Patent application title: Novel Carboxylesterase Nucleic Acid Molecules, Proteins and Uses Thereof
Inventors:
Gary M. Silver (Fort Collins, CO, US)
Nancy Wisnewski (Fort Collins, CO, US)
Kevin S. Brandt (Fort Collins, CO, US)
IPC8 Class: AC12Q144FI
USPC Class:
435 19
Class name: Measuring or testing process involving enzymes or micro-organisms; composition or test strip therefore; processes of forming such composition or test strip involving hydrolase involving esterase
Publication date: 2009-07-02
Patent application number: 20090170146
Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
Patent application title: Novel Carboxylesterase Nucleic Acid Molecules, Proteins and Uses Thereof
Inventors:
Nancy Wisnewski
Kevin S. Brandt
Gary M. Silver
Agents:
SHERIDAN ROSS P.C.
Assignees:
Origin: DENVER, CO US
IPC8 Class: AC12Q144FI
USPC Class:
435 19
Abstract:
The present invention relates to arthropod esterase proteins; to arthropod
esterase nucleic acid molecules, including those that encode such
esterase proteins; to antibodies raised against such esterase proteins;
and to other compounds that inhibit arthropod esterase activity. The
present invention also includes methods to obtain such proteins, nucleic
acid molecules, antibodies, and inhibitory compounds. Also included in
the present invention are therapeutic compositions comprising such
proteins, nucleic acid molecules, antibodies and/or inhibitory compounds
as well as the use of such therapeutic compositions to protect animals
from hematophagous arthropod infestation.Claims:
1-59. (canceled)
60. An isolated nucleic acid molecule selected from the group consisting of: (a) a nucleic acid molecule that encodes a protein at least 95% identical to a protein selected from the group consisting of SEQ ID NO:2, SEQ ID NO:5, SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:19, SEQ ID NO:25, SEQ ID NO:31, SEQ ID NO:37, SEQ ID NO:39, SEQ ID NO:40, SEQ ID NO:41, SEQ ID NO:42, SEQ ID NO:43, SEQ ID NO:44, SEQ ID NO:53, SEQ ID NO:54, SEQ ID NO:55, SEQ ID NO:58, SEQ ID NO:73, SEQ ID NO:74; and (b) an isolated nucleic acid molecule fully complementary to a nucleic acid molecule of (a).
61. The isolated nucleic acid molecule of claim 60, wherein said encoded protein has esterase activity.
62. A recombinant molecule comprising a nucleic acid molecule as set forth in claim 60 operatively linked to a transcription control sequence.
63. A recombinant virus comprising a nucleic acid molecule as set forth in claim 60.
64. A recombinant cell comprising a nucleic acid molecule as set forth in claim 60.
65. A method to produce a carboxylesterase protein, said method comprising culturing a cell capable of expressing said protein, said protein being encoded by a nucleic acid molecule of claim 60, part (a).
66. An isolated protein comprising an amino acid sequence at least 95% identical to a protein selected from the group consisting of SEQ ID NO:2, SEQ ID NO:5, SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:19, SEQ ID NO:25, SEQ ID NO:31, SEQ ID NO:37, SEQ ID NO:39, SEQ ID NO:40, SEQ ID NO:41, SEQ ID NO:42, SEQ ID NO:43, SEQ ID NO:44, SEQ ID NO:53, SEQ ID NO:54, SEQ ID NO:55, SEQ ID NO:58, SEQ ID NO:73, SEQ ID NO:74.
67. The isolated protein of claim 66, wherein said isolated protein has carboxylesterase activity.
68. The protein of claim 66, wherein said protein, when administered to an animal, elicits an immune response against a carboxylesterase protein.
69. A method to identify a compound capable of inhibiting flea carboxylesterase activity, said method comprising:(a) contacting a protein comprising an amino acid sequence at least 95% identical to a protein selected from the group consisting of SEQ ID NO:2, SEQ ID NO:5, SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:19, SEQ ID NO:25, SEQ ID NO:31, SEQ ID NO:37, SEQ ID NO:39, SEQ ID NO:40, SEQ ID NO:41, SEQ ID NO:42, SEQ ID NO:43, SEQ ID NO:44, SEQ ID NO:53, SEQ ID NO:54, SEQ ID NO:55, SEQ ID NO:58, SEQ ID NO:73, SEQ ID NO:74. with a putative inhibitory compound under conditions in which, in the absence of said compound, said protein has carboxylesterase activity; and(b) determining if said putative inhibitory compound inhibits said activity.
Description:
CROSS-REFERENCE TO RELATED APPLICATIONS
[0001]This application is a Continuation of co-pending U.S. patent application Ser. No. 10/678,521, filed Oct. 2, 2003; which is a Divisional of U.S. patent application Ser. No. 09/403,942, filed May 2, 2000, and issued as U.S. Pat. No. 6,664,090; which is a 371 filing of International Patent Application No. PCT/US97/20598, filed Nov. 10, 1997, which is a continuation-in-part of U.S. Ser. No. 08/747,221, filed Nov. 12, 1996, which issued as U.S. Pat. No. 6,063,610 on May 16, 2000; all of which are entitled "NOVEL CARBOXYLESTERASE NUCLEIC ACID MOLECULES, PROTEINS AND USES THEREOF" and are incorporated herein by reference.
FIELD OF THE INVENTION
[0002]The present invention relates to arthropod esterase nucleic acid molecules, proteins encoded by such nucleic acid molecules, antibodies raised against such proteins, and inhibitors of such proteins. The present invention also includes therapeutic compositions comprising such nucleic acid molecules, proteins, antibodies, and/or other inhibitors, as well as their use to protect an animal from hematophagous arthropod infestation.
BACKGROUND OF THE INVENTION
[0003]Hematophagous arthropod infestation of animals is a health and economic concern because hematophagous arthropods are known to cause and/or transmit a variety of diseases. Hematophagous arthropods directly cause a variety of diseases, including allergies, and also carry a variety of infectious agents including, but not limited to, endoparasites (e.g., nematodes, cestodes, trematodes and protozoa), bacteria and viruses. In particular, the bites of hematophagous arthropods are a problem for animals maintained as pets because the infestation becomes a source of annoyance not only for the pet but also for the pet owner who may find his or her home generally contaminated with insects. As such, hematophagous arthropods are a problem not only when they are on an animal but also when they are in the general environment of the animal.
[0004]Bites from hematophagous arthropods are a particular problem because they not only can lead to disease transmission but also can cause a hypersensitive response in animals which is manifested as disease. For example, bites from fleas can cause an allergic disease called flea allergic (or allergy) dermatitis (FAD). A hypersensitive response in animals typically results in localized tissue inflammation and damage, causing substantial discomfort to the animal.
[0005]The medical importance of arthropod infestation has prompted the development of reagents capable of controlling arthropod infestation. Commonly encountered methods to control arthropod infestation are generally focused on use of insecticides. While some of these products are efficacious, most, at best, offer protection of a very limited duration. Furthermore, many of the methods are often not successful in reducing arthropod populations. In particular, insecticides have been used to prevent hematophagous arthropod infestation of animals by adding such insecticides to shampoos, powders, collars, sprays, foggers and liquid bath treatments (i.e., dips). Reduction of hematophagous arthropod infestation on the pet has been unsuccessful for one or more of the following reasons: (1) failure of owner compliance (frequent administration is required); (2) behavioral or physiological intolerance of the pet to the pesticide product or means of administration; and (3) the emergence of hematophagous arthropod populations resistant to the prescribed dose of pesticide. However, hematophagous arthropod populations have been found to become resistant to insecticides.
[0006]Prior investigators have described insect carboxylesterase (CE) protein biochemistry, for example, Chen et al., Insect Biochem. Molec. Biol., 24:347-355, 1994; Whyard et al., Biochemical Genetics, 32:924, 1994 and Argentine et al., Insect Biochem. Molec Biol, 25:621-630, 1995. Other investigators have disclosed certain insect CE amino acid sequences, for example, Mouches et al., Proc Natl Acad Sci USA, 87:2574-2578, 1990 and Cooke et al., Proc Natl Acad Sci USA, 86:1426-1430, 1989, and nucleic acid sequence (Vaughn et al., J. Biol. Chem., 270:17044-17049, 1995). [0007]Prior investigators have described certain insect juvenile hormone esterase (JHE) nucleic acid and amino acid sequences: for example, sequence for Heliothis virescens is disclosed by Hanzlik et al., J. Biol. Chem., 264:12419-12425, 1989; Eldridge et al., App Environ Microbiol, 58:1583-1591, 1992; Bonning et al., Insect Biochem. Molec. Biol., 22:453-458, 1992; Bonning et al., Natural and Engineered Pest Management Agents, pp. 368-383, 1994 and Harshman et al., Insect Biochem. Molec. Biol, 24:671-676, 1994; sequence for Manduca sexta is disclosed by Vankatesh et al., J Biol Chem, 265:21727-21732, 1990; sequence for Trichoplusia ni is disclosed by Venkatararnan et al., Dev. Genet., 15:391-400, 1994 and Jones et al., Biochem. J, 302:827-835, 1994; and sequence for Lymantria dispar is disclosed by Valaitis, Insect Biochem. Molec. Biol., 22:639-648, 1992.
[0008]Identification of an esterase of the present invention is unexpected, however, because even the most similar nucleic acid sequence identified by previous investigators could not be used to identify an esterase of the present invention. In addition, identification of an esterase protein of the present invention is unexpected because a protein fraction from flea prepupal larvae that was obtained by monitoring for serine protease activity surprisingly also contained esterase proteins of the present invention.
[0009]In summary, there remains a need to develop a reagent and a method to protect animals or plants from hematophagous arthropod infestation.
SUMMARY OF THE INVENTION
[0010]The present invention relates to a novel product and process for protection of animals or plants from arthropod infestation. According to the present invention there are provided arthropod esterase proteins and mimetopes thereof; arthropod nucleic acid molecules, including those that encode such proteins; antibodies raised against such esterase proteins (i.e., anti-arthropod esterase antibodies); and compounds that inhibit arthropod esterase activity (i.e, inhibitory compounds or inhibitors).
[0011]The present invention also includes methods to obtain such proteins, mimetopes, nucleic acid molecules, antibodies and inhibitory compounds. Also included in the present invention are therapeutic compositions comprising such proteins, mimetopes, nucleic acid molecules, antibodies, and/or inhibitory compounds, as well as use of such therapeutic compositions to protect animals from arthropod infestation.
[0012]Identification of an esterase of the present invention is unexpected, however, because the most similar nucleic acid sequence identified by previous investigators could not be used to identify an esterase of the present invention. In addition, identification of an esterase protein of the present invention is unexpected because a protein fraction from flea prepupal larvae that was obtained by monitoring for serine protease activity surprisingly also contained esterase proteins of the present invention.
[0013]One embodiment of the present invention is an isolated nucleic acid molecule that hybridizes under stringent hybridization conditions with a gene comprising a nucleic acid sequence including SEQ ID NO:1, SEQ ID NO:3, SEQ ID NO:4, SEQ ID NO:6, SEQ ID NO:7, SEQ ID NO:9, SEQ ID NO:10, SEQ ID NO:12, SEQ ID NO:13, SEQ ID NO:15, SEQ ID NO:16, SEQ ID NO:17, SEQ ID NO:18, SEQ ID NO:20, SEQ ID NO:21, SEQ ID NO:22, SEQ ID NO:23, SEQ ID NO:24, SEQ ID NO:26, SEQ ID NO:27, SEQ ID NO:28, SEQ ID NO:29, SEQ ID NO:30, SEQ ID NO:32, SEQ ID NO:33, SEQ ID NO:34, SEQ ID NO:35, SEQ ID NO:36, SEQ ID NO:38, SEQ ID NO:51, SEQ ID NO:52, SEQ ID NO:57, SEQ ID NO:59, SEQ ID NO:60, SEQ ID NO:61, SEQ ID NO:67, SEQ ID NO:69, SEQ ID NO:70, SEQ ID NO:71, SEQ ID NO:72, SEQ ID NO:76 and/or a nucleic acid molecule encoding a protein comprising amino acid sequence SEQ ID NO:74.
[0014]The present invention also includes a nucleic acid molecule that hybridizes under stringent hybridization conditions with a nucleic acid molecule encoding a protein comprising at least one of the following amino acid sequences: SEQ ID NO:2, SEQ ID NO:5, SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:19, SEQ ID NO:25, SEQ ID NO:31, SEQ ID NO:37, SEQ ID NO:39, SEQ ID NO:40, SEQ ID NO:41, SEQ ID NO:42, SEQ ID NO:43, SEQ ID NO:44, SEQ ID NO:53, SEQ ID NO:54, SEQ ID NO:55, SEQ ID NO:58, SEQ ID NO:68, SEQ ID NO:73 and/or SEQ ID NO:74; and particularly a nucleic acid molecule that hybridizes with a nucleic acid sequence that is a complement of a nucleic acid sequence encoding any of the amino acid sequences. A preferred nucleic acid molecule of the present invention includes a nucleic acid molecule comprising a nucleic acid sequence including SEQ ID NO: 1, SEQ ID NO:3, SEQ ID NO:4, SEQ ID NO:6, SEQ ID NO:7, SEQ ID NO:9, SEQ ID NO:10, SEQ ID NO:12, SEQ ID NO:13, SEQ ID NO:15, SEQ ID NO:16, SEQ ID NO:17, SEQ ID NO:18, SEQ ID NO:20, SEQ ID NO:21, SEQ ID NO:22, SEQ ID NO:23, SEQ ID NO:24, SEQ ID NO:26, SEQ ID NO:27, SEQ ID NO:28, SEQ ID NO:29, SEQ ID NO:30, SEQ ID NO:32, SEQ ID NO:33, SEQ ID NO:34, SEQ ID NO:35, SEQ ID NO:36, SEQ ID NO:38, SEQ ID NO:51, SEQ ID NO:52, SEQ ID NO:57, SEQ ID NO:59, SEQ ID NO:60, SEQ ID NO:61, SEQ ID NO:67, SEQ ID NO:69, SEQ ID NO:70, SEQ ID NO:71, SEQ ID NO:72, SEQ ID NO:76 and/or a nucleic acid molecule encoding a protein comprising amino acid sequence SEQ ID NO:74, and allelic variants thereof.
[0015]The present invention also includes an isolated carboxylesterase nucleic acid molecule comprising a nucleic acid sequence encoding a protein comprising an amino acid sequence including SEQ ID NO:5, SEQ ID NO:19, SEQ ID NO:39, SEQ ID NO:40, SEQ ID NO:41, SEQ ID NO:42, SEQ ID NO:43, SEQ ID NO:44 and/or SEQ ID NO:53. SEQ ID NO:39, SEQ ID NO:40, SEQ ID NO:41, SEQ ID NO:42, SEQ ID NO:43 and SEQ ID NO:44 represent N-terminal amino acid sequences of carboxylesterases isolated from prepupal flea larvae, the production of which are described in the Examples of the present application.
[0016]The present invention also relates to recombinant molecules, recombinant viruses and recombinant cells that include a nucleic acid molecule of the present invention. Also included are methods to produce such nucleic acid molecules, recombinant molecules, recombinant viruses and recombinant cells.
[0017]Another embodiment of the present invention includes an isolated esterase protein that is encoded by a nucleic acid molecule that hybridizes under stringent hybridization conditions to (a) a nucleic acid molecule that includes at least one of the following nucleic acid sequences: SEQ ID NO:3, SEQ ID NO:6, SEQ ID NO:9, SEQ ID NO:12, SEQ ID NO:15, SEQ ID NO:17, SEQ ID NO:20, SEQ ID NO:22, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, SEQ ID NO:38, SEQ ID NO:52, SEQ ID NO:59, SEQ ID NO:61, SEQ ID NO:69, and SEQ ID NO:71; and/or (b) a nucleic acid molecule encoding a protein including at least one of the following amino acid sequences: SEQ ID NO:39, SEQ ID NO:40, SEQ ID NO:41, SEQ ID NO:42, SEQ ID NO:43, SEQ ID NO:44, SEQ ID NO:53, SEQ ID NO:54, SEQ ID NO:55 and SEQ ID NO:74. One embodiment is a carboxylesterase protein encoded by a nucleic acid molecule that hybridizes under stringent hybridization conditions to a nucleic acid molecule that encodes a protein comprising at least one of the following amino acid sequences: SEQ ID NO:5, SEQ ID NO:19, SEQ ID NO:39, SEQ ID NO:40, SEQ ID NO:41, SEQ ID NO:42, SEQ ID NO:43, SEQ ID NO:44 and/or SEQ ID NO:53. Preferred proteins of the present invention are isolated flea proteins including at least one of the following amino acid sequences: SEQ ID NO:2, SEQ ID NO:5, SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:19, SEQ ID NO:25, SEQ ID NO:31, SEQ ID NO:37, SEQ ID NO:39, SEQ ID NO:40, SEQ ID NO:41, SEQ ID NO:42, SEQ ID NO:43, SEQ ID NO:44, SEQ ID NO:53, SEQ ID NO:54, SEQ ID NO:55, SEQ ID NO:58, SEQ ID NO:68, SEQ ID NO:73 and SEQ ID NO:74; also included are proteins encoded by allelic variants of nucleic acid molecules encoding proteins comprising any of the above-listed amino acid sequences.
[0018]The present invention also relates to mimetopes of arthropod esterase proteins as well as to isolated antibodies that selectively bind to arthropod esterase proteins or mimetopes thereof. Also included are methods, including recombinant methods, to produce proteins, mimetopes and antibodies of the present invention.
[0019]The present invention also includes a formulation of flea carboxylesterase proteins, in which the proteins, when submitted to 14% Tris-glycine SDS-PAGE, comprise a fractionation profile as depicted in FIG. 3, in which the proteins have carboxylesterase activity.
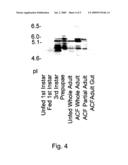
[0020]Also included in the present invention is a formulation of flea carboxylesterase proteins, in which the proteins, when submitted to IEF-PAGE, comprise a fractionation profile as depicted in FIG. 4, lane 3, lane 4, lane 5, lane 6 and/or lane 7, wherein the proteins have carboxylesterase activity.
[0021]Another embodiment of the present invention is an isolated flea protein or a formulation of flea proteins that hydrolyzes α-napthyl acetate to produce α-napthol, when the protein is incubated in the presence of α-napthyl acetate contained in 20 mM Tris at pH 8.0 for about 15 minutes at about 37° C.
[0022]Yet another embodiment of the present invention is an isolated flea protein or a formulation of flea proteins that hydrolyzes the methyl ester group of juvenile hormone to produce a juvenile hormone acid.
[0023]Another embodiment of the present invention is a method to identify a compound capable of inhibiting flea carboxylesterase activity, the method comprising: (a) contacting an isolated flea carboxylesterase with a putative inhibitory compound under conditions in which, in the absence of the compound, the protein has carboxylesterase activity; and (b) determining if the putative inhibitory compound inhibits the activity. Also included in the present invention is a test kit to identify a compound capable of inhibiting flea carboxylesterase activity, the test kit comprising an isolated flea carboxylesterase protein having esterase activity and a means for determining the extent of inhibition of the activity in the presence of a putative inhibitory compound.
[0024]Yet another embodiment of the present invention is a therapeutic composition that is capable of reducing hematophagous ectoparasite infestation. Such a therapeutic composition includes at least one of the following protective compounds: an isolated hematophagous ectoparasite carboxylesterase protein or a mimetope thereof, an isolated carboxylesterase nucleic acid molecule that hybridizes under stringent hybridization conditions with a Ctenocephalides felis carboxylesterase gene, an isolated antibody that selectively binds to a hematophagous ectoparasite carboxylesterase protein, and an inhibitor of carboxylesterase activity identified by its ability to inhibit the activity of a flea carboxylesterase. A therapeutic composition of the present invention can also include an excipient, an adjuvant and/or a carrier. Preferred esterase nucleic acid molecule compounds of the present invention include naked nucleic acid vaccines, recombinant virus vaccines and recombinant cell vaccines. Also included in the present invention is a method to protect an animal from hematophagous ectoparasite infestation, comprising the step of administering to the animal a therapeutic composition of the present invention.
BRIEF DESCRIPTION OF THE FIGURES
[0025]FIG. 1 depicts SDS-PAGE analysis of DFP-labeled esterase proteins.
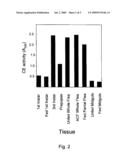
[0026]FIG. 2 depicts carboxylesterase activity of certain esterase proteins of the present invention.
[0027]FIG. 3 depicts SDS-PAGE analysis of carboxylesterase activity of certain esterase proteins of the present invention.
[0028]FIG. 4 depicts IEF analysis of certain esterase proteins of the present invention.
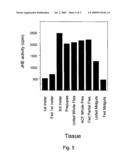
[0029]FIG. 5 depicts juvenile hormone esterase activity of certain esterase proteins of the present invention.
DETAILED DESCRIPTION OF THE INVENTION
[0030]The present invention provides for isolated arthropod esterase proteins, isolated arthropod esterase nucleic acid molecules, antibodies directed against arthropod esterase proteins and other inhibitors of arthropod esterase activity. As used herein, the terms isolated arthropod esterase proteins and isolated arthropod esterase nucleic acid molecules refers to esterase proteins and esterase nucleic acid molecules derived from arthropods and, as such, can be obtained from their natural source or can be produced using, for example, recombinant nucleic acid technology or chemical synthesis. Also included in the present invention is the use of these proteins, nucleic acid molecules, antibodies and inhibitors as therapeutic compositions to protect animals from hematophagous ectoparasite infestation as well as in other applications, such as those disclosed below.
[0031]Arthropod esterase proteins and nucleic acid molecules of the present invention have utility because they represent novel targets for anti-arthropod vaccines and drugs. The products and processes of the present invention are advantageous because they enable the inhibition of arthropod development, metamorphosis, feeding, digestion and reproduction processes that involve esterases. While not being bound by theory, it is believed that expression of arthropod esterase proteins are developmentally regulated, thereby suggesting that esterase proteins are involved in arthropod development and/or reproduction. The present invention is particularly advantageous because the proteins of the present invention were identified in larval fleas, thereby suggesting the importance of the proteins as developmental proteins.
[0032]One embodiment of the present invention is an esterase formulation that includes one or more esterase proteins capable of binding to diisopropylfluorophosphate (DFP). A preferred embodiment of an esterase formulation of the present invention comprises one or more arthropod esterase proteins that range in molecular weight from about 20 kilodaltons (kD) to about 200 kD, more preferably from about 40 kD to about 100 kD, and even more preferably from about 60 kD to about 75 kD, as determined by SDS-PAGE (sodium dodecyl sulfate polyacrylamide gel electrophoresis). An even more preferred formulation includes one or more flea esterase proteins having elution (or migration) patterns as shown in FIG. 1.
[0033]Another embodiment of the present invention is a formulation comprising one or more hematophagous ectoparasite carboxylesterase (CE) proteins. The present invention includes the discovery that such a formulation has general CE activity. General CE activity can be identified using methods known to those of skill in the art and described in the Examples section herein. A suitable formulation of the present invention comprises one or more flea proteins capable of hydrolyzing α-napthyl acetate to produce α-napthyl when the proteins are incubated in the presence of α-napthyl acetate contained in 20 mm Tris at pH 8.0 for about 15 minutes at about 37° C. General CE activity can be identified following such incubation by detecting the production of from about 0.3 to about 2.5 absorbance units in the presence of Fast Blue when measured at 590 nm.
[0034]A preferred CE formulation of the present invention includes one or more flea CE proteins having acidic to neutral isoelectric points, or pI values. An isoelectric pH, or pI, value refers to the pH value at which a molecule has no net electric charge and fails to move in an electric field. A preferred formulation of the present invention includes one or more proteins having a pI value ranging from about pI 2 to about 10, more preferably from about pI 3 to about 8, and even more preferably from about pI 4.7 to about 5.2, as determined by IEF-PAGE.
[0035]An esterase formulation, including a CE formulation, of the present invention can be prepared by a method that includes the steps of: (a) preparing an extract by isolating flea tissue, homogenizing the tissue by sonication and clarifying the extract by centrifugation at a low speed spin, e.g., about 18,000 rpm for about 30 minutes; (b) recovering soluble proteins from said centrifuged extract and applying the proteins to a p-aminobenzamidine agarose bead column; (c) recovering unbound protein from the column and clarifying by filtration; (d) applying the clarified protein to a gel filtration column and eluting and collecting fractions with esterase activity; (e) dilayzing the eluate against 20 mM MES buffer, pH 6.0, containing 10 mM NaCl; (f) applying the dialysate to a cation exchange chromatography column, eluting protein bound to the column with a linear gradient of from about 10 mM NaCl to about 1 M NaCl in 20 mM MES buffer, pH 6, and collecting fractions having esterase activity; (g) adjusting the pH of the resulting fractions to pH 7 and applying the fractions to an anion exchange chromatography column; (h) eluting protein bound to the column with a linear gradient of from about 0 to about 1 M NaCl in 25 mM Tris buffer, pH 6.8 and collecting fractions having esterase activity, such activity elutes from the column at about 170 mM NaCl.
[0036]Tissue can be obtained from unfed fleas or from fleas that recently consumed a blood meal (i.e., blood-fed fleas). Such flea tissues are referred to herein as, respectively, unfed flea and fed flea tissue. Preferred flea tissue from which to obtain an esterase formulation of the present invention includes pre-pupal larval tissue, wandering flea larvae, 3rd instar tissue, fed adult tissue and unfed adult tissue.
[0037]In a preferred embodiment, a CE formulation of the present invention comprises a flea protein comprising amino acid sequence SEQ ID NO:5, SEQ ID NO:19, SEQ ID NO:39, SEQ ID NO:40, SEQ ID NO:41, SEQ ID NO:42, SEQ ID NO:43, SEQ ID NO:44 and/or SEQ ID NO:53.
[0038]Another embodiment of the present invention is a juvenile hormone esterase (JHE) formulation comprising one or more arthropod JHE proteins, the arthropod being of the order Hemiptera, Anoplura, Mallophaga, Diptera, Siphonaptera, Parasitiformes, Acariformes and Acarina. The present invention includes the discovery that such a formulation has JHE activity. JHE activity can be identified using methods known to those of skill in the art and described in the Examples section herein. A suitable formulation of the present invention comprises one or more arthropod proteins capable of hydrolyzing a methyl ester group of juvenile hormone to produce a juvenile hormone acid. Preferably, such a protein is capable of releasing of at least about 120 counts per minute when such a protein is incubated in the presence of 3H-juvenile hormone to create a reaction mixture, the reaction mixture is combined with isooctane, the aqueous phase is recovered and the amount of 3H-juvenile hormone present in that phase is determined. Such a protein is also preferably capable of causing release of methane thiol when such protein is incubated in the presence of methyl 1-heptylthioacetothioate (HEPTAT) using the method generally disclosed in McCutchen et al., Insect Biochem. Molec. Biol., Vol. 25, No. 1, pg 119-126, 1995, which is incorporated in its entirety by this reference.
[0039]In one embodiment, a juvenile hormone esterase formulation of the present invention comprises a protein comprising amino acid sequence SEQ ID NO:74.
[0040]According to the present invention, an arthropod that is not of the order lepidoptera includes an arthropod of the order Hemiptera, Anoplura, Mallophaga, Diptera, Siphonaptera, Parasitiformes, Acariformes and Acarina. Preferred arthropods include Hemiptera cimicidae, Hemiptera reduviidae, Anoplura pediculidae, Anoplura pthiridae, Diptera culicidae, Diptera simuliidae, Diptera psychodidae, Diptera ceratopogonidae, Diptera chaoboridae, Diptera tabanidae, Diptera rhagionidae, athericidae, Diptera chloropidae, Diptera muscidae, Diptera hippoboscidae, Diptera calliphoridae, Diptera sarcophagidae, Diptera oestridae, Diptera gastrophilidae, Diptera cuterebridae, Siphonaptera ceratophyllidae, Siphonaptera leptopsyllidae, Siphonaptera pulicidae, Siphonaptera tungidae, Parasitiformes dermanyssidae, Acariformes tetranychidae, Acariformes cheyletide, Acariformes demodicidae, Acariformes erythraeidae, Acariformes trombiculidae, Acariformes psoroptidae, Acariformes sarcoptidae, Acarina argasidae and Acarina ixodidae. Preferred Diptera muscidae include Musca, Hydrotaea, Stomoxys Haematobia. Preferred Siphonaptera include Ceratophyllidae nosopsyllus, Ceratophyllidae diamanus, Ceratophyllidae ceratophyllus, Leptopsyllidae leptopsylla, Pulicidae pulex, Pulicidae ctenocephalides, Pulicidae xenopsylla, Pulicidae echidnophaga and Tungidae tunga. Preferred Parasitiformes dermanyssidae include Ornithonyssus and Lilponyssoides. Preferred Acarina include Argasidae argas, Argasidae ornithodoros, Argasidae otobius, Ixodidae ixodes, Ixodidae hyalomma, Ixodidae nosomma, Ixodidae rhipicephalus, Ixodidae boophilus, Ixodidae dermacentor, Ixodidae haemaphysalus, Ixodidae amblyomma and Ixodidae anocentor.
[0041]One embodiment of a JHE formulation of the present invention is one or more arthropod JHE proteins that range in molecular weight from about 20 kD to about 200 kD, more preferably from about 40 kD to about 100 kD, and even more preferably from about 60 kD to about 75 kD, as determined by SDS-PAGE (sodium dodecyl sulfate polyacrylamide gel electrophoresis).
[0042]A JHE formulation of the present invention can be prepared by a method that includes the steps of: (a) preparing soluble proteins from arthropod extracts as described above for CE purification and purifying such soluble proteins by gel filtration; (b) collecting fractions having JHE activity from the gel filtration step, loading the fractions onto a cation exchange column, eluting the cation exchange column with a linear gradient of from about 10 mM NaCl to about 1 M NaCl in 20 mM MES buffer, pH 6 and collecting fractions having JHE activity; (c) adjusting the pH of the collected fractions to about pH 7 are dialyzed against about 10 mM phosphate buffer (pH 7.2) containing about 10 mM NaCl; (d) applying the dialysate to a hydroxyapatite column, eluting protein bound to the column with a linear gradient of from about 10 mM phosphate buffer (pH 7.2) containing 10 mM NaCl to about 0.5 M phosphate buffer (pH 6.5) containing 10 mM NaCl and collecting fractions having JHE activity; (e) dialyzing the fractions against 20 mM Tris buffer (pH 8.0) containing 10 mM NaCl; (f) applying the dialysate an anion exchange chromatography column and eluting protein bound to the column with a linear gradient of from about 10 mM to about 1 M NaCl in 20 mM Tris buffer, pH 8 and collecting fractions containing SHE activity.
[0043]A JHE formulation of the present invention can be prepared by a method that includes the steps of: (a) preparing flea extracts as described herein in the Examples section and applying the extract to p-aminobenzamidine linked agarose beads and collecting protein not bound to the beads; (b) applying the unbound protein to a Superdex 200 HR gel filtration column and collecting fractions having JHE activity; (c) applying the fractions to an anion exchange chromatography column, eluting the anion exchange column with a linear gradient of 0 to 1 M NaCl in 25 mM Tris buffer, pH 6.8 and collecting fractions having JHE activity; (d) dialyzing the fractions overnight against about 1 L of 20 mM Tris buffer, pH 8.0, containing 10 mM NaCl; (e) applying the dialysate to a Poros 10 HQ anion exchange column, eluting the column with buffer containing about 120 mM NaCl and collecting fractions having JHE activity.
[0044]Suitable arthropods from which to isolate a JHE formulation of the present invention include, but are not limited to agricultural pests, stored product pests, forest pests, structural pests or animal health pests. Suitable agricultural pests of the present invention include, but are not limited to Colorado potato beetles, corn earworms, fleahoppers, weevils, pink boll worms, cotton aphids, beet armyworms, lygus bugs, hessian flies, sod webworms, whites grubs, diamond back moths, white flies, planthoppers, leafhoppers, mealy bugs, mormon crickets and mole crickets. Suitable stored product pests of the present invention include, but are not limited to dermestids, anobeids, saw toothed grain beetles, indian mealmoths, flour beetles, long-horn wood boring beetles and metallic wood boring beetles. Suitable forest pests of the present invention include, but are not limited to southern pine bark bettles, gypsy moths, elm beetles, ambrosia bettles, bag worms, tent worms and tussock moths. Suitable structural pests of the present invention include, but are not limited to, bess beetles, termites, fire ants, carpenter ants, wasps, hornets, cockroaches, silverfish, Musca domestics and Musca autumnalis. Suitable animal health pests of the present invention include, but are not limited to fleas, ticks, mosquitoes, black flies, lice, true bugs, sand flies, Psychodidae, tsetse flies, sheep blow flies, cattle grub, mites, horn flies, heel flies, deer flies, Culicoides and warble flies. Preferred arthropods from which to isolate a JHE formulation of the present invention include fleas, midges, mosquitoes, sand flies, black flies, horse flies, snipe flies, louse flies, horn flies, deer flies, tsetse flies, buffalo flies, blow flies, stable flies, meiosis-causing flies, biting gnats, lice, mites, bee, wasps, ants, true bugs and ticks, preferably fleas, ticks and blow flies, and more preferably fleas. Preferred fleas from which to isolate JHE proteins include Ctenocephalides, Ceratophyllus, Diamanus, Echidnophaga, Nosopsyllus, Pulex, Tunga, Oropsylla, Orchopeus and Xenopsylla. More preferred fleas include Ctenocephalidesfelis, Ctenocephalides canis, Ceratophyllus pulicidae, Pulex irritans, Oropsylla (Thrassis) bacchi, Oropsylla (Diamanus) montana, Orchopeus howardi, Xenopsylla cheopis and Pulex simulans, with C. felis being even more preferred.
[0045]Suitable tissue from which to isolate a JHE formulation of the present invention includes unfed fleas or fleas that recently consumed a blood meal (i.e., blood-fed fleas). Such flea tissues are referred to herein as, respectively, unfed flea and fed flea tissue. Preferred flea tissue from which to obtain a JHE formulation of the present invention includes pre-pupal larval tissue, 3rd instar tissue, fed or unfed adult tissue, with unfed adult gut tissue being more preferred than fed or unfed whole adult tissue. It is of note that a JHE formulation of the present invention obtained from pre-pupal larval tissue does not hydrolyze α-napthyl acetate.
[0046]Another embodiment of the present invention is an esterase formulation comprising a combination of one or more arthropod CE and JHE proteins of the present invention. Suitable arthropods from which to isolate a combined CE and JHE formulation include those arthropods described herein for the isolation of a JHE formulation of the present invention. Preferred arthropods from which to isolate a combined CE and JHE formulation include fleas, midges, mosquitoes, sand flies, black flies, horse flies, horn flies, deer flies, tsetse flies, buffalo flies, blow flies, stable flies, meiosis-causing flies, biting gnats, lice, bee, wasps, ants, true bugs and ticks, preferably fleas, ticks and blow flies, and more preferably fleas. Suitable flea tissue from which to isolate a combined CE and JHE formulation of the present invention includes 3rd instar tissue, fed or unfed adult tissue and unfed adult tissue, with unfed adult gut tissue being more preferred than fed or unfed whole adult tissue.
[0047]In one embodiment, a formulation of the present invention comprises an esterase having both CE and JHE activity. Preferably, a formulation of the present invention that comprises an esterase having both CE and JHE activity comprises a flea protein comprising amino acid sequence SEQ ID NO:8 and/or SEQ ID NO:37.
[0048]Another embodiment of the present invention is an isolated protein comprising an arthropod esterase protein. It is to be noted that the term "a" or "an" entity refers to one or more of that entity; for example, a protein refers to one or more proteins or at least one protein. As such, the terms "a" (or "an"), "one or more" and "at least one" can be used interchangeably herein. It is also to be noted that the terms "comprising", "including", and "having" can be used interchangeably. Furthermore, a compound "selected from the group consisting of" refers to one or more of the compounds in the list that follows, including mixtures (i.e., combinations) of two or more of the compounds. According to the present invention, an isolated, or biologically pure, protein, is a protein that has been removed from its natural milieu. As such, "isolated" and "biologically pure" do not necessarily reflect the extent to which the protein has been purified. An isolated protein of the present invention can be obtained from its natural source, can be produced using recombinant DNA technology or can be produced by chemical synthesis.
[0049]As used herein, an isolated arthropod esterase protein can be a full-length protein or any homolog of such a protein. An isolated protein of the present invention, including a homolog, can be identified in a straight-forward manner by the protein's ability to elicit an immune response against arthropod esterase proteins, to hydrolyze α-napthyl acetate, to hydrolyze the methyl ester group of juvenile hormone or bind to DFP. Esterase proteins of the present invention include CE and JHE proteins. As such, an esterase protein of the present invention can comprise a protein capable of hydrolyzing α-napthyl acetate, hydrolyzing the methyl ester group of juvenile hormone and/or binding to DFP. Examples of esterase homologs include esterase proteins in which amino acids have been deleted (e.g., a truncated version of the protein, such as a peptide), inserted, inverted, substituted and/or derivatized (e.g., by glycosylation, phosphorylation, acetylation, myristoylation, prenylation, palmitoylation, amidation and/or addition of glycerophosphatidyl inositol) such that the homolog includes at least one epitope capable of eliciting an immune response against an arthropod esterase protein. That is, when the homolog is administered to an animal as an immunogen, using techniques known to those skilled in the art, the animal will produce an immune response against at least one epitope of a natural arthropod esterase protein. The ability of a protein to effect an immune response, can be measured using techniques known to those skilled in the art. Esterase protein homologs of the present invention also include esterase proteins that hydrolyze α-napthyl acetate and/or that hydrolyze the methyl ester group of juvenile hormone.
[0050]Arthropod esterase protein homologs can be the result of natural allelic variation or natural mutation. Esterase protein homologs of the present invention can also be produced using techniques known in the art including, but not limited to, direct modifications to the protein or modifications to the gene encoding the protein using, for example, classic or recombinant nucleic acid techniques to effect random or targeted mutagenesis.
[0051]Isolated esterase proteins of the present invention have the further characteristic of being encoded by nucleic acid molecules that hybridize under stringent hybridization conditions to a gene encoding a Ctenocephalides felis protein (i.e., a C. felis esterase gene). As used herein, stringent hybridization conditions refer to standard hybridization conditions under which nucleic acid molecules, including oligonucleotides, are used to identify similar nucleic acid molecules. Such standard conditions are disclosed, for example, in Sambrook et al., Molecular Cloning: A Laboratory Manual, Cold Spring Harbor Labs Press, 1989; Sambrook et al., ibid., is incorporated by reference herein in its entirety. Stringent hybridization conditions typically permit isolation of nucleic acid molecules having at least about 70% nucleic acid sequence identity with the nucleic acid molecule being used to probe in the hybridization reaction. Formulae to calculate the appropriate hybridization and wash conditions to achieve hybridization permitting 30% or less mismatch of nucleotides are disclosed, for example, in Meinkoth et al., 1984, Anal. Biochem. 138, 267-284; Meinkoth et al., ibid., is incorporated by reference herein in its entirety.
[0052]As used herein, a C. felis esterase gene includes all nucleic acid sequences related to a natural C. felis esterase gene such as regulatory regions that control production of the C. felis esterase protein encoded by that gene (such as, but not limited to, transcription, translation or post-translation control regions) as well as the coding region itself. In one embodiment, a C. felis esterase gene of the present invention includes the nucleic acid sequence SEQ ID NO:1, SEQ ID NO:3, SEQ ID NO:4, SEQ ID NO:6, SEQ ID NO:7, SEQ ID NO:9, SEQ ID NO:1, SEQ ID NO:12, SEQ ID NO:13, SEQ ID NO:15, SEQ ID NO:16, SEQ ID NO:17, SEQ ID NO:18, SEQ ID NO:20, SEQ ID NO:21, SEQ ID NO:22, SEQ ID NO:23, SEQ ID NO:24, SEQ ID NO:26, SEQ ID NO:27, SEQ ID NO:28, SEQ ID NO:29, SEQ ID NO:30, SEQ ID NO:32, SEQ ID NO:33, SEQ ID NO:34, SEQ ID NO:35, SEQ ID NO:36, SEQ ID NO:38, SEQ ID NO:51, SEQ ID NO:52, SEQ ID NO:57, SEQ ID NO:59, SEQ ID NO:60, SEQ ID NO:61, SEQ ID NO:67, SEQ ID NO:69, SEQ ID NO:70, SEQ ID NO:71, SEQ ID NO:72, SEQ ID NO:76 and/or a nucleic acid molecule encoding a protein comprising amino acid sequence SEQ ID NO:74. Nucleic acid sequence SEQ ID NO:1 represents the deduced sequence of the coding strand of a PCR amplified nucleic acid molecule denoted herein as nfE1401, the production of which is disclosed in the Examples. The complement of SEQ ID NO:1 (represented herein by SEQ ID NO:3) refers to the nucleic acid sequence of the strand complementary to the strand having SEQ ID NO:1, which can easily be determined by those skilled in the art. Likewise, a nucleic acid sequence complement of any nucleic acid sequence of the present invention refers to the nucleic acid sequence of the nucleic acid strand that is complementary to (i.e., can form a complete double helix with) the strand for which the sequence is cited.
[0053]Nucleic acid sequence SEQ ID NO:4 represents the deduced sequence of the coding strand of a PCR amplified nucleic acid molecule denoted herein as nfE2364, the production of which is disclosed in the Examples. The complement of SEQ ID NO:4 is represented herein by SEQ ID NO:6.
[0054]Nucleic acid sequence SEQ ID NO:7 represents the deduced sequence of the coding strand of a PCR amplified nucleic acid molecule denoted herein as nfE3421, the production of which is disclosed in the Examples. The complement of SEQ ID NO:7 is represented herein by SEQ ID NO:9.
[0055]Nucleic acid sequence SEQ ID NO:10 represents the deduced sequence of the coding strand of a PCR amplified nucleic acid molecule denoted herein as nfE4524, the production of which is disclosed in the Examples. The complement of SEQ ID NO:10 is represented herein by SEQ ID NO:12.
[0056]Nucleic acid sequence SEQ ID NO:13 represents the deduced sequence of the coding strand of an apparent coding region of a complementary DNA (cDNA) nucleic acid molecule denoted herein as nfE51982, the production of which is disclosed in the Examples. The complement of SEQ ID NO:13 is represented herein by SEQ ID NO:15.
[0057]Nucleic acid sequence SEQ ID NO:18 represents the deduced sequence of the coding strand of an apparent coding region of a cDNA nucleic acid molecule denoted herein as nfE61792, the production of which is disclosed in the Examples. The complement of SEQ ID NO:18 is represented herein by SEQ ID NO:20.
[0058]Nucleic acid sequence SEQ ID NO:24 represents the deduced sequence of the coding strand of an apparent coding region of a cDNA nucleic acid molecule denoted herein as nfE72836, the production of which is disclosed in the Examples. The complement of SEQ ID NO:24 is represented herein by SEQ ID NO:26.
[0059]Nucleic acid sequence SEQ ID NO:30 represents the deduced sequence of the coding strand of an apparent coding region of a cDNA nucleic acid molecule denoted herein as nfE82801, the production of which is disclosed in the Examples. The complement of SEQ ID NO:30 is represented herein by SEQ ID NO:32.
[0060]Nucleic acid sequence SEQ ID NO:36 represents the deduced sequence of the coding strand of an apparent coding region of a cDNA nucleic acid molecule denoted herein as nfE92007, the production of which is disclosed in the Examples. The complement of SEQ ID NO:36 is represented herein by SEQ ID NO:38.
[0061]Nucleic acid sequence SEQ ID NO:57 represents the deduced sequence of the coding strand of an apparent coding region of a cDNA nucleic acid molecule denoted herein as nfE52144, the production of which is disclosed in the Examples. The complement of SEQ ID NO:57 is represented herein by SEQ ID NO:59.
[0062]Nucleic acid sequence SEQ ID NO:67 represents the deduced sequence of the coding strand of an apparent coding region of a cDNA nucleic acid molecule denoted herein as nfE101987, the production of which is disclosed in the Examples. The complement of SEQ ID NO:67 is represented herein by SEQ ID NO:69.
[0063]It should be noted that since nucleic acid sequencing technology is not entirely error-free, the nucleic acid sequences and amino acid sequences presented herein represent, respectively, apparent nucleic acid sequences of nucleic acid molecules of the present invention and apparent amino acid sequences of esterase proteins of the present invention.
[0064]In another embodiment, a C. felis esterase gene can be an allelic variant that includes a similar but not identical sequence to SEQ ID NO:1, SEQ ID NO:3, SEQ ID NO:4, SEQ ID NO:6, SEQ ID NO:7, SEQ ID NO:9, SEQ ID NO:10, SEQ ID NO:12, SEQ ID NO:13, SEQ ID NO:15, SEQ ID NO:16, SEQ ID NO:17, SEQ ID NO:18, SEQ ID NO:20, SEQ ID NO:21, SEQ ID NO:22, SEQ ID NO:23, SEQ ID NO:24, SEQ ID NO:26, SEQ ID NO:27, SEQ ID NO:28, SEQ ID NO:29, SEQ ID NO:30, SEQ ID NO:32, SEQ ID NO:33, SEQ ID NO:34, SEQ ID NO:35, SEQ ID NO:36, SEQ ID NO:38, SEQ ID NO:51, SEQ ID NO:52, SEQ ID NO:57, SEQ ID NO:59, SEQ ID NO:60, SEQ ID NO:61, SEQ ID NO:67, SEQ ID NO:69, SEQ ID NO:70, SEQ ID NO:71, SEQ ID NO:72, SEQ ID NO:76 and/or a nucleic acid molecule encoding a protein comprising amino acid sequence SEQ ID NO:74. An allelic variant of a C. felis esterase gene is a gene that occurs at essentially the same locus (or loci) in the genome as the gene including SEQ ID NO:1, SEQ ID NO:3, SEQ ID NO:4, SEQ ID NO:6, SEQ ID NO:7, SEQ ID NO:9, SEQ ID NO:10, SEQ ID NO:12, SEQ ID NO:13, SEQ ID NO:15, SEQ ID NO:16, SEQ ID NO:17, SEQ ID NO:18, SEQ ID NO:20, SEQ ID NO:21, SEQ ID NO:22, SEQ ID NO:23, SEQ ID NO:24, SEQ ID NO:26, SEQ ID NO:27, SEQ ID NO:28, SEQ ID NO:29, SEQ ID NO:30, SEQ ID NO:32, SEQ ID NO:33, SEQ ID NO:34, SEQ ID NO:35, SEQ ID NO:36, SEQ ID NO:38, SEQ ID NO:51, SEQ ID NO:52, SEQ ID NO:57, SEQ ID NO:59, SEQ ID NO:60, SEQ ID NO:61, SEQ ID NO:67, SEQ ID NO:69, SEQ ID NO:70, SEQ ID NO:71, SEQ ID NO:72, SEQ ID NO:76 and/or a nucleic acid molecule encoding a protein comprising amino acid sequence SEQ ID NO:74, but which, due to natural variations caused by, for example, mutation or recombination, has a similar but not identical sequence. Allelic variants typically encode proteins having similar activity to that of the protein encoded by the gene to which they are being compared. Allelic variants can also comprise alterations in the 5' or 3' untranslated regions of the gene (e.g., in regulatory control regions). Allelic variants are well known to those skilled in the art and would be expected to be found within a given arthropod since the genome is diploid and/or among a group of two or more arthropods.
[0065]The minimal size of an esterase protein homolog of the present invention is a size sufficient to be encoded by a nucleic acid molecule capable of forming a stable hybrid (i.e., hybridize under stringent hybridization conditions) with the complementary sequence of a nucleic acid molecule encoding the corresponding natural protein. As such, the size of the nucleic acid molecule encoding such a protein homolog is dependent on nucleic acid composition and percent homology between the nucleic acid molecule and complementary sequence. It should also be noted that the extent of homology required to form a stable hybrid can vary depending on whether the homologous sequences are interspersed throughout the nucleic acid molecules or are clustered (i.e., localized) in distinct regions on the nucleic acid molecules. The minimal size of such nucleic acid molecules is typically at least about 12 to about 15 nucleotides in length if the nucleic acid molecules are GC-rich and at least about 15 to about 17 bases in length if they are AT-rich. As such, the minimal size of a nucleic acid molecule used to encode an esterase protein homolog of the present invention is from about 12 to about 18 nucleotides in length. Thus, the minimal size of an esterase protein homolog of the present invention is from about 4 to about 6 amino acids in length. There is no limit, other than a practical limit, on the maximal size of such a nucleic acid molecule in that the nucleic acid molecule can include a portion of a gene, an entire gene, multiple genes, or portions thereof. The preferred size of a protein encoded by a nucleic acid molecule of the present invention depends on whether a full-length, fusion, multivalent, or functional portion of such a protein is desired.
[0066]One embodiment of the present invention includes an arthropod esterase protein having CE enzyme activity. Such a CE protein preferably includes: a catalytic triad of serine--histidine--glutamic acid as well as the essential amino acids arginine and aspartic acid at positions similar to those described for juvenile hormone esterase, for example by Ward et al., 1992, Int J Biochem 24: 1933-1941; this reference is incorporated by reference herein in its entirety. Analysis of the apparent full-length protein sequences disclosed herein indicates that each of these amino acid sequences includes these amino acid motifs, as well as surrounding consensus sequences.
[0067]Suitable arthropods from which to isolate esterase proteins having general CE activity of the present invention (including isolation of the natural protein or production of the protein by recombinant or synthetic techniques) preferably include insects and acarines but not Culicidae, Drosophilidae, Calliphoridae, Sphingidae, Lymantriidae, Noctuidae, Fulgoroidae and Aphididae. Preferred arthropods from which to isolate CE proteins having general CE activity include fleas, ticks, black flies, lice, true bugs, sand flies, Psychodidae, tsetse flies, cattle grub, mites, horn flies, heel flies, deer flies, Culicoides and warble flies. Preferred arthropods from which to isolate an esterase proteins having general CE activity include fleas, midges, sand flies, black flies, horse flies, snipe flies, louse flies, horn flies, deer flies, tsetse flies, buffalo flies, blow flies, stable flies, meiosis-causing flies, biting gnats, lice, mites, bee, wasps, ants, true bugs and ticks, preferably fleas, ticks and blow flies, and more preferably fleas. Preferred fleas from which to isolate esterase proteins having general CE activity include Ctenocephalides, Ceratophyllus, Diamanus, Echidnophaga, Nosopsyllus, Pulex, Tunga, Oropsylla, Orchopeus and Xenopsylla. More preferred fleas include Ctenocephalides felis, Ctenocephalides canis, Ceratophyllus pulicidae, Pulex irritans, Oropsylla (Thrassis) bacchi, Oropsylla (Diamanus) montana, Orchopeus howardi, Xenopsylla cheopis and Pulex simulans, with C. felis being even more preferred.
[0068]A preferred arthropod esterase protein of the present invention is a compound that when administered to an animal in an effective manner, is capable of protecting that animal from hematophagous ectoparasite infestation. In accordance with the present invention, the ability of an esterase protein of the present invention to protect an animal from hematophagous ectoparasite infestation refers to the ability of that protein to, for example, treat, ameliorate and/or prevent infestation caused by hematophagous arthropods. In particular, the phrase "to protect an animal from hematophagous ectoparasite infestation" refers to reducing the potential for hematophagous ectoparasite population expansion on and around the animal (i.e., reducing the hematophagous ectoparasite burden). Preferably, the hematophagous ectoparasite population size is decreased, optimally to an extent that the animal is no longer bothered by hematophagous ectoparasites. A host animal, as used herein, is an animal from which hematophagous ectoparasites can feed by attaching to and feeding through the skin of the animal. Hematophagous ectoparasites, and other ectoparasites, can live on a host animal for an extended period of time or can attach temporarily to an animal in order to feed. At any given time, a certain percentage of a hematophagous ectoparasite population can be on a host animal whereas the remainder can be in the environment of the animal. Such an environment can include not only adult hematophagous ectoparasites, but also hematophagous ectoparasite eggs and/or hematophagous ectoparasite larvae. The environment can be of any size such that hematophagous ectoparasites in the environment are able to jump onto and off of a host animal. For example, the environment of an animal can include plants, such as crops, from which hematophagous ectoparasites infest an animal. As such, it is desirable not only to reduce the hematophagous ectoparasite burden on an animal per se, but also to reduce the hematophagous ectoparasite burden in the environment of the animal. In one embodiment, an esterase protein of the present invention can elicit an immune response (including a humoral and/or cellular immune response) against a hematophagous ectoparasite.
[0069]Suitable hematophagous ectoparasites to target include any hematophagous ectoparasite that is essentially incapable of infesting an animal administered an esterase protein of the present invention. As such, a hematophagous ectoparasite to target includes any hernatophagous ectoparasite that produces a protein having one or more epitopes that can be targeted by a humoral and/or cellular immune response against an esterase protein of the present invention and/or that can be targeted by a compound that otherwise inhibits esterase activity (e.g., a compound that inhibits hydrolysis of α-napthyl acetate, hydrolysis of the methyl ester group of juvenile hormone, and/or binds to DFP), thereby resulting in the decreased ability of the hematophagous ectoparasite to infest an animal. Preferred hematophagous ectoparasite to target include ectoparasites disclosed herein as being useful in the production of esterase proteins of the present invention.
[0070]The present invention also includes mimetopes of esterase proteins of the present invention. As used herein, a mimetope of an esterase protein of the present invention refers to any compound that is able to mimic the activity of such a protein (e.g., ability to elicit an immune response against an arthropod esterase protein of the present invention and/or ability to inhibit esterase activity), often because the mimetope has a structure that mimics the esterase protein. It is to be noted, however, that the mimetope need not have a structure similar to an esterase protein as long as the mimetope functionally mimics the protein. Mimetopes can be, but are not limited to: peptides that have been modified to decrease their susceptibility to degradation; anti-idiotypic and/or catalytic antibodies, or fragments thereof; non-proteinaceous immunogenic portions of an isolated protein (e.g., carbohydrate structures); synthetic or natural organic or inorganic molecules, including nucleic acids; and/or any other peptidomimetic compounds. Mimetopes of the present invention can be designed using computer-generated structures of esterase proteins of the present invention. Mimetopes can also be obtained by generating random samples of molecules, such as oligonucleotides, peptides or other organic molecules, and screening such samples by affinity chromatography techniques using the corresponding binding partner, (e.g., an esterase substrate, an esterase substrate analog, or an anti-esterase antibody). A preferred mimetope is a peptidomimetic compound that is structurally and/or functionally similar to an esterase protein of the present invention, particularly to the active site of the esterase protein.
[0071]The present invention also includes mimetopes of esterase proteins of the present invention. As used herein, a mimetope of an esterase protein of the present invention refers to any compound that is able to mimic the activity of such an esterase protein, often because the mimetope has a structure that mimics the esterase protein. Mimetopes can be, but are not limited to: peptides that have been modified to decrease their susceptibility to degradation; anti-idiotypic and/or catalytic antibodies, or fragments thereof; non-proteinaceous immunogenic portions of an isolated protein (e.g., carbohydrate structures); and synthetic or natural organic molecules, including nucleic acids. Such mimetopes can be designed using computer-generated structures of proteins of the present invention. Mimetopes can also be obtained by generating random samples of molecules, such as oligonucleotides, peptides or other organic molecules, and screening such samples by affinity chromatography techniques using the corresponding binding partner.
[0072]One embodiment of an arthropod esterase protein of the present invention is a fusion protein that includes an arthropod esterase protein-containing domain attached to one or more fusion segments. Suitable fusion segments for use with the present invention include, but are not limited to, segments that can: enhance a protein's stability; act as an immunopotentiator to enhance an immune response against an esterase protein; and/or assist purification of an esterase protein (e.g., by affinity chromatography). A suitable fusion segment can be a domain of any size that has the desired function (e.g., imparts increased stability, imparts increased immunogenicity to a protein, and/or simplifies purification of a protein). Fusion segments can be joined to amino and/or carboxyl termini of the esterase-containing domain of the protein and can be susceptible to cleavage in order to enable straight-forward recovery of an esterase protein. Fusion proteins are preferably produced by culturing a recombinant cell transformed with a fusion nucleic acid molecule that encodes a protein including the fusion segment attached to either the carboxyl and/or amino terminal end of an esterase-containing domain. Preferred fusion segments include a metal binding domain (e.g., a poly-histidine segment); an immunoglobulin binding domain (e.g., Protein A; Protein G; T cell; B cell; Fc receptor or complement protein antibody-binding domains); a sugar binding domain (e.g., a maltose binding domain); and/or a "tag" domain (e.g., at least a portion of β-galactosidase, a strep tag peptide, other domains that can be purified using compounds that bind to the domain, such as monoclonal antibodies). More preferred fusion segments include metal binding domains, such as a poly-histidine segment; a maltose binding domain; a strep tag peptide, such as that available from Biometra in Tampa, Fla.; and an S10 peptide. Examples of particularly preferred fusion proteins of the present invention include PHIS-PfE6540, PHIS-PfE7275, PHIS-PfE7570, PHIS-PfE8570 and PHIS-PfE9528, production of which are disclosed herein.
[0073]In another embodiment, an arthropod esterase protein of the present invention also includes at least one additional protein segment that is capable of protecting an animal from hematophagous ectoparasite infestations. Such a multivalent protective protein can be produced by culturing a cell transformed with a nucleic acid molecule comprising two or more nucleic acid domains joined together in such a manner that the resulting nucleic acid molecule is expressed as a multivalent protective compound containing at least two protective compounds, or portions thereof, capable of protecting an animal from hematophagous ectoparasite infestation by, for example, targeting two different arthropod proteins.
[0074]Examples of multivalent protective compounds include, but are not limited to, an esterase protein of the present invention attached to one or more compounds protective against one or more arthropod compounds. Preferred second compounds are proteinaceous compounds that effect active immunization (e.g., antigen vaccines), passive immunization (e.g., antibodies), or that otherwise inhibit a arthropod activity that when inhibited can reduce hematophagous ectoparasite burden on and around an animal. Examples of second compounds include a compound that inhibits binding between an arthropod protein and its ligand (e.g., a compound that inhibits flea ATPase activity or a compound that inhibits binding of a peptide or steroid hormone to its receptor), a compound that inhibits hormone (including peptide or steroid hormone) synthesis, a compound that inhibits vitellogenesis (including production of vitellin and/or transport and maturation thereof into a major egg yolk protein), a compound that inhibits fat body function, a compound that inhibits muscle action, a compound that inhibits the nervous system, a compound that inhibits the immune system and/or a compound that inhibits hematophagous ectoparasite feeding. Examples of second compounds also include proteins obtained from different stages of hematophagous ectoparasite development. Particular examples of second compounds include, but are not limited to, serine proteases, cysteine proteases, aminopeptidases, serine protease inhibitor proteins, calreticulins, larval serum proteins and echdysone receptors, as well as antibodies to and inhibitors of such proteins. In one embodiment, an arthropod esterase protein of the present invention is attached to one or more additional compounds protective against hematophagous ectoparasite infestation. In another embodiment, one or more protective compounds, such as those listed above, can be included in a multivalent vaccine comprising an arthropod esterase protein of the present invention and one or more other protective molecules as separate compounds.
[0075]A preferred isolated protein of the present invention is a protein encoded by a nucleic acid molecule that hybridizes under stringent hybridization conditions with nucleic acid molecules nfE1401, nfE2364, nfE3421, nfE4524, nfE51982, nfE51515, nfE52144, nfE51650, nfE61488, nfE61792, nfE61650, nfE72836, nfE71788, nfE71710, nfE7650, nfE82801, nfE81785, nfE81710, nfE92007, nfE91584, nfE91540, nfE101987 and/or nfE101590. A further preferred isolated protein is encoded by a nucleic acid molecule that hybridizes under stringent hybridization conditions with a nucleic acid molecule having nucleic acid sequence SEQ ID NO:3, SEQ ID NO:6, SEQ ID NO:9, SEQ ID NO:12, SEQ ID NO:15, SEQ ID NO: 17, SEQ ID NO:20, SEQ ID NO:22, SEQ ID NO:26, SEQ ID NO:29, SEQ ID NO:32, SEQ ID NO:35, SEQ ID NO:38, SEQ ID NO:52, SEQ ID NO:59, SEQ ID NO:61, SEQ ID NO:69 and/or SEQ ID NO:71.
[0076]Translation of SEQ ID NO:1 suggests that nucleic acid molecule nfE1401 encodes a non-full-length arthropod esterase protein of about 103 amino acids, referred to herein as PfE1103, represented by SEQ ID NO:2, assuming the first codon spans from nucleotide 92 through nucleotide 94 of SEQ ID NO:1.
[0077]Comparison of amino acid sequence SEQ ID NO:2 (i.e., the amino acid sequence of PfE1103) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:2, showed the most homology, i.e., about 33% identity, between SEQ ID NO:2 and alpha esterase protein from Drosophila melanogaster.
[0078]Translation of SEQ ID NO:4 suggests that nucleic acid molecule nfE2364 encodes a non-full-length arthropod esterase protein of about 121 amino acids, referred to herein as PfE2121, represented by SEQ ID NO:5, assuming the first codon spans from nucleotide 2 through nucleotide 4 of SEQ ID NO:4.
[0079]Comparison of amino acid sequence SEQ ID NO:5 (i.e., the amino acid sequence of PfE2121) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:5, showed the most homology, i.e., about 38% identity, between SEQ ID NO:5 and alpha esterase protein from Drosophila melanogaster.
[0080]Translation of SEQ ID NO:7 suggests that nucleic acid molecule nfE3421 encodes a non-full-length arthropod esterase protein of about 103 amino acids, referred to herein as PfE3103, represented by SEQ ID NO:8, assuming the first codon spans from nucleotide 113 through nucleotide 115 of SEQ ID NO:7.
[0081]Comparison of amino acid sequence SEQ ID NO:8 (i.e., the amino acid sequence of PfE3103) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:8, showed the most homology, i.e., about 39% identity, between SEQ ID NO:8 and alpha esterase protein from Drosophila melanogaster.
[0082]Translation of SEQ ID NO:10 suggests that nucleic acid molecule nfE4524 encodes a non-full-length arthropod esterase protein of about 137 amino acids, referred to herein as PfF4137, represented by SEQ ID NO:11, assuming the first codon spans from nucleotide 113 through nucleotide 115 of SEQ ID NO:10.
[0083]Comparison of amino acid sequence SEQ ID NO:11 (i.e., the amino acid sequence of PfE4137) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:11, showed the most homology, i.e., about 30% identity, between SEQ ID NO:11 and Leptinotarsa decemlineata acetylcholinesterase.
[0084]Translation of SEQ ID NO:57 suggests that nucleic acid molecule nfE52144 encodes a full-length arthropod esterase protein of about 550 amino acids, referred to herein as PfE5550, represented by SEQ ID NO:58, assuming an open reading frame in which the initiation codon spans from nucleotide 30 through nucleotide 32 of SEQ ID NO:57 and the termination (stop) codon spans from nucleotide 1680 through nucleotide 1682 of SEQ ID NO:57. The complement of SEQ ID NO:57 is represented herein by SEQ ID NO:59. The coding region encoding PfE5550 is represented by the nucleic acid molecule nfE51650, having a coding strand with the nucleic acid sequence represented by SEQ ID NO:60 and a complementary strand with nucleic acid sequence SEQ ID NO:61. The deduced amino acid sequence of PfE5550 (i.e., SEQ ID NO:58) predicts that PfE5550 has an estimated molecular weight of about 61.8 kD and an estimated pI of about 5.5.
[0085]Comparison of amino acid sequence SEQ ID NO:58 (i.e., the amino acid sequence of PfE5550) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:58 showed the most homology, i.e., about 36% identity between SEQ ID NO:58 and Drosophila melanogaster alpha esterase protein.
[0086]Translation of SEQ ID NO:18 suggests that nucleic acid molecule nfE61792 encodes a full-length arthropod esterase protein of about 550 amino acids, referred to herein as PfE6550, represented by SEQ ID NO:19, assuming an open reading frame having an initiation codon spanning from nucleotide 49 through nucleotide 51 of SEQ ID NO:18 and a stop codon spanning from nucleotide 1699 through nucleotide 1701 of SEQ ID NO:18. The coding region encoding PfE6550, is represented by nucleic acid molecule nfE61650, having a coding strand with the nucleic acid sequence represented by SEQ ID NO:21 and a complementary strand with nucleic acid sequence SEQ ID NO:22. The proposed mature protein, denoted herein as PfE6530, contains about 530 amino acids which is represented herein as SEQ ID NO:53. The nucleic acid molecule encoding PfE6530 is denoted herein as nfE61590 and has a coding strand having the nucleic acid sequence SEQ ID NO:23. The deduced amino acid sequence SEQ ID NO:19 suggests a protein having a molecular weight of about 61.8 kD and an estimated pI of about 5.5.
[0087]Comparison of amino acid sequence SEQ ID NO:19 (i.e., the amino acid sequence of PfE6550) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:19 showed the most homology, i.e., about 28% identity between SEQ ID NO:19 and Drosophila melanogaster alpha esterase protein.
[0088]Translation of SEQ ID NO:24 suggests that nucleic acid molecule nfE72836 encodes a full-length arthropod esterase protein of about 596 amino acids, referred to herein as PfE7596, represented by SEQ ID NO:25, assuming an open reading frame having an initiation codon spanning from nucleotide 99 through nucleotide 101 of SEQ ID NO:24 and a stop codon spanning from nucleotide 1887 through nucleotide 1889 of SEQ ID NO:24. The coding region encoding PfE7596, is represented by nucleic acid molecule nfE71788, having a coding strand with the nucleic acid sequence represented by SEQ ID NO:28 and a complementary strand with nucleic acid sequence SEQ ID NO:29. The proposed mature protein, denoted herein as PfE7570, contains about 570 amino acids which is represented herein as SEQ ID NO:54. The nucleic acid molecule encoding PfE7570 is denoted herein as nfE71710 and has a coding strand having the nucleic acid sequence SEQ ID NO:27. The deduced amino acid sequence SEQ ID NO:25 suggests a protein having a molecular weight of about 68.7 kD and an estimated pI of about 6.1.
[0089]Comparison of amino acid sequence SEQ ID NO:25 (i.e., the amino acid sequence of PfE7596) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:25 showed the most homology, i.e., about 27% identity between SEQ ID NO:25 and Drosophila melanogaster alpha esterase protein.
[0090]Translation of SEQ ID NO:30 suggests that nucleic acid molecule nfE82801 encodes a full-length arthropod esterase protein of about 595 amino acids, referred to herein as PfE8595, represented by SEQ ID NO:31, assuming an open reading frame having an initiation codon spanning from nucleotide 99 through nucleotide 101 of SEQ ID NO:30 and a stop codon spanning from nucleotide 1884 through nucleotide 1886 of SEQ ID NO:30. The coding region encoding PfE8595, is represented by nucleic acid molecule nfE81785, having a coding strand with the nucleic acid sequence represented by SEQ ID NO:34 and a complementary strand with nucleic acid sequence SEQ ID NO:35. The proposed mature protein, denoted herein as PfE8570, contains about 570 amino acids which is represented herein as SEQ ID NO:55. The nucleic acid molecule encoding PfE8570 is denoted herein as nfE81710 and has a coding strand having the nucleic acid sequence SEQ ID NO:33. The deduced amino acid sequence SEQ ID NO:31 suggests a protein having a molecular weight of about 68.6 kD and an estimated pI of about 6.1.
[0091]Comparison of amino acid sequence SEQ ID NO:31 (i.e., the amino acid sequence of PfE8595) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:31 showed the most homology, i.e., about 28% identity between SEQ ID NO:31 and estalpha-2 esterase of Culex pipiens quinquefasciatus.
[0092]Translation of SEQ ID NO:36 suggests that nucleic acid molecule nfE92007 encodes a full-length arthropod esterase protein of about 528 amino acids, referred to herein as PfE9528, represented by SEQ ID NO:37, assuming an open reading frame having an initiation codon spanning from nucleotide 11 through nucleotide 13 of SEQ ID NO:36 and a stop codon spanning from nucleotide 1595 through nucleotide 1597 of SEQ ID NO:36. The coding region encoding PfE9528, is represented by nucleic acid molecule nfE91584, having a coding strand with the nucleic acid sequence represented by SEQ ID NO:51 and a complementary strand with nucleic acid sequence SEQ ID NO:52. The deduced amino acid sequence SEQ ID NO:37 suggests a protein having a molecular weight of about 60 kD and an estimated pI of about 5.43.
[0093]Comparison of amino acid sequence SEQ ID NO:37 (i.e., the amino acid sequence of PfE9528) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:37 showed the most homology, i.e., about 37% identity between SEQ ID NO:37 and alpha esterase protein from Drosophila melanogaster.
[0094]Translation of SEQ ID NO:67 suggests that nucleic acid molecule nfE101987 encodes a full-length flea esterase protein of about 530 amino acids, referred to herein as PfE10530, having amino acid sequence SEQ ID NO:68, assuming an open reading frame in which the initiation codon spans from nucleotide 231 through nucleotide 233 of SEQ ID NO:67 and a stop codon spanning from nucleotide 1821 through nucleotide 1823 of SEQ ID NO:67. The complement of SEQ ID NO:67 is represented herein by SEQ ID NO:69. The coding region encoding PfE10530, is represented by nucleic acid molecule nfE101590, having a coding strand with the nucleic acid sequence represented by SEQ ID NO:70 and a complementary strand with nucleic acid sequence SEQ ID NO:71. The amino acid sequence of PfE10530 (i.e., SEQ ID NO:68) predicts that PfE10530 has an estimated molecular weight of about 59.5 kD and an estimated pI of about 5.5.
[0095]Comparison of amino acid sequence SEQ ID NO:68 (i.e., the amino acid sequence of PfE10530) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:68 showed the most homology, i.e., about 30% identity between SEQ ID NO:68 and Culex pipens esterase b1 precurser protein (SWISSPROT® # P16854).
[0096]More preferred arthropod esterase proteins of the present invention include proteins comprising amino acid sequences that are at least about 40%, preferably at least about 45%, more preferably at least about 50%, even more preferably at least about 55%, even more preferably at least about 60%, even more preferably at least about 70%, even more preferably at least about 80%, even more preferably at least about 90%, and even more preferably at least about 95%, identical to amino acid sequence SEQ ID NO:2, SEQ ID NO:5, SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:19, SEQ ID NO:25, SEQ ID NO:31, SEQ ID NO:37, SEQ ID NO:39, SEQ ID NO:40, SEQ ID NO:41, SEQ ID NO:42, SEQ ID NO:43, SEQ ID NO:44, SEQ ID NO:53, SEQ ID NO:54, SEQ ID NO:55, SEQ ID NO:58, SEQ ID NO:68, SEQ ID NO:73 and/or SEQ ID NO:74.
[0097]More preferred arthropod esterase proteins of the present invention include proteins encoded by a nucleic acid molecule comprising at least a portion of nfE1401, nfE2364, nfE3421, nfE4524, nfE51982, nfE51515, nfE52144, nfE51650, nfE61488, nfE61792, nfE61650, nfE72836, nfE71788, nfE71710, nfE7650, nfE82801, nfE81785, nfE81710, nfE92007, nfE91584, nfE91540, nfE101987 and/or nfE101590, or of allelic variants of such nucleic acid molecules. More preferred is an esterase protein encoded by nfE1401, nfE2364, nfE3421, nfE4524, nfE51982, nfE51515, nfE52144, nfE51650, nfE61488, nfE61792, nfE61650, nfE72836, nfE71788, nfE71710, nfE7650, nfE82801, nfE81785, nfE81710, nfE92007, nfE91584, nfE91540, nfE101987 and/or nfE101590, or by an allelic variant of such nucleic acid molecules. Particularly preferred arthropod esterase proteins are PfE1103, PfE2121, PfE3103, PfE4137, PfE5505, PfE5550, PfE6550, PfE6530, PfE7596, PfE7570, PfE8595, PfE8570, PfE9528 and PfE10530.
[0098]In one embodiment, a preferred esterase protein of the present invention is encoded by at least a portion of SEQ ID NO:1, SEQ ID NO:4, SEQ ID NO:7, SEQ ID NO:10, SEQ ID NO:13, SEQ ID NO:16, SEQ ID NO:18, SEQ ID NO:21, SEQ ID NO:23, SEQ ID NO:24, SEQ ID NO:27, SEQ ID NO:28, SEQ ID NO:30, SEQ ID NO:33, SEQ ID NO:34, SEQ ID NO:36, SEQ ID NO:51, SEQ ID NO:57, SEQ ID NO:60 and/or SEQ ID NO:67, and, as such, has an amino acid sequence that includes at least a portion of SEQ ID NO:2, SEQ ID NO:5, SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:19, SEQ ID NO:25, SEQ ID NO:31, SEQ ID NO:37, SEQ ID NO:53, SEQ ID NO:54, SEQ ID NO:55, SEQ ID NO:58 and/or SEQ ID NO:68. Also preferred is a protein encoded by an allelic variant of a nucleic acid molecule comprising at least a portion of the above-listed nucleic acid sequences.
[0099]Particularly preferred esterase proteins of the present invention include SEQ ID NO:2, SEQ ID NO:5, SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:19, SEQ ID NO:25, SEQ ID NO:31, SEQ ID NO:37, SEQ ID NO:39, SEQ ID NO:40, SEQ ID NO:41, SEQ ID NO:42, SEQ ID NO:43, SEQ ID NO:44, SEQ ID NO:53, SEQ ID NO:54, SEQ ID NO:55, SEQ ID NO:58, SEQ ID NO:68, SEQ ID NO:73 and/or SEQ ID NO:74. (including, but not limited to, the proteins consisting of such sequences, fusion proteins and multivalent proteins) and proteins encoded by allelic variants of SEQ ID NO:1, SEQ ID NO:4, SEQ ID NO:7, SEQ ID NO:10, SEQ ID NO:13, SEQ ID NO:16, SEQ ID NO:18, SEQ ID NO:21, SEQ ID NO:23, SEQ ID NO:24, SEQ ID NO:27, SEQ ID NO:28, SEQ ID NO:30, SEQ ID NO:33, SEQ ID NO:34, SEQ ID NO:36, SEQ ID NO:51, SEQ ID NO:57, SEQ ID NO:60 and/or SEQ ID NO:67.
[0100]Another embodiment of the present invention is an isolated nucleic acid molecule that hybridizes under stringent hybridization conditions with a C. felis esterase gene. The identifying characteristics of such a gene are heretofore described. A nucleic acid molecule of the present invention can include an isolated natural arthropod esterase gene or a homolog thereof, the latter of which is described in more detail below. A nucleic acid molecule of the present invention can include one or more regulatory regions, full-length or partial coding regions, or combinations thereof. The minimal size of a nucleic acid molecule of the present invention is the minimal size that can form a stable hybrid with a C. felis esterase gene under stringent hybridization conditions.
[0101]In accordance with the present invention, an isolated nucleic acid molecule is a nucleic acid molecule that has been removed from its natural milieu (i.e., that has been subject to human manipulation) and can include DNA, RNA, or derivatives of either DNA or RNA. As such, "isolated" does not reflect the extent to which the nucleic acid molecule has been purified. An isolated arthropod esterase nucleic acid molecule of the present invention can be isolated from its natural source or can be produced using recombinant DNA technology (e.g., polymerase chain reaction (PCR) amplification, cloning) or chemical synthesis. Isolated esterase nucleic acid molecules can include, for example, natural allelic variants and nucleic acid molecules modified by nucleotide insertions, deletions, substitutions, and/or inversions in a manner such that the modifications do not substantially interfere with the nucleic acid molecule's ability to encode an esterase protein of the present invention or to form stable hybrids under stringent conditions with natural gene isolates.
[0102]An arthropod esterase nucleic acid molecule homolog can be produced using a number of methods known to those skilled in the art (see, for example, Sambrook et al, ibid.). For example, nucleic acid molecules can be modified using a variety of techniques including, but not limited to, classic mutagenesis and recombinant DNA techniques (e.g., site-directed mutagenesis, chemical treatment, restriction enzyme cleavage, ligation of nucleic acid fragments and/or PCR amplification), synthesis of oligonucleotide mixtures and ligation of mixture groups to "build" a mixture of nucleic acid molecules and combinations thereof. Nucleic acid molecule homologs can be selected by hybridization with a C. felis esterase gene or by screening for the function of a protein encoded by the nucleic acid molecule (e.g., ability to elicit an immune response against at least one epitope of an arthropod esterase protein, hydrolyze α-napthyl acetate, hydrolyze the methyl ester group of juvenile hormone and/or bind to DFP).
[0103]An isolated nucleic acid molecule of the present invention can include a nucleic acid sequence that encodes at least one arthropod esterase protein of the present invention, examples of such proteins being disclosed herein. Although the phrase "nucleic acid molecule" primarily refers to the physical nucleic acid molecule and the phrase "nucleic acid sequence" primarily refers to the sequence of nucleotides on the nucleic acid molecule, the two phrases can be used interchangeably, especially with respect to a nucleic acid molecule, or a nucleic acid sequence, being capable of encoding an arthropod esterase protein.
[0104]A preferred nucleic acid molecule of the present invention, when administered to an animal, is capable of protecting that animal from infestation by a hematophagous ectoparasite. As will be disclosed in more detail below, such a nucleic acid molecule can be, or can encode, an antisense RNA, a molecule capable of triple helix formation, a ribozyme, or other nucleic acid-based drug compound. In additional embodiments, a nucleic acid molecule of the present invention can encode a protective esterase protein (e.g., an esterase protein of the present invention), the nucleic acid molecule being delivered to the animal, for example, by direct injection (i.e, as a naked nucleic acid) or in a vehicle such as a recombinant virus vaccine or a recombinant cell vaccine.
[0105]One embodiment of the present invention is an esterase nucleic acid molecule that hybridizes under stringent hybridization conditions with nucleic acid molecule nfE1401 and preferably with a nucleic acid molecule having nucleic acid sequence SEQ ID NO:1 and/or SEQ ID NO:3.
[0106]Another embodiment of the present invention is an esterase nucleic acid molecule that hybridizes under stringent hybridization conditions with nucleic acid molecule nfE2364 and preferably with a nucleic acid molecule having nucleic acid sequence SEQ ID NO:4 and/or SEQ ID NO:6.
[0107]Another embodiment of the present invention is an esterase nucleic acid molecule that hybridizes under stringent hybridization conditions with nucleic acid molecule nfE3421 and preferably with a nucleic acid molecule having nucleic acid sequence SEQ ID NO:7 and/or SEQ ID NO:9.
[0108]Another embodiment of the present invention is an esterase nucleic acid molecule that hybridizes under stringent hybridization conditions with nucleic acid molecule nfE4524 and preferably with a nucleic acid molecule having nucleic acid sequence SEQ ID NO:10 and/or SEQ ID NO:12.
[0109]Another embodiment of the present invention is an esterase nucleic acid molecule that hybridizes under stringent hybridization conditions with nucleic acid molecule nfE52144 and preferably with a nucleic acid molecule having nucleic acid sequence SEQ ID NO:57 and/or SEQ ID NO:59.
[0110]Another embodiment of the present invention is an esterase nucleic acid molecule that hybridizes under stringent hybridization conditions with nucleic acid molecule nfE61792 and preferably with a nucleic acid molecule having nucleic acid sequence SEQ ID NO:18 and/or SEQ ID NO:20.
[0111]Another embodiment of the present invention is an esterase nucleic acid molecule that hybridizes under stringent hybridization conditions with nucleic acid molecule nfE72836 and preferably with a nucleic acid molecule having nucleic acid sequence SEQ ID NO:24 and/or SEQ ID NO:26.
[0112]Another embodiment of the present invention is an esterase nucleic acid molecule that hybridizes under stringent hybridization conditions with nucleic acid molecule nfE82801 and preferably with a nucleic acid molecule having nucleic acid sequence SEQ ID NO:30 and/or SEQ ID NO:32.
[0113]Another embodiment of the present invention is an esterase nucleic acid molecule that hybridizes under stringent hybridization conditions with nucleic acid molecule nfE92007 and preferably with a nucleic acid molecule having nucleic acid sequence SEQ ID NO:36 and/or SEQ ID NO:38.
[0114]Another embodiment of the present invention is an esterase nucleic acid molecule that hybridizes under stringent hybridization conditions with nucleic acid molecule nfE101987 and preferably with a nucleic acid molecule having nucleic acid sequence SEQ ID NO:67 and/or SEQ ID NO:69.
[0115]Comparison of nucleic acid sequence SEQ ID NO:1 (i.e., the nucleic acid sequence of nfE1401) with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:1 showed no identifiable identity with any sequence reported in GENBANK®.
[0116]Comparison of nucleic acid sequence SEQ ID NO:4 (i.e., the coding strand of nucleic acid sequence of nfE2364) with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:4 showed the most homolog, i.e., about 43% identity, between SEQ ID NO:4 and a H. virescens juvenile hormone esterase gene.
[0117]Comparison of nucleic acid sequence SEQ ID NO:7 (i.e., the coding strand of nucleic acid sequence of nfE3421) with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:7 showed the most homolog, i.e., about 53% identity, between SEQ ID NO:7 and a Torpedo marmorata acetylcholinesterase gene.
[0118]Comparison of nucleic acid sequence SEQ ID NO:10 (i.e., the coding strand of nucleic acid sequence of nfE4524) with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:10 showed the most homolog, i.e., about 47% identity, between SEQ ID NO: 10 and an Anas platyrhyncos thioesterase B gene.
[0119]Comparison of nucleic acid sequence SEQ ID NO:57 (i.e., the coding strand of nucleic acid sequence of nfE52144) with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:57 showed the most homolog, i.e., about 41% identity, between SEQ ID NO:57 and a esterase mRNA from Myzus persicae.
[0120]Comparison of nucleic acid sequence SEQ ID NO:18 (i.e., the coding strand of nucleic acid sequence of nfE61792) with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:18 showed the most homolog, i.e., about 41% identity, between SEQ ID NO:18 and a esterase gene from Myzus persicae.
[0121]Comparison of nucleic acid sequence SEQ ID NO:24 (i.e., the coding strand of nucleic acid sequence of nfE72836) with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:24 showed the most homolog, i.e., about 48% identity, between SEQ ID NO:24 and an Anas platychyncos thioesterase B gene.
[0122]Comparison of nucleic acid sequence SEQ ID NO:30 (i.e., the coding strand of nucleic acid sequence of nfE82801) with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:30 showed the most homolog, i.e., about 46% identity, between SEQ ID NO:30 and a Mus musculus carboxyl ester lipase gene.
[0123]Comparison of nucleic acid sequence SEQ ID NO:36 (i.e., the coding strand of nucleic acid sequence of nfE92007) with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:36 showed the most homolog, i.e., about 47% identity, between SEQ ID NO:36 and a hamster mRNA for CE precursor gene.
[0124]Comparison of nucleic acid sequence SEQ ID NO:67 with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:67 showed the most homology, i.e., about 48% identity, between SEQ ID NO:67 and a Lucilia cuprina alpha esterase gene (genembl # U56636) gene.
[0125]Preferred arthropod esterase nucleic acid molecules include nucleic acid molecules having a nucleic acid sequence that is at least about 55%, preferably at least about 60%, more preferably at least about 65%, more preferably at least about 70%, more preferably at least about 75%, more preferably at least about 80%, more preferably at least about 90%, and even more preferably at least about 95% identical to nucleic acid sequence SEQ ID NO:1, SEQ ID NO:3, SEQ ID NO:4, SEQ ID NO:6, SEQ ID NO:7, SEQ ID NO:9, SEQ ID NO:10, SEQ ID NO:12, SEQ ID NO:13, SEQ ID NO:15, SEQ ID NO:16, SEQ ID NO:17, SEQ ID NO:18, SEQ ID NO:20, SEQ ID NO:21, SEQ ID NO:22, SEQ ID NO:23, SEQ ID NO:24, SEQ ID NO:26, SEQ ID NO:27, SEQ ID NO:28, SEQ ID NO:29, SEQ ID NO:30, SEQ ID NO:32, SEQ ID NO:33, SEQ ID NO:34, SEQ ID NO:35, SEQ ID NO:36, SEQ ID NO:38, SEQ ID NO:51, SEQ ID NO:52, SEQ ID NO:57, SEQ ID NO:59, SEQ ID NO:60, SEQ ID NO:61, SEQ ID NO:67, SEQ ID NO:69, SEQ ID NO:70, SEQ ID NO:71, SEQ ID NO:72, SEQ ID NO:76 and/or a nucleic acid molecule encoding a protein comprising amino acid sequence SEQ ID NO:74.
[0126]Another preferred nucleic acid molecule of the present invention includes at least a portion of nucleic acid sequence SEQ ID NO:1, SEQ ID NO:3, SEQ ID NO:4, SEQ ID NO:6, SEQ ID NO:7, SEQ ID NO:9, SEQ ID NO:10, SEQ ID NO:12, SEQ ID NO:13, SEQ ID NO:15, SEQ ID NO:16, SEQ ID NO:17, SEQ ID NO:18, SEQ ID NO:20, SEQ ID NO:21, SEQ ID NO:22, SEQ ID NO:23, SEQ ID NO:24, SEQ ID NO:26, SEQ ID NO:27, SEQ ID NO:28, SEQ ID NO:29, SEQ ID NO:30, SEQ ID NO:32, SEQ ID NO:33, SEQ ID NO:34, SEQ ID NO:35, SEQ ID NO:36, SEQ ID NO:38, SEQ ID NO:51, SEQ ID NO:52, SEQ ID NO:57, SEQ ID NO:59, SEQ ID NO:60, SEQ ID NO:61, SEQ ID NO:67, SEQ ID NO:69, SEQ ID NO:70, SEQ ID NO:71, SEQ ID NO:72, SEQ ID NO:76 and/or a nucleic acid molecule encoding a protein comprising amino acid sequence SEQ ID NO:74, that is capable of hybridizing to a C. felis esterase gene of the present invention, as well as allelic variants thereof. A more preferred nucleic acid molecule includes the nucleic acid sequence SEQ ID NO:1, SEQ ID NO:3, SEQ ID NO:4, SEQ ID NO:6, SEQ ID NO:7, SEQ ID NO:9, SEQ ID NO:10, SEQ ID NO:12, SEQ ID NO:13, SEQ ID NO:15, SEQ ID NO:16, SEQ ID NO:17, SEQ ID NO:18, SEQ ID NO:20, SEQ ID NO:21, SEQ ID NO:22, SEQ ID NO:23, SEQ ID NO:24, SEQ ID NO:26, SEQ ID NO:27, SEQ ID NO:28, SEQ ID NO:29, SEQ ID NO:30, SEQ ID NO:32, SEQ ID NO:33, SEQ ID NO:34, SEQ ID NO:35, SEQ ID NO:36, SEQ ID NO:38, SEQ ID NO:51, SEQ ID NO:52, SEQ ID NO:57, SEQ ID NO:59, SEQ ID NO:60, SEQ ID NO:61, SEQ ID NO:67, SEQ ID NO:69, SEQ ID NO:70, SEQ ID NO:71, SEQ ID NO:72, SEQ ID NO:76 and/or a nucleic acid molecule encoding a protein comprising amino acid sequence SEQ ID NO:74, as well as allelic variants thereof. Such nucleic acid molecules can include nucleotides in addition to those included in the SEQ ID NOs, such as, but not limited to, a full-length gene, a full-length coding region, a nucleic acid molecule encoding a fusion protein, or a nucleic acid molecule encoding a multivalent protective compound. Particularly preferred nucleic acid molecules include nfE1401, nfE2364, nfE3421, nfE4524, nfE51982, nfE51515, nfE52144, nfE51650, nfE61488, nfE61792, nfE61650, nfE72836, nfE71788, nfE71710, nfE7650, nfE82801, nfE81785, nfE81710, nfE92007, nfE91584, nfE91540, nfE11987 and nfE11590.
[0127]The present invention also includes a nucleic acid molecule encoding a protein having at least a portion of SEQ ID NO:2, SEQ ID NO:5, SEQ ID NO:8, SEQ ID NO:11, SEQ ID NO:14, SEQ ID NO:19, SEQ ID NO:25, SEQ ID NO:31, SEQ ID NO:37, SEQ ID NO:39, SEQ ID NO:40, SEQ ID NO:41, SEQ ID NO:42, SEQ ID NO:43, SEQ ID NO:44, SEQ ID NO:53, SEQ ID NO:54, SEQ ID NO:55, SEQ ID NO:58, SEQ ID NO:68, SEQ ID NO:73 and/or SEQ ID NO:74, including nucleic acid molecules that have been modified to accommodate codon usage properties of the cells in which such nucleic acid molecules are to be expressed.
[0128]Knowing the nucleic acid sequences of certain arthropod esterase nucleic acid molecules of the present invention allows one skilled in the art to, for example, (a) make copies of those nucleic acid molecules, (b) obtain nucleic acid molecules including at least a portion of such nucleic acid molecules (e.g., nucleic acid molecules including full-length genes, full-length coding regions, regulatory control sequences, truncated coding regions), and (c) obtain esterase nucleic acid molecules from other arthropods. Such nucleic acid molecules can be obtained in a variety of ways including screening appropriate expression libraries with antibodies of the present invention; traditional cloning techniques using oligonucleotide probes of the present invention to screen appropriate libraries or DNA; and PCR amplification of appropriate libraries or DNA using oligonucleotide primers of the present invention. Preferred libraries to screen or from which to amplify nucleic acid molecule include flea pre-pupal, 3rd instar or adult cDNA libraries as well as genomic DNA libraries. Similarly, preferred DNA sources to screen or from which to amplify nucleic acid molecules include flea pre-pupal, 3rd instar or adult cDNA and genomic DNA. Techniques to clone and amplify genes are disclosed, for example, in Sambrook et al., ibid.
[0129]The present invention also includes nucleic acid molecules that are oligonucleotides capable of hybridizing, under stringent hybridization conditions, with complementary regions of other, preferably longer, nucleic acid molecules of the present invention such as those comprising arthropod esterase genes or other arthropod esterase nucleic acid molecules. Oligonucleotides of the present invention can be RNA, DNA, or derivatives of either. The minimum size of such oligonucleotides is the size required for formation of a stable hybrid between an oligonucleotide and a complementary sequence on a nucleic acid molecule of the present invention. Minimal size characteristics are disclosed herein. The present invention includes oligonucleotides that can be used as, for example, probes to identify nucleic acid molecules, primers to produce nucleic acid molecules or therapeutic reagents to inhibit esterase protein production or activity (e.g., as antisense-, triplex formation-, ribozyme- and/or RNA drug-based reagents). The present invention also includes the use of such oligonucleotides to protect animals from disease using one or more of such technologies. Appropriate oligonucleotide-containing therapeutic compositions can be administered to an animal using techniques known to those skilled in the art.
[0130]One embodiment of the present invention includes a recombinant vector, which includes at least one isolated nucleic acid molecule of the present invention, inserted into any vector capable of delivering the nucleic acid molecule into a host cell. Such a vector contains heterologous nucleic acid sequences, that is nucleic acid sequences that are not naturally found adjacent to nucleic acid molecules of the present invention and that preferably are derived from a species other than the species from which the nucleic acid molecule(s) are derived. The vector can be either RNA or DNA, either prokaryotic or eukaryotic, and typically is a virus or a plasmid. Recombinant vectors can be used in the cloning, sequencing, and/or otherwise manipulation of arthropod esterase nucleic acid molecules of the present invention.
[0131]One type of recombinant vector, referred to herein as a recombinant molecule, comprises a nucleic acid molecule of the present invention operatively linked to an expression vector. The phrase operatively linked refers to insertion of a nucleic acid molecule into an expression vector in a manner such that the molecule is able to be expressed when transformed into a host cell. As used herein, an expression vector is a DNA or RNA vector that is capable of transforming a host cell and of effecting expression of a specified nucleic acid molecule. Preferably, the expression vector is also capable of replicating within the host cell. Expression vectors can be either prokaryotic or eukaryotic, and are typically viruses or plasmids. Expression vectors of the present invention include any vectors that function (i.e., direct gene expression) in recombinant cells of the present invention, including in bacterial, fungal, endoparasite, insect, other animal, and plant cells. Preferred expression vectors of the present invention can direct gene expression in bacterial, yeast, insect and mammalian cells and more preferably in the cell types disclosed herein.
[0132]In particular, expression vectors of the present invention contain regulatory sequences such as transcription control sequences, translation control sequences, origins of replication, and other regulatory sequences that are compatible with the recombinant cell and that control the expression of nucleic acid molecules of the present invention. In particular, recombinant molecules of the present invention include transcription control sequences. Transcription control sequences are sequences which control the initiation, elongation, and termination of transcription. Particularly important transcription control sequences are those which control transcription initiation, such as promoter, enhancer, operator and repressor sequences. Suitable transcription control sequences include any transcription control sequence that can function in at least one of the recombinant cells of the present invention. A variety of such transcription control sequences are known to those skilled in the art. Preferred transcription control sequences include those which function in bacterial, yeast, insect and mammalian cells, such as, but not limited to, tac, lac, trp, trc, oxy-pro, omp/lpp, rrnB, bacteriophage lambda (such as lambda pL and lambda pR and fusions that include such promoters), bacteriophage T7, T7lac, bacteriophage T3, bacteriophage SP6, bacteriophage SP01, metallothionein, alpha-mating factor, Pichia alcohol oxidase, alphavirus subgenomic promoters (such as Sindbis virus subgenomic promoters), antibiotic resistance gene, baculovirus, Heliothis zea insect virus, vaccinia virus, herpesvirus, raccoon poxvirus, other poxvirus, adenovirus, cytomegalovirus (such as intermediate early promoters), simian virus 40, retrovirus, actin, retroviral long terminal repeat, Rous sarcoma virus, heat shock, phosphate and nitrate transcription control sequences as well as other sequences capable of controlling gene expression in prokaryotic or eukaryotic cells. Additional suitable transcription control sequences include tissue-specific promoters and enhancers as well as lymphokine-inducible promoters (e.g., promoters inducible by interferons or interleukins). Transcription control sequences of the present invention can also include naturally occurring transcription control sequences naturally associated with arthropods, such as, C. felis.
[0133]Suitable and preferred nucleic acid molecules to include in recombinant vectors of the present invention are as disclosed herein. Preferred nucleic acid molecules to include in recombinant vectors, and particularly in recombinant molecules, include nfE1401, nfE2364, nfE3421, nfE4524, nfE51982, nfE51515, nfE52144, nfE51650, nfE61488, nfE61792, nfE61650, nfE72836, nfE71788, nfE71710, nfE7650, nfE82801, nfE81785, nfE81710, nfE92007, nfE91584, nfE91540, nfE101987 and/or nfE101590. Particularly preferred recombinant molecules of the present invention include pCro-nfE61488, pTrc-nfE7650, pTrc-nfE71710, pTrc-nfE81710, pTrc-nfE51650, pTrc-nfE91540, pFB-nfE61679, pVL-nfE71802, pVL-fE81792 and pVL-nfE91600, the production of which are described in the Examples section.
[0134]Recombinant molecules of the present invention may also (a) contain secretory signals (i.e., signal segment nucleic acid sequences) to enable an expressed arthropod protein of the present invention to be secreted from the cell that produces the protein and/or (b) contain fusion sequences which lead to the expression of nucleic acid molecules of the present invention as fusion proteins. Examples of suitable signal segments include any signal segment capable of directing the secretion of a protein of the present invention. Preferred signal segments include, but are not limited to, tissue plasminogen activator (t-PA), interferon, interleukin, growth hormone, histocompatibility and viral envelope glycoprotein signal segments, as well as natural signal sequences. Suitable fusion segments encoded by fusion segment nucleic acids are disclosed herein. In addition, a nucleic acid molecule of the present invention can be joined to a fusion segment that directs the encoded protein to the proteosome, such as a ubiquitin fusion segment. Recombinant molecules may also include intervening and/or untranslated sequences surrounding and/or within the nucleic acid sequences of nucleic acid molecules of the present invention.
[0135]Another embodiment of the present invention includes a recombinant cell comprising a host cell transformed with one or more recombinant molecules of the present invention. Transformation of a nucleic acid molecule into a cell can be accomplished by any method by which a nucleic acid molecule can be inserted into the cell. Transformation techniques include, but are not limited to, transfection, electroporation, microinjection, lipofection, adsorption, and protoplast fusion. A recombinant cell may remain unicellular or may grow into a tissue, organ or a multicellular organism. Transformed nucleic acid molecules of the present invention can remain extrachromosomal or can integrate into one or more sites within a chromosome of the transformed (i.e., recombinant) cell in such a manner that their ability to be expressed is retained. Preferred nucleic acid molecules with which to transform a cell include arthropod esterase nucleic acid molecules disclosed herein. Particularly preferred nucleic acid molecules with which to transform a cell include nfE1401, nfE2364, nfE3421, nfE4524, nfE51982, nfE51515, nfE52144, nfE51650, nfE61488, nfE61792, nfE61650, nfE72836, nfE71788, nfE71710, nfE7650, nfE82801, nfE81785, nfE81710, nfE92007, nfE91584, nfE91540, nfE101987 and/or nfE101590.
[0136]Suitable host cells to transform include any cell that can be transformed with a nucleic acid molecule of the present invention. Host cells can be either untransformed cells or cells that are already transformed with at least one nucleic acid molecule (e.g., nucleic acid molecules encoding one or more proteins of the present invention and/or other proteins useful in the production of multivalent vaccines). Host cells of the present invention either can be endogenously (i.e., naturally) capable of producing arthropod esterase proteins of the present invention or can be capable of producing such proteins after being transformed with at least one nucleic acid molecule of the present invention. Host cells of the present invention can be any cell capable of producing at least one protein of the present invention, and include bacterial, fungal (including yeast), parasite, other insect, other animal and plant cells. Preferred host cells include bacterial, mycobacterial, yeast, insect and mammalian cells. More preferred host cells include Salmonella, Escherichia, Bacillus, Listeria, Saccharomyces, Spodoptera, Mycobacteria, Trichoplusia, BHK (baby hamster kidney) cells, MDCK cells (normal dog kidney cell line for canine herpesvirus cultivation), CRFK cells (normal cat kidney cell line for feline herpesvirus cultivation), CV-1 cells (African monkey kidney cell line used, for example, to culture raccoon poxvirus), COS (e.g., COS-7) cells, and Vero cells. Particularly preferred host cells are Escherichia coli, including E. coli K-12 derivatives; Salmonella typhi; Salmonella typhimurium, including attenuated strains such as UK-1 .sub.X3987 and SR-11 .sub.X4072; Spodoptera frugiperda; Trichoplusia ni; BHK cells; MDCK cells; CRFK cells; CV-1 cells; COS cells; Vero cells; and non-tumorigenic mouse myoblast G8 cells (e.g., ATCC CRL 1246). Additional appropriate mammalian cell hosts include other kidney cell lines, other fibroblast cell lines (e.g., human, murine or chicken embryo fibroblast cell lines), myeloma cell lines, Chinese hamster ovary cells, mouse NIH/3T3 cells, LMTK31 cells and/or HeLa cells. In one embodiment, the proteins may be expressed as heterologous proteins in myeloma cell lines employing immunoglobulin promoters.
[0137]A recombinant cell is preferably produced by transforming a host cell with one or more recombinant molecules, each comprising one or more nucleic acid molecules of the present invention operatively linked to an expression vector containing one or more transcription control sequences. The phrase operatively linked refers to insertion of a nucleic acid molecule into an expression vector in a manner such that the molecule is able to be expressed when transformed into a host cell.
[0138]A recombinant molecule of the present invention is a molecule that can include at least one of any nucleic acid molecule heretofore described operatively linked to at least one of any transcription control sequence capable of effectively regulating expression of the nucleic acid molecule(s) in the cell to be transformed, examples of which are disclosed herein. Particularly preferred recombinant molecules include pCro-nfE61488, pTrc-nfE7650, pTrc-nfE71710, pTrc-nfE81710, pTrc-nfE51650, pTrc-nfE91540, pFB-nfE61679, pVL-nfE71802, pVL-fE81792 and pVL-nfE91600.
[0139]A recombinant cell of the present invention includes any cell transformed with at least one of any nucleic acid molecule of the present invention. Suitable and preferred nucleic acid molecules as well as suitable and preferred recombinant molecules with which to transform cells are disclosed herein. Particularly preferred recombinant cells include E. coli:pCro-nfE61488, E. coli:pTrc-nfE71710, E coli:pTrc-nfE7650, E. coli:pTrc-nfE81710, E. coli:pTrc-nfE51650, E. coli:pTrc-nfE91540, S. frugiperda:pVL-nfE71802, S. frugiperda:pVL-nfE81792, S. frugiperda:pVL-nfE91600 and S. frugiperda:pVL-nfE61679. Details regarding the production of these recombinant cells are disclosed herein.
[0140]Recombinant cells of the present invention can also be co-transformed with one or more recombinant molecules including arthropod esterase nucleic acid molecules encoding one or more proteins of the present invention and one or more other nucleic acid molecules encoding other protective compounds, as disclosed herein (e.g., to produce multivalent vaccines).
[0141]Recombinant DNA technologies can be used to improve expression of transformed nucleic acid molecules by manipulating, for example, the number of copies of the nucleic acid molecules within a host cell, the efficiency with which those nucleic acid molecules are transcribed, the efficiency with which the resultant transcripts are translated, and the efficiency of post-translational modifications. Recombinant techniques useful for increasing the expression of nucleic acid molecules of the present invention include, but are not limited to, operatively linking nucleic acid molecules to high-copy number plasmids, integration of the nucleic acid molecules into one or more host cell chromosomes, addition of vector stability sequences to plasmids, substitutions or modifications of transcription control signals (e.g., promoters, operators, enhancers), substitutions or modifications of translational control signals (e.g., ribosome binding sites, Shine-Dalgarno sequences), modification of nucleic acid molecules of the present invention to correspond to the codon usage of the host cell, deletion of sequences that destabilize transcripts, and use of control signals that temporally separate recombinant cell growth from recombinant enzyme production during fermentation. The activity of an expressed recombinant protein of the present invention may be improved by fragmenting, modifying, or derivatizing nucleic acid molecules encoding such a protein.
[0142]Isolated esterase proteins of the present invention can be produced in a variety of ways, including production and recovery of natural proteins, production and recovery of recombinant proteins, and chemical synthesis of the proteins. In one embodiment, an isolated protein of the present invention is produced by culturing a cell capable of expressing the protein under conditions effective to produce the protein, and recovering the protein. A preferred cell to culture is a recombinant cell of the present invention. Effective culture conditions include, but are not limited to, effective media, bioreactor, temperature, pH and oxygen conditions that permit protein production. An effective medium refers to any medium in which a cell is cultured to produce an arthropod esterase protein of the present invention. Such medium typically comprises an aqueous medium having assimilable carbon, nitrogen and phosphate sources, and appropriate salts, minerals, metals and other nutrients, such as vitamins. Cells of the present invention can be cultured in conventional fermentation bioreactors, shake flasks, test tubes, microtiter dishes, and petri plates. Culturing can be carried out at a temperature, pH and oxygen content appropriate for a recombinant cell. Such culturing conditions are within the expertise of one of ordinary skill in the art. Examples of suitable conditions are included in the Examples section.
[0143]Depending on the vector and host system used for production, resultant proteins of the present invention may either remain within the recombinant cell; be secreted into the fermentation medium; be secreted into a space between two cellular membranes, such as the periplasmic space in E. coli; or be retained on the outer surface of a cell or viral membrane. The phrase "recovering the protein", as well as similar phrases, refers to collecting the whole fermentation medium containing the protein and need not imply additional steps of separation or purification. Proteins of the present invention can be purified using a variety of standard protein purification techniques, such as, but not limited to, affinity chromatography, ion exchange chromatography, filtration, electrophoresis, hydrophobic interaction chromatography, gel filtration chromatography, reverse phase chromatography, concanavalin A chromatography, chromatofocusing and differential solubilization. Proteins of the present invention are preferably retrieved in "substantially pure" form. As used herein, "substantially pure" refers to a purity that allows for the effective use of the protein as a therapeutic composition or diagnostic. A therapeutic composition for animals, for example, should exhibit no substantial toxicity and preferably should be capable of stimulating the production of antibodies in a treated animal.
[0144]The present invention also includes isolated (i.e., removed from their natural milieu) antibodies that selectively bind to an arthropod esterase protein of the present invention or a mimetope thereof (i.e., anti-arthropod esterase antibodies). As used herein, the term "selectively binds to" an esterase protein refers to the ability of antibodies of the present invention to preferentially bind to specified proteins and mimetopes thereof of the present invention. Binding can be measured using a variety of methods standard in the art including enzyme immunoassays (e.g., ELISA), immunoblot assays, etc.; see, for example, Sambrook et al., ibid. An anti-arthropod esterase antibody preferably selectively binds to an arthropod esterase protein in such a way as to reduce the activity of that protein.
[0145]Isolated antibodies of the present invention can include antibodies in a bodily fluid (such as, but not limited to, serum), or antibodies that have been purified to varying degrees. Antibodies of the present invention can be polyclonal or monoclonal, functional equivalents such as antibody fragments and genetically-engineered antibodies, including single chain antibodies or chimeric antibodies that can bind to more than one epitope.
[0146]A preferred method to produce antibodies of the present invention includes (a) administering to an animal an effective amount of a protein, peptide or mimetope thereof of the present invention to produce the antibodies and (b) recovering the antibodies. In another method, antibodies of the present invention are produced recombinantly using techniques as heretofore disclosed to produce arthropod esterase proteins of the present invention. Antibodies raised against defined proteins or mimetopes can be advantageous because such antibodies are not substantially contaminated with antibodies against other substances that might otherwise cause interference in a diagnostic assay or side effects if used in a therapeutic composition.
[0147]Antibodies of the present invention have a variety of potential uses that are within the scope of the present invention. For example, such antibodies can be used (a) as therapeutic compounds to passively immunize an animal in order to protect the animal from arthropods susceptible to treatment by such antibodies and/or (b) as tools to screen expression libraries and/or to recover desired proteins of the present invention from a mixture of proteins and other contaminants. Furthermore, antibodies of the present invention can be used to target cytotoxic agents to hematophagous ectoparasites such as those discloses herein, in order to directly kill such hematophagous ectoparasites. Targeting can be accomplished by conjugating (i.e., stably joining) such antibodies to the cytotoxic agents using techniques known to those skilled in the art. Suitable cytotoxic agents are known to those skilled in the art.
[0148]One embodiment of the present invention is a therapeutic composition that, when administered to an animal in an effective manner, is capable of protecting that animal from infestation by hematophagous ectoparasite. Therapeutic compositions of the present invention include at least one of the following protective compounds: an isolated hematophagous arthropod esterase protein (including a peptide); a mimetope of such a protein; an isolated nucleic acid molecule that hybridizes under stringent hybridization conditions with a Ctenocephalides felis esterase gene; an isolated antibody that selectively binds to an hematophagous arthropod esterase protein; and inhibitors of hematophagous arthropod esterase activity (including esterase substrate analogs). As used herein, a protective compound refers to a compound that, when administered to an animal in an effective manner, is able to treat, ameliorate, and/or prevent disease caused by an arthropod of the present invention. Preferred arthropods to target are heretofore disclosed. Examples of proteins, nucleic acid molecules, antibodies and inhibitors of the present invention are disclosed herein.
[0149]A preferred therapeutic composition of the present invention includes at least one of the following protective compounds: an isolated hematophagous ectoparasite carboxylesterase protein (including a peptide); a mimetope of such a protein; an isolated hematophagous ectoparasite carboxylesterase nucleic acid molecule that hybridizes under stringent hybridization conditions with a Ctenocephalides felis carboxylesterase gene; an isolated antibody that selectively binds to a hematophagous ectoparasite carboxylesterase protein; and an inhibitor of carboxylesterase activity identified by its ability to inhibit the activity of a flea carboxylesterase (including a substrate analog).
[0150]Suitable inhibitors of esterase activity are compounds that interact directly with an esterase protein's active site, thereby inhibiting that esterase's activity, usually by binding to or otherwise interacting with or otherwise modifying the esterase's active site. Esterase inhibitors can also interact with other regions of the esterase protein to inhibit esterase activity, for example, by allosteric interaction. Inhibitors of esterases are usually relatively small compounds and as such differ from anti-esterase antibodies. Preferably, an esterase inhibitor of the present invention is identified by its ability to bind to, or otherwise interact with, a flea esterase protein, thereby inhibiting the activity of the flea esterase.
[0151]Esterase inhibitors can be used directly as compounds in compositions of the present invention to treat animals as long as such compounds are not harmful to host animals being treated. Esterase inhibitors can also be used to identify preferred types of arthropod esterases to target using compositions of the present invention, for example by affinity chromatography. Preferred esterase inhibitors of the present invention include, but are not limited to, flea esterase substrate analogs, and other molecules that bind to a flea esterase (e.g., to an allosteric site) in such a manner that esterase activity of the flea esterase is inhibited; examples include, but are not limited to, juvenile hormone analogs and cholinesterase inhibitors as well as other neural transmission inhibitors. An esterase substrate analog refers to a compound that interacts with (e.g., binds to, associates with, modifies) the active site of an esterase protein. A preferred esterase substrate analog inhibits esterase activity. Esterase substrate analogs can be of any inorganic or organic composition, and, as such, can be, but are not limited to, peptides, nucleic acids, and peptidomimetic compounds. Esterase substrate analogs can be, but need not be, structurally similar to an esterase's natural substrate as long as they can interact with the active site of that esterase protein. Esterase substrate analogs can be designed using computer-generated structures of esterase proteins of the present invention or computer structures of esterases' natural substrates. Substrate analogs can also be obtained by generating random samples of molecules, such as oligonucleotides, peptides, peptidomimetic compounds, or other inorganic or organic molecules, and screening such samples by affinity chromatography techniques using the corresponding binding partner, (e.g., a flea esterase). A preferred esterase substrate analog is a peptidomimetic compound (i.e., a compound that is structurally and/or functionally similar to a natural substrate of an esterase of the present invention, particularly to the region of the substrate that interacts with the esterase active site, but that inhibits esterase activity upon interacting with the esterase active site).
[0152]Esterase peptides, mimetopes and substrate analogs, as well as other protective compounds, can be used directly as compounds in compositions of the present invention to treat animals as long as such compounds are not harmful to the animals being treated.
[0153]The present invention also includes a therapeutic composition comprising at least one arthropod esterase-based compound of the present invention in combination with at least one additional compound protective against hematophagous ectoparasite infestation. Examples of such compounds are disclosed herein.
[0154]In one embodiment, a therapeutic composition of the present invention can be used to protect an animal from hematophagous ectoparasite infestation by administering such composition to a hematophagous ectoparasite, such as to a flea, in order to prevent infestation. Such administration could be oral, or by application to the environment (e.g., spraying). Examples of such compositions include, but are not limited to, transgenic vectors capable of producing at least one therapeutic composition of the present invention. In another embodiment, a hematophagous ectoparasite, such as a flea, can ingest therapeutic compositions, or products thereof, present in the blood of a host animal that has been administered a therapeutic composition of the present invention.
[0155]Compositions of the present invention can be administered to any animal susceptible to hematophagous ectoparasite infestation (i.e., a host animal), including warm-blooded animals. Preferred animals to treat include mammals and birds, with cats, dogs, humans, cattle, chinchillas, ferrets, goats, mice, minks, rabbits, raccoons, rats, sheep, squirrels, swine, chickens, ostriches, quail and turkeys as well as other furry animals, pets, zoo animals, work animals and/or food animals, being more preferred. Particularly preferred animals to protect are cats and dogs.
[0156]In accordance with the present invention, a host animal (i.e., an animal that is or is capable of being infested with a hematophagous ectoparasite) is treated by administering to the animal a therapeutic composition of the present invention in such a manner that the composition itself (e.g., an esterase inhibitor, an esterase synthesis suppressor (i.e., a compound that decreases the production of esterase in the hematophagous ectoparasite), an esterase mimetope, or an anti-esterase antibody) or a product generated by the animal in response to administration of the composition (e.g., antibodies produced in response to administration of an arthropod esterase protein or nucleic acid molecule, or conversion of an inactive inhibitor "prodrug" to an active esterase inhibitor) ultimately enters the hematophagous ectoparasite. A host animal is preferably treated in such a way that the compound or product thereof enters the blood stream of the animal. Hematophagous ectoparasites are then exposed to the composition or product when they feed from the animal. For example, flea esterase inhibitors administered to an animal are administered in such a way that the inhibitors enter the blood stream of the animal, where they can be taken up by feeding fleas. In another embodiment, when a host animal is administered an arthropod esterase protein or nucleic acid molecule, the treated animal mounts an immune response resulting in the production of antibodies against the esterase (i.e., anti-esterase antibodies) which circulate in the animal's blood stream and are taken up by hematophagous ectoparasites upon feeding. Blood taken up by hematophagous ectoparasites enters the hematophagous ectoparasites where compounds of the present invention, or products thereof, such as anti-esterase antibodies, esterase inhibitors, esterase mimetopes and/or esterase synthesis suppressors, interact with, and reduce esterase activity in the hematophagous ectoparasite.
[0157]The present invention also includes the ability to reduce larval hematophagous ectoparasite infestation in that when hematophagous ectoparasites feed from a host animal that has been administered a therapeutic composition of the present invention, at least a portion of compounds of the present invention, or products thereof in the blood taken up by the hematophagous ectoparasite are excreted by the hematophagous ectoparasite in feces, which is subsequently ingested by hematophagous ectoparasite larvae. In particular, it is of note that flea larvae obtain most, if not all, of their nutrition from flea feces.
[0158]In accordance with the present invention, reducing esterase activity in a hematophagous ectoparasite can lead to a number of outcomes that reduce hematophagous ectoparasite burden on treated animals and their surrounding environments. Such outcomes include, but are not limited to, (a) reducing the viability of hematophagous ectoparasites that feed from the treated animal, (b) reducing the fecundity of female hematophagous ectoparasites that feed from the treated animal, (c) reducing the reproductive capacity of male hematophagous ectoparasites that feed from the treated animal, (d) reducing the viability of eggs laid by female hematophagous ectoparasites that feed from the treated animal, (e) altering the blood feeding behavior of hematophagous ectoparasites that feed from the treated animal (e.g., hematophagous ectoparasites take up less volume per feeding or feed less frequently), (f) reducing the viability of hematophagous ectoparasite larvae, for example due to the feeding of larvae from feces of hematophagous ectoparasites that feed from the treated animal and/or (g) altering the development of hematophagous ectoparasite larvae (e.g., by decreasing feeding behavior, inhibiting growth, inhibiting (e.g., slowing or blocking) molting, and/or otherwise inhibiting maturation to adults).
[0159]Therapeutic compositions of the present invention also include excipients in which protective compounds are formulated. An excipient can be any material that the animal to be treated can tolerate. Examples of such excipients include water, saline, Ringer's solution, dextrose solution, Hank's solution, and other aqueous physiologically balanced salt solutions. Nonaqueous vehicles, such as fixed oils, sesame oil, ethyl oleate, or triglycerides may also be used. Other useful formulations include suspensions containing viscosity enhancing agents, such as sodium carboxymethylcellulose, sorbitol, or dextran. Excipients can also contain minor amounts of additives, such as substances that enhance isotonicity and chemical stability. Examples of buffers include phosphate buffer, bicarbonate buffer and Tris buffer, while examples of preservatives include thimerosal or o-cresol, formalin and benzyl alcohol. Standard formulations can either be liquid injectables or solids which can be taken up in a suitable liquid as a suspension or solution for injection. Thus, in a non-liquid formulation, the excipient can comprise dextrose, human serum albumin, dog serum albumin, cat serum albumin, preservatives, etc., to which sterile water or saline can be added prior to administration.
[0160]In one embodiment of the present invention, a therapeutic composition can include an adjuvant. Adjuvants are agents that are capable of enhancing the immune response of an animal to a specific antigen. Suitable adjuvants include, but are not limited to, cytokines, chemokines, and compounds that induce the production of cytokines and chemokines (e.g., granulocyte macrophage colony stimulating factor (GM-CSF), granulocyte colony stimulating factor (G-CSF), macrophage colony stimulating factor (M-CSF), colony stimulating factor (CSF), erythropoietin (EPO), interleukin 2 (IL-2), interleukin-3 (IL-3), interleukin 4 (IL-4), interleukin 5 (IL-5), interleukin 6 (IL-6), interleukin 7 (IL-7), interleukin 8 (IL-8), interleukin 10 (IL-10), interleukin 12 (IL-12), interferon gamma, interferon gamma inducing factor I (IGIF), transforming growth factor beta, RANTES (regulated upon activation, normal T cell expressed and presumably secreted), macrophage inflammatory proteins (e.g., MIP-1 alpha and MIP-1 beta), and Leishmania elongation initiating factor (LEIF); bacterial components (e.g., endotoxins, in particular superantigens, exotoxins and cell wall components); aluminum-based salts; calcium-based salts; silica; polynucleotides; toxoids; serum proteins, viral coat proteins; block copolymer adjuvants (e.g., Hunter's TITERMAX® adjuvant (Vaxcel, Inc. Norcross, Ga.), Ribi adjuvants (Ribi Immunochem Research, Inc., Hamilton, Mont.); and saponins and their derivatives (e.g., Quil A (Superfos Biosector A/S, Denmark). Protein adjuvants of the present invention can be delivered in the form of the protein themselves or of nucleic acid molecules encoding such proteins using the methods described herein.
[0161]In one embodiment of the present invention, a therapeutic composition can include a carrier. Carriers include compounds that increase the half-life of a therapeutic composition in the treated animal. Suitable carriers include, but are not limited to, polymeric controlled release vehicles, biodegradable implants, liposomes, bacteria, viruses, other cells, oils, esters, and glycols.
[0162]One embodiment of the present invention is a controlled release formulation that is capable of slowly releasing a composition of the present invention into an animal. As used herein, a controlled release formulation comprises a composition of the present invention in a controlled release vehicle. Suitable controlled release vehicles include, but are not limited to, biocompatible polymers, other polymeric matrices, capsules, microcapsules, microparticles, bolus preparations, osmotic pumps, diffusion devices, liposomes, lipospheres, and transdermal delivery systems. Other controlled release formulations of the present invention include liquids that, upon administration to an animal, form a solid or a gel in situ. Preferred controlled release formulations are biodegradable (i.e., bioerodible).
[0163]A preferred controlled release formulation of the present invention is capable of releasing a composition of the present invention into the blood of an animal at a constant rate sufficient to attain therapeutic dose levels of the composition to protect an animal from hematophagous ectoparasite infestation. The therapeutic composition is preferably released over a period of time ranging from about 1 to about 12 months. A preferred controlled release formulation of the present invention is capable of effecting a treatment preferably for at least about 1 month, more preferably for at least about 3 months, even more preferably for at least about 6 months, even more preferably for at least about 9 months, and even more preferably for at least about 12 months.
[0164]Acceptable protocols to administer therapeutic compositions of the present invention in an effective manner include individual dose size, number of doses, frequency of dose administration, and mode of administration. Determination of such protocols can be accomplished by those skilled in the art. A suitable single dose is a dose that is capable of protecting an animal from disease when administered one or more times over a suitable time period. For example, a preferred single dose of a protein, mimetope or antibody therapeutic composition is from about 1 microgram (μg) to about 10 milligrams (mg) of the therapeutic composition per kilogram body weight of the animal. Booster vaccinations can be administered from about 2 weeks to several years after the original administration. Booster administrations preferably are administered when the immune response of the animal becomes insufficient to protect the animal from disease. A preferred administration schedule is one in which from about 10 μg to about 1 mg of the therapeutic composition per kg body weight of the animal is administered from about one to about two times over a time period of from about 2 weeks to about 12 months. Modes of administration can include, but are not limited to, subcutaneous, intradermal, intravenous, intranasal, oral, transdermal, intraocular and intramuscular routes.
[0165]According to one embodiment, a nucleic acid molecule of the present invention can be administered to an animal in a fashion to enable expression of that nucleic acid molecule into a protective protein or protective RNA (e.g., antisense RNA, ribozyme, triple helix forms or RNA drug) in the animal. Nucleic acid molecules can be delivered to an animal in a variety of methods including, but not limited to, (a) administering a naked (i.e., not packaged in a viral coat or cellular membrane) nucleic acid vaccine (e.g., as naked DNA or RNA molecules, such as is taught, for example in Wolff et al., 1990, Science 247, 1465-1468) or (b) administering a nucleic acid molecule packaged as a recombinant virus vaccine or as a recombinant cell vaccine (i.e., the nucleic acid molecule is delivered by a viral or cellular vehicle).
[0166]A naked nucleic acid vaccine of the present invention includes a nucleic acid molecule of the present invention and preferably includes a recombinant molecule of the present invention that preferably is replication, or otherwise amplification, competent. A naked nucleic acid vaccine of the present invention can comprise one or more nucleic acid molecules of the present invention in the form of, for example, a bicistronic recombinant molecule having, for example one or more internal ribosome entry sites. Preferred naked nucleic acid vaccines include at least a portion of a viral genome (i.e., a viral vector). Preferred viral vectors include those based on alphaviruses, poxviruses, adenoviruses, herpesviruses, and retroviruses, with those based on alphaviruses (such as Sindbis or Semliki virus), species-specific herpesviruses and species-specific poxviruses being particularly preferred. Any suitable transcription control sequence can be used, including those disclosed as suitable for protein production. Particularly preferred transcription control sequence include cytomegalovirus intermediate early (preferably in conjunction with Intron-A), Rous Sarcoma Virus long terminal repeat, and tissue-specific transcription control sequences, as well as transcription control sequences endogenous to viral vectors if viral vectors are used. The incorporation of "strong" poly(A) sequences are also preferred.
[0167]Naked nucleic acid vaccines of the present invention can be administered in a variety of ways, with intramuscular, subcutaneous, intradermal, transdermal, intranasal and oral routes of administration being preferred. A preferred single dose of a naked nucleic acid vaccines ranges from about 1 nanogram (ng) to about 100 μg, depending on the route of administration and/or method of delivery, as can be determined by those skilled in the art. Suitable delivery methods include, for example, by injection, as drops, aerosolized and/or topically. Naked DNA of the present invention can be contained in an aqueous excipient (e.g., phosphate buffered saline) alone or a carrier (e.g., lipid-based vehicles).
[0168]A recombinant virus vaccine of the present invention includes a recombinant molecule of the present invention that is packaged in a viral coat and that can be expressed in an animal after administration. Preferably, the recombinant molecule is packaging-deficient and/or encodes an attenuated virus. A number of recombinant viruses can be used, including, but not limited to, those based on alphaviruses, poxviruses, adenoviruses, herpesviruses, and retroviruses. Preferred recombinant virus vaccines are those based on alphaviruses (such as Sindbis virus), raccoon poxviruses, species-specific herpesviruses and species-specific poxviruses. An example of methods to produce and use alphavirus recombinant virus vaccines is disclosed in PCT Publication No. WO 94/17813, by Xiong et al., published Aug. 18, 1994, which is incorporated by reference herein in its entirety.
[0169]When administered to an animal, a recombinant virus vaccine of the present invention infects cells within the immunized animal and directs the production of a protective protein or RNA nucleic acid molecule that is capable of protecting the animal from hematophagous ectoparasite infestation. For example, a recombinant virus vaccine comprising an arthropod CE nucleic acid molecule of the present invention is administered according to a protocol that results in the animal producing a sufficient immune response to protect itself from hematophagous ectoparasite infestation. A preferred single dose of a recombinant virus vaccine of the present invention is from about 1×104 to about 1×107 virus plaque forming units (pfu) per kilogram body weight of the animal. Administration protocols are similar to those described herein for protein-based vaccines, with subcutaneous, intramuscular, intranasal and oral administration routes being preferred.
[0170]A recombinant cell vaccine of the present invention includes recombinant cells of the present invention that express at least one protein of the present invention. Preferred recombinant cells for this embodiment include Salmonella, E. coli, Listeria, Mycobacterium, S. frugiperda, yeast, (including Saccharomyces cerevisiae), BHK, CV-1, myoblast G8, COS (e.g., COS-7), Vero, MDCK and CRFK recombinant cells. Recombinant cell vaccines of the present invention can be administered in a variety of ways but have the advantage that they can be administered orally, preferably at doses ranging from about 108 to about 1012 cells per kilogram body weight. Administration protocols are similar to those described herein for protein-based vaccines. Recombinant cell vaccines can comprise whole cells, cells stripped of cell walls or cell lysates.
[0171]The efficacy of a therapeutic composition of the present invention to protect an animal from hematophagous ectoparasite infestation can be tested in a variety of ways including, but not limited to, detection of anti-arthropod esterase antibodies (using, for example, proteins or mimetopes of the present invention), detection of cellular immunity within the treated animal, or challenge of the treated animal with hematophagous ectoparasites to determine whether, for example, the feeding, fecundity or viability of hematophagous ectoparasites feeding from the treated animal is disrupted. Challenge studies can include attachment of chambers containing hematophagous ectoparasites onto the skin of the treated animal. In one embodiment, therapeutic compositions can be tested in animal models such as mice. Such techniques are known to those skilled in the art.
[0172]One preferred embodiment of the present invention is the use of arthropod protective compounds, such as proteins, mimetopes, nucleic acid molecules, antibodies and inhibitory compounds of the present invention, to protect an animal from hematophagous ectoparasite, and particularly flea, infestation. Preferred protective compounds of the present invention include, but are not limited to, C. felis esterase nucleic acid molecules, C. felis esterase proteins and mimetopes thereof, anti-C. felis esterase antibodies, and inhibitors of C. felis esterase activity. More preferred protective compounds of the present invention include, but are not limited to, CE or JHE formulations of the present invention, C. felis CE nucleic acid molecules, C. felis CE proteins and mimetopes thereof, anti-flea CE antibodies, anti-flea JHE antibodies, inhibitors of C. felis CE activity and inhibitors of flea JHE activity. Additional protection may be obtained by administering additional protective compounds, including other proteins, mimetopes, nucleic acid molecules, antibodies and inhibitory compounds, as disclosed herein.
[0173]One therapeutic composition of the present invention includes an inhibitor of arthropod esterase activity, i.e., a compound capable of substantially interfering with the function of an arthropod esterase susceptible to inhibition by an inhibitor of arthropod esterase activity. An inhibitor of esterase activity can be identified using arthropod esterase proteins of the present invention. One embodiment of the present invention is a method to identify a compound capable of inhibiting esterase activity of an arthropod. Such a method includes the steps of (a) contacting (e.g., combining, mixing) an isolated flea esterase protein, preferably a C. felis esterase protein of the present invention, with a putative inhibitory compound under conditions in which, in the absence of the compound, the protein has esterase activity, and (b) determining if the putative inhibitory compound inhibits the esterase activity. Putative inhibitory compounds to screen include small organic molecules, antibodies (including mimetopes thereof) and substrate analogs. Methods to determine esterase activity are known to those skilled in the art; see, for example, the Examples section of the present application.
[0174]The present invention also includes a test kit to identify a compound capable of inhibiting esterase activity of an arthropod. Such a test kit includes an isolated flea esterase protein, preferably a C. felis esterase protein, having esterase activity and a means for determining the extent of inhibition of esterase activity in the presence of (i.e., effected by) a putative inhibitory compound. Such compounds are also screened to identify those that are substantially not toxic in host animals.
[0175]Esterase inhibitors isolated by such a method, and/or test kit, can be used to inhibit any esterase that is susceptible to such an inhibitor. Preferred esterase proteins to inhibit are those produced by arthropods. A particularly preferred esterase inhibitor of the present invention is capable of protecting an animal from hematophagous ectoparasite infestation. Effective amounts and dosing regimens can be determined using techniques known to those skilled in the art.
[0176]The following examples are provided for the purposes of illustration and are not intended to limit the scope of the present invention.
EXAMPLES
[0177]It is to be noted that the Examples include a number of molecular biology, microbiology, immunology and biochemistry techniques considered to be known to those skilled in the art. Disclosure of such techniques can be found, for example, in Sambrook et al., ibid., Borovsky, Arch Insect Biochem. and Phys., 7:187-210, 1988, and related references.
Example 1
[0178]This example describes labeling of proteases and esterases with radiolabeled diisopropylfluorophosphate.
[0179]Tissue samples were isolated from unfed or bovine blood-fed 1st instar Ctenocephalides felis flea larvae; bovine blood-fed or cat blood-fed 3rd instar Ctenocephalides felis flea larvae; bovine blood-fed or cat blood-fed Ctenocephalides felis prepupal flea larvae; bovine blood-fed or cat blood-fed adult Ctenocephalides felis flea midgut tissue, and whole unfed, bovine blood-fed or cat blood-fed adult Ctenocephalides felis fleas. The 1st instar, 3rd instar, prepupal and adult midgut tissues were then homogenized by freeze-fracture and sonicated in a Tris buffer comprising 50 mM Tris, pH 8.0 and 100 mM CaCl2. The whole adult flea sample was then homogenized by freeze-fracture and ground with a microtube mortar and pestle. The extracts were centrifuged at about 14,000×g for 20 minutes (min.) and the soluble material recovered. The soluble material was then diluted to a final concentration of about 1 to about 1.2 tissue equivalents per microliter (μl) of Tris buffer. Each sample was labeled with [1,3-3H]-diisopropylfluorophosphate (3H-DFP) (available from DuPont-NEN, Wilmington, Del.) using the method generally described in Borovsky, ibid. About 20 tissue equivalents of each tissue sample were mixed with about 1 μCi of 3H-DFP and incubated for about 18 hours at 4° C. Proteins contained in each sample were then resolved using a 14% Tris-glycine sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) (available from Novex, San Diego, Calif.) under reducing conditions. The gel was soaked in Entensify (available from DuPont-NEN) according to manufacturers instructions, and exposed to X-ray film (available from Kodak X-0mat AR, Rochester, N.Y.) for about 3 days at -70° C.
[0180]Analysis of the resulting autoradiogram (shown in FIG. 1) indicated that tissue samples from 3rd instar, prepupal larvae and whole adult flea contained proteins that labeled with DFP, having a molecular weight (MW) of about 60 kilodalton (kD). No proteins of this MW were labeled in tissue samples from unfed or fed 1st instar larvae and adult midgut. The results indicated preferred tissue distribution and stage-specific expression of DFP-labeled serine esterases in fleas.
Example 2
[0181]This example describes the identification of general CE activity in flea tissue extracts.
[0182]Tissue samples and soluble extracts were prepared as described above in Example 1, except not labelled, from unfed (UF) and bovine blood-fed 1st instar flea larvae, bovine blood-fed 3rd instar flea larvae, bovine blood-fed prepupal flea larvae, unfed whole adult fleas, cat blood-fed adult (ACF) whole fleas, cat blood-fed adult fleas that have had their heads and midguts removed (referred to herein as fed adult partial fleas), unfed adult flea midguts and cat blood-fed adult flea midguts. About 5 tissue equivalents of each tissue were assayed for general CE activity using the following method. Tissue samples of about 5 μl were added to separate wells of flat-bottomed microtiter plate (available from Becton Dickinson, Lincoln Park, N.J.). A control well was prepared by adding about 5 μl of Tris buffer to an empty well of the plate. About 95 μl of 25 mM Tris-HCl (pH 8.0) was then added to each sample to increase the volume in each well to about 100 μl. About 100 μl of 0.25 mM α-napthyl acetate (available from Sigma-Aldrich, St. Louis, Mo.) dissolved in 25 mM Tris-HCl (pH 8.0) was then added to each well. The plate was then incubated for about 15 min. at 37° C. Following the incubation, about 40 μl of 0.3% Fast Blue salt BN (tetrazotized o-dianisidine; available from Signa-Aldrich) dissolved in 3.3% SDS in water was added to each well.
[0183]The microtiter plate was then analyzed using a Cambridge Technology, Inc. (Watertown, Pa.) model 7500 Microplate Reader set to 590 nm. The absorbance value for the control sample was subtracted from absorbance values of experimental samples, such that the background value was zero.
[0184]The results shown in FIG. 2 indicated that general CE activity was detected in all tissue samples. The level of activity varied, with unfed and fed 1st instar larvae, unfed adult flea midguts, and fed adult flea midguts having relatively lower activity than in the other tissues. Thus, the results indicated preferred tissue distribution and stage-specific expression of general CE activity in fleas.
Example 3
[0185]This example describes the determination of general CE activity using isoelectric focusing (JEF)-PAGE and non-reducing SDS-PAGE.
[0186]A. Non-Reducing SDS-PAGE.
[0187]Soluble extracts from unfed and bovine blood-fed 1st instar flea larvae, bovine blood-fed 3rd instar flea larvae, bovine blood-fed prepupal flea larvae, bovine blood-fed adult (ABF) whole fleas and cat blood-fed adult whole fleas were prepared using the method described in Example 1. Each soluble extract sample was combined with SDS sample buffer (available from Novex) and proteins in the samples were resolved by gel electrophoresis using 14% Tris-glycine SDS electrophoresis gels (available from Novex). The gels were run at room temperature for about 1 hour at 200 volts. After electrophoresis, the gels were soaked for about for 30 minutes in 50 mM Tris, pH 8.0, containing 2.5% Triton X-100 to renature the proteins. The gels were then soaked in 50 mM Tris, pH 8.0, for about 5 minutes and then stained for about 5 min. in 50 milliliters (ml) of 25 mM Tris, pH 8.0, containing 50 mg Fast blue salt BN and 10 mg α-napthyl acetate (dissolved in 1 ml acetone). Once protein was detected on the stained gels, the gels were rinsed with water and photographed.
[0188]B. IEF-PAGE.
[0189]Soluble extracts from unfed and bovine blood-fed 1st instar flea larvae, bovine blood-fed 3rd instar flea larvae, bovine blood-fed prepupal flea larvae, unfed and cat blood-fed whole fleas, cat blood-fed adult partial fleas and cat blood-fed adult midguts were prepared as described above in Section A. The extracts were each combined with IEF sample buffer pH 3-7 (available from Novex) and loaded onto pH 3-7 IEF electrophoresis gels (available from Novex). The gels were electrophoresed at room temperature first for about 1 hour at about 100 volts, then for about 1 hour at about 200 volts, and then for about 30 min. at about 500 volts. Following electrophoresis, the gels were soaked in 25 mM Tris buffer, pH 8.0, for about 5 min. and then stained for about 1-5 min. in 50 ml of 25 mM Tris buffer, pH 8.0, containing 50 mg Fast blue salt BN and 10 mg α-napthyl acetate (dissolved in 1 ml acetone). Once protein was detected on the stained gels, the gels were rinsed with water and photographed.
[0190]C. Results.
[0191]The results from gel electrophoresis experiments described above in Sections A and B are shown in FIGS. 3 and 4. The results indicated that certain flea tissues contain proteins having MW's of from about 60 to about 70 kD and native pI values of from about 4.7 to about 5.2 that have CE activity. In particular, CE activity was identified in prepupal larvae and fed adult flea extracts resolved by non-reduced SDS-PAGE. No CE activity was identified in unfed and fed 1st instar larvae or fed 3rd instar larvae extracts (see FIG. 3). When extracts were resolved by native IEF-PAGE, CE activity was identified in fed 3rd instar larvae, prepupal larvae, unfed and fed whole adult flea, and fed adult partial flea extracts (see FIG. 4, lanes 3-7)). No CE activity was identified in unfed or fed 1st instar larvae, or in fed adult flea midgut extracts (see FIG. 4, lanes 1, 2, and 8).
Example 4
[0192]This example describes the purification of CE protein from prepupal flea larvae.
[0193]About 15,000 bovine blood-fed prepupal flea larvae were collected and the larvae were homogenized in TBS by sonication in 50 ml Oak Ridge centrifuge tubes (available from Nalgene Co., Rochester, N.Y.) by sonicating 4 times 20 seconds each at a setting of 5 of a model W-380 Sonicator (available from Heat Systems-Ultrasonics, Inc.). The sonicates were clarified by centrifugation at 18,000 RPM for 30 minutes to produce an extract. Soluble protein in the extract was removed by aspiration and diluted to a volume of about 20 ml in TBS (equivalent to about 1 larva per μl TBS). The extract was then added to a column containing about 5 ml of p-aminobenzamidine linked to agarose beads (available from Sigma-Aldrich, St. Louis, Mo.) and incubated overnight at 4° C. The column was then washed with about 30 ml TBS to remove unbound protein. The collected unbound protein was then concentrated to a volume of about 20 ml using a Macrosep 10 centrifugal protein concentrator (Filtron Technology Corp., Northborough, Mass.) and filtered sequentially through a 1 μm syringe filter and then through a 0.2 μm syringe filter to clarify the sample for chromatography.
[0194]Aliquots of about 0.5 ml were loaded onto a 20 ml Superdex 200 HR gel filtration column (available from Pharmacia, Piscataway, N.J.) equilibrated in TBS, operated on a BioLogic liquid chromatography system (available from BioRad, Burlingame, Calif.). About 1 ml fractions were then collected. Repetitive runs were performed until about 30 ml of each fraction was collected. The fractions were analyzed for CE activity using the assay described above in Example 2. In preparation for cation exchange chromatography, fractions having CE activity (Ve=16-18 ml) were combined and dialyzed against about 2 liters of 20 mM MES buffer (2-(N-morpholino)ethanesulfonic acid), pH 6.0, containing 10 mM NaCl, for about 1.5 hours, and then against about 1 liter of the same buffer overnight at 4° C. Prior to loading onto the cation exchange chromatography column, the sample was again filtered through a 0.2 μm syringe filter to remove precipitated proteins. The sample was then applied to a Bio-Scale S2 cation exchange column (available from BioRad) at a rate of about 0.5 ml/min. The column was washed with MES buffer until all unbound protein was removed. Protein bound to the column was then eluted with a linear gradient from 10 mM to 1 M NaCl in 20 mM MES buffer, pH 6. Fractions were assayed for CE activity using the assay described above in Example 2. The results indicated that CE activity was not retained on the cation exchange column using the above conditions, and all of the activity was found in the flow-through fractions.
[0195]Fractions containing CE activity were pooled and adjusted to pH 7 using 0.5 M Tris, pH 8.0, in preparation for anion exchange chromatography. The pooled fractions were then loaded onto a 4.5 mm×50 mm Poros 10 HQ anion exchange chromatography column (available from PerSeptive Biosystems, Cambridge, Mass.) equilibrated in 25 mM Tris buffer, pH 6.8. The column was washed with the loading buffer, and bound proteins were eluted with a linear gradient of 0 to 1 M NaCl in 25 mM Tris buffer, pH 6.8. Fractions were tested for CE activity using the assay described above in Example 2. The results indicated that CE activity was eluted at about 170 mM NaCl. Fractions containing CE activity were pooled and diafiltered into TBS.
Example 5
[0196]This example describes the determination of N-terminal amino acid sequences of carboxylesterases isolated from prepupal flea larvae.
[0197]A. Anion Exchange Chromatography Fractions.
[0198]Anion exchange chromatography fractions described above in Example 4 that contained proteins having CE activity were pooled, diafiltered into TBS buffer and concentrated 3-fold in a SPEEDVAC® Concentrator (available from Savant Instruments, Holbrook, N.Y.). Proteins in the concentrated samples were then resolved on a reducing, 10% SDS-PAGE Tris-glycine gel (available from Novex) for 1 hour at about 200 V. The proteins on the gel were then blotted onto a polyvinylidene difluoride (PVDF) membrane (available from Novex) for about 70 min in 10 mM CAPS buffer (3-[cyclohexylamino]-1-propanesulfonic acid; available from Sigma-Aldrich), pH 11, with 0.5 mM dithiothreitol (DTT). The membrane was then stained for 1 minute in 0.1% Coomassie Blue R-250 dissolved in 40% methanol and 1% acetic acid. The membrane was destained in 50% methanol for about 10 minutes, rinsed with MilliQ water and air dried. Three stained protein bands were identified having apparent molecular weights of about 64 kD, 65 kD, and 66 kD, respectively. The portion of the membrane containing each band was excised separately. Protein contained in each membrane segment was subjected to N-terminal amino acid sequencing using a 473A Protein Sequencer (available from Applied Biosystems, Foster City, Calif.) and using standard techniques.
[0199]The results indicated that the N-terminal amino acid sequence of the putative 64 kD protein was DPPTVTLPQGEL (denoted SEQ ID NO:39); the N-terminal amino acid sequence of the putative 65 kD protein was DPPTVTLPQGELVGKATNEnxk (denoted SEQ ID NO:40); and the N-terminal amino acid sequence of the putative 66 kD protein was DppTVTLPQGEL (denoted SEQ ID NO:41), in which the lower case letters designate uncertainties and "x" designates an undetermined residue.
[0200]B. Proteins Resolved by Native IEF-PAGE.
[0201]Proteins isolated by anion exchange chromatography as described above in Section A were further resolved by native IEF-PAGE. Proteins were loaded onto a pH 3-10 IEF gel (available from Novex) and separated in Novex's IEF buffers according to Novex's standard procedure (60 min at 100 V; then 60 min at 200 V; and then 30 min at 500 V). Following electrophoresis, part of the gel was stained for CE activity using the method described above in Example 2. The remaining portion of the gel was blotted onto PVDF membrane by reversing the orientation of the gel and membrane so that positively charged proteins migrated to the membrane, electrophoresing the protein for 60 min at 10 V, using 0.7% acetic acid as the transfer buffer. The membrane was stained as described above in Section A. After the membrane was dried, stained protein bands on the membrane were compared to bands on the gel tested for CE activity to identify corresponding bands. Protein bands on the membrane corresponding to proteins having CE activity were excised and submitted to N-terminal sequencing as described in Section A.
[0202]N-terminal amino acid sequence was obtained for protein contained in two bands having pI values of about pI 4.8 and about pI 4.9. N-terminal amino acid sequence of the pI 4.8 band was DPPTVTLPQGELVGKALSNen (denoted SEQ ID NO:42) and N-terminal amino acid sequence of the pI 4.9 band was DPPTVTLP (denoted SEQ ID NO:43). A comparison of the N-terminal amino acid sequences identified here and described in Section A indicates closely related proteins having a consensus sequence of DPPTVTLPQGELVGKALTNEnGk (denoted SEQ ID NO:44).
[0203]The amino acid sequences of SEQ ID NO:39, SEQ ID NO:40, SEQ ID NO:41, SEQ ID NO:42, SEQ ID NO:43 and SEQ ID NO:44 are substantially contained within SEQ ID NO:5, SEQ ID NO:19 and SEQ ID NO:53, which are described below in Example 11.
Example 6
[0204]This example describes partial purification of CE from 3rd instar flea larvae.
[0205]Using the extract preparation methods described in Example 1 without labelling, extracts were prepared from about 50,000 bovine blood-fed 3rd instar flea larvae. The extract was then further purified over a p-aminobenzamidine linked agarose bead column using the method also described in Example 1. Collected unbound protein was concentrated to about 70 ml using a 200 ml stirred cell fitted with a YM-10 membrane (available from Amicon, Beverly, Mass.). Seven ml (about 5,000 3rd instar flea larval equivalents) of the concentrated extract was used for the remainder of the purification scheme described in Example 4. Resulting fractions from the anion exchange chromatography column were tested for CE activity using the assay described above in Example 2.
[0206]The results indicated that CE activity was eluted in two overlapping peaks at about 120 mM and about 210 mM NaCl.
Example 7
[0207]This example describes the identification of JHE activity in different flea tissues.
[0208]Tissue samples were prepared as described above in Example 1 from unfed and bovine blood-fed 1st instar flea larvae, bovine blood-fed 3rd instar flea larvae, bovine blood-fed prepupal flea larvae, unfed and cat blood-fed whole adult fleas, cat blood-fed adult partial fleas and cat blood-fed adult flea midguts. About 5 tissue equivalents of each tissue was assayed for JHE activity as follows.
[0209]Unlabeled juvenile hormone (JH; available from ICN Biomedicals, Inc., Aurora, Ohio) was diluted in hexane to concentration of about 0.025 M. Labeled 10-3H-juvenile hormone (3H-JH; available from Dupont-NEN) was diluted in hexane to concentration of about 80,000 cpm/μl. A JH substrate mixture was prepared by mixing about 20 μl of unlabeled JH with about 80 μl of 3H-JH (about 5 μCi) in a 4 ml screw cap vial. The substrate mixture was then covered with nitrogen (i.e., "blanketed") and the solvent contained in the mixture was evaporated by heating the mixture at 35° C. When just dry, about 1 ml of absolute anhydrous ethanol (final concentration 5×10-4 M, or 6400 cpm/μl) was added to the vial. The substrate mixture was then stored at -20° C.
[0210]About 5 equivalents of each tissue (about 5 μl of protein) was added into the bottom of a small glass autosampler vial. About 95 μl of Tris-buffered saline (TBS) was added to each vial to bring the final volume in each vial to about 100 μl. Two control samples were also prepared by adding 100 μl TBS to two separate vials. About 1 μl of the substrate mixture described above was added to all of the vials including the control samples. The final JH concentration in each vial was about 5×10-6 M. The vials were then capped and spun in a microfuge to bring all of the liquid to the bottom of each vial. The vials were then transferred to a heat block and incubated at 35° C. for about 30 minutes. Following the incubation, enzyme activity was stopped by adding about 50 μl of methanol buffer (methanol:water:concentrated ammonium hydroxide at a 10:9:1 ratio, respectively) to each vial and removing the vials from the heat block.
[0211]To measure labeled juvenile hormone acid, about 250 μl isooctane was added to each vial. Each vial was vortexed for about 15 seconds or until an emulsion formed. Each vial was then centrifuged in a microfuge for about 1 minute to separate aqueous and organic phases. About 75 μl of the aqueous layer was removed from each vial and added to about 2 ml Eco-lume scintillation fluid (available from ICN). The amount of 3H-juvenile hormone acid contained in each vial was determined using a Beckman LS-1801 liquid scintillation counter (available from Beckman, Fullerton, Calif.).
[0212]The results shown in FIG. 5 indicated that all flea tissues tested contain active JHE. Referring to Example 2, the level of CE activity differed from JHE activity in various tissue samples. The combined JHE and CE data indicated the differential expression of these two enzymatic activities during the development of a flea.
Example 8
[0213]This example describes the purification of JHE protein from cat blood-fed adult midguts.
[0214]About 23,000 cat blood-fed adult midguts were collected and prepared using the method described in Example 1. The extract was then added in 4 aliquots to columns containing about 3 to about 5 ml of p-aminobenzamidine linked agarose beads (available from Sigma-Aldrich), equilibrated in 50 mM Tris (pH 8.0), 100 mM CaCl2, 400 mM NaCl, and incubated overnight at 4° C. The columns were then washed with about 15 to about 125 ml of the equilibration Tris buffer to remove unbound protein. The collected unbound protein was pooled and then concentrated to a volume of about 5 ml using an Ultrafree-20 10 kD centrifugal concentrator (available from Millipore, Bedford, Mass.) and filtered sequentially through a 0.2 μm centrifugal ultrafiltration membrane (available from Lida, Kenosha, Wis.) to clarify the sample for chromatography.
[0215]Aliquots of about 0.5 ml were loaded onto a Superdex 200 HR gel filtration column using the method described in Example 4. Repeated runs were performed until about 10 ml of each fraction was collected. The fractions were analyzed for JHE activity using the assay described in Example 7. In preparation for anion exchange chromatography, fractions having JHE activity (Ve=17-18 ml) were combined and dialyzed overnight against about 1 L of 20 mM Tris buffer, pH 8.0, containing 10 mM NaCl. The sample was then loaded onto a Poros 10 HQ anion exchange column using the method described in Example 4. Resulting fractions were tested for JHE activity as described in Example 7.
[0216]The results indicated that midgut JHE activity was eluted from the anion exchange column in a single peak at about 120 mM NaCl.
Example 9
[0217]This example describes partial purification of JHE from prepupal flea larvae and 3rd instar larvae.
[0218]A. JHE Purification from Prepupal Tissue.
[0219]Using the extract preparation methods described in Example 1, gel filtration fractions were obtained using a Superdex 200 HR gel filtration column (available from Pharmacia) using the method described in Example 4, from about 15,000 bovine blood-fed prepupal flea larvae. The fractions were analyzed for JHE activity using the assay described above in Example 7. Those fractions containing protein having JHE activity (Ve=16-18 ml) were combined and dialyzed using the method described in Example 8.
[0220]The fractions were then further purified by passing the fractions over a Bio-Scale S2 cation exchange column (available from BioRad) at a rate of about 0.5 ml/min. The column was washed with MES until all unbound protein was eluted. Bound protein was then eluted with a linear gradient of 20 mM MES buffer, pH 6.0, containing 10 mM NaCl to 1 M NaCl. Resulting fractions were assayed for JHE activity using the method described in Example 7. The results indicated that proteins having JHE activity using prepupal tissue eluted from the column in about 200 to 300 mM NaCl.
[0221]The fractions containing JHE activity were combined and the pH adjusted to pH 7 using 0.5 M Tris buffer (pH 8.0). The fractions were then dialyzed twice against about 1 liter of 10 mM phosphate buffer (pH 7.2) containing 10 mM NaCl at 4° C. The resulting dialyzed fractions were then loaded onto a Bio-Scale CHT2-I Hydroxyapatite Column (available from BioRad) at a rate of about 0.5 ml/min. Unbound protein was washed from the column using the dialysis buffer. Bound protein was then eluted with a linear gradient of from 10 mM phosphate buffer, pH 7.2, containing 10 mM NaCl to 0.5 M phosphate buffer pH 6.5 containing 10 mM NaCl. One ml fractions were collected and each tested for JHE activity by the method described in Example 7.
[0222]The results indicated that JHE eluted in 2 overlapping peaks at about 100 mM and 150 mM phosphate. These two JHE activities were designated PP JHE I and PP JHE II, and were kept separate for the remainder of the purification. Both JHE samples were dialyzed overnight against 20 mM Tris buffer (pH 8.0) containing 10 mM NaCl. The two samples were then loaded, separately, onto a 4.5 mm×50 mm Poros 10 HQ anion exchange chromatography column (available from PerSeptive Biosystems) equilibrated with 20 mM Tris buffer, pH 8.0, containing 10 mM NaCl. Unbound proteins were washed from the column using the same buffer. Bound proteins were eluted with a linear gradient of from 10 mM to 1 M NaCl in 20 mM Tris buffer, pH 8.0. Resulting fractions were tested for JHE activity using the method described in Example 7.
[0223]The results indicated that in both samples, JHE activity was eluted from the column in a single peak at about 100 mM NaCl.
[0224]B. JHE Purification from 3rd Instar Tissue
[0225]Using the procedure described above in Section A, proteins having JHE activity were obtained using about 5,000 bovine blood-fed 3rd instar flea larvae. Following purification by cation exchange, proteins having JHE activity using 3rd instar tissue were found to elute in 2 peaks. The first peak having JHE activity was not retained on the column and also exhibited CE activity (referred to herein as CE/JHE fractions). The second peak having JHE activity eluted from the column in about 100-200 mM NaCl and did not contain CE activity.
[0226]The CE/JHE fractions were pooled and adjusted to about pH 7 using 0.5 M Tris, pH 8.0. The fractions were then loaded onto a 4.5 mm×50 mm Poros 10 HQ anion exchange chromatography column (available from PerSeptive Biosystems) and the column was equilibrated in 25 mM Tris buffer, pH 6.8. The column was washed with the same buffer and bound proteins were eluted with a linear gradient of 0 to 1 M NaCl in 25 mM Tris buffer, pH 6.8. Fractions were then tested for JHE activity using the method described in Example 7. JHE activity was eluted in two overlapping peaks at about 120 mM and 210 mM NaCl. The fraction containing JHE activity also contained CE activity when tested using the method described in Example 2.
[0227]Fractions from the cation exchange column containing only JHE activity were combined, diluted in 20 mM Tris buffer, pH 8.0 containing 10 mM NaCl, and concentrated to about 5 ml. The fractions were purified on a Poros 10 HQ anion exchange chromatography column as described immediately above. Fractions were then tested for JHE activity using the method described in Example 7. The JHE activity was eluted in a single peak at about 120 mM. The peak contained no detectable CE activity.
Example 10
[0228]This example describes the purification of JHE protein from unfed adult midguts.
[0229]About 16,000 unfed adult midguts were collected in 20 mM Tris buffer (pH 7.7), containing 130 mM NaCl, 1 mM sodium EDTA, 1 mM PEFABLOC® (available from Boehringer Mannheim, Indianapolis, Ind.), 1 microgram/ml (μg/ml) leupeptin and 1 μg/ml pepstatin. The midguts were homogenized by freeze-fracture and sonication, and then centrifuged at about 14,000×g for 20 min. The soluble material from the centrifugation step was recovered. The soluble material was then concentrated to about 1 ml using an Ultrafree-20 10 kD centrifugal concentrator (available from Millipore) and filtered sequentially through a 0.2 μm centrifugal ultrafiltration membrane to clarify the sample for chromatography. Aliquots of about 0.5 ml were loaded onto a Superdex 200 HR gel filtration column using the method described in Example 4. Repeated column runs were performed until about 2 ml of each fraction was collected. The fractions were analyzed for JHE activity using the assay described in Example 7. In preparation for cation exchange chromatography, fractions having JHE activity (Ve=15-17 ml) were combined and dialyzed overnight against about 1 L of 20 mM MES buffer, pH 6.0, containing 10 mM NaCl. The sample was then applied to a Bio-Scale S2 cation exchange column using the method described in Example 4. Fractions of eluate were assayed for JHE activity using the method described in Example 7.
[0230]The results indicate that JHE is present in unfed midguts in two forms, one that is not retained on the cation exchange column and one that is bound to the column under low salt conditions at about 100 mM NaCl. The form that was not retained under low salt conditions was shown to have general CE activity using the methods described in Example 2.
Example 11
[0231]This example describes the identification of certain esterase nucleic acid molecules of the present invention.
[0232]Several flea esterase nucleic acid molecules, representing one or more partial flea esterase genes, were PCR amplified from a flea mixed instar cDNA library or a flea prepupal cDNA library. The flea mixed instar cDNA library was produced using unfed 1st instar, bovine blood-fed 1st instar, bovine blood-fed 2nd instar and bovine blood-fed 3rd instar flea larvae (this combination of tissues is referred to herein as mixed instar larval tissues for purposes of this example). The flea prepupal cDNA library was produced using prepupal flea larvae. For each library, total RNA was extracted from mixed instar or prepupal tissue, respectfully, using an acid-guanidinium-phenol-chloroform method similar to that described by Chomczynski et al., 1987, Anal. Biochem. 162, p. 156-159. Approximately 5,164 mixed instar larvae or 3,653 prepupal larvae were used in each RNA preparation. Poly A+ selected RNA was separated from each total RNA preparation by oligo-dT cellulose chromatography using POLY(A)QUICK® mRNA isolation kits (available from Stratagene Cloning Systems, La Jolla, Calif.), according to the method recommended by the manufacturer.
[0233]A mixed instar cDNA expression library and a prepupal cDNA expression library were constructed in lambda (λ) UNI-ZAP®XR vector (available from Stratagene Cloning Systems) using STRATAGENE® ZAP-cDNA® Synthesis Kit protocol. About 6.34 μg of mixed instar poly A+ RNA were used to produce the mixed instar library and about 6.72 μg of prepupal poly A+ RNA were used to produce the prepupal library. The resultant mixed instar library was amplified to a titer of about 2.17×1010 pfu/ml with about 97% recombinants. The resultant prepupal library was amplified to a titer of about 3.5×1010 pfu/ml with about 97% recombinants.
[0234]A pair of primers was used to amplify DNA from the cDNA libraries. A sense vector primer T-3X (corresponding to the vector in which nucleic acid molecules of the present invention had been ligated), having the nucleic acid sequence AATTAACCCT CACTAAAGGG (available from Gibco BRL, Gaithersburg, Md.; denoted SEQ ID NO:45), was used in combination with a degenerate primer, the design of which was based on a highly conserved esterase amino acid sequence (disclosed in Hanzlik et al., J. Biol. Chem. 264:12419-12423, 1989; I Y/H G G G F/L) located in a region downstream from the mature amino terminus in a number of known esterases. The degenerate primer, referred to herein as FCEF, is an anti-sense primer having the nucleic acid sequence ARDCCDCCDC CRTRDAT (R indicating an A or G; and D indicating an A, G or T; denoted SEQ ID NO:46). The resultant PCR products from the mixed instar cDNA library, obtained using standard PCR conditions (e.g., Sambrook et al., ibid.), were about 550 nucleotides. The resultant PCR products from the prepupal cDNA library were from about 500 nucleotides to about 860 nucleotides.
[0235]A. PCR Products.
[0236]PCR products were gel purified and cloned into the TA VECTOR® (available from Invitrogen Corp., San Diego, Calif.). Approximately 8 clones were identified from the prepupal library and 6 clones were identified from the mixed instar library. These nucleic acid molecules were subjected to nucleic acid sequencing using the Sanger dideoxy chain termination method, as described in Sambrook et al., ibid.
[0237]1. Flea esterase clone 1 isolated from the mixed instar cDNA library was determined to comprise nucleic acid molecule nfE1401, the nucleic acid sequence of the coding strand which is denoted herein as SEQ ID NO:1. Translation of SEQ ID NO:1 suggests that nucleic acid molecule nfE1401 encodes a non-full-length flea esterase protein of about 103 amino acids, referred to herein as PfE1103, having amino acid sequence SEQ ID NO:2, assuming an initiation codon spanning from nucleotide 92 through nucleotide 94 of SEQ ID NO:1. The complement of SEQ ID NO:1 is represented herein by SEQ ID NO:3. Comparison of amino acid sequence SEQ ID NO:2 (i.e., the amino acid sequence of PfE1103) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:2, showed the most homology, i.e., about 33% identity, between SEQ ID NO:2 and alpha esterase protein from Drosophila melanogaster.
[0238]2. Flea esterase clone 2 isolated from the mixed instar cDNA library was determined to comprise nucleic acid molecule nfE2364, the nucleic acid sequence of the coding strand which is denoted herein as SEQ ID NO:4. Translation of SEQ ID NO:4 suggests that nucleic acid molecule nfE2364 encodes a non-full-length flea esterase protein of about 121 amino acids, referred to herein as PfE2121, having amino acid sequence SEQ ID NO:5, assuming the first codon spans from nucleotide 2 through nucleotide 4 of SEQ ID NO:4. The complement of SEQ ID NO:4 is represented herein by SEQ ID NO:6. Comparison of nucleic acid sequence SEQ ID NO:4 with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:4 showed the most homology, i.e., about 43% identity, between SEQ ID NO:4 and a H. virescens JHE gene. Comparison of amino acid sequence SEQ ID NO:5 (i.e., the amino acid sequence of PfE2121) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:5, showed the most homology, i.e., about 38% identity, between SEQ ID NO:5 and alpha esterase protein from Drosophila melanogaster.
[0239]3. Flea esterase clone 3 isolated from the prepupal cDNA library was determined to comprise nucleic acid molecule nfE3421, the nucleic acid sequence of the coding strand which is denoted herein as SEQ ID NO:7. Translation of SEQ ID NO:7 suggests that nucleic acid molecule nfE3421 encodes a non-full-length flea esterase protein of about 103 amino acids, referred to herein as PfE3103, having amino acid sequence SEQ ID NO:8, assuming an initiation codon spanning from nucleotide 113 through nucleotide 115 of SEQ ID NO:7. The complement of SEQ ID NO:7 is represented herein by SEQ ID NO:9. Comparison of nucleic acid sequence SEQ ID NO:7 with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:7 showed the most homology, i.e., about 53% identity, between SEQ ID NO:7 and a Torpedo marmorata acetylcholinesterase gene. Comparison of amino acid sequence SEQ ID NO:8 (i.e., the amino acid sequence of PfE3103) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:8, showed the most homology, i.e., about 39% identity, between SEQ ID NO:5 and alpha esterase protein from Drosophila melanogaster.
[0240]4. Flea esterase clone 4 isolated from the prepupal cDNA library was determined to comprise nucleic acid molecule nfE4524, the nucleic acid sequence of the coding strand which is denoted herein as SEQ ID NO:10. Translation of SEQ ID NO:10 suggests that nucleic acid molecule nfE4524 encodes a non-full-length flea esterase protein of about 137 amino acids, referred to herein as PfE4137, having amino acid sequence SEQ ID NO:11, assuming an initiation codon spanning from nucleotide 113 through nucleotide 115 of SEQ ID NO:10. The complement of SEQ ID NO:10 is represented herein by SEQ ID NO:12. Comparison of nucleic acid sequence SEQ ID NO:10 with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:10 showed the most homology, i.e., about 47% identity, between SEQ ID NO:10 and an Anas platyrhyncos thioesterase B gene. Comparison of amino acid sequence SEQ ID NO:11(i.e., the amino acid sequence of PfE4137) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:11, showed the most homology, i.e., about 30% identity, between SEQ ID NO: 11 and Leptinotarsa decemlineata acetylcholinesterase.
[0241]B. cDNA Clones.
[0242]Certain amplified PCR fragments were used as probes to identify full-length flea esterase genes in the prepupal cDNA library.
[0243]1. Nucleic acid molecule nfE2364 was labeled with 32P and used as a probe to screen the mixed instar cDNA library described in Section A, using standard hybridization techniques. Two clones were isolated. A first clone included about a 2300-nucleotide insert, referred to herein as nfE52300. Nucleic acid sequence was obtained using standard techniques from nfE52300, to yield a flea esterase nucleic acid molecule named nfE51982 having a nucleic acid sequence of the coding strand which is denoted herein as SEQ ID NO:13. Translation of SEQ ID NO:13 suggests that nucleic acid molecule nfE51982 encodes a non-full-length flea esterase protein of about 505 amino acids, referred to herein as PfE5505 having amino acid sequence SEQ ID NO:14, assuming the first codon spans from nucleotide 1 through nucleotide 3 of SEQ ID NO:13 and the stop codon spans from nucleotide 1518 through nucleotide 1520 of SEQ ID NO:13. The complement of SEQ ID NO:13 is represented herein by SEQ ID NO:15. The amino acid sequence of PfE5505 (i.e., SEQ ID NO:14) predicts that PfE5505 has an estimated molecular weight of about 56.8 kD and an estimated pI of about 5.5. The nucleic acid molecule representing the coding region for PfE5505 is referred to herein as nfE51515; the nucleic acid sequences of the coding strand and the complementary strand are represented by SEQ ID NO:16 and SEQ ID NO:17, respectively.
[0244]The nucleic acid sequence of nfE51982 was used to design primers to use in combination with a vector primer to PCR amplify the 5' terminal fragment of the remainder of the flea esterase coding region from the flea mixed instar cDNA library. A pair of primers was used to amplify DNA from the cDNA library. A sense vector primer T3-X (corresponding to the vector in which nucleic acid molecules of the present invention had been ligated), having the nucleic acid sequence 5' AATTAACCCT CACTAAAGGG 3' (denoted SEQ ID NO:45), was used in combination with an anti-sense primer M6/M265', having the nucleic acid sequence 5' GTGCGTACAC GTTTACTACC 3' (denoted SEQ ID NO:56). The resultant PCR product from the mixed instar cDNA library, obtained using standard PCR conditions (e.g., Sambrook et al., ibid.), were about 354 nucleotides.
[0245]The PCR product was subjected to DNA sequencing analysis, and a composite sequence representing a full-length flea esterase coding region was deduced. The nucleic acid sequence of the composite nucleic acid molecule, referred to herein as nfE52144 is denoted herein as SEQ ID NO:57+Translation of SEQ ID NO:57 suggests that nucleic acid molecule nfE52144 encodes a full-length flea esterase protein of about 550 amino acids, referred to herein as PfE5550, having amino acid sequence SEQ ID NO:58, assuming an open reading frame in which the initiation codon spans from nucleotide 30 through nucleotide 32 of SEQ ID NO:57 and the stop codon spans from nucleotide 1680 through nucleotide 1682 of SEQ ID NO:57. The complement of SEQ ID NO:57 is represented herein by SEQ ID NO:59. The coding region encoding PfE5550 is represented by the nucleic acid molecule nfE51650, having a coding strand with the nucleic acid sequence represented by SEQ ID NO:60 and a complementary strand with nucleic acid sequence SEQ ID NO:61. The amino acid sequence of PfE5550 (i.e., SEQ ID NO:58) predicts that PfE5550 has an estimated molecular weight of about 61.8 kD and an estimated pI of about 5.5.
[0246]Comparison of nucleic acid sequence SEQ ID NO:57 with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:57 showed the most homology, i.e., about 41% identity, between SEQ ID NO:57 and a M. persicae esterase FE4 mRNA sequence. Comparison of amino acid sequence SEQ ID NO:58 (i.e., the amino acid sequence of PfE5550) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:58 showed the most homology, i.e., about 36% identity between SEQ ID NO:58 and Drosophila melanogster alpha esterase protein.
[0247]A second clone included about a 1900 nucleotide insert, referred to herein as nfE61900. Nucleic acid sequence was obtained using standard techniques from nfE61900, to yield a flea esterase nucleic acid molecule named nfE61792 having a nucleic acid sequence of the coding strand which is denoted herein as SEQ ID NO:18. Translation of SEQ ID NO:18 suggests that nucleic acid molecule nfE61792 encodes a full-length flea esterase protein of about 550 amino acids, referred to herein as PfE6550, having amino acid sequence SEQ ID NO:19, assuming an open reading frame in which the initiation codon spans from nucleotide 49 through nucleotide 51 of SEQ ID NO:18 and a stop codon spanning from nucleotide 1699 through nucleotide 1701 of SEQ ID NO:18. The complement of SEQ ID NO: 8 is represented herein by SEQ ID NO:20. The coding region encoding PfE6550, is represented by nucleic acid molecule nfE61650, having a coding strand with the nucleic acid sequence represented by SEQ ID NO:21 and a complementary strand with nucleic acid sequence SEQ ID NO:22. The proposed mature protein, denoted herein as PfE6530, contains about 530 amino acids which is represented herein as SEQ ID NO:53. The nucleic acid molecule encoding PfE6530 is denoted herein as nfE61590 and has a coding strand having the nucleic acid sequence SEQ ID NO:23. The amino acid sequence of PfE6550 (i.e., SEQ ID NO:19) predicts that PfE6550 has an estimated molecular weight of about 61.8 kD and an estimated pI of about 5.5.
[0248]Comparison of nucleic acid sequence SEQ ID NO:18 with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:18 showed the most homology, i.e., about 41% identity, between SEQ ID NO:18 and a Myzus pericae esterase gene. Comparison of amino acid sequence SEQ ID NO:19 (i.e., the amino acid sequence of PfE6550) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:19 showed the most homology, i.e., about 28% identity between SEQ ID NO:19 and Drosophila melanogaster alpha esterase protein.
[0249]2. Nucleic acid molecule nfE4524 was labeled with 32P and used as a probe to screen the prepupal cDNA library described in Example 11, using standard hybridization techniques (e.g., Sambrook et al., ibid.). Two clones were isolated. A first clone included about a 3000 nucleotide insert, referred to herein as nfE73000. Nucleic acid sequence was obtained using standard techniques from nfE73000, to yield a flea esterase nucleic acid molecule named nfE72836 having a nucleic acid sequence of the coding strand which is denoted herein as SEQ ID NO:24. Translation of SEQ ID NO:24 suggests that nucleic acid molecule nfE72836 encodes a full-length flea esterase protein of about 596 amino acids, referred to herein as PfE7596, having amino acid sequence SEQ ID NO:25, assuming an open reading frame in which the initiation codon spans from nucleotide 99 through nucleotide 101 of SEQ ID NO:24 and a stop codon spanning from nucleotide 1887 through nucleotide 1889 of SEQ ID NO:25. The complement of SEQ ID NO:24 is represented herein by SEQ ID NO:26. The coding region encoding PfE7596, is represented by nucleic acid molecule nfE71788, having a coding strand with the nucleic acid sequence represented by SEQ ID NO:28 and a complementary strand with nucleic acid sequence SEQ ID NO:29. The proposed mature protein, denoted herein as PfE7570, contains about 570 amino acids which is represented herein as SEQ ID NO:54. The nucleic acid molecule encoding PfE7570 is denoted herein as nfE71710 and has a coding strand having the nucleic acid sequence SEQ ID NO:27. The amino acid sequence of PfE7596 (i.e., SEQ ID NO:25) predicts that PfE7596 has an estimated molecular weight of about 68.7 kD and an estimated pI of about 6.1.
[0250]Comparison of nucleic acid sequence SEQ ID NO:24 with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:24 showed the most homology, i.e., about 48% identity, between SEQ ID NO:24 and an Anas platychyncos thioesterase B gene. Comparison of amino acid sequence SEQ ID NO:25 (i.e., the amino acid sequence of PfE7596) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:25 showed the most homology, i.e., about 27% identity between SEQ ID NO:25 and Drosophila melanogaster alpha esterase protein.
[0251]A second clone included about a 3000 nucleotide insert, referred to herein as nfE83000. Nucleic acid sequence was obtained using standard techniques from nfE83000, to yield a flea esterase nucleic acid molecule named nfE82801 having a nucleic acid sequence of the coding strand which is denoted herein as SEQ ID NO:30. Translation of SEQ ID NO:30 suggests that nucleic acid molecule nfE82801 encodes a full-length flea esterase protein of about 595 amino acids, referred to herein as PfE8595, having amino acid sequence SEQ ID NO:31, assuming an open reading frame in which the initiation codon spans from nucleotide 99 through nucleotide 101 of SEQ ID NO:30 and a stop codon spanning from nucleotide 1884 through nucleotide 1886 of SEQ ID NO:30. The complement of SEQ ID NO:30 is represented herein by SEQ ID NO:32. The coding region encoding PfE8595, is represented by nucleic acid molecule nfE81785, having a coding strand with the nucleic acid sequence represented by SEQ ID NO:34 and a complementary strand with nucleic acid sequence SEQ ID NO:35. The proposed mature protein, denoted herein as PfE8570, contains about 570 amino acids which is represented herein as SEQ ID NO:55. The nucleic acid molecule encoding PfE8570 is denoted herein as nfE81710 and has a coding strand having the nucleic acid sequence SEQ ID NO:33. The amino acid sequence of PfE8595 (i.e., SEQ ID NO:31) predicts that PfE8595 has an estimated molecular weight of about 68.6 kD and an estimated pI of about 6.1.
[0252]Comparison of nucleic acid sequence SEQ ID NO:30 with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:30 showed the most homology, i.e., about 46% identity, between SEQ ID NO:30 and a Mus musculus carboxyl ester lipase gene. Comparison of amino acid sequence SEQ ID NO:31 (i.e., the amino acid sequence of PfE8595) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:31 showed the most homology, i.e., about 28% identity between SEQ ID NO:31 and estalpha-2 esterase of Culex pipiens quinque fasciatus.
[0253]3. Nucleic acid molecule nfE3421 was labeled with 32P and used as a probe to screen the prepupal cDNA library using standard hybridization techniques (e.g., Sambrook et al., ibid.). Two clones were isolated. One clone included about a 1900 nucleotide insert, referred to herein as nfE91900. Nucleic acid sequence was obtained using standard techniques from nfE91900, to yield a flea esterase nucleic acid molecule named nfE92007 having nucleic acid sequence of the coding strand which is denoted herein as SEQ ID NO:36. Translation of SEQ ID NO:36 suggests that nucleic acid molecule nfE92007 encodes a full-length flea esterase protein of about 528 amino acids, referred to herein as PfE9528, having amino acid sequence SEQ ID NO:37, assuming an open reading frame in which the initiation codon spans from nucleotide 11 through nucleotide 13 of SEQ ID NO:36 and a stop codon spanning from nucleotide 1595 through nucleotide 1597 of SEQ ID NO:36. The complement of SEQ ID NO:36 is represented herein by SEQ ID NO:38. The coding region encoding PfE9528, is represented by nucleic acid molecule nfE91584, having a coding strand with the nucleic acid sequence represented by SEQ ID NO:51 and a complementary strand with nucleic acid sequence SEQ ID NO:52. The amino acid sequence of PfE9528 (i.e., SEQ ID NO:37) predicts that PfE9528 has an estimated molecular weight of about 60 kD and an estimated pI of about 5.43.
[0254]Comparison of nucleic acid sequence SEQ ID NO:36 with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:36 showed the most homology, i.e., about 47% identity, between SEQ ID NO:36 and a hamster mRNA for carboxylesterase precursor gene. Comparison of amino acid sequence SEQ ID NO:37 (i.e., the amino acid sequence of PfE9528) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:37 showed the most homology, i.e., about 37% identity between SEQ ID NO:37 and alpha esterase protein from Drosophila melanogaster.
[0255]As is the case for any of the nucleic acid molecules described in this example, variations between sequences may be due to a number of factors, such as but not limited to, sequencing errors or allelic variation.
[0256]4. Nucleic acid molecule nfE1401 was labeled with 32P and used as a probe to screen the mixed instar cDNA library using standard hybridization techniques (e.g., Sambrook et al., ibid.). A clone was isolated that included about a 2000 nucleotide insert, referred to herein as nfE102000. Nucleic acid sequence was obtained using standard techniques from nfE102000, to yield a flea esterase nucleic acid molecule named nfE101987 having nucleic acid sequence of the coding strand which is denoted herein as SEQ ID NO:67. Translation of SEQ ID NO:67 suggests that nucleic acid molecule nfE101987 encodes a full-length flea esterase protein of about 530 amino acids, referred to herein as PfE10530, having amino acid sequence SEQ ID NO:68, assuming an open reading frame in which the initiation codon spans from nucleotide 231 through nucleotide 233 of SEQ ID NO:67 and a stop codon spanning from nucleotide 1821 through nucleotide 1823 of SEQ ID NO:67. The complement of SEQ ID NO:67 is represented herein by SEQ ID NO:69. The coding region encoding PfE10530, is represented by nucleic acid molecule nfE101590, having a coding strand with the nucleic acid sequence represented by SEQ ID NO:70 and a complementary strand with nucleic acid sequence SEQ ID NO:71. The amino acid sequence of PfE10530 (i.e., SEQ ID NO:68) predicts that PfE10530 has an estimated molecular weight of about 59.5 kD and an estimated pI of about 5.5.
[0257]Comparison of nucleic acid sequence SEQ ID NO:67 with nucleic acid sequences reported in GENBANK® indicates that SEQ ID NO:67 showed the most homology, i.e., about 48% identity, between SEQ ID NO:67 and a lucilia cuprina alpha esterase gene (genembl #U56636) gene. Comparison of amino acid sequence SEQ ID NO:68 (i.e., the amino acid sequence of PfE10530) with amino acid sequences reported in GENBANK® indicates that SEQ ID NO:68 showed the most homology, i.e., about 30% identity between SEQ ID NO:68 and Culex pipens esterase b1 precurser protein (SWISSPROT®# P16854).
[0258]As is the case for any of the nucleic acid molecules described in this example, variations between sequences may be due to a number of factors, such as but not limited to, sequencing errors or allelic variation.
Example 12
[0259]This Example demonstrates the production of esterase proteins of the present invention in E. coli cells.
[0260]A. Flea esterase protein PHIS-PfE7570 and flea esterase protein PHIS-PfE8570 were produced in the following manner. A pair of primers was used to amplify DNA from flea esterase nucleic acid molecule nfE72836 or nfE82801 produced as described in Example 11. A sense primer containing an XhoI site (shown in bold) having the nucleic acid sequence 59 TGTGCTCGAG ATGGGATAAC CTAGATCAGC ATTTGTGC 3' (denoted SEQ ID NO:47), was used in combination with an anti-sense primer containing a KpnI site (shown in bold) having the nucleic acid sequence 5' TTAAGGTACC TCATCTAATA CTTCCTTCAT TACAG 3' (denoted SEQ ID NO:48). A PCR product was derived from nfE72836, and is referred to herein as nfE71710, having nucleic acid sequence SEQ ID NO:27. The PCR product was digested with XhoI and KpnI restriction endonucleases, gel purified and subcloned into expression vector pTrcHisB (available from Invitrogen). The resultant recombinant molecule, referred to herein as pTrc-nfE71710, was transformed into E. coli HB101 competent cells (available from Gibco BRL) to form recombinant cell E. coli:pTrc-nfE71710.
[0261]The PCR product derived from nfE82801 using the primers is referred to herein as nfE81710, having nucleic acid sequence SEQ ID NO:33. PCR product nfE81710 was digested with XhoI and KpnI restriction endonucleases, gel purified and subcloned into expression vector pTrcHisB. The resultant recombinant molecule, referred to herein as pTrc-nfE81710, was transformed into E. coli HB101 competent cells to form recombinant cell E. coli:pTrc-nfE81710.
[0262]The recombinant cells were cultured in enriched bacterial growth medium containing 0.1 mg/ml ampicillin and 0.1% glucose at about 32° C. When the cells reached an OD600 of about 0.4-0.5, expression of recombinant protein was induced by the addition of 0.5 mM isopropyl-B-D-thiogalactoside (IPTG), and the cells were cultured for about 2 hours at about 32° C. Immunoblot analysis of recombinant cell E. coli:pTrc-nfE71710 and E. coli:pTrc-nfE81710 lysates using a T7 tag monoclonal antibody (available from Novagen, Inc., Madison, Wis.) directed against the fusion portion of the recombinant PHIS-PfE7570 and PHIS-PfE8570 fusion proteins identified proteins of appropriate size, namely an about 65 kD protein for each fusion protein.
[0263]B. Flea esterase protein PHIS-PfE6540 was produced in the following manner.
[0264]A pair of primers was used to amplify DNA from flea esterase nucleic acid molecule nfE61792 produced as described in Example 11. A sense primer containing an XhoI site having the nucleic acid sequence 5' AAACTCGAGT CCCCCGACTG TAACTTTGC 3' (denoted SEQ ID NO:62; XhoI site shown in bold), was used in combination with an anti-sense primer containing a PstI site having the nucleic acid sequence 5' TCATCTGCAG TTATTGACTG TGCAAAGTTT TTGTGG 3' (denoted SEQ ID NO:63; PstI site shown in bold). A PCR product was derived from nfE61792, and is referred to herein as nfE61488, having nucleic acid sequence SEQ ID NO:76. The PCR product was digested with XhoI and PstI restriction endonucleases, gel purified and subcloned into expression vector lambdaPR/T2ori/S10HIS-RSET-A9, that had been digested with XhoI and PstI and dephosphorylated. The resultant recombinant molecule, referred to herein as pCro-nfE61488, was transformed into E. coli HB101 competent cells (available from Gibco BRL) to form recombinant cell E. coli:pCro-nfE61488.
[0265]The recombinant cells were cultured using the method generally described in Section A of this example, except that the cells were grown under heat shift conditions rather than in the presence of IPTG. The cells were grown at 32° C. for about 2 hours, and then grown at 42° C. Immunoblot analysis of recombinant cell E. coli:pCro-nfE61488 lysate using a T7 tag monoclonal antibody directed against the fusion portion of the recombinant PHIS-PfE6540 fusion protein identified proteins of appropriate size, namely an about 60 kD protein for each fusion protein.
[0266]Expression of the recombinant PHIS-PfE6540 fusion protein was improved by transforming supercoiled plasmid pCro-nfE61488 DNA harvested from E. coli:pCro-nfE61488 cells into the BL-21 strain of E. coli (available from Novagen). The amount of expression PHIS-PfE6540 was confirmed by immunoblot using the method described immediately above.
[0267]E. coli cells expressing PHIS-PfE6540 protein were harvested from about 2 liters of media and suspended in about 140 ml of 50 mM Tris, pH 8.0, 50 mM NaCl, 0.1 mM phenylmethylsulfonylfluoride (PMSF) (Solubilization Buffer). The cells were broken by passage through a microfluidizer at 30 psi for 30 cycles. The sample was centrifuged at about 16,000×g for 30 min at 4° C. The supernatant S1) was recovered and the pellet was resuspended in about 80 ml of Solubilization Buffer and centrifuged at about 16,000×g for 30 min at 4° C. The supernatant (S2) was recovered and the pellet was resuspended in about 80 ml of Solubilization Buffer containing 0.1% Triton-X100 and centrifuged at about 16,000×g for 30 min at 4° C. The supernatant (S3) was recovered and the pellet was resuspended in about 140 mls 50 mM Tris, pH 8.0, 8 M Urea, 0.1 M PMSF and centrifuged at about 16,000×g. The supernatant (S4) was recovered and the pellet was resuspended in 40 mls 50 mM Tris, 8 M Urea, 0.1 M PMSF. Aliquots of each pellet and supernatant were analyzed by SDS-PAGE and immunoblot using the T7 tag monoclonal antibody described above. The results indicated that the PHIS PfE6540 protein was located in the final supernatant (S4). The PHIS-PfE6540 protein was loaded onto a 5.0 ml, Metal chelating HiTrap column charged with NiCl2 (obtained from Pharmacia Biotech Inc., Piscataway, N.J.), previously equilibrated with 50 mM Tris, 1 mM PMSF, 1 mM β-mercaptoethanol (βME), 8 M urea, pH 8.0 (Buffer A). The column was washed with 10 column volumes (CV) Buffer A and then with 10 cv with 50 mM Tris, 25 mM sodium acetate, 1 mM PMSF, 1 mM βME, 8 M urea, pH 6.0 (Buffer B) to remove loosely bound proteins. Bound PHIS-PfE6540 protein was eluted with 10 cv of 50 mM Tris, 25 mM sodium acetate, 1 mM PMSF, 1 mM βME, 8 M urea, pH 4.0 (Buffer C). Column fractions were analyzed for the presence of PHIS-PfE6540 protein by immunoblot using the T7 tag monoclonal antibody as described above. The results indicated that the majority of the PHIS-PfE6540 protein was eluted by Buffer C. The fractions containing the PHIS-PfE6540 protein were combined and loaded onto a 5 ml SP-Sepharose HiTrap column (obtained from Pharmacia Biotech Inc.) previously equilibrated with 50 mM Tris, 25 mM Sodium Acetate, 1 mM PMSF, 1 mM βME, 8 M Urea, pH 4.5 (SP-Sepharose Buffer). The column was washed with SP-Sepharose Buffer until most of the unbound protein was removed. Bound protein was eluted with an increasing salt gradient to 1 M NaCl over 100 ml (20 cv) in SP-sepharose buffer. Column fractions were analyzed for the presence of PHIS-PfE6540 protein by immunoblot using the T7 tag monoclonal antibody as described above. The results indicated that the PHIS-PfE6540 protein was eluted at about 0.75 M NaCl.
[0268]The purified PHIS-PfE61488 protein was used to produce an anti-M6 polyclonal antiserum as follows. Rabbits were immunized with PHIS-PfE61488 protein diluted to a concentration of about 0.1 mg/ml in PBS. One milliliter of the dilution was mixed 1:1 mix with Complete Freunds Adjuvant. In the primary immunization, about 500 μl of the 1:1 mmix was injected subcutaneously into 5 different sites (0.1 ml/site) and 500 μl was injected intradermally into 5 different sites (0.1 ml/site) on the rabbit. Booster shots were administered to the rabbit intramuscularly in 4 sites using 250 μl/site of a 1:1 mix of PHIS-PfE61488 protein with Incomplete Freunds Adjuvant. The booster shots were administered at days 14 and 35. Serum samples were obtained prior to immunization (pre-bleed), and at day 14 after primary immunization and day 14 after the first and second boost.
[0269]C. Flea esterase protein PHIS-PfE9528 was produced in the following manner.
[0270]A pair of primers was used to amplify DNA from flea esterase nucleic acid molecule nfE92007 produced as described in Example 11. A sense primer containing an BamHI site having the nucleic acid sequence 5'-TTC CGG ATC CGG CTG ATC TAC AAG TGA CTT TG-3' (denoted SEQ ID NO:64; BamHI site shown in bold), was used in combination with an anti-sense primer containing a XhoI site having the nucleic acid sequence 5' TGG TAC TCG AGT CAT AAA AAT TTA TTC CAA AAT C 3' (denoted SEQ ID NO:65; XhoI site shown in bold). A PCR product was derived from nfE92007, and is referred to herein as nfE91540, having nucleic acid sequence SEQ ID NO:51. The PCR product was digested with BamI and XhoI restriction endonucleases, gel purified and subcloned into expression vector pTrcHisB (available from Invitrogen). The resultant recombinant molecule, referred to herein as pTrc-nfE91540, was transformed into E. coli HB101 competent cells (available from Gibco BRL) to form recombinant cell E. coli:pTrc-nfE9540.
[0271]The recombinant cells were cultured using the method described in Section A of this example. Immunoblot analysis of recombinant cell E. coli:pTrc-nfE91540 lysate using a T7 tag monoclonal antibody directed against the fusion portion of the recombinant PHIS-PfE9528 fusion protein identified proteins of appropriate size, namely an about 59 kD protein for each fusion protein.
[0272]Expression of the recombinant PHIS-PfE9528 fusion protein was improved by transforming supercoiled plasmid pTrc-nfE91584 DNA harvested from E. coli:pTrc-nfE91540 cells into the BL-21 strain of E. coli. The amount of expression PHIS-PfE9528 was confirmed by immunoblot using the method described immediately above.
[0273]Two liters of media from cultures of E. coli cells expressing PHIS PfE9528 protein were harvested and S4 supernatant was prepared using the method described above in section B. PHIS PfE9528 protein contained in the S4 supernatant was loaded onto a 5.0 ml, Metal chelating HiTrap column, charged with NiCl2 (available from Pharmacia Biotech Inc., Piscataway, N.J.), previously equilibrated with 50 mM Tris, 1 mM PMSF, 1 mM βME, 8 M urea, pH 8.0 (Buffer A). The column was washed with 5 cv of Buffer A until all unbound protein was removed. Bound protein was eluted with a linear gradient from Buffer A to 50 mM Tris, 1 mM PMSF, 1 mM βME, 8 M urea, 1 M NaCl, pH 4.0. Column fractions were analyzed for the presence of PHIS pfE9528 protein by immunoblot using the T7 tag monoclonal antibody as described above. The results indicated that the majority of the PHIS pfE9528 protein was eluted at about 250 mM NaCl. The fractions containing the PHIS pfE9528 protein were combined and loaded onto a C4-reversed phase column (obtained from Vydak, Hesperia, Calif.), previously equilibrated with 0.05% trifluoroacetic acid (TFA). The column was washed with 0.05% TFA until all unbound protein was removed. Bound proteins were eluted with a linear gradient from 0.05% TFA to 0.05% TFA in acetonitrile. Column fractions were analyzed for the presence of PHIS pfE9528 protein by immunoblot using the T7 tag monoclonal antibody as described above. The results indicated that the PHIS pfE9528 protein was eluted at about 40% acetonitrile. The fractions containing the PHIS pfE9528 protein were combined and loaded onto a 5 ml Q-Sepharose HiTrap column previously equilibrated with 50 mM Tris, 25 mM Sodium Acetate, 1 mM PMSF, 1 mM βME, 8 M Urea, pH 8.5 (Q-Sepharose Buffer). The column was washed with Q-Sepharose Buffer until all unbound protein was removed. Bound protein was eluted with an increasing salt gradient to 1 M NaCl over 100 ml (20 cv) in Q-sepharose buffer. Column fractions were analyzed for the presence of PHIS-PfE9528 protein by immunoblot using the T7 tag monoclonal antibody as described above. The results indicated that the PHIS-PfE9528 protein was eluted at about 0.3 M NaCl.
[0274]The purified PHIS-PfE9528 protein was used to produce an anti-P1 polyclonal antiserum as follows. Rabbits were immunized with PHIS-PfE9528 protein diluted to a concentration of about 0.1 mg/ml in PBS. One milliliter of the dilution was mixed 1:1 mix with Complete Freunds Adjuvant. In the primary immunization, about 500 μl of the 1:1 mix was injected subcutaneously into 5 different sites (0.1 ml/site) and 500 μl was injected intradermally into 5 different sites (0.1 ml/site) on the rabbit. Booster shots were administered to the rabbit intramuscularly in 4 sites using 250 μl/site of a 1:1 mix of PHIS-PfE9528 protein with Incomplete Freunds Adjuvant. The booster shots were administered at days 14 and 35. Serum samples were obtained prior to immunization (pre-bleed), and at day 14 after primary immunization and day 14 after the first and second boost.
[0275]D. Flea esterase protein PHIS-PfE7275 was produced in the following manner.
[0276]A 650-bp fragment was produced by digesting nfE72836 DNA with the restriction enzymes BamHI and BgIII. The BamHI and BgIII fragment derived from nfE72836 is referred to herein as nfE7650, having nucleic acid sequence SEQ ID NO:72 and amino acid SEQ ID NO:73. The fragment was purified using a QIAQUICK® Kit (available from Qiagen, Santa Clarita, Calif.), according to methods provided by the manufacturer. The purified fragment was subcloned into expression vector pTrcH is C which had been digested with BamHI and BgIII. The resultant recombinant molecule, referred to herein as pTrc-nfE7650 was transformed into E. coli DH-5a competent cells (available from Gibco BRL) to form recombinant cell E. coli:pTrc-nfE7650.
[0277]The recombinant cells were cultured using the method described above in section A. Immunoblot analysis of recombinant cell E. coli:pTrc-nfE7650 lysate using a T7 tag monoclonal antibody directed against the fusion portion of the recombinant PHIS-PfE7275 fusion protein identified proteins of appropriate size, namely an about 35 kD protein for each fusion protein.
[0278]Expression of the recombinant fusion protein was improved by transforming supercoiled plasmid pTrc-nfE7650 DNA harvested from E. coli:pTrc-nfE7650 cells into the BL-21 strain of E. coli. The amount of expression E. coli:pTrc-nfE7650 was confirmed by immunoblot using the method described immediately above.
Example 13
[0279]This Example demonstrates the production of esterase proteins of the present invention in eukaryotic cells.
[0280]A. Recombinant molecule pBv-nfE71788, containing a flea esterase nucleic acid molecule spanning nucleotides from about 99 through about 1886 of SEQ ID NO:24, and pBv-nfE81785, containing a flea esterase nucleic acid molecule spanning nucleotides from about 99 through about 1883 of SEQ ID NO:30 each, operatively linked to baculovirus polyhedron transcription control sequences were produced in the following manner. In order to subclone a flea esterase nucleic acid molecule into baculovirus expression vectors, flea esterase nucleic acid molecule-containing fragments were separately PCR amplified from nfE72836 or nfE82801 DNA. A PCR fragment of 1858 nucleotides, named nfE71858, was amplified from nfE72836 using a sense primer E1113 FWD having the nucleic acid sequence 5'-AAAACTGCAG TATAAATATG TTACCTCACA GTAGTG-3' (SEQ ID NO:49; PstI site shown in bold) and an antisense primer E 1113/2212 REV having the nucleic acid sequence 5'-TGCTCTAGAT TATCTAATAC TTCCTTCATT ACAG (SEQ ID NO:50; XbaI site shown in bold). A PCR fragment of 1858 nucleotides, named nfE81858, was amplified from nfE82801 using a sense primer E2212 FWD having the nucleic acid sequence 5'-AAAACTGCAG TATAAATATG TTACCTCACA GTGCATTAG-3' (SEQ ID NO:66; PstI site shown in bold), and the antisense primer E 1113/2212 REV. The N-terminal primer was designed from the pol h sequence of baculovirus with modifications to enhance expression in the baculovirus system.
[0281]In order to produce a baculovirus recombinant molecule capable of directing the production of .sub.PfE7596, the about 1,802 base pair PCR product (referred to as Bv-nfE71802) was digested with PstI and XbaI and subcloned into unique PstI and XbaI sites of pVL1392 baculovirus shuttle plasmid (available from Pharmingen, San Diego, Calif.) to produce the recombinant molecule referred to herein as pVL-nfE71802.
[0282]In order to produce a baculovirus recombinant molecule capable of directing the production of PfE8595, the about 1,792 base pair PCR product (referred to as Bv-nfE81792) was digested with PstI and XbaI and subcloned into PstI and XbaI digested to produce the recombinant molecule referred to herein as pVL-nfE81792.
[0283]The resultant recombinant molecules, pVL-nfE71802 and pVL-nfE8792, were verified for proper insert orientation by restriction mapping. Such a recombinant molecule can be co-transfected with a linear Baculogold baculovirus DNA (available from Pharmingen) into S. frugiperda Sf9 cells (available from Invitrogen) to form the recombinant cells denoted S. frugiperda:pVL-nfE71802 and S. frugiperda:pVL-fE81792. S. frugiperda:pVL-nfE71802 can be cultured in order to produce a flea esterase protein PfE7596. S. frugiperda:pVL-nfE81792 can be cultured in order to produce a flea esterase protein PfE8595.
[0284]B. Recombinant molecule pBv-PfE9528, containing a flea esterase nucleic acid molecule spanning nucleotides from 14 through 1595 of SEQ ID NO:36, operatively linked to baculovirus polyhedron transcription control sequences were produced in the following manner. In order to subclone a flea esterase nucleic acid molecule into baculovirus expression vectors, a flea esterase nucleic acid molecule-containing fragment was PCR amplified from nfE92007 DNA. A PCR fragment of about 1600 nucleotides, named nfE91600, was amplified from nfE92007 using a sense primer P121B1 Sense having the nucleic acid sequence 5'-CGC GGA TCC GCT GAT CTA CAA GTG ACT TTG C-3' (SEQ ID NO:75; BamHI site shown in bold) and an antisense primer P121B1 Anti having the nucleic acid sequence 5'-CCG AGC GGC CGC ATA AAA ATT TAT TCC AAA ATC TAA GTC G-3' (SEQ ID NO:76; NotI site shown in bold). The N-terminal primer was designed from the pol h sequence of baculovirus with modifications to enhance expression in the baculovirus system.
[0285]In order to produce a baculovirus recombinant molecule capable of directing the production of PfE9528, the about 1,600 base pair PCR product (referred to as Bv-nfE91600) was digested with BamHI and NotI and subcloned into unique BamHI and NotI sites of pVL1393 baculovirus shuttle plasmid (available from Pharmingen, San Diego, Calif.) to produce the recombinant molecule referred to herein as pVL-nfE91600.
[0286]The resultant recombinant molecule, pVL-nfE91600, was verified for proper insert orientation by restriction mapping. Such a recombinant molecule can be co-transfected with a linear Baculogold baculovirus DNA into S. frugiperda Sf9 cells to form the recombinant cells denoted S. frugiperda:pVL-nfE91600. S. frugiperda:pVL-nfE91600 can be cultured in order to produce a flea esterase protein PfE9528.
[0287]An immunoblot of supernatant from cultures of S. frugiperda:pVL-nfE91600 cells producing the flea esterase protein PfE9528 was performed using the anti-P1 polyclonal antiserum described in detail in Example 12. Blots were incubated using serum samples from the pre-bleed or from serum collected 14 days after the first boost of the rabbit. Analysis of the supernatant from cultures of S. frugiperda:pVL-nfE91600 cells identified an about 66 kD protein.
[0288]C. Recombinant molecule pBv-PfE6530, containing a flea esterase nucleic acid molecule spanning nucleotides from 50 through 1701 of SEQ ID NO:18, operatively linked to baculovirus polyhedron transcription control sequences were produced in the following manner. In order to subclone a flea esterase nucleic acid molecule into baculovirus expression vectors, a flea esterase nucleic acid molecule-containing fragment was PCR amplified from nfE61792 DNA. A PCR fragment of about 1679 nucleotides, named nfE101679, was amplified from nfE61792 using a sense primer M6M32 Sense having the nucleic acid sequence 5'-GCG AGG CCT TAT AAA TAT GTC TCG TGT TAT TTT TTT AAG TTG-3' (SEQ ID NO:75; StuI site shown in bold) and an antisense primer M6M32 Anti having the nucleic acid sequence 5'-GCA CTG CAG TTA TTG ACT GTG CAA AGT TTT TGT GG-3' (SEQ ID NO:76; PstI site shown in bold). The N-terminal primer was designed from the pol h sequence of baculovirus with modifications to enhance expression in the baculovirus system.
[0289]In order to produce a baculovirus recombinant molecule capable of directing the production of PfE6530, the about 1,679 base pair PCR product (referred to as Bv-nfE61679) was digested with StuI and PstI and subcloned into unique StuI and PstI sites of FAST BAC® baculovirus shuttle plasmid (obtained from Gibco-BRL) to produce the recombinant molecule referred to herein as pFB-nfE61679.
[0290]The resultant recombinant molecule, pFB-nfE61679, was verified for proper insert orientation by restriction mapping. Such a recombinant molecule can be transformed into E. coli strain DH10 (obtained from Gibco-BRL) according to the manufacturer's instructions. The pFB-nfE61679 isolated from the transformed DH10 cells can then be co-transfected with a linear Baculogold baculovirus DNA into S. frugiperda Sf9 cells to form the recombinant cells denoted S. frugiperda:pFB-nfE61679.
S. frugiperda:pFB-nfE61679 can be cultured in order to produce a flea esterase protein PfE6530.
[0291]An immunoblot of supernatant from cultures of S. frugiperda:pFB-nfE61679 cells producing the flea esterase FfE6530 was performed using the anti-M6 polyclonal antiserum described in detail in Example 12. Blots were incubated using serum samples from the pre-bleed or from serum collected 14 days after the first boost of the rabbit. Analysis of the supernatant from cultures of S. frugiperda:pFB-nfE61679 cells identified an about 66 kD protein.
[0292]N-terminal amino acid sequence was obtained using standard methods for the about 66 kD protein identified using the anti-M6 polyclonal antiserum. The N-terminal amino acid sequence was determined to be identical to the N-terminal amino acid sequence of SEQ ID NO:44.
Example 14
[0293]This example describes the purification of carboxylesterase protein from fed flea midguts.
[0294]About 43,000 cat blood-fed adult flea midguts were collected and prepared as previously described in Example 1. The extract was then added in 2 aliquots to columns containing about 1 to about 2 ml of p-aminobenzamidine linked agarose beads (available from Sigma-Aldrich), equilibrated in 50 mM Tris (pH 8.0), 400 mM NaCl, and incubated overnight at 4° C. The columns were then drained to remove unbound protein and the two aliquots of unbound protein were combined. The collected unbound protein was then concentrated and diafiltered into a total volume of about 16 ml of 25 mM Tris (pH 8), 10 mM NaCl using an Ultrafree-20 10 kD centrifugal concentrator (available from Millipore, Bedford, Mass.).
[0295]Aliquots of about 8 ml were loaded onto an Uno Q6 anion exchange column (available from Bio-Rad, Hercules, Calif.) equilibrated in 25 mM Tris (pH 8), 10 mM NaCl, operated on a BioLogic liquid chromatography system (available from Bio-Rad). The column was washed with 25 mM Tris (pH 8), 10 mM NaCl until all unbound protein was removed. Protein bound to the column was then eluted with a linear gradient from 10 mM to 1 M NaCl in 25 mM Tris, pH 8. Fractions were assayed for CE activity using the assay described previously. The results indicated that CE activity was eluted at about 220 mM NaCl.
[0296]Fractions containing CE activity were pooled and diafiltered into a total volume of about 3 ml of 20 mM MES buffer (2-(N-morpholino)ethanesulfonic acid), pH 6.0, containing 10 mM NaCl, in preparation for cation exchange chromatography. The sample was then applied to an Uno S1 cation exchange column (available from Bio-Rad) equilibrated in MES buffer. The column was washed with MES buffer until all unbound protein was removed. Protein bound to the column was then eluted with a linear gradient from 10 mM to 1 M NaCl in 20 mM MES buffer, pH 6. Fractions were assayed for CE activity using the assay described previously. The results indicated that CE activity was not retained on the cation exchange column using the above conditions, and all of the activity was found in the flow-through fractions.
[0297]Fractions containing CE activity were pooled and diafiltered into a total volume of about 3 ml of 25 mM Tris (pH 8), 10 mM NaCl, in preparation for an additional anion exchange chromatography step. The sample was then applied to a Bio-Scale Q2 anion exchange column (available from Bio-Rad). The column was washed with 25 mM Tris (pH 8), 10 mM NaCl until all unbound protein was removed. Protein bound to the column was then eluted with a linear gradient from 10 mM to 1 M NaCl in 25 mM Tris, pH 8. Fractions were assayed for CE activity using the assay described previously. The results indicated that CE activity was eluted at about 130 mM NaCl.
[0298]A fraction containing CE activity was diluted into a total volume of about 4 ml of 10 mM phosphate buffer, pH 7.2 containing 10 mM NaCl, in preparation for hydroxyapatite chromatography. The sample was then applied to a Bio-Scale CHT2-I column (available from Bio-Rad) at a flow rate of about 0.5 ml/min. The column was washed with 10 mM phosphate buffer, pH 7.2 containing 10 mM NaCl until all unbound protein was removed. Protein bound to the column was then eluted with a linear gradient from 10 mM phosphate buffer, pH 7.2 containing 10 mM NaCl to 0.5 M 10 mM phosphate buffer, pH 6.5 containing 10 mM NaCl. Fractions were assayed for CE activity using the assay described previously. The results indicated that CE activity was eluted at about 200 mM phosphate.
Example 15
[0299]This example describes the purification of a carboxylesterase protein from wandering flea larvae.
[0300]About 120,000 bovine blood-fed adult wandering flea larvae were homogenized in 3 batches of about 40,000 wandering larvae in each batch, in Tris buffered saline (TBS), pH 8.0 as previously described, except that about 1.2 mg of phenylthiourea was added to each ml of TBS during the extraction procedure to inhibit cross linking reactions. The extracts were dialyzed against 2 changes of about 2 L of 10 mM phosphate buffer, pH 7.2 containing 10 mM NaCl in preparation for hydroxyapatite batch chromatography. The samples were then filtered through glass ACRODISCS® (available from Gelman Sciences, Ann Arbor, Mich.) and added to 14 g of Macro-Prep Ceramic Hydroxyapatite, Type I, 40 μm beads (available from Bio-Rad), previously equilibrated in 10 mM phosphate buffer, pH 7.2 containing 10 mM NaCl. The extracts and beads were rocked at room temperature for about 30 minutes. Following incubation, the beads were centrifuged for about 5 minutes at 500×g and the supernatants removed. The beads were washed with about 40 ml 10 mM phosphate buffer, pH 7.2 containing 10 mM NaCl, centrifuged as above, and washed and centrifuged again to eliminate all unbound protein. Bound proteins were eluted by washing the beads with about 40 ml of each of 100 mM, 200 mM, 300 mM, and 400 mM phosphate buffer, pH 6.5 containing 10 mM NaCl. Following elution, the supernatants from each concentration of phosphate buffer were tested for juvenile hormone esterase activity as described previously in Example 7. The juvenile hormone esterase activity eluted at different phosphate concentrations in each batch, but the activity was generally found in the 200 mM to 300 mM phosphate fractions.
[0301]The fractions that contained the highest juvenile hormone esterase activity were combined and diafiltered into a total volume of about 50 ml of 10 mM phosphate buffer, pH 7.2 containing 10 mM NaCl using a stirred cell concentrator fitted with a YM10 ultrafiltration membrane (available from Amicon, Beverly, Mass.). Aliquots of about 5 ml to 10 ml were applied to a chromatography column containing about 10 ml of Macro-Prep Ceramic Hydroxyapatite, Type I, 20 μm beads, previously equilibrated with 10 mM phosphate buffer, pH 7.2 containing 10 mM NaCl. The column was washed with 10 mM phosphate buffer, pH 7.2 containing 10 mM NaCl until all unbound protein was removed. Protein bound to the column was then eluted with a linear gradient from 10 mM phosphate buffer, pH 7.2 containing 10 mM NaCl to 0.5 M 10 mM phosphate buffer, pH 6.5 containing 10 mM NaCl. Fractions were assayed for carboxylesterase activity using the assay described previously. The results indicated that carboxylesterase activity was eluted at about 160 mM phosphate.
[0302]The fractions that contained the highest carboxylesterase activity were combined and diafiltered into a total volume of about 15 ml of 20 mM sodium acetate buffer, pH 4.0 in preparation for cation exchange chromatography. Aliquots of about 3 ml were applied to a PolyCat A cation exchange column (available from PolyLC, Columbia, Md.) equilibrated in 20 mM sodium acetate buffer, pH 6.0, operated on a Waters high performance liquid chromatography system (available from Waters Corporation, Milford, Mass.). The column was washed with 20 mM sodium acetate buffer, pH 6.0 until all unbound protein was removed. Protein bound to the column was then eluted with a linear gradient from 20 mM sodium acetate buffer, pH 6.0 to 20 mM sodium acetate buffer, pH 6.0 containing 1 M NaCl. Fractions were assayed for CE activity using the assay described previously. The results indicated that there were two pools of CE activity. The first pool was not retained on the cation exchange column, and the second pool was eluted at about 170 mM NaCl.
[0303]The fractions from the second pool that contained the highest carboxylesterase activity were combined and diafiltered into a total volume of about 10 ml of 25 mM Tris (pH 8), 1 mM NaCl, in preparation for anion exchange chromatography. The sample was then applied to a Bio-Scale Q2 anion exchange column (available from Bio-Rad). The column was washed with 25 mM Tris (pH 8), 10 mM NaCl until all unbound protein was removed. Protein bound to the column was then eluted with a linear gradient from 10 mM to 1 M NaCl in 25 mM Tris, pH 8. Fractions were assayed for carboxylesterase activity using the assay described previously. The results indicated that carboxylesterase activity was eluted at about 350 mM NaCl.
[0304]Fractions containing carboxylesterase activity were combined and concentrated to about 175 ml using a Centricon 10 centrifugal concentrator (available from Amicon, Beverly, Mass.) in preparation for size exclusion chromatography. The sample was applied to a Bio-Select SEC 125-5 size exclusion chromatography column (available from Bio-Rad), previously equilibrated in TBS, pH 7.2. About 250 μl fractions were then collected. The fractions were assayed for carboxylesterase activity using the assay described previously. The results indicated that the carboxylesterase activity was eluted in about 5.5 to 6 ml of buffer, corresponding to a molecular weight of about 40 to 100 kDa based on the elution volumes of gel filtration molecular weight standard proteins (available from Sigma-Aldrich, St. Louis, Mo.).
Example 16
[0305]This example describes the purification of juvenile hormone esterase activity from unfed adult flea midguts by affinity chromatography.
[0306]About 16,000 unfed adult flea midguts were collected in 20 mM Tris buffer (pH 7.7), containing 130 mM NaCl, 1 mM sodium EDTA, 1 mM PEFABLOC® (available from Boehringer Mannheim, Indianapolis, Ind.), 1 microgram/ml (μg/ml) leupeptin and 1 μg/ml pepstatin. The midguts were homogenized by freeze-fracture and sonication, and then centrifuged at about 14,000×g for 20 min. The soluble material from the centrifugation step was recovered, diafiltered into Tris buffered saline (TBS), and applied to a disposable plastic column containing about 1 ml of 3-[(4'-mercapto)butylthio]-1,1,1-trifluoropropane-2-one linked Sepharose 6B beads, prepared similarly to the method described by Venkatesh et al. (J. Biol. Chem., Vol. 265, No. 35, 21727-21732, 1990) (the 3-[(4'-mercapto)butylthio]-1,1,1-trifluoropropane-2-one was a gift from Novartis Corp., Basel, Switzerland; and the Epoxy-activated Sepharose 6B is available from Pharmacia Biotech Inc., Piscataway, N.J.). After overnight incubation at 4° C., the column was drained and the beads were washed with about 10 ml TBS, then about 10 ml TBS containing 0.1% (w/v) n-octylglucoside (OG; available from Boehringer Mannheim). The pre-column, flow-through, and wash fractions were tested for juvenile hormone esterase activity by the method previously described above in Example 7. The results indicate that the flow-through fraction contained approximately 40% less juvenile hormone esterase activity than the pre-column material, and that the washes contained very little activity.
[0307]Bound protein was eluted from the beads by adding about 10 ml of TBS containing 0.1% (w/v) OG and 1 mM 3-octylthio-1,1,1-trifluoropropane-2-one (OTFP; a gift from Novartis Corp.). After a 2 hour incubation at 4° C., about 5 ml of the eluate was collected, and the remaining 5 ml was incubated with the beads overnight at 4° C. The following day, the beads were drained, the eluate collected, and an additional 10 ml of TBS containing 0.1% (w/v) OG and 1 mM OTFP was added to the beads. After an overnight incubation at 4° C., the beads were drained and the eluate collected. The final 10 ml elution step was repeated 3 additional times so that we had 6 eluted fractions. The first elution fraction was dialyzed overnight twice against 1 liter of fresh TBS to remove excess OTFP. The second elution fraction was also dialyzed overnight against 1 liter of fresh TBS to remove OTFP. The third through sixth elution fractions were not dialyzed. All six eluted fractions were tested for juvenile hormone esterase activity by the method previously described above in Example 7. The results indicate that only the third elution fraction contained detectable juvenile hormone esterase activity. Analysis of the eluted fractions by silver-stained SDS-PAGE indicated that several proteins were specifically bound to the affinity beads and were eluted by OTFP. The apparent molecular weights of these proteins, as determined by SDS-PAGE, were about 66 kDa, 55 kDa, and 33 kDa. About 3.5 ml of each elution fraction were combined and concentrated to about 110 μl using a Centriplus 10 centrifugal concentrator (available from Amicon, Beverly, Mass.). This pool was separated by SDS-PAGE and blotted onto a polyvinylidene difluoride (PVDF) membrane as described previously in Example 5. The stained protein band at about 66 kDa was excised and subjected to N-terminal sequence analysis as described previously.
[0308]The results indicated that the N-terminal amino acid sequence of the putative 66 kDa juvenile hormone esterase protein was DL y/g V k/y/g v/q/n LQGTLKGKE (denoted herein as SEQ ID NO:74), in which the lower case letters designate uncertainties. Below is shown a comparison between different esterase amino acid sequences of the present invention.
TABLE-US-00001 SEQ ID NO:74: DL (y/g) V (k/y/g) (v/q/n) LQGTLKGKE SEQ ID NO:37: DL Q V T L LQGTLKGKE (Residues 3-17)
Example 17
[0309]This example describes the purification of an active recombinant juvenile hormone esterase protein from baculovirus supernatants.
[0310]About 1 liter of supernatant from cultures of S. frugiperda:pVL-nfE91600 cells producing the flea esterase protein PfE9528 was brought to about 50% saturation with ammonium sulfate and centrifuged at about 20000×g for about 30 minutes at 4° C. to pellet the precipitated material. After centrifugation, the pellet was retained and the supernatant was brought to about 100% saturation with ammonium sulfate and centrifuged as above. The material in both pellets were resuspended separately in about 35 ml of Tris buffered saline (TBS), pH 8.0. The resuspended pellets were assayed for the presence of flea esterase protein PfE9528 using standard Western blot techniques and a polyclonal antiserum that binds specifically to PfE9528 protein. Briefly, a rabbit was immunized with PHIS-PfE9528 protein purified from E. coli:pTrc-nfE91584 cells (described above in Example 12C) and boosted using standard procedures. The results indicated that the flea esterase protein PfE9528 was present in the S. frugiperda:pVL-nfE91600 supernatants and the protein was precipitated by adjusting the ammonium sulfate concentration from about 50% saturation to about 100% saturation.
[0311]The resuspended flea protein PfE9528 was diafiltered into about 10 ml of 25 mM Tris (pH 8.0), 10 mM NaCl using an Ultrafree-20 10 kD centrifugal concentrator in preparation for anion exchange chromatography. Aliquots of about 5 ml were loaded onto an Uno Q6 anion exchange column equilibrated in 25 mM Tris (pH 8.0), 10 mM NaCl. The column was washed with 25 mM Tris pH 8.0), 10 mM NaCl until most of the unbound protein was removed. Protein bound to the column was then eluted with a linear gradient from 10 mM to 1 M NaCl in 25 mM Tris buffer pH 8.0). Fractions were assayed for the presence of flea esterase protein PfE9528 by the immunoblot method described above. The results indicated that the flea esterase protein PfE9528 was eluted at about 200 mM NaCl.
[0312]Fractions containing the flea esterase protein PfE9528 were pooled and concentrated to about 440 μl using a Centricon 10 kD centrifugal concentrator in preparation for size exclusion chromatography. The sample was applied in 3 aliquots to a Bio-Select SEC 125-5 size exclusion chromatography column (available from Bio-Rad), previously equilibrated in TBS, pH 7.2. The column was eluted with TBS, pH 7.2 at a flow rate of about 0.5 ml/min, and fractions of about 250 μl were collected. The fractions were assayed for the presence of flea esterase protein PfE9528 by the immunoblot method described above. The results indicated that the flea esterase protein PfE9528 was eluted with about 6 ml of buffer, corresponding to a molecular weight of about 40 to 100 kDa based on the elution volumes of gel filtration molecular weight standard proteins (available from Sigma-Aldrich, St. Louis, Mo.).
[0313]Fractions containing flea esterase protein PfE9528 were then assayed for juvenile hormone esterase activity as described in Example 7 and carboxylesterase activity as described in Example 2. The results indicated that the purified flea esterase protein PfE9528 had both juvenile hormone esterase activity and carboxylesterase activity.
Sequence CWU
1
761401DNACtenocephalides felisCDS(92)..(400)misc_feature(219)..(219)At
nucleotide 219, n = unknown At amino acid residue 43, Xaa = Ile,
Thr, Lys or Arg 1tttacatcat taataaacat aaatctaata aatcttgtgg atcaagatca
agtttattag 60tgagagtgtt ggatttgtga aatatttcaa a atg aat tct tta att
gta aaa 112Met Asn Ser Leu Ile Val Lys1 5att tct caa
gga gct att gag ggg aag gaa atg att aat gat aat gga 160Ile Ser Gln
Gly Ala Ile Glu Gly Lys Glu Met Ile Asn Asp Asn Gly10 15
20aag tcg ttt aga gga ttt ttg ggt ata cct tat gct aaa
ccg cct ata 208Lys Ser Phe Arg Gly Phe Leu Gly Ile Pro Tyr Ala Lys
Pro Pro Ile25 30 35gga aat ctt ana ttt
aag cct cct caa aag cct gat gat tgg aat gat 256Gly Asn Leu Xaa Phe
Lys Pro Pro Gln Lys Pro Asp Asp Trp Asn Asp40 45
50 55gtt cga cca gct act gaa naa gca aat ggt
tgt aga tcg aaa cat atg 304Val Arg Pro Ala Thr Glu Xaa Ala Asn Gly
Cys Arg Ser Lys His Met60 65 70ctg cag
cat cat att att gga gac naa nat tgt cta tac cta aac gtn 352Leu Gln
His His Ile Ile Gly Asp Xaa Xaa Cys Leu Tyr Leu Asn Val75
80 85tat gtt cca ttg act tcc aaa ttg gag aaa cta cca
gta atg ttc tgg g 401Tyr Val Pro Leu Thr Ser Lys Leu Glu Lys Leu Pro
Val Met Phe Trp90 95
1002103PRTCtenocephalides felismisc_feature(43)..(43)The 'Xaa' at
location 43 stands for Lys, Arg, Thr, or Ile. 2Met Asn Ser Leu Ile
Val Lys Ile Ser Gln Gly Ala Ile Glu Gly Lys1 5
10 15Glu Met Ile Asn Asp Asn Gly Lys Ser Phe Arg
Gly Phe Leu Gly Ile20 25 30Pro Tyr Ala
Lys Pro Pro Ile Gly Asn Leu Xaa Phe Lys Pro Pro Gln35 40
45Lys Pro Asp Asp Trp Asn Asp Val Arg Pro Ala Thr Glu
Xaa Ala Asn50 55 60Gly Cys Arg Ser Lys
His Met Leu Gln His His Ile Ile Gly Asp Xaa65 70
75 80Xaa Cys Leu Tyr Leu Asn Val Tyr Val Pro
Leu Thr Ser Lys Leu Glu85 90 95Lys Leu
Pro Val Met Phe Trp1003401DNACtenocephalides felismisc_feature(50)..(50)n
= unknown 3cccagaacat tactggtagt ttctccaatt tggaagtcaa tggaacatan
acgtttaggt 60atagacaatn ttngtctcca ataatatgat gctgcagcat atgtttcgat
ctacaaccat 120ttgcttnttc agtagctggt cgaacatcat tccaatcatc aggcttttga
ggaggcttaa 180atntaagatt tcctataggc ggtttagcat aaggtatacc caaaaatcct
ctaaacgact 240ttccattatc attaatcatt tccttcccct caatagctcc ttgagaaatt
tttacaatta 300aagaattcat tttgaaatat ttcacaaatc caacactctc actaataaac
ttgatcttga 360tccacaagat ttattagatt tatgtttatt aatgatgtaa a
4014364DNACtenocephalides felisCDS(2)..(364) 4g tct cgt gtt
att ttt tta agt tgt att ttt ttg ttt agt ttt aat ttt 49Ser Arg Val Ile
Phe Leu Ser Cys Ile Phe Leu Phe Ser Phe Asn Phe1 5
10 15ata aac tgt gat tcc ccg act gta act ttg
ccc caa ggc gaa ttg gtt 97Ile Asn Cys Asp Ser Pro Thr Val Thr Leu
Pro Gln Gly Glu Leu Val20 25 30gga aaa
gct ttg acg aac gaa aat gga aaa gag tat ttt agc tac aca 145Gly Lys
Ala Leu Thr Asn Glu Asn Gly Lys Glu Tyr Phe Ser Tyr Thr35
40 45ggt gta cct tat gct aaa cct cct gtt gga gaa ctt
aga ttt aag cct 193Gly Val Pro Tyr Ala Lys Pro Pro Val Gly Glu Leu
Arg Phe Lys Pro50 55 60cca cag aaa gct
gag cca tgg caa ggt gtt ttc aac gcc aca tta tac 241Pro Gln Lys Ala
Glu Pro Trp Gln Gly Val Phe Asn Ala Thr Leu Tyr65 70
75 80gga aat gtg tgt aaa tct tta aat ttc
ttc ttg aag aaa att gaa gga 289Gly Asn Val Cys Lys Ser Leu Asn Phe
Phe Leu Lys Lys Ile Glu Gly85 90 95gac
gaa gac tgc ttg gta gta aac gtg tac gca cca aaa aca act tct 337Asp
Glu Asp Cys Leu Val Val Asn Val Tyr Ala Pro Lys Thr Thr Ser100
105 110gat aaa aaa ctt cca gta ttt ttc tgg
364Asp Lys Lys Leu Pro Val Phe Phe Trp115
1205121PRTCtenocephalides felis 5Ser Arg Val Ile Phe Leu Ser Cys
Ile Phe Leu Phe Ser Phe Asn Phe1 5 10
15Ile Asn Cys Asp Ser Pro Thr Val Thr Leu Pro Gln Gly Glu
Leu Val20 25 30Gly Lys Ala Leu Thr Asn
Glu Asn Gly Lys Glu Tyr Phe Ser Tyr Thr35 40
45Gly Val Pro Tyr Ala Lys Pro Pro Val Gly Glu Leu Arg Phe Lys Pro50
55 60Pro Gln Lys Ala Glu Pro Trp Gln Gly
Val Phe Asn Ala Thr Leu Tyr65 70 75
80Gly Asn Val Cys Lys Ser Leu Asn Phe Phe Leu Lys Lys Ile
Glu Gly85 90 95Asp Glu Asp Cys Leu Val
Val Asn Val Tyr Ala Pro Lys Thr Thr Ser100 105
110Asp Lys Lys Leu Pro Val Phe Phe Trp115
1206364DNACtenocephalides felis 6ccagaaaaat actggaagtt ttttatcaga
agttgttttt ggtgcgtaca cgtttactac 60caagcagtct tcgtctcctt caattttctt
caagaagaaa tttaaagatt tacacacatt 120tccgtataat gtggcgttga aaacaccttg
ccatggctca gctttctgtg gaggcttaaa 180tctaagttct ccaacaggag gtttagcata
aggtacacct gtgtagctaa aatactcttt 240tccattttcg ttcgtcaaag cttttccaac
caattcgcct tggggcaaag ttacagtcgg 300ggaatcacag tttataaaat taaaactaaa
caaaaaaata caacttaaaa aaataacacg 360agac
3647421DNACtenocephalides
felisCDS(113)..(421) 7tttacattac atcaaatcat atttttatta gtatattttt
tagaagaacc tagccaaaaa 60atatggactt tagactgtga ttaatttatt ttacctgaga
ttttccttta ca atg ggt 118Met Gly1gat ctt caa gtg act ttg tta caa ggt
tct ttg aga gga aaa gag caa 166Asp Leu Gln Val Thr Leu Leu Gln Gly
Ser Leu Arg Gly Lys Glu Gln5 10 15att
aat gaa aag gga aat gtg ttt tat agt tat tct gga att cca tat 214Ile
Asn Glu Lys Gly Asn Val Phe Tyr Ser Tyr Ser Gly Ile Pro Tyr20
25 30gcc aaa cct cca gtt ggt gat cta aga ttc aag
cca cct caa cct gca 262Ala Lys Pro Pro Val Gly Asp Leu Arg Phe Lys
Pro Pro Gln Pro Ala35 40 45
50gaa cct tgg tca ggt gtc ctt gat gct act aaa gaa ggg aat agt tgt
310Glu Pro Trp Ser Gly Val Leu Asp Ala Thr Lys Glu Gly Asn Ser Cys55
60 65aga tct gta cat ttt att aaa aag att
aaa gta ggg gct gaa gat tgt 358Arg Ser Val His Phe Ile Lys Lys Ile
Lys Val Gly Ala Glu Asp Cys70 75 80cta
tac ctc aat gtc tat gta cca aaa aca tca gag aaa tcc ctt ctt 406Leu
Tyr Leu Asn Val Tyr Val Pro Lys Thr Ser Glu Lys Ser Leu Leu85
90 95cca gta atg gta tgg
421Pro Val Met Val Trp1008103PRTCtenocephalides
felis 8Met Gly Asp Leu Gln Val Thr Leu Leu Gln Gly Ser Leu Arg Gly Lys1
5 10 15Glu Gln Ile Asn Glu
Lys Gly Asn Val Phe Tyr Ser Tyr Ser Gly Ile20 25
30Pro Tyr Ala Lys Pro Pro Val Gly Asp Leu Arg Phe Lys Pro Pro
Gln35 40 45Pro Ala Glu Pro Trp Ser Gly
Val Leu Asp Ala Thr Lys Glu Gly Asn50 55
60Ser Cys Arg Ser Val His Phe Ile Lys Lys Ile Lys Val Gly Ala Glu65
70 75 80Asp Cys Leu Tyr Leu
Asn Val Tyr Val Pro Lys Thr Ser Glu Lys Ser85 90
95Leu Leu Pro Val Met Val Trp1009421DNACtenocephalides felis
9ccataccatt actggaagaa gggatttctc tgatgttttt ggtacataga cattgaggta
60tagacaatct tcagccccta ctttaatctt tttaataaaa tgtacagatc tacaactatt
120cccttcttta gtagcatcaa ggacacctga ccaaggttct gcaggttgag gtggcttgaa
180tcttagatca ccaactggag gtttggcata tggaattcca gaataactat aaaacacatt
240tcccttttca ttaatttgct cttttcctct caaagaacct tgtaacaaag tcacttgaag
300atcacccatt gtaaaggaaa atctcaggta aaataaatta atcacagtct aaagtccata
360ttttttggct aggttcttct aaaaaatata ctaataaaaa tatgatttga tgtaatgtaa
420a
42110524DNACtenocephalides felisCDS(113)..(523) 10gaacgttgat acgatagaca
tgtcgtcttc aaaacgtcta ttttatcata aacaaaacga 60gataaataat aacaattaag
caaccaaaat gcattaaaaa acacaataaa aa atg tta 118Met Leu1cct cac agt agt
gca tta gtt tta ttt tta ttt ttt tta ttt ttc tta 166Pro His Ser Ser
Ala Leu Val Leu Phe Leu Phe Phe Leu Phe Phe Leu5 10
15ttt aca cct atc ttg tgc ata cta tgg gat aac cta gat cag
cat ttg 214Phe Thr Pro Ile Leu Cys Ile Leu Trp Asp Asn Leu Asp Gln
His Leu20 25 30tgc aga gtt caa ttt aac
agg atc acg gaa gga aaa ccg ttc cga tat 262Cys Arg Val Gln Phe Asn
Arg Ile Thr Glu Gly Lys Pro Phe Arg Tyr35 40
45 50aaa gat cat agg aat gat gta tat tgt tct tat
ttg gga att cct tat 310Lys Asp His Arg Asn Asp Val Tyr Cys Ser Tyr
Leu Gly Ile Pro Tyr55 60 65gcc gaa ccg
cct att gga cca tta cga ttt cag tct cca aaa cca ata 358Ala Glu Pro
Pro Ile Gly Pro Leu Arg Phe Gln Ser Pro Lys Pro Ile70 75
80tca aat cca aaa aca gga ttc gta cag gct cga act ttg
gga gac aaa 406Ser Asn Pro Lys Thr Gly Phe Val Gln Ala Arg Thr Leu
Gly Asp Lys85 90 95tgt ttc cag gaa agt
cta ata tat tct tat gca gga agc gaa gat tgc 454Cys Phe Gln Glu Ser
Leu Ile Tyr Ser Tyr Ala Gly Ser Glu Asp Cys100 105
110tta tat ctg aat ata ttc acg cca gag act gtt aat tct gcg aac
aat 502Leu Tyr Leu Asn Ile Phe Thr Pro Glu Thr Val Asn Ser Ala Asn
Asn115 120 125 130aca aaa
tat cct gta atg ttc t 524Thr Lys
Tyr Pro Val Met Phe13511137PRTCtenocephalides felis 11Met Leu Pro His Ser
Ser Ala Leu Val Leu Phe Leu Phe Phe Leu Phe1 5
10 15Phe Leu Phe Thr Pro Ile Leu Cys Ile Leu Trp
Asp Asn Leu Asp Gln20 25 30His Leu Cys
Arg Val Gln Phe Asn Arg Ile Thr Glu Gly Lys Pro Phe35 40
45Arg Tyr Lys Asp His Arg Asn Asp Val Tyr Cys Ser Tyr
Leu Gly Ile50 55 60Pro Tyr Ala Glu Pro
Pro Ile Gly Pro Leu Arg Phe Gln Ser Pro Lys65 70
75 80Pro Ile Ser Asn Pro Lys Thr Gly Phe Val
Gln Ala Arg Thr Leu Gly85 90 95Asp Lys
Cys Phe Gln Glu Ser Leu Ile Tyr Ser Tyr Ala Gly Ser Glu100
105 110Asp Cys Leu Tyr Leu Asn Ile Phe Thr Pro Glu Thr
Val Asn Ser Ala115 120 125Asn Asn Thr Lys
Tyr Pro Val Met Phe130 13512524DNACtenocephalides felis
12agaacattac aggatatttt gtattgttcg cagaattaac agtctctggc gtgaatatat
60tcagatataa gcaatcttcg cttcctgcat aagaatatat tagactttcc tggaaacatt
120tgtctcccaa agttcgagcc tgtacgaatc ctgtttttgg atttgatatt ggttttggag
180actgaaatcg taatggtcca ataggcggtt cggcataagg aattcccaaa taagaacaat
240atacatcatt cctatgatct ttatatcgga acggttttcc ttccgtgatc ctgttaaatt
300gaactctgca caaatgctga tctaggttat cccatagtat gcacaagata ggtgtaaata
360agaaaaataa aaaaaataaa aataaaacta atgcactact gtgaggtaac attttttatt
420gtgtttttta atgcattttg gttgcttaat tgttattatt tatctcgttt tgtttatgat
480aaaatagacg ttttgaagac gacatgtcta tcgtatcaac gttc
524131982DNACtenocephalides felisCDS(3)..(1517)misc_feature(300)..(300)At
nucleotide 300, r = a or g At amino acid residue 100, Xaa = Asn or
Asp 13at ttt agc tac aca ggt gta cct tat gct aaa cct cct gtt gga gaa
47Phe Ser Tyr Thr Gly Val Pro Tyr Ala Lys Pro Pro Val Gly Glu1
5 10 15ctt aga ttt aag cct cca
cag aaa gct gag cca tgg caa ggt gtt ttc 95Leu Arg Phe Lys Pro Pro
Gln Lys Ala Glu Pro Trp Gln Gly Val Phe20 25
30aac gcc aca tta tac gga aat gtg tgt aaa tct tta aat ttc ttc ttg
143Asn Ala Thr Leu Tyr Gly Asn Val Cys Lys Ser Leu Asn Phe Phe Leu35
40 45aag aaa att gaa gga gac gaa gac tgc
ttg gta gta aac gtg tac gca 191Lys Lys Ile Glu Gly Asp Glu Asp Cys
Leu Val Val Asn Val Tyr Ala50 55 60cca
aaa aca act tct gat aaa aaa ctt cca gta ttt ttc tgg gtt cat 239Pro
Lys Thr Thr Ser Asp Lys Lys Leu Pro Val Phe Phe Trp Val His65
70 75ggt ggt ggt ttt gtg act gga tcc gga aat tta
gaa ttc caa agc cca 287Gly Gly Gly Phe Val Thr Gly Ser Gly Asn Leu
Glu Phe Gln Ser Pro80 85 90
95gat tat tta gta rat ttt gat gtt att ttc gta act ttc aat tac cga
335Asp Tyr Leu Val Xaa Phe Asp Val Ile Phe Val Thr Phe Asn Tyr Arg100
105 110ttg gga cct ctc gga ttt ctg aat ttg
gag ttg gag ggt gct cca gga 383Leu Gly Pro Leu Gly Phe Leu Asn Leu
Glu Leu Glu Gly Ala Pro Gly115 120 125aat
gta gga tta ttg gat cag gtg gca gct ctg aaa tgg acc aaa gaa 431Asn
Val Gly Leu Leu Asp Gln Val Ala Ala Leu Lys Trp Thr Lys Glu130
135 140aac att gag aaa ttt ggt gga gat cca gaa aat
att aca att ggt ggt 479Asn Ile Glu Lys Phe Gly Gly Asp Pro Glu Asn
Ile Thr Ile Gly Gly145 150 155gtt tct gct
ggt gga gca agt gtt cat tat ctt ttg tta tct cat aca 527Val Ser Ala
Gly Gly Ala Ser Val His Tyr Leu Leu Leu Ser His Thr160
165 170 175acc act gga ctt tac aaa agg
gca att gct caa agt gga agt gct ttt 575Thr Thr Gly Leu Tyr Lys Arg
Ala Ile Ala Gln Ser Gly Ser Ala Phe180 185
190aat cca tgg gcc ttc caa aga cat cca gta aag cgt agt ctt caa ctt
623Asn Pro Trp Ala Phe Gln Arg His Pro Val Lys Arg Ser Leu Gln Leu195
200 205gct gag ata ttg ggt cat ccc aca aac
aat act caa gat gct tta gaa 671Ala Glu Ile Leu Gly His Pro Thr Asn
Asn Thr Gln Asp Ala Leu Glu210 215 220ttc
tta caa aaa gcc ccc gta gac agt ctc ctg aag aaa atg cca gct 719Phe
Leu Gln Lys Ala Pro Val Asp Ser Leu Leu Lys Lys Met Pro Ala225
230 235gaa aca gaa ggt gaa ata ata gaa gag ttt gtc
ttc gta cca tca att 767Glu Thr Glu Gly Glu Ile Ile Glu Glu Phe Val
Phe Val Pro Ser Ile240 245 250
255gaa aaa gtt ttc cca tcc cac caa cct ttc ttg gaa gaa tca cca ttg
815Glu Lys Val Phe Pro Ser His Gln Pro Phe Leu Glu Glu Ser Pro Leu260
265 270gcc aga atg aaa tcc gga tcc ttt aac
aaa gta cct tta tta gtt gga 863Ala Arg Met Lys Ser Gly Ser Phe Asn
Lys Val Pro Leu Leu Val Gly275 280 285ttt
aac agt gca gaa gga ctt ttg ttc aaa ttc ttc atg aaa gaa aaa 911Phe
Asn Ser Ala Glu Gly Leu Leu Phe Lys Phe Phe Met Lys Glu Lys290
295 300cca gag atg ctg aac caa gct gaa gca gat ttt
gaa aga ctc gta cca 959Pro Glu Met Leu Asn Gln Ala Glu Ala Asp Phe
Glu Arg Leu Val Pro305 310 315gcc gaa ttt
gaa tta gtc cat gga tca gag gaa tcg aaa aaa ctt gca 1007Ala Glu Phe
Glu Leu Val His Gly Ser Glu Glu Ser Lys Lys Leu Ala320
325 330 335gaa aaa atc agg aag ttt tac
ttt gac gat aaa ccc gtt cca gaa aat 1055Glu Lys Ile Arg Lys Phe Tyr
Phe Asp Asp Lys Pro Val Pro Glu Asn340 345
350gaa cag aaa ttt att gac ttg ata gga gat att tgg ttt act aga ggt
1103Glu Gln Lys Phe Ile Asp Leu Ile Gly Asp Ile Trp Phe Thr Arg Gly355
360 365gtt gac aag cat gtc aag ttg tct gtg
gag aaa caa gac gaa cca gtt 1151Val Asp Lys His Val Lys Leu Ser Val
Glu Lys Gln Asp Glu Pro Val370 375 380tat
tat tat gaa tat tcc ttc tcg gaa agt cat cct gca aaa gga aca 1199Tyr
Tyr Tyr Glu Tyr Ser Phe Ser Glu Ser His Pro Ala Lys Gly Thr385
390 395ttt ggt gat cat aat ctg act ggt gca tgc cat
gga gaa gaa ctt gtg 1247Phe Gly Asp His Asn Leu Thr Gly Ala Cys His
Gly Glu Glu Leu Val400 405 410
415aat tta ttc aaa gtc gag atg atg aag ctg gaa aaa gat aaa cct aat
1295Asn Leu Phe Lys Val Glu Met Met Lys Leu Glu Lys Asp Lys Pro Asn420
425 430gtt cta tta aca aaa gat aga gta ctt
gcc atg tgg act aac ttc atc 1343Val Leu Leu Thr Lys Asp Arg Val Leu
Ala Met Trp Thr Asn Phe Ile435 440 445aaa
aat gga aat cct act cct gaa gta aca gaa tta ttg cca gtt aaa 1391Lys
Asn Gly Asn Pro Thr Pro Glu Val Thr Glu Leu Leu Pro Val Lys450
455 460tgg gaa cct gcc aca aaa gac aag ttg aat tat
ttg aac att gat gcc 1439Trp Glu Pro Ala Thr Lys Asp Lys Leu Asn Tyr
Leu Asn Ile Asp Ala465 470 475acc tta act
ttg gga aca aat cct gag gca aac cga gtc aaa ttt tgg 1487Thr Leu Thr
Leu Gly Thr Asn Pro Glu Ala Asn Arg Val Lys Phe Trp480
485 490 495gaa gac gcc aca aaa tct ttg
cac ggt caa taataattta tgaaaattgt 1537Glu Asp Ala Thr Lys Ser Leu
His Gly Gln500 505tttaaatact ttaggtaata tattaggtaa
ataaaaatta aaaaataaca atttttatgt 1597tttatgtatt ggcttatgtg tatcagttct
aattttattt atttattctt gttttgcttg 1657ttttgaaata tcatggtttt aattttcaaa
acacaacgtc gtttgttttt agcaaaattt 1717ccaatagata tgttatatta agtactctga
agtattttta tatatacact aaaatcagta 1777aaaatacatt aactaaaaat ataagatatt
ttcaataatt ttttttaaag aaaataccaa 1837aaataaagta aaattccaaa cggaattttt
gtttaactta aaaataaaat taactcttca 1897ataattttga taattagtat ttctgatatc
attagtgaaa attatatttt gataatacgt 1957atttatattt aaaataaaat tatgt
198214505PRTCtenocephalides
felismisc_feature(100)..(100)The 'Xaa' at location 100 stands for Asp, or
Asn. 14Phe Ser Tyr Thr Gly Val Pro Tyr Ala Lys Pro Pro Val Gly Glu
Leu1 5 10 15Arg Phe Lys
Pro Pro Gln Lys Ala Glu Pro Trp Gln Gly Val Phe Asn20 25
30Ala Thr Leu Tyr Gly Asn Val Cys Lys Ser Leu Asn Phe
Phe Leu Lys35 40 45Lys Ile Glu Gly Asp
Glu Asp Cys Leu Val Val Asn Val Tyr Ala Pro50 55
60Lys Thr Thr Ser Asp Lys Lys Leu Pro Val Phe Phe Trp Val His
Gly65 70 75 80Gly Gly
Phe Val Thr Gly Ser Gly Asn Leu Glu Phe Gln Ser Pro Asp85
90 95Tyr Leu Val Xaa Phe Asp Val Ile Phe Val Thr Phe
Asn Tyr Arg Leu100 105 110Gly Pro Leu Gly
Phe Leu Asn Leu Glu Leu Glu Gly Ala Pro Gly Asn115 120
125Val Gly Leu Leu Asp Gln Val Ala Ala Leu Lys Trp Thr Lys
Glu Asn130 135 140Ile Glu Lys Phe Gly Gly
Asp Pro Glu Asn Ile Thr Ile Gly Gly Val145 150
155 160Ser Ala Gly Gly Ala Ser Val His Tyr Leu Leu
Leu Ser His Thr Thr165 170 175Thr Gly Leu
Tyr Lys Arg Ala Ile Ala Gln Ser Gly Ser Ala Phe Asn180
185 190Pro Trp Ala Phe Gln Arg His Pro Val Lys Arg Ser
Leu Gln Leu Ala195 200 205Glu Ile Leu Gly
His Pro Thr Asn Asn Thr Gln Asp Ala Leu Glu Phe210 215
220Leu Gln Lys Ala Pro Val Asp Ser Leu Leu Lys Lys Met Pro
Ala Glu225 230 235 240Thr
Glu Gly Glu Ile Ile Glu Glu Phe Val Phe Val Pro Ser Ile Glu245
250 255Lys Val Phe Pro Ser His Gln Pro Phe Leu Glu
Glu Ser Pro Leu Ala260 265 270Arg Met Lys
Ser Gly Ser Phe Asn Lys Val Pro Leu Leu Val Gly Phe275
280 285Asn Ser Ala Glu Gly Leu Leu Phe Lys Phe Phe Met
Lys Glu Lys Pro290 295 300Glu Met Leu Asn
Gln Ala Glu Ala Asp Phe Glu Arg Leu Val Pro Ala305 310
315 320Glu Phe Glu Leu Val His Gly Ser Glu
Glu Ser Lys Lys Leu Ala Glu325 330 335Lys
Ile Arg Lys Phe Tyr Phe Asp Asp Lys Pro Val Pro Glu Asn Glu340
345 350Gln Lys Phe Ile Asp Leu Ile Gly Asp Ile Trp
Phe Thr Arg Gly Val355 360 365Asp Lys His
Val Lys Leu Ser Val Glu Lys Gln Asp Glu Pro Val Tyr370
375 380Tyr Tyr Glu Tyr Ser Phe Ser Glu Ser His Pro Ala
Lys Gly Thr Phe385 390 395
400Gly Asp His Asn Leu Thr Gly Ala Cys His Gly Glu Glu Leu Val Asn405
410 415Leu Phe Lys Val Glu Met Met Lys Leu
Glu Lys Asp Lys Pro Asn Val420 425 430Leu
Leu Thr Lys Asp Arg Val Leu Ala Met Trp Thr Asn Phe Ile Lys435
440 445Asn Gly Asn Pro Thr Pro Glu Val Thr Glu Leu
Leu Pro Val Lys Trp450 455 460Glu Pro Ala
Thr Lys Asp Lys Leu Asn Tyr Leu Asn Ile Asp Ala Thr465
470 475 480Leu Thr Leu Gly Thr Asn Pro
Glu Ala Asn Arg Val Lys Phe Trp Glu485 490
495Asp Ala Thr Lys Ser Leu His Gly Gln500
505151982DNACtenocephalides felis 15acataatttt attttaaata taaatacgta
ttatcaaaat ataattttca ctaatgatat 60cagaaatact aattatcaaa attattgaag
agttaatttt atttttaagt taaacaaaaa 120ttccgtttgg aattttactt tatttttggt
attttcttta aaaaaaatta ttgaaaatat 180cttatatttt tagttaatgt atttttactg
attttagtgt atatataaaa atacttcaga 240gtacttaata taacatatct attggaaatt
ttgctaaaaa caaacgacgt tgtgttttga 300aaattaaaac catgatattt caaaacaagc
aaaacaagaa taaataaata aaattagaac 360tgatacacat aagccaatac ataaaacata
aaaattgtta ttttttaatt tttatttacc 420taatatatta cctaaagtat ttaaaacaat
tttcataaat tattattgac cgtgcaaaga 480ttttgtggcg tcttcccaaa atttgactcg
gtttgcctca ggatttgttc ccaaagttaa 540ggtggcatca atgttcaaat aattcaactt
gtcttttgtg gcaggttccc atttaactgg 600caataattct gttacttcag gagtaggatt
tccatttttg atgaagttag tccacatggc 660aagtactcta tcttttgtta atagaacatt
aggtttatct ttttccagct tcatcatctc 720gactttgaat aaattcacaa gttcttctcc
atggcatgca ccagtcagat tatgatcacc 780aaatgttcct tttgcaggat gactttccga
gaaggaatat tcataataat aaactggttc 840gtcttgtttc tccacagaca acttgacatg
cttgtcaaca cctctagtaa accaaatatc 900tcctatcaag tcaataaatt tctgttcatt
ttctggaacg ggtttatcgt caaagtaaaa 960cttcctgatt ttttctgcaa gttttttcga
ttcctctgat ccatggacta attcaaattc 1020ggctggtacg agtctttcaa aatctgcttc
agcttggttc agcatctctg gtttttcttt 1080catgaagaat ttgaacaaaa gtccttctgc
actgttaaat ccaactaata aaggtacttt 1140gttaaaggat ccggatttca ttctggccaa
tggtgattct tccaagaaag gttggtggga 1200tgggaaaact ttttcaattg atggtacgaa
gacaaactct tctattattt caccttctgt 1260ttcagctggc attttcttca ggagactgtc
tacgggggct ttttgtaaga attctaaagc 1320atcttgagta ttgtttgtgg gatgacccaa
tatctcagca agttgaagac tacgctttac 1380tggatgtctt tggaaggccc atggattaaa
agcacttcca ctttgagcaa ttgccctttt 1440gtaaagtcca gtggttgtat gagataacaa
aagataatga acacttgctc caccagcaga 1500aacaccacca attgtaatat tttctggatc
tccaccaaat ttctcaatgt tttctttggt 1560ccatttcaga gctgccacct gatccaataa
tcctacattt cctggagcac cctccaactc 1620caaattcaga aatccgagag gtcccaatcg
gtaattgaaa gttacgaaaa taacatcaaa 1680atytactaaa taatctgggc tttggaattc
taaatttccg gatccagtca caaaaccacc 1740accatgaacc cagaaaaata ctggaagttt
tttatcagaa gttgtttttg gtgcgtacac 1800gtttactacc aagcagtctt cgtctccttc
aattttcttc aagaagaaat ttaaagattt 1860acacacattt ccgtataatg tggcgttgaa
aacaccttgc catggctcag ctttctgtgg 1920aggcttaaat ctaagttctc caacaggagg
tttagcataa ggtacacctg tgtagctaaa 1980at
1982161515DNACtenocephalides
felisexon(1)..(1515)misc_feature(298)..(298)At nucleotide 298, r = a or g
At amino acid residue 100, Xaa = Asn or Asp 16ttt agc tac aca ggt
gta cct tat gct aaa cct cct gtt gga gaa ctt 48Phe Ser Tyr Thr Gly
Val Pro Tyr Ala Lys Pro Pro Val Gly Glu Leu1 5
10 15aga ttt aag cct cca cag aaa gct gag cca tgg
caa ggt gtt ttc aac 96Arg Phe Lys Pro Pro Gln Lys Ala Glu Pro Trp
Gln Gly Val Phe Asn20 25 30gcc aca tta
tac gga aat gtg tgt aaa tct tta aat ttc ttc ttg aag 144Ala Thr Leu
Tyr Gly Asn Val Cys Lys Ser Leu Asn Phe Phe Leu Lys35 40
45aaa att gaa gga gac gaa gac tgc ttg gta gta aac gtg
tac gca cca 192Lys Ile Glu Gly Asp Glu Asp Cys Leu Val Val Asn Val
Tyr Ala Pro50 55 60aaa aca act tct gat
aaa aaa ctt cca gta ttt ttc tgg gtt cat ggt 240Lys Thr Thr Ser Asp
Lys Lys Leu Pro Val Phe Phe Trp Val His Gly65 70
75 80ggt ggt ttt gtg act gga tcc gga aat tta
gaa ttc caa agc cca gat 288Gly Gly Phe Val Thr Gly Ser Gly Asn Leu
Glu Phe Gln Ser Pro Asp85 90 95tat tta
gta rat ttt gat gtt att ttc gta act ttc aat tac cga ttg 336Tyr Leu
Val Xaa Phe Asp Val Ile Phe Val Thr Phe Asn Tyr Arg Leu100
105 110gga cct ctc gga ttt ctg aat ttg gag ttg gag ggt
gct cca gga aat 384Gly Pro Leu Gly Phe Leu Asn Leu Glu Leu Glu Gly
Ala Pro Gly Asn115 120 125gta gga tta ttg
gat cag gtg gca gct ctg aaa tgg acc aaa gaa aac 432Val Gly Leu Leu
Asp Gln Val Ala Ala Leu Lys Trp Thr Lys Glu Asn130 135
140att gag aaa ttt ggt gga gat cca gaa aat att aca att ggt
ggt gtt 480Ile Glu Lys Phe Gly Gly Asp Pro Glu Asn Ile Thr Ile Gly
Gly Val145 150 155 160tct
gct ggt gga gca agt gtt cat tat ctt ttg tta tct cat aca acc 528Ser
Ala Gly Gly Ala Ser Val His Tyr Leu Leu Leu Ser His Thr Thr165
170 175act gga ctt tac aaa agg gca att gct caa agt
gga agt gct ttt aat 576Thr Gly Leu Tyr Lys Arg Ala Ile Ala Gln Ser
Gly Ser Ala Phe Asn180 185 190cca tgg gcc
ttc caa aga cat cca gta aag cgt agt ctt caa ctt gct 624Pro Trp Ala
Phe Gln Arg His Pro Val Lys Arg Ser Leu Gln Leu Ala195
200 205gag ata ttg ggt cat ccc aca aac aat act caa gat
gct tta gaa ttc 672Glu Ile Leu Gly His Pro Thr Asn Asn Thr Gln Asp
Ala Leu Glu Phe210 215 220tta caa aaa gcc
ccc gta gac agt ctc ctg aag aaa atg cca gct gaa 720Leu Gln Lys Ala
Pro Val Asp Ser Leu Leu Lys Lys Met Pro Ala Glu225 230
235 240aca gaa ggt gaa ata ata gaa gag ttt
gtc ttc gta cca tca att gaa 768Thr Glu Gly Glu Ile Ile Glu Glu Phe
Val Phe Val Pro Ser Ile Glu245 250 255aaa
gtt ttc cca tcc cac caa cct ttc ttg gaa gaa tca cca ttg gcc 816Lys
Val Phe Pro Ser His Gln Pro Phe Leu Glu Glu Ser Pro Leu Ala260
265 270aga atg aaa tcc gga tcc ttt aac aaa gta cct
tta tta gtt gga ttt 864Arg Met Lys Ser Gly Ser Phe Asn Lys Val Pro
Leu Leu Val Gly Phe275 280 285aac agt gca
gaa gga ctt ttg ttc aaa ttc ttc atg aaa gaa aaa cca 912Asn Ser Ala
Glu Gly Leu Leu Phe Lys Phe Phe Met Lys Glu Lys Pro290
295 300gag atg ctg aac caa gct gaa gca gat ttt gaa aga
ctc gta cca gcc 960Glu Met Leu Asn Gln Ala Glu Ala Asp Phe Glu Arg
Leu Val Pro Ala305 310 315
320gaa ttt gaa tta gtc cat gga tca gag gaa tcg aaa aaa ctt gca gaa
1008Glu Phe Glu Leu Val His Gly Ser Glu Glu Ser Lys Lys Leu Ala Glu325
330 335aaa atc agg aag ttt tac ttt gac gat
aaa ccc gtt cca gaa aat gaa 1056Lys Ile Arg Lys Phe Tyr Phe Asp Asp
Lys Pro Val Pro Glu Asn Glu340 345 350cag
aaa ttt att gac ttg ata gga gat att tgg ttt act aga ggt gtt 1104Gln
Lys Phe Ile Asp Leu Ile Gly Asp Ile Trp Phe Thr Arg Gly Val355
360 365gac aag cat gtc aag ttg tct gtg gag aaa caa
gac gaa cca gtt tat 1152Asp Lys His Val Lys Leu Ser Val Glu Lys Gln
Asp Glu Pro Val Tyr370 375 380tat tat gaa
tat tcc ttc tcg gaa agt cat cct gca aaa gga aca ttt 1200Tyr Tyr Glu
Tyr Ser Phe Ser Glu Ser His Pro Ala Lys Gly Thr Phe385
390 395 400ggt gat cat aat ctg act ggt
gca tgc cat gga gaa gaa ctt gtg aat 1248Gly Asp His Asn Leu Thr Gly
Ala Cys His Gly Glu Glu Leu Val Asn405 410
415tta ttc aaa gtc gag atg atg aag ctg gaa aaa gat aaa cct aat gtt
1296Leu Phe Lys Val Glu Met Met Lys Leu Glu Lys Asp Lys Pro Asn Val420
425 430cta tta aca aaa gat aga gta ctt gcc
atg tgg act aac ttc atc aaa 1344Leu Leu Thr Lys Asp Arg Val Leu Ala
Met Trp Thr Asn Phe Ile Lys435 440 445aat
gga aat cct act cct gaa gta aca gaa tta ttg cca gtt aaa tgg 1392Asn
Gly Asn Pro Thr Pro Glu Val Thr Glu Leu Leu Pro Val Lys Trp450
455 460gaa cct gcc aca aaa gac aag ttg aat tat ttg
aac att gat gcc acc 1440Glu Pro Ala Thr Lys Asp Lys Leu Asn Tyr Leu
Asn Ile Asp Ala Thr465 470 475
480tta act ttg gga aca aat cct gag gca aac cga gtc aaa ttt tgg gaa
1488Leu Thr Leu Gly Thr Asn Pro Glu Ala Asn Arg Val Lys Phe Trp Glu485
490 495gac gcc aca aaa tct ttg cac ggt caa
1515Asp Ala Thr Lys Ser Leu His Gly
Gln500 505171515DNACtenocephalides felis 17ttgaccgtgc
aaagattttg tggcgtcttc ccaaaatttg actcggtttg cctcaggatt 60tgttcccaaa
gttaaggtgg catcaatgtt caaataattc aacttgtctt ttgtggcagg 120ttcccattta
actggcaata attctgttac ttcaggagta ggatttccat ttttgatgaa 180gttagtccac
atggcaagta ctctatcttt tgttaataga acattaggtt tatctttttc 240cagcttcatc
atctcgactt tgaataaatt cacaagttct tctccatggc atgcaccagt 300cagattatga
tcaccaaatg ttccttttgc aggatgactt tccgagaagg aatattcata 360ataataaact
ggttcgtctt gtttctccac agacaacttg acatgcttgt caacacctct 420agtaaaccaa
atatctccta tcaagtcaat aaatttctgt tcattttctg gaacgggttt 480atcgtcaaag
taaaacttcc tgattttttc tgcaagtttt ttcgattcct ctgatccatg 540gactaattca
aattcggctg gtacgagtct ttcaaaatct gcttcagctt ggttcagcat 600ctctggtttt
tctttcatga agaatttgaa caaaagtcct tctgcactgt taaatccaac 660taataaaggt
actttgttaa aggatccgga tttcattctg gccaatggtg attcttccaa 720gaaaggttgg
tgggatggga aaactttttc aattgatggt acgaagacaa actcttctat 780tatttcacct
tctgtttcag ctggcatttt cttcaggaga ctgtctacgg gggctttttg 840taagaattct
aaagcatctt gagtattgtt tgtgggatga cccaatatct cagcaagttg 900aagactacgc
tttactggat gtctttggaa ggcccatgga ttaaaagcac ttccactttg 960agcaattgcc
cttttgtaaa gtccagtggt tgtatgagat aacaaaagat aatgaacact 1020tgctccacca
gcagaaacac caccaattgt aatattttct ggatctccac caaatttctc 1080aatgttttct
ttggtccatt tcagagctgc cacctgatcc aataatccta catttcctgg 1140agcaccctcc
aactccaaat tcagaaatcc gagaggtccc aatcggtaat tgaaagttac 1200gaaaataaca
tcaaaatyta ctaaataatc tgggctttgg aattctaaat ttccggatcc 1260agtcacaaaa
ccaccaccat gaacccagaa aaatactgga agttttttat cagaagttgt 1320ttttggtgcg
tacacgttta ctaccaagca gtcttcgtct ccttcaattt tcttcaagaa 1380gaaatttaaa
gatttacaca catttccgta taatgtggcg ttgaaaacac cttgccatgg 1440ctcagctttc
tgtggaggct taaatctaag ttctccaaca ggaggtttag cataaggtac 1500acctgtgtag
ctaaa
1515181792DNACtenocephalides
felisCDS(49)..(1701)misc_feature(1758)..(1758)n = unknown 18actgtgtgct
aataattcag tacacacagt caatagtcta gatccaag atg tct cgt 57Met Ser
Arg1gtt att ttt tta agt tgt att ttt ttg ttt agt ttt aat ttt ata aaa
105Val Ile Phe Leu Ser Cys Ile Phe Leu Phe Ser Phe Asn Phe Ile Lys5
10 15tgt gat ccc ccg act gta act ttg ccc cag
ggc gaa ttg gtt gga aaa 153Cys Asp Pro Pro Thr Val Thr Leu Pro Gln
Gly Glu Leu Val Gly Lys20 25 30
35gct ttg acg aac gaa aat gga aaa gag tat ttt agc tac aca ggt
gtg 201Ala Leu Thr Asn Glu Asn Gly Lys Glu Tyr Phe Ser Tyr Thr Gly
Val40 45 50cct tat gct aaa cct cca gtt
gga gaa ctt aga ttt aag cct cca cag 249Pro Tyr Ala Lys Pro Pro Val
Gly Glu Leu Arg Phe Lys Pro Pro Gln55 60
65aaa gct gag cca tgg aat ggt gtt ttc aac gcc aca tca cat gga aat
297Lys Ala Glu Pro Trp Asn Gly Val Phe Asn Ala Thr Ser His Gly Asn70
75 80gtg tgc aaa gct ttg aat ttc ttc ttg aaa
aaa att gaa gga gac gaa 345Val Cys Lys Ala Leu Asn Phe Phe Leu Lys
Lys Ile Glu Gly Asp Glu85 90 95gac tgc
ttg ttg gtg aat gtg tac gca cca aaa aca act tct gac aaa 393Asp Cys
Leu Leu Val Asn Val Tyr Ala Pro Lys Thr Thr Ser Asp Lys100
105 110 115aaa ctt cca gta ttt ttc tgg
gtt cat ggt ggc ggt ttt gtg act gga 441Lys Leu Pro Val Phe Phe Trp
Val His Gly Gly Gly Phe Val Thr Gly120 125
130tcc gga aat tta gaa ttt caa agc cca gat tat tta gta aat tat gat
489Ser Gly Asn Leu Glu Phe Gln Ser Pro Asp Tyr Leu Val Asn Tyr Asp135
140 145gtt att ttt gta act ttc aat tac cga
ttg gga cca ctc gga ttt ttg 537Val Ile Phe Val Thr Phe Asn Tyr Arg
Leu Gly Pro Leu Gly Phe Leu150 155 160aat
ttg gag ttg gaa ggt gct cct gga aat gta gga tta ttg gat cag 585Asn
Leu Glu Leu Glu Gly Ala Pro Gly Asn Val Gly Leu Leu Asp Gln165
170 175gta gca gct ttg aaa tgg acc aaa gaa aat att
gag aaa ttt ggt gga 633Val Ala Ala Leu Lys Trp Thr Lys Glu Asn Ile
Glu Lys Phe Gly Gly180 185 190
195gat cca gaa aat att aca att ggt ggt gtt tct gct ggt gga gca agt
681Asp Pro Glu Asn Ile Thr Ile Gly Gly Val Ser Ala Gly Gly Ala Ser200
205 210gtt cat tat ctt tta ttg tca cat aca
acc act gga ctt tac aaa agg 729Val His Tyr Leu Leu Leu Ser His Thr
Thr Thr Gly Leu Tyr Lys Arg215 220 225gca
att gct caa agt gga agt gct tta aat cca tgg gcc ttc caa aga 777Ala
Ile Ala Gln Ser Gly Ser Ala Leu Asn Pro Trp Ala Phe Gln Arg230
235 240cat cca gta aag cgt agt ctt caa ctt gct gag
ata tta ggt cat ccc 825His Pro Val Lys Arg Ser Leu Gln Leu Ala Glu
Ile Leu Gly His Pro245 250 255aca aac aac
act caa gat gct tta gaa ttc tta caa aaa gcc cca gta 873Thr Asn Asn
Thr Gln Asp Ala Leu Glu Phe Leu Gln Lys Ala Pro Val260
265 270 275gac agt ctc ctg aaa aaa atg
cca gct gaa aca gaa ggt gaa ata ata 921Asp Ser Leu Leu Lys Lys Met
Pro Ala Glu Thr Glu Gly Glu Ile Ile280 285
290gaa gag ttc gtc ttc gta cca tca att gaa aaa gtt ttc cca tcc cac
969Glu Glu Phe Val Phe Val Pro Ser Ile Glu Lys Val Phe Pro Ser His295
300 305caa cct ttc ttg gaa gaa tca cca ttg
gcc aga atg aaa tct gga tcc 1017Gln Pro Phe Leu Glu Glu Ser Pro Leu
Ala Arg Met Lys Ser Gly Ser310 315 320ttt
aac aaa gta cct tta tta gtt gga ttc aac agc gca gaa gga ctt 1065Phe
Asn Lys Val Pro Leu Leu Val Gly Phe Asn Ser Ala Glu Gly Leu325
330 335ttg tac aaa ttc ttt atg aaa gaa aaa cca gag
atg ctg aac caa gct 1113Leu Tyr Lys Phe Phe Met Lys Glu Lys Pro Glu
Met Leu Asn Gln Ala340 345 350
355gaa gca gat ttc gaa aga ctc gta cca gcc gaa ttt gaa tta gcc cat
1161Glu Ala Asp Phe Glu Arg Leu Val Pro Ala Glu Phe Glu Leu Ala His360
365 370gga tca gaa gaa tcg aaa aaa ctt gca
gaa aaa atc agg aag ttt tac 1209Gly Ser Glu Glu Ser Lys Lys Leu Ala
Glu Lys Ile Arg Lys Phe Tyr375 380 385ttt
gac gat aaa ccc gtt cct gaa aat gag cag aaa ttt att gac ttg 1257Phe
Asp Asp Lys Pro Val Pro Glu Asn Glu Gln Lys Phe Ile Asp Leu390
395 400ata gga gat att tgg ttt act aga ggc att gac
aag cat gtc aag ttg 1305Ile Gly Asp Ile Trp Phe Thr Arg Gly Ile Asp
Lys His Val Lys Leu405 410 415tct gta gaa
aaa caa gac gag cca gta tat tat tat gaa tat tct ttc 1353Ser Val Glu
Lys Gln Asp Glu Pro Val Tyr Tyr Tyr Glu Tyr Ser Phe420
425 430 435tct gaa agt cat cct gca aaa
gga aca ttt ggt gac cat aac ttg act 1401Ser Glu Ser His Pro Ala Lys
Gly Thr Phe Gly Asp His Asn Leu Thr440 445
450gga gca tgt cat ggt gaa gaa ctt gtg aat tta ttc aaa gtc gag atg
1449Gly Ala Cys His Gly Glu Glu Leu Val Asn Leu Phe Lys Val Glu Met455
460 465atg aag ctg gaa aaa gat aaa ccg aat
gtt tta tta aca aaa gat agg 1497Met Lys Leu Glu Lys Asp Lys Pro Asn
Val Leu Leu Thr Lys Asp Arg470 475 480gta
ctt gct atg tgg acg aac ttc atc aaa aat gga aat cct act cct 1545Val
Leu Ala Met Trp Thr Asn Phe Ile Lys Asn Gly Asn Pro Thr Pro485
490 495gaa gta act gaa tta ttg cca gtt aaa tgg gaa
cct gcc aca aaa gac 1593Glu Val Thr Glu Leu Leu Pro Val Lys Trp Glu
Pro Ala Thr Lys Asp500 505 510
515aag ttg aat tat ttg aac att gat gcc acc tta act ttg gga aca aat
1641Lys Leu Asn Tyr Leu Asn Ile Asp Ala Thr Leu Thr Leu Gly Thr Asn520
525 530cca gaa gaa acc cga gtc aaa tty tgg
gaa gat gcc aca aaa act ttg 1689Pro Glu Glu Thr Arg Val Lys Phe Trp
Glu Asp Ala Thr Lys Thr Leu535 540 545cac
agt caa taa aaatgtatga aaattgtttt aattatttta ggtaatacat 1741His
Ser Gln550taggtaaata aaaattnaaa aataacnaaa aaaaaaaaaa aaaaaaaaaa a
179219550PRTCtenocephalides felis 19Met Ser Arg Val Ile Phe Leu Ser
Cys Ile Phe Leu Phe Ser Phe Asn1 5 10
15Phe Ile Lys Cys Asp Pro Pro Thr Val Thr Leu Pro Gln Gly
Glu Leu20 25 30Val Gly Lys Ala Leu Thr
Asn Glu Asn Gly Lys Glu Tyr Phe Ser Tyr35 40
45Thr Gly Val Pro Tyr Ala Lys Pro Pro Val Gly Glu Leu Arg Phe Lys50
55 60Pro Pro Gln Lys Ala Glu Pro Trp Asn
Gly Val Phe Asn Ala Thr Ser65 70 75
80His Gly Asn Val Cys Lys Ala Leu Asn Phe Phe Leu Lys Lys
Ile Glu85 90 95Gly Asp Glu Asp Cys Leu
Leu Val Asn Val Tyr Ala Pro Lys Thr Thr100 105
110Ser Asp Lys Lys Leu Pro Val Phe Phe Trp Val His Gly Gly Gly
Phe115 120 125Val Thr Gly Ser Gly Asn Leu
Glu Phe Gln Ser Pro Asp Tyr Leu Val130 135
140Asn Tyr Asp Val Ile Phe Val Thr Phe Asn Tyr Arg Leu Gly Pro Leu145
150 155 160Gly Phe Leu Asn
Leu Glu Leu Glu Gly Ala Pro Gly Asn Val Gly Leu165 170
175Leu Asp Gln Val Ala Ala Leu Lys Trp Thr Lys Glu Asn Ile
Glu Lys180 185 190Phe Gly Gly Asp Pro Glu
Asn Ile Thr Ile Gly Gly Val Ser Ala Gly195 200
205Gly Ala Ser Val His Tyr Leu Leu Leu Ser His Thr Thr Thr Gly
Leu210 215 220Tyr Lys Arg Ala Ile Ala Gln
Ser Gly Ser Ala Leu Asn Pro Trp Ala225 230
235 240Phe Gln Arg His Pro Val Lys Arg Ser Leu Gln Leu
Ala Glu Ile Leu245 250 255Gly His Pro Thr
Asn Asn Thr Gln Asp Ala Leu Glu Phe Leu Gln Lys260 265
270Ala Pro Val Asp Ser Leu Leu Lys Lys Met Pro Ala Glu Thr
Glu Gly275 280 285Glu Ile Ile Glu Glu Phe
Val Phe Val Pro Ser Ile Glu Lys Val Phe290 295
300Pro Ser His Gln Pro Phe Leu Glu Glu Ser Pro Leu Ala Arg Met
Lys305 310 315 320Ser Gly
Ser Phe Asn Lys Val Pro Leu Leu Val Gly Phe Asn Ser Ala325
330 335Glu Gly Leu Leu Tyr Lys Phe Phe Met Lys Glu Lys
Pro Glu Met Leu340 345 350Asn Gln Ala Glu
Ala Asp Phe Glu Arg Leu Val Pro Ala Glu Phe Glu355 360
365Leu Ala His Gly Ser Glu Glu Ser Lys Lys Leu Ala Glu Lys
Ile Arg370 375 380Lys Phe Tyr Phe Asp Asp
Lys Pro Val Pro Glu Asn Glu Gln Lys Phe385 390
395 400Ile Asp Leu Ile Gly Asp Ile Trp Phe Thr Arg
Gly Ile Asp Lys His405 410 415Val Lys Leu
Ser Val Glu Lys Gln Asp Glu Pro Val Tyr Tyr Tyr Glu420
425 430Tyr Ser Phe Ser Glu Ser His Pro Ala Lys Gly Thr
Phe Gly Asp His435 440 445Asn Leu Thr Gly
Ala Cys His Gly Glu Glu Leu Val Asn Leu Phe Lys450 455
460Val Glu Met Met Lys Leu Glu Lys Asp Lys Pro Asn Val Leu
Leu Thr465 470 475 480Lys
Asp Arg Val Leu Ala Met Trp Thr Asn Phe Ile Lys Asn Gly Asn485
490 495Pro Thr Pro Glu Val Thr Glu Leu Leu Pro Val
Lys Trp Glu Pro Ala500 505 510Thr Lys Asp
Lys Leu Asn Tyr Leu Asn Ile Asp Ala Thr Leu Thr Leu515
520 525Gly Thr Asn Pro Glu Glu Thr Arg Val Lys Phe Trp
Glu Asp Ala Thr530 535 540Lys Thr Leu His
Ser Gln545 550201792DNACtenocephalides
felismisc_feature(25)..(25)n = unknown 20tttttttttt tttttttttt ttttngttat
ttttnaattt ttatttacct aatgtattac 60ctaaaataat taaaacaatt ttcatacatt
tttattgact gtgcaaagtt tttgtggcat 120cttcccaraa tttgactcgg gtttcttctg
gatttgttcc caaagttaag gtggcatcaa 180tgttcaaata attcaacttg tcttttgtgg
caggttccca tttaactggc aataattcag 240ttacttcagg agtaggattt ccatttttga
tgaagttcgt ccacatagca agtaccctat 300cttttgttaa taaaacattc ggtttatctt
tttccagctt catcatctcg actttgaata 360aattcacaag ttcttcacca tgacatgctc
cagtcaagtt atggtcacca aatgttcctt 420ttgcaggatg actttcagag aaagaatatt
cataataata tactggctcg tcttgttttt 480ctacagacaa cttgacatgc ttgtcaatgc
ctctagtaaa ccaaatatct cctatcaagt 540caataaattt ctgctcattt tcaggaacgg
gtttatcgtc aaagtaaaac ttcctgattt 600tttctgcaag ttttttcgat tcttctgatc
catgggctaa ttcaaattcg gctggtacga 660gtctttcgaa atctgcttca gcttggttca
gcatctctgg tttttctttc ataaagaatt 720tgtacaaaag tccttctgcg ctgttgaatc
caactaataa aggtactttg ttaaaggatc 780cagatttcat tctggccaat ggtgattctt
ccaagaaagg ttggtgggat gggaaaactt 840tttcaattga tggtacgaag acgaactctt
ctattatttc accttctgtt tcagctggca 900tttttttcag gagactgtct actggggctt
tttgtaagaa ttctaaagca tcttgagtgt 960tgtttgtggg atgacctaat atctcagcaa
gttgaagact acgctttact ggatgtcttt 1020ggaaggccca tggatttaaa gcacttccac
tttgagcaat tgcccttttg taaagtccag 1080tggttgtatg tgacaataaa agataatgaa
cacttgctcc accagcagaa acaccaccaa 1140ttgtaatatt ttctggatct ccaccaaatt
tctcaatatt ttctttggtc catttcaaag 1200ctgctacctg atccaataat cctacatttc
caggagcacc ttccaactcc aaattcaaaa 1260atccgagtgg tcccaatcgg taattgaaag
ttacaaaaat aacatcataa tttactaaat 1320aatctgggct ttgaaattct aaatttccgg
atccagtcac aaaaccgcca ccatgaaccc 1380agaaaaatac tggaagtttt ttgtcagaag
ttgtttttgg tgcgtacaca ttcaccaaca 1440agcagtcttc gtctccttca atttttttca
agaagaaatt caaagctttg cacacatttc 1500catgtgatgt ggcgttgaaa acaccattcc
atggctcagc tttctgtgga ggcttaaatc 1560taagttctcc aactggaggt ttagcataag
gcacacctgt gtagctaaaa tactcttttc 1620cattttcgtt cgtcaaagct tttccaacca
attcgccctg gggcaaagtt acagtcgggg 1680gatcacattt tataaaatta aaactaaaca
aaaaaataca acttaaaaaa ataacacgag 1740acatcttgga tctagactat tgactgtgtg
tactgaatta ttagcacaca gt 1792211650DNACtenocephalides
felisexon(1)..(1650) 21atg tct cgt gtt att ttt tta agt tgt att ttt ttg
ttt agt ttt aat 48Met Ser Arg Val Ile Phe Leu Ser Cys Ile Phe Leu
Phe Ser Phe Asn1 5 10
15ttt ata aaa tgt gat ccc ccg act gta act ttg ccc cag ggc gaa ttg
96Phe Ile Lys Cys Asp Pro Pro Thr Val Thr Leu Pro Gln Gly Glu Leu20
25 30gtt gga aaa gct ttg acg aac gaa aat gga
aaa gag tat ttt agc tac 144Val Gly Lys Ala Leu Thr Asn Glu Asn Gly
Lys Glu Tyr Phe Ser Tyr35 40 45aca ggt
gtg cct tat gct aaa cct cca gtt gga gaa ctt aga ttt aag 192Thr Gly
Val Pro Tyr Ala Lys Pro Pro Val Gly Glu Leu Arg Phe Lys50
55 60cct cca cag aaa gct gag cca tgg aat ggt gtt ttc
aac gcc aca tca 240Pro Pro Gln Lys Ala Glu Pro Trp Asn Gly Val Phe
Asn Ala Thr Ser65 70 75
80cat gga aat gtg tgc aaa gct ttg aat ttc ttc ttg aaa aaa att gaa
288His Gly Asn Val Cys Lys Ala Leu Asn Phe Phe Leu Lys Lys Ile Glu85
90 95gga gac gaa gac tgc ttg ttg gtg aat gtg
tac gca cca aaa aca act 336Gly Asp Glu Asp Cys Leu Leu Val Asn Val
Tyr Ala Pro Lys Thr Thr100 105 110tct gac
aaa aaa ctt cca gta ttt ttc tgg gtt cat ggt ggc ggt ttt 384Ser Asp
Lys Lys Leu Pro Val Phe Phe Trp Val His Gly Gly Gly Phe115
120 125gtg act gga tcc gga aat tta gaa ttt caa agc cca
gat tat tta gta 432Val Thr Gly Ser Gly Asn Leu Glu Phe Gln Ser Pro
Asp Tyr Leu Val130 135 140aat tat gat gtt
att ttt gta act ttc aat tac cga ttg gga cca ctc 480Asn Tyr Asp Val
Ile Phe Val Thr Phe Asn Tyr Arg Leu Gly Pro Leu145 150
155 160gga ttt ttg aat ttg gag ttg gaa ggt
gct cct gga aat gta gga tta 528Gly Phe Leu Asn Leu Glu Leu Glu Gly
Ala Pro Gly Asn Val Gly Leu165 170 175ttg
gat cag gta gca gct ttg aaa tgg acc aaa gaa aat att gag aaa 576Leu
Asp Gln Val Ala Ala Leu Lys Trp Thr Lys Glu Asn Ile Glu Lys180
185 190ttt ggt gga gat cca gaa aat att aca att ggt
ggt gtt tct gct ggt 624Phe Gly Gly Asp Pro Glu Asn Ile Thr Ile Gly
Gly Val Ser Ala Gly195 200 205gga gca agt
gtt cat tat ctt tta ttg tca cat aca acc act gga ctt 672Gly Ala Ser
Val His Tyr Leu Leu Leu Ser His Thr Thr Thr Gly Leu210
215 220tac aaa agg gca att gct caa agt gga agt gct tta
aat cca tgg gcc 720Tyr Lys Arg Ala Ile Ala Gln Ser Gly Ser Ala Leu
Asn Pro Trp Ala225 230 235
240ttc caa aga cat cca gta aag cgt agt ctt caa ctt gct gag ata tta
768Phe Gln Arg His Pro Val Lys Arg Ser Leu Gln Leu Ala Glu Ile Leu245
250 255ggt cat ccc aca aac aac act caa gat
gct tta gaa ttc tta caa aaa 816Gly His Pro Thr Asn Asn Thr Gln Asp
Ala Leu Glu Phe Leu Gln Lys260 265 270gcc
cca gta gac agt ctc ctg aaa aaa atg cca gct gaa aca gaa ggt 864Ala
Pro Val Asp Ser Leu Leu Lys Lys Met Pro Ala Glu Thr Glu Gly275
280 285gaa ata ata gaa gag ttc gtc ttc gta cca tca
att gaa aaa gtt ttc 912Glu Ile Ile Glu Glu Phe Val Phe Val Pro Ser
Ile Glu Lys Val Phe290 295 300cca tcc cac
caa cct ttc ttg gaa gaa tca cca ttg gcc aga atg aaa 960Pro Ser His
Gln Pro Phe Leu Glu Glu Ser Pro Leu Ala Arg Met Lys305
310 315 320tct gga tcc ttt aac aaa gta
cct tta tta gtt gga ttc aac agc gca 1008Ser Gly Ser Phe Asn Lys Val
Pro Leu Leu Val Gly Phe Asn Ser Ala325 330
335gaa gga ctt ttg tac aaa ttc ttt atg aaa gaa aaa cca gag atg ctg
1056Glu Gly Leu Leu Tyr Lys Phe Phe Met Lys Glu Lys Pro Glu Met Leu340
345 350aac caa gct gaa gca gat ttc gaa aga
ctc gta cca gcc gaa ttt gaa 1104Asn Gln Ala Glu Ala Asp Phe Glu Arg
Leu Val Pro Ala Glu Phe Glu355 360 365tta
gcc cat gga tca gaa gaa tcg aaa aaa ctt gca gaa aaa atc agg 1152Leu
Ala His Gly Ser Glu Glu Ser Lys Lys Leu Ala Glu Lys Ile Arg370
375 380aag ttt tac ttt gac gat aaa ccc gtt cct gaa
aat gag cag aaa ttt 1200Lys Phe Tyr Phe Asp Asp Lys Pro Val Pro Glu
Asn Glu Gln Lys Phe385 390 395
400att gac ttg ata gga gat att tgg ttt act aga ggc att gac aag cat
1248Ile Asp Leu Ile Gly Asp Ile Trp Phe Thr Arg Gly Ile Asp Lys His405
410 415gtc aag ttg tct gta gaa aaa caa gac
gag cca gta tat tat tat gaa 1296Val Lys Leu Ser Val Glu Lys Gln Asp
Glu Pro Val Tyr Tyr Tyr Glu420 425 430tat
tct ttc tct gaa agt cat cct gca aaa gga aca ttt ggt gac cat 1344Tyr
Ser Phe Ser Glu Ser His Pro Ala Lys Gly Thr Phe Gly Asp His435
440 445aac ttg act gga gca tgt cat ggt gaa gaa ctt
gtg aat tta ttc aaa 1392Asn Leu Thr Gly Ala Cys His Gly Glu Glu Leu
Val Asn Leu Phe Lys450 455 460gtc gag atg
atg aag ctg gaa aaa gat aaa ccg aat gtt tta tta aca 1440Val Glu Met
Met Lys Leu Glu Lys Asp Lys Pro Asn Val Leu Leu Thr465
470 475 480aaa gat agg gta ctt gct atg
tgg acg aac ttc atc aaa aat gga aat 1488Lys Asp Arg Val Leu Ala Met
Trp Thr Asn Phe Ile Lys Asn Gly Asn485 490
495cct act cct gaa gta act gaa tta ttg cca gtt aaa tgg gaa cct gcc
1536Pro Thr Pro Glu Val Thr Glu Leu Leu Pro Val Lys Trp Glu Pro Ala500
505 510aca aaa gac aag ttg aat tat ttg aac
att gat gcc acc tta act ttg 1584Thr Lys Asp Lys Leu Asn Tyr Leu Asn
Ile Asp Ala Thr Leu Thr Leu515 520 525gga
aca aat cca gaa gaa acc cga gtc aaa tty tgg gaa gat gcc aca 1632Gly
Thr Asn Pro Glu Glu Thr Arg Val Lys Phe Trp Glu Asp Ala Thr530
535 540aaa act ttg cac agt caa
1650Lys Thr Leu His Ser Gln545
550221650DNACtenocephalides felis 22ttgactgtgc aaagtttttg tggcatcttc
ccaraatttg actcgggttt cttctggatt 60tgttcccaaa gttaaggtgg catcaatgtt
caaataattc aacttgtctt ttgtggcagg 120ttcccattta actggcaata attcagttac
ttcaggagta ggatttccat ttttgatgaa 180gttcgtccac atagcaagta ccctatcttt
tgttaataaa acattcggtt tatctttttc 240cagcttcatc atctcgactt tgaataaatt
cacaagttct tcaccatgac atgctccagt 300caagttatgg tcaccaaatg ttccttttgc
aggatgactt tcagagaaag aatattcata 360ataatatact ggctcgtctt gtttttctac
agacaacttg acatgcttgt caatgcctct 420agtaaaccaa atatctccta tcaagtcaat
aaatttctgc tcattttcag gaacgggttt 480atcgtcaaag taaaacttcc tgattttttc
tgcaagtttt ttcgattctt ctgatccatg 540ggctaattca aattcggctg gtacgagtct
ttcgaaatct gcttcagctt ggttcagcat 600ctctggtttt tctttcataa agaatttgta
caaaagtcct tctgcgctgt tgaatccaac 660taataaaggt actttgttaa aggatccaga
tttcattctg gccaatggtg attcttccaa 720gaaaggttgg tgggatggga aaactttttc
aattgatggt acgaagacga actcttctat 780tatttcacct tctgtttcag ctggcatttt
tttcaggaga ctgtctactg gggctttttg 840taagaattct aaagcatctt gagtgttgtt
tgtgggatga cctaatatct cagcaagttg 900aagactacgc tttactggat gtctttggaa
ggcccatgga tttaaagcac ttccactttg 960agcaattgcc cttttgtaaa gtccagtggt
tgtatgtgac aataaaagat aatgaacact 1020tgctccacca gcagaaacac caccaattgt
aatattttct ggatctccac caaatttctc 1080aatattttct ttggtccatt tcaaagctgc
tacctgatcc aataatccta catttccagg 1140agcaccttcc aactccaaat tcaaaaatcc
gagtggtccc aatcggtaat tgaaagttac 1200aaaaataaca tcataattta ctaaataatc
tgggctttga aattctaaat ttccggatcc 1260agtcacaaaa ccgccaccat gaacccagaa
aaatactgga agttttttgt cagaagttgt 1320ttttggtgcg tacacattca ccaacaagca
gtcttcgtct ccttcaattt ttttcaagaa 1380gaaattcaaa gctttgcaca catttccatg
tgatgtggcg ttgaaaacac cattccatgg 1440ctcagctttc tgtggaggct taaatctaag
ttctccaact ggaggtttag cataaggcac 1500acctgtgtag ctaaaatact cttttccatt
ttcgttcgtc aaagcttttc caaccaattc 1560gccctggggc aaagttacag tcgggggatc
acattttata aaattaaaac taaacaaaaa 1620aatacaactt aaaaaaataa cacgagacat
1650231590DNACtenocephalides
felisexon(1)..(1590) 23gat ccc ccg act gta act ttg ccc cag ggc gaa ttg
gtt gga aaa gct 48Asp Pro Pro Thr Val Thr Leu Pro Gln Gly Glu Leu
Val Gly Lys Ala1 5 10
15ttg acg aac gaa aat gga aaa gag tat ttt agc tac aca ggt gtg cct
96Leu Thr Asn Glu Asn Gly Lys Glu Tyr Phe Ser Tyr Thr Gly Val Pro20
25 30tat gct aaa cct cca gtt gga gaa ctt aga
ttt aag cct cca cag aaa 144Tyr Ala Lys Pro Pro Val Gly Glu Leu Arg
Phe Lys Pro Pro Gln Lys35 40 45gct gag
cca tgg aat ggt gtt ttc aac gcc aca tca cat gga aat gtg 192Ala Glu
Pro Trp Asn Gly Val Phe Asn Ala Thr Ser His Gly Asn Val50
55 60tgc aaa gct ttg aat ttc ttc ttg aaa aaa att gaa
gga gac gaa gac 240Cys Lys Ala Leu Asn Phe Phe Leu Lys Lys Ile Glu
Gly Asp Glu Asp65 70 75
80tgc ttg ttg gtg aat gtg tac gca cca aaa aca act tct gac aaa aaa
288Cys Leu Leu Val Asn Val Tyr Ala Pro Lys Thr Thr Ser Asp Lys Lys85
90 95ctt cca gta ttt ttc tgg gtt cat ggt ggc
ggt ttt gtg act gga tcc 336Leu Pro Val Phe Phe Trp Val His Gly Gly
Gly Phe Val Thr Gly Ser100 105 110gga aat
tta gaa ttt caa agc cca gat tat tta gta aat tat gat gtt 384Gly Asn
Leu Glu Phe Gln Ser Pro Asp Tyr Leu Val Asn Tyr Asp Val115
120 125att ttt gta act ttc aat tac cga ttg gga cca ctc
gga ttt ttg aat 432Ile Phe Val Thr Phe Asn Tyr Arg Leu Gly Pro Leu
Gly Phe Leu Asn130 135 140ttg gag ttg gaa
ggt gct cct gga aat gta gga tta ttg gat cag gta 480Leu Glu Leu Glu
Gly Ala Pro Gly Asn Val Gly Leu Leu Asp Gln Val145 150
155 160gca gct ttg aaa tgg acc aaa gaa aat
att gag aaa ttt ggt gga gat 528Ala Ala Leu Lys Trp Thr Lys Glu Asn
Ile Glu Lys Phe Gly Gly Asp165 170 175cca
gaa aat att aca att ggt ggt gtt tct gct ggt gga gca agt gtt 576Pro
Glu Asn Ile Thr Ile Gly Gly Val Ser Ala Gly Gly Ala Ser Val180
185 190cat tat ctt tta ttg tca cat aca acc act gga
ctt tac aaa agg gca 624His Tyr Leu Leu Leu Ser His Thr Thr Thr Gly
Leu Tyr Lys Arg Ala195 200 205att gct caa
agt gga agt gct tta aat cca tgg gcc ttc caa aga cat 672Ile Ala Gln
Ser Gly Ser Ala Leu Asn Pro Trp Ala Phe Gln Arg His210
215 220cca gta aag cgt agt ctt caa ctt gct gag ata tta
ggt cat ccc aca 720Pro Val Lys Arg Ser Leu Gln Leu Ala Glu Ile Leu
Gly His Pro Thr225 230 235
240aac aac act caa gat gct tta gaa ttc tta caa aaa gcc cca gta gac
768Asn Asn Thr Gln Asp Ala Leu Glu Phe Leu Gln Lys Ala Pro Val Asp245
250 255agt ctc ctg aaa aaa atg cca gct gaa
aca gaa ggt gaa ata ata gaa 816Ser Leu Leu Lys Lys Met Pro Ala Glu
Thr Glu Gly Glu Ile Ile Glu260 265 270gag
ttc gtc ttc gta cca tca att gaa aaa gtt ttc cca tcc cac caa 864Glu
Phe Val Phe Val Pro Ser Ile Glu Lys Val Phe Pro Ser His Gln275
280 285cct ttc ttg gaa gaa tca cca ttg gcc aga atg
aaa tct gga tcc ttt 912Pro Phe Leu Glu Glu Ser Pro Leu Ala Arg Met
Lys Ser Gly Ser Phe290 295 300aac aaa gta
cct tta tta gtt gga ttc aac agc gca gaa gga ctt ttg 960Asn Lys Val
Pro Leu Leu Val Gly Phe Asn Ser Ala Glu Gly Leu Leu305
310 315 320tac aaa ttc ttt atg aaa gaa
aaa cca gag atg ctg aac caa gct gaa 1008Tyr Lys Phe Phe Met Lys Glu
Lys Pro Glu Met Leu Asn Gln Ala Glu325 330
335gca gat ttc gaa aga ctc gta cca gcc gaa ttt gaa tta gcc cat gga
1056Ala Asp Phe Glu Arg Leu Val Pro Ala Glu Phe Glu Leu Ala His Gly340
345 350tca gaa gaa tcg aaa aaa ctt gca gaa
aaa atc agg aag ttt tac ttt 1104Ser Glu Glu Ser Lys Lys Leu Ala Glu
Lys Ile Arg Lys Phe Tyr Phe355 360 365gac
gat aaa ccc gtt cct gaa aat gag cag aaa ttt att gac ttg ata 1152Asp
Asp Lys Pro Val Pro Glu Asn Glu Gln Lys Phe Ile Asp Leu Ile370
375 380gga gat att tgg ttt act aga ggc att gac aag
cat gtc aag ttg tct 1200Gly Asp Ile Trp Phe Thr Arg Gly Ile Asp Lys
His Val Lys Leu Ser385 390 395
400gta gaa aaa caa gac gag cca gta tat tat tat gaa tat tct ttc tct
1248Val Glu Lys Gln Asp Glu Pro Val Tyr Tyr Tyr Glu Tyr Ser Phe Ser405
410 415gaa agt cat cct gca aaa gga aca ttt
ggt gac cat aac ttg act gga 1296Glu Ser His Pro Ala Lys Gly Thr Phe
Gly Asp His Asn Leu Thr Gly420 425 430gca
tgt cat ggt gaa gaa ctt gtg aat tta ttc aaa gtc gag atg atg 1344Ala
Cys His Gly Glu Glu Leu Val Asn Leu Phe Lys Val Glu Met Met435
440 445aag ctg gaa aaa gat aaa ccg aat gtt tta tta
aca aaa gat agg gta 1392Lys Leu Glu Lys Asp Lys Pro Asn Val Leu Leu
Thr Lys Asp Arg Val450 455 460ctt gct atg
tgg acg aac ttc atc aaa aat gga aat cct act cct gaa 1440Leu Ala Met
Trp Thr Asn Phe Ile Lys Asn Gly Asn Pro Thr Pro Glu465
470 475 480gta act gaa tta ttg cca gtt
aaa tgg gaa cct gcc aca aaa gac aag 1488Val Thr Glu Leu Leu Pro Val
Lys Trp Glu Pro Ala Thr Lys Asp Lys485 490
495ttg aat tat ttg aac att gat gcc acc tta act ttg gga aca aat cca
1536Leu Asn Tyr Leu Asn Ile Asp Ala Thr Leu Thr Leu Gly Thr Asn Pro500
505 510gaa gaa acc cga gtc aaa tty tgg gaa
gat gcc aca aaa act ttg cac 1584Glu Glu Thr Arg Val Lys Phe Trp Glu
Asp Ala Thr Lys Thr Leu His515 520 525agt
caa 1590Ser
Gln530242836DNACtenocephalides
felisCDS(99)..(1889)misc_feature(2278)..(2278)n = unknown 24tagacatgtc
gtcttcaaaa cgtctatttt atcataaaca aaacgagata aataataaca 60attaagcaac
caaaatgcat taaaaaacac aataaaaa atg tta cct cac agt agt 116Met Leu Pro
His Ser Ser1 5gca tta gtt tta ttt tta ttt ttt tta ttt ttc
tta ttt aca cct atc 164Ala Leu Val Leu Phe Leu Phe Phe Leu Phe Phe
Leu Phe Thr Pro Ile10 15 20ttg tgc ata
cta tgg gat aac cta gat cag cat ttg tgc aga gtt caa 212Leu Cys Ile
Leu Trp Asp Asn Leu Asp Gln His Leu Cys Arg Val Gln25 30
35ttt aac ggg atc acg gaa gga aaa ccg ttc cga tat aaa
gat cat agg 260Phe Asn Gly Ile Thr Glu Gly Lys Pro Phe Arg Tyr Lys
Asp His Arg40 45 50aat gat gta tat tgt
tct tat ttg gga att cct tat gcc gaa ccg cct 308Asn Asp Val Tyr Cys
Ser Tyr Leu Gly Ile Pro Tyr Ala Glu Pro Pro55 60
65 70ttt gga cca tta cga ttt cag tct cca aaa
cca ata tca aat cca aaa 356Phe Gly Pro Leu Arg Phe Gln Ser Pro Lys
Pro Ile Ser Asn Pro Lys75 80 85aca gga
ttc gta cag gct cga act ttg gga gac aaa tgt ttc cag gaa 404Thr Gly
Phe Val Gln Ala Arg Thr Leu Gly Asp Lys Cys Phe Gln Glu90
95 100agt cta ata tat tct tat gca gga agc gaa gat tgc
tta tat ctg aat 452Ser Leu Ile Tyr Ser Tyr Ala Gly Ser Glu Asp Cys
Leu Tyr Leu Asn105 110 115ata ttc acg cca
gag act gtt aat tct gcg aac aat aca aaa tat cct 500Ile Phe Thr Pro
Glu Thr Val Asn Ser Ala Asn Asn Thr Lys Tyr Pro120 125
130gta atg ttc tgg atc cat gga ggc gca ttc aac caa gga tca
gga tct 548Val Met Phe Trp Ile His Gly Gly Ala Phe Asn Gln Gly Ser
Gly Ser135 140 145 150tat
aat ttt ttt gga cct gat tat ttg atc agg gaa gga att att ttg 596Tyr
Asn Phe Phe Gly Pro Asp Tyr Leu Ile Arg Glu Gly Ile Ile Leu155
160 165gtc act atc aac tat aga tta gga gtt ttc ggt
ttt cta tca gcg ccg 644Val Thr Ile Asn Tyr Arg Leu Gly Val Phe Gly
Phe Leu Ser Ala Pro170 175 180gaa tgg gat
atc cat gga aat atg ggt cta aaa gac cag aga ttg gca 692Glu Trp Asp
Ile His Gly Asn Met Gly Leu Lys Asp Gln Arg Leu Ala185
190 195cta aaa tgg gtt tac gac aac atc gaa aag ttt ggt
gga gac aga gaa 740Leu Lys Trp Val Tyr Asp Asn Ile Glu Lys Phe Gly
Gly Asp Arg Glu200 205 210aaa att aca att
gct gga gaa tct gct gga gca gca agt gtc cat ttt 788Lys Ile Thr Ile
Ala Gly Glu Ser Ala Gly Ala Ala Ser Val His Phe215 220
225 230ctg atg atg gac aac tcg act aga aaa
tac tac caa agg gcc att ttg 836Leu Met Met Asp Asn Ser Thr Arg Lys
Tyr Tyr Gln Arg Ala Ile Leu235 240 245cag
agt ggg aca tta cta aat ccg act gct aat caa att caa ctt ctg 884Gln
Ser Gly Thr Leu Leu Asn Pro Thr Ala Asn Gln Ile Gln Leu Leu250
255 260cat aga ttt gaa aaa ctc aaa caa gtg cta aac
atc acg caa aaa caa 932His Arg Phe Glu Lys Leu Lys Gln Val Leu Asn
Ile Thr Gln Lys Gln265 270 275gaa ctc cta
aac ctg gat aaa aac cta att tta cga gca gcc tta aac 980Glu Leu Leu
Asn Leu Asp Lys Asn Leu Ile Leu Arg Ala Ala Leu Asn280
285 290aga gtt cct gat agc aac gac cat gac cga gac aca
gta cca gta ttt 1028Arg Val Pro Asp Ser Asn Asp His Asp Arg Asp Thr
Val Pro Val Phe295 300 305
310aat cca gtc tta gaa tca cca gaa tct cca gat cca ata aca ttt cca
1076Asn Pro Val Leu Glu Ser Pro Glu Ser Pro Asp Pro Ile Thr Phe Pro315
320 325tct gcc ttg gaa aga atg aga aat ggt
gaa ttt cct gat gtc gat gtc 1124Ser Ala Leu Glu Arg Met Arg Asn Gly
Glu Phe Pro Asp Val Asp Val330 335 340atc
att ggt ttc aat agt gct gaa ggt tta aga tct atg gca aga gta 1172Ile
Ile Gly Phe Asn Ser Ala Glu Gly Leu Arg Ser Met Ala Arg Val345
350 355acc aga gga aac atg gaa gtt cac aag act ttg
aca aat ata gaa agg 1220Thr Arg Gly Asn Met Glu Val His Lys Thr Leu
Thr Asn Ile Glu Arg360 365 370gct ata cct
aga gat gct aat att tgg aaa aat cca aat ggt att gag 1268Ala Ile Pro
Arg Asp Ala Asn Ile Trp Lys Asn Pro Asn Gly Ile Glu375
380 385 390gag aaa aaa cta ata aaa atg
ctt aca gag ttt tat gac caa gtg aaa 1316Glu Lys Lys Leu Ile Lys Met
Leu Thr Glu Phe Tyr Asp Gln Val Lys395 400
405gaa caa aac gat gac att gaa gcc tac gtc caa cta aaa ggc gat gct
1364Glu Gln Asn Asp Asp Ile Glu Ala Tyr Val Gln Leu Lys Gly Asp Ala410
415 420ggt tac ctc caa gga atc tac cgt acc
ttg aaa gcc ata ttt ttc aat 1412Gly Tyr Leu Gln Gly Ile Tyr Arg Thr
Leu Lys Ala Ile Phe Phe Asn425 430 435gaa
ttc aga agg aat tcc aat ttg tat ttg tac agg tta tca gac gat 1460Glu
Phe Arg Arg Asn Ser Asn Leu Tyr Leu Tyr Arg Leu Ser Asp Asp440
445 450acg tat agt gta tat aaa agt tat atc ttg ccc
tat cga tgg ggt tcc 1508Thr Tyr Ser Val Tyr Lys Ser Tyr Ile Leu Pro
Tyr Arg Trp Gly Ser455 460 465
470ttg cca gga gtt agt cat ggt gat gat tta gga tat ctt ttt gca aac
1556Leu Pro Gly Val Ser His Gly Asp Asp Leu Gly Tyr Leu Phe Ala Asn475
480 485tcg ttg gat gtt cct att ttg gga aca
acg cac att tct ata ccg caa 1604Ser Leu Asp Val Pro Ile Leu Gly Thr
Thr His Ile Ser Ile Pro Gln490 495 500gat
gct atg cag act ctg gaa agg atg gtc agg atc tgg acc aat ttt 1652Asp
Ala Met Gln Thr Leu Glu Arg Met Val Arg Ile Trp Thr Asn Phe505
510 515gta aag aat gga aaa cct aca tca aac act gaa
gat gca tca tgt gat 1700Val Lys Asn Gly Lys Pro Thr Ser Asn Thr Glu
Asp Ala Ser Cys Asp520 525 530aca aaa aga
cat tta aac gac att ttt tgg gaa cca tac aac gac gaa 1748Thr Lys Arg
His Leu Asn Asp Ile Phe Trp Glu Pro Tyr Asn Asp Glu535
540 545 550gaa cca aaa tat ttg gac atg
gga aaa gaa aat ttt gaa atg aaa aat 1796Glu Pro Lys Tyr Leu Asp Met
Gly Lys Glu Asn Phe Glu Met Lys Asn555 560
565att ttg gaa cta aaa cgc atg atg ctt tgg gat gaa gtt tat aga aat
1844Ile Leu Glu Leu Lys Arg Met Met Leu Trp Asp Glu Val Tyr Arg Asn570
575 580gcg aat ttg cgg ttt aga gtc tgt aat
gaa gaa agt att aga tga 1889Ala Asn Leu Arg Phe Arg Val Cys Asn
Glu Glu Ser Ile Arg585 590 595gtttttttaa
ttttacatac agccgagagg aaacatgact aaaattggaa agaaaaatca 1949gaaaaagaaa
aatcacatgg accatagtaa ctttattaca tgatttagtt tcaagtgtat 2009caagaaaact
tattgcatca aagaaaatat tattttgcca aaattcttgg aaaaacactt 2069tttatgactg
acatggccca taattgaagc tttttcttct tttaccaaat cgccaaattt 2129tgtagcgtca
gacacattta tttatgacat ggcaattaat gtgttaaaca ttcaactcta 2189tattaaaaat
ggtagtattt tctaataaga aggttatata aaaagacttg aaaataataa 2249gatttgctcg
gctatatata aaaacttanc gtctcgttat gctaaacttt tttgatggta 2309aaaatatgtt
gattttccta ataatctaag atattatatt ttagattaaa ttaaaatatg 2369atattttcaa
ttaattaatt ttagttttaa atgtactata tttaccagta ctatgaaact 2429attttaaata
tattttttat tacaatattt atttctcaaa aatgtttagt gtaacaagac 2489cattaaatta
gagttaatgt tgtaaattaa actatttttt atctatcaca accgcttaat 2549tggtgcaaag
aaaaatttta ctgtgataat atttgacatt tacacaatat tacgaattgt 2609aaactcacaa
ttatgtgaat attgtttttt gttaaaaaaa catacatgac ttttctatat 2669cattttatat
tacggtgata tggattaatg tcaacatgta aaatacaaat gcggttgtta 2729aaaataatct
gtattaaaat tgttatataa aatctgaata aatgtacttt taagtaaaaa 2789aaaaaaaaaa
aaaaaaaaaa aaaaaaaaaa aaaaaaaaaa aaaaaaa
283625596PRTCtenocephalides felis 25Met Leu Pro His Ser Ser Ala Leu Val
Leu Phe Leu Phe Phe Leu Phe1 5 10
15Phe Leu Phe Thr Pro Ile Leu Cys Ile Leu Trp Asp Asn Leu Asp
Gln20 25 30His Leu Cys Arg Val Gln Phe
Asn Gly Ile Thr Glu Gly Lys Pro Phe35 40
45Arg Tyr Lys Asp His Arg Asn Asp Val Tyr Cys Ser Tyr Leu Gly Ile50
55 60Pro Tyr Ala Glu Pro Pro Phe Gly Pro Leu
Arg Phe Gln Ser Pro Lys65 70 75
80Pro Ile Ser Asn Pro Lys Thr Gly Phe Val Gln Ala Arg Thr Leu
Gly85 90 95Asp Lys Cys Phe Gln Glu Ser
Leu Ile Tyr Ser Tyr Ala Gly Ser Glu100 105
110Asp Cys Leu Tyr Leu Asn Ile Phe Thr Pro Glu Thr Val Asn Ser Ala115
120 125Asn Asn Thr Lys Tyr Pro Val Met Phe
Trp Ile His Gly Gly Ala Phe130 135 140Asn
Gln Gly Ser Gly Ser Tyr Asn Phe Phe Gly Pro Asp Tyr Leu Ile145
150 155 160Arg Glu Gly Ile Ile Leu
Val Thr Ile Asn Tyr Arg Leu Gly Val Phe165 170
175Gly Phe Leu Ser Ala Pro Glu Trp Asp Ile His Gly Asn Met Gly
Leu180 185 190Lys Asp Gln Arg Leu Ala Leu
Lys Trp Val Tyr Asp Asn Ile Glu Lys195 200
205Phe Gly Gly Asp Arg Glu Lys Ile Thr Ile Ala Gly Glu Ser Ala Gly210
215 220Ala Ala Ser Val His Phe Leu Met Met
Asp Asn Ser Thr Arg Lys Tyr225 230 235
240Tyr Gln Arg Ala Ile Leu Gln Ser Gly Thr Leu Leu Asn Pro
Thr Ala245 250 255Asn Gln Ile Gln Leu Leu
His Arg Phe Glu Lys Leu Lys Gln Val Leu260 265
270Asn Ile Thr Gln Lys Gln Glu Leu Leu Asn Leu Asp Lys Asn Leu
Ile275 280 285Leu Arg Ala Ala Leu Asn Arg
Val Pro Asp Ser Asn Asp His Asp Arg290 295
300Asp Thr Val Pro Val Phe Asn Pro Val Leu Glu Ser Pro Glu Ser Pro305
310 315 320Asp Pro Ile Thr
Phe Pro Ser Ala Leu Glu Arg Met Arg Asn Gly Glu325 330
335Phe Pro Asp Val Asp Val Ile Ile Gly Phe Asn Ser Ala Glu
Gly Leu340 345 350Arg Ser Met Ala Arg Val
Thr Arg Gly Asn Met Glu Val His Lys Thr355 360
365Leu Thr Asn Ile Glu Arg Ala Ile Pro Arg Asp Ala Asn Ile Trp
Lys370 375 380Asn Pro Asn Gly Ile Glu Glu
Lys Lys Leu Ile Lys Met Leu Thr Glu385 390
395 400Phe Tyr Asp Gln Val Lys Glu Gln Asn Asp Asp Ile
Glu Ala Tyr Val405 410 415Gln Leu Lys Gly
Asp Ala Gly Tyr Leu Gln Gly Ile Tyr Arg Thr Leu420 425
430Lys Ala Ile Phe Phe Asn Glu Phe Arg Arg Asn Ser Asn Leu
Tyr Leu435 440 445Tyr Arg Leu Ser Asp Asp
Thr Tyr Ser Val Tyr Lys Ser Tyr Ile Leu450 455
460Pro Tyr Arg Trp Gly Ser Leu Pro Gly Val Ser His Gly Asp Asp
Leu465 470 475 480Gly Tyr
Leu Phe Ala Asn Ser Leu Asp Val Pro Ile Leu Gly Thr Thr485
490 495His Ile Ser Ile Pro Gln Asp Ala Met Gln Thr Leu
Glu Arg Met Val500 505 510Arg Ile Trp Thr
Asn Phe Val Lys Asn Gly Lys Pro Thr Ser Asn Thr515 520
525Glu Asp Ala Ser Cys Asp Thr Lys Arg His Leu Asn Asp Ile
Phe Trp530 535 540Glu Pro Tyr Asn Asp Glu
Glu Pro Lys Tyr Leu Asp Met Gly Lys Glu545 550
555 560Asn Phe Glu Met Lys Asn Ile Leu Glu Leu Lys
Arg Met Met Leu Trp565 570 575Asp Glu Val
Tyr Arg Asn Ala Asn Leu Arg Phe Arg Val Cys Asn Glu580
585 590Glu Ser Ile Arg595262836DNACtenocephalides
felismisc_feature(559)..(559)n = unknown 26tttttttttt tttttttttt
tttttttttt tttttttttt tttttttttt ttacttaaaa 60gtacatttat tcagatttta
tataacaatt ttaatacaga ttatttttaa caaccgcatt 120tgtattttac atgttgacat
taatccatat caccgtaata taaaatgata tagaaaagtc 180atgtatgttt ttttaacaaa
aaacaatatt cacataattg tgagtttaca attcgtaata 240ttgtgtaaat gtcaaatatt
atcacagtaa aatttttctt tgcaccaatt aagcggttgt 300gatagataaa aaatagttta
atttacaaca ttaactctaa tttaatggtc ttgttacact 360aaacattttt gagaaataaa
tattgtaata aaaaatatat ttaaaatagt ttcatagtac 420tggtaaatat agtacattta
aaactaaaat taattaattg aaaatatcat attttaattt 480aatctaaaat ataatatctt
agattattag gaaaatcaac atatttttac catcaaaaaa 540gtttagcata acgagacgnt
aagtttttat atatagccga gcaaatctta ttattttcaa 600gtctttttat ataaccttct
tattagaaaa tactaccatt tttaatatag agttgaatgt 660ttaacacatt aattgccatg
tcataaataa atgtgtctga cgctacaaaa tttggcgatt 720tggtaaaaga agaaaaagct
tcaattatgg gccatgtcag tcataaaaag tgtttttcca 780agaattttgg caaaataata
ttttctttga tgcaataagt tttcttgata cacttgaaac 840taaatcatgt aataaagtta
ctatggtcca tgtgattttt ctttttctga tttttctttc 900caattttagt catgtttcct
ctcggctgta tgtaaaatta aaaaaactca tctaatactt 960tcttcattac agactctaaa
ccgcaaattc gcatttctat aaacttcatc ccaaagcatc 1020atgcgtttta gttccaaaat
atttttcatt tcaaaatttt cttttcccat gtccaaatat 1080tttggttctt cgtcgttgta
tggttcccaa aaaatgtcgt ttaaatgtct ttttgtatca 1140catgatgcat cttcagtgtt
tgatgtaggt tttccattct ttacaaaatt ggtccagatc 1200ctgaccatcc tttccagagt
ctgcatagca tcttgcggta tagaaatgtg cgttgttccc 1260aaaataggaa catccaacga
gtttgcaaaa agatatccta aatcatcacc atgactaact 1320cctggcaagg aaccccatcg
atagggcaag atataacttt tatatacact atacgtatcg 1380tctgataacc tgtacaaata
caaattggaa ttccttctga attcattgaa aaatatggct 1440ttcaaggtac ggtagattcc
ttggaggtaa ccagcatcgc cttttagttg gacgtaggct 1500tcaatgtcat cgttttgttc
tttcacttgg tcataaaact ctgtaagcat ttttattagt 1560tttttctcct caataccatt
tggatttttc caaatattag catctctagg tatagccctt 1620tctatatttg tcaaagtctt
gtgaacttcc atgtttcctc tggttactct tgccatagat 1680cttaaacctt cagcactatt
gaaaccaatg atgacatcga catcaggaaa ttcaccattt 1740ctcattcttt ccaaggcaga
tggaaatgtt attggatctg gagattctgg tgattctaag 1800actggattaa atactggtac
tgtgtctcgg tcatggtcgt tgctatcagg aactctgttt 1860aaggctgctc gtaaaattag
gtttttatcc aggtttagga gttcttgttt ttgcgtgatg 1920tttagcactt gtttgagttt
ttcaaatcta tgcagaagtt gaatttgatt agcagtcgga 1980tttagtaatg tcccactctg
caaaatggcc ctttggtagt attttctagt cgagttgtcc 2040atcatcagaa aatggacact
tgctgctcca gcagattctc cagcaattgt aattttttct 2100ctgtctccac caaacttttc
gatgttgtcg taaacccatt ttagtgccaa tctctggtct 2160tttagaccca tatttccatg
gatatcccat tccggcgctg atagaaaacc gaaaactcct 2220aatctatagt tgatagtgac
caaaataatt ccttccctga tcaaataatc aggtccaaaa 2280aaattataag atcctgatcc
ttggttgaat gcgcctccat ggatccagaa cattacagga 2340tattttgtat tgttcgcaga
attaacagtc tctggcgtga atatattcag atataagcaa 2400tcttcgcttc ctgcataaga
atatattaga ctttcctgga aacatttgtc tcccaaagtt 2460cgagcctgta cgaatcctgt
ttttggattt gatattggtt ttggagactg aaatcgtaat 2520ggtccaaaag gcggttcggc
ataaggaatt cccaaataag aacaatatac atcattccta 2580tgatctttat atcggaacgg
ttttccttcc gtgatcccgt taaattgaac tctgcacaaa 2640tgctgatcta ggttatccca
tagtatgcac aagataggtg taaataagaa aaataaaaaa 2700aataaaaata aaactaatgc
actactgtga ggtaacattt tttattgtgt tttttaatgc 2760attttggttg cttaattgtt
attatttatc tcgttttgtt tatgataaaa tagacgtttt 2820gaagacgaca tgtcta
2836271710DNACtenocephalides
felisexon(1)..(1710) 27tgg gat aac cta gat cag cat ttg tgc aga gtt caa
ttt aac ggg atc 48Trp Asp Asn Leu Asp Gln His Leu Cys Arg Val Gln
Phe Asn Gly Ile1 5 10
15acg gaa gga aaa ccg ttc cga tat aaa gat cat agg aat gat gta tat
96Thr Glu Gly Lys Pro Phe Arg Tyr Lys Asp His Arg Asn Asp Val Tyr20
25 30tgt tct tat ttg gga att cct tat gcc gaa
ccg cct ttt gga cca tta 144Cys Ser Tyr Leu Gly Ile Pro Tyr Ala Glu
Pro Pro Phe Gly Pro Leu35 40 45cga ttt
cag tct cca aaa cca ata tca aat cca aaa aca gga ttc gta 192Arg Phe
Gln Ser Pro Lys Pro Ile Ser Asn Pro Lys Thr Gly Phe Val50
55 60cag gct cga act ttg gga gac aaa tgt ttc cag gaa
agt cta ata tat 240Gln Ala Arg Thr Leu Gly Asp Lys Cys Phe Gln Glu
Ser Leu Ile Tyr65 70 75
80tct tat gca gga agc gaa gat tgc tta tat ctg aat ata ttc acg cca
288Ser Tyr Ala Gly Ser Glu Asp Cys Leu Tyr Leu Asn Ile Phe Thr Pro85
90 95gag act gtt aat tct gcg aac aat aca aaa
tat cct gta atg ttc tgg 336Glu Thr Val Asn Ser Ala Asn Asn Thr Lys
Tyr Pro Val Met Phe Trp100 105 110atc cat
gga ggc gca ttc aac caa gga tca gga tct tat aat ttt ttt 384Ile His
Gly Gly Ala Phe Asn Gln Gly Ser Gly Ser Tyr Asn Phe Phe115
120 125gga cct gat tat ttg atc agg gaa gga att att ttg
gtc act atc aac 432Gly Pro Asp Tyr Leu Ile Arg Glu Gly Ile Ile Leu
Val Thr Ile Asn130 135 140tat aga tta gga
gtt ttc ggt ttt cta tca gcg ccg gaa tgg gat atc 480Tyr Arg Leu Gly
Val Phe Gly Phe Leu Ser Ala Pro Glu Trp Asp Ile145 150
155 160cat gga aat atg ggt cta aaa gac cag
aga ttg gca cta aaa tgg gtt 528His Gly Asn Met Gly Leu Lys Asp Gln
Arg Leu Ala Leu Lys Trp Val165 170 175tac
gac aac atc gaa aag ttt ggt gga gac aga gaa aaa att aca att 576Tyr
Asp Asn Ile Glu Lys Phe Gly Gly Asp Arg Glu Lys Ile Thr Ile180
185 190gct gga gaa tct gct gga gca gca agt gtc cat
ttt ctg atg atg gac 624Ala Gly Glu Ser Ala Gly Ala Ala Ser Val His
Phe Leu Met Met Asp195 200 205aac tcg act
aga aaa tac tac caa agg gcc att ttg cag agt ggg aca 672Asn Ser Thr
Arg Lys Tyr Tyr Gln Arg Ala Ile Leu Gln Ser Gly Thr210
215 220tta cta aat ccg act gct aat caa att caa ctt ctg
cat aga ttt gaa 720Leu Leu Asn Pro Thr Ala Asn Gln Ile Gln Leu Leu
His Arg Phe Glu225 230 235
240aaa ctc aaa caa gtg cta aac atc acg caa aaa caa gaa ctc cta aac
768Lys Leu Lys Gln Val Leu Asn Ile Thr Gln Lys Gln Glu Leu Leu Asn245
250 255ctg gat aaa aac cta att tta cga gca
gcc tta aac aga gtt cct gat 816Leu Asp Lys Asn Leu Ile Leu Arg Ala
Ala Leu Asn Arg Val Pro Asp260 265 270agc
aac gac cat gac cga gac aca gta cca gta ttt aat cca gtc tta 864Ser
Asn Asp His Asp Arg Asp Thr Val Pro Val Phe Asn Pro Val Leu275
280 285gaa tca cca gaa tct cca gat cca ata aca ttt
cca tct gcc ttg gaa 912Glu Ser Pro Glu Ser Pro Asp Pro Ile Thr Phe
Pro Ser Ala Leu Glu290 295 300aga atg aga
aat ggt gaa ttt cct gat gtc gat gtc atc att ggt ttc 960Arg Met Arg
Asn Gly Glu Phe Pro Asp Val Asp Val Ile Ile Gly Phe305
310 315 320aat agt gct gaa ggt tta aga
tct atg gca aga gta acc aga gga aac 1008Asn Ser Ala Glu Gly Leu Arg
Ser Met Ala Arg Val Thr Arg Gly Asn325 330
335atg gaa gtt cac aag act ttg aca aat ata gaa agg gct ata cct aga
1056Met Glu Val His Lys Thr Leu Thr Asn Ile Glu Arg Ala Ile Pro Arg340
345 350gat gct aat att tgg aaa aat cca aat
ggt att gag gag aaa aaa cta 1104Asp Ala Asn Ile Trp Lys Asn Pro Asn
Gly Ile Glu Glu Lys Lys Leu355 360 365ata
aaa atg ctt aca gag ttt tat gac caa gtg aaa gaa caa aac gat 1152Ile
Lys Met Leu Thr Glu Phe Tyr Asp Gln Val Lys Glu Gln Asn Asp370
375 380gac att gaa gcc tac gtc caa cta aaa ggc gat
gct ggt tac ctc caa 1200Asp Ile Glu Ala Tyr Val Gln Leu Lys Gly Asp
Ala Gly Tyr Leu Gln385 390 395
400gga atc tac cgt acc ttg aaa gcc ata ttt ttc aat gaa ttc aga agg
1248Gly Ile Tyr Arg Thr Leu Lys Ala Ile Phe Phe Asn Glu Phe Arg Arg405
410 415aat tcc aat ttg tat ttg tac agg tta
tca gac gat acg tat agt gta 1296Asn Ser Asn Leu Tyr Leu Tyr Arg Leu
Ser Asp Asp Thr Tyr Ser Val420 425 430tat
aaa agt tat atc ttg ccc tat cga tgg ggt tcc ttg cca gga gtt 1344Tyr
Lys Ser Tyr Ile Leu Pro Tyr Arg Trp Gly Ser Leu Pro Gly Val435
440 445agt cat ggt gat gat tta gga tat ctt ttt gca
aac tcg ttg gat gtt 1392Ser His Gly Asp Asp Leu Gly Tyr Leu Phe Ala
Asn Ser Leu Asp Val450 455 460cct att ttg
gga aca acg cac att tct ata ccg caa gat gct atg cag 1440Pro Ile Leu
Gly Thr Thr His Ile Ser Ile Pro Gln Asp Ala Met Gln465
470 475 480act ctg gaa agg atg gtc agg
atc tgg acc aat ttt gta aag aat gga 1488Thr Leu Glu Arg Met Val Arg
Ile Trp Thr Asn Phe Val Lys Asn Gly485 490
495aaa cct aca tca aac act gaa gat gca tca tgt gat aca aaa aga cat
1536Lys Pro Thr Ser Asn Thr Glu Asp Ala Ser Cys Asp Thr Lys Arg His500
505 510tta aac gac att ttt tgg gaa cca tac
aac gac gaa gaa cca aaa tat 1584Leu Asn Asp Ile Phe Trp Glu Pro Tyr
Asn Asp Glu Glu Pro Lys Tyr515 520 525ttg
gac atg gga aaa gaa aat ttt gaa atg aaa aat att ttg gaa cta 1632Leu
Asp Met Gly Lys Glu Asn Phe Glu Met Lys Asn Ile Leu Glu Leu530
535 540aaa cgc atg atg ctt tgg gat gaa gtt tat aga
aat gcg aat ttg cgg 1680Lys Arg Met Met Leu Trp Asp Glu Val Tyr Arg
Asn Ala Asn Leu Arg545 550 555
560ttt aga gtc tgt aat gaa gaa agt att aga
1710Phe Arg Val Cys Asn Glu Glu Ser Ile Arg565
570281788DNACtenocephalides felisexon(1)..(1788) 28atg tta cct cac agt
agt gca tta gtt tta ttt tta ttt ttt tta ttt 48Met Leu Pro His Ser
Ser Ala Leu Val Leu Phe Leu Phe Phe Leu Phe1 5
10 15ttc tta ttt aca cct atc ttg tgc ata cta tgg
gat aac cta gat cag 96Phe Leu Phe Thr Pro Ile Leu Cys Ile Leu Trp
Asp Asn Leu Asp Gln20 25 30cat ttg tgc
aga gtt caa ttt aac ggg atc acg gaa gga aaa ccg ttc 144His Leu Cys
Arg Val Gln Phe Asn Gly Ile Thr Glu Gly Lys Pro Phe35 40
45cga tat aaa gat cat agg aat gat gta tat tgt tct tat
ttg gga att 192Arg Tyr Lys Asp His Arg Asn Asp Val Tyr Cys Ser Tyr
Leu Gly Ile50 55 60cct tat gcc gaa ccg
cct ttt gga cca tta cga ttt cag tct cca aaa 240Pro Tyr Ala Glu Pro
Pro Phe Gly Pro Leu Arg Phe Gln Ser Pro Lys65 70
75 80cca ata tca aat cca aaa aca gga ttc gta
cag gct cga act ttg gga 288Pro Ile Ser Asn Pro Lys Thr Gly Phe Val
Gln Ala Arg Thr Leu Gly85 90 95gac aaa
tgt ttc cag gaa agt cta ata tat tct tat gca gga agc gaa 336Asp Lys
Cys Phe Gln Glu Ser Leu Ile Tyr Ser Tyr Ala Gly Ser Glu100
105 110gat tgc tta tat ctg aat ata ttc acg cca gag act
gtt aat tct gcg 384Asp Cys Leu Tyr Leu Asn Ile Phe Thr Pro Glu Thr
Val Asn Ser Ala115 120 125aac aat aca aaa
tat cct gta atg ttc tgg atc cat gga ggc gca ttc 432Asn Asn Thr Lys
Tyr Pro Val Met Phe Trp Ile His Gly Gly Ala Phe130 135
140aac caa gga tca gga tct tat aat ttt ttt gga cct gat tat
ttg atc 480Asn Gln Gly Ser Gly Ser Tyr Asn Phe Phe Gly Pro Asp Tyr
Leu Ile145 150 155 160agg
gaa gga att att ttg gtc act atc aac tat aga tta gga gtt ttc 528Arg
Glu Gly Ile Ile Leu Val Thr Ile Asn Tyr Arg Leu Gly Val Phe165
170 175ggt ttt cta tca gcg ccg gaa tgg gat atc cat
gga aat atg ggt cta 576Gly Phe Leu Ser Ala Pro Glu Trp Asp Ile His
Gly Asn Met Gly Leu180 185 190aaa gac cag
aga ttg gca cta aaa tgg gtt tac gac aac atc gaa aag 624Lys Asp Gln
Arg Leu Ala Leu Lys Trp Val Tyr Asp Asn Ile Glu Lys195
200 205ttt ggt gga gac aga gaa aaa att aca att gct gga
gaa tct gct gga 672Phe Gly Gly Asp Arg Glu Lys Ile Thr Ile Ala Gly
Glu Ser Ala Gly210 215 220gca gca agt gtc
cat ttt ctg atg atg gac aac tcg act aga aaa tac 720Ala Ala Ser Val
His Phe Leu Met Met Asp Asn Ser Thr Arg Lys Tyr225 230
235 240tac caa agg gcc att ttg cag agt ggg
aca tta cta aat ccg act gct 768Tyr Gln Arg Ala Ile Leu Gln Ser Gly
Thr Leu Leu Asn Pro Thr Ala245 250 255aat
caa att caa ctt ctg cat aga ttt gaa aaa ctc aaa caa gtg cta 816Asn
Gln Ile Gln Leu Leu His Arg Phe Glu Lys Leu Lys Gln Val Leu260
265 270aac atc acg caa aaa caa gaa ctc cta aac ctg
gat aaa aac cta att 864Asn Ile Thr Gln Lys Gln Glu Leu Leu Asn Leu
Asp Lys Asn Leu Ile275 280 285tta cga gca
gcc tta aac aga gtt cct gat agc aac gac cat gac cga 912Leu Arg Ala
Ala Leu Asn Arg Val Pro Asp Ser Asn Asp His Asp Arg290
295 300gac aca gta cca gta ttt aat cca gtc tta gaa tca
cca gaa tct cca 960Asp Thr Val Pro Val Phe Asn Pro Val Leu Glu Ser
Pro Glu Ser Pro305 310 315
320gat cca ata aca ttt cca tct gcc ttg gaa aga atg aga aat ggt gaa
1008Asp Pro Ile Thr Phe Pro Ser Ala Leu Glu Arg Met Arg Asn Gly Glu325
330 335ttt cct gat gtc gat gtc atc att ggt
ttc aat agt gct gaa ggt tta 1056Phe Pro Asp Val Asp Val Ile Ile Gly
Phe Asn Ser Ala Glu Gly Leu340 345 350aga
tct atg gca aga gta acc aga gga aac atg gaa gtt cac aag act 1104Arg
Ser Met Ala Arg Val Thr Arg Gly Asn Met Glu Val His Lys Thr355
360 365ttg aca aat ata gaa agg gct ata cct aga gat
gct aat att tgg aaa 1152Leu Thr Asn Ile Glu Arg Ala Ile Pro Arg Asp
Ala Asn Ile Trp Lys370 375 380aat cca aat
ggt att gag gag aaa aaa cta ata aaa atg ctt aca gag 1200Asn Pro Asn
Gly Ile Glu Glu Lys Lys Leu Ile Lys Met Leu Thr Glu385
390 395 400ttt tat gac caa gtg aaa gaa
caa aac gat gac att gaa gcc tac gtc 1248Phe Tyr Asp Gln Val Lys Glu
Gln Asn Asp Asp Ile Glu Ala Tyr Val405 410
415caa cta aaa ggc gat gct ggt tac ctc caa gga atc tac cgt acc ttg
1296Gln Leu Lys Gly Asp Ala Gly Tyr Leu Gln Gly Ile Tyr Arg Thr Leu420
425 430aaa gcc ata ttt ttc aat gaa ttc aga
agg aat tcc aat ttg tat ttg 1344Lys Ala Ile Phe Phe Asn Glu Phe Arg
Arg Asn Ser Asn Leu Tyr Leu435 440 445tac
agg tta tca gac gat acg tat agt gta tat aaa agt tat atc ttg 1392Tyr
Arg Leu Ser Asp Asp Thr Tyr Ser Val Tyr Lys Ser Tyr Ile Leu450
455 460ccc tat cga tgg ggt tcc ttg cca gga gtt agt
cat ggt gat gat tta 1440Pro Tyr Arg Trp Gly Ser Leu Pro Gly Val Ser
His Gly Asp Asp Leu465 470 475
480gga tat ctt ttt gca aac tcg ttg gat gtt cct att ttg gga aca acg
1488Gly Tyr Leu Phe Ala Asn Ser Leu Asp Val Pro Ile Leu Gly Thr Thr485
490 495cac att tct ata ccg caa gat gct atg
cag act ctg gaa agg atg gtc 1536His Ile Ser Ile Pro Gln Asp Ala Met
Gln Thr Leu Glu Arg Met Val500 505 510agg
atc tgg acc aat ttt gta aag aat gga aaa cct aca tca aac act 1584Arg
Ile Trp Thr Asn Phe Val Lys Asn Gly Lys Pro Thr Ser Asn Thr515
520 525gaa gat gca tca tgt gat aca aaa aga cat tta
aac gac att ttt tgg 1632Glu Asp Ala Ser Cys Asp Thr Lys Arg His Leu
Asn Asp Ile Phe Trp530 535 540gaa cca tac
aac gac gaa gaa cca aaa tat ttg gac atg gga aaa gaa 1680Glu Pro Tyr
Asn Asp Glu Glu Pro Lys Tyr Leu Asp Met Gly Lys Glu545
550 555 560aat ttt gaa atg aaa aat att
ttg gaa cta aaa cgc atg atg ctt tgg 1728Asn Phe Glu Met Lys Asn Ile
Leu Glu Leu Lys Arg Met Met Leu Trp565 570
575gat gaa gtt tat aga aat gcg aat ttg cgg ttt aga gtc tgt aat gaa
1776Asp Glu Val Tyr Arg Asn Ala Asn Leu Arg Phe Arg Val Cys Asn Glu580
585 590gaa agt att aga
1788Glu Ser Ile
Arg595291788DNACtenocephalides felis 29tctaatactt tcttcattac agactctaaa
ccgcaaattc gcatttctat aaacttcatc 60ccaaagcatc atgcgtttta gttccaaaat
atttttcatt tcaaaatttt cttttcccat 120gtccaaatat tttggttctt cgtcgttgta
tggttcccaa aaaatgtcgt ttaaatgtct 180ttttgtatca catgatgcat cttcagtgtt
tgatgtaggt tttccattct ttacaaaatt 240ggtccagatc ctgaccatcc tttccagagt
ctgcatagca tcttgcggta tagaaatgtg 300cgttgttccc aaaataggaa catccaacga
gtttgcaaaa agatatccta aatcatcacc 360atgactaact cctggcaagg aaccccatcg
atagggcaag atataacttt tatatacact 420atacgtatcg tctgataacc tgtacaaata
caaattggaa ttccttctga attcattgaa 480aaatatggct ttcaaggtac ggtagattcc
ttggaggtaa ccagcatcgc cttttagttg 540gacgtaggct tcaatgtcat cgttttgttc
tttcacttgg tcataaaact ctgtaagcat 600ttttattagt tttttctcct caataccatt
tggatttttc caaatattag catctctagg 660tatagccctt tctatatttg tcaaagtctt
gtgaacttcc atgtttcctc tggttactct 720tgccatagat cttaaacctt cagcactatt
gaaaccaatg atgacatcga catcaggaaa 780ttcaccattt ctcattcttt ccaaggcaga
tggaaatgtt attggatctg gagattctgg 840tgattctaag actggattaa atactggtac
tgtgtctcgg tcatggtcgt tgctatcagg 900aactctgttt aaggctgctc gtaaaattag
gtttttatcc aggtttagga gttcttgttt 960ttgcgtgatg tttagcactt gtttgagttt
ttcaaatcta tgcagaagtt gaatttgatt 1020agcagtcgga tttagtaatg tcccactctg
caaaatggcc ctttggtagt attttctagt 1080cgagttgtcc atcatcagaa aatggacact
tgctgctcca gcagattctc cagcaattgt 1140aattttttct ctgtctccac caaacttttc
gatgttgtcg taaacccatt ttagtgccaa 1200tctctggtct tttagaccca tatttccatg
gatatcccat tccggcgctg atagaaaacc 1260gaaaactcct aatctatagt tgatagtgac
caaaataatt ccttccctga tcaaataatc 1320aggtccaaaa aaattataag atcctgatcc
ttggttgaat gcgcctccat ggatccagaa 1380cattacagga tattttgtat tgttcgcaga
attaacagtc tctggcgtga atatattcag 1440atataagcaa tcttcgcttc ctgcataaga
atatattaga ctttcctgga aacatttgtc 1500tcccaaagtt cgagcctgta cgaatcctgt
ttttggattt gatattggtt ttggagactg 1560aaatcgtaat ggtccaaaag gcggttcggc
ataaggaatt cccaaataag aacaatatac 1620atcattccta tgatctttat atcggaacgg
ttttccttcc gtgatcccgt taaattgaac 1680tctgcacaaa tgctgatcta ggttatccca
tagtatgcac aagataggtg taaataagaa 1740aaataaaaaa aataaaaata aaactaatgc
actactgtga ggtaacat 1788302801DNACtenocephalides
felisCDS(99)..(1886)misc_feature(2275)..(2275)n = unknown 30gacatgtcgt
cttcaaaacg tctattttat cataaacaaa acgagataaa taataacaat 60taagcatcca
aaatgcatta aaaaaaacat cataaaaa atg tta cct cac agt gca 116Met Leu Pro
His Ser Ala1 5tta gtt tta ttt tta ttt ttt tta ttt ttc tta
ttt aca cct gtc ttg 164Leu Val Leu Phe Leu Phe Phe Leu Phe Phe Leu
Phe Thr Pro Val Leu10 15 20tgc ata cta
tgg gat aac cta gat cag cat ttg tgc aga gtt caa ttt 212Cys Ile Leu
Trp Asp Asn Leu Asp Gln His Leu Cys Arg Val Gln Phe25 30
35aac ggg atc acg gaa gga aaa ccg ttc cga tat aaa gat
cat aaa aat 260Asn Gly Ile Thr Glu Gly Lys Pro Phe Arg Tyr Lys Asp
His Lys Asn40 45 50gat gta tat tgt tcc
tat ttg gga att cct tat gca gaa ccg cct att 308Asp Val Tyr Cys Ser
Tyr Leu Gly Ile Pro Tyr Ala Glu Pro Pro Ile55 60
65 70gga cca ttg cga ttt cag tct cca aaa cca
ata tca aat cca aaa aca 356Gly Pro Leu Arg Phe Gln Ser Pro Lys Pro
Ile Ser Asn Pro Lys Thr75 80 85gga ttc
gtt cag gct cgg tct tta gga gac aaa tgt ttc cag gaa agt 404Gly Phe
Val Gln Ala Arg Ser Leu Gly Asp Lys Cys Phe Gln Glu Ser90
95 100cta ata tat tct tat gca gga agc gaa gat tgc tta
tat ctg aat ata 452Leu Ile Tyr Ser Tyr Ala Gly Ser Glu Asp Cys Leu
Tyr Leu Asn Ile105 110 115ttc acg cca gag
act gtt aat tct gcg aac aat aca aaa tat cct gta 500Phe Thr Pro Glu
Thr Val Asn Ser Ala Asn Asn Thr Lys Tyr Pro Val120 125
130atg ttc tgg atc cat gga ggc gca ttc aac caa gga tca gga
tct tat 548Met Phe Trp Ile His Gly Gly Ala Phe Asn Gln Gly Ser Gly
Ser Tyr135 140 145 150aat
ttt ttt gga cct gat tat ttg atc agg gaa gga att att ttg gtc 596Asn
Phe Phe Gly Pro Asp Tyr Leu Ile Arg Glu Gly Ile Ile Leu Val155
160 165act atc aac tat aga tta gga gtt ttc ggt ttt
cta tca gcg ccg gaa 644Thr Ile Asn Tyr Arg Leu Gly Val Phe Gly Phe
Leu Ser Ala Pro Glu170 175 180tgg gat atc
cat gga aat atg ggt cta aaa gac cag aga ttg gca cta 692Trp Asp Ile
His Gly Asn Met Gly Leu Lys Asp Gln Arg Leu Ala Leu185
190 195aaa tgg gtt tat gac aac atc gaa aaa ttt ggt gga
gac aga gat aaa 740Lys Trp Val Tyr Asp Asn Ile Glu Lys Phe Gly Gly
Asp Arg Asp Lys200 205 210atc act ata gct
gga gaa tct gct gga gca gca agt gtt cat ttt ctg 788Ile Thr Ile Ala
Gly Glu Ser Ala Gly Ala Ala Ser Val His Phe Leu215 220
225 230atg atg gac aat tct act aga aaa tac
tac caa agg gca att ttg cag 836Met Met Asp Asn Ser Thr Arg Lys Tyr
Tyr Gln Arg Ala Ile Leu Gln235 240 245agt
ggg aca tta ctc aat ccg act gct aat caa att caa cct ctg cat 884Ser
Gly Thr Leu Leu Asn Pro Thr Ala Asn Gln Ile Gln Pro Leu His250
255 260aga ttt gaa aaa cta aaa caa gtg ctg aac atc
acg caa aaa caa gaa 932Arg Phe Glu Lys Leu Lys Gln Val Leu Asn Ile
Thr Gln Lys Gln Glu265 270 275ctc cta aat
ctg gac aaa aat caa att ttg cga gca gcc tta aac aga 980Leu Leu Asn
Leu Asp Lys Asn Gln Ile Leu Arg Ala Ala Leu Asn Arg280
285 290gtc cca gat aac aac gac cac gaa agg gac aca gta
cca gta ttt aat 1028Val Pro Asp Asn Asn Asp His Glu Arg Asp Thr Val
Pro Val Phe Asn295 300 305
310cca gtc cta gaa tca cca gaa tct cca gac cca ata aca ttt cca tct
1076Pro Val Leu Glu Ser Pro Glu Ser Pro Asp Pro Ile Thr Phe Pro Ser315
320 325gct tta gaa aga atg aga aat ggt gaa
ttt cct gac gtt gat gtc atc 1124Ala Leu Glu Arg Met Arg Asn Gly Glu
Phe Pro Asp Val Asp Val Ile330 335 340att
gga ttc aat agt gct gaa ggt tta aga tct atg cca aga gta acc 1172Ile
Gly Phe Asn Ser Ala Glu Gly Leu Arg Ser Met Pro Arg Val Thr345
350 355aga gga aac atg gaa gtt tac aag act ttg aca
aat ata gag aga gct 1220Arg Gly Asn Met Glu Val Tyr Lys Thr Leu Thr
Asn Ile Glu Arg Ala360 365 370ata cct aga
gat gct aat att tgg aaa aat cct aat ggc att gag gag 1268Ile Pro Arg
Asp Ala Asn Ile Trp Lys Asn Pro Asn Gly Ile Glu Glu375
380 385 390aaa aaa ctt ata aaa atg ctt
aca gag ttt tat gac caa gtt aaa gaa 1316Lys Lys Leu Ile Lys Met Leu
Thr Glu Phe Tyr Asp Gln Val Lys Glu395 400
405caa aac gat gac atc gaa gcc tat gtc caa cta aaa ggc gat gct ggt
1364Gln Asn Asp Asp Ile Glu Ala Tyr Val Gln Leu Lys Gly Asp Ala Gly410
415 420tat ctc caa gga att tac cgt acc ttg
aaa gcc ata ttt ttc aat gaa 1412Tyr Leu Gln Gly Ile Tyr Arg Thr Leu
Lys Ala Ile Phe Phe Asn Glu425 430 435atc
aaa aga aat tcc aac ttg tat ttg tat agg tta tca gat gat acg 1460Ile
Lys Arg Asn Ser Asn Leu Tyr Leu Tyr Arg Leu Ser Asp Asp Thr440
445 450tat agt gta tat aaa agt tat atc ttg ccc tat
cga tgg ggt tcc ttg 1508Tyr Ser Val Tyr Lys Ser Tyr Ile Leu Pro Tyr
Arg Trp Gly Ser Leu455 460 465
470cca gga gtt agt cat ggt gat gat tta gga tat ctt ttt gca aac tct
1556Pro Gly Val Ser His Gly Asp Asp Leu Gly Tyr Leu Phe Ala Asn Ser475
480 485ttg gat gtt cct att ttg gga aca acg
cac att tct ata ccg caa gat 1604Leu Asp Val Pro Ile Leu Gly Thr Thr
His Ile Ser Ile Pro Gln Asp490 495 500gct
atg cag act ctg gaa agg atg gtc agg atc tgg acc aat ttt gta 1652Ala
Met Gln Thr Leu Glu Arg Met Val Arg Ile Trp Thr Asn Phe Val505
510 515aag aat gga aaa cct aca tca aac act gaa gat
gca tca tgt gat aca 1700Lys Asn Gly Lys Pro Thr Ser Asn Thr Glu Asp
Ala Ser Cys Asp Thr520 525 530aaa aga cat
tta aac gac att ttt tgg gaa cca tac aac gac gaa gaa 1748Lys Arg His
Leu Asn Asp Ile Phe Trp Glu Pro Tyr Asn Asp Glu Glu535
540 545 550cca aaa tat ttg gac atg gga
aaa gaa cat ttt gaa atg aaa aat att 1796Pro Lys Tyr Leu Asp Met Gly
Lys Glu His Phe Glu Met Lys Asn Ile555 560
565ttg gaa cta aaa cgc atg atg ctt tgg gat gaa gtt tat aga aat gcg
1844Leu Glu Leu Lys Arg Met Met Leu Trp Asp Glu Val Tyr Arg Asn Ala570
575 580aat ttg cgg ttt aga gtc tgt aat gaa
gaa agt att aga tga 1886Asn Leu Arg Phe Arg Val Cys Asn Glu
Glu Ser Ile Arg585 590 595gtttttttaa
ttttacatac agccgagagg aaacatgact aaaattggaa agaaaaatca 1946gaaaaagaaa
aatcacatgg accatagtaa ctttattaca tgatttagtt tcaagtgtat 2006caagaaaact
tattgcatca aagaaaatat tattttgcca aaattcttgg aaaaacactt 2066tttatgactg
acatggccca taattgaagc tttttcttct tttaccaaat cgccaaattt 2126tgtagcgtca
gacacattta tttatgacat ggcaattaat gtgttaaaca ttcaactcta 2186tattaaaaat
ggtagtattt tctaataaga aggttatata aaaagacttg aaaataataa 2246gatttgctcg
gctatatata aaaacttanc gtctcgttat gctaaacttt tttgatggta 2306aaaatatgtt
gattttccta ataatctaag atattatatt ttagattaaa ttaaaatatg 2366atattttcaa
ttaattaatt ttagttttaa atgtactata tttaccagta ctatgaaact 2426attttaaata
tattttttat tacaatattt atttctcaaa aatgtttagt gtaacaagac 2486cattaaatta
gagttaatgt tgtaaattaa actatttttt atctatcaca accgcttaat 2546tggtgcaaag
aaaaatttta ctgtgataat atttgacatt tacacaatat tacgaattgt 2606aaactcacaa
ttatgtgaat attgtttttt gttaaaaaaa catacatgac ttttctatat 2666cattttatat
tacggtgata tggattaatg tcaacatgta aaatacaaat gcggttgtta 2726aaaataatct
gtattaaaat tgttatataa aatctgaata aatgtacttt taagtaaaaa 2786aaaaaaaaaa
aaaaa
280131595PRTCtenocephalides felis 31Met Leu Pro His Ser Ala Leu Val Leu
Phe Leu Phe Phe Leu Phe Phe1 5 10
15Leu Phe Thr Pro Val Leu Cys Ile Leu Trp Asp Asn Leu Asp Gln
His20 25 30Leu Cys Arg Val Gln Phe Asn
Gly Ile Thr Glu Gly Lys Pro Phe Arg35 40
45Tyr Lys Asp His Lys Asn Asp Val Tyr Cys Ser Tyr Leu Gly Ile Pro50
55 60Tyr Ala Glu Pro Pro Ile Gly Pro Leu Arg
Phe Gln Ser Pro Lys Pro65 70 75
80Ile Ser Asn Pro Lys Thr Gly Phe Val Gln Ala Arg Ser Leu Gly
Asp85 90 95Lys Cys Phe Gln Glu Ser Leu
Ile Tyr Ser Tyr Ala Gly Ser Glu Asp100 105
110Cys Leu Tyr Leu Asn Ile Phe Thr Pro Glu Thr Val Asn Ser Ala Asn115
120 125Asn Thr Lys Tyr Pro Val Met Phe Trp
Ile His Gly Gly Ala Phe Asn130 135 140Gln
Gly Ser Gly Ser Tyr Asn Phe Phe Gly Pro Asp Tyr Leu Ile Arg145
150 155 160Glu Gly Ile Ile Leu Val
Thr Ile Asn Tyr Arg Leu Gly Val Phe Gly165 170
175Phe Leu Ser Ala Pro Glu Trp Asp Ile His Gly Asn Met Gly Leu
Lys180 185 190Asp Gln Arg Leu Ala Leu Lys
Trp Val Tyr Asp Asn Ile Glu Lys Phe195 200
205Gly Gly Asp Arg Asp Lys Ile Thr Ile Ala Gly Glu Ser Ala Gly Ala210
215 220Ala Ser Val His Phe Leu Met Met Asp
Asn Ser Thr Arg Lys Tyr Tyr225 230 235
240Gln Arg Ala Ile Leu Gln Ser Gly Thr Leu Leu Asn Pro Thr
Ala Asn245 250 255Gln Ile Gln Pro Leu His
Arg Phe Glu Lys Leu Lys Gln Val Leu Asn260 265
270Ile Thr Gln Lys Gln Glu Leu Leu Asn Leu Asp Lys Asn Gln Ile
Leu275 280 285Arg Ala Ala Leu Asn Arg Val
Pro Asp Asn Asn Asp His Glu Arg Asp290 295
300Thr Val Pro Val Phe Asn Pro Val Leu Glu Ser Pro Glu Ser Pro Asp305
310 315 320Pro Ile Thr Phe
Pro Ser Ala Leu Glu Arg Met Arg Asn Gly Glu Phe325 330
335Pro Asp Val Asp Val Ile Ile Gly Phe Asn Ser Ala Glu Gly
Leu Arg340 345 350Ser Met Pro Arg Val Thr
Arg Gly Asn Met Glu Val Tyr Lys Thr Leu355 360
365Thr Asn Ile Glu Arg Ala Ile Pro Arg Asp Ala Asn Ile Trp Lys
Asn370 375 380Pro Asn Gly Ile Glu Glu Lys
Lys Leu Ile Lys Met Leu Thr Glu Phe385 390
395 400Tyr Asp Gln Val Lys Glu Gln Asn Asp Asp Ile Glu
Ala Tyr Val Gln405 410 415Leu Lys Gly Asp
Ala Gly Tyr Leu Gln Gly Ile Tyr Arg Thr Leu Lys420 425
430Ala Ile Phe Phe Asn Glu Ile Lys Arg Asn Ser Asn Leu Tyr
Leu Tyr435 440 445Arg Leu Ser Asp Asp Thr
Tyr Ser Val Tyr Lys Ser Tyr Ile Leu Pro450 455
460Tyr Arg Trp Gly Ser Leu Pro Gly Val Ser His Gly Asp Asp Leu
Gly465 470 475 480Tyr Leu
Phe Ala Asn Ser Leu Asp Val Pro Ile Leu Gly Thr Thr His485
490 495Ile Ser Ile Pro Gln Asp Ala Met Gln Thr Leu Glu
Arg Met Val Arg500 505 510Ile Trp Thr Asn
Phe Val Lys Asn Gly Lys Pro Thr Ser Asn Thr Glu515 520
525Asp Ala Ser Cys Asp Thr Lys Arg His Leu Asn Asp Ile Phe
Trp Glu530 535 540Pro Tyr Asn Asp Glu Glu
Pro Lys Tyr Leu Asp Met Gly Lys Glu His545 550
555 560Phe Glu Met Lys Asn Ile Leu Glu Leu Lys Arg
Met Met Leu Trp Asp565 570 575Glu Val Tyr
Arg Asn Ala Asn Leu Arg Phe Arg Val Cys Asn Glu Glu580
585 590Ser Ile Arg595322801DNACtenocephalides
felismisc_feature(527)..(527)n = unknown at position 527 32tttttttttt
tttttttttt acttaaaagt acatttattc agattttata taacaatttt 60aatacagatt
atttttaaca accgcatttg tattttacat gttgacatta atccatatca 120ccgtaatata
aaatgatata gaaaagtcat gtatgttttt ttaacaaaaa acaatattca 180cataattgtg
agtttacaat tcgtaatatt gtgtaaatgt caaatattat cacagtaaaa 240tttttctttg
caccaattaa gcggttgtga tagataaaaa atagtttaat ttacaacatt 300aactctaatt
taatggtctt gttacactaa acatttttga gaaataaata ttgtaataaa 360aaatatattt
aaaatagttt catagtactg gtaaatatag tacatttaaa actaaaatta 420attaattgaa
aatatcatat tttaatttaa tctaaaatat aatatcttag attattagga 480aaatcaacat
atttttacca tcaaaaaagt ttagcataac gagacgntaa gtttttatat 540atagccgagc
aaatcttatt attttcaagt ctttttatat aaccttctta ttagaaaata 600ctaccatttt
taatatagag ttgaatgttt aacacattaa ttgccatgtc ataaataaat 660gtgtctgacg
ctacaaaatt tggcgatttg gtaaaagaag aaaaagcttc aattatgggc 720catgtcagtc
ataaaaagtg tttttccaag aattttggca aaataatatt ttctttgatg 780caataagttt
tcttgataca cttgaaacta aatcatgtaa taaagttact atggtccatg 840tgatttttct
ttttctgatt tttctttcca attttagtca tgtttcctct cggctgtatg 900taaaattaaa
aaaactcatc taatactttc ttcattacag actctaaacc gcaaattcgc 960atttctataa
acttcatccc aaagcatcat gcgttttagt tccaaaatat ttttcatttc 1020aaaatgttct
tttcccatgt ccaaatattt tggttcttcg tcgttgtatg gttcccaaaa 1080aatgtcgttt
aaatgtcttt ttgtatcaca tgatgcatct tcagtgtttg atgtaggttt 1140tccattcttt
acaaaattgg tccagatcct gaccatcctt tccagagtct gcatagcatc 1200ttgcggtata
gaaatgtgcg ttgttcccaa aataggaaca tccaaagagt ttgcaaaaag 1260atatcctaaa
tcatcaccat gactaactcc tggcaaggaa ccccatcgat agggcaagat 1320ataactttta
tatacactat acgtatcatc tgataaccta tacaaataca agttggaatt 1380tcttttgatt
tcattgaaaa atatggcttt caaggtacgg taaattcctt ggagataacc 1440agcatcgcct
tttagttgga cataggcttc gatgtcatcg ttttgttctt taacttggtc 1500ataaaactct
gtaagcattt ttataagttt tttctcctca atgccattag gatttttcca 1560aatattagca
tctctaggta tagctctctc tatatttgtc aaagtcttgt aaacttccat 1620gtttcctctg
gttactcttg gcatagatct taaaccttca gcactattga atccaatgat 1680gacatcaacg
tcaggaaatt caccatttct cattctttct aaagcagatg gaaatgttat 1740tgggtctgga
gattctggtg attctaggac tggattaaat actggtactg tgtccctttc 1800gtggtcgttg
ttatctggga ctctgtttaa ggctgctcgc aaaatttgat ttttgtccag 1860atttaggagt
tcttgttttt gcgtgatgtt cagcacttgt tttagttttt caaatctatg 1920cagaggttga
atttgattag cagtcggatt gagtaatgtc ccactctgca aaattgccct 1980ttggtagtat
tttctagtag aattgtccat catcagaaaa tgaacacttg ctgctccagc 2040agattctcca
gctatagtga ttttatctct gtctccacca aatttttcga tgttgtcata 2100aacccatttt
agtgccaatc tctggtcttt tagacccata tttccatgga tatcccattc 2160cggcgctgat
agaaaaccga aaactcctaa tctatagttg atagtgacca aaataattcc 2220ttccctgatc
aaataatcag gtccaaaaaa attataagat cctgatcctt ggttgaatgc 2280gcctccatgg
atccagaaca ttacaggata ttttgtattg ttcgcagaat taacagtctc 2340tggcgtgaat
atattcagat ataagcaatc ttcgcttcct gcataagaat atattagact 2400ttcctggaaa
catttgtctc ctaaagaccg agcctgaacg aatcctgttt ttggatttga 2460tattggtttt
ggagactgaa atcgcaatgg tccaataggc ggttctgcat aaggaattcc 2520caaataggaa
caatatacat catttttatg atctttatat cggaacggtt ttccttccgt 2580gatcccgtta
aattgaactc tgcacaaatg ctgatctagg ttatcccata gtatgcacaa 2640gacaggtgta
aataagaaaa ataaaaaaaa taaaaataaa actaatgcac tgtgaggtaa 2700cattttttat
gatgtttttt ttaatgcatt ttggatgctt aattgttatt atttatctcg 2760ttttgtttat
gataaaatag acgttttgaa gacgacatgt c
2801331710DNACtenocephalides felisexon(1)..(1710) 33tgg gat aac cta gat
cag cat ttg tgc aga gtt caa ttt aac ggg atc 48Trp Asp Asn Leu Asp
Gln His Leu Cys Arg Val Gln Phe Asn Gly Ile1 5
10 15acg gaa gga aaa ccg ttc cga tat aaa gat cat
aaa aat gat gta tat 96Thr Glu Gly Lys Pro Phe Arg Tyr Lys Asp His
Lys Asn Asp Val Tyr20 25 30tgt tcc tat
ttg gga att cct tat gca gaa ccg cct att gga cca ttg 144Cys Ser Tyr
Leu Gly Ile Pro Tyr Ala Glu Pro Pro Ile Gly Pro Leu35 40
45cga ttt cag tct cca aaa cca ata tca aat cca aaa aca
gga ttc gtt 192Arg Phe Gln Ser Pro Lys Pro Ile Ser Asn Pro Lys Thr
Gly Phe Val50 55 60cag gct cgg tct tta
gga gac aaa tgt ttc cag gaa agt cta ata tat 240Gln Ala Arg Ser Leu
Gly Asp Lys Cys Phe Gln Glu Ser Leu Ile Tyr65 70
75 80tct tat gca gga agc gaa gat tgc tta tat
ctg aat ata ttc acg cca 288Ser Tyr Ala Gly Ser Glu Asp Cys Leu Tyr
Leu Asn Ile Phe Thr Pro85 90 95gag act
gtt aat tct gcg aac aat aca aaa tat cct gta atg ttc tgg 336Glu Thr
Val Asn Ser Ala Asn Asn Thr Lys Tyr Pro Val Met Phe Trp100
105 110atc cat gga ggc gca ttc aac caa gga tca gga tct
tat aat ttt ttt 384Ile His Gly Gly Ala Phe Asn Gln Gly Ser Gly Ser
Tyr Asn Phe Phe115 120 125gga cct gat tat
ttg atc agg gaa gga att att ttg gtc act atc aac 432Gly Pro Asp Tyr
Leu Ile Arg Glu Gly Ile Ile Leu Val Thr Ile Asn130 135
140tat aga tta gga gtt ttc ggt ttt cta tca gcg ccg gaa tgg
gat atc 480Tyr Arg Leu Gly Val Phe Gly Phe Leu Ser Ala Pro Glu Trp
Asp Ile145 150 155 160cat
gga aat atg ggt cta aaa gac cag aga ttg gca cta aaa tgg gtt 528His
Gly Asn Met Gly Leu Lys Asp Gln Arg Leu Ala Leu Lys Trp Val165
170 175tat gac aac atc gaa aaa ttt ggt gga gac aga
gat aaa atc act ata 576Tyr Asp Asn Ile Glu Lys Phe Gly Gly Asp Arg
Asp Lys Ile Thr Ile180 185 190gct gga gaa
tct gct gga gca gca agt gtt cat ttt ctg atg atg gac 624Ala Gly Glu
Ser Ala Gly Ala Ala Ser Val His Phe Leu Met Met Asp195
200 205aat tct act aga aaa tac tac caa agg gca att ttg
cag agt ggg aca 672Asn Ser Thr Arg Lys Tyr Tyr Gln Arg Ala Ile Leu
Gln Ser Gly Thr210 215 220tta ctc aat ccg
act gct aat caa att caa cct ctg cat aga ttt gaa 720Leu Leu Asn Pro
Thr Ala Asn Gln Ile Gln Pro Leu His Arg Phe Glu225 230
235 240aaa cta aaa caa gtg ctg aac atc acg
caa aaa caa gaa ctc cta aat 768Lys Leu Lys Gln Val Leu Asn Ile Thr
Gln Lys Gln Glu Leu Leu Asn245 250 255ctg
gac aaa aat caa att ttg cga gca gcc tta aac aga gtc cca gat 816Leu
Asp Lys Asn Gln Ile Leu Arg Ala Ala Leu Asn Arg Val Pro Asp260
265 270aac aac gac cac gaa agg gac aca gta cca gta
ttt aat cca gtc cta 864Asn Asn Asp His Glu Arg Asp Thr Val Pro Val
Phe Asn Pro Val Leu275 280 285gaa tca cca
gaa tct cca gac cca ata aca ttt cca tct gct tta gaa 912Glu Ser Pro
Glu Ser Pro Asp Pro Ile Thr Phe Pro Ser Ala Leu Glu290
295 300aga atg aga aat ggt gaa ttt cct gac gtt gat gtc
atc att gga ttc 960Arg Met Arg Asn Gly Glu Phe Pro Asp Val Asp Val
Ile Ile Gly Phe305 310 315
320aat agt gct gaa ggt tta aga tct atg cca aga gta acc aga gga aac
1008Asn Ser Ala Glu Gly Leu Arg Ser Met Pro Arg Val Thr Arg Gly Asn325
330 335atg gaa gtt tac aag act ttg aca aat
ata gag aga gct ata cct aga 1056Met Glu Val Tyr Lys Thr Leu Thr Asn
Ile Glu Arg Ala Ile Pro Arg340 345 350gat
gct aat att tgg aaa aat cct aat ggc att gag gag aaa aaa ctt 1104Asp
Ala Asn Ile Trp Lys Asn Pro Asn Gly Ile Glu Glu Lys Lys Leu355
360 365ata aaa atg ctt aca gag ttt tat gac caa gtt
aaa gaa caa aac gat 1152Ile Lys Met Leu Thr Glu Phe Tyr Asp Gln Val
Lys Glu Gln Asn Asp370 375 380gac atc gaa
gcc tat gtc caa cta aaa ggc gat gct ggt tat ctc caa 1200Asp Ile Glu
Ala Tyr Val Gln Leu Lys Gly Asp Ala Gly Tyr Leu Gln385
390 395 400gga att tac cgt acc ttg aaa
gcc ata ttt ttc aat gaa atc aaa aga 1248Gly Ile Tyr Arg Thr Leu Lys
Ala Ile Phe Phe Asn Glu Ile Lys Arg405 410
415aat tcc aac ttg tat ttg tat agg tta tca gat gat acg tat agt gta
1296Asn Ser Asn Leu Tyr Leu Tyr Arg Leu Ser Asp Asp Thr Tyr Ser Val420
425 430tat aaa agt tat atc ttg ccc tat cga
tgg ggt tcc ttg cca gga gtt 1344Tyr Lys Ser Tyr Ile Leu Pro Tyr Arg
Trp Gly Ser Leu Pro Gly Val435 440 445agt
cat ggt gat gat tta gga tat ctt ttt gca aac tct ttg gat gtt 1392Ser
His Gly Asp Asp Leu Gly Tyr Leu Phe Ala Asn Ser Leu Asp Val450
455 460cct att ttg gga aca acg cac att tct ata ccg
caa gat gct atg cag 1440Pro Ile Leu Gly Thr Thr His Ile Ser Ile Pro
Gln Asp Ala Met Gln465 470 475
480act ctg gaa agg atg gtc agg atc tgg acc aat ttt gta aag aat gga
1488Thr Leu Glu Arg Met Val Arg Ile Trp Thr Asn Phe Val Lys Asn Gly485
490 495aaa cct aca tca aac act gaa gat gca
tca tgt gat aca aaa aga cat 1536Lys Pro Thr Ser Asn Thr Glu Asp Ala
Ser Cys Asp Thr Lys Arg His500 505 510tta
aac gac att ttt tgg gaa cca tac aac gac gaa gaa cca aaa tat 1584Leu
Asn Asp Ile Phe Trp Glu Pro Tyr Asn Asp Glu Glu Pro Lys Tyr515
520 525ttg gac atg gga aaa gaa cat ttt gaa atg aaa
aat att ttg gaa cta 1632Leu Asp Met Gly Lys Glu His Phe Glu Met Lys
Asn Ile Leu Glu Leu530 535 540aaa cgc atg
atg ctt tgg gat gaa gtt tat aga aat gcg aat ttg cgg 1680Lys Arg Met
Met Leu Trp Asp Glu Val Tyr Arg Asn Ala Asn Leu Arg545
550 555 560ttt aga gtc tgt aat gaa gaa
agt att aga 1710Phe Arg Val Cys Asn Glu Glu
Ser Ile Arg565 570341785DNACtenocephalides
felisexon(1)..(1785) 34atg tta cct cac agt gca tta gtt tta ttt tta ttt
ttt tta ttt ttc 48Met Leu Pro His Ser Ala Leu Val Leu Phe Leu Phe
Phe Leu Phe Phe1 5 10
15tta ttt aca cct gtc ttg tgc ata cta tgg gat aac cta gat cag cat
96Leu Phe Thr Pro Val Leu Cys Ile Leu Trp Asp Asn Leu Asp Gln His20
25 30ttg tgc aga gtt caa ttt aac ggg atc acg
gaa gga aaa ccg ttc cga 144Leu Cys Arg Val Gln Phe Asn Gly Ile Thr
Glu Gly Lys Pro Phe Arg35 40 45tat aaa
gat cat aaa aat gat gta tat tgt tcc tat ttg gga att cct 192Tyr Lys
Asp His Lys Asn Asp Val Tyr Cys Ser Tyr Leu Gly Ile Pro50
55 60tat gca gaa ccg cct att gga cca ttg cga ttt cag
tct cca aaa cca 240Tyr Ala Glu Pro Pro Ile Gly Pro Leu Arg Phe Gln
Ser Pro Lys Pro65 70 75
80ata tca aat cca aaa aca gga ttc gtt cag gct cgg tct tta gga gac
288Ile Ser Asn Pro Lys Thr Gly Phe Val Gln Ala Arg Ser Leu Gly Asp85
90 95aaa tgt ttc cag gaa agt cta ata tat tct
tat gca gga agc gaa gat 336Lys Cys Phe Gln Glu Ser Leu Ile Tyr Ser
Tyr Ala Gly Ser Glu Asp100 105 110tgc tta
tat ctg aat ata ttc acg cca gag act gtt aat tct gcg aac 384Cys Leu
Tyr Leu Asn Ile Phe Thr Pro Glu Thr Val Asn Ser Ala Asn115
120 125aat aca aaa tat cct gta atg ttc tgg atc cat gga
ggc gca ttc aac 432Asn Thr Lys Tyr Pro Val Met Phe Trp Ile His Gly
Gly Ala Phe Asn130 135 140caa gga tca gga
tct tat aat ttt ttt gga cct gat tat ttg atc agg 480Gln Gly Ser Gly
Ser Tyr Asn Phe Phe Gly Pro Asp Tyr Leu Ile Arg145 150
155 160gaa gga att att ttg gtc act atc aac
tat aga tta gga gtt ttc ggt 528Glu Gly Ile Ile Leu Val Thr Ile Asn
Tyr Arg Leu Gly Val Phe Gly165 170 175ttt
cta tca gcg ccg gaa tgg gat atc cat gga aat atg ggt cta aaa 576Phe
Leu Ser Ala Pro Glu Trp Asp Ile His Gly Asn Met Gly Leu Lys180
185 190gac cag aga ttg gca cta aaa tgg gtt tat gac
aac atc gaa aaa ttt 624Asp Gln Arg Leu Ala Leu Lys Trp Val Tyr Asp
Asn Ile Glu Lys Phe195 200 205ggt gga gac
aga gat aaa atc act ata gct gga gaa tct gct gga gca 672Gly Gly Asp
Arg Asp Lys Ile Thr Ile Ala Gly Glu Ser Ala Gly Ala210
215 220gca agt gtt cat ttt ctg atg atg gac aat tct act
aga aaa tac tac 720Ala Ser Val His Phe Leu Met Met Asp Asn Ser Thr
Arg Lys Tyr Tyr225 230 235
240caa agg gca att ttg cag agt ggg aca tta ctc aat ccg act gct aat
768Gln Arg Ala Ile Leu Gln Ser Gly Thr Leu Leu Asn Pro Thr Ala Asn245
250 255caa att caa cct ctg cat aga ttt gaa
aaa cta aaa caa gtg ctg aac 816Gln Ile Gln Pro Leu His Arg Phe Glu
Lys Leu Lys Gln Val Leu Asn260 265 270atc
acg caa aaa caa gaa ctc cta aat ctg gac aaa aat caa att ttg 864Ile
Thr Gln Lys Gln Glu Leu Leu Asn Leu Asp Lys Asn Gln Ile Leu275
280 285cga gca gcc tta aac aga gtc cca gat aac aac
gac cac gaa agg gac 912Arg Ala Ala Leu Asn Arg Val Pro Asp Asn Asn
Asp His Glu Arg Asp290 295 300aca gta cca
gta ttt aat cca gtc cta gaa tca cca gaa tct cca gac 960Thr Val Pro
Val Phe Asn Pro Val Leu Glu Ser Pro Glu Ser Pro Asp305
310 315 320cca ata aca ttt cca tct gct
tta gaa aga atg aga aat ggt gaa ttt 1008Pro Ile Thr Phe Pro Ser Ala
Leu Glu Arg Met Arg Asn Gly Glu Phe325 330
335cct gac gtt gat gtc atc att gga ttc aat agt gct gaa ggt tta aga
1056Pro Asp Val Asp Val Ile Ile Gly Phe Asn Ser Ala Glu Gly Leu Arg340
345 350tct atg cca aga gta acc aga gga aac
atg gaa gtt tac aag act ttg 1104Ser Met Pro Arg Val Thr Arg Gly Asn
Met Glu Val Tyr Lys Thr Leu355 360 365aca
aat ata gag aga gct ata cct aga gat gct aat att tgg aaa aat 1152Thr
Asn Ile Glu Arg Ala Ile Pro Arg Asp Ala Asn Ile Trp Lys Asn370
375 380cct aat ggc att gag gag aaa aaa ctt ata aaa
atg ctt aca gag ttt 1200Pro Asn Gly Ile Glu Glu Lys Lys Leu Ile Lys
Met Leu Thr Glu Phe385 390 395
400tat gac caa gtt aaa gaa caa aac gat gac atc gaa gcc tat gtc caa
1248Tyr Asp Gln Val Lys Glu Gln Asn Asp Asp Ile Glu Ala Tyr Val Gln405
410 415cta aaa ggc gat gct ggt tat ctc caa
gga att tac cgt acc ttg aaa 1296Leu Lys Gly Asp Ala Gly Tyr Leu Gln
Gly Ile Tyr Arg Thr Leu Lys420 425 430gcc
ata ttt ttc aat gaa atc aaa aga aat tcc aac ttg tat ttg tat 1344Ala
Ile Phe Phe Asn Glu Ile Lys Arg Asn Ser Asn Leu Tyr Leu Tyr435
440 445agg tta tca gat gat acg tat agt gta tat aaa
agt tat atc ttg ccc 1392Arg Leu Ser Asp Asp Thr Tyr Ser Val Tyr Lys
Ser Tyr Ile Leu Pro450 455 460tat cga tgg
ggt tcc ttg cca gga gtt agt cat ggt gat gat tta gga 1440Tyr Arg Trp
Gly Ser Leu Pro Gly Val Ser His Gly Asp Asp Leu Gly465
470 475 480tat ctt ttt gca aac tct ttg
gat gtt cct att ttg gga aca acg cac 1488Tyr Leu Phe Ala Asn Ser Leu
Asp Val Pro Ile Leu Gly Thr Thr His485 490
495att tct ata ccg caa gat gct atg cag act ctg gaa agg atg gtc agg
1536Ile Ser Ile Pro Gln Asp Ala Met Gln Thr Leu Glu Arg Met Val Arg500
505 510atc tgg acc aat ttt gta aag aat gga
aaa cct aca tca aac act gaa 1584Ile Trp Thr Asn Phe Val Lys Asn Gly
Lys Pro Thr Ser Asn Thr Glu515 520 525gat
gca tca tgt gat aca aaa aga cat tta aac gac att ttt tgg gaa 1632Asp
Ala Ser Cys Asp Thr Lys Arg His Leu Asn Asp Ile Phe Trp Glu530
535 540cca tac aac gac gaa gaa cca aaa tat ttg gac
atg gga aaa gaa cat 1680Pro Tyr Asn Asp Glu Glu Pro Lys Tyr Leu Asp
Met Gly Lys Glu His545 550 555
560ttt gaa atg aaa aat att ttg gaa cta aaa cgc atg atg ctt tgg gat
1728Phe Glu Met Lys Asn Ile Leu Glu Leu Lys Arg Met Met Leu Trp Asp565
570 575gaa gtt tat aga aat gcg aat ttg cgg
ttt aga gtc tgt aat gaa gaa 1776Glu Val Tyr Arg Asn Ala Asn Leu Arg
Phe Arg Val Cys Asn Glu Glu580 585 590agt
att aga 1785Ser
Ile Arg595351785DNACtenocephalides felis 35tctaatactt tcttcattac
agactctaaa ccgcaaattc gcatttctat aaacttcatc 60ccaaagcatc atgcgtttta
gttccaaaat atttttcatt tcaaaatgtt cttttcccat 120gtccaaatat tttggttctt
cgtcgttgta tggttcccaa aaaatgtcgt ttaaatgtct 180ttttgtatca catgatgcat
cttcagtgtt tgatgtaggt tttccattct ttacaaaatt 240ggtccagatc ctgaccatcc
tttccagagt ctgcatagca tcttgcggta tagaaatgtg 300cgttgttccc aaaataggaa
catccaaaga gtttgcaaaa agatatccta aatcatcacc 360atgactaact cctggcaagg
aaccccatcg atagggcaag atataacttt tatatacact 420atacgtatca tctgataacc
tatacaaata caagttggaa tttcttttga tttcattgaa 480aaatatggct ttcaaggtac
ggtaaattcc ttggagataa ccagcatcgc cttttagttg 540gacataggct tcgatgtcat
cgttttgttc tttaacttgg tcataaaact ctgtaagcat 600ttttataagt tttttctcct
caatgccatt aggatttttc caaatattag catctctagg 660tatagctctc tctatatttg
tcaaagtctt gtaaacttcc atgtttcctc tggttactct 720tggcatagat cttaaacctt
cagcactatt gaatccaatg atgacatcaa cgtcaggaaa 780ttcaccattt ctcattcttt
ctaaagcaga tggaaatgtt attgggtctg gagattctgg 840tgattctagg actggattaa
atactggtac tgtgtccctt tcgtggtcgt tgttatctgg 900gactctgttt aaggctgctc
gcaaaatttg atttttgtcc agatttagga gttcttgttt 960ttgcgtgatg ttcagcactt
gttttagttt ttcaaatcta tgcagaggtt gaatttgatt 1020agcagtcgga ttgagtaatg
tcccactctg caaaattgcc ctttggtagt attttctagt 1080agaattgtcc atcatcagaa
aatgaacact tgctgctcca gcagattctc cagctatagt 1140gattttatct ctgtctccac
caaatttttc gatgttgtca taaacccatt ttagtgccaa 1200tctctggtct tttagaccca
tatttccatg gatatcccat tccggcgctg atagaaaacc 1260gaaaactcct aatctatagt
tgatagtgac caaaataatt ccttccctga tcaaataatc 1320aggtccaaaa aaattataag
atcctgatcc ttggttgaat gcgcctccat ggatccagaa 1380cattacagga tattttgtat
tgttcgcaga attaacagtc tctggcgtga atatattcag 1440atataagcaa tcttcgcttc
ctgcataaga atatattaga ctttcctgga aacatttgtc 1500tcctaaagac cgagcctgaa
cgaatcctgt ttttggattt gatattggtt ttggagactg 1560aaatcgcaat ggtccaatag
gcggttctgc ataaggaatt cccaaatagg aacaatatac 1620atcattttta tgatctttat
atcggaacgg ttttccttcc gtgatcccgt taaattgaac 1680tctgcacaaa tgctgatcta
ggttatccca tagtatgcac aagacaggtg taaataagaa 1740aaataaaaaa aataaaaata
aaactaatgc actgtgaggt aacat
1785362007DNACtenocephalides felisCDS(11)..(1594) 36agttccaacg atg gct
gat cta caa gtg act ttg ctt caa ggt act tta 49Met Ala Asp Leu Gln
Val Thr Leu Leu Gln Gly Thr Leu1 5 10aaa
gga aaa gag caa att agt gaa aaa gga aat gtg ttc cat agt tat 97Lys
Gly Lys Glu Gln Ile Ser Glu Lys Gly Asn Val Phe His Ser Tyr15
20 25tct gga att cca tat gcc aaa cct cct gta ggt
gat cta aga ttt aag 145Ser Gly Ile Pro Tyr Ala Lys Pro Pro Val Gly
Asp Leu Arg Phe Lys30 35 40
45cca cct caa cct gca gaa cct tgg tca ggt gtt ctt gat gct agt aaa
193Pro Pro Gln Pro Ala Glu Pro Trp Ser Gly Val Leu Asp Ala Ser Lys50
55 60gaa ggg aat agt tgt aga tca gta cat
ttt att aaa aaa att aaa gta 241Glu Gly Asn Ser Cys Arg Ser Val His
Phe Ile Lys Lys Ile Lys Val65 70 75ggg
gct gaa gat tgt tta tac ctc aat gtc tat gta cca aaa aca tca 289Gly
Ala Glu Asp Cys Leu Tyr Leu Asn Val Tyr Val Pro Lys Thr Ser80
85 90gag aaa tca ctt ctt cca gta atg gta tgg ata
cat gga gga ggc ttc 337Glu Lys Ser Leu Leu Pro Val Met Val Trp Ile
His Gly Gly Gly Phe95 100 105ttc atg gga
tct gga aat agt gat atg tat ggt cct gaa tat ttg atg 385Phe Met Gly
Ser Gly Asn Ser Asp Met Tyr Gly Pro Glu Tyr Leu Met110
115 120 125gat tat gga att gtt ctg gtt
act ttc aat tat cga tta ggt gtt ttg 433Asp Tyr Gly Ile Val Leu Val
Thr Phe Asn Tyr Arg Leu Gly Val Leu130 135
140gga ttt ttg aac ctg gga ata gaa gaa gcg cct ggc aat gtt ggt ttg
481Gly Phe Leu Asn Leu Gly Ile Glu Glu Ala Pro Gly Asn Val Gly Leu145
150 155atg gac cag gtt gaa gct cta aaa tgg
gta aaa aac aat att gca tcc 529Met Asp Gln Val Glu Ala Leu Lys Trp
Val Lys Asn Asn Ile Ala Ser160 165 170ttt
ggt ggt gac ccc aac aat gtg act att ttt gga gaa tca gca ggt 577Phe
Gly Gly Asp Pro Asn Asn Val Thr Ile Phe Gly Glu Ser Ala Gly175
180 185ggt gca agt gtt cat tat ttg atg tta tca gat
ctt tcc aaa gga ctt 625Gly Ala Ser Val His Tyr Leu Met Leu Ser Asp
Leu Ser Lys Gly Leu190 195 200
205ttt cat aaa gcg atc tca caa agt gga agt gct ttt aat cct tgg gca
673Phe His Lys Ala Ile Ser Gln Ser Gly Ser Ala Phe Asn Pro Trp Ala210
215 220ctt caa cat gat aat aat aaa gaa aat
gca ttc cgc ctc tgc aaa ctt 721Leu Gln His Asp Asn Asn Lys Glu Asn
Ala Phe Arg Leu Cys Lys Leu225 230 235ctg
ggt cat cct gtc gat aac gag aca gaa gct cta aaa atc ctt cgt 769Leu
Gly His Pro Val Asp Asn Glu Thr Glu Ala Leu Lys Ile Leu Arg240
245 250caa gcc ccc ata gat gat ctt ata gac aac aga
ata aaa cca aaa gac 817Gln Ala Pro Ile Asp Asp Leu Ile Asp Asn Arg
Ile Lys Pro Lys Asp255 260 265aaa ggc caa
ctt att ata gac tat cct ttt cta cca aca ata gaa aaa 865Lys Gly Gln
Leu Ile Ile Asp Tyr Pro Phe Leu Pro Thr Ile Glu Lys270
275 280 285cgt tat caa aat ttt gaa cca
ttc ttg gac cag tct cca tta tca aaa 913Arg Tyr Gln Asn Phe Glu Pro
Phe Leu Asp Gln Ser Pro Leu Ser Lys290 295
300atg caa tca ggc aat ttc aca aaa gtc cca ttt ata tgt gga tac aac
961Met Gln Ser Gly Asn Phe Thr Lys Val Pro Phe Ile Cys Gly Tyr Asn305
310 315agt gct gaa gga att tta ggt tta atg
gac ttc aag gat gac cca aat 1009Ser Ala Glu Gly Ile Leu Gly Leu Met
Asp Phe Lys Asp Asp Pro Asn320 325 330ata
ttt gag aag ttt gaa gct gat ttt gaa aga ttt gta cca gta gat 1057Ile
Phe Glu Lys Phe Glu Ala Asp Phe Glu Arg Phe Val Pro Val Asp335
340 345ttg aat cta act tta agg tct aag gaa tct aaa
aaa ttg gct gaa gaa 1105Leu Asn Leu Thr Leu Arg Ser Lys Glu Ser Lys
Lys Leu Ala Glu Glu350 355 360
365atg aga aag ttt tat tac caa gac gaa cct gtt tct tca gac aac aaa
1153Met Arg Lys Phe Tyr Tyr Gln Asp Glu Pro Val Ser Ser Asp Asn Lys370
375 380gaa aaa ttt gtc agt gtt att agt gat
act tgg ttt ttg aga ggg att 1201Glu Lys Phe Val Ser Val Ile Ser Asp
Thr Trp Phe Leu Arg Gly Ile385 390 395aaa
aat act gca aga tat ata att gaa cat tcc tca gaa ccg tta tat 1249Lys
Asn Thr Ala Arg Tyr Ile Ile Glu His Ser Ser Glu Pro Leu Tyr400
405 410tta tat gtt tat agt ttt gat gat ttt ggt ttt
ttg aag aaa ctt gta 1297Leu Tyr Val Tyr Ser Phe Asp Asp Phe Gly Phe
Leu Lys Lys Leu Val415 420 425tta gat cct
aat att gaa gga gca gct cat gga gat gag ctg gga tat 1345Leu Asp Pro
Asn Ile Glu Gly Ala Ala His Gly Asp Glu Leu Gly Tyr430
435 440 445ctt ttc aag atg agt ttt aca
gaa ttt cca aaa gat tta cca agt gca 1393Leu Phe Lys Met Ser Phe Thr
Glu Phe Pro Lys Asp Leu Pro Ser Ala450 455
460gtg gtg aat agg gaa cga ttg ttg caa ctt tgg aca aat ttt gca aaa
1441Val Val Asn Arg Glu Arg Leu Leu Gln Leu Trp Thr Asn Phe Ala Lys465
470 475aca gga aat ccc act cct gaa atc aat
gat gtt ata aca aca aaa tgg 1489Thr Gly Asn Pro Thr Pro Glu Ile Asn
Asp Val Ile Thr Thr Lys Trp480 485 490gat
aaa gct act gag gaa aaa tca gat cat atg gat atc gat aat act 1537Asp
Lys Ala Thr Glu Glu Lys Ser Asp His Met Asp Ile Asp Asn Thr495
500 505ttg aga atg att cca gat cct gat gca aaa cga
ctt aga ttt tgg aat 1585Leu Arg Met Ile Pro Asp Pro Asp Ala Lys Arg
Leu Arg Phe Trp Asn510 515 520
525aaa ttt tta tgataaatat accaattatc gattttatta tagagtttct
1634Lys Phe Leugtattagtat aattatcacg tttagatgta cgagattcaa ttggctctaa
ttgaagtata 1694tttcgatttc aaatttactc tgattattgg aaaaaaagct tttacagttg
taataatcaa 1754gaagtaggtg gtaaatttag aacaaattct gttttagtga tttgcgcatt
caacagatgg 1814tgtactgtgc ctaaatttgt cgctcttctt gaagaactga actaaaaatg
tgattaatgg 1874acgccacatt atttatattt gatattatta ccatctttgt atcatatttg
cttttatttt 1934ttcatttttt ttttatttca aatatattgt ttttttataa aaaaaaaaaa
aaaaaaaaaa 1994aaaaaaaaaa aaa
200737528PRTCtenocephalides felis 37Met Ala Asp Leu Gln Val
Thr Leu Leu Gln Gly Thr Leu Lys Gly Lys1 5
10 15Glu Gln Ile Ser Glu Lys Gly Asn Val Phe His Ser
Tyr Ser Gly Ile20 25 30Pro Tyr Ala Lys
Pro Pro Val Gly Asp Leu Arg Phe Lys Pro Pro Gln35 40
45Pro Ala Glu Pro Trp Ser Gly Val Leu Asp Ala Ser Lys Glu
Gly Asn50 55 60Ser Cys Arg Ser Val His
Phe Ile Lys Lys Ile Lys Val Gly Ala Glu65 70
75 80Asp Cys Leu Tyr Leu Asn Val Tyr Val Pro Lys
Thr Ser Glu Lys Ser85 90 95Leu Leu Pro
Val Met Val Trp Ile His Gly Gly Gly Phe Phe Met Gly100
105 110Ser Gly Asn Ser Asp Met Tyr Gly Pro Glu Tyr Leu
Met Asp Tyr Gly115 120 125Ile Val Leu Val
Thr Phe Asn Tyr Arg Leu Gly Val Leu Gly Phe Leu130 135
140Asn Leu Gly Ile Glu Glu Ala Pro Gly Asn Val Gly Leu Met
Asp Gln145 150 155 160Val
Glu Ala Leu Lys Trp Val Lys Asn Asn Ile Ala Ser Phe Gly Gly165
170 175Asp Pro Asn Asn Val Thr Ile Phe Gly Glu Ser
Ala Gly Gly Ala Ser180 185 190Val His Tyr
Leu Met Leu Ser Asp Leu Ser Lys Gly Leu Phe His Lys195
200 205Ala Ile Ser Gln Ser Gly Ser Ala Phe Asn Pro Trp
Ala Leu Gln His210 215 220Asp Asn Asn Lys
Glu Asn Ala Phe Arg Leu Cys Lys Leu Leu Gly His225 230
235 240Pro Val Asp Asn Glu Thr Glu Ala Leu
Lys Ile Leu Arg Gln Ala Pro245 250 255Ile
Asp Asp Leu Ile Asp Asn Arg Ile Lys Pro Lys Asp Lys Gly Gln260
265 270Leu Ile Ile Asp Tyr Pro Phe Leu Pro Thr Ile
Glu Lys Arg Tyr Gln275 280 285Asn Phe Glu
Pro Phe Leu Asp Gln Ser Pro Leu Ser Lys Met Gln Ser290
295 300Gly Asn Phe Thr Lys Val Pro Phe Ile Cys Gly Tyr
Asn Ser Ala Glu305 310 315
320Gly Ile Leu Gly Leu Met Asp Phe Lys Asp Asp Pro Asn Ile Phe Glu325
330 335Lys Phe Glu Ala Asp Phe Glu Arg Phe
Val Pro Val Asp Leu Asn Leu340 345 350Thr
Leu Arg Ser Lys Glu Ser Lys Lys Leu Ala Glu Glu Met Arg Lys355
360 365Phe Tyr Tyr Gln Asp Glu Pro Val Ser Ser Asp
Asn Lys Glu Lys Phe370 375 380Val Ser Val
Ile Ser Asp Thr Trp Phe Leu Arg Gly Ile Lys Asn Thr385
390 395 400Ala Arg Tyr Ile Ile Glu His
Ser Ser Glu Pro Leu Tyr Leu Tyr Val405 410
415Tyr Ser Phe Asp Asp Phe Gly Phe Leu Lys Lys Leu Val Leu Asp Pro420
425 430Asn Ile Glu Gly Ala Ala His Gly Asp
Glu Leu Gly Tyr Leu Phe Lys435 440 445Met
Ser Phe Thr Glu Phe Pro Lys Asp Leu Pro Ser Ala Val Val Asn450
455 460Arg Glu Arg Leu Leu Gln Leu Trp Thr Asn Phe
Ala Lys Thr Gly Asn465 470 475
480Pro Thr Pro Glu Ile Asn Asp Val Ile Thr Thr Lys Trp Asp Lys
Ala485 490 495Thr Glu Glu Lys Ser Asp His
Met Asp Ile Asp Asn Thr Leu Arg Met500 505
510Ile Pro Asp Pro Asp Ala Lys Arg Leu Arg Phe Trp Asn Lys Phe Leu515
520 525382007DNACtenocephalides felis
38tttttttttt tttttttttt tttttttttt tttttataaa aaaacaatat atttgaaata
60aaaaaaaaat gaaaaaataa aagcaaatat gatacaaaga tggtaataat atcaaatata
120aataatgtgg cgtccattaa tcacattttt agttcagttc ttcaagaaga gcgacaaatt
180taggcacagt acaccatctg ttgaatgcgc aaatcactaa aacagaattt gttctaaatt
240taccacctac ttcttgatta ttacaactgt aaaagctttt tttccaataa tcagagtaaa
300tttgaaatcg aaatatactt caattagagc caattgaatc tcgtacatct aaacgtgata
360attatactaa tacagaaact ctataataaa atcgataatt ggtatattta tcataaaaat
420ttattccaaa atctaagtcg ttttgcatca ggatctggaa tcattctcaa agtattatcg
480atatccatat gatctgattt ttcctcagta gctttatccc attttgttgt tataacatca
540ttgatttcag gagtgggatt tcctgttttt gcaaaatttg tccaaagttg caacaatcgt
600tccctattca ccactgcact tggtaaatct tttggaaatt ctgtaaaact catcttgaaa
660agatatccca gctcatctcc atgagctgct ccttcaatat taggatctaa tacaagtttc
720ttcaaaaaac caaaatcatc aaaactataa acatataaat ataacggttc tgaggaatgt
780tcaattatat atcttgcagt atttttaatc cctctcaaaa accaagtatc actaataaca
840ctgacaaatt tttctttgtt gtctgaagaa acaggttcgt cttggtaata aaactttctc
900atttcttcag ccaatttttt agattcctta gaccttaaag ttagattcaa atctactggt
960acaaatcttt caaaatcagc ttcaaacttc tcaaatatat ttgggtcatc cttgaagtcc
1020attaaaccta aaattccttc agcactgttg tatccacata taaatgggac ttttgtgaaa
1080ttgcctgatt gcatttttga taatggagac tggtccaaga atggttcaaa attttgataa
1140cgtttttcta ttgttggtag aaaaggatag tctataataa gttggccttt gtcttttggt
1200tttattctgt tgtctataag atcatctatg ggggcttgac gaaggatttt tagagcttct
1260gtctcgttat cgacaggatg acccagaagt ttgcagaggc ggaatgcatt ttctttatta
1320ttatcatgtt gaagtgccca aggattaaaa gcacttccac tttgtgagat cgctttatga
1380aaaagtcctt tggaaagatc tgataacatc aaataatgaa cacttgcacc acctgctgat
1440tctccaaaaa tagtcacatt gttggggtca ccaccaaagg atgcaatatt gttttttacc
1500cattttagag cttcaacctg gtccatcaaa ccaacattgc caggcgcttc ttctattccc
1560aggttcaaaa atcccaaaac acctaatcga taattgaaag taaccagaac aattccataa
1620tccatcaaat attcaggacc atacatatca ctatttccag atcccatgaa gaagcctcct
1680ccatgtatcc ataccattac tggaagaagt gatttctctg atgtttttgg tacatagaca
1740ttgaggtata aacaatcttc agcccctact ttaatttttt taataaaatg tactgatcta
1800caactattcc cttctttact agcatcaaga acacctgacc aaggttctgc aggttgaggt
1860ggcttaaatc ttagatcacc tacaggaggt ttggcatatg gaattccaga ataactatgg
1920aacacatttc ctttttcact aatttgctct tttcctttta aagtaccttg aagcaaagtc
1980acttgtagat cagccatcgt tggaact
20073912PRTCtenocephalides felis 39Asp Pro Pro Thr Val Thr Leu Pro Gln
Gly Glu Leu1 5 104022PRTCtenocephalides
felisMISC_FEATURE(21)..(21)Xaa = unknown 40Asp Pro Pro Thr Val Thr Leu
Pro Gln Gly Glu Leu Val Gly Lys Ala1 5 10
15Thr Asn Glu Asn Xaa Lys204112PRTCtenocephalides felis
41Asp Pro Pro Thr Val Thr Leu Pro Gln Gly Glu Leu1 5
104221PRTCtenocephalides felis 42Asp Pro Pro Thr Val Thr Leu
Pro Gln Gly Glu Leu Val Gly Lys Ala1 5 10
15Leu Ser Asn Glu Asn20438PRTCtenocephalides felis 43Asp
Pro Pro Thr Val Thr Leu Pro1 54423PRTCtenocephalides felis
44Asp Pro Pro Thr Val Thr Leu Pro Gln Gly Glu Leu Val Gly Lys Ala1
5 10 15Leu Thr Asn Glu Asn Gly
Lys204520DNAArtificial sequenceSynthetic Primer 45aattaaccct cactaaaggg
204617DNAArtificial
sequenceSynthetic Primer 46ardccdccdc crtrdat
174738DNAArtificial sequenceSynthetic Primer
47tgtgctcgag atgggataac ctagatcagc atttgtgc
384835DNAArtificial sequenceSynthetic Primer 48ttaaggtacc tcatctaata
cttccttcat tacag 354936DNAArtificial
sequenceSynthetic Primer 49aaaactgcag tataaatatg ttacctcaca gtagtg
365034DNAArtificial sequenceSynthetic Primer
50tgctctagat tatctaatac ttccttcatt acag
34511584DNACtenocephalides felisexon(1)..(1584) 51atg gct gat cta caa gtg
act ttg ctt caa ggt act tta aaa gga aaa 48Met Ala Asp Leu Gln Val
Thr Leu Leu Gln Gly Thr Leu Lys Gly Lys1 5
10 15gag caa att agt gaa aaa gga aat gtg ttc cat agt
tat tct gga att 96Glu Gln Ile Ser Glu Lys Gly Asn Val Phe His Ser
Tyr Ser Gly Ile20 25 30cca tat gcc aaa
cct cct gta ggt gat cta aga ttt aag cca cct caa 144Pro Tyr Ala Lys
Pro Pro Val Gly Asp Leu Arg Phe Lys Pro Pro Gln35 40
45cct gca gaa cct tgg tca ggt gtt ctt gat gct agt aaa gaa
ggg aat 192Pro Ala Glu Pro Trp Ser Gly Val Leu Asp Ala Ser Lys Glu
Gly Asn50 55 60agt tgt aga tca gta cat
ttt att aaa aaa att aaa gta ggg gct gaa 240Ser Cys Arg Ser Val His
Phe Ile Lys Lys Ile Lys Val Gly Ala Glu65 70
75 80gat tgt tta tac ctc aat gtc tat gta cca aaa
aca tca gag aaa tca 288Asp Cys Leu Tyr Leu Asn Val Tyr Val Pro Lys
Thr Ser Glu Lys Ser85 90 95ctt ctt cca
gta atg gta tgg ata cat gga gga ggc ttc ttc atg gga 336Leu Leu Pro
Val Met Val Trp Ile His Gly Gly Gly Phe Phe Met Gly100
105 110tct gga aat agt gat atg tat ggt cct gaa tat ttg
atg gat tat gga 384Ser Gly Asn Ser Asp Met Tyr Gly Pro Glu Tyr Leu
Met Asp Tyr Gly115 120 125att gtt ctg gtt
act ttc aat tat cga tta ggt gtt ttg gga ttt ttg 432Ile Val Leu Val
Thr Phe Asn Tyr Arg Leu Gly Val Leu Gly Phe Leu130 135
140aac ctg gga ata gaa gaa gcg cct ggc aat gtt ggt ttg atg
gac cag 480Asn Leu Gly Ile Glu Glu Ala Pro Gly Asn Val Gly Leu Met
Asp Gln145 150 155 160gtt
gaa gct cta aaa tgg gta aaa aac aat att gca tcc ttt ggt ggt 528Val
Glu Ala Leu Lys Trp Val Lys Asn Asn Ile Ala Ser Phe Gly Gly165
170 175gac ccc aac aat gtg act att ttt gga gaa tca
gca ggt ggt gca agt 576Asp Pro Asn Asn Val Thr Ile Phe Gly Glu Ser
Ala Gly Gly Ala Ser180 185 190gtt cat tat
ttg atg tta tca gat ctt tcc aaa gga ctt ttt cat aaa 624Val His Tyr
Leu Met Leu Ser Asp Leu Ser Lys Gly Leu Phe His Lys195
200 205gcg atc tca caa agt gga agt gct ttt aat cct tgg
gca ctt caa cat 672Ala Ile Ser Gln Ser Gly Ser Ala Phe Asn Pro Trp
Ala Leu Gln His210 215 220gat aat aat aaa
gaa aat gca ttc cgc ctc tgc aaa ctt ctg ggt cat 720Asp Asn Asn Lys
Glu Asn Ala Phe Arg Leu Cys Lys Leu Leu Gly His225 230
235 240cct gtc gat aac gag aca gaa gct cta
aaa atc ctt cgt caa gcc ccc 768Pro Val Asp Asn Glu Thr Glu Ala Leu
Lys Ile Leu Arg Gln Ala Pro245 250 255ata
gat gat ctt ata gac aac aga ata aaa cca aaa gac aaa ggc caa 816Ile
Asp Asp Leu Ile Asp Asn Arg Ile Lys Pro Lys Asp Lys Gly Gln260
265 270ctt att ata gac tat cct ttt cta cca aca ata
gaa aaa cgt tat caa 864Leu Ile Ile Asp Tyr Pro Phe Leu Pro Thr Ile
Glu Lys Arg Tyr Gln275 280 285aat ttt gaa
cca ttc ttg gac cag tct cca tta tca aaa atg caa tca 912Asn Phe Glu
Pro Phe Leu Asp Gln Ser Pro Leu Ser Lys Met Gln Ser290
295 300ggc aat ttc aca aaa gtc cca ttt ata tgt gga tac
aac agt gct gaa 960Gly Asn Phe Thr Lys Val Pro Phe Ile Cys Gly Tyr
Asn Ser Ala Glu305 310 315
320gga att tta ggt tta atg gac ttc aag gat gac cca aat ata ttt gag
1008Gly Ile Leu Gly Leu Met Asp Phe Lys Asp Asp Pro Asn Ile Phe Glu325
330 335aag ttt gaa gct gat ttt gaa aga ttt
gta cca gta gat ttg aat cta 1056Lys Phe Glu Ala Asp Phe Glu Arg Phe
Val Pro Val Asp Leu Asn Leu340 345 350act
tta agg tct aag gaa tct aaa aaa ttg gct gaa gaa atg aga aag 1104Thr
Leu Arg Ser Lys Glu Ser Lys Lys Leu Ala Glu Glu Met Arg Lys355
360 365ttt tat tac caa gac gaa cct gtt tct tca gac
aac aaa gaa aaa ttt 1152Phe Tyr Tyr Gln Asp Glu Pro Val Ser Ser Asp
Asn Lys Glu Lys Phe370 375 380gtc agt gtt
att agt gat act tgg ttt ttg aga ggg att aaa aat act 1200Val Ser Val
Ile Ser Asp Thr Trp Phe Leu Arg Gly Ile Lys Asn Thr385
390 395 400gca aga tat ata att gaa cat
tcc tca gaa ccg tta tat tta tat gtt 1248Ala Arg Tyr Ile Ile Glu His
Ser Ser Glu Pro Leu Tyr Leu Tyr Val405 410
415tat agt ttt gat gat ttt ggt ttt ttg aag aaa ctt gta tta gat cct
1296Tyr Ser Phe Asp Asp Phe Gly Phe Leu Lys Lys Leu Val Leu Asp Pro420
425 430aat att gaa gga gca gct cat gga gat
gag ctg gga tat ctt ttc aag 1344Asn Ile Glu Gly Ala Ala His Gly Asp
Glu Leu Gly Tyr Leu Phe Lys435 440 445atg
agt ttt aca gaa ttt cca aaa gat tta cca agt gca gtg gtg aat 1392Met
Ser Phe Thr Glu Phe Pro Lys Asp Leu Pro Ser Ala Val Val Asn450
455 460agg gaa cga ttg ttg caa ctt tgg aca aat ttt
gca aaa aca gga aat 1440Arg Glu Arg Leu Leu Gln Leu Trp Thr Asn Phe
Ala Lys Thr Gly Asn465 470 475
480ccc act cct gaa atc aat gat gtt ata aca aca aaa tgg gat aaa gct
1488Pro Thr Pro Glu Ile Asn Asp Val Ile Thr Thr Lys Trp Asp Lys Ala485
490 495act gag gaa aaa tca gat cat atg gat
atc gat aat act ttg aga atg 1536Thr Glu Glu Lys Ser Asp His Met Asp
Ile Asp Asn Thr Leu Arg Met500 505 510att
cca gat cct gat gca aaa cga ctt aga ttt tgg aat aaa ttt tta 1584Ile
Pro Asp Pro Asp Ala Lys Arg Leu Arg Phe Trp Asn Lys Phe Leu515
520 525521584DNACtenocephalides felis 52taaaaattta
ttccaaaatc taagtcgttt tgcatcagga tctggaatca ttctcaaagt 60attatcgata
tccatatgat ctgatttttc ctcagtagct ttatcccatt ttgttgttat 120aacatcattg
atttcaggag tgggatttcc tgtttttgca aaatttgtcc aaagttgcaa 180caatcgttcc
ctattcacca ctgcacttgg taaatctttt ggaaattctg taaaactcat 240cttgaaaaga
tatcccagct catctccatg agctgctcct tcaatattag gatctaatac 300aagtttcttc
aaaaaaccaa aatcatcaaa actataaaca tataaatata acggttctga 360ggaatgttca
attatatatc ttgcagtatt tttaatccct ctcaaaaacc aagtatcact 420aataacactg
acaaattttt ctttgttgtc tgaagaaaca ggttcgtctt ggtaataaaa 480ctttctcatt
tcttcagcca attttttaga ttccttagac cttaaagtta gattcaaatc 540tactggtaca
aatctttcaa aatcagcttc aaacttctca aatatatttg ggtcatcctt 600gaagtccatt
aaacctaaaa ttccttcagc actgttgtat ccacatataa atgggacttt 660tgtgaaattg
cctgattgca tttttgataa tggagactgg tccaagaatg gttcaaaatt 720ttgataacgt
ttttctattg ttggtagaaa aggatagtct ataataagtt ggcctttgtc 780ttttggtttt
attctgttgt ctataagatc atctatgggg gcttgacgaa ggatttttag 840agcttctgtc
tcgttatcga caggatgacc cagaagtttg cagaggcgga atgcattttc 900tttattatta
tcatgttgaa gtgcccaagg attaaaagca cttccacttt gtgagatcgc 960tttatgaaaa
agtcctttgg aaagatctga taacatcaaa taatgaacac ttgcaccacc 1020tgctgattct
ccaaaaatag tcacattgtt ggggtcacca ccaaaggatg caatattgtt 1080ttttacccat
tttagagctt caacctggtc catcaaacca acattgccag gcgcttcttc 1140tattcccagg
ttcaaaaatc ccaaaacacc taatcgataa ttgaaagtaa ccagaacaat 1200tccataatcc
atcaaatatt caggaccata catatcacta tttccagatc ccatgaagaa 1260gcctcctcca
tgtatccata ccattactgg aagaagtgat ttctctgatg tttttggtac 1320atagacattg
aggtataaac aatcttcagc ccctacttta atttttttaa taaaatgtac 1380tgatctacaa
ctattccctt ctttactagc atcaagaaca cctgaccaag gttctgcagg 1440ttgaggtggc
ttaaatctta gatcacctac aggaggtttg gcatatggaa ttccagaata 1500actatggaac
acatttcctt tttcactaat ttgctctttt ccttttaaag taccttgaag 1560caaagtcact
tgtagatcag ccat
158453530PRTCtenocephalides felis 53Asp Pro Pro Thr Val Thr Leu Pro Gln
Gly Glu Leu Val Gly Lys Ala1 5 10
15Leu Thr Asn Glu Asn Gly Lys Glu Tyr Phe Ser Tyr Thr Gly Val
Pro20 25 30Tyr Ala Lys Pro Pro Val Gly
Glu Leu Arg Phe Lys Pro Pro Gln Lys35 40
45Ala Glu Pro Trp Asn Gly Val Phe Asn Ala Thr Ser His Gly Asn Val50
55 60Cys Lys Ala Leu Asn Phe Phe Leu Lys Lys
Ile Glu Gly Asp Glu Asp65 70 75
80Cys Leu Leu Val Asn Val Tyr Ala Pro Lys Thr Thr Ser Asp Lys
Lys85 90 95Leu Pro Val Phe Phe Trp Val
His Gly Gly Gly Phe Val Thr Gly Ser100 105
110Gly Asn Leu Glu Phe Gln Ser Pro Asp Tyr Leu Val Asn Tyr Asp Val115
120 125Ile Phe Val Thr Phe Asn Tyr Arg Leu
Gly Pro Leu Gly Phe Leu Asn130 135 140Leu
Glu Leu Glu Gly Ala Pro Gly Asn Val Gly Leu Leu Asp Gln Val145
150 155 160Ala Ala Leu Lys Trp Thr
Lys Glu Asn Ile Glu Lys Phe Gly Gly Asp165 170
175Pro Glu Asn Ile Thr Ile Gly Gly Val Ser Ala Gly Gly Ala Ser
Val180 185 190His Tyr Leu Leu Leu Ser His
Thr Thr Thr Gly Leu Tyr Lys Arg Ala195 200
205Ile Ala Gln Ser Gly Ser Ala Leu Asn Pro Trp Ala Phe Gln Arg His210
215 220Pro Val Lys Arg Ser Leu Gln Leu Ala
Glu Ile Leu Gly His Pro Thr225 230 235
240Asn Asn Thr Gln Asp Ala Leu Glu Phe Leu Gln Lys Ala Pro
Val Asp245 250 255Ser Leu Leu Lys Lys Met
Pro Ala Glu Thr Glu Gly Glu Ile Ile Glu260 265
270Glu Phe Val Phe Val Pro Ser Ile Glu Lys Val Phe Pro Ser His
Gln275 280 285Pro Phe Leu Glu Glu Ser Pro
Leu Ala Arg Met Lys Ser Gly Ser Phe290 295
300Asn Lys Val Pro Leu Leu Val Gly Phe Asn Ser Ala Glu Gly Leu Leu305
310 315 320Tyr Lys Phe Phe
Met Lys Glu Lys Pro Glu Met Leu Asn Gln Ala Glu325 330
335Ala Asp Phe Glu Arg Leu Val Pro Ala Glu Phe Glu Leu Ala
His Gly340 345 350Ser Glu Glu Ser Lys Lys
Leu Ala Glu Lys Ile Arg Lys Phe Tyr Phe355 360
365Asp Asp Lys Pro Val Pro Glu Asn Glu Gln Lys Phe Ile Asp Leu
Ile370 375 380Gly Asp Ile Trp Phe Thr Arg
Gly Ile Asp Lys His Val Lys Leu Ser385 390
395 400Val Glu Lys Gln Asp Glu Pro Val Tyr Tyr Tyr Glu
Tyr Ser Phe Ser405 410 415Glu Ser His Pro
Ala Lys Gly Thr Phe Gly Asp His Asn Leu Thr Gly420 425
430Ala Cys His Gly Glu Glu Leu Val Asn Leu Phe Lys Val Glu
Met Met435 440 445Lys Leu Glu Lys Asp Lys
Pro Asn Val Leu Leu Thr Lys Asp Arg Val450 455
460Leu Ala Met Trp Thr Asn Phe Ile Lys Asn Gly Asn Pro Thr Pro
Glu465 470 475 480Val Thr
Glu Leu Leu Pro Val Lys Trp Glu Pro Ala Thr Lys Asp Lys485
490 495Leu Asn Tyr Leu Asn Ile Asp Ala Thr Leu Thr Leu
Gly Thr Asn Pro500 505 510Glu Glu Thr Arg
Val Lys Phe Trp Glu Asp Ala Thr Lys Thr Leu His515 520
525Ser Gln53054570PRTCtenocephalides felis 54Trp Asp Asn Leu
Asp Gln His Leu Cys Arg Val Gln Phe Asn Gly Ile1 5
10 15Thr Glu Gly Lys Pro Phe Arg Tyr Lys Asp
His Arg Asn Asp Val Tyr20 25 30Cys Ser
Tyr Leu Gly Ile Pro Tyr Ala Glu Pro Pro Phe Gly Pro Leu35
40 45Arg Phe Gln Ser Pro Lys Pro Ile Ser Asn Pro Lys
Thr Gly Phe Val50 55 60Gln Ala Arg Thr
Leu Gly Asp Lys Cys Phe Gln Glu Ser Leu Ile Tyr65 70
75 80Ser Tyr Ala Gly Ser Glu Asp Cys Leu
Tyr Leu Asn Ile Phe Thr Pro85 90 95Glu
Thr Val Asn Ser Ala Asn Asn Thr Lys Tyr Pro Val Met Phe Trp100
105 110Ile His Gly Gly Ala Phe Asn Gln Gly Ser Gly
Ser Tyr Asn Phe Phe115 120 125Gly Pro Asp
Tyr Leu Ile Arg Glu Gly Ile Ile Leu Val Thr Ile Asn130
135 140Tyr Arg Leu Gly Val Phe Gly Phe Leu Ser Ala Pro
Glu Trp Asp Ile145 150 155
160His Gly Asn Met Gly Leu Lys Asp Gln Arg Leu Ala Leu Lys Trp Val165
170 175Tyr Asp Asn Ile Glu Lys Phe Gly Gly
Asp Arg Glu Lys Ile Thr Ile180 185 190Ala
Gly Glu Ser Ala Gly Ala Ala Ser Val His Phe Leu Met Met Asp195
200 205Asn Ser Thr Arg Lys Tyr Tyr Gln Arg Ala Ile
Leu Gln Ser Gly Thr210 215 220Leu Leu Asn
Pro Thr Ala Asn Gln Ile Gln Leu Leu His Arg Phe Glu225
230 235 240Lys Leu Lys Gln Val Leu Asn
Ile Thr Gln Lys Gln Glu Leu Leu Asn245 250
255Leu Asp Lys Asn Leu Ile Leu Arg Ala Ala Leu Asn Arg Val Pro Asp260
265 270Ser Asn Asp His Asp Arg Asp Thr Val
Pro Val Phe Asn Pro Val Leu275 280 285Glu
Ser Pro Glu Ser Pro Asp Pro Ile Thr Phe Pro Ser Ala Leu Glu290
295 300Arg Met Arg Asn Gly Glu Phe Pro Asp Val Asp
Val Ile Ile Gly Phe305 310 315
320Asn Ser Ala Glu Gly Leu Arg Ser Met Ala Arg Val Thr Arg Gly
Asn325 330 335Met Glu Val His Lys Thr Leu
Thr Asn Ile Glu Arg Ala Ile Pro Arg340 345
350Asp Ala Asn Ile Trp Lys Asn Pro Asn Gly Ile Glu Glu Lys Lys Leu355
360 365Ile Lys Met Leu Thr Glu Phe Tyr Asp
Gln Val Lys Glu Gln Asn Asp370 375 380Asp
Ile Glu Ala Tyr Val Gln Leu Lys Gly Asp Ala Gly Tyr Leu Gln385
390 395 400Gly Ile Tyr Arg Thr Leu
Lys Ala Ile Phe Phe Asn Glu Phe Arg Arg405 410
415Asn Ser Asn Leu Tyr Leu Tyr Arg Leu Ser Asp Asp Thr Tyr Ser
Val420 425 430Tyr Lys Ser Tyr Ile Leu Pro
Tyr Arg Trp Gly Ser Leu Pro Gly Val435 440
445Ser His Gly Asp Asp Leu Gly Tyr Leu Phe Ala Asn Ser Leu Asp Val450
455 460Pro Ile Leu Gly Thr Thr His Ile Ser
Ile Pro Gln Asp Ala Met Gln465 470 475
480Thr Leu Glu Arg Met Val Arg Ile Trp Thr Asn Phe Val Lys
Asn Gly485 490 495Lys Pro Thr Ser Asn Thr
Glu Asp Ala Ser Cys Asp Thr Lys Arg His500 505
510Leu Asn Asp Ile Phe Trp Glu Pro Tyr Asn Asp Glu Glu Pro Lys
Tyr515 520 525Leu Asp Met Gly Lys Glu Asn
Phe Glu Met Lys Asn Ile Leu Glu Leu530 535
540Lys Arg Met Met Leu Trp Asp Glu Val Tyr Arg Asn Ala Asn Leu Arg545
550 555 560Phe Arg Val Cys
Asn Glu Gly Ser Ile Arg565 57055570PRTCtenocephalides
felis 55Trp Asp Asn Leu Asp Gln His Leu Cys Arg Val Gln Phe Asn Gly Ile1
5 10 15Thr Glu Gly Lys
Pro Phe Arg Tyr Lys Asp His Lys Asn Asp Val Tyr20 25
30Cys Ser Tyr Leu Gly Ile Pro Tyr Ala Glu Pro Pro Ile Gly
Pro Leu35 40 45Arg Phe Gln Ser Pro Lys
Pro Ile Ser Asn Pro Lys Thr Gly Phe Val50 55
60Gln Ala Arg Ser Leu Gly Asp Lys Cys Phe Gln Glu Ser Leu Ile Tyr65
70 75 80Ser Tyr Ala Gly
Ser Glu Asp Cys Leu Tyr Leu Asn Ile Phe Thr Pro85 90
95Glu Thr Val Asn Ser Ala Asn Asn Thr Lys Tyr Pro Val Met
Phe Trp100 105 110Ile His Gly Gly Ala Phe
Asn Gln Gly Ser Gly Ser Tyr Asn Phe Phe115 120
125Gly Pro Asp Tyr Leu Ile Arg Glu Gly Ile Ile Leu Val Thr Ile
Asn130 135 140Tyr Arg Leu Gly Val Phe Gly
Phe Leu Ser Ala Pro Glu Trp Asp Ile145 150
155 160His Gly Asn Met Gly Leu Lys Asp Gln Arg Leu Ala
Leu Lys Trp Val165 170 175Tyr Asp Asn Ile
Glu Lys Phe Gly Gly Asp Arg Asp Lys Ile Thr Ile180 185
190Ala Gly Glu Ser Ala Gly Ala Ala Ser Val His Phe Leu Met
Met Asp195 200 205Asn Ser Thr Arg Lys Tyr
Tyr Gln Arg Ala Ile Leu Gln Ser Gly Thr210 215
220Leu Leu Asn Pro Thr Ala Asn Gln Ile Gln Pro Leu His Arg Phe
Glu225 230 235 240Lys Leu
Lys Gln Val Leu Asn Ile Thr Gln Lys Gln Glu Leu Leu Asn245
250 255Leu Asp Lys Asn Gln Ile Leu Arg Ala Ala Leu Asn
Arg Val Pro Asp260 265 270Asn Asn Asp His
Glu Arg Asp Thr Val Pro Val Phe Asn Pro Val Leu275 280
285Glu Ser Pro Glu Ser Pro Asp Pro Ile Thr Phe Pro Ser Ala
Leu Glu290 295 300Arg Met Arg Asn Gly Glu
Phe Pro Asp Val Asp Val Ile Ile Gly Phe305 310
315 320Asn Ser Ala Glu Gly Leu Arg Ser Met Pro Arg
Val Thr Arg Gly Asn325 330 335Met Glu Val
Tyr Lys Thr Leu Thr Asn Ile Glu Arg Ala Ile Pro Arg340
345 350Asp Ala Asn Ile Trp Lys Asn Pro Asn Gly Ile Glu
Glu Lys Lys Leu355 360 365Ile Lys Met Leu
Thr Glu Phe Tyr Asp Gln Val Lys Glu Gln Asn Asp370 375
380Asp Ile Glu Ala Tyr Val Gln Leu Lys Gly Asp Ala Gly Tyr
Leu Gln385 390 395 400Gly
Ile Tyr Arg Thr Leu Lys Ala Ile Phe Phe Asn Glu Ile Lys Arg405
410 415Asn Ser Asn Leu Tyr Leu Tyr Arg Leu Ser Asp
Asp Thr Tyr Ser Val420 425 430Tyr Lys Ser
Tyr Ile Leu Pro Tyr Arg Trp Gly Ser Leu Pro Gly Val435
440 445Ser His Gly Asp Asp Leu Gly Tyr Leu Phe Ala Asn
Ser Leu Asp Val450 455 460Pro Ile Leu Gly
Thr Thr His Ile Ser Ile Pro Gln Asp Ala Met Gln465 470
475 480Thr Leu Glu Arg Met Val Arg Ile Trp
Thr Asn Phe Val Lys Asn Gly485 490 495Lys
Pro Thr Ser Asn Thr Glu Asp Ala Ser Cys Asp Thr Lys Arg His500
505 510Leu Asn Asp Ile Phe Trp Glu Pro Tyr Asn Asp
Glu Glu Pro Lys Tyr515 520 525Leu Asp Met
Gly Lys Glu His Phe Glu Met Lys Asn Ile Leu Glu Leu530
535 540Lys Arg Met Met Leu Trp Asp Glu Val Tyr Arg Asn
Ala Asn Leu Arg545 550 555
560Phe Arg Val Cys Asn Glu Gly Ser Ile Arg565
5705620DNAArtificial sequenceSynthetic Primer 56gtgcgtacac gtttactacc
20572144DNACtenocephalides
felisCDS(30)..(1682)misc_feature(462)..(462)At nucleotide 462, r = a or g
At amino acid residue 145, Xaa = Asn or Asp 57gtacacatag tcaatagtct
agatccaag atg tct cgt gtt att ttt tta agt 53Met Ser Arg Val Ile Phe
Leu Ser1 5ytgt att ttt ttg ttt agt ttt aat ttt ata aaa tgt
gat tcc ccg act 101Cys Ile Phe Leu Phe Ser Phe Asn Phe Ile Lys Cys
Asp Ser Pro Thr10 15 20ygta act ttg ccc
caa ggc gaa ttg gtt gga aaa gct ttg acg aac gaa 149Val Thr Leu Pro
Gln Gly Glu Leu Val Gly Lys Ala Leu Thr Asn Glu25 30
35 40yaat gga aaa gag tat ttt agc tac aca
ggt gta cct tat gct aaa cct 197Asn Gly Lys Glu Tyr Phe Ser Tyr Thr
Gly Val Pro Tyr Ala Lys Pro45 50 55ycct
gtt gga gaa ctt aga ttt aag cct cca cag aaa gct gag cca tgg 245Pro
Val Gly Glu Leu Arg Phe Lys Pro Pro Gln Lys Ala Glu Pro Trp60
65 70ycaa ggt gtt ttc aac gcc aca tta tac gga aat
gtg tgt aaa tct tta 293Gln Gly Val Phe Asn Ala Thr Leu Tyr Gly Asn
Val Cys Lys Ser Leu75 80 85yaat ttc ttc
ttg aag aaa att gaa gga gac gaa gac tgc ttg gta gta 341Asn Phe Phe
Leu Lys Lys Ile Glu Gly Asp Glu Asp Cys Leu Val Val90 95
100yaac gtg tac gca cca aaa aca act tct gat aaa aaa ctt
cca gta ttt 389Asn Val Tyr Ala Pro Lys Thr Thr Ser Asp Lys Lys Leu
Pro Val Phe105 110 115
120yttc tgg gtt cat ggt ggt ggt ttt gtg act gga tcc gga aat tta gaa
437Phe Trp Val His Gly Gly Gly Phe Val Thr Gly Ser Gly Asn Leu Glu125
130 135yttc caa agc cca gat tat tta gta rat
ttt gat gtt att ttc gta act 485Phe Gln Ser Pro Asp Tyr Leu Val Xaa
Phe Asp Val Ile Phe Val Thr140 145
150yttc aat tac cga ttg gga cct ctc gga ttt ctg aat ttg gag ttg gag
533Phe Asn Tyr Arg Leu Gly Pro Leu Gly Phe Leu Asn Leu Glu Leu Glu155
160 165yggt gct cca gga aat gta gga tta ttg
gat cag gtg gca gct ctg aaa 581Gly Ala Pro Gly Asn Val Gly Leu Leu
Asp Gln Val Ala Ala Leu Lys170 175
180ytgg acc aaa gaa aac att gag aaa ttt ggt gga gat cca gaa aat att
629Trp Thr Lys Glu Asn Ile Glu Lys Phe Gly Gly Asp Pro Glu Asn Ile185
190 195 200yaca att ggt ggt
gtt tct gct ggt gga gca agt gtt cat tat ctt ttg 677Thr Ile Gly Gly
Val Ser Ala Gly Gly Ala Ser Val His Tyr Leu Leu205 210
215ytta tct cat aca acc act gga ctt tac aaa agg gca att gct
caa agt 725Leu Ser His Thr Thr Thr Gly Leu Tyr Lys Arg Ala Ile Ala
Gln Ser220 225 230ygga agt gct ttt aat
cca tgg gcc ttc caa aga cat cca gta aag cgt 773Gly Ser Ala Phe Asn
Pro Trp Ala Phe Gln Arg His Pro Val Lys Arg235 240
245yagt ctt caa ctt gct gag ata ttg ggt cat ccc aca aac aat act
caa 821Ser Leu Gln Leu Ala Glu Ile Leu Gly His Pro Thr Asn Asn Thr
Gln250 255 260ygat gct tta gaa ttc tta
caa aaa gcc ccc gta gac agt ctc ctg aag 869Asp Ala Leu Glu Phe Leu
Gln Lys Ala Pro Val Asp Ser Leu Leu Lys265 270
275 280yaaa atg cca gct gaa aca gaa ggt gaa ata ata
gaa gag ttt gtc ttc 917Lys Met Pro Ala Glu Thr Glu Gly Glu Ile Ile
Glu Glu Phe Val Phe285 290 295ygta cca
tca att gaa aaa gtt ttc cca tcc cac caa cct ttc ttg gaa 965Val Pro
Ser Ile Glu Lys Val Phe Pro Ser His Gln Pro Phe Leu Glu300
305 310gaa tca cca ttg gcc aga atg aaa tcc gga tcc ttt
aac aaa gta cct 1013Glu Ser Pro Leu Ala Arg Met Lys Ser Gly Ser Phe
Asn Lys Val Pro315 320 325tta tta gtt gga
ttt aac agt gca gaa gga ctt ttg ttc aaa ttc ttc 1061Leu Leu Val Gly
Phe Asn Ser Ala Glu Gly Leu Leu Phe Lys Phe Phe330 335
340atg aaa gaa aaa cca gag atg ctg aac caa gct gaa gca gat
ttt gaa 1109Met Lys Glu Lys Pro Glu Met Leu Asn Gln Ala Glu Ala Asp
Phe Glu345 350 355 360aga
ctc gta cca gcc gaa ttt gaa tta gtc cat gga tca gag gaa tcg 1157Arg
Leu Val Pro Ala Glu Phe Glu Leu Val His Gly Ser Glu Glu Ser365
370 375aaa aaa ctt gca gaa aaa atc agg aag ttt tac
ttt gac gat aaa ccc 1205Lys Lys Leu Ala Glu Lys Ile Arg Lys Phe Tyr
Phe Asp Asp Lys Pro380 385 390gtt cca gaa
aat gaa cag aaa ttt att gac ttg ata gga gat att tgg 1253Val Pro Glu
Asn Glu Gln Lys Phe Ile Asp Leu Ile Gly Asp Ile Trp395
400 405ttt act aga ggt gtt gac aag cat gtc aag ttg tct
gtg gag aaa caa 1301Phe Thr Arg Gly Val Asp Lys His Val Lys Leu Ser
Val Glu Lys Gln410 415 420gac gaa cca gtt
tat tat tat gaa tat tcc ttc tcg gaa agt cat cct 1349Asp Glu Pro Val
Tyr Tyr Tyr Glu Tyr Ser Phe Ser Glu Ser His Pro425 430
435 440gca aaa gga aca ttt ggt gat cat aat
ctg act ggt gca tgc cat gga 1397Ala Lys Gly Thr Phe Gly Asp His Asn
Leu Thr Gly Ala Cys His Gly445 450 455gaa
gaa ctt gtg aat tta ttc aaa gtc gag atg atg aag ctg gaa aaa 1445Glu
Glu Leu Val Asn Leu Phe Lys Val Glu Met Met Lys Leu Glu Lys460
465 470gat aaa cct aat gtt cta tta aca aaa gat aga
gta ctt gcc atg tgg 1493Asp Lys Pro Asn Val Leu Leu Thr Lys Asp Arg
Val Leu Ala Met Trp475 480 485act aac ttc
atc aaa aat gga aat cct act cct gaa gta aca gaa tta 1541Thr Asn Phe
Ile Lys Asn Gly Asn Pro Thr Pro Glu Val Thr Glu Leu490
495 500ttg cca gtt aaa tgg gaa cct gcc aca aaa gac aag
ttg aat tat ttg 1589Leu Pro Val Lys Trp Glu Pro Ala Thr Lys Asp Lys
Leu Asn Tyr Leu505 510 515
520aac att gat gcc acc tta act ttg gga aca aat cct gag gca aac cga
1637Asn Ile Asp Ala Thr Leu Thr Leu Gly Thr Asn Pro Glu Ala Asn Arg525
530 535gtc aaa ttt tgg gaa gac gcc aca aaa
tct ttg cac ggt caa taa 1682Val Lys Phe Trp Glu Asp Ala Thr Lys
Ser Leu His Gly Gln540 545 550taatttatga
aaattgtttt aaatacttta ggtaatatat taggtaaata aaaattaaaa 1742aataacaatt
tttatgtttt atgtattggc ttatgtgtat cagttctaat tttatttatt 1802tattcttgtt
ttgcttgttt tgaaatatca tggttttaat tttcaaaaca caacgtcgtt 1862tgtttttagc
aaaatttcca atagatatgt tatattaagt actctgaagt atttttatat 1922atacactaaa
atcagtaaaa atacattaac taaaaatata agatattttc aataattttt 1982tttaaagaaa
ataccaaaaa taaagtaaaa ttccaaacgg aatttttgtt taacttaaaa 2042ataaaattaa
ctcttcaata attttgataa ttagtatttc tgatatcatt agtgaaaatt 2102atattttgat
aatacgtatt tatatttaaa ataaaattat gt
214458550PRTCtenocephalides felismisc_feature(145)..(145)The 'Xaa' at
location 145 stands for Asp, or Asn. 58Met Ser Arg Val Ile Phe Leu
Ser Cys Ile Phe Leu Phe Ser Phe Asn1 5 10
15Phe Ile Lys Cys Asp Ser Pro Thr Val Thr Leu Pro Gln
Gly Glu Leu20 25 30Val Gly Lys Ala Leu
Thr Asn Glu Asn Gly Lys Glu Tyr Phe Ser Tyr35 40
45Thr Gly Val Pro Tyr Ala Lys Pro Pro Val Gly Glu Leu Arg Phe
Lys50 55 60Pro Pro Gln Lys Ala Glu Pro
Trp Gln Gly Val Phe Asn Ala Thr Leu65 70
75 80Tyr Gly Asn Val Cys Lys Ser Leu Asn Phe Phe Leu
Lys Lys Ile Glu85 90 95Gly Asp Glu Asp
Cys Leu Val Val Asn Val Tyr Ala Pro Lys Thr Thr100 105
110Ser Asp Lys Lys Leu Pro Val Phe Phe Trp Val His Gly Gly
Gly Phe115 120 125Val Thr Gly Ser Gly Asn
Leu Glu Phe Gln Ser Pro Asp Tyr Leu Val130 135
140Xaa Phe Asp Val Ile Phe Val Thr Phe Asn Tyr Arg Leu Gly Pro
Leu145 150 155 160Gly Phe
Leu Asn Leu Glu Leu Glu Gly Ala Pro Gly Asn Val Gly Leu165
170 175Leu Asp Gln Val Ala Ala Leu Lys Trp Thr Lys Glu
Asn Ile Glu Lys180 185 190Phe Gly Gly Asp
Pro Glu Asn Ile Thr Ile Gly Gly Val Ser Ala Gly195 200
205Gly Ala Ser Val His Tyr Leu Leu Leu Ser His Thr Thr Thr
Gly Leu210 215 220Tyr Lys Arg Ala Ile Ala
Gln Ser Gly Ser Ala Phe Asn Pro Trp Ala225 230
235 240Phe Gln Arg His Pro Val Lys Arg Ser Leu Gln
Leu Ala Glu Ile Leu245 250 255Gly His Pro
Thr Asn Asn Thr Gln Asp Ala Leu Glu Phe Leu Gln Lys260
265 270Ala Pro Val Asp Ser Leu Leu Lys Lys Met Pro Ala
Glu Thr Glu Gly275 280 285Glu Ile Ile Glu
Glu Phe Val Phe Val Pro Ser Ile Glu Lys Val Phe290 295
300Pro Ser His Gln Pro Phe Leu Glu Glu Ser Pro Leu Ala Arg
Met Lys305 310 315 320Ser
Gly Ser Phe Asn Lys Val Pro Leu Leu Val Gly Phe Asn Ser Ala325
330 335Glu Gly Leu Leu Phe Lys Phe Phe Met Lys Glu
Lys Pro Glu Met Leu340 345 350Asn Gln Ala
Glu Ala Asp Phe Glu Arg Leu Val Pro Ala Glu Phe Glu355
360 365Leu Val His Gly Ser Glu Glu Ser Lys Lys Leu Ala
Glu Lys Ile Arg370 375 380Lys Phe Tyr Phe
Asp Asp Lys Pro Val Pro Glu Asn Glu Gln Lys Phe385 390
395 400Ile Asp Leu Ile Gly Asp Ile Trp Phe
Thr Arg Gly Val Asp Lys His405 410 415Val
Lys Leu Ser Val Glu Lys Gln Asp Glu Pro Val Tyr Tyr Tyr Glu420
425 430Tyr Ser Phe Ser Glu Ser His Pro Ala Lys Gly
Thr Phe Gly Asp His435 440 445Asn Leu Thr
Gly Ala Cys His Gly Glu Glu Leu Val Asn Leu Phe Lys450
455 460Val Glu Met Met Lys Leu Glu Lys Asp Lys Pro Asn
Val Leu Leu Thr465 470 475
480Lys Asp Arg Val Leu Ala Met Trp Thr Asn Phe Ile Lys Asn Gly Asn485
490 495Pro Thr Pro Glu Val Thr Glu Leu Leu
Pro Val Lys Trp Glu Pro Ala500 505 510Thr
Lys Asp Lys Leu Asn Tyr Leu Asn Ile Asp Ala Thr Leu Thr Leu515
520 525Gly Thr Asn Pro Glu Ala Asn Arg Val Lys Phe
Trp Glu Asp Ala Thr530 535 540Lys Ser Leu
His Gly Gln545 550592144DNACtenocephalides felis
59acataatttt attttaaata taaatacgta ttatcaaaat ataattttca ctaatgatat
60cagaaatact aattatcaaa attattgaag agttaatttt atttttaagt taaacaaaaa
120ttccgtttgg aattttactt tatttttggt attttcttta aaaaaaatta ttgaaaatat
180cttatatttt tagttaatgt atttttactg attttagtgt atatataaaa atacttcaga
240gtacttaata taacatatct attggaaatt ttgctaaaaa caaacgacgt tgtgttttga
300aaattaaaac catgatattt caaaacaagc aaaacaagaa taaataaata aaattagaac
360tgatacacat aagccaatac ataaaacata aaaattgtta ttttttaatt tttatttacc
420taatatatta cctaaagtat ttaaaacaat tttcataaat tattattgac cgtgcaaaga
480ttttgtggcg tcttcccaaa atttgactcg gtttgcctca ggatttgttc ccaaagttaa
540ggtggcatca atgttcaaat aattcaactt gtcttttgtg gcaggttccc atttaactgg
600caataattct gttacttcag gagtaggatt tccatttttg atgaagttag tccacatggc
660aagtactcta tcttttgtta atagaacatt aggtttatct ttttccagct tcatcatctc
720gactttgaat aaattcacaa gttcttctcc atggcatgca ccagtcagat tatgatcacc
780aaatgttcct tttgcaggat gactttccga gaaggaatat tcataataat aaactggttc
840gtcttgtttc tccacagaca acttgacatg cttgtcaaca cctctagtaa accaaatatc
900tcctatcaag tcaataaatt tctgttcatt ttctggaacg ggtttatcgt caaagtaaaa
960cttcctgatt ttttctgcaa gttttttcga ttcctctgat ccatggacta attcaaattc
1020ggctggtacg agtctttcaa aatctgcttc agcttggttc agcatctctg gtttttcttt
1080catgaagaat ttgaacaaaa gtccttctgc actgttaaat ccaactaata aaggtacttt
1140gttaaaggat ccggatttca ttctggccaa tggtgattct tccaagaaag gttggtggga
1200tgggaaaact ttttcaattg atggtacgaa gacaaactct tctattattt caccttctgt
1260ttcagctggc attttcttca ggagactgtc tacgggggct ttttgtaaga attctaaagc
1320atcttgagta ttgtttgtgg gatgacccaa tatctcagca agttgaagac tacgctttac
1380tggatgtctt tggaaggccc atggattaaa agcacttcca ctttgagcaa ttgccctttt
1440gtaaagtcca gtggttgtat gagataacaa aagataatga acacttgctc caccagcaga
1500aacaccacca attgtaatat tttctggatc tccaccaaat ttctcaatgt tttctttggt
1560ccatttcaga gctgccacct gatccaataa tcctacattt cctggagcac cctccaactc
1620caaattcaga aatccgagag gtcccaatcg gtaattgaaa gttacgaaaa taacatcaaa
1680atytactaaa taatctgggc tttggaattc taaatttccg gatccagtca caaaaccacc
1740accatgaacc cagaaaaata ctggaagttt tttatcagaa gttgtttttg gtgcgtacac
1800gtttactacc aagcagtctt cgtctccttc aattttcttc aagaagaaat ttaaagattt
1860acacacattt ccgtataatg tggcgttgaa aacaccttgc catggctcag ctttctgtgg
1920aggcttaaat ctaagttctc caacaggagg tttagcataa ggtacacctg tgtagctaaa
1980atactctttt ccattttcgt tcgtcaaagc ttttccaacc aattcgcctt ggggcaaagt
2040tacagtcggg gaatcacatt ttataaaatt aaaactaaac aaaaaaatac aacttaaaaa
2100aataacacga gacatcttgg atctagacta ttgactatgt gtac
2144601650DNACtenocephalides
felisexon(1)..(1650)misc_feature(433)..(433)At nucleotide 433, r = a or g
At amino acid residue 145, Xaa = Asn or Asp 60atg tct cgt gtt att
ttt tta agt tgt att ttt ttg ttt agt ttt aat 48Met Ser Arg Val Ile
Phe Leu Ser Cys Ile Phe Leu Phe Ser Phe Asn1 5
10 15ttt ata aaa tgt gat tcc ccg act gta act ttg
ccc caa ggc gaa ttg 96Phe Ile Lys Cys Asp Ser Pro Thr Val Thr Leu
Pro Gln Gly Glu Leu20 25 30gtt gga aaa
gct ttg acg aac gaa aat gga aaa gag tat ttt agc tac 144Val Gly Lys
Ala Leu Thr Asn Glu Asn Gly Lys Glu Tyr Phe Ser Tyr35 40
45aca ggt gta cct tat gct aaa cct cct gtt gga gaa ctt
aga ttt aag 192Thr Gly Val Pro Tyr Ala Lys Pro Pro Val Gly Glu Leu
Arg Phe Lys50 55 60cct cca cag aaa gct
gag cca tgg caa ggt gtt ttc aac gcc aca tta 240Pro Pro Gln Lys Ala
Glu Pro Trp Gln Gly Val Phe Asn Ala Thr Leu65 70
75 80tac gga aat gtg tgt aaa tct tta aat ttc
ttc ttg aag aaa att gaa 288Tyr Gly Asn Val Cys Lys Ser Leu Asn Phe
Phe Leu Lys Lys Ile Glu85 90 95gga gac
gaa gac tgc ttg gta gta aac gtg tac gca cca aaa aca act 336Gly Asp
Glu Asp Cys Leu Val Val Asn Val Tyr Ala Pro Lys Thr Thr100
105 110tct gat aaa aaa ctt cca gta ttt ttc tgg gtt cat
ggt ggt ggt ttt 384Ser Asp Lys Lys Leu Pro Val Phe Phe Trp Val His
Gly Gly Gly Phe115 120 125gtg act gga tcc
gga aat tta gaa ttc caa agc cca gat tat tta gta 432Val Thr Gly Ser
Gly Asn Leu Glu Phe Gln Ser Pro Asp Tyr Leu Val130 135
140rat ttt gat gtt att ttc gta act ttc aat tac cga ttg gga
cct ctc 480Xaa Phe Asp Val Ile Phe Val Thr Phe Asn Tyr Arg Leu Gly
Pro Leu145 150 155 160gga
ttt ctg aat ttg gag ttg gag ggt gct cca gga aat gta gga tta 528Gly
Phe Leu Asn Leu Glu Leu Glu Gly Ala Pro Gly Asn Val Gly Leu165
170 175ttg gat cag gtg gca gct ctg aaa tgg acc aaa
gaa aac att gag aaa 576Leu Asp Gln Val Ala Ala Leu Lys Trp Thr Lys
Glu Asn Ile Glu Lys180 185 190ttt ggt gga
gat cca gaa aat att aca att ggt ggt gtt tct gct ggt 624Phe Gly Gly
Asp Pro Glu Asn Ile Thr Ile Gly Gly Val Ser Ala Gly195
200 205gga gca agt gtt cat tat ctt ttg tta tct cat aca
acc act gga ctt 672Gly Ala Ser Val His Tyr Leu Leu Leu Ser His Thr
Thr Thr Gly Leu210 215 220tac aaa agg gca
att gct caa agt gga agt gct ttt aat cca tgg gcc 720Tyr Lys Arg Ala
Ile Ala Gln Ser Gly Ser Ala Phe Asn Pro Trp Ala225 230
235 240ttc caa aga cat cca gta aag cgt agt
ctt caa ctt gct gag ata ttg 768Phe Gln Arg His Pro Val Lys Arg Ser
Leu Gln Leu Ala Glu Ile Leu245 250 255ggt
cat ccc aca aac aat act caa gat gct tta gaa ttc tta caa aaa 816Gly
His Pro Thr Asn Asn Thr Gln Asp Ala Leu Glu Phe Leu Gln Lys260
265 270gcc ccc gta gac agt ctc ctg aag aaa atg cca
gct gaa aca gaa ggt 864Ala Pro Val Asp Ser Leu Leu Lys Lys Met Pro
Ala Glu Thr Glu Gly275 280 285gaa ata ata
gaa gag ttt gtc ttc gta cca tca att gaa aaa gtt ttc 912Glu Ile Ile
Glu Glu Phe Val Phe Val Pro Ser Ile Glu Lys Val Phe290
295 300cca tcc cac caa cct ttc ttg gaa gaa tca cca ttg
gcc aga atg aaa 960Pro Ser His Gln Pro Phe Leu Glu Glu Ser Pro Leu
Ala Arg Met Lys305 310 315
320tcc gga tcc ttt aac aaa gta cct tta tta gtt gga ttt aac agt gca
1008Ser Gly Ser Phe Asn Lys Val Pro Leu Leu Val Gly Phe Asn Ser Ala325
330 335gaa gga ctt ttg ttc aaa ttc ttc atg
aaa gaa aaa cca gag atg ctg 1056Glu Gly Leu Leu Phe Lys Phe Phe Met
Lys Glu Lys Pro Glu Met Leu340 345 350aac
caa gct gaa gca gat ttt gaa aga ctc gta cca gcc gaa ttt gaa 1104Asn
Gln Ala Glu Ala Asp Phe Glu Arg Leu Val Pro Ala Glu Phe Glu355
360 365tta gtc cat gga tca gag gaa tcg aaa aaa ctt
gca gaa aaa atc agg 1152Leu Val His Gly Ser Glu Glu Ser Lys Lys Leu
Ala Glu Lys Ile Arg370 375 380aag ttt tac
ttt gac gat aaa ccc gtt cca gaa aat gaa cag aaa ttt 1200Lys Phe Tyr
Phe Asp Asp Lys Pro Val Pro Glu Asn Glu Gln Lys Phe385
390 395 400att gac ttg ata gga gat att
tgg ttt act aga ggt gtt gac aag cat 1248Ile Asp Leu Ile Gly Asp Ile
Trp Phe Thr Arg Gly Val Asp Lys His405 410
415gtc aag ttg tct gtg gag aaa caa gac gaa cca gtt tat tat tat gaa
1296Val Lys Leu Ser Val Glu Lys Gln Asp Glu Pro Val Tyr Tyr Tyr Glu420
425 430tat tcc ttc tcg gaa agt cat cct gca
aaa gga aca ttt ggt gat cat 1344Tyr Ser Phe Ser Glu Ser His Pro Ala
Lys Gly Thr Phe Gly Asp His435 440 445aat
ctg act ggt gca tgc cat gga gaa gaa ctt gtg aat tta ttc aaa 1392Asn
Leu Thr Gly Ala Cys His Gly Glu Glu Leu Val Asn Leu Phe Lys450
455 460gtc gag atg atg aag ctg gaa aaa gat aaa cct
aat gtt cta tta aca 1440Val Glu Met Met Lys Leu Glu Lys Asp Lys Pro
Asn Val Leu Leu Thr465 470 475
480aaa gat aga gta ctt gcc atg tgg act aac ttc atc aaa aat gga aat
1488Lys Asp Arg Val Leu Ala Met Trp Thr Asn Phe Ile Lys Asn Gly Asn485
490 495cct act cct gaa gta aca gaa tta ttg
cca gtt aaa tgg gaa cct gcc 1536Pro Thr Pro Glu Val Thr Glu Leu Leu
Pro Val Lys Trp Glu Pro Ala500 505 510aca
aaa gac aag ttg aat tat ttg aac att gat gcc acc tta act ttg 1584Thr
Lys Asp Lys Leu Asn Tyr Leu Asn Ile Asp Ala Thr Leu Thr Leu515
520 525gga aca aat cct gag gca aac cga gtc aaa ttt
tgg gaa gac gcc aca 1632Gly Thr Asn Pro Glu Ala Asn Arg Val Lys Phe
Trp Glu Asp Ala Thr530 535 540aaa tct ttg
cac ggt caa 1650Lys Ser Leu
His Gly Gln545 550611650DNACtenocephalides felis
61ttgaccgtgc aaagattttg tggcgtcttc ccaaaatttg actcggtttg cctcaggatt
60tgttcccaaa gttaaggtgg catcaatgtt caaataattc aacttgtctt ttgtggcagg
120ttcccattta actggcaata attctgttac ttcaggagta ggatttccat ttttgatgaa
180gttagtccac atggcaagta ctctatcttt tgttaataga acattaggtt tatctttttc
240cagcttcatc atctcgactt tgaataaatt cacaagttct tctccatggc atgcaccagt
300cagattatga tcaccaaatg ttccttttgc aggatgactt tccgagaagg aatattcata
360ataataaact ggttcgtctt gtttctccac agacaacttg acatgcttgt caacacctct
420agtaaaccaa atatctccta tcaagtcaat aaatttctgt tcattttctg gaacgggttt
480atcgtcaaag taaaacttcc tgattttttc tgcaagtttt ttcgattcct ctgatccatg
540gactaattca aattcggctg gtacgagtct ttcaaaatct gcttcagctt ggttcagcat
600ctctggtttt tctttcatga agaatttgaa caaaagtcct tctgcactgt taaatccaac
660taataaaggt actttgttaa aggatccgga tttcattctg gccaatggtg attcttccaa
720gaaaggttgg tgggatggga aaactttttc aattgatggt acgaagacaa actcttctat
780tatttcacct tctgtttcag ctggcatttt cttcaggaga ctgtctacgg gggctttttg
840taagaattct aaagcatctt gagtattgtt tgtgggatga cccaatatct cagcaagttg
900aagactacgc tttactggat gtctttggaa ggcccatgga ttaaaagcac ttccactttg
960agcaattgcc cttttgtaaa gtccagtggt tgtatgagat aacaaaagat aatgaacact
1020tgctccacca gcagaaacac caccaattgt aatattttct ggatctccac caaatttctc
1080aatgttttct ttggtccatt tcagagctgc cacctgatcc aataatccta catttcctgg
1140agcaccctcc aactccaaat tcagaaatcc gagaggtccc aatcggtaat tgaaagttac
1200gaaaataaca tcaaaatyta ctaaataatc tgggctttgg aattctaaat ttccggatcc
1260agtcacaaaa ccaccaccat gaacccagaa aaatactgga agttttttat cagaagttgt
1320ttttggtgcg tacacgttta ctaccaagca gtcttcgtct ccttcaattt tcttcaagaa
1380gaaatttaaa gatttacaca catttccgta taatgtggcg ttgaaaacac cttgccatgg
1440ctcagctttc tgtggaggct taaatctaag ttctccaaca ggaggtttag cataaggtac
1500acctgtgtag ctaaaatact cttttccatt ttcgttcgtc aaagcttttc caaccaattc
1560gccttggggc aaagttacag tcggggaatc acattttata aaattaaaac taaacaaaaa
1620aatacaactt aaaaaaataa cacgagacat
16506229DNAArtificial sequenceSynthetic Primer 62aaactcgagt cccccgactg
taactttgc 296336DNAArtificial
sequenceSynthetic Primer 63tcatctgcag ttattgactg tgcaaagttt ttgtgg
366432DNAArtificial sequenceSynthetic Primer
64ttccggatcc ggctgatcta caagtgactt tg
326534DNAArtificial sequenceSynthetic Primer 65tggtactcga gtcataaaaa
tttattccaa aatc 346639DNAArtificial
sequenceSynthetic Primer 66aaaactgcag tataaatatg ttacctcaca gtgcattag
39671987DNACtenocephalides felisCDS(231)..(1820)
67aattcacagt gtaaataatt ttatttgata taaatgtatt taatttttat tttaatctaa
60ttttaattta aatatatata gttttattta taaaaaaata ttttttttat gatcgaaaag
120aaatttttat ttatgtttat gagtgtgtgt tttggctatg atttacatta tttttgagct
180agtataaaat taaaccatat tatattttgg atatataata acattttata atg tgt
236Met Cys1gat cca tta cta aaa aca aca aca tat gga att ctg aaa ggc aag
aaa 284Asp Pro Leu Leu Lys Thr Thr Thr Tyr Gly Ile Leu Lys Gly Lys
Lys5 10 15gtt gta aac gaa aat ggt aaa
att tac tat agt tac aca ggt ata ccc 332Val Val Asn Glu Asn Gly Lys
Ile Tyr Tyr Ser Tyr Thr Gly Ile Pro20 25
30tat gca aaa tct cct gta aat gat ctc aga ttc aag cca cca caa aaa
380Tyr Ala Lys Ser Pro Val Asn Asp Leu Arg Phe Lys Pro Pro Gln Lys35
40 45 50ctt gat cct tgg aat
ggt gtt ttt gac gcc act cag tat gga aat aat 428Leu Asp Pro Trp Asn
Gly Val Phe Asp Ala Thr Gln Tyr Gly Asn Asn55 60
65tgt gct gct ggg aaa tgg ttt ttg aaa tca gct ggg ggt tgc gaa
gat 476Cys Ala Ala Gly Lys Trp Phe Leu Lys Ser Ala Gly Gly Cys Glu
Asp70 75 80tgc ctt tac tta aat atc tat
gtc cca caa aac act tca gaa aat cct 524Cys Leu Tyr Leu Asn Ile Tyr
Val Pro Gln Asn Thr Ser Glu Asn Pro85 90
95ttg cca gta atg ttt tgg att cat gga gga gca ttt gtg gtc gga tca
572Leu Pro Val Met Phe Trp Ile His Gly Gly Ala Phe Val Val Gly Ser100
105 110gga aat tct gat ata cat ggt cct gat
tat tta ata gaa tat gat att 620Gly Asn Ser Asp Ile His Gly Pro Asp
Tyr Leu Ile Glu Tyr Asp Ile115 120 125
130atc tta gta act att aat tat cgt cta gga cca ctt ggt ttt
ctt aat 668Ile Leu Val Thr Ile Asn Tyr Arg Leu Gly Pro Leu Gly Phe
Leu Asn135 140 145ttg gaa atc gaa gat gcg
cct ggg aat gtt gga ttg atg gat caa gtt 716Leu Glu Ile Glu Asp Ala
Pro Gly Asn Val Gly Leu Met Asp Gln Val150 155
160gca gcc cta aaa tgg gta aat gaa aat att gca acc ttt agt gga gac
764Ala Ala Leu Lys Trp Val Asn Glu Asn Ile Ala Thr Phe Ser Gly Asp165
170 175cca aaa aat att aca att tgt gga gca
act gct gga gct gca agt gta 812Pro Lys Asn Ile Thr Ile Cys Gly Ala
Thr Ala Gly Ala Ala Ser Val180 185 190cat
tat cac att ttg tca caa ctt acc aaa ggt tta ttc cac aag gct 860His
Tyr His Ile Leu Ser Gln Leu Thr Lys Gly Leu Phe His Lys Ala195
200 205 210ata gca caa agt gga agt
gct ttt aat ccc tgg gct ttc caa aaa aat 908Ile Ala Gln Ser Gly Ser
Ala Phe Asn Pro Trp Ala Phe Gln Lys Asn215 220
225cct gtt aag aat gca ctt cga cta tgc aaa acc tta ggc ctt acc aca
956Pro Val Lys Asn Ala Leu Arg Leu Cys Lys Thr Leu Gly Leu Thr Thr230
235 240aac aac ctt caa gaa gcc ttg gat ttt
ttg aaa aac cta cca gta gaa 1004Asn Asn Leu Gln Glu Ala Leu Asp Phe
Leu Lys Asn Leu Pro Val Glu245 250 255aca
ttg tta aat acc aaa tta ccc caa gaa att gat ggt caa ctg ctg 1052Thr
Leu Leu Asn Thr Lys Leu Pro Gln Glu Ile Asp Gly Gln Leu Leu260
265 270gat gac ttc gtg ttt gta cct tcg att gaa aaa
aca ttt cca gaa caa 1100Asp Asp Phe Val Phe Val Pro Ser Ile Glu Lys
Thr Phe Pro Glu Gln275 280 285
290gat tcg tac tta act gac ttg cca ata cca ata ata aat tca gga aaa
1148Asp Ser Tyr Leu Thr Asp Leu Pro Ile Pro Ile Ile Asn Ser Gly Lys295
300 305ttc cac aaa gtt cca ttg ttg aca ggt
tac aac agt gcc gaa ggc aat 1196Phe His Lys Val Pro Leu Leu Thr Gly
Tyr Asn Ser Ala Glu Gly Asn310 315 320cta
ttt ttc atg tac tta aaa aca gat cca gat tta tta aat aaa ttt 1244Leu
Phe Phe Met Tyr Leu Lys Thr Asp Pro Asp Leu Leu Asn Lys Phe325
330 335gaa gct gat ttt gaa aga ttt ata cca act gac
tta gaa tta cct ttg 1292Glu Ala Asp Phe Glu Arg Phe Ile Pro Thr Asp
Leu Glu Leu Pro Leu340 345 350cga tca caa
aaa tct att gca ctg ggt gaa gca atc agg gaa ttt tat 1340Arg Ser Gln
Lys Ser Ile Ala Leu Gly Glu Ala Ile Arg Glu Phe Tyr355
360 365 370ttc caa aac aaa acc ata tca
gaa aat atg cag aat ttt gta gat gtt 1388Phe Gln Asn Lys Thr Ile Ser
Glu Asn Met Gln Asn Phe Val Asp Val375 380
385tta agt gat aat tgg ttt aca cgt gga att gat gag caa gta aag tta
1436Leu Ser Asp Asn Trp Phe Thr Arg Gly Ile Asp Glu Gln Val Lys Leu390
395 400act gtt aaa aat cag gaa gaa cca gtt
ttt tat tat gtt tat aat ttt 1484Thr Val Lys Asn Gln Glu Glu Pro Val
Phe Tyr Tyr Val Tyr Asn Phe405 410 415gat
gaa aat tct cca agt cgg aaa gtt ttt ggt gat ttt gga ata aaa 1532Asp
Glu Asn Ser Pro Ser Arg Lys Val Phe Gly Asp Phe Gly Ile Lys420
425 430ggc ggt ggt cat gct gat gaa ttg ggt aat ata
ttt aaa gcc aaa agt 1580Gly Gly Gly His Ala Asp Glu Leu Gly Asn Ile
Phe Lys Ala Lys Ser435 440 445
450gca aat ttt ggg aag gaa aca cca aat gct gtg ttg gtt cag aga agg
1628Ala Asn Phe Gly Lys Glu Thr Pro Asn Ala Val Leu Val Gln Arg Arg455
460 465atg ctg gag atg tgg act aat ttt gct
aaa ttt gga aat cct act cca 1676Met Leu Glu Met Trp Thr Asn Phe Ala
Lys Phe Gly Asn Pro Thr Pro470 475 480gct
att acg gat aca ctt cca ata aaa tgg gaa cct gct ttt aaa gaa 1724Ala
Ile Thr Asp Thr Leu Pro Ile Lys Trp Glu Pro Ala Phe Lys Glu485
490 495aat atg act ttt gtt caa att gac att gat tta
aat ttg agt act gat 1772Asn Met Thr Phe Val Gln Ile Asp Ile Asp Leu
Asn Leu Ser Thr Asp500 505 510cca cta aaa
agt cgt atg gaa ttt ggg aat aaa ata aaa tta tta aaa 1820Pro Leu Lys
Ser Arg Met Glu Phe Gly Asn Lys Ile Lys Leu Leu Lys515
520 525 530taagtaacta tacttagcta
aaccataata taccaaataa tagtatagga atacttcaca 1880attttttgtt acttcgttaa
gtaaatttaa ttttttataa aaccaacttt tacgaataaa 1940aaatgtaatt attttggaaa
aaaaaaagaa aaaaaaaaaa aaaaaac 198768530PRTCtenocephalides
felis 68Met Cys Asp Pro Leu Leu Lys Thr Thr Thr Tyr Gly Ile Leu Lys Gly1
5 10 15Lys Lys Val Val
Asn Glu Asn Gly Lys Ile Tyr Tyr Ser Tyr Thr Gly20 25
30Ile Pro Tyr Ala Lys Ser Pro Val Asn Asp Leu Arg Phe Lys
Pro Pro35 40 45Gln Lys Leu Asp Pro Trp
Asn Gly Val Phe Asp Ala Thr Gln Tyr Gly50 55
60Asn Asn Cys Ala Ala Gly Lys Trp Phe Leu Lys Ser Ala Gly Gly Cys65
70 75 80Glu Asp Cys Leu
Tyr Leu Asn Ile Tyr Val Pro Gln Asn Thr Ser Glu85 90
95Asn Pro Leu Pro Val Met Phe Trp Ile His Gly Gly Ala Phe
Val Val100 105 110Gly Ser Gly Asn Ser Asp
Ile His Gly Pro Asp Tyr Leu Ile Glu Tyr115 120
125Asp Ile Ile Leu Val Thr Ile Asn Tyr Arg Leu Gly Pro Leu Gly
Phe130 135 140Leu Asn Leu Glu Ile Glu Asp
Ala Pro Gly Asn Val Gly Leu Met Asp145 150
155 160Gln Val Ala Ala Leu Lys Trp Val Asn Glu Asn Ile
Ala Thr Phe Ser165 170 175Gly Asp Pro Lys
Asn Ile Thr Ile Cys Gly Ala Thr Ala Gly Ala Ala180 185
190Ser Val His Tyr His Ile Leu Ser Gln Leu Thr Lys Gly Leu
Phe His195 200 205Lys Ala Ile Ala Gln Ser
Gly Ser Ala Phe Asn Pro Trp Ala Phe Gln210 215
220Lys Asn Pro Val Lys Asn Ala Leu Arg Leu Cys Lys Thr Leu Gly
Leu225 230 235 240Thr Thr
Asn Asn Leu Gln Glu Ala Leu Asp Phe Leu Lys Asn Leu Pro245
250 255Val Glu Thr Leu Leu Asn Thr Lys Leu Pro Gln Glu
Ile Asp Gly Gln260 265 270Leu Leu Asp Asp
Phe Val Phe Val Pro Ser Ile Glu Lys Thr Phe Pro275 280
285Glu Gln Asp Ser Tyr Leu Thr Asp Leu Pro Ile Pro Ile Ile
Asn Ser290 295 300Gly Lys Phe His Lys Val
Pro Leu Leu Thr Gly Tyr Asn Ser Ala Glu305 310
315 320Gly Asn Leu Phe Phe Met Tyr Leu Lys Thr Asp
Pro Asp Leu Leu Asn325 330 335Lys Phe Glu
Ala Asp Phe Glu Arg Phe Ile Pro Thr Asp Leu Glu Leu340
345 350Pro Leu Arg Ser Gln Lys Ser Ile Ala Leu Gly Glu
Ala Ile Arg Glu355 360 365Phe Tyr Phe Gln
Asn Lys Thr Ile Ser Glu Asn Met Gln Asn Phe Val370 375
380Asp Val Leu Ser Asp Asn Trp Phe Thr Arg Gly Ile Asp Glu
Gln Val385 390 395 400Lys
Leu Thr Val Lys Asn Gln Glu Glu Pro Val Phe Tyr Tyr Val Tyr405
410 415Asn Phe Asp Glu Asn Ser Pro Ser Arg Lys Val
Phe Gly Asp Phe Gly420 425 430Ile Lys Gly
Gly Gly His Ala Asp Glu Leu Gly Asn Ile Phe Lys Ala435
440 445Lys Ser Ala Asn Phe Gly Lys Glu Thr Pro Asn Ala
Val Leu Val Gln450 455 460Arg Arg Met Leu
Glu Met Trp Thr Asn Phe Ala Lys Phe Gly Asn Pro465 470
475 480Thr Pro Ala Ile Thr Asp Thr Leu Pro
Ile Lys Trp Glu Pro Ala Phe485 490 495Lys
Glu Asn Met Thr Phe Val Gln Ile Asp Ile Asp Leu Asn Leu Ser500
505 510Thr Asp Pro Leu Lys Ser Arg Met Glu Phe Gly
Asn Lys Ile Lys Leu515 520 525Leu
Lys530691987DNACtenocephalides felis 69gttttttttt tttttttttc tttttttttt
ccaaaataat tacatttttt attcgtaaaa 60gttggtttta taaaaaatta aatttactta
acgaagtaac aaaaaattgt gaagtattcc 120tatactatta tttggtatat tatggtttag
ctaagtatag ttacttattt taataatttt 180attttattcc caaattccat acgacttttt
agtggatcag tactcaaatt taaatcaatg 240tcaatttgaa caaaagtcat attttcttta
aaagcaggtt cccattttat tggaagtgta 300tccgtaatag ctggagtagg atttccaaat
ttagcaaaat tagtccacat ctccagcatc 360cttctctgaa ccaacacagc atttggtgtt
tccttcccaa aatttgcact tttggcttta 420aatatattac ccaattcatc agcatgacca
ccgcctttta ttccaaaatc accaaaaact 480ttccgacttg gagaattttc atcaaaatta
taaacataat aaaaaactgg ttcttcctga 540tttttaacag ttaactttac ttgctcatca
attccacgtg taaaccaatt atcacttaaa 600acatctacaa aattctgcat attttctgat
atggttttgt tttggaaata aaattccctg 660attgcttcac ccagtgcaat agatttttgt
gatcgcaaag gtaattctaa gtcagttggt 720ataaatcttt caaaatcagc ttcaaattta
tttaataaat ctggatctgt ttttaagtac 780atgaaaaata gattgccttc ggcactgttg
taacctgtca acaatggaac tttgtggaat 840tttcctgaat ttattattgg tattggcaag
tcagttaagt acgaatcttg ttctggaaat 900gttttttcaa tcgaaggtac aaacacgaag
tcatccagca gttgaccatc aatttcttgg 960ggtaatttgg tatttaacaa tgtttctact
ggtaggtttt tcaaaaaatc caaggcttct 1020tgaaggttgt ttgtggtaag gcctaaggtt
ttgcatagtc gaagtgcatt cttaacagga 1080tttttttgga aagcccaggg attaaaagca
cttccacttt gtgctatagc cttgtggaat 1140aaacctttgg taagttgtga caaaatgtga
taatgtacac ttgcagctcc agcagttgct 1200ccacaaattg taatattttt tgggtctcca
ctaaaggttg caatattttc atttacccat 1260tttagggctg caacttgatc catcaatcca
acattcccag gcgcatcttc gatttccaaa 1320ttaagaaaac caagtggtcc tagacgataa
ttaatagtta ctaagataat atcatattct 1380attaaataat caggaccatg tatatcagaa
tttcctgatc cgaccacaaa tgctcctcca 1440tgaatccaaa acattactgg caaaggattt
tctgaagtgt tttgtgggac atagatattt 1500aagtaaaggc aatcttcgca acccccagct
gatttcaaaa accatttccc agcagcacaa 1560ttatttccat actgagtggc gtcaaaaaca
ccattccaag gatcaagttt ttgtggtggc 1620ttgaatctga gatcatttac aggagatttt
gcatagggta tacctgtgta actatagtaa 1680attttaccat tttcgtttac aactttcttg
cctttcagaa ttccatatgt tgttgttttt 1740agtaatggat cacacattat aaaatgttat
tatatatcca aaatataata tggtttaatt 1800ttatactagc tcaaaaataa tgtaaatcat
agccaaaaca cacactcata aacataaata 1860aaaatttctt ttcgatcata aaaaaaatat
ttttttataa ataaaactat atatatttaa 1920attaaaatta gattaaaata aaaattaaat
acatttatat caaataaaat tatttacact 1980gtgaatt
1987701590DNACtenocephalides
felisexon(1)..(1590) 70atg tgt gat cca tta cta aaa aca aca aca tat gga
att ctg aaa ggc 48Met Cys Asp Pro Leu Leu Lys Thr Thr Thr Tyr Gly
Ile Leu Lys Gly1 5 10
15aag aaa gtt gta aac gaa aat ggt aaa att tac tat agt tac aca ggt
96Lys Lys Val Val Asn Glu Asn Gly Lys Ile Tyr Tyr Ser Tyr Thr Gly20
25 30ata ccc tat gca aaa tct cct gta aat gat
ctc aga ttc aag cca cca 144Ile Pro Tyr Ala Lys Ser Pro Val Asn Asp
Leu Arg Phe Lys Pro Pro35 40 45caa aaa
ctt gat cct tgg aat ggt gtt ttt gac gcc act cag tat gga 192Gln Lys
Leu Asp Pro Trp Asn Gly Val Phe Asp Ala Thr Gln Tyr Gly50
55 60aat aat tgt gct gct ggg aaa tgg ttt ttg aaa tca
gct ggg ggt tgc 240Asn Asn Cys Ala Ala Gly Lys Trp Phe Leu Lys Ser
Ala Gly Gly Cys65 70 75
80gaa gat tgc ctt tac tta aat atc tat gtc cca caa aac act tca gaa
288Glu Asp Cys Leu Tyr Leu Asn Ile Tyr Val Pro Gln Asn Thr Ser Glu85
90 95aat cct ttg cca gta atg ttt tgg att cat
gga gga gca ttt gtg gtc 336Asn Pro Leu Pro Val Met Phe Trp Ile His
Gly Gly Ala Phe Val Val100 105 110gga tca
gga aat tct gat ata cat ggt cct gat tat tta ata gaa tat 384Gly Ser
Gly Asn Ser Asp Ile His Gly Pro Asp Tyr Leu Ile Glu Tyr115
120 125gat att atc tta gta act att aat tat cgt cta gga
cca ctt ggt ttt 432Asp Ile Ile Leu Val Thr Ile Asn Tyr Arg Leu Gly
Pro Leu Gly Phe130 135 140ctt aat ttg gaa
atc gaa gat gcg cct ggg aat gtt gga ttg atg gat 480Leu Asn Leu Glu
Ile Glu Asp Ala Pro Gly Asn Val Gly Leu Met Asp145 150
155 160caa gtt gca gcc cta aaa tgg gta aat
gaa aat att gca acc ttt agt 528Gln Val Ala Ala Leu Lys Trp Val Asn
Glu Asn Ile Ala Thr Phe Ser165 170 175gga
gac cca aaa aat att aca att tgt gga gca act gct gga gct gca 576Gly
Asp Pro Lys Asn Ile Thr Ile Cys Gly Ala Thr Ala Gly Ala Ala180
185 190agt gta cat tat cac att ttg tca caa ctt acc
aaa ggt tta ttc cac 624Ser Val His Tyr His Ile Leu Ser Gln Leu Thr
Lys Gly Leu Phe His195 200 205aag gct ata
gca caa agt gga agt gct ttt aat ccc tgg gct ttc caa 672Lys Ala Ile
Ala Gln Ser Gly Ser Ala Phe Asn Pro Trp Ala Phe Gln210
215 220aaa aat cct gtt aag aat gca ctt cga cta tgc aaa
acc tta ggc ctt 720Lys Asn Pro Val Lys Asn Ala Leu Arg Leu Cys Lys
Thr Leu Gly Leu225 230 235
240acc aca aac aac ctt caa gaa gcc ttg gat ttt ttg aaa aac cta cca
768Thr Thr Asn Asn Leu Gln Glu Ala Leu Asp Phe Leu Lys Asn Leu Pro245
250 255gta gaa aca ttg tta aat acc aaa tta
ccc caa gaa att gat ggt caa 816Val Glu Thr Leu Leu Asn Thr Lys Leu
Pro Gln Glu Ile Asp Gly Gln260 265 270ctg
ctg gat gac ttc gtg ttt gta cct tcg att gaa aaa aca ttt cca 864Leu
Leu Asp Asp Phe Val Phe Val Pro Ser Ile Glu Lys Thr Phe Pro275
280 285gaa caa gat tcg tac tta act gac ttg cca ata
cca ata ata aat tca 912Glu Gln Asp Ser Tyr Leu Thr Asp Leu Pro Ile
Pro Ile Ile Asn Ser290 295 300gga aaa ttc
cac aaa gtt cca ttg ttg aca ggt tac aac agt gcc gaa 960Gly Lys Phe
His Lys Val Pro Leu Leu Thr Gly Tyr Asn Ser Ala Glu305
310 315 320ggc aat cta ttt ttc atg tac
tta aaa aca gat cca gat tta tta aat 1008Gly Asn Leu Phe Phe Met Tyr
Leu Lys Thr Asp Pro Asp Leu Leu Asn325 330
335aaa ttt gaa gct gat ttt gaa aga ttt ata cca act gac tta gaa tta
1056Lys Phe Glu Ala Asp Phe Glu Arg Phe Ile Pro Thr Asp Leu Glu Leu340
345 350cct ttg cga tca caa aaa tct att gca
ctg ggt gaa gca atc agg gaa 1104Pro Leu Arg Ser Gln Lys Ser Ile Ala
Leu Gly Glu Ala Ile Arg Glu355 360 365ttt
tat ttc caa aac aaa acc ata tca gaa aat atg cag aat ttt gta 1152Phe
Tyr Phe Gln Asn Lys Thr Ile Ser Glu Asn Met Gln Asn Phe Val370
375 380gat gtt tta agt gat aat tgg ttt aca cgt gga
att gat gag caa gta 1200Asp Val Leu Ser Asp Asn Trp Phe Thr Arg Gly
Ile Asp Glu Gln Val385 390 395
400aag tta act gtt aaa aat cag gaa gaa cca gtt ttt tat tat gtt tat
1248Lys Leu Thr Val Lys Asn Gln Glu Glu Pro Val Phe Tyr Tyr Val Tyr405
410 415aat ttt gat gaa aat tct cca agt cgg
aaa gtt ttt ggt gat ttt gga 1296Asn Phe Asp Glu Asn Ser Pro Ser Arg
Lys Val Phe Gly Asp Phe Gly420 425 430ata
aaa ggc ggt ggt cat gct gat gaa ttg ggt aat ata ttt aaa gcc 1344Ile
Lys Gly Gly Gly His Ala Asp Glu Leu Gly Asn Ile Phe Lys Ala435
440 445aaa agt gca aat ttt ggg aag gaa aca cca aat
gct gtg ttg gtt cag 1392Lys Ser Ala Asn Phe Gly Lys Glu Thr Pro Asn
Ala Val Leu Val Gln450 455 460aga agg atg
ctg gag atg tgg act aat ttt gct aaa ttt gga aat cct 1440Arg Arg Met
Leu Glu Met Trp Thr Asn Phe Ala Lys Phe Gly Asn Pro465
470 475 480act cca gct att acg gat aca
ctt cca ata aaa tgg gaa cct gct ttt 1488Thr Pro Ala Ile Thr Asp Thr
Leu Pro Ile Lys Trp Glu Pro Ala Phe485 490
495aaa gaa aat atg act ttt gtt caa att gac att gat tta aat ttg agt
1536Lys Glu Asn Met Thr Phe Val Gln Ile Asp Ile Asp Leu Asn Leu Ser500
505 510act gat cca cta aaa agt cgt atg gaa
ttt ggg aat aaa ata aaa tta 1584Thr Asp Pro Leu Lys Ser Arg Met Glu
Phe Gly Asn Lys Ile Lys Leu515 520 525tta
aaa 1590Leu
Lys530711590DNACtenocephalides felis 71ttttaataat tttattttat tcccaaattc
catacgactt tttagtggat cagtactcaa 60atttaaatca atgtcaattt gaacaaaagt
catattttct ttaaaagcag gttcccattt 120tattggaagt gtatccgtaa tagctggagt
aggatttcca aatttagcaa aattagtcca 180catctccagc atccttctct gaaccaacac
agcatttggt gtttccttcc caaaatttgc 240acttttggct ttaaatatat tacccaattc
atcagcatga ccaccgcctt ttattccaaa 300atcaccaaaa actttccgac ttggagaatt
ttcatcaaaa ttataaacat aataaaaaac 360tggttcttcc tgatttttaa cagttaactt
tacttgctca tcaattccac gtgtaaacca 420attatcactt aaaacatcta caaaattctg
catattttct gatatggttt tgttttggaa 480ataaaattcc ctgattgctt cacccagtgc
aatagatttt tgtgatcgca aaggtaattc 540taagtcagtt ggtataaatc tttcaaaatc
agcttcaaat ttatttaata aatctggatc 600tgtttttaag tacatgaaaa atagattgcc
ttcggcactg ttgtaacctg tcaacaatgg 660aactttgtgg aattttcctg aatttattat
tggtattggc aagtcagtta agtacgaatc 720ttgttctgga aatgtttttt caatcgaagg
tacaaacacg aagtcatcca gcagttgacc 780atcaatttct tggggtaatt tggtatttaa
caatgtttct actggtaggt ttttcaaaaa 840atccaaggct tcttgaaggt tgtttgtggt
aaggcctaag gttttgcata gtcgaagtgc 900attcttaaca ggattttttt ggaaagccca
gggattaaaa gcacttccac tttgtgctat 960agccttgtgg aataaacctt tggtaagttg
tgacaaaatg tgataatgta cacttgcagc 1020tccagcagtt gctccacaaa ttgtaatatt
ttttgggtct ccactaaagg ttgcaatatt 1080ttcatttacc cattttaggg ctgcaacttg
atccatcaat ccaacattcc caggcgcatc 1140ttcgatttcc aaattaagaa aaccaagtgg
tcctagacga taattaatag ttactaagat 1200aatatcatat tctattaaat aatcaggacc
atgtatatca gaatttcctg atccgaccac 1260aaatgctcct ccatgaatcc aaaacattac
tggcaaagga ttttctgaag tgttttgtgg 1320gacatagata tttaagtaaa ggcaatcttc
gcaaccccca gctgatttca aaaaccattt 1380cccagcagca caattatttc catactgagt
ggcgtcaaaa acaccattcc aaggatcaag 1440tttttgtggt ggcttgaatc tgagatcatt
tacaggagat tttgcatagg gtatacctgt 1500gtaactatag taaattttac cattttcgtt
tacaactttc ttgcctttca gaattccata 1560tgttgttgtt tttagtaatg gatcacacat
159072650DNACtenocephalides
felisCDS(3)..(650) 72gg atc cat gga ggc gca ttc aac caa gga tca gga tct
tat aat ttt 47Ile His Gly Gly Ala Phe Asn Gln Gly Ser Gly Ser Tyr
Asn Phe1 5 10 15ttt gga
cct gat tat ttg atc agg gaa gga att att ttg gtc act atc 95Phe Gly
Pro Asp Tyr Leu Ile Arg Glu Gly Ile Ile Leu Val Thr Ile20
25 30aac tat aga tta gga gtt ttc ggt ttt cta tca gcg
ccg gaa tgg gat 143Asn Tyr Arg Leu Gly Val Phe Gly Phe Leu Ser Ala
Pro Glu Trp Asp35 40 45atc cat gga aat
atg ggt cta aaa gac cag aga ttg gca cta aaa tgg 191Ile His Gly Asn
Met Gly Leu Lys Asp Gln Arg Leu Ala Leu Lys Trp50 55
60gtt tac gac aac atc gaa aag ttt ggt gga gac aga gaa aaa
att aca 239Val Tyr Asp Asn Ile Glu Lys Phe Gly Gly Asp Arg Glu Lys
Ile Thr65 70 75att gct gga gaa tct gct
gga gca gca agt gtc cat ttt ctg atg atg 287Ile Ala Gly Glu Ser Ala
Gly Ala Ala Ser Val His Phe Leu Met Met80 85
90 95gac aac tcg act aga aaa tac tac caa agg gcc
att ttg cag agt ggg 335Asp Asn Ser Thr Arg Lys Tyr Tyr Gln Arg Ala
Ile Leu Gln Ser Gly100 105 110aca tta cta
aat ccg act gct aat caa att caa ctt ctg cat aga ttt 383Thr Leu Leu
Asn Pro Thr Ala Asn Gln Ile Gln Leu Leu His Arg Phe115
120 125gaa aaa ctc aaa caa gtg cta aac atc acg caa aaa
caa gaa ctc cta 431Glu Lys Leu Lys Gln Val Leu Asn Ile Thr Gln Lys
Gln Glu Leu Leu130 135 140aac ctg gat aaa
aac cta att tta cga gca gcc tta aac aga gtt cct 479Asn Leu Asp Lys
Asn Leu Ile Leu Arg Ala Ala Leu Asn Arg Val Pro145 150
155gat agc aac gac cat gac cga gac aca gta cca gta ttt aat
cca gtc 527Asp Ser Asn Asp His Asp Arg Asp Thr Val Pro Val Phe Asn
Pro Val160 165 170 175tta
gaa tca cca gaa tct cca gat cca ata aca ttt cca tct gcc ttg 575Leu
Glu Ser Pro Glu Ser Pro Asp Pro Ile Thr Phe Pro Ser Ala Leu180
185 190gaa aga atg aga aat ggt gaa ttt cct gat gtc
gat gtc atc att ggt 623Glu Arg Met Arg Asn Gly Glu Phe Pro Asp Val
Asp Val Ile Ile Gly195 200 205ttc aat agt
gct gaa ggt tta aga tct 650Phe Asn Ser
Ala Glu Gly Leu Arg Ser210 21573216PRTCtenocephalides
felis 73Ile His Gly Gly Ala Phe Asn Gln Gly Ser Gly Ser Tyr Asn Phe Phe1
5 10 15Gly Pro Asp Tyr
Leu Ile Arg Glu Gly Ile Ile Leu Val Thr Ile Asn20 25
30Tyr Arg Leu Gly Val Phe Gly Phe Leu Ser Ala Pro Glu Trp
Asp Ile35 40 45His Gly Asn Met Gly Leu
Lys Asp Gln Arg Leu Ala Leu Lys Trp Val50 55
60Tyr Asp Asn Ile Glu Lys Phe Gly Gly Asp Arg Glu Lys Ile Thr Ile65
70 75 80Ala Gly Glu Ser
Ala Gly Ala Ala Ser Val His Phe Leu Met Met Asp85 90
95Asn Ser Thr Arg Lys Tyr Tyr Gln Arg Ala Ile Leu Gln Ser
Gly Thr100 105 110Leu Leu Asn Pro Thr Ala
Asn Gln Ile Gln Leu Leu His Arg Phe Glu115 120
125Lys Leu Lys Gln Val Leu Asn Ile Thr Gln Lys Gln Glu Leu Leu
Asn130 135 140Leu Asp Lys Asn Leu Ile Leu
Arg Ala Ala Leu Asn Arg Val Pro Asp145 150
155 160Ser Asn Asp His Asp Arg Asp Thr Val Pro Val Phe
Asn Pro Val Leu165 170 175Glu Ser Pro Glu
Ser Pro Asp Pro Ile Thr Phe Pro Ser Ala Leu Glu180 185
190Arg Met Arg Asn Gly Glu Phe Pro Asp Val Asp Val Ile Ile
Gly Phe195 200 205Asn Ser Ala Glu Gly Leu
Arg Ser210 2157415PRTCtenocephalides
felisMISC_FEATURE(3)..(3)Xaa = unknown 74Asp Leu Xaa Val Xaa Xaa Leu Gln
Gly Thr Leu Lys Gly Lys Glu1 5 10
157531DNAArtificial sequenceSynthetic Primer 75cgcggatccg
ctgatctaca agtgactttg c
31761488DNACtenocephalides felisexon(3)..(1487) 76cc cag ggc gaa ttg gtt
gga aaa gct ttg acg aac gaa aat gga aaa 47Gln Gly Glu Leu Val Gly
Lys Ala Leu Thr Asn Glu Asn Gly Lys1 5 10
15gag tat ttt agc tac aca ggt gtg cct tat gct aaa cct
cca gtt gga 95Glu Tyr Phe Ser Tyr Thr Gly Val Pro Tyr Ala Lys Pro
Pro Val Gly20 25 30gaa ctt aga ttt aag
cct cca cag aaa gct gag cca tgg aat ggt gtt 143Glu Leu Arg Phe Lys
Pro Pro Gln Lys Ala Glu Pro Trp Asn Gly Val35 40
45ttc aac gcc aca tca cat gga aat gtg tgc aaa gct ttg aat ttc
ttc 191Phe Asn Ala Thr Ser His Gly Asn Val Cys Lys Ala Leu Asn Phe
Phe50 55 60ttg aaa aaa att gaa gga gac
gaa gac tgc ttg ttg gtg aat gtg tac 239Leu Lys Lys Ile Glu Gly Asp
Glu Asp Cys Leu Leu Val Asn Val Tyr65 70
75gca cca aaa aca act tct gac aaa aaa ctt cca gta ttt ttc tgg gtt
287Ala Pro Lys Thr Thr Ser Asp Lys Lys Leu Pro Val Phe Phe Trp Val80
85 90 95cat ggt ggc ggt ttt
gtg act gga tcc gga aat tta gaa ttt caa agc 335His Gly Gly Gly Phe
Val Thr Gly Ser Gly Asn Leu Glu Phe Gln Ser100 105
110cca gat tat tta gta aat tat gat gtt att ttt gta act ttc aat
tac 383Pro Asp Tyr Leu Val Asn Tyr Asp Val Ile Phe Val Thr Phe Asn
Tyr115 120 125cga ttg gga cca ctc gga ttt
ttg aat ttg gag ttg gaa ggt gct cct 431Arg Leu Gly Pro Leu Gly Phe
Leu Asn Leu Glu Leu Glu Gly Ala Pro130 135
140gga aat gta gga tta ttg gat cag gta gca gct ttg aaa tgg acc aaa
479Gly Asn Val Gly Leu Leu Asp Gln Val Ala Ala Leu Lys Trp Thr Lys145
150 155gaa aat att gag aaa ttt ggt gga gat
cca gaa aat att aca att ggt 527Glu Asn Ile Glu Lys Phe Gly Gly Asp
Pro Glu Asn Ile Thr Ile Gly160 165 170
175ggt gtt tct gct ggt gga gca agt gtt cat tat ctt tta ttg
tca cat 575Gly Val Ser Ala Gly Gly Ala Ser Val His Tyr Leu Leu Leu
Ser His180 185 190aca acc act gga ctt tac
aaa agg gca att gct caa agt gga agt gct 623Thr Thr Thr Gly Leu Tyr
Lys Arg Ala Ile Ala Gln Ser Gly Ser Ala195 200
205tta aat cca tgg gcc ttc caa aga cat cca gta aag cgt agt ctt caa
671Leu Asn Pro Trp Ala Phe Gln Arg His Pro Val Lys Arg Ser Leu Gln210
215 220ctt gct gag ata tta ggt cat ccc aca
aac aac act caa gat gct tta 719Leu Ala Glu Ile Leu Gly His Pro Thr
Asn Asn Thr Gln Asp Ala Leu225 230 235gaa
ttc tta caa aaa gcc cca gta gac agt ctc ctg aaa aaa atg cca 767Glu
Phe Leu Gln Lys Ala Pro Val Asp Ser Leu Leu Lys Lys Met Pro240
245 250 255gct gaa aca gaa ggt gaa
ata ata gaa gag ttc gtc ttc gta cca tca 815Ala Glu Thr Glu Gly Glu
Ile Ile Glu Glu Phe Val Phe Val Pro Ser260 265
270att gaa aaa gtt ttc cca tcc cac caa cct ttc ttg gaa gaa tca cca
863Ile Glu Lys Val Phe Pro Ser His Gln Pro Phe Leu Glu Glu Ser Pro275
280 285ttg gcc aga atg aaa tct gga tcc ttt
aac aaa gta cct tta tta gtt 911Leu Ala Arg Met Lys Ser Gly Ser Phe
Asn Lys Val Pro Leu Leu Val290 295 300gga
ttc aac agc gca gaa gga ctt ttg tac aaa ttc ttt atg aaa gaa 959Gly
Phe Asn Ser Ala Glu Gly Leu Leu Tyr Lys Phe Phe Met Lys Glu305
310 315aaa cca gag atg ctg aac caa gct gaa gca gat
ttc gaa aga ctc gta 1007Lys Pro Glu Met Leu Asn Gln Ala Glu Ala Asp
Phe Glu Arg Leu Val320 325 330
335cca gcc gaa ttt gaa tta gcc cat gga tca gaa gaa tcg aaa aaa ctt
1055Pro Ala Glu Phe Glu Leu Ala His Gly Ser Glu Glu Ser Lys Lys Leu340
345 350gca gaa aaa atc agg aag ttt tac ttt
gac gat aaa ccc gtt cct gaa 1103Ala Glu Lys Ile Arg Lys Phe Tyr Phe
Asp Asp Lys Pro Val Pro Glu355 360 365aat
gag cag aaa ttt att gac ttg ata gga gat att tgg ttt act aga 1151Asn
Glu Gln Lys Phe Ile Asp Leu Ile Gly Asp Ile Trp Phe Thr Arg370
375 380ggc att gac aag cat gtc aag ttg tct gta gaa
aaa caa gac gag cca 1199Gly Ile Asp Lys His Val Lys Leu Ser Val Glu
Lys Gln Asp Glu Pro385 390 395gta tat tat
tat gaa tat tct ttc tct gaa agt cat cct gca aaa gga 1247Val Tyr Tyr
Tyr Glu Tyr Ser Phe Ser Glu Ser His Pro Ala Lys Gly400
405 410 415aca ttt ggt gac cat aac ttg
act gga gca tgt cat ggt gaa gaa ctt 1295Thr Phe Gly Asp His Asn Leu
Thr Gly Ala Cys His Gly Glu Glu Leu420 425
430gtg aat tta ttc aaa gtc gag atg atg aag ctg gaa aaa gat aaa ccg
1343Val Asn Leu Phe Lys Val Glu Met Met Lys Leu Glu Lys Asp Lys Pro435
440 445aat gtt tta tta aca aaa gat agg gta
ctt gct atg tgg acg aac ttc 1391Asn Val Leu Leu Thr Lys Asp Arg Val
Leu Ala Met Trp Thr Asn Phe450 455 460atc
aaa aat gga aat cct act cct gaa gta act gaa tta ttg cca gtt 1439Ile
Lys Asn Gly Asn Pro Thr Pro Glu Val Thr Glu Leu Leu Pro Val465
470 475aaa tgg gaa cct gcc aca aaa gac aag ttg aat
tat ttg aac att gat g 1488Lys Trp Glu Pro Ala Thr Lys Asp Lys Leu Asn
Tyr Leu Asn Ile Asp480 485 490
495
User Contributions:
comments("1"); ?> comment_form("1"); ?>Inventors list |
Agents list |
Assignees list |
List by place |
Classification tree browser |
Top 100 Inventors |
Top 100 Agents |
Top 100 Assignees |
Usenet FAQ Index |
Documents |
Other FAQs |
User Contributions:
Comment about this patent or add new information about this topic: