESCRIPTION OF THE DRAWINGS
[0004] The accompanying drawings illustrate one or more embodiments of the invention and, together with the written description, serve to explain the principles of the invention. Wherever possible, the same reference numbers are used throughout the drawings to refer to the same or like elements of an embodiment.
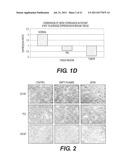
[0005] FIG. 1. QRT-PCR analysis of DEFB1 Expression. In order to verify induction of DEFB1 expression, QRT-PCR was performed. A, DEFB1 relative expression levels were compared in clinical samples from 6 patients that underwent radical prostatectomies. B, DEFB1 relative expression levels were compared in benign and malignant prostatic clinical samples, hPrEC cells and in prostate cancer cell lines before and after DEFB1 induction. C, DEFB1 relative expression levels were analyzed in benign tissue, malignant tissue and PIN in a single tissue section. D, DEFB1 expression in benign tissue, malignant tissue and PIN in one patient was compared to the average DEFB1 expression level found in benign tissue.
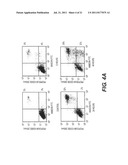
[0006] FIG. 2. Microscopic analysis of DEFB1 induced changes in membrane integrity and cell morphology. Cell morphology of DU145, PC3 and LNCaP was analyzed by phase contrast microscopy after 48 hours of DEFB1 induction. Membrane ruffling is indicated by black arrows and apoptotic bodies are indicated white arrows.
[0007] FIG. 3. Analysis of DEFB1 Cytotoxicity in Prostate Cancer Cells. The prostate cell lines DU145, PC3 and LNCaP were treated with PonA to induce DEFB1 expression for 1-3 days after which MTT assay was performed to determine cell viability. Results represent mean±s.d., n=9.
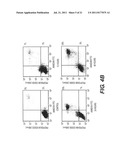
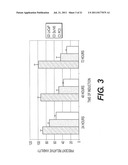
[0008] FIG. 4. Induction of cell death in DU145 and PC3 cells by DEFB1. DEFB1 expression was induced in prostate cancer cell lines DU145 (A) and PC3 (B) and then subjected to annexin V/FITC/propidium iodide staining and flow cytometric analysis. Cells positive for propidium iodide and annexin V were considered apoptotic. Times of induction are shown under each panel. Numbers next to the boxes for each time point represent the percentages of propidium iodide (PI).sup.- annexin V.sup.+ cells (lower right quadrant), and PI.sup.+ annexin V.sup.+ cells (upper right quadrant). The data are from a single experiment that is representative of three separate experiments.
[0009] FIG. 5. Pan-caspase analysis following DEFB1 induction. DU145 and PC3 cells were stained with FAM-VAD-FMK-labeled fluoromethyl ketone to detect caspase activity. Cells were visible under DIC for each condition. Confocal microscopic analysis revealed no caspase staining in control DU145 (B), PC3 cells (F) and LNCaP (J). Cells treated with PonA for 24 hours to induce DEFB1 revealed caspase activity in DU145 (D) and PC3 (H). No caspase activity was detected in LNCaP (L).
[0010] FIG. 6. Silencing of PAX2 Protein Expression Following PAX2 siRNA Treatment. (a) Western blot analysis of PC3 and DU145 cells transfected with PAX2 siRNA duplex at day zero (lane 1), day two (lane 2), and day four (lane 3). (b) Western blot analysis of PC3 and DU145 cells transfected with PAX2 siRNA duplex at day zero (lane 1), day two (lane 2), day four (lane 3) and day 6 (lane 4). PAX2 protein was undetectable as early as after four days of treatment (lane 3) in DU145 cells and after six days of treatment in PC3. Blots were stripped and re-probed for β-actin as an internal control.
[0011] FIG. 7. Analysis of Prostate Cancer Cells Growth after Treatment with Pax 2 siRNA. Phase contrast microscopic analysis of DU145, PC3 and LNCaP at 6 days in the presence of normal growth media. Treatment with negative control siRNA had no effect on the cells. However, there was a significant reduction in cell number in all three lines following treatment with PAX2 siRNA.
[0012] FIG. 8. Analysis of Cell Death Following siRNA Silencing of PAX2. Prostate cancer cell lines PC3, DU145, and LNCaP were treated with 0.5 μg of a pool of four PAX2 siRNA's or four non-specific control siRNA's for 2, 4 or 6 days after which MTT assay was done to determine cell viability. Results represent mean±s.d., n=9.
[0013] FIG. 9. Analysis of Caspase Activity. DU145, PC3 and LNCaP cells were stained with carboxyfluorescein-labeled fluoromethyl ketone to detected caspase activity following treatment with PAX2 siRNA. Confocal microscopic analysis of untreated and treated cells show cells were visible with DIC. Analysis under fluorescence revealed no caspase staining in control DU145 (B), PC3 cells (F) and LNCaP cells (J). However, cell treated with PAX2 siRNA induced caspase activity in DU145 (D), PC3 (H) and LNCaP (L).
[0014] FIG. 10. Analysis of Apoptotic Factors Following PAX2 siRNA Treatment. Changes in expression of pro-apoptotic factors were compared in untreated control cells and in cells treated for six days with PAX2 siRNA. A, BAX expression levels increased in DU145, PC3 and LNCaP. B, BID expression increased in DU145 and LNCaP, but change in PC3. C, BAD expression levels increased in all three cell lines.
[0015] FIG. 11. Model of PAX2 Binding to DNA Recognition Sequence. The PAX2 transcriptional repressor binds to a CCTTG (SEQ ID NO:1) recognition site immediately adjacent to the DEFB1 TATA box preventing transcription and DEFB1 protein expression. Inhibition of PAX2 protein expression allows normal DEFB1 expression.
[0016] FIG. 12. Illustration of the DEFB1 Reporter Construct. The DEFB1 promoter consisting of the first 160 bases upstream of the mRNA start site was PCR amplified from DU145 cell and ligated into the pGL3 luciferase reporter plasmid.
[0017] FIG. 13. Inhibition of PAX2 Results in DEFB1 Expression. DU145, PC3, LNCaP and HPrEC were treated for 48 hours with PAX2 siRNA. QRT-PCR analysis before treatment showed no DEFB1 expression in DU145, PC3 and LNCaP. However, DEFB1 expression was restored following treatment in all lines. There was no change in DEFB1 expression following siRNA treatment of PAX2-null HprEC.
[0018] FIG. 14. Inhibition of PAX2 Results in Increased DEFB1 Promoter Activity. PC3 promoter/pGL3 and DU145 promoter/pGL3 construct were generated and were transfected into PC3 and DU145 cells, respectively. Promoter activity was compared before and after PAX2 inhibition by siRNA treatment. DEFB1 promoter activity increased 2.65-fold in DU145 and 3.78 fold in PC3 following treatment.
[0019] FIG. 15. DEFB1 Causes Loss of Membrane Integrity. Membrane integrity of PC3 and DU145 cells was analyzed by confocal laser microscopy following the induction of DEFB1 expression for 48 hours. Green staining was indicative of the localization of AO, and red staining represents EtBr. Yellow staining represents the co-localization of both AO and EtBr in the nucleus.
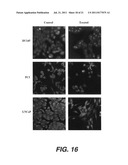
[0020] FIG. 16. PAX2 Inhibition Results in Loss of Membrane Integrity. Cells were treated for 48 hours with PAX2 siRNA and membrane integrity was analyzed by confocal laser microscopy.
[0021] FIG. 17. ChIP Analysis of PAX2 binding to DEFB1 Promoter. ChIP analysis was performed on DU145 and PC3 cells. Following immunoprecipitation with an anti-PAX2 antibody, PCR was performed to detect the DEFB1 promoter region containing the GTTCC (SEQ ID NO: 2) PAX2 recognition site. This demonstrates that the PAX2 transcriptional repressor is bound to the DEFB1 promoter in prostate cancer cell lines.
[0022] FIG. 18. Predicted Structure of the PrdPD and PrdHD with DNA. The coordinates of the structures of the PrdPD bound to DNA (Xu et al., 1995 and the PrdHD bound to DNA (Wilson et al., 1995) were used to construct a model of the two domains as they bound to a PH0 site. The individual binding sites are abutted next to each other with a specific orientation as indicated. The PAI binding site is in red, the HD binding site is in blue, and the corresponding PAI domain is in turquoise, and HD is in yellow. The RED domain is oriented based on the PrdPD crystal structure.
[0023] FIG. 19. Comparison of Consensus Sequences of Different Paired Domains. At the top of the Figure is drawn a schematic representation of protein±DNA contacts described in the crystallographic analysis of the Prd-paired-domain±DNA complex [9]. Empty boxes indicate a-helices, shaded boxes indicates b-sheets and a thick line indicate a b-turn. Contacting amino acids are shown by single-letter code. Only direct amino acid±base contacts are shown. Empty circles indicate major groove contacts while red arrows indicate minor groove contacts. This scheme is aligned to all known consensus sequences for paired-domain proteins (top strands only are shown). Vertical lines between consensus sequences indicate conserved base-pairs. Numbering of the positions is shown at the bottom of the Figure and it is the same as that used in [9].
DETAILED DESCRIPTION
[0024] As shown herein, PAX2 inhibits expression of DEFB1, and DEFB1 is shown to have tumor cell killing activity. Thus, provided is a method of treating cancer in a subject by inhibiting expression of PAX2. An example of a cancer treated by the present method is prostate cancer. The present methods are particularly effective for treatment of late stage prostate cancer.
[0025] In the cancer treatment methods disclosed, the method of inhibiting expression of PAX 2 can be by administration of a nucleic acid encoding a siRNA for PAX 2. Dharmachon is a commercial source for such siRNAs.
[0026] The siRNA for use in the methods can be selected from the group consisting of:
TABLE-US-00001 AUAGACUCGACUUGACUUCUU (SEQ ID NO: 3) AUCUUCAUCACGUUUCCUCUU (SEQ ID NO: 4) GUAUUCAGCAAUCUUGUCCUU (SEQ ID NO: 5) GAUUUGAUGUGCUCUGAUGUU (SEQ ID NO: 6)
[0027] The following table illustrates the above antisense sequences and their corresponding sense sequences.
TABLE-US-00002 Sense (5'-3') Antisense (5'-3') Sequence A 5'-GAAGUCAAGUC 5'-AUAGACUCGAC GAGUCUAUUU-3' UUGACUUCUU-3' (SEQ ID NO: 7) (SEQ ID NO: 3) Sequence B 5'-GAGGAAACGUG 5'-AUCUUCAUCAC AUGAAGAUUU-3' GUUUCCUCUU-3' (SEQ ID NO: 8) (SEQ ID NO: 4) Sequence C 5'-GGACAAGAUUG 5'-GUAUUCAGCAA CUGAAUACUU-3' UCUUGUCCUU-3' (SEQ ID NO: 9) (SEQ ID NO: 5) Sequence D 5'-CAUCAGAGCA- 5'-GAUUUGAUGUG CAUCAAAUCUU-3' CUCUGAUGUU-3' (SEQ ID NO: 10) (SEQ ID NO: 6)
[0028] Further examples of molecules that inhibit PAX2 include:
[0029] #1 ACCCGACTATGTTCGCCTGG (SEQ ID NO: 11),
[0030] #2 AAGCTCTGGATCGAGTCTTTG (SEQ ID NO: 12),
[0031] and #4 ATGTGTCAGGCACACAGACG (SEQ ID NO: 13). #4 was shown to inhibit PAX2 (Davies et al., Hum. Mol. Gen Jan. 15, 13 (2); 235).
[0032] Another paper (Muratovska et al., Paired-Box genes are frequently expressed in cancer and often required for cancer cell survival Oncogene (2003) 22, 7989-7997) discloses the following siRNAs: GUCGAGUCUAUCUGCAUCCUU (SEQ ID NO: 14) and GGAUGCAGAUAGACUCGACUU (SEQ ID NO: 15).
[0033] To down-regulate Pax2 expression, Fonsato et al. transfected tumor-derived endothelial cells with an anti-sense PAX2 vector. See Fonsato V. et al (Expression of Pax2 in human renal tumor-derived endothelial cells sustains apoptosis resistance and angiogenesis, Am J Pathol. 2006 February; 168(2):706-1), incorporated herein by reference for its description of this molecule. Similarly, Hueber et al. teach that PAX2 antisense cDNA and PAX2-small interfering RNA (100 nM) reduce endogenous PAX2 protein. See Hueber et al. PAX2 inactivation enhances cisplatin-induced apoptosis in renal carcinoma cells, Kidney Int. 2006 April; 69(7):1139-45 incorporated herein for its teaching of PAX2 antisense and PAX2 siRNA.
[0034] Additional inhibitors of PAX2 expression or the binding of PAX2 to the DEFB1 promoter are provided to increase DEFB1 expression in the presently disclosed methods. For example, small molecules and antibodies are designed based on the present studies to interfere with or inhibit the binding of PAX2 to the DEFB1 promoter.
[0035] As shown herein, PAX2 inhibits expression of DEFB1, and DEFB1 is shown to have tumor cell killing activity. Thus, a method of treating cancer in a subject by administering DEFB1 is also provided. An example of a cancer treated by the present method is prostate cancer.
[0036] Similarly, provided is a method of treating cancer in a subject by increasing expression of DEFB1 in the subject. The present methods of administering or increasing the expression of DEFB1 are particularly effective for treatment of late stage prostate cancer.
[0037] In one embodiment of the methods of the invention for treating cancer by administering DEFB1 or increasing DEFB1 expression (e.g., by inhibiting expression or binding of PAX2), the subject is a subject diagnosed with prostate cancer. In a further embodiment of the methods of the invention for treating cancer by administering DEFB1 or increasing DEFB1 expression (e.g., by inhibiting expression or binding of PAX2), the subject is a subject diagnosed with advanced (late stage) prostate cancer.
[0038] In the method wherein the expression of DEFB1 is increased, it can be increased by blocking the binding of PAX2 to the DEFB1 promoter. The blocking of binding of PAX2 to the DEFB1 promoter can be by administration of an oligonucleotide containing the PAX2 DNA binding site of DEFB1. This oligonucleotide can be complementary to the sequence of PAX2 that binds to the DEFB1 promoter. Alternatively, the oligonucleotide can interact with the PAX2 in a way that inhibits binding to DEFB1. This interaction can be based on three-dimensional structure rather than primary nucleotide sequence.
[0039] PAX proteins are a family of transcription factors conserved during evolution and able to bind specific DNA sequences through a domains called a "paired domain" and a "homeodomain". The paired domain (PD) is a consensus sequence shared by certain PAX proteins (e.g., PAX2 and PAX6). The PD directs DNA binding of amino acids located in the α3-helix forming a DNA-Protein complex. For PAX2, the amino acids in the HD recognize and interact specifically with a CCTTG (SEQ ID NO: 1) DNA core sequence. Therefore, the critical region for PAX2 binding to DEFB1 would be AAGTTCACCCTTGACTGTG (SEQ ID NO: 16). Oligonucleotides up to and exceeding 64 bases in length, which include this sequence or its complement are expected to be inhibitors.
[0040] The DNA-binding specificity of the PAX-8 paired domain was investigated. Site selection experiments indicate that PAX-8 binds to a consensus sequence similar to those bound by PAX-2 and PAX-5. When consensus sequences of various paired domains are observed in light of recent structural studies describing paired-domain-DNA interaction [Xu, Rould, Jun, Desplan and Pabo (1995) Cell 80, 639-650], it appears that base-pairs contacted in the minor groove are conserved, while most of the base-pairs contacted in the major groove are not. Therefore a network of specific minor groove contacts is a common characteristic of paired-domain-DNA interactions. The functional importance of such a network can be successfully tested by analyzing the effect of consensus-based mutations on the PAX2 binding site of the DEFB1 promoter.
[0041] The PAX2 DNA binding site of DEFB1 can comprise SEQ ID NO:1 (CCTTG).
[0042] The oligonucleotide comprising to the PAX2 DNA binding site of DEFB1 is selected from the group consisting of X1 CCTTG X2 (SEQ ID NO: 17), wherein X1=NNNNNNN NNNNNNNNNNNNNNNNNNNNNNNNNNNN and is/are from 1 to 35 contiguous flanking nucleotides of DEFB1 and X2=NNNNNNNNNNNNNNNNNNNNNNNNNN NNNNNNNNN and is/are from 1 to 35 contiguous flanking nucleotides of DEFB1. The nucleotides can be contiguous nucleotides that normally flank the PAX2 DNA binding site of DEFB1. Alternatively, they can be unrelated to DEFB1, and selected routinely to avoid interference with the recognition sequence.
[0043] For example, the inhibitory oligonucleotides can be selected from the group consisting of:
TABLE-US-00003 (SEQ ID NO: 18) CTCCCTTCAGTTCCGTCGAC (SEQ ID NO: 19) CTCCCTTCACCTTGGTCGAC (SEQ ID NO: 20) ACTGTGGCACCTCCCTTCAGTTCCGTCGACGAGGTTGTGC (SEQ ID NO: 21) ACTGTGGCACCTCCCTTCACCTTGGTCGACGAGGTTGTGC.
[0044] The disclosed compositions can be used to treat any disease where uncontrolled cellular proliferation occurs such as cancers. A non-limiting list of different types of cancers is as follows: lymphomas (Hodgkins and non-Hodgkins), leukemias, carcinomas, carcinomas of solid tissues, squamous cell carcinomas, adenocarcinomas, sarcomas, gliomas, high grade gliomas, blastomas, neuroblastomas, plasmacytomas, histiocytomas, melanomas, adenomas, hypoxic tumors, myelomas, AIDS-related lymphomas or sarcomas, metastatic cancers, or cancers in general.
[0045] A representative but non-limiting list of cancers that the disclosed compositions can be used to treat is the following: lymphoma, B cell lymphoma, T cell lymphoma, mycosis fungoides, Hodgkin's Disease, myeloid leukemia, bladder cancer, brain cancer, nervous system cancer, head and neck cancer, squamous cell carcinoma of head and neck, kidney cancer, lung cancers such as small cell lung cancer and non-small cell lung cancer, neuroblastoma/glioblastoma, ovarian cancer, pancreatic cancer, prostate cancer, skin cancer, liver cancer, melanoma, squamous cell carcinomas of the mouth, throat, larynx, and lung, colon cancer, cervical cancer, cervical carcinoma, breast cancer, and epithelial cancer, renal cancer, genitourinary cancer, pulmonary cancer, esophageal carcinoma, head and neck carcinoma, large bowel cancer, hematopoietic cancers; testicular cancer; colon and rectal cancers, prostatic cancer, or pancreatic cancer. Compounds disclosed herein may also be used for the treatment of precancer conditions such as cervical and anal dysplasias, other dysplasias, severe dysplasias, hyperplasias, atypical hyperplasias, and neoplasias. Further, a number of diseases stemming from chronic inflammation, e.g., prostatitis and Benign Prostatic Hypertrophy (BPH), as well as various cancers of the prostate, can be impacted by the present methods and compounds.
[0046] DEFB1's gene locus (8p23.3) is a hotspot for deletions and has been linked to patients with poorer prognosis. Thus, DEFB1 (and perhaps PAX2) can be used as a biomarker, e.g., in a screening for the early detection of prostate cancer. Furthermore, data presented here indicate that its loss may occur as early as PIN (or even before), and may be a major contributing factor to the onset of prostate cancer.
Nucleic Acid Homology/Identity/Similarity
[0047] It is understood that as discussed herein the use of the terms homology and identity mean the same thing as similarity. Thus, for example, if the use of the word homology is used between two non-natural sequences it is understood that this is not necessarily indicating an evolutionary relationship between these two sequences, but rather is looking at the similarity or relatedness between their nucleic acid sequences. Many of the methods for determining homology between two evolutionarily related molecules are routinely applied to any two or more nucleic acids or proteins for the purpose of measuring sequence similarity regardless of whether they are evolutionarily related or not.
[0048] In general, it is understood that one way to define any known variants and derivatives or those that might arise, of the disclosed genes and proteins herein, is through defining the variants and derivatives in terms of homology to specific known sequences. This identity of particular sequences disclosed herein is also discussed elsewhere herein. In general, variants of genes and proteins herein disclosed typically have at least, about 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99 percent homology to the stated sequence or the native sequence. Those of skill in the art readily understand how to determine the homology of two proteins or nucleic acids, such as genes. For example, the homology can be calculated after aligning the two sequences so that the homology is at its highest level.
[0049] Another way of calculating homology can be performed by published algorithms. Optimal alignment of sequences for comparison may be conducted by the local homology algorithm of Smith and Waterman Adv. Appl. Math. 2: 482 (1981), by the homology alignment algorithm of Needleman and Wunsch, J. MoL Biol. 48: 443 (1970), by the search for similarity method of Pearson and Lipman, Proc. Natl. Acad. Sci. U.S.A. 85: 2444 (1988), by computerized implementations of these algorithms (GAP, BESTFIT, FASTA, and TFASTA in the Wisconsin Genetics Software Package, Genetics Computer Group, 575 Science Dr., Madison, Wis.), or by inspection.
[0050] The same types of homology can be obtained for nucleic acids by for example the algorithms disclosed in Zuker, M. Science 244:48-52, 1989, Jaeger et al. Proc. Natl. Acad. Sci. USA 86:7706-7710, 1989, Jaeger et al. Methods Enzymol. 183:281-306, 1989 which are herein incorporated by reference for at least material related to nucleic acid alignment. It is understood that any of the methods typically can be used and that in certain instances the results of these various methods may differ, but the skilled artisan understands if identity is found with at least one of these methods, the sequences would be said to have the stated identity, and be disclosed herein.
[0051] For example, as used herein, a sequence recited as having a particular percent homology to another sequence refers to sequences that have the recited homology as calculated by any one or more of the calculation methods described above. For example, a first sequence has 80 percent homology, as defined herein, to a second sequence if the first sequence is calculated to have 80 percent homology to the second sequence using the Zuker calculation method even if the first sequence does not have 80 percent homology to the second sequence as calculated by any of the other calculation methods. As another example, a first sequence has 80 percent homology, as defined herein, to a second sequence if the first sequence is calculated to have 80 percent homology to the second sequence using both the Zuker calculation method and the Pearson and Lipman calculation method even if the first sequence does not have 80 percent homology to the second sequence as calculated by the Smith and Waterman calculation method, the Needleman and Wunsch calculation method, the Jaeger calculation methods, or any of the other calculation methods. As yet another example, a first sequence has 80 percent homology, as defined herein, to a second sequence if the first sequence is calculated to have 80 percent homology to the second sequence using each of calculation methods (although, in practice, the different calculation methods will often result in different calculated homology percentages).
Hybridization/Selective Hybridization
[0052] The term hybridization typically means a sequence driven interaction between at least two nucleic acid molecules, such as an oligonucleotide inhibitor, a primer or a probe and a gene. Sequence driven interaction means an interaction that occurs between two nucleotides or nucleotide analogs or nucleotide derivatives in a nucleotide specific manner. For example, G interacting with C or A interacting with T are sequence driven interactions. Typically sequence driven interactions occur on the Watson-Crick face or Hoogsteen face of the nucleotide. The hybridization of two nucleic acids is affected by a number of conditions and parameters known to those of skill in the art. For example, the salt concentrations, pH, and temperature of the reaction all affect whether two nucleic acid molecules will hybridize.
[0053] Parameters for selective hybridization between two nucleic acid molecules are well known to those of skill in the art. For example, in some embodiments selective hybridization conditions can be defined as stringent hybridization conditions. For example, stringency of hybridization is controlled by both temperature and salt concentration of either or both of the hybridization and washing steps. For example, the conditions of hybridization to achieve selective hybridization may involve hybridization in high ionic strength solution (6×SSC or 6×SSPE) at a temperature that is about 12-25° C. below the Tm (the melting temperature at which half of the molecules dissociate from their hybridization partners) followed by washing at a combination of temperature and salt concentration chosen so that the washing temperature is about 5° C. to 20° C. below the Tm. The temperature and salt conditions are readily determined empirically in preliminary experiments in which samples of reference DNA immobilized on filters are hybridized to a labeled nucleic acid of interest and then washed under conditions of different stringencies. Hybridization temperatures are typically higher for DNA-RNA and RNA-RNA hybridizations. The conditions can be used as described above to achieve stringency, or as is known in the art. (Sambrook et al., Molecular Cloning: A Laboratory Manual, 2nd Ed., Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y., 1989; Kunkel et al. Methods Enzymol. 1987:154:367, 1987 which is herein incorporated by reference for material at least related to hybridization of nucleic acids). A preferable stringent hybridization condition for a DNA:DNA hybridization can be at about 68° C. (in aqueous solution) in 6×SSC or 6×SSPE followed by washing at 68° C. Stringency of hybridization and washing, if desired, can be reduced accordingly as the degree of complementarity desired is decreased, and further, depending upon the G-C or A-T richness of any area wherein variability is searched for. Likewise, stringency of hybridization and washing, if desired, can be increased accordingly as homology desired is increased, and further, depending upon the G-C or A-T richness of any area wherein high homology is desired, all as known in the art.
[0054] Another way to define selective hybridization is by looking at the amount (percentage) of one of the nucleic acids bound to the other nucleic acid. For example, in some embodiments selective hybridization conditions would be when at least about, 60, 65, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 100 percent of the limiting nucleic acid is bound to the non-limiting nucleic acid. Typically, the non-limiting nucleic acid is in for example, 10 or 100 or 1000 fold excess. This type of assay can be performed at under conditions where both the limiting and non-limiting nucleic acid are for example, 10 fold or 100 fold or 1000 fold below their kd, or where only one of the nucleic acid molecules is 10 fold or 100 fold or 1000 fold or where one or both nucleic acid molecules are above their kd.
[0055] Another way to define selective hybridization is by looking at the percentage of nucleic acid that gets enzymatically manipulated under conditions where hybridization is required to promote the desired enzymatic manipulation, e.g., for primers. For example, in some embodiments selective hybridization conditions would be when at least about, 60, 65, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 100 percent of the primer is enzymatically manipulated under conditions which promote the enzymatic manipulation, for example if the enzymatic manipulation is DNA extension, then selective hybridization conditions would be when at least about 60, 65, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 100 percent of the primer molecules are extended. Preferred conditions also include those suggested by the manufacturer or indicated in the art as being appropriate for the enzyme performing the manipulation.
[0056] Just as with homology, it is understood that there are a variety of methods herein disclosed for determining the level of hybridization between two nucleic acid molecules. It is understood that these methods and conditions may provide different percentages of hybridization between two nucleic acid molecules, but unless otherwise indicated meeting the parameters of any of the methods would be sufficient. For example if 80% hybridization was required and as long as hybridization occurs within the required parameters in any one of these methods it is considered disclosed herein.
[0057] It is understood that those of skill in the art understand that if a composition or method meets any one of these criteria for determining hybridization either collectively or singly it is a composition or method that is disclosed herein.
Nucleotides and Related Molecules
[0058] A nucleotide is a molecule that contains a base moiety, a sugar moiety and a phosphate moiety. Nucleotides can be linked together through their phosphate moieties and sugar moieties creating an internucleoside linkage. The base moiety of a nucleotide can be adenin-9-yl (A), cytosin-1-yl (C), guanin-9-yl (G), uracil-1-yl (U), and thymin-1-yl (T). The sugar moiety of a nucleotide is a ribose or a deoxyribose. The phosphate moiety of a nucleotide is pentavalent phosphate. A non-limiting example of a nucleotide would be 3'-AMP (3'-adenosine monophosphate) or 5'-GMP (5'-guanosine monophosphate).
[0059] A nucleotide analog is a nucleotide which contains some type of modification to any of the base, sugar, or phosphate moieties. Modifications to the base moiety would include natural and synthetic modifications of A, C, G, and T/U as well as different purine or pyrimidine bases, such as uracil-5-yl (ψ), hypoxanthin-9-yl (I), and 2-aminoadenin-9-yl. A modified base includes but is not limited to 5-methylcytosine (5-me-C), 5-hydroxymethyl cytosine, xanthine, hypoxanthine, 2-aminoadenine, 6-methyl and other alkyl derivatives of adenine and guanine, 2-propyl and other alkyl derivatives of adenine and guanine, 2-thiouracil, 2-thiothymine and 2-thiocytosine, 5-halouracil and cytosine, 5-propynyl uracil and cytosine, 6-azo uracil, cytosine and thymine, 5-uracil (pseudouracil), 4-thiouracil, 8-halo, 8-amino, 8-thiol, 8-thioalkyl, 8-hydroxyl and other 8-substituted adenines and guanines, 5-halo particularly 5-bromo, 5-trifluoromethyl and other 5-substituted uracils and cytosines, 7-methylguanine and 7-methyladenine, -azaguanine and 8-azaadenine, -deazaguanine and 7-deazaadenine and 3-deazaguanine and 3-deazaadenine. Additional base modifications can be found for example in U.S. Pat. No. 3,687,808, Englisch et al., Angewandte Chemie, International Edition, 1991, 30, 613, and Sanghvi, Y. S., Chapter 15, Antisense Research and Applications, pages 289-302, Crooke, S. T. and Lebleu, B. ed., CRC Press, 1993. Certain nucleotide analogs, such as 5-substituted pyrimidines, 6-azapyrimidines and N-2, N-6 and O-6 substituted purines, including 2-aminopropyladenine, 5-propynyluracil and 5-propynylcytosine. 5-methylcytosine can increase the stability of duplex formation. Often time base modifications can be combined with for example a sugar modification, such as 2'-O-methoxyethyl, to achieve unique properties such as increased duplex stability. There are numerous United States patents such as U.S. Pat. Nos. 4,845,205; 5,130,302; 5,134,066; 5,175,273; 5,367,066; 5,432,272; 5,457,187; 5,459,255; 5,484,908; 5,502,177; 5,525,711; 5,552,540; 5,587,469; 5,594,121, 5,596,091; 5,614,617; and 5,681,941, which detail and describe a range of base modifications. Each of these patents is herein incorporated by reference.
[0060] Nucleotide analogs can also include modifications of the sugar moiety. Modifications to the sugar moiety would include natural modifications of the ribose and deoxyribose as well as synthetic modifications. Sugar modifications include but are not limited to the following modifications at the 2' position: OH; F; O-, S-, or N-alkyl; O-, S-, or N-alkenyl; O-, S- or N-alkynyl; or O-alkyl-O-alkyl, wherein the alkyl, alkenyl and alkynyl may be substituted or unsubstituted C1 to C10, alkyl or C2 to C10 alkenyl and alkynyl. 2' sugar modifications also include but are not limited to --O[(CH2)nO]mCH3, --O(CH2)nOCH3, --O(CH2)nNH2, --O(CH2)nCH3, --O(CH2)n--ONH2, and --O(CH2)nON[(CH2)nCH3)]2, where n and m are from 1 to about 10.
[0061] Other modifications at the 2' position include but are not limited to: C1 to C10 lower alkyl, substituted lower alkyl, alkaryl, aralkyl, O-alkaryl or O-aralkyl, SH, SCH3, OCN, Cl, Br, CN, CF3, OCF3, SOCH3, SO2CH3, ONO2, NO2, N3, NH2, heterocycloalkyl, heterocycloalkaryl, aminoalkylamino, polyalkylamino, substituted silyl, an RNA cleaving group, a reporter group, an intercalator, a group for improving the pharmacokinetic properties of an oligonucleotide, or a group for improving the pharmacodynamic properties of an oligonucleotide, and other substituents having similar properties. Similar modifications may also be made at other positions on the sugar, particularly the 3' position of the sugar on the 3' terminal nucleotide or in 2'-5' linked oligonucleotides and the 5' position of 5' terminal nucleotide. Modified sugars would also include those that contain modifications at the bridging ring oxygen, such as CH2 and S, Nucleotide sugar analogs may also have sugar mimetics such as cyclobutyl moieties in place of the pentofuranosyl sugar. There are numerous United States patents that teach the preparation of such modified sugar structures such as U.S. Pat. Nos. 4,981,957; 5,118,800; 5,319,080; 5,359,044; 5,393,878; 5,446,137; 5,466,786; 5,514,785; 5,519,134; 5,567,811; 5,576,427; 5,591,722; 5,597,909; 5,610,300; 5,627,053; 5,639,873; 5,646,265; 5,658,873; 5,670,633; and 5,700,920, each of which is herein incorporated by reference in its entirety.
[0062] Nucleotide analogs can also be modified at the phosphate moiety. Modified phosphate moieties include but are not limited to those that can be modified so that the linkage between two nucleotides contains a phosphorothioate, chiral phosphorothioate, phosphorodithioate, phosphotriester, aminoalkylphosphotriester, methyl and other alkyl phosphonates including 3'-alkylene phosphonate and chiral phosphonates, phosphinates, phosphoramidates including 3'-amino phosphoramidate and aminoalkylphosphoramidates, thionophosphoramidates, thionoalkylphosphonates, thionoalkylphosphotriesters, and boranophosphates. It is understood that these phosphate or modified phosphate linkage between two nucleotides can be through a linkage or a 2'-5' linkage, and the linkage can contain inverted polarity such as 3'-5' to 5'-3' or 2'-5' to 5'-2'. Various salts, mixed salts and free acid forms are also included. Numerous United States patents teach how to make and use nucleotides containing modified phosphates and include but are not limited to, U.S. Pat. Nos. 3,687,808; 4,469,863; 4,476,301; 5,023,243; 5,177,196; 5,188,897; 5,264,423; 5,276,019; 5,278,302; 5,286,717; 5,321,131; 5,399,676; 5,405,939; 5,453,496; 5,455,233; 5,466,677; 5,476,925; 5,519,126; 5,536,821; 5,541,306; 5,550,111; 5,563,253; 5,571,799; 5,587,361; and 5,625,050, each of which is herein incorporated by reference.
[0063] It is understood that nucleotide analogs need only contain a single modification, but may also contain multiple modifications within one of the moieties or between different moieties.
[0064] Nucleotide substitutes are molecules having similar functional properties to nucleotides, but which do not contain a phosphate moiety, such as peptide nucleic acid (PNA). Nucleotide substitutes are molecules that will recognize nucleic acids in a Watson-Crick or Hoogsteen manner, but which are linked together through a moiety other than a phosphate moiety. Nucleotide substitutes are able to conform to a double helix type structure when interacting with the appropriate target nucleic acid.
[0065] Nucleotide substitutes are nucleotides or nucleotide analogs that have had the phosphate moiety and/or sugar moieties replaced. Nucleotide substitutes do not contain a standard phosphorus atom. Substitutes for the phosphate can be for example, short chain alkyl or cycloalkyl internucleoside linkages, mixed heteroatom and alkyl or cycloalkyl internucleoside linkages, or one or more short chain heteroatomic or heterocyclic internucleoside linkages. These include those having morpholino linkages (formed in part from the sugar portion of a nucleoside); siloxane backbones; sulfide, sulfoxide and sulfone backbones; formacetyl and thioformacetyl backbones; methylene formacetyl and thioformacetyl backbones; alkene containing backbones; sulfamate backbones; methyleneimino and methylenehydrazino backbones; sulfonate and sulfonamide backbones; amide backbones; and others having mixed N, O, S and CH2 component parts. Numerous United States patents disclose how to make and use these types of phosphate replacements and include but are not limited to U.S. Pat. Nos. 5,034,506; 5,166,315; 5,185,444; 5,214,134; 5,216,141; 5,235,033; 5,264,562; 5,264,564; 5,405,938; 5,434,257; 5,466,677; 5,470,967; 5,489,677; 5,541,307; 5,561,225; 5,596,086; 5,602,240; 5,610,289; 5,602,240; 5,608,046; 5,610,289; 5,618,704; 5,623,070; 5,663,312; 5,633,360; 5,677,437; and 5,677,439, each of which is herein incorporated by reference.
[0066] It is also understood in a nucleotide substitute that both the sugar and the phosphate moieties of the nucleotide can be replaced, by for example an amide type linkage (aminoethylglycine) (PNA). U.S. Pat. Nos. 5,539,082; 5,714,331; and 5,719,262 teach how to make and use PNA molecules, each of which is herein incorporated by reference. (See also Nielsen et al., Science, 1991, 254, 1497-1500).
[0067] It is also possible to link other types of molecules (conjugates) to nucleotides or nucleotide analogs to enhance for example, cellular uptake. Conjugates can be chemically linked to the nucleotide or nucleotide analogs. Such conjugates include but are not limited to lipid moieties such as a cholesterol moiety (Letsinger et al., Proc. Natl. Acad. Sci. USA, 1989, 86, 6553-6556), cholic acid (Manoharan et al., Bioorg. Med. Chem. Let., 1994, 4, 1053-1060), a thioether, e.g., hexyl-5-tritylthiol (Manoharan et al., Ann. N.Y. Acad. Sci., 1992, 660, 306-309; Manoharan et al., Bioorg. Med. Chem. Let., 1993, 3, 2765-2770), a thiocholesterol (Oberhauser et al., Nucl. Acids Res., 1992, 20, 533-538), an aliphatic chain, e.g., dodecandiol or undecyl residues (Saison-Behmoaras et al., EMBO J., 1991, 10, 1111-1118; Kabanov et al., FEBS Lett., 1990, 259, 327-330; Svinarchuk et al., Biochimie, 1993, 75, 49-54), a phospholipid, e.g., di-hexadecyl-rac-glycerol or triethylammonium 1,2-di-O-hexadecyl-rac-glycero-3-H-phosphonate (Manoharan et al., Tetrahedron Lett., 1995, 36, 3651-3654; Shea et al., Nucl. Acids Res., 1990, 18, 3777-3783), a polyamine or a polyethylene glycol chain (Manoharan et al., Nucleosides & Nucleotides, 1995, 14, 969-973), or adamantane acetic acid (Manoharan et al., Tetrahedron Lett., 1995, 36, 3651-3654), a palmityl moiety (Mishra et al., Biochim. Biophys. Acta, 1995, 1264, 229-237), or an octadecylamine or hexylamino-carbonyl-oxycholesterol moiety (Crooke et al., J. Pharmacol. Exp. Ther., 1996, 277, 923-937. Numerous United States patents teach the preparation of such conjugates and include, but are not limited to U.S. Pat. Nos. 4,828,979; 4,948,882; 5,218,105; 5,525,465; 5,541,313; 5,545,730; 5,552,538; 5,578,717, 5,580,731; 5,580,731; 5,591,584; 5,109,124; 5,118,802; 5,138,045; 5,414,077; 5,486,603; 5,512,439; 5,578,718; 5,608,046; 4,587,044; 4,605,735; 4,667,025; 4,762,779; 4,789,737; 4,824,941; 4,835,263; 4,876,335; 4,904,582; 4,958,013; 5,082,830; 5,112,963; 5,214,136; 5,082,830; 5,112,963; 5,214,136; 5,245,022; 5,254,469; 5,258,506; 5,262,536; 5,272,250; 5,292,873; 5,317,098; 5,371,241, 5,391,723; 5,416,203, 5,451,463; 5,510,475; 5,512,667; 5,514,785; 5,565,552; 5,567,810; 5,574,142; 5,585,481; 5,587,371; 5,595,726; 5,597,696; 5,599,923; 5,599,928 and 5,688,941, each of which is herein incorporated by reference.
[0068] A Watson-Crick interaction is at least one interaction with the Watson-Crick face of a nucleotide, nucleotide analog, or nucleotide substitute. The Watson-Crick face of a nucleotide, nucleotide analog, or nucleotide substitute includes the C2, N1, and C6 positions of a purine based nucleotide, nucleotide analog, or nucleotide substitute and the C2, N3, C4 positions of a pyrimidine based nucleotide, nucleotide analog, or nucleotide substitute.
[0069] A Hoogsteen interaction is the interaction that takes place on the Hoogsteen face of a nucleotide or nucleotide analog, which is exposed in the major groove of duplex DNA. The Hoogsteen face includes the N7 position and reactive groups (NH2 or O) at the C6 position of purine nucleotides.
Sequences
[0070] There are a variety of sequences related to the DEFB1 gene and to the PAX2 transcriptional factor, respectively, having the following GenBank Accession Numbers: U50930 and NM--003989.1. These sequences and others are herein incorporated by reference in their entireties as well as for individual subsequences contained therein.
[0071] The one particular sequence set forth in SEQ ID NO: 64 and having GenBank accession number U50930 is used herein as an example to exemplify a source for the disclosed DEFB1 nucleic acids. The one particular sequence set forth in SEQ ID NO: 46 and having GenBank accession number NM--003989.1 is used herein as an example, to exemplify a source for the disclosed PAX2 nucleic acids. Other examples of PAX2 sequences, based on alternative splicing are also found in GenBank. These are variants a-e, shown in Appendices B-F. It is understood that the description related to this sequence is applicable to any sequence related to unless specifically indicated otherwise. Those of skill in the art understand how to resolve sequence discrepancies and differences and to adjust the compositions and methods relating to a particular sequence to other related sequences. siRNA molecules, competitive inhibitors of DEFB1 promoter-PAX2, and primers and/or probes can be designed for any DEFB1 or PAX2 sequence given the information disclosed herein and known in the art.
Nucleic Acid Synthesis
[0072] The nucleic acids, such as, the oligonucleotides to be used as inhibitors can be made using standard chemical synthesis methods or can be produced using enzymatic methods or any other known method. Such methods can range from standard enzymatic digestion followed by nucleotide fragment isolation (see for example, Sambrook et al., Molecular Cloning: A Laboratory Manual, 2nd Edition (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y., 1989) Chapters 5, 6) to purely synthetic methods, for example, by the cyanoethyl phosphoramidite method using a Milligen or Beckman System 1Plus DNA synthesizer (for example, Model 8700 automated synthesizer of Milligen-Biosearch, Burlington, Mass. or ABI Model 380B). Synthetic methods useful for making oligonucleotides are also described by Ikuta et al., Ann. Rev. Biochem. 53:323-356 (1984), (phosphotriester and phosphite-triester methods), and Narang et al., Methods Enzymol., 65:610-620 (1980), (phosphotriester method). Protein nucleic acid molecules can be made using known methods such as those described by Nielsen et al., Bioconjug. Chem. 5:3-7 (1994).
Primers and Probes
[0073] Disclosed are compositions including primers and probes, which are capable of interacting with the DEFB1 gene as disclosed herein. In certain embodiments the primers are used to support DNA amplification reactions. Typically the primers will be capable of being extended in a sequence specific manner. Extension of a primer in a sequence specific manner includes any methods wherein the sequence and/or composition of the nucleic acid molecule to which the primer is hybridized or otherwise associated directs or influences the composition or sequence of the product produced by the extension of the primer. Extension of the primer in a sequence specific manner therefore includes, but is not limited to, PCR, DNA sequencing, DNA extension, DNA polymerization, RNA transcription, or reverse transcription. Techniques and conditions that amplify the primer in a sequence specific manner are preferred. In certain embodiments the primers are used for the DNA amplification reactions, such as PCR or direct sequencing. It is understood that in certain embodiments the primers can also be extended using non-enzymatic techniques, where for example, the nucleotides or oligonucleotides used to extend the primer are modified such that they will chemically react to extend the primer in a sequence specific manner.
[0074] The size of the primers or probes for interaction with the DEFB1 or PAX2 gene in certain embodiments can be any size that supports the desired enzymatic manipulation of the primer, such as DNA amplification or the simple hybridization of the probe or primer. A typical primer or probe would be at least 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 125, 150, 175, 200, 225, 250, 275, 300, 325, 350, 375, 400, 425, 450, 475, 500, 550, 600, 650, 700, 750, 800, 850, 900, 950, 1000, 1250, 1500, 1750, 2000, 2250, 2500, 2750, 3000, 3500, or 4000 nucleotides long.
[0075] In other embodiments a primer or probe can be less than or equal to 6, 7, 8, 9, 10, 11, 12 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 125, 150, 175, 200, 225, 250, 275, 300, 325, 350, 375, 400, 425, 450, 475, 500, 550, 600, 650, 700, 750, 800, 850, 900, 950, 1000, 1250, 1500, 1750, 2000, 2250, 2500, 2750, 3000, 3500, or 4000 nucleotides long.
[0076] The primers for the DEFB1 or PAX2 gene typically will be used to produce an amplified DNA product that contains the region of the DEFB1 gene to which PAX2 binds. In general, typically the size of the product will be such that the size can be accurately determined to within 3, or 2 or 1 nucleotides.
[0077] In certain embodiments this product is at least 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 125, 150, 175, 200, 225, 250, 275, 300, 325, 350, 375, 400, 425, 450, 475, 500, 550, 600, 650, 700, 750, 800, 850, 900, 950, 1000, 1250, 1500, 1750, 2000, 2250, 2500, 2750, 3000, 3500, or 4000 nucleotides long.
[0078] In other embodiments the product is less than or equal to 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 125, 150, 175, 200, 225, 250, 275, 300, 325, 350, 375, 400, 425, 450, 475, 500, 550, 600, 650, 700, 750, 800, 850, 900, 950, 1000, 1250, 1500, 1750, 2000, 2250, 2500, 2750, 3000, 3500, or 4000 nucleotides long.
Functional Nucleic Acids
[0079] RNAi
[0080] It is also understood that the disclosed nucleic acids can be used for RNAi or RNA interference. It is thought that RNAi involves a two-step mechanism for RNA interference (RNAi): an initiation step and an effector step. For example, in the first step, input double-stranded (ds) RNA (siRNA) is processed into small fragments, such as 21-23-nucleotide `guide sequences`. RNA amplification occurs in whole animals. Typically then, the guide RNAs can be incorporated into a protein RNA complex which is capable of degrading RNA, the nuclease complex, which has been called the RNA-induced silencing complex (RISC). This RISC complex acts in the second effector step to destroy mRNAs that are recognized by the guide RNAs through base-pairing interactions. RNAi involves the introduction by any means of double stranded RNA into the cell which triggers events that cause the degradation of a target RNA. RNAi is a form of post-transcriptional gene silencing. In addition to the siRNAs disclosed herein, disclosed are RNA hairpins that can act in RNAi. For description of making and using RNAi molecules see, e.g., Hammond et al., Nature Rev Gen 2: 110-119 (2001); Sharp, Genes Dev 15: 485-490 (2001), Waterhouse et al., Proc. Natl. Acad. Sci. USA 95(23): 13959-13964 (1998) all of which are incorporated herein by reference in their entireties and at least form material related to delivery and making of RNAi molecules.
[0081] RNAi has been shown to work in many types of cells, including mammalian cells. For work in mammalian cells it is preferred that the RNA molecules which will be used as targeting sequences within the RISC complex are shorter. For example, less than or equal to 50 or 40 or 30 or 29, 28, 27, 26, 25, 24, 23, 22, 21, 20, 19, 18, 17, 16, 15, 14, 13, 12, 11, or 10 nucleotides in length. These RNA molecules can also have overhangs on the 3' or 5' ends relative to the target RNA which is to be cleaved. These overhangs can be at least or less than or equal to 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 15, or 20 nucleotides long. RNAi works in mammalian stem cells, such as mouse ES cells. Examples of siRNAs can be found in Table 4.
[0082] Functional nucleic acids are nucleic acid molecules that have a specific function, such as binding a target molecule or catalyzing a specific reaction. Functional nucleic acid molecules can be divided into the following categories, which are not meant to be limiting. For example, functional nucleic acids include antisense molecules, aptamers, ribozymes, triplex forming molecules, and external guide sequences. The functional nucleic acid molecules can act as effectors, inhibitors, modulators, and stimulators of a specific activity possessed by a target molecule, or the functional nucleic acid molecules can possess a de novo activity independent of any other molecules.
[0083] Functional nucleic acid molecules can interact with any macromolecule, such as DNA, RNA, polypeptides, or carbohydrate chains. Thus, functional nucleic acids can interact with mRNA or the genomic DNA of PAX2. Often functional nucleic acids are designed to interact with other nucleic acids based on sequence homology between the target molecule and the functional nucleic acid molecule. In other situations, the specific recognition between the functional nucleic acid molecule and the target molecule is not based on sequence homology between the functional nucleic acid molecule and the target molecule, but rather is based on the formation of tertiary structure that allows specific recognition to take place.
[0084] Antisense molecules are designed to interact with a target nucleic acid molecule through either canonical or non-canonical base pairing. The interaction of the antisense molecule and the target molecule is designed to promote the destruction of the target molecule through, for example, RNAseH mediated RNA-DNA hybrid degradation. Alternatively the antisense molecule is designed to interrupt a processing function that normally would take place on the target molecule, such as transcription or replication. Antisense molecules can be designed based on the sequence of the target molecule. Numerous methods for optimization of antisense efficiency by finding the most accessible regions of the target molecule exist. Exemplary methods would be in vitro selection experiments and DNA modification studies using DMS and DEPC. It is preferred that antisense molecules bind the target molecule with a dissociation constant (kd) less than or equal to 10-6, 10-8, 10-10, or 10-12. A representative sample of methods and techniques which aid in the design and use of antisense molecules can be found in the following non-limiting list of U.S. Pat. Nos. 5,135,917, 5,294,533, 5,627,158, 5,641,754, 5,691,317, 5,780,607, 5,786,138, 5,849,903, 5,856,103, 5,919,772, 5,955,590, 5,990,088, 5,994,320, 5,998,602, 6,005,095, 6,007,995, 6,013,522, 6,017,898, 6,018,042, 6,025,198, 6,033,910, 6,040,296, 6,046,004, 6,046,319, and 6,057,437.
[0085] Aptamers are molecules that interact with a target molecule, preferably in a specific way. Typically aptamers are small nucleic acids ranging from 15-50 bases in length that fold into defined secondary and tertiary structures, such as stem-loops or G-quartets. Aptamers can bind small molecules, such as ATP (U.S. Pat. No. 5,631,146) and theophiline (U.S. Pat. No. 5,580,737), as well as large molecules, such as reverse transcriptase (U.S. Pat. No. 5,786,462) and thrombin (U.S. Pat. No. 5,543,293). Aptamers can bind very tightly with kds from the target molecule of less than 10-12 M. It is preferred that the aptamers bind the target molecule with a kd less than 10-6, 10-8, 10-10, or 10-12 Aptamers can bind the target molecule with a very high degree of specificity. For example, aptamers have been isolated that have greater than a 10000 fold difference in binding affinities between the target molecule and another molecule that differ at only a single position on the molecule (U.S. Pat. No. 5,543,293). It is preferred that the aptamer have a kd with the target molecule at least 10, 100, 1000, 10,000, or 100,000 fold lower than the kd with a background binding molecule. Representative examples of how to make and use aptamers to bind a variety of different target molecules can be found in the following non-limiting list of U.S. Pat. Nos. 5,476,766, 5,503,978, 5,631,146, 5,731,424, 5,780,228, 5,792,613, 5,795,721, 5,846,713, 5,858,660, 5,861,254, 5,864,026, 5,869,641, 5,958,691, 6,001,988, 6,011,020, 6,013,443, 6,020,130, 6,028,186, 6,030,776, and 6,051,698.
[0086] Ribozymes are nucleic acid molecules that are capable of catalyzing a chemical reaction, either intramolecularly or intermolecularly. Ribozymes are thus catalytic nucleic acid. It is preferred that the ribozymes catalyze intermolecular reactions. There are a number of different types of ribozymes that catalyze nuclease or nucleic acid polymerase type reactions which are based on ribozymes found in natural systems, such as hammerhead ribozymes, (for example, but not limited to the following U.S. Pat. Nos. 5,334,711, 5,436,330, 5,616,466, 5,633,133, 5,646,020, 5,652,094, 5,712,384, 5,770,715, 5,856,463, 5,861,288, 5,891,683, 5,891,684, 5,985,621, 5,989,908, 5,998,193, 5,998,203, WO 9858058 by Ludwig and Sproat, WO 9858057 by Ludwig and Sproat, and WO 9718312 by Ludwig and Sproat) hairpin ribozymes (for example, but not limited to the following U.S. Pat. Nos. 5,631,115, 5,646,031, 5,683,902, 5,712,384, 5,856,188, 5,866,701, 5,869,339, and 6,022,962), and tetrahymena ribozymes (for example, but not limited to the following U.S. Pat. Nos. 5,595,873 and 5,652,107). There are also a number of ribozymes that are not found in natural systems, but which have been engineered to catalyze specific reactions de novo (for example, but not limited to the following U.S. Pat. Nos. 5,580,967, 5,688,670, 5,807,718, and 5,910,408). Preferred ribozymes cleave RNA or DNA substrates, and more preferably cleave RNA substrates. Ribozymes typically cleave nucleic acid substrates through recognition and binding of the target substrate with subsequent cleavage. This recognition is often based mostly on canonical or non-canonical base pair interactions. This property makes ribozymes particularly good candidates for target specific cleavage of nucleic acids because recognition of the target substrate is based on the target substrates sequence. Representative examples of how to make and use ribozymes to catalyze a variety of different reactions can be found in the following non-limiting list of U.S. Pat. Nos. 5,646,042, 5,693,535, 5,731,295, 5,811,300, 5,837,855, 5,869,253, 5,877,021, 5,877,022, 5,972,699, 5,972,704, 5,989,906, and 6,017,756.
[0087] Triplex forming functional nucleic acid molecules are molecules that can interact with either double-stranded or single-stranded nucleic acid. When triplex molecules interact with a target region, a structure called a triplex is formed, in which there three strands of DNA are forming a complex dependant on both Watson-Crick and Hoogsteen base-pairing. Triplex molecules are preferred because they can bind target regions with high affinity and specificity. It is preferred that the triplex forming molecules bind the target molecule with a kd less than 10-6, 10-8, 10-10, or 10-12. Representative examples of how to make and use triplex forming molecules to bind a variety of different target molecules can be found in the following non-limiting list of U.S. Pat. Nos. 5,176,996, 5,645,985, 5,650,316, 5,683,874, 5,693,773, 5,834,185, 5,869,246, 5,874,566, and 5,962,426.
[0088] External guide sequences (EGSs) are molecules that bind a target nucleic acid molecule forming a complex, and this complex is recognized by RNase P, which cleaves the target molecule. EGSs can be designed to specifically target a RNA molecule of choice. RNAse P aids in processing transfer RNA (tRNA) within a cell. Bacterial RNAse P can be recruited to cleave virtually any RNA sequence by using an EGS that causes the target RNA:EGS complex to mimic the natural tRNA substrate. (WO 92/03566 by Yale, and Forster and Altman, Science 238:407-409 (1990)).
[0089] Similarly, eukaryotic EGS/RNAse P-directed cleavage of RNA can be utilized to cleave desired targets within eukaryotic cells. (Yuan et al., Proc. Natl. Acad. Sci. USA 89:8006-8010 (1992); WO 93/22434 by Yale; WO 95/24489 by Yale; Yuan and Altman, EMBO J 14:159-168 (1995), and Carrara et al., Proc. Natl. Acad. Sci. (USA) 92:2627-2631 (1995)). Representative examples of how to make and use EGS molecules to facilitate cleavage of a variety of different target molecules be found in the following non-limiting list of U.S. Pat. Nos. 5,168,053, 5,624,824, 5,683,873, 5,728,521, 5,869,248, and 5,877,162.
Delivery of the Compositions to Cells
[0090] There are a number of compositions and methods which can be used to deliver nucleic acids to cells, either in vitro or in vivo. These methods and compositions can largely be broken down into two classes: viral based delivery systems and non-viral based delivery systems. For example, the nucleic acids can be delivered through a number of direct delivery systems such as, electroporation, lipofection, calcium phosphate precipitation, plasmids, viral vectors, viral nucleic acids, phage nucleic acids, phages, cosmids, or via transfer of genetic material in cells or carriers such as cationic liposomes. Appropriate means for transfection, including viral vectors, chemical transfectants, or physico-mechanical methods such as electroporation and direct diffusion of DNA, are described by, for example, Wolff, J. A., et al., Science, 247, 1465-1468, (1990); and Wolff, J. A. Nature, 352, 815-818, (1991) Such methods are well known in the art and readily adaptable for use with the compositions and methods described herein. In certain cases, the methods will be modified to specifically function with large DNA molecules. Further, these methods can be used to target certain diseases and cell populations by using the targeting characteristics of the carrier.
Nucleic Acid Based Delivery Systems
[0091] Transfer vectors can be any nucleotide construction used to deliver genes into cells (e.g., a plasmid), or as part of a general strategy to deliver genes, e.g., as part of recombinant retrovirus or adenovirus (Ram et al. Cancer Res. 53:83-88, (1993)).
[0092] As used herein, plasmid or viral vectors are agents that transport the disclosed nucleic acids, such as DEFB1 coding sequences, PAX2 siRNAs or other antisense molecules into the cell without degradation and include a promoter yielding expression of the gene in the cells into which it is delivered. Viral vectors are, for example, Adenovirus, Adeno-associated virus, Herpes virus, Vaccinia virus, Polio virus, AIDS virus, neuronal trophic virus, Sindbis and other RNA viruses, including these viruses with the HIV backbone. Also preferred are any viral families which share the properties of these viruses which make them suitable for use as vectors. Retroviruses include Murine Maloney Leukemia virus, MMLV, and retroviruses that express the desirable properties of MMLV as a vector. Retroviral vectors are able to carry a larger genetic payload, i.e., a transgene or marker gene, than other viral vectors, and for this reason are a commonly used vector. However, they are not as useful in non-proliferating cells. Adenovirus vectors are relatively stable and easy to work with, have high titers, and can be delivered in aerosol formulation, and can transfect non-dividing cells. Pox viral vectors are large and have several sites for inserting genes, they are thermostable and can be stored at room temperature. A preferred embodiment is a viral vector which has been engineered so as to suppress the immune response of the host organism, elicited by the viral antigens. Preferred vectors of this type will carry coding regions for Interleukin 8 or 10.
[0093] Viral vectors can have higher transaction (ability to introduce genes) abilities than chemical or physical methods to introduce genes into cells. Typically, viral vectors contain, nonstructural early genes, structural late genes, an RNA polymerase III transcript, inverted terminal repeats necessary for replication and encapsidation, and promoters to control the transcription and replication of the viral genome. When engineered as vectors, viruses typically have one or more of the early genes removed and a gene or gene/promotor cassette is inserted into the viral genome in place of the removed viral DNA. Constructs of this type can carry up to about 8 kb of foreign genetic material. The necessary functions of the removed early genes are typically supplied by cell lines which have been engineered to express the gene products of the early genes in trans.
Retroviral Vectors
[0094] A retrovirus is an animal virus belonging to the virus family of Retroviridae, including any types, subfamilies, genus, or tropisms. Retroviral vectors, in general, are described by Verma, I. M., Retroviral vectors for gene transfer. In Microbiology-1985, American Society for Microbiology, pp. 229-232, Washington, (1985), which is incorporated by reference herein. Examples of methods for using retroviral vectors for gene therapy are described in U.S. Pat. Nos. 4,868,116 and 4,980,286; PCT applications WO 90/02806 and WO 89/07136; and Mulligan, (Science 260:926-932 (1993)); the teachings of which are incorporated herein by reference.
[0095] A retrovirus is essentially a package which has packed into it nucleic acid cargo. The nucleic acid cargo carries with it a packaging signal, which ensures that the replicated daughter molecules will be efficiently packaged within the package coat. In addition to the package signal, there are a number of molecules which are needed in cis, for the replication, and packaging of the replicated virus. Typically a retroviral genome, contains the gag, pol, and env genes which are involved in the making of the protein coat. It is the gag, pol, and env genes which are typically replaced by the foreign DNA that it is to be transferred to the target cell. Retrovirus vectors typically contain a packaging signal for incorporation into the package coat, a sequence which signals the start of the gag transcription unit, elements necessary for reverse transcription, including a primer binding site to bind the tRNA primer of reverse transcription, terminal repeat sequences that guide the switch of RNA strands during DNA synthesis, a purine rich sequence 5' to the 3' LTR that serve as the priming site for the synthesis of the second strand of DNA synthesis, and specific sequences near the ends of the LTRs that enable the insertion of the DNA state of the retrovirus to insert into the host genome. The removal of the gag, pol, and env genes allows for about 8 kb of foreign sequence to be inserted into the viral genome, become reverse transcribed, and upon replication be packaged into a new retroviral particle. This amount of nucleic acid is sufficient for the delivery of a one to many genes depending on the size of each transcript. It is preferable to include either positive or negative selectable markers along with other genes in the insert.
[0096] Since the replication machinery and packaging proteins in most retroviral vectors have been removed (gag, pol, and env), the vectors are typically generated by placing them into a packaging cell line. A packaging cell line is a cell line which has been transfected or transformed with a retrovirus that contains the replication and packaging machinery, but lacks any packaging signal. When the vector carrying the DNA of choice is transfected into these cell lines, the vector containing the gene of interest is replicated and packaged into new retroviral particles, by the machinery provided in cis by the helper cell. The genomes for the machinery are not packaged because they lack the necessary signals.
Adenoviral Vectors
[0097] The construction of replication-defective adenoviruses has been described (Berkner et al., J. Virology 61:1213-1220 (1987); Massie et al., Mol. Cell. Biol. 6:2872-2883 (1986); Haj-Ahmad et al., J. Virology 57:267-274 (1986); Davidson et al., J. Virology 61:1226-1239 (1987); Zhang "Generation and identification of recombinant adenovirus by liposome-mediated transfection and PCR analysis" BioTechniques 15:868-872 (1993)). The benefit of the use of these viruses as vectors is that they are limited in the extent to which they can spread to other cell types, since they can replicate within an initial infected cell, but are unable to form new infectious viral particles. Recombinant adenoviruses have been shown to achieve high efficiency gene transfer after direct, in vivo delivery to airway epithelium, hepatocytes, vascular endothelium, CNS parenchyma and a number of other tissue sites (Morsy, J. Clin. Invest. 92:1580-1586 (1993); Kirshenbaum, J. Clin. Invest. 92:381-387 (1993); Roessler, J. Clin. Invest. 92:1085-1092 (1993); Moullier, Nature Genetics 4:154-159 (1993); La Salle, Science 259:988-990 (1993); Gomez-Foix, J. Biol. Chem. 267:25129-25134 (1992); Rich, Human Gene Therapy 4:461-476 (1993); Zabner, Nature Genetics 6:75-83 (1994); Guzman, Circulation Research 73:1201-1207 (1993); Bout, Human Gene Therapy 5:3-10 (1994); Zabner, Cell 75:207-216 (1993); Caillaud, Eur. J. Neuroscience 5:1287-1291 (1993); and Ragot, J. Gen. Virology 74:501-507 (1993)). Recombinant adenoviruses achieve gene transduction by binding to specific cell surface receptors, after which the virus is internalized by receptor-mediated endocytosis, in the same manner as wild type or replication-defective adenovirus (Chardonnet and Dales, Virology 40:462-477 (1970); Brown and Burlingham, J. Virology 12:386-396 (1973); Svensson and Persson, J. Virology 55:442-449 (1985); Seth, et al., J. Virol. 51:650-655 (1984); Seth, et al., Mol. Cell. Biol. 4:1528-1533 (1984); Varga et al., J. Virology 65:6061-6070 (1991); Wickham et al., Cell 73:309-319 (1993)).
[0098] A viral vector can be one based on an adenovirus which has had the E1 gene removed and these virions are generated in a cell line such as the human 293 cell line. In another preferred embodiment both the E1 and E3 genes are removed from the adenovirus genome.
Adeno-Associated Viral Vectors
[0099] Another type of viral vector is based on an adeno-associated virus (AAV). This defective parvovirus is a preferred vector because it can infect many cell types and is nonpathogenic to humans. AAV type vectors can transport about 4 to 5 kb and wild type AAV is known to stably insert into chromosome 19. Vectors which contain this site specific integration property are preferred. An especially preferred embodiment of this type of vector is the P4.1 C vector produced by Avigen, San Francisco, Calif., which can contain the herpes simplex virus thymidine kinase gene, HSV-tk, and/or a marker gene, such as the gene encoding the green fluorescent protein, GFP.
[0100] In another type of AAV virus, the AAV contains a pair of inverted terminal repeats (ITRs) which flank at least one cassette containing a promoter which directs cell-specific expression operably linked to a heterologous gene. Heterologous in this context refers to any nucleotide sequence or gene which is not native to the AAV or B19 parvovirus.
[0101] Typically the AAV and B19 coding regions have been deleted, resulting in a safe, noncytotoxic vector. The AAV ITRs, or modifications thereof, confer infectivity and site-specific integration, but not cytotoxicity, and the promoter directs cell-specific expression. U.S. Pat. No. 6,261,834 is herein incorporated by reference for material related to the AAV vector.
[0102] 1. The disclosed vectors thus provide DNA molecules which are capable of integration into a mammalian chromosome without substantial toxicity.
[0103] 2. The inserted genes in viral and retroviral usually contain promoters, and/or enhancers to help control the expression of the desired gene product. A promoter is generally a sequence or sequences of DNA that function when in a relatively fixed location in regard to the transcription start site. A promoter contains core elements required for basic interaction of RNA polymerase and transcription factors, and may contain upstream elements and response elements.
Large Payload Viral Vectors
[0104] Molecular genetic experiments with large human herpes viruses have provided a means whereby large heterologous DNA fragments can be cloned, propagated and established in cells permissive for infection with herpes viruses (Sun et al., Nature genetics 8: 33-41, 1994; Cotter and Robertson, Curr Opin Mol Ther 5: 633-644, 1999). These large DNA viruses (herpes simplex virus (HSV) and Epstein-Barr virus (EBV), have the potential to deliver fragments of human heterologous DNA >150 kb to specific cells. EBV recombinants can maintain large pieces of DNA in the infected B-cells as episomal DNA. Individual clones carried human genomic inserts up to 330 kb appeared genetically stable. The maintenance of these episomes requires a specific EBV nuclear protein, EBNA1, constitutively expressed during infection with EBV. Additionally, these vectors can be used for transfection, where large amounts of protein can be generated transiently in vitro. Herpesvirus amplicon systems are also being used to package pieces of DNA >220 kb and to infect cells that can stably maintain DNA as episomes.
[0105] Other useful systems include, for example, replicating and host-restricted non-replicating vaccinia virus vectors.
Non-Nucleic Acid Based Systems
[0106] The disclosed compositions can be delivered to the target cells in a variety of ways. For example, the compositions can be delivered through electroporation, or through lipofection, or through calcium phosphate precipitation. The delivery mechanism chosen will depend in part on the type of cell targeted and whether the delivery is occurring for example in vivo or in vitro.
[0107] Thus, the compositions can comprise, in addition to the disclosed vectors for example, lipids such as liposomes, such as cationic liposomes (e.g., DOTMA, DOPE, and DC-cholesterol) or anionic liposomes. Liposomes can further comprise proteins to facilitate targeting a particular cell, if desired. Administration of a composition comprising a compound and a cationic liposome can be administered to the blood afferent to a target organ or inhaled into the respiratory tract to target cells of the respiratory tract. Regarding liposomes, see, e.g., Brigham et al. Am. J. Resp. Cell. Mol. Biol. 1:95-100 (1989); Feigner et al. Proc. Natl. Acad. Sci USA 84:7413-7417 (1987); U.S. Pat. No. 4,897,355. Furthermore, the compound can be administered as a component of a microcapsule that can be targeted to specific cell types, such as macrophages, or where the diffusion of the compound or delivery of the compound from the microcapsule is designed for a specific rate or dosage.
[0108] In the methods described above which include the administration and uptake of exogenous DNA into the cells of a subject (i.e., gene transduction or transfection), delivery of the compositions to cells can be via a variety of mechanisms. As one example, delivery can be via a liposome, using commercially available liposome preparations such as LIPOFECTIN, LIPOFECTAMINE (GIBCO-BRL, Inc., Gaithersburg, Md.), SUPERFECT (Qiagen, Inc. Hilden, Germany) and TRANSFECTAM (Promega Biotec, Inc., Madison, Wis.), as well as other liposomes developed according to procedures standard in the art. In addition, the disclosed nucleic acid or vector can be delivered in vivo by electroporation, the technology for which is available from Genetronics, Inc. (San Diego, Calif.) as well as by means of a SONOPORATION machine (ImaRx Pharmaceutical Corp., Tucson, Ariz.).
[0109] The materials may be in solution, suspension (for example, incorporated into microparticles, liposomes, or cells). These may be targeted to a particular cell type via antibodies, receptors, or receptor ligands. The following references are examples of the use of this technology to target specific proteins to tumor tissue (Senter, et al., Bioconjugate Chem., 2:447-451, (1991); Bagshawe, K. D., Br. J. Cancer, 60:275-281, (1989); Bagshawe, et al., Br. J. Cancer, 58:700-703, (1988); Senter, et al., Bioconjugate Chem., 4:3-9, (1993); Battelli, et al., Cancer Immunol. Immunother., 35:421-425, (1992); Pietersz and McKenzie, Immunolog. Reviews, 129:57-80, (1992); and Roffler, et al., Biochem. Pharmacol, 42:2062-2065, (1991)). These techniques can be used for a variety of other specific cell types. Vehicles such as "stealth" and other antibody conjugated liposomes (including lipid mediated drug targeting to colonic carcinoma), receptor mediated targeting of DNA through cell specific ligands, lymphocyte directed tumor targeting, and highly specific therapeutic retroviral targeting of murine glioma cells in vivo. The following references are examples of the use of this technology to target specific proteins to tumor tissue (Hughes et al., Cancer Research, 49:6214-6220, (1989); and Litzinger and Huang, Biochimica et Biophysica Acta, 1104:179-187, (1992)). In general, receptors are involved in pathways of endocytosis, either constitutive or ligand induced. These receptors cluster in clathrin-coated pits, enter the cell via clathrin-coated vesicles, pass through an acidified endosome in which the receptors are sorted, and then either recycle to the cell surface, become stored intracellular, or are degraded in lysosomes. The internalization pathways serve a variety of functions, such as nutrient uptake, removal of activated proteins, clearance of macromolecules, opportunistic entry of viruses and toxins, dissociation and degradation of ligand, and receptor-level regulation. Many receptors follow more than one intracellular pathway, depending on the cell type, receptor concentration, type of ligand, ligand valency, and ligand concentration. Molecular and cellular mechanisms of receptor-mediated endocytosis has been reviewed (Brown and Greene, DNA and Cell Biology 10:6, 399-409 (1991)).
[0110] Nucleic acids that are delivered to cells which are to be integrated into the host cell genome, typically contain integration sequences. These sequences are often viral related sequences, particularly when viral based systems are used. These viral integration systems can also be incorporated into nucleic acids which are to be delivered using a non-nucleic acid based system of deliver, such as a liposome, so that the nucleic acid contained in the delivery system can be come integrated into the host genome.
[0111] Other general techniques for integration into the host genome include, for example, systems designed to promote homologous recombination with the host genome. These systems typically rely on sequence flanking the nucleic acid to be expressed that has enough homology with a target sequence within the host cell genome that recombination between the vector nucleic acid and the target nucleic acid takes place, causing the delivered nucleic acid to be integrated into the host genome. These systems and the methods necessary to promote homologous recombination are known to those of skill in the art.
In Vivo/Ex Vivo
[0112] As described herein, the compositions can be administered in a pharmaceutically acceptable carrier and can be delivered to the subject's cells in vivo and/or ex vivo by a variety of mechanisms well known in the art (e.g., uptake of naked DNA, liposome fusion, intramuscular injection of DNA via a gene gun, endocytosis and the like).
[0113] If ex vivo methods are employed, cells or tissues can be removed and maintained outside the body according to standard protocols well known in the art. The compositions can be introduced into the cells via any gene transfer mechanism, such as, for example, calcium phosphate mediated gene delivery, electroporation, microinjection or proteoliposomes. The transduced cells can then be infused (e.g., in a pharmaceutically acceptable carrier) or homotopically transplanted back into the subject per standard methods for the cell or tissue type. Standard methods are known for transplantation or infusion of various cells into a subject.
Expression Systems
[0114] The nucleic acids that are delivered to cells typically contain expression controlling systems. For example, the inserted genes in viral and retroviral systems usually contain promoters, and/or enhancers to help control the expression of the desired gene product. A promoter is generally a sequence or sequences of DNA that function when in a relatively fixed location in regard to the transcription start site. A promoter contains core elements required for basic interaction of RNA polymerase and transcription factors, and may contain upstream elements and response elements.
Viral Promoters and Enhancers
[0115] Preferred promoters controlling transcription from vectors in mammalian host cells may be obtained from various sources, for example, the genomes of viruses such as: polyoma, Simian Virus 40 (SV40), adenovirus, retroviruses, hepatitis-B virus and most preferably cytomegalovirus, or from heterologous mammalian promoters, e.g. beta actin promoter. The early and late promoters of the SV40 virus are conveniently obtained as an SV40 restriction fragment which also contains the SV40 viral origin of replication (Fiers et al., Nature, 273: 113 (1978)). The immediate early promoter of the human cytomegalovirus is conveniently obtained as a HindIII E restriction fragment (Greenway, P. J. et al., Gene 18: 355-360 (1982)). Of course, promoters from the host cell or related species also are useful herein.
[0116] Enhancer generally refers to a sequence of DNA that functions at no fixed distance from the transcription start site and can be either 5' (Laimins, L. et al., Proc. Natl. Acad. Sci. 78: 993 (1981)) or 3' (Lusky, M. L., et al., Mol. Cell Bio. 3: 1108 (1983)) to the transcription unit. Furthermore, enhancers can be within an intron (Banerji, J. L. et al., Cell 33: 729 (1983)) as well as within the coding sequence itself (Osborne, T. F., et al., Mol. Cell Bio. 4: 1293 (1984)). They are usually between 10 and 300 bp in length, and they function in cis. Enhancers function to increase transcription from nearby promoters. Enhancers also often contain response elements that mediate the regulation of transcription. Promoters can also contain response elements that mediate the regulation of transcription. Enhancers often determine the regulation of expression of a gene. While many enhancer sequences are now known from mammalian genes (globin, elastase, albumin, -fetoprotein and insulin), typically one will use an enhancer from a eukaryotic cell virus for general expression. Preferred examples are the SV40 enhancer on the late side of the replication origin (bp 100-270), the cytomegalovirus early promoter enhancer, the polyoma enhancer on the late side of the replication origin, and adenovirus enhancers.
[0117] The promotor and/or enhancer may be specifically activated either by light or specific chemical events which trigger their function. Systems can be regulated by reagents such as tetracycline and dexamethasone. There are also ways to enhance viral vector gene expression by exposure to irradiation, such as gamma irradiation, or alkylating chemotherapy drugs.
[0118] In certain embodiments the promoter and/or enhancer region can act as a constitutive promoter and/or enhancer to maximize expression of the region of the transcription unit to be transcribed. In certain constructs the promoter and/or enhancer region be active in all eukaryotic cell types, even if it is only expressed in a particular type of cell at a particular time. A preferred promoter of this type is the CMV promoter (650 bases). Other preferred promoters are SV40 promoters, cytomegalovirus (full length promoter), and retroviral vector LTR.
[0119] It has been shown that all specific regulatory elements can be cloned and used to construct expression vectors that are selectively expressed in specific cell types such as melanoma cells. The glial fibrillary acetic protein (GFAP) promoter has been used to selectively express genes in cells of glial origin.
[0120] Expression vectors used in eukaryotic host cells (yeast, fungi, insect, plant, animal, human or nucleated cells) may also contain sequences necessary for the termination of transcription which may affect mRNA expression. These regions are transcribed as polyadenylated segments in the untranslated portion of the mRNA encoding tissue factor protein. The 3' untranslated regions also include transcription termination sites. It is preferred that the transcription unit also contain a polyadenylation region. One benefit of this region is that it increases the likelihood that the transcribed unit will be processed and transported like mRNA. The identification and use of polyadenylation signals in expression constructs is well established. It is preferred that homologous polyadenylation signals be used in the transgene constructs. In certain transcription units, the polyadenylation region is derived from the SV40 early polyadenylation signal and consists of about 400 bases. It is also preferred that the transcribed units contain other standard sequences alone or in combination with the above sequences improve expression from, or stability of, the construct.
Protein Variants
[0121] Variants of the DEFB1 protein are provided. Derivatives of the DEFB1 protein function in the disclosed methods and compositions. Protein variants and derivatives are well understood to those of skill in the art and in can involve amino acid sequence modifications. For example, amino acid sequence modifications typically fall into one or more of three classes: substitutional, insertional or deletional variants. Insertions include amino and/or carboxyl terminal fusions as well as intrasequence insertions of single or multiple amino acid residues. Insertions ordinarily will be smaller insertions than those of amino or carboxyl terminal fusions, for example, on the order of one to four residues. Immunogenic fusion protein derivatives, such as those described in the examples, are made by fusing a polypeptide sufficiently large to confer immunogenicity to the target sequence by cross-linking in vitro or by recombinant cell culture transformed with DNA encoding the fusion. Deletions are characterized by the removal of one or more amino acid residues from the protein sequence. Typically, no more than about from 2 to 6 residues are deleted at any one site within the protein molecule. These variants ordinarily are prepared by site specific mutagenesis of nucleotides in the DNA encoding the protein, thereby producing DNA encoding the variant, and thereafter expressing the DNA in recombinant cell culture. Techniques for making substitution mutations at predetermined sites in DNA having a known sequence are well known, for example M13 primer mutagenesis and PCR mutagenesis. Amino acid substitutions are typically of single residues, but can occur at a number of different locations at once; insertions usually will be on the order of about from 1 to 10 amino acid residues; and deletions will range about from 1 to 30 residues. Deletions or insertions preferably are made in adjacent pairs, i.e. a deletion of 2 residues or insertion of 2 residues. Substitutions, deletions, insertions or any combination thereof may be combined to arrive at a final construct. The mutations must not place the sequence out of reading frame and preferably will not create complementary regions that could produce secondary mRNA structure. Substitutional variants are those in which at least one residue has been removed and a different residue inserted in its place. Such substitutions generally are made in accordance with the following Tables 1 and 2 and are referred to as conservative substitutions.
TABLE-US-00004 TABLE 1 Amino Acid Abbreviations Amino Acid Abbreviations alanine AlaA allosoleucine AIle arginine ArgR asparagine AsnN aspartic acid AspD cysteine CysC glutamic acid GluE glutamine GlnK glycine GlyG histidine HisH isolelucine IleI leucine LeuL lysine LysK phenylalanine PheF proline ProP pyroglutamic Glu acidp serine SerS threonine ThrT tyrosine TyrY tryptophan TrpW valine ValV
TABLE-US-00005 TABLE 2 Amino Acid Substitutions Original Residue Exemplary Conservative Substitutions, others are known in the art. Alaser Arglys, gln Asngln; his Aspglu Cysser Glnasn, lys Gluasp Glypro Hisasn; gln Ileleu; val Leuile; val Lysarg; gln; MetLeu; ile Phemet; leu; tyr Serthr Thrser Trptyr Tyrtrp; phe Valile; leu
[0122] Substantial changes in function or immunological identity are made by selecting substitutions that are less conservative than those in Table 2, i.e., selecting residues that differ more significantly in their effect on maintaining (a) the structure of the polypeptide backbone in the area of the substitution, for example as a sheet or helical conformation, (b) the charge or hydrophobicity of the molecule at the target site or (c) the bulk of the side chain. The substitutions which in general are expected to produce the greatest changes in the protein properties will be those in which (a) a hydrophilic residue, e.g. seryl or threonyl, is substituted for (or by) a hydrophobic residue, e.g. leucyl, isoleucyl, phenylalanyl, valyl or alanyl; (b) a cysteine or proline is substituted for (or by) any other residue; (c) a residue having an electropositive side chain, e.g., lysyl, arginyl, or histidyl, is substituted for (or by) an electronegative residue, e.g., glutamyl or aspartyl; or (d) a residue having a bulky side chain, e.g., phenylalanine, is substituted for (or by) one not having a side chain, e.g., glycine, in this case, (e) by increasing the number of sites for sulfation and/or glycosylation.
[0123] For example, the replacement of one amino acid residue with another that is biologically and/or chemically similar is known to those skilled in the art as a conservative substitution. For example, a conservative substitution would be replacing one hydrophobic residue for another, or one polar residue for another. The substitutions include combinations such as, for example, Gly, Ala; Val, Ile, Leu; Asp, Glu; Asn, Gln; Ser, Thr; Lys, Arg; and Phe, Tyr. Such conservatively substituted variations of each explicitly disclosed sequence are included within the mosaic polypeptides provided herein.
[0124] Substitutional or deletional mutagenesis can be employed to insert sites for N-glycosylation (Asn-X-Thr/Ser) or O-glycosylation (Ser or Thr). Deletions of cysteine or other labile residues also may be desirable. Deletions or substitutions of potential proteolysis sites, e.g. Arg, is accomplished for example by deleting one of the basic residues or substituting one by glutaminyl or histidyl residues.
[0125] Certain post-translational derivatizations are the result of the action of recombinant host cells on the expressed polypeptide. Glutaminyl and asparaginyl residues are frequently post-translationally deamidated to the corresponding glutamyl and asparyl residues. Alternatively, these residues are deamidated under mildly acidic conditions. Other post-translational modifications include hydroxylation of proline and lysine, phosphorylation of hydroxyl groups of seryl or threonyl residues, methylation of the o-amino groups of lysine, arginine, and histidine side chains (T. E. Creighton, Proteins: Structure and Molecular Properties, W. H. Freeman & Co., San Francisco pp 79-86 [1983]), acetylation of the N-terminal amine and, in some instances, amidation of the C-terminal carboxyl.
[0126] It is understood that one way to define the variants and derivatives of the disclosed proteins herein is through defining the variants and derivatives in terms of homology/identity to specific known sequences. For example, SEQ ID NO: 63 sets forth a particular sequence of DEFB1 and SEQ ID NO: 45 sets forth a particular sequence of PAX2. Specifically disclosed are variants of these and other proteins herein disclosed which have at least, 70% or 75% or 80% or 85% or 90% or 95% homology to the stated sequence. Those of skill in the art readily understand how to determine the homology of two proteins. For example, the homology can be calculated after aligning the two sequences so that the homology is at its highest level.
[0127] Another way of calculating homology can be performed by published algorithms. Optimal alignment of sequences for comparison may be conducted by the local homology algorithm of Smith and Waterman Adv. Appl. Math. 2: 482 (1981), by the homology alignment algorithm of Needleman and Wunsch, J. MoL Biol. 48: 443 (1970), by the search for similarity method of Pearson and Lipman, Proc. Natl. Acad. Sci. U.S.A. 85: 2444 (1988), by computerized implementations of these algorithms (GAP, BESTFIT, FASTA, and TFASTA in the Wisconsin Genetics Software Package, Genetics Computer Group, 575 Science Dr., Madison, Wis.), or by inspection.
[0128] The same types of homology can be obtained for nucleic acids by for example the algorithms disclosed in Zuker, M. Science 244:48-52, 1989, Jaeger et al. Proc. Natl. Acad. Sci. USA 86:7706-7710, 1989, Jaeger et al. Methods Enzymol. 183:281-306, 1989 which are herein incorporated by reference for at least material related to nucleic acid alignment.
[0129] It is understood that the description of conservative mutations and homology can be combined together in any combination, such as embodiments that have at least 70% homology to a particular sequence wherein the variants are conservative mutations.
[0130] As this specification discusses various proteins and protein sequences it is understood that the nucleic acids that can encode those protein sequences are also disclosed. This would include all degenerate sequences related to a specific protein sequence, i.e. all nucleic acids having a sequence that encodes one particular protein sequence as well as all nucleic acids, including degenerate nucleic acids, encoding the disclosed variants and derivatives of the protein sequences. Thus, while each particular nucleic acid sequence may not be written out herein, it is understood that each and every sequence is in fact disclosed and described herein through the disclosed protein sequence. For example, one of the many nucleic acid sequences that can encode the DEFB1 protein sequence set forth in SEQ ID NO: 63 is set forth in SEQ ID NO: 64. It is also understood that while no amino acid sequence indicates what particular DNA sequence encodes that protein within an organism, where particular variants of a disclosed protein are disclosed herein, the known nucleic acid sequence that encodes that protein in the particular DEFB1 from which that protein arises is also known and herein disclosed and described.
[0131] It is understood that there are numerous amino acid and peptide analogs which can be incorporated into the disclosed compositions. For example, there are numerous D amino acids or amino acids which have a different functional substituent then the amino acids shown in Table 1 and Table 2. The opposite stereo isomers of naturally occurring peptides are disclosed, as well as the stereo isomers of peptide analogs. These amino acids can readily be incorporated into polypeptide chains by charging tRNA molecules with the amino acid of choice and engineering genetic constructs that utilize, for example, amber codons, to insert the analog amino acid into a peptide chain in a site specific way (Thorson et al., Methods in Molec. Biol. 77:43-73 (1991), Zoller, Current Opinion in Biotechnology, 3:348-354 (1992); Ibba, Biotechnology & Genetic Engineering Reviews 13:197-216 (1995), Cahill et al., TIBS, 14(10):400-403 (1989); Benner, TIB Tech, 12:158-163 (1994); Ibba and Hennecke, Bio/technology, 12:678-682 (1994) all of which are herein incorporated by reference at least for material related to amino acid analogs).
[0132] Molecules can be produced that resemble peptides, but which are not connected via a natural peptide linkage. For example, linkages for amino acids or amino acid analogs can include CH2NH--, --CH2S--, --CH2--CH2--, --CH═CH-- (cis and trans), --COCH2--, --CH(OH)CH2--, and --CHH2SO-- (These and others can be found in Spatola, A. F. in Chemistry and Biochemistry of Amino Acids, Peptides, and Proteins, B. Weinstein, eds Marcel Dekker, New York, p. 267 (1983); Spatola, A. F., Vega Data (March 1983), Vol. 1, Issue 3, Peptide Backbone Modifications (general review); Morley, Trends Pharm Sci (1980) pp. 463-468; Hudson, D. et al., Int J Pept Prot Res 14:177-185 (1979) (--CH2NH--, CH2CH2--); Spatola et al. Life Sci 38:1243-1249 (1986) (--CHH2--S); Hann J. Chem. Soc Perkin Trans. 1307-314 (1982) (--CH--CH--, cis and trans); Almquist et al. J. Med. Chem. 23:1392-1398 (1980) (--COCH2--); Jennings-White et al. Tetrahedron Lett 23:2533 (1982) (--COCH2--); Szelke et al. European Appln, EP 45665 CA (1982): 97:39405 (1982) (--CH(OH)CH2--); Holladay et al. Tetrahedron. Lett 24:4401-4404 (1983) (--C(OH)CH2--); and Hruby Life Sci 31:189-199 (1982) (--CH2--S--); each of which is incorporated herein by reference. A particularly preferred non-peptide linkage is --CH2NH--. It is understood that peptide analogs can have more than one atom between the bond atoms, such as b-alanine, g-aminobutyric acid, and the like.
[0133] Amino acid analogs and analogs and peptide analogs often have enhanced or desirable properties, such as, more economical production, greater chemical stability, enhanced pharmacological properties (half-life, absorption, potency, efficacy, etc.), altered specificity (e.g., a broad-spectrum of biological activities), reduced antigenicity, and others.
[0134] D-amino acids can be used to generate more stable peptides, because D amino acids are not recognized by peptidases and such. Systematic substitution of one or more amino acids of a consensus sequence with a D-amino acid of the same type (e.g., D-lysine in place of L-lysine) can be used to generate more stable peptides. Cysteine residues can be used to cyclize or attach two or more peptides together. This can be beneficial to constrain peptides into particular conformations. (Rizo and Gierasch Ann. Rev. Biochem. 61:387 (1992), incorporated herein by reference).
Pharmaceutical Carriers/Delivery of Pharmaceutical Products
[0135] As described above, the compositions can also be administered in vivo in a pharmaceutically acceptable carrier. By "pharmaceutically acceptable" is meant a material that is not biologically or otherwise undesirable, i.e., the material may be administered to a subject, along with the nucleic acid or vector, without causing any undesirable biological effects or interacting in a deleterious manner with any of the other components of the pharmaceutical composition in which it is contained. The carrier would naturally be selected to minimize any degradation of the active ingredient and to minimize any adverse side effects in the subject, as would be well known to one of skill in the art.
[0136] The compositions may be administered orally, parenterally (e.g., intravenously), by intramuscular injection, by intraperitoneal injection, transdermally, extracorporeally, topically or the like, including topical intranasal administration or administration by inhalant. As used herein, "topical intranasal administration" means delivery of the compositions into the nose and nasal passages through one or both of the nares and can comprise delivery by a spraying mechanism or droplet mechanism, or through aerosolization of the nucleic acid or vector. Administration of the compositions by inhalant can be through the nose or mouth via delivery by a spraying or droplet mechanism. Delivery can also be directly to any area of the respiratory system (e.g., lungs) via intubation. The exact amount of the compositions required will vary from subject to subject, depending on the species, age, weight and general condition of the subject, the severity of the allergic disorder being treated, the particular nucleic acid or vector used, its mode of administration and the like. Thus, it is not possible to specify an exact amount for every composition. However, an appropriate amount can be determined by one of ordinary skill in the art using only routine experimentation given the teachings herein.
[0137] Parenteral administration of the composition, if used, is generally characterized by injection. Injectables can be prepared in conventional forms, either as liquid solutions or suspensions, solid forms suitable for solution of suspension in liquid prior to injection, or as emulsions. A more recently revised approach for parenteral administration involves use of a slow release or sustained release system such that a constant dosage is maintained. See, e.g., U.S. Pat. No. 3,610,795, which is incorporated by reference herein.
[0138] The materials may be in solution, suspension (for example, incorporated into microparticles, liposomes, or cells). These may be targeted to a particular cell type via antibodies, receptors, or receptor ligands. The following references are examples of the use of this technology to target specific proteins to tumor tissue (Senter, et al., Bioconjugate Chem., 2:447-451, (1991); Bagshawe, K. D., Br. J. Cancer, 60:275-281, (1989); Bagshawe, et al., Br. J. Cancer, 58:700-703, (1988); Senter, et al., Bioconjugate Chem., 4:3-9, (1993); Battelli, et al., Cancer Immunol. Immunother., 35:421-425, (1992); Pietersz and McKenzie, Immunolog. Reviews, 129:57-80, (1992); and Roffler, et al., Biochem. Pharmacol, 42:2062-2065, (1991)). Vehicles such as "stealth" and other antibody conjugated liposomes (including lipid mediated drug targeting to colonic carcinoma), receptor mediated targeting of DNA through cell specific ligands, lymphocyte directed tumor targeting, and highly specific therapeutic retroviral targeting of murine glioma cells in vivo. The following references are examples of the use of this technology to target specific proteins to tumor tissue (Hughes et al., Cancer Research, 49:6214-6220, (1989); and Litzinger and Huang, Biochimica et Biophysica Acta, 1104:179-187, (1992)). In general, receptors are involved in pathways of endocytosis, either constitutive or ligand induced. These receptors cluster in clathrin-coated pits, enter the cell via clathrin-coated vesicles, pass through an acidified endosome in which the receptors are sorted, and then either recycle to the cell surface, become stored intracellularly, or are degraded in lysosomes. The internalization pathways serve a variety of functions, such as nutrient uptake, removal of activated proteins, clearance of macromolecules, opportunistic entry of viruses and toxins, dissociation and degradation of ligand, and receptor-level regulation. Many receptors follow more than one intracellular pathway, depending on the cell type, receptor concentration, type of ligand, ligand valency, and ligand concentration. Molecular and cellular mechanisms of receptor-mediated endocytosis has been reviewed (Brown and Greene, DNA and Cell Biology 10:6, 399-409 (1991)).
Pharmaceutically Acceptable Carriers
[0139] The compositions, including DEFB1, DEFB1-encoding nucleic acids, oligonucleotide inhibitors of PAX2 binding, can be used therapeutically in combination with a pharmaceutically acceptable carrier.
[0140] Suitable carriers and their formulations are described in Remington: The Science and Practice of Pharmacy (19th ed.) ed. A. R. Gennaro, Mack Publishing Company, Easton, Pa. 1995. Typically, an appropriate amount of a pharmaceutically-acceptable salt is used in the formulation to render the formulation isotonic. Examples of the pharmaceutically-acceptable carrier include, but are not limited to, saline, Ringer's solution and dextrose solution. The pH of the solution is preferably from about 5 to about 8, and more preferably from about 7 to about 7.5. Further carriers include sustained release preparations such as semipermeable matrices of solid hydrophobic polymers containing the antibody, which matrices are in the form of shaped articles, e.g., films, liposomes or microparticles. It will be apparent to those persons skilled in the art that certain carriers may be more preferable depending upon, for instance, the route of administration and concentration of composition being administered.
[0141] Pharmaceutical carriers are known to those skilled in the art. These most typically would be standard carriers for administration of drugs to humans, including solutions such as sterile water, saline, and buffered solutions at physiological pH. The compositions can be administered intramuscularly or subcutaneously. Other compounds will be administered according to standard procedures used by those skilled in the art.
[0142] Pharmaceutical compositions may include carriers, thickeners, diluents, buffers, preservatives, surface active agents and the like in addition to the molecule of choice. Pharmaceutical compositions may also include one or more active ingredients such as antimicrobial agents, antiinflammatory agents, anesthetics, and the like.
[0143] The pharmaceutical composition may be administered in a number of ways depending on whether local or systemic treatment is desired, and on the area to be treated. Administration may be topically (including ophthalmically, vaginally, rectally, intranasally), orally, by inhalation, or parenterally, for example by intravenous drip, subcutaneous, intraperitoneal or intramuscular injection. The disclosed antibodies can be administered intravenously, intraperitoneally, intramuscularly, subcutaneously, intracavity, or transdermally.
[0144] Preparations for parenteral administration include sterile aqueous or non-aqueous solutions, suspensions, and emulsions. Examples of non-aqueous solvents are propylene glycol, polyethylene glycol, vegetable oils such as olive oil, and injectable organic esters such as ethyl oleate. Aqueous carriers include water, alcoholic/aqueous solutions, emulsions or suspensions, including saline and buffered media. Parenteral vehicles include sodium chloride solution, Ringer's dextrose, dextrose and sodium chloride, lactated Ringer's, or fixed oils. Intravenous vehicles include fluid and nutrient replenishers, electrolyte replenishers (such as those based on Ringer's dextrose), and the like. Preservatives and other additives may also be present such as, for example, antimicrobials, anti-oxidants, chelating agents, and inert gases and the like.
[0145] Formulations for topical administration may include ointments, lotions, creams, gels, drops, suppositories, sprays, liquids and powders. Conventional pharmaceutical carriers, aqueous, powder or oily bases, thickeners and the like may be necessary or desirable.
[0146] Compositions for oral administration include powders or granules, suspensions or solutions in water or non-aqueous media, capsules, sachets, or tablets. Thickeners, flavorings, diluents, emulsifiers, dispersing aids or binders may be desirable.
[0147] Some of the compositions may potentially be administered as a pharmaceutically acceptable acid- or base-addition salt, formed by reaction with inorganic acids such as hydrochloric acid, hydrobromic acid, perchloric acid, nitric acid, thiocyanic acid, sulfuric acid, and phosphoric acid, and organic acids such as formic acid, acetic acid, propionic acid, glycolic acid, lactic acid, pyruvic acid, oxalic acid, malonic acid, succinic acid, maleic acid, and fumaric acid, or by reaction with an inorganic base such as sodium hydroxide, ammonium hydroxide, potassium hydroxide, and organic bases such as mono-, di-, trialkyl and aryl amines and substituted ethanolamines.
Therapeutic Uses
[0148] Effective dosages and schedules for administering the compositions may be determined empirically, and making such determinations is within the skill in the art. The dosage ranges for the administration of the compositions are those large enough to produce the desired effect in which the symptoms disorder are effected. The dosage should not be so large as to cause adverse side effects, such as unwanted cross-reactions, anaphylactic reactions, and the like. Generally, the dosage will vary with the age, condition, sex and extent of the disease in the patient, route of administration, or whether other drugs are included in the regimen, and can be determined by one of skill in the art. The dosage can be adjusted by the individual physician in the event of any counterindications. Dosage can vary, and can be administered in one or more dose administrations daily, for one or several days. Guidance can be found in the literature for appropriate dosages for given classes of pharmaceutical products. For example, guidance in selecting appropriate doses for antibodies can be found in the literature on therapeutic uses of antibodies, e.g., Handbook of Monoclonal Antibodies, Ferrone et al., eds., Noges Publications, Park Ridge, N.J., (1985) ch. 22 and pp. 303-357; Smith et al., Antibodies in Human Diagnosis and Therapy, Haber et al., eds., Raven Press, New York (1977) pp. 365-389. A typical daily dosage of the oligonucleotide used alone might range from about 1 μg/kg to up to 100 mg/kg of body weight or more per day, depending on the factors mentioned above. In a more specific example, 5 to 7 mg/kg/day can be used. This is appropriate for i.v. administration and higher dosages, up to about 150 mg/m2/day s.c. are appropriate. This can be administered daily for a week in from 1 to 24 courses. See for example A Phase I Pharmacokinetic and Biological Correlative Study of Oblimersen Sodium (Genasense, G3139), an Antisense Oligonucleotide to the Bcl-2 mRNA, and of Docetaxel in Patients with Hormone-Refractory Prostate Cancer, Clinical Cancer Research, Vol. 10, 5048-5057, Aug. 1, 2004, incorporated herein for it's teaching of dosages for oligonucleotides.
[0149] Following administration of a disclosed composition for treating cancer, the efficacy of the therapeutic composition can be assessed in various ways well known to the skilled practitioner. For instance, one of ordinary skill in the art will understand that a composition, DEFB1, DEFB1-encoding nucleic acid, inhibitor of PAX2, disclosed herein is efficacious in treating cancer in a subject by observing that the composition reduces tumor load or prevents a further increase in tumor load. Methods of assessing tumor load are known in the art
[0150] The compositions that inhibit the interactions between PAX2 and the DEFB1 promoter can be administered prophylactically to patients or subjects who are at risk for cancer.
[0151] Other molecules that interact with PAX2 to inhibit its interaction with the DEFB1 promoter can be delivered in ways similar to those described for the pharmaceutical products.
[0152] The disclosed compositions and methods can also be used for example as tools to isolate and test new drug candidates for a variety of related diseases. Thus, a method of identifying inhibitors of the binding of PAX2 to the DEFB1 promoter is provided. The method can comprise contacting a system that expresses DEFB1 with a putative inhibitor in the presence and/or absence of PAX2 to determine whether there is an inhibitory effect on this interaction.
Compositions Identified by Screening with Disclosed Compositions/Combinatorial Chemistry
[0153] Combinatorial Chemistry
[0154] The disclosed compositions can be used as targets for any combinatorial technique to identify molecules or macromolecular molecules that interact with the disclosed compositions in a desired way. Also disclosed are the compositions that are identified through combinatorial techniques or screening techniques in which the compositions disclosed as the PAX2 sequence or portions thereof (e.g., PAX2 DNA-binding domain), are used as the target in a combinatorial or screening protocol.
[0155] It is understood that when using the disclosed compositions in combinatorial techniques or screening methods, molecules, such as macromolecular molecules, will be identified that have particular desired properties such as inhibition or stimulation or the target molecule's function. The molecules identified and isolated when using the disclosed compositions, such as, DEFB1 or PAX2, are also disclosed. Thus, the products produced using the combinatorial or screening approaches that involve the disclosed compositions are also considered herein disclosed.
[0156] It is understood that the disclosed methods for identifying molecules that inhibit the interactions between, for example, DEFB1 promoter and PAX2 can be performed using high through put means. For example, putative inhibitors can be identified using Fluorescence Resonance Energy Transfer (FRET) to quickly identify interactions. The underlying theory of the techniques is that when two molecules are close in space, ie, interacting at a level beyond background, a signal is produced or a signal can be quenched. Then, a variety of experiments can be performed, including, for example, adding in a putative inhibitor. If the inhibitor competes with the interaction between the two signaling molecules, the signals will be removed from each other in space, and this will cause a decrease or an increase in the signal, depending on the type of signal used. This decrease or increasing signal can be correlated to the presence or absence of the putative inhibitor. Any signaling means can be used. For example, disclosed are methods of identifying an inhibitor of the interaction between any two of the disclosed molecules comprising, contacting a first molecule and a second molecule together in the presence of a putative inhibitor, wherein the first molecule or second molecule comprises a fluorescence donor, wherein the first or second molecule, typically the molecule not comprising the donor, comprises a fluorescence acceptor; and measuring Fluorescence Resonance Energy Transfer (FRET), in the presence of the putative inhibitor and the in absence of the putative inhibitor, wherein a decrease in FRET in the presence of the putative inhibitor as compared to FRET measurement in its absence indicates the putative inhibitor inhibits binding between the two molecules. This type of method can be performed with a cell system as well.
[0157] Combinatorial chemistry includes but is not limited to all methods for isolating small molecules or macromolecules that are capable of binding either a small molecule or another macromolecule, typically in an iterative process. Proteins, oligonucleotides, and sugars are examples of macromolecules. For example, oligonucleotide molecules with a given function, catalytic or ligand-binding, can be isolated from a complex mixture of random oligonucleotides in what has been referred to as "in vitro genetics" (Szostak, TIBS 19:89, 1992). One synthesizes a large pool of molecules bearing random and defined sequences and subjects that complex mixture, for example, approximately 1015 individual sequences in 100 μg of a 100 nucleotide RNA, to some selection and enrichment process. Through repeated cycles of affinity chromatography and PCR amplification of the molecules bound to the ligand on the column, Ellington and Szostak (1990) estimated that 1 in 1010 RNA molecules folded in such a way as to bind a small molecule dyes. DNA molecules with such ligand-binding behavior have been isolated as well (Ellington and Szostak, 1992; Bock et al, 1992). Techniques aimed at similar goals exist for small organic molecules, proteins, antibodies and other macromolecules known to those of skill in the art. Screening sets of molecules for a desired activity whether based on small organic libraries, oligonucleotides, or antibodies is broadly referred to as combinatorial chemistry. Combinatorial techniques are particularly suited for defining binding interactions between molecules and for isolating molecules that have a specific binding activity, often called aptamers when the macromolecules are nucleic acids.
[0158] There are a number of methods for isolating proteins which either have de novo activity or a modified activity. For example, phage display libraries have been used to isolate numerous peptides that interact with a specific target. (See for example, U.S. Pat. Nos. 6,031,071; 5,824,520; 5,596,079; and 5,565,332 which are herein incorporated by reference at least for their material related to phage display and methods relate to combinatorial chemistry)
[0159] A preferred method for isolating proteins that have a given function is described by Roberts and Szostak (Roberts R. W. and Szostak J. W. Proc. Natl. Acad. Sci. USA, 94(23)12997-302 (1997). This combinatorial chemistry method couples the functional power of proteins and the genetic power of nucleic acids. An RNA molecule is generated in which a puromycin molecule is covalently attached to the 3'-end of the RNA molecule. An in vitro translation of this modified RNA molecule causes the correct protein, encoded by the RNA to be translated. In addition, because of the attachment of the puromycin, a peptidyl acceptor which cannot be extended, the growing peptide chain is attached to the puromycin which is attached to the RNA. Thus, the protein molecule is attached to the genetic material that encodes it. Normal in vitro selection procedures can now be done to isolate functional peptides. Once the selection procedure for peptide function is complete traditional nucleic acid manipulation procedures are performed to amplify the nucleic acid that codes for the selected functional peptides. After amplification of the genetic material, new RNA is transcribed with puromycin at the 3'-end, new peptide is translated and another functional round of selection is performed. Thus, protein selection can be performed in an iterative manner just like nucleic acid selection techniques. The peptide which is translated is controlled by the sequence of the RNA attached to the puromycin. This sequence can be anything from a random sequence engineered for optimum translation (i.e. no stop codons etc.) or it can be a degenerate sequence of a known RNA molecule to look for improved or altered function of a known peptide. The conditions for nucleic acid amplification and in vitro translation are well known to those of ordinary skill in the art and are preferably performed as in Roberts and Szostak (Roberts R. W. and Szostak J. W. Proc. Natl. Acad. Sci. USA, 94(23)12997-302 (1997)).
[0160] Another preferred method for combinatorial methods designed to isolate peptides is described in Cohen et al. (Cohen B. A., et al., Proc. Natl. Acad. Sci. USA 95(24):14272-7 (1998)). This method utilizes and modifies two-hybrid technology. Yeast two-hybrid systems are useful for the detection and analysis of protein:protein interactions. The two-hybrid system, initially described in the yeast Saccharomyces cerevisiae, is a powerful molecular genetic technique for identifying new regulatory molecules, specific to the protein of interest (Fields and Song, Nature 340:245-6 (1989)). Cohen et al., modified this technology so that novel interactions between synthetic or engineered peptide sequences could be identified which bind a molecule of choice. The benefit of this type of technology is that the selection is done in an intracellular environment. The method utilizes a library of peptide molecules that attached to an acidic activation domain.
[0161] Using methodology well known to those of skill in the art, in combination with various combinatorial libraries, one can isolate and characterize those small molecules or macromolecules, which bind to or interact with the desired target. The relative binding affinity of these compounds can be compared and optimum compounds identified using competitive binding studies, which are well known to those of skill in the art.
[0162] Techniques for making combinatorial libraries and screening combinatorial libraries to isolate molecules which bind a desired target are well known to those of skill in the art. Representative techniques and methods can be found in but are not limited to U.S. Pat. Nos. 5,084,824, 5,288,514, 5,449,754, 5,506,337, 5,539,083, 5,545,568, 5,556,762, 5,565,324, 5,565,332, 5,573,905, 5,618,825, 5,619,680, 5,627,210, 5,646,285, 5,663,046, 5,670,326, 5,677,195, 5,683,899, 5,688,696, 5,688,997, 5,698,685, 5,712,146, 5,721,099, 5,723,598, 5,741,713, 5,792,431, 5,807,683, 5,807,754, 5,821,130, 5,831,014, 5,834,195, 5,834,318, 5,834,588, 5,840,500, 5,847,150, 5,856,107, 5,856,496, 5,859,190, 5,864,010, 5,874,443, 5,877,214, 5,880,972, 5,886,126, 5,886,127, 5,891,737, 5,916,899, 5,919,955, 5,925,527, 5,939,268, 5,942,387, 5,945,070, 5,948,696, 5,958,702, 5,958,792, 5,962,337, 5,965,719, 5,972,719, 5,976,894, 5,980,704, 5,985,356, 5,999,086, 6,001,579, 6,004,617, 6,008,321, 6,017,768, 6,025,371, 6,030,917, 6,040,193, 6,045,671, 6,045,755, 6,060,596, and 6,061,636.
[0163] Combinatorial libraries can be made from a wide array of molecules using a number of different synthetic techniques. For example, libraries containing fused 2,4-pyrimidinediones (U.S. Pat. No. 6,025,371) dihydrobenzopyrans (U.S. Pat. Nos. 6,017,768 and 5,821,130), amide alcohols (U.S. Pat. No. 5,976,894), hydroxy-amino acid amides (U.S. Pat. No. 5,972,719) carbohydrates (U.S. Pat. No. 5,965,719), 1,4-benzodiazepin-2,5-diones (U.S. Pat. No. 5,962,337), cyclics (U.S. Pat. No. 5,958,792), biaryl amino acid amides (U.S. Pat. No. 5,948,696), thiophenes (U.S. Pat. No. 5,942,387), tricyclic Tetrahydroquinolines (U.S. Pat. No. 5,925,527), benzofurans (U.S. Pat. No. 5,919,955), isoquinolines (U.S. Pat. No. 5,916,899), hydantoin and thiohydantoin (U.S. Pat. No. 5,859,190), indoles (U.S. Pat. No. 5,856,496), imidazol-pyrido-indole and imidazol-pyrido-benzothiophenes (U.S. Pat. No. 5,856,107) substituted 2-methylene-2,3-dihydrothiazoles (U.S. Pat. No. 5,847,150), quinolines (U.S. Pat. No. 5,840,500), PNA (U.S. Pat. No. 5,831,014), containing tags (U.S. Pat. No. 5,721,099), polyketides (U.S. Pat. No. 5,712,146), morpholino-subunits (U.S. Pat. Nos. 5,698,685 and 5,506,337), sulfamides (U.S. Pat. No. 5,618,825), and benzodiazepines (U.S. Pat. No. 5,288,514).
[0164] Screening molecules similar to the disclosed siRNA molecules for inhibition of PAX2 suppression of DEFB1 expression is a method of isolating desired compounds.
[0165] Molecules isolated which can either be competitive inhibitors or non-competitive inhibitors.
[0166] In another embodiment the inhibitors are non-competitive inhibitors. One type of non-competitive inhibitor will cause allosteric rearrangements.
[0167] As used herein combinatorial methods and libraries included traditional screening methods and libraries as well as methods and libraries used in iterative processes.
Computer Assisted Drug Design
[0168] The disclosed compositions can be used as targets for any molecular modeling technique to identify either the structure of the disclosed compositions or to identify potential or actual molecules, such as small molecules, which interact in a desired way with the disclosed compositions. The nucleic acids, peptides, and related molecules disclosed herein can be used as targets in any molecular modeling program or approach.
[0169] It is understood that when using the disclosed compositions in modeling techniques, molecules, such as macromolecular molecules, will be identified that have particular desired properties such as inhibition or stimulation of the target molecule's function. The molecules identified and isolated when using the disclosed compositions, such as, SEQ ID NO:1, are also disclosed. Thus, the products produced using the molecular modeling approaches that involve the disclosed compositions, such as, SEQ ID NO:1, are also considered herein disclosed.
[0170] Thus, one way to isolate molecules that bind a molecule of choice is through rational design. This is achieved through structural information and computer modeling. Computer modeling technology allows visualization of the three-dimensional atomic structure of a selected molecule and the rational design of new compounds that will interact with the molecule. The three-dimensional construct typically depends on data from x-ray crystallographic analyses or NMR imaging of the selected molecule. The molecular dynamics require force field data. The computer graphics systems enable prediction of how a new compound will link to the target molecule and allow experimental manipulation of the structures of the compound and target molecule to perfect binding specificity. Prediction of what the molecule-compound interaction will be when small changes are made in one or both requires molecular mechanics software and computationally intensive computers, usually coupled with user-friendly, menu-driven interfaces between the molecular design program and the user.
[0171] Examples of molecular modeling systems are the CHARMm and QUANTA programs, Polygen Corporation, Waltham, Mass. CHARMm performs the energy minimization and molecular dynamics functions. QUANTA performs the construction, graphic modeling and analysis of molecular structure. QUANTA allows interactive construction, modification, visualization, and analysis of the behavior of molecules with each other.
[0172] A number of articles review computer modeling of drugs interactive with specific proteins, such as Rotivinen, et al., 1988 Acta Pharmaceutica Fennica 97, 159-166; Ripka, New Scientist 54-57 (Jun. 16, 1988); McKinaly and Rossmann, 1989 Annu. Rev. Pharmacol. Toxiciol. 29, 111-122; Perry and Davies, QSAR: Quantitative Structure-Activity Relationships in Drug Design pp. 189-193 (Alan R. Liss, Inc. 1989); Lewis and Dean, 1989 Proc. R. Soc. Lond. 236, 125-140 and 141-162; and, with respect to a model enzyme for nucleic acid components, Askew, et al., 1989 J. Am. Chem. Soc. 111, 1082-1090. Other computer programs that screen and graphically depict chemicals are available from companies such as BioDesign, Inc., Pasadena, Calif., Allelix, Inc, Mississauga, Ontario, Canada, and Hypercube, Inc., Cambridge, Ontario. Although these are primarily designed for application to drugs specific to particular proteins, they can be adapted to design of molecules specifically interacting with specific regions of DNA or RNA, once that region is identified.
[0173] Although described above with reference to design and generation of compounds which could alter binding, one could also screen libraries of known compounds, including natural products or synthetic chemicals, and biologically active materials, including proteins, for compounds which alter substrate binding or enzymatic activity.
Computer Readable Mediums
[0174] It is understood that the disclosed nucleic acids and proteins can be represented as a sequence consisting of the nucleotides of amino acids. There are a variety of ways to display these sequences, for example the nucleotide guanosine can be represented by G or g. Likewise the amino acid valine can be represented by Val or V. Those of skill in the art understand how to display and express any nucleic acid or protein sequence in any of the variety of ways that exist, each of which is considered herein disclosed. Specifically contemplated herein is the display of these sequences on computer readable mediums, such as, commercially available floppy disks, tapes, chips, hard drives, compact disks, and video disks, or other computer readable mediums. Also disclosed are the binary code representations of the disclosed sequences. Those of skill in the art understand what computer readable mediums. Thus, computer readable mediums on which the nucleic acids or protein sequences are recorded, stored, or saved.
[0175] Disclosed are computer readable mediums comprising the sequences and information regarding the sequences set forth herein. Also disclosed are computer readable mediums comprising the sequences and information regarding the sequences set forth.
[0176] The following examples are put forth so as to provide those of ordinary skill in the art with a complete disclosure and description of how the compounds, compositions, articles, devices and/or methods claimed herein are made and evaluated, and are intended to be purely exemplary and are not intended to limit the disclosure. Efforts have been made to ensure accuracy with respect to numbers (e.g., amounts, temperature, etc.), but some errors and deviations should be accounted for. Unless indicated otherwise, parts are parts by weight, temperature is in ° C. or is at ambient temperature, and pressure is at or near atmospheric.
[0177] The following examples are put forth so as to provide those of ordinary skill in the art with a complete disclosure and description of how the compounds, compositions, articles, devices and/or methods claimed herein are made and evaluated, and are intended to be purely exemplary of the invention and are not intended to limit the scope of what the inventors regard as their invention. Efforts have been made to ensure accuracy with respect to numbers (e.g., amounts, temperature, etc.), but some errors and deviations should be accounted for. Unless indicated otherwise, parts are parts by weight, temperature is in C or is at ambient temperature, and pressure is at or near atmospheric.
Example I
Human Beta Defensin-1 is Cytotoxic to Late-Stage Prostate Cancer and Plays a Role in Prostate Cancer Tumor Immunity
[0178] Abstract
[0179] DEFB1 was cloned into an inducible expression system to examine what effect it had on normal prostate epithelial cells, as well as androgen receptor positive (AR.sup.+) and androgen receptor negative (AR.sup.-) prostate cancer cell lines. Induction of DEFB1 expression resulted in a decrease in cellular growth in AR.sup.- cells DU145 and PC3, but had no effect on the growth of the AR.sup.+ prostate cancer cells LNCaP. DEFB1 also caused rapid induction of caspase-mediated apoptosis. Data presented here are the first to provide evidence of its role in innate tumor immunity and indicate that its loss contributes to tumor progression in prostate cancer.
Materials and Methods
[0180] Cell Lines
[0181] The cell lines DU145 were cultured in DMEM medium, PC3 were grown in F12 medium, and LNCaP were grown in RPMI medium (Life Technologies, Inc., Grand Island, N.Y.). Growth media for all three lines was supplemented with 10% (v/v) fetal bovine serum (Life Technologies). The hPrEC cells were cultured in prostate epithelium basal media (Cambrex Bio Science, Inc., Walkersville, Md.). All cell lines were maintained at 37° C. and 5% CO2.
[0182] Tissue Samples and Laser Capture Microdissection
[0183] Prostate tissues obtained from consented patients that underwent radical prostatectomy were acquired through the Hollings Cancer Center tumor bank in accordance with an Institutional Review Board-approved protocol. This included guidelines for the processing, sectioning, histological characterization, RNA purification and PCR amplification of samples. Following pathologic examination of frozen tissue sections, laser capture microdissection (LCM) was performed to ensure that the tissue samples assayed consisted of pure populations of benign prostate cells. For each tissue section analyzed, LCM was performed at three different regions containing benign tissue and the cells collected were then pooled.
[0184] Cloning of DEFB1 Gene
[0185] DEFB1 cDNA was generated from RNA by reverse transcription-PCR. The PCR primers were designed to contain ClaI and KpnI restriction sites. DEFB1 PCR products were restriction digested with ClaI and KpnI and ligated into a TA cloning vector. The TA/DEFB1 vector was then transfected into E. coli by heat shock and individual clones were selected and expanded. Plasmids were isolated by Cell Culture DNA Midiprep (Qiagen, Valencia, Calif.) and sequence integrity verified by automated sequencing. The DEFB1 gene fragment was then ligated into the pTRE2 digested with ClaI and KpnI, which served as an intermediate vector for orientation purposes. Then the pTRE2/DEFB1 construct was digested with ApaI and KpnI to excise the DEFB1 insert, which was ligated into pIND vector of the Ecdysone Inducible Expression System (Invitrogen, Carlsbad, Calif.) also double digested with ApaI and KpnI. The construct was again transfected into E. coli and individual clones were selected and expanded. Plasmids were isolated and sequence integrity of pIND/DEFB1 was again verified by automated sequencing.
[0186] Transfection
[0187] Cells (1×106) were seeded onto 100-mm Petri dishes and grown overnight. Then the cells were co-transfected using Lipofectamine 2000 (Invitrogen, Carlsbad, Calif.) with 1 μg of pVgRXR plasmid, which expresses the heterodimeric ecdysone receptor, and 1 μg of the pIND/DEFB1 vector construct or empty pIND control vector in Opti-MEM media (Life Technologies, Inc., Grand Island, N.Y.).
[0188] RNA Isolation and Quantitative RT-PCR
[0189] In order to verify DEFB1 protein expression in the cells transfected with DEFB1 construct, RNA was collected after a 24 hour induction period with Ponasterone A (Pon A). Briefly, total RNA was isolated using the SV Total RNA Isolation System (Promega, Madison, Wis.) from approximately 1×106 cells harvested by trypsinizing. Here, cells were lysed and total RNA was isolated by centrifugation through spin columns. For cells collected by LCM, total RNA was isolated using the PicoPure RNA Isolation Kit (Arcturus Biosciences, Mt. View, Calif.) following the manufacturer's protocol. Total RNA (0.5 μg per reaction) from both sources was reverse transcribed into cDNA utilizing random primers (Promega). AMV Reverse Transcriptase II enzyme (500 units per reaction; Promega) was used for first strand synthesis and Tfl DNA Polymerase for second strand synthesis (500 units per reaction; Promega) as per the manufacturer's protocol. In each case, 50 pg of cDNA was used per ensuing PCR reaction. Two-step QRT-PCR was performed on cDNA generated using the MultiScribe Reverse Transcripatase from the TaqMan Reverse Transcription System and the SYBR Green PCR Master Mix (Applied Biosystems).
[0190] The primer pair for DEFB1 (Table 3) was generated from the published DEFB1 sequence (GenBank Accession No. U50930)10. Forty cycles of PCR were performed under standard conditions using an annealing temperature of 56° C. In addition, β-actin (Table 3) was amplified as a housekeeping gene to normalize the initial content of total cDNA. DEFB1 expression was calculated as the relative expression ratio between DEFB1 and β-actin and was compared in cells lines induced and uninduced for DEFB1 expression, as well as LCM benign prostatic tissue. As a negative control, QRT-PCR reactions without cDNA template were also performed. All reactions were run three times in triplicate.
[0191] MTT Cell Viability Assay
[0192] To examine the effects of DEFB1 on cell growth, metabolic 3-[4,5-dimethylthiazol-2yl]-2,5 diphenyl tetrazolium bromide (MTT) assays were performed. PC3, DU145 and LNCaP cells co-transfected with pVgRXR plasmid and pIND/DEFB1 construct or empty pIND vector were seeded onto a 96-well plate at 1-5×103 cells per well. Twenty-four hours after seeding, fresh growth medium was added containing 10 μM Ponasterone A daily to induce DEFB1 expression for 24-, 48- and 72 hours after which the MTT assay was performed according to the manufacturer's instructions (Promega). Reactions were performed three times in triplicate.
[0193] Flow Cytometry
[0194] PC3 and DU145 cells co-transfected with the DEFB1 expression system were grown in 60-mm dishes and induced for 12, 24, and 48 hours with 10 μM Ponasterone A. Following each incubation period, the medium was collected from the plates (to retain any detached cells) and combined with PBS used to wash the plates. The remaining attached cells were harvested by trypsinization and combined with the detached cells and PBS. The cells were then pelleted at 4° C. (500×g) for 5 min, washed twice in PBS, and resuspended in 100 ul of 1× Annexin binding buffer (0.1 M Hepes/NaOH at pH 7.4, 1.4 M NaCl, 25 mM CaCl2) containing 5 μl of Annexin V-FITC and 5 μl of PI. The cells were incubated at RT for 15 min in the dark, then diluted with 400 μl of 1× Annexin binding buffer and analyzed by FACscan (Becton Dickinson, San Jose, Calif.). All reactions were performed three times.
[0195] Microscopic Analysis
[0196] Cell morphology was analyzed by phase contrast microscopy. DU145, PC3 and LNCaP cells containing no vector, empty plasmid or DEFB1 plasmid were seeded onto 6 well culture plates (BD Falcon, USA). The following day plasmid-containing cells were induced for a period of 48 h with media containing 10 μM Ponasterone A, while control cells received fresh media. The cells were then viewed under an inverted Zeiss IM 35 microscope (Carl Zeiss, Germany). Phase contrast pictures of a field of cells were obtained using the SPOT Insight Mosaic 4.2 camera (Diagnostic Instruments, USA). Cells were examined by phase contrast microscopy under 32× magnification and digital images were stored as uncompressed TIFF files and exported into Photoshop CS software (Adobe Systems, San Jose, Calif.) for image processing and hard copy presentation.
[0197] Caspase Detection
[0198] Detection of caspase activity in the prostate cancer cell lines was performed using APO LOGIX® Carboxyfluorescin Caspase detection kit (Cell Technology, Mountain View, Calif.). Active caspases were detected through the use of a FAM-VAD-FMK inhibitor that irreversibly binds to active caspases. Briefly, DU145 and PC3 cells (1.5-3×105) containing the DEFB1 expression system were plated in 35 mm glass bottom microwell dishes (Matek, Ashland, Mass.) and treated for 24 hours with media only or with media containing PonA as previously described. Next, 10 μl of a 30× working dilution of carboxyfluorescein labeled peptide fluoromethyl ketone (FAM-VAD-FMK) was added to 300 μl of media and added to each 35 mm dish. Cells were then incubated for 1 hour at 37° C. under 5% CO2. Then, the medium was aspirated and the cells were washed twice with 2 ml of a 1× Working dilution Wash Buffer. Cells were viewed under differential interference contrast (DIC) or under laser excitation at 488 nm. The fluorescent signal was analyzed using a confocal microscope (Zeiss LSM 5 Pascal) and a 63×DIC oil lens with a Vario 2 RGB Laser Scanning Module.
[0199] Statistical Analysis
[0200] Statistical differences were evaluated using the Student's t-test for unpaired values. P values were determined by a two-sided calculation, and a P value of less than 0.05 was considered statistically significant.
Results
[0201] DEFB1 Expression in Prostate Tissue and Cell Lines
[0202] DEFB1 expression levels were measured by QRT-PCR in benign and malignant prostatic tissue, hPrEC prostate epithelial cells and DU145, PC3 and LNCaP prostate cancer cells. DEFB1 expression was detected in all of the benign clinical samples. The average amount of DEFB1 relative expression was 0.0073. In addition, DEFB1 relative expression in hPrEC cells was 0.0089. There was no statistical difference in DEFB1 expression detected in the benign prostatic tissue samples and hPrEC (FIG. 1A). Analysis of the relative DEFB1 expression levels in the prostate cancer cell lines revealed significantly lower levels in DU145, PC3 and LNCaP. As a further point of reference, relative DEFB1 expression was measured in the adjacent malignant section of prostatic tissue from patient #1215. There were no significant differences in the level of DEFB1 expression observed in the three prostate cancer lines compared to malignant prostatic tissue from patient #1215 (FIG. 1B). In addition, expression levels in all four samples were close to the no template negative controls which confirmed little to no endogenous DEFB1 expression (data not shown). QRT-PCR was also performed on the prostate cancer cell lines transfected with the DEFB1 expression system. Following a 24 hour induction period, relative expression levels were 0.01360 in DU145, 0.01503 in PC3 and 0.138 in LNCaP. Amplification products were verified by gel electrophoresis.
[0203] QRT-PCR was performed on LCM tissues regions containing benign, PIN and cancer. DEFB1 relative expression was 0.0146 in the benign region compared to 0.0009 in the malignant region (FIG. 1C.). This represents a 94% decrease which again demonstrates a significant down-regulation of expression. Furthermore, analysis of PIN revealed that DEFB1 expression level was 0.044 which was a 70% decrease. Comparing expression in patient #1457 to the average expression level found in benign regions of six other patients (FIG. 1A.) revealed a ratio of 1.997 representing almost twice as much expression (FIG. 1D.). However, the expression ratio was 0.0595 in PIN and was 0.125 in malignant tissue compared to average expression levels in benign tissue.
[0204] DEFB1 Causes Cell Membrane Permeability and Ruffling
[0205] Induction of DEFB1 in the prostate cancer cell lines resulted in a significant reduction in cell number in DU145 and PC3, but had no effect on cell proliferation in LNCaP (FIG. 2). As a negative control, cell proliferation was monitored in all three lines containing empty plasmid. There were no observable changes in cell morphology in DU145, PC3 or LNCaP cells following the addition of PonA. In addition, DEFB1 induction resulted in morphological changes in both DU145 and PC3. Here cells appeared more rounded and exhibited membrane ruffling indicative of cell death. Apoptotic bodies were also present in both lines.
[0206] Expression of DEFB1 Results in Decreased Cell Viability
[0207] The MTT assay showed a reduction in cell viability by DEFB1 in PC3 and DU145 cells, but no significant effect on LNCaP cells (FIG. 3). After 24 hours, relative cell viability was 72% in DU145 and 56% in PC3. Analysis 48 hours after induction revealed 49% cell viability in DU145 and 37% cell viability in PC3. After 72 hours of DEFB1 expression resulted in 44% and 29% relative cell viability in DU145 and PC3 cells, respectively.
[0208] DEFB1 Causes Rapid Caspase-Mediated Apoptosis in Late-Stage Prostate Cancer Cells
[0209] In order to determine whether the effects of DEFB1 on PC3 and DU145 were cytostatic or cytotoxic, FACS analysis was performed. Under normal growth conditions, more than 90% of PC3 and DU145 cultures were viable and non-apoptotic (lower left quadrant) and did not stain with annexin V or PI (FIG. 4). After inducing DEFB1 expression in PC3 cells, the number of apoptotic cells (lower and upper right quadrants) totaled 10% at 12 hours, 20% at 24 hours, and 44% at 48 hours. For DU145 cells, the number of apoptotic cells totaled 12% after 12 hours, 34% at 24 hours, and 59% after 48 hours of induction. There was no increase in apoptosis observed in cells containing empty plasmid following induction with PonA (data not shown).
[0210] Caspase activity was determined by confocal laser microscopic analysis (FIG. 5). DU145 and PC3 cell were induced for DEFB1 expression and activity was monitored based on the binding of green fluorescing FAM-VAD-FMK to caspases in cells actively undergoing apoptosis. Analysis of cells under DIC showed the presence of viable control DU145 (A), PC3 (E) and LNCaP (I) cells at 0 hours. Excitation by the confocal laser at 488 nm produced no detectable green staining which indicates no caspase activity in DU145 (B), PC3 (F) or LNCaP (J). Following induction for 24 hours, DU145 (C), PC3 (G) and LNCaP (K) cells were again visible under DIC. Confocal analysis under fluorescence revealed green staining in DU145 (D) and PC3 (H) cell indicating caspase activity. However, there was no green staining in LNCaP (L), indicating no induction of apoptosis by DEFB1.
Conclusion
[0211] To assess its functional role, the DEFB1 gene was cloned into the ecdysone inducible expression system and examined its effect on prostate cancer cells. The present data demonstrate DEFB1 cytotoxic activity against late-stage androgen receptor negative hormone refractory prostate cancer cells. In conclusion, this study provides the functional role of DEFB1 in prostate cancer. Furthermore, these findings show that DEFB1 is part of an innate immune system involved in tumor immunity. Data presented here demonstrate that DEFB1 expressed at physiological levels is cytotoxic to AR.sup.- hormone refractory prostate cancer cells, but not to AR+ hormone sensitive prostate cancer cell nor to normal prostate epithelial cells. Given that DEFB1 is constitutively expressed in normal prostate cells without cytotoxicity, it may be that late-stage ARprostate cancer cells possess distinct phenotypic characteristics that render them sensitive to DEFB1 cytotoxicity. Thus, DEFB1 is a viable therapeutic agent for the treatment of late-stage prostate cancer.
Example II
siRNA Mediated Knockdown of PAX2 Expression Results in Prostate Cancer Cell Death Independent of p53 Status
Abstract
[0212] This example examines the effects of inhibiting PAX2 expression by RNA interference in prostate cancer cells which differ in p53 gene status. These results demonstrate that the inhibition of PAX2 results in cell death irrespective of p53 status, indicating that there are additional tumor suppressor genes or cell death pathways inhibited by PAX2 in prostate cancer.
Materials and Methods
[0213] Cell Lines
[0214] The cell lines PC3, DU145 and LNCaP were obtained from the American Type Culture Collection (Rockville, Md., USA). PC3 cell were grown in F-12 media, DU145 in DMEM, and LNCaP in RPMI all supplemented with 10% (v/v) fetal bovine serum. Cell were maintained at 37° C. in 5% CO2.
[0215] siRNA Silencing of PAX2
[0216] In order to achieve efficient gene silencing, a pool of four complementary short interfering ribonucleotides (siRNAs) targeting human PAX2 mRNA (Accession No. NM--003989.1), were synthesized (Dharmacon Research, Lafayette, Colo., USA). A second pool of four siRNAs were used as an internal control to test for the specificity of PAX2 siRNAs. Two of the sequences synthesized target the GL2 luciferase mRNA (Accession No. X65324), and two were non-sequence-specific (Table 4). For annealing of siRNAs, 35 M of single strands were incubated in annealing buffer (100 mM potassium acetate, 30 mM HEPES-KOH at pH 7.4, 2 mM magnesium acetate) for 1 min at 90° C. followed by 1 h incubation at 37° C.
[0217] Western Analysis
[0218] Briefly, cells were harvested by trypsinization and washed twice with PBS. Lysis buffer was prepared according to the manufacturer's instructions (Sigma), and was then added to the cells. Following a 15 minute incubation period at 4° C. on an orbital shaker, cell lysate were then collected and centrifuged for 10 minutes at 12000×g to pellet cellular debris. The protein-containing supernatant were then collected and quantitated. Next, 25 μg protein extract was loaded onto an 8-16% gradient SDS-PAGE (Novex). Following electrophoresis, proteins were transferred to PVDF membranes, and then blocked with 5% nonfat dry milk in TTBS (0.05% Tween 20 and 100 mM Tris-Cl) for 1 hour. Blots were then probed with rabbit anti-Pax2 primary antibody (Zymed, San Francisco, Calif.) at a 1:2000 dilution. After washing, the membranes were incubated with anti-rabbit antibody conjugated to horseradish peroxidase (HRP) (dilution 1:5000; Sigma), and signal detection was visualized using chemiluminescence reagents (Pierce) on an Alpha Innotech Fluorchem 8900. As a control, blots were stripped and reprobed with mouse anti-β-actin primary antibody (1:5000; Sigma-Aldrich) and HRP-conjugated anti-mouse secondary antibody (1:5000; Sigma-Aldrich) and signal detection was again visualized.
[0219] Phase Contrast Microscopy
[0220] The effect of PAX2 knock-down on cell growth was analyzed by phase contrast microscopy. Here, 1-2×104 cells were seeded onto 6 well culture plates (BD Falcon, USA).
[0221] The following day cells were treated with media only, negative control non-specific siRNA or PAX2 siRNA and allowed to incubate for six days. The cells were then viewed under an inverted Zeiss IM 35 microscope (Carl Zeiss, Germany) at 32× magnification. Phase contrast pictures of a field of cells were obtained using the SPOT Insight Mosaic 4.2 camera (Diagnostic Instruments, USA).
[0222] MTT Cytotoxicity Assay
[0223] DU145, PC3 and LNCaP cells (1×105) were transfected with 0.5 μg of the PAX2 siRNA pool or control siRNA pool using Codebreaker transfection reagent according to the manufacturer's protocol (Promega). Next, cell suspensions were diluted and seeded onto a 96-well plate at 1-5×103 cells per well and allowed to grow for 2-, 4- or 6 days. After culture, cell viability was determined by measuring the conversion of 3-[4,5-dimethylthiazol-2yl]-2,5 diphenyl tetrazolium bromide, MTT (Promega), to a colored formazan product. Absorbance was read at 540 nm on a scanning multiwell spectrophotometer.
[0224] Pan-Caspase Detection
[0225] Detection of caspase activity in the prostate cancer cell lines was performed using APO LOGIX® Carboxyfluorescin Caspase detection kit (Cell Technology, Mountain View, Calif.). Active caspases were detected through the use of a FAM-VAD-FMK inhibitor that irreversibly binds to active caspases. Briefly, cells (1-2×104) onto 35 mm glass bottom microwell dishes (Matek, Ashland, Mass.) and treated with media only or PAX2 siRNA as previously described. Next, 10 μl of a 30× working dilution of carboxyfluorescein labeled peptide fluoromethyl ketone (FAM-VAD-FMK) was added to 300 μl of media and added to each 35 mm dish. Cells were then incubated for 1 hour at 37° C. under 5% CO2. Then, the medium was aspirated and the cells were washed twice with 2 ml of a 1× Working dilution Wash Buffer. Cells were viewed under differential interference contrast (DIC) or under laser excitation at 488 nm. The fluorescent signal was analyzed using a confocal microscope (Zeiss LSM 5 Pascal) and a 63×DIC oil lens with a Vario 2 RGB Laser Scanning Module.
[0226] Quantitative Real-Time RT-PCR
[0227] Quantitative real-time RT-PCR was performed in order to verify gene expression after PAX2 siRNA treatment in PC3, DU145 and LNCaP cell lines. Total RNA was isolated using the SV Total RNA Isolation System (Promega). Briefly, approximately 1×106 cells were harvested by trypsinizing, and rinsed in PBS. Cells were then lysed and total RNA was isolated by centrifugation through spin columns. Total RNA (0.5 μg per reaction) was reverse transcribed into cDNA utilizing Oligo (dT) 15 primer (Promega) and AMV Reverse Transcriptase II enzyme (500 units per reaction; Promega) for first strand synthesis and Tfl DNA Polymerase for second strand synthesis (500 units per reaction; Promega) as per the manufacturers' protocol, with identical control samples treated without RT enzyme. Typically, 50 pg of each cDNA was used per ensuing PCR reaction Two-step QRT-PCR was performed on cDNA generated using the MultiScribe Reverse Transcripatase from the TaqMan Reverse Transcription System and the SYBR Green PCR Master Mix (PE Biosystems). The primer pairs for BAX, BID and BAD were generated from the published sequences (Table 3). Reactions were performed in MicroAmp Optical 96-well Reaction Plate (PE Biosystems). Forty cycles of PCR were performed under standard conditions using an annealing temperature of 60° C. Quantification was determined by the cycle number where exponential amplification began (threshold value) and averaged from the values obtained from the triplicate repeats. There was an inverse relationship between message level and threshold value. In addition, GAPDH was used as a housekeeping gene to normalize the initial content of total cDNA. Gene expression was calculated as the relative expression ratio between the pro-apoptotic genes and GAPDH. All reactions were carried out in triplicate.
Results
[0228] siRNA Inhibition of PAX2 Protein
[0229] In order to confirm that the siRNA effective targeted the PAX2 mRNA, Western Analysis was performed to monitor PAX2 protein expression levels over a six day treatment period. Cells were given a single round of transfection with the pool of PAX2 siRNA. The results confirmed specific targeting of PAX2 mRNA by showing knock-down of PAX2 protein by day four in DU145 (FIG. 6a) and by day six in PC3 (FIG. 6b).
[0230] Knock-Down of PAX2 Inhibit Prostate Cancer Cell Growth
[0231] Cells were analyzed following a six day treatment period with media only, negative control non-specific siRNA or PAX2 siRNA (FIG. 7). DU145 (a), PC3 (d) and LNCaP (g) cells all reached at least 90% confluency in the culture dishes containing media only. Treatment of DU145 (b), PC3 (e) and LNCaP (h) with negative control non-specific siRNA had no effect on cell growth, and cells again reached confluency after six days. However, treatment with PAX2 siRNA resulted in a significant decrease in cell number. DU145 cells were approximately 15% confluent (c) and PC3 cells were only 10% confluent (f). LNCaP cell were 5% confluent following siRNA treatment.
[0232] Cytotoxicity Assays
[0233] Cell viability was measured after two-, four-, and six-day exposure times, and is expressed as a ratio of the 570-630 nm absorbance of treated cells divided by that of the untreated control cells (FIG. 8). Relative cell viability following 2 days of treatment was 77% in LNCaP, 82% in DU145 and 78% in PC3. After four days, relative cell viability was 46% in LNCaP, 53% in DU145 and 63% in PC3. After six days of treatment, relative cell viability decreased to 31% in LNCaP, 37% in PC3, and was 53% in DU145. As negative controls, cell viability was measured in after a six day treatment period with negative control non-specific siRNA or transfection reagent alone. For both conditions, there was no statistically significant change in cell viability compared to normal growth media (data not shown).
[0234] Pan-Caspase Detection
[0235] Caspase activity was detected by confocal laser microscopic analysis. DU145, PC3 and LNCaP cells were treated with PAX2 siRNA and activity was monitored based on the binding of FAM-labeled peptide to caspases in cells actively undergoing apoptosis which will fluoresce green. Analysis of cells with media only under DIC shows the presence of viable DU145 (A), PC3 (E) and LNCaP (I) cells at 0 hours (FIG. 9). Excitation by the confocal laser at 488 nm produced no detectable green staining which indicates no caspase activity in untreated DU145 (B), PC3 (F) or LNCaP (J). Following four days of treatment with PAX2 siRNA, DU145 (C), PC3 (G) and LNCaP (K) cells were again visible under DIC. Under fluorescence, the treated DU145 (D), PC3 (H) and LNCaP (L) cells presented green staining indicating caspase activity.
[0236] Effect of Pax2 Inhibition on Pro-Apoptotic Factors
[0237] DU145, PC3 and LNCaP cells were treated with siRNA against PAX2 for six days and expression of pro-apoptotic genes dependent and independent of p53 transcription regulation were measured to monitor cell death pathways. For BAX, there was a 1.81-fold increase in LNCaP, a 2.73-fold increase in DU145, and a 1.87-fold increase in PC3 (FIG. 10a). Expression levels of BID increased by 1.38-fold in LNCaP and 1.77-fold in DU145 (FIG. 10b). However, BID expression levels decreased by 1.44-fold in PC3 following treatment (FIG. 10c). Analysis of BAD revealed a 2.0-fold increase in expression in LNCaP, a 1.38-fold increase in DU145, and a 1.58-fold increase in PC3.
Conclusion
[0238] Despite significant advances in cancer therapy there is still little progress in the treatment of advanced disease. Successful drug treatment of prostate cancer requires the use of therapeutics with specific effects on target cells while maintaining minimal clinical effects on the host. The goal of cancer therapy is to trigger tumor-selective cell death. Therefore, understanding the mechanisms in such death is critical in determining the efficacy of a specific treatment.
[0239] The dependency of prostate cancer cell survival on PAX2 expression is shown here. In order to distinguish between death observed in the p53-expressing cell line LNCaP, the p53-mutated line DU145, and the p53-null line PC3 downstream events that follow p53 activation as a result of PAX2 knock-down were examined. Caspase activity was detected in all three lines indicative of the initiation of programmed cell death. With this, changes in the expression of pro-apoptotic genes were examined. Here, BAX expression was upregulated in all three cell lines independent of p53 status. The expression of pro-apoptotic factor BAD was increased in all three lines following PAX2 inhibition. Following treatment with PAX2 siRNA, BID expression was increased in LNCaP and DU145, but actually decreased in PC3. This indicates that cell death observed in prostate cancer is influenced by but is not dependent on p53 expression. The initiation of apoptosis in prostate cancer cells through different cell death pathways irrespective of p53 status indicates that PAX2 inhibits other tumor suppressors
Example III
Inhibition of PAX2 Oncogene Results in DEFB1-Mediated Death of Prostate Cancer Cells
[0240] Abstract
[0241] The identification of tumor-specific molecules that serve as targets for the development of new cancer drugs is considered to be a major goal in cancer research. Example I demonstrated that there is a high frequency of DEFB1 expression loss in prostate cancer, and that induction of DEFB1 expression results in rapid apoptosis in androgen receptor negative-stage prostate cancer. These data show that DEFB1 plays a role in prostate tumor suppression. In addition, given that it is a naturally occurring component of the immune system of normal prostate epithelium, DEFB1 is expected to be a viable therapeutic agent with little to no side effects. Example II demonstrated that inhibition of PAX2 expression results in prostate cancer cell death independent of p53. These data indicate that there is an addition pro-apoptotic factor or tumor suppressor that is inhibited by PAX2. In addition, the data show that the oncogenic factor PAX2, which is over-expressed in prostate cancer, is a transcriptional repressor of DEFB1. The purpose of this study is to determine if DEFB1 loss of expression is due to aberrant expression of the PAX2 oncogene, and whether inhibiting PAX2 results in DEFB1-mediated cell death.
[0242] The data show that loss of DEFB1 expression occurs at the transcriptional level. Furthermore, computational analysis of the DEFB1 promoter revealed the presence of a GTTCC (SEQ ID NO: 2) DNA binding site for the PAX2 transcriptional repressor next to the DEFB1 TATA box (FIG. 1). The results presented here show that PAX2 and DEFB1 exhibit several attributes of suitable cancer targets, including a role in the suppression of cell death. Therefore, DEFB1 plays a role in tumor immunity and its expression is modulated through therapeutic down-regulation of the PAX2 oncogene.
Materials and Methods
[0243] RNA Isolation and Quantitative RT-PCR
[0244] In order to verify changes in DEFB1 expression levels RNA was collected after 4 days of PAX2 siRNA treatment. Briefly, total RNA was isolated using the SV Total RNA Isolation System (Promega, Madison, Wis.) from approximately 1×106 cells harvested by trypsinizing. Here, cells were lysed and total RNA was isolated by centrifugation through spin columns. Total RNA (0.5 μg per reaction) from both sources was reverse transcribed into cDNA utilizing random primers (Promega). AMV Reverse Transcriptase II enzyme (500 units per reaction; Promega) was used for first strand synthesis and Tfl DNA Polymerase for second strand synthesis (500 units per reaction; Promega) as per the manufacturer's protocol. In each case, 50 pg of cDNA was used per ensuing PCR reaction. Two-step QRT-PCR was performed on cDNA generated using the MultiScribe Reverse Transcripatase from the TaqMan Reverse Transcription System and the SYBR Green PCR Master Mix (Applied Biosystems).
[0245] The primer pair for DEFB1 was generated from the published DEFB1 sequence (Accession No. U50930). Forty cycles of PCR were performed under standard conditions using an annealing temperature of 56° C. In addition, GAPDH was amplified as a housekeeping gene to normalize the initial content of total cDNA. DEFB1 expression was calculated as the relative expression ratio between DEFB1 and GAPDH and was compared in cells lines before and after siRNA knock-down of PAX2 expression. All reactions were run three times in triplicate.
[0246] Generation of the DEFB1 Reporter Construct
[0247] The pGL3 luciferase reporter plasmid was used to monitor DEFB1 reporter activity. Here, a region 160 bases upstream of the DEFB1 transcription initiation site and included the DEFB1 TATA box. The region also included the GTTCC (SEQ ID NO: 2) sequence which is necessary for PAX2 binding. The PCR primers were designed to contain KpnI and NheI restriction sites. The DEFB1 promoter PCR products were restriction digested Kpn I and NheI and ligated into a similarly restriction digested pGL3 plasmid (FIG. 2). The constructs were transfected into E. coli and individual clones were selected and expanded. Plasmids were isolated and sequence integrity of the DEFB1/pGL3 construct was verified by automated sequencing.
[0248] Luciferase Reporter Assay
[0249] Here, 1 μg of the DEFB1 reporter construct or the control pGL3 plasmid was transfected into 1×106 DU145 cells. Next, 0.5×103 cells were seeded onto each well of a 96-well plate and allowed to grow overnight. Then fresh medium was added containing PAX2 siRNA or media only and the cells were incubated for 48 hours. Luciferase was detected by the BrightGlo kit according to the manufacturer's protocol (Promega) and the plates were read on a Veritas automated 96-well luminometer. Promoter activity was expressed as relative luminescence.
[0250] Analysis of Membrane Permeability
[0251] Acridine orange (AO)/ethidium bromide (EtBr) dual staining was performed to identify changes in cell membrane integrity, as well as apoptotic cells by staining the condensed chromatin. AO stains viable cells as well as early apoptotic cells, whereas EtBr stains late stage apoptotic cells that have lost membrane permeability. Briefly, cells were seeded into 2 chamber culture slides (BD Falcon, USA). Cells transfected with empty pIND plasmid/pvgRXR or pIND DEFB1/pvgRXR were induced for 24 or 48 h with media containing 10 μM Ponasterone A. Control cells were provided fresh media at 24 and 48 h. In order to determine the effect of PAX2 inhibition on membrane integrity, separate culture slides containing DU145, PC3 and LNCaP were treated with PAX2 siRNA and incubated for 4 days. Following this, cells were washed once with PBS and stained with 2 ml of a mixture (1:1) of AO (Sigma, USA) and EtBr (Promega, USA) (5 ug/ml) solution for 5 min. Following staining, the cells were again washed with PBS. Fluorescence was viewed by a Zeiss LSM 5 Pascal Vario 2 Laser Scanning Confocal Microscope (Carl Zeiss Jena, Germany). The excitation color wheel contain BS505-530 (green) and LP560 (red) filter blocks which allowed for the separation of emitted green light from AO into the green channel and red light from EtBr into the red channel. The laser power output and gain control settings within each individual experiment were identical between control and DEFB1 induced cells. The excitation was provided by a Kr/Ar mixed gas laser at wavelengths of 543 nm for AO and 488 nm for EtBr. Slides were analyzed under 40× magnification and digital images were stored as uncompressed TIFF files and exported into Photoshop CS software (Adobe Systems, San Jose, Calif.) for image processing and hard copy presentation.
[0252] ChIP Analysis of PAX2
[0253] Chromatin immunoprecipitation (ChIP) allows the identification of binding sites for DNA-binding proteins based upon in vivo occupancy of a promoter by a transcription factor and enrichment of transcription factor bound chromatin by immunoprecipitation (66). A modification of the protocol described by the Farnham laboratory ((67, 68) was used; also on line at http://mcardle.oncology.wisc.edu/farnham/). The DU145 and PC3 cell lines over-expresses the PAX2 protein, but does not express DEFB1. Cells were incubated with PBS containing 1.0% formaldehyde for 10 minutes to crosslink proteins to DNA. Samples were then sonicated to yield DNA with an average length of 600 bp. Sonicated chromatin precleared with Protein A Dynabeads was incubated with PAX2-specific antibody or "no antibody" control [isotype-matched control antibodies]. Washed immunoprecipitates were then collected. After reversal of the crosslinks, DNA was analyzed by PCR using promoter-specific primers to determine whether DEFB1 is represented in the PAX2-immunoprecipitated samples. Primers were designed to amplify the 160 bp region immediately upstream of the DEFB1 mRNA start site which contained the DEFB1 TATA box and the functional GTTCC (SEQ ID NO: 2) PAX2 recognition site. For these studies, positive controls included PCR of an aliquot of the input chromatin (prior to immunoprecipitation, but crosslinks reversed). All steps were performed in the presence of protease inhibitors.
Results
[0254] siRNA Inhibition of PAX2 Increases DEFB1 Expression
[0255] QRT-PCR analysis of DEFB1 expression before siRNA treatment revealed relative expression levels of 0.00097 in DU145, 0.00001 in PC3, and 0.00004 LNCaP (FIG. 13). Following siRNA knock-down of PAX2, relative expression was 0.03294 (338-fold increase) in DU145, 0.00020 (22.2-fold increase) in PC3 and 0.00019 (4.92-fold increase) in LNCaP. As a negative control, the human prostate epithelial cell line (hPrEC) which is PAX2 null, revealed expression levels at 0.00687 before treatment and 0.00661 following siRNA treatment confirming no statistical change in DEFB1 expression.
[0256] DEFB1 Causes Cell Membrane Permeability
[0257] Membrane integrity was monitored by confocal analysis (FIG. 14). Here, intact cells stain green due to AO which is membrane permeable. In addition, cells with compromised plasma membranes would stain red by EtBr which is membrane impermeable. Here, uninduced DU145 (A) and PC3 (D) cells stained positively with AO and emitted green color, but did not stain with EtBr. However, DEFB1 induction in both DU145 (B) and PC3 (E) resulted in the accumulation of EtBr in the cytoplasm at 24 hours indicated by the red staining. By 48 hours, DU145 (C) and PC3 (F) possessed condensed nuclei and appeared yellow, which was due to the presence of both green and red staining resulting from the accumulation of AO and EtBr, respectively.
[0258] Inhibition of PAX2 Results in Membrane Permeability
[0259] Cells were treated with PAX2 siRNA for 4 days and membrane integrity was monitored again by confocal analysis (FIG. 15). Here, both DU145 (B) and PC3 (E) possessed condensed nuclei and appeared yellow. However, LNCaP cells' cytoplasm and nuclei remained green following siRNA treatment. Also red staining at the cell periphery indicates the maintenance of cell membrane integrity. These findings indicate that the inhibition of PAX2 results in specifically DEFB1-mediated cell death in DU1145 and PC3, but not LNCaP cells. Death observed in LNCaP (refer to Chapter II) is due to the transactivation of the existing wild-type p53 in LNCap following PAX2 inhibition.
[0260] siRNA Inhibition of PAX2 Increases DEFB1 Promoter Activity
[0261] Analysis of DEFB1 promoter activity in DU145 cells containing the DEFB1/pGL3 construct revealed a 2.65 fold increase in relative light units following 48 hours of treatment compared to untreated cells (FIG. 16). In PC3 cells, there was a 3.78-fold increase in relative light units compared to untreated cells.
[0262] PAX2 Binds to the DEFB1 Promoter
[0263] ChIP analysis was performed on DU145 and PC3 cells to determine if the PAX2 transcriptional repressor is bound to the DEFB1 promoter (FIG. 17). Lane 1 contains a 100 bp molecular weight marker. Lane 2 is a positive control representing 160 bp region of the DEFB1 promoter amplified from DU145 before cross-linking and immunoprecipitation. Lane 3 is a negative control representing PCR performed without DNA. Lane 4 and 5 are negative controls representing PCR from immunoprecipitations performed with IgG from cross-linked DU145 and PC3, respectively. PCR amplification of 25 pg of DNA (lane 6 and 8) and 50 pg of DNA (lane 7 and 9) immunoprecitipated with anti-PAX2 antibody after crosslinking show 160 bp promoter fragment in DU145 and PC3, respectively.
Conclusion
[0264] The present novel data are the first to disclose the role of DEFB1 in prostate cancer tumor immunity. The data also show that the oncogenic factor PAX2 suppresses DEFB1 expression. One of the hallmarks of defensin cytotoxicity is the disruption of membrane integrity. The present results show that ectopic expression of DEFB1 in prostate cancer cells results in a loss of membrane potential due to compromised cell membranes. The same phenomenon is observed after inhibiting PAX2 protein expression. ChIP analysis was also performed and confirmed that PAX2 is bound to the DEFB1 promoter resulting in the repression of DEFB1 expression. Therefore, suppression of PAX2 expression or function, results in the re-establishment of DEFB1 expression and subsequently DEFB1-mediated cell death. Also, the present data establish the utility of DEFB1 as a directed therapy for prostate cancer treatment through innate immunity.
Example IV
Expression of DEFB1 Results in Tumor Shrinkage
[0265] The anti-tumoral ability of DEFB1 is evaluated by injecting tumor cells that overexpress DEFB1 into nude mice. DEFB1 is cloned into pBI-EGFP vector, which has a bidirectional tetracycline responsible promoter. Tet-Off Cell lines are generated by transfecting pTet-Off into DU145, PC3 and LNCaP cells and selecting with G418. The pBI-EGFP-DEFB1 plasmid is co-transfected with pTK-Hyg into the Tet-off cell lines and selected with hygromycin. Only single-cell suspensions with a viability of >90% are used. Each animal receives approximately 500,000 cells administered subcutaneously into the right flank of female nude mice. There are two groups, a control group injected with vector only clones and a group injected with the DEFB1 over-expressing clones. 35 mice are in each group as determined by a statistician. Animals are weighed twice weekly, tumor growth monitored by calipers and tumor volumes determined using the following formula: volume=0.5×(width)2×length. All animals are sacrificed by CO2 overdose when tumor size reaches 2 mm3 or 6 months following implantation; tumors are excised, weighed and stored in neutral buffered formalin for pathological examination. Differences in tumor growth between the groups are descriptively characterized through summary statistics and graphical displays. Statistical significance is evaluated with either the t-test or non-parametric equivalent.
Example V
Expression of PAX2 siRNA Results in Up-Regulation of DEFB1 Expression and Tumor Shrinkage In Vivo
[0266] Hairpin PAX2 siRNA template oligonucleotides utilized in the in vitro studies are utilized to examine the effect of the up-regulation of DEFB1 expression in vivo. The sense and antisense strand (see Table 4) are annealed and cloned into pSilencer 2.1 U6 hygro siRNA expression vector (Ambion) under the control of the human U6 RNA pol III promoter. The cloned plasmid is sequenced, verified and transfected into PC3, Du145, and LNCap cell lines. Scrambled shRNA is cloned and used as a negative control in this study. Hygromycin resistant colonies are selected, cells are introduced into the mice subcutaneously and tumor growth is monitored as described above.
Example VI
Small Molecule Inhibitors of PAX2 Binding Results in Up-Regulation of DEFB1 Expression and Tumor Shrinkage In Vivo
[0267] The DNA recognition sequence for PAX2 binding resides in the DEFB1 promoter between nucleotides -75 and -71 [+1 refers to the transcriptional start site]. Short oligonucleotides complementary to the PAX2 DNA-binding domain are provided. Examples of such oligonucleotides include the 20-mer and 40-mer oligonucleotides containing the GTTCC (SEQ ID NO: 2) recognition sequence provided below. These lengths were randomly selected, and other lengths are expected to be effective as blockers of binding. As a negative control, oligonucleotides with a scrambled sequence (CTCTG) (SEQ ID NO: 22) were designed to verify specificity. The oligonucleotides are transfected into the prostate cancer cells and the HPrEC cells with lipofectamine reagent or Codebreaker transfection reagent (Promega, Inc). In order to confirm DNA-protein interactions, double stranded oligonucleotides will be labeled with [32P] dCTP and electrophoretic mobility shift assays are performed. In addition, DEFB1 expression is monitored by QRT-PCR and Western analysis following treatment with oligonucleotides. Finally, cell death is detected by MTT assay and flow cytometry as previously described.
TABLE-US-00006 Recognition Sequence #1: (SEQ ID NO: 18) CTCCCTTCA TCGAC Recognition Sequence #2: (SEQ ID NO: 19) CTCCCTTCACCTTGGTCGAC Scramble Sequence #1: (SEQ ID NO: 23) CTCCCTTCACTCTGGTCGAC Recognition Sequence #3: (SEQ ID NO: 20) ACTGTGGCACCTCCCTTCA GTCGACGAGGTTGTGC Recognition Sequence #4: (SEQ ID NO: 21) ACTGTGGCACCTCCCTTCACCTTGGTCGACGAGGTTGTGC Scramble Sequence #2: (SEQ ID NO: 24) ACTGTGGCACCTCCCTTCACTCTGGTCGACGAGGTTGTGC
[0268] Further examples of oligonucleotides of the invention include:
TABLE-US-00007 Recognition Sequence #1: (SEQ ID No: 25) 5'-AGAAGTTCAC ACTGT-3' Recognition Sequence #2: (SEQ ID No: 26) 5'-AGAAGTTCAC ACTGT-3' Scramble Sequence #1: (SEQ ID No: 27) 5'-AGAAGTTCACGCTCTACTGT-3' Recognition Sequence #3: (SEQ ID No: 28) 5'-TTAGCGATTAGAAGTTCAC ACTGTGGCACCTCCC-3' Recognition Sequence #4: (SEQ ID No: 29) 5'-GTTAGCGATTAGAAGTTCAC ACTGTGGCACCTCCC-3' Scramble Sequence #2: (SEQ ID No: 30) 5'-GTTAGCGATTAGAAGTTCACGCTCTACTGTGGCACCTCCC-3'
[0269] This set of alternative inhibitory oligonucleotides represents the recognition sequence (along with the CCTTG (SEQ ID NO: 1) core sequence) for the PAX2 binding domain and homeobox. These include actual sequences from the DEFB1 promoter.
[0270] The PAX2 gene is required for the growth and survival of various cancer cells including prostate. In addition, the inhibition of PAX2 expression results in cell death mediated by the innate immunity component DEFB1. Suppression of DEFB1 expression and activity is accomplished by binding of the PAX2 protein to a GTTCC (SEQ ID NO: 2) recognition site in the DEFB1 promoter. Therefore, this pathway provides a viable therapeutic target for the treatment of prostate cancer. In this method, the sequences bind to the PAX2 DNA binding site and block PAX2 binding to the DEFB1 promoter thus allowing DEFB1 expression and activity. The oligonucleotide sequences and experiment described above are examples of and demonstrate a model for the design of additional PAX2 inhibitor drugs.
[0271] Given that the GTTCC (SEQ ID NO: 2) sequence exists in interleukin-3, interleukin-4, the insulin receptor and others, PAX2 regulates their expression and activity as well. Therefore the PAX2 inhibitors disclosed herein have utility in a number of other diseases including those directed related to inflammation including prostatitis and benign prostatic hypertrophy (BPH).
Example VII
Loss of DEFB1 Expression Results in Increased Tumorigenesis
[0272] Generation of Loss of Function Mice
[0273] The Cre/loxP system has been useful in elucidating the molecular mechanisms underlying prostate carcinogenesis. Here a DEFB1 Cre conditional KO is used for inducible disruption within the prostate. The DEFB1 Cre conditional KO involves the generation of a targeting vector containing loxP sites flanking DEFB1 coding exons, targeted ES cells with this vector and the generation of germline chimeric mice from these targeted ES cells. Heterozygotes are mated to prostate-specific Cre transgenics and heterozygous intercross is used to generate prostate-specific DEFB1 KO mice. Four genotoxic chemical compounds have been found to induce prostate carcinomas in rodents: N-methyl-N-nitrosourea (MNU), N-nitrosobis 2-oxopropyl. amine (BOP), 3,2X-dimethyl-4-amino-biphenyl (MAB) and 2-amino-1-methyl-6-phenylimidazow 4,5-bxpyridine (PhIP). DEFB1-transgenic mice are treated with these carcinogenic compounds via intra-gastric administration or i.v. injection for prostate adenoma and adenocarcinoma induction studies. Prostate samples are studied for differences in tumor growth and changes gene expression though histological, immunohistological, mRNA and protein analyses.
[0274] Generation of GOF Mice
[0275] For PAX2 inducible GOF mice, PAX2 GOF (bi-transgenic) and wild-type (mono-transgenic) littermates are administered doxycycline (Dox) from 5 weeks of age to induce prostate-specific PAX2 expression. Briefly, PROBASIN-rtTA mono-transgenic mice (prostate cell-specific expression of tet-dependent rtTA inducer) are crossed to our PAX2 transgenic responder lines. For induction, bi-transgenic mice are fed Dox via the drinking water (500 mg/L freshly prepared twice a week). Initial experiments verify low background levels, good inducibility and cell-type specific expression of PAX2 and the EGFP reporter using transgenic founder line in bi-transgenic mice. Regarding experimental group sizes, 5-7 age- and sex-matched individuals in each group (wild-type and GOF) allow for statistical significance. For all animals in this study, prostate tissues are collected initially at weekly intervals for analysis and comparison, to determine carcinogenic time parameters.
[0276] PCR Genotyping, RT-PCR and qPCR
[0277] PROBASIN-rtTA transgenic mice are genotyped using the following PCR primers and conditions: PROBASINS (forward) 5'-ACTGCCCATTGCCCAAACAC-3' (SEQ ID NO: 31); RTTA3 (reverse) 5'-AAAATCTTGCCAGCTTTCCCC-3' (SEQ ID NO: 32); 95° C. denaturation for 5 min, followed by 30 cycles of 95° C. for 30 sec, 57° C. for 30 sec, 72° C. for 30 sec, followed by a 5 min extension at 72° C., yielding a 600 bp product. PAX2 inducible transgenic mice are genotyped using the following PCR primers and conditions: PAX2For 5'-GTCGGTTACGGAGCGGACCGGAG-3'(SEQ ID NO: 33); Rev5'IRES 5'-TAACATATAGACAAACGCACACCG-3' (SEQ ID NO: 34); 95° C. denaturation for 5 min, followed by 34 cycles of 95° C. for 30 sec, 63° C. for 30 sec, 72° C. for 30 sec, followed by a 5 min extension at 72° C., yielding a 460 bp product. Immortomouse hemizygotes are be genotyped using the following PCR primers and conditions: Immol1, 5'-GCGCTTGTGTC GCCATTGTATTC-3' (SEQ ID NO: 35); Immol2, 5'-GTCACACCACAGAAGTAAGGTTCC-3' (SEQ ID NO: 36); 94° C. 30 sec, 58° C. 1 min, 72° C. 1 min 30 sec, 30 cycles to yield a 1 kb transgene band. For genotyping PAX2 knockout mice, the following PCR primers and conditions are used: PAX2 For 5'-GTCGGTTACGGAGCGGACCGGAG-3' (SEQ ID NO: 37); PAX2Rev 5'-CACAGAGCATTGGCGATCTCGATGC-3' (SEQ ID NO: 38); 94° C. 1 min, 65° C. 1 min, 72° C. 30 sec, 36 cycles to yield a 280 bp band.
[0278] DEFB1 Peptide Animal Studies
[0279] Six-week-old male athymic (nude) mice purchased from Charles River Laboratories are injected sub-cutaneously over the scapula with 106 viable PC3 cells. One week after injection, the animals are randomly allocated to one of three groups--group I: control; group II: intraperitoneal injections of DEFB1, 100 μg/day, 5 days a week, for weeks 2-14; group III: intraperitoneal injections of DEFB1, 100 mg/day, 5 days a week, for weeks 8-14. Animals are maintained in sterile housing, four animals to a cage, and observed on a daily basis. At 10-day intervals, the tumors are measured by using calipers, and the volumes of the tumors are calculated by using V=(L×W2)/2.
TABLE-US-00008 TABLE 3 Sequences of QRT-PCR Primers. Sense (5'-3') Antisense (5'-3') β-actin 5'-CCTGGCACCCAGCACAAT-3' 5'-GCC GATCCACACGGAGTACT-3' (SEQ ID NO: 51) (SEQ ID NO: 52) DEFB1 5'-GTTGCCTGCCAGTC GCCAT GAGAACTTCCTAC-3' 5'-TGGCCTTCCCTCTGTA ACAGGTGCCTTGAATT-3' (SEQ ID NO: 53) (SEQ ID NO: 54)
TABLE-US-00009 TABLE 4 PAX2 siRNA Sequences: a pool of four siRNA was utilized to inhibit PAX2 protein expression. Sense (5'-3') Antisense (5'-3') Sequence A 5'-GAAGUCAAGUCGAGUCUAUUU-3' 5'-AUAGACUCGACUUGACUUCUU-3' (SEQ ID NO: 7) (SEQ ID NO: 3) Sequence B 5'-GAGGAAACGUGAUGAAGAUUU-3' 5'-AUCUUCAUCACGUUUCCUCUU-3' (SEQ ID NO: 8) (SEQ ID NO: 4) Sequence C 5'-GGACAAGAUUGCUGAAUACUU-3' 5'-GUAUUCAGCAAUCUUGUCCUU-3' (SEQ ID NO: 9) (SEQ ID NO: 5) Sequence D 5'-CAUCAGAGCACAUCAAAUCUU-3' 5'-GAUUUGAUGUGCUCUGAUGUU-3' (SEQ ID NO: 10) (SEQ ID NO: 6)
TABLE-US-00010 TABLE 5 Quantitative RT-PCR Primers: nucleotide sequences of primers used to amplify PAX2 and GAPDH. Sense (5'-3') Antisense (5'-3') GAPDH 5'-CCACCCATGGCAAATTCCATGGCA-3' 5'-TCTAGACGGCAGGTCAGGTCAACC-3' (SEQ ID NO: 55) (SEQ ID NO: 56) BAD 5'-CTCAGGCCTATGCAAAAAGAGGA-3' 5'-GCCCTCCCTCCAAAGGAGAC-3' (SEQ ID NO: 57) (SEQ ID NO: 58) BID 5'-AACCTACGCACCTACGTGAGGAG-3' 5'-CGTTCAGTCCATCCCATTTCTG-3' (SEQ ID NO: 59) (SEQ ID NO: 60) BAX 5'-GACACCTGAGCTGACCTTGG-3' 5'-GAGGAAGTCCAGTGTCCAGC-3' (SEQ ID NO: 61) (SEQ ID NO: 62)
[0280] Throughout this application, various publications are referenced. The disclosures of these publications in their entireties are hereby incorporated by reference into this application in order to more fully describe the state of the art to which this invention pertains.
REFERENCES
[0281] 1. Jemal, A.; Tiwari, R. C.; Murray, T.; Ghafoor, A.; Samuels, A.; Ward, E.; Feuer, E. J.; Thun, M. J., Cancer statistics, 2004. CA Cancer J Clin 2004, 54, (1), 8-29. [0282] 2. Prasad, M. A.; Trybus, T. M.; Wojno, K. J.; Macoska, J. A., Homozygous and frequent deletion of proximal 8p sequences in human prostate cancers: identification of a potential tumor suppressor gene site. Genes Chromosomes Cancer 1998, 23, (3), 255-62. [0283] 3. McNeel, D. G.; Malkovsky, M., Immune-based therapies for prostate cancer. Immunology Letters 2005, 96, (1), 3. [0284] 4. Tien, A. H.; Xu, L.; Helgason, C. D., Altered Immunity Accompanies Disease Progression in a Mouse Model of Prostate Dysplasia. Cancer Res 2005, 65, (7), 2947-2955. [0285] 5. Banchereau, J.; Palucka, A. K.; Dhodapkar, M.; Burkeholder, S.; Taquet, N.; Rolland, A.; Taquet, S.; Coquery, S.; Wittkowski, K. M.; Bhardwaj, N.; Pineiro, L.; Steinman, R.; Fay, J., Immune and Clinical Responses in Patients with Metastatic Melanoma to CD34+ Progenitor-derived Dendritic Cell Vaccine. Cancer Res 2001, 61, (17), 6451-6458. [0286] 6. Fong, L.; Brockstedt, D.; Benike, C.; Breen, J. K.; Strang, G.; Ruegg, C. L.; Engleman, E. G., Dendritic Cell-Based Xenoantigen Vaccination for Prostate Cancer Immunotherapy. J Immunol 2001, 167, (12), 7150-7156. [0287] 7. Linzmeier, R.; Ho, C. H.; Hoang, B. V.; Ganz, T., A 450-kb contig of defensin genes on human chromosome 8p23. Gene 1999, 233, (1-2), 205-11. [0288] 8. Yang, D.; Biragyn, A.; Hoover, D. M.; Lubkowski, J.; Oppenheim, J. J., Multiple Roles of Antimicrobial Defensins, Cathelicidins, and Eosinophil-Derived Neurotoxin in Host Defense. Annual Review of Immunology 2004, 22, (1), 181-215. [0289] 9. Donald, C. D.; Sun, C. Q.; Lim, S. D.; Macoska, J.; Cohen, C.; Amin, M. B.; Young, A. N.; Ganz, T. A.; Marshall, F. F.; Petros, J. A., Cancer-specific loss of beta-defensin 1 in renal and prostatic carcinomas. Lab Invest 2003, 83, (4), 501-5. [0290] 10. Ganz, T., Defensins: antimicrobial peptides of vertebrates. C R Biol 2004, 327, (6), 539-49. [0291] 11. Mazzucchelli, R.; Barbisan, F.; Tarquini, L. M.; Galosi, A. B.; Stramazzotti, D., Molecular mechanisms in prostate cancer. A review. Anal Quant Cytol Histol 2004, 26, (3), 127-33. [0292] 12. Ganz, T., Defensins and host defense. Science 1999, 286, (5439), 420-1. [0293] 13. Ganz, T., Immunology. Versatile defensins. Science 2002, 298, (5595), 977-9. [0294] 14. Braida, L.; Boniotto, M.; Pontillo, A.; Tovo, P. A.; Amoroso, A.; Crovella, S., A single-nucleotide polymorphism in the human beta-defensin 1 gene is associated with HIV-1 infection in Italian children. Aids 2004, 18, (11), 1598-600. [0295] 15. Gropp, R.; Frye, M.; Wagner, T. O.; Bargon, J., Epithelial defensins impair adenoviral infection: implication for adenovirus-mediated gene therapy. Hum Gene Ther 1999, 10, (6), 957-64. [0296] 16. Catalano, M. G.; Pfeffer, U.; Raineri, M.; Ferro, P.; Curto, A.; Capuzzi, P.; Como, F.; Berta, L.; Fortunati, N., Altered expression of androgen-receptor isoforms in human colon-cancer tissues. Int J Cancer 2000, 86, (3), 325-30. [0297] 17. Takeuchi, S.; Iida, M.; Kobayashi, S.; Jin, K.; Matsuda, T.; Kojima, H., Differential effects of phthalate esters on transcriptional activities via human estrogen receptors alpha and beta, and androgen receptor. Toxicology 2005, 210, (2-3), 223-33. [0298] 18. Zucht, H. D.; Grabowsky, J.; Schrader, M.; Liepke, C.; Jurgens, M.; Schulz-Knappe, P.; Forssmann, W. G., Human beta-defensin-1: A urinary peptide present in variant molecular forms and its putative functional implication. Eur J Med Res 1998, 3, (7), 315-23. [0299] 19. Nishimura, M.; Abiko, Y.; Kurashige, Y.; Takeshima, M.; Yamazaki, M.; Kusano, K.; Saitoh, M.; Nakashima, K.; Inoue, T.; Kaku, T., Effect of defensin peptides on eukaryotic cells: primary epithelial cells, fibroblasts and squamous cell carcinoma cell lines. Journal of Dermatological Science 2004, 36, (2), 87. [0300] 20. Fromont, G.; Joulin, V.; Chantrel-Groussard, K.; Vallancien, G.; Guillonneau, B.; Validire, P.; Latil, A.; Cussenot, O., Allelic losses in localized prostate cancer: association with prognostic factors. J Urol 2003, 170, (4 Pt 1), 1394-7. [0301] 21. Wang, Z.; Lai, F. M., [Analysis of loss of heterozygosity on chromosome 8 in human prostate carcinoma and high grade prostatic intraepithelial neoplasia]. Zhonghua Nan Ke Xue 2004, 10, (1), 26-8, 31. [0302] 22. Hugel, A.; Wernert, N., Loss of heterozygosity (LOH), malignancy grade and clonality in microdissected prostate cancer. Br J Cancer 1999, 79, (3-4), 551-7. [0303] 23. Bockmuhl, U.; Ishwad, C. S.; Ferrell, R. E.; Gollin, S. M., Association of 8p23 deletions with poor survival in head and neck cancer. Otolaryngol Head Neck Surg 2001, 124, (4), 451-5. [0304] 24. Macoska, J. A.; Paris, P.; Collins, C.; Andaya, A.; Beheshti, B.; Chaib, H.; Kant, R.; Begley, L.; MacDonald, J. W.; Squire, J. A., Evolution of 8p loss in transformed human prostate epithelial cells. Cancer Genet Cytogenet 2004, 154, (1), 36-43. [0305] 25. Chaib, H.; MacDonald, J. W.; Vessella, R. L.; Washburn, J. G.; Quinn, J. E.; Odman, A.; Rubin, M. A.; Macoska, J. A., Haploinsufficiency and reduced expression of genes localized to the 8p chromosomal region in human prostate tumors. Genes Chromosomes Cancer 2003, 37, (3), 306-13. [0306] 26. Teixeira, M. R.; Ribeiro, F. R.; Eknaes, M.; Waehre, H.; Stenwig, A. E.; Giercksky, K. E.; Heim, S.; Lothe, R. A., Genomic analysis of prostate carcinoma specimens obtained via ultrasound-guided needle biopsy may be of use in preoperative decision-making. Cancer 2004, 101, (8), 1786-93. [0307] 27. Vecchione, A.; Ishii, H.; Baldassarre, G.; Bassi, P.; Trapasso, F.; Alder, H.; Pagano, F.; Gomella, L. G.; Croce, C. M.; Baffa, R., FEZ1/LZTS1 is down-regulated in high-grade bladder cancer, and its restoration suppresses tumorigenicity in transitional cell carcinoma cells. Am J Pathol 2002, 160, (4), 1345-52. [0308] 28. Valore, E. V.; Park, C. H.; Quayle, A. J.; Wiles, K. R.; McCray, P. B., Jr.; Ganz, T., Human beta-defensin-1: an antimicrobial peptide of urogenital tissues. J Clin Invest 1998, 101, (8), 1633-42. [0309] 29. Gunther, M.; Wagner, E.; Ogris, M., Specific targets in tumor tissue for the delivery of therapeutic genes. Curr Med Chem Anti-Canc Agents 2005, 5, (2), 157-71. [0310] 30. Dressler, G. R., Pax-2, kidney development, and oncogenesis. Med Pediatr Oncol 1996, 27, (5), 440-4. [0311] 31. Eccles, M. R.; He, S.; Legge, M.; Kumar, R.; Fox, J.; Zhou, C.; French, M.; Tsai, R. W., PAX genes in development and disease: the role of PAX2 in urogenital tract development. Int J Dev Biol 2002, 46, (4), 535-44. [0312] 32. Dressler, G. R.; Woolf, A. S., Pax2 in development and renal disease. Int J Dev Biol 1999, 43, (5), 463-8. [0313] 33. Khoubehi, B.; Kessling, A. M.; Adshead, J. M.; Smith, G. L.; Smith, R. D.; Ogden, C. W., Expression of the developmental and oncogenic PAX2 gene in human prostate cancer. J Urol 2001, 165, (6 Pt 1), 2115-20. [0314] 34. Havik, B.; Ragnhildstveit, E.; Lorens, J. B.; Saelemyr, K.; Fauske, O.; Knudsen, L. K.; Fjose, A., A novel paired domain DNA recognition motif can mediate Pax2 repression of gene transcription. Biochem Biophys Res Commun 1999, 266, (2), 532-41. [0315] 35. Discenza, M. T.; He, S.; Lee, T. H.; Chu, L. L.; Bolon, B.; Goodyer, P.; Eccles, M.; Pelletier, J., WT1 is a modifier of the Pax2 mutant phenotype: cooperation and interaction between WT1 and Pax2. Oncogene 2003, 22, (50), 8145-55. [0316] 36. McConnell, M. J.; Cunliffe, H. E.; Chua, L. J.; Ward, T. A.; Eccles, M. R., Differential regulation of the human Wilms tumour suppressor gene (WT1) promoter by two isoforms of PAX2. Oncogene 1997, 14, (22), 2689-700. [0317] 37. Yuan, S. S.; Yeh, Y. T.; Lee, E. Y., Pax-2 interacts with RB and reverses its repression on the promoter of Rig-1, a Robo member. Biochem Biophys Res Commun 2002, 296, (4), 1019-25. [0318] 38. Stuart, E. T.; Haffner, R.; Oren, M.; Gruss, P., Loss of p53 function through PAX-mediated transcriptional repression. Embo J 1995, 14, (22), 5638-45. [0319] 39. Michalak, E.; Villunger, A.; Erlacher, M.; Strasser, A., Death squads enlisted by the tumour suppressor p53. Biochem Biophys Res Commun 2005, 331, (3), 786-98. [0320] 40. Tokino, T.; Nakamura, Y., The role of p53-target genes in human cancer. Crit Rev Oncol Hematol 2000, 33, (1), 1-6. [0321] 41. Muratovska, A.; Zhou, C.; He, S.; Goodyer, P.; Eccles, M. R., Paired-Box genes are frequently expressed in cancer and often required for cancer cell survival. Oncogene 2003, 22, (39), 7989-97. [0322] 42. Tagge, E. P.; Hanson, P.; Re, G. G.; Othersen, H. B., Jr.; Smith, C. D.; Garvin, A. J., Paired box gene expression in Wilms' tumor. J Pediatr Surg 1994, 29, (2), 134-41. [0323] 43. Murer, L.; Caridi, G.; Della Vella, M.; Montini, G.; Carasi, C.; Ghiggeri, G.; Zacchello, G., Expression of nuclear transcription factor PAX2 in renal biopsies of juvenile nephronophthisis. Nephron 2002, 91, (4), 588-93. [0324] 44. Eccles, M. R.; Wallis, L. J.; Fidler, A. E.; Spun, N. K.; Goodfellow, P. J.; Reeve, A. E., Expression of the PAX2 gene in human fetal kidney and Wilms' tumor. Cell Growth Differ 1992, 3, (5), 279-89. [0325] 45. Ogata, T.; Muroya, K.; Sasagawa, I.; Kosho, T.; Wakui, K.; Sakazume, S.; Ito, K.; Matsuo, N.; Ohashi, H.; Nagai, T., Genetic evidence for a novel gene(s) involved in urogenital development on 10q26. Kidney Int 2000, 58, (6), 2281-90. [0326] 46. Ostrom, L.; Tang, M. J.; Gruss, P.; Dressler, G. R., Reduced Pax2 gene dosage increases apoptosis and slows the progression of renal cystic disease. Dev Biol 2000, 219, (2), 250-8. [0327] 47. Mazal, P. R.; Stichenwirth, M.; Koller, A.; Blach, S.; Haitel, A.; Susani, M., Expression of aquaporins and PAX-2 compared to CD10 and cytokeratin 7 in renal neoplasms: a tissue microarray study. Mod Pathol 2005, 18, (4), 535-40. [0328] 48. Buttiglieri, S.; Deregibus, M. C.; Bravo, S.; Cassoni, P.; Chiarle, R.; Bussolati, B.; Camussi, G., Role of Pax2 in apoptosis resistance and proinvasive phenotype of Kaposi's sarcoma cells. J Biol Chem 2004, 279, (6), 4136-43. [0329] 49. Gnarra, J. R.; Dressler, G. R., Expression of Pax-2 in human renal cell carcinoma and growth inhibition by antisense oligonucleotides. Cancer Res 1995, 55, (18), 4092-8. [0330] 50. Strasser, A., The role of BH3-only proteins in the immune system. Nat Rev Immunol 2005, 5, (3), 189-200. [0331] 51. Lin, S.; Ying, S. Y., Differentially expressed genes in activin-induced apoptotic LNCaP cells. Biochem Biophys Res Commun 1999, 257, (1), 187-92. [0332] 52. Perfettini, J. L.; Kroemer, R. T.; Kroemer, G., Fatal liaisons of p53 with Bax and Bak. Nat Cell Biol 2004, 6, (5), 386-8. [0333] 53. Coultas, L.; Strasser, A., The role of the Bcl-2 protein family in cancer. Semin Cancer Biol 2003, 13, (2), 115-23. [0334] 54. Margue, C. M.; Bernasconi, M.; Barr, F. G.; Schafer, B. W., Transcriptional modulation of the anti-apoptotic protein BCL-XL by the paired box transcription factors PAX3 and PAX3/FKHR. Oncogene 2000, 19, (25), 2921-9. [0335] 55. Nakamura, Y., Isolation of p53-target genes and their functional analysis. Cancer Sci 2004, 95, (1), 7-11. [0336] 56. Perfettini, J. L.; Roumier, T.; Kroemer, G., Mitochondrial fusion and fission in the control of apoptosis. Trends Cell Biol 2005, 15, (4), 179-83. [0337] 57. Orlando, V. Mapping chromosomal proteins in vivo by formaldehyde-crosslinked-chromatin immunoprecipitation. Trends Biochem Sci, 25: 99-104., 2000. [0338] 58. Boyd, K. E. and Farnham, P. J. Coexamination of site-specific transcription factor binding and promoter activity in living cells. Mol Cell Biol, 19: 8393-8399., 1999. [0339] 59. Wells, J. and Farnham, P. J. Characterizing transcription factor binding sites using formaldehyde crosslinking and immunoprecipitation. Methods, 26: 48-56., 2002. [0340] 60. Sikorski, R. S. and Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics, 122: 19-27., 1989. [0341] 61. Nigro, J. M., Sikorski, R., Reed, S. I., and Vogelstein, B. Human p53 and CDC2Hs genes combine to inhibit the proliferation of Saccharomyces cerevisiae. Mol Cell Biol, 12: 1357-1365, 1992. [0342] 62. Wilson, T. E., Fahrner, T. J., Johnston, M., and Milbrandt, J. Identification of the DNA binding site for NGFI-B by genetic selection in yeast. Science, 252: 1296-1300., 1991. [0343] 63. Liu, J., Wilson, T. E., Milbrandt, J., and Johnsen, M. Identifying DNA-binding sites and analyzing DNA-binding domains using a yeast selection system. METHODS: A companion to Methods in Enzymology, 5: 125-137, 1993. [0344] 64. Jackers, P., Szalai, G., and Watson, D. K. Ets-dependent regulation of target gene expression during megakaryopoiesis. in preparation, 2003.
[0345] It will be apparent to those skilled in the art that various modifications and variations can be made in the present invention without departing from the scope or spirit of the invention. Other embodiments of the invention will be apparent to those skilled in the art from consideration of the specification and practice of the invention disclosed herein. It is intended that the specification and examples be considered as exemplary only, with a true scope and spirit of the invention being indicated by the following claims.
Sequence CWU
1
6415DNAHomo sapiens 1ccttg
525DNAHomo sapiens 2gttcc
5321RNAHomo sapiens 3auagacucga
cuugacuucu u 21421RNAHomo
sapiens 4aucuucauca cguuuccucu u
21521RNAHomo sapiens 5guauucagca aucuuguccu u
21621RNAHomo sapiens 6gauuugaugu gcucugaugu u
21721RNAHomo sapiens 7gaagucaagu
cgagucuauu u 21821RNAHomo
sapiens 8gaggaaacgu gaugaagauu u
21921RNAHomo sapiens 9ggacaagauu gcugaauacu u
211021RNAHomo sapiens 10caucagagca caucaaaucu u
211120DNAHomo sapiens
11acccgactat gttcgcctgg
201221DNAHomo sapiens 12aagctctgga tcgagtcttt g
211320DNAHomo sapiens 13atgtgtcagg cacacagacg
201421RNAHomo sapiens
14gucgagucua ucugcauccu u
211521RNAHomo sapiens 15ggaugcagau agacucgacu u
211619DNAHomo sapiens 16aagttcaccc ttgactgtg
19177DNAHomo
sapiensmisc_structure(1)..(1)each n independently represents 1 to 35
continuous flanking nucleotides of DEFB1 DNA core sequence of PAX2
protein binding site 17nccttgn
71820DNAHomo sapiens 18ctcccttcag ttccgtcgac
201920DNAHomo sapiens 19ctcccttcac
cttggtcgac 202040DNAHomo
sapiens 20actgtggcac ctcccttcag ttccgtcgac gaggttgtgc
402140DNAHomo sapiens 21actgtggcac ctcccttcac cttggtcgac gaggttgtgc
40225DNAHomo sapiens 22ctctg
52320DNAHomo sapiens
23ctcccttcac tctggtcgac
202440DNAHomo sapiens 24actgtggcac ctcccttcac tctggtcgac gaggttgtgc
402520DNAHomo sapiens 25agaagttcac ccttgactgt
202620DNAHomo sapiens
26agaagttcac gttccactgt
202720DNAHomo sapiens 27agaagttcac gctctactgt
202839DNAHomo sapiens 28ttagcgatta gaagttcacc
cttgactgtg gcacctccc 392940DNAHomo sapiens
29gttagcgatt agaagttcac gttccactgt ggcacctccc
403040DNAHomo sapiens 30gttagcgatt agaagttcac gctctactgt ggcacctccc
403120DNAHomo sapiens 31actgcccatt gcccaaacac
203221DNAHomo sapiens
32aaaatcttgc cagctttccc c
213323DNAHomo sapiens 33gtcggttacg gagcggaccg gag
233424DNAHomo sapiens 34taacatatag acaaacgcac accg
243523DNAHomo sapiens
35gcgcttgtgt cgccattgta ttc
233624DNAHomo sapiens 36gtcacaccac agaagtaagg ttcc
243723DNAHomo sapiens 37gtcggttacg gagcggaccg gag
233825DNAHomo sapiens
38cacagagcat tggcgatctc gatgc
2539135PRTHomo sapiens 39Met Asp Met His Cys Lys Ala Asp Pro Phe Ser Ala
Met His Arg His1 5 10
15Gly Gly Val Asn Gln Leu Gly Gly Val Phe Val Asn Gly Arg Pro Leu
20 25 30Pro Asp Val Val Arg Gln Arg
Ile Val Glu Leu Ala His Gln Gly Val 35 40
45Arg Pro Cys Asp Ile Ser Arg Gln Leu Arg Val Ser His Gly Cys
Val 50 55 60Ser Lys Ile Leu Gly Arg
Tyr Tyr Glu Thr Gly Ser Ile Lys Pro Gly65 70
75 80Val Ile Gly Gly Ser Lys Pro Lys Val Ala Thr
Pro Lys Val Val Asp 85 90
95Lys Ile Ala Glu Tyr Lys Arg Gln Asn Pro Thr Met Phe Ala Trp Glu
100 105 110Ile Arg Asp Arg Leu Leu
Ala Glu Gly Ile Cys Asp Asn Asp Thr Val 115 120
125Pro Ser Val Ser Ser Ile Asn 130
135407331DNAHomo sapiensmisc_feature(1)..(7331)n= a, c, g, or t
40ttcccccttt ccangagggc ctaatccgtt gcgcgcgcgc acgcggacac acacacacac
60acacacacac acacacacac acacacggcc cccatagcca ccgcaactct cagcagcagn
120ncctagctcc tctgacccga ggccccaaga cggcgggcac aggaacccct gggacgtcct
180ggctccaggc tggacgtagg cggaggtggc aggagtggac aaacccaggc gggtcccacg
240acgccccttt cctcgggtct ctccttgttt cagccagccg ctctcgcccc tggtcccctc
300ttccctgcgt tagggtcctt tgtctccagc cacctcgcag cctgtccccg cctcggcggc
360cctgcccttt gggcctccca gatctctctg gcgggtcccc ctgccttacc agctcccggc
420tgtggcgcgc tcttcgcctg ctcctcacat ncacacagct gctgggagag gaggaaggaa
480aggcggncgc gccgcggatg gatccgagac ggtagatttg gtgccggctc gcaaactctg
540ggaaacttaa ngccggttct tccgcccctc tncaactatg nccagcgcgg cccggtcgcg
600cgcgctcacc ccgcggggac cctttccttt tcctgtattt cggctgcggc tgtttcgctt
660cctctggtct cccagccttt ggagtggctt ccctggccct gcactccgtt ccctttcggc
720cgcccccggc tgtcgcctgc ccccaccctc cgcaggtccc acggtcgcgg cggcgatgac
780tgtggaggta acgccgggga cgtcctgggt cagcctgcac cgtctccctc gaccacagcc
840cgatgaggcc gcgggctccg ggccggctgc taagagagtt aatcattact tcgccagcga
900cactcagcct ccccttccga ctctctcgcc cggcctaggg gaggagggga ggggacagct
960ggccaggtgg ggacttcggc ttcgcacaaa ccagcctctt caggcctccc agagacaggt
1020ggtggcttct cagttccctc ggcaactctc taaggtcctc tttcttcccc tcctgtctct
1080ccctccttcg agcctcctcc cagccaggcc tctccccacc gtctcctgtc cgctctggct
1140ttgactgatt aactgcaggt cctgggagaa ccaactttct ttgtttggaa ccggaccgga
1200cgggatttcc ttccctaggt ctccgccaat gggccagctc ctcccgacgg ttttggcgga
1260ctggctgaag aggaccgcgc ctgaggccac aattaacccg gctgttggtg gtggtggttg
1320gggggtgggc agtgaggaat ttaaccgatc ctctagcagc tgcgctggtg cagttgggag
1380gggggtgcag gaagtgggaa tggaggagtg gcaggaggta tagacagagg gaagaacgat
1440aaacctggac aggtgtggca tagccaatag aaggggaaac aaaataaaac aggaaggcgg
1500cgcggggagg aatccccagt aacctttata ggattgaagt tgggtggaaa acgccacctc
1560ctgccctacc ttagcactca gatccctcct ttacctcttt gtgaaagggt aagagttcag
1620aaagctggcc atttactcca taatctacta gagaaatgtc tgggtttgca aaatgcctat
1680tgattagctc catggagtag acaagacagg cgtaattatc cccattttac aggtgagaaa
1740actgagtctc aaagaagcaa agggactgtg tatgtagtgg ctgtcacttt ttcctgtagg
1800ctgtggggtg agtggcccct ttagctgtgc agaggtccat gggtatctag ggaggcggta
1860caggctgtgt ccaggtctga gccagaagta ccagggcctc acggggctcc tagccctttt
1920agcttgttct ctgttggaca ggaccttcac tcttactctc tagacctgct ggctgggttt
1980ctcccagctt cgctattttt tcagttccct agtagagtgg cccatgggcg gtagccacct
2040ggctggcccg tgccactaag aggcagcttt ggtggccaag tggcttgcat tgttgttgct
2100cctcaaaggg cctgtgaagg gctgggcagg tcgcaaagac ctcttgtgag gggaaagcta
2160gattaaaggg ggtaaggatc ctggaggata aaggccaagc acgtgcgcct ggactccaca
2220ggaccaacag accgagcggg cggggccngc tgggagtcag gccccccggg cttcacgcag
2280ggagcccaaa tattgggaac aaaagcagga aaagaagagt gagagcagga gggagggagg
2340gagcgaggaa gcagaaatta gggggtctta gatgaaaaaa aaaagaaagt agctttaggg
2400ggaatgtgct gtggagtgtg aaattgcagc ccatggtgct ccatattgta ccagaagctc
2460ttccaaaaaa aaaaaaaaaa accatcctcc aacgtgacca gagggccagg cagggggaag
2520ggcggggaga gaatggggag gaggaggggg aaaggccggg caggagccgg tcaggccttt
2580ctgcggaagg ggctggggtg taagtttcgg ctccctggga tctgacagcc gagggtatgc
2640gccctggggt gcgccgggac ccagagggcg agtgagcctc ggttggtcgg ctctggagtt
2700cggttgtcag aagaactttt atttttcttt ttggtggtga cttctaaaag tgggaataat
2760ccagaaatga agctcagctg cggagctgca gctctgttct ccctctctcc cctgcctttc
2820tgcttctctt cccttcggac tacttttctc cccttggttc taaatagctt tttcccctct
2880gaactttaat gcatttaatt tggtccgcgc tgtggggagc atttcctggg gagatgcatt
2940taatttcgga atttctaatc ccctccctca gaccccggtc ctagctcccc tagccgctcc
3000ccgggaagtg gaaggaggaa ggcaggtccc ggccacgggg gaggggcgcg gctgggatgc
3060tcccgcggcc ccctccgtct caccaaggct cagccgcctt cccaagctac tggaggccgg
3120gcgcctgggc cccgggtcag ggccctgcan gaagaagaga ggcaaccccc gctttctgcc
3180ttttcttcgc ctgggcaaga aaacgctggg ccagggaact ggaaaccgga aaacaggaga
3240aagggtttnt ggaaggcanc gggagcgggt ggcagncggg gcancgggca ntggactagg
3300tctacaccgg cacttcactt ttgcacaaca tgcccagaaa cgcatttgag agccctggag
3360tcgcgcttgg cttggcttgg ggcgccggtg cgtgggtaca ctcgaggtcg gggtgcctat
3420ccgccacccc gacacctaca cccagtgcag agcaggcgcg gcccagccag acaaccaggc
3480cggcagtagc tcggcctgga gggcggaggc aaggttgggg gccgccaggc gcctgggcaa
3540gcctggcagg gaagggagcc gagaaggcaa aggagccgag atccacaagg aagattnntt
3600gggcagatca gatgcacaga ggcggctaat gaagcaaatc ccgagatggg tttcagagca
3660actccccaaa agtttatttt gcctttaaat ttccgcaggg aggcgggctc cttgtttgaa
3720gtgtaaatgc ccctaggttg gggggtggaa gggccgcttt gaaaacacca gagagaaaag
3780gttcatttag aggcggacgg gaaaagcaac caaccctgac aggtcggagc ccgggtagtg
3840tttggggttg ggtngttttc tttctttctc tttcttttcc cctttcctct tctttcttcc
3900cttttgtgnn ttttnnttgt tttttttntn ttntttttnt ttaantggct ttcttgcttc
3960cccccacccc tctactagac tctatagaag aaagagaaca gaaaaggggg agtcagagga
4020gcggccagtg actggatgaa ggccagccct tcatcctgga gccccaggag aaggcagagc
4080tttggagaaa aggggttcct aatctccagg gagcattact ctttgactct ctagacccag
4140gaatgggctg gacgctaatg gggaagcggc caggaacccg gcctggcgga agagtgagtg
4200tccagctagt gcagtgctgg gaagacgatc ccaggagcag gggggactct caggggctac
4260ctgggaatgg gactatcaga agggtcttta ctcctcanaa ggtgcatgtg aaggacaggt
4320gtgtgaggac aacttccagc acacttggcg cattaagtcc ccttctctac aaaatggaaa
4380atccttctcg cccaacatgt gaaaatgctt gttgtgggca cccacatttc atggtacttg
4440taacatagga catgtctagc tggttctaga aaaatctgtg tctgtgtgga aggggggggg
4500tttactcaca gctttcttcc ttcaatagtt cacacacccc gagacaaatt cctggatgac
4560caacttggag agacctgggg caaaggttac tttagttctg agctcctcta aataaggacc
4620ctttctcaac gttcctttca ccccagttct gggttaatta cttccagtta gtgcgtgttc
4680gtggggttgt gaggccaaag caaacccggg agcgccatct gcaggcctca agaggaagag
4740actgacctta gaggctaggc cctgcgtctt caacctctag cccaagggaa ccaacctgcc
4800tagccaccca agggaagtgg gataggggct gggaggggca ggcggtgagg agtgttttcc
4860tcccagactt taccccgcag gtggattaag cttattgggc tctggaggat acaggaggga
4920gggcaaatgc caggatccca gcggacccag gccccacagg agtgagaggc tcagaacctc
4980gtcccgctga gcctggcctg agctcctcct gaggaataag ggcatcccaa aaacccgggt
5040acaagacgcc cagtagtagt agttaggctg agtcaggcag gtgcatctct ccccatggta
5100tctgccgccc aggctccggc cagagggagg ggagcgcgag tccgcggcgc ttccgcgggg
5160cgcccggaac tgcagacggg ggctggagga atctcggatt cgggctgcaa gagcgctgcg
5220caagcttcgc cgagccgccc tttcgcagac ccagggaagc ggggggaggg agcgaaggag
5280ggagagagag ttaaaacatc agcttgaaag tgcccaagat gattttatta agaccgaggg
5340gaaaattatt ttcatgaaag attctccccg gaatatttct tgtacttaac ccagttagga
5400agacaaaggg cttctttctg cctggtgcgg tgcgagcgga ccccagcgag caagggagct
5460agtgccaaag agaactgcgg aggctccggc aggagtgggg acgtccccgt ggttgcgcct
5520cctgcgctcg ccccggatcc accgagctag cagcgggcgg cgctcagccg cgtccgcagc
5580ctcctcttct ccccagccgg ggagagccag cctcgtctcc cacatcctct gccgccagcg
5640acctgcagct ccgcactgtt tccctcccct gtaccccctt cccagtcacc cgagggttca
5700gaaaccaagt cccccggctc tcccgccatc cgctgggtcc caccgaggca ggtgggtact
5760cgccggaggt cttcagctcg attctgaacc aagcgttctg gactgcccag acccggtggg
5820caaggggact ggggaggccc tgcgcacagt cgcgtggaac gggaggggac aagacaaact
5880gctggacact tttccgtgga atgagaagtg gggggtgcgt gggtgggaag gtacctccgg
5940agggaaaggc caaagggaag gaccagaaag agaggaagga agagccggga aggaacggaa
6000gggaactcag agccgagggt ggtggggttg gggctaggga tgcgcactgg gcccggggcc
6060gcgcggccca ggcgggcact ggccagtgga tggcagggct gggcgagtta gaactgagag
6120cccggcttca cagcgcagcg cgctccgagg ccctctgtcg ttacctgaat attcattaga
6180ctgaccgctc tttatcctta tctaacgttt atcttatcgg cgagtttcgt ttctcagtgt
6240agttttaatc ccgggctccc attccccctc ccccggtccg ctcccctccc tccctcttcc
6300ttcgccggct gctccctccc tccctccctc ccatttctcc ctcccctgcc ctccccttgc
6360cggcaccgga gtgacaggct cggggccctc ctcgccgaag ctcggggctc cagcgctggc
6420gaatcacaga gtggtggaat ctattgcctt tgtctgacaa gtcatccatc tcccggcgcg
6480gggaggggga ggaggtctgg agggggcttt gcagctttta gagagacaca caccgggagc
6540cgaggctcca gtctccggcc gagtcttcta gcagccgcaa cccacctggg gccagcccag
6600agctgccagc gccgctcggc tccctccctc cctcccggcc cttcggccgc ggcggcgtgc
6660gcctgccttt tccgggggcg ggggcctggc ccgcgcgctc ccctcccgca ggcgccacct
6720cggacatccc cgggattgct acttctctgc caacttcgcc aactcgccag cacttggaga
6780ggcccggctc ccctcccggc gccctctgac cgcccccgcc ccgcgcgctc tccgaccacc
6840gcctctcgga tgaacaggtt ccaggggagc tgagcgagtc gcctcccccg cccagcttca
6900gccctggctg cagctgcagc gcgagccatg cgcccccagt gcaccccggc ccggcccacc
6960gccccggggc cattctgctg accgcccagc cccgagcccc gacagtggca agttgcggct
7020actgcggttg caagctccgg ccaacccgga ggagccccag cggggagcgc agtgttgcgc
7080cccccgcccc cgcgcgcgcc gcagcagccg ggcgttcact catcctccct cccccaccgt
7140ccctcccttt tctcctcaag tcctgaagtt gagtttgaga ggcgacacgg cggcggcggc
7200cgcgctgctc ccgctcctct gcctccccat ggatatgcac tgcaaagcag accccttctc
7260cgcgatgcac cgtgagtacc cgcgcccggc tcctgtcccg gctcgggctc tccgtcccaa
7320ccctgtccag t
733141416PRTHomo sapiens 41Met Asp Met His Cys Lys Ala Asp Pro Phe Ser
Ala Met His Pro Gly1 5 10
15His Gly Gly Val Asn Gln Leu Gly Gly Val Phe Val Asn Gly Arg Pro
20 25 30Leu Pro Asp Val Val Arg Gln
Arg Ile Val Glu Leu Ala His Gln Gly 35 40
45Val Arg Pro Cys Asp Ile Ser Arg Gln Leu Arg Val Ser His Gly
Cys 50 55 60Val Ser Lys Ile Leu Gly
Arg Tyr Tyr Glu Thr Gly Ser Ile Lys Pro65 70
75 80Gly Val Ile Gly Gly Ser Lys Pro Lys Val Ala
Thr Pro Lys Val Val 85 90
95Asp Lys Ile Ala Glu Tyr Lys Arg Gln Asn Pro Thr Met Phe Ala Trp
100 105 110Glu Ile Arg Asp Arg Leu
Leu Ala Glu Gly Ile Cys Asp Asn Asp Thr 115 120
125Val Pro Ser Val Ser Ser Ile Asn Arg Ile Ile Arg Thr Lys
Val Gln 130 135 140Gln Pro Phe His Pro
Thr Pro Asp Gly Ala Gly Thr Gly Val Thr Ala145 150
155 160Pro Gly His Thr Ile Val Pro Ser Thr Ala
Ser Pro Pro Val Ser Ser 165 170
175Ala Ser Asn Asp Pro Val Gly Ser Tyr Ser Ile Asn Gly Ile Leu Gly
180 185 190Ile Pro Arg Ser Asn
Gly Glu Lys Arg Lys Arg Asp Glu Val Glu Val 195
200 205Tyr Thr Asp Pro Ala His Ile Arg Gly Gly Gly Gly
Leu His Leu Val 210 215 220Trp Thr Leu
Arg Asp Val Ser Glu Gly Ser Val Pro Asn Gly Asp Ser225
230 235 240Gln Ser Gly Val Asp Ser Leu
Arg Lys His Leu Arg Ala Asp Thr Phe 245
250 255Thr Gln Gln Gln Leu Glu Ala Leu Asp Arg Val Phe
Glu Arg Pro Ser 260 265 270Tyr
Pro Asp Val Phe Gln Ala Ser Glu His Ile Lys Ser Glu Gln Gly 275
280 285Asn Glu Tyr Ser Leu Pro Ala Leu Thr
Pro Gly Leu Asp Glu Val Lys 290 295
300Ser Ser Leu Ser Ala Ser Thr Asn Pro Glu Leu Gly Ser Asn Val Ser305
310 315 320Gly Thr Gln Thr
Tyr Pro Val Val Thr Gly Arg Asp Met Ala Ser Thr 325
330 335Thr Leu Pro Gly Tyr Pro Pro His Val Pro
Pro Thr Gly Gln Gly Ser 340 345
350Tyr Pro Thr Ser Thr Leu Ala Gly Met Val Pro Gly Ser Glu Phe Ser
355 360 365Gly Asn Pro Tyr Ser His Pro
Gln Tyr Thr Ala Tyr Asn Glu Ala Trp 370 375
380Arg Phe Ser Asn Pro Ala Leu Leu Ser Ser Pro Tyr Tyr Tyr Ser
Ala385 390 395 400Ala Pro
Arg Ser Ala Pro Ala Ala Ala Ala Ala Ala Tyr Asp Arg His
405 410 415424276DNAHomo sapiens 42
aggctccagt ctccggccga gtcttctcgc agccgcaacc cacctggggc cagcccagag
60ctgccagcgc cgctcggctc cctccctccc tcccggccct tcggccgcgg cggcgtgcgc
120ctgccttttc cgggggcggg ggcctggccc gcgcgctccc ctcccgcagg cgccacctcg
180gacatccccg ggattgctac ttctctgcca acttcgccaa ctcgccagca cttggagagg
240cccggctccc ctcccggcgc cctctgaccg cccccgcccc gcgcgctctc cgaccaccgc
300ctctcggatg accaggttcc aggggagctg agcgagtcgc ctcccccgcc cagcttcagc
360cctggctgca gctgcagcgc gagccatgcg cccccagtgc accccggccc ggcccaccgc
420cccggggcca ttctgctgac cgcccagccc cgagccccga cagtggcaag ttgcggctac
480tgcagttgca agctccggcc aacccggagg agccccagcg gggagcgcag tgttgcgccc
540cccgcccccg cgcgccccgc agcagccggg cgttcactca tcctccctcc cccaccgtcc
600ctcccttttc tcctcaagtc ctgaagttga gtttgagagg cgacacggcg gcggcggccg
660cgctgctccc gctcctctgc ctccccatgg atatgcactg caaagcagac cccttctccg
720cgatgcaccc agggcacggg ggtgtgaacc agctcggggg ggtgtttgtg aacggccggc
780ccctacccga cgtggtgagg cagcgcatcg tggagctggc ccaccagggt gtgcggccct
840gtgacatctc ccggcagctg cgggtcagcc acggctgtgt cagcaaaatc ctgggcaggt
900actacgagac cggcagcatc aagccgggtg tgatcggtgg ctccaagccc aaagtggcga
960cgcccaaagt ggtggacaag attgctgaat acaaacgaca gaacccgact atgttcgcct
1020gggagattcg agaccggctc ctggccgagg gcatctgtga caatgacaca gtgcccagcg
1080tctcttccat caacagaatc atccggacca aagttcagca gcctttccac ccaacgccgg
1140atggggctgg gacaggagtg accgcccctg gccacaccat tgttcccagc acggcctccc
1200ctcctgtttc cagcgcctcc aatgacccag tgggatccta ctccatcaat gggatcctgg
1260ggattcctcg ctccaatggt gagaagagga aacgtgatga agttgaggta tacactgatc
1320ctgcccacat tagaggaggt ggaggtttgc atctggtctg gactttaaga gatgtgtctg
1380agggctcagt ccccaatgga gattcccaga gtggtgtgga cagtttgcgg aagcacttgc
1440gagctgacac cttcacccag cagcagctgg aagctttgga tcgggtcttt gagcgtcctt
1500cctaccctga cgtcttccag gcatcagagc acatcaaatc agaacagggg aacgagtact
1560ccctcccagc cctgacccct gggcttgatg aagtcaagtc gagtctatct gcatccacca
1620accctgagct gggcagcaac gtgtcaggca cacagacata cccagttgtg actggtcgtg
1680acatggcgag caccactctg cctggttacc cccctcacgt gccccccact ggccagggaa
1740gctaccccac ctccaccctg gcaggaatgg tgcctgggag cgagttctcc ggcaacccgt
1800acagccaccc ccagtacacg gcctacaacg aggcttggag attcagcaac cccgccttac
1860taagttcccc ttattattat agtgccgccc cccggtccgc ccctgccgct gctgccgctg
1920cctatgaccg ccactagtta ccgcggggac cacatcaagc ttcaggccga cagcttcggc
1980ctccacatcg tccccgtctg accccacccc ggagggaggg aggaccgacg cgacgcgatg
2040cctcccggcc accgccccag cctcacccca tcccacgacc cccgcaaccc ttcacatcac
2100ccccctcgaa ggtcggacag gacgggtgga gccgtgggcg ggaccctcag gcccgggccc
2160gccgccccca gccccgcctg ccgcccctcc ccgcctgcct ggactgcgcg gcgccgtgag
2220ggggattcgg cccagctcgt cccggcctcc accaagccag ccccgaagcc cgccagccac
2280cctgccggac tcgggcgcga cctgctggcg cgcgccggat gtttctgtga cacacaatca
2340gcgcggaccg cagcgcggcc cagccccggg cacccgcctc ggacgctcgg gcgccaggag
2400gcttcgctgg aggggctggg ccaaggagat taagaagaaa acgactttct gcaggaggaa
2460gagcccgctg ccgaatccct gggaaaaatt cttttccccc agtgccagcc ggactgccct
2520cgccttccgg gtgtgccctg tcccagaaga tggaatgggg gtgtgggggt ccggctctag
2580gaacgggctt tgggggcgtc aggtctttcc aaggttggga cccaaggatc ggggggccca
2640gcagcccgca ccgatcgagc cggactctcg gctcttcact gctcctcctg gcctgcctag
2700ttccccaggg cccggcacct cctgctgcga gacccggctc tcagccctgc cttgccccta
2760cctcagcgtc tcttccacct gctggcctcc cagtttcccc tcctgccagt ccttcgcctg
2820tcccttgacg ccctgcatcc tcctccctga ctcgcagccc catcggacgc tctcccggga
2880ccgccgcagg accagtttcc atagactgcg gactggggtc ttcctccagc agttacttga
2940tgccccctcc cccgacacag actctcaatc tgccggtggt aagaaccggt tctgagctgg
3000cgtctgagct gctgcggggt ggaagtgggg ggctgcccac tccactcctc ccatcccctc
3060ccagcctcct cctccggcag gaactgaaca gaaccacaaa aagtctacat ttatttaata
3120tgatggtctt tgcaaaaagg aacaaaacaa cacaaaagcc caccaggctg ctgctttgtg
3180gaaagacggt gtgtgtcgtg tgaaggcgaa acccggtgta cataacccct ccccctccgc
3240cccgccccgc ccggccccgt agagtccctg tcgcccgccg gccctgcctg tagatacgcc
3300ccgctgtctg tgctgtgaga gtcgccgctc gctggggggg aaggggggga cacagctaca
3360cgcccattaa agcacagcac gtcctggggg aggggggcat tttttatgtt acaaaaaaaa
3420attacgaaag aaaagaaatc tctatgcaaa atgacgaaca tggtcctgtg gactcctctg
3480gcctgttttg ttggctcttt ctctgtaatt ccgtgttttc gctttttcct ccctgcccct
3540ctctccctct gcccctctct cctctccgct tctctccccc tctgtctctg tctctctccg
3600tctctgtcgc tcttgtctgt ctgtctctgc tctttcctcg gcctctctcc ccagacctgg
3660cccggccgcc ctgtctccgc aggctagatc cgaggtggca gctccagccc ccgggctcgc
3720cccctcgcgg gcgtgccccg cgcgccccgg gcggccgaag gccgggccgc cccgtcccgc
3780cccgtagttg ctctttcggt agtggcgatg cgccctgcat gtctcctcac ccgtggatcg
3840tgacgactcg aaataacaga aacaaagtca ataaagtgaa aataaataaa aatccttgaa
3900caaatccgaa aaggcttgga gtcctcgccc agatctctct cccctgcgag ccctttttat
3960ttgagaagga aaaagagaaa agagaatcgt ttaagggaac ccggcgccca gccaggctcc
4020agtggcccga acggggcggc gagggcggcg agggcgccga ggtccggccc atcccagtcc
4080tgtggggctg gccgggcaga gaccccggac ccaggcccag gcctaacctg ctaaatgtcc
4140ccggacggtt ctggtctcct cggccacttt cagtgcgtcg gttcgttttg attctttttc
4200ttttgtgcac ataagaaata aataataata ataaataaag aataaaattt tgtatgtcaa
4260aaaaaaaaaa aaaaaa
427643393PRTHomo sapiens 43Met Asp Met His Cys Lys Ala Asp Pro Phe Ser
Ala Met His Pro Gly1 5 10
15His Gly Gly Val Asn Gln Leu Gly Gly Val Phe Val Asn Gly Arg Pro
20 25 30Leu Pro Asp Val Val Arg Gln
Arg Ile Val Glu Leu Ala His Gln Gly 35 40
45Val Arg Pro Cys Asp Ile Ser Arg Gln Leu Arg Val Ser His Gly
Cys 50 55 60Val Ser Lys Ile Leu Gly
Arg Tyr Tyr Glu Thr Gly Ser Ile Lys Pro65 70
75 80Gly Val Ile Gly Gly Ser Lys Pro Lys Val Ala
Thr Pro Lys Val Val 85 90
95Asp Lys Ile Ala Glu Tyr Lys Arg Gln Asn Pro Thr Met Phe Ala Trp
100 105 110Glu Ile Arg Asp Arg Leu
Leu Ala Glu Gly Ile Cys Asp Asn Asp Thr 115 120
125Val Pro Ser Val Ser Ser Ile Asn Arg Ile Ile Arg Thr Lys
Val Gln 130 135 140Gln Pro Phe His Pro
Thr Pro Asp Gly Ala Gly Thr Gly Val Thr Ala145 150
155 160Pro Gly His Thr Ile Val Pro Ser Thr Ala
Ser Pro Pro Val Ser Ser 165 170
175Ala Ser Asn Asp Pro Val Gly Ser Tyr Ser Ile Asn Gly Ile Leu Gly
180 185 190Ile Pro Arg Ser Asn
Gly Glu Lys Arg Lys Arg Asp Glu Asp Val Ser 195
200 205Glu Gly Ser Val Pro Asn Gly Asp Ser Gln Ser Gly
Val Asp Ser Leu 210 215 220Arg Lys His
Leu Arg Ala Asp Thr Phe Thr Gln Gln Gln Leu Glu Ala225
230 235 240Leu Asp Arg Val Phe Glu Arg
Pro Ser Tyr Pro Asp Val Phe Gln Ala 245
250 255Ser Glu His Ile Lys Ser Glu Gln Gly Asn Glu Tyr
Ser Leu Pro Ala 260 265 270Leu
Thr Pro Gly Leu Asp Glu Val Lys Ser Ser Leu Ser Ala Ser Thr 275
280 285Asn Pro Glu Leu Gly Ser Asn Val Ser
Gly Thr Gln Thr Tyr Pro Val 290 295
300Val Thr Gly Arg Asp Met Ala Ser Thr Thr Leu Pro Gly Tyr Pro Pro305
310 315 320His Val Pro Pro
Thr Gly Gln Gly Ser Tyr Pro Thr Ser Thr Leu Ala 325
330 335Gly Met Val Pro Gly Ser Glu Phe Ser Gly
Asn Pro Tyr Ser His Pro 340 345
350Gln Tyr Thr Ala Tyr Asn Glu Ala Trp Arg Phe Ser Asn Pro Ala Leu
355 360 365Leu Ser Ser Pro Tyr Tyr Tyr
Ser Ala Ala Pro Arg Ser Ala Pro Ala 370 375
380Ala Ala Ala Ala Ala Tyr Asp Arg His385
390444207DNAHomo sapiens 44aggctccagt ctccggccga gtcttctcgc agccgcaacc
cacctggggc cagcccagag 60ctgccagcgc cgctcggctc cctccctccc tcccggccct
tcggccgcgg cggcgtgcgc 120ctgccttttc cgggggcggg ggcctggccc gcgcgctccc
ctcccgcagg cgccacctcg 180gacatccccg ggattgctac ttctctgcca acttcgccaa
ctcgccagca cttggagagg 240cccggctccc ctcccggcgc cctctgaccg cccccgcccc
gcgcgctctc cgaccaccgc 300ctctcggatg accaggttcc aggggagctg agcgagtcgc
ctcccccgcc cagcttcagc 360cctggctgca gctgcagcgc gagccatgcg cccccagtgc
accccggccc ggcccaccgc 420cccggggcca ttctgctgac cgcccagccc cgagccccga
cagtggcaag ttgcggctac 480tgcagttgca agctccggcc aacccggagg agccccagcg
gggagcgcag tgttgcgccc 540cccgcccccg cgcgccccgc agcagccggg cgttcactca
tcctccctcc cccaccgtcc 600ctcccttttc tcctcaagtc ctgaagttga gtttgagagg
cgacacggcg gcggcggccg 660cgctgctccc gctcctctgc ctccccatgg atatgcactg
caaagcagac cccttctccg 720cgatgcaccc agggcacggg ggtgtgaacc agctcggggg
ggtgtttgtg aacggccggc 780ccctacccga cgtggtgagg cagcgcatcg tggagctggc
ccaccagggt gtgcggccct 840gtgacatctc ccggcagctg cgggtcagcc acggctgtgt
cagcaaaatc ctgggcaggt 900actacgagac cggcagcatc aagccgggtg tgatcggtgg
ctccaagccc aaagtggcga 960cgcccaaagt ggtggacaag attgctgaat acaaacgaca
gaacccgact atgttcgcct 1020gggagattcg agaccggctc ctggccgagg gcatctgtga
caatgacaca gtgcccagcg 1080tctcttccat caacagaatc atccggacca aagttcagca
gcctttccac ccaacgccgg 1140atggggctgg gacaggagtg accgcccctg gccacaccat
tgttcccagc acggcctccc 1200ctcctgtttc cagcgcctcc aatgacccag tgggatccta
ctccatcaat gggatcctgg 1260ggattcctcg ctccaatggt gagaagagga aacgtgatga
agatgtgtct gagggctcag 1320tccccaatgg agattcccag agtggtgtgg acagtttgcg
gaagcacttg cgagctgaca 1380ccttcaccca gcagcagctg gaagctttgg atcgggtctt
tgagcgtcct tcctaccctg 1440acgtcttcca ggcatcagag cacatcaaat cagaacaggg
gaacgagtac tccctcccag 1500ccctgacccc tgggcttgat gaagtcaagt cgagtctatc
tgcatccacc aaccctgagc 1560tgggcagcaa cgtgtcaggc acacagacat acccagttgt
gactggtcgt gacatggcga 1620gcaccactct gcctggttac ccccctcacg tgccccccac
tggccaggga agctacccca 1680cctccaccct ggcaggaatg gtgcctggga gcgagttctc
cggcaacccg tacagccacc 1740cccagtacac ggcctacaac gaggcttgga gattcagcaa
ccccgcctta ctaagttccc 1800cttattatta tagtgccgcc ccccggtccg cccctgccgc
tgctgccgct gcctatgacc 1860gccactagtt accgcgggga ccacatcaag cttcaggccg
acagcttcgg cctccacatc 1920gtccccgtct gaccccaccc cggagggagg gaggaccgac
gcgacgcgat gcctcccggc 1980caccgcccca gcctcacccc atcccacgac ccccgcaacc
cttcacatca cccccctcga 2040aggtcggaca ggacgggtgg agccgtgggc gggaccctca
ggcccgggcc cgccgccccc 2100agccccgcct gccgcccctc cccgcctgcc tggactgcgc
ggcgccgtga gggggattcg 2160gcccagctcg tcccggcctc caccaagcca gccccgaagc
ccgccagcca ccctgccgga 2220ctcgggcgcg acctgctggc gcgcgccgga tgtttctgtg
acacacaatc agcgcggacc 2280gcagcgcggc ccagccccgg gcacccgcct cggacgctcg
ggcgccagga ggcttcgctg 2340gaggggctgg gccaaggaga ttaagaagaa aacgactttc
tgcaggagga agagcccgct 2400gccgaatccc tgggaaaaat tcttttcccc cagtgccagc
cggactgccc tcgccttccg 2460ggtgtgccct gtcccagaag atggaatggg ggtgtggggg
tccggctcta ggaacgggct 2520ttgggggcgt caggtctttc caaggttggg acccaaggat
cggggggccc agcagcccgc 2580accgatcgag ccggactctc ggctcttcac tgctcctcct
ggcctgccta gttccccagg 2640gcccggcacc tcctgctgcg agacccggct ctcagccctg
ccttgcccct acctcagcgt 2700ctcttccacc tgctggcctc ccagtttccc ctcctgccag
tccttcgcct gtcccttgac 2760gccctgcatc ctcctccctg actcgcagcc ccatcggacg
ctctcccggg accgccgcag 2820gaccagtttc catagactgc ggactggggt cttcctccag
cagttacttg atgccccctc 2880ccccgacaca gactctcaat ctgccggtgg taagaaccgg
ttctgagctg gcgtctgagc 2940tgctgcgggg tggaagtggg gggctgccca ctccactcct
cccatcccct cccagcctcc 3000tcctccggca ggaactgaac agaaccacaa aaagtctaca
tttatttaat atgatggtct 3060ttgcaaaaag gaacaaaaca acacaaaagc ccaccaggct
gctgctttgt ggaaagacgg 3120tgtgtgtcgt gtgaaggcga aacccggtgt acataacccc
tccccctccg ccccgccccg 3180cccggccccg tagagtccct gtcgcccgcc ggccctgcct
gtagatacgc cccgctgtct 3240gtgctgtgag agtcgccgct cgctgggggg gaaggggggg
acacagctac acgcccatta 3300aagcacagca cgtcctgggg gaggggggca ttttttatgt
tacaaaaaaa aattacgaaa 3360gaaaagaaat ctctatgcaa aatgacgaac atggtcctgt
ggactcctct ggcctgtttt 3420gttggctctt tctctgtaat tccgtgtttt cgctttttcc
tccctgcccc tctctccctc 3480tgcccctctc tcctctccgc ttctctcccc ctctgtctct
gtctctctcc gtctctgtcg 3540ctcttgtctg tctgtctctg ctctttcctc ggcctctctc
cccagacctg gcccggccgc 3600cctgtctccg caggctagat ccgaggtggc agctccagcc
cccgggctcg ccccctcgcg 3660ggcgtgcccc gcgcgccccg ggcggccgaa ggccgggccg
ccccgtcccg ccccgtagtt 3720gctctttcgg tagtggcgat gcgccctgca tgtctcctca
cccgtggatc gtgacgactc 3780gaaataacag aaacaaagtc aataaagtga aaataaataa
aaatccttga acaaatccga 3840aaaggcttgg agtcctcgcc cagatctctc tcccctgcga
gcccttttta tttgagaagg 3900aaaaagagaa aagagaatcg tttaagggaa cccggcgccc
agccaggctc cagtggcccg 3960aacggggcgg cgagggcggc gagggcgccg aggtccggcc
catcccagtc ctgtggggct 4020ggccgggcag agaccccgga cccaggccca ggcctaacct
gctaaatgtc cccggacggt 4080tctggtctcc tcggccactt tcagtgcgtc ggttcgtttt
gattcttttt cttttgtgca 4140cataagaaat aaataataat aataaataaa gaataaaatt
ttgtatgtca aaaaaaaaaa 4200aaaaaaa
420745396PRTHomo sapiens 45Met Asp Met His Cys Lys
Ala Asp Pro Phe Ser Ala Met His Pro Gly1 5
10 15His Gly Gly Val Asn Gln Leu Gly Gly Val Phe Val
Asn Gly Arg Pro 20 25 30Leu
Pro Asp Val Val Arg Gln Arg Ile Val Glu Leu Ala His Gln Gly 35
40 45Val Arg Pro Cys Asp Ile Ser Arg Gln
Leu Arg Val Ser His Gly Cys 50 55
60Val Ser Lys Ile Leu Gly Arg Tyr Tyr Glu Thr Gly Ser Ile Lys Pro65
70 75 80Gly Val Ile Gly Gly
Ser Lys Pro Lys Val Ala Thr Pro Lys Val Val 85
90 95Asp Lys Ile Ala Glu Tyr Lys Arg Gln Asn Pro
Thr Met Phe Ala Trp 100 105
110Glu Ile Arg Asp Arg Leu Leu Ala Glu Gly Ile Cys Asp Asn Asp Thr
115 120 125Val Pro Ser Val Ser Ser Ile
Asn Arg Ile Ile Arg Thr Lys Val Gln 130 135
140Gln Pro Phe His Pro Thr Pro Asp Gly Ala Gly Thr Gly Val Thr
Ala145 150 155 160Pro Gly
His Thr Ile Val Pro Ser Thr Ala Ser Pro Pro Val Ser Ser
165 170 175Ala Ser Asn Asp Pro Val Gly
Ser Tyr Ser Ile Asn Gly Ile Leu Gly 180 185
190Ile Pro Arg Ser Asn Gly Glu Lys Arg Lys Arg Asp Glu Asp
Val Ser 195 200 205Glu Gly Ser Val
Pro Asn Gly Asp Ser Gln Ser Gly Val Asp Ser Leu 210
215 220Arg Lys His Leu Arg Ala Asp Thr Phe Thr Gln Gln
Gln Leu Glu Ala225 230 235
240Leu Asp Arg Val Phe Glu Arg Pro Ser Tyr Pro Asp Val Phe Gln Ala
245 250 255Ser Glu His Ile Lys
Ser Glu Gln Gly Asn Glu Tyr Ser Leu Pro Ala 260
265 270Leu Thr Pro Gly Leu Asp Glu Val Lys Ser Ser Leu
Ser Ala Ser Thr 275 280 285Asn Pro
Glu Leu Gly Ser Asn Val Ser Gly Thr Gln Thr Tyr Pro Val 290
295 300Val Thr Gly Arg Asp Met Ala Ser Thr Thr Leu
Pro Gly Tyr Pro Pro305 310 315
320His Val Pro Pro Thr Gly Gln Gly Ser Tyr Pro Thr Ser Thr Leu Ala
325 330 335Gly Met Val Pro
Glu Ala Ala Val Gly Pro Ser Ser Ser Leu Met Ser 340
345 350Lys Pro Gly Arg Lys Leu Ala Glu Val Pro Pro
Cys Val Gln Pro Thr 355 360 365Gly
Ala Ser Ser Pro Ala Thr Arg Thr Ala Thr Pro Ser Thr Arg Pro 370
375 380Thr Thr Arg Leu Gly Asp Ser Ala Thr Pro
Pro Tyr385 390 395464290DNAHomo sapiens
46aggctccagt ctccggccga gtcttctcgc agccgcaacc cacctggggc cagcccagag
60ctgccagcgc cgctcggctc cctccctccc tcccggccct tcggccgcgg cggcgtgcgc
120ctgccttttc cgggggcggg ggcctggccc gcgcgctccc ctcccgcagg cgccacctcg
180gacatccccg ggattgctac ttctctgcca acttcgccaa ctcgccagca cttggagagg
240cccggctccc ctcccggcgc cctctgaccg cccccgcccc gcgcgctctc cgaccaccgc
300ctctcggatg accaggttcc aggggagctg agcgagtcgc ctcccccgcc cagcttcagc
360cctggctgca gctgcagcgc gagccatgcg cccccagtgc accccggccc ggcccaccgc
420cccggggcca ttctgctgac cgcccagccc cgagccccga cagtggcaag ttgcggctac
480tgcagttgca agctccggcc aacccggagg agccccagcg gggagcgcag tgttgcgccc
540cccgcccccg cgcgccccgc agcagccggg cgttcactca tcctccctcc cccaccgtcc
600ctcccttttc tcctcaagtc ctgaagttga gtttgagagg cgacacggcg gcggcggccg
660cgctgctccc gctcctctgc ctccccatgg atatgcactg caaagcagac cccttctccg
720cgatgcaccc agggcacggg ggtgtgaacc agctcggggg ggtgtttgtg aacggccggc
780ccctacccga cgtggtgagg cagcgcatcg tggagctggc ccaccagggt gtgcggccct
840gtgacatctc ccggcagctg cgggtcagcc acggctgtgt cagcaaaatc ctgggcaggt
900actacgagac cggcagcatc aagccgggtg tgatcggtgg ctccaagccc aaagtggcga
960cgcccaaagt ggtggacaag attgctgaat acaaacgaca gaacccgact atgttcgcct
1020gggagattcg agaccggctc ctggccgagg gcatctgtga caatgacaca gtgcccagcg
1080tctcttccat caacagaatc atccggacca aagttcagca gcctttccac ccaacgccgg
1140atggggctgg gacaggagtg accgcccctg gccacaccat tgttcccagc acggcctccc
1200ctcctgtttc cagcgcctcc aatgacccag tgggatccta ctccatcaat gggatcctgg
1260ggattcctcg ctccaatggt gagaagagga aacgtgatga agatgtgtct gagggctcag
1320tccccaatgg agattcccag agtggtgtgg acagtttgcg gaagcacttg cgagctgaca
1380ccttcaccca gcagcagctg gaagctttgg atcgggtctt tgagcgtcct tcctaccctg
1440acgtcttcca ggcatcagag cacatcaaat cagaacaggg gaacgagtac tccctcccag
1500ccctgacccc tgggcttgat gaagtcaagt cgagtctatc tgcatccacc aaccctgagc
1560tgggcagcaa cgtgtcaggc acacagacat acccagttgt gactggtcgt gacatggcga
1620gcaccactct gcctggttac ccccctcacg tgccccccac tggccaggga agctacccca
1680cctccaccct ggcaggaatg gtgcctgagg ctgcagttgg tccctcatcc tccctcatga
1740gcaagccggg gaggaagctt gcagaagtgc ccccttgtgt gcaacccact ggagcgagtt
1800ctccggcaac ccgtacagcc acccccagta cacggcctac aacgaggctt ggagattcag
1860caaccccgcc ttactaagtt ccccttatta ttatagtgcc gccccccggt ccgcccctgc
1920cgctgctgcc gctgcctatg accgccacta gttaccgcgg ggaccacatc aagcttcagg
1980ccgacagctt cggcctccac atcgtccccg tctgacccca ccccggaggg agggaggacc
2040gacgcgacgc gatgcctccc ggccaccgcc ccagcctcac cccatcccac gacccccgca
2100acccttcaca tcacccccct cgaaggtcgg acaggacggg tggagccgtg ggcgggaccc
2160tcaggcccgg gcccgccgcc cccagccccg cctgccgccc ctccccgcct gcctggactg
2220cgcggcgccg tgagggggat tcggcccagc tcgtcccggc ctccaccaag ccagccccga
2280agcccgccag ccaccctgcc ggactcgggc gcgacctgct ggcgcgcgcc ggatgtttct
2340gtgacacaca atcagcgcgg accgcagcgc ggcccagccc cgggcacccg cctcggacgc
2400tcgggcgcca ggaggcttcg ctggaggggc tgggccaagg agattaagaa gaaaacgact
2460ttctgcagga ggaagagccc gctgccgaat ccctgggaaa aattcttttc ccccagtgcc
2520agccggactg ccctcgcctt ccgggtgtgc cctgtcccag aagatggaat gggggtgtgg
2580gggtccggct ctaggaacgg gctttggggg cgtcaggtct ttccaaggtt gggacccaag
2640gatcgggggg cccagcagcc cgcaccgatc gagccggact ctcggctctt cactgctcct
2700cctggcctgc ctagttcccc agggcccggc acctcctgct gcgagacccg gctctcagcc
2760ctgccttgcc cctacctcag cgtctcttcc acctgctggc ctcccagttt cccctcctgc
2820cagtccttcg cctgtccctt gacgccctgc atcctcctcc ctgactcgca gccccatcgg
2880acgctctccc gggaccgccg caggaccagt ttccatagac tgcggactgg ggtcttcctc
2940cagcagttac ttgatgcccc ctcccccgac acagactctc aatctgccgg tggtaagaac
3000cggttctgag ctggcgtctg agctgctgcg gggtggaagt ggggggctgc ccactccact
3060cctcccatcc cctcccagcc tcctcctccg gcaggaactg aacagaacca caaaaagtct
3120acatttattt aatatgatgg tctttgcaaa aaggaacaaa acaacacaaa agcccaccag
3180gctgctgctt tgtggaaaga cggtgtgtgt cgtgtgaagg cgaaacccgg tgtacataac
3240ccctccccct ccgccccgcc ccgcccggcc ccgtagagtc cctgtcgccc gccggccctg
3300cctgtagata cgccccgctg tctgtgctgt gagagtcgcc gctcgctggg ggggaagggg
3360gggacacagc tacacgccca ttaaagcaca gcacgtcctg ggggaggggg gcatttttta
3420tgttacaaaa aaaaattacg aaagaaaaga aatctctatg caaaatgacg aacatggtcc
3480tgtggactcc tctggcctgt tttgttggct ctttctctgt aattccgtgt tttcgctttt
3540tcctccctgc ccctctctcc ctctgcccct ctctcctctc cgcttctctc cccctctgtc
3600tctgtctctc tccgtctctg tcgctcttgt ctgtctgtct ctgctctttc ctcggcctct
3660ctccccagac ctggcccggc cgccctgtct ccgcaggcta gatccgaggt ggcagctcca
3720gcccccgggc tcgccccctc gcgggcgtgc cccgcgcgcc ccgggcggcc gaaggccggg
3780ccgccccgtc ccgccccgta gttgctcttt cggtagtggc gatgcgccct gcatgtctcc
3840tcacccgtgg atcgtgacga ctcgaaataa cagaaacaaa gtcaataaag tgaaaataaa
3900taaaaatcct tgaacaaatc cgaaaaggct tggagtcctc gcccagatct ctctcccctg
3960cgagcccttt ttatttgaga aggaaaaaga gaaaagagaa tcgtttaagg gaacccggcg
4020cccagccagg ctccagtggc ccgaacgggg cggcgagggc ggcgagggcg ccgaggtccg
4080gcccatccca gtcctgtggg gctggccggg cagagacccc ggacccaggc ccaggcctaa
4140cctgctaaat gtccccggac ggttctggtc tcctcggcca ctttcagtgc gtcggttcgt
4200tttgattctt tttcttttgt gcacataaga aataaataat aataataaat aaagaataaa
4260attttgtatg tcaaaaaaaa aaaaaaaaaa
429047408PRTHomo sapiens 47Met Asp Met His Cys Lys Ala Asp Pro Phe Ser
Ala Met His Pro Gly1 5 10
15His Gly Gly Val Asn Gln Leu Gly Gly Val Phe Val Asn Gly Arg Pro
20 25 30Leu Pro Asp Val Val Arg Gln
Arg Ile Val Glu Leu Ala His Gln Gly 35 40
45Val Arg Pro Cys Asp Ile Ser Arg Gln Leu Arg Val Ser His Gly
Cys 50 55 60Val Ser Lys Ile Leu Gly
Arg Tyr Tyr Glu Thr Gly Ser Ile Lys Pro65 70
75 80Gly Val Ile Gly Gly Ser Lys Pro Lys Val Ala
Thr Pro Lys Val Val 85 90
95Asp Lys Ile Ala Glu Tyr Lys Arg Gln Asn Pro Thr Met Phe Ala Trp
100 105 110Glu Ile Arg Asp Arg Leu
Leu Ala Glu Gly Ile Cys Asp Asn Asp Thr 115 120
125Val Pro Ser Val Ser Ser Ile Asn Arg Ile Ile Arg Thr Lys
Val Gln 130 135 140Gln Pro Phe His Pro
Thr Pro Asp Gly Ala Gly Thr Gly Val Thr Ala145 150
155 160Pro Gly His Thr Ile Val Pro Ser Thr Ala
Ser Pro Pro Val Ser Ser 165 170
175Ala Ser Asn Asp Pro Val Gly Ser Tyr Ser Ile Asn Gly Ile Leu Gly
180 185 190Ile Pro Arg Ser Asn
Gly Glu Lys Arg Lys Arg Asp Glu Asp Val Ser 195
200 205Glu Gly Ser Val Pro Asn Gly Asp Ser Gln Ser Gly
Val Asp Ser Leu 210 215 220Arg Lys His
Leu Arg Ala Asp Thr Phe Thr Gln Gln Gln Leu Glu Ala225
230 235 240Leu Asp Arg Val Phe Glu Arg
Pro Ser Tyr Pro Asp Val Phe Gln Ala 245
250 255Ser Glu His Ile Lys Ser Glu Gln Gly Asn Glu Tyr
Ser Leu Pro Ala 260 265 270Leu
Thr Pro Gly Leu Asp Glu Val Lys Ser Ser Leu Ser Ala Ser Thr 275
280 285Asn Pro Glu Leu Gly Ser Asn Val Ser
Gly Thr Gln Thr Tyr Pro Val 290 295
300Val Thr Gly Arg Asp Met Ala Ser Thr Thr Leu Pro Gly Tyr Pro Pro305
310 315 320His Val Pro Pro
Thr Gly Gln Gly Ser Tyr Pro Thr Ser Thr Leu Ala 325
330 335Gly Met Val Pro Gly Ser Glu Phe Ser Gly
Asn Pro Tyr Ser His Pro 340 345
350Gln Tyr Thr Ala Tyr Asn Glu Ala Trp Arg Phe Ser Asn Pro Ala Leu
355 360 365Leu Met Pro Pro Pro Gly Pro
Pro Leu Pro Leu Leu Pro Leu Pro Met 370 375
380Thr Ala Thr Ser Tyr Arg Gly Asp His Ile Lys Leu Gln Ala Asp
Ser385 390 395 400Phe Gly
Leu His Ile Val Pro Val 405484188DNAHomo sapiens
48aggctccagt ctccggccga gtcttctcgc agccgcaacc cacctggggc cagcccagag
60ctgccagcgc cgctcggctc cctccctccc tcccggccct tcggccgcgg cggcgtgcgc
120ctgccttttc cgggggcggg ggcctggccc gcgcgctccc ctcccgcagg cgccacctcg
180gacatccccg ggattgctac ttctctgcca acttcgccaa ctcgccagca cttggagagg
240cccggctccc ctcccggcgc cctctgaccg cccccgcccc gcgcgctctc cgaccaccgc
300ctctcggatg accaggttcc aggggagctg agcgagtcgc ctcccccgcc cagcttcagc
360cctggctgca gctgcagcgc gagccatgcg cccccagtgc accccggccc ggcccaccgc
420cccggggcca ttctgctgac cgcccagccc cgagccccga cagtggcaag ttgcggctac
480tgcagttgca agctccggcc aacccggagg agccccagcg gggagcgcag tgttgcgccc
540cccgcccccg cgcgccccgc agcagccggg cgttcactca tcctccctcc cccaccgtcc
600ctcccttttc tcctcaagtc ctgaagttga gtttgagagg cgacacggcg gcggcggccg
660cgctgctccc gctcctctgc ctccccatgg atatgcactg caaagcagac cccttctccg
720cgatgcaccc agggcacggg ggtgtgaacc agctcggggg ggtgtttgtg aacggccggc
780ccctacccga cgtggtgagg cagcgcatcg tggagctggc ccaccagggt gtgcggccct
840gtgacatctc ccggcagctg cgggtcagcc acggctgtgt cagcaaaatc ctgggcaggt
900actacgagac cggcagcatc aagccgggtg tgatcggtgg ctccaagccc aaagtggcga
960cgcccaaagt ggtggacaag attgctgaat acaaacgaca gaacccgact atgttcgcct
1020gggagattcg agaccggctc ctggccgagg gcatctgtga caatgacaca gtgcccagcg
1080tctcttccat caacagaatc atccggacca aagttcagca gcctttccac ccaacgccgg
1140atggggctgg gacaggagtg accgcccctg gccacaccat tgttcccagc acggcctccc
1200ctcctgtttc cagcgcctcc aatgacccag tgggatccta ctccatcaat gggatcctgg
1260ggattcctcg ctccaatggt gagaagagga aacgtgatga agatgtgtct gagggctcag
1320tccccaatgg agattcccag agtggtgtgg acagtttgcg gaagcacttg cgagctgaca
1380ccttcaccca gcagcagctg gaagctttgg atcgggtctt tgagcgtcct tcctaccctg
1440acgtcttcca ggcatcagag cacatcaaat cagaacaggg gaacgagtac tccctcccag
1500ccctgacccc tgggcttgat gaagtcaagt cgagtctatc tgcatccacc aaccctgagc
1560tgggcagcaa cgtgtcaggc acacagacat acccagttgt gactggtcgt gacatggcga
1620gcaccactct gcctggttac ccccctcacg tgccccccac tggccaggga agctacccca
1680cctccaccct ggcaggaatg gtgcctggga gcgagttctc cggcaacccg tacagccacc
1740cccagtacac ggcctacaac gaggcttgga gattcagcaa ccccgcctta ctaatgccgc
1800cccccggtcc gcccctgccg ctgctgccgc tgcctatgac cgccactagt taccgcgggg
1860accacatcaa gcttcaggcc gacagcttcg gcctccacat cgtccccgtc tgaccccacc
1920ccggagggag ggaggaccga cgcgacgcga tgcctcccgg ccaccgcccc agcctcaccc
1980catcccacga cccccgcaac ccttcacatc acccccctcg aaggtcggac aggacgggtg
2040gagccgtggg cgggaccctc aggcccgggc ccgccgcccc cagccccgcc tgccgcccct
2100ccccgcctgc ctggactgcg cggcgccgtg agggggattc ggcccagctc gtcccggcct
2160ccaccaagcc agccccgaag cccgccagcc accctgccgg actcgggcgc gacctgctgg
2220cgcgcgccgg atgtttctgt gacacacaat cagcgcggac cgcagcgcgg cccagccccg
2280ggcacccgcc tcggacgctc gggcgccagg aggcttcgct ggaggggctg ggccaaggag
2340attaagaaga aaacgacttt ctgcaggagg aagagcccgc tgccgaatcc ctgggaaaaa
2400ttcttttccc ccagtgccag ccggactgcc ctcgccttcc gggtgtgccc tgtcccagaa
2460gatggaatgg gggtgtgggg gtccggctct aggaacgggc tttgggggcg tcaggtcttt
2520ccaaggttgg gacccaagga tcggggggcc cagcagcccg caccgatcga gccggactct
2580cggctcttca ctgctcctcc tggcctgcct agttccccag ggcccggcac ctcctgctgc
2640gagacccggc tctcagccct gccttgcccc tacctcagcg tctcttccac ctgctggcct
2700cccagtttcc cctcctgcca gtccttcgcc tgtcccttga cgccctgcat cctcctccct
2760gactcgcagc cccatcggac gctctcccgg gaccgccgca ggaccagttt ccatagactg
2820cggactgggg tcttcctcca gcagttactt gatgccccct cccccgacac agactctcaa
2880tctgccggtg gtaagaaccg gttctgagct ggcgtctgag ctgctgcggg gtggaagtgg
2940ggggctgccc actccactcc tcccatcccc tcccagcctc ctcctccggc aggaactgaa
3000cagaaccaca aaaagtctac atttatttaa tatgatggtc tttgcaaaaa ggaacaaaac
3060aacacaaaag cccaccaggc tgctgctttg tggaaagacg gtgtgtgtcg tgtgaaggcg
3120aaacccggtg tacataaccc ctccccctcc gccccgcccc gcccggcccc gtagagtccc
3180tgtcgcccgc cggccctgcc tgtagatacg ccccgctgtc tgtgctgtga gagtcgccgc
3240tcgctggggg ggaagggggg gacacagcta cacgcccatt aaagcacagc acgtcctggg
3300ggaggggggc attttttatg ttacaaaaaa aaattacgaa agaaaagaaa tctctatgca
3360aaatgacgaa catggtcctg tggactcctc tggcctgttt tgttggctct ttctctgtaa
3420ttccgtgttt tcgctttttc ctccctgccc ctctctccct ctgcccctct ctcctctccg
3480cttctctccc cctctgtctc tgtctctctc cgtctctgtc gctcttgtct gtctgtctct
3540gctctttcct cggcctctct ccccagacct ggcccggccg ccctgtctcc gcaggctaga
3600tccgaggtgg cagctccagc ccccgggctc gccccctcgc gggcgtgccc cgcgcgcccc
3660gggcggccga aggccgggcc gccccgtccc gccccgtagt tgctctttcg gtagtggcga
3720tgcgccctgc atgtctcctc acccgtggat cgtgacgact cgaaataaca gaaacaaagt
3780caataaagtg aaaataaata aaaatccttg aacaaatccg aaaaggcttg gagtcctcgc
3840ccagatctct ctcccctgcg agcccttttt atttgagaag gaaaaagaga aaagagaatc
3900gtttaaggga acccggcgcc cagccaggct ccagtggccc gaacggggcg gcgagggcgg
3960cgagggcgcc gaggtccggc ccatcccagt cctgtggggc tggccgggca gagaccccgg
4020acccaggccc aggcctaacc tgctaaatgt ccccggacgg ttctggtctc ctcggccact
4080ttcagtgcgt cggttcgttt tgattctttt tcttttgtgc acataagaaa taaataataa
4140taataaataa agaataaaat tttgtatgtc aaaaaaaaaa aaaaaaaa
418849431PRTHomo sapiens 49Met Asp Met His Cys Lys Ala Asp Pro Phe Ser
Ala Met His Pro Gly1 5 10
15His Gly Gly Val Asn Gln Leu Gly Gly Val Phe Val Asn Gly Arg Pro
20 25 30Leu Pro Asp Val Val Arg Gln
Arg Ile Val Glu Leu Ala His Gln Gly 35 40
45Val Arg Pro Cys Asp Ile Ser Arg Gln Leu Arg Val Ser His Gly
Cys 50 55 60Val Ser Lys Ile Leu Gly
Arg Tyr Tyr Glu Thr Gly Ser Ile Lys Pro65 70
75 80Gly Val Ile Gly Gly Ser Lys Pro Lys Val Ala
Thr Pro Lys Val Val 85 90
95Asp Lys Ile Ala Glu Tyr Lys Arg Gln Asn Pro Thr Met Phe Ala Trp
100 105 110Glu Ile Arg Asp Arg Leu
Leu Ala Glu Gly Ile Cys Asp Asn Asp Thr 115 120
125Val Pro Ser Val Ser Ser Ile Asn Arg Ile Ile Arg Thr Lys
Val Gln 130 135 140Gln Pro Phe His Pro
Thr Pro Asp Gly Ala Gly Thr Gly Val Thr Ala145 150
155 160Pro Gly His Thr Ile Val Pro Ser Thr Ala
Ser Pro Pro Val Ser Ser 165 170
175Ala Ser Asn Asp Pro Val Gly Ser Tyr Ser Ile Asn Gly Ile Leu Gly
180 185 190Ile Pro Arg Ser Asn
Gly Glu Lys Arg Lys Arg Asp Glu Val Glu Val 195
200 205Tyr Thr Asp Pro Ala His Ile Arg Gly Gly Gly Gly
Leu His Leu Val 210 215 220Trp Thr Leu
Arg Asp Val Ser Glu Gly Ser Val Pro Asn Gly Asp Ser225
230 235 240Gln Ser Gly Val Asp Ser Leu
Arg Lys His Leu Arg Ala Asp Thr Phe 245
250 255Thr Gln Gln Gln Leu Glu Ala Leu Asp Arg Val Phe
Glu Arg Pro Ser 260 265 270Tyr
Pro Asp Val Phe Gln Ala Ser Glu His Ile Lys Ser Glu Gln Gly 275
280 285Asn Glu Tyr Ser Leu Pro Ala Leu Thr
Pro Gly Leu Asp Glu Val Lys 290 295
300Ser Ser Leu Ser Ala Ser Thr Asn Pro Glu Leu Gly Ser Asn Val Ser305
310 315 320Gly Thr Gln Thr
Tyr Pro Val Val Thr Gly Arg Asp Met Ala Ser Thr 325
330 335Thr Leu Pro Gly Tyr Pro Pro His Val Pro
Pro Thr Gly Gln Gly Ser 340 345
350Tyr Pro Thr Ser Thr Leu Ala Gly Met Val Pro Gly Ser Glu Phe Ser
355 360 365Gly Asn Pro Tyr Ser His Pro
Gln Tyr Thr Ala Tyr Asn Glu Ala Trp 370 375
380Arg Phe Ser Asn Pro Ala Leu Leu Met Pro Pro Pro Gly Pro Pro
Leu385 390 395 400Pro Leu
Leu Pro Leu Pro Met Thr Ala Thr Ser Tyr Arg Gly Asp His
405 410 415Ile Lys Leu Gln Ala Asp Ser
Phe Gly Leu His Ile Val Pro Val 420 425
430504257DNAHomo sapiens 50aggctccagt ctccggccga gtcttctcgc
agccgcaacc cacctggggc cagcccagag 60ctgccagcgc cgctcggctc cctccctccc
tcccggccct tcggccgcgg cggcgtgcgc 120ctgccttttc cgggggcggg ggcctggccc
gcgcgctccc ctcccgcagg cgccacctcg 180gacatccccg ggattgctac ttctctgcca
acttcgccaa ctcgccagca cttggagagg 240cccggctccc ctcccggcgc cctctgaccg
cccccgcccc gcgcgctctc cgaccaccgc 300ctctcggatg accaggttcc aggggagctg
agcgagtcgc ctcccccgcc cagcttcagc 360cctggctgca gctgcagcgc gagccatgcg
cccccagtgc accccggccc ggcccaccgc 420cccggggcca ttctgctgac cgcccagccc
cgagccccga cagtggcaag ttgcggctac 480tgcagttgca agctccggcc aacccggagg
agccccagcg gggagcgcag tgttgcgccc 540cccgcccccg cgcgccccgc agcagccggg
cgttcactca tcctccctcc cccaccgtcc 600ctcccttttc tcctcaagtc ctgaagttga
gtttgagagg cgacacggcg gcggcggccg 660cgctgctccc gctcctctgc ctccccatgg
atatgcactg caaagcagac cccttctccg 720cgatgcaccc agggcacggg ggtgtgaacc
agctcggggg ggtgtttgtg aacggccggc 780ccctacccga cgtggtgagg cagcgcatcg
tggagctggc ccaccagggt gtgcggccct 840gtgacatctc ccggcagctg cgggtcagcc
acggctgtgt cagcaaaatc ctgggcaggt 900actacgagac cggcagcatc aagccgggtg
tgatcggtgg ctccaagccc aaagtggcga 960cgcccaaagt ggtggacaag attgctgaat
acaaacgaca gaacccgact atgttcgcct 1020gggagattcg agaccggctc ctggccgagg
gcatctgtga caatgacaca gtgcccagcg 1080tctcttccat caacagaatc atccggacca
aagttcagca gcctttccac ccaacgccgg 1140atggggctgg gacaggagtg accgcccctg
gccacaccat tgttcccagc acggcctccc 1200ctcctgtttc cagcgcctcc aatgacccag
tgggatccta ctccatcaat gggatcctgg 1260ggattcctcg ctccaatggt gagaagagga
aacgtgatga agttgaggta tacactgatc 1320ctgcccacat tagaggaggt ggaggtttgc
atctggtctg gactttaaga gatgtgtctg 1380agggctcagt ccccaatgga gattcccaga
gtggtgtgga cagtttgcgg aagcacttgc 1440gagctgacac cttcacccag cagcagctgg
aagctttgga tcgggtcttt gagcgtcctt 1500cctaccctga cgtcttccag gcatcagagc
acatcaaatc agaacagggg aacgagtact 1560ccctcccagc cctgacccct gggcttgatg
aagtcaagtc gagtctatct gcatccacca 1620accctgagct gggcagcaac gtgtcaggca
cacagacata cccagttgtg actggtcgtg 1680acatggcgag caccactctg cctggttacc
cccctcacgt gccccccact ggccagggaa 1740gctaccccac ctccaccctg gcaggaatgg
tgcctgggag cgagttctcc ggcaacccgt 1800acagccaccc ccagtacacg gcctacaacg
aggcttggag attcagcaac cccgccttac 1860taatgccgcc ccccggtccg cccctgccgc
tgctgccgct gcctatgacc gccactagtt 1920accgcgggga ccacatcaag cttcaggccg
acagcttcgg cctccacatc gtccccgtct 1980gaccccaccc cggagggagg gaggaccgac
gcgacgcgat gcctcccggc caccgcccca 2040gcctcacccc atcccacgac ccccgcaacc
cttcacatca cccccctcga aggtcggaca 2100ggacgggtgg agccgtgggc gggaccctca
ggcccgggcc cgccgccccc agccccgcct 2160gccgcccctc cccgcctgcc tggactgcgc
ggcgccgtga gggggattcg gcccagctcg 2220tcccggcctc caccaagcca gccccgaagc
ccgccagcca ccctgccgga ctcgggcgcg 2280acctgctggc gcgcgccgga tgtttctgtg
acacacaatc agcgcggacc gcagcgcggc 2340ccagccccgg gcacccgcct cggacgctcg
ggcgccagga ggcttcgctg gaggggctgg 2400gccaaggaga ttaagaagaa aacgactttc
tgcaggagga agagcccgct gccgaatccc 2460tgggaaaaat tcttttcccc cagtgccagc
cggactgccc tcgccttccg ggtgtgccct 2520gtcccagaag atggaatggg ggtgtggggg
tccggctcta ggaacgggct ttgggggcgt 2580caggtctttc caaggttggg acccaaggat
cggggggccc agcagcccgc accgatcgag 2640ccggactctc ggctcttcac tgctcctcct
ggcctgccta gttccccagg gcccggcacc 2700tcctgctgcg agacccggct ctcagccctg
ccttgcccct acctcagcgt ctcttccacc 2760tgctggcctc ccagtttccc ctcctgccag
tccttcgcct gtcccttgac gccctgcatc 2820ctcctccctg actcgcagcc ccatcggacg
ctctcccggg accgccgcag gaccagtttc 2880catagactgc ggactggggt cttcctccag
cagttacttg atgccccctc ccccgacaca 2940gactctcaat ctgccggtgg taagaaccgg
ttctgagctg gcgtctgagc tgctgcgggg 3000tggaagtggg gggctgccca ctccactcct
cccatcccct cccagcctcc tcctccggca 3060ggaactgaac agaaccacaa aaagtctaca
tttatttaat atgatggtct ttgcaaaaag 3120gaacaaaaca acacaaaagc ccaccaggct
gctgctttgt ggaaagacgg tgtgtgtcgt 3180gtgaaggcga aacccggtgt acataacccc
tccccctccg ccccgccccg cccggccccg 3240tagagtccct gtcgcccgcc ggccctgcct
gtagatacgc cccgctgtct gtgctgtgag 3300agtcgccgct cgctgggggg gaaggggggg
acacagctac acgcccatta aagcacagca 3360cgtcctgggg gaggggggca ttttttatgt
tacaaaaaaa aattacgaaa gaaaagaaat 3420ctctatgcaa aatgacgaac atggtcctgt
ggactcctct ggcctgtttt gttggctctt 3480tctctgtaat tccgtgtttt cgctttttcc
tccctgcccc tctctccctc tgcccctctc 3540tcctctccgc ttctctcccc ctctgtctct
gtctctctcc gtctctgtcg ctcttgtctg 3600tctgtctctg ctctttcctc ggcctctctc
cccagacctg gcccggccgc cctgtctccg 3660caggctagat ccgaggtggc agctccagcc
cccgggctcg ccccctcgcg ggcgtgcccc 3720gcgcgccccg ggcggccgaa ggccgggccg
ccccgtcccg ccccgtagtt gctctttcgg 3780tagtggcgat gcgccctgca tgtctcctca
cccgtggatc gtgacgactc gaaataacag 3840aaacaaagtc aataaagtga aaataaataa
aaatccttga acaaatccga aaaggcttgg 3900agtcctcgcc cagatctctc tcccctgcga
gcccttttta tttgagaagg aaaaagagaa 3960aagagaatcg tttaagggaa cccggcgccc
agccaggctc cagtggcccg aacggggcgg 4020cgagggcggc gagggcgccg aggtccggcc
catcccagtc ctgtggggct ggccgggcag 4080agaccccgga cccaggccca ggcctaacct
gctaaatgtc cccggacggt tctggtctcc 4140tcggccactt tcagtgcgtc ggttcgtttt
gattcttttt cttttgtgca cataagaaat 4200aaataataat aataaataaa gaataaaatt
ttgtatgtca aaaaaaaaaa aaaaaaa 42575118DNAHomo sapiens 51cctggcaccc
agcacaat 185220DNAHomo
sapiens 52gccgatccac acggagtact
205332DNAHomo sapiens 53gttgcctgcc agtcgccatg agaacttcct ac
325432DNAHomo sapiens 54tggccttccc tctgtaacag
gtgccttgaa tt 325524DNAHomo sapiens
55ccacccatgg caaattccat ggca
245624DNAHomo sapiens 56tctagacggc aggtcaggtc aacc
245723DNAHomo sapiens 57ctcaggccta tgcaaaaaga gga
235820DNAHomo sapiens
58gccctccctc caaaggagac
205923DNAHomo sapiens 59aacctacgca cctacgtgag gag
236022DNAHomo sapiens 60cgttcagtcc atcccatttc tg
226120DNAHomo sapiens
61gacacctgag ctgaccttgg
206220DNAHomo sapiens 62gaggaagtcc agtgtccagc
206368PRTHomo sapiens 63Met Arg Thr Ser Tyr Leu Leu
Leu Phe Thr Leu Cys Leu Leu Leu Ser1 5 10
15Glu Met Ala Ser Gly Gly Asn Phe Leu Thr Gly Leu Gly
His Arg Ser 20 25 30Asp His
Tyr Asn Cys Val Ser Ser Gly Gly Gln Cys Leu Tyr Ser Ala 35
40 45Cys Pro Ile Phe Thr Lys Ile Gln Gly Thr
Cys Tyr Arg Gly Lys Ala 50 55 60Lys
Cys Cys Lys6564914DNAHomo sapiensmisc_feature(1)..(914)n= a, c, g, or t
64ctgcagggtg ggcccaggct gggccnagac cctcaccctc caagggccac actgggggct
60cactttctga ggagtgccct ttggaaacgt cccaggaaca cgtctagtgg gaaaagagaa
120aagttggtcc atcgaggaga gtgttctgca taaggggaga gatgagaagg tagccttggc
180cagaggaaga aacttcatta caaccagctc tccttctsca agggaagagg gtgaagtttg
240agtttgtctt gcaggaagac aatcaaacta aagaggccaa caccagctta gagccgagcg
300gccccctgct cagagcttcc ctgtggctct cctccatgtg atccagaagg agggactcca
360gtgtgaactg cctgttccag aaaccccatc agaactgcct aacctagaaa accaaacagg
420aggagctggc accagggctc caggctgaaa gctaaatcca gcggcagcca gatggagaca
480atgtgccatg tgactgctga ctgctcaggg caaatgacac caggggttag cgattagaag
540ttcacccttg actgtggcac ctcccttcag ttccgtcgac gaggttgtgc aatccaccag
600tcttataaat acagtgacgc tccagcctct ggaagcctct gtcagctcag cctccaaagg
660agccagcctc tccccagttc ctgaaatcct gagtgttgcc tgccagtcgc catgagaact
720tcctaccttc tgctgtttac tctctgctta cttttgtctg agatggcctc aggtaagctc
780tggtacctgc tagagtttcc catccccagg gctggggaca atggggctga tgtgagtctc
840ggatggctgc ctccgtgtcc caagggacga ggaacaagca gcaggaaagc atcccgtggt
900tgagtggcct gcag
914
User Contributions:
Comment about this patent or add new information about this topic:
| People who visited this patent also read: | |
| Patent application number | Title |
|---|---|
| 20220032238 | Thin Metal/Ceramic Hybrid Membrane Sheet and Filter |
| 20220032237 | CMS MEMBRANE, METHOD FOR THE PRODUCTION THEREOF AND USE THEREOF |
| 20220032236 | FLUID SEPARATION ELEMENT |
| 20220032235 | DEVICE FOR BIND AND ELUTE CHROMATOGRAPHY USING MEMBRANES, AND METHOD OF MANUFACTURE |
| 20220032234 | METHOD FOR OPERATING MEMBRANE FILTRATION UNIT AND MEMBRANE FILTRATION UNIT |