Patent application title: Complexes of alpha (2) macroglobulin and antigenic molecules for immunotherapy
Inventors:
Pramod K. Srivastava (Avon, CT, US)
Pramod K. Srivastava (Avon, CT, US)
Robert J. Binder (Farmington, CT, US)
IPC8 Class: AA61K3816FI
USPC Class:
514 12
Class name: Designated organic active ingredient containing (doai) peptide containing (e.g., protein, peptones, fibrinogen, etc.) doai 25 or more peptide repeating units in known peptide chain structure
Publication date: 2009-10-01
Patent application number: 20090247456
Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
Patent application title: Complexes of alpha (2) macroglobulin and antigenic molecules for immunotherapy
Inventors:
Pramod K. Srivastava
Robert J. Binder
Agents:
JONES DAY
Assignees:
Origin: NEW YORK, NY US
IPC8 Class: AA61K3816FI
USPC Class:
514 12
Patent application number: 20090247456
Abstract:
The present invention relates to complexes of alpha (2) macroglobulin
associated with antigenic molecules for use in immunotherapy. The
invention relates to methods for using such compositions in the diagnosis
and treatment of immune disorders, proliferative disorders, and
infectious diseases.Claims:
1.-36. (canceled)
37. A method of treating cancer in an individual having a type of cancer, comprising administering to the individual a composition comprising a therapeutically effective amount of purified molecular complexes, which are at least 65% noncovalent complexes of an alpha-2-macroglobulin and an antigenic molecule, said purified molecular complexes comprising an alpha-2-macroglobulin associated with an antigenic molecule of an antigen overexpressed in a cancer cell of said type of cancer relative to its expression in a noncancerous cell of the same cell type as the cancer cell.
38. The method of claim 37, wherein the antigenic molecule is a tumor-specific antigen or a tumor-associated antigen.
39. The method of claim 37, wherein the cells of said type of cancer is a metastasis.
40. The method of claim 37, wherein said administering of said composition is repeated at weekly intervals.
41. The method of claim 37, wherein said administering of said composition is repeated at the same site of the individual.
42. The method of claim 37, wherein said administering of said composition is intradermally or subcutaneously.
43. The method of claim 37, wherein the individual is a human individual.
44. The method of claim 38, wherein the individual is a human individual.
45. The method of claim 37, wherein the type of cancer is fibrosarcoma, myxosarcoma, liposarcoma, chondrosarcoma, osteogenic sarcoma, chordoma, angiosarcoma, endotheliosarcoma, lymphangiosarcoma, lymphangioendotheliosarcoma, synovioma, mesothelioma, Ewing's tumor, leiomyosarcoma, rhabdomyosarcoma, colon carcinoma, pancreatic cancer, breast cancer, ovarian cancer, prostate cancer, squamous cell carcinoma, basal cell carcinoma, adenocarcinoma, sweat gland carcinoma, sebaceous gland carcinoma, papillary carcinoma, papillary adenocarcinomas, cystadenocarcinoma, medullary carcinoma, bronchogenic carcinoma, renal cell carcinoma, hepatoma, bile duct carcinoma, choriocarcinoma, seminoma, embryonal carcinoma, Wilms' tumor, cervical cancer, testicular tumor, lung carcinoma, small cell lung carcinoma, bladder carcinoma, epithelial carcinoma, glioma, astrocytoma, medulloblastoma, craniopharyngioma, ependymoma, pinealoma, hemangioblastoma, acoustic neuroma, oligodendroglioma, meningioma, melanoma, neuroblastoma, retinoblastoma, acute lymphocytic leukemia, myeloblastic, promyelocytic, myelomonocytic, monocytic, erythroleukemia, chronic myelocytic, leukemia, chronic lymphocytic leukemia, polycythemia vera, Hodgkin's disease lymphoma, non-Hodgkin's disease lymphoma, multiple myeloma, Waldenstrom's macroglobulinemia, or heavy chain disease.
46. The method of claim 37, wherein the composition comprises one or more adjuvants.
47. The method of claim 46, wherein the composition comprises an adjuvant selected from the group consisting of aluminum hydroxide, aluminum phosphate, calcium phosphate, lysolecithin, pluronic polyols, polyanions, peptides, oil emulsions, keyhole limpet hemocyanins, and dinitrophenol, cytokines, saponins, muramyl dipeptides, tripeptide derivatives, CpG dinucleotides, CpG oligonucleotides, monophosphoryl Lipid A, polyphosphazenes, emulsions, liposomes, virosomes, cochleates, Freund's complete adjuvant, Freund's incomplete adjuvant, bacille Calmette-Guerin, and corynebacterium parvum.
Description:
[0001]This application is a divisional of U.S. patent application Ser. No.
09/873,403 filed Jun. 4, 2001, now U.S. Pat. No. 7,449,557, which is a
continuation-in-part of U.S. patent application Ser. No. 09/625,139 filed
Jul. 25, 2000, now abandoned, which claims benefit under 35 U.S.C. §
119(e) of U.S. Provisional Patent Application No. 60/209,266 filed Jun.
2, 2000, each of which are incorporated by reference herein in their
entireties.
1. INTRODUCTION
[0003]The present invention relates to complexes of alpha (2) macroglobulin associated with antigenic molecules for use in immunotherapy. The invention relates to methods for using such compositions in the diagnosis and treatment of immune disorders, proliferative disorders, and infectious diseases.
2. BACKGROUND OF THE INVENTION
2.1 Heat Shock Proteins
[0004]Heat shock proteins (HSPs), also referred to as stress proteins, were first identified as proteins synthesized by cells in response to heat shock. Hsps have classified into five families, based on molecular weight, Hsp100, Hsp90, Hsp70, Hsp60, and smhsp. Many members of these families were found subsequently to be induced in response to other stressful stimuli including nutrient deprivation, metabolic disruption, oxygen radicals, and infection with intracellular pathogens (see Welch, May 1993, Scientific American 56-64; Young, 1990, Annu. Rev. Immunol. 8:401-420; Craig, 1993, Science 260:1902-1903; Gething et al., 1992, Nature 355:33-45; and Lindquist et al., 1988, Annu. Rev. Genetics 22:631-677).
[0005]Heat shock proteins are among the most highly conserved proteins in existence. For example, DnaK, the Hsp70 from E. coli has about 50% amino acid sequence identity with Hsp70 proteins from excoriates (Bardwell et al., 1984, Proc. Natl. Acad. Sci. 81:848-852). The Hsp60 and Hsp90 families also show similarly high levels of intra-family conservation (Hickey et al., 1989, Mol. Cell. Biol. 9:2615-2626; Jindal, 1989, Mol. Cell. Biol. 9:2279-2283). In addition, it has been discovered that the Hsp60, Hsp70 and Hsp90 families are composed of proteins that are related to the stress proteins in sequence, for example, having greater than 35% amino acid identity, but whose expression levels are not altered by stress.
[0006]Studies on the cellular response to heat shock and other physiological stresses revealed that the HSPs are involved not only in cellular protection against these adverse conditions, but also in essential biochemical and immunological processes in unstressed cells. Hsps accomplish different kinds of chaperoning functions. For example, members of the Hsp70 family, located in the cell cytoplasm, nucleus, mitochondria, or endoplasmic reticulum (Lindquist et al., 1988, Ann. Rev. Genetics 22:631-677), are involved in the presentation of antigens to the cells of the immune system, and are also involved in the transfer, folding and assembly of proteins in normal cells. Hsps are capable of binding proteins or peptides, and releasing the bound proteins or peptides in the presence of adenosine triphosphate (ATP) or low pH.
2.2 Immunogenicity of Hsp-Peptide Complexes
[0007]Srivastava et al. demonstrated immune response to methylcholanthrene-induced sarcomas of inbred mice (1988, Immunol. Today 9:78-83). In these studies, it was found that the molecules responsible for the individually distinct immunogenicity of these tumors were glycoproteins of 96 kDa (gp96) and intracellular proteins of 84 to 86 kDa (Srivastava et al., 1986, Proc. Natl. Acad. Sci. USA 83:3407-3411; Ullrich et al., 1986, Proc. Natl. Acad. Sci. USA 83:3121-3125). Immunization of mice with gp96 or p84/86 isolated from a particular tumor rendered the mice immune to that particular tumor, but not to antigenically distinct tumors. Isolation and characterization of genes encoding gp96 and p84/86 revealed significant homology between them, and showed that gp96 and p84/86 were, respectively, the endoplasmic reticular and cytosolic counterparts of the same heat shock proteins (Srivastava et al., 1988, Immunogenetics 28:205-207; Srivastava et al., 1991, Curr. Top. Microbiol. Immunol. 167:109-123). Further, Hsp70 was shown to elicit immunity to the tumor from which it was isolated but not to antigenically distinct tumors. However, Hsp70 depleted of peptides was found to lose its immunogenic activity (Udono and Srivastava, 1993, J. Exp. Med. 178:1391-1396). These observations suggested that the heat shock proteins are not immunogenic per se, but form noncovalent complexes with antigenic peptides, and the complexes can elicit specific immunity to the antigenic peptides (Srivastava, 1993, Adv. Cancer Res. 62:153-177; Udono et al., 1994, J. Immunol., 152:5398-5403; Suto et al., 1995, Science 269:1585-1588).
[0008]Noncovalent complexes of HSPs and peptide, purified from cancer cells, can be used for the treatment and prevention of cancer and have been described in PCT publications WO 96/10411, dated Apr. 11, 1996, and WO 97/10001, dated Mar. 20, 1997 (U.S. Pat. No. 5,750,119 issued Apr. 12, 1998, and U.S. Pat. No. 5,837,251 issued Nov. 17, 1998, respectively, each of which is incorporated by reference herein in its entirety). The isolation and purification of stress protein-antigen complexes has been described, for example, from pathogen-infected cells, and used for the treatment and prevention of infection caused by the pathogen, such as viruses, and other intracellular pathogens, including bacteria, protozoa, fungi and parasites (see, for example, PCT Publication WO 95/24923, dated Sep. 21, 1995). Immunogenic stress protein-antigen complexes can also be prepared by in vitro complexing of stress protein and antigenic peptides, and the uses of such complexes for the treatment and prevention of cancer and infectious diseases has been described in PCT publication WO 97/10000, dated Mar. 20, 1997 (U.S. Pat. No. 6,030,618 issued Feb. 29, 2000. The use of stress protein-antigen complexes for sensitizing antigen presenting cells in vitro for use in adoptive immunotherapy is described in PCT publication WO 97/10002, dated Mar. 20, 1997 (see also U.S. Pat. No. 5,985,270 issued Nov. 16, 1999).
2.3 Alpha (2) Macroglobulin Receptor
[0009]The α-macroglobulins are members of a protein superfamily of structurally related proteins which also comprises complement components C3, C4 and C5. The human plasma protein alpha (2) macroglobulin (α2M) is a 720 kDa homotetrameric protein primarily known as proteinase inhibitor and plasma and inflammatory fluid proteinase scavenger molecule (for review see Chu and Pizzo, 1994, Lab. Invest. 71:792). Alpha (2) macroglobulin is synthesized as a 1474 amino acid precursor, the first 23 of which function as a signal sequence that is cleaved to yield a 1451 amino acid mature protein (Kan et al., 1985, Proc. Natl. Acad. Sci. U.S.A. 82:2282-2286).
[0010]Alpha (2) macroglobulin promiscuously binds to proteins and peptides with nucleophilic amino acid side chains in a covalent manner (Chu et al., 1994, Ann. N.Y. Acad. Sci. 737:291-307) and targets them to cells which express the α2M receptor (α2MR) (Chu and Pizzo, 1993, J. Immunol. 150:48). Binding of α2M to the α2M receptor is mediated by the C-terminal portion of α2M (Holtet et al., 1994, FEBS Lett. 344:242-246) and key residues have been identified (Nielsen et al., 1996, J. Biol. Chem. 271:12909-12912).
[0011]Generally known for inhibiting protease activity, α2M binds to a variety of proteases thorough multiple binding sites (see, e.g., Hall et al., 1981, Biochem. Biophys. Res. Commun. 100(1):8-16). Protease interaction with α2M results in a complex structural rearrangement called transformation, which is the result of a cleavage within the "bait" region of α2M after the proteinase becomes "trapped" by thioesters. The conformational change exposes residues required for receptor binding, allowing the α2M-proteinase complex to bind to the α2MR. Methylamine can induce similar conformational changes and cleavage as that induced by proteinases. The uncleaved form of α2M, which is not recognized by the receptor, is often referred to as the "slow" form (s-α2M). The cleaved form is referred to as the "fast" form (f-α2M) (reviewed by Chu et al., 1994, Ann. N.Y. Acad. Sci. 737:291-307).
[0012]Studies have shown that in addition to its proteinase-inhibitory functions, α2M, when complexed to antigens, can enhance the antigens' ability to be taken up by antigen presenting cells such as macrophages and presented to T cell hybridomas in vitro by up to two orders of magnitude (Chu and Pizzo, 1994, Lab. Invest. 71:792), and induce T cell proliferation (Osada et al., 1987, Biochem. Biophys. Res. Commun. 146:26-31). Further evidence suggests that complexing antigen with α2M enhances antibody production by crude spleen cells in vitro (Osada et al., 1988, Biochem. Biophys. Res. Commun. 150:883) elicits an in vivo antibody responses in experimental rabbits (Chu et al., 1994, J. Immunol. 152:1538-1545) and mice (Mitsuda et al., 1993, Biochem. Biophys. Res. Commun. 101: 1326-1331). However, none of these studies have shown whether alphα2M-antigen complexes are capable of eliciting cytotoxic T cell responses in vivo.
2.4 Immunogenicity of Heat Shock/Stress Proteins
[0013]Srivastava et al. demonstrated immune response to methylcholanthrene-induced sarcomas of inbred mice (1988, Immunol. Today 9:78-83). In these studies, it was found that the molecules responsible for the individually distinct immunogenicity of these tumors were identified as cell-surface glycoproteins of 96 kDa (gp96) and intracellular proteins of 84 to 86 kDa (Srivastava et al., 1986, Proc. Natl. Acad. Sci. USA 83:3407-3411; Ullrich, S. J. et al., 1986, Proc. Natl. Acad. Sci. USA 83:3121-3125). Immunization of mice with gp96 or p84/86 isolated from a particular tumor rendered the mice immune to that particular tumor, but not to antigenically distinct tumors. Isolation and characterization of genes encoding gp96 and p84/86 revealed significant homology between them, and showed that gp96 and p84/86 were, respectively, the endoplasmic reticular and cytosolic counterparts of the same heat shock proteins (Srivastava et al., 1988, Immunogenetics 28:205-207; Srivastava et al., 1991, Curr. Top. Microbiol. Immunol. 167:109-123). Further, Hsp70 was shown to elicit immunity to the tumor from which it was isolated but not to antigenically distinct tumors. However, Hsp70 depleted of peptides was found to lose its immunogenic activity (Udono and Srivastava, 1993, J. Exp. Med. 178:1391-1396). These observations suggested that the heat shock proteins are not immunogenic per se, but form noncovalent complexes with antigenic peptides, and the complexes can elicit specific immunity to the antigenic peptides (Srivastava, 1993, Adv. Cancer Res. 62:153-177; Udono et al., 1994, J. Immunol., 152:5398-5403; Suto et al., 1995, Science, 269:1585-1588).
[0014]The use of noncovalent complexes of stress proteins and peptides, purified from cancer cells, for the treatment and prevention of cancer, as well as the use of such complexes in combination with adoptive immunotherapy, has been described (see U.S. Pat. No. 5,750,199; U.S. Pat. No. 5,830,464; Patent Cooperation Treaty ("PCT") publications WO 96/10411, dated Apr. 11, 1996; and WO 97/10001, dated Mar. 20, 1997; each of which is incorporated by reference herein in its entirety. The purification of stress protein-peptide complexes from cell lysates has been described previously; stress protein-peptide complexes can be isolated from pathogen-infected cells and used for the treatment and prevention of infection caused by pathogens, such as viruses and other intracellular pathogens, including bacteria, protozoa, fungi and parasites (see PCT publication WO 95/24923, dated Sep. 21, 1995).
[0015]Immunogenic stress protein-peptide complexes can also be prepared by in vitro complexing of stress protein and antigenic peptides, and the uses of such complexes for the treatment and prevention of infectious diseases and cancer has been described in PCT publication WO 97/10000, dated Mar. 20, 1997. The use of heat shock proteins in combination with a defined antigen for the treatment of infectious diseases and cancer have also been described in PCT publication WO 97/06821, dated Feb. 27, 1997. The administration of expressible polynucleotides encoding eukaryotic heat shock proteins to mammalian cells for stimulating an immune response, and for treatment of infectious diseases and cancer has been described in PCT publications, WO 97/06685 and WO 97/06828, both dated Feb. 27, 1997. The use of stress protein-peptide complexes for sensitizing antigen presenting cells in vitro for use in adoptive immunotherapy is described in PCT publication WO 97/10002, dated Mar. 20, 1997.
2.5 Antigen Presentation
[0016]Major histocompatibility complex (MHC) molecules present antigens on the cell surface of antigen-presenting cells. Cytotoxic T lymphocytes (CTLs) then recognize MHC molecules and their associated peptides and kill the target cell. Antigens are processed by two distinct antigen processing routes depending upon whether their origin is intracellular or extracellular. Intracellular or endogenous protein antigens, i.e., antigens synthesized within the antigen-presenting cell, are presented by MHC class I (MHC I) molecules to CD8+ cytotoxic T lymphocytes. On the other hand, extracellular or exogenously synthesized antigenic determinants are presented on the cell surface of "specialized" or "professional" APCs (macrophages, for example) by MHC class II molecules to CD4+ T cells (see, generally, Fundamental Immunology, W. E. Paul (ed.), New York: Raven Press, 1984). This compartmental segregation of antigen processing routes is important to prevent tissue destruction that could otherwise occur during an immune response as a result of shedding of neighboring cell MHC I antigens.
[0017]The heat shock protein gp96 chaperones a wide array of peptides, depending upon the source from which gp96 is isolated (for review, see Srivastava et al., 1998, Immunity 8: 657-665). Tumor-derived gp96 carries tumor-antigenic peptides (Ishii et al., 1999, J. Immunology 162:1303-1309); gp96 preparations from virus-infected cells carry viral epitopes (Suto and Srivastava, 1995, Science 269:1585-1588; Nieland et al., 1998, Proc. Natl. Acad. Sci. USA 95:1800-1805), and gp96 preparations from cells transfected with model antigens such as ovalbumin or β-galactosidase are associated with the corresponding epitopes (Arnold et al., 1995, J. Exp. Med. 182:885-889; Breloer et al., 1998, Eur. J. Immunol. 28:1016-1021). The association of gp96 with peptides occurs in vivo (Menoret and Srivastava, 1999, Biochem. Biophys. Research Commun. 262:813-818). Gp96-peptide complexes, whether isolated from cells (Tamura et al., 1997, Science 278:117-120), or reconstituted in vitro (Blachere et al., 1997, J. Exp. Med. 186:1183-1406) are excellent immunogens and have been used extensively to elicit CD8+ T cell responses specific for the gp96-chaperoned antigenic peptides.
[0018]The capacity of gp96-peptide complexes to elicit an immune response is dependent upon the transfer of the peptide to MHC class I molecules of antigen-presenting cells (Suto and Srivastava, 1995, supra). Endogenously synthesized antigens chaperoned by gp96 in the endoplasmic reticulum [ER] can prime antigen-specific CD8+ T cells (or MHC I-restricted CTLs) in vivo; this priming of CD8+ T cells requires macrophages. However, the process whereby exogenously introduced gp96-peptide complexes elicit the antigen-specific CD8+ T cell response is not completely understood since there is no established pathway for the translocation of extracellular antigens into the class I presentation machinery. Yet antigenic peptides of extracellular origin associated with HSPs are somehow salvaged by macrophages, channeled into the endogenous pathway, and presented by MHC I molecules to be recognized by CD8+ lymphocytes (Suto and Srivastava, 1995, supra; Blachere et al., 1997, J. Exp. Med. 186:1315-22).
[0019]Several models have been proposed to explain the delivery of extracellular peptides for antigen presentation. One proposal, known as the "direct transfer" model, suggests that HSP-chaperoned peptides are transferred to MHC I molecules on the cell surface of macrophages for presentation to CD8+ T lymphocytes. Another suggestion is that soluble extracellular proteins can be trafficked to the cytosol via constitutive macropinocytosis in bone marrow-derived macrophages and dendritic cells (Norbury et al., 1997, Eur. J. Immunol. 27:280-288). Yet another proposed mechanism is that HSPs are taken up by the MHC class Imolecules of the macrophage, which stimulate the appropriate T cells (Srivastava et al., 1994, Immunogenetics 39:93-98. Others have suggested that a novel intracellular trafficking pathway may be involved for the transport of peptides from the extracellular medium into the lumen of ER (Day et al., 1997, Proc. Natl. Acad. Sci. 94:8064-8069; Nicchitta, 1998, Curr. Opin. in Immunol. 10:103-109). Further suggestions include the involvement of phagocytes which (a) possess an ill-defined pathway to shunt protein from the phagosome into the cytosol where it would enter the normal class I pathway; (b) digest ingested material in lysosomes and regurgitate peptides for loading on the surface to class I molecules (Bevan, 1995, J. Exp. Med. 182:639-41).
[0020]Still others have proposed a receptor-mediated pathway for the delivery of extracellular peptides to the cell surface of APS for antigen presentation. In view of the extremely small quantity of gp96-chaperoned antigenic peptides required for immunization (Blachere et al., 1997, supra), and the strict dependence of immunogenicity of gp96-peptide complexes on functional antigen presenting cells (APCs) (Udono et al., 1994, Proc. Natl. Acad. Sci. U.S.A. 91:3077-3081), APCs had been proposed to possess receptors for gp96 (Srivastava et al., 1994, Immunogenetics 39:93-98). Preliminary microscopic evidence consistent with such receptors has been recently obtained (Binder et al., 1998, Cell Stress & Chaperones 3 (Supp.1):2; Arnold-Schild et al., 1999, J. Immunol. 162: 3757-3760; and Wassenberg et al., 1999, J. Cell Sci. 1:12). One hypothesis is that the mannose receptor is used in the uptake of gp96, but no mechanism has been proposed for the non-glycosylated HSPs, such as Hsp70 (Ciupitu et al., 1998, J. Exp. Med., 187:685-691).
[0021]The identification and characterization of specific molecules involved in HSP-mediated antigen presentation of peptides, could provide useful reagents and techniques for eliciting specific immunity by HSP and HSP-peptide complexes, and for developing novel diagnostic and therapeutic methods.
[0022]Citation or discussion of a reference herein shall not be construed as an admission that such is prior art to the present invention.
3. SUMMARY OF THE INVENTION
[0023]The present invention provides complexes comprising alpha (2) macroglobulin ("α2M") and methods for their use in immunotherapy. The invention is based, in part, on the Applicant's discovery that α2M directly competes for the binding of heat shock protein gp96 to the α2M receptor, indicating that α2M and HSPs may bind to a common recognition site on the alpha (2) macroglobulin receptor. Thus, because HSPs and α2M have a number of common functional attributes, such as the ability to bind peptides and the recognition and uptake by the alpha (2) macroglobulin receptor, the Applicants have discovered that α2M can be used in the methods described herein for immunotherapy against cancer and infectious disease. Alpha-2-macroglobulin can form complexes with antigens, which are taken up by antigen presenting cells ("APCs") via the alpha (2) macroglobulin receptor, also known as LDL (low-density lipoprotein) Receptor-Related Protein ("LRP") or CD91. Thus, the invention provides methods and compositions for using specific α2M-antigenic molecule complexes for targeting an immune response against immune disorders, proliferative disorders, and infectious diseases.
[0024]The invention encompasses complexes of alpha (2) macroglobulin noncovalently associated antigenic molecules, recombinant cells that express the complexes of α2M associated with antigenic molecules, and antibodies and other molecules that specifically recognize α2M-antigenic molecule complexes. The invention also provides methods for using these compositions in the diagnosis and treatment of immune disorders, proliferative disorders, and infectious diseases.
[0025]As used herein, an alpha (2) macroglobulin is associated with an antigenic molecule is bound to the antigenic molecule by a covalent or noncovalent bond. A covalent bond can be a peptide bond or a thioester linkage, for example. Thus, fusion proteins between alpha (2) macroglobulin and an antigenic molecule are within the scope of the invention.
[0026]The invention provides a pharmaceutical composition comprising an amount of a molecular complex effective for treatment or prevention of an infectious disease or cancer, and a pharmaceutically acceptable carrier, said molecular complex comprising an alpha (2) macroglobulin polypeptide noncovalently associated with an antigenic molecule which displays the antigenicity of an antigen of an infectious agent or an antigen overexpressed in a cancer cell relative to its expression in a noncancerous cell of said cell type. As used herein a cell type of a cancer cell, refers to the cell type of the tissue of origin, e.g., breast, lung, ovarian. In one embodiment, the antigenic molecule displays the antigenicity of an antigen of an infectious agent. In another embodiment, the antigenic molecule displays the antigenicity of an antigen overexpressed in a cancer cell relative to its expression in a noncancerous cell of said cell type. In another embodiment, the antigenic molecule is a tumor specific antigen or a tumor-associated antigen. In another embodiment, the antigenic molecule displays the antigenicity of an antigen overexpressed in a cancer cell relative to its expression in a noncancerous cell of said cell type.
[0027]In another embodiment, the molecular complex effective for treatment or prevention of an infectious disease or cancer, comprising the alpha (2) macroglobulin polypeptide noncovalently associated with the antigenic molecule is purified. In particular, the term "purified" molecular complexes refer to complexes which are at least 65% 75%, 80%, 85%, 90%, 95%, 98% or 100% noncovalent complexes of the alpha (2) macroglobulin polypeptide and the antigenic molecule. In another embodiment, the purified molecular complex comprising an alpha (2) macroglobulin polypeptide associated with an antigenic molecule of an infectious agent or an antigen overexpressed in a cancer cell relative to its expression in a noncancerous cell of said cell type.
[0028]The invention further provides a purified population of molecular complexes in which at least 65%, 70%, 75%, 80%, 85%, 90%, 95%, 98%, or 100% of the complexes comprise an alpha (2) macroglobulin noncovalently associated with an antigenic molecule. Also provided by the invention is a purified population of molecular complexes purified from a recombinant cell in which at least 65%, 70%, 75%, 80%, 85%, 90%, 95%, 98%, or 100% of the complexes comprise an alpha (2) macroglobulin noncovalently associated with an antigenic molecule.
[0029]The invention also provides a recombinant cell infected with a pathogen and transformed with a nucleic acid comprising a nucleotide sequence that (i) is operably linked to a promoter, and (ii) encodes an alpha (2) macroglobulin polypeptide, which alpha (2) macroglobulin polypeptide associates with an antigenic molecule, when said antigenic molecule is present, to form a complex that in sufficient amount is capable of eliciting an immune response to the antigenic molecule. The invention provides a recombinant cancer cell transformed with a nucleic acid comprising a nucleotide sequence that (i) is operably linked to a promoter, and (ii) encodes an alpha (2) macroglobulin polypeptide, which alpha (2) macroglobulin polypeptide associates with an antigenic molecule, when said antigen is present, to form a complex that in sufficient amount is capable of eliciting an immune response to the antigenic molecule. In another embodiment, the invention provides a recombinant cell transformed with (i) a first nucleic acid comprising a first nucleotide sequence that is operably linked to a first promoter and encodes an alpha (2) macroglobulin polypeptide, and (ii) a second nucleic acid comprising a second nucleotide sequence that is operably linked to a second promoter and encodes an antigenic molecule, such that the alpha (2) macroglobulin polypeptide and the antigenic molecule are expressed within the cell and associate with each other to form a complex that in sufficient amount is capable of eliciting an immune response to the antigenic molecule. In various embodiments, the recombinant cells are human cells. In various embodiments, the pharmaceutical composition comprises a recombinant cell and a pharmaceutically acceptable carrier.
[0030]In one embodiment, a method is provided for preparing a complex of an alpha (2) macroglobulin polypeptide noncovalently associated with an antigenic molecule, said alpha (2) macroglobulin polypeptide comprising: (a) culturing a cell transformed with a nucleic acid comprising a nucleotide sequence encoding the alpha (2) macroglobulin polypeptide, under conditions such that said alpha (2) macroglobulin polypeptide is expressed by the cells and associates with an antigenic molecule of the cell; and (b) recovering a population of complexes of the alpha (2) macroglobulin polypeptide noncovalently associated with antigenic molecules from the cells.
[0031]The invention further provides a method for preparing an alpha (2) macroglobulin polypeptide noncovalently associated with antigenic molecules derived from one or more antigens of an infectious agent, comprising: culturing infected cells, transformed with a nucleic acid comprising a nucleotide sequence encoding the alpha (2) macroglobulin polypeptide, or fragment, analog, or variant thereof, and operably linked to a promoter, under conditions such that the alpha (2) macroglobulin polypeptide is expressed by the cells and associates with peptides of the cells; and (b) recovering from the cells a population of complexes of the alpha (2) macroglobulin polypeptide noncovalently associated with peptides derived from the infectious agent. In one embodiment, the method further comprises purifying the complexes. In another embodiment, the method further comprising purifying the complexes by affinity chromatography.
[0032]The invention further provides a method of treating or preventing an infectious disease in an individual having an infectious disease comprising administering to the individual one or more immunogenic complexes of an alpha (2) macroglobulin polypeptide noncovalently associated with a first antigenic molecule, wherein the first antigenic molecule displays the antigenicity of an antigen of an infectious agent of the infectious disease. In another embodiment, the method further comprises, before, concurrently or after administration of the immunogenic complex, administering to the individual a composition comprising antigen presenting cells sensitized in vitro with a sensitizing amount of a second complex of alpha (2) macroglobulin polypeptide noncovalently associated with a second antigenic molecule, said second antigenic molecule displaying the antigenicity of a second antigenic molecule of said infectious agent.
[0033]The invention further provides a method of treating or preventing an infectious disease in a subject having an infectious disease comprising: a) culturing an infected cell transformed with a nucleic acid comprising a nucleotide sequence encoding an alpha (2) macroglobulin polypeptide, said infected cell displaying the antigenicity of an antigen of an infectious agent of the infectious disease, said nucleotide sequence being operably linked to a promoter, under conditions such that the alpha (2) macroglobulin polypeptide is expressed by the infected cells and associates with antigenic molecules of the cell; b) recovering complexes of the alpha (2) macroglobulin polypeptide noncovalently associated with antigenic molecules from the infected cell; and c) administering to the subject an amount of the recovered complexes effective to treat or prevent the infectious disease. In one embodiment, the method further comprises, prior to step (a), the step of obtaining infected cells from the subject and transforming the infected cells with the nucleic acid. In another embodiment, the method further comprises, prior to step (a), the step of obtaining the infected cell from one or more individuals and transforming the infected cells with the nucleic acid, said one or more individuals being different from the subject and having the same type of infectious disease as the subject.
[0034]The invention further provides a method of treating or preventing an infectious disease in a subject having an infectious disease comprising: a) culturing a recombinant cell transformed with (i) a first nucleic acid encoding an alpha (2) macroglobulin polypeptide and (ii) a second nucleic acid encoding an antigenic molecule displaying the antigenicity of an antigen of an infectious agent of the infectious disease; b) recovering complexes of the alpha (2) macroglobulin polypeptide noncovalently associated with the antigen; and c) administering to the subject an amount of the recovered complexes effective to treat or prevent the infectious disease. In various embodiments, the infectious disease is caused by an infectious agent selected from the group consisting of a virus, a bacterium, a fungus, and a parasite.
[0035]Also provide by the invention is a method of treating or preventing cancer in an individual having a type of cancer or in whom prevention of a type of cancer is desired comprising administering to the individual an immunogenic complex of an alpha (2) macroglobulin polypeptide noncovalently associated with a first antigenic molecule, wherein either (a) the first antigenic molecule displays antigenicity of said type of cancer or a metastasis thereof; or (b) the complex is obtained by recovering complexes from said type of cancer cells or a metastasis thereof that recombinantly express the alpha (2) macroglobulin polypeptide. In one embodiment, this method further comprises, before, concurrently or after administration of the immunogenic complex, administering to the individual a composition comprising antigen presenting cells sensitized in vitro with a sensitizing amount of a second complex of an alpha (2) macroglobulin noncovalently associated with a second antigenic molecule, said second antigenic molecule displaying the antigenicity of an antigen overexpressed in a cancer cell relative to its expression in a noncancerous cell of said cell type. an infectious agent of the infectious disease. In another embodiment, the first antigenic molecule is an antigen overexpressed in a cancer cell relative to its expression in a noncancerous cell of said cell type.
[0036]In one embodiment, a method is provided for treating or preventing cancer in a subject having a type of cancer or in whom prevention of a type of cancer is desired comprising: a) culturing a cancer cell of said type transformed with a nucleic acid comprising a nucleotide sequence encoding an alpha (2) macroglobulin polypeptide said nucleotide sequence being operably linked to a promoter, under conditions such that the alpha (2) macroglobulin polypeptide is expressed by the cancer cell and associates with at least one antigenic molecule of the cell; b) recovering complexes of the alpha (2) macroglobulin polypeptide noncovalently associated with at least one antigenic molecule from the cancer cell; and c) administering to the subject an amount of the recovered complexes effective to treat or prevent cancer. In one embodiment, the method further comprises, prior to step (a), the step of obtaining cancer cells from the subject and transforming the cancer cells with the nucleic acid. In another embodiment, the method further comprises, prior to step (a), the step of obtaining cancer cells from one or more individuals and transforming the cancer cells with the nucleic acid, said one or more individuals being different from the subject and having the same type of cancer as the subject.
[0037]The invention further provides a method of treating or preventing cancer in a subject having a type of cancer or in whom prevention of a type of cancer is desired comprising: a) culturing a recombinant cell transformed with (i) a first nucleic acid encoding an alpha (2) macroglobulin polypeptide, and (ii) a second nucleic acid encoding an antigenic molecule displaying the antigenicity of an antigen of a cancer cell; b) recovering complexes of the alpha (2) macroglobulin polypeptide noncovalently associated with the antigen; and c) administering to the subject an amount of the recovered complexes effective to treat or prevent cancer.
[0038]As used herein a "type of cancer" refers to e.g., melanoma, breast cancer, renal carcinoma, or a metastasis thereof, where a metastasis refers to the same type of cancer as the cell of origin. In various embodiments, the cancer is selected from the group consisting of fibrosarcoma, myxosarcoma, liposarcoma, chondrosarcoma, osteogenic sarcoma, chordoma, angiosarcoma, endotheliosarcoma, lymphangiosarcoma, lymphangioendotheliosarcoma, synovioma, mesothelioma, Ewing's tumor, leiomyosarcoma, rhabdomyosarcoma, colon carcinoma, pancreatic cancer, breast cancer, ovarian cancer, prostate cancer, squamous cell carcinoma, basal cell carcinoma, adenocarcinoma, sweat gland carcinoma, sebaceous gland carcinoma, papillary carcinoma, papillary adenocarcinomas, cystadenocarcinoma, medullary carcinoma, bronchogenic carcinoma, renal cell carcinoma, hepatoma, bile duct carcinoma, choriocarcinoma, seminoma, embryonal carcinoma, Wilms' tumor, cervical cancer, testicular tumor, lung carcinoma, small cell lung carcinoma, bladder carcinoma, epithelial carcinoma, glioma, astrocytoma, medulloblastoma, craniopharyngioma, ependymoma, pinealoma, hemangioblastoma, acoustic neuroma, oligodendroglioma, meningioma, melanoma, neuroblastoma, retinoblastoma, leukemia, lymphoma, multiple myeloma, Waldenstrom's macroglobulinemia, and heavy chain disease.
[0039]The invention also encompasses a method for treating an autoimmune disorder comprising administering to a mammal in need of such treatment an antibody specific for alpha (2) macroglobulin. In one embodiment, the antibody is purified.
4. BRIEF DESCRIPTION OF THE FIGURES
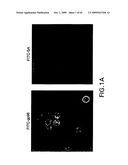
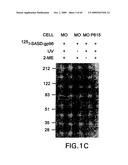
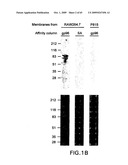
[0040]FIG. 1A-C. Identification of an 80 kDa polypeptide as a putative gp96 receptor. A. Confocal microscopy of re-presentation-competent RAW264.7 cells stained with gp96-FITC (left panel) and with albumin-FITC (right panel). B. SDS-PAGE analysis of detergent extracts of plasma membranes from surface biotinylated RAW264.7 (re-presentation-competent) or P815 cells (representation-incompetent) eluted from gp96 or albumin-Sepharose (SA) columns and stained with silver stain (top) or avidin-peroxidase (bottom). C. gp96-SASD-I125 was cross-linked to live peritoneal macrophages (MO) or P815 cells, and the cell lysates examined by SDS-PAGE and autoradiography. Various components were omitted as controls, as indicated.
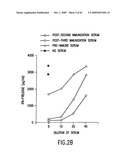
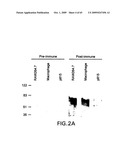
[0041]FIG. 2A-B. Anti-p80 antiserum detects an 80 kDa molecule and inhibits re-presentation of gp96-chaperoned AHI peptide by macrophage. A. Pre-immune and immune sera were used to probe blots of plasma membrane extracts of RAW264.7, peritoneal macrophages (both cell types re-presentation-competent), or P815 cells. B. Re-presentation of gp96-chaperoned peptide AHI. Sera were added at the final dilution indicated. The solid cross indicates the level of T cell stimulation when the APCs were pulsed directly with the AHi peptide. The open cross indicates the corresponding value with unpulsed APCs.
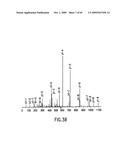
[0042]FIG. 3A-C. Protein microsequencing of the 80 kDa protein. A. Analysis of a single tryptic (GGALHIYHQR) (SEQ ID NO: 6)_peptide by tandem-mass spectrometry. All possible b- and y-ion series together with identified b-ion series (red) and y-ion series (blue) are shown. B. Collision-induced dissociation (CID) spectrum of this peptide is shown. C. Four identified peptides from the α2M receptor, (i.e., SGFSLGSDGK (SEQ ID NO: 9), GIALDPAMGK (SEQ ID NO: 10), GGALHIYHQR (SEQ ID NO: 11), and VFFTDYGQIPK (SEQ ID NO:12)), peptide mass, and sequence are shown.
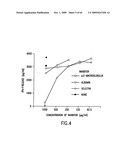
[0043]FIG. 4. α2-Macroglobulin inhibits re-presentation of gp96-chaperoned AH1 peptide by macrophage. The solid cross indicates the level of T cell stimulation when the APCs were pulsed directly with the AH1 peptide. The open cross indicates the corresponding value with unpulsed APCs.
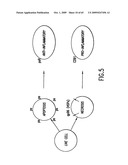
[0044]FIG. 5. α2M receptor is a sensor of necrotic cell death due to its ability to detect extracellular gp96. Conversely, receptors (psR) for phosphatidyl serine (ps) detect apoptotic cell death.
[0045]FIG. 6A-B. A. The mouse α2MR cDNA (SEQ ID NO:1) and predicted open reading frame of murine α2MR protein (Genbank accession no. CAA47817). B. The murine α2MR protein (SEQ ID NO:2), with residues identified by microsequencing an 80 kDa, gp96-interacting fragment of the receptor highlighted in bold.
[0046]FIG. 7A-B. A. Translated amino acid sequence of α2M (SEQ ID NO: 3) and nucleotide sequence of α2M (SEQ ID NO: 4).). B. Amino acid sequence of mature α2M (SEQ ID NO.: 8). The 138 amino acid sequence (SEQ ID NO.: 5) of the receptor binding domain from α2M is highlighted.
5. DETAILED DESCRIPTION OF THE INVENTION
[0047]The present invention provides compositions and methods for alpha (2) macroglobulin ("α2M") vaccines for use in immunotherapy. The invention is based, in part, on the Applicant's discovery that α2M blocks uptake of heat shock proteins by antigen presenting cells. In particular, the invention provides complexes of α2M associated with antigenic molecules, which are recognized by the alpha (2) macroglobulin receptor on antigen presenting cells ("APCs"), and are presented by such cells to the immune system. Thus, the invention provides methods and compositions for using specific α2M-antigenic molecule complexes for targeting an immune response against immune disorders, proliferative disorders, and infectious diseases.
[0048]The human plasma protein alpha (2) macroglobulin is a 720 kDa homotetrameric proteinase inhibitor primarily known as proteinase inhibitor and plasma and inflammatory fluid proteinase scavenger molecule (for review see Chu and Pizzo, 1994, Lab. Invest. 71:792). During proteolytic activation of α2M, non-proteolytic ligands can become incorporated, covalently and noncovalently, to the activated thioesters (see Osada et al., 1987, Biochem. Biophys. Res. Comm. 146:26-31; Osada et al., 1988, Biochem. Biophys. Res. Comm. 150:883-889; Chu and Pizzo, 1993, J. Immunology 150: 48-58; Chu et al., 1994, 152:1538-1545; Mitsuda et al., 1993, Biochem. Biophys. Res. Comm. 191:1326-1331). As described herein, when complexes formed between α2M and an antigenic molecule having the antigenicity of a cancer cell antigen or of a pathogen, such α2M-antigenic molecule complexes can be used to stimulate a cytotoxic T cell response directed against the α2M incorporated antigen. Such complexes can be used as immunotherapeutic agents to treat cancer and infectious diseases.
[0049]Described in detailed hereinbelow are methods and compositions for use in preparation and delivery of such therapeutic α2M-antigenic molecule complexes. The invention encompasses complexes of alpha (2) macroglobulin associated antigenic molecules, antigenic cells that express the α2M, and antibodies and other molecules that specifically recognize α2M-antigenic molecule complexes. The invention also relates to methods for using these compositions in the diagnosis and treatment of immune disorders, proliferative disorders, and infectious diseases.
5.1 Compositions of the Invention
[0050]The present invention provides compositions that can be used in immunotherapy against proliferative disorders, infectious diseases, and immune disorders. Such compositions include antibodies that specifically recognize α2M complexes, isolated antigenic cells that express α2M complexes, and recombinant cells that contain recombinant α2M and sequences encoding antigenic molecules.
[0051]It is contemplated that the definition of α2M, as used herein, embraces other polypeptide fragments, analogs, and variants of α2M having at least 35% to 55%, preferably 55% to 75%, and most preferably 75% to 85% amino acid identity with α2M, and is capable of forming a complex with an antigenic molecule, which complex is capable of being taken up by an antigen presenting cell and eliciting an immune response against the antigenic molecule. The determination of percent identity between two sequences can also be accomplished using a mathematical algorithm. A preferred, non-limiting example of a mathematical algorithm utilized for the comparison of two sequences is the algorithm of Karlin and Altschul, 1990, Proc. Natl. Acad. Sci. USA 87:2264-2268, modified as in Karlin and Altschul, 1993, Proc. Natl. Acad. Sci. USA 90:5873-5877. Such an algorithm is incorporated into the NBLAST and XBLAST programs of Altschul, et al., 1990, J. Mol. Biol. 215:403-410. BLAST nucleotide searches can be performed with the NBLAST program, score=100, wordlength=12 to obtain nucleotide sequences homologous to a nucleic acid molecules of the invention. BLAST protein searches can be performed with the XBLAST program, score=50, wordlength=3 to obtain amino acid sequences homologous to a protein molecules of the invention. To obtain gapped alignments for comparison purposes, Gapped BLAST can be utilized as described in Altschul et al., 1997, Nucleic Acids Res. 25:3389-3402. Alternatively, PSI-Blast can be used to perform an iterated search which detects distant relationships between molecules (Altschul et al., 1997, supra). When utilizing BLAST, Gapped BLAST, and PSI-Blast programs, the default parameters of the respective programs (e.g., XBLAST and NBLAST) can be used. Another preferred, non-limiting example of a mathematical algorithm utilized for the comparison of sequences is the algorithm of Myers and Miller, 1988, CABIOS 4:11-17. Such an algorithm is incorporated into the ALIGN program (version 2.0) which is part of the GCG sequence alignment software package. When utilizing the ALIGN program for comparing amino acid sequences, a PAM120 weight residue table, a gap length penalty of 12, and a gap penalty of 4 can be used.
[0052]The immunogenic α2M-antigenic molecule complexes of the invention may include any complex containing an α2M and an antigenic peptide that is capable of inducing an immune response in a mammal.
[0053]α2M and/or antigenic molecules can be purified from natural sources, chemically synthesized, or recombinantly produced.
[0054]5.1.1. α2M Polypeptides
[0055]The alpha (2) macroglobulin complex of the invention is comprised of an alpha (2) macroglobulin polypeptide associated with an antigenic peptide. Alpha (2) macroglobulin polypeptides may be produced by recombinant DNA techniques, synthetic methods, or by enzymatic or chemical cleavage of native α2M polypeptides. Described herein are methods for producing such α2M polypeptides.
5.1.1.1 Isolation of α2M Gene Sequences
[0056]In various aspects, the invention relates to compositions comprising amino acid sequences of α2M, and fragments, derivatives, analogs, and variants thereof. Nucleic acids encoding α2M are provided, as well as nucleic acids complementary to and capable of hybridizing to such nucleic acids.
[0057]Any eukaryotic cell may serve as the nucleic acid source for obtaining the coding region of an α2M gene. Nucleic acid sequences encoding α2M can be isolated from vertebrate, mammalian, as well as primate sources, including humans.
[0058]Amino acid sequences and nucleotide sequences of naturally occurring α2M polypeptides are generally available in sequence databases, such as GenBank. Non-limiting examples of α2M sequences that can be used for preparation of the α2M polypeptides of the invention are as follows: Genbank Accession Nos. M11313, P01023, AAA51551; Kan et al., 1985, Proc. Nat. Acad. Sci. 82: 2282-2286. Due to the degeneracy of the genetic code, the term "α2M gene", as used herein, refers not only to the naturally occurring nucleotide sequence but also encompasses all the other degenerate DNA sequences that encode an α2M polypeptide. Computer programs, such as Entrez, can be used to browse the database, and retrieve any amino acid sequence and genetic sequence data of interest by accession number. These databases can also be searched to identify sequences with various degrees of similarities to a query sequence using programs, such as FASTA and BLAST, which rank the similar sequences by alignment scores and statistics. BLAST nucleotide searches can be performed with the NBLAST program, score=100, wordlength=12 to obtain nucleotide sequences homologous to a nucleic acid molecules of the invention. BLAST protein searches can be performed with the XBLAST program, score=50, wordlength=3 to obtain amino acid sequences homologous to a protein molecules of the invention. To obtain gapped alignments for comparison purposes, Gapped BLAST can be utilized as described in Altschul et al., 1997, Nucleic Acids Res. 25:3389-3402. Alternatively, PSI-Blast can be used to perform an iterated search which detects distant relationships between molecules (Altschul et al., 1997, supra). When utilizing BLAST, Gapped BLAST, and PSI-Blast programs, the default parameters of the respective programs (e.g., XBLAST and NBLAST) can be used.
[0059]The DNA may be obtained by standard procedures known in the art by DNA amplification or molecular cloning directly from a tissue, cell culture, or cloned DNA (e.g., a DNA "library"). Clones derived from genomic DNA may contain regulatory and intron DNA regions in addition to coding regions; clones derived from cDNA will contain only exon sequences. Whatever the source, the α2M gene should be cloned into a suitable vector for propagation of the gene.
[0060]In a preferred embodiment, DNA can be amplified from genomic or cDNA by polymerase chain reaction (PCR) amplification using primers designed from the known sequence of a related or homologous α2M. PCR is used to amplify the desired sequence in DNA clone or a genomic or cDNA library, prior to selection. PCR can be carried out, e.g., by use of a thermal cycler and Taq polymerase (sold under the trademark GENE AMP). The DNA being amplified can include cDNA or genomic DNA from any species. Oligonucleotide primers representing known nucleic acid sequences of related HSPs can be used as primers in PCR. In a preferred aspect, the oligonucleotide primers represent at least part of the α2M gene that is highly conserved between α2M genes of different species. One can choose to synthesize several different degenerate primers, for use in the PCR reactions. It is also possible to vary the stringency of hybridization conditions used in priming the PCR reactions, to allow for greater or lesser degrees of nucleotide sequence similarity between the known α2M nucleotide sequence and the nucleic acid homolog being isolated. For cross species hybridization, low stringency conditions are preferred. For same species hybridization, moderately stringent conditions are preferred. After successful amplification, the sequence encoding an α2M may be cloned and sequenced. If the size of the coding region of the α2M gene being amplified is too large to be amplified in a single PCR, several PCR covering the entire gene, preferably with overlapping regions, may be carried out, and the products of the PCR ligated together to form the entire coding sequence. Alternatively, if a segment of an α2M gene is amplified, that segment may be cloned, and utilized as a probe to isolate a complete cDNA or genomic clone.
[0061]In another embodiment, for the molecular cloning of an α2M gene from genomic DNA, DNA fragments are generated to form a genomic library. Since some of the sequences encoding related α2Ms are available and can be purified and labeled, the cloned DNA fragments in the genomic DNA library may be screened by nucleic acid hybridization to the labeled probe (Benton and Davis, 1977, Science 196:180; Grunstein and Hogness, 1975, Proc. Natl. Acad. Sci. U.S.A. 72:3961). Those DNA fragments with substantial homology to the probe will hybridize. It is also possible to identify the appropriate fragment by restriction enzyme digestion(s) and comparison of fragment sizes with those expected according to a known restriction map if such is available.
[0062]Alternatives to isolating the α2M genomic DNA include, but are not limited to, chemically synthesizing the gene sequence itself from a known sequence or making cDNA to the mRNA which encodes α2M. For example, RNA for cDNA cloning of the α2M gene can be isolated from cells which express α2M. A cDNA library may be generated by methods known in the art and screened by methods, such as those disclosed for screening a genomic DNA library. If an antibody to α2M is available, α2M may be identified by binding of labeled antibody to the putatively α2M synthesizing clones.
[0063]Other specific embodiments for the cloning of a nucleotide sequence encoding an α2M, are presented as examples but not by way of limitation, as follows:
[0064]In a specific embodiment, nucleotide sequences encoding α2M proteins within a family can be identified and obtained by hybridization with a probe comprising nucleotide sequence encoding α2M under conditions of low to medium stringency.
[0065]By way of example and not limitation, procedures using such conditions of low stringency are as follows (see also Shilo and Weinberg, 1981, Proc. Natl. Acad. Sci. USA 78:6789-6792). Filters containing DNA are pretreated for 6 h at 40° C. in a solution containing 35% formamide, 5×SSC, 50 mM Tris-HCl (pH 7.5), 5 mM EDTA, 0.1% PVP, 0.1% Ficoll, 1% BSA, and 500 μg/ml denatured salmon sperm DNA. Hybridizations are carried out in the same solution with the following modifications: 0.02% PVP, 0.02% Ficoll, 0.2% BSA, 100 μg/ml salmon sperm DNA, 10% (wt/vol) dextran sulfate, and 5-20×106 cpm 32P-labeled probe is used. Filters are incubated in hybridization mixture for 18-20 h at 40° C., and then washed for 1.5 h at 55° C. in a solution containing 2×SSC, 25 mM Tris-HCl (pH 7.4), 5 mM EDTA, and 0.1% SDS. The wash solution is replaced with fresh solution and incubated an additional 1.5 h at 60° C. Filters are blotted dry and exposed for autoradiography. If necessary, filters are washed for a third time at 65-68° C. and reexposed to film. Other conditions of low stringency which may be used are well known in the art (e.g., as employed for cross-species hybridizations).
[0066]An α2M gene fragment can be inserted into an appropriate cloning vector and introduced into host cells so that many copies of the gene sequence are generated. A large number of vector-host systems known in the art may be used such as, but not limited to, bacteriophages such as lambda derivatives, or plasmids such as pBR322 or pUC plasmid derivatives or the Bluescript vector (Stratagene).
[0067]Any technique for mutagenesis known in the art can be used to modify individual nucleotides in a DNA sequence, for purpose of making amino acid substitution(s) in the expressed peptide sequence, or for creating/deleting restriction sites to facilitate further manipulations. Such techniques include but are not limited to, chemical mutagenesis, in vitro site-directed mutagenesis (Hutchinson et al., 1978, J. Biol. Chem. 253:6551), oligonucleotide-directed mutagenesis (Smith, 1985, Ann. Rev. Genet. 19:423-463; Hill et al., 1987, Methods Enzymol. 155:558-568), PCR-based overlap extension (Ho et al., 1989, Gene 77:51-59), PCR-based megaprimer mutagenesis (Sarkar et al., 1990, Biotechniques, 8:404-407), etc. Modifications can be confirmed by double stranded dideoxy DNA sequencing.
[0068]The polymerase chain reaction (PCR) is commonly used for obtaining genes or gene fragments of interest. For example, a nucleotide sequence encoding α2M polypeptide of any desired length can be generated using PCR primers that flank the nucleotide sequence encoding α2M, or the peptide-binding domain thereof. Alternatively, an α2M gene sequence can be cleaved at appropriate sites with restriction endonuclease(s) if such sites are available, releasing a fragment of DNA encoding α2M, or the peptide-binding domain thereof. If convenient restriction sites are not available, they may be created in the appropriate positions by site-directed mutagenesis and/or DNA amplification methods known in the art (see, for example, Shankarappa et al., 1992, PCR Method Appl. 1:277-278). The DNA fragment that encodes α2M, or the peptide-binding domain thereof, is then isolated, and ligated into an appropriate expression vector, care being taken to ensure that the proper translation reading frame is maintained.
[0069]Alpha (2) macroglobulin polypeptides of the invention may be expressed as fusion proteins to facilitate recovery and purification from the cells in which they are expressed. For example, an α2M polypeptide may contain a signal sequence leader peptide to direct its translocation across the ER membrane for secretion into culture medium. Further, an α2M polypeptide may contain an affinity label, such as a affinity label, fused to any portion of the α2M polypeptide not involved in binding antigenic peptide, such as for example, the carboxyl terminal. The affinity label can be used to facilitate purification of the protein, by binding to an affinity partner molecule.
[0070]Various methods for production of such fusion proteins are well known in the art. The manipulations which result in their production can occur at the gene or protein level, preferably at the gene level. For example, the cloned coding region of an α2M polypeptide may be modified by any of numerous recombinant DNA methods known in the art (Sambrook et al., 1990, Molecular Cloning, A Laboratory Manual, 2d ed., Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.; Ausubel et al., in Chapter 8 of Current Protocols in Molecular Biology, Greene Publishing Associates and Wiley Interscience, New York). It will be apparent from the following discussion that substitutions, deletions, insertions, or any combination thereof are introduced or combined to arrive at a final nucleotide sequence encoding an α2M polypeptide.
[0071]In various embodiments, fusion proteins comprising the α2M polypeptide may be made using recombinant DNA techniques. For example, a recombinant gene encoding an α2M polypeptide may be constructed by introducing an α2M gene fragment in the proper reading frame into a vector containing the sequence of an affinity label, such that the α2M polypeptide is expressed as a peptide-tagged fusion protein. Affinity labels, which may be recognized by specific binding partners, may be used for affinity purification of the α2M polypeptide.
[0072]In a preferred embodiment, the affinity label is fused at its amino terminal to the carboxyl terminal of α2M. The precise site at which the fusion is made in the carboxyl terminal is not critical. The optimal site can be determined by routine experimentation.
[0073]A variety of affinity labels known in the art may be used, such as, but not limited to, the immunoglobulin constant regions, polyhistidine sequence (Petty, 1996, Metal-chelate affinity chromatography, in Current Protocols in Molecular Biology, Vol. 2, Ed. Ausubel et al., Greene Publish. Assoc. & Wiley Interscience), glutathione S-transferase (GST; Smith, 1993, Methods Mol. Cell. Bio. 4:220-229), the E. coli maltose binding protein (Guan et al., 1987, Gene 67:21-30), and various cellulose binding domains (U.S. Pat. Nos. 5,496,934; 5,202,247; 5,137,819; Tomme et al., 1994, Protein Eng. 7:117-123), etc. Other affinity labels may impart fluorescent properties to an α2M polypeptide, e.g., portions of green fluorescent protein and the like. Other possible affinity labels are short amino acid sequences to which monoclonal antibodies are available, such as but not limited to the following well known examples, the FLAG epitope, the myc epitope at amino acids 408-439, the influenza virus hemagglutinin (HA) epitope. Other affinity labels are recognized by specific binding partners and thus facilitate isolation by affinity binding to the binding partner which can be immobilized onto a solid support. Some affinity labels may afford the α2M polypeptide novel structural properties, such as the ability to form multimers. Dimerization of an α2M polypeptide with a bound peptide may increase avidity of interaction between the α2M polypeptide and its partner in the course of antigen presentation. These affinity labels are usually derived from proteins that normally exist as homopolymers. Affinity labels such as the extracellular domains of CD8 (Shiue et al., 1988, J. Exp. Med. 168:1993-2005), or CD28 (Lee et al., 1990, J. Immunol. 145:344-352), or portions of the immunoglobulin molecule containing sites for interchain disulfide bonds, could lead to the formation of multimers. As will be appreciated by those skilled in the art, many methods can be used to obtain the coding region of the above-mentioned affinity labels, including but not limited to, DNA cloning, DNA amplification, and synthetic methods. Some of the affinity labels and reagents for their detection and isolation are available commercially.
[0074]A preferred affinity label is a non-variable portion of the immunoglobulin molecule. Typically, such portions comprise at least a functionally operative CH2 and CH3 domain of the constant region of an immunoglobulin heavy chain. Fusions are also made using the carboxyl terminus of the Fc portion of a constant domain, or a region immediately amino-terminal to the CH1 of the heavy or light chain. Suitable immunoglobulin-based affinity label may be obtained from IgG-1, -2, -3, or -4 subtypes, IgA, IgE, IgD, or IgM, but preferably IgG1. Preferably, a human immunoglobulin is used when the α2M polypeptide is intended for in vivo use for humans. Many DNA encoding immunoglobulin light or heavy chain constant regions is known or readily available from cDNA libraries. See, for example, Adams et al., Biochemistry, 1980, 19:2711-2719; Gough et al., 1980, Biochemistry, 19:2702-2710; Dolby et al., 1980, Proc. Natl. Acad. Sci. U.S.A., 77:6027-6031; Rice et al., 1982, Proc. Natl. Acad. Sci. U.S.A., 79:7862-7865; Falkner et al., 1982, Nature, 298:286-288; and Morrison et al., 1984, Ann. Rev. Immunol, 2:239-256. Because many immunological reagents and labeling systems are available for the detection of immunoglobulins, the α2M polypeptide-Ig fusion protein can readily be detected and quantified by a variety of immunological techniques known in the art, such as the use of enzyme-linked immunosorbent assay (ELISA), immunoprecipitation, fluorescence activated cell sorting (FACS), etc. Similarly, if the affinity label is an epitope with readily available antibodies, such reagents can be used with the techniques mentioned above to detect, quantitate, and isolate the α2M polypeptide containing the affinity label. In many instances, there is no need to develop specific antibodies to the α2M polypeptide.
[0075]A particularly preferred embodiment is a fusion of an α2M polypeptide to the hinge, the CH2 and CH3 domains of human immunoglobulin G-1 (IgG-1; see Bowen et al., 1996, J. Immunol. 156:442-49). This hinge region contains three cysteine residues which are normally involved in disulfide bonding with other cysteines in the Ig molecule. Since none of the cysteines are required for the peptide to function as a tag, one or more of these cysteine residues may optionally be substituted by another amino acid residue, such as for example, serine.
[0076]Various leader sequences known in the art can be used for the efficient secretion of α2M polypeptide from bacterial and mammalian cells (von Heijne, 1985, J. Mol. Biol. 184:99-105). Leader peptides are selected based on the intended host cell, and may include bacterial, yeast, viral, animal, and mammalian sequences. For example, the herpes virus glycoprotein D leader peptide is suitable for use in a variety of mammalian cells. A preferred leader peptide for use in mammalian cells can be obtained from the V-J2-C region of the mouse immunoglobulin kappa chain (Bernard et al., 1981, Proc. Natl. Acad. Sci. 78:5812-5816). Preferred leader sequences for targeting α2M polypeptide expression in bacterial cells include, but are not limited to, the leader sequences of the E. coli proteins OmpA (Hobom et al., 1995, Dev. Biol. Stand. 84:255-262), Pho A (Oka et al., 1985, Proc. Natl. Acad. Sci. 82:7212-16), OmpT (Johnson et al., 1996, Protein Expression 7:104-113), LamB and OmpF (Hoffman & Wright, 1985, Proc. Natl. Acad. Sci. USA 82:5107-5111), β-lactamase (Kadonaga et al., 1984, J. Biol. Chem. 259:2149-54), enterotoxins (Morioka-Fujimoto et al., 1991, J. Biol. Chem. 266:1728-32), and the Staphylococcus aureus protein A (Abrahmsen et al., 1986, Nucleic Acids Res. 14:7487-7500), and the B. subtilis endoglucanase (Lo et al., Appl. Environ. Microbiol. 54:2287-2292), as well as artificial and synthetic signal sequences (Maclntyre et al., 1990, Mol. Gen. Genet. 221:466-74; Kaiser et al., 1987, Science, 235:312-317).
[0077]DNA sequences encoding a desired affinity label or leader peptide, which may be readily obtained from libraries, produced synthetically, or may be available from commercial suppliers, are suitable for the practice of this invention. Such methods are well known in the art.
[0078]5.1.1.2 Recombinant Expression
[0079]In various embodiments of the invention, sequences encoding an α2M polypeptide are inserted into an expression vector for propagation and expression in recombinant cells.
[0080]An expression construct, as used herein, refers to a nucleotide sequence encoding an α2M polypeptide operably associated with one or more regulatory regions which allows expression of the α2M polypeptide in an appropriate host cell. "Operably-associated" refers to an association in which the regulatory regions and the α2M polypeptide sequence to be expressed are joined and positioned in such a way as to permit transcription, and ultimately, translation.
[0081]Vectors based on E. coli are the most popular and versatile systems for high level expression of foreign proteins (Makrides, 1996, Microbiol Rev, 60:512-538). Non-limiting examples of regulatory regions that can be used for expression in E. coli may include but not limited to lac, trp, lpp, phoa, recA, tac, XPL, and phage T3 and T7 promoters (Makrides, 1996, Microbiol Rev, 60:512-538). Non-limiting examples of prokaryotic expression vectors may include the λgt vector series such as λgt11 (Huynh et al., 1984 in "DNA Cloning Techniques", Vol. I: A Practical Approach (D. Glover, ed.), pp. 49-78, IRL Press, Oxford), and the pET vector series (Studier et al., 1990, Methods Enzymol., 185:60-89). However, a potential drawback of a prokaryotic host-vector system is the inability to perform many of the post-translational processing events of mammalian cells. Thus, an eukaryotic host-vector system is preferred, a mammalian host-vector system is more preferred, and a human host-vector system is the most preferred.
[0082]The regulatory regions necessary for transcription of the α2M polypeptide can be provided by the expression vector. A translation initiation codon (ATG) may also be provided to express a nucleotide sequence encoding an α2M polypeptide that lacks an initiation codon. In a compatible host-construct system, cellular proteins required for transcription, such as RNA polymerase and transcription factors, will bind to the regulatory regions on the expression construct to effect transcription of the α2M polypeptide sequence in the host organism. The precise nature of the regulatory regions needed for gene expression may vary from host cell to host cell. Generally, a promoter is required which is capable of binding RNA polymerase to initiate the transcription of an operably-associated nucleic acid sequence. Such regulatory regions may include those 5'-non-coding sequences involved with initiation of transcription and translation, such as the TATA box, the cap site, a CAAT box, and the like. The non-coding region 3' to the coding sequence may contain transcriptional termination regulatory sequences, such as terminators and polyadenylation sites.
[0083]Both constitutive and inducible regulatory regions may be used for expression of the α2M polypeptide. It may be desirable to use inducible promoters when the conditions optimal for growth of the recombinant cells and the conditions for high level expression of the α2M polypeptide are different. Examples of useful regulatory regions are provided in the next section below.
[0084]For expression of α2M polypeptides in mammalian host cells, a variety of regulatory regions can be used, for example, the SV40 early and late promoters, the cytomegalovirus (CMV) immediate early promoter, and the Rous sarcoma virus long terminal repeat (RSV-LTR) promoter. Inducible promoters that may be useful in mammalian cells include but are not limited to those associated with the metallothionein II gene, mouse mammary tumor virus glucocorticoid responsive long terminal repeats (MMTV-LTR), the β-interferon gene, and the α2M70 gene (Williams et al., 1989, Cancer Res. 49:2735-42; Taylor et al., 1990, Mol. Cell. Biol., 10:165-75).
[0085]The following animal regulatory regions, which exhibit tissue specificity and have been utilized in transgenic animals, can also be used in tumor cells of a particular tissue type: elastase I gene control region which is active in pancreatic acinar cells (Swift et al., 1984, Cell 38:639-646; Ornitz et al., 1986, Cold Spring Harbor Symp. Quant. Biol. 50:399-409; MacDonald, 1987, Hepatology 7:425-515); insulin gene control region which is active in pancreatic beta cells (Hanahan, 1985, Nature 315:115-122), immunoglobulin gene control region which is active in lymphoid cells (Grosschedl et al., 1984, Cell 38:647-658; Adames et al., 1985, Nature 318:533-538; Alexander et al., 1987, Mol. Cell. Biol. 7:1436-1444), mouse mammary tumor virus control region which is active in testicular, breast, lymphoid and mast cells (Leder et al., 1986, Cell 45:485-495), albumin gene control region which is active in liver (Pinkert et al., 1987, Genes and Devel. 1:268-276), alpha-fetoprotein gene control region which is active in liver (Krumlauf et al., 1985, Mol. Cell. Biol. 5:1639-1648; Hammer et al., 1987, Science 235:53-58; alpha 1-antitrypsin gene control region which is active in the liver (Kelsey et al., 1987, Genes and Devel. 1:161-171), beta-globin gene control region which is active in myeloid cells (Mogram et al., 1985, Nature 315:338-340; Kollias et al., 1986, Cell 46:89-94; myelin basic protein gene control region which is active in oligodendrocyte cells in the brain (Readhead et al., 1987, Cell 48:703-712); myosin light chain-2 gene control region which is active in skeletal muscle (Sani, 1985, Nature 314:283-286), and gonadotropic releasing hormone gene control region which is active in the hypothalamus (Mason et al., 1986, Science 234:1372-1378).
[0086]The efficiency of expression of the α2M polypeptide in a host cell may be enhanced by the inclusion of appropriate transcription enhancer elements in the expression vector, such as those found in SV40 virus, Hepatitis B virus, cytomegalovirus, immunoglobulin genes, metallothionein, β-actin (see Bittner et al., 1987, Methods in Enzymol. 153:516-544; Gorman, 1990, Curr. Op. in Biotechnol. 1:36-47).
[0087]The expression vector may also contain sequences that permit maintenance and replication of the vector in more than one type of host cell, or integration of the vector into the host chromosome. Such sequences may include but are not limited to replication origins, autonomously replicating sequences (ARS), centromere DNA, and telomere DNA. It may also be advantageous to use shuttle vectors that can be replicated and maintained in at least two types of host cells.
[0088]In addition, the expression vector may contain selectable or screenable marker genes for initially isolating or identifying host cells that contain DNA encoding an α2M polypeptide. For long term, high yield production of α2M polypeptide-antigenic molecule complexes, stable expression in mammalian cells is preferred. A number of selection systems may be used for mammalian cells, including, but not limited, to the Herpes simplex virus thymidine kinase (Wigler et al., 1977, Cell 11:223), hypoxanthine-guanine phosphoribosyltransferase (Szybalski and Szybalski, 1962, Proc. Natl. Acad. Sci. USA 48:2026), and adenine phosphoribosyltransferase (Lowy et al., 1980, Cell 22:817) genes can be employed in tk.sup.-, hgprt.sup.- or aprt.sup.- cells, respectively. Also, antimetabolite resistance can be used as the basis of selection for dihydrofolate reductase (dhfr), which confers resistance to methotrexate (Wigler et al., 1980, Natl. Acad. Sci. USA 77:3567; O'Hare et al., 1981, Proc. Natl. Acad. Sci. USA 78:1527); gpt, which confers resistance to mycophenolic acid (Mulligan & Berg, 1981, Proc. Natl. Acad. Sci. USA 78:2072); neomycin phosphotransferase (neo), which confers resistance to the aminoglycoside G-418 (Colberre-Garapin et al., 1981, J. Mol. Biol. 150:1); and hygromycin phosphotransferase (hyg), which confers resistance to hygromycin (Santerre et al., 1984, Gene 30:147). Other selectable markers, such as but not limited to histidinol and Zeocin® can also be used.
[0089]In order to insert the α2M polypeptide DNA sequence into the cloning site of a vector, DNA sequences with regulatory functions, such as promoters, must be attached to DNA sequences encoding the α2M peptide-binding region. To do this, linkers or adapters providing the appropriate compatible restriction sites may be ligated to the ends of cDNA or synthetic DNA encoding an α2M polypeptide, by techniques well known in the art (Wu et al., 1987, Methods in Enzymol 152:343-349). Cleavage with a restriction enzyme can be followed by modification to create blunt ends by digesting back or filling in single-stranded DNA termini before ligation. Alternatively, a desired restriction enzyme site can be introduced into a fragment of DNA by amplification of the DNA by use of PCR with primers containing the desired restriction enzyme site.
[0090]An expression construct comprising an α2M polypeptide sequence operably associated with regulatory regions can be directly introduced into appropriate host cells for expression and production of α2M polypeptide-antigenic molecule complexes without further cloning (see, for example, U.S. Pat. No. 5,580,859). The expression constructs may also contain DNA sequences that facilitate integration of the α2M polypeptide sequence into the genome of the host cell, e.g., via homologous recombination. In this instance, it is not necessary to employ an expression vector comprising a replication origin suitable for appropriate host cells in order to propagate and express the α2M polypeptide in the host cells.
[0091]Expression constructs containing cloned nucleotide sequence encoding α2M polypeptides can be introduced into the host cell by a variety of techniques known in the art, including but not limited to, for prokaryotic cells, bacterial transformation (Hanahan, 1985, in DNA Cloning, A Practical Approach, 1:109-136), and for eukaryotic cells, calcium phosphate mediated transfection (Wigler et al., 1977, Cell 11:223-232), liposome-mediated transfection (Schaefer-Ridder et al., 1982, Science 215:166-168), electroporation (Wolff et al., 1987, Proc Natl Acad Sci 84:3344), and microinjection (Cappechi, 1980, Cell 22:479-488). Co-expression of an α2M polypeptide and an antigenic molecule in the same host cell can be achieved by essentially the same methods.
[0092]For long term, high yield production of properly processed α2M polypeptides or α2M polypeptide-antigenic molecule complexes, stable expression in mammalian cells is preferred. Cell lines that stably express α2M polypeptides or α2M polypeptide-antigenic molecule complexes may be engineered by using a vector that contains a selectable marker. By way of example but not limitation, following the introduction of the expression constructs, engineered cells may be allowed to grow for 1-2 days in an enriched media, and then are switched to a selective media. The selectable marker in the expression construct confers resistance to the selection and optimally allows cells to stably integrate the expression construct into their chromosomes and to grow in culture and to be expanded into cell lines. Such cells can be cultured for a long period of time while α2M polypeptide is expressed continuously.
[0093]The recombinant cells may be cultured under standard conditions of temperature, incubation time, optical density, and media composition. Alternatively, a recombinant antigenic cells may be cultured under conditions emulating the nutritional and physiological requirements of the cancer cell or infected cell. However, conditions for growth of recombinant cells may be different from those for expression of α2M polypeptides and antigenic proteins. Modified culture conditions and media may also be used to enhance production of α2M-antigenic molecule complexes. Any techniques known in the art may be applied to establish the optimal conditions for producing α2M polypeptide or α2M polypeptide-antigenic molecule complexes.
[0094]5.1.1.3 Purification Methods for Recombinant α2M Polypeptides
[0095]Generally, the α2M polypeptides of the invention can be recovered and purified from recombinant cell cultures by known methods, including ammonium sulfate precipitation, acid extraction, anion or cation exchange chromatography, phosphocellulose chromatography, immunoaffinity chromatography, hydroxyapatite chromatography, and lectin chromatography.
[0096]The invention provides methods for purification of recombinant α2M polypeptides by affinity purification, based on the properties of the affinity label present on the α2M polypeptide. One approach is based on specific molecular interactions between a tag and its binding partner. The other approach relies on the immunospecific binding of an antibody to an epitope present on the tag. The principle of affinity chromatography well known in the art is generally applicable to both of these approaches.
[0097]Described below are several methods based on specific molecular interactions of a tag and its binding partner.
[0098]A method that is generally applicable to purifying recombinant α2Ms that are fused to the constant regions of immunoglobulin is protein A affinity chromatography, a technique that is well known in the art. Staphylococcus protein A is a 42 kD polypeptide that binds specifically to a region located between the second and third constant regions of heavy chain immunoglobulins. Because of the Fc domains of different classes, subclasses and species of immunoglobulins, affinity of protein A for human Fc regions is strong, but may vary with other species. Subclasses that are less preferred include human IgG-3, and most rat subclasses. For certain subclasses, protein G (of Streptococci) may be used in place of protein A in the purification. Protein-A sepharose (Pharmacia or Biorad) is a commonly used solid phase for affinity purification of antibodies, and can be used essentially in the same manner for the purification of α2M polypeptide fused to an immunoglobulin Fc fragment. Secreted α2M polypeptide present in cell supernatant binds specifically to protein A on the solid phase, while the contaminants are washed away. Bound α2M polypeptide can be eluted by various buffer systems known in the art, including a succession of citrate, acetate and glycine-HCl buffers which gradually lowers the pH. This method is less preferred if the recombinant cells also produce antibodies which will be copurified with the α2M polypeptide. See, for example, Langone, 1982, J. Immunol. meth. 51:3; Wilchek et al., 1982, Biochem. Intl. 4:629; Sjobring et al., 1991, J. Biol. Chem. 26:399; page 617-618, in Antibodies A Laboratory Manual, edited by Harlow and Lane, Cold Spring Harbor laboratory, 1988.
[0099]Alternatively, a polyhistidine tag may be used, in which case, the α2M polypeptide can be purified by metal chelate chromatography. The polyhistidine tag, usually a sequence of six histidines, has a high affinity for divalent metal ions, such as nickel ions (Ni2+), which can be immobilized on a solid phase, such as nitrilotriacetic acid matrices. Polyhistidine has a well characterized affinity for Ni2+-NTA-agarose, and can be eluted with either of two mild treatments: imidazole (0.1-0.2 M) will effectively compete with the resin for binding sites; or lowering the pH just below 6.0 will protonate the histidine side-chains and disrupt the binding. The purification method comprises loading the cell culture supernatant onto the Ni2+-NTA-agarose column, washing the contaminants through, and eluting the α2M polypeptide with imidazole or weak acid. Ni2+-NTA-agarose can be obtained from commercial suppliers such as Sigma (St. Louis) and Qiagen. Antibodies that recognize the polyhistidine tag are also available which can be used to detect and quantify the α2M polypeptide.
[0100]Another exemplary affinity label that can be used is the glutathione-S-transferase (GST) sequence, originally cloned from the helminth, Schistosoma japonicum. In general, an α2M-GST fusion expressed in a prokaryotic host cell, such as E. coli, can be purified from the cell culture supernatant by absorption with glutathione agarose beads, followed by elution in the presence of free reduced glutathione at neutral pH. Denaturing conditions are not required at any stage during purification, and therefore, it may be desirable for use in the loading of immobilized α2M polypeptides with antigenic peptides. Moreover, since GST is known to form dimers under certain conditions, dimeric α2M polypeptides may be obtained. See, Smith, 1993, Methods Mol. Cell. Bio. 4:220-229.
[0101]Another useful affinity label that can be used is the maltose binding protein (MBP) of E. coli, which is encoded by the malE gene. The secreted α2M polypeptide-MBP present in the cell supernatant binds to amylose resin while contaminants are washed away. The bound α2M polypeptide-MBP is eluted from the amylose resin by maltose. See, for example, Guan et al., 1987, Gene 67:21-30.
[0102]The second approach for purifying α2M polypeptide is applicable to affinity labels that contain an epitope for which polyclonal or monoclonal antibodies are available. Various methods known in the art for purification of protein by immunospecific binding, such as immunoaffinity chromatography, and immunoprecipitation, can be used. See, for example, Chapter 13 in "Antibodies A Laboratory Manual", 1988, Harlow and Lane, (eds.), Cold Spring Harbor Laboratory, N.Y. and Chapter 8, Sections I and II, in "Current Protocols in Immunology", 1991, Coligan et al. (eds.), John Wiley, the disclosure of which are both incorporated by reference herein.
[0103]The embodiments described above may be used to recover and purify α2M polypeptide-antigenic molecule complexes from the cell culture medium of mammalian cells, such as human cells expressing an α2M polypeptide of the invention. The methods can be adapted to perform medium and large scale purification of an α2M polypeptide and/or α2M-antigenic molecule complexes. Methods that do not require lowering pH or denaturing conditions are most preferred for purification of α2M polypeptide-antigenic molecule complexes. The methods may be used to isolate α2M polypeptides from eukaryotic cells, for example, cancer cells, tissues, isolated cells, or immortalized eukaryote cell lines infected with an intracellular pathogen, or cells obtained from a subject infected with a pathogen.
[0104]5.1.1.4 Host-Vector Systems
[0105]Described herein are systems of vectors and host cells that can be used for the expression of α2M polypeptides. A variety of expression vectors may be used in the present invention which include, but are not limited to, plasmids, cosmids, phage, phagemids, or modified viruses. Typically, such expression vectors comprise a functional origin of replication for propagation of the vector in an appropriate host cell, one or more restriction endonuclease sites for insertion of the α2M polypeptide gene sequence, and one or more selection markers. The expression vector must be used with a compatible host cell which may be derived from a prokaryotic or an eukaryotic organism including but not limited to bacteria, yeasts, insects, mammals, and humans.
[0106]Expression constructs and vectors are introduced into host cells for the purpose of producing an α2M polypeptide. Any cell type that can produce α2Ms and is compatible with the expression vector may be used, including those that have been cultured in vitro or genetically engineered. Host cells may be obtained from normal or affected subjects, including healthy humans, cancer patients, and patients with an infectious disease, private laboratory deposits, public culture collections such as the American Type Culture Collection, or from commercial suppliers.
[0107]Different host cells have characteristic and specific mechanisms for the post-translational processing and modification of proteins. A host cell may be chosen which modifies and processes the expressed gene products in a specific fashion similar to the way the recipient processes α2Ms. For the purpose of producing large amounts of α2M, it is preferable that the type of host cell used in the present invention has been used for expression of heterologous genes, and is reasonably well characterized and developed for large-scale production processes. In a specific embodiment, the host cells are from the same patient to whom α2M polypeptide-antigenic molecule complexes or recombinant cells expressing α2M polypeptide-antigenic molecule complexes are going to be administered. Otherwise said, the cells used to express the α2M polypeptide and used subsequently to administer immunotherapy to a subject are autologous to the subject.
[0108]Preferred mammalian host cells include but are not limited to those derived from humans, monkeys and rodents, (see, for example, Kriegler M. in "Gene Transfer and Expression: A Laboratory Manual", New York, Freeman & Co. 1990), such as monkey kidney cell line transformed by SV40 (COS-7, ATCC CRL 1651); human embryonic kidney line (293, 293-EBNA, or 293 cells subcloned for growth in suspension culture, Graham et al., J. Gen. Virol., 36:59, 1977; baby hamster kidney cells (BHK, ATCC CCL 10); chinese hamster ovary-cells-DHFR (CHO, Urlaub and Chasin. Proc. Natl. Acad. Sci. 77; 4216, 1980); mouse sertoli cells (Mather, Biol. Reprod. 23:243-251, 1980); mouse fibroblast cells (NIH-3T3), monkey kidney cells (CV1 ATCC CCL 70); african green monkey kidney cells (VERO-76, ATCC CRL-1587); human cervical carcinoma cells (HELA, ATCC CCL 2); canine kidney cells (MDCK, ATCC CCL 34); buffalo rat liver cells (BRL 3A, ATCC CRL 1442); human lung cells (WI 38, ATCC CCL 75); human liver cells (Hep G2, HB 8065); and mouse mammary tumor cells (MMT 060562, ATCC CCL51). Exemplary cancer cell types used for demonstrating the utility of recombinant cells (producing α2M polypeptide-antigenic molecule complexes) as a cancer vaccine are provided as follows: mouse fibroblast cell line, NIH3T3, mouse Lewis lung carcinoma cell line, LLC, mouse mastocytoma cell line, P815, mouse lymphoma cell line, EL4 and its ovalbumin transfectant, E.G7, mouse melanoma cell line, B16F10, mouse fibrosarcoma cell line, MC57, and human small cell lung carcinoma cell lines, SCLC#2 and SCLC#7.
[0109]A number of viral-based expression systems may also be utilized with mammalian cells to produce α2M polypeptides. Vectors using DNA virus backbones have been derived from simian virus 40 (SV40) (Hamer et al., 1979, Cell 17:725), adenovirus (Van Doren et al., 1984, Mol Cell Biol 4:1653), adeno-associated virus (McLaughlin et al., 1988, J Virol 62:1963), and bovine papillomas virus (Zinn et al., 1982, Proc Natl Acad Sci 79:4897). In cases where an adenovirus is used as an expression vector, the donor DNA sequence may be ligated to an adenovirus transcription/translation control region, e.g., the late promoter and tripartite leader sequence. This chimeric gene may then be inserted in the adenovirus genome by in vitro or in vivo recombination. Insertion in a non-essential region of the viral genome (e.g., region E1 or E3) will result in a recombinant virus that is viable and capable of expressing heterologous products in infected hosts (see e.g., Logan and Shenk, 1984, Proc. Natl. Acad. Sci. (USA) 81:3655-3659).
[0110]Bovine papillomavirus (BPV) can infect many higher vertebrates, including man, and its DNA replicates as an episome. A number of shuttle vectors have been developed for recombinant gene expression which exist as stable, multicopy (20-300 copies/cell) extrachromosomal elements in mammalian cells. Typically, these vectors contain a segment of BPV DNA (the entire genome or a 69% transforming fragment), a promoter with a broad host range, a polyadenylation signal, splice signals, a selectable marker, and "poisonless" plasmid sequences that allow the vector to be propagated in E. coli. Following construction and amplification in bacteria, the expression gene construct is transfected into cultured mammalian cells, for example, by the techniques of calcium phosphate coprecipitation or electroporation. For those host cells that do not manifest a transformed phenotype, selection of transformants is achieved by use of a dominant selectable marker, such as histidinol and G418 resistance. For example, BPV vectors such as pBCMGSNeo and pBCMGHis may be used to express α2M polypeptide sequences (Karasuyama et al., Eur. J. Immunol. 18:97-104; Ohe et al., Human Gene Therapy, 6:325-33) which may then be transfected into a diverse range of cell types for expression of the α2M polypeptide.
[0111]Alternatively, the vaccinia 7.5K promoter may be used (see, e.g., Mackett et al., 1982, Proc. Natl. Acad. Sci. (USA) 79:7415-7419; Mackett et al., 1984, J. Virol. 49:857-864; Panicali et al., 1982, Proc. Natl. Acad. Sci. 79:4927-4931.) In cases where a human host cell is used, vectors based on the Epstein-Barr virus (EBV) origin (OriP) and EBV nuclear antigen 1 (EBNA-1; a trans-acting replication factor) may be used. Such vectors can be used with a broad range of human host cells, e.g., EBO-pCD (Spickofsky et al., 1990, DNA Prot Eng Tech 2:14-18), pDR2 and λDR2 (available from Clontech Laboratories).
[0112]α2M polypeptides may also be made with a retrovirus-based expression system. In contrast to transfection, retroviruses can efficiently infect and transfer genes to a wide range of cell types including, for example, primary hematopoietic cells. In retroviruses such as Moloney murine leukemia virus, most of the viral gene sequences can be removed and replaced with nucleic acid sequences encoding α2M, while the missing viral functions can be supplied in trans. The host range for infection by a retroviral vector can also be manipulated by the choice of envelope used for vector packaging.
[0113]For example, a retroviral vector can comprise a 5' long terminal repeat (LTR), a 3' LTR, a packaging signal, a bacterial origin of replication, and a selectable marker. The α2M polypeptide DNA is inserted into a position between the 5' LTR and 3' LTR, such that transcription from the 5' LTR promoter transcribes the cloned DNA. The 5' LTR comprises a promoter, including but not limited to an LTR promoter, an R region, a U5 region and a primer binding site, in that order. Nucleotide sequences of these LTR elements are well known in the art. A heterologous promoter as well as multiple drug selection markers may also be included in the expression vector to facilitate selection of infected cells (see McLauchlin et al., 1990, Prog. Nucleic Acid Res. and Molec. Biol. 38:91-135; Morgenstern et al., 1990, Nucleic Acid Res. 18:3587-3596; Choulika et al., 1996, J. Virol 70:1792-1798; Boesen et al., 1994, Biotherapy 6:291-302; Salmons and Gunzberg, 1993, Human Gene Therapy 4:129-141; and Grossman and Wilson, 1993, Curr. Opin. in Genetics and Devel. 3:110-114).
[0114]Other useful eukaryotic host-vector system may include yeast and insect systems. In yeast, a number of vectors containing constitutive or inducible promoters may be used with Saccharomyces cerevisiae (baker's yeast), Schizosaccharomyces pombe (fission yeast), Pichia pastoris, and Hansenula polymorpha (methylotropic yeasts). For a review see, "Current Protocols in Molecular Biology", Vol. 2, 1988, Ausubel et al. (eds.), Greene Publish. Assoc. & Wiley Interscience, Ch. 13; Grant et al., 1987, Expression and Secretion Vectors for Yeast, 1987, in "Methods in Enzymology", Wu and Grossman (eds.), Acad. Press, N.Y., Vol. 153, pp. 516-544; Glover, 1986, DNA Cloning, Vol. II, IRL Press, Wash., D.C., Ch. 3; and Bitter, 1987, Heterologous Gene Expression in Yeast, in "Methods in Enzymology", Berger and Kimmel (eds.), Acad. Press, N.Y., Vol. 152, pp. 673-684; and "The Molecular Biology of the Yeast Saccharomyces", 1982, Strathern et al. (eds.), Cold Spring Harbor Press, Vols. I and II.
[0115]In an insect system a baculovirus, Autographa californica nuclear polyhidrosis virus (AcNPV), can be used as a vector to express an α2M polypeptide in Spodoptera frugiperda cells. The α2M polypeptide DNA may be cloned into non-essential regions (for example the polyhedrin gene) of the virus and placed under control of an AcNPV promoter (for example the polyhedrin promoter). These recombinant viruses are then used to infect host cells in which the inserted DNA is expressed (see, e.g., Smith et al., 1983, J. Virol. 46:584; Smith, U.S. Pat. No. 4,215,051).
[0116]Any of the cloning and expression vectors described herein may be synthesized and assembled from known DNA sequences by techniques well known in the art. The regulatory regions and enhancer elements can be of a variety of origins, both natural and synthetic. Some vectors and host cells may be obtained commercially. Non-limiting examples of useful vectors are described in Appendix 5 of Current Protocols in Molecular Biology, 1988, ed. Ausubel et al., Greene Publish. Assoc. & Wiley Interscience, which is incorporated herein by reference; and the catalogs of commercial suppliers such as Clontech Laboratories, Stratagene Inc., and Invitrogen, Inc.
[0117]5.1.1.5 Synthetic Production
[0118]An alternative to producing α2M by recombinant techniques is peptide synthesis. For example, a peptide corresponding to a portion of an α2M comprising the substrate-binding domain, or which binds peptides in vitro, can be synthesized by use of a peptide synthesizer. Conventional peptide synthesis may be used or other synthetic protocols well known in the art.
[0119]In addition, analogs and derivatives of α2M polypeptides can be chemically synthesized. Furthermore, if desired, nonclassical amino acids or chemical amino acid analogs can be introduced as a substitution or addition into the α2M sequence. Non-classical amino acids include but are not limited to the D-isomers of the common amino acids, α-amino isobutyric acid, 4-aminobutyric acid, Abu, 2-amino butyric acid, β-Abu, ε-Ahx, 6-amino hexanoic acid, Aib, 2-amino isobutyric acid, 3-amino propionic acid, ornithine, norleucine, norvaline, hydroxyproline, sarcosine, citrulline, cysteic acid, t-butylglycine, t-butylalanine, phenylglycine, cyclohexylalanine, β-alanine, fluoro-amino acids, designer amino acids such as β-methyl amino acids, Cα-methyl amino acids, Nα-methyl amino acids, and amino acid analogs in general.
[0120]Peptides having α2M amino acid sequences, or a fragment, analog, mutant or derivative thereof, may be synthesized by solid-phase peptide synthesis using procedures similar to those described by Merrifield, 1963, J. Am. Chem. Soc., 85:2149. During synthesis, N-α-protected amino acids having protected side chains are added stepwise to a growing polypeptide chain linked by its C-terminal and to an insoluble polymeric support i.e., polystyrene beads. The peptides are synthesized by linking an amino group of an N-α-deprotected amino acid to an α-carboxyl group of an N-α-protected amino acid that has been activated by reacting it with a reagent such as dicyclohexylcarbodiimide. The attachment of a free amino group to the activated carboxyl leads to peptide bond formation. The most commonly used N-α-protecting groups include Boc which is acid labile and Fmoc which is base labile. Details of appropriate chemistries, resins, protecting groups, protected amino acids and reagents are well known in the art and so are not discussed in detail herein (See, Atherton, et al., 1989, Solid Phase Peptide Synthesis: A Practical Approach, IRL Press, and Bodanszky, 1993, Peptide Chemistry, A Practical Textbook, 2nd Ed., Springer-Verlag).
[0121]Purification of the resulting α2M polypeptides accomplished using conventional procedures, such as preparative HPLC using gel permeation, partition and/or ion exchange chromatography. The choice of appropriate matrices and buffers are well known in the art and so are not described in detail herein.
5.2 Antigenic Complexes Comprising α2M Polypeptides
[0122]5.2.1. Isolation of Intracellular Complexes of α2M Polypeptides with Antigenic Molecules
[0123]Described herein are methods for purifying α2M polypeptides or α2M polypeptide-antigenic molecule complexes of the invention from recombinant cells, and, with minor modifications known in the art, the α2M polypeptide or α2M-antigenic molecule complexes from the cell culture. Recombinant cells include, for example, cells expressing antigenic molecules and recombinantly expressing an α2M polypeptide. Such cells may be derived from a variety of sources, including, but not limited to, cells infected with an infectious agent and cancer cells.
[0124]The invention provides methods for purification of recombinant α2M polypeptide-antigenic molecule complexes by affinity purification, based on the properties of the affinity label present on the α2M polypeptide. One approach is based on specific molecular interactions between a tag and its binding partner. The other approach relies on the immunospecific binding of an antibody to an epitope present on the tag. The principle of affinity chromatography well known in the art is generally applicable to both of these approaches.
[0125]To produce α2M polypeptide-antigenic molecule complexes, a nucleotide sequence encoding an α2M polypeptide can be introduced into a cell. When an antigenic molecule is present in the cell, the α2M polypeptide can associate intracellularly with the antigenic molecule, forming a covalent or a noncovalent complex of α2M polypeptide and the antigenic molecule. Cells into which an α2M polypeptide-encoding nucleotide sequence can be introduced, include, but are not limited to, epithelial cells, endothelial cells, keratinocytes, fibroblasts, muscle cells, hepatocytes; blood cells such as T lymphocytes, B lymphocytes, monocytes, macrophages, neutrophils, eosinophils, megakaryocytes, granulocytes; various stem or progenitor cells, in particular hematopoietic stem or progenitor cells, e.g., as obtained from bone marrow, umbilical cord blood, peripheral blood, fetal liver, etc. The choice of cell type depends on the type of tumor or infectious disease being treated or prevented, and can be determined by one of skill in the art. In a specific embodiment, an expression construct comprising a nucleic acid sequence encoding the α2M polypeptide is introduced into an antigenic cell. As used herein, antigenic cells may include cells that are infected with an infectious agent or pathogen, cells infected with non-infectious or non-pathogenic forms of an infectious agent or pathogen (e.g., by use of a helper infectious agent), cells infected by or engineered to express an attenuated form of an infectious agent or a non-pathogenic or replication-deficient variant of a pathogen, pre-neoplastic cells that are infected with a cancer-causing infectious agent, such as a virus, but which are not yet neoplastic; or antigenic cells that have been exposed to a mutagen or cancer-causing agent, such as, for example DNA-damaging agents, radiation, etc. Other cells that can be used are pre-neoplastic cells which are in transition from a normal to a neoplastic form as characterized by morphology, physiological or biochemical functions. Preferably, the cancer cells and pre-neoplastic cells used in the methods of the invention are of mammalian origin. Mammals contemplated by this aspect of the invention include humans, companion animals (e.g., dogs and cats), livestock animals (e.g., sheep, cattle, goats, pigs and horses), laboratory animals (e.g., mice, rats and rabbits), and captive or free wild animals.
[0126]In various embodiments, any cancer cell, preferably a human cancer cell, can be used in the present methods for producing α2M polypeptide-antigenic molecule complexes. The cancer cells provide the antigenic peptides which become associated covalently or noncovalently with the expressed α2M polypeptide. α2M polypeptide-antigenic molecule complexes are then purified from the cells and used to treat such cancers. Cancers which can be treated or prevented with immunogenic compositions prepared by methods of the invention include, but are not limited to, tumors such as sarcomas and carcinomas. Examples of cancers that are amenable to the methods of the invention are listed in Section 5.6. Accordingly, any tissues or cells isolated from a pre-neoplastic lesion, a cancer, including cancer that has metastasized to multiple remote sites, can be used in the present method. For example, cells found in abnormally growing tissue, circulating leukemic cells, metastatic lesions as well as solid tumor tissue can be used.
[0127]In another embodiment, cell lines derived from a pre-neoplastic lesion, cancer tissues or cancer cells can also be used, provided that the cells of the cell line have at least one or more antigenic determinants in common with antigens on the target cancer cells. Cancer tissues, cancer cells, cells infected with a cancer-causing agent, other pre-neoplastic cells, and cell lines of human origin are preferred.
[0128]Cancer and pre-neoplastic cells can be identified by any method known in the art. For example, cancer cells can be identified by morphology, enzyme assays, proliferation assays, cytogenetic characterization, DNA mapping, DNA sequencing, the presence of cancer-causing virus, or a history of exposure to mutagen or cancer-causing agent, imaging, etc. Cancer cells may also be obtained by surgery, endoscopy, or other biopsy techniques. If some distinctive characteristics of the cancer cells are known, they can also be obtained or purified by any biochemical or immunological methods known in the art, such as but not limited to affinity chromatography, and fluorescence activated cell sorting (e.g., with fluorescently tagged antibody against an antigen expressed by the cancer cells).
[0129]Cancer tissues, cancer cells or cell lines may be obtained from a single individual or pooled from several individuals. It is not essential that clonal, homogeneous, or purified population of cancer cells be used. It is also not necessary to use cells of the ultimate target in vivo (e.g., cells from the tumor of the intended recipient), so long as at least one or more antigenic determinants on the target cancer cells is present on the cells used for expression of the α2M polypeptide. In addition, cells derived from distant metastases may be used to prepare an immunogenic composition against the primary cancer. A mixture of cells can be used provided that a substantial number of cells in the mixture are cancer cells and share at least one antigenic determinant with the target cancer cell. In a specific embodiment, the cancer cells to be used in expressing an α2M polypeptide are purified.
[0130]5.2.2. In Vitro Complexing
[0131]In another embodiment, complexes of α2M polypeptides and antigenic molecules are produced in vitro. Immunogenic α2M polypeptide-antigenic molecule complexes can be generated in vitro by any method known in the art for forming α2M polypeptide-antigenic molecule complexes. Procedures for forming such α2M-antigenic molecule complexes and methods for isolating antigenic peptides are described in detail herein.
[0132]Methods for formation in vitro of noncovalent immunogenic complexes are well known in the art. For example, such complexes can be generated in vitro by noncovalent complexing of an α2M polypeptide with an antigenic molecule using methods which have been previously described for noncovalent coupling of an HSP with an antigenic molecule (see e.g., Blachere et al., 1997, supra; PCT publication WO 97/10000, dated Mar. 20, 1997). Preferably, the immunogenic molecular complex is not prepared by treatment with a protease, or with an activating agent such as ammonia or methyamine. In another preferred embodiment, the α2M molecule of the immunogenic molecular complex is not cleaved within the "bait" region. In yet another embodiment, the α2M polypeptide is not covalently associated with the antigenic molecule through a thioester linkage.
[0133]Methods for covalent coupling are also well known in the art (see, e.g., Osada et al., 1987, supra; Osada et al., 1988, supra; Chu and Pizzo 1993, supra; Chu et al., 1994, supra; Mitsuda et al., 1993, supra). In one embodiment, for example, when an α2M polypeptide is mixed with protease, During proteolytic activation of α2M, non-proteolytic ligands can become covalently bound to the activated thioesters. Non-proteolytic ligands can also be incorporated into the activated α2M molecule by ammonia or methylamine during reversal of the nucleophilic activation, employing heat (Gron and Pizzo, 1998, Biochemistry, 37: 6009-6014). Such conditions that allow fortuitous trapping of peptides by α2M are employed to prepare the α2M polypeptide-antigenic molecule complexes of the invention.
[0134]For example, in various embodiments of the invention, an α2M polypeptide may be mixed with antigenic molecule in the presence of a protease, ammonia or other small amine nucleophiles such as methylamine and ethylamine. Non-limiting examples of proteases which may be used include trypsin, porcine pancreatic elastase (PEP), human neutrophil elastase, cathepsin G, S. aureus V-8 proteinase trypsin, a-chymotrypsin, V8 protease, papain, and proteinase K (see Ausubel et al., (eds.), in "Current Protocols in Molecular Biology", Greene Publishing Associates and Wiley Interscience, New York, 17.4.6-17.4.8).
[0135]In another embodiment for preparation of covalent α2M polypeptide-antigenic molecule complexes, α2M polypeptides and antigenic molecules are prepared, and then covalently coupled using, for example, chemical crosslinking. Chemical crosslinking methods are well known in the art. For example, in a preferred embodiment, glutaraldehyde crosslinking may be used. Glutaradehyde crosslinking has been used for formation of covalent complexes of peptides and hsps (see Barrios et al., 1992, Eur. J. Immunol. 22: 1365-1372). In one embodiment, the following protocol is used. Optionally, α2M polypeptides may be pretreated with ATP or low pH prior to complexing, in order to remove any peptides that may be associated with the α2M polypeptide. Preferably, 1 mg of α2M polypeptide is crosslinked to 1 mg of peptide in the presence of 0.002% glutaraldehyde for 2 hours. Glutaraldehyde is removed by dialysis against phosphate buffered saline (PBS) overnight (Lussow et al., 1991, Eur. J. Immunol. 21: 2297-2302).
[0136]Other methods for chemical crosslinking may also be used, in addition other methods for covalent attachment of proteins, such as photocrosslinking (see Current Protocols in Molecular Biology, Ausubel et al. (eds.), Greene Publishing Associates and Wiley Interscience, New York).
[0137]Antigenic molecules for covalent or noncovalent α2M polypeptide-antigenic molecule complexes may be isolated from various sources, chemically synthesized, or produced recombinantly. Such methods can be readily adapted for medium or large scale production of the immunotherapeutic or prophylactic vaccines of the invention.
[0138]Following complexing, the immunogenic α2M-antigenic molecule complexes can optionally be purified. In a preferred embodiment, such complexes are at least 65%, 70%, 75%, 80%, 85%, 90%, 95%, 98%, or 100% nonconvalent complexes of α2M and the antigenic molecule. Such complexes may be assayed in vitro using, for example, the mixed lymphocyte target cell assay (MLTC) described below. Once immunogenic complexes have been isolated they can be optionally characterized further in animal models using the preferred administration protocols and excipients discussed below.
[0139]5.2.3. α2M-Antigenic Molecule Fusion Proteins
[0140]In another embodiment, recombinant fusion proteins, comprised of α2M sequences linked to antigenic molecule sequences, may be used for immunotherapy. To produce such a recombinant fusion protein, an expression vector is constructed using nucleic acid sequences encoding α2M fused to sequences encoding an antigenic molecule, using recombinant methods known in the art, such as those described in Sections 5.1.1.1 and 5.1.1.2, above (see Suzue et al., 1997, Proc. Natl. Acad. Sci. U.S.A. 94: 13146-51). α2M-antigenic peptide fusions are then expressed and isolated. By specifically designing the antigenic peptide portion of the molecule, such fusion proteins can be used to elicit an immune response and in immunotherapy against target cancer and infectious diseases.
[0141]5.2.4. Sources of Antigenic Molecules
[0142]Antigenic molecules, or antigenic portions thereof, specific to one or more types of cancer or infected cells, can be chosen from among those known in the art. Alternatively, such antigenic molecules can be selected for their antigenicity or their immunogenicity, as determined by immunoassays or by their ability to generate an immune response.
[0143]5.2.4.1 Exogenous Antigenic Molecules
[0144]Preferably, where it is desired to treat or prevent cancer, known tumor-specific antigenic molecules or fragments or derivatives thereof are used. For example, such tumor specific or tumor-associated antigenic molecules include but are not limited to KS 1/4 pan-carcinoma antigen (Perez and Walker, 1990, J. Immunol. 142:3662-3667; Bumal, 1988, Hybridoma 7(4):407-415); ovarian carcinoma antigen (CA125) (Yu, et al., 1991, Cancer Res. 51(2):468-475); prostatic acid phosphate (Tailer, et al., 1990, Nucl. Acids Res. 18(16):4928); prostate specific antigen (Henttu and Vihko, 1989, Biochem. Biophys. Res. Comm. 160(2):903-910; Israeli, et al., 1993, Cancer Res. 53:227-230); melanoma-associated antigen p97 (Estin, et al., 1989, J. Natl. Cancer Inst. 81(6):445-446); melanoma antigen gp75 (Vijayasardahl, et al., 1990, J. Exp. Med. 171(4):1375-1380); high molecular weight melanoma antigen (Natali, et al., 1987, Cancer 59:55-63) and prostate specific membrane antigen.
[0145]In a specific embodiment, an antigenic molecule or fragment or derivative thereof specific to a certain tumor is selected for complexing to α2M polypeptide and subsequent administration to a patient having that tumor.
[0146]In a preferred embodiment, where it is desired to treat or prevent viral diseases, molecules comprising epitopes of known viruses are used. For example, such antigenic epitopes may be prepared from viruses including, but not limited to, hepatitis type A, hepatitis type B, hepatitis type C, influenza, varicella, adenovirus, herpes simplex type I (HSV-I), herpes simplex type II (H SV-II), rinderpest, rhinovirus, echovirus, rotavirus, respiratory syncytial virus, papilloma virus, papova virus, cytomegalovirus, echinovirus, arbovirus, huntavirus, coxsackie virus, mumps virus, measles virus, rubella virus, polio virus, human immunodeficiency virus type I (HIV-I), and human immunodeficiency virus type II (HIV-II).
[0147]In another preferred embodiment, where it is desired to treat or prevent bacterial infections, molecules comprising epitopes of known bacteria are used. For example, such antigenic epitopes may be prepared from bacteria including, but not limited to, mycobacteria rickettsia, mycoplasma, neisseria and legionella.
[0148]In another preferred embodiment, where it is desired to treat or prevent protozoal infections, molecules comprising epitopes of known protozoa are used. For example, such antigenic epitopes may be prepared from protozoa including, but not limited to, leishmania, kokzidioa, and trypanosoma.
[0149]In yet another preferred embodiment, where it is desired to treat or prevent parasitic infections, molecules comprising epitopes of known parasites are used. For example, such antigenic epitopes may be from parasites including, but not limited to, chlamydia and rickettsia.
[0150]To determine immunogenicity or antigenicity of a putative antigen by detecting binding to antibody, various immunoassays known in the art can be used, including but not limited to competitive and non-competitive assay systems using techniques such as radioimmunoassays, ELISA (enzyme linked immunosorbent assay), "sandwich" immunoassays, immunoradiometric assays, gel diffusion precipitin reactions, immunodiffusion assays, in vivo immunoassays (using colloidal gold, enzyme or radioisotope labels, for example), western blots, immunoprecipitation reactions, agglutination assays (e.g., gel agglutination assays, hemagglutination assays), complement fixation assays, immunofluorescence assays, protein A assays, and immunoelectrophoresis assays, etc. In one aspect, antibody binding is detected by detecting a label on the primary antibody. In another aspect, the primary antibody is detected by detecting binding of a secondary antibody or reagent to the primary antibody. In a further aspect, the secondary antibody is labeled. Many means are known in the art for detecting binding in an immunoassay and are envisioned for use. In one embodiment for detecting immunogenicity, T cell-mediated responses can be assayed by standard methods, e.g., in vitro cytoxicity assays or in vivo delayed-type hypersensitivity assays.
[0151]Potentially useful antigenic molecules, or derivatives thereof, can be identified by various criteria, such as the antigen's involvement in neutralization of a pathogen's infectivity (wherein it is desired to treat or prevent infection by such a pathogen) (Norrby, 1985, Summary, in Vaccines 85, Lerner, et al. (eds.), Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y., pp. 388-389), type or group specificity, recognition by patients' antisera or immune cells, and/or the demonstration of protective effects of antisera or immune cells specific for the antigen. In addition, where it is desired to treat or prevent a disease caused by pathogen, the antigen's encoded epitope should preferably display a small or no degree of antigenic variation in time or amongst different isolates of the same pathogen.
[0152]5.2.4.2 Antigenic Molecules from α2M Complexes
[0153]Antigenic peptides for complexing in vitro to α2M polypeptides of the invention can also be obtained from endogenous complexes of peptides and α2Ms. Two methods may be used to elute the peptide from an α2M-antigenic molecule complex. One approach involves incubating the α2M-antigenic molecule complex in the presence of ATP. The other approach involves incubating the complexes in a low pH buffer.
[0154]Briefly, the complex of interest is centrifuged through a Centricon 10 assembly (Millipore) to remove any low molecular weight material loosely associated with the complex. The large molecular weight fraction may be removed and analyzed by SDS-PAGE while the low molecular weight may be analyzed by HPLC as described below. In the ATP incubation protocol, the α2M-antigenic molecule complex in the large molecular weight fraction is incubated with 10 mM ATP for 30 minutes at room temperature. In the low pH protocol, acetic acid or trifluoroacetic acid (TFA) is added to the α2M-antigenic molecule complex to give a final concentration of 10% (vol/vol) and the mixture incubated at room temperature or in a boiling water bath or any temperature in between, for 10 minutes.
[0155]The resulting samples are centrifuged through a Centricon10 assembly as mentioned previously. The high and low molecular weight fractions are recovered. The remaining large molecular weight α2M-antigenic molecule complexes can be reincubated with ATP or low pH to remove any remaining peptides.
[0156]The resulting lower molecular weight fractions are pooled, concentrated by evaporation and dissolved in 0.1% TFA. The dissolved material is then fractionated by reverse phase high pressure liquid chromatography (HPLC) using for example a VYDAC C18 reverse phase column equilibrated with 0.1% TFA. The bound material is then eluted at a flow rate of about 0.8 ml/min by developing the column with a linear gradient of 0 to 80% acetonitrile in 0.1% TFA. The elution of the peptides can be monitored by OD210 and the fractions containing the peptides collected.
[0157]5.2.4.3 Peptide Antigens from Mhc Complexes
[0158]Peptides bound to MHC molecules in vivo can also be used in vitro to form complexes with α2M polypeptides of the invention. The isolation of potentially immunogenic peptides from MHC molecules is well known in the art and so is not described in detail herein (see, Falk, et al., 1990, Nature 348:248-251; Rotzsche, at al., 1990, Nature 348:252-254; Elliott, et al., 1990, Nature 348:191-197; Falk, et al., 1991, Nature 351:290-296; Demotz, et al., 1989, Nature 343:682-684; Rotzsche, et al., 1990, Science 249:283-287), the disclosures of which are incorporated herein by reference.
[0159]Briefly, MHC-antigenic molecule complexes may be isolated by a conventional immunoaffinity procedure. The peptides then may be eluted from the MHC-antigenic molecule complex by incubating the complexes in the presence of about 0.1% TFA in acetonitrile. The eluted peptides may be fractionated and purified by reverse phase HPLC, as before.
[0160]5.2.4.4 Synthetic Antigenic Molecules
[0161]The amino acid sequences of the peptides eluted from MHC molecules or α2M may be determined either by manual or automated amino acid sequencing techniques well known in the art. Once the amino acid sequence of a potentially protective peptide has been determined, the peptide may be synthesized in using conventional peptide synthesis or other protocols well known in the art.
[0162]Peptides having the same amino acid sequence as those isolated above may be synthesized by solid-phase peptide synthesis using procedures similar to those described by Merrifield, 1963, J. Am. Chem. Soc., 85:2149. During synthesis, N-α-protected amino acids having protected side chains are added stepwise to a growing polypeptide chain linked by its C-terminal and to an insoluble polymeric support i.e., polystyrene beads. The peptides are synthesized by linking an amino group of an N-α-deprotected amino acid to an α-carboxyl group of an N-α-protected amino acid that has been activated by reacting it with a reagent such as dicyclohexylcarbodiimide. The attachment of a free amino group to the activated carboxyl leads to peptide bond formation. The most commonly used N-α-protecting groups include Boc which is acid labile and Fmoc which is base labile. Details of appropriate chemistries, resins, protecting groups, protected amino acids and reagents are well known in the art and so are not discussed in detail herein (See, Atherton, et al., 1989, Solid Phase Peptide Synthesis: A Practical Approach, IRL Press, and Bodanszky, 1993, Peptide Chemistry, A Practical Textbook, 2nd Ed., Springer-Verlag).
[0163]Purification of the resulting peptides is accomplished using conventional procedures, such as preparative HPLC using gel permeation, partition and/or ion exchange chromatography. The choice of appropriate matrices and buffers are well known in the art and so are not described in detail herein.
[0164]5.2.4.5 Recombinantly Produced Antigenic Molecules
[0165]In a particular embodiment of the invention, a nucleotide sequence encoding a protein antigenic molecule or portions thereof can be introduced into a host cell for production of the antigenic molecule. The nucleotide sequence encoding any antigenic protein can be obtained and cloned into an expression vector for expression essentially by the same methods described for the cloning and expression of a nucleotide sequence encoding an α2M polypeptide. The techniques are described in Sections 5.1.1.1 and 5.1.1.2, and are well known in the art. The recombinant antigenic protein or portions thereof can be purified by any methods appropriate for the protein, and then used to form complexes with α2M polypeptides in vitro as described in Section 5.2.2. Such an α2M polypeptide-antigenic molecule complex can be used as a vaccine to stimulate an immune response against the antigenic protein in a subject for the purpose of treatment or prevention of infectious diseases or cancer.
[0166]5.3 Therapeutic Applications for α2M Complexes
[0167]The present invention encompasses the use of α2M polypeptides in methods for treatment of and prevention of infectious diseases and cancer. In various embodiments described in detail herein, an effective amount of a α2M polypeptide in a covalent or noncovalent complex with an antigenic molecule is administered to a patient for therapeutic purposes.
[0168]5.3.1. Prevention and Treatment of Infectious Diseases
[0169]For treatment and prevention of infectious disease, α2M-antigenic molecule complexes are prepared from a cell that displays the antigenicity of an antigen of an infectious agent or pathogenic agent, and used as vaccines against the infectious disease. As will be appreciated by those skilled in the art, the protocols described herein may be used to isolate α2M polypeptide-antigenic molecule complexes from any cell that displays the antigenicity of an antigen of the infectious agent. For example, cells may be infected by the infectious agent itself, or alternatively, cells may be infected by or engineered to express an attenuated form of the infectious agent or a non-pathogenic or replication-deficient variant of the pathogen. In one embodiment, α2M-antigenic molecule complexes can be prepared from cells infected with non-infectious or non-pathogenic forms of the infectious agent (e.g., by use of a helper infectious agent). In another embodiment, the α2M-antigenic molecule complexes of the invention may be prepared from cells infected with an intracellular pathogen. In another embodiment, α2M polypeptide-complexes can be prepared from cells that have been transformed by an intracellular pathogen. For example, immunogenic α2M polypeptide-antigenic molecule complexes may be isolated from eukaryotic cells transformed with a transforming virus such as SV40.
[0170]A preferred method for treatment or prevention of an infectious disease comprises introducing into a cell that displays the antigenicity of an infectious agent an expressible α2M polypeptide gene sequence, preferably as an expression gene construct. The α2M polypeptide gene sequence is manipulated by recombinant methods, such as those described above in Sections 5.1.1.1 and 5.1.1.2 above, so that the α2M polypeptide gene sequence, in the form of an expression construct, located extrachromosomally or integrated in the chromosome, is suitable for expression of the α2M polypeptide in the recombinant cells. The recombinant cells containing the expression gene constructs are cultured under conditions such that α2M polypeptides encoded by the expression gene construct are expressed. Complexes of α2M polypeptides covalently or noncovalently associated with antigenic molecules of the infectious agent are purified from the cell culture or culture medium by the methods described in Section 5.2.1.
[0171]In various embodiments, α2M-antigenic molecule complexes are prepared from a cell genetically manipulated to express an α2M polypeptide, for example, tissues, isolated cells or immortalized eukaryotic cell lines infected with an intracellular pathogen. When immortalized animal cell lines are used as a source of the α2M polypeptide-antigenic molecule complex, it is important to use cell lines that can be infected with the pathogen of interest. In addition, it is preferable to use cells that are derived from the same species as the intended recipient of the vaccine. Techniques for introducing an expressible form of the α2M polypeptide gene sequences into these cell lines are described above in Section 5.1.1.2. If a pathogen is expected to cause lysis of the host cells, it is preferred to introduce the expressible α2M polypeptide gene sequence into the host cell prior to infecting the cells with the pathogen. For example, in order to prepare an α2M polypeptide-antigenic molecule complex for administration to humans that may be effective against HIV-1, the virus may be propagated in human cells which include, but are not limited to, human CD4+ T cells, HepG2 cells, and U937 promonocytic cells, which have already been transfected with an expressible α2M polypeptide sequence. Similarly, influenza viruses may be propagated in, for example, transfected human fibroblast cell lines and MDCK cells, and mycobacteria may be cultured in, for example, transfected human Schwaan cells. The cell supernatant containing α2M-antigenic molecule complex may be collected just prior to lysis of the host cell.
[0172]In a preferred aspect of the invention, the purified α2M-antigenic molecule complex vaccines may have particular utility in the treatment of human diseases caused by intracellular pathogens. It is appreciated, however, that the vaccines developed using the principles described herein will be useful in treating diseases of other mammals, for example, farm animals including: cattle; horses; sheep; goats; and pigs, and household pets including: cats; and dogs, that similarly are caused by intracellular pathogens.
[0173]In accordance with the methods described herein, vaccines may be prepared that stimulate an immune response, in particular a cytotoxic T cell responses, against cells infected with viruses including, but not limited to, hepatitis type A, hepatitis type B, hepatitis type C, influenza, varicella, adenovirus, HSV-I, HSV-II, rinderpest rhinovirus, echovirus, rotavirus, respiratory syncytial virus, papilloma virus, papova virus, cytomegalovirus, echinovirus, arbovirus, huntavirus, coxsachie virus, mumps virus, measles virus, rubella virus, polio virus, HIV-I, and HIV-II. Similarly, vaccines may also be prepared that stimulate cytotoxic T cell responses against cells infected with intracellular bacteria, including, but not limited to, Mycobacteria, Rickettsia, Mycoplasma, Neisseria and Legionella. In addition, vaccines may also be prepared that stimulate cytotoxic T cell responses against cells infected with intracellular protozoa, including, but not limited to, Leishmani, Kokzidioa, and Trypanosoma. Furthermore, vaccines may be prepared that stimulate cytotoxic T cell responses against cells infected with intracellular parasites including, but not limited to, Chlamydia and Rickettsia.
[0174]The effect of immunotherapy with modified α2M polypeptide-antigenic molecule complexes on progression of infectious diseases can be monitored by any methods known to one skilled in the art.
[0175]5.3.2. Prevention and Treatment of Cancer
[0176]There are many reasons why immunotherapy as provided by the covalent or noncovalent α2M polypeptide-antigenic molecule complexes or recombinant cells expressing α2M polypeptides prepared by the present invention is desired for use in cancer patients. First, if cancer patients are immunosuppressed, and surgery with anesthesia, and subsequent chemotherapy, may worsen the immunosuppression, then with appropriate immunotherapy in the preoperative period, this immunosuppression may be prevented or reversed. This could lead to fewer infectious complications and to accelerated wound healing. Second, tumor bulk is minimal following surgery and immunotherapy is most likely to be effective in this situation. A third reason is the possibility that tumor cells are shed into the circulation at surgery and effective immunotherapy applied at this time can eliminate these cells.
[0177]In a specific embodiment, the preventive and therapeutic utility of the invention is directed at enhancing the immunocompetence of the cancer patient either before surgery, at or after surgery, and at inducing tumor-specific immunity to cancer cells, with the objective being inhibition of cancer, and with the ultimate clinical objective being total cancer regression and eradication.
[0178]According to the invention, preferred methods of treatment or prevention of cancer comprise isolating cancer cells from one or more individual, preferably the individual in need of treatment, and introducing into such cells an expressible α2M polypeptide gene sequence, preferably as an expression gene construct. The α2M polypeptide gene sequence is manipulated by methods described above in Sections 5.1.1.1 and 5.1.1.2, such that the α2M polypeptide gene sequence, in the form of an expression construct, or intrachromosomally integrated, are suitable for expression of the α2M polypeptide in the recombinant cells. The recombinant cells containing the expression gene constructs are cultured under conditions such that α2M polypeptides encoded by the expression gene construct are expressed by the recombinant host cells. Complexes of α2M polypeptides covalently or noncovalently associated with antigenic molecules of the cancer cell are purified from the cell culture or culture medium by the methods described in Section 5.2.1. Depending on the route of administration, the α2M polypeptide-antigenic molecule complexes are formulated accordingly as described in Section 5.7, and administered to the individual autologously (e.g., to treat the primary cancer or metastases thereof), or to other individuals who are in need of treatment for cancer of a similar tissue type, or to individuals at enhanced risk of cancer due to familial history or environmental risk factors.
[0179]For example, treatment with α2M polypeptide-antigenic molecule complexes prepared as described above may be started any time after surgery. However, if the patient has received chemotherapy, α2M-antigenic molecule complexes are usually administered after an interval of four weeks or more so as to allow the immune system to recover. The therapeutic regimen may include weekly injections of the α2M polypeptide-antigenic molecule complex, dissolved in saline or other physiologically compatible solution. The route and site of injection is varied each time, for example, the first injection is given subcutaneously on the left arm, the second injection on the right arm, the third injection on the left abdominal region, the fourth injection on the right abdominal region, the fifth injection on the left thigh, the sixth injection on the right thigh, etc. The same site is repeated after a gap of one or more injections. In addition, injections are split and each half of the dose is administered at a different site on the same day. Overall, the first four to six injections are given at weekly intervals. Subsequently, two injections are given at two-week intervals, followed by a regimen of injections at monthly intervals.
[0180]Alternatively, recombinant tumor cells expressing α2M-antigenic molecule complexes can be used as a vaccine for injection into a patient to stimulate an immune response against the tumor cells or cells bearing tumor antigens. Autologous recombinant tumor cells stably expressing α2M polypeptide-antigenic molecule complexes are preferred. To determine the appropriate dose, the amount of α2M polypeptide-antigenic molecule complex produced by the recombinant cells is quantitated, and the number of recombinant cells used for vaccination is adjusted accordingly to assure a consistent level of expression in vivo. A preferred dose is the number of recombinant cells that can produce about 100 ng α2M polypeptide per 24 hours. For the safety of the patient, the recombinant tumor cells can be irradiated (12000 rad) immediately prior to injection into a patient. Irradiated cells do not proliferate, and can continue to express α2M polypeptide-antigenic molecule complexes for about 7-10 days which is sufficient to induce an immune response.
[0181]Cancers that can be treated or prevented by using covalent or noncovalent α2M-antigenic molecule complexes prepared by the methods of the present invention include, but not limited to human sarcomas and carcinomas, e.g., fibrosarcoma, myxosarcoma, liposarcoma, chondrosarcoma, osteogenic sarcoma, chordoma, angiosarcoma, endotheliosarcoma, lymphangiosarcoma, lymphangioendotheliosarcoma, synovioma, mesothelioma, Ewing's tumor, leiomyosarcoma, rhabdomyosarcoma, colon carcinoma, pancreatic cancer, breast cancer, ovarian cancer, prostate cancer, squamous cell carcinoma, basal cell carcinoma, adenocarcinoma, sweat gland carcinoma, sebaceous gland carcinoma, papillary carcinoma, papillary adenocarcinomas, cystadenocarcinoma, medullary carcinoma, bronchogenic carcinoma, renal cell carcinoma, hepatoma, bile duct carcinoma, choriocarcinoma, seminoma, embryonal carcinoma, Wilms' tumor, cervical cancer, testicular tumor, lung carcinoma, small cell lung carcinoma, bladder carcinoma, epithelial carcinoma, glioma, astrocytoma, medulloblastoma, craniopharyngioma, ependymoma, pinealoma, hemangioblastoma, acoustic neuroma, oligodendroglioma, meningioma, melanoma, neuroblastoma, retinoblastoma; leukemias, e.g., acute lymphocytic leukemia and acute myelocytic leukemia (myeloblastic, promyelocytic, myelomonocytic, monocytic and erythroleukemia); chronic leukemia (chronic myelocytic (granulocytic) leukemia and chronic lymphocytic leukemia); and polycythemia vera, lymphoma (Hodgkin's disease and non-Hodgkin's disease), multiple myeloma, Waldenstrom's macroglobulinemia, and heavy chain disease.
[0182]In a specific embodiment, the cancer is metastatic. In another specific embodiment, the patient having a cancer is immunosuppressed by reason of having undergone anti-cancer therapy (e.g., chemotherapy radiation) prior to administration of the α2M-antigenic molecule complexes of the invention. In another specific embodiment, the cancer is a tumor.
[0183]The effect of immunotherapy with α2M polypeptide-antigenic molecule complexes on progression of neoplastic diseases can be monitored by any methods known to one skilled in the art, including but not limited to measuring: a) delayed hypersensitivity as an assessment of cellular immunity; b) activity of cytolytic T-lymphocytes in vitro; c) levels of tumor specific antigens, e.g., carcinoembryonic (CEA) antigens; d) changes in the morphology of tumors using techniques such as a computed tomographic (CT) scan; e) changes in levels of putative biomarkers of risk for a particular cancer in individuals at high risk, and f) changes in the morphology of tumors using a sonogram. Other techniques that can also be used include scintigraphy and endoscopy.
[0184]The preventive effect of immunotherapy using α2M polypeptide-antigenic molecule complexes may also be estimated by determining levels of a putative biomarker for risk of a specific cancer. For example, in individuals at enhanced risk for prostate cancer, serum prostate-specific antigen (PSA) is measured by the procedure described by Brawer et al., 1992, J. Urol. 147:841-845, and Catalona et al., 1993, JAMA 270:948-958; or in individuals at risk for colorectal cancer, CEA is measured by methods known in the art; and in individuals at enhanced risk for breast cancer, 16-α-hydroxylation of estradiol is measured by the procedure described by Schneider et al., 1982, Proc. Natl. Acad. Sci. USA 79:3047-3051. The references cited above are incorporated by reference herein in their entirety.
[0185]5.3.3. Combination with Adoptive Immunotherapy
[0186]Adoptive immunotherapy refers to a therapeutic approach for treating infectious diseases or cancer in which immune cells are administered to a host with the aim that the cells mediate specific immunity, either directly or indirectly, to the infected cells or tumor cells and/or antigenic components, and result in treatment of the infectious disease or regression of the tumor, as the case may be (see U.S. patent application Ser. No. 08/527,546, filed Sep. 13, 1995, which is incorporated by reference herein in its entirety). α2M polypeptides may be used to sensitize antigen presenting cells (APCs) using in covalent or noncovalent complexes with antigenic (or immunogenic) molecules, for adoptive immunotherapy.
[0187]According to the invention, therapy by administration of α2M polypeptide-antigenic molecule complexes, using any desired route of administration, is combined with adoptive immunotherapy using APC sensitized with α2M polypeptide-antigenic molecule complexes. The α2M polypeptide-antigenic molecule complex-sensitized APC can be administered concurrently with α2M polypeptide-antigenic molecule complexes, or before or after administration of α2M polypeptide-antigenic molecule complexes. Furthermore, the mode of administration can be varied, including but not limited to, e.g., subcutaneously, intravenously, intraperitoneally, intramuscularly, intradermally or mucosally.
[0188]5.3.3.1 Sensitization of Antigen Presenting Cells with α2M Complexes
[0189]The antigen-presenting cells, including but not limited to macrophages, dendritic cells and B-cells, are preferably obtained by production in vitro from stem and progenitor cells from human peripheral blood or bone marrow as described by Inaba et al., 1992, J. Exp. Med. 176:1693-1702. APC can be obtained by any of various methods known in the art. In a preferred aspect human macrophages are used, obtained from human blood cells.
[0190]By way of example, but not limitation, macrophages can be obtained as follows: Mononuclear cells are isolated from peripheral blood of a patient (preferably the patient to be treated), by Ficoll-Hypaque gradient centrifugation and are seeded on tissue culture dishes which are pre-coated with the patient's own serum or with other AB+ human serum. The cells are incubated at 37° C. for 1 hr, then non-adherent cells are removed by pipetting. To the adherent cells left in the dish, is added cold (4° C.) 1 mM EDTA in phosphate-buffered saline and the dishes are left at room temperature for 15 minutes. The cells are harvested, washed with RPMI buffer and suspended in RPMI buffer. Increased numbers of macrophages may be obtained by incubating at 37° C. with macrophage-colony stimulating factor (M-CSF); increased numbers of dendritic cells may be obtained by incubating with granulocyte-macrophage-colony stimulating factor (GM-CSF) as described in detail by Inaba, K., et al., 1992, J. Exp. Med. 176:1693-1702.
[0191]APC are sensitized with α2M polypeptides covalently or noncovalently bound to antigenic molecules by incubating the cells in vitro with the complexes. The APC are sensitized with complexes of α2M polypeptide and antigenic molecules preferably by incubating in vitro with the α2M polypeptide-complex at 37° C. for 15 minutes to 24 hours. By way of example but not limitation, 4×107 macrophages can be incubated with 10 microgram α2M-antigenic molecule complexes per ml or 100 microgram α2M-antigenic molecule complexes per ml at 37° C. for 15 mins to 24 hrs in 1 ml plain RPMI medium. The cells are washed three times and resuspended in a physiological medium preferably sterile, at a convenient concentration (e.g, 1×107/ml) for injection in a patient. In a preferred embodiment, the antigen presenting cells are autologous to the patient, that is, the patient into which the sensitized APCs are injected is the patient from which the APC were originally isolated.
[0192]Optionally, the ability of sensitized APC to stimulate, for example, the antigen-specific, class I-restricted cytotoxic T-lymphocytes (CTL) can be monitored by their ability to stimulate CTLs to release tumor necrosis factor, and by their ability to act as targets of such CTLs.
[0193]5.3.3.2 Reinfusion of Sensitized APC
[0194]The α2M polypeptide-antigen-sensitized APC are reinfused into the patient systemically, preferably intravenously, by conventional clinical procedures. These activated cells are reinfused, preferentially by systemic administration into the autologous patient. Patients generally receive from about 106 to about 1012 sensitized macrophages, depending on the condition of the patient. In some regimens, patients may optionally receive in addition a suitable dosage of a biological response modifier including but not limited to the cytokines IFN-α, IFN-γ, IL-2, IL-4, IL-6, TNF or other cytokine growth factor.
[0195]5.3.4. Determination of Immunogenicity of α2M-Antigen Molecule Complexes
[0196]In an optional procedure, the purified α2M polypeptide-antigenic molecule complexes can be assayed for immunogenicity using the mixed lymphocyte target culture assay (MLTC) well known in the art.
[0197]By way of example but not limitation, the following procedure can be used. Briefly, mice are injected subcutaneously with the candidate α2M polypeptide-antigenic molecule complexes. As a positive control another set of mice are immunized with whole cancer cells of the type from which the α2M polypeptides are derived. As a negative control, mice are injected with either α2M-antigenic molecule complexes isolated from normal, non-recombinant cells or whole cells (i.e., antigenically distinct from the type of cell from which the α2M polypeptides are derived). The mice are injected twice, 7-10 days apart. The mice are injected twice, 7-10 days apart. Ten days after the last immunization, the spleens are removed and the lymphocytes released. The released lymphocytes may be restimulated subsequently in vitro by the addition of dead cells that expressed the complex of interest.
[0198]For example, 8×106 immune spleen cells may be stimulated with 4×104 mitomycin C treated or γ-irradiated (5-10,000 rads) pathogen-infected cells (or cells transfected with a gene encoding an antigen of the infectious agent, as the case may be), or tumor cells in 3 ml RPMI medium containing 10% fetal calf serum. In certain cases 33% secondary mixed lymphocyte culture supernatant or interleukin 2 (IL-2) may be included in the culture medium as a source of T cell growth factors (See, Glasebrook et al., 1980, J. Exp. Med. 151:876). To test the primary cytotoxic T cell response after immunization, spleen cells may be cultured without stimulation. In some experiments spleen cells of the immunized mice may also be restimulated with antigenically distinct cells, to determine the specificity of the cytotoxic T cell response.
[0199]Six days later the cultures are tested for cytotoxicity in a 4 hour 51Cr-release assay (See, Palladino et al., 1987, Cancer Res. 47:5074-5079 and Blachere, at al., 1993, J. Immunotherapy 14:352-356). In this assay, the mixed lymphocyte culture is added to a target cell suspension to give different effector:target (E:T) ratios (usually 1:1 to 40:1). The target cells are prelabelled by incubating 1×106 target cells in culture medium containing 200 mCi 51Cr/ml for one hour at 37° C. The cells are washed three times following labeling. Each assay point (E:T ratio) is performed in triplicate and the appropriate controls incorporated to measure spontaneous 51Cr release (no lymphocytes added to assay) and 100% release (cells lysed with detergent). After incubating the cell mixtures for 4 hours, the cells are pelleted by centrifugation at 200 g for 5 minutes. The amount of 51Cr released into the supernatant is measured by a gamma counter. The percent cytotoxicity is measured as cpm in the test sample minus spontaneously released cpm divided by the total detergent released cpm minus spontaneously released cpm.
[0200]In order to block the MHC class I cascade a concentrated hybridoma supernatant derived from K-44 hybridoma cells (an anti-MHC class I hybridoma) is added to the test samples to a final concentration of 12.5% (Heike et al., 1994, J. Immunotherapy 15:165-174).
[0201]An alternative to the chromium-release assay is the ELISPOT assay which measures cytokine release by cytotoxic T cells in vitro after stimulation with specific antigen. Cytokine release is detected by antibodies which are specific for a particular cytokine, such as interleukin-2, tumor necrosis factor α or interferon-γ (for example, see Scheibenbogen et al., 1997, Int. J. Cancer, 71:932-936). The assay is carried out in a microtiter plate which has been pre-coated with an antibody specific for a cytokine of interest which captures the cytokine secreted by T cells. After incubation of T cells for 24-48 hours in the coated wells, the cytotoxic T cells are removed and replaced with a second labeled antibody that recognizes a different epitope on the cytokine. After extensive washing to remove unbound antibody, an enzyme substrate which produces a colored reaction product is added to the plate. The number of cytokine-producing cells is counted under a microscope. This method has the advantages of short assay time, and sensitivity without the need of a large number of cytotoxic T cells.
[0202]5.3.5. Monitoring of Effects During Immunotherapy
[0203]The effect of immunotherapy with α2M polypeptide-antigenic molecule complexes can be monitored by any methods known to one skilled in the art, including but not limited to measuring: a) delayed hypersensitivity as an assessment of cellular immunity; b) activity of cytolytic T-lymphocytes in vitro; c) levels of infective agent-agent or tumor-specific antigens, e.g., carcinoembryonic (CEA) antigens. In the case of the use of α2M-antigenic molecule complexes for prevention or treatment of cancer, the effect can additionally be monitored by: d) changes in the morphology of tumors using techniques such as a computed tomographic (CT) scan; and e) changes in levels of putative biomarkers of risk for a particular cancer in individuals at high risk, and f) changes in the morphology of tumors using a sonogram.
[0204]5.3.5.1 Delayed Hypersensitivity Skin Test
[0205]Delayed hypersensitivity skin tests are of great value in the overall immunocompetence and cellular immunity to an antigen. Inability to react to a battery of common skin antigens is termed anergy (Sato et al., 1995, Clin. Immunol. Pathol. 74:35-43).
[0206]Proper technique of skin testing requires that the antigens be stored sterile at 4° C., protected from light and reconstituted shorted before use. A 25- or 27-gauge needle ensures intradermal, rather than subcutaneous, administration of antigen. Twenty-four and 48 hours after intradermal administration of the antigen, the largest dimensions of both erythema and induration are measured with a ruler. Hypoactivity to any given antigen or group of antigens is confirmed by testing with higher concentrations of antigen or, in ambiguous circumstances, by a repeat test with an intermediate test.
[0207]5.3.5.2 In Vitro Activation of Cytotoxic T Cells
[0208]The activity of cytotoxic T-lymphocytes can be assessed in vitro using the following method. Eight×106 peripheral blood-derived T lymphocytes isolated by the Ficoll-Hypaque centrifugation gradient technique, are restimulated with 4×104 mitomycinC-treated tumor cells in 3 ml RPMI medium containing 10% fetal calf serum. In some experiments, 33% secondary mixed lymphocyte culture supernatant or IL-2, is included in the culture medium as a source of T cell growth factors.
[0209]In order to measure the primary response of cytolytic T-lymphocytes after immunization, T cells are cultured without the stimulator tumor cells. In other experiments, T cells are restimulated with antigenically distinct cells. After six days, the cultures are tested for cytotoxicity in a 4 hour 51Cr-release assay. The spontaneous 51Cr-release of the targets should reach a level less than 20%. For the anti-MHC class I blocking activity, a tenfold concentrated supernatant of W6/32 hybridoma is added to the test at a final concentration of 12.5% (Heike M., et al., J Immunotherapy 15:165-174).
[0210]5.3.5.3 Levels of Tumor Specific Antigens
[0211]Although it may not be possible to detect unique tumor antigens on all tumors, many tumors display antigens that distinguish them from normal cells. The monoclonal antibody reagents have permitted the isolation and biochemical characterization of the antigens and have been invaluable diagnostically for distinction of transformed from nontransformed cells and for definition of the cell lineage of transformed cells. The best-characterized human tumor-associated antigens are the oncofetal antigens. These antigens are expressed during embryogenesis, but are absent or very difficult to detect in normal adult tissue. The prototype antigen is carcinoembryonic antigen (CEA), a glycoprotein found on fetal gut and human colon cancer cells, but not on normal adult colon cells. Since CEA is shed from colon carcinoma cells and found in the serum, it was originally thought that the presence of this antigen in the serum could be used to screen patients for colon cancer. However, patients with other tumors, such as pancreatic and breast cancer, also have elevated serum levels of CEA. Therefore, monitoring the fall and rise of CEA levels in cancer patients undergoing therapy has proven useful for predicting tumor progression and responses to treatment.
[0212]Several other oncofetal antigens have been useful for diagnosing and monitoring human tumors, e.g., alpha-fetoprotein, an alpha-globulin normally secreted by fetal liver and yolk sac cells, is found in the serum of patients with liver and germinal cell tumors and can be used as a marker of disease status.
[0213]5.3.5.4 Computed Tomographic (CT) Scan
[0214]CT remains the choice of techniques for the accurate staging of cancers. CT has proved more sensitive and specific than any other imaging techniques for the detection of metastases. A sonogram remains an alternative choice of technique for the accurate staging of cancers.
[0215]5.3.5.5 MEASUREMENT OF PUTATIVE BIOMARKERS
[0216]The levels of a putative biomarker for risk of a specific cancer are measured to monitor the effect of α2M covalently or noncovalently bound to antigenic molecule complexes. For example, in individuals at enhanced risk for prostate cancer, serum prostate-specific antigen (PSA) is measured by the procedure described by Brawer et. al., 1992, J. Urol. 147:841-845, and Catalona et al., 1993, JAMA 270:948-958; and in individuals at enhanced risk for breast cancer, 16-α-hydroxylation of estradiol is measured by the procedure described by Schneider et al., 1982, Proc. Natl. Acad. Sci. ISA 79:3047-3051.
[0217]5.4 Target Autoimmune Diseases
[0218]Autoimmune diseases that can be treated by the methods of the present invention include, but are not limited to, insulin dependent diabetes mellitus (i.e., IDDM, or autoimmune diabetes), multiple sclerosis, systemic lupus erythematosus, Sjogren's syndrome, scleroderma, polymyositis, chronic active hepatitis, mixed connective tissue disease, primary biliary cirrhosis, pernicious anemia, autoimmune thyroiditis, idiopathic Addison's disease, vitiligo, gluten-sensitive enteropathy, Graves' disease, myasthenia gravis, autoimmune neutropenia, idiopathic thrombocytopenia purpura, rheumatoid arthritis, cirrhosis, pemphigus vulgaris, autoimmune infertility, Goodpasture's disease, bullous pemphigoid, discoid lupus, ulcerative colitis, and dense deposit disease. The diseases set forth above, as referred to herein, include those exhibited by animal models for such diseases, such as, for example non-obese diabetic (NOD) mice for IDDM and experimental autoimmune encephalomyelitis (EAE) mice for multiple sclerosis.
[0219]The methods of the present invention can be used to treat such autoimmune diseases by reducing or eliminating the immune response to the patient's own (self) tissue, or, alternatively, by reducing or eliminating a pre-existing autoimmune response directed at tissues or organs transplanted to replace self tissues or organs damaged by the autoimmune response.
5.5 Target Infectious Diseases
[0220]The infectious diseases that can be treated or prevented using the methods and compositions of the present invention include those caused by intracellular pathogens such as viruses, bacteria, protozoans, and intracellular parasites. Viruses include, but are not limited to viral diseases such as those caused by hepatitis type B virus, parvoviruses, such as adeno-associated virus and cytomegalovirus, papovaviruses such as papilloma virus, polyoma viruses, and SV40, adenoviruses, herpes viruses such as herpes simplex type I (HSV-I), herpes simplex type II (HSV-II), and Epstein-Barr virus, poxviruses, such as variola (smallpox) and vaccinia virus, RNA viruses, including but not limited to human immunodeficiency virus type I (HIV-I), human immunodeficiency virus type II (HIV-II), human T-cell lymphotropic virus type I (HTLV-I), and human T-cell lymphotropic virus type II (HTLV-II); influenza virus, measles virus, rabies virus, Sendai virus, picornaviruses such as poliomyelitis virus, coxsackieviruses, rhinoviruses, reoviruses, togaviruses such as rubella virus (German measles) and Semliki forest virus, arboviruses, and hepatitis type A virus.
[0221]In another embodiment, bacterial infections can be treated or prevented such as, but not limited to disorders caused by pathogenic bacteria including, but not limited to, Streptococcus pyogenes, Streptococcus pneumoniae, Neisseria gonorrhoea, Neisseria meningitidis, Corynebacterium diphtheriae, Clostridium botulinum, Clostridium perfringens, Clostridium tetani, Haemophilus influenzae, Klebsiella pneumoniae, Klebsiella ozaenae, Klebsiella rhinoscleromotis, Staphylococcus aureus, Vibrio cholerae, Escherichia coli, Pseudomonas aeruginosa, Campylobacter (Vibrio) fetus, Campylobacter jejuni, Aeromonas hydrophila, Bacillus cereus, Edwardsiella tarda, Yersinia enterocolitica, Yersinia pestis, Yersinia pseudotuberculosis, Shigella dysenteriae, Shigella flexneri, Shigella sonnei, Salmonella typhiimurium, Salmonella typhii, Treponema pallidum, Treponema pertenue, Treponema carateneum, Borrelia vincentii, Borrelia burgdorferi, Leptospira icterohemorrhagiae, Mycobacterium tuberculosis, Toxoplasma gondii, Pneumocystis carinii, Francisella tularensis, Brucella abortus, Brucella suis, Brucella melitensis, Mycoplasma spp., Rickettsia prowazeki, Rickettsia tsutsugumushi, Chlamydia spp., and Helicobacter pylori.
[0222]In another preferred embodiment, the methods can be used to treat or prevent infections caused by pathogenic protozoans such as, but not limited to, Entomoeba histolytica, Trichomonas tenas, Trichomonas hominis, Trichomonas vaginalis, Trypanosoma gambiense, Trypanosoma rhodesiense, Trypanosoma cruzi, Leishmania donovani, Leishmania tropica, Leishmania braziliensis, Pneumocystis pneumonia, Plasmodium vivax, Plasmodium falciparum, and Plasmodium malaria.
5.6 Target Proliferative Cell Disorders
[0223]With respect to specific proliferative and oncogenic disease associated with α2M-α2M activity, the diseases that can be treated or prevented by the methods of the present invention include, but are not limited to: human sarcomas and carcinomas, e.g., fibrosarcoma, myxosarcoma, liposarcoma, chondrosarcoma, osteogenic sarcoma, chordoma, angiosarcoma, endotheliosarcoma, lymphangiosarcoma, lymphangioendotheliosarcoma, synovioma, mesothelioma, Ewing's tumor, leiomyosarcoma, rhabdomyosarcoma, colon carcinoma, pancreatic cancer, breast cancer, ovarian cancer, prostate cancer, squamous cell carcinoma, basal cell carcinoma, adenocarcinoma, sweat gland carcinoma, sebaceous gland carcinoma, papillary carcinoma, papillary adenocarcinomas, cystadenocarcinoma, medullary carcinoma, bronchogenic carcinoma, renal cell carcinoma, hepatoma, bile duct carcinoma, choriocarcinoma, seminoma, embryonal carcinoma, Wilms' tumor, cervical cancer, testicular tumor, lung carcinoma, small cell lung carcinoma, bladder carcinoma, epithelial carcinoma, glioma, astrocytoma, medulloblastoma, craniopharyngioma, ependymoma, pinealoma, hemangioblastoma, acoustic neuroma, oligodendroglioma, meningioma, melanoma, neuroblastoma, retinoblastoma; leukemias, e.g., acute lymphocytic leukemia and acute myelocytic leukemia (myeloblastic, promyelocytic, myelomonocytic, monocytic and erythroleukemia); chronic leukemia (chronic myelocytic (granulocytic) leukemia and chronic lymphocytic leukemia); and polycythemia vera, lymphoma (Hodgkin's disease and non-Hodgkin's disease), multiple myeloma, Waldenstrom's macroglobulinemia, and heavy chain disease.
[0224]Diseases and disorders involving a deficiency in cell proliferation or in which cell proliferation is desired for treatment or prevention, and that can be treated or prevented by inhibiting the α2M function, include but are not limited to degenerative disorders, growth deficiencies, hypoproliferative disorders, physical trauma, lesions, and wounds; for example, to promote wound healing, or to promote regeneration in degenerated, lesioned or injured tissues, etc.
5.7 Dosage Regimens and Formulation
[0225]Covalent or noncovalent complexes of α2M polypeptides and antigenic molecules of the invention may be formulated into pharmaceutical preparations for administration to mammals for treatment or prevention of infectious diseases or cancer at therapeutically effective doses for immunotherapy.
[0226]Selection of the preferred effective dose will be determined by a skilled artisan based upon considering several factors which will be known to one of ordinary skill in the art. Such factors include the particular form of α2M, and its pharmacokinetic parameters such as bioavailability, metabolism, half-life, etc., which will have been established during the usual development procedures typically employed in obtaining regulatory approval for a pharmaceutical compound. Further factors in considering the dose include the condition or disease to be treated or the benefit to be achieved in a normal individual, the body mass of the patient, the route of administration, whether administration is acute or chronic, concomitant medications, and other factors well known to affect the efficacy of administered pharmaceutical agents. Thus the precise dosage should be decided according to the judgment of the practitioner and each patient's circumstances, e.g., depending upon the condition and the immune status of the individual patient, according to standard clinical techniques.
[0227]Covalent or noncovalent complexes of α2M polypeptides and antigenic molecules of the invention may be formulated into pharmaceutical preparations for administration to mammals for treatment or prevention of infectious diseases or cancer. Drug solubility and the site of absorption are factors which should be considered when choosing the route of administration of a therapeutic agent.
[0228]α2M polypeptide-antigenic molecule complexes of the invention may optionally be administered with one or more adjuvants in order to enhance the immunological response. For example, depending on the host species, adjuvants which may be used include, but are not limited to: mineral salts or mineral gels such as aluminum hydroxide, aluminum phosphate, and calcium phosphate; surface active substances such as lysolecithin, pluronic polyols, polyanions, peptides, oil emulsions, keyhole limpet hemocyanins, and dinitrophenol; immunostimulatory molecules, such as cytokines, saponins, muramyl dipeptides and tripeptide derivatives, CpG dinucleotides, CpG oligonucleotides, monophosphoryl Lipid A, and polyphosphazenes; particulate and microparticulate adjuvant, such as emulsions, liposomes, virosomes, cochleates; or an immune stimulating complex mucosal adjuvants, Freund's (complete and incomplete, and potentially useful human adjuvants such as BCG (bacille Calmette-Guerin) and corynebacterium parvum.).
[0229]α2M polypeptide-antigenic molecule complexes of the invention may be administered using any desired route of administration, including but not limited to, e.g., subcutaneously, intravenously or intramuscularly, although intradermally or mucosally is preferred. Advantages of intradermal or mucosal administration include use of lower doses and rapid absorption, respectively. Mucosal routes of administration include, but are not limited to, oral, rectal and nasal administration. Preparations for mucosal administrations are suitable in various formulations as described below. The route of administration can be varied during a course of treatment.
[0230]The doses recited above are preferably given once weekly for a period of about 4-6 weeks, and the mode or site of administration is preferably varied with each administration. In a preferred example, subcutaneous administrations are given, with each site of administration varied sequentially. Thus, by way of example and not limitation, the first injection may be given subcutaneously on the left arm, the second on the right arm, the third on the left belly, the fourth on the right belly, the fifth on the left thigh, the sixth on the right thigh, etc. The same site may be repeated after a gap of one or more injections. Also, split injections may be given. Thus, for example, half the dose may be given in one site and the other half on an other site on the same day.
[0231]Alternatively, the mode of administration is sequentially varied, e.g., weekly injections are given in sequence subcutaneously, intramuscularly, intravenously or intraperitoneally.
[0232]After 4-6 weeks, further injections are preferably given at two-week intervals over a period of time of one month. Later injections may be given monthly. The pace of later injections may be modified, depending upon the patient's clinical progress and responsiveness to the immunotherapy.
[0233]Compositions comprising covalent or noncovalent complexes formulated in a compatible pharmaceutical carrier may be prepared, packaged, and labeled for treatment of the indicated infectious disease or tumor. In preferred aspects, an amount of α2M polypeptide-antigenic molecule complex is administered to a human that is in the range of about 2 to 150 μg, preferably 2 to 50 μg, most preferably about 25 μg, given once weekly for about 4-6 weeks, intradermally with the site of administration varied sequentially.
[0234]If the complex is water-soluble, then it may be formulated in an appropriate buffer, for example, phosphate buffered saline or other physiologically compatible solutions. Alternatively, if the resulting complex has poor solubility in aqueous solvents, then it may be formulated with a non-ionic surfactant such as Tween, or polyethylene glycol. Thus, the covalent or noncovalent complexes and their physiologically acceptable solvates may be formulated for administration by inhalation or insufflation (either through the mouth or the nose) or oral, buccal, parenteral, rectal administration or, in the case of tumors, directly injected into a solid tumor.
[0235]For oral administration, the pharmaceutical preparation may be in liquid form, for example, solutions, syrups or suspensions, or may be presented as a drug product for reconstitution with water or other suitable vehicle before use. Such liquid preparations may be prepared by conventional means with pharmaceutically acceptable additives such as suspending agents (e.g., sorbitol syrup, cellulose derivatives or hydrogenated edible fats); emulsifying agents (e.g., lecithin or acacia); non-aqueous vehicles (e.g., almond oil, oily esters, or fractionated vegetable oils); and preservatives (e.g., methyl or propyl-p-hydroxybenzoates or sorbic acid). The pharmaceutical compositions may take the form of, for example, tablets or capsules prepared by conventional means with pharmaceutically acceptable excipients such as binding agents (e.g., pregelatinized maize starch, polyvinyl pyrrolidone or hydroxypropyl methylcellulose); fillers (e.g., lactose, microcrystalline cellulose or calcium hydrogen phosphate); lubricants (e.g., magnesium stearate, talc or silica); disintegrants (e.g., potato starch or sodium starch glycolate); or wetting agents (e.g., sodium lauryl sulphate). The tablets may be coated by methods well-known in the art.
[0236]Preparations for oral administration may be suitably formulated to give controlled release of the complexes. Such compositions may take the form of tablets or lozenges formulated in conventional manner.
[0237]For administration by inhalation, the complexes may be conveniently delivered in the form of an aerosol spray presentation from pressurized packs or a nebulizer, with the use of a suitable propellant, e.g., dichlorodifluoromethane, trichlorofluoromethane, dichlorotetrafluoroethane, carbon dioxide or other suitable gas. In the case of a pressurized aerosol the dosage unit may be determined by providing a valve to deliver a metered amount. Capsules and cartridges of, e.g., gelatin for use in an inhaler or insufflator may be formulated containing a powder mix of the complexes and a suitable powder base such as lactose or starch.
[0238]The complexes may be formulated for parenteral administration by injection, e.g., by bolus injection or continuous infusion. Formulations for injection may be presented in unit dosage form, e.g., in ampoules or in multi-dose containers, with an added preservative. The compositions may take such forms as suspensions, solutions or emulsions in oily or aqueous vehicles, and may contain formulatory agents such as suspending, stabilizing and/or dispersing agents. Alternatively, the active ingredient may be in powder form for constitution with a suitable vehicle, e.g., sterile pyrogen-free water, before use.
[0239]The complexes may also be formulated in rectal compositions such as suppositories or retention enemas, e.g., containing conventional suppository bases such as cocoa butter or other glycerides.
[0240]In addition to the formulations described previously, the complexes may also be formulated as a depot preparation. Such long acting formulations may be administered by implantation (for example, subcutaneously or intramuscularly) or by intramuscular injection. Thus, for example, the complexes may be formulated with suitable polymeric or hydrophobic materials (for example, as an emulsion in an acceptable oil) or ion exchange resins, or as sparingly soluble derivatives, for example, as a sparingly soluble salt. Liposomes and emulsions are well known examples of delivery vehicles or carriers for hydrophilic drugs.
[0241]The complexes may, if desired, be presented in a pack or dispenser device which may contain one or more unit dosage forms containing the covalent or noncovalent complexes. The pack may for example comprise metal or plastic foil, such as a blister pack. The pack or dispenser device may be accompanied by instructions for administration.
[0242]The invention also provides kits for carrying out the therapeutic regimens of the invention. Such kits comprise in one or more containers therapeutically or prophylactically effective amounts of the covalent or noncovalent α2M polypeptide-antigenic molecule complexes in pharmaceutically acceptable form. The α2M polypeptide-antigenic molecule complexes in a vial of a kit of the invention may be in the form of a pharmaceutically acceptable solution, e.g., in combination with sterile saline, dextrose solution, or buffered solution, or other pharmaceutically acceptable sterile fluid. Alternatively, the complex may be lyophilized or desiccated; in this instance, the kit optionally further comprises in a container a pharmaceutically acceptable solution (e.g., saline, dextrose solution, etc.), preferably sterile, to reconstitute the complex to form a solution for injection purposes.
[0243]In another embodiment, a kit of the invention further comprises a needle or syringe, preferably packaged in sterile form, for injecting the complex, and/or a packaged alcohol pad. Instructions are optionally included for administration of α2M polypeptide-antigenic molecule complexes by a clinician or by the patient.
6. EXAMPLE
α2M Antagonizes HSP-Mediated Antigen Presentation Via the α2M Receptor
6.1 INTRODUCTION
[0244]The Example presented herein describes the successful identification of an interaction between gp96 and the α2M receptor present in macrophages and dendritic cells in vivo, and the blocking of this interaction by α2M. The experiments presented herein form the basis for the compositions and therapeutic methods of the present invention which relate to the use of α2M polypeptide-antigenic molecule complexes as immunotherapeutic agents for treatment of immune disorders, proliferative disorders, and infectious diseases.
[0245]The Applicant of the present invention noted that certain observations were inconsistent with a "direct transfer" model of HSP-chaperoned peptide antigen presentation. First, the immunogenicity of HSP preparations is dependent on the presence of functional phagocytic cells but not B cells or other nonprofessional antigen-presenting cells (Udono and Srivastava, 1993, supra; Suto and Srivastava, 1995, supra), whereas free peptides can sensitize all cell types. Second, extremely small quantities of HSP-peptide complexes were effective in eliciting specific immunity, i.e., gp96-chaperoned peptides are several hundred times as effective as free peptides in sensitizing macrophages for CTL recognition, suggesting the possibility of a specific uptake mechanism. Third, gp96-chaperoned peptides elicited an MHC I response that was not limited by the size of peptide. Finally, the processing of gp96-peptide complexes in macrophage was found to be sensitive to Brefeldin A (BFA), which blocks transport through the Golgi apparatus, suggesting that processing occurred through an intercellular mechanism. These observations led to the hypothesis that HSP-chaperoned peptides may be processed internally and re-presented by MHC class I molecules on the cell surfaces of macrophages (Suto and Srivastava, 1995, supra). There is also the hypothesis that the mannose receptor is used in the uptake of gp96 but no mechanism has been proposed for the non-glycosylated HSPs, such as HSP70 (Ciupitu et al., 1998, J. Exp. Med., 187: 685-691). Others suggested that a novel intracellular trafficking pathway may be involved for the transport of peptides from the extracellular medium into the lumen of ER) Day et al., 1997, Proc. Natl. Acad. Sci. 94:8065-8069; Nicchitta, 1998, Curr. Opin. in Immunol. 10:103-109). Further suggestions include the involvement of phagocytes which (a) possess an ill-defined pathway to shunt protein from the phagosome into thecytosol where it would enter the normal class I pathway; (b) digest ingested material in lysosomes and regurgitate peptides for loading on the surface to class Imolecules (Bevan, 1995, J. Exp. Med. 192:639-41). The discovery of a receptor for heat shock protein as disclosed herein helps to resolve the paradox of how extracellular antigenic peptides complexed to HSPs can be presented by MHC class Imolecules on antigen presenting cells.
6.2 Materials and Methods
[0246]Affinity chromatography. Proteins (1 mg) in 2 ml volume were incubated with 2 ml of equilibrated AminoLink beads in PBS with a reductant (NaCNBH3) for 1 hour. Uncoupled protein was removed by extensive washing of the column and unreactive groups quenched. Immobilization yields were typically >92% of the starting amount of protein. Columns were stored at 4° C. until used. Such columns were made with gp96 (purified as described in Srivastava et al., 1986, Proc. Natl. Acad. Sci., U.S.A. 83:3407-3411) and albumin. For membrane purification, cells were lysed by dounce homogenization in hypotonic buffer containing PMSF. Unlyzed cells and nuclei were removed by centrifugation at 1000 g for 5 mm. The postnuclear supernatant was centrifuged at 100,000 g for 90 mins. The pellet contains total membranes and was fractionated by aqueous two-phase partition with a dextran/polyethylene glycol biphase. Briefly membranes were resuspended in PEG (33% wt/wt in 0.22 M sodium phosphate buffer, pH 6.5) and underlaid gently with dextran (20% wt/wt in 0.22M sodium phosphate buffer, pH 6.5). The two phases were mixed gently and centrifuged at 2000 g for 15 mins. The white material at the interphase was enriched for plasma membranes, whose proteins were extracted by 2 hr incubation in 20 mM Tris buffer (pH8, containing 0.08% octylglucoside) at 4° C.
[0247]Photo cross-linking of gp96 to putative receptor. The cross-linker (SASD, (Pierce) was labeled with I125 using iodobeads (Pierce). Radiolabeled SASD was covalently attached to gp96 by incubation at room temperature for 1 hr. Free SASD and I125 were removed by size exclusion column (KwikSep columns, Pierce). For cross-linking studies, I125-SASD-gp96 (50 μg gp96) was incubated with purified CD11b+ cells. Unbound protein was removed by washing. All procedures to this point were carried out in very dim light. Proteins were cross-linked with UV light. Cells were lysed with lysis buffer (0.5% NP4O, 10 mM Tris, ImMEDTA, 150 mM NaCl) and treated with 100 mM 2-mercaptoethanol to cleave the cross-linker. Cell lysates were analyzed by SDS-PAGE and autoradiography.
[0248]Re-presentation assays. Re-presentation assays were carried out as described (Suto and Srivastava, 1995, Science 269:1585-1588). Antigen presenting cells (RAW264.7 macrophage cell line) were plated at a 1:1 ratio with AH I-specific T cells in complete RPMI. Approximately 10,000 cells of each type were used. Gp96 (10 μg/ml) chaperoning the AH1-20 mer peptide (RVTYHSPSYVYHQFERRAK, SEQ ID NO: 7) was added to the cells and the entire culture was incubated for 20 hrs. Stimulation of T cells was measured by quantifying the amount of IFN-γ released into the supernatants by ELISA (Endogen).
[0249]Protein Microsequencing. Proteins identified by affinity chromatography were analyzed on SDS-PAGE and stained with coomasie blue or transferred onto PVDF membrane and stained with coomasie blue, all of it under keratin-free conditions. Protein bands were excised with a razor from the gel or membrane. Tryptic peptides from an 80 kDa faint coomassie band were extracted by 50% acetonitrile, 5% formic acid, dried, and loaded onto a 75 m 10 cm, reverse-phase C18, microcapillary column (3 μl vol) and tryptic peptides were separated by on-line microcapillary liquid chromatography-tendem mass spectrometry followed by database searching using the SEQUEST program as previously described. (Gatlin et al., 2000, Anal. Chem. 72:757-63; Link et al., 1999, Nat. Biotechnol. 17:676-82). The analysis was carried out in a data-dependent auto-MS/MS fashion using a Finnigan LCQ iontrap Mass Spectrometer.
6.3 RESULTS
[0250]Identification of an 80 kDa protein as a potential gp96 receptor. Homogenous preparations of gp96 were coupled to FITC and the gp96-FITC was used to stain RAW264.7 cells, shown to be functionally capable of re-presenting gp96-chaperoned peptides. Gp96-FITC but not control albumin-FITC preparations stained the cell surface of RAW264.7 cells (FIG. 1A). Plasma membrane preparations of cell surface-biotinylated RAW264.7 cells were solubilized in 0.08% octyl-glucoside and the soluble extract was applied to a gp96-Sepharose column. The bound proteins were eluted with 3M sodium chloride. SDS-PAGE analysis of the eluate showed 2 major bands of ˜75-80 kDa size (FIG. 1B, top left). Blotting of this gel with avidin-peroxidase showed that both bands were biotinylated, indicating their surface localization (FIG. 1B, bottom left). Affinity purification of membrane extracts of RAW264.7 cells over control serum albumin affinity columns did not result in isolation of any proteins, nor did probing of immunoblots of such gels with avidin peroxidase detect any albumin-binding surface proteins (FIG. 1B, top and bottom center lanes). As an additional control, chromatography of membrane extracts of P815 cells which do not bind gp96-FITC and which do not re-present gp96-chaperoned peptides, on gp96 affinity columns did not result in elution of any gp96-binding proteins (FIG. 1B, top and bottom right lanes).
[0251]In parallel experiments, gp96 molecules were coupled to the radio-iodinated linker sulfosuccinimidyl (4-azidosalicylamido) hexanoate (SASD) which contains a photo cross-linkable group. Gp96-SASD-I125 was pulsed onto peritoneal macrophages, which have been shown previously to re-present gp96-chaperoned peptides (Suto and Srivastava, 1995, Science 269:1585-1588). Excess gp96-SASD was removed by multiple rounds of washing of the cells and photoactivation was carried out by exposure of cells to UV light for 10 mm. Cell lysates were reduced in order to transfer the I125 group to the putative gp96 ligand and were analyzed by SDS-PAGE followed by autoradiography. The gp96 molecule was observed to cross-link to an ˜80 kDa band specifically present in re-presentation-competent macrophage but not in the re-presentation-incompetent P815 cells (FIG. 1C). This band appears to correspond in size to the larger of the two bands seen in eluates of gp96 affinity columns (FIG. 1D). No band corresponding to the lower band in that preparation is seen in the photo cross-linked preparation. The observation of a specific binding of gp96 to an 80 kDa protein in two different re-presentation-competent cell types, but not in a re-presentation-incompetent cell line, and by two independent assays supported the candidacy of the 80 kDa molecule for the gp96 receptor.
[0252]Antiserum against the 80 kDa protein inhibits re-presentation of a gp96-chaperoned antigenicpeptide. The eluates containing the 75-80 kDa proteins were used to immunize a New Zealand white rabbit, and pre-immune and immune sera were used to probe blots of plasma membrane extracts of the re-presentation-competent RAW264.7 and primary peritoneal macrophages and the re-presentation-incompetent P815 cells. The immune but not the pre-immune serum detected the 80 kDa band (and a faint lower 75 kDa band) in plasma membrane extracts of primary macrophage and the RAW264.7 membranes but not of P815 cells (FIG. 2A). The pre-immune and immune sera were tested in a functional assay for their ability to block re-presentation of gp96-chaperoned peptides. The Ld-restricted epitope AH 1 derived from the gp70 antigen of murine colon carcinoma CT26 (Huang et al., 1996, Proc. Natl. Acad. Sci. U.S.A. 93:9730-9735) was used as the model system. Complexes of gp96 with an AH1 precursor (used to inhibit direct presentation) were pulsed onto RAW264.7 cells which were used to stimulate a Ld/AH1-specific CD8+ T cell clone. Release of interferon-γ by the clones was measured as a marker of their activation. RAW264.7 cells were able to re-present gp96-chaperoned AH1 precursor effectively in this assay. It was observed that at the highest concentration, the immune sera inhibited re-presentation completely (FIG. 2B). Although the pre-immune serum was ineffective in inhibiting representation as compared to the immune sera, it did inhibit re-presentation significantly at higher concentrations. The significance of this observation became clear later when we determined the identity of the gp96 receptor. Repeated immunizations with the affinity-purified gp96-binding proteins did not result in corresponding increase in antibody titers.
[0253]Identification of the 80 kDa protein as an amino terminal fragment of the heavy chain of the α2M receptor. The 80 kDa protein eluted from the gp96 affinity column was resolved on SDS-PAGE and visualized by staining with Coomassie Brilliant Blue. The protein band was subjected to in-gel trypsin digestion and mass spectrometry-based protein microsequencing as described in the methods in Section 6.2. Four independent tryptic peptides corresponding to N-terminal region of the α2-macroglobulin (α2M) receptor, designated by immunologists as CD91, were identified (FIG. 3C).
[0254]α2M inhibits re-presentation of a gp96-chaperoned antigenic peptide by RAW264.7. α2M receptor is one of the known natural ligands for the α2M receptor. Its ability to inhibit re-presentation of gp96-chaperoned antigenic peptide AH1 was tested in the assay described in FIG. 2. α2M but not control proteins selectin (CD62) or serum albumin was observed to inhibit re-presentation completely and titratably (FIG. 4). This observation was also consistent with the result in FIG. 2 that while the pre-immune serum did not detect an 80 kDa band in plasma membranes of RAW264.7 cells, it did inhibit re-presentation to some degree at high concentrations. Thus, by structural as well as functional criteria, the α2M receptor was determined to fulfill the criteria essential for a receptor for gp96.
6.4 Discussion
[0255]The α2M receptor, which is also designated CD91, was initially identified as a protein related to the low density lipoprotein (LDL) receptor Related Protein (LRP) (Strickland et al., 1990, J. Biol. Chem. 265:17401-17404; Kristensen et al., 1990, FEBS Lett. 276:151-155). The protein consists of an ˜420 kDa α subunit, an 85 kDa β subunit and a 39 kDa tightly associated molecule (RAP). The α and β subunits are encoded by a single transcript of ˜15 Kb in size (Van Leuven et al., 1993, Biochim. Biophys. Acta. 1173:71-74. The receptor has been shown to be present in cells of the monocytic lineage and in hepatocytes, fibroblasts and keratinocytes. CD91 has been shown previously to bind the activated form of the plasma glycoprotein α2M, which binds to and inhibits a wide variety of endoproteinases. α2M receptor also binds to other ligands such as transforming growth factor β (O'Connor-McCourt et al., 1987, J. Biol. Chem. 262:14090-14099), platelet-derived growth factor (Huang et al., 1984, Proc. Natl. Acad. Sci. U.S.A. 81:342-346), and fibroblast growth factor (Dennis et al., 1989, J. Biol. Chem. 264:7210-7216). α2M is thus believed to regulate, and specifically diminish, the activities of its various ligands. Complexed with these various ligands, α2M binds α2M receptor on the cell surface and is internalized through receptor-mediated endocytosis. Uptake of α2M-complexed ligands has been assumed thus far to be the primary function of the α2M receptor, although a role for it in lipid metabolism is also assumed. α2M receptor ligands other than α2M, such as tissue-specific plasminogen activator-inhibitor complex (Orth et al., 1992, Proc. Natl. Acad. Sci. U.S.A. 89:7422-7426) and urokinase-PAI1 complex (Nykjaer et al., 1992, J. Biol. Chem. 267:14543-14546), have been identified. These ligands attest to a role for α2M receptor in clearing a range of extracellular, plasma products.
[0256]The studies reported here show that the heat shock protein gp96 is an additional ligand for the α2M receptor. The human gp96-coding gene has been mapped previously by us at chromosome 12 (q24.2→q24.3) (Maki et al., 1993, Somatic Cell Mol. Gen. 19:73-81). It is of interest in this regard that the α2M receptor gene has been mapped to the same chromosome and at a not too distant location (q13→q14) (Hilliker et al. Genomics 13:472-474). Gp96 binds α2M receptor directly and not through other ligands such as α2M. Homogenous preparations of gp96, in solution, or cross-linked to a solid matrix, bind to the α2M receptor. Indeed, the major ligand for the α2M receptor, α2M, actually inhibits interaction of gp96 with α2M receptor, instead of promoting it, providing evidence that gp96 is a direct ligand for the α2M receptor. The 80 kDa protein shown to bind gp96 is clearly an amino terminal degradation product of the α subunit of the α2M receptor. Degradation products of the α2M receptor in this size range have also been observed in previous studies (Jensen et al., 1989, Biochem. Arch. 5:171-176), and may indicate the existence of a discrete ectodomain in the α2M receptor which may be particularly sensitive to proteolytic cleavage.
[0257]As shown here, the gp96-α2M receptor interaction provides a new type of function for α2M receptor, a function of a sensor, not only of the extracellular environment with its previously known plasma-based ligands, but also a sensor of the intracellular milieu as well. HSPs such as gp96 are obligate intracellular molecules and are released into the extracellular milieu only under conditions of necrotic (but not apoptotic) cell death. Thus, the α2M receptor may act as a sensor for necrotic cell death (see FIG. 5), just as the scavenger receptor CD36 and the recently identified phosphatidyl serine-binding protein act as sensors of apoptotic cell death and receptors for apoptotic cells (Savill et al., 1992, J. Clin. Invest. 90:1513-1522; Fadok et al., 2000, Nature 405:85-90). Interaction of the macrophages with the apoptotic cells leads to a down-regulation of the inflammatory cytokines such as TNF (Fadok et al., 2000, supra), while gp96-APC interaction leads to re-presentation of gp96-chaperoned peptides by MHC I molecules of the APC, followed by stimulation of antigen-specific T cells (Suto and Srivastava, 1995, supra) and, in addition, secretion of pro-inflammatory cytokines such as TNF, GM-CSF and IL-12. Interestingly, α2M, an independent ligand for the α2M receptor, inhibits representation of gp96-chaperoned peptides by macrophages. This observation suggests that re-presentation of gp96-chaperoned peptides can not occur physiologically in blood, but only within tissues as a result of localized necrotic cell death. This is consistent with the complete absence of gp96 or other HSPs in blood under all conditions tested. Together, these observations point towards a possible mechanism whereby the release of HSPs in the blood as a result of severe tissue injury and lysis will not lead to a systemic and lethal pro-inflammatory cytokine cascade.
[0258]It is possible, therefore, that the α2M receptor renders it possible for the APCs to sample (i) the extracellular milieu of the blood through α2M and other plasma ligands and (ii) the intracellular milieu of the tissues through HSPs, particularly of the gp96 family. The former permits APCs to implement their primordial phagocytic function, while the latter allows them to execute its innate and adaptive immunological functions. Viewed in another perspective, recognition of apoptotic cells by APCs through CD36 or phophatidyl serine, leads to anti-inflammatory signals, while interaction of the APC with necrotic cells through α2M receptor leads to pro-inflammatory innate and adaptive immune responses (see Srivastava et al., 1998, Immunity 8: 657-665).
[0259]The invention is not to be limited in scope by the specific embodiments described which are intended as single illustrations of individual aspects of the invention, and functionally equivalent methods and components are within the scope of the invention. Indeed various modifications of the invention, in addition to those shown and described herein will become apparent to those skilled in the art from the foregoing description and accompanying drawings. Such modifications are intended to fall within the scope of the appended claims.
[0260]All references cited herein, including patent applications, patents, and other publications, are incorporated by reference herein in their entireties for all purposes.
Sequence CWU
1
12114849DNAMus sp.CDS(442)..(14079) 1cgctgctccc cgccagtgca ctgaggaggc
ggaaacgggg gagcccctag tgctccatca 60ggcccctacc aaggcacccc catcgggtcc
acgcccccca ccccccaccc cgcctcctcc 120caattgtgca tttttgcagc cggagtcggc
tccgagatgg ggctgtgagc ttcgccctgg 180gagggggaga ggagcgagga gtaaagcagg
ggtgaagggt tcgaatttgg gggcaggggg 240cgcacccgcg tcagcaggcc cttcccaggg
ggctcggaac tgtaccattt cacctatgcc 300cctggttcgc tttgcttaag gaaggataag
atagaagagt cggggagagg aagataaagg 360gggacccccc aattgggggg ggcgaggaca
agaagtaaca ggaccagagg gtgggggctg 420ctgtttgcat cggcccacac c atg ctg
acc ccg ccg ttg ctg ctg ctc gtg 471Met Leu Thr Pro Pro Leu Leu Leu
Leu Val1 5 10ccg ctg ctt tca gct ctg gtc
tcc ggg gcc act atg gat gcc cct aaa 519Pro Leu Leu Ser Ala Leu Val
Ser Gly Ala Thr Met Asp Ala Pro Lys 15 20
25act tgc agc cct aag cag ttt gcc tgc aga gac caa atc
acc tgt atc 567Thr Cys Ser Pro Lys Gln Phe Ala Cys Arg Asp Gln Ile
Thr Cys Ile 30 35 40tca aag
ggc tgg cgg tgt gac ggt gaa aga gat tgc ccc gac ggc tct 615Ser Lys
Gly Trp Arg Cys Asp Gly Glu Arg Asp Cys Pro Asp Gly Ser 45
50 55gat gaa gcc cct gag atc tgt cca cag agt
aaa gcc cag aga tgc ccg 663Asp Glu Ala Pro Glu Ile Cys Pro Gln Ser
Lys Ala Gln Arg Cys Pro 60 65 70cca
aat gag cac agt tgt ctg ggg act gag cta tgt gtc ccc atg tct 711Pro
Asn Glu His Ser Cys Leu Gly Thr Glu Leu Cys Val Pro Met Ser75
80 85 90cgt ctc tgc aac ggg atc
cag gac tgc atg gat ggc tca gac gag ggt 759Arg Leu Cys Asn Gly Ile
Gln Asp Cys Met Asp Gly Ser Asp Glu Gly 95
100 105gct cac tgc cga gag ctc cga gcc aac tgt tct cga
atg ggt tgt caa 807Ala His Cys Arg Glu Leu Arg Ala Asn Cys Ser Arg
Met Gly Cys Gln 110 115 120cac
cat tgt gta cct aca ccc agt ggg ccc acg tgc tac tgt aac agc 855His
His Cys Val Pro Thr Pro Ser Gly Pro Thr Cys Tyr Cys Asn Ser 125
130 135agc ttc cag ctc gag gca gat ggc aag
acg tgc aaa gat ttt gac gag 903Ser Phe Gln Leu Glu Ala Asp Gly Lys
Thr Cys Lys Asp Phe Asp Glu 140 145
150tgt tcc gtg tat ggc acc tgc agc cag ctt tgc acc aac aca gat ggc
951Cys Ser Val Tyr Gly Thr Cys Ser Gln Leu Cys Thr Asn Thr Asp Gly155
160 165 170tcc ttc aca tgt
ggc tgt gtt gaa ggc tac ctg ctg caa ccg gac aac 999Ser Phe Thr Cys
Gly Cys Val Glu Gly Tyr Leu Leu Gln Pro Asp Asn 175
180 185cgc tcc tgc aag gcc aag aat gag cca gta
gat cgg ccg cca gtg cta 1047Arg Ser Cys Lys Ala Lys Asn Glu Pro Val
Asp Arg Pro Pro Val Leu 190 195
200ctg att gcc aac tct cag aac atc cta gct acg tac ctg agt ggg gcc
1095Leu Ile Ala Asn Ser Gln Asn Ile Leu Ala Thr Tyr Leu Ser Gly Ala
205 210 215caa gtg tct acc atc aca ccc
acc agc acc cga caa acc acg gcc atg 1143Gln Val Ser Thr Ile Thr Pro
Thr Ser Thr Arg Gln Thr Thr Ala Met 220 225
230gac ttc agt tat gcc aat gag acc gta tgc tgg gtg cac gtt ggg gac
1191Asp Phe Ser Tyr Ala Asn Glu Thr Val Cys Trp Val His Val Gly Asp235
240 245 250agt gct gcc cag
aca cag ctc aag tgt gcc cgg atg cct ggc ctg aag 1239Ser Ala Ala Gln
Thr Gln Leu Lys Cys Ala Arg Met Pro Gly Leu Lys 255
260 265ggc ttt gtg gat gag cat acc atc aac atc
tcc ctc agc ctg cac cac 1287Gly Phe Val Asp Glu His Thr Ile Asn Ile
Ser Leu Ser Leu His His 270 275
280gtg gag cag atg gca atc gac tgg ctg acg gga aac ttc tac ttt gtc
1335Val Glu Gln Met Ala Ile Asp Trp Leu Thr Gly Asn Phe Tyr Phe Val
285 290 295gac gac att gac gac agg atc
ttt gtc tgt aac cga aac ggg gac acc 1383Asp Asp Ile Asp Asp Arg Ile
Phe Val Cys Asn Arg Asn Gly Asp Thr 300 305
310tgt gtc act ctg ctg gac ctg gaa ctc tac aac ccc aaa ggc atc gcc
1431Cys Val Thr Leu Leu Asp Leu Glu Leu Tyr Asn Pro Lys Gly Ile Ala315
320 325 330ttg gac ccc gcc
atg ggg aag gtg ttc ttc act gac tac ggg cag atc 1479Leu Asp Pro Ala
Met Gly Lys Val Phe Phe Thr Asp Tyr Gly Gln Ile 335
340 345cca aag gtg gag cgc tgt gac atg gat gga
cag aac cgc acc aag ctg 1527Pro Lys Val Glu Arg Cys Asp Met Asp Gly
Gln Asn Arg Thr Lys Leu 350 355
360gtg gat agc aag atc gtg ttt cca cac ggc atc acc ctg gac ctg gtc
1575Val Asp Ser Lys Ile Val Phe Pro His Gly Ile Thr Leu Asp Leu Val
365 370 375agc cgc ctc gtc tac tgg gcg
gac gcc tac cta gac tac atc gag gtg 1623Ser Arg Leu Val Tyr Trp Ala
Asp Ala Tyr Leu Asp Tyr Ile Glu Val 380 385
390gta gac tac gaa ggg aag ggt cgg cag acc atc atc caa ggc atc ctg
1671Val Asp Tyr Glu Gly Lys Gly Arg Gln Thr Ile Ile Gln Gly Ile Leu395
400 405 410atc gag cac ctg
tac ggc ctg acc gtg ttt gag aac tat ctc tac gcc 1719Ile Glu His Leu
Tyr Gly Leu Thr Val Phe Glu Asn Tyr Leu Tyr Ala 415
420 425acc aac tcg gac aat gcc aac acg cag cag
aag acg agc gtg atc cga 1767Thr Asn Ser Asp Asn Ala Asn Thr Gln Gln
Lys Thr Ser Val Ile Arg 430 435
440gtg aac cgg ttc aac agt act gag tac cag gtc gtc acc cgt gtg gac
1815Val Asn Arg Phe Asn Ser Thr Glu Tyr Gln Val Val Thr Arg Val Asp
445 450 455aag ggt ggt gcc ctg cat atc
tac cac cag cga cgc cag ccc cga gtg 1863Lys Gly Gly Ala Leu His Ile
Tyr His Gln Arg Arg Gln Pro Arg Val 460 465
470cgg agt cac gcc tgt gag aat gac cag tac ggg aag cca ggt ggc tgc
1911Arg Ser His Ala Cys Glu Asn Asp Gln Tyr Gly Lys Pro Gly Gly Cys475
480 485 490tcc gac atc tgc
ctc ctg gcc aac agt cac aag gca agg acc tgc agg 1959Ser Asp Ile Cys
Leu Leu Ala Asn Ser His Lys Ala Arg Thr Cys Arg 495
500 505tgc agg tct ggc ttc agc ctg gga agt gat
ggg aag tct tgt aag aaa 2007Cys Arg Ser Gly Phe Ser Leu Gly Ser Asp
Gly Lys Ser Cys Lys Lys 510 515
520cct gaa cat gag ctg ttc ctc gtg tat ggc aag ggc cga cca ggc atc
2055Pro Glu His Glu Leu Phe Leu Val Tyr Gly Lys Gly Arg Pro Gly Ile
525 530 535att aga ggc atg gac atg ggg
gcc aag gtc cca gat gag cac atg atc 2103Ile Arg Gly Met Asp Met Gly
Ala Lys Val Pro Asp Glu His Met Ile 540 545
550ccc atc gag aac ctt atg aat cca cgc gct ctg gac ttc cac gcc gag
2151Pro Ile Glu Asn Leu Met Asn Pro Arg Ala Leu Asp Phe His Ala Glu555
560 565 570acc ggc ttc atc
tac ttt gct gac acc acc agc tac ctc att ggc cgc 2199Thr Gly Phe Ile
Tyr Phe Ala Asp Thr Thr Ser Tyr Leu Ile Gly Arg 575
580 585cag aaa att gat ggc acg gag aga gag act
atc ctg aag gat ggc atc 2247Gln Lys Ile Asp Gly Thr Glu Arg Glu Thr
Ile Leu Lys Asp Gly Ile 590 595
600cac aat gtg gag ggc gta gcc gtg gac tgg atg gga gac aat ctt tac
2295His Asn Val Glu Gly Val Ala Val Asp Trp Met Gly Asp Asn Leu Tyr
605 610 615tgg act gat gat ggc ccc aag
aag acc att agt gtg gcc agg ctg gag 2343Trp Thr Asp Asp Gly Pro Lys
Lys Thr Ile Ser Val Ala Arg Leu Glu 620 625
630aaa gcc gct cag acc cgg aag act cta att gag ggc aag atg aca cac
2391Lys Ala Ala Gln Thr Arg Lys Thr Leu Ile Glu Gly Lys Met Thr His635
640 645 650ccc agg gcc att
gta gtg gat cca ctc aat ggg tgg atg tac tgg aca 2439Pro Arg Ala Ile
Val Val Asp Pro Leu Asn Gly Trp Met Tyr Trp Thr 655
660 665gac tgg gag gag gac ccc aag gac agt cgg
cga ggg cgg ctc gag agg 2487Asp Trp Glu Glu Asp Pro Lys Asp Ser Arg
Arg Gly Arg Leu Glu Arg 670 675
680gct tgg atg gac ggc tca cac cga gat atc ttt gtc acc tcc aag aca
2535Ala Trp Met Asp Gly Ser His Arg Asp Ile Phe Val Thr Ser Lys Thr
685 690 695gtg ctt tgg ccc aat ggg cta
agc ctg gat atc cca gcc gga cgc ctc 2583Val Leu Trp Pro Asn Gly Leu
Ser Leu Asp Ile Pro Ala Gly Arg Leu 700 705
710tac tgg gtg gat gcc ttc tat gac cga att gag acc ata ctg ctc aat
2631Tyr Trp Val Asp Ala Phe Tyr Asp Arg Ile Glu Thr Ile Leu Leu Asn715
720 725 730ggc aca gac cgg
aag att gta tat gag ggt cct gaa ctg aat cat gcc 2679Gly Thr Asp Arg
Lys Ile Val Tyr Glu Gly Pro Glu Leu Asn His Ala 735
740 745ttc ggc ctg tgt cac cat ggc aac tac ctc
ttt tgg acc gag tac cgg 2727Phe Gly Leu Cys His His Gly Asn Tyr Leu
Phe Trp Thr Glu Tyr Arg 750 755
760agc ggc agc gtc tac cgc ttg gaa cgg ggc gtg gca ggc gca ccg ccc
2775Ser Gly Ser Val Tyr Arg Leu Glu Arg Gly Val Ala Gly Ala Pro Pro
765 770 775act gtg acc ctt ctg cgc agc
gag aga ccg cct atc ttt gag atc cga 2823Thr Val Thr Leu Leu Arg Ser
Glu Arg Pro Pro Ile Phe Glu Ile Arg 780 785
790atg tac gac gcg cac gag cag caa gtg ggt acc aac aaa tgc cgg gta
2871Met Tyr Asp Ala His Glu Gln Gln Val Gly Thr Asn Lys Cys Arg Val795
800 805 810aat aac gga ggc
tgc agc agc ctg tgc ctc gcc acc ccc ggg agc cgc 2919Asn Asn Gly Gly
Cys Ser Ser Leu Cys Leu Ala Thr Pro Gly Ser Arg 815
820 825cag tgt gcc tgt gcc gag gac cag gtg ttg
gac aca gat ggt gtc acc 2967Gln Cys Ala Cys Ala Glu Asp Gln Val Leu
Asp Thr Asp Gly Val Thr 830 835
840tgc ttg gcg aac cca tcc tac gtg ccc cca ccc cag tgc cag ccg ggc
3015Cys Leu Ala Asn Pro Ser Tyr Val Pro Pro Pro Gln Cys Gln Pro Gly
845 850 855cag ttt gcc tgt gcc aac aac
cgc tgc atc cag gag cgc tgg aag tgt 3063Gln Phe Ala Cys Ala Asn Asn
Arg Cys Ile Gln Glu Arg Trp Lys Cys 860 865
870gac gga gac aac gac tgt ctg gac aac agc gat gag gcc cca gca ctg
3111Asp Gly Asp Asn Asp Cys Leu Asp Asn Ser Asp Glu Ala Pro Ala Leu875
880 885 890tgc cat caa cac
acc tgt ccc tcg gac cga ttc aag tgt gag aac aac 3159Cys His Gln His
Thr Cys Pro Ser Asp Arg Phe Lys Cys Glu Asn Asn 895
900 905cgg tgt atc ccc aac cgc tgg ctc tgt gat
ggg gat aat gat tgt ggc 3207Arg Cys Ile Pro Asn Arg Trp Leu Cys Asp
Gly Asp Asn Asp Cys Gly 910 915
920aac agc gag gac gaa tcc aat gcc acg tgc tca gcc cgc acc tgt cca
3255Asn Ser Glu Asp Glu Ser Asn Ala Thr Cys Ser Ala Arg Thr Cys Pro
925 930 935ccc aac cag ttc tcc tgt gcc
agt ggc cga tgc att cct atc tca tgg 3303Pro Asn Gln Phe Ser Cys Ala
Ser Gly Arg Cys Ile Pro Ile Ser Trp 940 945
950acc tgt gat ctg gat gat gac tgt ggg gac cgg tcc gat gag tca gcc
3351Thr Cys Asp Leu Asp Asp Asp Cys Gly Asp Arg Ser Asp Glu Ser Ala955
960 965 970tca tgc gcc tac
ccc acc tgc ttc ccc ctg act caa ttt acc tgc aac 3399Ser Cys Ala Tyr
Pro Thr Cys Phe Pro Leu Thr Gln Phe Thr Cys Asn 975
980 985aat ggc aga tgt att aac atc aac tgg cgg
tgt gac aac gac aat gac 3447Asn Gly Arg Cys Ile Asn Ile Asn Trp Arg
Cys Asp Asn Asp Asn Asp 990 995
1000tgt ggg gac aac agc gac gaa gcc ggc tgc agt cac tcc tgc tcc
3492Cys Gly Asp Asn Ser Asp Glu Ala Gly Cys Ser His Ser Cys Ser
1005 1010 1015agt acc cag ttc aag tgc
aac agt ggc aga tgc atc ccc gag cac 3537Ser Thr Gln Phe Lys Cys
Asn Ser Gly Arg Cys Ile Pro Glu His 1020 1025
1030tgg acg tgt gat ggg gac aat gat tgt ggg gac tac agc
gac gag 3582Trp Thr Cys Asp Gly Asp Asn Asp Cys Gly Asp Tyr Ser
Asp Glu 1035 1040 1045aca cac gcc
aac tgt acc aac cag gct aca aga cct cct ggt ggc 3627Thr His Ala
Asn Cys Thr Asn Gln Ala Thr Arg Pro Pro Gly Gly 1050
1055 1060tgc cac tcg gat gag ttc cag tgc ccg cta
gat ggc ctg tgc atc 3672Cys His Ser Asp Glu Phe Gln Cys Pro Leu
Asp Gly Leu Cys Ile 1065 1070
1075ccc ctg agg tgg cgc tgc gac ggg gac acc gac tgc atg gat tcc
3717Pro Leu Arg Trp Arg Cys Asp Gly Asp Thr Asp Cys Met Asp Ser
1080 1085 1090agc gat gag aag agc tgt
gag ggc gtg acc cat gtt tgt gac ccg 3762Ser Asp Glu Lys Ser Cys
Glu Gly Val Thr His Val Cys Asp Pro 1095 1100
1105aat gtc aag ttt ggc tgc aag gac tcc gcc cgg tgc atc
agc aag 3807Asn Val Lys Phe Gly Cys Lys Asp Ser Ala Arg Cys Ile
Ser Lys 1110 1115 1120gcg tgg gtg
tgt gat ggc gac agc gac tgt gaa gat aac tcc gac 3852Ala Trp Val
Cys Asp Gly Asp Ser Asp Cys Glu Asp Asn Ser Asp 1125
1130 1135gag gag aac tgt gag gcc ctg gcc tgc agg
cca ccc tcc cat ccc 3897Glu Glu Asn Cys Glu Ala Leu Ala Cys Arg
Pro Pro Ser His Pro 1140 1145
1150tgc gcc aac aac acc tct gtc tgc ctg cct cct gac aag ctg tgc
3942Cys Ala Asn Asn Thr Ser Val Cys Leu Pro Pro Asp Lys Leu Cys
1155 1160 1165gac ggc aag gat gac tgt
gga gac ggc tcg gat gag ggc gag ctc 3987Asp Gly Lys Asp Asp Cys
Gly Asp Gly Ser Asp Glu Gly Glu Leu 1170 1175
1180tgt gac cag tgt tct ctg aat aat ggt ggc tgt agt cac
aac tgc 4032Cys Asp Gln Cys Ser Leu Asn Asn Gly Gly Cys Ser His
Asn Cys 1185 1190 1195tca gtg gcc
cct ggt gaa ggc atc gtg tgc tct tgc cct ctg ggc 4077Ser Val Ala
Pro Gly Glu Gly Ile Val Cys Ser Cys Pro Leu Gly 1200
1205 1210atg gag ctg ggc tct gac aac cac acc tgc
cag atc cag agc tac 4122Met Glu Leu Gly Ser Asp Asn His Thr Cys
Gln Ile Gln Ser Tyr 1215 1220
1225tgt gcc aag cac ctc aaa tgc agc cag aag tgt gac cag aac aag
4167Cys Ala Lys His Leu Lys Cys Ser Gln Lys Cys Asp Gln Asn Lys
1230 1235 1240ttc agt gtg aag tgc tcc
tgc tac gag ggc tgg gtc ttg gag cct 4212Phe Ser Val Lys Cys Ser
Cys Tyr Glu Gly Trp Val Leu Glu Pro 1245 1250
1255gac ggg gaa acg tgc cgc agt ctg gat ccc ttc aaa ctg
ttc atc 4257Asp Gly Glu Thr Cys Arg Ser Leu Asp Pro Phe Lys Leu
Phe Ile 1260 1265 1270atc ttc tcc
aac cgc cac gag atc agg cgc att gac ctt cac aag 4302Ile Phe Ser
Asn Arg His Glu Ile Arg Arg Ile Asp Leu His Lys 1275
1280 1285ggg gac tac agc gtc cta gtg cct ggc ctg
cgc aac act att gcc 4347Gly Asp Tyr Ser Val Leu Val Pro Gly Leu
Arg Asn Thr Ile Ala 1290 1295
1300ctg gac ttc cac ctc agc cag agt gcc ctc tac tgg acc gac gcg
4392Leu Asp Phe His Leu Ser Gln Ser Ala Leu Tyr Trp Thr Asp Ala
1305 1310 1315gta gag gac aag atc tac
cgt ggg aaa ctc ctg gac aac gga gcc 4437Val Glu Asp Lys Ile Tyr
Arg Gly Lys Leu Leu Asp Asn Gly Ala 1320 1325
1330ctg acc agc ttt gag gtg gtg att cag tat ggc ttg gcc
aca cca 4482Leu Thr Ser Phe Glu Val Val Ile Gln Tyr Gly Leu Ala
Thr Pro 1335 1340 1345gag ggc ctg
gct gta gat tgg att gca ggc aac atc tac tgg gtg 4527Glu Gly Leu
Ala Val Asp Trp Ile Ala Gly Asn Ile Tyr Trp Val 1350
1355 1360gag agc aac ctg gac cag atc gaa gtg gcc
aag ctg gac gga acc 4572Glu Ser Asn Leu Asp Gln Ile Glu Val Ala
Lys Leu Asp Gly Thr 1365 1370
1375ctc cga acc act ctg ctg gcg ggt gac att gag cac ccg agg gcc
4617Leu Arg Thr Thr Leu Leu Ala Gly Asp Ile Glu His Pro Arg Ala
1380 1385 1390atc gct ctg gac cct cgg
gat ggg att ctg ttt tgg aca gac tgg 4662Ile Ala Leu Asp Pro Arg
Asp Gly Ile Leu Phe Trp Thr Asp Trp 1395 1400
1405gat gcc agc ctg cca cga atc gag gct gca tcc atg agt
gga gct 4707Asp Ala Ser Leu Pro Arg Ile Glu Ala Ala Ser Met Ser
Gly Ala 1410 1415 1420ggc cgc cga
acc atc cac cgg gag aca ggc tct ggg ggc tgc gcc 4752Gly Arg Arg
Thr Ile His Arg Glu Thr Gly Ser Gly Gly Cys Ala 1425
1430 1435aat ggg ctc acc gtg gat tac ctg gag aag
cgc atc ctc tgg att 4797Asn Gly Leu Thr Val Asp Tyr Leu Glu Lys
Arg Ile Leu Trp Ile 1440 1445
1450gat gct agg tca gat gcc atc tat tca gcc cgg tat gac ggc tcc
4842Asp Ala Arg Ser Asp Ala Ile Tyr Ser Ala Arg Tyr Asp Gly Ser
1455 1460 1465ggc cac atg gag gtg ctt
cgg gga cac gag ttc ctg tca cac cca 4887Gly His Met Glu Val Leu
Arg Gly His Glu Phe Leu Ser His Pro 1470 1475
1480ttt gcc gtg aca ctg tac ggt ggg gag gtg tac tgg acc
gac tgg 4932Phe Ala Val Thr Leu Tyr Gly Gly Glu Val Tyr Trp Thr
Asp Trp 1485 1490 1495cga aca aat
aca ctg gct aag gcc aac aag tgg act ggc cac aac 4977Arg Thr Asn
Thr Leu Ala Lys Ala Asn Lys Trp Thr Gly His Asn 1500
1505 1510gtc acc gtg gta cag agg acc aac acc cag
ccc ttc gac ctg cag 5022Val Thr Val Val Gln Arg Thr Asn Thr Gln
Pro Phe Asp Leu Gln 1515 1520
1525gtg tat cac cct tcc cgg cag ccc atg gct cca aac cca tgt gag
5067Val Tyr His Pro Ser Arg Gln Pro Met Ala Pro Asn Pro Cys Glu
1530 1535 1540gcc aat ggc ggc cgg ggc
ccc tgt tcc cat ctg tgc ctc atc aac 5112Ala Asn Gly Gly Arg Gly
Pro Cys Ser His Leu Cys Leu Ile Asn 1545 1550
1555tac aac cgg acc gtc tcc tgg gcc tgt ccc cac ctc atg
aag ctg 5157Tyr Asn Arg Thr Val Ser Trp Ala Cys Pro His Leu Met
Lys Leu 1560 1565 1570cac aag gac
aac acc acc tgc tat gag ttt aag aag ttc ctg ctg 5202His Lys Asp
Asn Thr Thr Cys Tyr Glu Phe Lys Lys Phe Leu Leu 1575
1580 1585tac gca cgt cag atg gag atc cgg ggc gtg
gac ctg gat gcc ccg 5247Tyr Ala Arg Gln Met Glu Ile Arg Gly Val
Asp Leu Asp Ala Pro 1590 1595
1600tac tac aat tat atc atc tcc ttc acg gtg cct gat atc gac aat
5292Tyr Tyr Asn Tyr Ile Ile Ser Phe Thr Val Pro Asp Ile Asp Asn
1605 1610 1615gtc acg gtg ctg gac tat
gat gcc cga gag cag cga gtt tac tgg 5337Val Thr Val Leu Asp Tyr
Asp Ala Arg Glu Gln Arg Val Tyr Trp 1620 1625
1630tct gat gtg cgg act caa gcc atc aaa agg gca ttt atc
aac ggc 5382Ser Asp Val Arg Thr Gln Ala Ile Lys Arg Ala Phe Ile
Asn Gly 1635 1640 1645act ggc gtg
gag acc gtt gtc tct gca gac ttg ccc aac gcc cac 5427Thr Gly Val
Glu Thr Val Val Ser Ala Asp Leu Pro Asn Ala His 1650
1655 1660ggg ctg gct gtg gac tgg gtc tcc cga aat
ctg ttt tgg aca agt 5472Gly Leu Ala Val Asp Trp Val Ser Arg Asn
Leu Phe Trp Thr Ser 1665 1670
1675tac gac acc aac aag aag cag att aac gtg gcc cgg ctg gac ggc
5517Tyr Asp Thr Asn Lys Lys Gln Ile Asn Val Ala Arg Leu Asp Gly
1680 1685 1690tcc ttc aag aat gcg gtg
gtg cag ggc ctg gag cag ccc cac ggc 5562Ser Phe Lys Asn Ala Val
Val Gln Gly Leu Glu Gln Pro His Gly 1695 1700
1705ctg gtc gtc cac ccg ctt cgt ggc aag ctc tac tgg act
gat ggg 5607Leu Val Val His Pro Leu Arg Gly Lys Leu Tyr Trp Thr
Asp Gly 1710 1715 1720gac aac atc
agc atg gcc aac atg gat ggg agc aac cac act ctg 5652Asp Asn Ile
Ser Met Ala Asn Met Asp Gly Ser Asn His Thr Leu 1725
1730 1735ctc ttc agt ggc cag aag ggc cct gtg ggg
ttg gcc att gac ttc 5697Leu Phe Ser Gly Gln Lys Gly Pro Val Gly
Leu Ala Ile Asp Phe 1740 1745
1750cct gag agc aaa ctc tac tgg atc agc tct ggg aac cac aca atc
5742Pro Glu Ser Lys Leu Tyr Trp Ile Ser Ser Gly Asn His Thr Ile
1755 1760 1765aac cgt tgc aat ctg gat
ggg agc gag ctg gag gtc atc gac acc 5787Asn Arg Cys Asn Leu Asp
Gly Ser Glu Leu Glu Val Ile Asp Thr 1770 1775
1780atg cgg agc cag ctg ggc aag gcc act gcc ctg gcc atc
atg ggg 5832Met Arg Ser Gln Leu Gly Lys Ala Thr Ala Leu Ala Ile
Met Gly 1785 1790 1795gac aag ctg
tgg tgg gca gat cag gtg tca gag aag atg ggc acg 5877Asp Lys Leu
Trp Trp Ala Asp Gln Val Ser Glu Lys Met Gly Thr 1800
1805 1810tgc aac aaa gcc gat ggc tct ggg tcc gtg
gtg ctg cgg aac agt 5922Cys Asn Lys Ala Asp Gly Ser Gly Ser Val
Val Leu Arg Asn Ser 1815 1820
1825acc acg ttg gtt atg cac atg aag gtg tat gac gag agc atc cag
5967Thr Thr Leu Val Met His Met Lys Val Tyr Asp Glu Ser Ile Gln
1830 1835 1840cta gag cat gag ggc acc
aac ccc tgc agt gtc aac aac gga gac 6012Leu Glu His Glu Gly Thr
Asn Pro Cys Ser Val Asn Asn Gly Asp 1845 1850
1855tgt tcc cag ctc tgc ctg cca aca tca gag acg act cgc
tcc tgt 6057Cys Ser Gln Leu Cys Leu Pro Thr Ser Glu Thr Thr Arg
Ser Cys 1860 1865 1870atg tgt aca
gcc ggt tac agc ctc cgg agc gga cag cag gcc tgt 6102Met Cys Thr
Ala Gly Tyr Ser Leu Arg Ser Gly Gln Gln Ala Cys 1875
1880 1885gag ggt gtg ggc tct ttt ctc ctg tac tct
gta cat gag gga att 6147Glu Gly Val Gly Ser Phe Leu Leu Tyr Ser
Val His Glu Gly Ile 1890 1895
1900cgg ggg att cca cta gat ccc aat gac aag tcg gat gcc ctg gtc
6192Arg Gly Ile Pro Leu Asp Pro Asn Asp Lys Ser Asp Ala Leu Val
1905 1910 1915cca gtg tcc gga act tca
ctg gct gtc gga atc gac ttc cat gcc 6237Pro Val Ser Gly Thr Ser
Leu Ala Val Gly Ile Asp Phe His Ala 1920 1925
1930gaa aat gac act att tat tgg gtg gat atg ggc cta agc
acc atc 6282Glu Asn Asp Thr Ile Tyr Trp Val Asp Met Gly Leu Ser
Thr Ile 1935 1940 1945agc agg gcc
aag cgt gac cag aca tgg cga gag gat gtg gtg acc 6327Ser Arg Ala
Lys Arg Asp Gln Thr Trp Arg Glu Asp Val Val Thr 1950
1955 1960aac ggt att ggc cgt gtg gag ggc atc gcc
gtg gac tgg atc gca 6372Asn Gly Ile Gly Arg Val Glu Gly Ile Ala
Val Asp Trp Ile Ala 1965 1970
1975ggc aac ata tac tgg acg gac cag ggc ttc gat gtc atc gag gtt
6417Gly Asn Ile Tyr Trp Thr Asp Gln Gly Phe Asp Val Ile Glu Val
1980 1985 1990gcc cgg ctc aat ggc tct
ttt cgt tat gtg gtc att tcc cag ggt 6462Ala Arg Leu Asn Gly Ser
Phe Arg Tyr Val Val Ile Ser Gln Gly 1995 2000
2005ctg gac aag cct cgg gcc atc act gtc cac cca gag aag
ggg tac 6507Leu Asp Lys Pro Arg Ala Ile Thr Val His Pro Glu Lys
Gly Tyr 2010 2015 2020ttg ttc tgg
acc gag tgg ggt cat tac cca cgt att gag cgg tct 6552Leu Phe Trp
Thr Glu Trp Gly His Tyr Pro Arg Ile Glu Arg Ser 2025
2030 2035cgc ctt gat ggc aca gag aga gtg gtg ttg
gtt aat gtc agc atc 6597Arg Leu Asp Gly Thr Glu Arg Val Val Leu
Val Asn Val Ser Ile 2040 2045
2050agc tgg ccc aat ggc atc tca gta gac tat cag ggc ggc aag ctc
6642Ser Trp Pro Asn Gly Ile Ser Val Asp Tyr Gln Gly Gly Lys Leu
2055 2060 2065tac tgg tgt gat gct cgg
atg gac aag atc gag cgc atc gac ctg 6687Tyr Trp Cys Asp Ala Arg
Met Asp Lys Ile Glu Arg Ile Asp Leu 2070 2075
2080gaa acg ggc gag aac cgg gag gtg gtc ctg tcc agc aat
aac atg 6732Glu Thr Gly Glu Asn Arg Glu Val Val Leu Ser Ser Asn
Asn Met 2085 2090 2095gat atg ttc
tcc gtg tcc gtg ttt gag gac ttc atc tac tgg agt 6777Asp Met Phe
Ser Val Ser Val Phe Glu Asp Phe Ile Tyr Trp Ser 2100
2105 2110gac aga act cac gcc aat ggc tcc atc aag
cgc ggc tgc aaa gac 6822Asp Arg Thr His Ala Asn Gly Ser Ile Lys
Arg Gly Cys Lys Asp 2115 2120
2125aat gct aca gac tcc gtg cct ctg agg aca ggc att ggt gtt cag
6867Asn Ala Thr Asp Ser Val Pro Leu Arg Thr Gly Ile Gly Val Gln
2130 2135 2140ctt aaa gac atc aag gtc
ttc aac agg gac agg cag aag ggt acc 6912Leu Lys Asp Ile Lys Val
Phe Asn Arg Asp Arg Gln Lys Gly Thr 2145 2150
2155aat gtg tgc gcg gta gcc aac ggc ggg tgc cag cag ctc
tgc ttg 6957Asn Val Cys Ala Val Ala Asn Gly Gly Cys Gln Gln Leu
Cys Leu 2160 2165 2170tat cgg ggt
ggc gga cag cga gcc tgt gcc tgt gcc cac ggg atg 7002Tyr Arg Gly
Gly Gly Gln Arg Ala Cys Ala Cys Ala His Gly Met 2175
2180 2185ctg gca gaa gac ggg gcc tca tgc cga gag
tac gct ggc tac ctg 7047Leu Ala Glu Asp Gly Ala Ser Cys Arg Glu
Tyr Ala Gly Tyr Leu 2190 2195
2200ctc tac tca gag cgg acc atc ctc aag agc atc cac ctg tcg gat
7092Leu Tyr Ser Glu Arg Thr Ile Leu Lys Ser Ile His Leu Ser Asp
2205 2210 2215gag cgt aac ctc aac gca
ccg gtg cag ccc ttt gaa gac ccc gag 7137Glu Arg Asn Leu Asn Ala
Pro Val Gln Pro Phe Glu Asp Pro Glu 2220 2225
2230cac atg aaa aat gtc atc gcc ctg gcc ttt gac tac cga
gca ggc 7182His Met Lys Asn Val Ile Ala Leu Ala Phe Asp Tyr Arg
Ala Gly 2235 2240 2245acc tcc ccg
ggg acc cct aac cgc atc ttc ttc agt gac atc cac 7227Thr Ser Pro
Gly Thr Pro Asn Arg Ile Phe Phe Ser Asp Ile His 2250
2255 2260ttt ggg aac atc cag cag atc aat gac gat
ggc tcg ggc agg acc 7272Phe Gly Asn Ile Gln Gln Ile Asn Asp Asp
Gly Ser Gly Arg Thr 2265 2270
2275acc atc gtg gaa aat gtg ggc tct gtg gaa ggc ctg gcc tat cac
7317Thr Ile Val Glu Asn Val Gly Ser Val Glu Gly Leu Ala Tyr His
2280 2285 2290cgt ggc tgg gac aca ctg
tac tgg aca agc tac acc aca tcc acc 7362Arg Gly Trp Asp Thr Leu
Tyr Trp Thr Ser Tyr Thr Thr Ser Thr 2295 2300
2305atc acc cgc cac acc gtg gac cag act cgc cca ggg gcc
ttc gag 7407Ile Thr Arg His Thr Val Asp Gln Thr Arg Pro Gly Ala
Phe Glu 2310 2315 2320agg gag aca
gtc atc acc atg tcc gga gac gac cac ccg aga gcc 7452Arg Glu Thr
Val Ile Thr Met Ser Gly Asp Asp His Pro Arg Ala 2325
2330 2335ttt gtg ctg gat gag tgc cag aac ctg atg
ttc tgg acc aat tgg 7497Phe Val Leu Asp Glu Cys Gln Asn Leu Met
Phe Trp Thr Asn Trp 2340 2345
2350aac gag ctc cat cca agc atc atg cgg gca gcc cta tcc gga gcc
7542Asn Glu Leu His Pro Ser Ile Met Arg Ala Ala Leu Ser Gly Ala
2355 2360 2365aac gtc ctg acc ctc att
gag aag gac atc cgc acg ccc aat ggg 7587Asn Val Leu Thr Leu Ile
Glu Lys Asp Ile Arg Thr Pro Asn Gly 2370 2375
2380ttg gcc atc gac cac cgg gcg gag aag ctg tac ttc tcg
gat gcc 7632Leu Ala Ile Asp His Arg Ala Glu Lys Leu Tyr Phe Ser
Asp Ala 2385 2390 2395acc ttg gac
aag atc gag cgc tgc gag tac gac ggc tcc cac cgc 7677Thr Leu Asp
Lys Ile Glu Arg Cys Glu Tyr Asp Gly Ser His Arg 2400
2405 2410tat gtg atc cta aag tcg gag ccc gtc cac
ccc ttt ggg ttg gcg 7722Tyr Val Ile Leu Lys Ser Glu Pro Val His
Pro Phe Gly Leu Ala 2415 2420
2425gtg tac gga gag cac att ttc tgg act gac tgg gtg cgg cgg gct
7767Val Tyr Gly Glu His Ile Phe Trp Thr Asp Trp Val Arg Arg Ala
2430 2435 2440gtg cag cga gcc aac aag
tat gtg ggc agc gac atg aag ctg ctt 7812Val Gln Arg Ala Asn Lys
Tyr Val Gly Ser Asp Met Lys Leu Leu 2445 2450
2455cgg gtg gac att ccc cag caa ccc atg ggc atc atc gcc
gtg gcc 7857Arg Val Asp Ile Pro Gln Gln Pro Met Gly Ile Ile Ala
Val Ala 2460 2465 2470aat gac acc
aac agc tgt gaa ctc tcc ccc tgc cgt atc aac aat 7902Asn Asp Thr
Asn Ser Cys Glu Leu Ser Pro Cys Arg Ile Asn Asn 2475
2480 2485gga ggc tgc cag gat ctg tgt ctg ctc acc
cac caa ggc cac gtc 7947Gly Gly Cys Gln Asp Leu Cys Leu Leu Thr
His Gln Gly His Val 2490 2495
2500aac tgt tcc tgt cga ggg ggc cgg atc ctc cag gag gac ttc acc
7992Asn Cys Ser Cys Arg Gly Gly Arg Ile Leu Gln Glu Asp Phe Thr
2505 2510 2515tgc cgg gct gtg aac tcc
tct tgt cgg gca caa gat gag ttt gag 8037Cys Arg Ala Val Asn Ser
Ser Cys Arg Ala Gln Asp Glu Phe Glu 2520 2525
2530tgt gcc aat ggg gaa tgt atc agc ttc agc ctc acc tgt
gat ggc 8082Cys Ala Asn Gly Glu Cys Ile Ser Phe Ser Leu Thr Cys
Asp Gly 2535 2540 2545gtc tcc cac
tgc aag gac aag tcc gat gag aag ccc tcc tac tgc 8127Val Ser His
Cys Lys Asp Lys Ser Asp Glu Lys Pro Ser Tyr Cys 2550
2555 2560aac tca cgc cgc tgc aag aag act ttc cgc
cag tgt aac aat ggc 8172Asn Ser Arg Arg Cys Lys Lys Thr Phe Arg
Gln Cys Asn Asn Gly 2565 2570
2575cgc tgt gta tcc aac atg ctg tgg tgc aat ggg gtg gat tac tgt
8217Arg Cys Val Ser Asn Met Leu Trp Cys Asn Gly Val Asp Tyr Cys
2580 2585 2590ggg gat ggc tct gat gag
ata cct tgc aac aag act gcc tgt ggt 8262Gly Asp Gly Ser Asp Glu
Ile Pro Cys Asn Lys Thr Ala Cys Gly 2595 2600
2605gtg ggt gag ttc cgc tgc cgg gat ggg tcc tgc atc ggg
aac tcc 8307Val Gly Glu Phe Arg Cys Arg Asp Gly Ser Cys Ile Gly
Asn Ser 2610 2615 2620agt cgc tgc
aac cag ttt gtg gat tgt gag gat gcc tcg gat gag 8352Ser Arg Cys
Asn Gln Phe Val Asp Cys Glu Asp Ala Ser Asp Glu 2625
2630 2635atg aat tgc agt gcc aca gac tgc agc agc
tat ttc cgc ctg ggc 8397Met Asn Cys Ser Ala Thr Asp Cys Ser Ser
Tyr Phe Arg Leu Gly 2640 2645
2650gtg aaa ggt gtc ctc ttc cag ccg tgc gag cgg aca tcc ctg tgc
8442Val Lys Gly Val Leu Phe Gln Pro Cys Glu Arg Thr Ser Leu Cys
2655 2660 2665tac gca cct agc tgg gtg
tgt gat ggc gcc aac gac tgt gga gac 8487Tyr Ala Pro Ser Trp Val
Cys Asp Gly Ala Asn Asp Cys Gly Asp 2670 2675
2680tac agc gat gaa cgt gac tgt cca ggt gtg aag cgc cct
agg tgc 8532Tyr Ser Asp Glu Arg Asp Cys Pro Gly Val Lys Arg Pro
Arg Cys 2685 2690 2695ccg ctc aat
tac ttt gcc tgc ccc agc ggg cgc tgt atc ccc atg 8577Pro Leu Asn
Tyr Phe Ala Cys Pro Ser Gly Arg Cys Ile Pro Met 2700
2705 2710agc tgg acg tgt gac aag gag gat gac tgt
gag aac ggc gag gat 8622Ser Trp Thr Cys Asp Lys Glu Asp Asp Cys
Glu Asn Gly Glu Asp 2715 2720
2725gag acc cac tgc aac aag ttc tgc tca gag gca cag ttc gag tgc
8667Glu Thr His Cys Asn Lys Phe Cys Ser Glu Ala Gln Phe Glu Cys
2730 2735 2740cag aac cac cgg tgt atc
tcc aag cag tgg ctg tgt gac ggt agc 8712Gln Asn His Arg Cys Ile
Ser Lys Gln Trp Leu Cys Asp Gly Ser 2745 2750
2755gat gat tgc ggg gat ggc tcc gat gag gca gct cac tgt
gaa ggc 8757Asp Asp Cys Gly Asp Gly Ser Asp Glu Ala Ala His Cys
Glu Gly 2760 2765 2770aag aca tgt
ggc ccc tcc tcc ttc tcc tgt ccc ggc acc cac gtg 8802Lys Thr Cys
Gly Pro Ser Ser Phe Ser Cys Pro Gly Thr His Val 2775
2780 2785tgt gtc cct gag cgc tgg ctc tgt gat ggc
gac aag gac tgt acc 8847Cys Val Pro Glu Arg Trp Leu Cys Asp Gly
Asp Lys Asp Cys Thr 2790 2795
2800gat ggc gcg gat gag agt gtc act gct ggc tgc ctg tac aac agc
8892Asp Gly Ala Asp Glu Ser Val Thr Ala Gly Cys Leu Tyr Asn Ser
2805 2810 2815acc tgt gat gac cgt gag
ttc atg tgc cag aac cgc ttg tgt att 8937Thr Cys Asp Asp Arg Glu
Phe Met Cys Gln Asn Arg Leu Cys Ile 2820 2825
2830ccc aag cat ttc gtg tgc gac cat gac cgt gac tgt gct
gat ggc 8982Pro Lys His Phe Val Cys Asp His Asp Arg Asp Cys Ala
Asp Gly 2835 2840 2845tct gat gaa
tcc cct gag tgt gag tac cca acc tgc ggg ccc aat 9027Ser Asp Glu
Ser Pro Glu Cys Glu Tyr Pro Thr Cys Gly Pro Asn 2850
2855 2860gaa ttc cgc tgt gcc aat ggg cgt tgt ctg
agc tcc cgt cag tgg 9072Glu Phe Arg Cys Ala Asn Gly Arg Cys Leu
Ser Ser Arg Gln Trp 2865 2870
2875gaa tgt gat ggg gag aat gac tgt cac gac cac agc gat gag gct
9117Glu Cys Asp Gly Glu Asn Asp Cys His Asp His Ser Asp Glu Ala
2880 2885 2890ccc aag aac cca cac tgc
acc agc cca gag cac aaa tgc aat gcc 9162Pro Lys Asn Pro His Cys
Thr Ser Pro Glu His Lys Cys Asn Ala 2895 2900
2905tca tca cag ttc ctg tgc agc agc ggg cgc tgc gtg gct
gag gcg 9207Ser Ser Gln Phe Leu Cys Ser Ser Gly Arg Cys Val Ala
Glu Ala 2910 2915 2920ttg ctc tgc
aac ggc cag gac gac tgt ggg gac ggt tca gac gaa 9252Leu Leu Cys
Asn Gly Gln Asp Asp Cys Gly Asp Gly Ser Asp Glu 2925
2930 2935cgc ggg tgc cat gtc aac gag tgt ctc agc
cgc aag ctc agt ggc 9297Arg Gly Cys His Val Asn Glu Cys Leu Ser
Arg Lys Leu Ser Gly 2940 2945
2950tgc agt cag gac tgc gag gac ctc aag ata ggc ttt aag tgc cgc
9342Cys Ser Gln Asp Cys Glu Asp Leu Lys Ile Gly Phe Lys Cys Arg
2955 2960 2965tgt cgc ccg ggc ttc cgg
cta aag gac gat ggc agg acc tgt gcc 9387Cys Arg Pro Gly Phe Arg
Leu Lys Asp Asp Gly Arg Thr Cys Ala 2970 2975
2980gac ctg gat gag tgc agc acc acc ttc ccc tgc agc cag
ctc tgc 9432Asp Leu Asp Glu Cys Ser Thr Thr Phe Pro Cys Ser Gln
Leu Cys 2985 2990 2995atc aac acc
cac gga agt tac aag tgt ctg tgt gtg gag ggc tat 9477Ile Asn Thr
His Gly Ser Tyr Lys Cys Leu Cys Val Glu Gly Tyr 3000
3005 3010gca ccc cgt ggc ggt gac ccc cac agc tgc
aaa gct gtg acc gat 9522Ala Pro Arg Gly Gly Asp Pro His Ser Cys
Lys Ala Val Thr Asp 3015 3020
3025gag gag cca ttt ctc atc ttt gcc aac cgg tac tac ctg cgg aag
9567Glu Glu Pro Phe Leu Ile Phe Ala Asn Arg Tyr Tyr Leu Arg Lys
3030 3035 3040ctc aac ctg gac ggc tcc
aac tac aca ctg ctt aag cag ggc ctg 9612Leu Asn Leu Asp Gly Ser
Asn Tyr Thr Leu Leu Lys Gln Gly Leu 3045 3050
3055aac aat gcg gtc gcc ttg gca ttt gac tac cga gag cag
atg atc 9657Asn Asn Ala Val Ala Leu Ala Phe Asp Tyr Arg Glu Gln
Met Ile 3060 3065 3070tac tgg acg
ggc gtg acc acc cag ggc agc atg att cgc agg atg 9702Tyr Trp Thr
Gly Val Thr Thr Gln Gly Ser Met Ile Arg Arg Met 3075
3080 3085cac ctc aac ggc agc aac gtg cag gtt ctg
cac cgg acg ggc ctt 9747His Leu Asn Gly Ser Asn Val Gln Val Leu
His Arg Thr Gly Leu 3090 3095
3100agt aac cca gat ggg ctc gct gtg gac tgg gtg ggt ggc aac ctg
9792Ser Asn Pro Asp Gly Leu Ala Val Asp Trp Val Gly Gly Asn Leu
3105 3110 3115tac tgg tgt gac aag ggc
aga gat acc att gag gtg tcc aag ctt 9837Tyr Trp Cys Asp Lys Gly
Arg Asp Thr Ile Glu Val Ser Lys Leu 3120 3125
3130aac ggg gcc tat cgg aca gtg ctg gtc agc tct ggc ctc
cgg gag 9882Asn Gly Ala Tyr Arg Thr Val Leu Val Ser Ser Gly Leu
Arg Glu 3135 3140 3145ccc aga gct
ctg gta gtg gat gta cag aat ggg tac ctg tac tgg 9927Pro Arg Ala
Leu Val Val Asp Val Gln Asn Gly Tyr Leu Tyr Trp 3150
3155 3160aca gac tgg ggt gac cac tca ctg atc ggc
cgg att ggc atg gat 9972Thr Asp Trp Gly Asp His Ser Leu Ile Gly
Arg Ile Gly Met Asp 3165 3170
3175gga tct ggc cgc agc atc atc gtg gac act aag atc aca tgg ccc
10017Gly Ser Gly Arg Ser Ile Ile Val Asp Thr Lys Ile Thr Trp Pro
3180 3185 3190aat ggc ctg acc gtg gac
tac gtc acg gaa cgc atc tac tgg gct 10062Asn Gly Leu Thr Val Asp
Tyr Val Thr Glu Arg Ile Tyr Trp Ala 3195 3200
3205gac gcc cgt gag gac tac atc gag ttc gcc agc ctg gat
ggc tcc 10107Asp Ala Arg Glu Asp Tyr Ile Glu Phe Ala Ser Leu Asp
Gly Ser 3210 3215 3220aac cgt cac
gtt gtg ctg agc caa gac atc cca cac atc ttt gcg 10152Asn Arg His
Val Val Leu Ser Gln Asp Ile Pro His Ile Phe Ala 3225
3230 3235ctg acc cta ttt gaa gac tac gtc tac tgg
aca gac tgg gaa acg 10197Leu Thr Leu Phe Glu Asp Tyr Val Tyr Trp
Thr Asp Trp Glu Thr 3240 3245
3250aag tcc atc aac cgg gcc cac aag acc acg ggt gcc aac aaa aca
10242Lys Ser Ile Asn Arg Ala His Lys Thr Thr Gly Ala Asn Lys Thr
3255 3260 3265ctc ctc atc agc acc ctg
cac cgg ccc atg gac tta cat gta ttc 10287Leu Leu Ile Ser Thr Leu
His Arg Pro Met Asp Leu His Val Phe 3270 3275
3280cac gcc ctg cgc cag cca gat gtg ccc aat cac ccc tgc
aaa gtc 10332His Ala Leu Arg Gln Pro Asp Val Pro Asn His Pro Cys
Lys Val 3285 3290 3295aac aat ggt
ggc tgc agc aac ctg tgc ctg ctg tcc cct ggg ggt 10377Asn Asn Gly
Gly Cys Ser Asn Leu Cys Leu Leu Ser Pro Gly Gly 3300
3305 3310ggt cac aag tgc gcc tgc ccc acc aac ttc
tat ctg ggt ggc gat 10422Gly His Lys Cys Ala Cys Pro Thr Asn Phe
Tyr Leu Gly Gly Asp 3315 3320
3325ggc cgt acc tgt gtg tcc aac tgc aca gca agc cag ttt gtg tgc
10467Gly Arg Thr Cys Val Ser Asn Cys Thr Ala Ser Gln Phe Val Cys
3330 3335 3340aaa aat gac aag tgc atc
ccc ttc tgg tgg aag tgt gac acg gag 10512Lys Asn Asp Lys Cys Ile
Pro Phe Trp Trp Lys Cys Asp Thr Glu 3345 3350
3355gac gac tgt ggg gat cac tca gac gag cct cca gac tgt
ccc gag 10557Asp Asp Cys Gly Asp His Ser Asp Glu Pro Pro Asp Cys
Pro Glu 3360 3365 3370ttc aag tgc
cgc cca ggc cag ttc cag tgc tcc acc ggc atc tgc 10602Phe Lys Cys
Arg Pro Gly Gln Phe Gln Cys Ser Thr Gly Ile Cys 3375
3380 3385acc aac cct gcc ttc atc tgt gat ggg gac
aat gac tgc caa gac 10647Thr Asn Pro Ala Phe Ile Cys Asp Gly Asp
Asn Asp Cys Gln Asp 3390 3395
3400aat agt gac gag gcc aat tgc gac att cac gtc tgc ttg ccc agc
10692Asn Ser Asp Glu Ala Asn Cys Asp Ile His Val Cys Leu Pro Ser
3405 3410 3415caa ttc aag tgc acc aac
acc aac cgc tgc att cct ggc atc ttc 10737Gln Phe Lys Cys Thr Asn
Thr Asn Arg Cys Ile Pro Gly Ile Phe 3420 3425
3430cgt tgc aat ggg cag gac aac tgc ggg gac ggc gag gat
gag cgg 10782Arg Cys Asn Gly Gln Asp Asn Cys Gly Asp Gly Glu Asp
Glu Arg 3435 3440 3445gat tgc cct
gag gtg acc tgc gcc ccc aac cag ttc cag tgc tcc 10827Asp Cys Pro
Glu Val Thr Cys Ala Pro Asn Gln Phe Gln Cys Ser 3450
3455 3460atc acc aag cgc tgc atc cct cgc gtc tgg
gtc tgt gac agg gat 10872Ile Thr Lys Arg Cys Ile Pro Arg Val Trp
Val Cys Asp Arg Asp 3465 3470
3475aat cac tgt gtg gac ggc agt gat gag cct gcc aac tgt acc caa
10917Asn His Cys Val Asp Gly Ser Asp Glu Pro Ala Asn Cys Thr Gln
3480 3485 3490atg acc tgt gga gtg gat
gag ttc cgc tgc aag gat tct ggc cgc 10962Met Thr Cys Gly Val Asp
Glu Phe Arg Cys Lys Asp Ser Gly Arg 3495 3500
3505tgc atc ccc gcg cgc tgg aag tgt gac gga gaa gat gac
tgt ggg 11007Cys Ile Pro Ala Arg Trp Lys Cys Asp Gly Glu Asp Asp
Cys Gly 3510 3515 3520gat ggt tca
gat gag ccc aag gaa gag tgt gat gag cgc acc tgt 11052Asp Gly Ser
Asp Glu Pro Lys Glu Glu Cys Asp Glu Arg Thr Cys 3525
3530 3535gag cca tac cag ttc cgc tgc aaa aac aac
cgc tgt gtc cca ggc 11097Glu Pro Tyr Gln Phe Arg Cys Lys Asn Asn
Arg Cys Val Pro Gly 3540 3545
3550cgt tgg caa tgt gac tac gac aac gac tgc gga gat aac tcg gac
11142Arg Trp Gln Cys Asp Tyr Asp Asn Asp Cys Gly Asp Asn Ser Asp
3555 3560 3565gag gag agc tgc aca cct
cgg ccc tgc tct gag agt gag ttt ttc 11187Glu Glu Ser Cys Thr Pro
Arg Pro Cys Ser Glu Ser Glu Phe Phe 3570 3575
3580tgt gcc aat ggc cgc tgc atc gct ggg cgc tgg aag tgt
gat ggg 11232Cys Ala Asn Gly Arg Cys Ile Ala Gly Arg Trp Lys Cys
Asp Gly 3585 3590 3595gac cat gac
tgt gcc gac ggc tca gac gag aaa gac tgc acc ccc 11277Asp His Asp
Cys Ala Asp Gly Ser Asp Glu Lys Asp Cys Thr Pro 3600
3605 3610cgc tgt gat atg gac cag ttc cag tgc aag
agt ggc cac tgc atc 11322Arg Cys Asp Met Asp Gln Phe Gln Cys Lys
Ser Gly His Cys Ile 3615 3620
3625ccc ctg cgc tgg ccg tgt gac gcg gat gct gac tgt atg gac ggc
11367Pro Leu Arg Trp Pro Cys Asp Ala Asp Ala Asp Cys Met Asp Gly
3630 3635 3640agt gac gag gaa gcc tgt
ggc act ggg gtg agg acc tgc cca ttg 11412Ser Asp Glu Glu Ala Cys
Gly Thr Gly Val Arg Thr Cys Pro Leu 3645 3650
3655gat gag ttt caa tgt aac aac acc ttg tgc aag ccg ctg
gcc tgg 11457Asp Glu Phe Gln Cys Asn Asn Thr Leu Cys Lys Pro Leu
Ala Trp 3660 3665 3670aag tgt gat
gga gag gac gac tgt ggg gac aac tca gat gag aac 11502Lys Cys Asp
Gly Glu Asp Asp Cys Gly Asp Asn Ser Asp Glu Asn 3675
3680 3685ccc gag gaa tgc gcc cgg ttc atc tgc cct
ccc aac cgg cct ttc 11547Pro Glu Glu Cys Ala Arg Phe Ile Cys Pro
Pro Asn Arg Pro Phe 3690 3695
3700cgc tgc aag aat gac cga gtc tgc ctg tgg att ggg cgc cag tgt
11592Arg Cys Lys Asn Asp Arg Val Cys Leu Trp Ile Gly Arg Gln Cys
3705 3710 3715gat ggc gtg gac aac tgt
gga gat ggg act gac gag gag gac tgt 11637Asp Gly Val Asp Asn Cys
Gly Asp Gly Thr Asp Glu Glu Asp Cys 3720 3725
3730gag ccc ccc acg gcc cag aac ccc cac tgc aaa gac aag
aag gag 11682Glu Pro Pro Thr Ala Gln Asn Pro His Cys Lys Asp Lys
Lys Glu 3735 3740 3745ttc ctg tgc
cga aac cag cgc tgt cta tca tcc tcc ctg cgc tgt 11727Phe Leu Cys
Arg Asn Gln Arg Cys Leu Ser Ser Ser Leu Arg Cys 3750
3755 3760aac atg ttc gat gac tgc ggc gat ggc tcc
gat gaa gaa gat tgc 11772Asn Met Phe Asp Asp Cys Gly Asp Gly Ser
Asp Glu Glu Asp Cys 3765 3770
3775agc atc gac ccc aag ctg acc agc tgt gcc acc aat gcc agc atg
11817Ser Ile Asp Pro Lys Leu Thr Ser Cys Ala Thr Asn Ala Ser Met
3780 3785 3790tgt ggg gac gaa gct cgt
tgt gtg cgc act gag aaa gct gcc tac 11862Cys Gly Asp Glu Ala Arg
Cys Val Arg Thr Glu Lys Ala Ala Tyr 3795 3800
3805tgt gcc tgc cgc tcg ggc ttc cat act gtg ccg ggc cag
ccc gga 11907Cys Ala Cys Arg Ser Gly Phe His Thr Val Pro Gly Gln
Pro Gly 3810 3815 3820tgc cag gac
atc aac gag tgc ctg cgc ttt ggt acc tgc tct cag 11952Cys Gln Asp
Ile Asn Glu Cys Leu Arg Phe Gly Thr Cys Ser Gln 3825
3830 3835ctc tgg aac aaa ccc aag gga ggc cac ctc
tgc agc tgt gcc cgc 11997Leu Trp Asn Lys Pro Lys Gly Gly His Leu
Cys Ser Cys Ala Arg 3840 3845
3850aac ttc atg aag aca cac aac acc tgc aaa gct gaa ggc tcc gag
12042Asn Phe Met Lys Thr His Asn Thr Cys Lys Ala Glu Gly Ser Glu
3855 3860 3865tac cag gtg cta tac atc
gcg gat gac aac gag atc cgc agc ttg 12087Tyr Gln Val Leu Tyr Ile
Ala Asp Asp Asn Glu Ile Arg Ser Leu 3870 3875
3880ttc ccg ggc cac ccc cac tca gcc tac gag cag aca ttc
cag ggc 12132Phe Pro Gly His Pro His Ser Ala Tyr Glu Gln Thr Phe
Gln Gly 3885 3890 3895gat gag agt
gtc cgc ata gat gcc atg gat gtc cat gtc aag gcc 12177Asp Glu Ser
Val Arg Ile Asp Ala Met Asp Val His Val Lys Ala 3900
3905 3910ggc cgt gtc tac tgg act aac tgg cac acg
ggc aca atc tcc tac 12222Gly Arg Val Tyr Trp Thr Asn Trp His Thr
Gly Thr Ile Ser Tyr 3915 3920
3925agg agc ctg ccc cct gcc gcc cct cct acc act tcc aac cgc cac
12267Arg Ser Leu Pro Pro Ala Ala Pro Pro Thr Thr Ser Asn Arg His
3930 3935 3940cgg agg cag atc gac cgg
ggt gtc acc cac ctc aat att tca ggg 12312Arg Arg Gln Ile Asp Arg
Gly Val Thr His Leu Asn Ile Ser Gly 3945 3950
3955ctg aag atg ccg agg ggt atc gct atc gac tgg gtg gcc
ggg aat 12357Leu Lys Met Pro Arg Gly Ile Ala Ile Asp Trp Val Ala
Gly Asn 3960 3965 3970gtg tac tgg
acc gat tcc ggc cga gac gtg att gag gtg gcg caa 12402Val Tyr Trp
Thr Asp Ser Gly Arg Asp Val Ile Glu Val Ala Gln 3975
3980 3985atg aag ggc gag aac cgc aag acg ctc atc
tcg ggc atg att gat 12447Met Lys Gly Glu Asn Arg Lys Thr Leu Ile
Ser Gly Met Ile Asp 3990 3995
4000gag ccc cat gcc atc gtg gtg gac cct ctg agg ggc acc atg tac
12492Glu Pro His Ala Ile Val Val Asp Pro Leu Arg Gly Thr Met Tyr
4005 4010 4015tgg tca gac tgg ggg aac
cac ccc aag att gaa aca gca gcg atg 12537Trp Ser Asp Trp Gly Asn
His Pro Lys Ile Glu Thr Ala Ala Met 4020 4025
4030gat ggc acc ctt cgg gag act ctc gtg caa gac aac att
cag tgg 12582Asp Gly Thr Leu Arg Glu Thr Leu Val Gln Asp Asn Ile
Gln Trp 4035 4040 4045cct aca ggg
ctg gct gtg gac tat cac aat gaa cgg ctc tac tgg 12627Pro Thr Gly
Leu Ala Val Asp Tyr His Asn Glu Arg Leu Tyr Trp 4050
4055 4060gca gat gcc aag ctt tcg gtc atc ggc agc
atc cgg ctc aac ggc 12672Ala Asp Ala Lys Leu Ser Val Ile Gly Ser
Ile Arg Leu Asn Gly 4065 4070
4075act gac ccc att gtg gct gct gac agc aaa cga ggc cta agt cac
12717Thr Asp Pro Ile Val Ala Ala Asp Ser Lys Arg Gly Leu Ser His
4080 4085 4090ccc ttc agc atc gat gtg
ttt gaa gac tac atc tac gga gtc act 12762Pro Phe Ser Ile Asp Val
Phe Glu Asp Tyr Ile Tyr Gly Val Thr 4095 4100
4105tac atc aat aat cgt gtc ttc aag atc cac aag ttt gga
cac agc 12807Tyr Ile Asn Asn Arg Val Phe Lys Ile His Lys Phe Gly
His Ser 4110 4115 4120ccc ttg tac
aac cta act ggg ggc ctg agc cat gcc tct gat gta 12852Pro Leu Tyr
Asn Leu Thr Gly Gly Leu Ser His Ala Ser Asp Val 4125
4130 4135gtc ctt tac cat caa cac aag cag cct gaa
gtg acc aac ccc tgt 12897Val Leu Tyr His Gln His Lys Gln Pro Glu
Val Thr Asn Pro Cys 4140 4145
4150gac cgc aag aaa tgc gaa tgg ctg tgt ctg ctg agc ccc agc ggg
12942Asp Arg Lys Lys Cys Glu Trp Leu Cys Leu Leu Ser Pro Ser Gly
4155 4160 4165cct gtc tgc acc tgt ccc
aat gga aag agg ctg gat aat ggc acc 12987Pro Val Cys Thr Cys Pro
Asn Gly Lys Arg Leu Asp Asn Gly Thr 4170 4175
4180tgt gtg cct gtg ccc tct cca aca ccc cct cca gat gcc
cct agg 13032Cys Val Pro Val Pro Ser Pro Thr Pro Pro Pro Asp Ala
Pro Arg 4185 4190 4195cct gga acc
tgc act ctg cag tgc ttc aat ggt ggt agt tgt ttc 13077Pro Gly Thr
Cys Thr Leu Gln Cys Phe Asn Gly Gly Ser Cys Phe 4200
4205 4210ctc aac gct cgg agg cag ccc aag tgc cgt
tgc cag ccc cgt tac 13122Leu Asn Ala Arg Arg Gln Pro Lys Cys Arg
Cys Gln Pro Arg Tyr 4215 4220
4225aca ggc gat aag tgt gag ctg gat cag tgc tgg gaa tac tgt cac
13167Thr Gly Asp Lys Cys Glu Leu Asp Gln Cys Trp Glu Tyr Cys His
4230 4235 4240aac gga ggc acc tgt gcg
gct tcc cca tct ggc atg ccc acg tgc 13212Asn Gly Gly Thr Cys Ala
Ala Ser Pro Ser Gly Met Pro Thr Cys 4245 4250
4255cgc tgt ccc act ggc ttc acg ggc ccc aaa tgc acc gca
cag gtg 13257Arg Cys Pro Thr Gly Phe Thr Gly Pro Lys Cys Thr Ala
Gln Val 4260 4265 4270tgt gca ggc
tac tgc tct aac aac agc acc tgc acc gtc aac cag 13302Cys Ala Gly
Tyr Cys Ser Asn Asn Ser Thr Cys Thr Val Asn Gln 4275
4280 4285ggc aac cag ccc cag tgc cga tgt cta cct
ggc ttc ctg ggc gac 13347Gly Asn Gln Pro Gln Cys Arg Cys Leu Pro
Gly Phe Leu Gly Asp 4290 4295
4300cgt tgc cag tac cgg cag tgc tct ggc ttc tgt gag aac ttt ggc
13392Arg Cys Gln Tyr Arg Gln Cys Ser Gly Phe Cys Glu Asn Phe Gly
4305 4310 4315acc tgt cag atg gct gct
gat ggc tcc cga caa tgt cgc tgc acc 13437Thr Cys Gln Met Ala Ala
Asp Gly Ser Arg Gln Cys Arg Cys Thr 4320 4325
4330gtc tac ttt gag gga cca agg tgt gag gtg aac aag tgt
agt cgc 13482Val Tyr Phe Glu Gly Pro Arg Cys Glu Val Asn Lys Cys
Ser Arg 4335 4340 4345tgt ctc caa
ggc gcc tgt gtg gtc aat aag cag acc gga gat gtc 13527Cys Leu Gln
Gly Ala Cys Val Val Asn Lys Gln Thr Gly Asp Val 4350
4355 4360aca tgc aac tgc act gat ggc cgg gta gcc
ccc agt tgt ctc acc 13572Thr Cys Asn Cys Thr Asp Gly Arg Val Ala
Pro Ser Cys Leu Thr 4365 4370
4375tgc atc gat cac tgt agc aat ggt ggc tcc tgc acc atg aac agc
13617Cys Ile Asp His Cys Ser Asn Gly Gly Ser Cys Thr Met Asn Ser
4380 4385 4390aag atg atg cct gag tgc
cag tgc ccg ccc cat atg aca gga ccc 13662Lys Met Met Pro Glu Cys
Gln Cys Pro Pro His Met Thr Gly Pro 4395 4400
4405cgg tgc cag gag cag gtt gtt agt cag caa cag cct ggg
cat atg 13707Arg Cys Gln Glu Gln Val Val Ser Gln Gln Gln Pro Gly
His Met 4410 4415 4420gcc tcc atc
ctg atc cct ctg ctg ctg ctt ctc ctg ctg ctt ctg 13752Ala Ser Ile
Leu Ile Pro Leu Leu Leu Leu Leu Leu Leu Leu Leu 4425
4430 4435gtg gct ggc gtg gtg ttc tgg tat aag cgg
cga gtc cga ggg gct 13797Val Ala Gly Val Val Phe Trp Tyr Lys Arg
Arg Val Arg Gly Ala 4440 4445
4450aag ggc ttc cag cac cag cgg atg acc aat ggg gcc atg aat gtg
13842Lys Gly Phe Gln His Gln Arg Met Thr Asn Gly Ala Met Asn Val
4455 4460 4465gaa att gga aac cct acc
tac aag atg tat gaa ggt gga gag ccc 13887Glu Ile Gly Asn Pro Thr
Tyr Lys Met Tyr Glu Gly Gly Glu Pro 4470 4475
4480gat gat gtc ggg ggc cta ctg gat gct gat ttt gcc ctt
gac cct 13932Asp Asp Val Gly Gly Leu Leu Asp Ala Asp Phe Ala Leu
Asp Pro 4485 4490 4495gac aag cct
acc aac ttc acc aac cca gtg tat gcc acg ctc tac 13977Asp Lys Pro
Thr Asn Phe Thr Asn Pro Val Tyr Ala Thr Leu Tyr 4500
4505 4510atg ggg ggc cac ggc agc cgc cat tcc ctg
gcc agc acg gac gag 14022Met Gly Gly His Gly Ser Arg His Ser Leu
Ala Ser Thr Asp Glu 4515 4520
4525aag cga gaa ctg ctg ggc cgg gga cct gaa gac gag ata gga gat
14067Lys Arg Glu Leu Leu Gly Arg Gly Pro Glu Asp Glu Ile Gly Asp
4530 4535 4540ccc ttg gca tag
ggccctgccc cgacggatgt ccccagaaag ccccctgcca 14119Pro Leu Ala
4545catgagtctt tcaatgaacc ccctccccag ccggcccttc tccggccctg ccgggtgtac
14179aaatgtaaaa atgaaggaat tactttttat atgtgagcga gcaagcgagc aagcacagta
14239ttatctcttt gcatttcctt cctgcctgct cctcagtatc ccccccatgc tgccttgagg
14299gggcggggag ggctttgtgg ctcaaaggta tgaaggagtc cacatgttcc ctaccgagca
14359tacccctgga agcctggcgg cacggcctcc ccaccacgcc tgtgcaagac actcaacggg
14419gctccgtgtc ccagctttcc tttccttggc tctctggggt tagttcaggg gaggtggagt
14479cctctgctga ccctgtctgg aagatttggc tctagctgag gaaggagtct tttagttgag
14539ggaagtcacc ccaaacccca gctcccactt tcaggggcac ctctcagatg gccatgctca
14599gtatcccttc cagacaggcc ctcccctctc tagcgccccc tctgtggctc ctagggctga
14659acacattctt tggtaactgt cccccaagcc tcccatcccc ctgagggcca ggaagagtcg
14719gggcacacca aggaagggca agcgggcagc cccattttgg ggacgtgaac gttttaataa
14779tttttgctga attcctttac aactaaataa cacagatatt gttataaata aaattgtaaa
14839aaaaaaaaaa
1484924545PRTMus sp. 2Met Leu Thr Pro Pro Leu Leu Leu Leu Val Pro Leu Leu
Ser Ala Leu1 5 10 15Val
Ser Gly Ala Thr Met Asp Ala Pro Lys Thr Cys Ser Pro Lys Gln 20
25 30Phe Ala Cys Arg Asp Gln Ile Thr
Cys Ile Ser Lys Gly Trp Arg Cys 35 40
45Asp Gly Glu Arg Asp Cys Pro Asp Gly Ser Asp Glu Ala Pro Glu Ile
50 55 60Cys Pro Gln Ser Lys Ala Gln Arg
Cys Pro Pro Asn Glu His Ser Cys65 70 75
80Leu Gly Thr Glu Leu Cys Val Pro Met Ser Arg Leu Cys
Asn Gly Ile 85 90 95Gln
Asp Cys Met Asp Gly Ser Asp Glu Gly Ala His Cys Arg Glu Leu
100 105 110Arg Ala Asn Cys Ser Arg Met
Gly Cys Gln His His Cys Val Pro Thr 115 120
125Pro Ser Gly Pro Thr Cys Tyr Cys Asn Ser Ser Phe Gln Leu Glu
Ala 130 135 140Asp Gly Lys Thr Cys Lys
Asp Phe Asp Glu Cys Ser Val Tyr Gly Thr145 150
155 160Cys Ser Gln Leu Cys Thr Asn Thr Asp Gly Ser
Phe Thr Cys Gly Cys 165 170
175Val Glu Gly Tyr Leu Leu Gln Pro Asp Asn Arg Ser Cys Lys Ala Lys
180 185 190Asn Glu Pro Val Asp Arg
Pro Pro Val Leu Leu Ile Ala Asn Ser Gln 195 200
205Asn Ile Leu Ala Thr Tyr Leu Ser Gly Ala Gln Val Ser Thr
Ile Thr 210 215 220Pro Thr Ser Thr Arg
Gln Thr Thr Ala Met Asp Phe Ser Tyr Ala Asn225 230
235 240Glu Thr Val Cys Trp Val His Val Gly Asp
Ser Ala Ala Gln Thr Gln 245 250
255Leu Lys Cys Ala Arg Met Pro Gly Leu Lys Gly Phe Val Asp Glu His
260 265 270Thr Ile Asn Ile Ser
Leu Ser Leu His His Val Glu Gln Met Ala Ile 275
280 285Asp Trp Leu Thr Gly Asn Phe Tyr Phe Val Asp Asp
Ile Asp Asp Arg 290 295 300Ile Phe Val
Cys Asn Arg Asn Gly Asp Thr Cys Val Thr Leu Leu Asp305
310 315 320Leu Glu Leu Tyr Asn Pro Lys
Gly Ile Ala Leu Asp Pro Ala Met Gly 325
330 335Lys Val Phe Phe Thr Asp Tyr Gly Gln Ile Pro Lys
Val Glu Arg Cys 340 345 350Asp
Met Asp Gly Gln Asn Arg Thr Lys Leu Val Asp Ser Lys Ile Val 355
360 365Phe Pro His Gly Ile Thr Leu Asp Leu
Val Ser Arg Leu Val Tyr Trp 370 375
380Ala Asp Ala Tyr Leu Asp Tyr Ile Glu Val Val Asp Tyr Glu Gly Lys385
390 395 400Gly Arg Gln Thr
Ile Ile Gln Gly Ile Leu Ile Glu His Leu Tyr Gly 405
410 415Leu Thr Val Phe Glu Asn Tyr Leu Tyr Ala
Thr Asn Ser Asp Asn Ala 420 425
430Asn Thr Gln Gln Lys Thr Ser Val Ile Arg Val Asn Arg Phe Asn Ser
435 440 445Thr Glu Tyr Gln Val Val Thr
Arg Val Asp Lys Gly Gly Ala Leu His 450 455
460Ile Tyr His Gln Arg Arg Gln Pro Arg Val Arg Ser His Ala Cys
Glu465 470 475 480Asn Asp
Gln Tyr Gly Lys Pro Gly Gly Cys Ser Asp Ile Cys Leu Leu
485 490 495Ala Asn Ser His Lys Ala Arg
Thr Cys Arg Cys Arg Ser Gly Phe Ser 500 505
510Leu Gly Ser Asp Gly Lys Ser Cys Lys Lys Pro Glu His Glu
Leu Phe 515 520 525Leu Val Tyr Gly
Lys Gly Arg Pro Gly Ile Ile Arg Gly Met Asp Met 530
535 540Gly Ala Lys Val Pro Asp Glu His Met Ile Pro Ile
Glu Asn Leu Met545 550 555
560Asn Pro Arg Ala Leu Asp Phe His Ala Glu Thr Gly Phe Ile Tyr Phe
565 570 575Ala Asp Thr Thr Ser
Tyr Leu Ile Gly Arg Gln Lys Ile Asp Gly Thr 580
585 590Glu Arg Glu Thr Ile Leu Lys Asp Gly Ile His Asn
Val Glu Gly Val 595 600 605Ala Val
Asp Trp Met Gly Asp Asn Leu Tyr Trp Thr Asp Asp Gly Pro 610
615 620Lys Lys Thr Ile Ser Val Ala Arg Leu Glu Lys
Ala Ala Gln Thr Arg625 630 635
640Lys Thr Leu Ile Glu Gly Lys Met Thr His Pro Arg Ala Ile Val Val
645 650 655Asp Pro Leu Asn
Gly Trp Met Tyr Trp Thr Asp Trp Glu Glu Asp Pro 660
665 670Lys Asp Ser Arg Arg Gly Arg Leu Glu Arg Ala
Trp Met Asp Gly Ser 675 680 685His
Arg Asp Ile Phe Val Thr Ser Lys Thr Val Leu Trp Pro Asn Gly 690
695 700Leu Ser Leu Asp Ile Pro Ala Gly Arg Leu
Tyr Trp Val Asp Ala Phe705 710 715
720Tyr Asp Arg Ile Glu Thr Ile Leu Leu Asn Gly Thr Asp Arg Lys
Ile 725 730 735Val Tyr Glu
Gly Pro Glu Leu Asn His Ala Phe Gly Leu Cys His His 740
745 750Gly Asn Tyr Leu Phe Trp Thr Glu Tyr Arg
Ser Gly Ser Val Tyr Arg 755 760
765Leu Glu Arg Gly Val Ala Gly Ala Pro Pro Thr Val Thr Leu Leu Arg 770
775 780Ser Glu Arg Pro Pro Ile Phe Glu
Ile Arg Met Tyr Asp Ala His Glu785 790
795 800Gln Gln Val Gly Thr Asn Lys Cys Arg Val Asn Asn
Gly Gly Cys Ser 805 810
815Ser Leu Cys Leu Ala Thr Pro Gly Ser Arg Gln Cys Ala Cys Ala Glu
820 825 830Asp Gln Val Leu Asp Thr
Asp Gly Val Thr Cys Leu Ala Asn Pro Ser 835 840
845Tyr Val Pro Pro Pro Gln Cys Gln Pro Gly Gln Phe Ala Cys
Ala Asn 850 855 860Asn Arg Cys Ile Gln
Glu Arg Trp Lys Cys Asp Gly Asp Asn Asp Cys865 870
875 880Leu Asp Asn Ser Asp Glu Ala Pro Ala Leu
Cys His Gln His Thr Cys 885 890
895Pro Ser Asp Arg Phe Lys Cys Glu Asn Asn Arg Cys Ile Pro Asn Arg
900 905 910Trp Leu Cys Asp Gly
Asp Asn Asp Cys Gly Asn Ser Glu Asp Glu Ser 915
920 925Asn Ala Thr Cys Ser Ala Arg Thr Cys Pro Pro Asn
Gln Phe Ser Cys 930 935 940Ala Ser Gly
Arg Cys Ile Pro Ile Ser Trp Thr Cys Asp Leu Asp Asp945
950 955 960Asp Cys Gly Asp Arg Ser Asp
Glu Ser Ala Ser Cys Ala Tyr Pro Thr 965
970 975Cys Phe Pro Leu Thr Gln Phe Thr Cys Asn Asn Gly
Arg Cys Ile Asn 980 985 990Ile
Asn Trp Arg Cys Asp Asn Asp Asn Asp Cys Gly Asp Asn Ser Asp 995
1000 1005Glu Ala Gly Cys Ser His Ser Cys
Ser Ser Thr Gln Phe Lys Cys 1010 1015
1020Asn Ser Gly Arg Cys Ile Pro Glu His Trp Thr Cys Asp Gly Asp
1025 1030 1035Asn Asp Cys Gly Asp Tyr
Ser Asp Glu Thr His Ala Asn Cys Thr 1040 1045
1050Asn Gln Ala Thr Arg Pro Pro Gly Gly Cys His Ser Asp Glu
Phe 1055 1060 1065Gln Cys Pro Leu Asp
Gly Leu Cys Ile Pro Leu Arg Trp Arg Cys 1070 1075
1080Asp Gly Asp Thr Asp Cys Met Asp Ser Ser Asp Glu Lys
Ser Cys 1085 1090 1095Glu Gly Val Thr
His Val Cys Asp Pro Asn Val Lys Phe Gly Cys 1100
1105 1110Lys Asp Ser Ala Arg Cys Ile Ser Lys Ala Trp
Val Cys Asp Gly 1115 1120 1125Asp Ser
Asp Cys Glu Asp Asn Ser Asp Glu Glu Asn Cys Glu Ala 1130
1135 1140Leu Ala Cys Arg Pro Pro Ser His Pro Cys
Ala Asn Asn Thr Ser 1145 1150 1155Val
Cys Leu Pro Pro Asp Lys Leu Cys Asp Gly Lys Asp Asp Cys 1160
1165 1170Gly Asp Gly Ser Asp Glu Gly Glu Leu
Cys Asp Gln Cys Ser Leu 1175 1180
1185Asn Asn Gly Gly Cys Ser His Asn Cys Ser Val Ala Pro Gly Glu
1190 1195 1200Gly Ile Val Cys Ser Cys
Pro Leu Gly Met Glu Leu Gly Ser Asp 1205 1210
1215Asn His Thr Cys Gln Ile Gln Ser Tyr Cys Ala Lys His Leu
Lys 1220 1225 1230Cys Ser Gln Lys Cys
Asp Gln Asn Lys Phe Ser Val Lys Cys Ser 1235 1240
1245Cys Tyr Glu Gly Trp Val Leu Glu Pro Asp Gly Glu Thr
Cys Arg 1250 1255 1260Ser Leu Asp Pro
Phe Lys Leu Phe Ile Ile Phe Ser Asn Arg His 1265
1270 1275Glu Ile Arg Arg Ile Asp Leu His Lys Gly Asp
Tyr Ser Val Leu 1280 1285 1290Val Pro
Gly Leu Arg Asn Thr Ile Ala Leu Asp Phe His Leu Ser 1295
1300 1305Gln Ser Ala Leu Tyr Trp Thr Asp Ala Val
Glu Asp Lys Ile Tyr 1310 1315 1320Arg
Gly Lys Leu Leu Asp Asn Gly Ala Leu Thr Ser Phe Glu Val 1325
1330 1335Val Ile Gln Tyr Gly Leu Ala Thr Pro
Glu Gly Leu Ala Val Asp 1340 1345
1350Trp Ile Ala Gly Asn Ile Tyr Trp Val Glu Ser Asn Leu Asp Gln
1355 1360 1365Ile Glu Val Ala Lys Leu
Asp Gly Thr Leu Arg Thr Thr Leu Leu 1370 1375
1380Ala Gly Asp Ile Glu His Pro Arg Ala Ile Ala Leu Asp Pro
Arg 1385 1390 1395Asp Gly Ile Leu Phe
Trp Thr Asp Trp Asp Ala Ser Leu Pro Arg 1400 1405
1410Ile Glu Ala Ala Ser Met Ser Gly Ala Gly Arg Arg Thr
Ile His 1415 1420 1425Arg Glu Thr Gly
Ser Gly Gly Cys Ala Asn Gly Leu Thr Val Asp 1430
1435 1440Tyr Leu Glu Lys Arg Ile Leu Trp Ile Asp Ala
Arg Ser Asp Ala 1445 1450 1455Ile Tyr
Ser Ala Arg Tyr Asp Gly Ser Gly His Met Glu Val Leu 1460
1465 1470Arg Gly His Glu Phe Leu Ser His Pro Phe
Ala Val Thr Leu Tyr 1475 1480 1485Gly
Gly Glu Val Tyr Trp Thr Asp Trp Arg Thr Asn Thr Leu Ala 1490
1495 1500Lys Ala Asn Lys Trp Thr Gly His Asn
Val Thr Val Val Gln Arg 1505 1510
1515Thr Asn Thr Gln Pro Phe Asp Leu Gln Val Tyr His Pro Ser Arg
1520 1525 1530Gln Pro Met Ala Pro Asn
Pro Cys Glu Ala Asn Gly Gly Arg Gly 1535 1540
1545Pro Cys Ser His Leu Cys Leu Ile Asn Tyr Asn Arg Thr Val
Ser 1550 1555 1560Trp Ala Cys Pro His
Leu Met Lys Leu His Lys Asp Asn Thr Thr 1565 1570
1575Cys Tyr Glu Phe Lys Lys Phe Leu Leu Tyr Ala Arg Gln
Met Glu 1580 1585 1590Ile Arg Gly Val
Asp Leu Asp Ala Pro Tyr Tyr Asn Tyr Ile Ile 1595
1600 1605Ser Phe Thr Val Pro Asp Ile Asp Asn Val Thr
Val Leu Asp Tyr 1610 1615 1620Asp Ala
Arg Glu Gln Arg Val Tyr Trp Ser Asp Val Arg Thr Gln 1625
1630 1635Ala Ile Lys Arg Ala Phe Ile Asn Gly Thr
Gly Val Glu Thr Val 1640 1645 1650Val
Ser Ala Asp Leu Pro Asn Ala His Gly Leu Ala Val Asp Trp 1655
1660 1665Val Ser Arg Asn Leu Phe Trp Thr Ser
Tyr Asp Thr Asn Lys Lys 1670 1675
1680Gln Ile Asn Val Ala Arg Leu Asp Gly Ser Phe Lys Asn Ala Val
1685 1690 1695Val Gln Gly Leu Glu Gln
Pro His Gly Leu Val Val His Pro Leu 1700 1705
1710Arg Gly Lys Leu Tyr Trp Thr Asp Gly Asp Asn Ile Ser Met
Ala 1715 1720 1725Asn Met Asp Gly Ser
Asn His Thr Leu Leu Phe Ser Gly Gln Lys 1730 1735
1740Gly Pro Val Gly Leu Ala Ile Asp Phe Pro Glu Ser Lys
Leu Tyr 1745 1750 1755Trp Ile Ser Ser
Gly Asn His Thr Ile Asn Arg Cys Asn Leu Asp 1760
1765 1770Gly Ser Glu Leu Glu Val Ile Asp Thr Met Arg
Ser Gln Leu Gly 1775 1780 1785Lys Ala
Thr Ala Leu Ala Ile Met Gly Asp Lys Leu Trp Trp Ala 1790
1795 1800Asp Gln Val Ser Glu Lys Met Gly Thr Cys
Asn Lys Ala Asp Gly 1805 1810 1815Ser
Gly Ser Val Val Leu Arg Asn Ser Thr Thr Leu Val Met His 1820
1825 1830Met Lys Val Tyr Asp Glu Ser Ile Gln
Leu Glu His Glu Gly Thr 1835 1840
1845Asn Pro Cys Ser Val Asn Asn Gly Asp Cys Ser Gln Leu Cys Leu
1850 1855 1860Pro Thr Ser Glu Thr Thr
Arg Ser Cys Met Cys Thr Ala Gly Tyr 1865 1870
1875Ser Leu Arg Ser Gly Gln Gln Ala Cys Glu Gly Val Gly Ser
Phe 1880 1885 1890Leu Leu Tyr Ser Val
His Glu Gly Ile Arg Gly Ile Pro Leu Asp 1895 1900
1905Pro Asn Asp Lys Ser Asp Ala Leu Val Pro Val Ser Gly
Thr Ser 1910 1915 1920Leu Ala Val Gly
Ile Asp Phe His Ala Glu Asn Asp Thr Ile Tyr 1925
1930 1935Trp Val Asp Met Gly Leu Ser Thr Ile Ser Arg
Ala Lys Arg Asp 1940 1945 1950Gln Thr
Trp Arg Glu Asp Val Val Thr Asn Gly Ile Gly Arg Val 1955
1960 1965Glu Gly Ile Ala Val Asp Trp Ile Ala Gly
Asn Ile Tyr Trp Thr 1970 1975 1980Asp
Gln Gly Phe Asp Val Ile Glu Val Ala Arg Leu Asn Gly Ser 1985
1990 1995Phe Arg Tyr Val Val Ile Ser Gln Gly
Leu Asp Lys Pro Arg Ala 2000 2005
2010Ile Thr Val His Pro Glu Lys Gly Tyr Leu Phe Trp Thr Glu Trp
2015 2020 2025Gly His Tyr Pro Arg Ile
Glu Arg Ser Arg Leu Asp Gly Thr Glu 2030 2035
2040Arg Val Val Leu Val Asn Val Ser Ile Ser Trp Pro Asn Gly
Ile 2045 2050 2055Ser Val Asp Tyr Gln
Gly Gly Lys Leu Tyr Trp Cys Asp Ala Arg 2060 2065
2070Met Asp Lys Ile Glu Arg Ile Asp Leu Glu Thr Gly Glu
Asn Arg 2075 2080 2085Glu Val Val Leu
Ser Ser Asn Asn Met Asp Met Phe Ser Val Ser 2090
2095 2100Val Phe Glu Asp Phe Ile Tyr Trp Ser Asp Arg
Thr His Ala Asn 2105 2110 2115Gly Ser
Ile Lys Arg Gly Cys Lys Asp Asn Ala Thr Asp Ser Val 2120
2125 2130Pro Leu Arg Thr Gly Ile Gly Val Gln Leu
Lys Asp Ile Lys Val 2135 2140 2145Phe
Asn Arg Asp Arg Gln Lys Gly Thr Asn Val Cys Ala Val Ala 2150
2155 2160Asn Gly Gly Cys Gln Gln Leu Cys Leu
Tyr Arg Gly Gly Gly Gln 2165 2170
2175Arg Ala Cys Ala Cys Ala His Gly Met Leu Ala Glu Asp Gly Ala
2180 2185 2190Ser Cys Arg Glu Tyr Ala
Gly Tyr Leu Leu Tyr Ser Glu Arg Thr 2195 2200
2205Ile Leu Lys Ser Ile His Leu Ser Asp Glu Arg Asn Leu Asn
Ala 2210 2215 2220Pro Val Gln Pro Phe
Glu Asp Pro Glu His Met Lys Asn Val Ile 2225 2230
2235Ala Leu Ala Phe Asp Tyr Arg Ala Gly Thr Ser Pro Gly
Thr Pro 2240 2245 2250Asn Arg Ile Phe
Phe Ser Asp Ile His Phe Gly Asn Ile Gln Gln 2255
2260 2265Ile Asn Asp Asp Gly Ser Gly Arg Thr Thr Ile
Val Glu Asn Val 2270 2275 2280Gly Ser
Val Glu Gly Leu Ala Tyr His Arg Gly Trp Asp Thr Leu 2285
2290 2295Tyr Trp Thr Ser Tyr Thr Thr Ser Thr Ile
Thr Arg His Thr Val 2300 2305 2310Asp
Gln Thr Arg Pro Gly Ala Phe Glu Arg Glu Thr Val Ile Thr 2315
2320 2325Met Ser Gly Asp Asp His Pro Arg Ala
Phe Val Leu Asp Glu Cys 2330 2335
2340Gln Asn Leu Met Phe Trp Thr Asn Trp Asn Glu Leu His Pro Ser
2345 2350 2355Ile Met Arg Ala Ala Leu
Ser Gly Ala Asn Val Leu Thr Leu Ile 2360 2365
2370Glu Lys Asp Ile Arg Thr Pro Asn Gly Leu Ala Ile Asp His
Arg 2375 2380 2385Ala Glu Lys Leu Tyr
Phe Ser Asp Ala Thr Leu Asp Lys Ile Glu 2390 2395
2400Arg Cys Glu Tyr Asp Gly Ser His Arg Tyr Val Ile Leu
Lys Ser 2405 2410 2415Glu Pro Val His
Pro Phe Gly Leu Ala Val Tyr Gly Glu His Ile 2420
2425 2430Phe Trp Thr Asp Trp Val Arg Arg Ala Val Gln
Arg Ala Asn Lys 2435 2440 2445Tyr Val
Gly Ser Asp Met Lys Leu Leu Arg Val Asp Ile Pro Gln 2450
2455 2460Gln Pro Met Gly Ile Ile Ala Val Ala Asn
Asp Thr Asn Ser Cys 2465 2470 2475Glu
Leu Ser Pro Cys Arg Ile Asn Asn Gly Gly Cys Gln Asp Leu 2480
2485 2490Cys Leu Leu Thr His Gln Gly His Val
Asn Cys Ser Cys Arg Gly 2495 2500
2505Gly Arg Ile Leu Gln Glu Asp Phe Thr Cys Arg Ala Val Asn Ser
2510 2515 2520Ser Cys Arg Ala Gln Asp
Glu Phe Glu Cys Ala Asn Gly Glu Cys 2525 2530
2535Ile Ser Phe Ser Leu Thr Cys Asp Gly Val Ser His Cys Lys
Asp 2540 2545 2550Lys Ser Asp Glu Lys
Pro Ser Tyr Cys Asn Ser Arg Arg Cys Lys 2555 2560
2565Lys Thr Phe Arg Gln Cys Asn Asn Gly Arg Cys Val Ser
Asn Met 2570 2575 2580Leu Trp Cys Asn
Gly Val Asp Tyr Cys Gly Asp Gly Ser Asp Glu 2585
2590 2595Ile Pro Cys Asn Lys Thr Ala Cys Gly Val Gly
Glu Phe Arg Cys 2600 2605 2610Arg Asp
Gly Ser Cys Ile Gly Asn Ser Ser Arg Cys Asn Gln Phe 2615
2620 2625Val Asp Cys Glu Asp Ala Ser Asp Glu Met
Asn Cys Ser Ala Thr 2630 2635 2640Asp
Cys Ser Ser Tyr Phe Arg Leu Gly Val Lys Gly Val Leu Phe 2645
2650 2655Gln Pro Cys Glu Arg Thr Ser Leu Cys
Tyr Ala Pro Ser Trp Val 2660 2665
2670Cys Asp Gly Ala Asn Asp Cys Gly Asp Tyr Ser Asp Glu Arg Asp
2675 2680 2685Cys Pro Gly Val Lys Arg
Pro Arg Cys Pro Leu Asn Tyr Phe Ala 2690 2695
2700Cys Pro Ser Gly Arg Cys Ile Pro Met Ser Trp Thr Cys Asp
Lys 2705 2710 2715Glu Asp Asp Cys Glu
Asn Gly Glu Asp Glu Thr His Cys Asn Lys 2720 2725
2730Phe Cys Ser Glu Ala Gln Phe Glu Cys Gln Asn His Arg
Cys Ile 2735 2740 2745Ser Lys Gln Trp
Leu Cys Asp Gly Ser Asp Asp Cys Gly Asp Gly 2750
2755 2760Ser Asp Glu Ala Ala His Cys Glu Gly Lys Thr
Cys Gly Pro Ser 2765 2770 2775Ser Phe
Ser Cys Pro Gly Thr His Val Cys Val Pro Glu Arg Trp 2780
2785 2790Leu Cys Asp Gly Asp Lys Asp Cys Thr Asp
Gly Ala Asp Glu Ser 2795 2800 2805Val
Thr Ala Gly Cys Leu Tyr Asn Ser Thr Cys Asp Asp Arg Glu 2810
2815 2820Phe Met Cys Gln Asn Arg Leu Cys Ile
Pro Lys His Phe Val Cys 2825 2830
2835Asp His Asp Arg Asp Cys Ala Asp Gly Ser Asp Glu Ser Pro Glu
2840 2845 2850Cys Glu Tyr Pro Thr Cys
Gly Pro Asn Glu Phe Arg Cys Ala Asn 2855 2860
2865Gly Arg Cys Leu Ser Ser Arg Gln Trp Glu Cys Asp Gly Glu
Asn 2870 2875 2880Asp Cys His Asp His
Ser Asp Glu Ala Pro Lys Asn Pro His Cys 2885 2890
2895Thr Ser Pro Glu His Lys Cys Asn Ala Ser Ser Gln Phe
Leu Cys 2900 2905 2910Ser Ser Gly Arg
Cys Val Ala Glu Ala Leu Leu Cys Asn Gly Gln 2915
2920 2925Asp Asp Cys Gly Asp Gly Ser Asp Glu Arg Gly
Cys His Val Asn 2930 2935 2940Glu Cys
Leu Ser Arg Lys Leu Ser Gly Cys Ser Gln Asp Cys Glu 2945
2950 2955Asp Leu Lys Ile Gly Phe Lys Cys Arg Cys
Arg Pro Gly Phe Arg 2960 2965 2970Leu
Lys Asp Asp Gly Arg Thr Cys Ala Asp Leu Asp Glu Cys Ser 2975
2980 2985Thr Thr Phe Pro Cys Ser Gln Leu Cys
Ile Asn Thr His Gly Ser 2990 2995
3000Tyr Lys Cys Leu Cys Val Glu Gly Tyr Ala Pro Arg Gly Gly Asp
3005 3010 3015Pro His Ser Cys Lys Ala
Val Thr Asp Glu Glu Pro Phe Leu Ile 3020 3025
3030Phe Ala Asn Arg Tyr Tyr Leu Arg Lys Leu Asn Leu Asp Gly
Ser 3035 3040 3045Asn Tyr Thr Leu Leu
Lys Gln Gly Leu Asn Asn Ala Val Ala Leu 3050 3055
3060Ala Phe Asp Tyr Arg Glu Gln Met Ile Tyr Trp Thr Gly
Val Thr 3065 3070 3075Thr Gln Gly Ser
Met Ile Arg Arg Met His Leu Asn Gly Ser Asn 3080
3085 3090Val Gln Val Leu His Arg Thr Gly Leu Ser Asn
Pro Asp Gly Leu 3095 3100 3105Ala Val
Asp Trp Val Gly Gly Asn Leu Tyr Trp Cys Asp Lys Gly 3110
3115 3120Arg Asp Thr Ile Glu Val Ser Lys Leu Asn
Gly Ala Tyr Arg Thr 3125 3130 3135Val
Leu Val Ser Ser Gly Leu Arg Glu Pro Arg Ala Leu Val Val 3140
3145 3150Asp Val Gln Asn Gly Tyr Leu Tyr Trp
Thr Asp Trp Gly Asp His 3155 3160
3165Ser Leu Ile Gly Arg Ile Gly Met Asp Gly Ser Gly Arg Ser Ile
3170 3175 3180Ile Val Asp Thr Lys Ile
Thr Trp Pro Asn Gly Leu Thr Val Asp 3185 3190
3195Tyr Val Thr Glu Arg Ile Tyr Trp Ala Asp Ala Arg Glu Asp
Tyr 3200 3205 3210Ile Glu Phe Ala Ser
Leu Asp Gly Ser Asn Arg His Val Val Leu 3215 3220
3225Ser Gln Asp Ile Pro His Ile Phe Ala Leu Thr Leu Phe
Glu Asp 3230 3235 3240Tyr Val Tyr Trp
Thr Asp Trp Glu Thr Lys Ser Ile Asn Arg Ala 3245
3250 3255His Lys Thr Thr Gly Ala Asn Lys Thr Leu Leu
Ile Ser Thr Leu 3260 3265 3270His Arg
Pro Met Asp Leu His Val Phe His Ala Leu Arg Gln Pro 3275
3280 3285Asp Val Pro Asn His Pro Cys Lys Val Asn
Asn Gly Gly Cys Ser 3290 3295 3300Asn
Leu Cys Leu Leu Ser Pro Gly Gly Gly His Lys Cys Ala Cys 3305
3310 3315Pro Thr Asn Phe Tyr Leu Gly Gly Asp
Gly Arg Thr Cys Val Ser 3320 3325
3330Asn Cys Thr Ala Ser Gln Phe Val Cys Lys Asn Asp Lys Cys Ile
3335 3340 3345Pro Phe Trp Trp Lys Cys
Asp Thr Glu Asp Asp Cys Gly Asp His 3350 3355
3360Ser Asp Glu Pro Pro Asp Cys Pro Glu Phe Lys Cys Arg Pro
Gly 3365 3370 3375Gln Phe Gln Cys Ser
Thr Gly Ile Cys Thr Asn Pro Ala Phe Ile 3380 3385
3390Cys Asp Gly Asp Asn Asp Cys Gln Asp Asn Ser Asp Glu
Ala Asn 3395 3400 3405Cys Asp Ile His
Val Cys Leu Pro Ser Gln Phe Lys Cys Thr Asn 3410
3415 3420Thr Asn Arg Cys Ile Pro Gly Ile Phe Arg Cys
Asn Gly Gln Asp 3425 3430 3435Asn Cys
Gly Asp Gly Glu Asp Glu Arg Asp Cys Pro Glu Val Thr 3440
3445 3450Cys Ala Pro Asn Gln Phe Gln Cys Ser Ile
Thr Lys Arg Cys Ile 3455 3460 3465Pro
Arg Val Trp Val Cys Asp Arg Asp Asn His Cys Val Asp Gly 3470
3475 3480Ser Asp Glu Pro Ala Asn Cys Thr Gln
Met Thr Cys Gly Val Asp 3485 3490
3495Glu Phe Arg Cys Lys Asp Ser Gly Arg Cys Ile Pro Ala Arg Trp
3500 3505 3510Lys Cys Asp Gly Glu Asp
Asp Cys Gly Asp Gly Ser Asp Glu Pro 3515 3520
3525Lys Glu Glu Cys Asp Glu Arg Thr Cys Glu Pro Tyr Gln Phe
Arg 3530 3535 3540Cys Lys Asn Asn Arg
Cys Val Pro Gly Arg Trp Gln Cys Asp Tyr 3545 3550
3555Asp Asn Asp Cys Gly Asp Asn Ser Asp Glu Glu Ser Cys
Thr Pro 3560 3565 3570Arg Pro Cys Ser
Glu Ser Glu Phe Phe Cys Ala Asn Gly Arg Cys 3575
3580 3585Ile Ala Gly Arg Trp Lys Cys Asp Gly Asp His
Asp Cys Ala Asp 3590 3595 3600Gly Ser
Asp Glu Lys Asp Cys Thr Pro Arg Cys Asp Met Asp Gln 3605
3610 3615Phe Gln Cys Lys Ser Gly His Cys Ile Pro
Leu Arg Trp Pro Cys 3620 3625 3630Asp
Ala Asp Ala Asp Cys Met Asp Gly Ser Asp Glu Glu Ala Cys 3635
3640 3645Gly Thr Gly Val Arg Thr Cys Pro Leu
Asp Glu Phe Gln Cys Asn 3650 3655
3660Asn Thr Leu Cys Lys Pro Leu Ala Trp Lys Cys Asp Gly Glu Asp
3665 3670 3675Asp Cys Gly Asp Asn Ser
Asp Glu Asn Pro Glu Glu Cys Ala Arg 3680 3685
3690Phe Ile Cys Pro Pro Asn Arg Pro Phe Arg Cys Lys Asn Asp
Arg 3695 3700 3705Val Cys Leu Trp Ile
Gly Arg Gln Cys Asp Gly Val Asp Asn Cys 3710 3715
3720Gly Asp Gly Thr Asp Glu Glu Asp Cys Glu Pro Pro Thr
Ala Gln 3725 3730 3735Asn Pro His Cys
Lys Asp Lys Lys Glu Phe Leu Cys Arg Asn Gln 3740
3745 3750Arg Cys Leu Ser Ser Ser Leu Arg Cys Asn Met
Phe Asp Asp Cys 3755 3760 3765Gly Asp
Gly Ser Asp Glu Glu Asp Cys Ser Ile Asp Pro Lys Leu 3770
3775 3780Thr Ser Cys Ala Thr Asn Ala Ser Met Cys
Gly Asp Glu Ala Arg 3785 3790 3795Cys
Val Arg Thr Glu Lys Ala Ala Tyr Cys Ala Cys Arg Ser Gly 3800
3805 3810Phe His Thr Val Pro Gly Gln Pro Gly
Cys Gln Asp Ile Asn Glu 3815 3820
3825Cys Leu Arg Phe Gly Thr Cys Ser Gln Leu Trp Asn Lys Pro Lys
3830 3835 3840Gly Gly His Leu Cys Ser
Cys Ala Arg Asn Phe Met Lys Thr His 3845 3850
3855Asn Thr Cys Lys Ala Glu Gly Ser Glu Tyr Gln Val Leu Tyr
Ile 3860 3865 3870Ala Asp Asp Asn Glu
Ile Arg Ser Leu Phe Pro Gly His Pro His 3875 3880
3885Ser Ala Tyr Glu Gln Thr Phe Gln Gly Asp Glu Ser Val
Arg Ile 3890 3895 3900Asp Ala Met Asp
Val His Val Lys Ala Gly Arg Val Tyr Trp Thr 3905
3910 3915Asn Trp His Thr Gly Thr Ile Ser Tyr Arg Ser
Leu Pro Pro Ala 3920 3925 3930Ala Pro
Pro Thr Thr Ser Asn Arg His Arg Arg Gln Ile Asp Arg 3935
3940 3945Gly Val Thr His Leu Asn Ile Ser Gly Leu
Lys Met Pro Arg Gly 3950 3955 3960Ile
Ala Ile Asp Trp Val Ala Gly Asn Val Tyr Trp Thr Asp Ser 3965
3970 3975Gly Arg Asp Val Ile Glu Val Ala Gln
Met Lys Gly Glu Asn Arg 3980 3985
3990Lys Thr Leu Ile Ser Gly Met Ile Asp Glu Pro His Ala Ile Val
3995 4000 4005Val Asp Pro Leu Arg Gly
Thr Met Tyr Trp Ser Asp Trp Gly Asn 4010 4015
4020His Pro Lys Ile Glu Thr Ala Ala Met Asp Gly Thr Leu Arg
Glu 4025 4030 4035Thr Leu Val Gln Asp
Asn Ile Gln Trp Pro Thr Gly Leu Ala Val 4040 4045
4050Asp Tyr His Asn Glu Arg Leu Tyr Trp Ala Asp Ala Lys
Leu Ser 4055 4060 4065Val Ile Gly Ser
Ile Arg Leu Asn Gly Thr Asp Pro Ile Val Ala 4070
4075 4080Ala Asp Ser Lys Arg Gly Leu Ser His Pro Phe
Ser Ile Asp Val 4085 4090 4095Phe Glu
Asp Tyr Ile Tyr Gly Val Thr Tyr Ile Asn Asn Arg Val 4100
4105 4110Phe Lys Ile His Lys Phe Gly His Ser Pro
Leu Tyr Asn Leu Thr 4115 4120 4125Gly
Gly Leu Ser His Ala Ser Asp Val Val Leu Tyr His Gln His 4130
4135 4140Lys Gln Pro Glu Val Thr Asn Pro Cys
Asp Arg Lys Lys Cys Glu 4145 4150
4155Trp Leu Cys Leu Leu Ser Pro Ser Gly Pro Val Cys Thr Cys Pro
4160 4165 4170Asn Gly Lys Arg Leu Asp
Asn Gly Thr Cys Val Pro Val Pro Ser 4175 4180
4185Pro Thr Pro Pro Pro Asp Ala Pro Arg Pro Gly Thr Cys Thr
Leu 4190 4195 4200Gln Cys Phe Asn Gly
Gly Ser Cys Phe Leu Asn Ala Arg Arg Gln 4205 4210
4215Pro Lys Cys Arg Cys Gln Pro Arg Tyr Thr Gly Asp Lys
Cys Glu 4220 4225 4230Leu Asp Gln Cys
Trp Glu Tyr Cys His Asn Gly Gly Thr Cys Ala 4235
4240 4245Ala Ser Pro Ser Gly Met Pro Thr Cys Arg Cys
Pro Thr Gly Phe 4250 4255 4260Thr Gly
Pro Lys Cys Thr Ala Gln Val Cys Ala Gly Tyr Cys Ser 4265
4270 4275Asn Asn Ser Thr Cys Thr Val Asn Gln Gly
Asn Gln Pro Gln Cys 4280 4285 4290Arg
Cys Leu Pro Gly Phe Leu Gly Asp Arg Cys Gln Tyr Arg Gln 4295
4300 4305Cys Ser Gly Phe Cys Glu Asn Phe Gly
Thr Cys Gln Met Ala Ala 4310 4315
4320Asp Gly Ser Arg Gln Cys Arg Cys Thr Val Tyr Phe Glu Gly Pro
4325 4330 4335Arg Cys Glu Val Asn Lys
Cys Ser Arg Cys Leu Gln Gly Ala Cys 4340 4345
4350Val Val Asn Lys Gln Thr Gly Asp Val Thr Cys Asn Cys Thr
Asp 4355 4360 4365Gly Arg Val Ala Pro
Ser Cys Leu Thr Cys Ile Asp His Cys Ser 4370 4375
4380Asn Gly Gly Ser Cys Thr Met Asn Ser Lys Met Met Pro
Glu Cys 4385 4390 4395Gln Cys Pro Pro
His Met Thr Gly Pro Arg Cys Gln Glu Gln Val 4400
4405 4410Val Ser Gln Gln Gln Pro Gly His Met Ala Ser
Ile Leu Ile Pro 4415 4420 4425Leu Leu
Leu Leu Leu Leu Leu Leu Leu Val Ala Gly Val Val Phe 4430
4435 4440Trp Tyr Lys Arg Arg Val Arg Gly Ala Lys
Gly Phe Gln His Gln 4445 4450 4455Arg
Met Thr Asn Gly Ala Met Asn Val Glu Ile Gly Asn Pro Thr 4460
4465 4470Tyr Lys Met Tyr Glu Gly Gly Glu Pro
Asp Asp Val Gly Gly Leu 4475 4480
4485Leu Asp Ala Asp Phe Ala Leu Asp Pro Asp Lys Pro Thr Asn Phe
4490 4495 4500Thr Asn Pro Val Tyr Ala
Thr Leu Tyr Met Gly Gly His Gly Ser 4505 4510
4515Arg His Ser Leu Ala Ser Thr Asp Glu Lys Arg Glu Leu Leu
Gly 4520 4525 4530Arg Gly Pro Glu Asp
Glu Ile Gly Asp Pro Leu Ala 4535 4540
454531474PRTHomo sapiens 3Met Gly Lys Asn Lys Leu Leu His Pro Ser Leu Val
Leu Leu Leu Leu1 5 10
15Val Leu Leu Pro Thr Asp Ala Ser Val Ser Gly Lys Pro Gln Tyr Met
20 25 30Val Leu Val Pro Ser Leu Leu
His Thr Glu Thr Thr Glu Lys Gly Cys 35 40
45Val Leu Leu Ser Tyr Leu Asn Glu Thr Val Thr Val Ser Ala Ser
Leu 50 55 60Glu Ser Val Arg Gly Asn
Arg Ser Leu Phe Thr Asp Leu Glu Ala Glu65 70
75 80Asn Asp Val Leu His Cys Val Ala Phe Ala Val
Pro Lys Ser Ser Ser 85 90
95Asn Glu Glu Val Met Phe Leu Thr Val Gln Val Lys Gly Pro Thr Gln
100 105 110Glu Phe Lys Lys Arg Thr
Thr Val Met Val Lys Asn Glu Asp Ser Leu 115 120
125Val Phe Val Gln Thr Asp Lys Ser Ile Tyr Lys Pro Gly Gln
Thr Val 130 135 140Lys Phe Arg Val Val
Ser Met Asp Glu Asn Phe His Pro Leu Asn Glu145 150
155 160Leu Ile Pro Leu Val Tyr Ile Gln Asp Pro
Lys Gly Asn Arg Ile Ala 165 170
175Gln Trp Gln Ser Phe Gln Leu Glu Gly Gly Leu Lys Gln Phe Ser Phe
180 185 190Pro Leu Ser Ser Glu
Pro Phe Gln Gly Ser Tyr Lys Val Val Val Gln 195
200 205Lys Lys Ser Gly Gly Arg Thr Glu His Pro Phe Thr
Val Glu Glu Phe 210 215 220Val Leu Pro
Lys Phe Glu Val Gln Val Thr Val Pro Lys Ile Ile Thr225
230 235 240Ile Leu Glu Glu Glu Met Asn
Val Ser Val Cys Gly Leu Tyr Thr Tyr 245
250 255Gly Lys Pro Val Pro Gly His Val Thr Val Ser Ile
Cys Arg Lys Tyr 260 265 270Ser
Asp Ala Ser Asp Cys His Gly Glu Asp Ser Gln Ala Phe Cys Glu 275
280 285Lys Phe Ser Gly Gln Leu Asn Ser His
Gly Cys Phe Tyr Gln Gln Val 290 295
300Lys Thr Lys Val Phe Gln Leu Lys Arg Lys Glu Tyr Glu Met Lys Leu305
310 315 320His Thr Glu Ala
Gln Ile Gln Glu Glu Gly Thr Val Val Glu Leu Thr 325
330 335Gly Arg Gln Ser Ser Glu Ile Thr Arg Thr
Ile Thr Lys Leu Ser Phe 340 345
350Val Lys Val Asp Ser His Phe Arg Gln Gly Ile Pro Phe Phe Gly Gln
355 360 365Val Arg Leu Val Asp Gly Lys
Gly Val Pro Ile Pro Asn Lys Val Ile 370 375
380Phe Ile Arg Gly Asn Glu Ala Asn Tyr Tyr Ser Asn Ala Thr Thr
Asp385 390 395 400Glu His
Gly Leu Val Gln Phe Ser Ile Asn Thr Thr Asn Val Met Gly
405 410 415Thr Ser Leu Thr Val Arg Val
Asn Tyr Lys Asp Arg Ser Pro Cys Tyr 420 425
430Gly Tyr Gln Trp Val Ser Glu Glu His Glu Glu Ala His His
Thr Ala 435 440 445Tyr Leu Val Phe
Ser Pro Ser Lys Ser Phe Val His Leu Glu Pro Met 450
455 460Ser His Glu Leu Pro Cys Gly His Thr Gln Thr Val
Gln Ala His Tyr465 470 475
480Ile Leu Asn Gly Gly Thr Leu Leu Gly Leu Lys Lys Leu Ser Phe Tyr
485 490 495Tyr Leu Ile Met Ala
Lys Gly Gly Ile Val Arg Thr Gly Thr His Gly 500
505 510Leu Leu Val Lys Gln Glu Asp Met Lys Gly His Phe
Ser Ile Ser Ile 515 520 525Pro Val
Lys Ser Asp Ile Ala Pro Val Ala Arg Leu Leu Ile Tyr Ala 530
535 540Val Leu Pro Thr Gly Asp Val Ile Gly Asp Ser
Ala Lys Tyr Asp Val545 550 555
560Glu Asn Cys Leu Ala Asn Lys Val Asp Leu Ser Phe Ser Pro Ser Gln
565 570 575Ser Leu Pro Ala
Ser His Ala His Leu Arg Val Thr Ala Ala Pro Gln 580
585 590Ser Val Cys Ala Leu Arg Ala Val Asp Gln Ser
Val Leu Leu Met Lys 595 600 605Pro
Asp Ala Glu Leu Ser Ala Ser Ser Val Tyr Asn Leu Leu Pro Glu 610
615 620Lys Asp Leu Thr Gly Phe Pro Gly Pro Leu
Asn Asp Gln Asp Asp Glu625 630 635
640Asp Cys Ile Asn Arg His Asn Val Tyr Ile Asn Gly Ile Thr Tyr
Thr 645 650 655Pro Val Ser
Ser Thr Asn Glu Lys Asp Met Tyr Ser Phe Leu Glu Asp 660
665 670Met Gly Leu Lys Ala Phe Thr Asn Ser Lys
Ile Arg Lys Pro Lys Met 675 680
685Cys Pro Gln Leu Gln Gln Tyr Glu Met His Gly Pro Glu Gly Leu Arg 690
695 700Val Gly Phe Tyr Glu Ser Asp Val
Met Gly Arg Gly His Ala Arg Leu705 710
715 720Val His Val Glu Glu Pro His Thr Glu Thr Val Arg
Lys Tyr Phe Pro 725 730
735Glu Thr Trp Ile Trp Asp Leu Val Val Val Asn Ser Ala Gly Val Ala
740 745 750Glu Val Gly Val Thr Val
Pro Asp Thr Ile Thr Glu Trp Lys Ala Gly 755 760
765Ala Phe Cys Leu Ser Glu Asp Ala Gly Leu Gly Ile Ser Ser
Thr Ala 770 775 780Ser Leu Arg Ala Phe
Gln Pro Phe Phe Val Glu Leu Thr Met Pro Tyr785 790
795 800Ser Val Ile Arg Gly Glu Ala Phe Thr Leu
Lys Ala Thr Val Leu Asn 805 810
815Tyr Leu Pro Lys Cys Ile Arg Val Ser Val Gln Leu Glu Ala Ser Pro
820 825 830Ala Phe Leu Ala Val
Pro Val Glu Lys Glu Gln Ala Pro His Cys Ile 835
840 845Cys Ala Asn Gly Arg Gln Thr Val Ser Trp Ala Val
Thr Pro Lys Ser 850 855 860Leu Gly Asn
Val Asn Phe Thr Val Ser Ala Glu Ala Leu Glu Ser Gln865
870 875 880Glu Leu Cys Gly Thr Glu Val
Pro Ser Val Pro Glu His Gly Arg Lys 885
890 895Asp Thr Val Ile Lys Pro Leu Leu Val Glu Pro Glu
Gly Leu Glu Lys 900 905 910Glu
Thr Thr Phe Asn Ser Leu Leu Cys Pro Ser Gly Gly Glu Val Ser 915
920 925Glu Glu Leu Ser Leu Lys Leu Pro Pro
Asn Val Val Glu Glu Ser Ala 930 935
940Arg Ala Ser Val Ser Val Leu Gly Asp Ile Leu Gly Ser Ala Met Gln945
950 955 960Asn Thr Gln Asn
Leu Leu Gln Met Pro Tyr Gly Cys Gly Glu Gln Asn 965
970 975Met Val Leu Phe Ala Pro Asn Ile Tyr Val
Leu Asp Tyr Leu Asn Glu 980 985
990Thr Gln Gln Leu Thr Pro Glu Val Lys Ser Lys Ala Ile Gly Tyr Leu
995 1000 1005Asn Thr Gly Tyr Gln Arg
Gln Leu Asn Tyr Lys His Tyr Asp Gly 1010 1015
1020Ser Tyr Ser Thr Phe Gly Glu Arg Tyr Gly Arg Asn Gln Gly
Asn 1025 1030 1035Thr Trp Leu Thr Ala
Phe Val Leu Lys Thr Phe Ala Gln Ala Arg 1040 1045
1050Ala Tyr Ile Phe Ile Asp Glu Ala His Ile Thr Gln Ala
Leu Ile 1055 1060 1065Trp Leu Ser Gln
Arg Gln Lys Asp Asn Gly Cys Phe Arg Ser Ser 1070
1075 1080Gly Ser Leu Leu Asn Asn Ala Ile Lys Gly Gly
Val Glu Asp Glu 1085 1090 1095Val Thr
Leu Ser Ala Tyr Ile Thr Ile Ala Leu Leu Glu Ile Pro 1100
1105 1110Leu Thr Val Thr His Pro Val Val Arg Asn
Ala Leu Phe Cys Leu 1115 1120 1125Glu
Ser Ala Trp Lys Thr Ala Gln Glu Gly Asp His Gly Ser His 1130
1135 1140Val Tyr Thr Lys Ala Leu Leu Ala Tyr
Ala Phe Ala Leu Ala Gly 1145 1150
1155Asn Gln Asp Lys Arg Lys Glu Val Leu Lys Ser Leu Asn Glu Glu
1160 1165 1170Ala Val Lys Lys Asp Asn
Ser Val His Trp Glu Arg Pro Gln Lys 1175 1180
1185Pro Lys Ala Pro Val Gly His Phe Tyr Glu Pro Gln Ala Pro
Ser 1190 1195 1200Ala Glu Val Glu Met
Thr Ser Tyr Val Leu Leu Ala Tyr Leu Thr 1205 1210
1215Ala Gln Pro Ala Pro Thr Ser Glu Asp Leu Thr Ser Ala
Thr Asn 1220 1225 1230Ile Val Lys Trp
Ile Thr Lys Gln Gln Asn Ala Gln Gly Gly Phe 1235
1240 1245Ser Ser Thr Gln Asp Thr Val Val Ala Leu His
Ala Leu Ser Lys 1250 1255 1260Tyr Gly
Ala Ala Thr Phe Thr Arg Thr Gly Lys Ala Ala Gln Val 1265
1270 1275Thr Ile Gln Ser Ser Gly Thr Phe Ser Ser
Lys Phe Gln Val Asp 1280 1285 1290Asn
Asn Asn Arg Leu Leu Leu Gln Gln Val Ser Leu Pro Glu Leu 1295
1300 1305Pro Gly Glu Tyr Ser Met Lys Val Thr
Gly Glu Gly Cys Val Tyr 1310 1315
1320Leu Gln Thr Ser Leu Lys Tyr Asn Ile Leu Pro Glu Lys Glu Glu
1325 1330 1335Phe Pro Phe Ala Leu Gly
Val Gln Thr Leu Pro Gln Thr Cys Asp 1340 1345
1350Glu Pro Lys Ala His Thr Ser Phe Gln Ile Ser Leu Ser Val
Ser 1355 1360 1365Tyr Thr Gly Ser Arg
Ser Ala Ser Asn Met Ala Ile Val Asp Val 1370 1375
1380Lys Met Val Ser Gly Phe Ile Pro Leu Lys Pro Thr Val
Lys Met 1385 1390 1395Leu Glu Arg Ser
Asn His Val Ser Arg Thr Glu Val Ser Ser Asn 1400
1405 1410His Val Leu Ile Tyr Leu Asp Lys Val Ser Asn
Gln Thr Leu Ser 1415 1420 1425Leu Phe
Phe Thr Val Leu Gln Asp Val Pro Val Arg Asp Leu Lys 1430
1435 1440Pro Ala Ile Val Lys Val Tyr Asp Tyr Tyr
Glu Thr Asp Glu Phe 1445 1450 1455Ala
Ile Ala Glu Tyr Asn Ala Pro Cys Ser Lys Asp Leu Gly Asn 1460
1465 1470Ala44577DNAHomo sapiens 4gctacaatcc
atctggtctc ctccagctcc ttctttctgc aacatgggga agaacaaact 60ccttcatcca
agtctggttc ttctcctctt ggtcctcctg cccacagacg cctcagtctc 120tggaaaaccg
cagtatatgg ttctggtccc ctccctgctc cacactgaga ccactgagaa 180gggctgtgtc
cttctgagct acctgaatga gacagtgact gtaagtgctt ccttggagtc 240tgtcagggga
aacaggagcc tcttcactga cctggaggcg gagaatgacg tactccactg 300tgtcgccttc
gctgtcccaa agtcttcatc caatgaggag gtaatgttcc tcactgtcca 360agtgaaagga
ccaacccaag aatttaagaa gcggaccaca gtgatggtta agaacgagga 420cagtctggtc
tttgtccaga cagacaaatc aatctacaaa ccagggcaga cagtgaaatt 480tcgtgttgtc
tccatggatg aaaactttca ccccctgaat gagttgattc cactagtata 540cattcaggat
cccaaaggaa atcgcatcgc acaatggcag agtttccagt tagagggtgg 600cctcaagcaa
ttttcttttc ccctctcatc agagcccttc cagggctcct acaaggtggt 660ggtacagaag
aaatcaggtg gaaggacaga gcaccctttc accgtggagg aatttgttct 720tcccaagttt
gaagtacaag taacagtgcc aaagataatc accatcttgg aagaagagat 780gaatgtatca
gtgtgtggcc tatacacata tgggaagcct gtccctggac atgtgactgt 840gagcatttgc
agaaagtata gtgacgcttc cgactgccac ggtgaagatt cacaggcttt 900ctgtgagaaa
ttcagtggac agctaaacag ccatggctgc ttctatcagc aagtaaaaac 960caaggtcttc
cagctgaaga ggaaggagta tgaaatgaaa cttcacactg aggcccagat 1020ccaagaagaa
ggaacagtgg tggaattgac tggaaggcag tccagtgaaa tcacaagaac 1080cataaccaaa
ctctcatttg tgaaagtgga ctcacacttt cgacagggaa ttcccttctt 1140tgggcaggtg
cgcctagtag atgggaaagg cgtccctata ccaaataaag tcatattcat 1200cagaggaaat
gaagcaaact attactccaa tgctaccacg gatgagcatg gccttgtaca 1260gttctctatc
aacaccacca acgttatggg tacctctctt actgttaggg tcaattacaa 1320ggatcgtagt
ccctgttacg gctaccagtg ggtgtcagaa gaacacgaag aggcacatca 1380cactgcttat
cttgtgttct ccccaagcaa gagctttgtc caccttgagc ccatgtctca 1440tgaactaccc
tgtggccata ctcagacagt ccaggcacat tatattctga atggaggcac 1500cctgctgggg
ctgaagaagc tctcctttta ttatctgata atggcaaagg gaggcattgt 1560ccgaactggg
actcatggac tgcttgtgaa gcaggaagac atgaagggcc atttttccat 1620ctcaatccct
gtgaagtcag acattgctcc tgtcgctcgg ttgctcatct atgctgtttt 1680acctaccggg
gacgtgattg gggattctgc aaaatatgat gttgaaaatt gtctggccaa 1740caaggtggat
ttgagcttca gcccatcaca aagtctccca gcctcacacg cccacctgcg 1800agtcacagcg
gctcctcagt ccgtctgcgc cctccgtgct gtggaccaaa gcgtgctgct 1860catgaagcct
gatgctgagc tctcggcgtc ctcggtttac aacctgctac cagaaaagga 1920cctcactggc
ttccctgggc ctttgaatga ccaggacgat gaagactgca tcaatcgtca 1980taatgtctat
attaatggaa tcacatatac tccagtatca agtacaaatg aaaaggatat 2040gtacagcttc
ctagaggaca tgggcttaaa ggcattcacc aactcaaaga ttcgtaaacc 2100caaaatgtgt
ccacagcttc aacagtatga aatgcatgga cctgaaggtc tacgtgtagg 2160tttttatgag
tcagatgtaa tgggaagagg ccatgcacgc ctggtgcatg ttgaagagcc 2220tcacacggag
accgtacgaa agtacttccc tgagacatgg atctgggatt tggtggtggt 2280aaactcagca
ggggtggctg aggtaggagt aacagtccct gacaccatca ccgagtggaa 2340ggcaggggcc
ttctgcctgt ctgaagatgc tggacttggt atctcttcca ctgcctctct 2400ccgagccttc
cagcccttct ttgtggagct tacaatgcct tactctgtga ttcgtggaga 2460ggccttcaca
ctcaaggcca cggtcctaaa ctaccttccc aaatgcatcc gggtcagtgt 2520gcagctggaa
gcctctcccg ccttccttgc tgtcccagtg gagaaggaac aagcgcctca 2580ctgcatctgt
gcaaacgggc ggcaaactgt gtcctgggca gtaaccccaa agtcattagg 2640aaatgtgaat
ttcactgtga gcgcagaggc actagagtct caagagctgt gtgggactga 2700ggtgccttca
gttcctgaac acggaaggaa agacacagtc atcaagcctc tgttggttga 2760acctgaagga
ctagagaagg aaacaacatt caactcccta ctttgtccat caggtggtga 2820ggtttctgaa
gaattatccc tgaaactgcc accaaatgtg gtagaagaat ctgcccgagc 2880ttctgtctca
gttttgggag acatattagg ctctgccatg caaaacacac aaaatcttct 2940ccagatgccc
tatggctgtg gagagcagaa tatggtcctc tttgctccta acatctatgt 3000actggattat
ctaaatgaaa cacagcagct tactccagag gtcaagtcca aggccattgg 3060ctatctcaac
actggttacc agagacagtt gaactacaaa cactatgatg gctcctacag 3120cacctttggg
gagcgatatg gcaggaacca gggcaacacc tggctcacag cctttgttct 3180gaagactttt
gcccaagctc gagcctacat cttcatcgat gaagcacaca ttacccaagc 3240cctcatatgg
ctctcccaga ggcagaagga caatggctgt ttcaggagct ctgggtcact 3300gctcaacaat
gccataaagg gaggagtaga agatgaagtg accctctccg cctatatcac 3360catcgccctt
ctggagattc ctctcacagt cactcaccct gttgtccgca atgccctgtt 3420ttgcctggag
tcagcctgga agacagcaca agaaggggac catggcagcc atgtatatac 3480caaagcactg
ctggcctatg cttttgccct ggcaggtaac caggacaaga ggaaggaagt 3540actcaagtca
cttaatgagg aagctgtgaa gaaagacaac tctgtccatt gggagcgccc 3600tcagaaaccc
aaggcaccag tggggcattt ttacgaaccc caggctccct ctgctgaggt 3660ggagatgaca
tcctatgtgc tcctcgctta tctcacggcc cagccagccc caacctcgga 3720ggacctgacc
tctgcaacca acatcgtgaa gtggatcacg aagcagcaga atgcccaggg 3780cggtttctcc
tccacccagg acacagtggt ggctctccat gctctgtcca aatatggagc 3840cgccacattt
accaggactg ggaaggctgc acaggtgact atccagtctt cagggacatt 3900ttccagcaaa
ttccaagtgg acaacaacaa tcgcctgtta ctgcagcagg tctcattgcc 3960agagctgcct
ggggaataca gcatgaaagt gacaggagaa ggatgtgtct acctccagac 4020ctccttgaaa
tacaatattc tcccagaaaa ggaagagttc ccctttgctt taggagtgca 4080gactctgcct
caaacttgtg atgaacccaa agcccacacc agcttccaaa tctccctaag 4140tgtcagttac
acagggagcc gctctgcctc caacatggcg atcgttgatg tgaagatggt 4200ctctggcttc
attcccctga agccaacagt gaaaatgctt gaaagatcta accatgtgag 4260ccggacagaa
gtcagcagca accatgtctt gatttacctt gataaggtgt caaatcagac 4320actgagcttg
ttcttcacgg ttctgcaaga tgtcccagta agagatctca aaccagccat 4380agtgaaagtc
tatgattact acgagacgga tgagtttgca atcgctgagt acaatgctcc 4440ttgcagcaaa
gatcttggaa atgcttgaag accacaaggc tgaaaagtgc tttgctggag 4500tcctgttctc
tgagctccac agaagacacg tgtttttgta tctttaaaga cttgatgaat 4560aaacactttt
tctggtc 45775138PRTHomo
sapiens 5Glu Glu Phe Pro Phe Ala Leu Gly Val Gln Thr Leu Pro Gln Thr Cys
1 5 10 15Asp Glu Pro Lys
Ala His Thr Ser Phe Gln Ile Ser Leu Ser Val Ser 20
25 30Tyr Thr Gly Ser Arg Ser Ala Ser Asn Met Ala
Ile Val Asp Val Lys 35 40 45Met
Val Ser Gly Phe Ile Pro Leu Lys Pro Thr Val Lys Met Leu Glu 50
55 60Arg Ser Asn His Val Ser Arg Thr Glu Val
Ser Ser Asn His Val Leu65 70 75
80Ile Tyr Leu Asp Lys Val Ser Asn Gln Thr Leu Ser Leu Phe Phe
Thr 85 90 95Val Leu Gln
Asp Val Pro Val Arg Asp Leu Lys Pro Ala Ile Val Lys 100
105 110Val Tyr Asp Tyr Tyr Glu Thr Asp Glu Phe
Ala Ile Ala Glu Tyr Asn 115 120
125Ala Pro Cys Ser Lys Asp Leu Gly Asn Ala 130
135610PRTHomo sapiens 6Gly Gly Ala Leu His Ile Tyr His Gln Arg1
5 10719PRTHomo sapiensAH-1 20-mer peptide 7Arg Val
Thr Tyr His Ser Pro Ser Tyr Val Tyr His Gln Phe Glu Arg1 5
10 15Arg Ala Lys81451PRTHomo
sapiensmature alpha-2M sequence 8Ser Val Ser Gly Lys Pro Gln Tyr Met Val
Leu Val Pro Ser Leu Leu 1 5 10
15His Thr Glu Thr Thr Glu Lys Gly Cys Val Leu Leu Ser Tyr Leu Asn
20 25 30Glu Thr Val Thr Val Ser
Ala Ser Leu Glu Ser Val Arg Gly Asn Arg 35 40
45Ser Leu Phe Thr Asp Leu Glu Ala Glu Asn Asp Val Leu His
Cys Val 50 55 60Ala Phe Ala Val Pro
Lys Ser Ser Ser Asn Glu Glu Val Met Phe Leu65 70
75 80Thr Val Gln Val Lys Gly Pro Thr Gln Glu
Phe Lys Lys Arg Thr Thr 85 90
95Val Met Val Lys Asn Glu Asp Ser Leu Val Phe Val Gln Thr Asp Lys
100 105 110Ser Ile Tyr Lys Pro
Gly Gln Thr Val Lys Phe Arg Val Val Ser Met 115
120 125Asp Glu Asn Phe His Pro Leu Asn Glu Leu Ile Pro
Leu Val Tyr Ile 130 135 140Gln Asp Pro
Lys Gly Asn Arg Ile Ala Gln Trp Gln Ser Phe Gln Leu145
150 155 160Glu Gly Gly Leu Lys Gln Phe
Ser Phe Pro Leu Ser Ser Glu Pro Phe 165
170 175Gln Gly Ser Tyr Lys Val Val Val Gln Lys Lys Ser
Gly Gly Arg Thr 180 185 190Glu
His Pro Phe Thr Val Glu Glu Phe Val Leu Pro Lys Phe Glu Val 195
200 205Gln Val Thr Val Pro Lys Ile Ile Thr
Ile Leu Glu Glu Glu Met Asn 210 215
220Val Ser Val Cys Gly Leu Tyr Thr Tyr Gly Lys Pro Val Pro Gly His225
230 235 240Val Thr Val Ser
Ile Cys Arg Lys Tyr Ser Asp Ala Ser Asp Cys His 245
250 255Gly Glu Asp Ser Gln Ala Phe Cys Glu Lys
Phe Ser Gly Gln Leu Asn 260 265
270Ser His Gly Cys Phe Tyr Gln Gln Val Lys Thr Lys Val Phe Gln Leu
275 280 285Lys Arg Lys Glu Tyr Glu Met
Lys Leu His Thr Glu Ala Gln Ile Gln 290 295
300Glu Glu Gly Thr Val Val Glu Leu Thr Gly Arg Gln Ser Ser Glu
Ile305 310 315 320Thr Arg
Thr Ile Thr Lys Leu Ser Phe Val Lys Val Asp Ser His Phe
325 330 335Arg Gln Gly Ile Pro Phe Phe
Gly Gln Val Arg Leu Val Asp Gly Lys 340 345
350Gly Val Pro Ile Pro Asn Lys Val Ile Phe Ile Arg Gly Asn
Glu Ala 355 360 365Asn Tyr Tyr Ser
Asn Ala Thr Thr Asp Glu His Gly Leu Val Gln Phe 370
375 380Ser Ile Asn Thr Thr Asn Val Met Gly Thr Ser Leu
Thr Val Arg Val385 390 395
400Asn Tyr Lys Asp Arg Ser Pro Cys Tyr Gly Tyr Gln Trp Val Ser Glu
405 410 415Glu His Glu Glu Ala
His His Thr Ala Tyr Leu Val Phe Ser Pro Ser 420
425 430Lys Ser Phe Val His Leu Glu Pro Met Ser His Glu
Leu Pro Cys Gly 435 440 445His Thr
Gln Thr Val Gln Ala His Tyr Ile Leu Asn Gly Gly Thr Leu 450
455 460Leu Gly Leu Lys Lys Leu Ser Phe Tyr Tyr Leu
Ile Met Ala Lys Gly465 470 475
480Gly Ile Val Arg Thr Gly Thr His Gly Leu Leu Val Lys Gln Glu Asp
485 490 495Met Lys Gly His
Phe Ser Ile Ser Ile Pro Val Lys Ser Asp Ile Ala 500
505 510Pro Val Ala Arg Leu Leu Ile Tyr Ala Val Leu
Pro Thr Gly Asp Val 515 520 525Ile
Gly Asp Ser Ala Lys Tyr Asp Val Glu Asn Cys Leu Ala Asn Lys 530
535 540Val Asp Leu Ser Phe Ser Pro Ser Gln Ser
Leu Pro Ala Ser His Ala545 550 555
560His Leu Arg Val Thr Ala Ala Pro Gln Ser Val Cys Ala Leu Arg
Ala 565 570 575Val Asp Gln
Ser Val Leu Leu Met Lys Pro Asp Ala Glu Leu Ser Ala 580
585 590Ser Ser Val Tyr Asn Leu Leu Pro Glu Lys
Asp Leu Thr Gly Phe Pro 595 600
605Gly Pro Leu Asn Asp Gln Asp Asp Glu Asp Cys Ile Asn Arg His Asn 610
615 620Val Tyr Ile Asn Gly Ile Thr Tyr
Thr Pro Val Ser Ser Thr Asn Glu625 630
635 640Lys Asp Met Tyr Ser Phe Leu Glu Asp Met Gly Leu
Lys Ala Phe Thr 645 650
655Asn Ser Lys Ile Arg Lys Pro Lys Met Cys Pro Gln Leu Gln Gln Tyr
660 665 670Glu Met His Gly Pro Glu
Gly Leu Arg Val Gly Phe Tyr Glu Ser Asp 675 680
685Val Met Gly Arg Gly His Ala Arg Leu Val His Val Glu Glu
Pro His 690 695 700Thr Glu Thr Val Arg
Lys Tyr Phe Pro Glu Thr Trp Ile Trp Asp Leu705 710
715 720Val Val Val Asn Ser Ala Gly Val Ala Glu
Val Gly Val Thr Val Pro 725 730
735Asp Thr Ile Thr Glu Trp Lys Ala Gly Ala Phe Cys Leu Ser Glu Asp
740 745 750Ala Gly Leu Gly Ile
Ser Ser Thr Ala Ser Leu Arg Ala Phe Gln Pro 755
760 765Phe Phe Val Glu Leu Thr Met Pro Tyr Ser Val Ile
Arg Gly Glu Ala 770 775 780Phe Thr Leu
Lys Ala Thr Val Leu Asn Tyr Leu Pro Lys Cys Ile Arg785
790 795 800Val Ser Val Gln Leu Glu Ala
Ser Pro Ala Phe Leu Ala Val Pro Val 805
810 815Glu Lys Glu Gln Ala Pro His Cys Ile Cys Ala Asn
Gly Arg Gln Thr 820 825 830Val
Ser Trp Ala Val Thr Pro Lys Ser Leu Gly Asn Val Asn Phe Thr 835
840 845Val Ser Ala Glu Ala Leu Glu Ser Gln
Glu Leu Cys Gly Thr Glu Val 850 855
860Pro Ser Val Pro Glu His Gly Arg Lys Asp Thr Val Ile Lys Pro Leu865
870 875 880Leu Val Glu Pro
Glu Gly Leu Glu Lys Glu Thr Thr Phe Asn Ser Leu 885
890 895Leu Cys Pro Ser Gly Gly Glu Val Ser Glu
Glu Leu Ser Leu Lys Leu 900 905
910Pro Pro Asn Val Val Glu Glu Ser Ala Arg Ala Ser Val Ser Val Leu
915 920 925Gly Asp Ile Leu Gly Ser Ala
Met Gln Asn Thr Gln Asn Leu Leu Gln 930 935
940Met Pro Tyr Gly Cys Gly Glu Gln Asn Met Val Leu Phe Ala Pro
Asn945 950 955 960Ile Tyr
Val Leu Asp Tyr Leu Asn Glu Thr Gln Gln Leu Thr Pro Glu
965 970 975Val Lys Ser Lys Ala Ile Gly
Tyr Leu Asn Thr Gly Tyr Gln Arg Gln 980 985
990Leu Asn Tyr Lys His Tyr Asp Gly Ser Tyr Ser Thr Phe Gly
Glu Arg 995 1000 1005Tyr Gly Arg
Asn Gln Gly Asn Thr Trp Leu Thr Ala Phe Val Leu Lys 1010
1015 1020Thr Phe Ala Gln Ala Arg Ala Tyr Ile Phe Ile Asp
Glu Ala His Ile1025 1030 1035
1040Thr Gln Ala Leu Ile Trp Leu Ser Gln Arg Gln Lys Asp Asn Gly Cys
1045 1050 1055Phe Arg Ser Ser Gly
Ser Leu Leu Asn Asn Ala Ile Lys Gly Gly Val 1060
1065 1070Glu Asp Glu Val Thr Leu Ser Ala Tyr Ile Thr Ile
Ala Leu Leu Glu 1075 1080 1085Ile
Pro Leu Thr Val Thr His Pro Val Val Arg Asn Ala Leu Phe Cys 1090
1095 1100Leu Glu Ser Ala Trp Lys Thr Ala Gln Glu
Gly Asp His Gly Ser His1105 1110 1115
1120Val Tyr Thr Lys Ala Leu Leu Ala Tyr Ala Phe Ala Leu Ala Gly
Asn 1125 1130 1135Gln Asp
Lys Arg Lys Glu Val Leu Lys Ser Leu Asn Glu Glu Ala Val 1140
1145 1150Lys Lys Asp Asn Ser Val His Trp Glu
Arg Pro Gln Lys Pro Lys Ala 1155 1160
1165Pro Val Gly His Phe Tyr Glu Pro Gln Ala Pro Ser Ala Glu Val Glu
1170 1175 1180Met Thr Ser Tyr Val Leu Leu
Ala Tyr Leu Thr Ala Gln Pro Ala Pro1185 1190
1195 1200Thr Ser Glu Asp Leu Thr Ser Ala Thr Asn Ile Val
Lys Trp Ile Thr 1205 1210
1215Lys Gln Gln Asn Ala Gln Gly Gly Phe Ser Ser Thr Gln Asp Thr Val
1220 1225 1230Val Ala Leu His Ala Leu
Ser Lys Tyr Gly Ala Ala Thr Phe Thr Arg 1235 1240
1245Thr Gly Lys Ala Ala Gln Val Thr Ile Gln Ser Ser Gly Thr
Phe Ser 1250 1255 1260Ser Lys Phe Gln
Val Asp Asn Asn Asn Arg Leu Leu Leu Gln Gln Val1265 1270
1275 1280Ser Leu Pro Glu Leu Pro Gly Glu Tyr
Ser Met Lys Val Thr Gly Glu 1285 1290
1295Gly Cys Val Tyr Leu Gln Thr Ser Leu Lys Tyr Asn Ile Leu Pro
Glu 1300 1305 1310Lys Glu Glu
Phe Pro Phe Ala Leu Gly Val Gln Thr Leu Pro Gln Thr 1315
1320 1325Cys Asp Glu Pro Lys Ala His Thr Ser Phe Gln
Ile Ser Leu Ser Val 1330 1335 1340Ser
Tyr Thr Gly Ser Arg Ser Ala Ser Asn Met Ala Ile Val Asp Val1345
1350 1355 1360Lys Met Val Ser Gly Phe
Ile Pro Leu Lys Pro Thr Val Lys Met Leu 1365
1370 1375Glu Arg Ser Asn His Val Ser Arg Thr Glu Val Ser
Ser Asn His Val 1380 1385
1390Leu Ile Tyr Leu Asp Lys Val Ser Asn Gln Thr Leu Ser Leu Phe Phe
1395 1400 1405Thr Val Leu Gln Asp Val Pro
Val Arg Asp Leu Lys Pro Ala Ile Val 1410 1415
1420Lys Val Tyr Asp Tyr Tyr Glu Thr Asp Glu Phe Ala Ile Ala Glu
Tyr1425 1430 1435 1440Asn
Ala Pro Cys Ser Lys Asp Leu Gly Asn Ala 1445
1450910PRTHomo sapiens 9Ser Gly Phe Ser Leu Gly Ser Asp Gly Lys1
5 101010PRTHomo sapiens 10Gly Ile Ala Leu Asp Pro
Ala Met Gly Lys1 5 101110PRTHomo sapiens
11Gly Gly Ala Leu His Ile Tyr His Gln Arg1 5
101211PRTHomo sapiens 12Val Phe Phe Thr Asp Tyr Gly Gln Ile Pro Lys1
5 10
User Contributions:
comments("1"); ?> comment_form("1"); ?>Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
User Contributions:
Comment about this patent or add new information about this topic:
| People who visited this patent also read: | |
| Patent application number | Title |
|---|---|
| 20150100662 | UTILIZING MULTIPLE DATA STRUCTURES FOR SLICE STORAGE |
| 20150100661 | SYSTEMS AND METHODS FOR MAPPING AND ROUTING BASED ON CLUSTERING |
| 20150100660 | SYSTEMS AND METHODS FOR CACHING CONTENT WITH NOTIFICATION-BASED INVALIDATION |
| 20150100659 | CACHING MECHANISM FOR DETERMINING VALIDITY OF CACHED DATA USING WEB APPLICATION BUSINESS LOGIC |
| 20150100658 | Dual Mode Master/Slave Interface |